Abstract
Animal pigmentation is essential for concealment and communication in nature. Though selection acts on the phenotypes of an organism, these traits are often resolved to genes that shape pigmentation in populations. In contrast, plasticity in coloration can occur within individuals in response to ambient environmental changes that act through integrative processes. It is well known that some animals often change color to match their visual ecology. The cichlid fish, Astatotilapia burtoni, can switch between robust blue and yellow body morphs in the lab and the wild. Different colors result from the positioning and density of chromatophores between different morphs and patterns. We show that A. burtoni switches from yellow to blue depending on its visual environment by downregulating endothelin receptor B (EdnRBa) mRNA via DNA hypermethylation at a single cytosine residue within its promoter. EdnRBa functions in yellow chromatophores to signal the aggregation of yellow pigments, making yellow less visible. Taken together, the regulation of EdnRBa through DNA methylation in yellow chromatophores contributes to pigmentation changes from blue to yellow morphs and their overall phenotypic plasticity.
Similar content being viewed by others
Introduction
Natural and sexual selection amongst cichlid fish with conspicuously colorful males and cryptic females occurs in males of polygamous species and maternal brood care1. This combination made coloration critical for the adaptive radiation of cichlid fish species throughout Rift Valley lakes in East Africa1. Color differences in cichlids are typically sexually dimorphic, affecting life histories through male competition, female mate choice, and background adaptation2. Different colors can result from complex interactions of neural or hormonal signals3,4,5. While it is widely accepted that genes and their transcription underlie such traits, little is known of how the reversible regulation of genes parallels changes in animal coloration. Several mechanisms have been reported to control color changes, such as cis-regulatory elements6, alternative splicing7, or microRNAs8. Given the plastic nature of animal pigmentation and its susceptibility to the environment, we considered DNA methylation due to its sensitivity to seasonal and social phenomena9,10,11,12. DNA methylation is reversible in its biochemistry13 and targets CpG dinucleotides of gene promoters, leading to decreases in gene transcription14. Here, we asked whether environmental changes in the visual environment might shape gene transcription via DNA methylation.
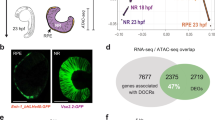
Adult male Astatotilapia burtoni can switch between blue and yellow-colored bodies (see Fig. 1A) in the wild15 and captivity16, though why this happens is unknown. In zebrafish, morphological darkening and lightening have been seen to be induced by ambient light17, suggesting a similar manipulation would also shape the outcome of color morphs in A. burtoni. We discovered this body color change can be induced in response to changes in substrate color (Fig. 1 & S2B, C). It has long been known that adult pigmentation patterns are biologically crucial for animals as countershading and for matching color patterns to habitat backgrounds18,19,20,21. In teleosts, as in many species, coloration is produced by a multilayer assemblage of pigment-containing cells and their aggregation and dispersal of pigments22,23. These pigment-bearing cells, called chromatophores, are categorized based on their pigment characteristics: In skin, there are black melanin-containing cells (melanophores) as well as yellow, orange, or red-colored cells (xanthophores) and reflective iridophores filled with crystalline chemochromes, which interfere, reflect, and scatter light into different wavelengths, such as blue22,24. In A. burtoni, changes in coloration have been documented under neurophysiological control (as seen in the black eye bar25,26) and morphological color change(as seen in body coloration), where reconfiguration of cellular substrates underlies changes in body color27 (See Fig. 2A–D).
A Wild-type (WT) blue and yellow male color morphs. B Pixel frequencies within WT yellow and blue morphs are plotted against significantly different hues. (Two-tailed t-test with Bonferroni correction, N = 9 fish, Error bars are shown as ± the standard error of the mean). C Schematic illustration of the experimental paradigm in which male A. burtoni were placed in blue (28 days), then yellow (28 days), then blue (28 days) visual environments in aquaria. C Photographs of one experimental fish at the end of each cycle showing body color change induced by exposure to different visual environments (Note: All experimental fish for the three color cycles are shown in Fig. S2A). D Pixel frequencies of experimentally induced blue and yellow morphs plotted against significantly different hues in a 256 RGB color range. (Kruskal-Wallis ANOVA, post-hoc two-tailed Mann-Whitney test, and Bonferroni correction. P < 0.05, N = 9 fish, error bars are shown as ± the standard error of the mean).
Changes in the visual environment shape the pigmentation of A. burtoni. Wild-type (WT) blue (A) and yellow (B) morphs; Tail fin sample insets at two magnifications: Scale bars are 600 μm (i) and 200 μm (ii) fin area following a 10−5 M norepinephrine treatment. Yellow xanthophore number/ total area of sampled micrograph (C) and % dispersal of yellow xanthophores (unpaired two-tailed t-test, t(8) = 6.07, p = 0.0003; mean difference = 0.373 ± 0.061 SEM; 95% CI [0.231, 0.515]; η² = 0.822; N = 5 fish per group; variances not significantly different by F(4,4) = 4.06, p = 0.204) (D) of WT blue and yellow color morphs (unpaired two-tailed t-test, t(8) = 4.81, p = 0.0013; mean difference = 0.000589 ± 0.000123 SEM; 95% CI [0.000307, 0.000872]; η² = 0.743; N = 5 fish per group). Variances were significantly different by F(4,4) = 19.40, p = 0.0139.). E Yellow xanthophore number / total area of sampled micrograph (repeated measures ANOVA with Geisser–Greenhouse correction, F(1.576, 12.61) = 21.59, p = 0.0002, η² = 0.73, N = 9 fish; Tukey’s post hoc tests: T1 Blue vs. T2 Yellow, p = 0.0036; T2 Yellow vs. T3 Blue, p = 0.0020) and (F) % dispersal of yellow xanthophores of induced color morphs (see Fig. 1C for environmental colors: blue-yellow-blue) at 28-day intervals. (repeated measures ANOVA with Geisser–Greenhouse correction, F(1.161, 9.284) = 190.9, p < 0.0001, η² = 0.96, N = 9 fish; Tukey’s post hoc tests). **P < 0.01, ***P < 0.001, ****P < 0.0001. Error bars are shown as the standard error of the mean. Source data are provided as a Source Data file.
