Abstract
Cofilin is a central regulator of actin filament turnover, traditionally thought to act through phosphorylation-dependent control of filament assembly. However, its mitochondrial functions remain poorly understood. Here we show that N-terminal α-amino SUMOylation, rather than phosphorylation or actin interaction, governs cofilin-1 translocation to mitochondria and activation of the apoptotic pathway. This modification strengthens the association of cofilin-1 with the mitochondrial import receptors Tom20 and Tom70 through the molecular chaperone HSP70, enabling its delivery to the mitochondrial matrix. Once imported, SUMO-modified cofilin-1 binds cytochrome c1, promotes the dissociation of cytochrome c from complex III, and initiates mitochondrial-mediated apoptosis. These findings redefine cofilin-1 as a regulator of mitochondrial integrity independent of its actin-related roles, uncovering a mechanism by which SUMOylation directs protein targeting and apoptotic signaling. This work broadens current understanding of mitochondrial regulation and may inform therapeutic strategies for diseases linked to defective cell death.
Similar content being viewed by others
Introduction
Cofilin (CFL), a key regulator of actin depolymerization, plays an essential role in the regulation of cytoskeletal dynamics. It exists in two primary isoforms, CFL1 (non-muscle type) and CFL2 (muscle type). CFL1 and CFL2 proteins show 81% sequence homology in mice and share all functional domains, indicating that they overlap in most of the functions identified1,2. Structurally, CFL consists of a six-stranded mixed β-sheet surrounded by seven α-helices. The flexible N-terminal region of CFL is critical for actin binding3, enabling it to facilitate the reversible disassembly of actin filaments. This disassembly orchestrates a variety of cellular processes, ranging from motility and cytokinesis to the complex mechanisms of morphogenesis4,5.
While its ancestral ties with the actin cytoskeleton are well-established, recent studies have broadened the understanding of CFL, linking it to mitochondrial functions6,7. Current views suggest that the regulation of mitochondrial function by CFL is primarily controlled by its post-translational modifications (PTMs), such as phosphorylation and oxidation. Among these, phosphorylation modifications have been most widely reported8. CFL is phosphorylated at Serine 3 (Ser3) by LIM kinase (LIMK) and TES kinase (TESK), and dephosphorylated by phosphatases such as slingshot homolog (SSH), chronophin (CIN), and protein phosphatases 1 and 2 A (PP1 and PP2A)9,10,11,12. Previous studies showed that dephosphorylation of CFL at the Ser3 site allows it to translocate and localize to mitochondria, triggering mitochondrial dysfunction and apoptosis13,14,15. Alternatively, recent studies suggest that the dephosphorylated form of CFL translocates to the outer mitochondrial membrane, causing mitochondrial fission and autophagy16,17. CFL phosphorylation regulates the assembly and disassembly of actin filaments at mitochondrial fission sites, participating in dynamin-related protein 1 (Drp1)-mediated mitochondrial fission18,19. Furthermore, only dephosphorylated Drp1 and dephosphorylated CFL translocate to the mitochondria. The interaction between CFL and Drp1 participates in mitochondrial fission and apoptosis20,21. Moreover, oxidation of CFL has also been reported to be associated with mitochondrial translocation and apoptosis. Human CFL contains four cysteine residues at positions 39, 80, 139, and 147, which are considered prone to oxidation. Oxidation of CFL leads to its translocation to the mitochondria, inducing apoptosis22, mitochondrial dysfunction, and mitochondrial fission23,24. Additionally, it has been reported that oxidation of cysteine residues at positions 39 or 80 induces CFL mitochondrial translocation, promoting caspase-independent, necrosis-like programmed cell death in T cells25.
Although the roles of CFL phosphorylation and oxidation in the regulation of mitochondrial function have been reported, the specific mechanisms by which they regulate CFL mitochondrial translocation and mitochondrial pathway apoptosis remain unclear. Furthermore, it is still unknown whether other PTMs of CFL, such as N-terminal α-amino SUMOylation, are involved in this process.
Here, we show that N-terminal α-amino SUMOylation governs CFL1 mitochondrial targeting and apoptosis. N-terminal α-amino SUMOylation, rather than phosphorylation or actin interaction, serves as the primary driver of CFL1 mitochondrial translocation. This modification enhances CFL1 interactions with the mitochondrial import receptors Tom20 and Tom70 in an HSP70-dependent manner, enabling efficient mitochondrial entry. Once localized to the mitochondrial matrix, N-terminal α-amino SUMOylated CFL1 interacts with cytochrome c1 (CYC1), disrupts the CYC1–cytochrome c (Cyt c) association, and triggers Cyt c release and apoptotic signaling. These findings establish N-terminal α-amino SUMOylation as a critical determinant of CFL1 mitochondrial localization and function, reveal a regulatory pathway governing mitochondrial apoptosis, and suggest that SUMOylation may represent a broader mechanism for directing cytosolic proteins to mitochondria in physiology and disease.
Results
N-terminal α-amino SUMOylation of CFL1 enhances its mitochondrial translocation
Previous studies have shown that activated CFL translocates to the mitochondria following staurosporine-induced apoptosis, where it triggers mitochondrial dysfunction and apoptotic cell death13,26. However, the molecular mechanisms governing CFL mitochondrial translocation remain unclear and are still under debate8. Our recent work demonstrated that CFL1 undergoes N-terminal α-amino SUMOylation, a post-translational modification essential for its actin-depolymerizing activity27. Building on this discovery, we investigated whether N-terminal α-amino SUMOylation influences the mitochondrial translocation of CFL1.
To mimic this modification, we generated two expression constructs: SUMO1-CFL1, in which SUMO1 was fused to the N-terminus of CFL1 via Gly97 of SUMO1, and SUMO1G97A-CFL1, in which the G97A mutation renders SUMO1 resistant to SENP1-mediated cleavage. HeLa cells with stable shRNA knockdown of endogenous CFL1 were transfected with wild-type (CFL1-HA), SUMO1-CFL1-HA, or SUMO1G97A-CFL1-HA plasmids, followed by mitochondrial isolation. Western blot analysis revealed that both SUMO1-CFL1-HA and SUMO1G97A-CFL1-HA exhibited significantly higher mitochondrial localization than CFL1-HA (Fig. 1a). Consistently, immunofluorescence microscopy confirmed enhanced colocalization of N-terminally SUMOylated CFL1 with mitochondrial markers, quantified using Pearson’s correlation coefficient (PCC)28 (Fig. 1b). These data demonstrate that N-terminal α-amino SUMOylation promotes CFL1 mitochondrial translocation.
a CFL1 fused with SUMO1 at the N-terminus exhibited increased mitochondrial translocation. CFL1 knockdown was established in HeLa cells using lentiviral shRNA followed by blasticidin selection. Cells were then transfected with CFL1-HA, SUMO1-CFL1-HA, or SUMO1G97A-CFL1-HA plasmids, followed by IP with HA beads. Subcellular localization was assessed by WB analysis of mitochondrial (Mito), cytoplasmic (Cyto), and total lysate fractions from the same cells. b Confocal images of colocalization between CFL1 and mitochondria. After transfection with CFL1-HA, SUMO1-CFL1-HA, or SUMO1G97A-CFL1-HA plasmids into HeLa cells with endogenous CFL1 knocked down, the cells were stained with anti-HA antibody (green) and MitoTracker (magenta). The PCC measured co-localization of magenta and green fluorescence. Data were presented as the mean ± SEM of six visual fields from 3 biological replicates. Statistical significance was determined by one-way ANOVA with Tukey’s multiple comparisons test. The p values were as follows: CFL1-HA vs. SUMO1-CFL1-HA, ****p < 0.0001; CFL1-HA vs. SUMO1G97A-CFL1-HA, ****p < 0.0001. c Overexpression of His-SUMO1 but not His-SUMO1G97A (conjugation-deficient mutant of SUMO1) in HeLa cells promoted SUMOylation and mitochondrial translocation of CFL1. IP with control IgG or anti-CFL1 antibody followed by WB. d Confocal images of colocalization between CFL1 and mitochondria in HeLa cells transfected mock, His-SUMO1, or His-SUMO1G97A plasmids. The cells were stained with anti-SUMO1 antibody (blue), anti-HA antibody (green) and MitoTracker (magenta). The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates. Statistical significance was determined by one-way ANOVA with Tukey’s multiple comparisons test. The p values were as follows: Mock vs. His-SUMO1, ***p = 0.0002; Mock vs. His-SUMO1G97A, *p = 0.0138. e SENP1 deficiency promoted SUMOylation and mitochondrial translocation of CFL1. Lysates from Senp1+/+ and Senp1−/− mice brain tissues at embryonic day (E) 13.5 were analyzed by IP and WB. f Senp1−/− MEF cells exhibited increased CFL1 SUMOylation and mitochondrial translocation. The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates; ****p < 0.0001, analyzed by two-tailed Student’s t-test. Source data are provided as a Source Data file.
To further validate this effect, HeLa cells were transiently transfected with His-SUMO1 or His-SUMO1G97A plasmids. Western blotting and immunofluorescence analyses showed that enhanced SUMOylation increased CFL1 accumulation within mitochondria (Fig. 1c, d). Because SENP1 is a SUMO-specific protease that counteracts global SUMOylation, we next examined whether SENP1 deficiency affects CFL1 mitochondrial translocation. Mitochondria isolated from brain tissues of Senp1−/− mouse embryos exhibited elevated CFL1 SUMOylation and significantly increased mitochondrial CFL1 levels compared with Senp1+/+ controls (Fig. 1e). Similarly, Senp1−/− mouse embryonic fibroblasts (MEFs) displayed greater CFL1-mitochondria colocalization than wild-type MEFs (Fig. 1f). Collectively, these findings establish that N-terminal α-amino SUMOylation markedly enhances the mitochondrial translocation of CFL1, suggesting that this modification may contribute to the regulation of CFL1 mitochondrial function.
Phosphorylation-independent mitochondrial translocation of CFL1 mediated by N-terminal α-amino SUMOylation
Phosphorylation of CFL1 at Ser3 is a well-established regulatory switch that suppresses actin depolymerization and mitochondrial translocation, whereas dephosphorylation promotes CFL mitochondrial translocation and apoptosis13,14,15. Given that our data demonstrated that N-terminal α-amino SUMOylation enhances the mitochondrial translocation of CFL1 (Fig. 1), we next investigated how this modification interacts with Ser3 phosphorylation.
Based on our previous findings27, the SUMOylation level of the CFL12KR (K112/114R) mutant was markedly reduced, whereas the CFL1K34R mutant exhibited enhanced SUMOylation. Mass spectrometry confirmed that SUMOylation in both mutants occurred at the N-terminus (Supplementary Fig. 1a, b). In HEK 293T cells depleted of endogenous CFL1, co-immunoprecipitation (Co-IP) assays revealed that CFL1K34R interacted more strongly with E1 (SAE1/SAE2) and E2 (Ubc9) enzymes and less with SENP1, explaining its elevated SUMOylation (Supplementary Fig. 1c). To examine the crosstalk between SUMOylation and phosphorylation, we expressed CFL1-HA, CFL12KR-HA, CFL1K34R-HA, CFL1S3A-HA (non-phosphorylatable)29 and CFL1S3E-HA (phosphomimetic)30 in HEK 293T cells. CFL1K34R displayed reduced Ser3 phosphorylation, whereas CFL12KR showed increased phosphorylation, suggesting that N-terminal SUMOylation negatively regulates Ser3 phosphorylation (Supplementary Fig. 1d). Consistently, titration of His-SUMO1 or its catalytically inactive mutant His-SUMO1G97A in CHO-K1 cells revealed a reciprocal relationship: increased SUMOylation correlated with reduced Ser3 phosphorylation, whereas inhibition of SUMO conjugation enhanced phosphorylation (Supplementary Fig. 1e). Expression of SUMO1-CFL1 or SUMO1G97A-CFL1 further supported the inhibitory effect of N-terminal SUMOylation on Ser3 phosphorylation (Supplementary Fig. 1f).
