Abstract
Protein dynamics play a crucial role in various physiological functions, including enzyme catalysis. To explore conformational changes during enzyme catalysis, we conduct mix-and-inject serial crystallography, an advanced technique to capture time-resolved protein structures in real time, using the microcrystals of bacterial copper amine oxidase containing a protein-derived quinone cofactor. Within 50 ms of mixing the microcrystals (<4 μm) with a preferred substrate (2-phenylethylamine) under anaerobic conditions (reductive half-reaction), we observe domain movements associated with substrate binding and formation of a metastable reaction intermediate, a product Schiff-base of the quinone cofactor. At 100–1000 ms after mixing, conformational transition from aminoresorcinol to the semiquinone radical forms of the reduced cofactor progresses gradually, likely depending on the replacement of the product aldehyde by the next-cycle amine substrate that triggers the cofactor conformational change. Overall, this study provides structural insight into enzyme catalysis accompanying the active-site conformational changes that are hardly scrutinized by studies in solution.
Similar content being viewed by others
Introduction
Proteins exhibit various motion modes over a wide range of time scales, including the thermal vibration of individual bonds in the femtosecond range, rotation or conformational changes of local structures in the pico–microsecond range, and domain/whole protein movement in the millisecond (ms) range1,2. Such dynamics play fundamental roles for proteins in exhibiting various cellular functions1. For receptors and enzymes, the motions induced by binding of ligands and substrates/effectors are essential for specific recognition. Thermal fluctuation of the proton/hydrogen acceptor/donor is linked to enzyme catalysis via hydrogen tunneling3. Adjustment of the protonation states of the active site via conformational changes in the surrounding residues is crucial to rearrange the hydrogen bonds or modulate the reaction efficiency. Enzymes optimize the environment of their active sites and accelerate specific reactions by modulating the protein dynamics.
Development of mix-and-inject serial crystallography (MISC), which is an application of serial femtosecond X-ray crystallography (SFX) using an X-ray free-electron laser (XFEL)4,5,6,7,8,9, has been a technical breakthrough to investigate the millisecond changes in the atomic structures associated with enzyme catalysis10,11,12,13,14. In MISC, a microcrystal suspension of an enzyme is mixed with the substrate and ejected to the XFEL interaction point, enabling the direct observation of the time-resolved structural changes during the catalytic reaction10,11,12,13,14. Namely, MISC is a crystallographic version of a classical continuous-flow or stopped-flow technique used for spectrophotometric detection of time-resolved reaction intermediates formed during enzyme catalysis in solution15,16. In addition, laser excitation of chromophores contained in the enzyme system, which is typically used for initiation of the reaction in most pump-probe experiments, is not required in MISC. These aspects underscore the versatility of MISC, making it applicable to various enzyme reactions.
We focus on copper amine oxidase, a ubiquitous enzyme that occurs widely in prokaryotes and eukaryotes and participates in various physiological functions such as microbial amine metabolism and plant germination17. Its peptidyl cofactor, 2,4,5-trihydroxyphenylalanine quinone (TPQ), is generated via a metal ion-dependent post-translational modification17,18,19. TPQ cofactor plays the central redox role in the oxidative deamination of primary amines via a ping-pong mechanism consisting of reductive and oxidative half-reactions (Supplementary Fig. 1). In the initial reductive half-reaction, oxidized form of the cofactor (TPQox) reacts with the substrate to form a substrate Schiff-base (TPQssb), which is converted into a product Schiff-base (TPQpsb) and hydrolyzed to form the first reaction product (a corresponding aldehyde) and 2e−-reduced form of the cofactor (TPQamr). TPQamr remains in equilibrium with the radical form of TPQ (TPQsq) by transferring 1e− to Cu(II). Subsequently, oxidative half-reaction starts with the dioxygen accepting 2e−/2H+ from the TPQamr/Cu(II) or TPQsq/Cu(I) couple depending on the enzyme source, forming the second product (H2O2) and an iminoquinone form (TPQimq), which finally reverts to initial TPQox via hydrolysis or TPQssb via transimination, releasing ammonia. We extensively study the mechanism of reductive half-reaction of the enzyme from Arthrobacter globiformis (AGAO) by conducting in-solution kinetic studies via stopped-flow measurements and determine the kinetic constant at each reaction step. In addition, the structures of reaction intermediates that form in the TPQamr/TPQsq equilibrium state are determined by X-ray and neutron crystallography at high resolution16,20,21,22,23,24,25,26,27 (Fig. 1). Taking advantage of these previous findings, we here show an MISC study of AGAO microcrystals conducted for further understanding of the interplay between the protein dynamics and catalytic mechanisms.
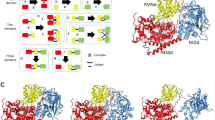
a Ribbon model of the dimeric structure of AGAO based on its highest-resolution structure22 (PDB entry: 3WA2). In the enlarged view, active site residues, including the TPQ cofactor (magenta) in an off-copper form, are represented by stick models with the copper atom (brown sphere). b Schematic of Cu(II)-dependent autoxidative TPQ generation from a specific Tyr residue18,19.
