Abstract
IgG antibodies have a conserved N-linked glycan on residue Asn297 of the homodimeric Fc region that modulates antibody-mediated effector functions. These effects on the immune response are reflected in correlations between IgG glycans and numerous infectious and autoimmune diseases. However, current studies fail to characterize these Asn297-linked glycans comprehensively, relying on glycan release methods. Here, we develop an intact LC/MS method to glycoprofile polyclonal IgG antibodies while preserving the Fc glycan spatial pairing. We analyze plasma samples from healthy and virally infected individuals and find that all individuals have asymmetrically glycosylated IgGs—the glycans on each of the Fc protomers are not identical. We find that the previously observed association between IgG afucosylation and severe dengue disease is due to asymmetric monofucosylation of IgGs, not symmetric afucosylation. Finally, we engineer monofucosylated IgG1s which are indistinguishable from afucosylated IgG1s in binding to FcγRIIIA in vitro and inducing effector functions in vivo.
Similar content being viewed by others
Introduction
N-linked glycosylation is an essential post-translational modification present on numerous human proteins, in which glycans are attached to the nitrogen atom of an asparagine (Asn) residue when the consensus sequence motif NXT/S—where X is any amino acid except a proline—is present1. These glycans play an important role in the intracellular and extracellular functions of the proteins that they modify. N-glycosylation has been shown to be linked to congenital diseases, cancer, immunity, and inflammation2.
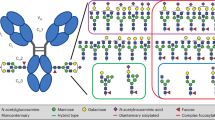
IgG antibodies have a conserved, but highly heterogeneous, N-linked glycan attached to Asn297 on the Fc region. Most commonly, this glycan is a complex biantennary glycan composed of a core structure of two β-linked N-acetylglucosamines (GlcNAcs), one β−1,4-linked mannose, and two α-linked mannoses that initiate the distinct α−1,3 and α−1,6 branches for further elaboration. The core structure can be further modified with additional sugars such as an α−1,6-linked fucose on the innermost GlcNAc and a bisecting GlcNAc. A β−1,4 galactose can be added to each of the α−1,3 and α−1,6 branches, and α−2,6 sialic acid can be further added when these galactose saccharides are present3.
The Asn297-linked glycans on IgG antibodies play an important role in regulating antibody-mediated effector functions such as antibody-dependent cellular cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP), and complement activation, mainly by modifying affinity to Fc γ receptors (FcγRs) and/or complement C1q4,5. For example, the absence of core fucose leads to increased affinity of FcγRIIIA, promoting ADCC6,7. On the other hand, the presence of sialic acid has each been shown to have an anti-inflammatory effect8,9, as it enables binding of the Fc domain to the type II FcR, DC-SIGN. Such interactions result in the upregulation of the inhibitory FcγRIIb, increasing the threshold of activation for the cells that express it10, including macrophages and DCs. Similarly, sialic acid binding to CD23 leads to an increase in FcγRIIb expression on the surface of B cells, thereby modulating B-cell selection11. Although galactose has been shown to increase C1q binding12, in vivo models suggest that this increases the threshold of complement activation, ultimately leading to anti-inflammatory properties9,13. Some studies also show that galactose increases activation of FcγRIIb14.
Due to the effect of the Asn297-linked glycans on antibody-mediated effector functions, the glycosylation of IgG Fcs has been intensively studied and shown to correlate with the severity of a multitude of autoimmune and infectious diseases, leading to a role in diagnosis and therapeutics5,15,16,17,18,19. In rheumatoid arthritis, a decrease in IgG galactosylation has been shown to directly precede disease onset20. Intravenous immunoglobulin (IVIG) is used as a therapeutic for autoimmune diseases to dampen inflammation21; an effect driven by the sialylated Fc glycoforms present in IVIG. In COVID, agalactosylated glycans with or without core fucose (G0F and G0) are upregulated in hospitalized patients, while vaccinated patients and those with mild symptoms display increased levels of galactosylation and sialylation22,23. In dengue, it is well established that afucosylation is a marker of severe disease24. Additionally, advances in glycoengineering of IgGs have enabled the engineering and approval of afucosylated monoclonal antibodies that have enhanced function as cancer therapeutics25.
However, the vast majority of glycoprofiling studies have been performed using glycan release workflows26. In these methods, enzymes such as PNGase F are used to cleave the glycans from IgGs, after which the glycans are labeled, and then detected and quantified using mass spectrometry27,28,29. While these methods are highly sensitive and provide information about the type and structure of glycans present in the sample, they miss an important aspect of IgG biology—IgG antibodies are homodimers with one Asn297-linked glycan on each of the two Fc protomers and these glycans can be asymmetric, i.e., the glycan on one of the Fc chains need not be the same as the glycan on the other. Intact LC/MS analysis of recombinantly expressed mAbs reveals that asymmetrically glycosylated IgG exists30. Additionally, a portion of human IgG antibodies from cord blood have been shown to be asymmetrically glycosylated using nano-ESI/MS31.
Structural studies show that IgG Fcs interact with Fc receptors asymmetrically, in which the contacts made with one Fc protomer are different from those made with the other protomer32, providing a basis for how asymmetrical glycosylation of IgG antibodies may translate to the induction of specific antibody-mediated effector functions. As a result, this raises the question of which glycan determines the affinity of the interaction in the case of asymmetrically glycosylated IgGs. We sought to investigate the relationship between the pairing of antibody glycosylation and Fc effector functions, focusing on core fucosylation, which is one of the most well-characterized determinants of affinity to FcγRIIIA. To do so, we developed an intact LC/MS method, Whole Immunoglobulin Glycoprofiling With Asymmetrical Monitoring (WIgGWAM), which enabled the glycoprofiling of polyclonal IgG1 antibodies while preserving the spatial pairing of the glycans. We used this method to confirm that peripheral human IgG1s, whether derived from healthy individuals or patients with COVID-19 exist in asymmetrically glycosylated forms. We also glycoprofiled IgG1 antibodies from the plasma of dengue patients and showed that core monofucosylation exists in the human body and is associated with severe and secondary dengue infections, whereas fully core-afucosylated IgG glycoforms are not. Finally, to investigate the mechanism of the correlation, we developed a strategy for engineering recombinant monofucosylated IgG1s and showed that they bind to FcγRIIIA in a high-affinity orientation in vitro and induce antibody-mediated effector functions in vivo as do fully afucosylated IgGs. Our studies highlight a layer of complexity regarding Fc glycan regulation of effector functions and demonstrate how asymmetrical glycosylation of IgG antibodies plays a role in human disease.
Results
WIgGWAM enables intact LC/MS glycoprofiling of polyclonal human IgG1 antibodies while preserving the spatial pairing of the Asn297-linked glycans
To glycoprofile polyclonal IgG antibodies while preserving the spatial pairing of the Asn297-linked glycans, we developed the intact LC/MS-based Whole Immunoglobulin Glycoprofiling With Asymmetric Monitoring (WIgGWAM) method. We initially incubate a polyclonal IgG sample, such as human plasma or serum, with papain, a protease that cleaves IgG antibodies in the upper hinge region, separating the Fabs from the Fcs33. We then selectively purify the Fcs, which contain the relevant Asn297-linked glycans, using protein A. Finally, we analyze the Fcs using intact mass spectrometry in which the Fc is denatured but not reduced, maintaining its homodimeric form30 (Fig. 1a). By removing the polyclonal Fabs from the sample, we reduce the mass heterogeneity to the glycoforms and the IgG Fc alleles present, and we can assign deconvoluted intact LC/MS spectra to specific glycan pairings (Fig. 1b). It is important to note that this list of glycoforms is not comprehensive. Indeed, with over 32 identified glycoforms, there are more than 1000 theoretical possibilities for IgG glycan pairings34. However, we have restricted our analysis to the 20 depicted in Fig. 1b based on the most common glycoforms that result from recombinant expression of human IgG1 antibodies in mammalian cell lines, as well as glycoforms that are most commonly identified in glycan release studies35,36.
a Schematic describing WIgGWAM workflow: IgG sample is incubated with papain at 37 °C for 30 min. Then, the cleaved Fcs are purified using protein A and analyzed with intact LC/MS with or without the addition of glycosidases. b Depictions of the 20 glycan pairs used for labeling the deconvoluted intact LC/MS spectra during data analysis of WIgGWAM. Glycoforms in blue text are asymmetric, in purple are symmetric, and in orange are ambiguous. Created in BioRender. D, J. (2025) https://BioRender.com/efko4f3.
To characterize the IgG1 Fcs after the WIgGWAM workflow, we treat a portion of the sample with EndoS2, an endoglycosidase that is highly specific for the Asn297-linked glycan on IgG Fcs and cleaves between the first and second GlcNAc37. The EndoS2-treated Fc contains 1 GlcNAc on each protomer and a fucose, if present. The removal of the heterogeneous glycans further simplifies the mass spectrum, allowing us to determine: (1) the IgG1 allele(s) present in the sample38,39,40; and (2) the fucosylation levels (Fig. 2a, b). While the less abundant monofucosylated and afucosylated Fcs are difficult to identify in a fully glycosylated mass spectrum, they are readily apparent in the EndoS2-treated mass spectrum. Additionally, the EndoS2-treated sample provides a reference mass from which we can calculate the masses of glycan pairs and assign them to specific peaks while allowing for a ±5 Da variability, consistent with other intact LC/MS analyses41 (Figs. 1b and 2c). The assigned peaks then go through a filtering step in which any peak less than 20% of the maximum peak height is excluded as noise (Fig. 2a, c, d); this conservative approach to assigning peaks in these mass spectra ensures that the corresponding IgG glycoforms are present in our sample.
a After WIgGWAM, a portion of the purified Fcs are treated with EndoS2 to determine fucosylation levels, the other portion of Fcs is analyzed with or without exoglycosidases and filtered according to a 20% of maximum peak height cutoff. Created in BioRender. D, J. (2025) https://BioRender.com/lybz9tn. b–f Deconvoluted intact LC/MS spectrum of Fcs purified with WIgGWAM from a healthy individual human plasma sample b treated with EndoS2. The dotted lines annotate the masses that correspond to the bifucosylated, monofucosylated, and afucosylated mass of the nG1m1 Fc allele c with no enzymes added. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of maximum peak height d with no enzymes added. The dotted lines annotate Fc-linked glycoforms whose peak heights were less than 20% of maximum peak height e treated with an exosialidase. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of maximum peak height f treated with an exogalactosidase. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak height. For all intact LC/MS deconvoluted spectra, the highest peak was normalized to 100. Source data are provided as a Source Data file.