We next tested whether phosphorylation reciprocally affects N-terminal SUMOylation. Using peptide-based modulation of LIMK and SSH activities31,32, cells were treated with control TAT peptide (TAT-Ctl), TAT-S3 (inhibits LIMK-mediated phosphorylation), or TAT-pS3 (inhibits SSH-mediated dephosphorylation). In parallel, CFL1-HA, CFL1S3A-HA, and CFL1S3E-HA were co-expressed with His-SUMO1. Both approaches confirmed that phosphorylation and N-terminal SUMOylation are mutually inhibitory, establishing a reciprocal regulatory relationship between these two modifications (Supplementary Fig. 1g, h).
To clarify which modification governs mitochondrial targeting, we analyzed mitochondria isolated from Senp1+/+ and Senp1−/− mouse embryos. As expected, increased SUMOylation in Senp1−/− embryos coincided with reduced Ser3 phosphorylation and enhanced mitochondrial localization of CFL1 (Fig. 2a). In Senp1+/+ and Senp1−/− MEFs, modulation of phosphorylation using TAT-Ctl, TAT-S3, or TAT-pS3 peptides altered phosphorylation levels but did not significantly affect mitochondrial localization in wild-type cells. In contrast, Senp1−/− cells retained robust mitochondrial translocation regardless of phosphorylation state, consistent with SUMOylation status (Fig. 2b).
a Senp1−/− mouse embryos exhibited increased CFL1 SUMOylation, mitochondrial translocation and decreased phosphorylation. b Senp1−/− MEF cells exhibited increased CFL1 SUMOylation and mitochondrial translocation, regardless of changes in CFL1 phosphorylation induced by TAT-S3 or TAT-pS3 peptide (10 μM, 16 h). The ratios of Mito CFL1, SUMOylated CFL1 and phospho-CFL1 (p-CFL1) to total CFL1 were presented as the mean ± SEM from 3 biological replicates; ****p < 0.0001, ***p = 0.0003, “ns” means not significant (p = 0.3613), analyzed by one-way ANOVA with Tukey’s multiple comparisons test. c Knockdown of SUMO1 and SUMO2/3 inhibited CFL1 mitochondrial translocation. WT or Sumo1 KO HeLa cells were transfected with either control siRNA or siRNA complex targeting SUMO2 and SUMO3. 48 h later, cell lysates were analyzed by IP and WB. d ML-792 inhibited CFL1 SUMOylation and mitochondrial translocation regardless of changes in CFL1 phosphorylation induced by peptides. HEK 293T cells were treated with TAT-Ctl, TAT-S3, or TAT-pS3 peptide (10 μM, 16 h), followed by treatment with DMSO or ML-792 (0.5 μM, 4 h). Cell lysates were analyzed by IP and WB. e Mitochondrial localization significantly increased for CFL1 S3A and S3E mutants when paired with the K34R mutation (CFL1K34R/S3A-HA, CFL1K34R/S3E-HA), but decreased with the 2KR mutation combined with S3A or S3E mutants (CFL12KR/S3A-HA, CFL12KR/S3E-HA). HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL12KR-HA, CFL1K34R-HA, CFL1S3A-HA, CFL1S3E-HA, CFL12KR/S3A-HA, CFL12KR/S3E-HA, CFL1K34R/S3A-HA, CFL1K34R/S3E-HA or CFL125KR-HA plasmids. His-SUMO1 plasmid was co-transfected. 24 h later, cell lysates were analyzed by IP and WB. f Confocal images of colocalization between CFL1 mutants and mitochondria. HeLa cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL12KR-HA, CFL1K34R-HA, CFL1S3A-HA, CFL1S3E-HA, CFL12KR/S3A-HA, CFL12KR/S3E-HA, CFL1K34R/S3A-HA, CFL1K34R/S3E-HA or CFL125KR-HA plasmids. 24 h later, cells were stained with anti-HA antibody and MitoTracker. The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates; ****p < 0.0001, analyzed by one-way ANOVA with Tukey’s multiple comparisons test. Source data are provided as a Source Data file.
To further establish necessity, we disrupted SUMOylation by generating SUMO1-knockout HeLa cells via CRISPR/Cas9 and silenced SUMO2/3 with siRNAs. CFL1 failed to translocate to mitochondria under these conditions (Fig. 2c). Similarly, pharmacological inhibition of SUMOylation by ML-792, a SUMO E1 (SAE) inhibitor33, prevented CFL1 mitochondrial accumulation despite altered phosphorylation states (Fig. 2d).
To test sufficiency in a phosphorylation-independent context, we combined SUMO-modifying (K34R or 2KR) and phosphorylation-modifying (S3A or S3E) mutations. In CFL1-depleted HEK 293T cells, mitochondrial localization increased in SUMOylation-enhanced mutants (CFL1K34R/S3A and CFL1K34R/S3E) but decreased in SUMOylation-reduced mutants (CFL12KR/S3A and CFL12KR/S3E) (Fig. 2e). Immunofluorescence analyses confirmed these patterns (Fig. 2f). Collectively, these results demonstrate that N-terminal α-amino SUMOylation, rather than Ser3 phosphorylation, is the key determinant of CFL1 mitochondrial translocation. This modification acts as a dominant signal directing CFL1 to mitochondria, independent of its phosphorylation state.
CFL1 translocates to mitochondria through interactions with Tom20, Tom70, and HSP70
Our data demonstrates that mitochondrial translocation of CFL1 depends on N-terminal α-amino SUMOylation rather than phosphorylation (Fig. 2). However, the underlying import mechanism remains unclear. Previous studies have reported that CFL, an actin-depolymerizing factor, localizes to the mitochondrial outer membrane to regulate actin depolymerization and mitochondrial fission16,19,24. Given our prior finding that CFL1 N-terminal α-amino SUMOylation enhances actin depolymerization27, we examined whether mitochondrial translocation of CFL1 depends on its actin-binding activity.
To this end, we generated two constructs: CFL1K96Q-HA, which is deficient in filamentous actin (F-actin) binding34, and CFL1Δ111–119-HA, which was designed to disrupt globular actin (G-actin) binding35,36. Mitochondrial fractionation revealed no significant difference in translocation among CFL1-HA, CFL1K96Q-HA, and CFL1Δ111–119-HA, indicating that mitochondrial translocation is independent of actin binding (Supplementary Fig. 2a). Treatment with cytochalasin D (Cyto D), which disrupts both F-actin and G-actin interactions37,38, abolished CFL1 binding to actin without affecting its mitochondrial localization (Supplementary Fig. 2b). Similarly, immunofluorescence confirmed that CFL1 still colocalized with mitochondria after Cyto D treatment (Supplementary Fig. 2c). These results collectively indicate that CFL1 mitochondrial translocation does not require actin interaction.
The translocase of the outer mitochondrial membrane (TOM) complex mediates the import of most mitochondrial proteins. Tom20 and Tom70 serve as major cytosolic receptors, cooperating with molecular chaperones such as HSP70 to deliver precursor proteins to the general import pore (GIP) complex39,40,41. To determine whether CFL1 utilizes this pathway, we performed co-immunoprecipitation/mass spectrometry (Co-IP/MS) in HEK 293T cells expressing CFL1-HA, which identified interactions with Tom20, Tom70, Tom40, and Tim23 (Supplementary Fig. 3a). We confirmed that endogenous CFL1 associates with Tom20 and Tom70 (Fig. 3a). Knockdown of either receptor decreased CFL1 mitochondrial localization, and dual knockdown completely abolished it (Fig. 3b, c), indicating that both receptors are required for translocation.
a Interaction of CFL1 and Tom20, Tom70. Lysates of HEK 293T cells were subjected to Co-IP with control IgG or anti-CFL1 antibody, which was followed by WB assay. b Knockdown of either Tom20 or Tom70 in HEK 293T cells decreased mitochondrial translocation of CFL1. c Confocal images showed knockdown of either Tom20 or Tom70 in HeLa cells reduced mitochondrial translocation of CFL1. The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates. Statistical significance was determined by one-way ANOVA with Tukey’s multiple comparisons test. The p values were as follows: shCtl vs. shTom20, ***p = 0.0004; shCtl vs. shTom70, ****p < 0.0001; shTom20 vs. shTom70, ***p = 0.0002; shTom70 vs. (shTom20+shTom70), *p = 0.0121. d The K13 and K73 residues on CFL1 were crucial for binding Tom20, while the 9–18 region of CFL1 was essential for binding Tom70. HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL1K13/73A-HA, CFL1Δ9–18-HA or CFL1K73A, Δ9–18-HA plasmids. After 24 h, the cell lysates were subjected to Co-IP using anti-HA beads and WB. e Tom20 binding site mutant (CFL1K13/73A-HA) and Tom70 binding site mutant (CFL1Δ9–18-HA) exhibited reduced CFL1 mitochondrial translocation. The combined Tom20 and Tom70 binding site mutant (CFL1K73A, Δ9–18-HA) completely abolished mitochondrial translocation of CFL1. f Endogenous HSP60, HSC70, HSP70, or HSP90 were knocked down in HEK 293T cells by shRNA Lentiviral. Only the knockdown of HSP70 significantly impaired CFL1 mitochondrial translocation. g HEK 293T cells with endogenous HSP70 knocked down significantly hindered CFL1’s interaction with Tom20 and Tom70. Source data are provided as a Source Data file.
Sequence analysis using MitoFates predicted no cleavable presequence of CFL1, but identified (i) a potential Tom20-binding motif and (ii) a positively charged α-helix (residues 9–18) consistent with Tom70-recognition motifs42,43,44,45,46 (Supplementary Fig. 3b). To test these predictions, we constructed CFL1K13/73A-HA (disrupting Tom20 recognition) and CFL1Δ9–18-HA (disrupting Tom70 recognition). Co-IP demonstrated that CFL1K13/73A lost Tom20 binding, whereas CFL1Δ9–18-HA lost Tom70 binding (Fig. 3d). Mitochondrial fractionation further showed reduced translocation for either single mutant (CFL1K13/73A, CFL1Δ9–18) and complete loss for the double mutant (CFL1K73A, Δ9–18, in which residues 9–18 (including K13) were deleted and K73 was substituted with Ala) (Fig. 3e). These findings establish that CFL1 is recognized by both Tom20 and Tom70 via distinct motifs.
We next explored how cytosolic CFL1 is delivered to these receptors. As molecular chaperones, HSP70/HSP90 maintain precursor proteins in an import-competent state and facilitate delivery to TOM complexes47,48. Co-IP/MS identified interactions between CFL1 and HSP60, HSC70, HSP70, and HSP90 (Supplementary Fig. 3c). Knockdown analysis revealed that only HSP70 depletion markedly reduced CFL1 mitochondrial translocation (Fig. 3f). Furthermore, HSP70 knockdown disrupted CFL1–Tom20/Tom70 interactions (Fig. 3g), demonstrating that HSP70 is essential for linking cytosolic CFL1 to the mitochondrial import machinery. Together, these data reveal that CFL1 translocates to mitochondria through interactions with Tom20 and Tom70, and that HSP70 acts as a critical mediator facilitating this recognition and import process.
Localization of CFL1 in the mitochondrial matrix
Mitochondria comprise four functional compartments: the outer membrane (OM), inner membrane (IM), intermembrane space (IMS), and matrix49. Protein import and compartment-specific localization are critical for mitochondrial homeostasis and cellular function50,51. Although previous studies reported that CFL1 accumulates at the mitochondrial outer membrane in response to apoptotic stimuli13,22,24, its precise sub-mitochondrial localization remains unclear.