Many genes are crucial for animal pigmentation across vertebrates28,29; however, we focused our screen on candidate genes unique to A. burtoni and related teleosts (Fig. 3A, B, S4). Prior studies of body color in fish species identified transcriptional differences that regulate pigmentation in natural environments in A. burtoni3, Lutjanus erythropterus (Red Snapper)30, and the common carp31. Across these species, endothelin signaling changed across the transcriptome, highlighting its relevance to plastic pigmentation. Additionally, endothelin signaling via G-protein receptors signals pigment translocation in yellow xanthophores3,29,30,31,32,33,34,35. Given its importance in the orange/yellow coloration of A. burtoni eggspots3, we further focus on the endothelin B receptor to explore DNA methylation as a process capable of shaping body color changes in A. burtoni. Here, we investigate transcriptional differences in endothelin signaling, examine their epigenetic plasticity, and test their functional role in regulating yellow and blue color morphs in an African cichlid.
A mRNA abundance of EdnRBa relative to Rpl7 (ribosomal housekeeping gene) as a function of environmental condition (See Fig. 1C) (one-way ANOVA, F(2,24) = 6.36, p = 0.0061, η² = 0.35; Tukey’s post hoc tests: T1 vs. T2, p = 0.0063; T2 vs. T3, p = 0.0391). B Schematic illustration of the CpG sites directly upstream of the transcriptional start site (TSS) for EdnRBa; (B-bottom) Corresponding DNA methylation levels for three sites upstream of the transcriptional start site. (Site +160: repeated measures ANOVA with Geisser–Greenhouse correction, F(1.270, 10.16) = 114.5, p < 0.0001, η² = 0.93, N = 9; Tukey’s post hoc tests). C Luciferase activity relative to the total protein of the EdnRBa promoter in pCpGl transfected into HEK293T cells; S-Sense; AS-antisense; mS-methylated sense; (one-way ANOVA, F(2,6) = 58.09, p = 0.0001, η² = 0.95, N = 3 independent experiments, each performed in triplicate; Dunnett’s post hoc tests: S vs. AS, p = 0.0018; S vs. mS, p < 0.0001). D Ex vivo treatment of caudal fin with PBS, Sarafotoxin (SRTX), and IRL1620; (one-way ANOVA, F(2,9) = 8.31, p = 0.0090, η² = 0.65, N = 3 per drug treatment and N = 6 of PBS treatment; Tukey’s post hoc tests: S6C vs. PBS, p = 0.0300; IRL1620 vs. PBS, p = 0.0163) (E) Ex vivo treatment of caudal fin with PBS, SRTX, and IRL1620. (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. ±Error bars are shown as the standard error of the mean.) Source data are provided as a Source Data file.
Results
Color differentiation and visual environment effects
We quantified the naturally occurring color differences between yellow and blue male A. burtoni from photographs, measuring pixel frequencies with 8-bit color analysis (See supplemental methods; Fig. 1B, D). Comparing whole body images, wildtype (WT) blue and yellow morphs housed on brown gravel had significant color differences in 134/256 colors, with each morph clustering independently (Fig. 1B & S1A). Next, we measured the response of male coloration to different visual environments by placing individuals sequentially into blue (T1), yellow (T2), and blue (T3) environments for 28 days each (Fig. 1C & S2). Males were housed alone in aquaria with yellow or blue gravel substrate, draped on three sides with either a blue or yellow plastic sheeting (Fig. 1C & S2B, D). Animals were photographed at the end of each exposure period, and their images were analyzed. We found differences across groups T1-T3 (Fig. 1D) with blue substrates showing matching profiles (see supplemental methods). Dissimilarities between animals in blue visual environments (T1/T3) compared to those in yellow visual environments (T2) were also evident in a chi-squared heat map for all hues across all images (Fig. S1B). Furthermore, differences between yellow and blue morphs were most notable in the range corresponding to orange-yellow coloration.
Cellular basis of induced color change
To discover how differences in fish color were generated, we sampled caudal fin tissue from the same animals at the end of each time point (T1-T3) (Fig. 1C). This allowed for the repeated measurement of tissue to assess cellular and molecular changes corresponding to whole-animal color change (Fig. 2), controlling for genetic variability among individuals. WT yellow males, in aquaria with brown gravel, had an increase in the number of yellow xanthophores compared to blue males under the same conditions (Fig. 2A–C). In addition to quantitative differences in chromatophore number, we observed that WT blue morphs had yellow xanthophores with their pigment robustly aggregated compared to yellow morphs, where the pigment was dispersed throughout the cell (Fig. 2D). In animals with induced body color changes, we found an increased number of yellow xanthophores in those housed in a yellow visual environment compared to those in a blue visual environment (Fig. 2E). Similarly, animals housed in a yellow visual environment had robust dispersal of yellow xanthophores compared to those housed in a blue visual environment (Fig. 2F). Despite a change in number (Fig. S3A, D), melanophores showed no shift in aggregation/dispersal in WT morphs or those induced by the visual environment (Fig. S3B, E). This data suggested that solely yellow xanthophores were shifting the function of pigment dispersal.
