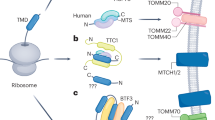
Abstract
The exact biological role of mitochondrial supercomplexes remains debated, particularly their role in guiding redox proteins across membranes during energy conversion. We integrate multiscale modeling and single particle cryo-electron microscopy (cryo-EM) to examine electron transfer in mitochondrial supercomplexes composed of complexes III and IV (CIII and CIV). Using bioinformatic and entropy-based methods, we generated structural ensembles capturing conformations of CIII’s disordered QCR6 hinge within the yeast CIII2CIV2 supercomplex. Molecular and Brownian Dynamics simulations reveal that these negatively charged hinge states electrostatically couple with redox proteins, promoting their binding and directional diffusion across the membrane on millisecond timescales. Rather than hindering transfer, disorder lowers the diffusion barrier. Anionic lipids reinforce this recognition by retaining a membrane pool of redox proteins when hinge length is critical. Cryo-EM models of ΔQCR6 show large rearrangements, yet maintain a robust electrostatic environment enabling surface-mediated transfer despite reduced charge. Overall, electron carriers confined on bioenergetic membranes follow a refolding-guided diffusion mechanism that enhances supercomplex energy conversion efficiency by nearly 30%.
Similar content being viewed by others
Introduction
Cellular bioenergetic pathways employ small diffusible redox proteins for the transport of single electrons between neighboring, as well as widely separated, integral-membrane enzymatic complexes. The movement of free electrons between solvated parts of the bioenergetic membranes is achieved only by special redox proteins, not by small molecules as the latter are converted through transfer of single electrons to chemical radicals that cannot be turned loose in the cell. Cytochrome c (cyt. c) is one such single electron carrier protein capable of protecting electrons from forming reactive chemical species. Cyt. c belongs to an exemplary family of redox proteins that participate in the mitochondrial electron transport chain, shuttling electrons, usually only over a short distance1, between the donor cytochrome bc1 complex (denoted respiratory Complex III or CIII) and acceptor cytochrome c oxidase (respiratory Complex IV or CIV). This electron current through the respiratory chain drives proton transport across the membrane (primarily via the respiratory Complex I or CI), generating an electrochemical gradient of ~200 meV that is ubiquitously employed to synthesize ATP and power all aerobic cells2.
The role of physical proximity of the charge donor and acceptor complexes in tuning the transport efficiency of the carrier proteins, and hence overall energy turnover of the cell has remained elusive over the past two decades now. Native gel electrophoresis together with cryo-tomography studies of bioenergetic membranes in mammals, yeast and plants revealed that the electron-transport chain components needed for ATP production are organized in a dynamic equilibrium between physically dispersed and closely packed states3,4. These clusters of the interacting respiratory complexes with variable stoichiometries are denoted as mitochondrial supercomplexes5. A general formula of the supercomplexes is written as CIxCIIIyCIVz, where x → 0, 1; y → 2; z → 1, 2. For the special case of x → 0, a supercomplex is reduced to CIIIyCIVz that we study here5,6. The fraction of CIII and CIV in the free vs. associated forms determines the redox state of the overall cyt. c pool, which is expected to tune the mechanisms of charge transfer in response to changes in environmental conditions and cellular redox-signaling pathways7,8. However, little is known about the detailed mechanism of such regulation.
Consequently, the biological relevance of supercomplexes has been questioned. A number of views have transpired. The generally agreed upon finding is that protein association reduces the probability of electron leakage, leading to a controlled formation of reactive oxygen species7,9. Also, the supercomplexes can avoid random aggregation of the proteins in the membrane10,11. However, the issue that has generated an exuberant discussion is whether the respiratory supercomplexes increase the efficiency of cellular energy conversion by reducing the physical distance between the individual complexes12,13. Central to addressing this puzzle is the definition of a supercomplex’s efficiency in terms of thermodynamic and kinetic parameters. Enhanced thermodynamic efficiency would entail an increase in the yield of ATP from the supercomplex-rich electron transport chain. But, despite several biophysical and biochemical studies, such enhanced ATP yields have not been observed3,14,15,16.
Alternatively, a combination of low-resolution electron microscopy data together with theoretical rate models have suggested kinetic advantages of supercomplex formation towards expediting cyt. c turnover between CIII and CIV if three-dimensional diffusion is the dominant mechanism of transport3,4,11,15,17,18,19. Even this interpretation becomes convoluted with the possibility of a two-dimensional cyt. c diffusion that can uniquely leverage the densely packed supercomplex environment4, and potentially compete with the three-dimensional diffusion pathway (Fig. 1A, B). In particular, the detailed molecular interactions underpinning the two- and three-dimensional diffusion of cyt. c are yet unresolved.
A Cyt. c (green) undergoes 3D diffusion where the protein, after visiting either complex III (purple) or complex IV (blue) (thick arrows), returns back to the bulk solution (thin arrows) before its next visit to either complexes87. B Cyt. c undergoes surface or 2D diffusion, where, as suggested in ref. 1, the protein stays in the vicinity of the supercomplex during its translocation between complex III (CIII) and IV (CIV). C Cyt. c can undergo guided diffusion, where the protein’s translocation is directed by the conformation of the disordered domain of QCR6 (orange), a hinge-like subunit of CIII. That is, the location for the localization of cyt. c, whether close to CIII or CIV, is directed by the folded state of QCR6 disordered domain. In figures below, the anionic mitochondria membrane is represented by our model membrane with anionic lipids (here, cardiolipin molecules) in red and charge-neutral lipids (here, POPC) in cyan. The dashed arrow in each figure indicates a QCR6-bound cyt. c can dissociate from QCR6 and interchanges with cyt. c in the bulk solution. Yet, the location of occupancy for this incoming cyt. c is still biased by the folded state of QCR6 to the location of the previously QCR6-bound cyt. c.
Two understudied molecular components that play a major role in the kinetics of diffusive cyt. c transport are (1) the anionic lipids e.g. cardiolipins in the membrane, and (2) the highly negatively charged QCR6 hinge of either an isolated or supercomplex-bound CIII. While the anionic lipids are already known to enhance the stability of mitochondrial complexes20,21,22,23,24, their organization within the yeast CIII2CIV2 supercomplex is revealed recently in our published cryo-EM models (PDB: 8E7S)25, and also in more recent models23,24,25,26. Cardiolipins offer a reconfigurable interaction network between individual complexes, which allow association of the complexes into supercomplex structures that may be required to regulate cyt. c dynamics in response to varying physiological conditions20. Hence, lower cardiolipin levels are associated with reduced formation of supercomplexes in a number of neurodegenerative diseases, ischemia followed by reperfusion, induction of apoptosis, heart failure, obesity and aging24,25,26,27.
QCR6, on the other hand, bears structural similarity to the twin CX9C proteins, having two canonical α-helices joined by only one disulfide bond into a ‘hinge’ shape. The N-terminal region of QCR6 is conformationally transient and has no reported structure thus far, even within the yeast supercomplex model at 3.2 Å resolution25; partial resolution is achieved with a more recent 2.4 Å model determined at low salinity28. It contributes to functional roles in preserving the heme environment of the cyt. c1 subunits of CIII and promotes interaction with cyt. c29,30. Consistent with this function, cells lacking QCR6 block maturation of the cyt. c1 subunit, attenuating CIII catalytic activity and growth29. Thus, despite not being a requisite for supercomplex formation, QCR6 directly contributes to the mechanism of cyt. c turnover. Recent findings and the identification of disease-associated mutations in specific CX9C motif-carrying proteins9 have highlighted members of this family of proteins as potential therapeutic targets in a variety of human disorders31.
Here, we seek to resolve the volume-dependent (3D) vs. surface-dependent (2D) mechanisms of cyt. c transport in the mitochondrial environment by modeling their hinge-mediated diffusion across the CIIIyCIVz-type supercomplexes under varying QCR6 composition and dynamics, anionic lipids concentration in the membrane, and association architectures. The approximately millisecond-scale computations offer a working model of how the supercomplex architecture tunes cyt. c movements between CIII to CIV from random 3D to 2D diffusion onto folding/unfolding-guided 2D diffusion (Fig. 1C). Integration of the simulated data into a master equation framework quantifies the kinetic advantage of energy conversion with surface-guided electron transport in a supercomplex environment over the volume-guided mechanism in freely dispersed complexes. We further investigate how supercomplexes reorganize the surface electrostatics potentials for charge-carrier recognition and transport even when QCR6 is deleted, hence maintaining a fairly robust environment for 2D diffusion dynamics to dominate biological energy conversion. Finally, our models offer insights on differences between conducting charge transport with soluble carriers in higher organisms over utilizing their membrane-anchored counterparts in more primitive bioenergetic membranes.
Results
In what follows, first our cryo-EM model of the CIII2CIV2 supercomplex is augmented with an ensemble refinement of the QCR6 structure. To this end, we employ a combination of cryo-electron microscopy (Cryo-EM) density maps with protein folding methods (Modeling Employing Limited Data or MELD), all atom and accelerated molecular dynamics (MD) simulations of the entire CIII2CIV2 construct32,33. Second, the dynamics of QCR6 is probed under different electrostatic environments as it undergoes conformational transitions to control the transport of cyt. c from bulk solution to CIII, and thereafter between CIII and CIV. Third, the interplay between supercomplex stoichiometry, membrane compositions and QCR6 dynamics is explored to determine the probability of 3D vs. 2D diffusion of cyt. c. This interplay between membrane interactions, and transient order and hinge-like movement of QCR6 is further interrogated with continuum electrostatics and Brownian Dynamics (BD) simulations34. Finally, in view of the existing structural, biochemical and computational data, a rate-kinetic model of soluble protein transport is proposed to outline advantages of energy transfer in confined spaces.
The QCR6 domain of yeast Complex III can exist in transiently folded conformations
There are no known structures of the complete QCR6 domain of CIII. It displays a diversity of lengths across species with yeast engendering one of the longest sequences (Supplementary Fig. 1)35. This unusual property of the yeast CIII prompted our ensemble refinement of the QCR6’s N-terminal hinge region, starting from the cryo-EM supercomplex model from Saccharomyces cerevisiae. The cryo-EM model of CIII2CIV2 used for this study was reported in the PDB: 8E7S, which is resolved at 3.2 Å and builds upon previously resolved CIII and supercomplexes (PDB: 6YMX; 1KB9; 3CX5; 6Q9E; 6GIQ). As in all the models, after purification, the yeast supercomplex has CIII bound to one or two CIV in a triangular trimeric or linear tetrameric shape, yet with comparable CIII-CIV interfaces. In the 8E7S model, these interfaces are found to be stabilized by anionic lipid molecules, with the density maps revealing ten distinct cardiolipin binding sites that were identified using POPG lipids.
To build a complete QCR6 structure within the supercomplex, first, a reduced cyt. c was docked on the p-side of the CIII2CIV2 supercomplex by leveraging existing X-ray crystallography data on the cyt. c-docked CIII structures (see Methods: Initial model preparation)36. Thereafter, the MELD-guided MD simulations (see Methods: MELD) revealed an ensemble of structures for the extended QCR6 N-terminal in the presence of the supercomplex-bound cyt. c (Supplementary Movie 1). Illustrated in Fig. 2A, the most probable folded state determined at room temperature, is composed of five helices with a total of 73 residues. Two of these helices are stabilized by positive charges on cyt. c, and are connected to the other three, which are proximal to the membrane via a 40-residue long loop.
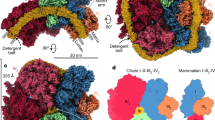
A An all-atom model of the yeast supercomplex (labeled SC) composed of electron donor CIII dimer (purple) associated with two copies of electron acceptor CIV (blue) and one carrier cyt. c (green) embedded in a POPC model membrane. B A starting model for the QCR6 N-terminal region is constructed by folding its first 73 residues using MELD-accelerated MD simulations in the presence of cyt. c and soluble residues from CIII that are within 14 Å of the cyt. c. Four snapshots (denoted states 0–3) representing the local interaction energy minima (see Supplementary Fig. 4B) along the protein folding simulation are illustrated, demonstrating the flexibility of the QCR6 hinge32. C (Left) Reconstruction of the supercomplex using non-uniform refinement shows strong density features (white map) located beneath the QCR6 (magenta chain). (Right) Supercomplex with an overlay of the disordered QCR6 hinge, encompassing models with an end-to-end distance of ~115 Å or shorter (see states between 2 and 3 in (B)) occupies a similar space in MELD-accelerated MD simulation snapshots as that observed in the refined density map. The impact of including our modeled and simulated QCR6 (with the acidic hinge region) to the partially resolved supercomplex was accessed by measuring the cross-correlation coefficients between the structure and 2 different cryo-EM maps. D Correlation between the supercomplex with/without QCR6 hinge region and the full cryo-EM map (shaded blue) was measured. The cross-correlation coefficients (ccc) with respect to the full cryo-EM map were computed over 270 sampled conformations of QCR6. This distribution is benchmarked against the ccc computed between SC without the QCR6 hinge domain (PDB: 8E7S)25 and the full cryo-EM map (vertical orange line). Because of the flexible nature of the QCR6 hinge region, a slight drop in correlation is observed upon including the QCR6 hinge region. E Correlation coefficient is recomputed with/without QCR6 hinge region and the fraction of cyro-EM map around the CIII-cyt. c binding interface (shaded red) is measured. Comparing against the ccc computed from PDB: 8E7S and the sourced cryo-EM map (vertical orange line), the inclusion of QCR6 hinge region noticeably improves the resulting ccc as the QCR6 hinge region fills the map region where not a single atom has been previously resolved (highlighted by a green rectangle). Source data are provided as a Source Data file.
