Abstract
Reversed seasonality and distinct navigation cues in the Earth’s two hemispheres may shape the evolution of migratory behaviour in animals. Migratory divides—contact zones where populations have evolved alternative migratory strategies—are well-documented in birds and typically occur longitudinally. We hypothesise that insect migratory divides are less likely to emerge longitudinally, but may exist latitudinally, driven by hemisphere-specific sensory adaptations that lead to spatial and temporal isolation. Here, we examine this hypothesis in the cosmopolitan painted lady butterfly (Vanessa cardui), whose Southern Hemisphere dynamics remain unexplored. Investigating the genomes of 300 individuals across Africa and Europe, we identify a 9 Mb chromosomal inversion on chromosome 8, which exhibits strong haplotype structure aligned with hemispheric origin, with a few potential heterozygotes near the equator. The inversion harbours 336 genes, including several directly relevant to migration. Notably, one inversion breakpoint intersects the gene encoding the GABA-B receptor, which responds to the neuropeptide γ-aminobutyric acid (GABA), crucial for insect navigation. Our findings provide genomic evidence of a migratory divide in insects and highlight the role of inverted seasonality in the two hemispheres and genomic rearrangements as isolating barriers for highly mobile species.
Similar content being viewed by others
Introduction
Movement plays a fundamental role in the evolution of life. Among various forms of movement, migration stands out as a particularly specialised and multifaceted behaviour – a choreography of finely tuned responses to fluctuating environmental conditions1. The evolutionary and molecular underpinnings of this complex behaviour remain largely unknown, as do the impacts of migration on species diversification2,3.
Seasonality is crucial in triggering migratory behaviour, as animals rely on environmental signals such as changes in photoperiod, plant phenology, and temperature to initiate, maintain, and terminate migration. These cyclical signals are thought to modulate transcriptional and epigenetic pathways that regulate migratory responses4,5. Once migration begins, navigational adaptations become essential for maintaining directional movement towards favourable destinations. Celestial cues, both diurnal and nocturnal, along with Earth’s magnetic field, influence navigation. These signals are integrated into neural networks that form an internal compass, guiding movement across vast distances6,7.
Seasonality patterns are generally consistent across longitude (east-west) but vary latitudinally (north-south), driving animal migratory movements predominantly in a north-south direction8. In vertebrates, migratory divides – zones where partially sympatric populations adopt distinct migratory strategies – have been widely described, particularly on philopatric species migrating north-south, where divides occur longitudinally9,10. These divides are often characterised by a clear separation between breeding and non-breeding seasons and areas, as well as by behavioural mechanisms such as learning and cultural transmission11.
The emergence of migratory divides can have important implications for early speciation processes in birds and other vertebrates12,13,14,15,16 by exposing sympatric populations to different selective pressures, such as different parasites, predator and climate regimes9,12,17. In addition to selection, assortative mating can also contribute to reproductive isolation12,15,18. Efforts to unravel the genetic basis of population differentiation in contrasting migratory phenotypes have primarily identified single-nucleotide polymorphisms (SNPs)19,20,21,22. However, with the acquisition of contiguous genomes, large genomic alterations such as structural variants have been found to underlie the genetic architecture of migratory divides. Recent studies in fish and birds highlight the importance of chromosomal inversions23,24,25,26,27, deletions28, and long repetitive-rich regions of transposable elements (TEs)29. These major structural variants can suppress recombination over large genomic regions, facilitating the formation of islands of differentiation even in the presence of ongoing gene flow30,31. However, the conditions under which these genomic islands might expand to generalised genome-wide differentiation are not fully understood32.
In insects, long-range cyclical migrations typically span multiple overlapping generations, with each generation completing only one leg of the migratory cycle, and where individuals rarely learn their routes from experience33. Each moving generation involves vast population numbers shaping complex reticular movements, formed by individuals travelling varying distances34,35,36,37. Moreover, because individuals tend to migrate and then reproduce within relatively short lifespans, breeding can take place in any suitable habitat along their migratory routes. Only a few well-known insect species, such as the monarch butterfly, seem to migrate to non-breeding destinations as part of their annual cycle. These dynamics disfavour the evolution of strict philopatry, instead resulting in broad north-south seasonal migratory movements that span extensive geographical ranges34,35,36,37. As a result, the formation of longitudinal migratory divides in insects seems unlikely, except for scenarios where major geographic barriers hinder gene flow37. Indeed, no migratory divide supported by genetic evidence has been identified in insects to date. Nevertheless, the possibility of latitudinal migratory divides with a low temporal overlap in the equator exists for species whose range extends to both hemispheres.
In this study, we investigate how living in the Northern or Southern Hemisphere may influence the evolution of insect migratory strategies – a question that remains largely unexplored. Most insights into migratory patterns in animals come from species from the Northern Hemisphere, while the Southern Hemisphere has been relatively neglected and rarely compared mechanistically. Although independent migratory circuits in each hemisphere are observed in some migratory vertebrates, such as certain birds38,39,40,41 and whales42,43,44, the potential role of hemispheric barriers in driving these patterns has not yet been proposed as a unifying hypothesis. Given that butterflies and moths possess magnetoreceptor organs that may aid in navigation45,46,47, we hypothesise that reversed seasonality, a necessarily opposite response to the magnetic field, and distinct celestial cues in the two hemispheres of Earth may shape the evolution of contrasting migratory strategies. Since environmental rhythmicity displays inverted pulses between hemispheres, populations of migratory insects may adjust their biological clocks, either through genetic changes or phenotypic plasticity, and respond differently to hemisphere-specific seasonal cues, potentially leading to spatial and temporal isolation between populations. We use genomics to address the hypothesis of a migratory divide aligned with hemispheric effects in a highly migratory insect species. The painted lady butterfly, Vanessa cardui, provides an ideal system to examine these dynamics, since its distribution in Africa encompasses the Northern and Southern hemispheres. In both hemispheres, painted ladies are well-known seasonal migrants, often among the most common butterflies in a wide diversity of habitats8. Recent studies have characterised ecological and genetic connectivity between Europe and equatorial Africa35,36,37,48,49,50,51, yet migratory patterns of populations south of the equator remain unknown.
Results And Discussion
Observational data suggest independent movement dynamics in the two hemispheres
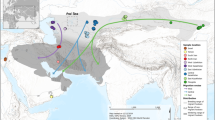
Insects migrate in multigenerational waves, resulting in a latitudinal progression of the species distribution through time. Earlier studies of V. cardui revealed two annual peaks of abundance in equatorial latitudes corresponding to the wet seasons, while only one major demographic peak is observed in the European part of the distribution48,50,52. Our analysis of population latitudinal progression throughout the year, based on 760,973 compiled observations of V. cardui in Europe and Africa (Supplementary Data S1), reveals similar patterns, with successive generations tracking locations with optimal conditions (Fig. 1A). As temperatures drop in the boreal autumn, V. cardui populations migrate from the Palaearctic to Africa, reaching as far south as the equatorial latitudes before migrating northwards again in early spring48,50,51. However, another peak of abundance is observed at equatorial latitudes from May to July (Fig. 1A). These individuals are unlikely to belong to the Northern population that has migrated to Europe during this period. We argue that they are part of a Southern African population expanding northward to avoid the austral winter and the progressive desiccation of subtropical zones (Fig. 1B). While these two populations might rarely overlap in time, they inhabit equatorial latitudes during the two different wet seasons. Overall, observational data suggest that individuals in the Southern Hemisphere constitute an independent migratory circuit.
A The central heatmap plot shows monthly frequencies of V. cardui occurrences by latitudinal divisions. Maps illustrate differing photoperiod and temperature amplitudes across the distributional range, factors likely influencing the migratory syndrome and denoting substantial differences between hemispheres. Occurrence data comprise 768,332 observations of painted ladies obtained from public repositories and authors’ field expeditions. Peaks of presence linking consecutive months can be observed. Monthly frequencies through the year are relative to each latitudinal division. B Summary of hypothesised migratory movements in both hemispheres. Synchrony in northwards and southwards movements in the two populations may lead to allochrony in the equatorial zone, with independent populations spatially overlapping but temporally isolated. The Sahara Desert (yellow) and the tropical forest (green) are regions where V. cardui seldom breeds. Data used for monthly observations are available in Supplementary Data 1. Data underlying the temperature amplitude map are deposited into the OSF repository and are available at the following URL: https://osf.io/6u32k/. Data underlying the photoperiod amplitude map can be obtained online using the provided code (https://doi.org/10.5281/zenodo.17113173). Maps were made using free vector and raster map data from naturalearthdata.com. Butterfly illustrations by Blanca Martí.