EdnRBa regulation and environmental response
Earlier reports showed that yellow xanthophore pigment aggregation results from EdnRBa agonizts in medaka34 and EdnRBa expression was paralleled by upregulated transcripts in the orange fin spots of A. burtoni6. Therefore, we considered EdnRBa a candidate gene regulating body color change in repeated caudal fin samples taken from animals before transfer. We found EdnRBa overexpressed in fin tissue from fish in a blue visual environment, decreased expression after transfer to a yellow visual environment, and increased when returning it to a blue environment (Fig. 3A).
We asked whether changes in DNA methylation of EdnRBa might be responsible for these differences by measuring methylation of 3 kb of the promoter and first exon (Fig. S4A, B). We found a robust change in DNA methylation at a CpG dinucleotide site corresponding to fish color (−160 bp upstream of the transcriptional start site; Fig. 3B & S4A, B). Specifically, DNA methylation at CpG −160 increased when fish were housed in a yellow environment (T2) and decreased when fish were housed in both blue environments (T1 and T3; Fig. 3B).
Next, we tested whether methylation of a putative cis-regulatory element containing three CpGs (−259, −160, and −125) could contribute to the transcription of EdnRBa within a CpG-less luciferase reporter construct36. When compared both to sense (S; Fig. 3C) and antisense (AS; Fig. 3C) controls, the in vitro methylated promoter (mS; Fig. 3C) showed a threefold decrease in luciferase activity compared to the sense probe in cultured HEK293T cells. To test the role of EdnRBa directly, we treated fin tissue with EdnRBa selective agonizts IRL1620 and Sarafotoxin 6c (SRTX), showing it decreased yellow coloration through the aggregation of pigment in yellow xanthophores (Fig. 3D, E).
Discussion
We show that A. burtoni males reversibly change between color morphs in response to their visual environment by changing the number and function of yellow xanthophores through EdnRBa signaling. While changes in the number of pigment cells occurred after two weeks, the variation of dispersal between blue and yellow males occurred after four weeks (Fig. S8). We showed that yellow pigment aggregation is controlled by the reversible DNA methylation of a cytosine residue in the promoter of the EdnRBa gene when the animal was housed in a yellow visual environment. Hypermethylation of this site decreased transcription of the EdnRBa gene in yellow morphs, while its hypomethylation resulted in increased transcription in blue morphs (Fig. 4). This CpG site may contribute to a cis-regulatory role in part demonstrated by in vitro methylation changes measured with luciferase expression and bisulfite sequencing data. We suspect it may present an interaction with tentative transcription factor binding sites for NFAT5, HOXC10, and MNT (Fig. S6C). Finally, direct stimulation of EdnRBa pharmacologically produced cell-specific aggregation of yellow xanthophores, confirming its role.
(Top) With CpG site −160 methylated, mRNA abundance (red) is reduced, and endothelin signaling is attenuated, permitting yellow pigment to spread through yellow xanthophores. (Bottom) DNA hypomethylation at CpG −160 increases EdnRBa mRNA (red) abundance, sensitizing yellow chromatophores to endogenous endothelins, resulting in increased aggregation.
Our data support a novel role for DNA methylation as a dynamic mechanism modulating morphological plasticity via yellow xanthophores through endothelin signaling. To our knowledge, this is the first example of epigenetic control of reversible body color change in vertebrate animal pigmentation, in contrast to extensive evidence confirming the genetic and developmental bases for animal pigmentation28,33,37. Neural mechanisms can produce rapid color changes important for behavioral interactions, as in the A.burtoni eyebar through chromatophore dispersal/aggregation25,26, as opposed to morphological changes that occur on longer time scales of days to weeks27. As shown here, changes in DNA methylation can alter colors via cellular and molecular substrates likely appropriate for longer-term cryptic2 or social strategies16. We suspect that the role of EdnRBa in yellow xanthophores is amplified by the time it takes to increase yellow xanthophores after 2 weeks (Fig. S8A) and to robustly disperse/aggregate yellow pigments after 4 weeks (Fig. 2, S8B). We show that yellow pigment dispersal more than doubles between three and four weeks of induction(Fig. S8B), suggesting that changes in the transcription and DNA methylation of EdnRBa occur after a month of tissue remodeling in the periphery. This is separate from the speed at which yellow xanthophores respond to Endothelin pharmacology (Fig. 3D, E), which only resolves receptor function to yellow xanthophores in WT fish.
This strong link between DNA methylation of the EdnRBa gene, producing reversible color morphs in A. burtoni, may be widespread among fish species that change color throughout their life. Our data suggest that hyper- and hypomethylation of CpG −160 may be subject to the recruitment of de novo DNA methyltransferases or TET hydroxylases that facilitate the addition or removal of methylation, respectively13. We suspect this is triggered by visual information entering the eye to the brain, signaling to neuroendocrine glands to modulate morphological changes in peripheral skin tissues via secondary messengers. For example, in A. burtoni, Djikstra et al. showed that α-MSH, known to be secreted from the pituitary, could increase the dispersal of yellow xanthophores38.
