Abstract
Retinal nerve fibre layer (RNFL) is a non-invasive structural biomarker of cardiometabolic health, yet its biological underpinnings remain unknown. Here, we integrate advanced retinal optical biopsy and artificial intelligence (AI) algorithms with two complementary metabolomic assays across ethnically diverse cohorts to elucidate the metabolic basis underlying RNFL degeneration and its link to cardiometabolic disease (CMD) in Western cohort and Eastern cohort (Guangzhou Diabetic Eye Study, GDES). We identify 26 metabolic biomarkers significantly associated with RNFL thickness, most of which (ranging from 19 to 26) are linked to HDL composition and lipid transport, mediating a substantial proportion of the RNFL–CMD association (e.g., 63.7% for type 2 diabetes and 44.7% for myocardial infarction). AI-driven RNFL metabolic state model stratifies CMD risk with up to 21.8-fold enrichment between risk deciles and augments prediction while translating into clinical utility across genetic and demographic strata, particularly within socially vulnerable populations. This integrated approach highlights RNFL metabolic states as a shared basis underlying retinal–cardiometabolic connections and as early indicators that inform equitable CMD management.
Similar content being viewed by others
Introduction
The eye with the retina as part of the central nervous system offers an anatomically unique opportunity for a non-invasive endoscopic view into cardiometabolic health1,2,3. Taking advantage of the optical transparency of the eye, state-of-the-art non-invasive in vivo retinal scanning now permits detection of subtle changes within the various retinal layers at an unprecedented micrometre-level resolution 4,5, generating new evidence suggesting the relevance of the retinal nerve fibre layer (RNFL) to cardiometabolic diseases (CMDs) proposed by us and others6,7,8,9,10,11. As these scans are risk-free, easy to administer, and require only minimal expertise and time, they are increasingly being conducted routinely in primary care settings and communities, poising RNFL as a scalable surrogate for monitoring cardiometabolic health12,13,14,15.
Despite this promise, the biological underpinnings of the retina–cardiometabolic connections have remained unclear. Elucidating the molecular mediators of this link is crucial not only for establishing RNFL as a valid biomarker but also for revealing early systemic perturbations that may be actionable. Previous research has shown that metabolic factors, such as serum concentrations of triglycerides, low-density lipoprotein (LDL) and high-density lipoprotein (HDL) cholesterol, and creatinine, play a significant role in the pathogenesis of CMDs16,17,18,19,20,21. These metabolic components exert harmful or protective effects on cardiometabolic health by modulating lipid metabolism or suggesting renal impairment18,21. Interestingly, these same circulating metabolic factors have also been found to affect retinal health, as evidenced by their associations with RNFL thickness (RNFLT)22,23,24,25,26,27. Given the reciprocal relationships observed between systemic metabolism and both RNFLT degeneration and CMD outcomes, we hypothesize that a specific metabolic basis may underlie the connection between RNFLT and CMDs, and that characterizing these intrinsic metabolic states may enable early risk stratification across CMD outcomes.
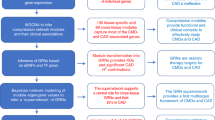
Recent advances in high-throughput metabolomics and machine learning have offered opportunities to characterize systemic biological states and uncover shared disease mechanisms28,29. Building on prior work decoding the metabolic basis underlying (1) pigment epithelium degeneration and type 2 diabetes (T2D) and (2) retinal aging and age-related multisystem disorders30,31, we extended this framework to interrogate retinal–cardiometabolic connections through systematically profiling RNFLT-associated metabolic states and their links with CMD risk. (Fig. 1). To explore the potential of the shared metabolic basis in informing CMD risk, we constructed predictive models integrating these metabolic insights and evaluated their performance and clinical utility across varying genetic susceptibility. Of significant interest is our focus on the benefits for socially vulnerable communities, aiming to mitigate entrenched health disparities in CMD management. Finally, this integrated framework was validated in an independent cohort with distinct ethnic and clinical backgrounds, highlighting the translational potential of RNFLT metabolic states as early indicators of CMD risk.
a The eligible study population was categorized into three distinct groups: population-I, for identifying RNFLT metabolic states; population-II, for unravelling revelations on CMD outcomes; and population-III, for independent concept validation. b To identify RNFLT metabolic states, we conducted retinal scanning and utilized two complementary metabolomic assays. Genotyping was performed to assess genetic susceptibility in individuals. ML algorithms were employed for comprehensive model construction and evaluation. c For CMD outcome risk modelling, the study populations were randomized into training and testing sets. Dataset balancing techniques were applied before feature selection and model training. d The outcomes examined in this study include incident T2D, myocardial infarction, heart failure, stroke, all-cause mortality, and CMD mortality. e Distinct risk stratification and improved predictability and clinical utility were observed for all studied outcomes. f Special attention was given to extending the benefits to women and socially vulnerable communities. g Comprehensive sets of predictors commonly used in the CMD primary prevention were incorporated as benchmark models. Parts of panels a–d and f were created from BioRender (https://BioRender.com/yu7axft) and Flaticon (https://flaticon.com). RNFLT retinal nerve fibre layer thickness, CMD cardiometabolic disease, UKB UK Biobank, GDES Guangzhou Diabetes Eye Study, T2D type 2 diabetes, FGCRS Framingham General Cardiovascular Risk Score, SCORE2 Systematic Coronary Risk Evaluation 2, WHO-CVD World Health Organization Cardiovascular Disease, AHA-ASCVD American Heart Association-Atherosclerotic Cardiovascular Disease, UKPDS UK Perspective Diabetes Study, NZ-DCS New Zealand Diabetes Cohort Study, WAN Wan’s model, BMI body mass index, eGFR estimated glomerular filtration rate, HDL-c high-density lipoprotein cholesterol.
Results
Participants’ characteristics
A total of 7824 participants who underwent both optical coherence tomography (OCT) scanning and metabolomic profiling (population-I), along with 86,014 who underwent metabolomic profiling alone at baseline (population-II), were included in this study (see the “Methods” section). The study population had a median age of 56.8 (8.1) years, of whom 54.6% were women, 10.8% were ever or current smokers, and 21.6% were on antihypertensive medication. Participants who underwent OCT scanning at baseline demonstrated a younger age, male sex, higher educational attainment, lower smoking prevalence, and a decreased likelihood of being on antihypertensive medications compared to those who did not (Table S1). The data of an additional 1286 participants who underwent both OCT scanning and metabolomic profiling from the Guangzhou Diabetic Eye Study (GDES) (population-III) are described in detail below (see the “Methods” section).
