Abstract
Digital pathology generates gigapixel whole-slide images that require extensive storage, limiting scalability and increasing costs. Traditional compression methods apply uniform ratios, disregarding variations in diagnostic importance across regions. Here we show Adaptive compression for gigapixel whole-slide images (AdaSlide), a framework that balances compression efficiency and diagnostic integrity. AdaSlide integrates a reinforcement learning-based Compression Decision Agent (CDA) that determines the optimal compression ratio per region, and a Foundational Image Enhancer (FIE) that restores visual fidelity after compression. In evaluations, pathologists achieved near-chance accuracy (55%) in distinguishing original from restored images, confirming high perceptual fidelity. Across 13 downstream tasks, AdaSlide maintained diagnostic performance in most tasks while reducing storage to 10-35% of the original size. By aligning compression strategies with clinical relevance, AdaSlide enables efficient, scalable, and reliable storage for digital pathology and future AI development.
Similar content being viewed by others
Introduction
Recently, the transformation from glass slide pathology to the digital pathology system (DPS) has led to the emergence of machine learning-integrated DPS, known as computational pathology (CPath)1,2,3,4. CPath has demonstrated strong performance in various pathological tasks, including cancer diagnosis 5,6,7,8,9,10,11, survival prediction12, lymph node invasion detection13,14, and cell segmentation15,16. CPath performance has significantly improved through the use of large-scale digitized slides, with robust foundation models recently proposed17,18,19,20,21,22,23. However, unlike natural images, whole-slide images (WSIs) are considerably large (3 or 4 GB), resulting in significant storage burdens for data archiving. For example, Linköping Hospital collects digital slides totaling 130 TB every six months 24. Due to storage limitations, slides older than six months are deleted, which is a temporary, impractical, and invasive solution. Simply expanding storage is also impractical, as it requires physical space and increases maintenance costs. Additionally, cloud servers may not be an optimal solution, as storage costs escalate cumulatively (see Supplementary Note 1 and Supplementary Fig. 1). However, most CPath studies remain focused on prediction accuracy, thus leaving the issue of efficient data storage unresolved.
To tackle this large-scale storage issue, several methods have been proposed, including neural image compression (NIC) that effectively compresses the original high-dimensional image into a compressed low-dimensional vector using neural networks25,26,27,28,29. These encoder-focused NIC methods utilize a supervised learning approach to maximize the task-specific prediction performance. However, the encoder-focused NICs are unsuitable for image data storage because, without a decoder, the compressed vectors cannot be reconstructed into the original images. Encoder-decoder-based NICs are more suitable for efficient storage, but limited methods are available for digital pathology applications30,31,32,33. A notable effort by Tellez et al.30 involved training Variational Autoencoders (VAE)34 and Bidirectional Generative Adversarial Networks (BiGAN)35 to encode and decode digital pathology images. They demonstrated the effectiveness of the NIC for digital pathology image compression and restoration, with minimal performance degradation compared to the original image-based methods. Nasr et al.33 focused on the compression ratio (CR) using the VAE model, and they reported a state-of-the-art (SOTA) CR (1/512) on cancer imaging data. Previous studies tessellated WSIs into 32 × 32 or 64 × 64 pixels. However, Keighley et al.31 employed patches of size 512 × 512 pixels and demonstrated successful compression and restoration using a Vector Quantized VAE (VQVAE)-236 model. With the matched CR (1/19.2), the outputs of VQVAE-2 (namely, VQVAE) showed high structural similarity (SSIM) to the JPEG-compressed images (0.8); however, the peak signal-to-noise ratio (PSNR) degraded (20 and 29, for VQVAE and JPEG, respectively). In contrast to NICs, Afshari et al.32 applied super-resolution (SR) methods for digital pathology images. The difference between NICs and SR is that NICs use compressed vectors, whereas SR relies on human-interpretable low-resolution images. In their empirical study, the SR models successfully reconstructed low-resolution images into their original high-resolution forms.
Previous studies30,31,32,33 have focused on compressing digital pathology images using specific models to achieve a higher CR with minimal information loss. However, even promising models suffer information loss during encoding and decoding steps due to structural limitations, which worsen at higher compression levels. Consequently, there is a risk of losing clinically significant information while aiming to maximize the CR. To address this concern, we focused on the inter-slide information variance in gigapixel WSIs, which we refer to as information disequilibrium (see “Information Disequilibrium” section for more details). Each WSI comprises numerous regions of interest (ROIs), some containing diagnostic information while others do not. By enabling the model to distinguish between informative and non-informative ROIs, we can optimize compression by heavily compressing only clinically uninformative ROIs, thereby minimizing critical information loss.
One recent work37 introduced a multiple instance learning (MIL)-based method for identifying informative regions. MIL learns the relationship between the labels provided during training and the instances within a bag, enabling the model to distinguish between more and less important instances. Based on this capability, the authors proposed applying different compression rates to important and less important regions, as determined by a trained MIL classifier. However, this approach has several limitations. Since the MIL classifier is trained in a supervised manner tailored to a specific task, (1) it learns to distinguish instances only in the context of that task, (2) making it difficult to generalize to other tasks with different objectives. Moreover, (3) the method is highly dependent on the performance of the MIL classifier itself and requires an appropriate dataset for MIL training, which may not always be available. Consequently, while this approach can be effective in certain controlled settings, its general applicability is limited, and its scalability is constrained.
To address these issues, we propose an Adaptive compression framework for gigapixel whole-slide images, namely AdaSlide, to account for information disequilibrium. AdaSlide is designed to meet the following constraints: (1) the lost information should be minimized, (2) each patch image must be maximally compressed, (3) the compression decision should be object-dependent, and (4) the compression decision and image enhancement should be agnostic to backbone-model architecture, organ, magnification, and downstream tasks.
AdaSlide consists of two core modules: a Compression Decision Agent (CDA) and a Foundational Image Enhancer (FIE). The CDA autonomously decides whether to compress an image based on its informational content. The images chosen for compression are downscaled from the D-dimension to the d-dimension (D > d), and later restored to the D-dimension using the FIE. The CDA is trained using reinforcement learning, eliminating the need for human annotation and allowing it to consider key factors in compression decisions based on the design of the reward function. The FIE is trained to handle multi-organ, multi-magnification scenarios and can be easily substituted with multiple backbones. In summary, our main contributions are as follows:
-
AdaSlide is an organ- and task-agnostic compression method.
-
AdaSlide provides flexible control over compression tendency and quantifies clinically relevant information.
-
AdaSlide applies information disequilibrium in a scalable, task-agnostic compression framework.
-
AdaSlide offers an optimal solution for mid- and long-term archiving, achieving over 65–90% storage reduction with minimal impact on clinical diagnosis.
Results
Overview of AdaSlide
AdaSlide trains in two stages: First, the FIE, designed to handle diverse magnification and cancer types, is trained. Subsequently, the CDA is trained using reinforcement learning to determine the optimal compression level for each patch image based on its information content. AdaSlide’s inference process consists of two stages: encoding and decoding. During encoding, the CDA decides action (keep or compress), and the original patch is compressed accordingly. Compressed patches are restored during decoding with the FIE. The entire AdaSlide pipeline is shown in Fig. 1B and Box 1.
A Patch distributions of the PanCancer dataset used to train AdaSlide. Using 930 WSIs from 31 TCGA projects, we extracted 40× and 20× magnification patch images (each magnification has equal proportions). B AdaSlide pipeline. A WSI is tessellated into multiple patch images, and the compression ratios are decided using the CDA. Patch images selected for compression undergo encoding and decoding steps using the FIE. Finally, the patch images excluded from compression, along with the enhanced images, were collected and reconstructed for further analysis. C Overall performance of AdaSlide on 13 datasets. The baseline indicates the conventional supervised learning using original images. The degraded performance implies information loss during compression and enhancement. AdaSlide showed comparable performance to the baseline, whereas the other compression models (ESRGAN, VQVAE, SwinIR, LDM) did not. WSI whole-slide image, TCGA The Cancer Genome Atlas, CDA Compression Decision Agent, FIE Foundational Image Enhancer.
Information disequilibrium
Information disequilibrium arises from the gigapixel scale of WSIs, where not all pixels contribute equally to clinical diagnosis. For instance, in identifying tumor regions within a WSI, areas densely packed with cells are more relevant than background, adipose tissue, or bone. Similarly, detecting lymphovascular invasion or perineural invasion requires focusing on specific tube-like structures, such as lymph nodes, vascular tubes, or perineural regions. This suggests that clinically important regions vary dynamically with diagnostic tasks and goals, rendering equal attention across all regions unnecessary.
For instance, MIL divides a WSI into multiple patch instances and applies attention mapping to selectively identify regions that significantly contribute to label prediction37,38,39. Advanced MIL models incorporate locality and global patterns40,41, prioritize key instances42, or leverage clustering loss to effectively differentiate label-relevant instances38. This study defines this dynamic allocation of importance across regions as information disequilibrium. Specifically, Li et al.37 proposed an adaptive decompression method that leverages MIL-based attention maps to assign different decompression depths to image patches. While the paper does not explicitly define the concept of information disequilibrium, its strategy effectively leverages the inherent information imbalance across regions to prioritize the decompression of diagnostically relevant areas.
Hypothesis of AdaSlide
Information disequilibrium is the cornerstone of this study because traditional uniform compression models fail to address it, leading to unavoidable information loss by compressing all regions equally. To design an information content-aware compression framework that accounts for information disequilibrium, we propose the following hypotheses:
-
Tumor-related diagnosis is one of the primary tasks.
-
Cellular information is clinically critical for tumor diagnosis.
-
Certain patch instances are more challenging to reconstruct than others.