Results
MISC measurement and data processing
To obtain sufficient diffraction data via XFEL irradiation, we prepared AGAO microcrystals with a uniform size of approximately 2.5 × 4 μm (parallelogrammic plates; thickness <1 μm) and suspended them in a crystallization buffer (approximately 108–109 mL−1), as previously described28. The crystal suspension and substrate solution containing 2-phenylethylamine (2-PEA; the most preferred substrate29 with Km = 2.5 μM, kcat = 76 s−1 determined at pH 6.8, 30 °C16) were prepared under anaerobic conditions at pH 9.0 so that the pH-dependent equilibrium between TPQamr and TPQsq shifts toward TPQsq24, and mixed inside the injector capillary to give the final substrate concentration of 4 mM. The reaction time elapsed after mixing the enzyme crystals with the substrate (hereafter defined as the delay time [Δtm]) was controlled using injector capillaries with different distances between the mixing and irradiation points and changing the total flow rate of the reaction mixture (see the Methods section for details). XFEL diffraction data were obtained at Δtm = 22, 25, 50, 100, 200, 300, 400, 500, and 1000 ms. Assuming that the reductive half-reaction in crystals proceeds rapidly as in solution with the decreasing order of rate constants (see Supplementary Fig. 1; k1 > k2 > k3 > k4)23, the Δtm was set at increasing time intervals to capture rapid structural changes in an early stage of the reaction. To obtain the initial enzyme structure (containing TPQox) before starting the reaction, single-flow SFX30,31 of the crystal suspension in a buffer without 2-PEA was conducted and the resultant data set was defined as Δtm = 0 ms. Subsequently, diffraction images collected at all Δtm points were subjected to data processing.
During the initial integration of diffraction data, we noted bimodal distributions of the lengths of a unit cell parameter, a-axis, among the indexed images at Δtm = 22, 25, and 50 ms (Fig. 2, middle), unlike the normal distributions of other parameters (b, c, α, β, and γ) with single peaks (Supplementary Fig. 2). Histograms of the number of images counted for a-axis values were best fitted to the sum of two normal distribution peaks. The a-axis lengths at the two peaks were estimated to differ by 0.68–0.69 Å in the curve fitting of the three data sets, indicating the presence of two microcrystal regions with unit lattices of different a-axis lengths. As the AGAO microcrystal size considerably exceeded the XFEL beam diameter (1.5 μm), various microcrystal regions were irradiated by the pulsed beam (Supplementary Fig. 3). Therefore, for subsequent structure refinement, we divided the diffraction data sets at the three Δtm points into two groups containing less than 10% of the counterpart distribution peaks (Fig. 2, bottom) designated as 22–50 ms (L) and (S) for long and short a-axes, respectively, and calculated them separately, together with the diffraction data sets at other Δtm points (0, 100, 200, 300, 400, 500, and 1000 ms), which exhibited single distribution peaks in all unit cell parameters. Structure models were refined by applying the structures of TPQ intermediates produced during the reductive half-reaction with 2-PEA (Fig. 3a). All data collection and structure refinement details are presented in Supplementary Tables 1 and 2. Differences in a-axis lengths between the (L) and (S) data sets at Δtm = 22, 25, and 50 ms were 0.81, 0.87, and 0.75 Å, respectively, after final data processing.
Number of indexed images obtained from the diffraction data at Δtm = 22 (a), 25 (b), and 50 (c) ms is plotted against the a-axis length. Histograms were fitted to two normal distribution curves (red) using the Igor Pro version 6 software (Wave Metrics, Inc.; middle), with residuals showing the difference between the observed number of images and fitted curve (top). Deconvoluted distribution curves are shown for the data sets at Δtm = 22–50 ms (S) and (L) in purple and orange, respectively (bottom). Colored thick bars indicate the areas of indexed images used for structure refinement, which are enclosed by dotted lines representing the cut-off a-axis lengths to minimize the contamination of the counterpart distribution peaks ( < 10%). The a-axis lengths recalculated for the peak of each species are 193.82 Å for 22 ms (L), 193.13 Å for 22 ms (S), 193.69 Å for 25 ms (L), 193.01 Å for 25 ms (S), 193.61 Å for 50 ms (L), and 192.92 Å for 50 ms (S). Source data are provided as a Source Data file.