Since the colors of a fish involve the coordinated dispersal/aggregation of various chromatophores, we would expect various changes in the transcriptome of different pigment cells to accommodate a whole organism’s change in color. We focus predominantly on yellow xanthophores as large contributors to yellow morphs; however, we suspect that decreases in the number of melanophores (and other chromatophores Fig. S7) in blue fish (Fig. S3A, B, D, E) also contribute to the “yellowness” of males. Similarly, the role of iridophores, their number, and dispersal may also contribute to the blue hues seen in blue males22,24. We suspect that neurophysiology modulates the earliest changes in color by aggregating all yellow xanthophores on yellow gravel (first few minutes), followed by the secretion of neuroendocrine signals that remodel chromatophore biology in the periphery (days-weeks) (Fig. S8A). Over a month, we see longer-term changes in xanthophores’ aggregation/dispersal function (Fig. S8B). Effectively, processes, such as DNA methylation may contribute to shifting the baseline of a reaction norm of an animal’s environmentally-triggered phenotypes.
Nuptial body coloration among male cichlids is vital for sexual selection27,39,40,41,42, with blue males being more sexually attractive to females43. Phenotypic plasticity in animal coloration may also contribute to antipredator camouflage, with blue males capable of concealing themselves from avian predators than yellow counterparts2. The transcription of various genes is likely to be involved and related to cell organization, pigment metabolism, and proliferation in chromatophores28. While there are various candidate genes to consider (see other screened candidates, Fig. S4C–E), we focused on EdnRBa due to robust changes in DNA methylation (Supp Fig. S4A, B). Transcriptional control of color morphs through DNA methylation in A. burtoni via endothelin signaling is consistent with its action in diverse pigmentation patterns in other species resulting from genetic mutations in EdnRBa34,37,44,45,46,47.
Waterland et al. reported that dietary folate, a methyl donor, given to pregnant mothers resulted in the DNA hypermethylation of the Agouti gene locus in offspring, generating grades in mottled coat colors48. In contrast to altering DNA methylation exogenously, we used repeated sampling of individuals to show that DNA methylation can be modulated by the visual environment in an animal. DNA methylation remains dynamic into adulthood and may be widespread in facilitating gene-by-environment interactions related to morphological plasticity10,27.
Considering the rapid speciation of cichlid species across the East African rift lakes, the phenotypic plasticity facilitated by DNA methylation may convey unique advantages for the fitness of A. burtoni males. Color polymorphism may underlie balancing selection during seasonal algal blooms49 and predation by avian predators2. In this environment, blue morphs might appear more cryptic while yellow morphs may better communicate social information related to male competition, possibly influencing female mate choice15,50. Since males are highly conspicuous during courtship, needing to be attractive to females while avoiding predation, different colors might aid in creating territories in novel terrain to increase camouflage. In light of recent evidence of high genetic conservation within diverse cichlid lineages51,52, diverse mechanisms of gene regulation in animals, including epigenetically influenced phenotype plasticity, likely affected cichlid diversification and speciation.
Methods
Animals and tissue collection
Laboratory-reared Astotilapia burtoni cichlid fish derived from wild-caught isogenic stock50,53. All animals were part of a bottleneck population with low genetic variation bred from 1979 until 2024, making this population well-suited to assay epigenetic variation. These stocks were used for all experimental procedures approved by the Stanford Animal Use Committee (Stanford APLAC protocol #9662) and Queens College Institutional Animal Care and Use Committee Protocol #198. Animals were reared in community tanks containing a social system typical of their natural habitats. The physical conditions matched their natural habitat of Lake Tanganyika (12:12 full spectrum light-dark regimen; pH of 7.8–8; temperature of 27 °C and water mineral conditions; Seachem, MFGR)50,53.
In nature, dominant A. burtoni can be either yellow or blue50,53, which is also true in laboratory16 where they live above a neutral brown substrate of sandy gravel. Naturally blue and yellow-colored males (N = 8 fish) were reared in community aquaria (113.5 L) until ~4 months old when they began to show body coloration15. All animals used in these experiments were chosen before any male-specific body coloration developed and were socially isolated when entering color-changing paradigms. As such, these males never had sex or socially interacted with other fish once they moved to induction tanks with either blue or yellow gravel colors. These males (N = 9 fish) taken from regular brown gravel substrate were individually housed for 28 days in aquaria (12.5 L) with blue gravel (#40508, Spectrastone). Three sides of each aquarium had a non-reflective blue covering (4010 BL NW Enterprises). This followed 28 days in aquaria with yellow gravel (#20505, Spectrastone) and three sides with a yellow non-reflective covering (4010 HY NW Enterprises). Following this, they were returned for 28 days to “blue” aquaria with blue gravel and blue cover as in the first condition. We used 28 days since it provided ample time for males to assume a stable blue or yellow morph without the likelihood of switching back or forming intermediary phenotypes. Caudal fin was used as a proxy for animal coloration since it was easily accessible to sample from the same animal and reliably represented coloration in comparisons between WT blue and yellow males (See Supplemental Figure S7).
Photographing fish and subsequent image analysis
At the end of each exposure to the blue or yellow environment, each fish was removed and photographed within 30 s on a white paper towel on a white laminate sheet. Tissue collection was carried out between 11 AM and 12 PM, 3–4 h after lights were turned on. Fin clips were taken and stored either in PBS for subsequent microscopy or snap-frozen on dry ice for storage at −80 °C for molecular analyses. Fin clips allowed repeated sampling from the same animals and were used as a proxy of body coloration for all molecular analyses. Photographs were taken (Sony Alpha NEX-C3; F stop. 4.5, exposure time 1/80 s, ISO 200) in a 12”x12”x12” lightbox (60121600, Cowboy Studio) with two full-spectrum 45 W 5500 K 2800Lumens 120 V LED lamps (1146-55-FS, ALZO) with the fish placed in an identical position each time. Aperture, shutter speed, and ISO were set manually to ensure constant conditions. All photos were taken between 11:00 AM and 12:30 PM. Figure S2A shows the individual fish as they were photographed after exposure to each condition.