Figure 2 depicts the analysis of a plasma sample from a healthy individual. After WIgGWAM, we treated a portion of the sample with EndoS2, enabling us to identify that this individual is a homozygote with the nG1m1 Fc allele (Fig. 2b). We performed intact LC/MS analysis on another portion of the sample, without any enzyme treatment. Using the filtering method described above, we assigned 9 glycoforms present in this sample: [1] G0F/G0F-GlcNAc, [2] G0F/G1F-GlcNAc, [3] G0F/G0F, [4] G0F/G1F, [5] G0F/G0F + bGlcNAc, [6] G2F/G0F or G1F/G1F, [7] G0F/G1F + bGlcNAc, [10] G1F/G2F, and [11] G1SF/G1F or G0F/G2SF (Figs. 1b and 2c, d). We confirmed the presence of these glycoforms by treating the sample with exoglycosidases. When we treated the sample with an exosialidase42, we identified only the non-sialylated glycoforms [1],[2],[3],[4],[5],[6],[7] and [10]. The sialylated glycoform [11] G1SF/G1F or G0F/G2SF was no longer detected (Figs. 1b and 2e). When we treated the sample with an exogalactosidase43, only agalactosylated glycoforms [1],[3], and [5] remained (Figs. 1b and 2f). Exoglycosidase treatment also provided a benchmark to confirm our chosen cutoff of 20% of maximum peak height, as there should be no sialylated glycoforms and no glycoforms with exposed galactoses, when treated with the respective enzyme. If we use a 5%, 10%, or 15% cutoff, peaks [9], [11], [13], [14], [15], [17], [18], [19], and/or [20] that contain sialic acids (Fig. 1b, and Supplementary Fig. 1A–C) or peaks [2], [4], [6], [7], and/or [10] that contain exposed galactoses (Fig. 1b, and Supplementary Fig. 1D–F) pass the threshold, even though such glycoforms are unlikely to be present in the sample. It is only when we use 20% as a cutoff that the analysis results in identifying peaks without sialic acids (Figs. 1b and 2e) or exposed galactoses (Figs. 1b and 2f).
WIgGWAM is an intact LC/MS-based method for glycoprofiling polyclonal IgG1 antibodies while preserving the spatial pairing of IgG glycoforms. The workflow provides information about the alleles and paired glycans present in the sample. We use a constant preset filtering method and exoglycosidase treatment to maximize the consistency and reproducibility of the method.
WIgGWAM identifies both symmetric and asymmetric IgG glycoforms
To ensure that the WIgGWAM workflow does not significantly alter the glycoforms present in the sample, we applied our method to Rituximab, a recombinant IgG144. Because it is a monoclonal antibody, Rituximab can be analyzed using intact LC/MS as a full-length IgG, allowing us to glycoprofile it before and after WIgGWAM. We observed six glycoforms of Rituximab IgG1 produced in Expi293F cells: [3] G0F/G0F, [4] G0F/G1F, [5] G0F/G0F+bGlcNAc, [6] G2F/G0F or G1F/G1F, [7] G0F/G1F+bGlcNAC, and [10] G1F/G2F when applying our 20% of maximum peak height cutoff (Figs. 1b, 2a, 3a, and Supplementary Fig. 2A, B). There are peaks that correspond to [1] G0F/G0F-GlcNAc and [2] G0F/G1F-GlcNAc; however, they do not pass the cutoff. After WIgGWAM, the Rituximab Fc consists of six glycoforms: [1] G0F/G0F-GlcNAc, [2] G0F/G1F-GlcNAc, [3] G0F/G0F, [4] G0F/G1F, [6] G2F/G0F or G1F/G1F, and [10] G1F/G2F. While there are peaks that correspond to [5] G0F/G0F+bGlcNAc and [7] G0F/G1F+bGlcNAc identified in the full-length IgG, they do not pass the cutoff (Figs. 1b and 3b, and Supplementary Fig. 2C, D). EndoS2 treatment of the Fc showed that the allele present is G1m1, which is consistent with the sequence of the plasmid we used to express Rituximab in Expi293F cells (Fig. 3c, and Supplementary Fig. 2E, F). Overall, the glycoforms and their relative percentages before and after WIgGWAM were not significantly different except for [5] G0F/G0F+bGlcNAc and [1] G0F/G0F-GlcNAc, highlighting inconsistencies that can appear for low-abundance glycoforms (Figs. 1b and 3d). The glycoforms before and after WIgGWAM were also largely consistent regardless of the antigen specificity of the monoclonal antibody or the system of expression. (Supplementary Figs. 3–6).
a Deconvoluted intact LC/MS spectrum of recombinant Rituximab IgG1 expressed in Expi293F cells. The dotted lines without an asterisk (*) annotate IgG-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak height. b, c Deconvoluted intact LC/MS spectrum of recombinant Rituximab Fcs after WIgGWAM (b) with no enzymes added. The dotted lines without an asterisk (*) annotate IgG-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak height (c) with EndoS2 added. The dotted lines annotate the masses corresponding to the bifucosylated, monofucosylated, and afucosylated mass of the G1m1 Fc allele. Lines marked with an asterisk (*) are peaks that did not pass the 20% cutoff for one spectrum but did in another. For all intact LC/MS deconvoluted spectra, the highest peak was normalized to 100. Legend created in BioRender. D, J. (2025) https://BioRender.com/2l4vf1u. d Comparison of the relative percentages of the glycoforms identified in recombinant Rituximab as an IgG1 before WIgGWAM (gray) and as an Fc after WIgGWAM (pink). Data are presented as mean values +/- SD. Multiple unpaired t-tests for 3 technical triplicates were used. (nd) or (*) indicate whether there was a significant difference using the two-stage step-up method for false discovery rate (FDR). e Bar graph showing the relative percentages of symmetric (purple), asymmetric (blue), and ambiguous (orange) glycoforms in recombinant Rituximab Fc after WIgGWAM for three technical triplicates. Data are presented as mean values +/- SD. f Table comparing the relative percentages of glycans released with PNGase F from recombinant Rituximab Fc (left column) and glycans identified from WIgGWAM of recombinant Rituximab IgG1 after a mock release calculation. Source data are provided as a Source Data file.
Our method also highlights that IgG1 glycoforms, when produced in Expi293F cells, can be symmetric (i.e., both Fc protomers have the same glycan) or asymmetric (i.e., the glycan on one of the Fc protomers is different from that on the other protomer) (Fig. 3e). When the symmetric and asymmetric pairing possibilities have the same mass, we are unable to distinguish between the two using our current method, and we label these glycoforms as ambiguous. In this sample, there exists ~51% asymmetrically glycosylated glycoforms and 30% symmetrically glycosylated glycoforms; 19% of glycoforms that are ambiguous are likely some mixture of asymmetric and symmetric glycoforms, but our current methods are unable to clarify this ambiguity.
The presence of asymmetric glycoforms emphasizes a feature of IgG glycosylation that can be readily quantified by WIgGWAM but cannot be observed in glycan release studies. To determine how the relative quantification of glycans using WIgGWAM compares to the widely used glycan release method, we compared the results of WIgGWAM on Rituximab IgG1 to the glycan release of Rituximab Fc, both of which we expressed recombinantly in Expi293F cells. First, we performed a mock glycan release for the WIgGWAM results, in which the relative percentage of each glycan pairing is divided equally between each of the two glycans, and the resulting percentages for each specific glycan are summed. For example, WIgGWAM of Rituximab suggests that G0F/G1F makes up ~30% of the glycan composition. In a mock release scenario, 15% would be assigned for the total of each G0F and G1F, and so forth. In some cases where the specific structure of the glycan is ambiguous, we consider a range of possible percentages for each glycan. Upon comparing the mock release and released glycan results, we found that the relative percentages of the majority of glycoforms are consistent between the WIgGWAM and glycan release methods (Fig. 3f). The only glycans that were inconsistent between the two methods were those of very low abundance. Our results indicate that WIgGWAM is a method that allows the identification of symmetric, asymmetric, and ambiguous glycoforms on IgG1 Fcs without significantly affecting the relative percentage of the existing glycoforms, especially for the most abundant glycans.
Asymmetric glycosylation is prevalent in the periphery of humans and IgG1 humanized mice
After confirming that WIgGWAM has a negligible effect on the glycan composition of the sample, we explored whether asymmetric glycosylation is present in peripheral human blood. After performing WIgGWAM on pooled human plasma and serum samples, we found that G1m1 and nG1m1 are the most prevalent IgG1 Fc alleles as is apparent in the EndoS2-treated spectrum, consistent with G1m1 alleles being found in African, European, and Asiatic populations and nG1m1 alleles being found in European populations45 (Fig. 4a, and Supplementary Figs. 7A, B, 8A–C). The untreated spectra for both alleles showed that a majority of glycoforms are asymmetric (Fig. 4b, c, and Supplementary Figs. 7C–F, 8D–I), similar to recombinant antibodies.
a, b Deconvoluted intact LC/MS spectrum of a pooled human plasma sample after WIgGWAM (a) treated with EndoS2. The peaks corresponding to the bifucosylated masses of the G1m1 and nG1m1 Fc alleles are labeled (b) with no enzymes added. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak height for the nG1m1 Fc allele (left) and the G1m1 Fc allele (right). For all intact LC/MS deconvoluted spectra, the highest peak was normalized to 100. c Bar graph showing the relative percentages of symmetric (purple), asymmetric (blue), and ambiguous (orange) glycoforms in pooled human plasma Fcs after WIgGWAM for three technical triplicates. Data are presented as mean values +/- SD. d, e Deconvoluted intact LC/MS spectrum of a serum sample from human IgG1 knock-in mice after WIgGWAM (d) treated with EndoS2. e with no enzymes added. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak height. f Bar graph showing the relative percentages of symmetric (purple), asymmetric (blue), and ambiguous (orange) glycoforms in hIgG1-KI mouse serum Fcs after WIgGWAM for three technical triplicates. Data are presented as mean values +/− SD. Source data are provided as a Source Data file.
We also glycoprofiled human IgG1 (hIgG1) antibodies produced in a mouse strain in which the hIgG1 constant region gene was knocked-in replacing the murine Ighg2c coding sequence using CRISPR-Cas9 gene editing46. The EndoS2-treated spectrum indicated that the hIgG1 is of the G1m1 Fc allele, consistent with only this one heavy chain gene being introduced into the mouse strain (Fig. 4d). Additionally, treating a portion of the sample with EndoS2 and AlfC, an exofucosidase47, indicates that, similar to humans, IgGs produced in mice are mostly bifucosylated, as the peak decreased by 292 Da, consistent with the loss of two fucoses (Supplementary Fig. 9A). The untreated spectrum suggested that 16 glycoforms, out of the 20 for which we scanned, were present in the sample. We further confirmed the glycoforms by treating them with exoglycosidases (Fig. 4e, and Supplementary Fig. 9B, C). When we added exosialidase to our sample, we could no longer detect any sialylated glycoforms [9], [11], [13], [14], [15], [16], [17] and [19] that were identified in the untreated spectrum (Fig. 1b, and Supplementary Fig. 9D). G1m1 IgG1 antibodies produced in mice appear to have increased sialylation compared to G1m1 IgG1 antibodies produced in humans. Additionally, when we added an exogalactosidase to our sample, we could detect the agalactosylated glycoforms [1] G0F/G0F-GlcNAc and [3] G0F/G0F (Fig. 1b, and Supplementary Fig. 9E). We also detected glycoforms [9] G0F/G1SF and [14] G1SF/G1SF or G0F/G2S2F which result from the cleavage of terminal galactosidases from glycoforms containing one or two sialic acids, respectively, i.e., glycoforms [11], [13], [14], [15], [16] and [17]. When both an exosialidase and exogalactosidase are added to the sample, only [1] G0F/G0F-GlcNAc and [3] G0F/G0F are detected (Fig. 1b, and Supplementary Fig. 9F). This analysis shows that at least 70% of hIgG1 glycoforms produced in these knock-in mice are asymmetric (Fig. 4f).