a Reaction sequence of the reductive half-reaction with 2-PEA as the substrate. TPQox, oxidized form (orange); TPQssb, substrate Schiff-base (black); TPQpsb, product Schiff-base (purple); TPQamr, aminoresorcinol form (magenta); TPQsq, semiquinone radical form (green). b Refined structures of TPQ, 2-PEA, and active site water molecules (W1, W2, W3, W4, Wax, and Weq; see also Supplementary Fig. 4). Structures obtained from the Δtm = 22–50 ms (L) and (S) data sets are shown above and below the line, respectively. Stick and sphere models are superimposed on the Fo–Fc polder omit maps (gray mesh) contoured at 5.5 σ (for 22 and 25 ms [L]) or 6.0 σ (others), except for Cu. c Relative amounts of time-resolved TPQ species are plotted against the delay time (Δtm) graduated in a logarithmic scale. Relative amounts of TPQox and TPQpsb were calculated from the total areas of the fitted curves for the 22–50 ms (L) and (S) data sets, respectively (Fig. 2), with standard error (SE) obtained in the curve fitting, and those of TPQamr and TPQsq were calculated from the average occupancies in the refined dimer structure at Δtm = 100–1000 ms with SE obtained in the structure refinement. Curves for TPQox and TPQpsb (100−TPQox) are the best fit to the equation of the 2D substrate-diffusion model described later, while those for TPQamr and TPQsq are the best fit to the exponential disappearance and appearance of TPQamr and TPQsq, respectively. Color codes are consistent in (a–c). Source data are provided as a Source Data file.
Structures captured in 50 ms
Structures determined for the 22–50 ms (L) data sets were identical to the initial enzyme structure before starting the reaction (0 ms), containing TPQox and four water molecules (W1–W4) in the substrate-binding pocket (Fig. 3b, above the line; Supplementary Fig. 4). Population of the TPQox structure without a bound substrate decreased in inverse proportion to Δtm (Fig. 3c). These results suggest that the data sets with a long a-axis are derived from the inner core region of the microcrystals (Supplementary Fig. 3) and that the diffusion of the amine substrate takes 50–100 ms until reaching the core region that remains in the initial substrate-free TPQox structure. In contrast, structures determined for the 22–50 ms (S) data sets, which possibly diffracted from the peripheral region of the microcrystals, showed the substrate-bound cofactor assignable to TPQpsb (Fig. 3b, below the line)16,21, as determined previously by conventional cryogenic crystallography32. Population of the TPQpsb structure increased in proportion to Δtm (Fig. 3c). Hence, time-resolved crystal structure containing TPQpsb was captured as a metastable synchronized catalytic intermediate in this study. However, the predicted first intermediate containing TPQssb (Fig. 3a) was not captured, possibly due to the much faster rates of its formation and decay compared to the rate of substrate diffusion into the microcrystals. The a-axis shortening associated with substrate binding is discussed later in relation to unidirectional protein shrinkage, including domain movement.
Structures captured in 100–1000 ms
Diffraction data sets at Δtm = 100–1000 ms had the a-axis lengths corresponding to those of the data sets at 22–50 ms (S) (Supplementary Table 1), confirming that all AGAO molecules in the microcrystals reacted with 2-PEA. The refined structures determined for these Δtm points, except for Δtm = 1000 ms, were essentially identical to previously reported structures24,26, containing an equilibrium mixture of TPQamr and TPQsq (Figs. 3b and 4a). TPQpsb hydrolysis to form TPQamr (Fig. 3a) proceeded slowly during 50–100 ms. The slower decay of TPQpsb compared to its formation, even at Δtm = 22 ms, is likely the reason for its capture as a metastable intermediate16,21. Occupancy of TPQamr relative to TPQsq, calculated after structural refinement, decreased gradually with the increase in Δtm and reached 0% at 1000 ms (Fig. 3c). Meanwhile, occupancy of conformer a of Tyr296, which stabilizes the TPQsq radical by forming a strong hydrogen bond27, concomitantly increased during 100–1000 ms (Fig. 4b). Hence, transition from TPQamr to TPQsq was visualized in real-time during the later stage (Δtm > 100 ms) of the reductive half-reaction. The order of magnitude of apparent rate constants of these steps (TPQpsb → TPQamr, 20 s−1 (1/[50 ms]); TPQamr → TPQsq, 1 s−1 (1/[1000 ms])) was compatible with that of the rate constants (102 and 39 s−1, respectively) determined in solution at 4 °C and pH 6.8 via stopped-flow measurements23, although the rates in crystals were slower by 5–40 fold even at a higher temperature (26 °C).
a Stick models of the active site residues and PAA/2-PEA in the refined structures at Δtm = 100 ms (for TPQamr, PAA, Tyr296b, Asp298, Asn381, His431, and His433) and 1000 ms (for TPQsq, 2-PEA, Tyr296a, Asp298, Asn381, His431, and His433) are shown along with the Fo–Fc polder omit maps (gray mesh) contoured at 6.0 σ for PAA, TPQamr/TPQsq, and conformers a/b of Tyr296. b B-factor values averaged for the nine atoms of bound 2-PEA molecules in the dimer structure (black square) and occupancies of TPQsq (green square) and Tyr296a (orange square) are plotted against the delay time with SE obtained in the structure refinement. Dotted line indicates the curve fitting of TPQsq formation to first-order kinetics. Notably, B-factor values of 2-PEA did not change significantly upon structure refinement with the PAA model (39.69 ± 3.96 Å2 in the refined structure at Δtm = 100 ms). Source data are provided as a Source Data file.