Pictures were converted from the Sony ARW format to compressed JPEG to assess whole-body coloration. Pixel density after conversion was 4,912 ×3,264 pixels. The fish’s body was cropped for image analysis in Adobe Photoshop CS6. RGB values of each image were transformed into HSV values (H-values) and binned into 256 different colors. All binned H-values were summed up for the images belonging to each group to create the histogram of group hue. Hue counts for each image were normalized to the total number of pixels making up the masked image of the whole fish. To illustrate the similarities and differences between groups, we plotted hue values as a heat map where each square represents the similarity between all normalized hues in collected images of whole fish54,55. Darker squares correspond to greater similarity between rows and columns (Figure S1).
Chromatophore characterization
We measured the base level of coloration while controlling for superficial changes by subjecting all fin clips to a 10−5 M norepinephrine (NE) (A7257-500MG, Sigma Aldrich) treatment, which aggregated all chromatophores from one another (Figs. 2, S3, 7, 8)34. Following aggregation, chromatophores could be accurately counted and characterized for their overall surface area (dispersal/aggregation). NE-treated cells were counted within a controlled area in each fin clip for each color morph in ImageJ. Images were thresholded within LAB color space where +/-B value could account for the yellow-blue variation to robustly label yellow xanthophores (See Figs. 3E and S5). These cells were then masked, counted, and with surface area quantified relative to the total surface area of the micrograph using basic ImageJ functions (Fig. 3D, E). Fin clips were mounted onto a slide with a coverslip (#18606-20, Polysciences). We focused solely on black melanocytes and yellow, red, and orange xanthophores. We excluded iridophores due to their plaque-like organization and lack of visible pigment, which made them particularly difficult to quantify using bright field microscopy.
Chromatophore types were initially surveyed within rays, inter-ray red spots, and between red spots of natural blue and yellow fish (Fig. S7) to establish a reliable counting procedure. We observed robust differences in melanophores and yellow xanthophores within the inter-ray membrane between red spots (Figure S7). This location was assayed in all subsequent measurements and designated within the boundaries of where the first red xanthophores appear adjacent to iridiphore plaques. We controlled for the area by randomly sampling three photos within the dorsal half closest to the trunk. Photos were taken of each fish between red spots on the upper half of the caudal fin closest to the trunk and counted manually by an observer blind to the condition.
Testing the function of EdnRBa
Whereas we measured overall dispersal/aggregation and cell counts in Fig. 2 using NE, for Fig. 3, we only sought to characterize aggregation/dispersal responses to EdnRBa pharmacology, and no NE was used at all. To understand the function of EdnRB transcription patterns in specific chromatophores, we used the EDNRB agonist, IRL1620 (1196, Tocris) and Sarafotoxin 6 C (SRTX) (S6545, Sigma-Aldrich) to identify which pigment cell populations are sensitive to endothelin signaling. For microscopy, caudal fin tissue was removed with a razor blade and cut lengthwise in thirds. Each third of tissue came from a single individual controlling chromatophore responses internally for all drug treatments. For pharmacology, each fin clip was kept in PBS, IRL1620, or SRTX for 30 min, after which tissue was mounted onto a glass slide and imaged. All photographs were collected on a microscope (Zeiss Axioimager M1 upright wide field fluorescence/DIC microscope with Zeiss CCD camera).
cDNA conversion and qRT-PCR
RNA was isolated from lysed tissue using Allprep kit (80004, Qiagen) with 1ug of RNA used for cDNA conversion and DNAse cleanup (iScript gDNA clear cDNA synthesis kit;1725034, Biorad) according to manufacturer’s guidelines. Primers were designed over exon boundaries and validated with gel electrophoresis, melting curves, and sequencing (See primer sequences in Supplementary Table 1). qPCR was carried out to measure relative EdnRBa (NCBI gene; Acc:102302496) mRNA abundance using RPL7 as reference genes in fin tissue since it has been used in previous studies and was not variable across compared groups6. Cycling conditions were as follows: 5 min 95 °C; 30 s 95 °C. 1 min 60 °C. 30 s 72 °C; 72 °C 10 min.
DNA methylation mapping
To map DNA methylation in the chromatophores of the tail fin clips, standard bisulfite conversion and pyrosequencing methods were used56,57. DNA was isolated from lysed tissue (Allprep kit 80004, Qiagen) with 3ug of DNA used for bisulfite conversion (EpiTect kit 59104, Qiagen) following the manufacturer’s protocols. Based on preliminary experiments, we examined the methylation state of EdnRBa. We assayed methylation within 3 kb of the EdnRBa transcriptional start site (TSS) across 5 PCR amplicons (Fig. S4A and Table S1). Due to issues of sequence context and distance from TSS, DNA methylation of individual CpGs in the EdnRBa promoter were considered independent variables (Figure S4A, B). In total, we mapped 20 CpG dinucleotides. These were chosen from a series of empirically tested amplicons designed on Qiagen’s Pyromark Assay Design software (Qiagen). Across this screen, we focused on robust changes in the R1 amplicon that mapped 4 of the first 5 CpG dinucleotides upstream of the TSS (See Figures S4, S6B). Using the fold change between blue and yellow fins, we noted two cytosines (CpG −160 and −1317) that deviated meaningfully from 1 (Figure S4B). From these two CpGs, we focused on CpG-160 as it was closer to the transcriptional start site than the more distal CpG-1317.