RNFLT-associated metabolites and incident CMD risk
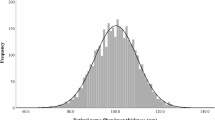
After adjusting for age, sex, ethnicity, assessment centre, household income, Townsend deprivation, education attainment, smoking status, alcohol consumption, body mass index, use of lipid-lowering medications, enzymatic HDL cholesterol, systolic blood pressure and HbA1c level, and applying multiple testing corrections, a total of 26 nuclear magnetic resonance (NMR)-based biomarkers were independently associated with the RNFLT in population-I (Fig. 2a and Table S2). Among these biomarkers, 24 exhibited negative associations, encompassing diverse constituents, such as total lipids, phospholipids, cholesterol, free cholesterol, and cholesteryl esters in various subclasses of HDL particles, as well as phosphatidylcholines, total cholines, and apolipoprotein A1. The adjusted β for these associations spanned a range from −0.201 (95% CI: −0.309 to −0.093) to −0.117 (95% CI: −0.229 to −0.005) per-standard deviation (SD) change. Two biomarkers exhibited positive associations with RNFLT, specifically, the phospholipids-to-total-lipids ratio in very low-density lipoprotein (β = 0.120; 95% CI: 0.015–0.224) and intermediate-density lipoprotein (β = 0.126; 95% CI: 0.026–0.227) subfractions. Sensitivity analyses yielded consistent results when further adjusting for prevalent CMDs (Table S3).
a Attribution of metabolic biomarkers to the outcome-specific ML-driven RNFLT metabolic state (n = 17,203). Individual attributions are aggregated by percentiles, with each dot representing one percentile. The distance of a dot from the circular baseline reflects the strength of the absolute attribution for that percentile. Deviations towards the centre and periphery indicate negative and positive contributions, respectively. Brick colours denote outcomes, while dot colours represent the normalized values for each metabolic biomarker. b–g Cumulative event rates throughout the observation period for CMD outcomes (n = 17,203), stratified by RNFLT metabolic state quantiles. Data are presented as observed event frequencies with 95% CIs shown as shading derived from survival proportions. Red represents the top decile, yellow represents the middle and blue represents the bottom. Illustrations of uniform risk scales are provided in Supplementary Figs. S1 and S2. Source data are provided as a Source Data file. RNFLT retinal nerve fibre layer thickness, ML machine learning, CMD cardiometabolic disease, HDL high-density lipoprotein, VLDL very low-density lipoprotein, T2D type 2 diabetes.
After a median time of 12.2 (interquartile range: 11.5–12.9) years of follow-up in population-II, a total of 6071 participants developed T2D, 2866 developed myocardial infarction, 2537 developed heart failure, 1578 developed stroke, and 6254 died, 1544 of whom succumbed to cardiovascular events. Following adjustment for the same covariates as above, most of the 26 RNFLT-associated metabolites were also associated with CMD outcomes (24 for T2D; 26 for myocardial infarction; 26 for heart failure; 19 for stroke; 19 for all-cause mortality; and 25 for cardiovascular mortality) (Table S4). These included 126 negative associations, with the adjusted hazard ratios (HRs) ranging from 0.688 (95% CI: 0.663–0.714) to 0.969 (95% CI: 0.942–0.996) per-SD increment, as well as 13 positive associations, with the adjusted HRs ranging from 1.048 (95% CI: 1.020–1.077) to 1.345 (95% CI: 1.308–1.383) per-SD increment. Stratified analyses by sex, Townsend deprivation, and educational attainment yielded consistent results (Tables S5–S7).
To assess whether these metabolic biomarkers mediate the relationship between RNFLT and CMD risk, we conducted a mediation analysis comparing the mediation effect of traditional risk factors and RNFL-associated metabolites (see the “Methods” section and Table 1). After accounting for traditional risk factors (same as above), most of the associations between RNFLT and CMD outcomes remained significant, suggesting that these factors explained only a proportion of the observed associations (e.g., 19.4% for myocardial infarction and 7.7% for all-cause mortality). In contrast, when RNFL-associated metabolites were included in the models, the direct associations between RNFLT and CMD outcomes were substantially attenuated toward the null. This change suggested that these metabolic biomarkers accounted for a major proportion of the relationship between RNFLT and CMD risk. The estimated proportions were 63.7% for T2D, 44.7% for myocardial infarction, 49.3% for heart failure, 35.8% for stroke, 17.0% for all-cause mortality, and 38.8% for CMD mortality.
RNFLT metabolic states for CMD prediction
The participants in population-II were split randomly into training and testing sets at an 8:2 ratio to evaluate the utility of RNFL-associated biomarkers in characterizing CMD risk (see the “Methods” section). Baseline characteristics were comparable between the two subsets (all P > 0.05) (Table S1). We first modelled RNFLT metabolic states and observed an increase in event rate trajectories over RNFLT metabolic states across all CMD endpoints (Fig. 2 and S1, S2). For T2D (Fig. 2b), participants in the top decile of RNFLT metabolic states exhibited event rates over 20-fold higher compared to those in the bottom decile (HR = 21.77, 95% CI: 14.18–33.42), suggesting substantial information encapsulated within RNFLT metabolic states for assessing the risk of T2D. For myocardial infarction (Fig. 2d), heart failure (Fig. 2f), and cardiovascular mortality (Fig. 2g), the ratios were greater than five (myocardial infarction, HR = 8.04; 95% CI: 5.15–12.54; heart failure, HR = 5.15; 95% CI: 3.38–7.85; cardiovascular mortality, HR = 5.72; 95% CI: 3.24–10.11). Distinct separation in risk trajectories was also evident for stroke (Fig. 2c) and all-cause mortality (Fig. 2e), with HRs of 4.23 (95% CI: 2.40–7.44) and 2.84 (95% CI: 2.22–3.63), respectively.
We then examined whether RNFLT metabolic states predict CMD risk (see the “Methods” section). Given the substantial class imbalance in CMD outcomes (Table S8), we quantified model performance using the area under the precision-recall curve (AUPRC) as it provides an unbiased assessment in imbalanced settings. C-statistics were additionally reported as a complementary metric to facilitate intuitive interpretability and cross-outcome comparison with differing event rates. In general, RNFLT metabolic states exhibited variable performance across CMD outcomes as assessed by C-statistics (Fig. 3a–f). For T2D, myocardial infarction, heart failure, and CMD mortality, the model fit solely on the RNFLT metabolic states outperformed the Age&Sex model and all individual conventional predictors, including age, blood pressure, enzymatic HDL cholesterol, and LDL cholesterol (Table S9). In the case of other CMD outcomes, the states also outperform most conventional predictors, demonstrating predictive performance competitive with or second only to age. When combined with minimal demographic characteristics (age and sex), RNFLT metabolic states already performed comparably to most established models (Fig. 3a–f). These findings were largely consistent when evaluated using C-statistics (Table S10 and Fig. S3).
a–f Comparison of model performance, including the Age&Sex model, established models, and models incorporating RNFLT metabolic states to predict CMD outcomes, estimated using AUPRC (n = 17,203). Data are presented as estimated coefficients (dots) with 95% CIs indicated by error bars. Different colours denote distinct models, with horizontal dashed lines indicating the performance benchmarks set by the Age&Sex model and four established models. g Comparison of the added benefits derived from integrating RNFLT metabolic states into established models for predicting CMD outcomes across varying genetic susceptibility (n = 17,203). Data are presented as estimated coefficients (dots) with 95% CIs indicated by error bars. Source data are provided as a Source Data file. CMD cardiometabolic disease, AUPRC area under precision-call curve, RNFLT retinal nerve fibre layer thickness, RNFLT MET RNFLT metabolic state, T2D type 2 diabetes, FGCRS Framingham General Cardiovascular Risk Score, SCORE2 Systematic Coronary Risk Evaluation 2, WHO-CVD World Health Organization Cardiovascular Disease, AHA-ASCVD American Heart Association-Atherosclerotic Cardiovascular Disease.