While tumor-related diagnosis is not the sole clinical priority, we focused on it due to our training data’s characteristics and the prominence of tumor-related research in CPath. Accordingly, we designed a CDA reward function tailored to tumor-related diagnosis. In this study, clinically informative regions are defined as tumor-related areas. However, AdaSlide is flexible and can redefine clinically informative regions by modifying the reward function.
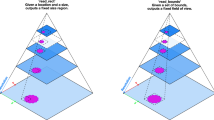
To quantify regions contributing to tumor-related diagnosis, we utilized cellular information, recognizing that the morphological features and distribution of cells are crucial to tumor diagnosis. Furthermore, we incorporated the compression difficulty of cellular information to balance the information loss and compression performance. Guided by hypotheses (2) and (3), we constructed a 2 × 2 conditional matrix (Fig. 2).
-
Clinically informative and easy to restore (Zone A).
-
Clinically informative and hard to restore (Zone B).
-
Clinically uninformative and easy to restore (Zone C).
-
Clinically uninformative and hard to restore (Zone D).
AdaSlide is trained to preserve Zone B (clinically informative and hard to restore) while compressing other zones, including Zones A, C, and D.
In this hypothesis field, the CDA is designed to prioritize Zone B by maintaining its original information as much as possible, while employing compression strategies for the remaining zones. This is achieved through a combination of compression rewards and information penalties.
Datasets
The PanCancer dataset, derived from 31 projects from The Cancer Genome Atlas (TCGA) dataset, has 930 WSIs, with 30 WSIs extracted from each project. It also includes 1.8 million patch images extracted at 20× and 40× magnifications Fig. 1A. The PanCancer dataset was divided into training (94.5%), validation (5%), and test (0.5%) sets, corresponding to 1,766,502, 93,307, and 9393 patch images, respectively. To further evaluate the FIEs’ reconstruction performance, we utilized datasets from the Clinical Proteomic Tumor Analysis Consortium (CPTAC). A total of 110 WSIs were selected, with 10 WSIs per project across 11 projects. Additionally, 29,861 patch images were extracted using the same patch generation pipeline and were used for performance evaluation. Details of the TCGA and CPTAC projects are summarized in Supplementary Note 2.
We evaluated the performance of AdaSlide on patch-level tasks (classification and segmentation) and slide-level tasks (classification) using 13 benchmark datasets, as summarized in Table 1. Detailed information, including preprocessing steps, is provided in the “Methods” section.
For the patch-level classification tasks, we used five datasets: NCT-CRC (9 classes; colon)43, MHIST (binary class; colon)44, LI (binary class; stomach)14, SICAPv2 (4 classes; prostate)45, and WSSS4LUAD (binary class; lung)46 datasets.
For patch-level segmentation tasks, we used the SNOW (binary class; breast)47, NuInsSeg (binary class; various organs)48, and PanNuke (binary class; various organs)49 datasets. The NuInsSeg and PanNuke datasets include multiple classes; however, for simplicity, we converted the multi-class masks into binary masks. Moreover, the NuInsSeg dataset contains mouse-derived H&E images and mask pairs, which we excluded from the analysis.
For the slide-level classification, we used the TCGA-RCC, TCGA-NSCLC, TCGA-BRCA, Camelyon16, and Children’s Brain Tumor Network (CBTN)50 datasets for slide-level classification analysis. The TCGA-RCC dataset combines multiple renal cell carcinoma (RCC) projects: TCGA-KICH, TCGA-KIRC, and TCGA-KIRP. Similarly, the TCGA-NSCLC dataset integrates non-small cell lung cancer (NSCLC) subtypes, including TCGA-LUAD and TCGA-LUSC. The TCGA-BRCA dataset was used for binary subtype classification, while the Camelyon16 dataset was used for tumor vs. non-tumor slide classification tasks. Lastly, the CBTN dataset was used for pediatric brain tumor subtype classification, including two pediatric-specific brain tumor cancer types: medulloblastoma and ependymoma.
Foundational Image Enhancer (FIE)
The image restoration performances of the FIEs are shown in Fig. 3A, and sample output images are illustrated in Fig. 3B and Supplementary Figs. 2 and3. We used the VAE34, VQVAE36, ESRGAN51, Swin Image Restoration (SwinIR)52, and Latent Diffusion Model (LDM)53 as FIE backbones, as these models have been widely adopted for digital pathology image compression tasks in previous studies30,31,33. In addition, we included Transformer-based and diffusion-based models that have demonstrated strong performance in image restoration tasks to provide a comprehensive comparison across different model families. Three evaluation metrics were used: Structural Similarity Index Measure (SSIM), Peak Signal-to-Noise Ratio (PSNR), and Learned Perceptual Image Patch Similarity (LPIPS)54.
A Quantitative results of enhanced images. The VAEs show lower performance in every metric; conversely, VQVAEs outperform all methods. ESRGAN shows comparable performance to VQVAE. B Example enhanced image outputs. The outputs of VAEs are notably blurry; conversely, the outputs of ESRGAN, VQVAE, SwinIR, and LDMs are clear. However, the VQVAEs exhibit unexpected noise patterns during the reconstruction of white background areas. Exact numerical values and statistics are provided in the Source Data file. FIE Foundational Image Enhancer.
The overall performance ranking was as follows: VQVAE, SwinIR, LDM, ESRGAN, and VAE. VQVAE demonstrated consistently strong reconstruction performance across both internal and external datasets. However, it occasionally produced artificial noise artifacts during reconstruction, as shown in Fig. 3B. SwinIR and LDM exhibited excellent performance on the internal dataset; however, their performance substantially degraded on the external dataset, indicating potential limitations in generalizability. In contrast, ESRGAN exhibited lower internal performance compared to SwinIR and LDM, but its performance degradation on the external dataset was less pronounced.
Although AdaSlide is designed to be agnostic to the choice of FIE model, it is necessary to fix the FIE during CDA training. This is because the reward function explicitly incorporates the quality of the enhanced images reconstructed from low-resolution inputs. Based on these considerations, we selected the FIE backbone by evaluating two key factors: (1) compatibility with compression, and (2) efficiency in training and inference.
VAE achieved the highest compression ratio, but the resulting image degradation was so severe that the reconstructed images were difficult to interpret visually. Therefore, this model was excluded from the candidate list. VQVAE demonstrated excellent compression and reconstruction performance. However, although the feature dimension of VQVAE (\({{\mathbb{R}}}^{128\times 128\times 2}+{{\mathbb{R}}}^{64\times 64\times 2}\)) is theoretically smaller than that of the original image (\({{\mathbb{R}}}^{512\times 512\times 3}\)), this only applies to the raw, uncompressed feature space. In practical applications, where image formats such as JPEG or PNG are used, the compressed features of VQVAE require larger storage space than image-based formats. Therefore, to support real-world applications such as demo systems, we focused on models that enable compression of low-resolution image files and subsequent reconstruction. As a result, the candidate models were SwinIR, LDM, and ESRGAN.
While LDM and SwinIR demonstrated strong performance, both required substantial computational resources and long processing times for training and inference. In particular, LDM, due to the inherent characteristics of diffusion models, was especially time-consuming. This makes LDM and SwinIR less practical for processing WSIs, which typically contain thousands to tens of thousands of image patches, as they would demand excessive computing resources and time. In contrast, ESRGAN offers the advantages of lower computational demands and the ability to leverage larger batch sizes during training, making it a more practical choice.
Based on these considerations, ESRGAN was selected as the FIE backbone for CDA training and for demonstration experiments, as it provided a good balance between performance and efficiency. However, it is important to note that this does not imply that AdaSlide is dependent on a specific FIE. To further assess the feasibility of FIE, we conducted a reader study involving a Visual Turing Test (VTT) with five pathology experts. The VTT evaluated whether the FIE-generated images could be distinguished from real images, using the internal dataset. The VTT results are summarized in Supplementary Table 1. VQVAE was excluded from this analysis due to its characteristic artificial noise Fig. 3B, which made it visually easy to differentiate. The VTT results indicated that FIE-generated images were challenging to distinguish from real ones, demonstrating the practicality of FIE-based reconstruction (56%; Z = 6, p = n.s.).
Compression Decision Agent (CDA)
To determine the compression level of an image, we employ a reinforcement learning-based agent that assesses the information content, reconstruction difficulty, and importance of each image. A conceptual diagram of the CDA is shown in Fig. 4A. We designed a reward function to compensate for the gain from compression and the loss of information (Equation (3)). This simplifies training by removing the need for human-annotated data and enhances model stability by eliminating inter-observer variation. To discourage the shortcut where the agent tries to compress every ROI, the agent receives a penalty for the amount of significant information lost during restoration. We defined the penalty as the difference between the cell segmentation output of the original image and the restored image, using HoverNet15. The concept of penalty based on HoverNet outputs is illustrated in Fig. 4B.
A Two possible outcomes depending on the CDA's binary decision. When the action is Keep (a = 0), the original image patch is stored directly, yielding zero reward. When the action is Compress (a = 1), the patch is encoded (f) and decoded (g), and the CDA receives a positive compression reward while incurring an information penalty proportional to the dissimilarity between the original and reconstructed segmentation masks (Sinfo = 1 − SDice). B Representative examples of successful and failed compression decisions, illustrating mask similarity and reconstruction fidelity. C Relationship between the hyperparameter λ and the CR. Increasing λ strengthens the information-preservation term, resulting in a lower CR and a negative association between λ and compression tendency. Exact numerical values and statistics are provided in the Source Data file. CDA Compression Decision Agent, CR compression ratio, SDice Dice score.
To control the tendency of CDA, we include a λ parameter. A higher λ value strengthens the penalty term, making the agent more conservative and less likely to compress images. We used grid search to find the optimal combination of λ and learning rate, observing that as λ increases, the learning rate and CR decrease (Fig. 4C). This indicates that a more conservative agent (higher λ) achieves a lower CR while likely preserving more image information.