PCR primers were designed to amplify the EdnRBa promoter with cycling conditions: 5 min 95 °C, 1 min 95 °C, 2.5 min 60 °C, 1 min 72 °C for 35 cycles; 72 °C 10 min. Biotinylated products were precipitated as described previously56,57. Briefly, products were incubated with 2ul of streptavidin sepharose high-performance beads (17-5113-01, GE Healthcare) in binding buffer (10 mM Tris-HCl 2 M NaCl 1 mM EDTA 0.1% Tween 20) for 10 min shaking. Products were denatured in 0.2 M NaOH for 5 s, then 70% Ethanol for 5 s, and washed in 10 mM Tris-Acetate. Beads were loaded onto Pyromark 24-well plates (978756, Qiagen) in annealing buffer (20 mM Tris-acetate, 2 mM MgAc2) and 3uM sequencing primer.
Luciferase Assay of EdnRBa Promoter Efficacy
To test the function of the EdnRBa promoter, we cloned 117 bp, including the differentially methylated site −160 along with two adjacent CpGs (−254 & −124) into a CpG-free luciferase reporter construct pCpGl36, performed in vitro methylation and subsequently used cell transfection to assay the ability of this 117 bp fragment to drive transcription comparing a methylated and unmethylated state(See Figure S6A). To validate in vitro methylation assays, we included a single CCGG site upstream of the cloned promoter with a BamHI (GGATCC) site in front of two cytosine nucleotides, making a CCGG site (See Figure S6A). This CCGG site was then targeted by an isoschizomer pair of methylation-sensitive and insensitive restriction enzymes (HpaII/MspI) to validate methylation status before transfection into HEK293T cells. Luciferase assays were done by cloning the EdnRBa promoter with HindIII and BamHI sites, flanking 117 bp from the differentially methylated CpG −160 (See Table S1). Plasmids were methylated in vitro using SssI methylase (#M0226L, New England Biolabs) for three hours, using S-adenosyl-methionine as a methyl donor twice during the reaction. Digestion with methylation-sensitive isoschizomers HpaII and MspI was used to assess the quality of the methylation reaction. 2.5 ug of plasmid was transfected into HEK293T cells using Lipofectamine 3000 in 6 well plates in triplicate following the manufacturer’s guidelines. Luciferase activity was measured by preparing protein extracts using a 1X luciferase lysis solution (E1531, Promega) and adding to 100ul of luciferase reagent (E1483, Promega) followed by measurements in 96 well format on a Cytation 3 multi-mode plate reader (Biotek) set to measure luminescence. Figure S6 was created in Biorender with TRANSFAC58 predicting transcription factor binding sites in the 117 bp amplicon used in luciferase assays.
Statistical analyses
Analyses between wildtype blue and yellow males regarding colors, cell number, and area(Fig. 2C, D, S4A, C–E, S7) were analyzed with a Student’s t-test. All analyses made between the induced coloration of males through changes in visual environment (T1-T3) regarding cell number, area, EdnRBa transcription, and EdnRBa DNA methylation used a repeated measures ANOVA with Tukey’s post-hoc test. Luciferase assays were analyzed with a one-way ANOVA followed by Tukey’s post-hoc test. In our color-changing paradigm, normalized values for each color from individual images in each group were compared using Kruskal-Wallis ANOVA in Matlab R2016b (p-value threshold <0.01), followed by a post-hoc Mann-Whitney test and Bonferroni correction for multiple comparisons. All significantly different colors were then plotted to show the color distribution differences among groups T1-T3.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study have been deposited in the Zenodo database under https://doi.org/10.5281/zenodo.17136613. Processed data from raw data is available in a source data file included at the provided DOI link. Raw data files for pyrograms and qPCR data are restricted under licenses to Pyromark Q24 and C1000 Manager software, respectively. Source data are provided with this paper.
Code availability
All code used in this study is publicly available at: https://github.com/QCAlvaradoLab/FishColorAnalysis. The repository contains the scripts and workflows necessary to reproduce the analyses and results presented in the manuscript. Detailed instructions for installation, usage, and data input formats are provided in the README file59. Any updates or revisions to the code will be maintained on the same repository.
References
Maan, M. E. & Sefc, K. M. Colour variation in cichlid fish: developmental mechanisms, selective pressures and evolutionary consequences. Semin Cell Dev. Biol. 24, 516–528 (2013).
Whitaker, K. W., Alvarez, M., Preuss, T., Cummings, M. E. & Hofmann, H. A. Courting danger: socially dominant fish adjust their escape behavior and compensate for increased conspicuousness to avian predators. Hydrobiologia 1–15, https://doi.org/10.1007/s10750-020-04475-9 (2021).
Santos, M. E. et al. Comparative transcriptomics of anal fin pigmentation patterns in cichlid fishes. Bmc Genomics 17, 712 (2016).
Kratochwil, C. F. et al. Agouti-related peptide 2 facilitates convergent evolution of stripe patterns across cichlid fish radiations. Sci. N. Y. N. Y 362, 457–460 (2018).
Liang, Y., Meyer, A. & Kratochwil, C. F. Neural innervation as a potential trigger of morphological color change and sexual dimorphism in cichlid fish. Sci. Rep. 10, 12329 (2020).
Santos, M. E. et al. The evolution of cichlid fish egg-spots is linked with a cis-regulatory change. Nat. Commun. 5, 5149 (2014).
Huang, J., Fang, W., Li, J., Cai, W. & Lu, J. Full-length transcriptome reveals alternative splicing regulation pattern of skin color variant in red tilapia (Oreochromis spp.). Aquaculture 598, 741963 (2025).