To assess the ability of RNFLT metabolic states to complement conventional models, we calculated the percent change of AUPRC (ΔAUPRC%) to unbiasedly quantify the predictive improvement achieved by incorporating RNFLT-associated metabolic information into established models (see the “Methods” section). Our findings showed that including these states resulted in a significant predictive improvement for the six CMD outcomes in all 30 models (Fig. 3a–f and Table S11). The most substantial improvement was observed for T2D, with a 117.2% improvement over Age&Sex model and an average improvement of 43.8% over all conventional models. However, the improvement became comparable to that observed for other hard cardiovascular outcomes when benchmarking against a model designated for predicting T2D risk (ΔAUPRC% = 9.7%). The average improvements over conventional models were 16.8% for myocardial infarction, 18.6% for heart failure, 17.2% for stroke, 10.6% for all-cause mortality, and 20.1% for CMD mortality. Findings from C-statistics (Fig. S3 and Table S12) and across alternative modelling approaches were largely consistent (Supplementary Results and Table S13).
By calculating polygenic risk scores (PRSs) using genotyped data and meta-analysed external genome-wide association study (GWAS) summary statistics, participants were stratified by genetic susceptibility to CMDs (see the “Methods” section). We prioritized C-statistics to facilitate cross-outcome comparison. In general, models integrating RNFLT metabolic states shared consistent improvement across varying risk categories (Table 2). In populations at high genetic risk, RNFLT metabolic states improved the predictability of most CMD outcomes over FGCRS, except for myocardial infarction (Fig. 3g). While these states provided greater incremental value for heart failure in high genetic risk populations, lower genetic risk populations typically experienced greater improvement in CMD outcome prediction. Additional analyses stratified by PRS for T2D generated from two external GWAS datasets32,33 yielded consistent findings in the T2D assessment (Table S14).
While improvement in discriminatory power is pivotal, it is imperative to evaluate the practical clinical utility of these models (Fig. 4). We observed that all models were well-calibrated for each CMD outcome (Fig. 4a–f). Decision curve analyses demonstrated that improvements in predictability generally lead to greater enhancements in clinical utility (Fig. 4g–l). The integration of RNFLT metabolic states either reduced or eradicated the discrepancy between the Age&Sex model and the established model (FGCRS). Furthermore, the integration of these RNFL metabolic states into the FGCRS contributed to additional improvements in clinical utility across all six CMD outcomes, spanning a wide range of clinical decision thresholds (Fig. 4g–l). Net reclassification improvement for integrating RNFLT metabolic states was moderate yet consistent for all CMD outcomes and models in Table 3.
a–f Calibration for Age&Sex (dotted line), the combination of Age&Sex with RNFLT metabolic states (sky blue), FGCRS (dashed line), and the combination of FGCRS with RNFLT metabolic states (red) in predicting CMD outcomes (n = 17,203). Flesh indicates optimal calibration. g–l Net benefit of clinical utility standardized by endpoint prevalence (n = 17,203), with horizontal dotted grey lines indicating ‘treat none’ and vertical solid grey lines indicating ‘treat all’. Shaded areas represent the incremental benefit of integrating RNFLT metabolic states into the Age&Sex model and FGCRS, respectively. Source data are provided as a Source Data file. RNFLT retinal nerve fibre layer thickness, RNFLT MET RNFLT metabolic state, FGCRS Framingham General Cardiovascular Risk Score, T2D type 2 diabetes.
Benefits to populations experiencing health inequalities
As health disparities exist in CMD outcomes and access to healthcare across diverse population groups34,35,36,37, we were interested in the potential benefits of incorporating RNFLT metabolic states for populations experiencing health inequalities. To address this, we quantified the disparities in discriminatory benefits across different groups with a particular focus on social health determinants, including sex, deprivation, and educational attainment (Fig. 5).
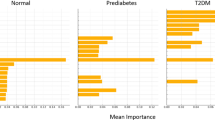
a–f Comparison across sexes (n = 9380 for females and n = 7823 for males). g–l Comparison across socioeconomic statuses (n = 8587 for high deprivation and n = 8591 for low deprivation). m–r Comparison across educational attainment (n = 6939 for university and n = 10,038 for non-university). Data are presented with bars representing the performance and benefits of various demographic groups, with 95% CIs indicated by error bars. Each bar represents a model-level C-index or ΔC-index calculated from all individuals within the corresponding subgroup. Source data are provided as a Source Data file. RNFLT MET RNFLT metabolic state, FGCRS Framingham General Cardiovascular Risk Score, T2D type 2 diabetes, TDI Townsend deprivation index.
In line with previous reports34,35, we observed sexual disparities in CMD risk prediction, as the models fitted on FGCRS predictors exhibited stronger discriminatory power for men compared to women across most CMD outcomes (Fig. 5a–f). However, with the inclusion of RNFLT metabolic states, we observed significantly greater improvements in women’s risk prediction across all CMD outcomes, even in cases where women did not exhibit a pre-existing disadvantage (Fig. 5c and d). This resulted in the eradication or narrowing of the performance gap for CMDs across sexes. In the case of T2D (Fig. 5a) and CMD mortality (Fig. 5f), the substantial improvements observed in women even led to a reversal of the performance gap, where women surpassed men in terms of discriminatory power. Furthermore, integrating RNFLT metabolic states also led to the eradication or narrowing of the prediction gap experienced by socioeconomically disadvantaged populations and those with reduced educational attainment (Fig. 5g–r and Supplementary Results). After employing data balancing techniques38 to address potential biases in the dataset, the main results were consistent (Tables S15 and S16). Taken together, RNFLT metabolic states may represent the common basis of early CMD pathogenesis across demographic strata, narrowing the prediction gap experienced by socially disadvantaged populations.
Extrapolation in an independent cohort
Leveraging data from the GDES39 (Table S17), we extrapolated our analytic framework to an ethnically distinct cohort of 1286 participants who underwent both OCT scanning and metabolomic profiling at baseline (population-III) (Fig. 6). Our findings demonstrated significant associations among 673 metabolites with RNFLT (Table S18), with the vast majority (n = 457 and 279) also associated with incident cardiovascular disease (CVD) (Fig. 6a and Table S19) and mortality (Table S20), indicating their relevance between RNFLT and CMD outcomes across multiple biological processes (Fig. S4). The panorama of these revelations encompassed the replication of phospholipids, phosphoglycerides, cholines and phosphatidylcholines across assays, while offering additional insights spanning a rich spectrum, including aminos, benzene derivatives, alcohols, amines, coenzymes, nucleotides, carbohydrates, organic acids, fatty acids, and heterocyclic compounds (Fig. 6a), which fell beyond the scope of assays in the UKB cohort. Enrichment analyses implicated several biological pathways related to these metabolites, such as aminoacyl-tRNA biosynthesis, arginine and phenylalanine metabolism, glutathione metabolism, as well as nicotinate and pantothenate metabolism (Fig. S4).