Glycoprofiling human IgG1 antibodies from peripheral human and IgG1 humanized murine blood using WIgGWAM showed that asymmetric glycosylation is prevalent in the periphery of humans and IgG1 humanized mice, suggesting that this is an inherent feature of the glycosylation machinery in mammals.
Asymmetric glycosylation is present in every individual human sample and is regulated by the B-cell differentiation pathway
After glycoprofiling IgG1 antibodies from pooled human plasma and serum samples, we investigated whether asymmetric glycosylation is present in every individual or if the asymmetric IgG glycoforms in pooled samples were derived from only a subset of individuals. Accordingly, we performed WIgGWAM on 31 individual plasma samples, 19 of which were diagnosed with COVID and 12 of which were healthy controls (Supplementary Table 1). The EndoS2-treated samples allow us to determine whether the individual was a homozygote, such as Individual G001, or heterozygote, such as Individual G013, for the IgG1 allele (Fig. 5a, d). We used the untreated spectra to glycoprofile the samples while retaining information about the pairing of the Asn297-linked glycans (Fig. 5b, c, e, f, and Supplementary Fig. 10). We found that, regardless of allele, zygosity, or infection status, asymmetric glycosylation is prevalent in every individual. On average, individuals in this study had ~27% symmetric glycosylation, 60% asymmetric glycosylation, and 14% ambiguous glycosylation (Fig. 5g). We observed no significant difference between the type of glycan pairing and health status, which suggests that while disease can change the type of glycans35 found on IgG1 antibodies, the glycoforms remain predominantly asymmetric. When additional individual samples (see below) are included, we have now analyzed serum or plasma from over 80 individual human subjects and have yet to identify a single individual who lacked asymmetrically glycosylated IgG antibodies. Indeed, our data suggests that asymmetrically glycosylated IgG1 antibodies are the predominant type of glycan pairing in the blood of every individual, regardless of health status.
Deconvoluted intact LC/MS spectrum of a plasma sample after WIgGWAM a from a COVID patient G001 and treated with EndoS2. The peak corresponding to the bifucosylated mass of the G1m1 Fc allele is labeled b from a COVID patient G001 with no enzymes added c from a healthy individual G020 with no enzyme added d from a COVID patient G013 with EndoS2 added. The peaks corresponding to the bifucosylated masses of the G1m1 and G1m1,27 Fc alleles are labeled e from a COVID patient G013 with no enzymes added f from a healthy individual G021 with no enzyme added. The dotted lines annotate Fc-linked glycoforms whose peak heights were at least greater than 20% of the maximum peak. For all intact LC/MS deconvoluted spectra, the highest peak was normalized to 100. g The relative percentage of symmetric (left), asymmetric (middle), and (right) ambiguous Fc-linked glycoforms after WIgGWAM of healthy (black) and COVID patient (pink) plasma samples (n = 31). The relative percentage of h total galactosylation i.e., glycan pairs with at least one galactose (left), and asymmetric galactosylation, i.e., glycan pairs with different number of galactoses on each glycan (right) and i total sialylation i.e., glycan pairs with at least one sialic acid (left), and asymmetric sialic acid, i.e., glycan pairs with different number of sialic acids on each glycan (right) of IgG Fcs after WIgGWAM of culture supernatants from memory B-cells stimulated with IL21 in combination with CD40L (black), R848 (pink), or CpG (green) (n = 11 for CD40L, n = 11 for R848, n = 9 for CpG, with 17 unique donors). 2way ANOVA with Tukey’s multiple comparisons test was used for the statistical analysis. Source data are provided as a Source Data file.
To determine whether asymmetric glycosylation of IgGs is a regulated process, we cultured memory B cells of healthy donors stimulated with CD40L (a costimulatory ligand), R848 (a TLR-7 agonist), or CpG (a TLR-9 agonist) in the presence of IL-21, which are conditions known to promote B-cell activation and IgG production. CD40L stimulation mimics T-cell-dependent activation of B cells. TLR stimulation, on the other hand, is associated with T-cell-independent extrafollicular activation of B-cells. We used WIgGWAM to analyze secreted IgGs from the culture supernatant using the protocol described earlier (Figs. 1 and 2b) with three notable deviations: (1) we increased the amount of papain added to account for the increased sample volume, (2) we added 11 glycan pairs to the analysis to account for the increased bisection in the secreted IgGs (Supplementary Fig. 11A), and (3) we treated the samples with carboxypeptidase B to remove the terminal lysines from the IgG Fcs. Our results show that under all conditions and for all donors, the secreted IgGs were predominantly asymmetrically glycosylated (Supplementary Fig. 11B–AG). We found that CD40L stimulation resulted in increased total galactosylation, i.e., glycan pairs with at least one galactose, and asymmetric galactosylation, i.e., glycan pairs with different number of galactoses on each glycan, of IgGs compared to R848 and CpG (Fig. 5h). Similarly, CD40L stimulation resulted in increased total and asymmetric IgG sialylation compared to R848 (Fig. 5i). Our results indicate that memory B cells secrete asymmetrically glycosylated IgGs that are regulated by the pathway taken by a B-cell to plasma cell differentiation.
Monofucosylation is a prevalent feature of peripheral hIgG1 antibodies and is associated with secondary and severe dengue disease
To investigate the role of asymmetric glycosylation of IgG1 antibodies in human disease, we performed WIgGWAM on 58 plasma samples from patients with dengue disease of varying severity (Supplementary Table 2). Previous studies have shown that afucosylation is increased in secondary compared to primary infections and in severe dengue compared to mild disease24. However, because those studies were performed using glycan release methods, no information exists concerning the symmetry of fucosylation or lack thereof. Thus, we investigated whether asymmetric fucosylation, which we and others call monofucosylation, but is also known as hemifucosylation48, exists in human IgG1 antibodies. We focused on the EndoS2-treated spectra, in which the only heterogeneity stems from the IgG1 allele(s) and fucosylation of the samples, and we determined the relative percentage of bifucosylation, monofucosylation, and afucosylation (Fig. 6a, b, and Supplementary Fig. 12). In Sample 23, the individual was a homozygote for the G1m1 Fc allele, and ~80% of their IgG1 Fcs were bifucosylated, 11% were monofucosylated, and 9% were afucosylated. In Sample 37, the individual was a heterozygote for the G1m1/nG1m1 alleles, and ~90% of their IgG1 Fcs were bifucosylated, 6% were monofucosylated, and 4% were afucosylated. We compared the results of fucosylation levels between WIgGWAM and traditional glycan release methods using the same mock release described above (Supplementary Fig. 13). Overall, both methods had consistent results. While the percentages of afucosylation were higher when determined by WIgGWAM, both methods similarly show that Sample 23 has higher afucosylation compared to Sample 37.
a, b Deconvoluted intact LC/MS spectrum of a dengue patient plasma sample after WIgGWAM and addition of EndoS2. The dotted lines annotate the masses corresponding to the bifucosylated, monofucosylated, and afucosylated mass of the G1m1 and nG1m1 Fc alleles. The bar graphs depict the relative percentage of bifucosylated (light purple), monofucosylated (blue), and afucosylated (dark purple) glycoforms. The relative percentage of bifucosylation, monofucosylation, and afucosylation present in dengue plasma samples when divided according to primary (black) vs secondary infection (pink) (c) or according to disease classification DF (black) vs DHF/DSS (pink) (d) (n = 58). Two-tailed Mann-Whitney test was used for the statistical analysis. Created in BioRender. D, J. (2025) https://BioRender.com/lybz9tn. Source data are provided as a Source Data file.
We then compared primary and secondary dengue infection, while differentiating between monofucosylation and complete afucosylation on IgG1 Fcs. We found that, consistent with previous studies24, bifucosylation is decreased in secondary infections. However, this reduction in bifucosylation is due to an increase in monofucosylation and not to an increase in afucosylation in secondary dengue infections, although the latter trends towards significance (Fig. 6c). Similarly, when comparing fucosylation levels between the less severe dengue fever (DF) classification and the more severe dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS) classifications, bifucosylation was significantly decreased in DHF/DSS relative to DF and only monofucosylation was significantly increased in the more severe cases of dengue (Fig. 6d). Overall, our results demonstrate that monofucosylated IgG1 antibodies exist in human blood and that they, rather than completely afucosylated IgG glycoforms, drive antibody-dependent enhancement of dengue disease49.
Monofucosylated IgG1 binds Fc receptors and induces effector functions similarly to fully afucosylated IgG1
Since data from patient sera suggested that monofucosylated IgG1s are associated with dengue disease severity, we hypothesized that monofucosylated IgG1s should bind to FcγRIIIA with a high affinity. To investigate this, we developed a chemoenzymatic strategy to glycoengineer a recombinant monofucosylated IgG1. We used knob-in-hole (KiH) mutations50 to generate a heterodimeric anti-DENV2 E protein antibody (C10) with two distinct Fc protomers. We also mutated residue A330 to Trp (W) in one of the protomers - a mutation that inhibits CU43 binding51. CU43 is an IgG-specific endoglycosidase that cleaves glycans between the first and second GlcNAc, similarly to EndoS252. As a result, when we added CU43 to the KiH heterodimeric antibody with the A330W mutation on only one protomer, one of the glycans is reduced to a GlcNAc and a fucose, while the other remains intact. Then, we used AlfC, an exofucosidase, to selectively cleave the exposed fucose47. This resulted in a monoglycosylated monofucosylated IgG1. We then used the glycosynthase variant EndoS2D184M with oxazoline-linked S2G2 glycans53 to form a diglycosylated monofucosylated IgG1. Finally, we used an exosialidase and exogalactosidase to reduce both glycans to G0 so that the only variation in the glycosylation of both protomers results from the presence or absence of the fucose (Fig. 7a). We generated an afucosylated IgG1 that contains the KiH and A330W mutations by expressing the antibody heavy and light chains in Expi293F cells in the presence of 2-deoxy-2-fluoro L-Fucose, a fucosyltransferase inhibitor54. We also expressed a bifucosylated KiH-A330W IgG1. We used an exogalactosidase to ensure that, similarly to the monofucosylated variant, the afucosylated and bifucosylated Abs were agalactosylated. We confirmed the glycoforms of all the generated antibodies using intact LC/MS (Supplementary Fig. 14).