To verify this predicted motif, we performed an Alphafold2 modeling of the QCR6 sequence both in the presence as well as absence of the bound cyt. c. In the absence of cyt. c there was minimal structure remaining in this highly negatively charged sequence, including the long disordered loop with –CE9C– residues. However, in the presence of cyt. c, the confidently resolved parts of the Alphafold2 solution, namely with a confidence threshold of pLDDT >90, were found to be within an RMSD of 2–6 Å of the converged MELD model (Supplementary Fig. 2). This analysis brings to light a transient nature of the QCR6 structure, particularly in the yeast-specific extended loop, which folds due to the presence of complementary cyt. c interfaces. Very recently, there is evidence of such cyt. c - induced QCR6 folding from cryo-EM studies28. Furthermore, kD estimates of the QCR6-updated CIII-cyt. c (determined from the PRODIGY server) is roughly 0.2 µM (Supplementary Fig. 3), very much in the range of values of ~0.3 µM determined from plasma resonance microscopy37. In the absence of QCR6 in our model, this value decreases to ~0.4 µM in our previous work, reinforcing the importance of this unresolved region in cyt. c binding to CIII38.
Going beyond the static Alphafold2 models, our MELD simulations further visit at least three partially folded low-energy conformations during the QCR6 folding (Fig. 2B). These intermediates elucidate the conformational space that QCR6 can thermally access at room temperature by varying its helical content (estimated in Supplementary Fig. 5). Within the folding intermediates seen by MELD, end-to-end distances of the extended QCR6 loop varies between 35 Å and 26 Å (denoted State 3), starting from a fully extended loop of length 175 Å (State 0, Fig. 2B). By comparing this stretch to the separation between the heme group of electron donor CIII and the copper group of electron acceptor CIV, which ranges up to 115 Å from the cryo-EM models of the supercomplex, we infer that a fully stretched QCR6 can potentially interact with CIV, and most certainly interact with the upper (p-side) leaflet of the mitochondrial membrane. Further processing of the cyt. c-free supercomplex data (EMD-27940) with a nonuniform refinement (Methods: Non-uniform refinement of WT supercomplex) brought to light delocalized unresolved density features grazing the surface of CIII and CIV, which cannot be interpreted by the structured subunits (Fig. 2C). Interestingly, a cross correlation analysis reveals that inclusion of the folded QCR6 N-terminal hinge from the MELD simulations improves match between the structural ensemble and the unresolved density features (Fig. 2D, E). Together with outcomes of the MELD simulation, now, we take this unresolved density for an ensemble representation of the dynamic QCR6 N-terminal.
Direct QCR6 interactions with anionic lipids regulate cyt. c movement between electron donor and acceptor complexes
The variance of QCR6 conformations cannot be accessed using brute-force MD, even with microsecond-long simulations. This is due to the longer timescales over which its folding-unfolding transitions and associated changes in the cyt. c structure occurs. Following an initial thermalization with 1 µs of conventional MD of the entire membrane-embedded CIII2CIV2 supercomplex, three copies of 500 ns long Steered MD (SMD) simulation were performed (see Methods: MD and SMD). Notably, the membrane is composed of POPC, and POPG lipids in the 8E7S model were replaced with cardiolipin (C18:2) molecules. During these SMD simulations, the biasing potential only acted on the distance between the center of mass of the alpha carbons of cyt. c and those of residues (TYR146, ASP150, GLU151, ASP162, ASP164) describing a putative cyt. c binding site on the CIV surface39. The QCR6 hinge (i.e. the flexible domain of QCR6) was, thus, capable of relaxing in any direction other than that of the steering. Such relaxation allowed the QCR6 to stretch and concomitantly interact with the membrane leaflet as well as with the cyt. c, which was translocating from CIII → CIV (Fig. 3). The membrane interactions of unstructured QCR6 (residue 1–73) facilitated its unfolding, marginally more in the presence of the cardiolipin headgroups40, which loses a few additional hydrogen bonds (Supplementary Fig. 6A - structures with 12 hydrogen bonds), while accompanying the cyt. c movement. In contrast, the structured part of QCR6, namely residues 74–147, gains hydrogen bonds (Supplementary Fig. 6B - structures with 12, 16 and 20 hydrogen bonds), and is therefore stabilized due to binding with the cardiolipin-containing membrane.
Structural snapshots from 500 ns of SMD simulation of cyt. c displacement from CIII → CIV in the absence (A) and presence (B) of cardiolipin. Movement of cyt. c on the surface of the supercomplex is accompanied with unfolding of the QCR6 hinge, which is exaggerated by the cardiolipin interactions with the QCR6 (Supplementary Movie 2). This unfolding event at different phases of the SMD is further verified using GaMD simulations (Supplementary Fig. 5). Hemes are shown in VdW representation inside CIII. C The basic residues (ARG104 and LYS106) on QCR6 hinge closest to the membrane surface form salt bridges with the anionic cardiolipins, while QCR6 N-terminal simultaneously aiding in cyt. c transport. D Favorable QCR6-CL interaction energies that emerge half way along the transport pathway, ~250 ns, and remain stable till the cyt. c reaches CIV. Source data are provided as a Source Data file.
Following translocation from CIII, binding of the cyt. c to the CuA pocket (or hotspot) of CIV was confirmed using dissociation constant (kD) computations of the cyt. c-CIV complex models resulting from SMD simulations (Supplementary Fig. 3). The computed affinities are ~0.1 μM in the range reported in the 0.01−1 μM range surface plasmon resonance measurements41,42. Such systematic underestimates of absolute protein affinities (closer to the 0.01 μM mark) have also been noted in the past43. Despite this discrepancy, the CIV binding energies are found to be at least two-fold stronger than those from cyt. c-CIII. This trend remains in line with the observation that cyt. c turnover at CIV (and not CIII where kD is almost twice weaker) offers one of the energy bottlenecks of electron transport1,19. Such stronger affinity of a reduced cyt. c for an oxidized CIV over oxidized CIII seen in our computations, further supports the forward directionality of electron flow in the ATP synthesis direction, and avoids backward leakage.
Remarkably, the cardiolipin headgroups in the membrane form an electrostatic interaction with the Arg104 and Lys106 residues of the folded QCR6 domain. The interaction saturates around −100 kcal/mol (Fig. 3). So, these interactions tether the QCR6 hinge to the membrane while the cyt. c approaches the CuA binding pocket of CIV. The SMD simulations were repeated in the absence of the cardiolipin (or CL) in the membrane. Now, the average work required to transport cyt. c from CIII to CIV increased by 25–30 kcal/mol (Supplementary Fig. 6C). Hence, the electrostatic coupling between the QCR6 and the anionic lipids of the membrane cuts down the cost of cyt. c transport across the supercomplex, for comparable displacement distance of this charge carrier. The CL-mediated membrane-QCR6 interactions add to the overall stability of the QCR6 to accompany cyt. c transport (Supplementary Fig. 6D). When the CL is removed, extension of the more flexible QCR6 (featuring a lesser number of hydrogen bonds) becomes more dissipative, and requires a larger energy to focus the cyt. c displacement precisely between CIII and CIV. A sequence alignment analysis further reveals that the ARG104 residues, which are found to interact with cardiolipin headgroups, are conserved across five different species, notwithstanding the heterogeneous length of the hinged region (Supplementary Fig. 1).
Conservation of the arginine residue (specifically ARG 104) further suggests that QCR6-assisted transport of cyt. c between the donor and the acceptor complexes can be ubiquitous to the electron transport chain, potentially beyond the yeast membrane. Also, the cardiolipin tails were already resolved in the supercomplex structure, highlighting the importance of protein-lipid interactions in offering stability13,22. Our simulation together with this structural information now suggests that the membrane does not just offer stability to the supercomplex, the anionic lipids directly interact with transient conformations of CIII’s disorder regions, namely the QCR6 loop, to energetically favor cyt. c transport across the supercomplex surface.
Surface-guided diffusion model for cyt. c transport
Gaussian accelerated MD (GaMD) simulations were performed to more exhaustively sample the secondary structures of the extended QCR6 during the cyt c transport from CIII to CIV in the anionic membrane. 400 ns of GaMD simulations, each starting from the SMD snapshots of Fig. 3 revealed that, in the absence of the cyt c, the QCR6 remains in a conformational pre-equilibrium between folded and unfolded states, revealing two distinct population of helicities (Supplementary Fig. 5). The helicity spontaneously switches anywhere between 60% and 80%. Such a dynamical pre-equilibrium, together with an overall high negative charge of QCR6’s N-terminal domain (−37 e), can underpin the disorder and radiation damage that has kept the piece of the protein unresolved via cryoEM. In contrast, in the presence of cyt. c, the QCR6 remains more tightly packed in the vicinity of CIII (population is skewed towards higher helicity bins). Indeed, very recent experimental evidence suggests a more tightly formed state of QCR6 in the presence of a bound cyt. c3. In the subsequent SMD snapshots, when the QCR6 approaches CIV accompanying cyt. c, GaMD further confirms the unwinding of the hinge. Starting with the helicity of 75–80% it systematically unfolds to an extended state with nearly 50% helicity, attaining its unfolded-like form seen in the pre-equilibrium in the absence of the cyt c. Altogether, the QCR6 can undergo order-disorder transition both in the presence and absence of the cyt c.
Almost 90% of the mitochondrial electron transfer proteins remain in supercomplex states within yeast, as opposed to only 35% of complexation propensity in mammals1,44,45. The longer QCR6 of the yeast CIII (Supplementary Fig. 1) offers a clear functional advantage in the densely packed supercomplex environments. A 2D movement of the charge carrier cyt. c was suggested based on low-resolution structural evidence1,4. Our model illustrates that such movement represents not just surface diffusion of the carrier, but explicit carrier-anionic lipid interactions mediated by the QCR6. Hence, the cardiolipins contribute on one hand to the stability of the supercomplexes, as highlighted in the cryo-EM structure, but more importantly, directly interact with transient conformations of CIII’s disordered regions to energetically favor cyt. c displacement across the supercomplex surface (Supplementary Movie 2). This interplay between protein-lipid and protein-protein interactions complements the utility of a relatively longer QCR6 in yeast supercomplex. The results, however, do not imply that the QCR6 enforces the cyt. c to act like a structural component of the supercomplex with any particular stoichiometry. Rather, the membrane interaction with the stretchable acidic region of QCR6 keeps the transporting cyt. c in the vicinity of both the donor and the acceptor, with exchanges of the cyt. c still possible between the surface and the bulk solvent (Supplementary Fig. 7).
Supercomplex recognition and surface localization of the cyt. c pool is reinforced by collective electrostatics of QCR6 and anionic lipids
It is now established that the supercomplex formation has minimal bearing on the rate-limiting step of the mitochondrial electron transfer chain1,4. Our results of Fig. 3 and Supplementary Fig. 3 are in line with such observation, in that we find the QCR6 in supercomplexes to only improve long-range interactions. Whereas, the rate-determining step associated with cyt. c or c2 binding is dominated by the entropy bottlenecks imposed via the short-range interactions, namely via a highly constrained half-ring distribution of complementary hydrophobic residues35,46. Such short-range interactions remain similar between the supercomplex and individually dispersed systems. The mechanism for molecular recognition, nonetheless, plays a major role to ensure that the cyt. c pool associates with the supercomplex and not unproductively interacts with the membrane surface, particularly the anionic lipids. So, we seek how QCR6-augmented and QCR6-removed (or ΔQCR6) supercomplex environments promote the feasibility of productive charge transfer.