Notably, substrate-derived electron densities in all structures refined for the data sets at Δtm = 100–1000 ms were disconnected from those of the reduced cofactors (Fig. 3b) and could be assigned as the reaction product, phenylacetaldehyde (PAA), or amine substrate, 2-PEA (Fig. 4a). The two ligands are almost indistinguishable by X-ray crystallography because of very similar electron densities derived from the aldehyde oxygen and amine nitrogen atoms, but our recent neutron crystallographic study27 unequivocally demonstrated the binding of an additional amine substrate replacing PAA, 2-PEA, to the reduced enzyme containing TPQsq. This newly bound 2-PEA possibly triggers the conformational change from TPQamr to TPQsq via electrostatic interactions27. Although the timing at which PAA is replaced by 2-PEA during Δtm = 100–1000 ms remains unknown, the substrate-derived electron densities possibly represent the mixture of PAA and 2-PEA. Atomic temperature factors (B-factors) of bound PAA/2-PEA also decreased gradually during Δtm = 100–400 ms and reached a constant level after 500 ms in inverse proportion to the relative occupancy of TPQsq (Fig. 4b), indicating the complete replacement of PAA by 2-PEA stably bound in the hydrophobic pocket.
Discussion
Time-dependent decrease in the number of diffraction images of 22–50 ms (L) and concomitant increase in that of 22–50 ms (S) (Fig. 3) reflect the progress of substrate diffusion into the microcrystals. As shown in Fig. 5, the map calculated by the program MAP_CHANNELS33 exhibited a planar solvent channel that expands to the direction parallel to the b- and c-axes, suggesting that the amine substrate easily diffuses in the two directions. Moreover, the entrances to the substrate channel (Supplementary Fig. 5) are connected to this solvent channel (shown by red arrows in Fig. 5). By contrast, the map indicated that the substrate can hardly diffuse in the direction parallel to the a-axis due to the discontinuous solvent channel interrupted by a layer of enzyme molecules. The a-axis of crystal lattice roughly coincides with the direction of crystal thin section, as confirmed by an X-ray diffraction analysis of crystal morphology using an AGAO macrocrystal (Supplementary Fig. 6). Thus, the slow crystal growth along the a-axis may also be explained by the poor contact of enzyme dimers separated by the solvent channel (Fig. 5). Assuming that substrate diffusion follows a two-dimensional model, in which diffusion across the crystal thin section is negligible (Supplementary Fig. 7), amine substrate requires approximately 120 ms to diffuse into the entire volume of the microcrystals. This diffusion time is markedly slower than the reported value (1 ms) for a crystal with a similar size (3 × 4 × 5 μm3)10. The amine substrate freely diffuses into the vacant spaces within the crystal lattice but must penetrate through the specific channel connecting the molecular surface and active site (Supplementary Fig. 5). Therefore, substrate diffusion with limited access and strict directionality to the substrate-binding pocket and also the associated domain movements caused by conformational changes of several residues described below become rate-limiting at an early stage of the reductive half-reaction, resulting in failure to capture the initial intermediates, including TPQssb. The slow rate of this substrate-binding process (32 s−1; estimated by first-order fitting in Supplementary Fig. 7c) is possibly due to the crystal packing effects, as the rate of substrate binding in solution is very rapid ( > 900 s−1; the rate of substrate binding should be at least higher than that of TPQssb formation23), occurring within the mixing dead time (2.3 ms) of the stopped-flow experiment at 4 °C23. The formations of TPQssb and TPQpsb in crystals were assumed to be very rapid as in solution (887, 206 s−1, respectively)23, which could be the reason for a relatively large structural change observed for a short period (3 ms) in an early stage (Δtm = 22–25 ms) of the reaction (Figs. 2 and 3c).
A solvent channel in the crystal lattice was visualized using a program MAP_CHANNELS33 with the initial enzyme structure (containing TPQox) before starting the reaction (Δtm = 0 ms). In the upper left panel, the solvent channel (cyan mesh map) is contoured with the AGAO dimer and its symmetric mates (light brown and gray surface models) aligned in the lattice. In the upper right panel, only the solvent channel is shown. A single crystal lattice is shown in thin yellow lines with labels indicating a-, b-, and c-axes. Small black arrows indicate the direction of the unit cell axes in the abc coordinate system. Red arrows in an enlarged view (lower panel) indicate the two entrances to the substrate channel of the AGAO dimmer.