a Schematic for glycoengineering monofucosylated agalactosylated recombinant antibodies. Created in BioRender. D, J. (2025) https://BioRender.com/3aevmgm. b Surface plasmon resonance (SPR) sensograms of bifucosylated, monofucosylated, and afucosylated anti-DENV2 E protein IgG1 binding to FcγRIIIA-F158. c Ifnar1−/− FcγR humanized mice were injected with 20 µg i.v. of afucosylated (n = 6 mice), monofucosylated (n = 5 mice), bifucosylated (n = 5 mice), or FcR null (G236R/L328R) (n = 5 mice) anti-DENV2 E protein IgG1s (C10) 6 h before challenge with DENV2 virus. Created in BioRender. D, J. (2025) https://BioRender.com/3aevmgm. d Change in the body weight of the mice from baseline. The dotted line represents the 80% weight loss threshold which defines death. e Comparison of areas under the curve (AUC) shown in (d) as a quantification of weight loss. Data are presented as mean values +/- SD. f Percentage survival of the mice, defined as a loss of greater than 20% of bodyweight. g Contour plot of CD3+ T cells gated on CD4+ or CD8+ in FcγR humanized mice (n = 6 per group) 24 h after injection with 10 µg i.v. of afucosylated, monofucosylated, or bifucosylated anti-CD4 IgG1 (YTS191). h Quantification of the frequency of CD4 + T cells over total CD3+ T cells after injection with afucosylated, monofucosylated, or bifucosylated anti-CD4 IgG1. Data are presented as mean values +/- SEM. Ordinary one-way ANOVA with Bonferroni’s multiple comparisons test was used for all the statistical analysis. For all in vivo studies, the results are from two independent experiments. Source data are provided as a Source Data file.
We used surface plasmon resonance (SPR) to test the binding of all three C10 glycovariants to human Fc γ receptors. The bifucosylated IgG1 bound to FcγRIIIA with a 129 nM affinity, consistent with previous literature55 (Fig. 7b, left). On the other hand, both monofucosylated and afucosylated IgG1s bound to FcγRIIIA with a higher affinity of 56 nM and 39 nM, respectively (Fig. 7b, middle and right). Overall, this demonstrates that the absence of only one fucose is sufficient to impart the higher affinity to FcγRIIIA that has been well established for afucosylated IgG1s6. All three glycoforms bound to the remaining Fc γ receptors, FcγRI, FcγRIIb, and FcγRIIa, with similar affinities (Supplementary Fig. 15A–C). Because the monofucosylated antibody was generated via a chemoenzymatic transglycosylation reaction that did not go to completion, there remained a small portion that was monoglycosylated (Supplementary Fig. 14E). To investigate the binding of this species, we generated monoglycosylated bifucosylated and monoglycosylated monofucosylated IgGs (Supplementary Fig. 15D–F). Both had sufficiently reduced binding to FcγRIIIA compared to diglycosylated antibodies with a KD of 1640 nM and 554 nM, respectively (Supplementary Fig. 15G, H) as to be unlikely to affect the binding affinities that we measured for the engineered diglycosylated IgG1 glycovariants.
To confirm whether the high affinity of monofucosylated IgGs to FcγRIIIA translated to increased pathogenicity in vivo, we tested the ability of the C10 glycovariants to enhance dengue infection in Ifnar1 knock-out FcγR humanized mice (Fig. 7c). Mice that were administered either the afucosylated or monofucosylated C10 exhibited significantly lower body weight and survival upon challenge with DENV2 virus compared to those receiving the bifucosylated or FcR null control C10 (Fig. 7d–f), indicating that both monofucosylation and afucosylation can drive severe dengue infection. These results also validate our finding that in human patients, ADE is driven by monofucosylated IgG1s.
Additionally, we engineered bifucosylated, monofucosylated, or afucosylated versions of the anti-CD4 IgG1 YTS191 (Supplementary Fig. 16A–F) and compared their efficacies in depleting CD4+ T cells in FcγR humanized mice. Monofucosylated YTS191 led to an almost complete depletion of CD4+ T cells, similar to that caused by afucosylated YTS191, whereas bifucosylated YTS191 led to no depletion (Fig. 7g, h, and Supplementary Fig. 16G). Overall, we show that monofucosylated IgG1s bind to Fcγ receptors in vitro and induce effector functions in vivo as do fully afucosylated IgG1s.
Monofucosylated IgG1 binds to FcγRIIIA in a high-affinity orientation
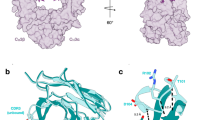
To investigate the mechanism by which monofucosylated IgG1s are able to bind to FcγRIIIA with a high affinity similar to that of afucosylated IgG1s, we examined published structures of bifucosylated and afucosylated Fcs in complex with FcγRIIIA. Previous studies have shown that in afucosylated Fcs, residue Y296 adopts a conformation that enables the formation of a hydrogen bond with residue K128 of the receptor56. Other studies also suggest that the absence of the fucose enables increased intermolecular carbohydrate interactions with the glycan on N162 of the receptor57,58. FcγRIIIA binds to the IgG Fc asymmetrically, in which the described side chain and glycan interactions are mainly driven by only one of the Fc protomers, depicted as chain A in Fig. 8a. Conversely, chain B plays a minor role in the binding, in which most of the intermolecular interactions are driven by the protein backbone. Accordingly, since the IgG is fully symmetrical in the bifucosylated and afucosylated states (i.e., the protein and glycan components on both Fc protomers are identical), they bind equivalently to FcγRIIIA in either orientation with predominant interactions with chain A or chain B, since these chains are indistinguishable in these antibodies (Fig. 8b). The only difference is that both orientations result in low-affinity FcγRIIIA binding for bifucosylated IgG but high-affinity FcγRIIIA binding for afucosylated IgG. We hypothesized that monofucosylated IgG Fcs bind in an orientation in which the afucosylated protomer is in the chain A position, while the fucosylated protomer is in the chain B position, leading to similar binding properties as when chain B also lacks a fucose. That is, monofucosylated IgG binds FcγRIIIA by restricting the possible orientations of Fc-FcγRIIIA complexes to the orientation that mimics the afucosylated IgG interaction, thereby resulting in high-affinity binding.
a Crystal structure of bifucosylated Fc with FcγRIIIA (PDB: 5XJE)57. The N-glycans are depicted as spheres with the Fc N297 core fucoses colored in red. Distances between E269 of chain A of the Fc and K131 of FcγRIIIA and E269 of chain B of the Fc and K114 of the receptor are highlighted. b Diagram depicting the potential binding orientations of bifucosylated, afucosylated, and monofucosylated IgGs to FcγRIIIA, shown as a gray oval. Surface plasmon resonance (SPR) sensograms of c bifucosylated A330W E269I KiH Rituximab IgG1, d afucosylated A330W E269I KiH Rituximab IgG1, e monofucosylated A330W E269I KiH Rituximab IgG1, f bifucosylated A330W E269M KiH Rituximab IgG1, g afucosylated A330W E269M KiH Rituximab IgG1, and h monofucosylated A330W E269M KiH Rituximab IgG1 binding to FcγRIIIA-F158. Created in BioRender. D, J. (2025) https://BioRender.com/2vqcv7e Source data are provided as a Source Data file.
To test this hypothesis, we utilized the same KiH-A330W strategy of engineering monofucosylated IgGs as described above. This time, we also mutated E269 to an Ile (I) in the afucosylated protomer (Supplementary Fig. 17A, B). Deep mutational scanning of the IgG1 Fc indicates that E269I leads to loss of binding to FcγRIIIA59. Examining the structure of the complex indicates that the glutamate at position 269 in chain A of the Fc forms side chain polar contacts with the receptor; however, it is dispensable in chain B as the side chain is pointing away from the binding interface and is positioned more than 10 Å from the nearest FcγRIIIA residue (Fig. 8a). So, if we introduce the E269I mutation in the afucosylated protomer, we predict that the IgG Fc will have to bind in a conformation where the fucosylated protomer is in the chain A position and the afucosylated protomer with the E269I mutation is in the chain B position. We also generated bifucosylated and afucosylated IgGs with the same mutations as controls (Supplementary Fig. 17C–F) and tested binding of these constructs to FcγRIIIA. The bifucosylated IgG had an affinity of 2070 nM, consistent with its lower overall binding affinity to the receptor (Fig. 8c). The afucosylated IgG had a higher affinity of 159 nM (Fig. 8d), while the monofucosylated IgG exhibited an affinity of 698 nM (Fig. 8e). These results suggest that this monofucosylated IgG construct binds with low affinity to FcγRIIIA because it is prohibited by the E269I mutation from accessing the high-affinity orientation defined by the afucosylated Fc-FcγRIIIA interaction.
We also designed a similar monofucosylated IgG construct, this time mutating E269 to a Met (M) in the fucosylated protomer, as well as the bifucosylated and afucosylated counterparts (Supplementary Fig. 18). Similar to E269I, E269M abolishes FcγRIIIA binding59. Since the E269M mutation is on the fucosylated protomer, we expect this monofucosylated construct to retain its high affinity to FcγRIIIA. Indeed, with this set of constructs, bifucosylated IgG again binds to the receptor with an affinity of 1480 nM (Fig. 8f). Both afucosylated and monofucosylated IgGs bind to the receptor with a similar high affinity of 152 nM (Fig. 8g) and 119 nM (Fig. 8h), respectively. Our results suggest that monofucosylated IgGs bind to FcγRIIIA in a high-affinity orientation, such that the afucosylated protomer is in a position that enables increased intermolecular protein and carbohydrate interactions similar to those in the afucosylated Fc-FcγRIIIA complexes.
Discussion
Glycosylation is an important post-translational modification of IgG antibodies that impacts their structure and function60. Specifically, the conserved Asn297-linked complex type glycan on the Fc region of IgG controls the binding affinities to activating and inhibitory Fc γ receptors and complement C1q. These differences in affinities lead to modified antibody effector functions such as ADCC, ADCP, and complement activation4,5, effector functions that enhance or dampen the immune response to pathogens and tumors61. ADCC is also an important mechanism by which some therapeutic mAbs function62.