Because OM proteins typically contain α-helical or β-barrel transmembrane domains that mediate membrane anchoring52, we first analyzed CFL1 using the TMHMM 2.0 transmembrane prediction tool and found no such domains (Fig. 4a), suggesting that CFL1 is not a membrane protein. Co-IP/MS had previously revealed CFL1 interactions with outer membrane receptors (Tom20, Tom70) and the inner membrane receptor Tim23 (Supplementary Fig. 3a), raising the possibility that CFL1 may be imported into the mitochondrial interior.
a Analysis using the TMHMM 2.0 webserver (https://services.healthtech.dtu.dk/services/TMHMM-2.0/) showed no segments were detected and highlighted in purple of mouse CFL1 sequence, indicated that CFL1 lacks of transmembrane helices. b Sub-mitochondrial localization of CFL1 analyzed by WB. Sub-mitochondrial compartments (OMM, IMS, IMM, and matrix) and Cyto fraction were obtained from HEK 293T cells treated with control or CFL1 shRNA. Tom70, Cyt c, COX IV, and HSP60 are respective markers of the OMM, IMS, IMM, and matrix. c WB analysis following PK protection assay showing that CFL1 was digested by PK only when Triton X-100 treated. Isolated mitochondria were suspended in isotonic mitochondrial buffer or subjected to swelling in hypotonic buffer to induce outer membrane rupture. Alternatively, they were suspended in isotonic mitochondrial buffer containing 0.5% Triton X-100 to disrupt all membranes. The mitochondrial proteins in these buffers were then exposed to PK digestion or not. d APEX2-enhanced brown particle localization in the mitochondrial matrix when transfected with CFL1-APEX2-HA fusion plasmid in CHO-K1 cells. e Interaction of CFL1 and Tim23. Lysates of HEK 293T cells were subjected to Co-IP with control IgG or anti-CFL1 antibody, followed by WB analysis. f Confocal images showed knockdown of Tim23 in HeLa cells reduced mitochondrial translocation of CFL1. The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates; ****p < 0.0001, analyzed by two-tailed Student’s t-test. g Knockdown of Tim23 or simultaneous knockdown of both Tom20 and Tom70 in HEK 293T cells completely prevented mitochondrial translocation of CFL1. Source data are provided as a Source Data file.
To determine its sub-mitochondrial localization, mitochondria and their subfractions were isolated from HEK 293T cells and analyzed by Western blotting using compartment-specific markers: Tom70 (OM), Cyt c (IMS), COX IV (IM), and HSP60 (matrix). CFL1 co-fractionated with the matrix marker HSP60, confirming its presence in the mitochondrial matrix (Fig. 4b). A proteinase K (PK) protection assay further validated this localization. Intact mitochondria were treated with PK under isotonic, hypotonic (outer membrane disrupted), or 0.5% Triton X-100 (inner membrane permeabilized) conditions. CFL1 was resistant to PK digestion unless the inner membrane was solubilized with Triton X-100, mirroring the degradation pattern of matrix protein HSP60 and IM protein COX IV (Fig. 4c). These results confirm that CFL1 resides within the mitochondrial matrix.
To visualize CFL1 distribution at ultrastructural resolution, we employed APEX2-based electron microscopy, which enables in situ peroxidase labeling53. CHO-K1 cells were transfected with APEX2-HA or CFL1-APEX2-HA plasmids. Upon incubation with diaminobenzidine (DAB) and H2O2, followed by osmium tetroxide staining, brown DAB polymers appeared specifically within the mitochondrial matrix of CFL1-APEX2-HA-expressing cells, confirming CFL1’s matrix localization (Fig. 4d).
Because Tim23, a key component of the inner membrane translocase, mediates the import of most matrix-targeted proteins54, we examined whether it is required for CFL1 import. Co-IP revealed an association between CFL1 and Tim23 (Fig. 4e). Lentiviral shRNA-mediated knockdown of Tim23 in HeLa cells reduced colocalization of CFL1 with mitochondria, as shown by immunofluorescence and PCC analysis (Fig. 4f).
Finally, knockdown of Tom20, Tom70, HSP70, or Tim23-individually or in combination, was performed in HEK 293T cells, followed by mitochondrial fractionation. Depletion of either Tom20 or Tom70 reduced CFL1 mitochondrial localization, whereas simultaneous knockdown of both or Tim23 alone completely abolished translocation (Fig. 4g).
Together, these findings demonstrate that CFL1 localizes to the mitochondrial matrix and that its import depends on the coordinated actions of HSP70, Tom20, Tom70, and Tim23.
N-terminal α-amino SUMOylation of CFL1 enhances its mitochondrial translocation through direct binding with HSP70, Tom20, and Tom70
Mitochondrial protein import typically relies on an N-terminal mitochondrial targeting sequence (MTS), which is cleaved by mitochondrial processing peptidase (MPP) upon entry. Recent studies, however, have identified internal MTS-like motifs (iMTS-Ls) that mediate protein recognition by the outer membrane receptor Tom7055,56. Our previous analyses showed that CFL1 lacks a canonical MTS (Supplementary Fig. 3b). To explore whether N-terminal α-amino SUMOylation influences CFL1 import, we analyzed the amino acid sequence of SUMO1-CFL1 fusion protein using the MitoFates prediction tool. MitoFates predicted no cleavable MTS (Supplementary Fig. 4a), but iMLP analysis (http://iMLP.bio.uni-kl.de) revealed that N-terminal SUMOylation markedly increased the iMTS-L propensity score compared with unmodified CFL1 (Supplementary Fig. 4b). These data suggest that N-terminal SUMO modification may act as a mitochondrial targeting signal for CFL1.
To test this hypothesis, we expressed CFL1-HA and CFL1K34R-HA (which exhibits enhanced SUMOylation) in HEK 293T cells with shRNA-mediated knockdown of endogenous CFL1. Co-IP/MS and differential analysis revealed that CFL1K34R displayed significantly stronger interactions with HSP70, Tom20, and Tom70 compared with wild-type CFL1 (Supplementary Fig. 4c), suggesting that N-terminal SUMOylation promotes these interactions.
To verify this relationship under physiological conditions, we analyzed brain tissues from Senp1+/+ and Senp1−/− mouse embryos under both denaturing and non-denaturing conditions. Immunoprecipitation using anti-CFL1 antibody followed by Western blotting showed increased overall SUMOylation, elevated SUMO-modified CFL1, and enhanced interactions with HSP70, Tom20, and Tom70 in Senp1−/− tissues (Fig. 5a). Conversely, pharmacological inhibition of SUMOylation by ML-792 in HEK 293T cells reduced CFL1 SUMOylation and weakened its binding to these proteins (Fig. 5b). Together, these data indicate that CFL1 N-terminal α-amino SUMOylation enhances its interactions with HSP70, Tom20, and Tom70 at the endogenous level.
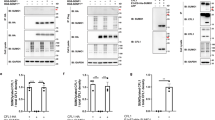
a Senp1−/− mouse embryos exhibited increased CFL1 SUMOylation and enhanced binding of CFL1 to HSP70, Tom20, and Tom70. b HEK 293T cells treated with ML-792 exhibited decreased CFL1 SUMOylation and reduced binding of CFL1 to HSP70, Tom20, and Tom70. c Overexpression of His-SUMO1 in HEK 293T cells exhibited enhanced binding of CFL1 to HSP70, Tom20, and Tom70. In contrast, overexpression of His-SUMO1G97A exhibited decreased binding of CFL1 to HSP70, Tom20, and Tom70. d The binding of CFL1 to HSP70, Tom20, and Tom70 was increased for CFL1 S3A and S3E mutants when paired with the K34R mutation (CFL1K34R/S3A-HA, CFL1K34R/S3E-HA), but decreased with the 2KR mutation combined with S3A or S3E mutants (CFL12KR/S3A-HA, CFL12KR/S3E-HA). HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL12KR-HA, CFL1K34R-HA, CFL1S3A-HA, CFL1S3E-HA, CFL12KR/S3A-HA, CFL12KR/S3E-HA, CFL1K34R/S3A-HA, CFL1K34R/S3E-HA or CFL125KR-HA plasmids. 24 h later, Co-IP with anti-HA beads, followed by WB analysis. e CFL1 SUMOylation promoted its mitochondrial translocation through the binding of CFL1 to Tom20, and Tom70. HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL1K73A, Δ9–18-HA, CFL1K34R-HA, CFL1K34R, K73A, Δ9–18-HA or CFL125KR-HA plasmids. 24 h later, Mito, Cyto, or total lysates fractions were prepared from the cells, followed by WB analysis. f Confocal images of colocalization between CFL1 mutants and mitochondria. HeLa cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL1K73A, Δ9–18-HA, CFL1K34R-HA, CFL1K34R, K73A, Δ9–18-HA or CFL125KR-HA plasmids. 24 h later, cells were stained with anti-HA antibody and MitoTracker. The PCC was presented as the mean ± SEM of six visual fields from 3 biological replicates. Statistical significance was determined by one-way ANOVA with Tukey’s multiple comparisons test. The p values were as follows: CFL1-HA vs. CFL1K73A, Δ9–18-HA, ***p = 0.0007; CFL1K34R-HA vs. CFL1K34R, K73A, Δ9–18-HA, ****p < 0.0001. Source data are provided as a Source Data file.
We next modulated SUMOylation directly by overexpressing His-SUMO1 or the SUMO1G97A mutant in HEK 293T cells. Overexpression of His-SUMO1 strengthened, whereas His-SUMO1G97A weakened, CFL1 binding to HSP70, Tom20, and Tom70 (Fig. 5c). To rule out indirect effects of global SUMOylation changes (e.g., via SENP1 deficiency or ML-792 treatment), we performed targeted mutagenesis of CFL1. Endogenous CFL1 was knocked down and replaced with CFL1 variants exhibiting differential SUMO or phosphorylation states: 2KR (SUMOylation-reduced), K34R (SUMOylation-enhanced), S3A (non-phosphorylatable), S3E (phosphomimetic), and their combinations (e.g., 2KR/S3A, 2KR/S3E, K34R/S3A, K34R/S3E and 25KR). Western blot analysis showed that SUMOylation-reduced variants (2KR, 2KR/S3A, 2KR/S3E and 25KR) exhibited significantly decreased binding to HSP70, Tom20, and Tom70, whereas SUMOylation-enhanced variants (K34R, K34R/S3A and K34R/S3E) displayed increased binding (Fig. 5d). These results confirm that N-terminal α-amino SUMOylation directly strengthens CFL1’s interaction with the mitochondrial import machinery.
To further test whether these interactions are required for translocation, we generated combined mutants with enhanced SUMOylation but disrupted receptor-binding motifs—CFL1K34R, K73A, Δ9–18. Mitochondrial fractionation and Western blotting showed significant loss of CFL1 import in cells expressing CFL1K73A, Δ9–18-HA or CFL1K34R, K73A, Δ9–18-HA (Fig. 5e), confirming that SUMOylation-dependent interactions with Tom20 and Tom70 are essential for mitochondrial entry. Immunofluorescence analysis further verified that these mutants abolished CFL1 mitochondrial localization (Fig. 5f). Collectively, these findings demonstrate that N-terminal α-amino SUMOylation promotes mitochondrial translocation of CFL1 by enhancing its direct binding to HSP70, Tom20, and Tom70, thereby establishing SUMOylation as a key determinant of CFL1 mitochondrial transportation.
N-terminal α-amino SUMOylation of CFL1 promotes mitochondrial pathway-induced apoptosis by regulating Cyt c release
CFL1 has been reported to translocate to mitochondria and regulate Cyt c release during apoptosis, a process previously attributed to Ser3 phosphorylation13. Our findings reveal instead that N-terminal α-amino SUMOylation is the principal determinant of CFL1 mitochondrial import, suggesting that this modification plays a key role in mitochondrial pathway-mediated apoptosis.