Role of QCR6 electrostatics
Atomic Resolution BD simulations were performed to simulate the diffusive dynamics of cyt. c in the vicinity of the supercomplex at different stages of its CIII → CIV transport (Fig. 4). Starting models for the BD simulations were extracted from the energetically favorable conformations of the SMD simulation (see Table 1 and Methods: Brownian Dynamics (BD)). The BD simulations offer a broader range of cyt. c-bound CIII and CIV models than SMD, which reveals a unimodal distribution of affinities for the former, and a bimodal distribution of relatively loosely bound models for the later (Supplementary Fig. 3). This result is in line with the presence of separate multiple electron-transfer complexes in CIV, the former being weaker with a kD of 0.4–0.7 μM and the latter with a stronger kD of 0.1–0.2 μM, while only a single productive complex is observed in CIII41,42.
A (left) Primary binding sites for diffusive cyt. c associations on CIII, in the presence of cardiolipins. Residues of CIII are colored according to the colorbar on the right of the molecular images, where the scale indicates percentage occupancies. The percentage occupancy of a residue is the ratio of its number of cyt. c contacts (i.e. the number of events when cyt. c is within 6 Å of the residue) with the maximum number of cyt. c - CIII contacts (cyt. c is within 6 Å of all the CIII surface residues) observed during simulations. So, when the percentage occupancy of a residue is 1, that residue is the most frequent cyt. c - interacting residue throughout the molecular trajectory. B (left) Similar molecular representation highlighting binding to the CuA group, the cofactor in CIV for receiving electrons from cyt. c, which also serves as the reference for distance measurements here. A (right), B (right) Distributions of the separation from each cyt. c - interacting residue, highlighted in the corresponding molecular image, to the respective cofactors for electron transfer are plotted. The data shown here for each residue is weighted by its percentage occupancy, where a higher percentage occupancy contributes a higher weight (see Methods: Analyzes details). As the QCR6 hinge domain unfolds away from CIII to approach CIV (corresponding to SMD snapshots evolving from t = 0 ns to 480 ns), BD revealed a decrease in the diffusive cyt. c occupancy closer to the CIII interface (distance of cyt. c heme to the heme c1 of CIII, ≤14 Å, indicated by the yellow arrow), and growth in the vicinity of CIV (distance of cyt. c heme to the CuA group of CIV, ≤20 Å39, indicated by the yellow arrow). See Supplementary Movie 3 for this shift of diffusive cyt. c occupancy on CIII and CIV. Source data are provided as a Source Data file.
Illustrated in Fig. 5A–C and Supplementary Fig. 9, we find that the cyt. c grazing on the surface of the supercomplex via at least three different pathways. This is facilitated by the additional negative charge on the QCR6 N-terminal that complements the seven Lys residues on cyt. c surface. Consequently, the number of cyt. c’s associations/Å2 surface area of the supercomplex (Methods: Computations of binding propensity per unit area) is 2–3 fold more than that of the cyt. c-membrane association, leaving only ~10% of the cyt.c in the bulk (Fig. 5D, E) for exchanging with the surface-associated ones. The model therefore suggests that the cyt. c - QCR6 interactions are stronger than that of its membrane affinity. Hence, as the QCR6 unfolds and stretches towards CIV, a cloud of cyt. c (quantified as occupancy, Methods: per-residue percentage occupancy and percentage-occupancy weighted distributions) moves away from the hotspot residues of CIII, which are close to the protein’s cofactor (heme), towards the CuA group of CIV (Fig. 4). Consequently, cyt. c switches from being localized around CIII surface to being localized around CIV (Supplementary Fig. 10). This finding on surface localization of cyt. c is also in agreement with the kinetic experiments4,47, wherein exclusive 3D diffusion is excluded as the most probable mode of cyt. c transfer in the electron transport chain. In fact, the distribution of cyt. c poses that are amenable to electron transfer involving CIII and CIV follow the same trend (Supplementary Fig. 18) - the primary localization of cyt. c shifts from CIII to CIV. The stated directional bias in cyt. c transfer is affected when the anionic lipids are removed (Supplementary Fig. 11). Now, the hotspot residues for cyt. c occupancy is more evenly distributed between CIII and CIV, despite the presence of QCR6, creating ambiguity about the site of electron acceptance. So, the dependence of directional cyt.c occupancy on membrane composition alludes to a possible cooperativity between QCR6 activity and anionic lipids for sustaining the carrier protein’s surface accessibility, which will be further dissected below.
Cyt. c (green) diffusion follows multiple pathways during BD simulations: A diffusing entirely in the bulk solvent and arriving at the supercomplex; B diffusing to the membrane then to the supercomplex; and C encountering the supercomplex then translocating between CIV and CIII through subsequent diffusion over either the membrane or QCR6 or both. These different diffusion pathways are indicated in (A–C) by black arrows tracing snapshots of cyt. c (green, transparent) captured in corresponding BD simulations. Color-code for CIII (purple) and CIV (blue) are maintained, while POPC and cardiolipin are indicated in cyan and red. The number of cyt. c’s associations with the supercomplex and that with the cardiolipin-containing membrane, normalized to their respective solvent accessible surface area (SASA), are shown in (D) for the complete supercomplex model, namely the WT and is labeled QCR6 here, and for the one without QCR6s’ disordered hinge domains, labeled ΔQCR6. The fraction of cyt. c remaining in the bulk solution along the simulation time is given in the inset. The number of cyt. c associations with the supercomplex and that with the membrane for the supercomplex at different time points of the SMD simulation, as well as for a thermally averaged electrostatic maps from all of these QCR6 conformations called “Ensemble average electrostatic”, are shown in (E), where data for cyt. c - supercomplex associations are shown by black bars and those for cyt. c - membrane associations are given in white bars. A molecular representation of this cyt. c - membrane association average model is given in the inset (details at Supplementary Fig. 9). For measurements reported in (D) and (E), the corresponding standard deviations are shown here as error bars in the figures. The details in extracting these standard deviations from simulations are given at Methods: Error estimations for cyt. c associations with supercomplex (SC) and relevant observables. Source data are provided as a Source Data file.
The electron transfer rates computed from these poses suggest that the speed of transfer from CIII to cyt. c is commensurate with that of cyt. c to CIV (Supplementary Fig. 19), measuring up to 1 × 105 s−1, which corroborates48,49,50,51. A small bimodal distribution is observed for electron transfer between cyt. c and CIV, in line with multiple dissociation constants observed for the binding interface (Supplementary Fig. 3). Residues that may take part in the electron transfer pathway (Supplementary Fig. 20) are found to spatially overlap with the hotspot residues of Fig. 4 that stabilize the cyt. c binding interface with the supercomplex. The electron-transfer modulating residues are mostly located on CIV, including W124, Y125, Y146, and M232. Indeed, participation of protein residues to electron transfer involving CIV or cyt. c oxidase is not uncommon - the transfer from CuA to heme a then to heme a3 in CIV also involves protein residues52. Mutations on cyt. c - interacting residues in Paracoccus Denitrificans cyt. c oxidase further modulates the rate from cyt. c to CuA of CIV53. This does not imply that protein residues must engage in electron transfer, especially for cases such as transfer between CIII and cyt. c, where the edge-to-edge separation between the two heme groups can be as close as 9.4 Å to allow direct transfer49. Yet, it is worth mentioning that there are discussions suggesting a looser edge-to-edge separation of up to 11 Å to explain differences between structure-based calculations and experimentally measured rates49.
The ΔQCR6 reduces the CIII-CIV recognition surface of the cyt. c from 1.1 cyt.c/Å2 of surface in the wild-type supercomplex to 0.7 cyt.c/Å2. In ΔQCR6, marginally more cyt. c interacts with the membrane in the vicinity of the supercomplex (from 0.5 cyt.c/Å2 to 0.6 cyt.c/Å2 Fig. 5D), while very little change happens to the bulk cyt. c concentration (Fig. 5D-inset). Noting that the surface area of the membrane is 7.9-fold larger than the supercomplex model, even the minor increase in the probability is adequate to shift a significant population of the cyt. c towards the weakly interacting membranes. Also, cyt. c’s residence on the supercomplex at regions close to to the electron-transferring cofactors of CIII and CIV for forming productive complexes (Fig. 4) becomes shorter-lived in ΔQCR6 relative to WT (Supplementary Fig. 12A). Thus, productive cyt. c-supercomplex interaction is compromised in the ΔQCR6 models. On the contrary, cyt. c’ s residence time on other parts of the supercomplex as well as on the membrane are similar between ΔQCR6 and WT (Supplementary Fig. 12B, C).
Rate matrices determined from the BD simulation following partition of the space into four regions (CIII, CIV, membrane, and the bulk in Supplementary Fig. 8A) reveals that the transition of cyt. c from CIII to CIV occurs noticeably differently between ∆QCR6 and WT. ∆QCR6 is able to achieve an initial population transfer rate from bulk to the surface that is around 2-fold higher than that for WT. Yet, ∆QCR6 reduces the transfer of the cyt. c population between the donor and acceptor complexes, resulting in a lower equilibrium cyt. c population on CIV (Supplementary Fig. 8B). As a result, the effective transfer rate of cyt. c from CIII to CIV for ∆QCR6 (0.63 µs−1) is lesser than that of WT (0.96 µs−1) (Supplementary Fig. 8C). Altogether, the combined propensity of protein and membrane interactions is conserved between the WT and ΔQCR6 models, but the number of productive interactions goes down in the latter, highlighting the importance of QCR6 for charge carrier binding and transport.
Cooperativity of QCR6 and membrane electrostatics
Next we focus on the role of anionic lipids in cyt. c recognition of the supercomplex. In the presence of anionic cardiolipins, the binding propensity of cyt. c per unit area of the membrane, ~0.5 cyt.c/Å2, is only around 45% relative to that of the QCR6 binding propensity of 1.1 cyt.c/Å2 (Fig. 5D). So, for a given surface segment, cyt. c are much more localized around the supercomplex than being scattered over the membrane. Nonetheless, this relatively weaker cyt. c - membrane interaction still prevents the cyt. c pool from being lost into the bulk e.g. leaving only ~10% of the cyt. c in the bulk (Fig. 5D-inset). When the anionic lipids are removed, despite a robust rate of cyt. c approaching the surface from the bulk (Supplementary Fig. 14), the bulk cyt. c concentration increases to from 20 to 50% (Supplementary Fig. 15), dissipating the surface pool of carriers. Though the association of the remaining surface-bound cyt. c improves, 2.1 cyt.c/Å2 for the supercomplex and only 0.2 cyt.c/Å2 for the membrane (Supplementary Fig. 16) and have comparable residence times as in the anionic membrane (Supplementary Fig. 17), their occupancy at CIV sites is considerably reduced (Fig. 4vs. Supplementary Fig. 11), as most cyt. c molecules are in the bulk. Therefore, instead of competing with QCR6, the anionic cardiolipins complement the environment created by this hinge domain to enable cyt.c’s surface diffusion directed from CIII to CIV54.
An artificial system is created, denoted QCR6-trunc, where the sequence of the QCR6 domain is partially truncated from Met1 to Glu46, representing a length comparable to the four higher organisms (see sequence alignment of Supplementary Fig. 1). Here also, similar to ΔQCR6 and QCR6, we find that the addition of anionic lipids to the membrane improves recognition of the supercomplex by cyt. c on the supercomplex, even with the remaining QCR6 (Supplementary Fig. 17). Our comparative analysis of WT, ΔQCR6 and QCR6-trunc BD simulations verify that fields from anionic lipids and those from the heterogeneous QCR6 conformations mutually cooperate to keep the cyt. c pool in the vicinity of the supercomplex during electron transport. Consequently, QCR6 acts as a recognition element within the supercomplex for the cyt. c to identify the mitochondrial complexes within the crowded cellular milieu. Despite not modifying the mechanism of binding, supercomplexes enrich the cyt. c pool and increases the probability of productive binding.
QCR6 deletion potentially rigidifies supercomplex
Normally, supercomplexes remain in an equilibrium between CIII2CIV2 and CIII2CIV stoichiometries, both containing similar CIII-CIV interfaces (Fig. 6A, B)55. The QCR6 is not required for CIII–CIV supercomplex assembly, yet previous computational and cryo-EM studies have demonstrated its important role in cyt. c recognition and surface diffusion within the yeast respiratory chain. To investigate how the supercomplex compensates for the loss of this interaction element, we imaged and analyzed a ΔQCR6 variant using single particle cryo-EM (Supplementary Fig. 25). From ~81,000 particles, we obtained a reconstruction with a global resolution of 4.2 Å (see: Methods: Cryo-EM data/details for ΔQCR6, Supplementary Figs. 25A, 30). While this is lower than our previously reported wild-type structure at 3.2 Å resolution, local resolution analysis shows that the ΔQCR6 map contains better-resolved features in the CIV region when both datasets use approximately the same number of particles and are reconstructed to the same global resolution. This improvement in local resolution likely reflects a reduction in conformational heterogeneity at the CIII–CIV interface, consistent with the loss of the acidic, flexible N-terminal QCR6 hinge from both the CIII monomers. It is also possible that variable CIV occupancy contributed to differences in local resolution, as particles contributing to the ΔQCR6 map displayed more consistent CIV density than in the wild-type dataset. A summary concerning the data collection and processing processes for the cryo-EM density map of ΔQCR6 variant is provided at Table 2.