A chromosomal inversion segregates populations in the two hemispheres
To investigate whether V. cardui individuals in the Southern Hemisphere form an independent migratory population from that found in the Palaearctic-African region, we examined genome-wide variation in 290 specimens collected along the entire latitudinal extent of Africa and Europe (Supplementary Data S2, Fig. 2A) using a reduced representation approach (see Methods). Genetic differentiation (fixation index, FST) was minimal across most of the genome, consistent with patterns observed in other migratory insects53,54,55,56. However, a single large and contiguous 9 Mb region in chromosome 8 stood out with a sharp increase in genetic differentiation between northern and southern individuals (Table S1, Fig. 3A). This region, spanning about 60% of chromosome 8, also showed pronounced divergence (dXY) and low diversity (π) (Table S1), suggesting the presence of a chromosomal inversion. A PCA based on this region showed two clusters, each corresponding to a different hemisphere (Fig. 2C). This signal persisted after linkage disequilibrium (LD) pruning (Fig. S3).
A Collection of sites for 300 V. cardui specimens. B Results from fineRADstructure, a Bayesian co-ancestry based analysis, using 2855 SNPs belonging to the region with the largest FST differentiation in chromosome 8 (excluding repetitive regions). Two clusters of high shared ancestry were recovered, corresponding to the northern and southern hemispheres. Some individuals (dark blue) were shown to be heterozygous for the inversion. C Principal-component analysis (PCA) in the differentiated region in chromosome 8 for 290 individuals using 976 SNPs. The two main clusters obtained correspond to the North and South hemisphere populations. Samples in grey correspond to potential heterozygotes. Collection sites information is available in the Supplementary Data 2. Data used for B and C are deposited into the OSF repository and are available at the following URL: https://osf.io/6u32k/.
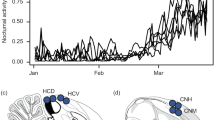
A Population statistics calculated across the genome (1,136,244 variants and 2,788,714 invariant sites) measuring differentiation (FST), absolute divergence (dXY) and diversity (π) between 60 V. cardui individuals from Northern and Southern populations. The plot illustrates the highly differentiated region in chromosome 8. Within this region, absolute divergence between these two populations shows the highest values at the beginning and end, whereas lower values of intra-population genetic diversity, measured as π, are observed for both populations. Window size was set to 100 bp for FST and 1 Mb for dXY and π. B Phylogenetic trees built with whole genome sequencing data for 5 V. cardui individuals representing major geographic regions of its distribution, and the sister species Vanessa kershawi. The tree on top represents the genome-wide tree with the inverted region excluded. The tree below is based on data exclusively from the inverted region. The representatives of the Northern hemisphere (in blue) form a monophyletic cluster, sister to the Southern Hemisphere representative (Namibia, in purple). C Heterozygosity values for the inverted region and for the rest of the genome, calculated as the ratio of variant sites to total genotyped sites. Heterozygosity varies significantly only within the inverted region, with lower values observed in the Southern cluster compared to the Northern cluster. Individuals located between these two main clusters exhibit higher heterozygosity levels, suggesting a heterozygous state (Fig. 3). Box plots depict the median (central line), the 25th and 75th percentiles (bounds of the box), and the whiskers represent the data points within 1.5 x the interquartile range from the box. Data points beyond the whiskers indicate outliers (large dots). Data underlying A–C are deposited into the OSF repository and are available at the following URL: https://osf.io/6u32k/.
Further evidence of population differentiation was provided by a co-ancestry Bayesian clustering analysis (fineRADstructure57), which showed higher levels of co-ancestry within individuals in each hemisphere (Fig. 2B), a pattern not observed when analysing the rest of the genome (Fig. S4). Phylogenetic analyses also support this hemispheric differentiation: while a tree inferred from the inverted region grouped individuals into well-supported clades corresponding to their hemisphere of origin, no phylogenetic structure following latitude arose from a tree inferred from the remainder of the genome (Fig. S5). Remarkably, phylogenetic divergence across the Northern clade, spanning a vast longitudinal range across the Holarctic, is lower than the divergence observed between hemispheres (Figs. 3B, S5), underscoring the pronounced role of the inversion and hemispheric barriers in shaping population structure.
To investigate in greater detail whether this region is an inversion, we analysed read mapping information (BreakDancer58) from a northern individual (reference genome assembly59) and a southern individual using previously published whole-genome data34 (sample 15D327, Namibia). This analysis identified inversion breakpoint coordinates at positions 4,402,570 and 13,379,476, which coincide with the region highlighted by the population genetic summary statistics. Additionally, several smaller inversions (<50 kb) were detected within the larger inverted region (Table S2), potentially reflecting artifacts introduced by mapping uncertainties60,61. However, two nested larger inversions of 3.2 Mb and 1.2 Mb (Table S2) suggest a complex history of structural rearrangements in this genomic region.
Interestingly, the PCA of this region on chromosome 8 revealed that some individuals do not cluster within the northern or the southern genotype groups (Fig. 2C), suggesting that these individuals are heterozygous for the inversion. While northern homozygous individuals formed a cohesive group at one extreme of the PCA, some individuals collected in the south and near the equator were located between this cluster and the southern homozygous type. Heterozygosity in the inverted region of these individuals was significantly higher than for the northern and southern homozygous groups (H = 0.056, Kruskal-Wallis χ2(2) = 64.15, P-value < 0.001, df = 2), but not significantly different in the rest of the genome (Kruskal-Wallis χ2(2) = 3.66, p-value = 0.160, df = 2). Most of these individuals were sampled from regions around equatorial latitudes in East Africa (e.g., Ethiopia, Kenya, Zambia), and outside peak abundance seasons. These findings suggest this area is a contact zone where gene flow can occur between alternative inversion haplotypes.
Chromosomal inversions lead to reduced recombination rates within the inverted region30,62. However, if genetic exchange between inversion variants occurs within an inversion via double crossovers or gene conversion, a U-shaped divergence profile is expected, with stronger divergence near the inversion breakpoints and less pronounced divergence towards the centre25,63,64. Consistent with this prediction, we observed the highest sequence divergence (dXY) between northern and southern populations near the breakpoints (Fig. 3A), supporting marginal gene exchange between migratory ranges.
Altogether, the existence of a single highly divergent genomic region between Northern and Southern Hemispheres, coupled with ecological observations, strongly indicates the presence of a migratory divide between populations of V. cardui butterflies. The lack of population differentiation outside the inversion and the U-shaped divergence within the inversion suggests that gene flow occurs extensively between populations on either side of the divide. This finding highlights the role of chromosomal inversions in maintaining sets of locally adapted alleles through recombination suppression65, acting as a key mechanism leading to local adaptation in highly mobile species66,67.
Recent divergence between hemispheres
We estimated a divergence time for the inversion of 0.55–0.69 mya, although this value is likely underestimated due to gene flow between the inverted and non-inverted haplotypes. In line with a recent split or ongoing gene flow between populations, none of the 32,719 SNPs were fixed between the haplotypes. These findings, combined with the fact that the divergence between V. cardui and its sister species, V. kershawi, is estimated to be much older – around 7 mya68– suggest that the inversion arose in V. cardui after the split between these two species of Vanessa.
Our analysis based on paired-end read mapping information revealed no chromosomal inversions between V. kershawi and the Northern V. cardui reference genome that match the coordinates and length of the inversion detected with population data between V. cardui of northern and southern origins. Therefore, V. kershawi shares collinearity with the Northern haplotype, supporting the Southern haplotype as the derived form. Under this scenario, the emergence of a rearranged haplotype, likely in southern Africa, may have facilitated the expansion of the ancestral haplotype across the Palaearctic and subsequent diversification in the Nearctic37, an interpretation that aligns with our phylogenomic analysis (Fig. 3B). A southern origin of the inversion is also supported by the asymmetrical distribution of haplotypes: both haplotypes as well as individuals presumably heterozygous for the inversion occur in the South, whereas only the Northern haplotype is found in the North (Fig. 2A). This pattern indicates that the inversion is still segregating in the Southern population. Furthermore, the lower heterozygosity observed in Southern individuals homozygous for the inversion, compared to their Northern counterparts (Fig. 3C), may result from a reduction in genetic diversity during the emergence of the Southern haplotype. Positive selection acting on the derived haplotype could also reinforce this pattern, and the higher heterozygosity in the Northern haplotype (apparently fixed) may be maintained through intra-population recombination.