a Landscape of the RNFLT metabolic state profile for cardiovascular disease captured by complementary LC–MS assays (n = 1286). Individual metabolite attributions are aggregated by percentiles, with each dot representing one percentile. The distance of a dot from the circular baseline reflects the strength of the absolute attribution for that percentile. Deviations towards the centre and periphery signify negative and positive contributions. Dot colours indicate the normalized values for each metabolite. b–e Comparison of predictability (b and d) and clinical utility (c and e) between established models and models incorporating RNFLT metabolic states for predicting cardiovascular disease across varying genetic susceptibility (n = 1286) (d and e). Data are presented as estimated performance for different models and genetic susceptibility contexts with 95% CIs indicated by error bars. Shaded areas illustrate the incremental net benefit of incorporating RNFLT metabolic states into established models. f–h Comparison of performance for predicting cardiovascular diseases across different demographic groups (n = 1286): sex (f), income (g), and educational attainment (h). Colours denote the absolute performance and benefits of various demographic groups. Source data are provided as a Source Data file. GDES Guangzhou Diabetic Eye Study, RNFLT MET RNFLT metabolic state, FGCRS Framingham General Cardiovascular Risk Score, UKPDS UK Perspective Diabetes Study, NZ-DCS New Zealand Diabetes Cohort Study, WAN Wan’s model.
RNFLT metabolic states remained the most powerful predictors for incident CVD and mortality (Tables S21 and S22), and their integration into both Age&Sex models and FGCRS yielded significant improvements across the overall population (Fig. 6b and Tables S23 and S24) and those with high genetic susceptibility (Fig. 6d and Table 3). Since the GDES cohort represents a diabetic population39, we included three additional benchmark models (UKPDS, NZ-DCS and WAN)40,41,42 customized for the diabetic population, alongside FGCRS for general population risk assessment, where consistent improvements were observed across all models (Fig. 6b, S5 and Tables S25, S26). The decision curve analysis revealed further improvement in clinical utility with the addition of these states (Fig. 6c and e). The findings of comparison analyses across sexes, social classes and education attainment were replicated, where prediction models for women, individuals with low income and those without university education yielded more substantial improvements compared to their respective counterparts (Fig. 6f–h and Table 4). As observed in the UKB, the substantial improvements observed in the case of women even led to a reversal of the performance gap, where women outperformed their men counterparts in terms of discriminative performance (Fig. 6f).
Discussion
Here, we propose and validate the concept of a shared metabolic basis that underlies the observed associations between RNFL degeneration and CMD risk, with the hypothesis that characterizing these metabolic states may enable early risk stratification across CMD outcomes. We demonstrate that (1) metabolomic approaches enable a comprehensive view onto the RNFLT metabolic landscape; (2) RNFLT metabolic states characterize risks of a spectrum of CMD outcomes, mediating a large proportion of the RNFLT–CMD connection; (3) CMD risk trajectories were distinctly separated when stratified by RNFLT metabolic states; (4) RNFLT metabolic states captured residual risk that eludes quantification by traditional risk factors; (5) the performance remained consistent across diverse genetic susceptibility; and (6) these states rectified prediction disparities among populations experiencing health inequalities. Replicated in ethnically distinct cohorts, our integrated approach demonstrates the potential to identify a shared RNFL–CMD metabolic basis that complements cardiometabolic health prediction across the societal spectrum.
RNFL represents the axons of retinal ganglion cells and has recently emerged as a structural proxy for cardiometabolic health6,7,8,9,10,11. Building on our prior work elucidating the metabolic basis of retinal pigment epithelium degeneration and retinal aging in relation to type 2 diabetes and age-related multisystem disorders30,31, we extended this framework to interrogate retinal–cardiometabolic connections. By integrating high-throughput metabolomics with high-resolution OCT B-scan datasets across two large cohorts, we identified distinct metabolic states that reflect both subtle neuroretinal alterations and future CMD risk. Notably, mediation analyses demonstrated that these metabolic states account for a substantial proportion of the RNFLT–CMD associations, suggesting a common metabolic basis linking RNFL degeneration and systemic cardiometabolic regulation. As expected, these states exhibited robust and generalizable predictive value across multiple CMD outcomes, highlighting unified risk stratification through a shared metabolic basis. Together, these findings position the RNFL not only as a valid biomarker of CMD risk but also as a metabolically informative tissue that captures subtle systemic perturbations in early CMD pathogenesis.
Since RNFLT metabolic states capture the shared biological basis of early CMDs beyond traditional risk factors, we hypothesize that they hold promise in bridging the extant gaps in cardiometabolic health inequities, as endorsed by the Lancet Commissions34,35. Our findings suggest that RNFLT metabolic states incorporate vital components of CMD risk that traditional approaches overlook, with a significant portion of which were particularly beneficial for women, as women exhibited remarkable improvement in CMD risk prediction, surpassing men and even reversing the prediction gap in certain outcomes. These findings were reinforced through external replication in an ethnically distinct cohort, aligning with prior evidence that highlights the stronger contribution of metabolic factors in women than men43. Notably, the sex-specific improvement cannot be attributed to “regression to the mean,” as women consistently manifested superior improvement to men, irrespective of any prior disadvantages or advantages. Moreover, the disparity is unlikely to result from dataset imbalance, as our results remained robust across various techniques employed to correct for this issue. An extended discussion of benefits for socially vulnerable populations is available in Supplementary Discussion.
Genetic susceptibility exerts substantial risk variability for developing CMDs over one’s lifespan44. To ascertain how RNFLT metabolic states inform CMD risk across different genetic profiles, we generated outcome-specific PRSs from four comprehensive meta-analysed external GWAS datasets32,33,45,46,47,48. Our results demonstrate that RNFLT metabolic states consistently improved CMD prediction across varying genetic susceptibility, a finding that was further confirmed in an ethnically distinct cohort. Notably, this improvement was particularly prominent in low genetic risk populations, whereas those at high genetic risk tended to benefit more from traditional predictors. One possible explanation is that in low genetic risk populations, traditional predictors may contribute modestly to their overall risk, while in high-risk populations, the improvement from RNFLT metabolic states may be overshadowed by the dominance of traditional predictors and the potential overlap of information provided. In contrast, in low-risk populations, RNFLT metabolic states offer more substantial improvement due to the relative absence of these traditional predictors. In addition, the consistent improvement across assays and modelling approaches further supports the robustness of our findings (Supplementary Discussion).
Previous studies have established the protective role of HDL concentration against both RNFL thinning and CMDs16,20,22. Our findings extend this by demonstrating that, even after accounting for enzymatic assay-based HDL concentration, all NMR-based findings remained robust. This suggests that variability in HDL particle properties (size, density, and chemical composition) may be at least as important as HDL concentration in retinal and cardiometabolic physiology. Unlike traditional enzymatic assays that measure concentration following chemical precipitation separation, NMR assays employ deconvolution based on diffusion and relaxation properties, generating composite metrics that integrate both particle properties and lipid concentration. For instance, the higher molecular mobility of cholesterol in small HDL particles extends their T2 relaxation times, resulting in reduced linewidth attenuation in the presaturated proton spectra and thus stronger signals than their larger counterparts. In addition to cholesterol metabolism, findings on alterations in other lipid components, including phospholipids, phosphoglycerides, cholines, and phosphatidylcholines, were also confirmed across assays.