The electrostatic potential map of CIII2CIV2 and CIII2CIV1 complex in the WT and ∆QCR6 supercomplex, where electrostatic surfaces at a potential of 0.5kBT per charge and a potential of −0.5kBT per charge are given in blue and red, respectively. Starting from the wild type CIII2CIV2 (A), the hypothetical impact of structural changes from CIII2CIV2 → CIII2CIV1 on the electrostatic properties is examined stepwise (B) by first removing a single copy of CIV (C) then deleting QCR6 (D), followed by adjustment on the remaining CIV’s position based on the cryo-EM information in Supplementary Fig. 26. Visual inspection of the four iso-surfaces show negligible differences in the vicinity of CIII and CIV. The cumulative density function (CDF) for negative electrostatic potential values, denoted as felec, was determined (see Methods: Analyzes details). CDFs for structures in panels (B), (C), and (E) are denoted as felecB, felecC, and felecE, respectively. The pairwise difference between these distributions, presented in (F), show that deletion of QCR6 expectedly reduces the strength of the electronegativity as from (B) to (C). The reorganization of the CIII2CIV structure (E) makes the assembly more electronegative than the CIV-deleted CIII2CIV2 structure in (C). Source data are provided as a Source Data file.
To further probe the structural basis of this difference, we examined the electrostatic surface potentials and conformational states of the ΔQCR6 supercomplex. In the wild-type, QCR6 contributes to a highly electronegative p-side surface that promotes dynamic cyt. c interactions but may introduce electrostatic repulsion between subunits. Upon QCR6 deletion, this surface negativity is diminished (Fig. 6C), leading to a structural rearrangement in which the CIV shifts into a more compact, triangular CIII2CIV1 configuration with a reduced inter-complex distance of 66 Å, compared to 73 Å in the wild-type (Fig. 6D, E). Such alternate packing was examined by Molecular Dynamics Flexible Fitting (MDFF) simulations, where displacement of CIV was observed in conjugation to the switch of the cryo-EM density map for WT CIII2CIV to that for ΔQCR6-CIII2CIV during the structural model relaxation process (Supplementary Fig. 26). This architecture partially restores electrostatic continuity across the CIII–CIV interface (Fig. 6F), and likely contributes to the observed improvement in local resolution. Although the ΔQCR6 complex retains a viable cyt. c binding sites, electrostatic field strength and surface coverage are reduced relative to the QCR6-containing wild-type supercomplex, consistent with impaired recognition dynamics56. This reorganization highlights how structural rigidity can partially compensate for the loss of QCR6-mediated surface targeting.
To quantify the role of QCR6 in supercomplex function, a rate-kinetic model is used. For a given stoichiometry of the donor and acceptor membrane proteins undergoing electron-exchange via carrier proteins, we have derived that the rate of ATP/sec generated from a given input pool of energy is inversely proportional to the turnover rate of the slower process57 (See Method: Estimating the ATP production rate) - in the current case, the turnover rate of CIV. The rate matrix of Supplementary Fig. 8 implicates this process as the timescale of CIII to CIV cyt. c transfer, which drops from 0.96 µs−1 in the native to 0.63 µs−1 in the ΔQCR6 supercomplex. Noting that respiration is ~50% efficient and at any instant mitochondria generates 4 Watt of power (so it assimilates λ = 8 Watt), the 1.5-fold drop in CIV turnover translates to about 30% drop in the ATP production rate in ΔQCR6, which saturates to a drop of a maximum 40% even at unphysically energy intake58 (Fig. 7A). Conversely, at very low energy intake values e.g. in aging cells, the native and ΔQCR6 supercomplex shows comparable efficiency, offering no general advantage to the surface guided charge transfer mechanism in the clustered environment. Recently, it has been argued that within crowded cellular environments the salinity is much lower (20 mM) than the 150 mM determined from osmosis experiments59. This lower salinity is expected to reduce the screening effect and further increase the kinetic gap between the native and the ΔQCR6 constructs. The kinetic effect is expected to be dramatic if multiple cyt. c bind to the supercomplex (as we see in Supplementary Movie 3)28. Indeed, the cyt. c activity dropped by ~50% in ΔQCR629, leading to petite phenotypes. Our simulations now reveal a potential molecular mechanism for this kinetic loss stemming from imprecise targeting of cyt. c within the supercomplex environment.
A The rates of ATP production for supercomplexes with and without the QCR6 hinge region, denoted respectively as \({K}_{{ATP}}^{{QCR}6}\) and \({K}_{{ATP}}^{\varDelta {QCR}6}\) were computed across a range of power input rate to the system, here yeast (See Methods: Estimating the ATP production rate). The ratio, \({K}_{{ATP}}^{\varDelta {QCR}6}/{K}_{{ATP}}^{{QCR}6}\), shown here reveals the removal of the QCR6 hinge region induces a noticeable decline at the high power input domain. The range of input power (3.4 W to 10 W) for which a percentage drop of such removal starts to go beyond 25% (0.75 for \({K}_{{ATP}}^{\varDelta {QCR}6}/{K}_{{ATP}}^{{QCR}6}\)) to at least 30% (0.7 for \({K}_{{ATP}}^{\varDelta {QCR}6}/{K}_{{ATP}}^{{QCR}6}\)) is highlighted in orange. B Simulated density of cyt. c diffusing between CIII and CIV in BD simulations is shown as a contour surface of an occupancy map (isovalue = 0.5). For the same region, density of cyt. c observed by cryo-EM experiment and analyzed by 3-dimensional variational analysis (3DVA) is shown in inset. Panel B is adapted from Moe, et al. Cryo-EM structure and kinetics reveal electron transfer by 2D diffusion of cytochrome c in the yeast III-IV respiratory supercomplex, Proc. Natl. Acad. Sci. U.S.A. 118 (11) e2021157118, https://doi.org/10.1073/pnas.2021157118 (2021). A cross-correlation analysis for this simulated diffusion profile of cyt. c and our experimentally obtained cryo-EM density map for SC is available in Supplementary Fig. 24. In (C), ΔΔG scores for individual residues of QCR6, which indicate the respective capability to stabilize QCR6 upon being mutated to ALA, are combined with respective residues’ percentage occupancies (Supplementary Fig. 21) to give a composite score for each residue (D). The top 7 residues with the highest composite scores are highlighted in the molecular image of QCR6, with acidic residues in red, basic residues in blue, polar residues are in green, and hydrophobic residues in white. Source data are provided as a Source Data file.
A working model of cyt. c transport within the yeast CIII2CIV2 supercomplex
A general picture of diffusive electron transport within the supercomplex environment arises: first, following the accelerated MD results, the reduced CIII within the supercomplex remains in equilibrium between folded and unfolded QCR6 conformations. Second, according to the BD simulations, recognition and binding of an oxidized cyt. c to CIII populates its folded QCR6 state. Third, following from our past investigations and dissociation constant measurements, electron transfer from CIII into cyt. c weakens the interface. Fourth, it is inferred from the steered MD simulations that the reduced cyt. c is dislodged from CIII, and leveraging the electrostatic environment uniquely created by the anionic lipids and the extending QCR6 hinge within the supercomplex assembly, cyt. c is transported to the oxidized CIV. Finally, since residual interactions of the stretched QCR6 with the CIV-bound cyt. c is negligible to those of either CIII-bound or CIV-bound cyt. c, it is expected that the QCR6 hinge will disengage with cyt. c, returning to its pre-equilibrium prior to the latter’s release from CIV into the medium, resetting the entire cycle (Fig. 1). The CIV-bound cyt. c releases electrons for acceptance by the CuA site to get reoxidized. Release mechanism of the oxidized cyt. c from CIV has been studied in the past60. Beyond our model, alternate mechanisms of cyt. c binding includes binding to the membrane and losing its electron to lipids, likely cardiolipin, potentially inducing oxidative stress. And finally, interaction with other proteins, such as cytochrome c peroxidase, which sources electrons from our reduced cyt. c to catalyze the reduction of hydrogen peroxide to water60, a way to eliminate toxic radical molecules and mitigate oxidative stress.
An occupancy map of cyt. c on the surface of the supercomplex (Fig. 7B), created by averaging over all cyt. c’s BD poses that make contact with either CIII or CIV (Supplementary Movie 3), matches qualitatively with the mask of cyt. c movements created from three dimensional variational analysis of the cryo-EM density by Brizenski et al. (Fig. 7B -inset)1. This agreement between our simulated occupancy maps and the processed experimental data suggests that the grazing of cyt. c on the supercomplex seen in our model can be interpreted using a combination of surface diffusion as well as QCR6-induced conformational changes.
Finally, using Rosetta alanine scanning61, we propose a list of mutations on the QCR6 region that will reduce its interaction with the cyt. c proteins, and decrease the conformationally coupled movements, relegating the charge transport primarily to diffusion. We primarily targeted the interface between the N-terminal hinge and cyt. c and chose residues with the top six most positive binding energies (ΔΔG) (Fig. 7C and Supplementary Fig. 21), followed by weighting them with their respective frequency of interacting with cyt. c. BD simulations of a hypothetical joint mutant brought to light the coupling between QCR6 conformations and diffusivity of the cyt. c proteins. The mutations collectively reduced the negative charge and weakened the attractive field in the vicinity of QCR6 (Supplementary Fig. 22A). Consequently, now, the cyt. c can diffuse more in the bulk solution, moving further away from the supercomplex (Supplementary Fig. 22C, D) and with less localization around the binding interface of CIII and CIV.
Discussion
Capitalizing on cryo-EM structures, we report all-atom models and multiscale simulation of an entire respiratory supercomplex. The cumulative simulation time is 6 µs of MD and 14.08 ms (\(0.64\) ms × 22) of BD simulations (Table 1). On one hand these computations push the envelope of modern exascale hardware such as on the Summit and Frontier supercomputers to achieve massively parallel biophysical computations, while on the other, it offers detailed insights on unresolved parts of the supercomplex and how they potentially contribute to membrane-wide electron transport. We found an electrostatic cooperativity between the negatively charged and flexible QCR6 loop of respiratory CIII and the anionic lipids in the membrane, which facilitates recognition and surface-bound transport of the cyt. c carrier proteins. Instead of competing for binding to cyt. c, the electrostatic potential from these two components, namely transient protein order and membrane surface charge, mutually reinforce each other to keep the electron carriers in the vicinity of the membrane. This finding is strengthened by the fact that about 8-fold higher concentrations of complex IV from cardiolipin-free mitochondria had to be used to obtain rates comparable with those in wild-type mitochondria17.
Unfolding or folding dynamics of the QCR6, specifically in yeast, prompts charge transport towards or away from CIII, supporting a directional CIII → CIV electron transfer model. Removal of either one of these recognition elements i.e. anionic lipids or QCR6 from the membrane model, makes the transport of cyt. c more three-dimensional, compromises the number or residence time of productive binding to the electron donors or acceptor complexes, or reorganizes the supercomplex to recover for the loss in surface potential. Electron transfer from CIII to CIV by two-dimensional diffusion of cyt. c along the supercomplex surface resembles a “substrate channeling” model, which has been criticized based on the finding of that cyt. c diffusion in S. cerevisiae is unrestricted62. However, our QCR6-guided model of surface diffusion of cyt. c is not in conflict with this finding because it elucidates electrostatic interactions between cyt. c and the supercomplex surface, and cyt. c remains in equilibrium with the cyt. c pool during the entire electron-transfer process. So, when the QCR6 is removed, the ATP activity drops, but still saturates to 60–70% of the original; the bulk cyt. c though slow still shuttles the electron to enable this reduced activity.