Functional significance of the inversion polymorphism: role in navigation and mating behaviour
Inversion polymorphisms can affect gene expression and subsequently phenotypes through various mechanisms. Primarily, breakpoints can directly affect fitness by disrupting coding and regulatory sequences or altering their positions in the genome69. Additionally, reduced recombination rates associated with chromosomal inversions facilitate faster accumulation of advantageous allele combinations and thus promote local adaptation, often leading to divergent behavioural phenotypes (e.g., refs. 70,71,72,73,74,75). Inversions have been linked to differences in migratory phenotypes in fish such as the rainbow trout76 and the Atlantic cod23,25, and birds such as the common quail26 and the willow warbler27,29,77, where an inversion and an expansion of a transposable element underlies a migratory divide in Europe.
Notably, here we found that both breakpoints overlapped with protein-coding genes. The distal breakpoint intersected the last intron of the subunit 2 of the type B receptor of the γ-aminobutyric acid (GABA) gene (GABAB2). GABA is a neurotransmitter involved in learning and memory that plays a key role in the neural network processing skylight information during navigation78. This network, extensively studied in migratory species like the Monarch butterfly6,7 and the desert locust79, integrates signals in the brain’s central complex to form an internal compass necessary for determining heading-direction during flight80,81,82. The neurons forming this network are GABAergic83,84,85, suggesting that alterations in GABA-B receptor patterns of expression might impact the interpretation of celestial cues. Interestingly, these neurons also receive circadian signals in fruit flies86. The proximal breakpoint was located in an uncharacterised protein from a leucine-rich repeat (LRR) family (Table S3), pointing at another intriguing candidate for future investigation. The fact that the estimated breakpoints of the inversion intersect with genes that are fully annotated and presumably intact in the Northern reference genome implies that these genes are likely disrupted in the Southern haplotype, providing additional support for the Southern haplotype being the derived form.
While the precise environmental cues triggering butterfly migration are not fully understood, synergistic factors like photoperiod, temperature shifts, and host plant availability likely play crucial roles87,88,89,90. To investigate whether the inverted region is rich in genes potentially involved in migratory responses, we first conducted a Gene Ontology (GO) term analysis. The analysis, encompassing the 336 genes within the inverted region, revealed enriched functions related to morphogenesis of locomotion-related tissues, fatty acid metabolism, sensory perception and mating behaviour (Figs. S6, S7).
Secondly, we investigated the association of highly differentiated SNPs within the inverted region with potentially altered gene functions. The inverted region contains 3342 highly differentiated SNPs (FST > 0.25) between southern and northern migratory ranges. Among these, 857 SNPs are located within 48 annotated protein-coding genes of varying degrees of putative impact in protein sequence and function (synonymous and non-synonymous substitutions) (Table S3). Below, we focus on functional roles of the candidate genes within the inverted region. However, we also report 10 genes containing isolated FST outlier SNPs outside the inversion in Table S4 to provide a complete overview and potential follow-up research, as some are associated with neural activity and metabolism. Among the genes containing FST outliers, several are involved in neuroendocrine regulation and behavioural control. This set comprises receptors regulating the main excitatory neuropeptides, such as the Vesicular glutamate transporter 1 and Sodium-dependent dopamine transporter. We also identified hormonal modulators of interest, like the Pituitary homeobox homologue (Ptx1), a transcription factor that activates the promoters of most pituitary hormone-coding genes91, the Juvenile hormone acid O-methyltransferase (JHAMT), a critical enzyme in the biosynthesis of juvenile hormone and repeatedly pinpointed by insect migration studies7,92,93,94 including V. cardui89,90, and the Pyrokinin-1 receptor (PK1R) of the diapause hormone, involved in pheromone biosynthesis95. The acyl-CoA desaturase Delta-9 gene, which produces the main component of the Spodoptera littoralis moth’s sex pheromone96, was also found in this gene set. In fact, several highly differentiated variants were identified in genes related to mating. Of particular interest are the dsf and siwi genes, which are encoding the dissatisfaction protein (DSF) affecting courtship behaviour, successful copulation and fertile egg-laying in Drosophila97, and the Siwi protein, which plays an essential role during spermatogenesis/oogenesis by repressing the mobility of transposable elements98. These genes are promising candidates for investigating premating mechanisms that may contribute to reproductive isolation67,99,100 and assortative mating between hemispheres.
We also found highly differentiated variants in genes related to energy metabolism, such as the main triglyceride-lipase in insect fat bodies (Phospholipase A1101), and the facilitated trehalose transporter (Tret1). Trehalose is the primary sugar in insect haemolymph, and its differential expression has been linked to migratory behaviour90,102,103. Additionally, we detected highly differentiated SNPs in some ABC transporter genes and Argonaute2 (AGO2) in the 3’ and 5’ UTR, respectively, which are associated with xenobiotic resistance. The protein AGO2 is crucial for antiviral defence by targeting and degrading viral RNAs104,105, and variation on its expression could contribute to differing innate immunity signatures across hemispheres, potentially due to variation in microbial community compositions.
Regulatory regions play a critical role in the evolution of novel traits, with most trait-associated loci falling in non-coding regions106. We found 1203 highly differentiated SNPs (FST > 0.25) in the vicinity of 120 genes (5 kb flanking regions), accounting for 36% of the total variants identified. However, our ability to detect genetic differentiation from RAD sequenced variants is limited, as the genes covered in our RAD sequences represent only 21% of the genes encoded within the 9 Mb inverted region. Comprehensive population analyses using whole-genome sequencing (WGS) data could elucidate the full extent of the diverged gene sequences.
Hemispheric migratory divides: a speciation driver in migratory species?
Migratory divides have been proposed as a driving force in speciation by exposing sympatric or parapatric populations to divergent ecological conditions or by limiting genetic exchange through assortative mating or selection against hybridization12,13,14,15,16,17,18,21,107. These divides represent boundaries between adjoined populations with locally adapted migratory traits, such as migration direction or length, often leading to geographic segregation and genetic differentiation. While this phenomenon is well-documented in many vertebrate species (despite its genetic basis being known only in a handful of species), evidence for similar patterns in migratory insects has remained elusive. Indeed, no genetic differentiation has been found between differentially migrating insect populations such as those of monarch butterflies108,109 (reviewed in110), the Bogong moth111 or the fall armyworm54.
However, our study provides evidence from Vanessa cardui suggesting that migratory divides may also play a significant role in insect speciation, despite the substantial differences between vertebrate and insect migration dynamics34. A key distinction in our findings is the identification of a latitudinal migratory divide spanning two hemispheres – a major departure from previously described longitudinal migratory divides that often arose from secondary contact following post-glacial colonisation from separate glacial refugia9. The biogeographic distributions of many migratory butterflies appear restricted to either hemisphere, with some species pairs showing evidence of speciation across hemispheric divides. An example is found in Danaus butterflies: D. plexippus (the monarch) is native to Central and North America, while its sister species, D. erippus, is confined to South America, engaging in extensive parallel migrations. Similarly, in Vanessa butterflies, the sister species V. carye and V. annabella complete their migratory cycles exclusively in South and North America, respectively, while V. kershawi, the painted lady sister species, migrates within Australia and New Zealand, where V. cardui is absent.
While these cases may represent instances involving complete speciation, other systems may shed light on earlier stages of the speciation continuum. Several migratory butterfly species have distributional ranges spanning both hemispheres, such as Ascia monuste, Aphrissa statira, Phoebis sennae in the Americas, and Belenois aurota and Catopsilia florella in the Afrotropical region. Although the migratory cycles and phylogeographic structures of most migratory insect species are still to be fully characterised, we hypothesise that gene flow across such extensive migratory ranges might be constrained by reversed seasonality and differential navigational cues in each hemisphere, eventually leading to adaptation to hemisphere-specific biological rhythms and cues.
This pattern likely extends beyond insects. For example, long-distance bird migrants (excluding shorebirds) breeding in the Northern Hemisphere and wintering in the Southern Hemisphere are proportionally a minority, and few Southern Hemisphere breeders are found to migrate north of the Equator38,39,40,41. Similarly, populations of humpback, fin whales, and several marine migratory fish species circulate almost exclusively within the Northern or Southern hemisphere, and rarely interact42,43,44.