To determine the backbone architecture for the CDA, we experimented with ResNet-18, ResNet-50, and Vision Transformer (ViT) models. Considering the alignment between the λ parameter and CR, as well as the efficiency of each model, ResNet-18 was selected as the final backbone for the CDA. The results are summarized in Supplementary Table 2.
Evaluation of information disequilibrium
We quantitatively evaluate the core assumption of information disequilibrium in CDA, based on its actual inference results. Specifically, we assess whether CDA effectively recognizes information imbalance across image regions through two complementary analyses: (1) whether cell-dense regions are appropriately prioritized, and (2) whether reconstruction difficulty is properly accounted for during compression decisions. Those results are summarized in Table 2.
To examine the first aspect, we leveraged cell segmentation results to implicitly guide CDA toward recognizing the importance of cell-rich regions. To validate this behavior, we utilized the zero-shot inference capability of the PLIP pathology vision-language model23. PLIP is a CLIP-based model trained on pathology image-caption pairs sourced from Twitter and is designed to capture the semantic relationships between images and textual descriptions within the pathology domain. For this analysis, we employed the following two query prompts to evaluate the relative similarity of each image:
-
A photo of densely packed cells.
-
A photo showing adipose tissue, stroma, or acellular background.
For each λ condition and compression decision, we measured the similarity of images to the first prompt (“A photo of densely packed cells.”) to assess whether CDA preferentially retained cell-rich regions. The findings indicate that in regions where compression was applied, the similarity to the cell-rich prompt was relatively lower, suggesting that CDA effectively deprioritizes semantically less informative regions during compression.
The second analysis aimed to verify whether the penalty design in CDA appropriately discourages indiscriminate compression by considering the difficulty of reconstructing compressed images. Specifically, we compared the pre- and post-reconstruction SSIM scores between images where CDA selected compression versus those where it selected retention (keep). The results show that as λ increases, images selected for compression exhibit higher SSIM scores compared to retained images. This trend suggests that CDA adapts its compression decisions to avoid high-penalty scenarios, demonstrating sensitivity to reconstruction difficulty and supporting the intended information-aware behavior.
Downstream tasks
We assessed AdaSlide across 13 downstream tasks, encompassing patch-level classification, patch-level segmentation, and slide-level classification. Information loss was quantified as the reduction in performance compared to results achieved with high-resolution image-based analysis. CR is a ratio of compression compared to the original high-resolution images. Hence, a lower CR value indicates a smaller file size (i.e., higher compression). Except for the training dataset, where images were compressed and enhanced using FIEs, all other conditions remained consistent. This ensured that any performance gap between original image-based analysis and processed image-based analysis was solely due to image information degradation during the image processing stages.
AdaSlide includes several variants depending on the backbone models (ESRGAN, VQVAE, SwinIR, LDM) and the compression tendency parameter λ (0.1, 0.25, 0.5, 0.75, 1.0). The average performance, best performance, and best CR of AdaSlide were reported in Fig. 5, while the detailed performances are summarized in Supplementary Tables 3–15. Since VAE’s performance was substantially lower, its results are omitted from Fig. 5 for clarity.
AdaSlide is presented in three parts: average, best performance, and best CR, respectively. A Performance difference from baseline across downstream tasks. The baseline represents performance using uncompressed images. AUROC scores were used for classification tasks (green and yellow border), and Dice scores for segmentation tasks (purple border). Positive values indicate improved performance compared to the baseline, while negative values indicate degradation. Detailed information is presented in Supplementary Tables 3–15. B Summary of CR. The baseline has a 100% CR; whereas, the other models have lower CRs than the baseline. AdaSlide achieves a balance between high CR and maintaining task performance, compared to ESRGAN and VQVAE. Exact numerical values and statistics are provided in the Source Data file. CR compression ratio, AUROC area under the receiver operating characteristic curve.
Supplementary Tables 3–7 summarize the patch-level classification results. Overall, the uniform compression models (VAE, ESRGAN, VQVAE, SwinIR, LDM) achieved the highest CR (smallest file sizes) but tended to degrade performance more than AdaSlide. Specifically, VAE exhibited a severe performance drop in every dataset (e.g., MHIST AUROC: 0.5552 vs. baseline 0.8437; LI AUROC: 0.5696 vs. baseline 0.8321), presumably due to the blurred reconstructions (Fig. 3B and Supplementary Fig. 2).
In contrast, AdaSlide outperformed the baseline on NCT-CRC, MHIST, LI, and SICAPv2, although it slightly underperformed in WSSS4LUAD. For instance, on NCT-CRC, VQVAE-based AdaSlide with a λ parameter of 0.5 (\({\,{\mbox{AdaSlide}}}_{{\mbox{VQVAE}}\,}^{0.5}\)) achieved the highest AUROC of 0.9943 (vs. baseline 0.9895). On MHIST, \({\,{\mbox{AdaSlide}}}_{{\mbox{ESRGAN}}\,}^{0.5}\) reached an AUROC of 0.8509, outperforming the baseline of 0.8437. The optimal λ and backbone model varied across datasets, indicating that compression preferences are dataset-specific.
Supplementary Tables 8–10 present the segmentation results (example images are in Supplementary Fig. 3). In contrast to classification tasks, segmentation requires high pixel-level fidelity for accurate mask delineation. As anticipated, VAE reconstructions often appeared too blurry, causing significant performance deterioration. Notably, in the NuInsSeg dataset, uniform compression models (VAE, ESRGAN, VQVAE without adaptive compressions) showed near-zero Dice scores, underscoring their difficulty in preserving crucial details.
Meanwhile, AdaSlide substantially mitigated information loss. For example, on the NuInsSeg dataset, \({\,{\mbox{AdaSlide}}}_{{\mbox{SwinIR}}\,}^{1.00}\) achieved a Dice score of 0.7068, with only a 2.12% drop from the baseline (0.7221). Conversely, in the same dataset, uniform compression models showed severely decreased performances (Dice: 0.0000). Interestingly, AdaSlide outperformed the baseline on certain datasets, including SNOW and PanNuke. For instance, on SNOW, \({\,{\mbox{AdaSlide}}}_{{\mbox{LDM}}\,}^{0.25}\) achieved a Dice of 0.9434, higher than the baseline (0.9334). Similarly, on PanNuke, \({\,{\mbox{AdaSlide}}}_{{\mbox{VQVAE}}\,}^{0.50}\) attained a Dice of 0.7304 (vs. baseline 0.7158).
For slide-level analysis, we employed the CLAM framework38, a widely used MIL method. As summarized in Supplementary Tables 11–15, AdaSlide slightly underperformed compared to the baseline in TCGA-RCC, Camelyon16, and CBTN but outperformed it in TCGA-NSCLC and TCGA-BRCA. For instance, in Camelyon16, AdaSlide ranked third (AUROC = 0.8676), slightly below the uniform compression VQVAE model (AUROC = 0.8679), and both were lower than the baseline (0.8852). Conversely, in TCGA-NSCLC, \({\,{\mbox{AdaSlide}}}_{{\mbox{SwinIR}}\,}^{1.00}\) achieved an AUROC of 0.8970, outperforming the baseline of 0.8774.
AdaSlide’s performance, information loss, optimal λ parameter, and FIE type vary based on the characteristics of the downstream datasets. This section reviews AdaSlide’s performance from the following perspectives: (1) regularization effects, (2) global and local patterns, (3) in-domain and out-of-domain datasets, and (4) dataset difficulty.
(1) Regularization Effects. A noteworthy observation across most datasets was that applying AdaSlide frequently improved performance compared to the baseline. As performance degradation relative to the baseline was defined as information loss, this improvement can be considered an information benefit. We hypothesize that this improvement stems from a regularization effect. Unlike uniform compression models, AdaSlide determines whether to pass an image through the FIE or use the original based on CDA’s decision. The FIE processes images only when the CDA predicts that it will not significantly degrade quality or affect performance. This selective approach likely enhances performance by reducing noise and normalizing colors in images, resulting in better outcomes for downstream tasks.
Key regions of high importance, where the FIE could potentially harm image quality, are left uncompressed by the CDA. This rationale explains why AdaSlide outperformed uniform compression models in most datasets, with the exception of Camelyon16. However, not all datasets exhibited positive effects, with some experiencing information loss, highlighting the limitations of the regularization hypothesis.
(2) Global and Local Patterns. The VAE model’s performance is constrained by its severe output blurriness. Nevertheless, this limitation offers insights into whether specific tasks rely more on global patterns or local details. Tasks that achieve acceptable performance despite blurry outputs suggest that while detailed information adds value, the primary directionality remains discernible.
For example, datasets such as NCT-CRC (patch-level classification) and slide-level classification datasets predominantly rely on global patterns, whereas datasets like NuInsSeg and other segmentation tasks depend heavily on local patterns. Tasks involving cell boundary segmentation or tumor infiltration detection rely heavily on fine-grained details (local patterns), while tasks focused on overall distribution and coloration (e.g., NCT-CRC) can be adequately performed using global patterns.
In tasks primarily driven by global patterns, even uniform compression models delivered satisfactory results. Conversely, for tasks dependent on local patterns, AdaSlide outperformed uniform compression models, effectively addressing information disequilibrium.
(3) In-domain and Out-of-domain Problem. Out-of-domain (OOD) challenges in AdaSlide arise from two primary sources of risk: (1) the image reconstruction performance of the FIE, and (2) the compression decision accuracy of the CDA. Since image reconstruction directly affects the amount of retained information, an FIE trained on in-domain data may fail to accurately reconstruct images from out-of-domain distributions, potentially introducing artifacts or blur. In parallel, the CDA, trained on cell information and image reconstruction difficulty, may misclassify important regions or overestimate reconstruction difficulty on unfamiliar data, resulting in information loss.