Beyond overlapping findings, the higher sensitivity of LC–MS enabled the detection of low-abundance metabolites that eluded NMR profiling. Phenylalanine and arginine metabolism are among the top pathways enriched. Confirmed in apoA-I knockout mice49, disruptions in phenylalanine metabolism have been linked to reduced apolipoprotein expression50, while arginine modulates endothelial function and supports HDL-mediated anti-inflammation and cholesterol transport51. Glutathione maintains HDL stability by supporting HDL-associated enzymes and the glyoxalase system52. Its depletion has been implicated in HDL structural contraction and particle shrinkage53, which aligns with NMR findings on shifts in HDL subclasses. In addition, enrichment of linoleic acid and vitamin metabolism indicates broader perturbations in membrane lipid composition and coenzyme biosynthesis, both of which are crucial for HDL’s structural integrity and phospholipid exchange. While specific metabolites were not directly matched across platforms, these findings provide complementary biochemical insights that converge on HDL remodelling, positioning RNFL as a downstream reflection of systemic metabolic perturbations in the early stages of CMD pathogenesis.
Several limitations pertain to our findings. First, the metabolic landscape of RNFLT is extensive, thus posing a challenge for any single metabolomic assay to comprehensively profile it. To address this, we integrated a complementary metabolomic assay, broadening our analysis beyond the scope of a solitary platform. Although more extensive metabolite coverage would be expected to further improve the model, our proof-of-concept is platform agnostic. Second, the predominance of European participants in the UKB and Chinese participants in the GDES limits generalizability to other ethnicities, particularly given the limited representation of Black individuals. Further studies in more diverse populations are warranted. Third, the availability of retinal scanning and metabolomic assays was restricted to a subset of UKB participants, thus limiting the sample size for identifying RNFLT metabolic states. Fourth, the limited number of participants longitudinally followed to a mortality outcome within the GDES cohort precluded replication of mortality across demographic groups. Fifth, this study relied on a single timepoint retinal scan and metabolomic assay, constraining our ability to capture dynamic RNFLT fluctuations and corresponding metabolic states over time. Finally, this study remains exploratory, pending further validation and mechanistic enquiry.
In conclusion, our integrated approach provided a framework for deciphering retinal–cardiometabolic connections. Through comprehensive assessments of the shared basis between the eyes and the body, our approach implicated RNFLT metabolic states in the early pathogenesis of multiple CMD outcomes, thereby augmenting risk prediction and clinical utility while bridging the prediction gaps among sexes and socially vulnerable populations. With the booming use of retinal scanning in primary healthcare settings, our findings provide evidence for a scalable, AI-driven oculomic approach that bridges ocular phenotypes with systemic health profiles.
Methods
Study population
The UKB study is a prospective, multicentre cohort study that recruited over 500,000 participants aged 40–69 years from 22 assessment centres across the United Kingdom54. Baseline assessments, including questionnaires, medical interviews, physical examinations and blood tests, were conducted between 2006 and 2010. The GDES is a community-based cohort study that recruited over 2500 patients with diabetes aged 35–85 in Guangzhou, China30. Baseline assessments were conducted from 2017 to 2019. The study was conducted in accordance with the principles of the Declaration of Helsinki and approved by the Northwest Multicenter Research Ethics Committee (11/NW/0382) and the Ethics Committee of Zhongshan Ophthalmic Center (2017KYPJ094). Prior to any engagement, written informed consent was obtained from all participants. The study adhered to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) and Standards for Reporting of Diagnostic Accuracy Studies (STARD) guidelines for reporting.
The UKB participants were categorized into three non-overlapping groups: (1) population-I comprised individuals who underwent both OCT scanning and metabolomic profiling (n = 7824); (2) population-II underwent metabolomic profiling only, without OCT scanning (n = 86,014); and (3) the remaining individuals were excluded from the analysis15. Participants with prevalent glaucoma, retinal disorders (including retinal detachment and break, retinal vascular occlusion, and others), and neurodegenerative diseases were further excluded from population-I because of the potential for secondary retinal destruction. Further information on the specific eligibility criteria for each population group can be found in the Supplementary Methods. The UKB population-I was used to identify RNFLT metabolic states, while population-II was used for downstream analyses. Replication in an independent cohort was conducted in a subset of the population that underwent both eligible OCT scanning and metabolomic profiling in the GDES cohort (population-III).
Metabolomic profiling
High-throughput proton NMR (1H-NMR) (Nightingale Health, Finland) was employed to quantify the metabolite concentrations from the plasma samples of the participants29,55,56,57. Cryopreserved EDTA plasma samples were thawed and subjected to centrifugation to separate the supernatant. This supernatant was then mixed with phosphate buffer to create the sample mixture. The prepared samples were loaded onto a specialized cooled sample changer. Two distinct NMR spectra were recorded for each plasma sample using a high-resolution 500 MHz NMR spectrometer (Bruker AVANCE IIIHD, USA). One of these spectra characterized resonances originating from proteins and lipid lipoprotein particles, while the other spectrum detected resonances from low-molecular-weight metabolites. Following quality control processes, metabolic biomarkers were quantified using the Nightingale Health Biomarker Quantification Library 2020. Detailed information can be found in the Supplementary Methods.
OCT imaging
Spectral-domain OCT was performed in an enclosed darkroom with a Topcon 3D OCT-1000 Mk II system (Topcon, Japan)58,59,60. The system had an axial resolution of 6 μm and an image acquisition rate of 18,000 A-scans/s. The imaging was performed using a 3D 6 × 6 mm macular volume scan mode centred at the fovea, with a scan density of 512 A-scans × 128 B-scans within 3.7 s. The Topcon Advanced Boundary Segmentation algorithm automatically segmented the retinal layers and determined the average RNFLT as the distance from the internal limiting membrane to the nerve fibre layer-ganglion cell layer junction61. Image quality scores, internal limiting membrane indicators, validity counts, and motion indicators were recorded for quality control (Supplementary Methods). Images with low signal strength (<45) or poor segmentation or centration (the worst 20% for each metric) were excluded from analysis. In cases in which both eyes of a participant met the eligibility criteria, the right eyes were selected for further analysis.
Outcomes and covariates
The Hospital Episode Statistics database, Scottish Morbidity Record and Patient Episode Database were used to record inpatient hospital records for England, Scotland, and Wales. Mortality data were obtained from national datasets with the National Health System (NHS) Digital and the NHS Central Register. The assessment of CMD outcomes and the recording of the primary causes of death were based on the International Classification of Diseases-10. The follow-up period was from 16 March 2006 to 31 March 2021. Person days for each participant were calculated from the date of baseline assessment to the date of disease onset, death, or the end of follow-up, whichever came first. Details of the covariate assessment are available in Supplementary Methods.