The majority of current methods employed in the field to study IgG glycosylation rely on glycan release workflows, in which the glycans are enzymatically cleaved from Asn297 on IgG Fcs26. Other studies have resorted to symmetrical glycoengineering, in which enzymes are used to produce IgGs in which glycans on both Fc homodimers are identical63. However, both methods miss a critical aspect of Fc glycobiology: the heterogeneity of glycosylation suggests that the glycan on each Fc monomer does not necessarily have to be identical; Fc glycosylation can be asymmetrical. Thus, we sought to explore how we can change and improve our understanding of the function of IgG Fc glycans by studying them as paired glycoforms, rather than independent glycans. Accordingly, we developed WIgGWAM, an intact LC/MS-based method for the glycoprofiling of polyclonal IgG1 antibodies that preserves the spatial pairing of the glycans. We ensured the reproducibility and consistency of our method by setting a predetermined filtering analysis process. Additionally, we used monoclonal antibodies to show that our workflow has no effect on the glycan composition of our sample. We demonstrated that human IgG1 antibodies are predominantly asymmetrically glycosylated regardless of allele, zygosity, or health status. We showed that IgG1 glycosylation is regulated by the pathway taken by a B-cell to plasma cell differentiation. Further, we showed that fucosylation can also be asymmetric and that monofucosylated IgG1 antibodies exist in every individual. Our analysis of dengue patient plasma samples indicates that increased monofucosylation, but not complete afucosylation, correlates with severity of disease. Our studies support and expand upon the hypothesis in the existing literature that enhanced dengue disease is a result of increased binding to FcγRIIIA24,49. However, this has usually been attributed to afucosylation, largely due to glycan release methods. Our results demonstrate that the threshold of increased FcγRIIIA binding necessary to induce enhanced disease can be achieved with monofucosylation. Thus, while both afucosylation and monofucosylation can cause ADE, the glycoform that requires the least deviation from the predominant bifucoyslation–monofucosylation–is sufficient and observed. Finally, we utilized antibody engineering and endoglycosidases to develop a monofucosylated recombinant IgG1. We determined via in vitro SPR and several in vivo FcγR humanized mouse models that monofucosylated IgG1 binds to Fc γ receptors with the same affinity as afucosylated IgG1, including having a high affinity to FcγRIIIA, resulting in identical effector functions. This positions monofucosylated mAbs as a therapeutic alternative to afucosylated mAbs, as they share the enhanced function that led to the clinical approval of some afucosylated anti-tumor mAbs25 but may be more cost and time-effective to produce.
Exploring the mechanism and role of monofucosylation is essential in furthering our understanding of how glycoform pairings impact Fc receptor affinity. Fucosylation is one of the best-characterized IgG glycan modifications. Studies have shown that the absence of fucose on IgG Fcs leads to increased FcγRIIIA binding64. Our results indicate that the loss of one fucose is sufficient to impart this increased affinity, as the monofucosylated Fc binds in a conformation that enables the increased intermolecular contacts that characterize the high-affinity interaction between afucosylated IgG Fcs and FcγRIIIA. Other studies also support the idea that monofucosylated IgG Fcs have an increased affinity to FcγRIIIA. When bifucosylated, monofucosylated, and afucosylated IgGs were purified from a mixture using an FcγRIIIA affinity column, the receptor affinity and ADCC activity of the predominantly monofucosylated fraction were very similar to that of the mostly afucosylated fraction48,65,66,67. Additionally, molecular dynamic simulations of FcγRIIIA binding to fucosylated and afucosylated IgGs show that upon the addition of a fucose to IgG Fcs, the receptor shifts contacts to the opposite Fc chain58. When IgG1 antibodies were purified from NK cells and their glycans released, approximately half of the glycans were afucosylated, which is what one would expect if these IgG1 antibodies were monofucosylated68. On the other hand, our studies and others suggest that fucosylation has minimal effect on binding to FcγRI, FcγRIIa, and FcγRIIb69. This can be further confirmed with functional studies that examine these receptor-mediated effector functions.
The high-affinity binding of monofucosylated IgG1s to FcγRIIIA supports the hypothesis that the affinity to the receptor is largely determined by the higher-affinity glycan. That is, monofucosylated IgGs exhibit Fc γ receptor binding that is hierarchical, in which one glycan dominates the Fc-Fc γ receptor interaction over the other. Whether all asymmetrically glycosylated IgG, and not just monofucosylated IgGs, bind Fc γ receptors hierarchically dependent on the glycan remains unclear. Asymmetric glycosylation is also likely to impact binding to C1q and Type II Fc receptors. Similar to Fc-Fc γ interactions, C1q binding to Fc receptors is asymmetric, in which different residues on each Fc protomer are essential for binding70. Since galactosylation of IgG has been shown to affect binding affinity to C1q12, it is reasonable to expect that asymmetric galactosylation can impact CDC. For Type II Fc receptors, such as DC-SIGN, direct binding with sialylated IgG remains contested71. However, recent studies suggest that sialylated IgGs interact indirectly with DC-SIGN via FcγRIIb72. Thus, asymmetric sialylation may impact binding to FcγRIIb, which is asymmetric, and indirectly affect DC-SIGN engagement. On the other hand, the binding between IgGs and the neonatal receptor FcRn is generally independent of the glycosylation status of the Fc73, so asymmetric glycosylation should have little impact on IgG recycling and serum half-life. As such, it is important to expand our studies to additional glycan modifications that play a role in disease, such as sialylation and galactosylation9,16 in order to improve our understanding of how biologically relevant glycan pairs contribute to antibody effector functions.
Understanding how asymmetrical glycosylation is regulated will further our mechanistic understanding of the correlation between glycoforms and antibody effector functions. Our studies suggest that T-cell-dependent activation of B-cells enhances asymmetric galactosylation and sialylation compared to T-cell-independent activation of B-cells. This shows the need for further studies that investigate the role of B-cell differentiation pathways in regulating asymmetric glycosylation. Additionally, studying the role of asymmetric glycosylation in non-viral contexts such as autoimmune diseases or cancer will help elucidate avenues for diagnostics and therapeutics.
Similarly, understanding how asymmetric glycosylation varies with demographic factors such as age and sex, as well as other pathological and metabolic factors, such as hormones and nutrition, will enable us to meaningfully and consistently separate differences between diseased and healthy populations from individual variations in glycosylation.
Limitations of the study
As described above, some fraction, typically about 20%, of IgG antibodies in our analysis are ambiguous as to whether they are symmetrically or asymmetrically glycosylated. This ambiguity results from certain glycan pairings (e.g., G1F/G1F [symmetrical] and G0F/G2F [asymmetrical]) having identical masses, making these glycoforms indistinguishable by the current WIgGWAM method. While this does not affect any of the findings reported here, fully defining IgG glycoforms may be important in future studies. It is likely that we will be able to break this ambiguity using various mass spectrometry ion fragmentation methods. Additionally, our 20% cutoff filtering method for data analysis can result in the exclusion of lower-abundance glycans. Other methods, such as glycan release, can be used in parallel to determine their presence. Also, our WIgGWAM analysis of IgG glycoforms is currently restricted to the IgG1 subclass, since IgG1 antibodies are by far the most abundant antibody type in blood, and mass spectrometry methods are highly sensitive to the relative abundancy of individual ions. In order to observe lower abundance ions, such as those derived from the IgG2, IgG3 and IgG4 subclasses, it will be necessary to deplete all IgG1 antibodies in a given sample prior to implementing the WIgGWAM workflow. This will likely require an increase in the minimal sample required for the analysis.
Methods
WIgGWAM workflow and EndoS2 and exoglycosidase reactions
For WIgGWAM, papain (Millipore Sigma 10108014001) was added to the IgG sample in a 1:10 (papain to IgG) ratio by mass. The reaction was diluted to 500 µL–1 mL using phosphate-buffered saline (PBS) and incubated at 37 °C for 30 min. The Fcs were purified using Protein A resin (Thermo Fisher) with PBS pH 7.4 being used as the binding buffer and 100 mM sodium citrate buffer pH 3.0 as the elution buffer. The fractions were neutralized with 1 M Tris pH 9.3. The Fcs were buffer exchanged into PBS and stored at 4 °C until ready for use.
A 20 µL 5 µM solution of the Fcs after WIgGWAM was prepared and either ran using intact LC/MS without any enzymes added or with 50 nM EndoS237, 50 µM AlfC47, 600 nM BgaA43, or 500 nM MvNA42.
Intact LC/MS analysis
The Fc samples were analyzed using an Agilent 1290 Infinity II LC System equipped with a 50 mm PLRP-S column from Agilent with 1000 Å pore size. The LC system is attached to an Agilent 6545XT Q-TOF mass spectrometer (Agilent, Santa Clara, CA). The mobile phases were 0.1% formic acid in water (Buffer A) and 0.1% formic acid in acetonitrile (Buffer B). Column temperature was 80 °C with a flow rate of 0.3 ml/min. 0.1 µL of sample were injected, and separation was performed using a 4.5 min method: 0 min: 98% Buffer A and 2% Buffer B (diverted to waste); 1.63 min: 80% Buffer A and 20% Buffer B; 3.4 min: 50% Buffer A and 50% Buffer B; 4 min: 10% Buffer A and 90% Buffer B.
The MS conditions were dual Agilent Jet Stream ESI source, positive mode, Sheath gas 400 °C at 11 L/min, dry gas 350 °C at 12 L/min, nebulizer pressure 60 psi, VCap 5500 V, Nozzle 2000 V, Fragmentor 380 V, m/z range 800–5000.
Relative amounts of each of the deglycosylated peaks were quantified after deconvolution of the raw data and identification of the corresponding peaks using BioConfirm 10.0 (Agilent, Santa Clara, CA).
Recombinant Rituximab and Trastuzumab IgG expression and purification
Rituximab or Trastuzumab heavy chain and light chain plasmids were ordered from Thermo Fisher Scientific in the pcDNA3.4-TOPO vector and transfected in a 1:2 ratio by mass in Expi293F cells. The plasmids were transfected as per manufacturers protocol (MAN0007814, Thermo Fisher Scientific) with the addition of Penicillin/Streptomycin mix 24 h post transfection. The cells were cultured for 96 h before harvesting. The proteins were purified using Protein A resin (Thermo Fisher) with PBS pH 7.4 used as the binding buffer and 100 mM sodium citrate buffer pH 3.0 as the elution buffer. The fractions were neutralized with 1 M Tris pH 9.3. The proteins were buffer exchanged into PBS and stored at 4 °C until ready for use.
WIgGWAM of commercial recombinant Trastuzumab
Commercial Trastuzumab expressed in CHO cells (Herceptin, Genentech Inc., 150 mg) was purchased from Emory University Hospital and reconstituted following package instructions. WIgGWAM analysis was performed as described earlier in 3 technical replicates.
WIgGWAM of pooled human plasma and serum samples
Pooled human plasma (Innovative Research Inc., VWR CAT# 102768-846) and serum (Innovative Research Inc., VWR CAT# 102768-938) were purchased and stored at −20 °C until ready for use. WIgGWAM analysis was performed as described earlier.
WIgGWAM from knock-in mouse serum
Serum was pooled from hIgG1 knock-in mice46 and WIgGWAM analysis was performed as described earlier. However, Fcs were purified using CaptureSelect™ FcXP Affinity Resin (ThermoFisher, CAT# 1943712005) in 3 technical replicates.