Among the 13 downstream tasks evaluated, AdaSlide exhibited notable performance degradation on three datasets: NuInsSeg, CBTN, and Camelyon16. These issues can be interpreted as follows: NuInsSeg represents a dataset shift in image brightness; CBTN introduces a domain shift to pediatric pathology; and Camelyon16 introduces a domain shift via lymph node sections, which differ from the tissue sections used in TCGA.
For NuInsSeg, performance degradation is primarily attributable to the FIE, as all FIE-based segmentation models failed (Dice score near zero), and the LDM-based model failed to generate adequate cell structures (Supplementary Fig. 3). This indicates that segmentation performance is highly dependent on the fidelity of the original high-resolution information, that is, on the compression ratio.
For CBTN, although performance degradation was less severe than for NuInsSeg, applying FIE alone resulted in noticeable performance drops. This is likely due to the pediatric-specific disease distribution, which diverges from the training domain of the FIE. However, when AdaSlide was applied, performance improved relative to using FIE alone, suggesting that the CDA mitigated some of the domain-induced reconstruction errors and that the primary issue lay with the FIE.
Camelyon16 exhibited a distinct pattern: it was the only case where FIE alone outperformed the full AdaSlide pipeline. Specifically, VQVAE, previously shown to generalize robustly-performed well, but adding CDA reduced performance. The CDA likely failed to operate effectively in this setting, as evidenced by the elevated CR and performance drop. Camelyon16 lymph node sections contain substantially higher cell densities than typical TCGA tissue sections. The CDA, trained to prioritize regions with high cell content, likely responded too aggressively in this context.
Despite these challenges, AdaSlide, composed of both FIE and CDA, outperformed FIE alone in most cases, suggesting that the CDA can compensate for FIE-induced information loss. Nevertheless, since both modules carry inherent OOD risks, careful experimental design and additional robustness strategies will be important considerations for future applications.
(4) Dataset Difficulty. The difficulty of each task played a significant role in determining the optimal λ parameter. For most datasets, λ = 0.50 produced the best results. For example, in patch-level segmentation tasks, dataset difficulty ranked as SNOW > PanNuke > NuInsSeg. This ranking considered both VAE performance and dataset characteristics, such as noise levels, with SNOW being a synthetic dataset with minimal noise. The optimal λ values were 0.25 for SNOW, 0.50 for PanNuke, and 1.00 for NuInsSeg. Higher λ values were more appropriate for out-of-domain datasets where the FIE struggled to reconstruct images accurately.
Slide-level classification showed relatively smaller performance differences compared to patch-level classification when selecting FIE and λ values. This was likely due to the reduced influence of individual patch instances in slide-level tasks, where classification relies more on shared global patterns. Although restoring fine-grained patterns remains important, the preservation of macroscopic patterns had a greater impact on slide-level classification. As a result, mid-range λ values typically achieved the best performance across most datasets, except for the CBTN dataset.
Best practice for selecting the optimal λ parameter
AdaSlide is not explicitly trained for any particular downstream task. Instead, its compression behavior is implicitly controlled through the choice of the λ parameter. Selecting an appropriate λ value is therefore important to balance compression efficiency and downstream task performance. Our experiments show that no single λ value performs optimally across all datasets and tasks. The optimal choice depends on several factors, including the target task’s sensitivity to fine-grained details, the relative importance of preserving subtle information, and the potential benefits of noise reduction introduced by the FIE. Because of this variability, the most reliable strategy is to conduct small-scale validation experiments that reflect the characteristics of the target task, as demonstrated in our downstream evaluations. Such experiments provide empirical guidance on selecting the λ value that best suits the intended application.
When performing dedicated validation is not feasible, either due to a lack of labeled data or limited computational resources, a practical heuristic can be used. In our experiments, we observed that λ = 0.50 generally provided a good balance between information preservation and compression across a wide range of datasets. Therefore, λ = 0.50 can be recommended as a reasonable default choice for general applications. For use cases where preserving fine-grained information is particularly important, such as tasks that are highly sensitive to local image patterns or applications where domain shift is expected between training and deployment data (for example, between TCGA and external cohorts), a more conservative setting such as λ = 1.00 is advisable. This choice reduces the risk of losing important information due to excessive compression. On the other hand, if storage efficiency is the primary concern and some level of information loss is acceptable, smaller λ values can be explored to maximize compression. More details can be found in Supplementary Note 4, and Supplementary Fig. 4.
Application example of AdaSlide
To evaluate compression performance at the WSI level, we implemented a real-world application of AdaSlide pipelines and validated both compression and enhancement. We selected one sample from TCGA-BRCA, TCGA-KIRC, and Camelyon16 datasets, respectively. The compression results are depicted in Fig. 6, and Supplementary Figs. 5 and 6.
Compression and information-preservation performance evaluated on the TCGA-BRCA sample TCGA-OL-A66K (original WSI size: 3.17 GB) at 20× magnification. A Original WSI with pathologist-annotated tumor regions. B CDA compression probability heatmap. Red regions indicate keep actions; blue regions indicate compress actions. Tumor regions exhibit a higher tendency for keep actions (AUROC = 0.6288). The overall compression ratio was 21.74%. C Representative examples of four regions based on information disequilibrium. Red borders: ESRGAN-reconstructed images; blue borders: original images. Clinically informative regions correspond to tumor areas. D Storage requirements by image format. The hybrid strategy required an average of 11.19% of the original WSI size. Exact numerical values and statistics are provided in the Source Data file. TCGA The Cancer Genome Atlas, BRCA breast invasive carcinoma, WSI whole-slide image, CDA Compression Decision Agent, AUROC area under the receiver operating characteristic curve.
Tumor regions exhibited a higher tendency for the Keep action, and the AUROC for distinguishing tumor versus non-tumor regions based on CDA action probabilities was 0.6288, indicating performance above chance level Fig. 6B. The table in Fig. 6D summarizes the effective compression performance when storing the compressed images. While JPEG format achieved the smallest storage footprint due to its inherent lossy compression, we observed notable image quality degradation during ESRGAN-based reconstruction, rendering it suboptimal for practical use (Supplementary Fig. 7). Conversely, PNG format provided the best reconstruction quality owing to its lossless nature, but resulted in substantially larger storage requirements. To balance these trade-offs, we adopted a hybrid strategy wherein high-resolution (keep) images were stored as JPEG to minimize storage demands, while compressed (compress) images were stored as PNG to preserve reconstruction quality. This approach provided an effective balance between storage efficiency and image fidelity.
Discussion
In this study, we developed AdaSlide, an adaptive compression framework that addresses the trade-offs between CR and information loss when compressing gigapixel WSIs. We assembled a PanCancer dataset from 31 TCGA projects, comprised of 1.8 million patches. Using this dataset, we trained two key modules: the FIE and the CDA. The FIE enhances the compressed images through encoding and decoding steps. The CDA, trained to handle information disequilibrium, autonomously determines the compression level based on the information content of the image. We evaluated the performance of AdaSlide across various downstream tasks, including patch-level classification, segmentation, and slide-level classification. The results indicated minimal information loss between the baseline and AdaSlide method, demonstrating its effectiveness in maintaining prediction accuracy while reducing the image storage capacity. AdaSlide defines information disequilibrium in a data-driven manner, which offers a key advantage over prior MIL-based approaches37 that rely on attention maps. In contrast to these methods, AdaSlide is task-agnostic and not reliant on task-specific datasets, enabling broader applicability across diverse clinical and research settings.
Notably, AdaSlide outperformed uniform compression models (i.e., ESRGAN, VQVAE, SwinIR, and LDM) in all downstream tasks except for Camelyon16. Furthermore, AdaSlide demonstrated performance comparable to that of baseline models. Specifically, on the NuInsSeg dataset, uniform compression models failed to properly segment cell masks (Dice: 0.0000), whereas AdaSlide preserved information more effectively than the baseline methods. Additionally, AdaSlide showed adaptability to out-of-domain datasets such as Camelyon16 and CBTN. Camelyon16 comprises lymph node section slides, which differ slightly from the TCGA PanCancer dataset. Meanwhile, CBTN focuses on pediatric tumor types, which are distinct from the adult cancer data in TCGA. While AdaSlide’s performance was lower than the baseline on these datasets, it still exceeded that of the uniform compression models. These findings suggest that AdaSlide effectively mitigates information loss caused by FIE limitations and addresses out-of-domain challenges to some extent. These findings highlight that improving the performance of the FIE is crucial for enhancing AdaSlide’s overall effectiveness. In the future, we plan to incorporate recent diffusion-based SOTA models55 to further improve performance.
The AdaSlide framework offers flexible options to accommodate various research objectives. For instance, researchers can regulate the compression tendency using the CDA’s λ parameter. Additionally, the reward function is easily modifiable. We used a reinforcement learning method to train the CDA, eliminating the need for human annotations. This process is scalable to larger training datasets and adaptable to different information criteria. By defining the penalty term, the CDA can learn the information contained in patch images. For example, we used cell segmentation masks as pseudo-ground truths to quantify information loss. However, diagnosing lipogenic neoplasms originating from adipocytic cells does not align with the current version of AdaSlide. Fat tissue, which generally exhibits low cellularity and scarce nuclei, is prone to being recognized as less informative by the CDA. Adjustments to the CDA’s reward function would be necessary to handle lipogenic neoplasms more effectively.