These eco-evolutionary patterns suggest that hemispheric divides could significantly influence the speciation of widespread migratory insect species. The existence of hemispheric divides and their segregating genetic diversity should be investigated across animal migratory lineages as a potentially overlooked pattern for speciation.
Methods
Observational and field data
Observational occurrences of V. cardui used to infer phenological patterns of presence and absence by latitude were retrieved from (1) the Global Biodiversity Information Facility (GBIF112, https://www.gbif.org/es/), (2) the African butterfly and moth mapping project (LepiMAP113, http://vmus.adu.org.za/), both accessed the 23rd of May 2023, and (3) the authors’ own field expeditions from 2006 to 2023 (e.g. ref. 50), including 802 observations from Africa. We excluded the downloaded observations recorded on the 1st of January, as this date appears to be disproportionately used and is likely a placeholder for unknown collection dates within a given year. We retrieved 69,134 observations in total throughout Europe and Africa, dating from 1800 to 2023 (Supplementary Data S1). Monthly data were compiled, and relative abundances for established latitudinal divisions were plotted using ggplot2114 R package (Fig. 1).
We calculated the maximum and minimum yearly photoperiod relative to each latitude using the R packages geosphere115 and terra116. Photoperiod amplitude was estimated as the subtraction of the maximum and minimum yearly day length. The temperature range was estimated as the subtraction of the lowest monthly average temperature from the highest monthly average temperature within each year, where monthly temperature averages were computed for a series of 20 years (2001–2021) using data from the Climate Research Unit (CRU v4.06) database117. Analyses were conducted using R v4.1.2, and R code is provided at the GitHub repository https://github.com/GTlabIBB/MigratoryDivide118. Both photoperiod and temperature amplitudes were projected to a map using QGIS v3.34.4119 to generate Fig. 1A.
Sampling
Painted ladies (V. cardui) are among the world’s most widespread butterfly species. We gathered 300 individuals across 38 countries in Europe and Africa (Fig. 1; Supplementary Data S2), as well as five individuals of V. kershawi, its sister species, which inhabits Australia and New Zealand (Supplementary Data S2). These specimens are deposited in the collections of the Botanical Institute of Barcelona (IBB, CSIC, Spain), the Institute of Evolutionary Biology (IBE, CSIC-Universitat Pompeu Fabra, Spain), and the Museum of Comparative Zoology (Harvard University, USA).
Sequencing and preprocessing of ddRAD data
For population genetic analysis, we prepared ddRAD genomic libraries for the 304 collected samples (Supplementary Data S2) following the protocol of Peterson et al.120. We used EcoRI and BfaI restriction enzymes and size selection of 300 bp using a PippinPrep instrument and 2% agarose cassettes (Sage Science, Beverly, MA, USA). Libraries were sequenced in 150 bp paired-end reads on Illumina HiSeq 2500 at the Harvard University Bauer Core Facility. Sequencing reads are deposited in the ENA (European Nucleotide Archive) database under project accession no. PRJEB80315. A total of 287,916,481 reads were generated, which were demultiplexed and assembled using ipyrad v0.9.81121 by mapping on the Vanessa cardui chromosome-level reference genome59 (GCA_905220365.1). Adapters were filtered using the cutadapt software integrated into the filter_adapters parameter of ipyrad. The clustering threshold was set to 0.85, resulting in a total of 188,408 loci across all 300 individuals. We used ipyrad default filtering parameters except for the minimum number of samples per locus, which was set to 100, i.e., one third of the samples (min_samples_locus parameter). After applying the filters, the number of assembled loci was 10,672. For downstream analysis, we subsequently filtered by variant missing data in VCFtools v0.1.16122, keeping SNPs that were present in at least 80% of the individuals (--max-missing parameter) and by minimum allele count, keeping alleles present in at least 2 individuals (--mac parameter). Last, in order to keep high-quality samples, we filtered out 3 individuals for which less than 40% of the total loci were recovered using the prop_typed_ind function in adegenet v2.1.5R package123, and removed 7 individuals that shared particularly high levels of coancestry according to fineRADstructure57, and thus were potentially related. These samples were indeed collected as larvae in the same locality, further reinforcing this inference. These filters resulted in a dataset, named dataset 1, comprising 290 samples and 32,719 SNPs. For local analysis of the inverted region on chromosome 8, we selected 976 SNPs located within this region using BCFtools v1.20124.
We constructed a second dataset for fineRADstructure analysis, dataset 2, using a reduced set of high quality samples. We selected the first 30 samples from each hemisphere with the lowest levels of missing data. We ran the ipyrad reference-aligned pipeline with default filtering parameters, but the minimum number of samples per locus was set to 30 (i.e., half of the dataset). Additionally, we filtered for missing data with VCFtools, retaining SNPs that were present in at least 80% of the individuals (--max-missing parameter). Initially, we recovered 109,614 loci and 546,458 SNPs across 60 individuals; after filtering, the dataset was reduced to 13,571 loci and 169,590 SNPs. For local analysis on chromosome 8, we selected 2855 SNPs.
A third dataset, named dataset 3, was generated for phylogenetic inference using the 60 samples from dataset 2, along with five additional samples from the sister species Vanessa kershawi as outgroup. The ipyrad assembly parameters and filtering steps were identical to those used for dataset 2. We recovered 117,546 loci across 65 samples, which were reduced to 12,973 loci and 569,090 SNPs after filtering steps. Summary statistics resulting from all ipyrad assembly steps for dataset 1, 2 and 3 are detailed in Supplementary Data S3.
Genome-wide genetic differentiation analysis
To assess population structure, we first performed a PCA analysis using the adegenet v2.1.5 R package123 on all samples (dataset 1, 290 samples, Supplementary Data S3) across three genomic partitions: (i) the whole genome (Fig. S1), (ii) the inverted region on chromosome 8 (Fig. 2B), and (iii) the whole genome excluding the inverted region (Fig. S2). Additional PCAs were performed in LD-pruned versions of these datasets to assess for potential linkage effects, using SNP pairwise comparisons in PLINK with r2 < 0.2 and r2 < 0.05 (Fig. S3).
We then used fineRADstructure to infer the number of genetic clusters and recent shared ancestry of individuals on each side of the migratory divide in dataset 2, both in the entire genome (60 samples, 169,590 SNPs, Fig. S1) and in the inverted region on chromosome 8 (Fig. 2A). In both analyses, sex chromosomes were excluded. Additionally, repetitive regions were filtered out using the annotation from Shipilina et al.125 with BEDTools v2.29.2126. No LD filtering was applied for fineRADstructure analyses, as the method specifically uses haplotype linkage information from RAD data to infer recent ancestry57.
We next explored patterns of genetic differentiation by calculating window-based estimates of FST, π and dXY along the genome for dataset 2. To account for the fragmented coverage of RAD loci we used Pixy v1.2.5.beta1127, which handles missing data by excluding regions lacking mapped reads, thus excluding regions without RAD loci from genome-wide calculations. In order to accommodate this strategy, we generated an AllSites VCF file from the dataset 2 alignments obtained from ipyrad. This variant calling step was implemented in SAMtools mpileup followed by BCFTools call. Additionally, we filtered the resulting file for variant missing data in VCFtools, retaining SNPs present in at least 80% of the individuals (--max-missing parameter). The resulting AllSites VCF contained 1,136,244 variant sites and 2,788,714 invariant sites. Window size was set to 100 bp for FST to enable dot plot visualisation, while 1 Mb was used for π and dXY for curve visualisation, for which no smoothing algorithm was used. Results were plotted using ggplot2.
Sequencing and preprocessing of WGS data
Short read data were used for phylogenetic inference and detection of inversion breakpoints. We used five V. cardui samples from a previous study34, collected in Ethiopia (14U392; 9°23′54.096′′ N, 38°49′25.323′′ E;), Namibia (15D327; 17°52′16.086′′ N, 19°24′22.611′′ E), Nepal (07F575; 28°25′0.635′′ N, 83°49′2.114′′ E), California, USA (15A205; 38°34′30.414′′ N, 121°34′38.117′′ E), and Spain (15A710; 40°15′29.16′′ N, 1°36′20.879′′ E). A Vanessa kershawi specimen, from Australia (AAM97U335; 34°19′30.63′′ S, 117°52′36.735′′ E), was sequenced for this study. A 350 bp insert paired-end library was prepared following the Illumina TruSeq DNA PCR-Free protocol. The library was sequenced at the Swedish National Genomics Infrastructure (NGI) facilities at 150 bp from both ends using a NovaSeq 6000 S4 sequencing platform, producing 140,660,532 reads (Table S5). Sequences are available in the ENA (European Nucleotide Archive) database under project accession no. PRJEB80315.