Genotyping and outcome-specific PRS
The PRSs for both T2D and CVD were computed to identify genetically high-risk individuals. Genotyping was conducted using the UK BiLEVE Axiom Array or the UKB Axiom Array. Quality control processes and imputation were conducted based on the Haplotype Reference Consortium and UK10K haplotype resources54. External GWAS summary statistic data were meta-analysed via fixed-effect inverse variance meta-analysis, with a correction for sample overlap where required62. The PRS algorithms were constructed based on outcome-specific meta-analyses using a Bayesian approach. Per-individual PRSs were computed as the genome-wide summation of per-variant posterior effect sizes multiplied by allele dosage. After generating raw PRSs using the derived weights, centring and standardization were applied to produce corrected PRSs (Supplementary Methods).
RNFLT-associated metabolites and their linkage to CMD outcomes
The associations of metabolic biomarkers with RNFLT were assessed using multilevel linear regression models after adjusting for age, sex, ethnicity, assessment centre, household income, Townsend deprivation index, education, smoking, alcohol consumption, body mass index, use of lipid-lowering medications, enzymatic assay-based HDL cholesterol, systolic blood pressure, and HbA1c level. Sensitivity analyses were performed by further adjusting for prevalent CMDs. The Benjamini–Hochberg method was employed to reduce the false-positive rate. Biomarkers that reached statistical significance after multiple testing were identified as RNFLT metabolic states.
The RNFLT-associated metabolites were further analysed using Cox proportional hazards models for the risk of six CMD outcomes, accounting for the same covariates while accounting for both time-event data and censored observations63. Participants with a baseline CMD diagnosis were excluded from the corresponding analysis (e.g., in the case of the T2D endpoint, participants with baseline T2D were excluded). A mediation analysis was conducted to estimate the mediation effect of traditional risk factors and RNFLT-associated metabolites on the association between RNFLT and CMD risk. We assessed it as the percentage change in adjusted Cox proportional hazards coefficients for each CMD outcome after adjusting for the metabolites of interest. The proportional hazard assumption was tested using the Schoenfeld residual method and was satisfied for each model.
Model development and evaluation
We employed Random Forest and several other traditional machine learning (extreme gradient boosting, support vector machine, least absolute shrinkage and selection operator, and ridge) and deep learning (multilayer perception and convolutional neural network) approaches to model CMD outcome risk. The datasets were randomly split into training and testing sets at an 8:2 ratio. In the case of deep learning modelling, an additional subset of the training data (20%) was set aside for validation, during which loss reduction monitoring was performed to ensure optimal model performance. Hyperparameters were fine-tuned through a grid search for optimal model configurations64. Detailed modelling procedures can be found in the Supplementary Methods.
Model evaluation was performed in the fully withheld testing set. Participants were stratified based on the predicted RNFLT metabolic states, which were derived from the respective RNFLT metabolic models for each CMD outcome, and cumulative event rates were compared across the top, middle and bottom 10% of states for each CMD outcome15,29. The AUPRC and Harrell’s C-statistic were calculated to assess the predictive value of the RNFLT metabolic states for each CMD outcome and were also benchmarked against individual conventional predictors. While the C-statistic provides an overall measure of discrimination, it assigns equal weight to false positives and false negatives and therefore distorts performance in imbalanced datasets. Given that the positive-to-negative ratios across CMD outcomes ranged from 1:99 to 7:93 (Table S10), AUPRC was prioritized to evaluate absolute and incremental performance for each individual outcome, as it provides an unbiased assessment in imbalanced settings. Notably, the baseline (no-skill) AUPRC varies by outcome and corresponds to the event rates in the dataset: 0.06 for T2D, 0.03 for myocardial infarction, 0.03 for heart failure, 0.02 for stroke, 0.07 for all-cause mortality, and 0.02 for CMD mortality. As such, interpretation of AUPRC values should be made relative to these outcome-specific baselines rather than on absolute magnitude alone. C-statistics were additionally reported as a complementary metric to facilitate intuitive interpretability and cross-outcome comparison with differing event rates.
We evaluated the incremental predictability of these states compared to the Age&Sex model and a spectrum of conventional algorithms for CMD prediction (FGCRS65, SCORE266, AHA/ASCVD67, WHO-CVD68, FRS for T2D69, NZ-DCS40, UKPDS41 and Wan’s model42), where applicable. Calibration plots were built to assess the goodness of model fit, and decision curve analyses were performed to estimate net benefits in clinical utility. The reclassification improvement was calculated by adding the percentage of patients with an event correctly reclassified to the percentage of patients without an event correctly reclassified, at a risk category of <5%, 5–10%, and >10%. Further subgroup analyses were conducted on different genetic susceptibility, sexes, social classes, and educational attainment.
Extrapolation in an independent cohort
At baseline, all GDES participants underwent retinal OCT scanning (SS-OCT, DRI OCT Triton; Topcon, Japan) using a 3D Macula Cube 7 × 7 mm scan mode centred on the fovea15,30. A built-in software (IMAGEnet 6, v1.22) was employed for automatic retinal segmentation (Supplementary Methods). Furthermore, a subset of the GDES participants underwent liquid chromatography tandem triple quadrupole mass spectrometry (LC, ExionLC AD, SCIEX, USA; MS, QTRAP System, SCIEX, USA) for metabolomic profiling with stringent quality control processes employed (Supplementary Methods)70,71. Incident CVD was defined as the development of coronary heart disease, heart failure, atrial fibrillation, stroke or related mortality during the follow-up period, determined by a combination of medical records, questionnaires, and verbal interviews. Fasting blood samples were collected to assess the serum concentration of HbA1c, total cholesterol, and HDL cholesterol (enzymatic), and to assess the glomerular filtration rate. The modelling details and analytical processes are as above.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used in this study are available from UKB via data access procedures (http://www.ukbiobank.ac.uk). Permission to use the UKB Resource was obtained via a material transfer agreement as part of Application 105658. Raw data from the GDES are not publicly available due to data privacy laws and were used for the purposes of this project with institutional permission from the Zhongshan Ophthalmic Center. All requests for access to in-house data should be addressed to the corresponding authors, Prof. Lingyi Liang (Email: lianglingyi@gzzoc.com) & Dr. Wei Wang (email: wangwei@gzzoc.com), and will be processed in accordance with Zhongshan Ophthalmic Center policies. The GDES group will evaluate all requests based on the purpose of the data request, and it may take ~90 days to process the request. A material-transfer and data-usage agreement will be required between Zhongshan Ophthalmic Center and the receiving organization, and the requesting organization must state the intended purpose of the data transfer and provide assurances that the transferred data will only be used for non-commercial academic and educational purposes in compliance with Zhongshan Ophthalmic Center institutional policies. Source data are provided with this paper.
Code availability
All software used in this study is publicly available. The code used in this study can be accessed at https://github.com/zocskl/RNFLTmetabolic-states-predict-CMD-outcomes72.
References
Flammer, J. et al. The eye and the heart. Eur. Heart J. 34, 1270–1278 (2013).
Cheung, C. Y., Ikram, M. K., Chen, C. & Wong, T. Y. Imaging retina to study dementia and stroke. Prog. Retin. Eye Res. 57, 89–107 (2017).