Feng, D. et al. Integrated analysis of microRNA and mRNA expression profiles in Crassostrea gigas to reveal functional miRNA and miRNA-targets regulating shell pigmentation. Sci. Rep. 10, 20238 (2020).
Alvarado, S., Mak, T., Liu, S., Storey, K. B. & Szyf, M. Dynamic changes in global and gene-specific DNA methylation during hibernation in adult thirteen-lined ground squirrels, Ictidomys tridecemlineatus. J. Exp. Biol. 218, 1787–1795 (2015).
Alvarado, S., Fernald, R. D., Storey, K. B. & Szyf, M. The dynamic nature of DNA methylation: a role in response to social and seasonal variation. Integr. Comp. Biol. 54, 68–76 (2014).
Alvarado, S. G., Lenkov, K., Williams, B. & Fernald, R. D. Social crowding during development causes changes in GnRH1 DNA methylation. Plos One 10, e0142043 (2015).
Herb, B. R. et al. Reversible switching between epigenetic states in honeybee behavioral subcastes. Nat. Neurosci. 15, 1371–1373 (2012).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Bio 20, 590–607 (2019).
Fernald, R. D. & Hirata, N. R. The Ontogeny of Social Behavior and Body Coloration in the African Cichlid Fish Haplochromis burtoni. Z. F.ür. Tierpsychologie 50, 180–187 (1979).
Korzan, W. J., Robison, R. R., Zhao, S. & Fernald, R. D. Color change as a potential behavioral strategy. Horm. Behav. 54, 463–470 (2008).
Sugimoto, M. Morphological color changes in fish: Regulation of pigment cell density and morphology. Microsc. Res. Tech. 58, 496–503 (2002).
van Eys, G. J. J. M. & Peters, P. T. W. Evidence for a direct role of α-MSH in morphological background adaptation of the skin in Sarotherodon mossambicus. Cell Tissue Res. 217, 361–372 (1981).
Longley, W. H. Studies upon the biological significance of animal coloration. I. The colors and color changes of West Indian reef-fishes. J. Exp. Zool. 23, 533–601 (1917).
Masazumi, S. Morphological color changes in the medaka, Oryzias latipes, after prolonged background adaptation—I. Changes in the population and morphology of melanophores. Comp. Biochem Physiol. Part Physiol. 104, 513–518 (1993).
Novales, R. R. Color Change Mechanisms of Cold-Blooded Vertebrates. H. Waring. Physiol. Zool. 37, 245–245 (1964).
Bagnara, J. T., Taylor, J. D. & Hadley, M. E. The dermal chromatophore unit. J. Cell Biol. 38, 67–79 (1968).
Ligon, R. A. & McCartney, K. L. Biochemical regulation of pigment motility in vertebrate chromatophores: a review of physiological color change mechanisms. Curr. Zool. 62, 237–252 (2016).
Spaeth, R. A. The physiology of the chromatophores of fishes. J. Exp. Zool. 15, 527–585 (1913).
Muske, L. E. & Fernald, R. D. Control of a teleost social signal: I. Neural basis for differential expression of a color pattern. J. Comp. Physiol. 160, 89–97 (1987).
Muske, L. E. & Fernald, R. D. Control of a teleost social signal: II. Anatomical and physiological specializations of chromatophores. J. Comp. Physiol. 160, 99–107 (1987).
Alvarado, S. G. Molecular plasticity in animal pigmentation: emerging processes underlying color changes. Integr. Comp. Biol. 60, 1531–1543 (2020).
Nüsslein-Volhard, C. & Singh, A. P. How fish color their skin: a paradigm for development and evolution of adult patterns. Bioessays 39, 1600231 (2017).
Singh, A. P. & Nüsslein-Volhard, C. Zebrafish stripes as a model for vertebrate colour pattern formation. Curr. Biol. 25, R81–R92 (2015).
Zhang, Y.-P. et al. Morphological characters and transcriptome profiles associated with black skin and red skin in crimson snapper (Lutjanus erythropterus). Int J. Mol. Sci. 16, 26991–27004 (2015).
Wang, C., Wachholtz, M., Wang, J., Liao, X. & Lu, G. Analysis of the skin transcriptome in two oujiang color varieties of common carp. Plos One 9, e90074 (2014).
Odenthal, J. et al. Mutations affecting xanthophore pigmentation in the zebrafish, Danio rerio. Dev. Camb. Engl. 123, 391–398 (1996).
Irion, U., Singh, A. P. & Nüsslein-Volhard, C. Chapter eight the developmental genetics of vertebrate color pattern formation lessons from Zebrafish. Curr. Top. Dev. Biol. 117, 141–169 (2016).
Murata, N. & Fujii, R. Pigment-aggregating action of endothelins on medaka xanthophores. Zool. Sci. 17, 853–862 (2000).
Spiewak, J. E. et al. Evolution of Endothelin signaling and diversification of adult pigment pattern in Danio fishes. Plos Genet 14, e1007538 (2018).
Klug, M. & Rehli, M. Functional analysis of promoter CPG-methylation using a CpG-free luciferase reporter vector. Epigenetics 1, 127–130 (2006).
Parichy, D. M. et al. Mutational analysis of endothelin receptor b1 (rose) during neural crest and pigment pattern development in the Zebrafish Danio rerio. Dev. Biol. 227, 294–306 (2000).
Dijkstra, P. D. et al. The melanocortin system regulates body pigmentation and social behaviour in a colour polymorphic cichlid fish†. Proc. R. Soc. B: Biol. Sci. 284, 20162838 (2017).