Faghani et al.56 introduced a lossless compression technique that identifies the tissue-containing foreground using an object detection-style binarization process and removes the background. Since this method does not alter the foreground (tissue) regions, it preserves the diagnostic information and does not degrade downstream performance. This represents an effective compression approach for certain scenarios. However, their method reconstructs the image by cropping the foreground regions into bounding boxes and reassembling them using a rectangle-packing strategy. As a result, it performs optimally when (1) the image contains repetitive patterns of similar size, (2) a large proportion of background is present, and (3) the tissue is predominantly organized along horizontal or vertical axes. In contrast, the method is less effective when the tissue exhibits diagonal structures or when the background is minimal, as occurs in many tumor-rich slides. Thus, the ideal application scenario for this method would be tissue microarrays (TMAs). Moreover, because the final reconstructed image disrupts the original spatial layout, direct spatial correspondence with the original WSI is lost, posing challenges for certain applications such as spatial omics alignment or visual inspection. In contrast, AdaSlide preserves the original spatial grid of the WSI and is not subject to the aforementioned constraints, enabling more flexible and generalizable compression.
This study had several limitations. First, we only considered a binary compression decision, determining whether to compress each image region. This approach could be extended to multi-class compression, where the CDA determines different compression levels for improved efficiency. Additionally, the CDA occasionally failed to converge stably during training. Future research will address this by exploring advanced policy-based reinforcement learning algorithms, such as proximal policy optimization (PPO)57, and conducting more sophisticated hyperparameter tuning. The λ parameter and the achieved CR were not always perfectly aligned with downstream task requirements, suggesting that the optimal λ value may vary depending on the dataset. Furthermore, the final compressed outputs could not be explicitly controlled, necessitating empirical experiments to identify the optimal λ value. To address this, we plan to modify the reward functions by incorporating explicit CR constraints to better control slide- or group-level CR. In addition, SSIM scores for tumor and non-tumor areas were relatively similar under the current CDA interpretation (Table 2). To address this, we plan to further refine the penalty design to more explicitly account for reconstruction difficulty. Incorporating semantic information from VLMs23 represents a promising extension to enhance this capability.
AdaSlide is currently limited to an ROI-level single-patch image-based model, although the final output is expected to operate at the slide level. As such, AdaSlide does not collectively consider related ROI information. To address this, we plan to enhance both the CDA and FIE to handle multiple ROIs in future studies. The CR of AdaSlide is based on the size of raw pixel images, which differs from the file sizes of compressed formats such as JPG or PNG. For example, in our demo using ESRGAN, tessellated patch instances saved in formats like JPG enabled compression to a smaller size than the original SVS image. However, using VQVAE, which operates with 32-bit floating-point values, resulted in file sizes exceeding those of typical image files. Future research should consider practical image formats and introduce techniques like quantization to address this issue. Currently, ESRGAN achieves optimal enhancement performance when applied to PNG-based images. As a result, in the current demonstration, a hybrid strategy was used, with Keep images stored in JPEG format and Compress images stored in PNG format. In future work, enabling ESRGAN to operate effectively on JPEG-compressed images would allow for maximal storage efficiency. This capability could potentially be combined with the foreground cropping method proposed by Faghani et al.56, further enhancing the overall compression effectiveness.
In summary, AdaSlide is a framework that adjusts the compression levels based on the image’s information content to effectively compress digital pathology images. AdaSlide demonstrated balanced performance between CR and information preservation in various downstream tasks. Additionally, AdaSlide offers flexibility in terms of backbone architecture, allows for adjustable CR, and includes a reward function design to define information penalties. AdaSlide stores digital pathology images more effectively, thereby reducing costs and contributing to large-scale digital pathology archiving for long-term storage.
Methods
PanCancer datasets
The main objective of the FIE is to achieve robustness across multiple domains, including different organs, scanner variations, and scanned image quality. To address these challenges, we collected data from TCGA, which encompasses 31 projects. Thirty diagnostic slides were randomly sampled from each project, which yielded a training set of 930 WSIs. Although the entire TCGA collection contained over 20,000 WSIs, we sampled 5% to reduce computational costs and facilitate efficient training. Each WSI was tessellated into multiple patch images. We considered two key objectives to generate diverse patches:
-
Multi-magnification representation: Patches were extracted from both 40× (0.25 μm/pixel) and 20× (0.5 μm/pixel) magnifications.
-
Spatial diversity: To promote adequate coverage of different WSI regions, we sampled 10% of the patches at 40× and 40% at 20×. The WSIs exhibited strong spatial correlations, implying that nearby patches often shared similar patterns. We employed a sampling-based patch selection instead of a sliding-window-based selection to encourage diverse patch selection.
To exclude non-tissue areas, we generated binary tissue masks using the Otsu thresholding with a validity threshold of 70%. Each patch was 512 × 512 pixels in size. We generated 1.8 million patches, ensuring equal representation at 40× and 20× magnifications. The PanCancer dataset encompasses diverse organs and cancer types. We divided PanCancer into training (94.5%), validation (5%), and test (0.5%) sets.
Datasets: Foundation Image Enhancer (FIE)
To train and validate the FIE, we started with high-resolution images (512 × 512 pixels) from the PanCancer dataset and artificially downscaled them to 128 × 128 pixels using bilinear interpolation. This simplified setting focused solely on the challenge of upscaling without the additional noise complexities encountered in real-world low-resolution images. Patch images were generated at both 40× and 20× magnifications to train and validate robustness across varying scales. The same preprocessing pipeline was applied to generate an external validation dataset sourced from CPTAC.
Datasets: Compress Decision Agent (CDA)
A validation split of the PanCancer dataset was used to train the CDA. For the pseudo-ground truth, the HoverNet15 outputs were acquired from both the original image and the compressed-and-restored image.
Downstream datasets
To validate the performance of the trained AdaSlide model, we leveraged several publicly available downstream datasets commonly used for slide-level and patch-level tasks. These datasets were specifically selected to evaluate the performance of AdaSlide in multi-magnification, multi-organ environments. A summary of each dataset is provided in Table 1.
The NCT-CRC dataset (National Center for Tumor Diseases Colorectal Cancer Type Classification) dataset consists of 100,000 training and 7180 test images extracted from H&E-stained WSIs of human CRC and normal tissue43. The dataset contains nine classes: adipose tissue, background, debris, lymphocytes, mucus, smooth muscle, normal colon mucosa, cancer-associated stroma, and NCT-CRC epithelium. All images were color-normalized using the Macenko method58. We split the original training set into a training and a validation set at a 7:3 ratio. The test dataset was then used according to a publicly available split set.
The MHIST (Minimalist Histopathology Image Analysis Dataset) dataset contains 2175 training and 977 test patch images of H&E-stained colorectal polyps (extracted from 328 FFPE WSIs) from Dartmouth-Hitchcock44. This dataset focused on the binary classification of benign hyperplastic polyps (HPs) vs. precancerous sessile serrated adenomas (SSAs). We split the original training set into a training and validation set at a 7:3 ratio. The test dataset was then used according to the publicly available split set.
The LI (Lymphatic Invasion Classification) dataset has 974 images extracted from H&E-stained WSIs of human stomach cancers14. It specifically targets the binary classification of LI regions versus normal regions. Unlike other datasets, the LI dataset concentrates on small regions containing lymphocytes and the presence of tumor invasions within those lymphocyte areas. Although the original LI dataset includes weakly-labeled negative cases, we excluded them from this study. The LI dataset was divided into training, validation, and test datasets at a ratio of 5:3:2.
The SICAPv2 (Sistema de Interpretación de Imágenes Histopatológicas para la Detección de Cáncer de Próstata) dataset comprises 155 biopsies from 95 patients, with labels based on primary Gleason grades ranging from non-cancerous (NC) to Grade 5 (GG5). The primary grade was used as the target label for classification. The public test split was utilized for validation, while the remaining data were divided into training and validation sets with a ratio of 0.8:0.2 based on the public training split set configuration.
The WSSS4LUAD (Weakly-supervised Semantic Segmentation Techniques for Histopathology Images of Lung Adenocarcinoma) dataset comprises LUAD image tiles ranging from 200 to 500 pixels in size, each annotated with labels such as tumor, tumor-associated stroma, or normal. For our analysis, we adopted the approach outlined in the UNI19, selecting only samples with a single ground-truth label. This resulted in a subset of 4693 images from the official training split. In this study, the dataset was further divided into training, validation, and test sets in a ratio of 0.6:0.2:0.2.
The SNOW (Synthetic Dataset for Human Breast Cancer Nuclei Segmentation) dataset contains 20,000 images of breast cancer patch images47. The patch images and nuclei annotation masks were generated from deep neural networks. We selected the first 5000 images and their corresponding mask pairs to simplify the experiments. These 5000 image-mask pairs were then divided into training, validation, and test datasets at a ratio of 9:0.5:0.5.
The NuInsSeg (Nuclei Instance Segmentation) dataset is a fully manually annotated dataset for nuclei segmentation in H&E images. It includes 665 image patches with over 30,000 manually segmented nuclei from 31 human and mouse organs. For this study, only the human H&E images (472 images) were used. The dataset was split into training, validation, and testing sets with a ratio of 0.6:0.2:0.2, respectively. Detailed instructions for generating the related segmentation masks are available in the public repository associated with the dataset.
The PanNuke (Pan-cancer Histology Dataset for Nuclei Instance Segmentation and Classification) dataset contains nuclei-level annotations derived from histopathology images spanning 19 tissue types. The dataset includes approximately 200,000 nuclei categorized into five clinically significant classes: neoplastic, inflammatory, connective tissue, necrotic/dead, and epithelial. In this study, we converted the dataset to binary segmentation masks for simplicity. The PanNuke dataset fold 1 was split into training, validation, and test datasets at a ratio of 0.6:0.2:0.2.