Glycan release and instant PC labeling and analysis
Glycans were released and labeled with instant PC according to the manual of Agilent AdvanceBio Gly-X N-Glycan Prep with InstantPC Kit (https://www.agilent.com/cs/library/usermanuals/public/5994-1231EN.pdf).
The labeled glycans were analyzed with an Agilent AdvanceBio Glycan Mapping column (1.8 µM, 2.1 × 150 mm) on an Agilent 1290 Infinity II LC System attached to an Agilent 6545XT Q-TOF mass spectrometer (Agilent, Santa Clara, CA). The mobile phases were 50 mM NH4F in water (Buffer A) and 100% acetonitrile (Buffer B). Column temperature was 40 °C with a flow rate of 0.25 mL/min. 2 µL of sample were injected and separation was performed using a 23-min method: 0 min: 18% Buffer A 82% Buffer B; 2 min: 18% Buffer A and 82% Buffer B; 2.5 min: 21% Buffer A and 79% Buffer B; 15 min: 60% Buffer A and 40% Buffer B; 15.5 min: 18% Buffer A and 82% Buffer B; 22 min:18% Buffer A and 82% Buffer B.
The MS conditions were dual Agilent Jet Stream ESI source, positive mode, Sheath gas 300 °C at 10 L/min, dry gas 150 °C at 1 L/min, nebulizer pressure 35 psig, VCap 2500 V, Nozzle 500 V, Fragmentor 120 V, m/z range 600–3200.
Glycan abundances were determined by summing the peak areas of all observable charge states and adducts for each glycan composition after manual inspection in Skyline 23.1.0.45574.
PBMC isolation and B cell enrichment
Fresh human leukopaks were obtained from the New York Blood Center. Leukopaks were evenly allocated into 50 mL conical tubes (around 20 mL per tube) and were diluted with Fetal Bovine Serum (FBS) buffer (5% FBS in HBSS) to achieve a final volume of 35 mL. Diluted cell suspension was then layered onto 15 mL of Ficoll (Fisher Scientific, Catalog No. 45-001-749) in new 50 mL conical tubes. Tubes were centrifuged at 935 × g for 20 min at room temperature (3 acceleration, 1 deceleration). The resulting buffy coat, rich in PBMCs, was carefully aspirated and aspirates from the same donor were pooled. After washing the pooled cells, the PBMC pellet was resuspended in 50 mL of 5% FBS buffer. A small 10 µL aliquot was removed for counting, while the rest of the cells were pelleted (526 × g, 7 min, room temp). Following this, B cells were enriched according to the EasySep™ Human B Cell Isolation Kit protocol (STEMCELL Technologies, Catalog No. 17954).
Staining and cell sorting
After their enrichment, B cells were centrifuged (526 × g, 7 min, 4 °C) and prepared for fluorescence-activated cell sorting (FACS). For identifying IgG Memory B Cells, a custom antibody cocktail was prepared with the following antibodies, along with their specified dilutions:
-
Fixable Viability Dye eFluor™ 506 (1/500, eBioscience, Catalog No. 65-0866-14).
-
CD19-PE-Cy7 (1/50, Biolegend, Catalog No. 302216).
-
CD27-PE (1/50, eBioscience, Catalog No. 12-0279-42).
-
CD38-PE-eFluor™ 610 (1/50, eBioscience, Catalog No. 61-0389-42).
-
IgA-APC (1/100, Miltenyi Biotec, Catalog No. 130-113-472).
-
IgM-BV421 (1/100, Biolegend, Catalog No. 314516).
The enriched B cell pellet was resuspended and incubated in the prepared antibody cocktail for 20 min on ice, protected from the light. Following a single wash with 2 mL of 5% FBS buffer and centrifugation at 526 × g RPM for 5 min at 4 °C, the pellet was resuspended in 500 µL of sorting buffer (5% FBS in PBS). Subsequently, IgG Memory B Cells (CD19+, CD27+, CD38-, IgM-, IgA-) were isolated using the BD FACSAria III Cell Sorter and collected into 5 mL polypropylene tubes containing 1 mL of Ex Vivo Serum-free Medium (Fisher Scientific, Catalog No. BW04-418Q).
B cell culture and stimulation
Following sorting, the collected cells were centrifuged at 526 × g for 7 min at 4 °C. The resulting pellet was resuspended in Ex Vivo medium. Cell concentration and viability were assessed by staining a 10 µL aliquot with 10 µL of Acridine Orange Propidium Iodide (AOPI) solution with a Nexcelom Cellometer. Cells were plated in a 96 well round bottom culture plate at 100,000 cells per well in 200 µL of Ex Vivo medium supplemented with one of three stimulation cocktails:
-
Interleukin 21 (IL21; 40 ng/mL, Thermo Fisher, Catalog No. 200-21-10UG) and CD40L (100 ng/mL, Enzo Life Sciences, Catalog No. ALX-522-110).
-
Interleukin 21 (IL21; 40 ng/mL, Thermo Fisher, Catalog No. 200-21-10UG) and R848 (Resiquimod; 2.5 µg/mL, Invivogen, Catalog No. tlrl-r848-10).
-
Interleukin 21 (IL21; 40 ng/mL, Thermo Fisher, Catalog No. 200-21-10UG) and CpG ODN 2006 (Class B human TLR9 agonist; 1.25 nM, Invivogen, Catalog No. tlrl-2006-1).
Cell Cultures were maintained in an incubator for 15–20 days (37 °C, humidified with 5% CO₂). A full medium exchange was performed every 3–4 days. For medium exchange, the culture plate was centrifuged (526 × g, 8 min, 4 °C) to pellet the cells. The conditioned supernatant was then carefully aspirated and replaced with 200 µL of fresh Ex Vivo medium containing the appropriate stimulation cocktail at its final 1× concentration. The cell pellet was gently resuspended before returning the plate to the incubator, and the collected medium was stored at −20 °C for WIgGWAM analysis.
WIgGWAM of B cell culture supernatant
0.5 mg (50 µl of a 10 mg/mL solution) papain (Millipore Sigma 10108014001) was added to 2–5 mL of B cell culture supernatant and incubated at 37 °C for 30 min. The Fcs were purified using Protein A resin (Thermo Fisher) with PBS pH 7.4 being used as the binding buffer and 100 mM sodium citrate buffer pH 3.0 as the elution buffer. The fractions were neutralized with 1 M Tris pH 9.3. The Fcs were buffer exchanged into PBS and stored at 4 °C until ready for use.
A 20 µL 5 µM solution of the Fcs after WIgGWAM was prepared and either ran using intact LC/MS with 0.5ul of 0.1 mg/mL rat carboxypeptidase-B (Thermofisher Cat # 400-00) added, or with both 50 nM EndoS237 and carboxypeptidase-B added.
Analysis of COVID and dengue samples
Prior to WIgGWAM analysis as described earlier, all the plasma samples from the COVID and dengue cohorts were blinded and analyzed in a randomized order.
Dengue patient recruitment and diagnosis
Blood samples were obtained from hospitalized children (≥2 years) who presented with dengue-like symptoms at the Kanta Bopha Hospital in Phnom Penh, Cambodia. Patients were classified according to the WHO 1997 criteria upon hospital discharge75. Platelet counts were determined by complete blood count at the hospital. Plasma was separated by density gradient centrifugation and tested for the presence of DENV by real-time reverse transcription polymerase chain reaction (RT-qPCR) as described before76. Patients were identified as having primary/secondary DENV infection using a hemagglutination inhibition test in line with the WHO criteria75.
Mice
All mice were maintained in a controlled temperature (20–25 °C) and humidity (30–70%) with a 12 h dark:light cycle at the Comparative Bioscience Center at Rockefeller University. FcγR humanized mice and Ifnar1−/−/FcγR humanized mice were developed and characterized as previously described77,78.
Measurement of FcγR affinity
Binding kinetics were measured using the Biacore T200 SPR system (Cytiva) as previously described78. Glycoengineered antibodies were immobilized on a protein G chip (Cytiva) at a density of 2000 response units (RU). Soluble human Fcγ receptor ectodomains were serially diluted at the indicated concentrations (ranging from 2000 to 0.78 nM; diluted 1:2 in HBS-EP+ buffer (Cytiva)) and injected through flow cells at a flow rate of 20 µl min−1 for 60 s (association time), followed by 300 s of dissociation. Chip surface was regenerated using 10 mM glycine, pH 1.5 for 30 s at 50 µl min−1 after each cycle. The Biacore T200 evaluation software (v 3.2.1., GE Healthcare) was used to calculate steady-state affinity constants (KD).
CD4 T cell depletion
FcγR humanized mice (males or females; 6–12 weeks old, described in ref. 77) were injected i.v. (10 µg) with afucosylated, monofucosylated, or bifucosylated IgG1 variants of the mouse-human chimeric anti-mouse CD4 mAb (clone YTS191). Mice were bled one day pre-injection and 24-h post mAb injection. Red blood cells were lysed with RBC lysis buffer (Biolegend) and blood samples were incubated with the following mixture of antibodies (1:250 dilution): FITC anti-mouse CD4 (clone RM4-4; Biolegend), BV510 anti-mouse/human CD11b (clone M1/70; Biolegend), BV421 anti-mouse CD3 (clone 145-2C11; Biolegend), BV605 anti-mouse CD8α (clone 53-6.7; Biolegend), APC anti-mouse CD45 (clone QA17A26; Biolegend). Samples were analyzed using the Attune NxT flow cytometer (ThermoFisher) with Attune NxT software v.3.1.2. Flow cytometry data were analyzed using FlowJo (v. 10.10.0) software.
In vivo antibody-dependent enhancement of dengue disease
DENV virus preparation and infection was completed as previously described78. Experiments were conducted in the Comparative Bioscience Center at Rockefeller University within animal biosafety level 2 (ABSL-2) containment in compliance with institutional and federal guidelines. Ifnar1−/−/FcγR humanized mice (male and female, 4–5 weeks old, described in ref. 78) were treated i.v. (20 μg) with the anti-DENV E protein mAb C10 (expressed as afucosylated, monofucosylated, or bifucosylated variants or as the G236R/L328R variant). 6 h post mAb administration, mice were challenged i.v. with DENV2 (New Guinea C strain, 4.4 × 105 IU, i.v.; 150 μl). Survival and weight loss were monitored and recorded for 7 days post infection. Death was defined as a 20% body weight loss threshold as authorized by the Rockefeller University Institutional Animal care and Use Committee.
Knob-in-hole IgG1 YTS191, C10, or Rituximab IgG1 expression and purification
A T366W mutation was introduced into the heavy chain of YTS191 and C10 IgG1. Separately, T366S/L368A/Y407V/A330W mutations were also introduced into the heavy chain of YTS191 and C10.
A T366W mutation alone or in combination with E269M was introduced into the Rituximab heavy chain. Separately, T366S/L368A/Y407V/A330W mutations alone or in combination with E269I were also introduced into the heavy chain of Rituximab.