To investigate this, HeLa cells were transfected with increasing concentrations (25, 50, or 100 nM) of siRNA targeting endogenous CFL1. Mitochondrial fractions were isolated to assess apoptotic markers and mitochondrial membrane potential. Quantification of the cytosolic-to-total Cyt c ratio (Cyto Cyt c / Total Cyt c) was performed using ImageJ, while caspase-3 activation was analyzed with an antibody recognizing both cleaved (C-Casp3) and pro-Casp3 forms. CFL1 knockdown markedly reduced cytosolic Cyt c levels and decreased the ratio of C-Casp3 to total Casp3 (Fig. 6a), consistent with reduced caspase activation. Western blot analysis confirmed diminished C-Casp3 and C-Casp9 expression and an increase in mitochondrial membrane potential (Fig. 6b and Supplementary Fig. 5a), indicating suppressed mitochondrial apoptosis and attenuated mitochondrial damage.
a Knockdown of CFL1 in HeLa cells reduced Cyt c release and caspase activation. Statistical significance of the cytoplasmic Cyt c to total Cyt c (Cyto + Mito) ratio: siCtl (25 nM) vs. siCtl (100 nM), “ns” (p = 0.8799); siCtl (100 nM) vs. siCFL1 (25 nM), “ns” (p = 0.1115); siCtl (100 nM) vs. siCFL1 (50 nM), **p = 0.0018; siCtl (100 nM) vs. siCFL1 (100 nM), ****p < 0.0001. Significance of the C-Casp3 to total Casp3 (the sum of C-Casp3 and Casp3) ratio: “ns” (p = 0.8454), **p = 0.0014, ****p < 0.0001. b CFL1 knockdown increased mitochondrial membrane potential. After siRNA transfection, HeLa cells grown in black 96-well plates were treated with 50 μM carbonyl cyanide m-chlorophenyl hydrazone (CCCP) for 20 min, and then incubated with tetramethylrhodamine ethyl ester (TMRE) for 30 min at 37 °C. After washing out unbound dye, TMRE fluorescence was measured using a SpectraMax i3X microplate reader (excitation at 550 nm, emission at 575 nm). “ns” (p > 0.999), **p = 0.004, ***p = 0.0002, ****p < 0.0001. c Brain tissue of Senp1−/− mouse embryos exhibited increased Cyt c release and caspase activation. d Senp1−/− MEF cells exhibited decreased mitochondrial membrane potential. ****p < 0.0001. e Expression of SUMOylation enhanced plasmids in HeLa cells increased Cyt c release and caspase activation. f Expression of SUMOylation enhanced plasmids in HeLa cells decreased mitochondrial membrane potential. ***p = 0.0003, ****p < 0.0001. g Expression of SUMO-fused CFL1 in HeLa cells increased Cyt c release and caspase activation. h Expression of SUMO-fused CFL1 in HEK 293T cells decreased mitochondrial membrane potential. ***p = 0.0002, ****p < 0.0001. i Brain tissue of homozygous KI mouse embryos exhibited increased Cyt c release and caspase activation. **p = 0.0055, ***p = 0.0002. One-way ANOVA with Tukey’s multiple comparisons test was used in (a, b, d, f, h). Two-tailed Student’s t-test was used in (i), all data were presented as the mean ± SEM from 3 biological replicates. Source data are provided as a Source Data file.
To test this mechanism in vivo, mitochondrial fractions from Senp1+/+ and Senp1−/− mouse embryos were analyzed. Senp1−/− tissues exhibited lower mitochondrial Cyt c retention and higher cytosolic Cyt c, C-Casp3, and C-Casp9 levels (Fig. 6c), as well as reduced mitochondrial membrane potential in corresponding MEF cells (Fig. 6d). These results suggest that elevated CFL1 SUMOylation in the absence of SENP1 promotes Cyt c release and mitochondrial apoptosis.
To directly assess the role of N-terminal SUMOylation, endogenous CFL1 was knocked down and replaced with SUMOylation-enhanced mutants (CFL1K34R, CFL1K34R/S3A, CFL1K34R/S3E) that exhibit higher mitochondrial localization (Fig. 2e, f). These mutants showed significantly increased Cyt c release and enhanced C-Casp3 and C-Casp9 levels (Fig. 6e), along with decreased mitochondrial membrane potential (Fig. 6f), demonstrating that SUMOylation-enhanced CFL1 augments mitochondrial apoptosis.
Similarly, cells expressing SUMO1-CFL1-HA or SUMO1G97A-CFL1-HA showed elevated Cyt c release, increased caspase activation, and reduced membrane potential compared with CFL1-HA controls (Fig. 6g, h). Hoechst 33258 and propidium iodide staining revealed a higher proportion of dead cells in SUMO-fused CFL1-expressing groups (Supplementary Fig. 5b), further confirming enhanced apoptotic activity driven by N-terminal SUMO modification.
To evaluate physiological relevance in vivo, we generated CFL1K34R knock-in mice using CRISPR/Cas9 gene editing (Supplementary Fig. 5c, d). While heterozygous CFL1WT/K34R mice were viable and fertile, no homozygous CFL1K34R/K34R (KI) offspring were recovered, indicating embryonic lethality (Supplementary Fig. 5e). KI embryos were smaller and developmentally delayed compared with wild-type littermates (Supplementary Fig. 5f). Western blotting of KI embryonic brain tissues revealed increased CFL1 SUMOylation, elevated mitochondrial translocation, enhanced Cyt c release, and higher C-Casp3 levels (Fig. 6i and Supplementary Fig. 5g), confirming that CFL1 N-terminal α-amino SUMOylation potentiates mitochondrial apoptosis in vivo.
N-terminal α-amino SUMOylation of CFL1 regulates Cyt c release through interaction with CYC1
Previous studies have reported that mitochondrial translocation of CFL1 facilitates Cyt c release, thereby activating caspase cascades and promoting apoptosis via the mitochondrial pathway13. However, the precise mechanism by which CFL1 triggers Cyt c release has remained unclear. Cyt c, located in the mitochondrial intermembrane space (IMS), can be released through two principal mechanisms: (i) pore formation in the outer mitochondrial membrane (OMM) via Bax/Bak oligomerization, or (ii) opening of the mitochondrial permeability transition pore (MPTP), which causes membrane rupture and Cyt c efflux into the cytosol57,58. These mechanisms can function independently or cooperatively during apoptosis. It has been proposed that CFL1 interacts with Bax to facilitate its recruitment to the OMM, leading to membrane depolarization and Cyt c release59. Alternatively, oxidation-activated CFL1 may induce MPTP opening and Cyt c release independently of Bax activation22.
To determine whether CFL1 N-terminal α-amino SUMOylation influences Cyt c release through either pathway, we examined Bax dimerization and MPTP opening in HEK 293T cells. Endogenous CFL1 was depleted by lentiviral shRNA, followed by expression of CFL1-HA, SUMO1-CFL1-HA, or SUMO1G97A-CFL1-HA. Bax dimerization and MPTP opening (assessed by calcein-AM fluorescence quenching) were unaffected by CFL1 SUMOylation (Supplementary Fig. 6a, b), indicating that SUMOylated CFL1 does not act through Bax or MPTP regulation. To investigate the mechanism by which N-terminal α-amino SUMOylation of CFL1 regulates Cyt c release, we overexpressed CFL1-HA or CFL1K34R-HA in HEK 293T cells with shRNA-mediated knockdown of endogenous CFL1. Under non-denaturing conditions, Co-IP/MS analysis and differential analysis using a volcano plot revealed that CFL1 interacts with CYC1 and that the N-terminal α-amino SUMO-enhanced CFL1K34R exhibits increased interaction with CYC1 (Supplementary Fig. 6c).
CYC1 mediates electron transfer from cytochrome b to Cyt c in the respiratory chain and modulates Cyt c release during apoptosis60,61. Endogenous co-immunoprecipitation confirmed the CFL1–CYC1 interaction, while no binding was detected between CFL1 and Bax or Cyt c (Fig. 7a). To test whether this interaction regulates Cyt c release, we used siRNA to knock down endogenous CFL1 or CYC1 in HEK 293T cells. CFL1 knockdown reduced the CFL1–CYC1 interaction, increased CYC1–Cyt c binding, and diminished Cyt c release to the cytosol. Conversely, partial knockdown of CYC1 decreased the CYC1-Cyt c interaction and enhanced Cyt c release (Fig. 7b), indicating that the CFL1–CYC1 complex negatively regulates CYC1–Cyt c association. Consistently, knockdown of SUMO1 and SUMO2/3 decreased CFL1 SUMOylation and weakened CFL1–CYC1 binding, while strengthening the CYC1–Cyt c interaction (Fig. 7c). In CFL1K34R knock-in mouse embryos, where SUMOylation is enhanced, CFL1–CYC1 binding increased, accompanied by reduced CYC1–Cyt c interaction and elevated Cyt c release (Fig. 7d).
a Co-IP experiments in HEK 293T cells exhibited the interaction between CFL1 and CYC1, as well as the interaction between CYC1 and Cyt c. b Knockdown of CFL1 in HEK 293T cells inhibited the interaction between CFL1 and CYC1, leading to increased binding of CYC1 to Cyt c and reduced Cyt c release. Statistical significance of the CFL1–CYC1 interaction: “ns” (p > 0.9999), ***p = 0.0001, ****p < 0.0001; CYC1–Cyt c interaction: “ns” (p > 0.9999), **p = 0.0017; cytoplasmic Cyt c to total Cyt c (Cyto + Mito) ratio: “ns” (p > 0.9999), ****p < 0.0001. All data were presented as the mean ± SEM from 3 biological replicates and analyzed by one-way ANOVA with Tukey’s multiple comparisons test. c CFL1 deSUMOylation impaired the interaction between CFL1 and CYC1. WT or Sumo1 KO HeLa cells were transfected with either control siRNA or siRNA complex targeting SUMO2 and SUMO3. 48 h later, cell lysates were subjected to Co-IP and WB. d Homozygous KI embryos exhibited increased binding of CFL1 to CYC1, decreased binding of CYC1 to Cyt c, and increased release of Cyt c. e Predictions based on AlphaFold3 and visual comparisons using PyMOL software demonstrated structure and spatial relationship between the CFL1 and CYC1. f The CFL1 K126A mutation impaired the binding between CFL1 and CYC1, while increasing the binding of CYC1 to Cyt c. HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL1K22A-HA, CFL1K126A-HA, CFL1K19A-HA, CFL1K132A-HA, CFL1V20A-HA, CFL1K96A-HA, or CFL1K95A-HA plasmids. 24 h later, cell lysates were subjected to Co-IP and WB. g CFL1 SUMOylation regulates Cyt c release through its interaction with CYC1. HEK 293T cells with endogenous CFL1 knocked down were transfected with CFL1-HA, CFL1K34A-HA, SUMO1-CFL1-HA, SUMO1G97A-CFL1-HA, CFL1K126A-HA, CFL1K34R, K126A-HA, SUMO1-CFL1K126A-HA or SUMO1G97A-CFL1K126A-HA plasmids. 24 h later, cell lysates were subjected to Co-IP and WB. Source data are provided as a Source Data file.
To probe the binding interface, we used AlphaFold3 structural modeling (Fig. 7e) to predict the interaction surface between CFL1 and CYC1. Guided by this model, we generated site-directed mutants of CFL1 in which putative CYC1-binding residues were substituted with alanine. In CFL1-depleted HEK 293T cells, mutating Lys126 (CFL1K126A) disrupted the CFL1–CYC1 interaction and enhanced the CYC1–Cyt c association (Fig. 7f). Furthermore, combining the K126A mutation with SUMOylation-enhanced constructs (CFL1K34R, SUMO1-CFL1, SUMO1G97A-CFL1) abolished the SUMO-dependent interaction with CYC1, increased CYC1–Cyt c binding, and reduced Cyt c release (Fig. 7g).
Collectively, these findings demonstrate that N-terminal α-amino SUMOylation of CFL1 promotes mitochondrial apoptosis by enhancing its interaction with CYC1, which in turn disrupts the CYC1–Cyt c complex, facilitating Cyt c release into the cytosol. This mechanism defines a SUMO-dependent pathway of CFL1-mediated mitochondrial apoptosis, distinct from the classical Bax- or MPTP-dependent model.