In primitive organisms such as the heliobacteria, the cyt. c remain tethered to the membrane by employing covalent linkers of sequence presented in Supplementary Fig. 23. These linkers diffuse with the membrane, and keep the charge carriers in the vicinity of the membrane. Consequently, the need to create a locally negative charge environment is partially avoided for keeping the cyt. c confined. Though this is a common confinement method for Gram-positive bacteria, employing these covalent linkers is not required to confine cyt. c, but one of many methods to ensure cyt. c is in close proximity to its charge carriers. Other organisms may have their own methods of achieving this confinement, with QCR6 being the case in yeast. Supercomplexes are yet to be observed in these species. The N-terminal domain of the mitochondrial QCR6 unit has a highly dissimilar sequence with the covalent tethers. This suggests that confinement of the charge carrier on the membrane surface is ubiquitous within multiple organisms, but the mechanism has evolved. In the absence of QCR6, covalent linkages may be used to confine charge carriers. QCR6 improves the probability of cyt. c recognition either in isolated CIII or as part of the supercomplex. It potentially transports cyt. c by directly interacting with the membrane, as can be seen in the yeast membrane. When it is removed, the supercomplex reorganizes to still maintain an optimal environment for confining the carriers that can still exchange between the bulk and the surface. In our studies, the electrostatic confinement of charge carriers is relevant to yeast supercomplexes that preclude Complex I, and similarly for purple bacteria where electron transfer occurs between light-harvesting reaction center complexes and CIII57. Plants and mammals possess much shorter QCR6 domains, but it has been surmised that presence of Complex I together with anionic lipids stabilizes the supercomplex assembly63,64, which still creates a local electrostatic environment to keep cyt. c in the vicinity of the donor and acceptor integral-membrane complexes. Hence, the need for a longer yeast-like QCR6 is avoided in higher eukaryotes. Nonetheless, the positive residues (e.g. Arg 104 in yeast) in the more structured regions remain conserved even in the shorter QCR6 domains. So, silencing of QCR6 attenuated the translocation of cyt. c65.
The observed differences between wild-type and ΔQCR6 supercomplex structures further underscore the functional and architectural role of QCR6 in modulating respiratory complex assembly and dynamics. Although QCR6 is not essential for supercomplex formation, its highly acidic and flexible N-terminal domain introduces localized structural heterogeneity and dynamic electrostatic interactions that support surface confinement and directional transport of cyt. c54. In its absence, the ΔQCR6 supercomplex reorganizes into a more compact architecture with reduced inter-complex spacing and enhanced local resolution, particularly at the CIII–CIV interface. This structural rearrangement likely reflects a narrower conformational ensemble and reduced electrostatic repulsion across the p-side surface. While these features contribute to a more rigid and apparently well-defined structural state, they do not indicate an inherent increase in biochemical or thermodynamic stability. Rather, they represent a shift in the balance between dynamic recognition and structural coherence. Our electrostatic analyses show that QCR6 deletion diminishes the surface potential that facilitates cyt. c recruitment and retention, and simulations reveal shorter cyt. c residence times on productive electron transfer sites in ΔQCR6 compared to wild-type. Thus, the improved local resolution in ΔQCR6 arises not from a gain in function, but from the loss of dynamic flexibility that normally supports surface-guided diffusion.
Overall, the ΔQCR6 supercomplex compensates for the loss of a flexible recognition domain by adopting a more compact and rigid structure that partially preserves electrostatic targeting and cyt. c accessibility. However, the loss of surface flexibility leads to a measurable decrease in electron transfer kinetics and ATP production efficiency. Together, these findings suggest that QCR6 promotes both electrostatic recognition and structural adaptability, which may be co-optimized to enhance energy conversion efficiency in the dynamic mitochondrial environment.
Any modification of QCR6 compromises the catalytic activity of CIII, which is indeed the case. Also, if QCR6 is removed, supercomplex formation is a tangible way of receiving cyt. c and hence CIII activity. We have now simulated the mechanism of this recovery of activity by describing the reorganization of the electrostatic environment, reinforcing that confinement of the charge carrier is an evolutionary requirement. Taken together, there can be three mechanisms to ensure confinement of charge carriers: covalent linkers, isolated complexes or supercomplexes. QCR6 on one hand, acts as a linker for guided surface diffusion when participating in the yeast supercomplex without driving complex formation, while on the other hand, still maintains its functional properties seen in CIII acting as a recognition element.
Methods
The entire modeling workflow is summarized in Supplementary Fig. 27.
Initial model construction
Though the flexible domain of QCR6 has thus far been unable to be resolved using experimental methods, the sequence is known (Supplementary Fig. 1). Five predicted structures for the flexible domain of QCR6 were generated through QUARK66. Using VMD, these structures were aligned with the resolved region of QCR6 in a Complex III with reduced cytochrome c bound (PDB: 3CX5). However, when aligning the QCR6, there were steric clashes between the flexible domain and Complex III that were unable to be resolved through conventional molecular dynamics. So, after some minimization of the model, the following MELD simulations were performed.
MELD
The Modeling Employing Limited Data (MELD) approach is useful to integrate sources of ambiguous and noisy information about the molecular system. In particular, we were interested in modeling QCR6, which is a subunit of complex III (CIII), and its interaction with the remaining CIII in the presence of a CIII-bound cytochrome c (cyt. c).
QCR6 is a small subunit of 147 residues. Among these residues, residues 74–147 have a well defined structure and are experimentally resolved. In contrast, residues 1–73 constitute a highly acidic region that is highly flexible and remains unsolved in both the crystal and cryo-EM density map. This flexible linker region is most likely unstructured or intrinsically disordered and is thus well posed for MELD, which is capable of modeling such flexibility while incorporating information from the resolved region. It should be noted that this flexible linker region, though interacting with CIII domains embedded in the membrane, is found outside the membrane.
For computational efficiency, we first constructed an initial model with CIII (PDB: 8E7S) domains (without QCR6) that were within 8 Å of QCR6 (chain Q). In this model, we excluded other proteins in the CIII2CIV2 supercomplex with which QCR6 might interact (chain F or W). We capped incomplete subunits (chains) of our initial model with ACE and NME residues. Then, we generated preliminary QCR6 models by QUARK. As all QUARK models had residues 1–73 of QCR6 overlapping with our starting model, which contained CIII, we removed residues 1–73 from these quark models and modeled residues 1–73 as a non-overlapping, extended chain outside our starting model. Lastly, we attached this extended chain to our initial model with the resolved part of QCR6 (residues 74–147) at its C-termini. This model with an extended chain of QCR6 formed the final form of our initial model.
We next modeled the information to guide the simulations. We chose two lists of residues from our initial model. List 1 consisted of residues (236 318 320 237 331 330 304 481 184 223 220 286 287) from cyt. c and list 2 consisted of all residues in the flexible region of QCR6 (residues 1–73) (Supplementary Fig. 28). We combinatorially created all possible contacts between the two lists (collection). Within the collection, we grouped restraints such that each CIII residue (without QCR6) would have a contact (pairwise alpha carbon distances less than 8 Å) to any residue of the flexible region. We satisfied one restraint in each group, and overall we satisfied 60% of the 13 groups in the collection. We further used cartesian restrains on residues belonging to CIII. Other aspects of the simulation were as in previous MELD simulations32, using the ff14SB side force field, a 4.5 fs timestep, and with the H,T-REMD protocol. The H,T-REMD protocol used 24,550 ns-long replicas that expanded a temperature range of 300 K to 450 K with the data (QCR6-CIII contacts) enforced strongly at low replica index and with force constants vanished at the highest replica index.
MD and Steered MD (SMD)
From the MELD results, interaction energies between cyt. c and Complex III were calculated using the NAMD Energy Plugin in VMD. Four conformational states were identified, with the first being the unfolded flexible domain of QCR6 (state 0 in Fig. 2), the last fully folded (state 3), and two intermediate states (states 1 and 2). A model from state 3, which had the lowest interaction energy was used to manually incorporate into the full CIII2CIV2 supercomplex. This model was set up in two membranes in CHARMM-GUI67: one with just POPC as a reference, and one with POPC + Cardiolipin (CL) to mimic the supercomplex environment. In particular, the POPG lipids binding to the supercomplex in the original 8E7S model were replaced with CL. The system was parameterized with CHARMM36 force fields and CHARMM-GUI was also used to solvate and ionize both systems. The system was thermalized under NPT conditions and explicit solvent molecular dynamics simulations were done using NAMD 2.14 for 1 µs68.
After explicit solvent molecular dynamics, SMD was used to probe the dynamics of QCR6 as a hinge when cyt. c moves from CIII to CIV. A distance collective variable was used, where one group was defined as the center of mass of alpha carbons of cyt. c and the second group defined five residues around the CIV binding pocket (TYR146, ASP150, GLU151, ASP162, ASP164) and the force constant used was 0.05 kcal/mol. Though no specific residues were identified in previous studies, a predicted binding site on CIV was used29, where similar residues were used as targets for the binding site. Following SMD, interaction energies were calculated between cyt. c-CIII and cytochrome c-CIV, using the NAMD Energy plugin in VMD. Dissociation constants (Kd) were calculated using the PRODIGY web-server69.
Brownian dynamics (BD)
BD simulations had been used to sample cyt. c’s diffusion over the yeast CIII2CIV2 supercomplex through an in-house software, the Atomic-Resolution Brownian Dynamics (ARBD)70. The starting models are extracted at times 0 ns, 220 ns, 400 ns, and 480 ns from the SMD simulations, which provided local energy minima of the cyt. c-QCR6 interaction energy profile of Fig. 4. The underlying mechanism for and the setup of BD simulations using ARBD had been discussed in detail previously54,71,72. Here, following this previous approach, we will outline our setup of BD simulations below.
First, we classified our biomolecules into 2 categories, namely the diffusing particle and the diffusion environment. In this project, cyt. c is our diffusing particle and it was dynamic during BD simulations. Its atoms were clustered into 4 representative atom types as in ref. 54. The particle distribution for each of these representative atom types and the charge distribution for the whole cyt. c were determined by VMD73. Our cyt. c was then represented by these distributions, each stored as a volumetric data grid. On the other hand, each of the different membrane systems, with the supercomplex embedded, was our diffusion environment. It was static during BD simulations, and its electrostatic potential and its potential of mean force (PMF) for each of the representative atom types of cyt. c were computed by Adaptive Poisson–Boltzmann Solver74,75 and Implicit ligand sampling76, respectively. Then, to mimic the fluctuating nature of atoms’ positions, a 3D-Gaussian filter with a half width of 1.5 Å, which resembles the RMSF of atoms belonging to the protein-membrane interface, was convoluted with each of these potentials. Finally, during BD simulations of each membrane system, the different volumetric data grids for cyt. c interacted with either the electrostatic potential or the corresponding PMFs of the membrane system. This interaction was converted to forces on cyt. c, as being prescribed in ref. 54.
After the volumetric data grids for cyt. c as well as the electrostatic potentials and PMFs for each membrane were computed, BD simulations were launched. For each replica of BD simulation, the membrane midplane of the membrane system was located at z = 0, and one cyt. c was initially positioned above the supercomplex at a height of 100 Å, which corresponded to z ~ 150 Å. The x and y coordinates for our cyt. c were randomized.
For each membrane system, there were 320 replicas of 2-microseconds long BD simulations. In the current study, we employed 21 different membrane systems, each differed from another in terms of either the lipid composition of the membrane or the conformation of QCR6 or both. The total simulation time for BD simulations in the current study was 1.12 milliseconds.
Gaussian accelerated molecular dynamics (GaMD)
GaMD simulations77 were performed at 3 time points for QCR6 with and without cytochrome c. All GaMD simulations were performed using AMBER 20. The proteins were parameterized using AMBER ff14SB force field, solvated in a TIP3P water box of dimension 20 Å, and K+ ions were added for neutralization. This system was minimized, heated to 300 K under NVT conditions, and equilibrated at 300 K under NPT conditions for 2 ns. Following this system preparation, GaMD equilibration was performed for 20 ns, including 2 ns of conventional MD. GaMD production was performed for 500 ns.
Non-uniform refinement of WT supercomplex
As described in ref. 25, 493,055 particle images of the wildtype supercomplex were reconstructed using standard single particle cryo-EM image processing routines in CryoSparc (v3.2) resulting in a final density map of the supercomplex at 3.2 Å resolution78. Using this same dataset, 3D variability analysis was carried out to identify heterogeneity in this data set. Of the 10 modes identified in the 3D variability analysis, the five modes containing the largest number of particles, a total of 85,455 particles, were selected for a heterogenous refinement and reconstruction. This refinement generated 2 C2-symmetry reconstructions: class 0, containing 19,447 particles, reconstructed to 9.3 Å and class 1, containing 66,008 particles, reconstructed to 6.6 Å resolution. Particles in class 1 were further processed imposing C1 symmetry and using non-uniform refinement, which were then reconstructed to 4.3 Å resolution. The wildtype model from Hryc et al. (PDB: 8E7S) was then fit to this 4.3 Å resolution density map using Chimera (v1.16), from which unmodeled density could be observed near the termini of QCR6.