Reads were mapped against the reference V. cardui genome59 (GCA_905220365.1) with BWA v0.7.12128, except for the V. kershawi specimen, where reads were mapped using Stampy v1.0.32129 and using an estimated substitution rate of 0.0406 to account for ~7my of divergence with V. cardui68. We obtained an average of 194 million mapped reads per sample (Table S5), which were sorted using SAMtools. The consensus sequences were obtained using SAMtools mpileup followed by BCFTools Call. The final fasta files were obtained with vcfutils vcf2fq122 command using a minimum coverage of 6x. The arthropod BUSCO v3.0.2130 analysis of completeness performed in the resulting assemblies using gvolante131 yielded an average of 98.63% completeness.
Identification of inversion breakpoints
To identify the inversion breakpoints, we used paired-end read mapping information from the southern V. cardui individual (Namibia) reads mapped to the northern V. cardui reference genome59 (GCA_905220365.1). The resulting alignment had a mean mapping coverage of 32.34x and mapping quality of 71.3 (Table S5), as calculated with Qualimap v2.2.1132. Structural variant detection was performed using BreakDancerMax, a pipeline to identify breakpoints implemented in the software BreakDancer58. BreakDancer identifies anomalous read pairs in order to predict structural variants, and estimates a confidence score for each variant based on a Poisson model that takes into consideration the number of supporting anomalous read-pairs, the size of the anchoring regions, and the coverage of the genome58. With the highest probability score, BreakDancer identified 14 paired-end reads with abnormally large insert sizes and inverted orientation, pinpointing the inversion breakpoints at coordinates 4,402,570 and 13,379,476 (Table S2). We used the established coordinates to extract the inverted region for all analyses.
Additionally, to investigate the synteny between V. cardui and its sister species, Vanessa kershawi on chromosome 8, we repeated the same procedure using WGS data from the sequenced V. kershawi specimen. The alignment had a mean coverage of 40.24x and a mean mapping quality of 43.61 (Table S5). No chromosomal inversions corresponding to the identified breakpoints with population data in V. cardui (positions 4,402,570–13,379,476) were detected in V. kershawi using read mapping information.
Divergence time estimation of the inverted region
We calculated the divergence time between the inverted and non-inverted haplotypes using the estimated absolute divergence (dXY) from Pixy. Absolute divergence was not consistent across the inversion length. Despite reduced crossover frequency, regions within large inversions can still pair in heterozygotes, allowing for gene conversion63,64. Stronger divergence near the inversion breakpoints and less pronounced divergence towards their centres are expected, showing a U-shaped profile25,26. Therefore, we focused on the first and last Mb of the inversion where recombination is lower, and which therefore provide the most reliable estimate of accumulated divergence since the origin of the inversion. We averaged the values obtained from the first and last Mb. Then, following the approach of Sánchez-Doñoso et al.26, we converted this divergence estimate to an estimate of divergence time using the equation133:
Where T is the divergence time in years, dXY is the absolute divergence, μ is the mutation rate in mutations per site per generation, and g is the number of generations per year. We used the estimated mutation rate for Heliconius melpomene134 (2.9 × 10−9), and considered a range of 7 to 10 generations per year50. Note that the divergence equation assumes a molecular clock, which implies neutral sequence divergence and a constant mutation rate. However, these regions are likely not neutral and might have complex demographic histories, so the time of divergence could be underestimated. Thus, it should be interpreted as an approximate indicator of the inversion’s origin and the onset of independent evolutionary trajectories between the arrangements.
Heterozygosity estimates
Heterozygosity was estimated both for the inverted region on chromosome 8 and along the genome (excluding the inverted region) in dataset 1. We calculated heterozygosity as the number of variant sites over the total number of genotyped sites for each sample. We determined the North group as the specimens clustered in the negative values of PC1, the South group as the specimens clustered at the other side of the PC1 with values > 6, and the specimens scattered between these two thresholds were considered as potential heterozygotes. The obtained heterozygosity estimates did not follow a normal Gaussian distribution (Shapiro-Wilk normality test; inverted region: W = 0.98641, p-value = 0.008075 and excluding inverted region: W = 0.98409, p-value = 0.002754). Therefore, we statistically searched for differences in heterozygosity between North, South, and potential heterozygous groups within each region using Kruskal-Wallis one-way analysis of variance.
Phylogenetic analyses
We investigated the phylogenetic history of the inverted region using whole-genome data from 5 specimens of Vanessa cardui across its worldwide distribution and one Vanessa kershawi specimen as an outgroup. The V. cardui and V. kershawi reference-based assemblies were aligned using MUSCLE135, implemented in Geneious Prime 2023.0.4136 (https://www.geneious.com). Gblocks v0.91b137 was then applied on the multiple sequence alignment to remove potentially ambiguously aligned regions using default parameters, generating concatenated alignments of 4,319,737 bp for the inverted region and 330,406,868 bp for the rest of the genome. Phylogenetic relationships were then inferred through a maximum likelihood (ML) analysis using RAxML-NG v1.1.0138, with a GTR + I model of nucleotide evolution. Nodal support was evaluated by 200 bootstrap iterations using the transfer bootstrap expectation (TBE).
Phylogenetic relationships were also inferred for RAD data using dataset 3. A total of 420 RAD loci alignments were obtained for the inverted region and 12,550 for the rest of the genome using the ipyrad assembly pipeline. We used Gblocks with default parameters in each alignment to discard highly ambiguously aligned regions. Sequences with more than 90% of unmapped or uncalled bases (“Ns”) were removed from the alignments using cutadapt v5.0139 --max-n. These two steps resulted in a total of 397 and 11,935 filtered RAD loci alignments for the inverted region and for the rest of the genome, which were concatenated into 88,704 bp and 2,677,821 bp alignments, respectively. We inferred phylogenetic trees with IQ-TREE v2.0140, using ModelFinder to select the best-fit evolutionary model for each alignment based on the Bayesian Information Criterion (BIC). The resulting phylogenetic trees were plotted using the ggtree141 R package.
Functional analysis
Enrichment of functional categories in the genes located within the inverted region was analysed using the Bioconductor package topGO v2.44.0142. The enrichment test was performed with Fisher’s exact test using the default algorithm (“weight01”), which accounts for the hierarchical structure of the GO-terms. The annotated gene set from Shipilina et al.125 was used as the functional database. We adjusted the p-values of the resulting tests using the Benjamini-Hochberg method143 to correct for multiple testing.
We explored whether SNPs showing the highest differentiation between southern and northern migratory ranges were located within protein-coding genes. We selected SNPs with FST > 0.25 (93.8th percentile) in dataset 2 and predicted effect strength and position in relation to known genes using SnpEff v5.2c144. We created the custom gene database in SnpEff based on the annotation version from Lohse et al.59. Information regarding the known biological functions of these proteins and functional effects is presented in Table S3 (inside the inversion) and Table S4 (outside the inversion).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Demultiplexed and adapter-trimmed RAD-sequencing reads, and whole genome sequencing reads generated in this study have been deposited in the European Nucleotide Archive under project accession number PRJEB80315. All accession numbers for samples used in this study are listed in Supplementary Data 2, including newly generated sequences and those from García-Berro et al.34 and Suchan et al.37 Data underlying Figs. 1–3 are available in Supplementary Data 1 and in the OSF repository at the following URL: https://osf.io/6u32k/ (https://doi.org/10.17605/OSF.IO/6U32K).
Code availability
Scripts used to process the data are available at: https://github.com/GTlabIBB/MigratoryDivide (https://doi.org/10.5281/zenodo.17113173).
References
Dingle, H. Migration: the biology of life on the move. Oxford University Press, USA. (2014).
Irwin, D. E. Speciation: new migratory direction provides route toward divergence. Curr. Biol. 19, R1111–R1113 (2009).
Liedvogel, M., Åkesson, S. & Bensch, S. The genetics of migration on the move. Trends Ecol. Evol. 26, 561–569 (2011).
Baerwald, M. R. et al. Migration-related phenotypic divergence is associated with epigenetic modifications in rainbow trout. Mol. Ecol. 25, 1785–1800 (2016).
Merlin, C. & Liedvogel, M. The genetics and epigenetics of animal migration and orientation: birds, butterflies and beyond. J. Exp. Biol. 222, jeb191890 (2019).