Farrah, T. E., Webb, D. J. & Dhaun, N. Retinal fingerprints for precision profiling of cardiovascular risk. Nat. Rev. Cardiol. 16, 379–381 (2019).
Aumann, S., Donner, S., Fischer, J. & Muller, F. Optical coherence tomography (OCT): principle and technical realization. In High Resolution Imaging in Microscopy and Ophthalmology 59–85 (ed. Bille, J.) (Springer, Cham, 2019).
Fujimoto, J. G. Optical coherence tomography for ultrahigh resolution in vivo imaging. Nat. Biotechnol. 21, 1361–1367 (2003).
Chen, Y. et al. Retinal nerve fiber layer thinning as a novel fingerprint for cardiovascular events: results from the prospective cohorts in UK and China. BMC Med. 21, 24 (2023).
Wang, D. et al. Localized retinal nerve fiber layer defects and stroke. Stroke 45, 1651–1656 (2014).
Chong, R. S. et al. Association of antihypertensive medication with retinal nerve fiber layer and ganglion cell-inner plexiform layer thickness. Ophthalmology 128, 393–400 (2021).
Majithia, S. et al. Retinal nerve fiber layer thickness and rim area profiles in Asians: pooled analysis from the Asian Eye Epidemiology Consortium. Ophthalmology 129, 552–561 (2022).
Lim, H. B., Shin, Y. I., Lee, M. W., Park, G. S. & Kim, J. Y. Longitudinal changes in the peripapillary retinal nerve fiber layer thickness of patients with type 2 diabetes. JAMA Ophthalmol. 137, 1125–1132 (2019).
Lee, M. W. et al. Effect of systemic hypertension on peripapillary RNFL thickness in patients with diabetes without diabetic retinopathy. Diabetes 70, 2663–2667 (2021).
Aumann, S., Donner, S., Fischer, J. & Müller, F. Optical coherence tomography (OCT): principle and technical realization. In High Resolution Imaging in Microscopy and Ophthalmology: New Frontiers in Biomedical Optics (ed. Bille, J. F.) 59–85 (Springer, 2019).
Fujimoto, J. & Swanson, E. The development, commercialization, and impact of optical coherence tomography. Investig. Ophthalmol. Vis. Sci. 57, OCT1–OCT13 (2016).
Wagner, S. K. et al. Insights into systemic disease through retinal imaging-based oculomics. Transl. Vis. Sci. Technol. 9, 6 (2020).
Yang, S. et al. Analysis of plasma metabolic profile on ganglion cell-inner plexiform layer thickness with mortality and common diseases. JAMA Netw. Open 6, e2313220 (2023).
Di Angelantonio, E. et al. Major lipids, apolipoproteins, and risk of vascular disease. JAMA 302, 1993–2000 (2009).
Pradhan, A. D. A new beginning for triglyceride-lowering therapies. Circulation 140, 167–169 (2019).
Khera, A. V. et al. Cholesterol efflux capacity, high-density lipoprotein function, and atherosclerosis. N. Engl. J. Med. 364, 127–135 (2011).
Sun, L. et al. Causal associations of blood lipids with risk of ischemic stroke and intracerebral hemorrhage in Chinese adults. Nat. Med. 25, 569–574 (2019).
Krauss, R. M. Lipids and lipoproteins in patients with type 2 diabetes. Diabetes Care 27, 1496–1504 (2004).
Schrier, R. W. Role of diminished renal function in cardiovascular mortality. J. Am. Coll. Cardiol. 47, 1–8 (2006).
Ho, H. et al. Retinal nerve fiber layer thickness in a multiethnic normal Asian population: the Singapore Epidemiology of Eye Diseases study. Ophthalmology 126, 702–711 (2019).
Rauscher, F. G. et al. Renal function and lipid metabolism are major predictors of circumpapillary retinal nerve fiber layer thickness-the LIFE-Adult Study. BMC Med. 19, 202 (2021).
Kardys, A. et al. Cholesterol affects retinal nerve fiber layer thickness in patients with multiple sclerosis with optic neuritis. Eur. J. Neurol. 20, 1264–1271 (2013).
Srivastav, K., Saxena, S., Mahdi, A. A., Kruzliak, P. & Khanna, V. K. Increased serum urea and creatinine levels correlate with decreased retinal nerve fibre layer thickness in diabetic retinopathy. Biomarkers 20, 470–473 (2015).
Karti, O., Nalbantoglu, O., Abali, S., Tunc, S. & Ozkan, B. The assessment of peripapillary retinal nerve fiber layer and macular ganglion cell layer changes in obese children: a cross-sectional study using optical coherence tomography. Int. Ophthalmol. 37, 1031–1038 (2017).
Ozen, B., Ozturk, H., Catli, G. & Dundar, B. An assessment of retinal nerve fiber layer thickness in non-diabetic obese children and adolescents. J. Clin. Res. Pediatr. Endocrinol. 10, 13–18 (2018).
Pietzner, M. et al. Plasma metabolites to profile pathways in noncommunicable disease multimorbidity. Nat. Med. 27, 471–479 (2021).
Buergel, T. et al. Metabolomic profiles predict individual multidisease outcomes. Nat. Med. 28, 2309–2320 (2022).
Yang, S. et al. Metabolic fingerprinting on retinal pigment epithelium thickness for individualized risk stratification of type 2 diabetes mellitus. Nat. Commun. 14, 6573 (2023).
Liu, R. et al. Metabolomic signature of retinal ageing, polygenetic susceptibility, and major health outcomes. Br. J. Ophthalmol. 109, 619–627 (2025).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Lavange, L. M. et al. Sample design and cohort selection in the Hispanic Community Health Study/Study of Latinos. Ann. Epidemiol. 20, 642–649 (2010).
The, L. Cardiology’s problem women. Lancet 393, 959 (2019).
Gerdts, E. & Regitz-Zagrosek, V. Sex differences in cardiometabolic disorders. Nat. Med. 25, 1657–1666 (2019).
O’Hearn, M., Lauren, B. N., Wong, J. B., Kim, D. D. & Mozaffarian, D. Trends and disparities in cardiometabolic health among U.S. adults, 1999–2018. J. Am. Coll. Cardiol. 80, 138–151 (2022).
Schultz, W. M. et al. Socioeconomic status and cardiovascular outcomes: challenges and interventions. Circulation 137, 2166–2178 (2018).
Kubát, M. & Matwin, S. Addressing the curse of imbalanced training sets: one-sided selection. In Proc. of the 14th International Conference on Machine Learning 179–186 (1997).
Zhang, S. et al. Design and baseline data of the diabetes registration Study: Guangzhou Diabetic Eye Study. Curr. Eye Res. 48, 591–599 (2023).
Elley, C. R., Robinson, E., Kenealy, T., Bramley, D. & Drury, P. L. Derivation and validation of a new cardiovascular risk score for people with type 2 diabetes: the new zealand diabetes cohort study. Diabetes Care 33, 1347–1352 (2010).