ΔQCR6 strain construction and purification
Mitochondria isolation and Supercomplex purification from ΔQCR6 strain were performed as described previously for the wild type strain25.
Mitochondria isolation
Cells were harvested at OD6002-3 by centrifugation (4700 × g for 10 min at 4 °C), washed with ice-cold 1X Tris-buffered saline. The pellet was resuspended in 100 mM TrisSO4 (pH 9.4) and 10 mM dithiothreitol (2 ml/g of cell pellet), incubated at 30 °C for 20 min with shaking, centrifuged at 750 × g at 4 °C for 10 min, washed with 1 M sorbitol. The cell wall was digested by zymolase-20T at 3 mg/g weight of cells in 1 M sorbitol and 20 mM Potassium phosphate buffer (pH 7.5) for 90 min at 37 °C. The spheroplasts were washed with 1 M Sorbitol (2 ml/g of cells) at 4 °C and disrupted with a Dounce homogenizer in the 10 mM Tris-HCl buffer (pH 7.2) with 0.5 M sorbitol, 0.02% bovine serum albumin, 1/1000 volume of protease inhibitor cocktail set III (Calbiochem Millipore) and 1 mM phenylmethylsulfonyl fluoride. The lysed cells were centrifuged at 1700 × g at 4 °C for 5 min, and the supernatant was centrifuged at 8500 × g at 4 °C for 20 min. The pellet was suspended in 1 ml of 10 mM Mops (pH 7.2) buffer with 250 mM sucrose and 1 mM EDTA and loaded onto a 15–60% sucrose gradient in 15 mM Tris-HCl (pH 7.4) and 20 mM KCl that was centrifuged at 111,000 × g at 4 °C for 90 min. The mitochondrial layer was aspirated, flash frozen with liquid nitrogen and stored at −80 °C.
Supercomplex purification
The Supercomplex from ΔQCR6 strain was purified by the method we previously developed for WT25. Isolated mitochondria (12 mg of protein) were suspended in the 1.5 ml of lysis buffer containing 2% (w/v) digitonin, 50 mM potassium acetate, 10% glycerol, 1:50 volume protease inhibitor cocktail, 1.5 mM phenylmethylsulfonyl fluoride and 30 mM HEPES-KOH, (pH 7.4) for 1 h at 4 °C with gentle shaking. After incubation the lysate was centrifuged at 4 °C for 20 min at 90,700 × g. Supernatant was incubated with magnetized Cobalt Beads (Dynabeads TALON catalog #101.02D, Invitrogen) for 45 min at 4 °C with shaking to remove FoF1-ATPase. After incubation the beads were removed using a magnetic separator. The supernatant (1.5 ml) was layered onto two equal 8 ml sucrose gradients (0.75 M to 1.5 M sucrose in 15 mM Tris-HCl (pH 7.2), 20 mM KCl, and 0.4% digitonin) and centrifuged at 4 °C for 22 h at 111,000 × g. Fractions (80–100 μL) from the gradients were analyzed by BN-PAGE and western Blot with antibodies to yeast CIII and CIV as described previously25. Fractions containing the purified SC were combined and used for cryo-EM. Protein concentrations were determined using the BCA protein assay kit (Thermo Scientific) according to manufacturer’s instructions.
Cryo-EM data/details for ΔQCR6
Data was collected with a Titan Krios microscope (Thermo Fisher Scientific) operated at 300 kV. A post-GIF K2 Summit direct electron detector (Gatan), operating in counting mode was used at a nominal magnification of 130,000× (pixel size of 1.07 Å) for image collection, and an energy slit with a width of 20 eV was used during data collection. A total dose of 56e−/Å2 fractionated over 35 frames; nominal defocus range set from −1.5 μm to −4 μm. Here, 7750 micrograph movies were collected of ΔQCR6.
The ΔQCR6 complex was processed with CryoSPARC78. Patch motion was used for frame alignment and exposure weighting with default parameters. Patch CTF was used to generate CTF parameters. Blob picker was initially used on a subset of images (200) to select a subset of particles, which were then used to generate a low-resolution template. This template was then used to train the Topaz particle picker79. A total of 369,297 particles were selected. Multiple rounds of 2D classification then followed, narrowing the dataset to 81,260 particles. An initial refinement was completed using C1 symmetry, which resulted in a 7.77 Å density map. A local refinement routine focusing on a CIII dimer and a CIV monomer then followed (using a mask to reduce the detergent band). This resulted in a 4.21 Å structure.
Analyses details
Computations of binding propensity per unit area
The per unit area binding propensity of cyt. c to a system A, labeled as \({p}_{A}\), is computed as,
Where \({Coun}{t}_{A}\) is the number of associations between cyt. c and A, and \({SAS}{A}_{A}\) is the corresponding solvent accessible surface area (SASA). In this work, A takes multiple forms, including membranes with various lipid compositions, the complete supercomplex, supercomplexes with different QCR6’s conformations, and mutants of supercomplex.
For each of the different systems (A) employed, \({Coun}{t}_{A}\) was sampled from BD simulations. An association between cyt. c and A was considered to occur if any heavy atom of cyt. c came within 6 Å of a heavy atom of A. It is worth recalling that a positional fluctuation of 1.5 Å (see Method: Brownian Dynamics (BD)) had been imposed in our BD simulations. This positional fluctuation extended the spatial regime in which a cyt. c could sense forces, including the steric repulsive force, from a static binding target A, subsequently rendering a cyt. c harder to come into extremely close proximity with A. Thus, to compensate for this phenomenon, a 6 Å cutoff was used in place of the smaller, commonly used 4 Å cutoff in analyzing contact formations in MD simulations. Similarly, for each system A, the corresponding \({SAS}{A}_{A}\) was computed using VMD73 through its internal measure sasa function with a 6 Å cutoff.
Per-residue percentage occupancy and percentage-occupancy weighted distributions
For each set of BD simulations, the number of events (\({N}_{i}\)) that a residue i makes contact with cyt. c was sampled. Assuming i is a member of domain I, where I can be CIII or CIV, the percentage occupancy (\({O}_{i}\)) of i is defined as,
Thus, \({O}_{i}\) measures how frequently i interacts (make contacts) with cyt. c with respect to the number of contacts achieved by the most frequent cyt. c - interacting residue from the same domain.
In Fig. 4, we are interested in the spatial distribution of cyt. c - interacting residues and their relevance to cyt. c - mediated electron transfer. To do so, for each cyt. c - interacting residue i in a domain I, we determined the minimal separation (\({d}_{i}\)) from i to the respective electron-transfer related cofactor. Then, we weighted the relevance of \({d}_{i}\) across the whole data set (\(D=\{{d}_{i}\}_{i\in I}\)) by the percentage occupancy of i, where a single data point \({d}_{i}\) is duplicated for \({O}_{i}\times {\max }_{i \in I} {N}_{i}\) times. After the weighting, a data point \({d}_{i}\) related to a residue i with a high \({O}_{i}\) will appear more frequently in D. This weighting scheme originates from the understanding that a residue i is spatially more relevant to electron transfer if it has both a small \({d}_{i}\) and a high \({O}_{i}\) simultaneously. The weighted data set D is then visualized in Fig. 4 to show how the populations of cyt. c - interacting residues changes across the unfolding of QCR6, as being characterized by their separations from electron-transfer linked cofactors, namely heme c of the cyt. c1 subunit of CIII and the CuA group of CIV.
Cumulative density function for voxel values recorded in an electrostatic potential (f elec)
The electrostatic potential for any system A employed in this study was computed using Adaptive Poisson Boltzmann Solver (APBS)71,72,73 with a resolution of 1 Å in each spatial dimension. The resulting electrostatic potential \({E}_{A}(x,y,z)\), where \(x,y,z\) span over the entire volume considered, was represented as a set of voxel values, \({E}_{1},{E}_{2},\cdots,{E}_{N}\) and was stored in the.dx file format. The index i of each of these values\({E}_{i}\), can be converted back to the \(x,y,z\) coordinates that it represents, and \(N\) is the total number of grid points employed for the calculation.
A system’s electrostatic field, once away from the source charges, decays quickly in a dielectric environment. For supercomplexes employed in our current study, spaces with electropositive potential below 0.05 kBT per charge are usually found outside the supercomplexes and are dispersed, behaving as fluctuations (Supplementary Fig. 29A). Spaces with the strength of electronegative potential below 0.05 kBT per charge are, on the other hand, space filling. Yet, these spaces are well separated from the surfaces of supercomplexes, with some of them located at a distance of around 40 Å away from the supercomplex’s surface (Supplementary Fig. 29B). Thus, voxel values lying between −0.05 kBT per charge and 0.05 kBT per charge primarily denote the non-atom-filling empty space involved in the electrostatic calculations for our systems. Also, it is worth noting that every atom in our systems is associated with a non-zero partial charge based on the Charmm36 force field80. These partial charges provide a quantum mechanical description for the classical behavior of atoms in molecular dynamics-like simulations and give non-zero electrostatic potential at the very close proximity of any atom. As a result, the number of voxel values lying between −0.05 kBT per charge and 0.05 kBT per charge is more likely to represent the amount of void spaces involved in our calculations rather than the extent of the electroneutral region over the system’s surface. Besides, a voxel value lying between −0.05 kBT per charge and 0.05 kBT per charge will only modify the energy of a unit charge by less than 0.05 kBT, which is 10% of the thermal energy associated with each degree of freedom for an atom. The impact of these voxel values on a charged object is thus relatively minor compared with the thermal fluctuations involved in Brownian dynamics. Therefore, with all these considerations, we discarded voxel values lying between −0.05 kBT per charge and 0.05 kBT per charge when computing the cumulative distribution for voxel values of an electrostatic potential (f elec).
After cleaning up our electrostatic data as mentioned above, we performed a histogram analysis on the remaining data. We computed the number of voxel values belonging to each bin of our histogram. The lower boundary value of a bin is −20 kBT per charge and the upper boundary value is 20 kBT per charge. The bin width is 0.01 kBT per charge. The resulting histogram is then normalized so that the value for any bin i (\({B}_{i}\)) with a maximum/boundary value of \({M}_{{B}_{i}}\) represents the probability (\(\Pr \{{E}_{j}\le {M}_{{B}_{i}}\}\)) to find a voxel value less than \({M}_{{B}_{i}}\). Then, following the definition of cumulative density function, we employ the following approximation,
to obtain the CDF for voxel values of the electrostatic potential for our system, where \({b}_{i}\) is the mid-point value of bin i, and \({b}_{i}\pm 0.005\) gives the upper and lower boundary respectively. Thus, \({M}_{{B}_{i}}={b}_{i}+0.005\).
Rate matrix computations and population transfer simulations
To compute the rate matrix for any particular cyt. c - supercomplex system, the number of occurrences for a cyt. c transfer from one cyt. c bound state to another was first sampled from the corresponding BD simulations. There are 8 cyt. c bound states: 1) c3m1 (cyt. c associated with CIII of monomer 1 of the supercomplex); 2) c3m2 (cyt. c associated with CIII of monomer 2 of the supercomplex); 3) c4m1 (cyt. c associated with CIV of monomer 1 of the supercomplex); 4) c4m2 (cyt. c associated with CIV of monomer 2 of the supercomplex); 5) qcm1 (cyt. c associated with QCR6 of monomer 1 of the supercomplex); 6) qcm2 (cyt. c associated with QCR6 of monomer 2 of the supercomplex); 7) Membrane (cyt. c associated with the membrane); and 8) Bulk (cyt. c remaining in the bulk). These sampled numbers were then laid out in a matrix format with the columns representing the current cyt. c bound state and the row representing the next cyt. c bound state. An example of such matrices, termed here as occurrence matrices to facilitate discussions, is given in Supplementary Fig. 8 for the WT.
To derive the rate matrix, modifications were applied to the corresponding occurrence matrix, which derivations are outlined above. For any occurrence matrix \(O\) with its elements \({O}_{{ij}}\), where \(i,{j}\) are indices for the row the column respectively, each matrix element \({R}_{{kl}}\) for the corresponding rate matrix \(R\) is given by
The change on the population of a bound state \(k\) at time \(t\), labeled as \({\rho }_{k}(t)\), over a time interval \(\Delta t\), which was used to sample the occurrence matrix, can be computed from the corresponding rate matrix as:
The above equation does not bear an analytical solution. Thus, we integrate the above equation numerically as,
The above integration was performed for 500 steps with a time interval of 20 ns, a value used in parsing the BD simulations into MD-like trajectories. As we want to focus the study on population transfer from CIII to CIV, an initial population condition where cyt. c is equally likely to be found across all different components of CIII was employed. Vector elements of this initial population condition are:
After obtaining \({\rho }_{k}(t)\) for all 8 different cyt. c bound states based on the above initial condition, the results were concatenated to highlight population changes for bound states related to CIII, CIV, the membrane, and the bulk solvent. The concatenation was done as:
The result of the above concatenation concerning population transfer starting from CIII to CIV is given in Supplementary Fig. 8.