Heinze, S. & Reppert, S. M. Sun compass integration of skylight cues in migratory monarch butterflies. Neuron 69, 345–358 (2011).
Reppert, S. M., Guerra, P. A. & Merlin, C. Neurobiology of monarch butterfly migration. Annu. Rev. Entomol. 61, 25–42 (2016).
Williams, C. B. The Migration of Butterflies (Oliver & Boyd, 1930).
Møller, A. P., Garamszegi, L. Z., Peralta-Sánchez, J. M. & Soler, J. J. Migratory divides and their consequences for dispersal, population size and parasite–host interactions. J. Evol. Biol. 24, 1744–1755 (2011).
Delmore, K. E., Kenyon, H. L., Germain, R. R. & Irwin, D. E. Phenotypic divergence during speciation is inversely associated with differences in seasonal migration. Proc. R. Soc. B Biol. Sci. 282, 20151921 (2015).
Aikens, E. O., Bontekoe, I. D., Blumenstiel, L., Schlicksupp, A. & Flack, A. Viewing animal migration through a social lens. Trends Ecol. Evol. 37, 985–996 (2022).
Rolshausen, G., Segelbacher, G., Hobson, K. A. & Schaefer, H. M. Contemporary evolution of reproductive isolation and phenotypic divergence in sympatry along a migratory divide. Curr. Biol. 19, 2097–2101 (2009).
Delmore, K. E. & Irwin, D. E. Hybrid songbirds employ intermediate routes in a migratory divide. Ecol. Lett. 17, 1211–1218 (2014).
Delmore, K. E. & Liedvogel, M. Investigating factors that generate and maintain variation in migratory orientation: a primer for recent and future work. Front. Behav. Neurosci. 10, 3 (2016).
Turbek, S. P., Scordato, E. S. & Safran, R. J. The role of seasonal migration in population divergence and reproductive isolation. Trends Ecol. Evol. 33, 164–175 (2018).
Uy, J. A. C., Irwin, D. E. & Webster, M. S. Behavioral isolation and incipient speciation in birds. Annu. Rev. Ecol. Evol. Syst. 49, 1–24 (2018).
von Rönn, J. A. C., Harrod, C., Bensch, S. & Wolf, J. B. W. Transcontinental migratory connectivity predicts parasite prevalence in breeding populations of the European barn swallow. J. Evol. Biol. 28, 535–546 (2015).
Bearhop, S. et al. Assortative mating as a mechanism for rapid evolution of a migratory divide. Science 310, 502–504 (2005).
Zhan, S. et al. The genetics of monarch butterfly migration and warning colouration. Nature 514, 317–321 (2014).
Lemopoulos, A., Uusi-Heikkilä, S., Huusko, A., Vasemägi, A. & Vainikka, A. Comparison of migratory and resident populations of brown trout reveals candidate genes for migration tendency. Genome Biol. Evol. 10, 1493–1503 (2018).
Scordato, E. S. et al. Migratory divides coincide with reproductive barriers across replicated avian hybrid zones above the Tibetan Plateau. Ecol. Lett. 23, 231–241 (2020).
Cavedon, M. et al. Genomic legacy of migration in endangered caribou. PLoS Genet. 18, e1009974 (2022).
Sinclair-Waters, M. et al. Ancient chromosomal rearrangement associated with local adaptation of a postglacially colonized population of Atlantic Cod in the northwest Atlantic. Mol. Ecol. 27, 339–351 (2018).
Pearse, D. E. et al. Sex-dependent dominance maintains migration supergene in rainbow trout. Nat. Ecol. Evol. 3, 1731–1742 (2019).
Matschiner, M. et al. Supergene origin and maintenance in Atlantic cod. Nat. Ecol. Evol. 6, 469–481 (2022).
Sánchez-Doñoso, I. et al. Massive genome inversion drives coexistence of divergent morphs in common quails. Curr. Biol. 32, 462–469 (2022).
Sokolovskis, K. et al. Migration direction in a songbird explained by two loci. Nat. Comm. 14, 165 (2023).
Delmore, K. E. et al. Structural genomic variation and migratory behavior in a wild songbird. Evol. lett. 7, 401–412 (2023).
Caballero-López, V., Lundberg, M., Sokolovskis, K. & Bensch, S. Transposable elements mark a repeat-rich region associated with migratory phenotypes of willow warblers (Phylloscopus trochilus). Mol. Ecol. 31, 1128–1141 (2022).
Kirkpatrick, M. How and why chromosome inversions evolve. PLoS Biol. 8, e1000501 (2010).
Mérot, C. et al Locally adaptive inversions modulate genetic variation at different geographic scales in a seaweed fly. Mol. Biol. Evol. 38, 3953–3971.
Ravinet, M. et al. Interpreting the genomic landscape of speciation: a road map for finding barriers to gene flow. J. Evol. Biol. 30, 1450–1477 (2017).
Chapman, J. W., Reynolds, D. R. & Wilson, K. Long-range seasonal migration in insects: mechanisms, evolutionary drivers and ecological consequences. Ecol. Lett. 18, 287–302 (2015).
García-Berro, A. et al. Migratory behaviour is positively associated with genetic diversity in butterflies. Mol. Ecol. 32, 560–574 (2023).
Reich, M. S. et al. Isotope geolocation and population genomics in Vanessa cardui: Short-and long-distance migrants are genetically undifferentiated. PNAS Nexus 4, pgae586 (2025).
Reich, M. S. et al. Trans-Saharan migratory patterns in Vanessa cardui and evidence for a southward leapfrog migration. iScience 27, 111342 (2024).
Suchan, T. et al. A trans-oceanic flight of over 4200 km by painted lady butterflies. Nat. Commun. 15, 5205 (2024).
Dingle, H. Bird migration in the southern hemisphere: a review comparing continents. Emu 108, 341–359 (2008).
Winkler, D. W. et al. Long-distance range expansion and rapid adjustment of migration in a newly established population of barn swallows breeding in Argentina. Curr. Biol. 27, 1080–1084 (2017).
Areta, J. I. et al. Rapid adjustments of migration and life history in hemisphere-switching cliff swallows. Curr. Biol. 31, 2914–2919 (2021).
Helm, B. & Muheim, R. Bird migration: clock and compass facilitate hemisphere switching. Curr. Biol. 31, R1058–R1061 (2021).
Jackson, J. A. et al. Global diversity and oceanic divergence of humpback whales (Megaptera novaeangliae). Proc. R. Soc. B 281, 20133222 (2014).
Pérez-Alvarez, M. et al. Contrasting phylogeographic patterns among Northern and Southern Hemisphere fin whale populations with new data from the Southern Pacific. Front. Mar. Sci. 8, 630233 (2021).
Relano, V. & Pauly, D. Philopatry as a tool to define tentative closed migration cycles and conservation areas for large pelagic fishes in the pacific. Sustainability 14, 5577 (2022).
Guerra, P. A., Gegear, R. J. & Reppert, S. M. A magnetic compass aids monarch butterfly migration. Nat. Commun. 5, 1–8 (2014).
Dreyer, D. et al. The Earth’s magnetic field and visual landmarks steer migratory flight behavior in the nocturnal Australian Bogong moth. Curr. Biol. 28, 2160–2166 (2018).
Wan, G., Hayden, A. N., Iiams, S. E. & Merlin, C. Cryptochrome 1 mediates light-dependent inclination magnetosensing in monarch butterflies. Nat. Commun. 12, 771 (2021).
Talavera, G. & Vila, R. Discovery of mass migration and breeding of the painted lady butterfly Vanessa cardui in the Sub-Sahara: the Europe–Africa migration revisited. Biol. J. Linn. Soc. 120, 274–285 (2017).
Talavera, G., Bataille, C., Benyamini, D., Gascoigne-Pees, M. & Vila, R. Round-trip across the Sahara: Afrotropical Painted Lady butterflies recolonize the Mediterranean in early spring. Biol. Lett. 14, 20180274 (2018).
Talavera, G. et al. The Afrotropical breeding grounds of the Palearctic-African migratory painted lady butterflies (Vanessa cardui). Proc. Natl. Acad. Sci. 120, e2218280120 (2023).
Gorki, J. L. et al. Pollen metabarcoding reveals the origin and multigenerational migratory pathway of an intercontinental-scale butterfly outbreak. Curr. Biol. 34, P2684–2692.E6 (2024).
Menchetti, M., Guéguen, M. & Talavera, G. Spatio-temporal ecological niche modelling of multigenerational insect migrations. Proc. Biol. Sci. 286, 20191583 (2019).