Stevens, R. J., Kothari, V., Adler, A. I. & Stratton, I. M. The UKPDS risk engine: a model for the risk of coronary heart disease in Type II diabetes (UKPDS 56). Clin. Sci. (Lond.) 101, 671–679 (2001).
Wan, E. et al. Development of a cardiovascular diseases risk prediction model and tools for Chinese patients with type 2 diabetes mellitus: a population-based retrospective cohort study. Diabetes Obes. Metab. 20, 309–318 (2018).
Paynter, N. P. et al. Metabolic predictors of incident coronary heart disease in women. Circulation 137, 841–853 (2018).
O’Sullivan, J. W. et al. Polygenic risk scores for cardiovascular disease: a scientific statement from the American Heart Association. Circulation 146, e93–e118 (2022).
Kolonel, L. N. et al. A multiethnic cohort in Hawaii and Los Angeles: baseline characteristics. Am. J. Epidemiol. 151, 346–357 (2000).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Bild, D. E. et al. Multi-ethnic study of atherosclerosis: objectives and design. Am. J. Epidemiol. 156, 871–881 (2002).
Olson, J. L., Bild, D. E., Kronmal, R. A. & Burke, G. L. Legacy of MESA. Glob. Heart 11, 269–274 (2016).
Liu, J. et al. LC–MS-based metabolomics and lipidomics study of high-density-lipoprotein-modulated glucose metabolism with an apoA-I knockout mouse model. J. Proteome Res. 18, 48–56 (2019).
Williams, R. A., Hooper, A. J., Bell, D. A., Mamotte, C. D. & Burnett, J. R. Plasma cholesterol in adults with phenylketonuria. Pathology 47, 134–137 (2015).
Riddell, D. R., Graham, A. & Owen, J. S. Apolipoprotein E inhibits platelet aggregation through the L-arginine:nitric oxide pathway. Implications for vascular disease. J. Biol. Chem. 272, 89–95 (1997).
Rosenblat, M., Volkova, N., Coleman, R. & Aviram, M. Anti-oxidant and anti-atherogenic properties of liposomal glutathione: studies in vitro, and in the atherosclerotic apolipoprotein E-deficient mice. Atherosclerosis 195, e61–e68 (2007).
Godfrey, L., Yamada-Fowler, N., Smith, J., Thornalley, P. J. & Rabbani, N. Arginine-directed glycation and decreased HDL plasma concentration and functionality. Nutr. Diabetes 4, e134 (2014).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Xu, Y. et al. An atlas of genetic scores to predict multi-omic traits. Nature 616, 123–131 (2023).
Julkunen, H. et al. Atlas of plasma NMR biomarkers for health and disease in 118,461 individuals from the UK Biobank. Nat. Commun. 14, 604 (2023).
Elliott, P. & Peakman, T. C. The UK Biobank sample handling and storage protocol for the collection, processing and archiving of human blood and urine. Int. J. Epidemiol. 37, 234–244 (2008).
Ko, F. et al. Association of retinal nerve fiber layer thinning with current and future cognitive decline: a study using optical coherence tomography. JAMA Neurol. 75, 1198–1205 (2018).
Ko, F. et al. Associations with retinal pigment epithelium thickness measures in a large cohort: results from the UK Biobank. Ophthalmology 124, 105–117 (2017).
Patel, P. J. et al. Spectral-domain optical coherence tomography imaging in 67 321 adults: associations with macular thickness in the UK Biobank study. Ophthalmology 123, 829–840 (2016).
Yang, Q. et al. Automated layer segmentation of macular OCT images using dual-scale gradient information. Opt. Express 18, 21293–21307 (2010).
Thompson, D. J. et al. A systematic evaluation of the performance and properties of the UK Biobank Polygenic Risk Score (PRS) Release. PLoS One 19, e0307270 (2024).
Cox, D. R. Regression models and life-tables. J. R. Stat. Soc.: Ser. B (Methodol.) 34, 187–202 (1972).
Feurer, M. & Hutter, F. Hyperparameter optimization. In Automated Machine Learning: Methods, Systems, Challenges (eds Hutter, F., Kotthoff, L. & Vanschoren, J.) 3–33 (Springer International Publishing, 2019).
D’Agostino, R. S. et al. General cardiovascular risk profile for use in primary care: the Framingham Heart Study. Circulation 117, 743–753 (2008).
SCORE2 risk prediction algorithms: new models to estimate 10-year risk of cardiovascular disease in Europe. Eur. Heart J. 42, 2439–2454 (2021).
Goff, D. J. et al. 2013 ACC/AHA guideline on the assessment of cardiovascular risk: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J. Am. Coll. Cardiol. 63, 2935–2959 (2014).
World Health Organization cardiovascular disease risk charts: revised models to estimate risk in 21 global regions. Lancet Glob. Health 7, e1332–e1345 (2019).
Bragg, F. et al. Predictive value of circulating NMR metabolic biomarkers for type 2 diabetes risk in the UK Biobank study. BMC Med. 20, 159 (2022).
Chen, W. et al. A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: application in the study of rice metabolomics. Mol. Plant. 6, 1769–1780 (2013).
Fraga, C. G., Clowers, B. H., Moore, R. J. & Zink, E. M. Signature-discovery approach for sample matching of a nerve-agent precursor using liquid chromatography–mass spectrometry, XCMS, and chemometrics. Anal. Chem. 82, 4165–4173 (2010).
zocskl. zocskl/RNFLT-metabolic-states-predict-CMD-outcomes: RNFL-metabolome and eye–heart connections (v1.0.0). Zenodo https://doi.org/10.5281/zenodo.17333465 (2025).
Acknowledgements
This study was funded by the Guangdong Basic Research Center of Excellence (GBRCE) for Major Blinding Eye Diseases Prevention and Treatment (2024-PIZC-002, 2024-RCPY-008, W.W.), the Hainan Province Clinical Medical Center, the National Natural Science Foundation of China (82371086, W.W.; 82171084, W.H.), the Natural Science Foundation of Guangdong Province (2023A1515011475, W.W.), and the Science and Technology Projects in Guangzhou (SL2024A03J00472; 2025A04J7150,2024A03J0253, W.W.), and the ZOC’s supporting funds (2025QZSPT02, 2025-PT0015, 2025-PT1069, W.W.). The authors thank all participants and staff involved in the UKB and GDES cohorts.
Author information
Authors and Affiliations
Contributions
Study concept and design: W.W., L.L. and C.Y.C.; Acquisition, analyses, or interpretation: S.Y., Z.X., H.L., Z.Z., LZ.Z. and X.S.; Drafting of the manuscript: S.Y., W.W. and Z.X.; Critical revision of the manuscript for important intellectual content: S.Y., Z.X., M.H., J.B.J., N.C. and C.Y.C.; Statistical analyses: W.W., S.Y. and Z.X.; Obtained funding: W.W., W.H.; Administrative, technical, or material support: L.L., S.Y., L.Z. and W.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, S., Xin, Z., Li, H. et al. Artificial intelligence-driven metabolomics of retinal nerve fibre layer to profile risks of mortality and cardiometabolic diseases. Nat Commun 16, 11039 (2025). https://doi.org/10.1038/s41467-025-66979-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66979-z