Estimating the ATP production rate
For a power input of \(\lambda\), the rate of ATP output (\({K}_{{ATP}}\)) for a linear electron transport chain is approximately given by81:
where \(N\) is the number of electron acceptor complex per donor complex (which is 1 in our case), \(\tau\) is the time scale of electron turnover. Thus, at large values of \(\lambda\) and constant donor-stoichiometry, \({K}_{{ATP}}\,\propto {\tau }^{-1}\), implying \(\frac{{K}_{{ATP},{WT}}}{{K}_{{ATP},\,\Delta {QCR}6}}\propto \frac{{\tau }_{\Delta {QCR}6}}{{\tau }_{{WT}}}\), which we present in Fig. 7A.
The value of \(\tau\) was computed through its inverse \({\tau }^{-1}\), which was the electron turnover rate. Here, the values of \({\tau }^{-1}\) were estimated for the WT supercomplex and the ΔQCR6-mutant by their respective rates for electrons transfer from CIII to CIV by cyt. c, namely, 0.96 µs−1 and 0.63 µs−1, as being given in Supplementary Fig. 8. These estimations were made based on the fact that the subsequent electron transfer rate from cyt. c to CIV was at an order of 1 ×105 s−1 (1 ×1011µs−1), suggesting the electron transfer from CIII to CIV by cyt. c as a key contributor to the time spent over electron turnover50.
Error estimations for cyt. c associations with supercomplex (SC) and relevant observables
For each of the BD simulations performed, 320 replicas of cyt. c were simulated simultaneously. To estimate the error of any observable, such as the number of cyt. c’s associations with SC, the following steps were taken:
-
1).
First, we randomly grouped our 320 replica simulations into 16 groups of replicas, each containing 20 replicas. This grouping mimics a situation where a simulation consisting of 20 replicas was repeated 16 times;
-
2).
Second, the examined observable was computed for each group of replicas;
-
3).
Third, the standard deviation of the computed values was calculated;
-
4).
Lastly, the computed standard deviation was scaled up by 16 if the involved observable was measured as counts, for instance, the number of cyt. c’s associations with SC. If the involved observable was measured as a fraction, for instance, the fraction of cyt. c associating with SC, no scaling was performed for the computed standard deviation.
Estimations of electron transfer rate (ETR)
The ETR concerning ET from CIII to cyt. c and that from cyt. c to CIV were estimated following the below procedure:
-
First, poses of cyt. c sampled by BD simulations to be associated with either CIII or CIV were examined by the closest separation between the iron atom of their respective heme groups and the corresponding ET-involved cofactors, namely heme c for CIII and CuA for CIV. Poses with proper orientations such that such separation were within values estimated or employed by earlier studies (17 Å for CIII48 and 20 Å39 for CIV) were selected. It should
-
Second, before processing selected poses, a reference structure for ET from CIII to cyt. c (PDB: 3XN5)48 was chosen, and a reference structure for ET from cyt. c to CIV, where cyt. c’ s heme group was estimated to be away from CIV’s CuA by 16 Å50, was constructed. These reference structures and the estimates of the order of magnitude for their corresponding ETRs, which were 1 ×106 for ET from CIII to cyt. c and order 1 ×105 for ET from cyt. c to CIV, were employed to estimate reference electronic couplings for subsequent calculations.
-
Third, selected poses, which were already associated with either CIII or CIV, were adjusted to mimic a tight binding condition with their binder partners by adjusting the separations between the iron atoms of their heme groups and the partners’ cofactors to values in the reference structures used in step 2, which were 17 Å for CIII and 16 Å for CIV. It is worth mentioning that, if sufficient computational resources are available, our adjustment made here will be best achieved through MD simulations of the cyt. c-SC complex, including the membrane embedding our SC and the surrounding solvent.
-
Fourth, pathways for adjusted cyt. c poses were surveyed by the VMD pathway tool82. The pathway tool makes use of the pathway model83 that utilizes graph theory to search for possible electron hopping paths connecting the iron atom of the cyt. c’s heme group to the involved cofactors (heme c for CIII and CuA for CIV). These electron hopping paths involved jump through vacuum, transfer over hydrogen bonds, and potentially covalent bonds. The calculations were performed for 10 iterations for each adjusted pose.
-
Fifth, the electronic coupling decay factor for each of the sampled paths for each pose was computed by the pathway tool. Calculations in step 4 were also repeated for the reference structures.
-
Sixth, our calculation results from step 5 for sampled cyt. c poses were fitted to those for reference structure to estimate the ETRs for our selected poses through attenuating the ETRs for reference structures by pose-dependent electronic coupling decay factor, an approach employed to study ETR between cyt. c2 and photosynthetic reaction centers84.
-
Lastly, statistics for any protein residue involved in sampled ET paths were aggregated to highlight residues that are potentially important to relay electron transfer between cyt. c’s heme and cofactors of CIII and CIV.
Molecular dynamics flexible fitting (MDFF)
MDFF simulations were performed for mutant SC (ΔQCR6-CIII2CIV), where QCR6s were fully removed, following the protocol detailed by MDFF developers85,86 as being shown below:
-
First, the cryo-EM density map shown in Supplementary Fig. 25 was converted into a MDFF potential. This potential functions as a grid force that will statistically bias simulated structures to fit the sourced density map when being implemented in NAMD68.
-
Second, we generated a set of constraint collective variables for CIV to retain the dihedral angles, secondary structures, and helical chirality of the complex during the simulations.
-
Third, we used the VMD MDFF plugin73,86 to dock a standalone CIV to our cryo-EM density map sourced in step 1. This docked CIV was utilized in our target molecular dynamics (TMD)68 routine inside MDFF with a force constant of 200 kcal/mol/Å2. Our CIV would be slowly driven towards its docked position if such motion was allowed by both the imposed constraints in step 2 and the sourced MDFF potential in step 1.
-
Lastly, our starting structure (PDB: 8E7S) was simulated in the presence of the MDFF potential computed in step 1 and the collective variables generated in step 2 in an implicit solvent environment for 500,000 steps under a time step of 1 femto-second.
The cross-correlation coefficient with respect to the cryo-EM density map and the corresponding root-mean-squared-deviation were computed for our simulated CIV, which was the primary SC domain undergoing positional rearrangements, along the simulation trajectory.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM density maps have been deposited in the Electron Microscopy Data Bank as entries EMD-71415 (Yeast Respiratory SuperComplex deltaQCR6) and EMD-71416 (Yeast Respiratory SuperComplex non-uniform refinement). All other data needed to interpret and replicate our study are available from the Figshare repository as entry 27740370. These data include: (i) modeled structures of QCR6 from MELD, which are central to our study; (ii) configuration files for setting up simulations performed in this study, including MD, GaMD, SMD, and ARBD; (iii) the wwwPDB EM validation reports for our employed cryo-EM maps and (iv) movies for animated simulations trajectories. Source data are provided with this paper.
Code availability
The data conversion process of converting ARBD simulation data into MD trajectories for subsequent analyses was performed by a currently developing, in-house package, namely SimpleARBD [https://doi.org/10.5281/zenodo.17371796].
References
Brzezinski, P., Moe, A. & Ädelroth, P. Structure and mechanism of respiratory III-IV supercomplexes in bioenergetic membranes. Chem. Rev. 121, 9644–9673 (2021).
Martin, D. R. & Matyushov, D. V. Electron-transfer chain in respiratory complex I. Sci. Rep. 7, 5495 (2017).
Moe, A., Di Trani, J., Rubinstein, J. L. & Brzezinski, P. Cryo-EM structure and kinetics reveal electron transfer by 2D diffusion of cytochrome c in the yeast III-IV respiratory supercomplex. Proc. Natl. Acad. Sci. 118, e2021157118 (2021).
Jha, P., Wang, X. & Auwerx, J. Analysis of mitochondrial respiratory chain supercomplexes using blue native polyacrylamide gel electrophoresis (BN-PAGE). Curr. Protoc. Mouse Biol. 6, 1–14 (2016).
Schägger, H. & Pfeiffer, K. Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J. 19, 1777–1783 (2000).
Mileykovskaya, E. et al. Arrangement of the respiratory chain complexes in Saccharomyces cerevisiae supercomplex III2IV2 revealed by single particle cryo-electron microscopy. J. Biol. Chem. 287, 23095–23103 (2012).
Maranzana, E., Barbero, G., Falasca, A. I., Lenaz, G. & Genova, M. L. Mitochondrial respiratory supercomplex association limits production of reactive oxygen species from complex I. Antioxid. Redox Signal 19, 1469–1480 (2013).
Rydström Lundin, C., von Ballmoos, C., Ott, M., Ädelroth, P. & Brzezinski, P. Regulatory role of the respiratory supercomplex factors in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. 113, E4476–E4485 (2016).
Javadov, S., Jang, S., Chapa-Dubocq, X. R., Khuchua, Z. & Camara, A. K. Mitochondrial respiratory supercomplexes in mammalian cells: structural versus functional role. J. Mol. Med. 99, 57–73 (2021).
Novack, G. V., Galeano, P., Castaño, E. M. & Morelli, L. Mitochondrial supercomplexes: physiological organization and dysregulation in age-related neurodegenerative disorders. Front. Endocrinol. 11, 600 (2020).
Blaza, J. N., Serreli, R., Jones, A. J., Mohammed, K. & Hirst, J. Kinetic evidence against partitioning of the ubiquinone pool and the catalytic relevance of respiratory-chain supercomplexes. Proc. Natl. Acad. Sci. USA 111, 15735–15740 (2014).
Berndtsson, J. et al. Respiratory supercomplexes enhance electron transport by decreasing cytochrome c diffusion distance. EMBO Rep. 21, e51015 (2020).
Kao, W.-C. et al. Structural basis for safe and efficient energy conversion in a respiratory supercomplex. Nat. Commun. 13, 545 (2022).
Wikström, M. & Springett, R. Thermodynamic efficiency, reversibility, and degree of coupling in energy conservation by the mitochondrial respiratory chain. Commun. Biol. 3, 451 (2020).
Hackenbrock, C. R., Chazotte, B. & Gupte, S. S. The random collision model and a critical assessment of diffusion and collision in mitochondrial electron transport. J. Bioenerg. Biomembr. 18, 331–368 (1986).
Genova, M. L. & Lenaz, G. Functional role of mitochondrial respiratory supercomplexes. Biochim. Biophys. Acta 1837, 427–443 (2014).
Pfeiffer, K. et al. Cardiolipin stabilizes respiratory chain supercomplexes. J. Biol. Chem. 278, 52873–52880 (2003).
Stuchebrukhov, A., Schäfer, J., Berg, J. & Brzezinski, P. Kinetic advantage of forming respiratory supercomplexes. Biochim. Biophys. Acta Bioenerg. 1861, 148193 (2020).
Pérez-Mejías, G., Guerra-Castellano, A., Díaz-Quintana, A., De la Rosa, M. A. & Díaz-Moreno, I. Cytochrome c: surfing off of the mitochondrial membrane on the tops of complexes III and IV. Comput. Struct. Biotechnol. J. 17, 654–660 (2019).
Bergstrom, C. L., Beales, P. A., Lv, Y., Vanderlick, T. K. & Groves, J. T. Cytochrome c causes pore formation in cardiolipin-containing membranes. Proc. Natl. Acad. Sci. USA 110, 6269–6274 (2013).
Böttinger, L. et al. Phosphatidylethanolamine and cardiolipin differentially affect the stability of mitochondrial respiratory chain supercomplexes. J. Mol. Biol. 423, 677–686 (2012).
Mileykovskaya, E. & Dowhan, W. Cardiolipin-dependent formation of mitochondrial respiratory supercomplexes. Chem. Phys. Lipids 179, 42–48 (2014).
Wú, F. et al. Structure of the II2-III2-IV2 mitochondrial supercomplex from the parasite Perkinsus marinus. bioRxiv, https://doi.org/10.1101/2024.05.25.595893 (2024).
Chicco, A. J. & Sparagna, G. C. Role of cardiolipin alterations in mitochondrial dysfunction and disease. Am. J. Physiol. Cell Physiol. 292, C33–C44 (2007).
Hryc, C. F. et al. Structural insights into cardiolipin replacement by phosphatidylglycerol in a cardiolipin-lacking yeast respiratory supercomplex. Nat. Commun. 14, 2783 (2023).
Hartley, A. M. et al. Structure of yeast cytochrome c oxidase in a supercomplex with cytochrome bc(1). Nat. Struct. Mol. Biol. 26, 78–83 (2019).