Discussion
Our study identifies N-terminal α-amino SUMOylation of CFL1 as a pivotal determinant of its mitochondrial translocation and apoptosis induction. Unlike the classical phosphorylation-dependent model, where Ser3 dephosphorylation governs CFL1 activity and mitochondrial targeting, our data reveal that SUMOylation alone is sufficient to drive mitochondrial import, independent of phosphorylation (Fig. 2). Mechanistically, SUMOylated CFL1 interacts directly with HSP70, Tom20, and Tom70, which facilitate its import into the mitochondrial matrix through the TOM/TIM machinery (Fig. 5). Once inside the matrix, SUMO-modified CFL1 binds to CYC1 to promote Cyt c release and apoptosis (Figs. 6, 7).
CFL has long been viewed as an actin-binding protein that regulates cytoskeletal turnover by severing F-actin and recycling G-actin1,2,12. Yet accumulating evidence indicates that CFL also exerts actin-independent functions, particularly in mitochondrial dynamics and apoptosis8. Early studies suggested that dephosphorylated CFL1 associates with Drp1 and F-actin at mitochondria-associated membranes (MAMs), modulating mitochondrial fission and apoptotic sensitivity16,21. Our findings refine this concept: even when actin binding is disrupted, SUMO-modified CFL1 is still efficiently imported into mitochondria, revealing an actin-independent route (Fig. 1 and Supplementary Fig. 2). This expands the functional landscape of CFL1 from a cytoskeletal modulator to a signal-responsive mitochondrial effector that integrates post-translational inputs to control cell fate.
Among the three ADF/CFL family members (CFL1, CFL2, and ADF), each exhibits distinct expression and physiological specialization. CFL1 is ubiquitous and essential for embryogenesis and neuronal morphogenesis62, CFL2 is enriched in muscle tissue and linked to congenital myopathies63, and ADF is implicated in epithelial cell migration and cancer metastasis64. The discovery that CFL1 undergoes N-terminal α-amino SUMOylation raises the possibility that this modification could confer family-specific regulation. Given that ADF/CFL, contains residues compatible with SUMO conjugation and Tom20/Tom70 binding motifs, N-terminal SUMOylation may represent an evolutionary innovation that equips ADF/CFL with a unique mitochondrial signaling function. Comparative studies across the ADF/CFL family will be essential to determine whether similar SUMO-mediated pathways operate in other contexts, such as muscle homeostasis or epithelial remodeling.
At the cellular level, CFL1 N-terminal SUMOylation functions as a molecular switch linking cytoskeletal dynamics to mitochondrial fate27. By enhancing CFL1 translocation into mitochondria, this modification amplifies Cyt c release, caspase-3/9 activation, and mitochondrial membrane depolarization (Figs. 1, 6). These results position SUMOylated CFL1 as a pro-apoptotic mediator acting independently of actin disassembly. Conceptually, N-terminal SUMOylation can be viewed as a mitochondrial targeting license that allows newly synthesized CFL1 molecules—before N-terminal methionine excision and acetylation are complete—to enter mitochondria and engage the apoptotic machinery.
Recent transcriptomic and clinical studies consistently show that elevated SENP1 expression correlates with higher tumor grade and poor prognosis in breast, renal, and prostate cancers65,66,67. SENP1 promotes tumorigenesis by de-SUMOylating substrates that normally restrain proliferation or promote apoptosis68,69. Our findings provide a mechanistic explanation for this correlation: excessive SENP1 activity would de-SUMOylate CFL1, thereby preventing its mitochondrial translocation and reducing apoptosis sensitivity in tumor cells. Conversely, restoring CFL1 SUMOylation or pharmacologically inhibiting SENP1 could resensitize cancer cells to intrinsic apoptotic cues. This interpretation aligns with the growing view that impaired SUMO homeostasis contributes to oncogenic survival pathways. Thus, the SENP1-CFL1 axis represents a potential therapeutic target linking post-translational regulation of cytoskeletal proteins to mitochondrial apoptosis and cancer progression.
Beyond oncology, CFL1 SUMOylation may influence other conditions where mitochondrial integrity and apoptosis are critical—such as neurodegeneration, ischemic injury, and metabolic disorders. Neurons, cardiomyocytes, and adipocytes all rely heavily on mitochondrial quality control, and CFL1 is abundantly expressed in these tissues. For instance, deregulated CFL1 activity has been implicated in Parkinson’s disease and ischemic neuronal death, contexts where mitochondrial stress and SENP1 up-regulation coexist70,71,72,73,74. It will be of interest to determine whether CFL1 N-terminal SUMOylation acts as a general sensor of cellular stress, dynamically linking cytoskeletal cues to mitochondrial responses across diverse pathophysiological settings.
In summary, our study establishes N-terminal α-amino SUMOylation as a post-translational modification that governs CFL1 mitochondrial targeting and apoptosis initiation. SUMOylated CFL1 engages HSP70, Tom20, and Tom70 for import into the mitochondrial matrix, where it interacts with CYC1 to displace Cyt c and activate the intrinsic apoptotic pathway (Supplementary Fig. 7). This mechanism operates independently of Ser3 phosphorylation or actin binding, revising the long-standing paradigm of CFL1 function. The integration of SUMO signaling with mitochondrial trafficking unveils a broader principle of cellular regulation, in which dynamic post-translational modifications on cytoskeletal proteins determine organelle behavior and cell fate. Future studies should investigate how CFL1 SUMOylation is modulated by metabolic or oxidative cues, and whether targeting the SENP1-CFL1 axis could provide therapeutic leverage in cancers and degenerative diseases driven by defective apoptosis.
Methods
Experimental animals
The mice were housed under conditions featuring a 12-h light/12-h dark cycle, with ad libitum access to food and water. The mouse strains used in this study were generated and maintained on the C57BL/6 background. The research included Senp1+/+ and Senp1−/− mice75. Embryos were removed from anaesthetized Senp1 heterozygous mice at embryonic day 13.5 (E13.5). CFL1K34R/K34R (KI) mice were generated by CRISPR/Cas9-mediated genome editing (Gempharmatech Co., Ltd., Nanjing, China). In brief, the target sequences of guide RNAs (gRNAs) for the CRISPR/Cas9 system were designed to the regions on intron 1 and intron 2 to promote DNA breaks and homologous recombination. The fragments for CFL1 gRNA1 (TGGAGATGAGACTATGGCCCAGG) and gRNA2 (GAGACTATGGCCCAGGAGTTTGG) were inserted into the gRNA cloning vector. The K34R (AAG to AGG) mutation site in the donor oligo was designed and synthesized. The in vitro-transcribed nuclease Cas9 mRNA (400 ng/μL), sgRNA (200 ng/μL), and the oligo donor (20 ng/μL) carrying the desired mutation (AAG to AGG) were co-injected into fertilized eggs for KI mice production. The genotype of mice was confirmed by PCR followed by sequence analysis using primers (CFL1 K34R-F: AATCTGTGTGGGGCTGAGAAGTGT; CFL1 K34R-R: CCTGAATACGGAGCTCACCAGA).
Cell culture and transfection
CHO-K1 cells (GNHa 7), HEK 293T cells (GNHu17), HeLa cells (TCHu187) were purchased from Cell Bank/Stem Cell Bank, Chinese Academy of Sciences (Shanghai, China). MEF cells were provided by J. Cheng (Shanghai Jiao Tong University School of Medicine, Shanghai, China). CHO-K1 cells were grown in Ham’s F12 medium containing 10% fetal bovine serum (FBS, Life Technologies, USA), and 100 μg/mL of penicillin/streptomycin (PS, BasalMedia, Shanghai, China). HEK 293T cells, HeLa cells, and MEF cells were grown in DMEM (high glucose) supplemented with 10% FBS and 100 μg/mL of PS. All cell cultures were maintained in a controlled environment with a humidified atmosphere of 5% CO2 at a temperature of 37 °C. Cells grown to ~80% confluence were transfected with the plasmids using Lipofectamine 3000 (L3000015, Invitrogen) or 1 mg/mL polyethylenimine (PEI, 23966, Polysciences) according to manufacturers’ instructions. Cells grown into ~60% confluence were transfected with the siRNA using Lipofectamine 3000.
Antibodies and reagents
Anti-CFL1 (66057-1-Ig, 1:1000) and anti-CFL1 (10960-1-AP, 1:1000) were purchased from Proteintech. Anti-SUMO1 (4940, 1:1000), anti-SAE1 (13585, 1:1000), anti-SAE2 (8688, 1:1000), anti-SENP3 (5591, 1:1000), anti-phospho-CFL1 (Ser3) (77G2, 1:1000), anti-pan-actin (4968, 1:1000), anti-HSP60 (12165, 1:1000), anti-Cyt c (4280, 1:1000), anti-Casp3 (9662, 1:1000), anti-C-Casp3 (9664, 1:1000), anti-C-Casp9 (7237, 1:1000), anti-Bax (2772, 1:1000) and anti-GAPDH (2118, 1:5000) were purchased from Cell Signaling Technology. Anti-HA (H6908, 1:3000) and anti-SUMO2/3 (SAB3500488, 1:1000) were purchased from Sigma-Aldrich. Anti-Ubc9 (ab33044, 1:1000), anti-SENP1 (ab108981, 1:1000), anti-SENP2 (ab58418, 1:1000) and anti-COX IV (ab33985, 1:1000) were purchased from Abcam. Anti-Tom20 (sc-36211, 1:200), anti-Tom70 (sc-390545, 1:200), anti-Tim23 (sc-514463, 1:200) and anti-HSC70 (sc-7298, 1:200) were purchased from Santa Cruz. Anti-HSP70 (T55496, 1:1000), anti-HSP90 (M20032, 1:1000), and anti-CYC1 (PH1515, 1:1000) were purchased from Abmart (Shanghai, China). Alexa Fluor 488-conjugated goat anti-mouse antibody (A11029), Alexa Fluor 633-conjugated goat anti-rabbit antibody (A21052), and MitoTracker Red CMXRos (M7512) were purchased from Invitrogen. Anti-HA agarose (26181), protein A/G agarose (20422), and disuccinimidyl suberate (DSS, 21555) were purchased from Thermo Fisher Scientific. Protease inhibitor cocktail (B14001) and phosphatase inhibitor cocktail (B15002) were purchased from Bimake. N-ethylmaleimide (NEM, E3876, 20 mM) and Percoll (GE17-0891-01) were purchased from Sigma-Aldrich. 3,3’-Diaminobenzidine (DAB, A690009) was purchased from Sangon Biotech (Shanghai, China). Cytochalasin D (BML-T109-0001) was purchased from Enzo Life Sciences. Latrunculin B (Lat B, MZ5810) was purchased from MKbio (Shanghai, China). ML-792 (S8697), MG132 (S2169), and MLN4924 (S7109) were purchased from Selleckchem. Polybrene (40804ES76), Hoechst 33258 (40729ES10,) and recombinant human HSP70 protein (95197ES25) were purchased from Yeasen Biotechnology (Shanghai, China). Glutaraldehyde (AS1063) was purchased from ASPEN. EMBed 812 (90529-77-4) was purchased from SPI. Propidium Iodide (PI, ST511), blasticidin S HCl (ST018), and mitochondrial membrane potential assay kit with TMRE (C2001S) were purchased from Beyotime Biotechnology (Shanghai, China). Mitochondrial permeability transition pore assay kit (KTA4002) was purchased from Abbkine Scientific Co., Ltd (Wuhan, China). TAT-Ctl peptide (RKKRRQRRR), TAT-S3 peptide (RKKRRQRRR-MASGVAVSDGVIKVFN), and phosphorylated TAT-pS3 peptide (RKKRRQRRR-MAS(p)GVAVSDGVIKVFN) were synthesized by Sangon Biotech (Shanghai, China).