The TCGA-RCC dataset comprises 884 WSIs from TCGA’s kidney chromophobe (TCGA-KICH), kidney renal clear cell carcinoma (TCGA-KIRC), and kidney renal papillary cell carcinoma (TCGA-KIRP) projects. TCGA-KICH contains 111 images from 99 patients, TCGA-KIRC contains 489 images from 483 patients, and TCGA-KIRP contains 284 images from 264 patients. Patches were generated at 20x magnification with a size of 512 × 512 pixels. The average number of patches per slide was approximately 802.66, with a standard deviation of 435.81. The maximum and minimum numbers of patch images per slide were 4001 and 12, respectively.
The TCGA-NSCLC dataset contains 993 WSIs from TCGA, comprising 507 lung squamous cell carcinoma (TCGA-LUAD) and 486 lung adenocarcinoma (TCGA-LUSC) cases across 896 patients. Patches were generated at 20× magnification and are 512 × 512 pixels in size. The average number of patches per slide was approximately 694.63, with a standard deviation of 490.80. The maximum and minimum patch images per slide were 2933 and 10, respectively.
The TCGA-BRCA dataset includes various cancer types; however, for this analysis, we focused on invasive ductal carcinoma (IDC) and invasive lobular carcinoma (ILC) subtypes. A total of 1007 cases across 948 patients were employed. Patches were generated at 20× magnification and are 512 × 512 pixels in size. The average number of patches per slide was approximately 712.82, with a standard deviation of 452.17. The maximum and minimum patch images per slide were 4252 and 12, respectively.
The Camelyon16 dataset (Cancer Metastases in Lymph Nodes Challenge 2016) contains 268 WSIs, including 111 slides from patients with lymph node metastases and 160 slides from normal lymph nodes. The dataset focuses on the binary classification of metastatic versus non-metastatic lymph nodes. Patches were extracted at 40× magnification with a size of 512 × 512 pixels. On average, each slide generated approximately 6021.86 patches, with a standard deviation of 4130.37. The maximum and minimum patch images per slide were 26,926 and 127, respectively.
The Children’s Brain Tumor Network (CBTN)50 contains 3607 WSIs across 41 diagnostic categories derived from pediatric samples. For this study, we narrowed the classification task to only include Medulloblastoma and Ependymoma diagnoses. 150 WSIs were selected from each class, and included 212 patients. Patches were generated at 20× magnification and are 512 × 512 pixels in size without overlaps. The average number of patches per slide was about 173, with a standard deviation of 127.12. The minimum and maximum number of patch images per slide were 4 and 721, respectively.
Model Developments: Foundation Image Enhancer (FIE)
The FIE was trained using three backbone architectures: VAE34, VQVAE36, ESRGAN51, SwinIR52, and LDM53. The FIE required two modules: an encoder f( ⋅ ) and a decoder g( ⋅ ). The encoder maps the original images x to the latent vectors, and the decoder maps the latent vectors to the original images. Therefore, the FIE works as follows:
where the FIE was trained to minimize the difference between x and \(\hat{x}\).
For the VAE34, the original study used 64 × 64 pixel images. We trained it with images of size 224 × 224 and 512 × 512 pixels. The architecture consisted of hidden dimensions with various sizes depending on the inputs: 224 × 224 (32, 64, 128, 256, 256, 512, 512) and 512 × 512 (32, 64, 128, 256, 512, 512, 512, 1024, 1024, 1024). The batch size was set to 128, and training used the Adam optimizer with a learning rate of 1 × 10−4. The training split of the PanCancer dataset was used and trained for five epochs. The checkpoints were selected based on the validation loss.
The VQVAE36 was proposed to replace the latent dimensions represented by a Gaussian distribution in VAEs. Instead, VQVAE employs quantized vector embeddings to represent the latent dimensions. In addition, VQVAE, particularly VQVAE-2, attempts to enhance the images in a two-stage manner. Similar to VAE, VQVAE exhibits low flexibility with respect to the input image shape. Therefore, we trained the VQVAE model twice, once for 224 × 224 pixels and again for 512 × 512 pixels. The architecture and hyperparameters are shared. The model hyperparameters and training parameters followed those of a previously proposed VQVAE model for histopathological image compression31. The batch size was set to 128 and the learning rate to 1 × 10−4, using the Adam optimizer with five epochs. Minimum validation loss was set as the checkpoint selection criterion.
Unlike the VQVAE models, the ESRGAN51 does not require trained encoders. The ESRGAN’s encoder f( ⋅ ) is a simple linear downscaling method. Unlike other methods, ESRGAN attempts to minimize perceptual differences. As numerical metrics such as SSIM and PSNR diverged from human expert selection, the optimal checkpoint was determined through visual inspection in collaboration with a pathologist. To train the ESRGAN model, a batch size of 256 was used, and the model was initialized with a pretrained ESRGAN model. The Adam optimizer with a learning rate of 1 × 10−4 was used, and training lasted for a total of 50,000 steps, with the first 2500 as a warm-up phase. All other hyperparameters were maintained at their default settings, as defined in the original ESRGAN model51.
The SwinIR52 is a Transformer-based image restoration model that utilizes the Swin Transformer architecture59. Compared to CNN-based models such as ESRGAN, SwinIR provides enhanced capability for modeling long-range dependencies and contextual information, which is particularly useful in reconstructing complex textures in histopathology images. In our implementation, we trained the SwinIR model to handle input sizes of 512 × 512 pixels. The architecture was initialized with a pretrained SwinIR model and fine-tuned using the PanCancer training split. A batch size of 12 and the Adam optimizer with a learning rate of 2 × 10−4 were used. The training was performed for 50,000 steps, with the multiple step scheduler with 5 milestones: 25,000, 40,000, 45,000, 47,500, 50,000. The checkpoint was selected based on the lowest validation loss. All other hyperparameters followed the original SwinIR implementation52. For training SwinIR, we employed the KAIR library (v1.1, https://github.com/cszn/KAIR).
The LDM53 is a generative diffusion model that performs denoising in a lower-dimensional latent space, enabling both efficient training and high-quality image reconstruction. Unlike other models in this study, LDM consists of three main components: a first-stage VQVAE to map images to latent space, a UNet-based denoising network that operates in latent space, and a conditioning mechanism to guide reconstruction. In our implementation, we adopted an LDM architecture customized for histopathology image reconstruction. The pretrained first-stage VQVAE was employed. The denoising network was trained with a noise schedule following the cosine beta schedule, with 1000 diffusion steps. We used a batch size of 8 and the Adam optimizer with a learning rate of 2 × 10−6. The training was conducted for 7 epochs. The best checkpoint was selected based on the lowest validation loss computed on reconstructed images. Training the LDM required approximately 10 GPU days.
The VAE and VQVAE depended on a fixed input size; therefore, these models were trained twice to handle the input sizes of 224 × 224 and 512 × 512 pixels, respectively. The ESRGAN, SwinIR, and LDM models were trained once to handle an input size of 128 × 128. Python (v3.8), PyTorch (v2.0.1), and a single RTX A6000 GPU were used to implement AdaSlide.
Model Developments: Compression Decision Agent (CDA)
The CDA takes a single image patch as input and makes a binary compression decision a ∈ {0, 1}, where a = 1 denotes Compress and a = 0 denotes Keep. The agent employs pretrained image encoders (ResNet-18, ResNet-50, ViT) and is optimized by policy-based learning using the REINFORCE algorithm60 (see Supplementary Note 3). Unlike conventional multi-step REINFORCE, we adopt a single-step setting with one patch per decision, eliminating the computational overhead of episode sampling.
We balance compression efficiency and information preservation. Let SDice ∈ [0, 1] be the Dice similarity between the segmentation mask derived from the ESRGAN-reconstructed (compressed) image and the reference mask obtained from the original image. The information penalty is
To avoid slow convergence, we introduce a quality threshold τ ∈ (0, 1) on SDice. The final reward is
where λ > 0 controls the strength of the information-preservation term.
We tune λ ∈ {0.1, 0.25, 0.5, 0.75, 1.0} and the learning rate ∈ {1 × 10−4, 5 × 10−5, 1 × 10−5, 5 × 10−6, 1 × 10−6} via grid search. To avoid over-/under-compression, models with CR outside 20–90% are discarded.
Downstream tasks
To compare performance, we evaluated 13 downstream tasks for each condition: supervised training using the original image, using only the enhanced image processed by the foundation image enhancer (when λ = 0), and applying adaptive compression with the AdaSlide method.
Patch-level downstream tasks are divided into classification and segmentation. The classification tasks include five datasets (NCT-CRC, MHIST, LI, SICAPv2, WSSS4LUAD), while the segmentation tasks include three datasets (SNOW, NuInsSeg, PanNuke). All experiments were repeated using five random seeds ({42, 43, 44, 45, 46}). For consistency, the same split sets were used across experiments, and public split sets were utilized whenever available. The checkpoints were selected based on the lowest validation loss. The Adam optimizer, paired with a cosine annealing learning rate scheduler, was applied throughout.
For patch-level classification tasks, a class-balanced sampler was employed to address class imbalance. Only resize and normalization augmentations were applied. In contrast, patch-level segmentation tasks, which generally involve smaller datasets, utilize various image augmentations, including flipping, random cropping, and color jittering. Detailed configurations for each downstream task are summarized in Table 3.
The slide-level classification datasets included TCGA-RCC, TCGA-NSCLC, TCGA-BRCA, Camelyon16, and CBTN. We utilized the CLAM pipeline38 for preprocessing and training the MIL models, with the CLAM-SB backbone model. Feature extraction for each patch image was performed using the ResNet-50 model. The batch size was set to 1, with training conducted over 50 epochs at a learning rate of 1 × 10−4. Early stopping was applied to prevent overfitting. The Adam optimizer was used without a learning rate scheduler.