Gómez, L. A. & Hagen, T. M. Age-related decline in mitochondrial bioenergetics: does supercomplex destabilization determine lower oxidative capacity and higher superoxide production?. Semin. Cell Dev. Biol. 23, 758–767 (2012).
Lobez, A. P. et al. Electron transfer in the respiratory chain at low salinity. Nat. Commun. 15, 8241 (2024).
Yang, M. & Trumpower, B. L. Deletion of QCR6, the gene encoding subunit six of the mitochondrial cytochrome bc1 complex, blocks maturation of cytochrome c1, and causes temperature-sensitive petite growth in Saccharomyces cerevisiae. J. Biol. Chem. 269, 1270–1275 (1994).
Nakai, M. et al. Acidic regions of cytochrome c1 are essential for ubiquinol-cytochrome c reductase activity in yeast cells lacking the acidic QCR6 protein. J. Biochem. 114, 919–925 (1993).
Modjtahedi, N., Tokatlidis, K., Dessen, P. & Kroemer, G. Mitochondrial proteins containing Coiled-Coil-Helix-Coiled-Coil-Helix (CHCH) domains in health and disease. Trends Biochem. Sci. 41, 245–260 (2016).
MacCallum, J. L., Perez, A. & Dill, K. A. Determining protein structures by combining semireliable data with atomistic physical models by Bayesian inference. Proc. Natl. Acad. Sci. 112, 6985–6990 (2015).
Shekhar, M. et al. CryoFold: determining protein structures and data-guided ensembles from cryo-EM density maps. Matter 4, 3195–3216 (2021).
Maffeo, C., Luan, B. & Aksimentiev, A. End-to-end attraction of duplex DNA. Nucleic Acids Res 40, 3812–3821 (2012).
Hunte, C., Koepke, J., Lange, C., Rossmanith, T. & Michel, H. Structure at 2.3 A resolution of the cytochrome bc(1) complex from the yeast Saccharomyces cerevisiae co-crystallized with an antibody Fv fragment. Structure 8, 669–684 (2000).
Solmaz, S. R. & Hunte, C. Structure of complex III with bound cytochrome c in reduced state and definition of a minimal core interface for electron transfer. J. Biol. Chem. 283, 17542–17549 (2008).
Devanathan, S. et al. Plasmon waveguide resonance spectroscopic evidence for differential binding of oxidized and reduced Rhodobacter capsulatus cytochrome c2 to the cytochrome bc1 complex mediated by the conformation of the Rieske iron-sulfur protein. Biochemistry 46, 7138–7145 (2007).
Singharoy, A., Barragan, A. M., Thangapandian, S., Tajkhorshid, E. & Schulten, K. Binding site recognition and docking dynamics of a single electron transport protein: cytochrome c2. J. Am. Chem. Soc. 138, 12077–12089 (2016).
Shimada, S. et al. Complex structure of cytochrome c-cytochrome c oxidase reveals a novel protein-protein interaction mode. EMBO J. 36, 291–300 (2017).
Dudek, J. Role of cardiolipin in mitochondrial signaling pathways. Front. Cell Dev. Biol. 5, 90 (2017).
Wilms, J., Veerman, E. C. I., König, B. W., Dekker, H. L. & van Gelder, B. F. Ionic strength effects on cytochrome aa3 kinetics. Biochim. Biophys. Acta Bioenerg. 635, 13–24 (1981).
Ferguson-Miller, S., Brautigan, D. L. & Margoliash, E. Correlation of the kinetics of electron transfer activity of various eukaryotic cytochromes c with binding to mitochondrial cytochrome c oxidase. J. Biol. Chem. 251, 1104–1115 (1976).
Malatesta, F. et al. Probing the high-affinity site of beef heart cytochrome c oxidase by cross-linking. Biochem. J. 315, 909–916 (1996).
Lobo-Jarne, T. & Ugalde, C. Respiratory chain supercomplexes: structures, function and biogenesis. Semin. Cell Dev. Biol. 76, 179–190 (2018).
Klusch, N. et al. Cryo-EM structure of the respiratory I + III2 supercomplex from Arabidopsis thaliana at 2Å resolution. Nat. Plants 9, 142–156 (2023).
Abresch, E. C., Gong, X. M., Paddock, M. L. & Okamura, M. Y. Electron transfer from cytochrome c(2) to the reaction center: a transition state model for ionic strength effects due to neutral mutations. Biochemistry 48, 11390–11398 (2009).
Zheng, W., Chai, P., Zhu, J. & Zhang, K. High-resolution in situ structures of mammalian respiratory supercomplexes. Nature 631, 232–239 (2024).
Hagras, M. A., Hayashi, T. & Stuchebrukhov, A. A. Quantum calculations of electron tunneling in respiratory complex III. J. Phys. Chem. B 119, 14637–14651 (2015).
Millett, F. & Durham, B. Kinetics of electron transfer within cytochrome bc1 and between cytochrome bc1 and cytochrome c. Photosynthesis Res. 82, 1–16 (2004).
Brzezinski, P. et al. Triplet-state quenching in complexes between Zn-cytochrome c and cytochrome oxidase or its CuA domain. Biophys. Chem. 54, 191–197 (1995).
Pan, L. P., Hibdon, S., Liu, R. Q., Durham, B. & Millett, F. Intracomplex electron transfer between ruthenium-cytochrome c derivatives and cytochrome c oxidase. Biochemistry 32, 8492–8498 (1993).
Lucas, M. F., Rousseau, D. L. & Guallar, V. Electron transfer pathways in cytochrome c oxidase. Biochim. Biophys. Acta Bioenerg. 1807, 1305–1313 (2011).
Lappalainen, P., Watmough, N. J., Greenwood, C. & Saraste, M. Electron transfer between cytochrome c and the isolated CuA domain: identification of substrate-binding residues in cytochrome c oxidase. Biochemistry 34, 5824–5830 (1995).
Chan, C. K., Singharoy, A. & Tajkhorshid, E. Anionic lipids confine cytochrome c(2) to the surface of bioenergetic membranes without compromising its interaction with redox partners. Biochemistry 61, 385–397 (2022).
Vercellino, I. & Sazanov, L. A. Structure and assembly of the mammalian mitochondrial supercomplex CIII(2)CIV. Nature 598, 364–367 (2021).
Smith, P. M., Fox, J. L. & Winge, D. R. Reprint of: Biogenesis of the cytochrome bc1 complex and role of assembly factors. Biochim. Biophys. Acta Bioenerg. 1817, 872–882 (2012).
Singharoy, A. et al. Atoms to phenotypes: molecular design principles of cellular energy metabolism. Cell 179, 1098–1111.e1023 (2019).
Pizzorno, J. Mitochondria-fundamental to life and health. Integr. Med. 13, 8–15 (2014).
Wennerström, H., Vallina Estrada, E., Danielsson, J. & Oliveberg, M. Colloidal stability of the living cell. Proc. Natl. Acad. Sci. USA 117, 10113–10121 (2020).
Erman, J. E. & Vitello, L. B. Yeast cytochrome c peroxidase: mechanistic studies via protein engineering. Biochim Biophys. Acta 1597, 193–220 (2002).
Kortemme, T.; Kim, D. E.; Baker, D. Computational alanine scanning of protein-protein interfaces. Sci STKE, 2004, pl2. https://doi.org/10.1126/stke.2192004pl2 (2004).
Trouillard, M., Meunier, B. & Rappaport, F. Questioning the functional relevance of mitochondrial supercomplexes by time-resolved analysis of the respiratory chain. Proc. Natl. Acad. Sci. 108, E1027–E1034 (2011).
Maldonado, M., Guo, F. & Letts, J. A. Atomic structures of respiratory complex III2, complex IV, and supercomplex III2-IV from vascular plants. eLife 10, e62047 (2021).
Heinemeyer, J., Braun, H. P., Boekema, E. J. & Kouril, R. A structural model of the cytochrome C reductase/oxidase supercomplex from yeast mitochondria. J. Biol. Chem. 282, 12240–12248 (2007).
Miyakuni, K. et al. Genome-wide analysis of DNA methylation identifies the apoptosis-related gene UQCRH as a tumor suppressor in renal cancer. Mol. Oncol. 16, 732–749 (2022).
Xu, D. & Zhang, Y. Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins 80, 1715–1735 (2012).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput Chem. 29, 1859–1865 (2008).
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Xue, L. C., Rodrigues, J. P., Kastritis, P. L., Bonvin, A. M. & Vangone, A. PRODIGY: a web server for predicting the binding affinity of protein–protein complexes. Bioinformatics 32, 3676–3678 (2016).
Comer, J. & Aksimentiev, A. Predicting the DNA sequence dependence of nanopore ion current using atomic-resolution Brownian dynamics. J. Phys. Chem. C. 116, 3376–3393 (2012).
Baker, A. T. et al. ChAdOx1 interacts with CAR and PF4 with implications for thrombosis with thrombocytopenia syndrome. Sci. Adv. 7, eabl8213 (2021).
Shekhar, M. et al. Revealing a Hidden Intermediate of Rotatory Catalysis with X-ray Crystallography and Molecular Simulations. ACS Central Science, 8, 915-925 (2022).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 33–38, 27-38 (1996).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Baker, N.A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc. Natl. Acade. Sci. 98, 10037-10041 (2001).
Cohen, J., Arkhipov, A., Braun, R. & Schulten, K. Imaging the migration pathways for O2, CO, NO, and Xe inside myoglobin. Biophys. J. 91, 1844–1857 (2006).
Miao, Y., Feher, V. A. & McCammon, J. A. Gaussian accelerated molecular dynamics: unconstrained enhanced sampling and free energy calculation. J. Chem. Theory Comput. 11, 3584–3595 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Huang, J. & MacKerell, A. D. Jr CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput Chem. 34, 2135–2145 (2013).
Sener, M., Strumpfer, J., Singharoy, A., Hunter, C. N. & Schulten, K. Overall energy conversion efficiency of a photosynthetic vesicle. eLife 5, e09541 (2016).
Balabin, I. A., Hu, X. & Beratan, D. N. Exploring biological electron transfer pathway dynamics with the Pathways plugin for VMD. J. Comput Chem. 33, 906–910 (2012).
Beratan, D. N., Betts, J. N. & Onuchic, J. N. Protein electron transfer rates set by the bridging secondary and tertiary structure. Science 252, 1285–1288 (1991).
Miyashita, O., Okamura, M. Y. & Onuchic, J. N. Interprotein electron transfer from cytochrome c2 to photosynthetic reaction center: tunneling across an aqueous interface. Proc. Natl. Acad. Sci. 102, 3558–3563 (2005).
Trabuco, L. G., Villa, E., Schreiner, E., Harrison, C. B. & Schulten, K. Molecular dynamics flexible fitting: a practical guide to combine cryo-electron microscopy and X-ray crystallography. Methods 49, 174–180 (2009).
Singharoy, A. et al. Molecular dynamics-based refinement and validation for sub-5 Å cryo-electron microscopy maps. eLife 5, e16105 (2016).
Volkov, A. N. & van Nuland, N. A. J. Electron transfer interactome of cytochrome c. PLOS Comput. Biol. 8, e1002807 (2012).
Acknowledgements
We acknowledge a CAREER award from NSF (MCB-1942763, A.S.). This work is also supported by the National Defense Education Program (NDEP) program under grant number HQ0034-21-S-F001 (A.S.), and DOE grants DE-SC0022956 (A.S.) and DE-SC0010575 (A.S.). The simulation work used resources of the Oak Ridge Leadership Computing Facility’s (OLCF) Frontier Supercomputer, which was awarded through the ASCR Leadership Computing Challenge (ALCC). OLCF is a DOE Office of Science User Facility supported under Contract DE-AC05-00OR22725 (A.S.). We further acknowledge John Rubinstein, Peter Brzezinski for permitting us to use their 3DVA analyzed density data in our analysis, and William Dowhan for fruitful discussions.
Author information
Authors and Affiliations
Contributions
C.K.C. and J.N. were responsible for the simulation results and analysis. C.F.H., M.B., W.D., and E.M. provided the Cryo-EM data. C.G. provided data on the GaMD simulations, and Alberto Perez performed the MELD simulations. K.R. provided valuable insights on the confinement of charge carriers in primitive organisms, such as heliobacteria. E.M. and A.S. are both corresponding authors and contributed to the science and writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chan, C.K., Nguyen, J., Hryc, C.F. et al. Transient protein structure guides surface diffusion pathways for electron transport in membrane supercomplexes. Nat Commun 17, 2892 (2026). https://doi.org/10.1038/s41467-025-67110-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67110-y