Chen, M. Z. et al. Migration trajectories of the diamondback moth Plutella xylostella in China inferred from population genomic variation. Pest Manag. Sci. 77, 1683–1693 (2021).
Schlum, K. A. et al. Whole genome comparisons reveal panmixia among fall armyworm (Spodoptera frugiperda) from diverse locations. BMC Genom. 22, 1–12 (2021).
Jin, M. et al. Adaptive evolution to the natural and anthropogenic environment in a global invasive crop pest, the cotton bollworm. Innov. 4 (2023).
Ahrens, C. W. et al. Stochastic wind-driven migration likely maintains panmixia in the endangered bogong moth, Agrotis infusa. Biol. Conserv. 302, 110993 (2025).
Malinsky, M., Trucchi, E., Lawson, D. J. & Falush, D. RADpainter and fineRADstructure: population inference from RADseq data. Mol. Evol. Biol. 35, 1284–1290 (2018).
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 6, 677–681 (2009).
Lohse, K. et al. The genome sequence of the painted lady, Vanessa cardui Linnaeus 1758. Wellcome Open Res. 6. (2021).
Alkan, C., Coe, B. & Eichler, E. Genome structural variation discovery and genotyping. Nat. Rev. Genet. 12, 363–376 (2011).
Kosugi, S. et al. Comprehensive evaluation of structural variation detection algorithms for whole genome sequencing. Genome Biol. 20, 117 (2019).
Sturtevant, A. H. & Beadle, G. W. The relations of inversions in the X chromosome of Drosophila melanogaster to crossing over and disjunction. Genetics 21, 554 (1936).
Navarro, A., Betrán, E., Barbadilla, A. & Ruiz, A. Recombination and gene flux caused by gene conversion and crossing over in inversion heterokaryotypes. Genetics 146, 695–709 (1997).
Reis, M., Vieira, C. P., Lata, R., Posnien, N. & Vieira, J. Origin and consequences of chromosomal inversions in the virilis group of Drosophila. Genome Biol. Evol. 10, 3152–3166 (2018).
Samuk, K. et al. Gene flow and selection interact to promote adaptive divergence in regions of low recombination. Mol. Ecol. 26, 4378–4390 (2017).
Rieseberg, L. H. Chromosomal rearrangements and speciation. Trends Ecol. Evol. 16, 351–358 (2001).
Faria, R., Johannesson, K., Butlin, R. K. & Westram, A. M. Evolving inversions. Trends Ecol. Evol. 34, 239–248 (2019).
Wahlberg, N. & Rubinoff, D. Vagility across Vanessa (Lepidoptera: Nymphalidae): mobility in butterfly species does not inhibit the formation and persistence of isolated sister taxa. Syst. Entomol. 36, 362–370 (2011).
Berdan, E. L. et al. How chromosomal inversions reorient the evolutionary process. J. Evol. Biol. 36, 1761–1782 (2023).
Gilburn, A. S. & Day, T. H. Female mating behaviour, sexual selection and chromosome I inversion karyotype in the seaweed fly, Coelopa frigida. Heredity 82, 276–281 (1999).
Thomas, J. W. et al. The chromosomal polymorphism linked to variation in social behavior in the white-throated sparrow (Zonotrichia albicollis) is a complex rearrangement and suppressor of recombination. Genetics 179, 1455–1468 (2008).
Ayala, D., Ullastres, A. & González, J. Adaptation through chromosomal inversions in Anopheles. Front. Genet. 5, 129 (2014).
Küpper, C. et al. A supergene determines highly divergent male reproductive morphs in the ruff. Nat. Genet. 48, 79–83 (2016).
Huang, Y. C., Dang, V. D., Chang, N. C. & Wang, J. Multiple large inversions and breakpoint rewiring of gene expression in the evolution of the fire ant social supergene. Proc. R. Soc. B 285, 20180221 (2018).
Koch, E. L. et al. Genetic variation for adaptive traits is associated with polymorphic inversions in Littorina saxatilis. Evol. Lett. 5, 196–213 (2021).
Arostegui, M. C., Quinn, T. P., Seeb, L. W., Seeb, J. E. & McKinney, G. J. Retention of a chromosomal inversion from an anadromous ancestor provides the genetic basis for alternative freshwater ecotypes in rainbow trout. Mol. Ecol. 28, 1412–1427 (2019).
Lundberg, M., Mackintosh, A., Petri, A. & Bensch, S. Inversions maintain differences between migratory phenotypes of a songbird. Nat. Commun. 14, 452 (2023).
Nguyen, T. A. T., Beetz, M. J., Merlin, C. & El Jundi, B. Sun compass neurons are tuned to migratory orientation in monarch butterflies. Proc. R. Soc. B 288, 20202988 (2021).
Homberg, U. et al. The sky compass network in the brain of the desert locust. J. Comp. Physiol. 209, 641–662 (2023).
Homberg, U., Heinze, S., Pfeiffer, K., Kinoshita, M. & El Jundi, B. Central neural coding of sky polarization in insects. Philos. Trans. R. Soc. B 366, 680–687 (2011).
Honkanen, A., Adden, A., da Silva Freitas, J. & Heinze, S. The insect central complex and the neural basis of navigational strategies. J. Exp. Biol. 222, jeb188854 (2019).
Merlin, C., Heinze, S. & Reppert, S. M. Unraveling navigational strategies in migratory insects. Curr. Opin. Neurobiol. 22, 353–361 (2012).
Homberg, U., Humberg, T. H., Seyfarth, J., Bode, K. & Perez, M. Q. GABA immunostaining in the central complex of dicondylian insects. J. Comp. Neurol. 526, 2301–2318 (2018).
von Hadeln, J. et al. Neuroarchitecture of the central complex of the desert locust: tangential neurons. J. Comp. Neurol. 528, 906–934 (2020).
Takahashi, N., Zittrell, F., Hensgen, R. & Homberg, U. Receptive field structures for two celestial compass cues at the input stage of the central complex in the locust brain. J. Exp. Biol. 225, jeb243858 (2022).
Lamaze, A., Krätschmer, P., Chen, K. F., Lowe, S. & Jepson, J. E. A wake-promoting circadian output circuit in Drosophila. Curr. Biol. 28, 3098–3105 (2018).
Boman, J. et al. Environmental stress during larval development induces DNA methylation shifts in the migratory painted lady butterfly (Vanessa cardui). Mol. Ecol. 32, 3513–3523 (2023).
Friberg, M., Lehmann, P. & Wiklund, C. Autumn mass change and winter mass loss differ between migratory and nonmigratory butterflies. Anim. Behav. 204, 67–75 (2023).
Näsvall K., Shipilina D., Vila R., Talavera G. & Backström N. Resource availability affects activity profiles of regulatory elements in a long-distance butterfly migrant. Authorea. https://doi.org/10.22541/au.167827909.99815237/v1 (2023).
Shipilina, D. et al. Gene expression responses to environmental cues shed light on components of the migratory syndrome in butterflies. bioRxiv, https://www.biorxiv.org/content/10.1101/2024.07.17.602486v1 (2024).
Tremblay, J. J., Lanctôt, C. & Drouin, J. The pan-pituitary activator of transcription, Ptx1 (pituitary homeobox 1), acts in synergy with SF-1 and Pit1 and is an upstream regulator of the Lim-homeodomain gene Lim3/Lhx3. Mol. Endocrinol. 12, 428–441 (1998).
Rankin, M. A. & Riddiford, L. M. Significance of haemolymph juvenile hormone titer changes in timing of migration and reproduction in adult Oncopeltus fasciatus. J. Insect Physiol. 24, 31–38 (1978).
Jones, C. M. et al. Genomewide transcriptional signatures of migratory flight activity in a globally invasive insect pest. Mol. Ecol. 24, 4901–4911 (2015).
Zhang, L. et al. Juvenile hormone regulates the shift from migrants to residents in adult oriental armyworm, Mythimna separata. Sci. Rep. 10, 11626 (2020).
Dou, X. & Jurenka, R. Pheromone biosynthesis activating neuropeptide family in insects: a review. Front. Endocrinol. 14, 1274750 (2023).
Rodríguez, S., Camps, F. & Fabriàs, G. Inhibition of the acyl-CoA desaturases involved in the biosynthesis of Spodoptera littoralis sex pheromone by analogs of 10, 11-methylene-10-tetradecenoic acid. Insect Biochem. Mol. Biol. 34, 283–289 (2004).