Hohenlohe, P. A. Ecological genomics in full colour. Mol. Ecol. 23, 5129–5131 (2014).
Selz, O. M., Thommen, R., Pierotti, M. E. R., Anaya-Rojas, J. M. & Seehausen, O. Differences in male coloration are predicted by divergent sexual selection between populations of a cichlid fish. Proc. R. Soc. B Biol. Sci. 283, 20160172 (2016).
Wagner, C. E., Harmon, L. J. & Seehausen, O. Ecological opportunity and sexual selection together predict adaptive radiation. Nature 487, 366–369 (2012).
Seehausen, O. & van Alphen, J. J. M. The effect of male coloration on female mate choice in closely related Lake Victoria cichlids (Haplochromis nyererei complex). Behav. Ecol. Sociobiol. 42, 1–8 (1998).
Dijkstra, P. D. et al. Sexual selection may support phenotypic plasticity in male coloration of an African cichlid fish. Proc. B 291, 20241127 (2024).
Ramanzini, G. C., Filadelfi, A. M. C. & Visconti, M. A. Chromatic effects of endothelin family peptides in non-innervated fish, Synbranchus marmoratus, melanophores. J. Exp. Zool. Part Comp. Exp. Biol. 305A, 551–558 (2006).
Kinoshita, K. et al. Endothelin receptor B2 (EDNRB2) is responsible for the tyrosinase-independent recessive white (mo(w)) and mottled (mo) plumage phenotypes in the chicken. Plos One 9, e86361 (2014).
Zhou, L. et al. Genetic characteristic and RNA-Seq analysis in transparent mutant of Carp–Goldfish nucleocytoplasmic hybrid. Genes 10, 704 (2019).
Hayashi, H., Nakamura, S. & Fujii, R. The endothelin receptors that mediate aggregation of pigment in fish melanophores. Comp. Biochem Physiol. Part B Biochem Mol. Biol. 115, 143–152 (1996).
Waterland, R. A. & Jirtle, R. L. Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol. Cell Biol. 23, 5293–5300 (2003).
Horion, S. et al. Optimized extraction of daily bio-optical time series derived from MODIS/Aqua imagery for Lake Tanganyika, Africa. Remote Sens Environ. 114, 781–791 (2010).
Fernald, R. D. & Hirata, N. R. Field study of Haplochromis burtoni: quantitative behavioural observations. Anim. Behav. 25, 964–975 (1977).
Brawand, D. et al. The genomic substrate for adaptive radiation in African cichlid fish. Nature 513, 375–381 (2014).
Malinsky, M. et al. Whole-genome sequences of Malawi cichlids reveal multiple radiations interconnected by gene flow. Nat. Ecol. Evol. 2, 1940–1955 (2018).
Fernald, R. D. Quantitative behavioural observations of Haplochromis burtoni under semi-natural conditions. Anim. Behav. 25, 643–653 (1977).
Schauerte, B. & Stiefelhagen, R. Learning robust color name models from web images. In Proc 21st International Conference on Pattern Recognition (ICPR2012) 3598–3601 (IEEE, 2012).
Schauerte, B. & Fink, G. A. Web-based learning of naturalized color models for human-machine interaction. Int. Conf Digital Image Comput Techniques Appl 498–503, https://doi.org/10.1109/dicta.2010.90 (2010).
Colella, S., Shen, L., Baggerly, K. A., Issa, J.-P. J. & Krahe, R. Sensitive and quantitative universal Pyrosequencing methylation analysis of CpG sites. Biotechniques 35, 146–150 (2003).
Alvarado, S., Rajakumar, R., Abouheif, E. & Szyf, M. Epigenetic variation in the Egfr gene generates quantitative variation in a complex trait in ants. Nat. Commun. 6, 6513 (2015).
Wingender, E. et al. TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res. 28, 316–319 (2000).
Lee, G. QCAlvaradoLab/FishColorAnalysis: MATLAB CODE for DNA methylation of the endothelin receptor B makes blue fish yellow, https://doi.org/10.5281/zenodo.17136614 (2025).
Acknowledgements
Thanks to Drs. M. Tajerian, K. Maruska, L. O’Connell, & R. Sapolsky for comments on earlier manuscript versions. We thank Drs. M. Tajerian and A. Olson for technical services and support (Stanford Neuroscience Microscopy Service, NIH NS069375). NSF Emerging Frontiers grant 1921773 supported W.F. and S.A. An A.P. Giannini Postdoctoral Fellowship supported S.A. VPUE of Stanford University, supported R.B. NIH-NINDS 3490 supported A.B., D.B., and R.D.F. NIH-GM101095 supported A.B., D.B., and R.D.F.
Author information
Authors and Affiliations
Contributions
W.F., S.G.A., and R.D.F. designed the study. S.G.A. and R.B. performed microscopy, color exposures, and cell counting. A.B. and D.B. collected tissue, photographed subjects, and conducted preliminary experiments. S.G.A. performed all molecular analyses measuring EdnRBa transcription, promoter methylation, and in vitro methylation assays. W.F., S.G.A., and G.L. performed image and statistical analyses. S.G.A. and performed pharmacological experiments on fin tissue. W.F., R.D.F., and S.G.A. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Patricia M. Schulte, and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available”.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, W., Blakkan, D., Byrne, A. et al. DNA methylation of the endothelin receptor B makes blue fish yellow. Nat Commun 17, 258 (2026). https://doi.org/10.1038/s41467-025-66936-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66936-w