The primers for mutagenesis were designed using the NEBaseChanger tool (https://nebasechanger.neb.com/) and PCR was carried out using the NEB Q5 High-Fidelity DNA Polymerase and the manufacturer’s instructions. All sequences were confirmed by Sanger sequencing at Genewiz (https://www.genewiz.com/).
For YTS191 and C10 KiH A330W expression: T366W heavy chain, T366S/L368A/Y407V/A330W heavy chain, and the appropriate light chain were transfected in a 1:1:1 ratio by mass in Expi293F cells.
For Rituximab KiH A330W E269I: T366W heavy chain, T366S/L368A/Y407V/A330W/E269I heavy chain, and the Rituximab light chain were transfected in a 1:1:1 ratio by mass in Expi293F cells.
For Rituximab KiH A330W E269M: T366W/E269M heavy chain, T366S/L368A/Y407V/A330W/heavy chain, and the Rituximab light chain were transfected in a 1:1:1 ratio by mass in Expi293F cells.
The plasmids were transfected as per manufacturers protocol (MAN0007814, Thermo Fisher Scientific) with the addition of Penicillin/Streptomycin mix 24 h post transfection. The cells were cultured for 96 h before harvesting. When making afucosylated IgGs, 250 μM of 2-deoxy-2-fluoro L-Fucose (Cayman Chemical) was added to the cultures.
The proteins were purified using Protein A resin (Thermo Fisher) with PBS pH 7.4 used as the binding buffer and 100 mM sodium citrate buffer pH 3.0 as the elution buffer. The fractions were neutralized with 1 M Tris pH 9.3. The proteins were buffer exchanged into PBS and stored at 4 °C until ready for use.
Engineering bifucosylated, afucosylated, and monofucosylated IgG1s
For the engineering of agalactosylated bifucosylated and afucosylated YTS191, C10, and Rituximab IgG1s: The bifucosylated or afucosylated IgG1s were purified as described above, an excess of the exogalactosidase BgaA was added, and then repurified with protein A, flash frozen, and stored at −80 °C until ready for use.
For the engineering of agalactosylated monofucosylated YTS191, C10, and Rituximab IgG1s: Bifucosylated IgG1 was purified as described above and treated with CU43 (1:1000 enzyme to antibody ratio by molar concentration) for 30 min at 37 °C. Then, the resulting monoglycosylated bifucosylated antibody was purified with protein A resin. Then, the exofucosidase AlfC was added in excess overnight at room temperature. The next day, transglycosylation reactions were carried out at 30 °C by adding 54 µl of 69 µM of the monoglycosylated monofucosylated antibody, 3 µl of 50 mg/mL oxazoline-linked S2G2, and 5.36 µg of EndoS2D184M. The reaction was dosed with 3 µl of ox-S2G2 every 30 min until completion as determined by continuous monitoring with intact LC/MS. The resulting diglycosylated monofucosylated antibody was then purified using Protein A. Finally, an excess of the exogalactosidase BgaA43 and the exosialidase MvNA42 were added to the remodeled antibody and incubated overnight at room temperature. The resulting agalactosylated monofucosylated antibody was purified once more with protein A, flash frozen, and stored at −80 °C until ready for use.
Ethical statement
Fresh human leukopaks were obtained from the New York Blood Center. The New York Blood Center receives written informed consent from donors upon receiving their donation, and this comes with the disclosure of potential use in research.
Pooled human plasma and serum were purchased from Innovative Research. Whole blood is collected from donors in an FDA-approved collection center.
Ethical approval of the COVID study was obtained from the Emory University Institutional Review Board under STUDY00001082. Written informed consent was obtained from all participants before inclusion in the study.
Ethical approval for the Dengue study was obtained from the National Ethics Committee of Health Research of Cambodia (nr 2018-310). Written informed consent was obtained from all participants or the guardians of participants under 16 years of age before inclusion in the study.
Animal care and experimentation were approved by the Rockefeller University Animal Care and Use Committee (23018-H) and consistent with federal laws.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper. All intact LC/MS data have been deposited to the ProteomeXchange Consortium via the PRIDE79 partner repository with the dataset identifier PXD070591. Source data are provided with this paper.
References
Breitling, J. & Aebi, M. N-linked protein glycosylation in the endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 5, a013359 (2013).
Reily, C., Stewart, T. J., Renfrow, M. B. & Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 15, 346–366 (2019).
Wang, T. T. IgG Fc glycosylation in human immunity. Curr. Top. Microbiol. Immunol. 423, 63–75 (2019).
Dekkers, G. et al. Decoding the human immunoglobulin G-glycan repertoire reveals a spectrum of Fc-receptor- and complement-mediated-effector activities. Front. Immunol. 8, 877 (2017).
Beyze, A., Larroque, C. & Le Quintrec, M. The role of antibody glycosylation in autoimmune and alloimmune kidney diseases. Nat. Rev. Nephrol. 20, 672–689 (2024).
Shields, R. L. et al. Lack of fucose on human IgG1 N-linked oligosaccharide improves binding to human Fcgamma RIII and antibody-dependent cellular toxicity. J. Biol. Chem. 277, 26733–26740 (2002).
Zou, G., Ochiai, H., Huang, W., Yang, Q., Li, C. & Wang, L. X. Chemoenzymatic synthesis and Fcgamma receptor binding of homogeneous glycoforms of antibody Fc domain. Presence of a bisecting sugar moiety enhances the affinity of Fc to FcgammaIIIa receptor. J. Am. Chem. Soc. 133, 18975–18991 (2011).
Vattepu, R., Sneed, S. L. & Anthony, R. M. Sialylation as an important regulator of antibody function. Front. Immunol. 13, 818736 (2022).
Dekkers, G., Rispens, T. & Vidarsson, G. Novel concepts of altered immunoglobulin G galactosylation in autoimmune diseases. Front. Immunol. 9, 553 (2018).
Smith, K. G. & Clatworthy, M. R. FcgammaRIIB in autoimmunity and infection: evolutionary and therapeutic implications. Nat. Rev. Immunol. 10, 328–343 (2010).
Wang, T. T. et al. Anti-HA glycoforms drive B cell affinity selection and determine influenza vaccine efficacy. Cell 162, 160–169 (2015).
Peschke, B., Keller, C. W., Weber, P., Quast, I. & Lunemann, J. D. Fc-Galactosylation of human immunoglobulin gamma isotypes improves C1q binding and enhances complement-dependent cytotoxicity. Front. Immunol. 8, 646 (2017).
Malhotra, R., Wormald, M. R., Rudd, P. M., Fischer, P. B., Dwek, R. A. & Sim, R. B. Glycosylation changes of IgG associated with rheumatoid arthritis can activate complement via the mannose-binding protein. Nat. Med. 1, 237–243 (1995).
Karsten, C. M. et al. Anti-inflammatory activity of IgG1 mediated by Fc galactosylation and association of FcgammaRIIB and dectin-1. Nat. Med. 18, 1401–1406 (2012).
Markina Y. V. et al. Sialylated immunoglobulins for the treatment of immuno-inflammatory diseases. Int. J. Mol. Sci. 21, 5472 (2020).
Li, D., Lou, Y., Zhang, Y., Liu, S., Li, J. & Tao, J. Sialylated immunoglobulin G: a promising diagnostic and therapeutic strategy for autoimmune diseases. Theranostics 11, 5430–5446 (2021).
Pagan, J. D., Kitaoka, M. & Anthony, R. M. Engineered sialylation of pathogenic antibodies in vivo attenuates autoimmune disease. Cell 172, 564–577 e513 (2018).
Irvine, E. B. & Alter, G. Understanding the role of antibody glycosylation through the lens of severe viral and bacterial diseases. Glycobiology 30, 241–253 (2020).
Haslund-Gourley B. S., Wigdahl B. & Comunale M. A. IgG N-glycan signatures as potential diagnostic and prognostic biomarkers. Diagnostics 13, 1016 (2023).
Ercan, A. et al. Aberrant IgG galactosylation precedes disease onset, correlates with disease activity, and is prevalent in autoantibodies in rheumatoid arthritis. Arthritis Rheum. 62, 2239–2248 (2010).
Schwab, I. & Nimmerjahn, F. Role of sialylation in the anti-inflammatory activity of intravenous immunoglobulin - F(ab’)(2) versus Fc sialylation. Clin. Exp. Immunol. 178, 97–99 (2014).
Chakraborty, S. et al. Proinflammatory IgG Fc structures in patients with severe COVID-19. Nat. Immunol. 22, 67–73 (2021).
Chakraborty, S. et al. Early non-neutralizing, afucosylated antibody responses are associated with COVID-19 severity. Sci. Transl. Med. 14, eabm7853 (2022).
Bournazos, S. et al. Antibody fucosylation predicts disease severity in secondary dengue infection. Science 372, 1102–1105 (2021).
Pereira, N. A., Chan, K. F., Lin, P. C. & Song, Z. The “less-is-more” in therapeutic antibodies: afucosylated anti-cancer antibodies with enhanced antibody-dependent cellular cytotoxicity. mAbs 10, 693–711 (2018).
Stockmann, H., Adamczyk, B., Hayes, J. & Rudd, P. M. Automated, high-throughput IgG-antibody glycoprofiling platform. Anal. Chem. 85, 8841–8849 (2013).
Chen, X. & Flynn, G. C. Analysis of N-glycans from recombinant immunoglobulin G by on-line reversed-phase high-performance liquid chromatography/mass spectrometry. Anal. Biochem. 370, 147–161 (2007).
Cao, L. et al. Global site-specific analysis of glycoprotein N-glycan processing. Nat. Protoc. 13, 1196–1212 (2018).
Helali, Y. & Delporte, C. Updates of the current strategies of labeling for N-glycan analysis. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 1237, 124068 (2024).
Senini, I. et al. Direct glycosylation analysis of intact monoclonal antibodies combining ESI MS of glycoforms and MALDI-in source decay MS of glycan fragments. Commun. Chem. 7, 203 (2024).
Masuda, K., Yamaguchi, Y., Kato, K., Takahashi, N., Shimada, I. & Arata, Y. Pairing of oligosaccharides in the Fc region of immunoglobulin G. FEBS Lett. 473, 349–357 (2000).
Liu, Z. et al. Asymmetrical Fc engineering greatly enhances antibody-dependent cellular cytotoxicity (ADCC) effector function and stability of the modified antibodies. J. Biol. Chem. 289, 3571–3590 (2014).
Wang, A. C. & Wang, I. Y. Cleavage sites of human IgG1 immunoglobulin by papain. Immunochemistry 14, 197–200 (1977).
Pucic, M. et al. High throughput isolation and glycosylation analysis of IgG-variability and heritability of the IgG glycome in three isolated human populations. Mol. Cell Proteom. 10, M111 010090 (2011).
Shkunnikova, S., Mijakovac, A., Sironic, L., Hanic, M., Lauc, G. & Kavur, M. M. IgG glycans in health and disease: Prediction, intervention, prognosis, and therapy. Biotechnol. Adv. 67, 108169 (2023).