Finley, K. D., Taylor, B. J., Milstein, M. & McKeown, M. dissatisfaction, a gene involved in sex-specific behavior and neural development of Drosophila melanogaster. Proc. Natl. Acad. Sci. 94, 913–918 (1997).
Kawaoka, S. et al. The Bombyx ovary-derived cell line endogenously expresses PIWI/PIWI-interacting RNA complexes. RNA 15, 1258–1264 (2009).
Dagilis, A. J. & Kirkpatrick, M. Prezygotic isolation, mating preferences, and the evolution of chromosomal inversions. Evolution 70, 1465–1472 (2016).
Huang, K. & Rieseberg, L. H. Frequency, origins, and evolutionary role of chromosomal inversions in plants. Front. Plant Sci. 11, 296 (2020).
Arrese, E. L., Patel, R. T. & Soulages, J. L. The main triglyceride-lipase from the insect fat body is an active phospholipase A1: identification and characterization. J. Lipid Res. 47, 2656–2667 (2006).
Doyle, T. et al. Genome-wide transcriptomic changes reveal the genetic pathways involved in insect migration. Mol. Ecol. 31, 5434332–5434350 (2022).
Jin, M. et al. Adaptive evolution to the natural and anthropogenic environment in a global invasive crop pest, the cotton bollworm. Innovation 4 (2023).
Van Mierlo, J. T. et al. Convergent evolution of argonaute-2 slicer antagonism in two distinct insect RNA viruses. PLOS Pathog. 8, e1002872 (2012).
Santos, D. et al. Insights into RNAi-based antiviral immunity in Lepidoptera: acute and persistent infections in Bombyx mori and Trichoplusia ni cell lines. Sci. Rep. 8, 2423 (2018).
Edwards, S. L., Beesley, J., French, J. D. & Dunning, A. M. Beyond GWASs: illuminating the dark road from association to function. Am. J. Hum. Genet. 93, 779–797 (2013).
Winker, K. On the origin of species through heteropatric differentiation: a review and a model of speciation in migratory animals. Ornithol. Monogr. 69, 1–30 (2010).
Lyons, J. I. et al. Lack of genetic differentiation between monarch butterflies with divergent migration destinations. Mol. Ecol. 21, 3433–3444 (2012).
Talla, V. et al. Genomic evidence for gene flow between monarchs with divergent migratory phenotypes and flight performance. Mol. Ecol. 29, 2567–2582 (2020).
Freedman, M. G. et al. Are eastern and western monarch butterflies distinct populations? A review of evidence for ecological, phenotypic, and genetic differentiation and implications for conservation. Conserv. Sci. Pract. 3, e432 (2021).
Wallace, J. R. Maleszka, R., & Warrant, E. J. Large-scale whole-genome sequencing of migratory Bogong moths Agrotis infusa reveals genetic variants associated with migratory direction in a panmictic population. bioRxiv, https://doi.org/10.1101/2022.05.27.493801 (2022).
GBIF.org (2024) GBIF Occurrence Download https://doi.org/10.15468/dl.z66kv5.
LepiMAP: Atlas of African Lepidoptera. Published by the FitzPatrick Institute of African Ornithology, University of Cape Town, http://vmus.adu.org.za/?vm=LepiMAP (2023)
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag). https://ggplot2.tidyverse.org (2009).
Hijmans, R. J., Williams, E., Vennes, C., & Hijmans, M. R. J. Package ‘geosphere’. Spherical trigonometry 1, 1-45. https://github.com/rspatial/geosphere (2017).
Hijmans, R. J. et al. Package ‘terra’. Maintainer: Vienna, Austria. https://rspatial.github.io/terra/ (2022).
Harris, I., Osborn, T. J., Jones, P. & Lister, D. Version 4 of the CRU TS monthly high-resolution gridded multivariate climate dataset. Sci. Data 7, 109 (2020).
García-Berro, A. et al. A north-south hemispheric migratory divide in the butterfly Vanessa cardui. https://github.com/GTlabIBB/MigratoryDivide, https://doi.org/10.5281/zenodo.17113173 (2025).
QGIS.org, QGIS Geographic Information System. Open Source Geospatial Foundation Project. http://qgis.org (2024).
Peterson, B. K., Weber, J. N., Kay, E. H., Fisher, H. S. & Hoekstra, H. E. Double digest RADseq: an inexpensive method for De Novo SNP discovery and genotyping in model and non-model species. PLoS One 7, e37135 (2012).
Eaton, D. A. & Overcast, I. ipyrad: Interactive assembly and analysis of RADseq datasets. Bioinform 36, 2592–2594 (2020).
Danecek, P. et al. The variant call format and VCFtools. Bioinform 27, 2156–2158 (2011).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinform 24, 1403–1405 (2008).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Shipilina, D. et al. Linkage mapping and genome annotation give novel insights into gene family expansions and regional recombination rate variation in the painted lady (Vanessa cardui) butterfly. Genomics 114, 110481 (2022).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinform 26, 841–842 (2010).
Korunes, K. L. & Samuk, K. pixy: Unbiased estimation of nucleotide diversity and divergence in the presence of missing data. Mol. Ecol. Res. 21, 1359–1368 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinform 25, 1754–1760 (2009).
Lunter, G. & Goodson, M. Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 21, 936–939 (2011).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinform 31, 3210–3212 (2015).
Nishimura, O., Hara, Y. & Kuraku, S. gVolante for standardizing completeness assessment of genome and transcriptome assemblies. Bioinform 33, 3635–3637 (2017).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinform 32, 292–294 (2016).
Kimura, M. Evolutionary rate at the molecular level. Nature 217, 624–626 (1968).
Keightley, P. D. et al. Estimation of the spontaneous mutation rate in Heliconius melpomene. Mol. Ecol. Evol. 32, 239–243 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinform 28, 1647–1649 (2012).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinform 35, 4453–4455 (2019).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Xu S. et al. “Ggtree: A serialized data object for visualization of a phylogenetic tree and annotation data.” iMeta, 1, e56 (2022).
Alexa, A., & Rahnenfuhrer, J. topGO: enrichment analysis for gene ontology. R package version, 2 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92 (2012).
Acknowledgements
We are grateful to our colleagues who contributed samples used in this study, including B. Acosta, A. Andersen, D. Benyamini, S. Cuvelier, L. Dapporto, V. Dincă, R. Eastwood, M. Espeland, M. Gascoigne-Pees, E. Hailu, A. Heath, J. Hernández-Roldán, J.C. Hinojosa, M. López-Munguira, V. Lukhtanov, E. Karolinskiy, M. Khaldi, R. Khellaf, D. Lohman, A. Marsiñach, A.A. Mignault, Y. Monasterio, S. Montagud, A. Morton, F. Páramo, L. Pokorny, I. Ribera, S. Scalercio, N. Shapoval and R. Vodă. This work was funded by the grants PID2020-117739GA-I00 and PID2023-152239NB-I00 (MCIN / AEI / 10.13039/501100011033), by the National Geographic Society (grant WW1-300R-18), by the grant LINKA20399 from the CSIC iLink program, and by grant 2021-SGR-01334 (Departament de Recerca i Universitats, Generalitat de Catalunya) to G.T. A.G.B is supported by the grant FPU-2019-01593. R.V. is supported by grant PID2022-139689NB-I00 (MICIU/ AEI/ 10.13039/501100011033 and ERDF, EU) and by grant 2021-SGR-00420 (Departament de Recerca i Universitats, Generalitat de Catalunya). G.T. and N.E.P. also acknowledge the Putnam Expeditionary Fund from the Museum of Comparative Zoology (MCZ, Harvard University). We thank Roger López-Mañas for support in environmental data visualisation, and Thomas Decroly and Dorcas Orengo for valuable discussion.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.G-B., D.S., N.B., N.E.P., R.V., G.T; Field Sampling: A.G-B., S.C.C., D.J.M., R.V., G.T; Data Collection: G.T.; Data analysis: A.G-B., D.S., N.B., T.S., A.P., G.T.; Writing – original draft: A.G-B., G.T.; Writing – review and editing: A.G-B., D.S., N.B., T.S., A.P., S.C.C., D.J.M., N.E.P., R.V., G.T. Funding: N.E.P., R.V., G.T.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
García-Berro, A., Shipilina, D., Backström, N. et al. A north-south hemispheric migratory divide in the butterfly Vanessa cardui. Nat Commun 16, 11341 (2025). https://doi.org/10.1038/s41467-025-67185-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67185-7