Visual Turing test (VTT)
Five expert pathologists participated in VTT. In each trial, participants were presented with two images: one original and the other restored using ESRGAN. They were instructed to identify which image appeared to be enhanced. A total of 30 questions were presented, and the chance level was set at 50%. The resulting accuracy scores were compared against the chance level using the Wilcoxon signed-rank test.
Application example of AdaSlide
Box 1 presents the AdaSlide pipeline. In this application, patch images were saved as JPEG for kept images and PNG for compressed images. The threshold τ was set to the optimal λ determined by downstream performance.
Using CLAM toolbox38, we preprocessed the WSIs by selecting foreground tissue areas using binarized images and applying median blurring to smooth the edges. For the Camelyon16 sample, patches were generated at level 0 (0.25 μm/pixel), with a size of 512 × 512 pixels. For TCGA-BRCA and TCGA-KIRC samples, the patch size was initially 1024 × 1024 at level 0, which was then resized to 512 × 512 pixels. The numbers of patches for Camelyon16, TCGA-BRCA, and TCGA-KIRC were 24,063, 9278, and 3720, respectively. Using the CDA, we made a patch-wise compression decision. Based on the decision made by the CDA, the patch images were either compressed to 128 × 128 pixels (compress) or left uncompressed (keep). In the decoding step, the compressed images are restored to their original pixel size and enhanced to the FIE. After the encoding-decoding step, the patch images were restored to their original WSI file format. The pyvips framework was used to generate a TIFF-format WSI file and JPEG compression with 75% quality. During reconstruction, the background images were not considered; therefore, the final size of the WSIs was reduced.
Evaluation Metrics: Foundation Image Enhancer (FIE)
The PSNR measures the extrinsic image quality based on the pixel values between the original and enhanced images. The PSNR is computed as follows:
where MAXI is the maximum pixel value of the image, and MSE is the mean squared error between the original and the enhanced images. Similarly, the SSIM measures the differences in structures and pixel values between the original and enhanced images. The SSIM is computed as follows:
where μx and μy are the average pixel values of the original and enhanced images, respectively. σx and σy are the standard deviations of the original and enhanced images, respectively. σxy is the covariance between the original and enhanced images. ϵ is a small constant that avoids a zero denominator.
The LPIPS measures the dissimilarity between the original and enhanced images in feature space using pretrained image encoders, such as VGG, ResNet-50 models. The LPIPS is computed as follows:
where L1 represents the mean absolute difference, and ϕ(x) and ϕ(y) are the feature vectors extracted from the original image (x) and the enhanced image (y) by a pretrained deep learning model designed to capture perceptual similarity. In this study, we used the ImageNet pretrained VGG 16 model.
Evaluation metrics: Compression Decision Agent (CDA)
CR refers to the theoretical amount of compression achieved through image compression. When the original image has an input size of 512 × 512, it is represented by a dimension of \({{\mathbb{R}}}^{512{\mbox{x}}512{\mbox{x}}3}\). However, for VAE512, the representation is \({{\mathbb{R}}}^{1,000}\), while VQVAE512 is represented by \({{\mathbb{R}}}^{128{\mbox{x}}128{\mbox{x}}2}+{{\mathbb{R}}}^{64{\mbox{x}}64{\mbox{x}}2}\), and ESRGAN by \({{\mathbb{R}}}^{128{\mbox{x}}128{\mbox{x}}3}\). The size of the dimension required by each method is expressed as the ratio of the CR. When adaptive compression was determined, the adaptive compression ratio was calculated by assuming the condition of no compression as the baseline, set to a value of one. Therefore, if only the original image was used, it was 100%, and if both were compressed using ESRGAN, it was 6.25%, equivalent to the compression ratio achieved by SwinIR and LDM.
Evaluation metrics: classification performance
The performance of the classification models was evaluated using various widely accepted metrics, including AUROC, AUPRC, accuracy, and F1 score.
-
AUROC: The AUROC evaluates a classifier’s ability to differentiate between positive and negative classes across varying decision thresholds. This provides a comprehensive measure of the discriminatory ability of the model.
-
AUPRC: The AUPRC summarizes the trade-off between precision and recall across different threshold values. This is particularly useful for imbalanced datasets where positive samples are scarce.
-
Accuracy: The proportion of correctly classified instances out of the total number of instances. Although it offers a straightforward assessment of the overall model performance, it may not be suitable for imbalanced datasets.
-
F1 score: The F1 score considers precision and recall, providing a balanced evaluation of the classifier’s performance. It is calculated as the harmonic mean of these two metrics and is particularly valuable when precision and recall are equally important.
Evaluation metrics: segmentation performance
The performance of the segmentation models was assessed using commonly employed metrics, namely the Dice coefficient and the Jaccard index.
-
Dice score: The Dice score measures the spatial overlap between the predicted segmentation mask and the ground-truth mask, expressed as the harmonic mean of precision and recall (Equation (7)).
-
Jaccard index: The Jaccard index evaluates the similarity between two sets by computing the ratio of intersections to unions, and is also known as the Intersection over Union (IoU) (Equation (8)).
The Dice coefficient was computed as:
The Jaccard index was computed as:
where TP, FP, and FN denote true positives, false positives, and false negatives, respectively.
Statistics and reproducibility
For pretraining the FIEs, 30 WSIs were randomly sampled from each TCGA project. During patch extraction, patches were also randomly selected within tissue regions to ensure representative sampling.
For the reconstruction performance of the FIE, SSIM, PSNR, and LPIPS were computed for each patch and then averaged across the evaluation set to obtain global scores. In the VTT, model performance was statistically compared against chance-level accuracy (0.5) using a two-sided Wilcoxon signed-rank test, with P < 0.05 considered statistically significant. No other formal statistical tests were applied.
For downstream classification and segmentation tasks, model performance was evaluated using AUROC, AUPRC, Dice score, and Jaccard index as the primary evaluation metrics. For patch-level analyses, these metrics were calculated per patch and averaged within slides, whereas for slide-level analyses, patch-level features were aggregated using attention weights to generate slide-level representations for classification. All downstream experiments were repeated five times with different random seeds (42, 43, 44, 45, and 46), and mean performance values were reported. All experiments were conducted under identical hyperparameter settings, except for the specific comparison variable being tested.
No statistical method was used to predetermine sample size, and no data were excluded from the analyses. The experiments were not randomized, and the investigators were not blinded to allocation during experiments or outcome assessment. All datasets used in this study are publicly available, and all codes, model configurations, and trained weights will be released to ensure reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
This study used publicly accessible WSI datasets. TCGA data (including TCGA-RCC, TCGA-NSCLC, TCGA-BRCA) can be accessed via the NIH Genomic Data Commons (https://portal.gdc.cancer.gov). CPTAC datasets are available through The Cancer Imaging Archive (https://www.cancerimagingarchive.net/). Additional publicly available datasets used in this work include Camelyon16 (https://camelyon16.grand-challenge.org/), NCT-CRC (https://zenodo.org/record/1214456), MHIST (https://bmirds.github.io/MHIST/), LI (https://zenodo.org/records/10020633), SICAPv2 (https://data.mendeley.com/datasets/9xxm58dvs3/1), WSSS4LUAD (https://wsss4luad.grand-challenge.org/), SNOW (https://zenodo.org/records/6633721), NuInsSeg (https://www.kaggle.com/datasets/ipateam/nuinsseg), and PanNuke (https://warwick.ac.uk/fac/cross_fac/tia/data/pannuke). The CBTN dataset is available under controlled access and can be requested from the Children’s Brain Tumor Network (https://cbtn.org/). Source data are provided with this paper.
Code availability
A demo version of the AdaSlide code, including example usage, is available for academic research through GitHub (https://github.com/PathfinderLab/AdaSlide_demo) and Zenodo (https://doi.org/10.5281/zenodo.17445388), and the corresponding demo outputs can be accessed at https://zenodo.org/records/15665900. The full AdaSlide training and evaluation code is provided at https://github.com/PathfinderLab/AdaSlide and also archived in Zenodo (https://doi.org/10.5281/zenodo.17445388). The trained FIE and CDA model weights are available via Zenodo (https://zenodo.org/record/11069591). Additional publicly available resources used in this study include HoverNet (https://github.com/vqdang/hover_net), ESRGAN (https://github.com/XPixelGroup/BasicSR), VQVAE (https://github.com/rosinality/vq-vae-2-pytorch), KAIR (https://github.com/cszn/KAIR), LDM (https://github.com/CompVis/latent-diffusion), and CLAM (https://github.com/mahmoodlab/CLAM).
References
Abels, E. et al. Computational pathology definitions, best practices, and recommendations for regulatory guidance: a white paper from the digital pathology association. J. Pathol. 249, 286–294 (2019).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology-new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Lipkova, J. et al. Artificial intelligence for multimodal data integration in oncology. Cancer Cell 40, 1095–1110 (2022).
Heinz, C. N., Echle, A., Foersch, S., Bychkov, A. & Kather, J. N. The future of artificial intelligence in digital pathology–results of a survey across stakeholder groups. Histopathology 80, 1121–1127 (2022).
Echle, A. et al. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br. J. Cancer 124, 686–696 (2021).
Mandair, D., Reis-Filho, J. S. & Ashworth, A. Biological insights and novel biomarker discovery through deep learning approaches in breast cancer histopathology. NPJ Breast Cancer 9, 21 (2023).
Tran, K. A. et al. Deep learning in cancer diagnosis, prognosis and treatment selection. Genome Med. 13, 1–17 (2021).
Baxi, V., Edwards, R., Montalto, M. & Saha, S. Digital pathology and artificial intelligence in translational medicine and clinical practice. Mod. Pathol. 35, 23–32 (2022).