Plasmids and RNA interference
Mouse CFL1 cDNA was amplified by PCR from mouse brain tissues and inserted into the pCMV-HA-C (Clontech) vector to obtain the CFL1-HA plasmid. SUMO1-CFL1-HA, SUMO1G97A-HA, APEX2-HA, and CFL1-APEX2-HA fusion plasmids were constructed by Tsingke Biotechnology (Beijing, China). Various CFL1-HA mutations: CFL1K112/114R-HA (CFL12KR-HA), CFL1K34R-HA, CFL1S3A-HA, CFL1S3E-HA, CFL12KR/S3A-HA, CFL12KR/S3E-HA, CFL1K34R/S3A-HA, CFL1K34R/S3E-HA, CFL125KR-HA, CFL1K96Q-HA, CFL1Δ111–119-HA, CFL1K13/73A-HA, CFL1Δ9–18-HA, CFL1K73A, Δ9–18-HA, CFL1K34R, K73A, Δ9–18-HA, CFL1K22A-HA, CFL1K126A-HA, CFL1K19A-HA, CFL1K132A-HA, CFL1V20A-HA, CFL1K96A-HA, or CFL1K95A-HA, CFL1K34R, K126A-HA, SUMO1-CFL1K126A-HA and SUMO1G97A-CFL1K126A-HA plasmids were generated using the KOD Plus mutagenesis kit (TOYOBO). His-SUMO1, His-SUMO1G97A plasmids were previously described76. All siRNAs were synthesized by Sangon Biotech (Shanghai, China), including control siRNA (5’- UUCUCCGAACGUGUCACGU-3’), CFL1 siRNA (5’-UGCCAACUUCUAACCGCAAUA-3’), SUMO2 siRNA (5’GACUGAGAACAACGAUCAUAU-3’), SUMO3 siRNA (5’-CAGUUATGUUGUCGUGUAU-3’, 5’-CAGAGAAUGACCACAUCAA-3’), si-CFL1 (5’-UGCCAACUUCUAACCGCAAUA-3’) and CYC1 siRNA (5’-CAGAUGUCUUAGAGUUUGA-3’, 5’-CCAGGGAAGCUGUUCGACUAU-3’).
shRNA knockdown experiments
Primers for shRNA were designed following the manufacturer’s instructions and cloned into the pLKO.1-blast Vector (Addgene), including control shRNA (targeting sequence: TTCTCCGAACGTGTCACGT), CFL1 shRNA (targeting sequence: TGCCAACTTCTAACCGCAATA), Tom20 shRNA (targeting sequence: CAAGTTACCTGACCTTAAA), Tom70 shRNA (targeting sequence: GCATGCTGTTAGCCGATAAAG), Tim23 shRNA (targeting sequence: CCAGCCTCTATGCACTATATA), HSP60 shRNA (targeting sequence: AGTCAAGGCTCCAGGGTTT), HSC70 shRNA (targeting sequence: GCCCGATTTGAAGAACTGAA), HSP70 shRNA (targeting sequence of HSPA1A/HSP70-1: GGCCAACAAGATCACCATC; targeting sequence of HSPA1B/HSP70-2: CACAAGAAGGACATCAGCC) and HSP90 shRNA (targeting sequence of HSP90AA1: GGCCAACAAGATCACCATC; targeting sequence of HSP90AB1: CACAAGAAGGACATCAGCC). Lentiviral vectors were co-transfected with psPAX2 and pMD2.G into HEK 293T cells. After 48 h, virus supernatant was collected. Cells were incubated in virus supernatant and polybrene (5 μg/mL) for 48 h. Stable knockdown stable cells were selected with blasticidin S HCl.
Immunoprecipitation and Western blotting
The denaturing IP and non-denaturing Co-IP differed in the lysis buffer and the procedure of making cell lysates27,76. For denaturing IP, cells and tissues were lysed in SDS lysis buffer containing 50 mM Tris-HCl (pH 6.8), 2% SDS, 40 mM DTT, and 5% glycerol. The lysates were boiled for 10 min at 95 °C to denature proteins and then diluted fivefold with Triton X-100 lysis buffer (20 mM NEM, 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% Triton X-100, 2 mM EDTA, 10% glycerol) with protease inhibitor cocktail, phosphatase inhibitor cocktail, and MEM. For non-denaturing Co-IP, cells and tissues were homogenized in Triton X-100 lysis buffer for 2 h. Following centrifugation at 13,000 g at 4 °C for 15 min, the supernatant was transferred to a new tube. The supernatant was then incubated with anti-HA beads for 4 h, or with anti-CFL1 antibody together with protein A/G beads overnight at 4 °C. After incubation, the beads were centrifuged at 3000 g for 1 min at 4 °C to precipitate the immunocomplexes. The beads were washed four times with Triton X-100 lysis buffer before being boiled in 1× SDS loading buffer at 95 °C, which released the proteins for SDS-PAGE analysis.
Protein samples were separated on 12% SDS-polyacrylamide gels and subsequently transferred onto PVDF membranes for further analysis. To minimize non-specific binding, the membranes were blocked by incubating them for 1 h at room temperature in a solution of 5% non-fat dry milk dissolved in Tris-buffered saline (TBS) containing 0.1% Tween-20. After blocking, the membranes were incubated with primary antibodies specific to the target proteins overnight at 4 °C, allowing the antibodies to bind to their respective antigens on the membrane. Following primary antibody incubation, the membranes were washed thoroughly and then incubated with suitable secondary antibodies for 1 h at room temperature. The secondary antibodies were conjugated to an enzyme, facilitating detection. Protein bands were visualized using an enhanced chemiluminescence (ECL) system (ImageQuant LAS 4000 Mini), according to the manufacturer’s protocol, with the appropriate ECL detection reagents. The presented data are representative of at least three independent experiments.
Immunoprecipitation-Mass spectrometry
To identify the SUMOylation site of CFL12KR and CFL1K34R, HEK 293T cells with CFL1 knockdown were transfected with either CFL12KR-HA or CFL1K34R-HA along with His-SUMO1E93R plasmids. 24 h later, followed by denaturing IP. After 4 h incubation at 4 °C, the beads were washed four times with Triton X-100 lysis, followed by 5 mM NH4HCO3 twice. Proteins were eluted from the beads with 0.15% trifluoroacetic acid (T6508, Sigma-Aldrich). After centrifugation at 4 °C for 15 min at 15,870 g, the supernatant was collected and dried in a vacuum. The dried samples were redissolved in 25 mM NH4HCO3 followed by digestion with trypsin (Promega) at 37 °C overnight, and were dried before LC-MS/MS analysis. For LC-MS/MS, the samples were resuspended with 20 mL buffer (water with 0.1% formic acid) and analyzed by online nanospray LC-MS/MS on a Q Exactive HF (Thermo Fisher Scientific) coupled to an Acquity UPLC M-class (Waters Corporation). Tandem mass spectra were processed by PEAKS Studio version X (Bioinformatics Solutions Inc.). PEAKS DB was set up to search the Mus musculus proteome database from UniProt (version 201907, 22290 entries), assuming trypsin as the digestion enzyme. PEAKS DB was searched with a fragment ion mass tolerance of 0.02 Da and a parent ion tolerance of 7 ppm. QTGG modifications were specified as the variable modifications. The peptides with −10lgP ≥ 20 and the proteins with −10lgP ≥ 20 and containing at least one unique peptide were filtered.
Protein-protein interactions were investigated by non-denaturing Co-IP and mass spectrometry assay (Co-IP/MS)77,78. To this end, HEK 293T cells with CFL1 knockdown were transfected with either CFL1-HA or CFL1K34R-HA plasmids. 24 h later, cells were lysed in Triton X-100 buffer containing protease inhibitor cocktail, phosphatase inhibitor cocktail, and MEM. After centrifugation at 13,000 g, 4 °C for 15 min, the supernatant was carefully collected and incubated with anti-HA beads for 4 h at 4 °C. Following incubation, the beads were washed three times with the initial Triton X-100 lysis buffer, and then twice with 5 mM NH4HCO3. Proteins were then eluted from the beads using 0.15% trifluoroacetic acid (TFA) and the eluted proteins were dried under vacuum to prepare them for LC-MS/MS analysis. For LC-MS/MS, peptides derived from the eluted proteins were meticulously analyzed. Tandem mass spectra obtained were processed using PEAKS Studio version X+ software (Bioinformatics Solutions Inc.), utilizing specific search parameters: a fragment ion mass tolerance of 0.02 Da and a parent ion tolerance of 7 ppm. PEAKS DB was set up to search the Homo sapiens proteome database from UniProt (version 2022, 20610 entries). The criteria for considering peptides and proteins significant were stringent: only those with a 10lgP score of ≥ 20 and proteins with a −10lgP score of ≥ 20, and containing at least one unique peptide, were included in the final dataset. The mass spectrometry data for each group were derived from three independent biological replicates.
Isolation of pure mitochondria and sub-mitochondrial compartments
Cells or tissues were homogenized and then centrifuged at 750 g for 5 min, repeated twice, to remove unbroken cells and nuclear contaminants. The supernatant was subsequently centrifuged at 10,000 g for 5 min to obtain the cytosolic fraction and the crude mitochondrial pellet. The crude mitochondria were layered onto 10 mL of 30% Percoll buffer in a 14 mL ultracentrifugation tube and centrifuged at 96,000 g for 30 min using a SW 40Ti rotor (Beckman). The resulting gradient typically contained two distinct white bands. The top band, corresponding to the MAMs fraction, and the bottom band, representing the pure mitochondria, were collected separately. The pure mitochondria were then diluted with mitochondria resuspension buffer (MRB) and subjected to repeated ultracentrifugation to further purify the mitochondrial fraction79.
To isolate sub-mitochondrial compartment fractionation, pure mitochondria were suspended in isolation medium (225 mM mannitol, 75 mM sucrose, 0.1 mM EGTA, 20 mM HEPES, pH 7.4) containing 0.12 mg digitonin per mg of mitochondria and gently stirred on ice for 15 min. After digitonin treatment, the samples were diluted with three volumes of isolation medium and centrifuged at 9000 g for 10 min to isolate the supernatant (supernatant A) and the pellet (pellet A). Supernatant A was ultracentrifuged at 144,000 g for 60 min to collect the pellet (outer membrane vesicles), and the remaining supernatant (inner membrane space fraction). Pellet A was resuspended in isolation buffer and sonicated for 30 sec on ice. The sonicated solution was then centrifuged at 6500 g for 10 min, followed by ultracentrifugation at 144,000 g for 60 min to collect the pellet (inner membrane vesicles) and the supernatant (matrix fraction)80.
Proteinase K protection assay
Pure mitochondria were resuspended in either isotonic buffer (50 mM mannitol, 0.5 mM EGTA, 5 mM HEPES, pH 7.4), hypotonic buffer (10 mM KCl, 2 mM HEPES, pH 7.2), or an isotonic solution containing 0.5% Triton X-100. PK was added at a final concentration of 7 μg/mL and incubated at 4 °C for 20 min. The reaction was then inhibited by adding 5 mM PMSF on ice for 15 min. Next, 1 mL of 10% trichloroacetic acid was added to precipitate the protein. The homogenates were centrifuged at 4 °C, 22,000 g for 10 min. The supernatant was discarded, and the protein pellet was washed twice with pre-cooled acetone. Afterward, the pellet was air-dried. The protein was resuspended in 20 μL PBS, and its concentration was measured using the BCA assay. WB analysis was then performed81.