Mesonzhnik N., Belushenko A., Novikova P., Kukharenko A. & Afonin M. Enhanced N-glycan profiling of therapeutic monoclonal antibodies through the application of upper-hinge middle-up level LC-HRMS analysis. Antibodies 13, 66 (2024).
Klontz, E. H. et al. Molecular basis of broad spectrum N-glycan specificity and processing of therapeutic IgG monoclonal antibodies by endoglycosidase S2. ACS Cent. Sci. 5, 524–538 (2019).
Jefferis, R. & Lefranc, M. P. Human immunoglobulin allotypes: possible implications for immunogenicity. mAbs 1, 332–338 (2009).
Lefranc, M. P. & Lefranc, G. Human Gm, Km, and Am allotypes and their molecular characterization: a remarkable demonstration of polymorphism. Methods Mol. Biol. 882, 635–680 (2012).
Goetze, A. M., Zhang, Z., Liu, L., Jacobsen, F. W. & Flynn, G. C. Rapid LC-MS screening for IgG Fc modifications and allelic variants in blood. Mol. Immunol. 49, 338–352 (2011).
Gadgil, H. S., Pipes, G. D., Dillon, T. M., Treuheit, M. J. & Bondarenko, P. V. Improving mass accuracy of high performance liquid chromatography/electrospray ionization time-of-flight mass spectrometry of intact antibodies. J. Am. Soc. Mass Spectrom. 17, 867–872 (2006).
Cheng, L. L., Shidmoossavee, F. S. & Bennet, A. J. Neuraminidase substrate promiscuity permits a mutant Micromonospora viridifaciens enzyme to synthesize artificial carbohydrates. Biochemistry 53, 3982–3989 (2014).
Singh, A. K. et al. Unravelling the multiple functions of the architecturally intricate Streptococcus pneumoniae beta-galactosidase, BgaA. PLoS Pathog. 10, e1004364 (2014).
Smith, M. R. Rituximab (monoclonal anti-CD20 antibody): mechanisms of action and resistance. Oncogene 22, 7359–7368 (2003).
Lefranc M.-P. Chapter 26 - IMGT® Immunoglobulin repertoire analysis and antibody humanization. in Molecular Biology of B Cells 2nd edn (eds Alt F. W., Honjo T., Radbruch A. & Reth M.) (Academic Press, 2015).
Gupta, A., Smith, P., Bournazos, S. & Ravetch, J. V. A novel mouse strain optimized for chronic human antibody administration. Proc. Natl. Acad. Sci. USA 119, e2123002119 (2022).
Klontz, E. H. et al. Structure and dynamics of an alpha-fucosidase reveal a mechanism for highly efficient IgG transfucosylation. Nat. Commun. 11, 6204 (2020).
Gstottner, C. et al. Benchmarking glycoform-resolved affinity separation - mass spectrometry assays for studying FcgammaRIIIa binding. Front. Immunol. 15, 1347871 (2024).
Yamin, R. et al. Human FcgammaRIIIa activation on splenic macrophages drives dengue pathogenesis in mice. Nat. Microbiol 8, 1468–1479 (2023).
Merchant, A. M. et al. An efficient route to human bispecific IgG. Nat. Biotechnol. 16, 677–681 (1998).
Sastre, D. E. et al. The mechanistic basis for interprotomer deglycosylation of antibodies by corynebacterial IgG-specific endoglycosidases. Nat. Commun. 16, 6147 (2025).
Sastre, D. E. et al. Potent efficacy of an IgG-specific endoglycosidase against IgG-mediated pathologies. Cell 187, 6994–7007 e6912 (2024).
Li, T., Tong, X., Yang, Q., Giddens, J. P. & Wang, L. X. Glycosynthase mutants of endoglycosidase S2 show potent transglycosylation activity and remarkably relaxed substrate specificity for antibody glycosylation remodeling. J. Biol. Chem. 291, 16508–16518 (2016).
Dekkers, G. et al. Multi-level glyco-engineering techniques to generate IgG with defined Fc-glycans. Sci. Rep. 6, 36964 (2016).
Bruhns, P. et al. Specificity and affinity of human Fcgamma receptors and their polymorphic variants for human IgG subclasses. Blood 113, 3716–3725 (2009).
Mizushima, T. et al. Structural basis for improved efficacy of therapeutic antibodies on defucosylation of their Fc glycans. Genes Cells 16, 1071–1080 (2011).
Sakae, Y. et al. Conformational effects of N-glycan core fucosylation of immunoglobulin G Fc region on its interaction with Fcgamma receptor IIIa. Sci. Rep. 7, 13780 (2017).
Falconer, D. J., Subedi, G. P., Marcella, A. M. & Barb, A. W. Antibody fucosylation lowers the FcgammaRIIIa/CD16a affinity by limiting the conformations sampled by the N162-glycan. ACS Chem. Biol. 13, 2179–2189 (2018).
Keith, A. D. et al. Mapping affinity and allostery in human IgG antibody Fc region-Fc gamma receptor interactions. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2025.03.28.645945v1 (2025).
Radovani, B. & Nimmerjahn, F. IgG glycosylation: biomarker, functional modulator, and structural component. J. Immunol. 213, 1573–1584 (2024).
Lu, L. L., Suscovich, T. J., Fortune, S. M. & Alter, G. Beyond binding: antibody effector functions in infectious diseases. Nat. Rev. Immunol. 18, 46–61 (2018).
Weiner, G. J. Building better monoclonal antibody-based therapeutics. Nat. Rev. Cancer 15, 361–370 (2015).
Wang, L. X., Tong, X., Li, C., Giddens, J. P. & Li, T. Glycoengineering of antibodies for modulating functions. Annu Rev. Biochem 88, 433–459 (2019).
Ferrara, C. et al. Unique carbohydrate-carbohydrate interactions are required for high affinity binding between FcgammaRIII and antibodies lacking core fucose. Proc. Natl. Acad. Sci. USA 108, 12669–12674 (2011).
Freimoser-Grundschober, A. et al. FcgammaRIIIa chromatography to enrich a-fucosylated glycoforms and assess the potency of glycoengineered therapeutic antibodies. J. Chromatogr. A 1610, 460554 (2020).
Lippold, S. et al. Fc gamma receptor IIIb binding of individual antibody proteoforms resolved by affinity chromatography-mass spectrometry. mAbs 13, 1982847 (2021).
Lippold, S. et al. Glycoform-resolved FcɣRIIIa affinity chromatography-mass spectrometry. mAbs 11, 1191–1196 (2019).
Patel, K. R., Nott, J. D. & Barb, A. W. primary human natural killer cells retain proinflammatory IgG1 at the cell surface and express CD16a glycoforms with donor-dependent variability. Mol. Cell Proteom. 18, 2178–2190 (2019).
Golay, J., Andrea, A. E. & Cattaneo, I. Role of Fc core fucosylation in the effector function of IgG1 antibodies. Front. Immunol. 13, 929895 (2022).
Schneider, S. & Zacharias, M. Atomic resolution model of the antibody Fc interaction with the complement C1q component. Mol. Immunol. 51, 66–72 (2012).
Temming, A. R. et al. Human DC-SIGN and CD23 do not interact with human IgG. Sci. Rep. 9, 9995 (2019).
Jones, A. T., Marino, A. E., Martynyuk, T., Bournazos, S., Ravetch, J. V. The anti-inflammatory activity of IgG is enhanced by co-engagement of type I and II Fc receptors. Science 390, eadv2927 (2025).
Kristic, J. & Lauc, G. The importance of IgG glycosylation-What did we learn after analyzing over 100,000 individuals. Immunol. Rev. 328, 143–170 (2024).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
World Health O. Dengue Haemorrhagic Fever: Diagnosis, Treatment, Prevention and Control. WHO, Geneva, (1997).
Ou, T. P. et al. Improved detection of dengue and Zika viruses using multiplex RT-qPCR assays. J. Virol. Methods 282, 113862 (2020).
Smith, P., DiLillo, D. J., Bournazos, S., Li, F. & Ravetch, J. V. Mouse model recapitulating human Fcγ receptor structural and functional diversity. Proc. Natl. Acad. Sci. USA 109, 6181–6186 (2012).
Yamin, R. et al. Human FcγRIIIa activation on splenic macrophages drives dengue pathogenesis in mice. Nat. Microbiol. 8, 1468–1479 (2023).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
We would like to thank the participating patients and doctors and nurses at Kantha Bopha Hospital, Phnom Penh, Cambodia, for patient enrollment and sample collection. We would like to thank Philippe Dussart and Sreymom Ken (Institut Pasteur du Cambodge) for their help in dengue diagnosis and sample characterization and Emily Lam and Ruben Peraza (Rockefeller University) for technical assistance and for maintaining and genotyping the mouse colonies used in this study. Dengue patient collection was funded by Wellcome Trust (208710/Z/17/Z). Research reported in this publication was supported in part by National Institute on Aging (NIA) grant R01AG085587 (to E.J.S., B.R.R., and J.J.D.), National Institute of Allergy and Infectious Diseases (NIAID) grants R21AI175869 (to E.J.S.), R01AI137276 (to S.B.), P01AI148102 (to B.D.), and 2U19AI144306 (to B.D.), and National Institute of Biomedical Imaging and Bioengineering (75N92019P00328, U54EB015408, and U54EB027690) as part of the Rapid Acceleration of Diagnostics (RADx) initiative. We acknowledge support from the Rockefeller University. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
Conceptualization: T.A., J.J.D., E.J.S. Methodology: T.A., S.B., H.G., T.A., D.E.S., F.Y.A., W.A.L., T.C., B.R.R., B.D., J.V.R., J.J.D., E.J.S. Investigation: T.A., S.B., H.G., M.W.F., E.G.N., V.D., T.J.B., A.M.R., D.E.S., J.J.D. Visualization: T.A., J.J.D., E.J.S. Funding acquisition: S.B., E.J.S., B.R.R., J.J.D., W.A.L., B.D. Supervision: E.J.S., J.V.R. Writing – original draft: T.A., E.J.S. Writing-review & editing: T.A., E.J.S., J.J.D., B.R.R., S.B., J.V.R., H.G., B.D.
Corresponding authors
Ethics declarations
Competing interests
T.A., D.E.S., J.J.D., and E.J.S. are inventors on a provisional patent application filed with the United States Patent and Trademark Office by Emory University relevant to the work in this manuscript. All other authors declare they have no competing interests.
Peer review
Peer review information
Nature Communications thanks Cristian Beccaria, Miles Holborough-Kerkvliet, and Ziv Shulman for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Azzam, T., Bournazos, S., Gunduz, H. et al. Asymmetrically glycosylated IgG1 antibodies are universal and drive human disease. Nat Commun 17, 383 (2026). https://doi.org/10.1038/s41467-025-67070-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67070-3