An unexpected finding of this study was the shortening of the a-axis associated with substrate binding. This a-axis shortening may be explained by the inspection of domain movement in the AGAO structure, which is composed of three domains in a monomeric unit: the large core domain D4 and two small peripheral domains D2 and D3 attached to D434 (labeled based on the four-domain structure of the Escherichia coli enzyme35). The active site containing the cofactor and adjacent substrate-binding pocket is located inside the protein in the boundary region of the three domains. In the refined dimer structure of 50 ms (S) superimposed on the core D4/D4 domains of the 50 ms (L) dimer by minimizing root-mean-square-deviation (RMSD) of the main-chain Cα positions, domains D2 and D3 indicated significant movements compared to the less mobile D4 domain (Fig. 6a). In the crystal lattice, while domain D3 mainly moved toward an empty space (−0.232 Å along b-axis), slightly affecting the a-axis length (−0.084 Å), domain D2 moved closer to D3 in the positive direction along a-axis (0.139 Å) so as to pack the substrate-binding pocket (Fig. 6b). Since these domain movements also occur on the opposite side of the dimer molecule in the reverse direction, the a-axis shortening per monomer (0.139 + 0.084 Å) would be magnified further in the crystal lattice by 4 folds (approximately 0.89 Å), which corresponds to an average value (0.81 Å) for the observed a-axis differences between the (L) and (S) data sets of Δtm = 22–50 ms (Supplementary Table 1).
Dimer structure of 50 ms (S) is superimposed on the core D4/D4 domains of the 50 ms (L) dimer by minimizing the root mean square deviation (RMSD) of the main-chain Cα positions. a RMSD values of Cα positions of the monomer (chain A) of the 50 ms (L) and (S) structures are plotted against the residue numbers. D2 (residues 9–91), D3 (residues 103–203), and D4 (residues 229–623) domains are indicated by blue, orange, and green bars, respectively, with residue numbers at both ends. Thin red lines indicates the average RMSD value (also mentioned in numbers) of each domain. Source data are provided as a Source Data file. b Array of dimers in the crystal lattice (thin gray line) is shown with the same color codes for each domain as indicated in panel (a), except for gray-colored domain D4, viewed in an orthoscopic manner with slight rotation (5°) around the a-axis. Four dimers in the unit cell are shown in darker colors. Small black arrows indicate the direction of the unit cell axes in the abc coordinate system. Enlarged window illustrates the movements of the D2 and D3 domains identified by comparing the 50 ms (L) and (S) structures. Two magenta arrows indicate the vectors of the mass center movements of D2 (blue; vector components: a = 0.139 Å, b = −0.053 Å, and c = 0.021 Å) and D3 (orange; a = −0.084 Å, b = −0.232 Å, and c = 0.0041 Å) against D4 (green) in the 50 ms (S) structure. To emphasize the moving direction, vector lengths are magnified by 40-fold at the mass centers calculated from the Cα positions of the domains. TPQpsb (magenta spheres) and substrate-binding pocket (white surface) are also overlaid.
Together with domain movement, several residues constituting the substrate channel and binding pocket underwent notable conformational changes. Namely, when the 22–50 ms (S) structures were compared with the 0 ms structure, Phe105, Ala135, Pro136, and Leu358* (belonging to another monomer) located in the entrance of the substrate channel (Supplementary Fig. 5) substantially changed their main-chain (N-Cα-CO) conformations, as represented by high RMSD values (Fig. 7a). As presented in Supplementary Table 3 for the changes in the side-chain dihedral angles, the side chain of Phe105 rotated by approximately 54° (an average value in the 22–50 ms (S) structures relative to the 0 ms structure) around the Cβ-Cγ bond (Fig. 7b). Furthermore, the side chain of Tyr302 located at the bottom of the substrate channel rotated by 43° on average around the Cβ-Cγ bond. Leu358* changed its side-chain conformation to close the substrate channel as a lid through Cα-Cβ (70°) and Cβ-Cγ rotation (117°) (Fig. 7b). Interestingly, the main-chain conformations and/or the side-chain dihedral angles of these residues were maintained after Δtm = 100 ms and did not return to their initial states during Δtm = 100–1000 ms (Fig. 7a–b and Supplementary Table 3). Collectively, the present study revealed that the binding of the amine substrate to AGAO leads to unidirectional protein shrinkage, which is associated with domain movement and conformational changes in relevant residues. This type of protein dynamics at less than an angstrom scale has rarely been scrutinized experimentally by other methods.
a RMSD values of the main chain conformations against the TPQox structure (Δtm = 0 ms) are plotted for active site residues (within 4 Å from TPQ; aligned according to the appearance in panel [b] from left to right) in the Δtm = 22 − 50 ms (L) (pink), 22 − 50 ms (S), and 100 − 1000 ms (gray-to-black gradient) structures. b Stick models of the active site residues drawn with the same color codes used in panel [a], with the residues mentioned in the main text shown as thick stick models. Source data are provided as a Source Data file.