Electron microscopy
CHO-K1 cells were transfected with either APEX2-HA or CFL1-APEX2-HA fusion plasmid fusion plasmid. After 24 h, cells were fixed using room temperature 2% glutaraldehyde 30 min, cells were rinsed five times for 2 min by PBS, then treated for 5 min in PBS containing 20 mM glycine to quench unreacted glutaraldehyde, followed by another five 2-min rinses by PBS. A freshly diluted solution of 0.5 mg/mL (1.4 mM) DAB was combined with 10 mM H2O2 in chilled buffer, and the solution was added to cells for 20 min. To halt the reaction, the DAB solution was removed, and cells were rinsed 5 × 2 min with PBS. Post-fixation staining was performed with 2% (w/v) osmium tetroxide for 30 min in chilled buffer. Cells were rinsed five times for 2 min each in chilled distilled water, then placed in chilled 2% (w/v) uranyl acetate in ddH2O overnight53,82. To improve the interpretability of APEX-EM labeling, we optimized the staining protocol by adding a lead citrate staining step after uranyl acetate, this modification enhanced the contrast of mitochondrial membranes and matrix structures83,84. Cells were brought to room temperature, washed in distilled water, then carefully scraped off the plastic, resuspended, and centrifuged at 700 g for 1 min to generate a cell pellet. The supernatant was removed, and the pellet was dehydrated in a graded ethanol series (50%, 70%, 80%, 90%, 95%, 100%, 100%), for 15 min each time, then infiltrated in EMBed 812 using 1:1 (v/v) resin and acetone overnight. Finally, the sample was exchanged once more with 100% resin before transfer to fresh resin and polymerization at 60 °C for 48 h. Embedded cell pellets were cut with a diamond knife into 60–80 nm sections and imaged on a JEOL JEM-1400Flash transmission electron microscope.
Isolation of G-actin and F-actin
HEK 293T cells were plated on 60 mm dishes. After 24 h of plasmid transfection, gently washed the adherent cells twice with ice-cold PBS and added 2 mL of F-actin stabilizing buffer (50 mM PIPES, 50 mM NaCl, 5 mM MgCl2, 5 mM EGTA, 1 mM ATP, 5% glycerol, 1% Triton X-100, 0.01% anti-foaming agent, pH 7.5, protease inhibitor cocktail, phosphatase inhibitor cocktail, and NEM). Using a syringe with a 25-gauge needle to repeatedly aspirate the cell lysate 20 times, and incubation on ice for 20 min to lysate the cells. Homogenates were centrifuged at 4 °C, 500 g for 5 min to remove nuclear contaminants and unbroken cells. Transferred the supernatant to an ultracentrifugation tube and centrifuged at 10,000 g for 1 h using a TLA 55 rotor (Beckman)85. The supernatant (G fraction) contains free, globular G-actin, while the pellet (F fraction) primarily contains F-actin and F-actin binding proteins.
Fluorescence staining
Cells were stained with 200 nM MitoTracker Red CMXRos for 50 min at 37 °C, and then fixed with 4% paraformaldehyde for 10 min86. After permeabilized, and blocked for 1 h at room temperature with blocking buffer (1× PBS with 10% normal goat serum and 0.3% Triton X-100), cells were rinsed three times with 0.01 M PBS (pH 7.4), incubated with the anti-CFL1 (1:100), anti-HA (1:500), or anti-SUMO1 (1:100) antibody overnight at 4 °C, washed thrice for 10 min, and incubated with Alexa Fluor 488-conjugated goat anti-mouse antibody (1:500) or Alexa Fluor 633-conjugated goat anti-rabbit antibody (1:500) for 1 h at room temperature. Cells were rinsed three times with PBS and then stained with DAPI. Confocal images were captured by a Leica SP8 confocal microscope (Leica) with a 63× oil objective.
For the Hoechst/PI double staining apoptosis assay, HEK 293T cells were seeded on 24-well plates and treated with 1 μg/mL of PI and 1 μg/mL of Hoechst. After incubation for 15 min at 37 °C under 5% CO2. Removed the dyes and washed the cells three times with PBS87. Images were taken under an Olympus 1×71 fluorescence microscope using a 10× objective lens.
Calcein fluorescence was measured in living cells using the mitochondria permeability transition pore assay kit following the protocol described88. Briefly, HEK 293T cells were seeded on 24-well plates and loaded with calcein AM (1 µM), MitoTracker Red CMXRos (200 nM), treated with or without CoCl2 (2 mM) in assay buffer, and incubated for 50 min at 37 °C under 5% CO2. After incubation, replace calcein loading solution with fresh culture medium preheated at 37 °C, and incubated for 30 min at 37 °C. Cells were then washed with PBS, and images were taken under an Olympus 1 × 71 fluorescence microscope using a 20× objective lens (excitation/emission wavelengths of 494/517 nm).
Statistical analyses
Cell counting from Hoechst/PI double staining and fluorescence intensity of calcein AM were quantified using ImageJ. Pearson’s correlation coefficient was calculated to quantify colocalization between CFL1 and mitochondria by using the Coloc 2 Plugin for ImageJ (https://imagej.net/plugins/coloc-2), which assessed colocalization between two channels on a pixel-by-pixel basis. We employed a bisection threshold regression and 1000 randomizations with a PSF of 3.0. The Pearson’s correlation coefficient was taken for each of the six fields of view. Statistical analyses in our research were performed using GraphPad Prism software, version 8.0.2. For thoroughness and accuracy, all data are represented as mean ± standard error of the mean (SEM), based on at least three independent experiments. No data were excluded from the analyses. The experiments were randomized. The investigators were blinded to group allocation during data collection and analysis. When data were normally distributed, the two-tailed Student’s t-test was utilized for comparisons between two groups. One-way ANOVA, accompanied by Tukey’s multiple comparison test was employed for analyzing differences among more than two groups. Throughout this study, p-value < 0.05 was deemed statistically significant, ensuring the reliability of the findings reported.
Ethical statement
The research presented in this study adheres to the ethical regulations at Shanghai Jiao Tong University School of Medicine. All experimental protocols with animals were carried out in accordance with the guidelines for the Care and Use of Laboratory Animals of Shanghai Jiao Tong University School of Medicine and approved by the Institutional Animal Care and Use Committee (IACUC) under Approval No. A-2022-040.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry data that support the findings of this study have been deposited in iProX with the accession code IPX0014133000 and proteomeXchange with accession code PXD070400. The data generated in this study are provided in the Supplementary Information and Source Data file. Source data are provided with this paper.
References
Ono, S., Minami, N., Abe, H. & Obinata, T. Characterization of a novel cofilin isoform that is predominantly expressed in mammalian skeletal muscle. J. Biol. Chem. 269, 15280–15286 (1994).
Hamill, S., Lou, H. J., Turk, B. E. & Boggon, T. J. Structural basis for noncanonical substrate recognition of cofilin/ADF proteins by LIM kinases. Mol. Cell 62, 397–408 (2016).
Sexton, J. A. et al. Distinct functional constraints driving conservation of the cofilin N-terminal regulatory tail. Nat. Commun. 15, 1426 (2024).
Gandhi, M., Achard, V., Blanchoin, L. & Goode, B. L. Coronin switches roles in actin disassembly depending on the nucleotide state of actin. Mol. Cell 34, 364–374 (2009).
Advedissian, T., Frémont, S. & Echard, A. Cytokinetic abscission requires actin-dependent microtubule severing. Nat. Commun. 15, 1949 (2024).
Beck, H. et al. Serum Response Factor (SRF)-cofilin-actin signaling axis modulates mitochondrial dynamics. Proc. Natl. Acad. Sci. USA. 109, E2523–E2532 (2012).
Hoffmann, L., Rust, M. B. & Culmsee, C. Actin(g) on mitochondria - a role for cofilin1 in neuronal cell death pathways. Biol. Chem. 400, 1089–1097 (2019).
Xu, J. et al. Cofilin: a promising protein implicated in cancer metastasis and apoptosis. Front. Cell Dev. Biol. 9, 599065 (2021).
Yang, N. et al. Cofilin phosphorylation by LIM-kinase 1 and its role in Rac-mediated actin reorganization. Nature 393, 809–812 (1998).
Ambach, A. et al. The serine phosphatases PP1 and PP2A associate with and activate the actin-binding protein cofilin in human T lymphocytes. Eur. J. Immunol. 30, 3422–3431 (2000).
Toshima, J., Toshima, J. Y., Takeuchi, K., Mori, R. & Mizuno, K. Cofilin phosphorylation and actin reorganization activities of testicular protein kinase 2 and its predominant expression in testicular Sertoli cells. J. Biol. Chem. 276, 31449–31458 (2001).
Niwa, R., Nagata-Ohashi, K., Takeichi, M., Mizuno, K. & Uemura, T. Control of actin reorganization by Slingshot, a family of phosphatases that dephosphorylate ADF/cofilin. Cell 108, 233–246 (2002).
Chua, B. T. et al. Mitochondrial translocation of cofilin is an early step in apoptosis induction. Nat. Cell Biol. 5, 1083–1089 (2003).
Roh, S. E. et al. Mitochondrial dysfunction and calcium deregulation by the RanBP9-cofilin pathway. FASEB J. 27, 4776–4789 (2013).
Woo, J. A. et al. Slingshot-Cofilin activation mediates mitochondrial and synaptic dysfunction via Aβ ligation to β1-integrin conformers. Cell Death Differ. 22, 921–934 (2015).
Li, G. B. et al. Mitochondrial fission and mitophagy depend on cofilin-mediated actin depolymerization activity at the mitochondrial fission site. Oncogene 37, 1485–1502 (2018).
Inoue, T. et al. Differentiation-inducing factor 1 activates cofilin through pyridoxal phosphatase and AMP-activated protein kinase, resulting in mitochondrial fission. J. Pharmacol. Sci. 152, 39–49 (2023).
Li, S. et al. Transient assembly of F-actin on the outer mitochondrial membrane contributes to mitochondrial fission. J. Cell Biol. 208, 109–123 (2015).
Rehklau, K. et al. Cofilin1-dependent actin dynamics control DRP1-mediated mitochondrial fission. Cell Death Dis. 8, e3063 (2017).
Li, G. et al. Mitochondrial translocation and interaction of cofilin and Drp1 are required for erucin-induced mitochondrial fission and apoptosis. Oncotarget 6, 1834–1849 (2015).
Hu, J. et al. ROCK1 activation-mediated mitochondrial translocation of Drp1 and cofilin are required for arnidiol-induced mitochondrial fission and apoptosis. J. Exp. Clin. Cancer Res. 39, 37 (2020).
Klamt, F. et al. Oxidant-induced apoptosis is mediated by oxidation of the actin-regulatory protein cofilin. Nat. Cell Biol. 11, 1241–1246 (2009).
Hoffmann, L. et al. Cofilin1 oxidation links oxidative distress to mitochondrial demise and neuronal cell death. Cell Death Dis. 12, 953 (2021).
Shen, L. W. et al. Cepharanthine sensitizes human triple negative breast cancer cells to chemotherapeutic agent epirubicin via inducing cofilin oxidation-mediated mitochondrial fission and apoptosis. Acta Pharmacol. Sin. 43, 177–193 (2022).
Wabnitz, G. H. et al. Mitochondrial translocation of oxidized cofilin induces caspase-independent necrotic-like programmed cell death of T cells. Cell Death Dis. 1, e58 (2010).
Wang, C., Zhou, G. L., Vedantam, S., Li, P. & Field, J. Mitochondrial shuttling of CAP1 promotes actin- and cofilin-dependent apoptosis. J. Cell Sci. 121, 2913–2920 (2008).
Weng, W. et al. N-terminal α-amino SUMOylation of cofilin-1 is critical for its regulation of actin depolymerization. Nat. Commun. 14, 5688 (2023).
Schiavon, C. R. et al. Actin chromobody imaging reveals sub-organellar actin dynamics. Nat. Methods 17, 917–921 (2020).
Moriyama, K., Iida, K. & Yahara, I. Phosphorylation of Ser-3 of cofilin regulates its essential function on actin. Genes Cells 1, 73–86 (1996).
Ghosh, M. et al. Cofilin promotes actin polymerization and defines the direction of cell motility. Science 304, 743–746 (2004).
Aizawa, H. et al. Phosphorylation of cofilin by LIM-kinase is necessary for semaphorin 3A-induced growth cone collapse. Nat. Neurosci. 4, 367–373 (2001).
Lee, K. Y. et al. Infantile spasms-linked Nedd4-2 mediates hippocampal plasticity and learning via cofilin signaling. EMBO Rep. 22, e52645 (2021).