Tolkach, Y., Dohmgörgen, T., Toma, M. & Kristiansen, G. High-accuracy prostate cancer pathology using deep learning. Nat. Mach. Intell. 2, 411–418 (2020).
Steyaert, S. et al. Multimodal data fusion for cancer biomarker discovery with deep learning. Nat. Mach. Intell. 5, 351–362 (2023).
Zhang, Z. et al. Pathologist-level interpretable whole-slide cancer diagnosis with deep learning. Nat. Mach. Intell. 1, 236–245 (2019).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer Cell 40, 865–878 (2022).
Ghosh, A. et al. The potential of artificial intelligence to detect lymphovascular invasion in testicular cancer. Cancers 13, 1325 (2021).
Lee, J., Ahn, S., Kim, H.-S., An, J. & Sim, J. A robust model training strategy using hard negative mining in a weakly labeled dataset for lymphatic invasion in gastric cancer. J. Pathol. Clin. Res. 10, e355 (2024).
Graham, S. et al. Hover-Net: simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 58, 101563 (2019).
Israel, U. et al. A foundation model for cell segmentation. Preprint at bioRxiv https://doi.org/10.1101/2023.11.17.567630 (2023).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Azizi, S. et al. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nat. Biomed. Eng. 7, 756–779 (2023).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Xiang, J. et al. A vision–language foundation model for precision oncology. Nature 638, 769–778 (2025).
Wang, X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature 634, 970–978 (2024).
Xu, H. et al. A whole-slide foundation model for digital pathology from real-world data. Nature 630, 181–188 (2024).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual–language foundation model for pathology image analysis using medical Twitter. Nat. Med. 29, 2307–2316 (2023).
Thorstenson, S., Molin, J. & Lundström, C. Implementation of large-scale routine diagnostics using whole slide imaging in Sweden: digital pathology experiences 2006-2013. J. Pathol. Inform. 5, 14 (2014).
Aswolinskiy, W. et al. Neural image compression for non-small cell lung cancer subtype classification in H&E stained whole-slide images. In Medical Imaging 2021: Digital Pathology Vol. 11603 1160304 (SPIE, 2021).
Chikontwe, P. et al. Weakly supervised segmentation on neural compressed histopathology with self-equivariant regularization. Med. Image Anal. 80, 102482 (2022).
Jewsbury, R., Bhalerao, A. & Rajpoot, N. M. A quadtree image representation for computational pathology. In Proc. IEEE/CVF International Conference on Computer Vision 648–656 (2021).
Wang, Y. et al. CWC-transformer: a visual transformer approach for compressed whole slide image classification. Neural Comput. Appl. 37, 7485–7497 (2025).
Bidgoli, A. A. et al. Evolutionary deep feature selection for compact representation of gigapixel images in digital pathology. Artif. Intell. Med. 132, 102368 (2022).
Tellez, D., Litjens, G., Van der Laak, J. & Ciompi, F. Neural image compression for gigapixel histopathology image analysis. IEEE Trans. Pattern Anal. Mach. Intell. 43, 567–578 (2019).
Keighley, J., de Kamps, M., Wright, A. & Treanor, D. Digital pathology whole slide image compression with vector quantized variational autoencoders. In Medical Imaging 2023: Digital and Computational Pathology Vol. 12471 344–353 (SPIE, 2023).
Afshari, M. et al. Single patch super-resolution of histopathology whole slide images: a comparative study. J. Med. Imaging 10, 017501–017501 (2023).
Nasr, M. S. et al. Clinically relevant latent space embedding of cancer histopathology slides through variational autoencoder based image compression. In 2023 IEEE 20th International Symposium on Biomedical Imaging (ISBI) 1–5 (IEEE, 2023).
Kingma, D. P. & Welling, M. Auto-encoding variational bayes. Preprint at https://arxiv.org/abs/1312.6114 (2013).
Donahue, J., Krähenbühl, P. & Darrell, T. Adversarial feature learning. Preprint at https://arXiv.org/abs/1605.09782 (2016).
Razavi, A., Van den Oord, A. & Vinyals, O. Generating diverse high-fidelity images with VQ-VAE-2. Adv. Neural Inf. Process. Syst. 32, 14866–14876 (2019).
Li, Z., Li, B., Eliceiri, K. W. & Narayanan, V. Computationally efficient adaptive decompression for whole slide image processing. Biomed. Opt. Express 14, 667–686 (2023).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. In International Conference on Machine Learning 2127–2136 (PMLR, 2018).
Shi, J., Li, C., Gong, T., Zheng, Y. & Fu, H. Vila-mil: Dual-scale vision-language multiple instance learning for whole slide image classification. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 11248–11258 (2024).
Li, B., Li, Y. & Eliceiri, K. W. Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 14318–14328 (2021).
Chen, Y.-C. & Lu, C.-S. RankMix: Data augmentation for weakly supervised learning of classifying whole slide images with diverse sizes and imbalanced categories. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 23936–23945 (2023).
Kather, J. N., Halama, N. & Marx, A. 100,000 histological images of human colorectal cancer and healthy tissue. Zenodo https://doi.org/10.5281/zenodo.1214456 (2018).
Wei, J. et al. A petri dish for histopathology image analysis. In Artificial Intelligence in Medicine: 19th International Conference on Artificial Intelligence in Medicine, AIME 2021, Virtual Event, June 15–18, 2021, Proceedings 11–24 (Springer, 2021).
Silva-Rodríguez, J., Colomer, A., Sales, M. A., Molina, R. & Naranjo, V. Going deeper through the Gleason scoring scale: An automatic end-to-end system for histology prostate grading and cribriform pattern detection. Comput. Methods Prog. Biomed. 195, 105637 (2020).
Han, C. et al. Wsss4luad: Grand challenge on weakly-supervised tissue semantic segmentation for lung adenocarcinoma. Preprint at https://arXiv.org/abs/2204.06455 (2022).
Ding, K. et al. A large-scale synthetic pathological dataset for deep learning-enabled segmentation of breast cancer. Sci. Data 10, 231 (2023).
Mahbod, A. et al. Nuinsseg: A fully annotated dataset for nuclei instance segmentation in H&E-stained histological images. Sci. Data 11, 295 (2024).
Gamper, J., Alemi Koohbanani, N., Benet, K., Khuram, A. & Rajpoot, N. Pannuke: an open pan-cancer histology dataset for nuclei instance segmentation and classification. In Digital Pathology: 15th European Congress, ECDP 2019, Warwick, UK, April 10–13, 2019, Proceedings 15 11–19 (Springer, 2019).
Lilly, J. V. et al. The children’s brain tumor network (CBTN)-accelerating research in pediatric central nervous system tumors through collaboration and open science. Neoplasia 35, 100846 (2023).
Wang, X. et al. Esrgan: Enhanced super-resolution generative adversarial networks. In Proc. European Conference on Computer Vision (ECCV) Workshops (2018).
Liang, J. et al. SwinIR: Image restoration using swin transformer. In Proc. IEEE/CVF International Conference on Computer Vision 1833–1844 (2021).
Rombach, R., Blattmann, A., Lorenz, D., Esser, P. & Ommer, B. High-resolution image synthesis with latent diffusion models. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 10684–10695 (2022).
Zhang, R., Isola, P., Efros, A. A., Shechtman, E. & Wang, O. The unreasonable effectiveness of deep features as a perceptual metric. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 586–595 (2018).
Xu, X., Kapse, S. & Prasanna, P. Histo-diffusion: A diffusion super-resolution method for digital pathology with comprehensive quality assessment. Preprint at https://arXiv.org/abs/2408.15218 (2024).
Faghani, S., Codipilly, D. C., Moassefi, M., Iyer, P. G. & Erickson, B. J. Optimizing storage and computational efficiency: an efficient algorithm for whole slide image size reduction. Mayo Clin. Proc. Digital Health 1, 419–424 (2023).
Schulman, J., Wolski, F., Dhariwal, P., Radford, A. & Klimov, O. Proximal policy optimization algorithms. Preprint at https://arXiv.org/abs/1707.06347 (2017).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. In 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro 1107–1110 (IEEE, 2009).
Liu, Z. et al. Swin transformer: hierarchical vision transformer using shifted windows. In Proc. IEEE/CVF International Conference on Computer Vision 10012–10022 (2021).
Sutton, R. S., McAllester, D., Singh, S. & Mansour, Y. Policy gradient methods for reinforcement learning with function approximation. Adv. Neural Inf. Process. Syst. 12, 1057–1063 (1999).
Acknowledgements
This research was supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health and Welfare, Republic of Korea (grant number RS-2021-KH113146; S.A., S.H.L.). It was also supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (grant number RS-2025-02215813; S.A.), as well as the Digital-Bio AI + X Global Innovative Talent Nurturing Project with Hands-on Experience of the NRF, funded by the Korean government (MSIT) (grant number RS-2024-00441029; S.A.). Additional support was provided by the High-Performance Computing Support Project, funded by the Government of the Republic of Korea (Ministry of Science and ICT) (grant number RQT-25-090105; S.A.).
Author information
Authors and Affiliations
Contributions
J.L. and L.T. carried out the experiments. J.L. wrote the manuscript with support from L.T., D.M.B., D.O., D.K., W.J., S.A. and S.H.L. conceived the original idea. D.O., D.K., S.A. and S.H.L. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare a related patent application covering the methodological aspects of this work (Applicants: Korea University and The Catholic University of Korea Industry-Academic Cooperation Foundations; Inventors: S.A., S.H.L., and J.L.; PCT/KR2025/007948; filed and pending). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, J., Takemaru, L., Bappy, D.M. et al. Adaptive compression framework for giga-pixel whole slide images. Nat Commun 17, 207 (2026). https://doi.org/10.1038/s41467-025-66889-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66889-0