TPQsq formation in the last step is equivalent to the conformational change of TPQamr, accompanied by 1e−-transfer to the bound Cu(II) (Supplementary Fig. 1), and can be regarded as a first-order process with an estimated rate constant of approximately 2.5 s−1 (Fig. 4b). The rate constants of TPQsq formation determined in solution at pH 6.8 by stopped-flow measurements of spectral changes are 92.9 s−1 at 15 °C21 and 39 s−1 at 4 °C23. Therefore, effect of crystal packing on the rate of TPQsq formation appeared to be much larger than the effects on the thermodynamic parameters of the TPQamr–TPQsq equilibrium determined at pH 6.0, which were less than 2 folds24. Meanwhile, rate of the equilibrium shift from TPQamr to TPQsq determined in solution by temperature-jump measurements is very fast: 325 s−1 at 32 °C and pH 7.236. Therefore, gradual formation of TPQsq over 100–1000 ms cannot be simply explained by the crystal packing effect. Based on 2-PEA binding in the TPQsq structure27, the first reaction product, PAA, must desorb from the substrate-binding pocket and be replaced by 2-PEA in the specified time range. Although the specific timing of PAA/2-PEA exchange remains unknown, it should be diffusion-controlled, similar to the initial substrate binding. If the TPQamr–TPQsq transition in crystals triggered by the binding of new 2-PEA27 is as rapid as in solution, gradual formation of TPQsq (Fig. 4b) represents the gradual replacement of PAA by 2-PEA, rather than the slow rate (2.5 s−1) itself. Considering the location of the active site inside the protein, which faces low crystal packing effects, rate of TPQsq formation in crystals is possibly also diffusion-controlled. The notable discrepancy in the rate constants of TPQsq formation determined in solution might also be attributed to the involvement of the invisible PAA/2-PEA replacement process in the stopped-flow measurements, but not in the temperature-jump measurements.
As PAA and 2-PEA are indistinguishable based on electron density (Fig. 4a), their replacement processes cannot be tracked via MISC. However, B-factor changes in ligands and surrounding residues provide important information on the binding and/or replacement processes. Here, B-factor of the bound ligand (PAA/2-PEA) decreased gradually during 100–400 ms, maintaining a nearly constant value from 500 ms onwards (Fig. 4b). This indicates that the PAA/2-PEA replacement initiates as early as Δtm = 100 ms and is completed at Δtm = 500 ms, yielding 2-PEA stably bound in the hydrophobic pocket. Although the average B-factors of the overall protein atoms were essentially identical throughout the measured time range (Supplementary Table 2), B-factors of the residues constituting the substrate channel, including Phe105, Ala135, Pro136, and Leu358*, were consistently higher than those of the active site residues throughout Δtm = 0–1000 ms, being highest at 100 ms, immediately after TPQpsb hydrolysis (Supplementary Fig. 8). The perspective outward view from the active site also revealed that the entrance of the substrate channel wide open in the initial structure (0 ms) became narrower during 22–100 ms and remained almost closed after 100 ms. Therefore, thermal fluctuations of the channel residues were elevated, representing the first binding of 2-PEA at 0–50 ms and desorption of product PAA followed by re-binding of secondary 2-PEA at 100–400 ms. After 500 ms, both bound 2-PEA (Fig. 4b) and residues in the substrate channel and binding pocket (Supplementary Fig. 8) maintained low B-factors. Collectively, these results suggest that the gradual transition from TPQamr to TPQsq during 100–1000 ms (Fig. 3c) is due to the slow replacement of PAA by 2-PEA depending on the diffusion-controlled rates of desorption of PAA from the substrate-binding pocket and re-binding of 2-PEA.
In conclusion, this study revealed the domain movements associated with substrate binding and captured time-resolved structures of catalytic intermediates formed during the reductive half-reaction of AGAO. Product desorption and the following substrate re-binding were also assessed indirectly from the conformational changes and thermal fluctuations of residues in the substrate channel. In contrast, the pump-probe SFX experiments give the time-dependent structural changes by subtracting the contribution of the unreacted species because only a part of molecules is activated in most cases (13–34% activation in ref. 4;40% in ref. 7; and 5,6,7,8,9,10,11,12,13,14,15. 2% in ref. 9 except 100% in ref. 8)4,7,8,9, although shorter time ranges are attainable than in MISC. Therefore, the changes in thermal fluctuations of active-site residues associated with structural changes are undetectable, clearly demonstrating a merit of MISC beyond the pump-probe SFX. Furthermore, the finding that the crystal packing effects result in significant perturbation of the rates of the catalytic reaction and ligand binding and release processes suggests that MISC might provide a general method to capture protein dynamics during enzyme catalysis. Finally, we have determined 12 time-resolved structures serially at 9 time points ranging from 22 milliseconds to 1 s formed during the reductive-half reaction of AGAO. Thus, this study provides a reference for the development of a flipbook-like movie of enzyme catalysis in the future. MISC is a simple form of kinetic X-ray crystallography applicable to various enzymes. Time-limiting diffusion of the substrate might be specific for AGAO, which is possibly overcome using nanocrystals10. The use of a designed mutant enzyme is an alternative way of lowering catalytic rates at a level detectable by MISC37. Moreover, high consumption of enzyme crystals, which is a limitation of MISC, can be prevented using a suitable delivery system, such as a conveyor belt system38.