He, X. et al. Probing the roles of SUMOylation in cancer cell biology by using a selective SAE inhibitor. Nat. Chem. Biol. 13, 1164–1171 (2017).
Tanaka, K. et al. Structural basis for cofilin binding and actin filament disassembly. Nat. Commun. 9, 1860 (2018).
Galkin, V. E. et al. Remodeling of actin filaments by ADF/cofilin proteins. Proc. Natl. Acad. Sci. Usa. 108, 20568–20572 (2011).
Jaswandkar, S. V., Katti, K. S. & Katti, D. R. Molecular and structural basis of actin filament severing by ADF/cofilin. Comput. Struct. Biotechnol. J. 20, 4157–4171 (2022).
Shoji, K., Ohashi, K., Sampei, K., Oikawa, M. & Mizuno, K. Cytochalasin D acts as an inhibitor of the actin-cofilin interaction. Biochem. Biophys. Res. Commun. 424, 52–57 (2012).
Wakatsuki, T., Schwab, B., Thompson, N. C. & Elson, E. L. Effects of cytochalasin D and latrunculin B on mechanical properties of cells. J. Cell Sci. 114, 1025–1036 (2001).
Craig, E. A. Hsp70 at the membrane: driving protein translocation. BMC Biol. 16, 11 (2018).
Sayyed, U. M. H. & Mahalakshmi, R. Mitochondrial protein translocation machinery: From TOM structural biogenesis to functional regulation. J. Biol. Chem. 298, 101870 (2022).
Chacinska, A., Koehler, C. M., Milenkovic, D., Lithgow, T. & Pfanner, N. Importing mitochondrial proteins: machineries and mechanisms. Cell 138, 628–644 (2009).
Fukasawa, Y. et al. MitoFates: improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol. Cell. Proteom. 14, 1113–1126 (2015).
Zhang, Y. et al. The molecular recognition mechanism for superoxide dismutase presequence binding to the mitochondrial protein import receptor Tom20 from Oryza sativa involves an LRTLA motif. J. Phys. Chem. B 114, 13839–13846 (2010).
Bykov, Y. S. et al. Widespread use of unconventional targeting signals in mitochondrial ribosome proteins. EMBO J. 41, e109519 (2022).
Araiso, Y., Imai, K. & Endo, T. Role of the TOM complex in protein Import into mitochondria: structural views. Annu. Rev. Biochem. 91, 679–703 (2022).
Busch, J. D., Fielden, L. F., Pfanner, N. & Wiedemann, N. Mitochondrial protein transport: Versatility of translocases and mechanisms. Mol. Cell 83, 890–910 (2023).
Young, J. C., Hoogenraad, N. J. & Hartl, F. U. Molecular chaperones Hsp90 and Hsp70 deliver preproteins to the mitochondrial import receptor Tom70. Cell 112, 41–50 (2003).
Endo, T. & Yamano, K. Transport of proteins across or into the mitochondrial outer membrane. Biochim. Biophys. Acta 1803, 706–714 (2010).
Pfanner, N., Warscheid, B. & Wiedemann, N. Mitochondrial proteins: from biogenesis to functional networks. Nat. Rev. Mol. Cell Biol. 20, 267–284 (2019).
den Brave, F., Schulte, U., Fakler, B., Pfanner, N. & Becker, T. Mitochondrial complexome and import network. Trends Cell Biol. 34, 578–594 (2024).
Xu, K. et al. A new paradigm in intracellular immunology: Mitochondria emerging as leading immune organelles. Redox Biol. 76, 103331 (2024).
Wiedemann, N. & Pfanner, N. Mitochondrial machineries for protein import and assembly. Annu. Rev. Biochem. 86, 685–714 (2017).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Sim, S. I., Chen, Y., Lynch, D. L., Gumbart, J. C. & Park, E. Structural basis of mitochondrial protein import by the TIM23 complex. Nature 621, 620–626 (2023).
Backes, S. et al. Tom70 enhances mitochondrial preprotein import efficiency by binding to internal targeting sequences. J. Cell Biol. 217, 1369–1382 (2018).
Schneider, K., Zimmer, D., Nielsen, H., Herrmann, J. M. & Mühlhaus, T. iMLP, a predictor for internal matrix targeting-like sequences in mitochondrial proteins. Biol. Chem. 402, 937–943 (2021).
Flores-Romero, H., Dadsena, S. & García-Sáez, A. J. Mitochondrial pores at the crossroad between cell death and inflammatory signaling. Mol. Cell 83, 843–856 (2023).
Victorelli, S. et al. Apoptotic stress causes mtDNA release during senescence and drives the SASP. Nature 622, 627–636 (2023).
Posadas, I., Pérez-Martínez, F. C., Guerra, J., Sánchez-Verdú, P. & Ceña, V. Cofilin activation mediates Bax translocation to mitochondria during excitotoxic neuronal death. J. Neurochem. 120, 515–527 (2012).
Zhu, Y. et al. Caspase cleavage of cytochrome c1 disrupts mitochondrial function and enhances cytochrome c release. Cell Res 22, 127–141 (2012).
Li, G. et al. CYC1 silencing sensitizes osteosarcoma cells to TRAIL-induced apoptosis. Cell. Physiol. Biochem. 34, 2070–2080 (2014).
Grego-Bessa, J., Hildebrand, J. & Anderson, K. V. Morphogenesis of the mouse neural plate depends on distinct roles of cofilin 1 in apical and basal epithelial domains. Development 142, 1305–1314 (2015).
Ockeloen, C. W. et al. Congenital myopathy caused by a novel missense mutation in the CFL2 gene. Neuromuscul. Disord. 22, 632–639 (2012).
Zhang, H. J. et al. Destrin contributes to lung adenocarcinoma progression by activating Wnt/β-catenin signaling pathway. Mol. Cancer Res. 18, 1789–1802 (2020).
Shangguan, X. et al. SUMOylation controls the binding of hexokinase 2 to mitochondria and protects against prostate cancer tumorigenesis. Nat. Commun. 12, 1812 (2021).
Gao, Y. et al. SENP1 promotes triple-negative breast cancer invasion and metastasis via enhancing CSN5 transcription mediated by GATA1 deSUMOylation. Int. J. Biol. Sci. 18, 2186–2201 (2022).
Lee, M. H. et al. The SUMO protease SENP1 promotes aggressive behaviors of high HIF2α expressing renal cell carcinoma cells. Oncogenesis 11, 65 (2022).
Sheng, Z. et al. SENP1-mediated deSUMOylation of YBX1 promotes colorectal cancer development through the SENP1-YBX1-AKT signaling axis. Oncogene 44, 1361–1374 (2025).
Gu, J. et al. Suppressing SENP1 inhibits esophageal squamous carcinoma cell growth via SIRT6 SUMOylation. Cell. Oncol. 48, 67–81 (2025).
Cimarosti, H. et al. Enhanced SUMOylation and SENP-1 protein levels following oxygen and glucose deprivation in neurones. J. Cereb. Blood Flow. Metab. 32, 17–22 (2012).
Zhang, H. et al. SUMO-specific protease 1 protects neurons from apoptotic death during transient brain ischemia/reperfusion. Cell Death Dis. 7, e2484 (2016).
He, J., Cheng, J. & Wang, T. SUMOylation-mediated response to mitochondrial stress. Int. J. Mol. Sci. 21, 5657 (2020).
Shehjar, F., Almarghalani, D. A., Mahajan, R., Hasan, S. A. & Shah, Z. A. The multifaceted role of cofilin in neurodegeneration and stroke: insights into pathogenesis and targeting as a therapy. Cells 13, 188–203 (2024).
Kovaleva, T., Gainullin, M., Mukhina, I., Pershin, V. & Matskova, L. Cofilin(s) and mitochondria: function beyond actin dynamics. Int. J. Mol. Sci. 26, 4094 (2025).
Sun, H. et al. Kainate receptor activation induces glycine receptor endocytosis through PKC deSUMOylation. Nat. Commun. 5, 4980 (2014).
Zhang, Q. et al. hnRNPA1 SUMOylation promotes cold hypersensitivity in chronic inflammatory pain by stabilizing TRPA1 mRNA. Cell Rep. 42, 113401 (2023).
Gao, Y. et al. TRPV1 SUMOylation suppresses itch by inhibiting TRPV1 interaction with H1 receptors. Cell Rep. 39, 110972 (2022).
Yang, Y. et al. SUMOylation-induced membrane localization of TRPV1 suppresses proliferation and migration in gastric cancer cells. Cell Commun. Signal 22, 465 (2024).
Wieckowski, M. R., Giorgi, C., Lebiedzinska, M., Duszynski, J. & Pinton, P. Isolation of mitochondria-associated membranes and mitochondria from animal tissues and cells. Nat. Protoc. 4, 1582–1590 (2009).
Wang, W. et al. The inhibition of TDP-43 mitochondrial localization blocks its neuronal toxicity. Nat. Med. 22, 869–878 (2016).
Coyne, L. P. et al. Mitochondrial protein import clogging as a mechanism of disease. Elife 12, e84330 (2023).
Zhang, Q., Lee, W. A., Paul, D. L. & Ginty, D. D. Multiplexed peroxidase-based electron microscopy labeling enables simultaneous visualization of multiple cell types. Nat. Neurosci. 22, 828–839 (2019).
Wong, M. & Munro, S. Membrane trafficking. The specificity of vesicle traffic to the Golgi is encoded in the golgin coiled-coil proteins. Science 346, 1256898 (2014).
Martell, J. D., Deerinck, T. J., Lam, S. S., Ellisman, M. H. & Ting, A. Y. Electron microscopy using the genetically encoded APEX2 tag in cultured mammalian cells. Nat. Protoc. 12, 1792–1816 (2017).
Liu, Z. et al. Vascular disease-causing mutation, smooth muscle α-actin R258C, dominantly suppresses functions of α-actin in human patient fibroblasts. Proc. Natl. Acad. Sci. USA. 114, E5569–e5578 (2017).
Ma, R. et al. DUSP6 SUMOylation protects cells from oxidative damage via direct regulation of Drp1 dephosphorylation. Sci. Adv. 6, eaaz0361 (2020).
McKeague, A. L., Wilson, D. J. & Nelson, J. Staurosporine-induced apoptosis and hydrogen peroxide-induced necrosis in two human breast cell lines. Br. J. Cancer 88, 125–131 (2003).
Bonora, M. et al. Comprehensive analysis of mitochondrial permeability transition pore activity in living cells using fluorescence-imaging-based techniques. Nat. Protoc. 11, 1067–1080 (2016).
Acknowledgements
This study was supported by the National Key Research and Development Program of China (2021ZD0203100 and 2020YFA0803602 to Y.L.) and grants from the National Natural Science Foundation of China (31830031, 82071510, 31761163002, and 31671053 to Y.L.) and the Innovative Research Team of High-Level Local Universities in Shanghai (SHSMU-ZDCX20ß211102).
Author information
Authors and Affiliations
Contributions
Y.L. conceived and designed the study. Q.D. conducted biochemical assays, immunofluorescence, and electron microscopy imaging. X.G., J.F., and J.X. provided assistance with biochemical assays. X.G., J.F., J.X., and X.L. performed immunoprecipitation and mass spectrometry (IP/MS) experiments. Q.D., X.G., and W.W. contributed to data analysis and interpretation. W.W., G.W.Z., O.H., and S.P. participated in discussions and provided critical insights into the results. O.H. and S.P. offered technical support. Q.D. and Y.L. drafted the manuscript, with contributions and critical revisions from all authors. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks James Bamburg, Thomas Kuhn, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, Q., Gu, X., Feng, J. et al. N-terminal α-amino SUMOylation promotes phosphorylation-independent cofilin-1 translocation to the mitochondrial matrix and induces apoptosis. Nat Commun 16, 11687 (2025). https://doi.org/10.1038/s41467-025-66859-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66859-6