Methods
Sample preparation and crystallization
Recombinant AGAO was purified in the inactive precursor form and converted into the Cu(II)/TPQ-containing active form, as previously described16. Protein concentrations were spectrophotometrically determined using the extinction coefficient at 280 nm (13.2) for a 1% solution of the active form16. AGAO microcrystals were prepared using a combination of micro-seeding and batch crystallization methods in 1.05 M potassium-sodium tartrate containing 25 mM HEPES buffer (pH 7.4), as previously described28. The pH of the microcrystal suspension was adjusted to 9.0 by repeatedly replacing the upper buffer with 1.05 M potassium-sodium tartrate containing 25 mM TAPS buffer (pH 9.0) after each microcrystal sedimentation. The final pH was measured using a pH meter. Microcrystal size and concentration were determined using the BZ-X710 all-in-one fluorescence microscope (Keyence, Osaka, Japan). Subsequently, microcrystal suspension was filtered through a nylon mesh (pore size: 20 μm; Sysmex, Kobe, Japan) via unforced gravitational sedimentation under atmospheric pressure. To establish fully anaerobic conditions, the microcrystal suspension (approximately 108–109 crystals mL−1; protein concentration, 30 mg mL−1) and substrate solution in the crystallization buffer (pH 9.0) were placed in a glovebox (AS-600PC; AS ONE, Osaka, Japan) filled with N2 gas for at least 24 h. Finally, plastic vessels containing the microcrystal suspension and substrate solution were doubly enclosed in an O2-impermeable plastic bag inside the glovebox and transported to the SPring-8 Angstrom Compact Linear Accelerator (SACLA) beamline, BL2 (Hyogo, Japan).
Data collection and processing
Diffraction data were collected at the SACLA beamline39 via MISC10,11,12,13,14. Two types of injectors were used for the measurements: A double-focusing mixing jet40 and gas dynamic virtual nozzle connected via a PEEK tube to a T-junction mixer11. Briefly, microcrystal suspension and substrate solution were fully loaded into 15-mL and 50-mL sample reservoirs, respectively, in an anaerobic glovebox (Coy Laboratory Products, MI, USA) at the beamline. The reservoir containing the microcrystal suspension was reciprocally and gently rotated to prevent microcrystal sedimentation. For MISC, the two reservoirs were connected to a high-performance liquid chromatography pump (LC-20AD; Shimadzu, Kyoto, Japan) with a water-filled PEEK tube. The microcrystal suspension and substrate solution were pushed out by pumping water via a piston in the reservoirs, avoiding direct contact of the samples with water, and delivered through the PEEK tube to the liquid injectors with an inner diameter of 75 μm. Flow rates of the microcrystal suspension and substrate solution were 0.02 and 0.23 mL min−1, respectively, with the double-focusing mixer (Δtm: 50–400 ms) and 0.125 mL min−1 (for both) with the T-junction mixer (Δtm: 500 and 1000 ms). The microcrystal suspension mixed with the substrate solution inside the injector was ejected to the XFEL irradiation point on a diverse application platform for hard X-ray diffraction at SACLA (DAPHNIS)31,41. Δtm was controlled by changing the flow distances inside the injector or T-junction system for Δtm = 50–1000 ms at a constant total flow rate (0.25 mL min−1) or setting the total flow rate at 0.57 and 0.50 mL min−1 for Δtm = 22 and 25 ms, respectively, using the injector with the shortest flow distance. Δtm was calculated based on the distance between the mixing and XFEL irradiation points, capillary or PEEK tube width, and flow rate. For single-flow SFX experiments (the data set was defined as Δtm = 0 ms), 5-fold diluted microcrystal suspension alone was delivered at a flow rate of 0.25 mL min−1 with the gas dynamic virtual nozzle26. Data sets for a single-flow SFX of the substrate-free initial enzyme (0 ms) and MISC at Δtm = 500 and 1000 ms were collected first (proposal number of the Japan Synchrotron Radiation Research Institute: 2020A8013), then those at Δtm = 22–50 ms (2021A8013), and lastly those at Δtm = 100–400 ms (2021B8029). As an average of the MISC measurements, operation time of about 8 h including the adjustment time and a total of 300 mg protein were required for collecting the diffraction images sufficient for a single data set.
XFEL diffraction patterns were observed at an ambient temperature (approximately 26 °C). A MultiPort Charged Coupled Device detector was set up with a short-working-distance octal sensor arrangement42 at a sample-to-detector distance of 50 mm. XFEL beam was focused at a beam diameter of 1.5 μm in full-width at half-maximum using a couple of elliptical mirrors in the Kirkpatrick–Baez geometry43. The pulse beam had a wavelength of 1.24 Å (10.0 keV), repetition rate of 30 Hz, temporal width within a 10-femtosecond duration, and pulse energy of approximately 640 μJ at the light source. Data collection was guided via real-time analysis using the SACLA data-processing pipeline44. Data were retrieved using the SACLA application programming interface45 and filtered using Cheetah46, with each image with over 20 spots accepted as a hit. Hit images were indexed and integrated using CrystFEL v 0.9.1 package47. Detector background estimated by averaging the dark images was subtracted from the diffraction patterns. Detector metrology was refined using Geoptimiser48. Indexing was performed using XGANDALF49. Data collection and processing details are presented in Supplementary Table 1.
Structure solution and refinement
Initial phase was determined via molecular replacement using Phaser50. The search model was based on the coordinates of the AGAO monomer (Protein Data Bank accession code: 3wa2) after removing all ligands, including water molecules and metal ions. Refinements, electron density map calculations, and solvent molecule assignments were performed using Phenix v 1.19.2 package51. Occupancies of alternate conformers found in several residues, including TPQ and Tyr296, were refined using occupancy refinement in Phenix, with the initial values set to equal occupancy ( = 0.5) for each conformer. Manual rebuilding was performed using Coot v 0.9.852. During refinement, water molecules and other ligands, such as Cu and TPQ, were added stepwise to the model. Specifically, TPQox, TPQpsb, TPQamr, and TPQsq models were constructed based on the Fo–Fc polder omit map. PyMOL (Schrödinger, New York, USA) was used to construct figures. Statistical information related to the refinement process are provided in Supplementary Table 2.
X-ray diffraction analysis of lattice orientation
To determine the lattice orientation in a large AGAO crystal, the oxidized form of enzyme was crystallized initially at pH 7.4 in dialysis buttons16. The crystals grown to a millimeter size were further dialyzed against 1.05 M potassium-sodium tartrate containing 25 mM TAPS buffer (pH 9.0). The buttons were then transferred to the crystallization buffer at pH 9.0 containing 45 % (w/v) glycerol as cryoprotectant and kept overnight. The crystals were frozen in liquid nitrogen. X-ray diffraction images of the crystals were measured under cryogenic conditions at a beamline station BL44XU in SPring-8 and were processed with DIALS53. The lattice orientation was determined with the directions of the respective unit vectors (a/|a|, b/|b|, and c/|c| ) of a-, b-, and c-axes of the real lattice, which were calculated with a*-, b*-, and c*-axes of the reciprocal lattice obtained from a crystal setting matrix A in a DIALS log using the following equations (also refer to the source file): a = (b*×c*)/V*, b = (c*×a*)/V*, c = (a*×b*)/V*, and a•a* = b•b* = c•c* = 1, where V* (volume of reciprocal lattice) = a*•(b*×c*).
Data availability
All crystallographic models (refined against the raw structure factors described in the article) and associated reflection data have been deposited into the Protein Data Bank (PDB) under accession codes 9LZD (structure of the TPQox form determined by single-flow SFX); 9LZ8 (structure at Δtm = 22 ms with long a-axis); 9LZ7 (structure at Δtm = 22 ms with short a-axis); 9LZA (structure at Δtm = 25 ms with long a-axis); 9LZ9 (structure at Δtm = 25 ms with short a-axis); 9LZC (structure at Δtm = 50 ms with long a-axis); 9LZB (structure at Δtm = 50 ms with short a-axis); 9LZ3 (structure at Δtm = 100 ms); 9LZ4 (structure at Δtm = 200 ms); 9LZ5 (structure at Δtm = 300 ms); 9LZ6 (structure at Δtm = 400 ms); 9LZE (structure at Δtm = 500 ms); and 9LZF (structure at Δtm = 1000 ms). PDB 3WA2 was initially used as the starting model for structure determination. The source data underlying Figs. 2, 3c, 4b, 6a and 7a, and Supplementary Figs. 2, 6, and 7c are provided with this paper as a Source Data file. Source data are provided with this paper.
References
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Ono, J., Matsumura, Y., Mori, T. & Saito, S. Conformational dynamics in proteins: entangled slow fluctuations and nonequilibrium reaction events. J. Phys. Chem. B 128, 20–32 (2024).
Klinman, J. P. & Kohen, A. Hydrogen tunneling links protein dynamics to enzyme catalysis. Annu. Rev. Biochem. 82, 471–496 (2013).
Nango, E. et al. A three-dimensional movie of structural changes in bacteriorhodopsin. Science 354, 1552–1557 (2016).
Neutze, R., Wouts, R., van der Spoel, D., Weckert, E. & Hajdu, J. Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 406, 752–757 (2000).
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77 (2011).
Tenboer, J. et al. Time-resolved serial crystallography captures high-resolution intermediates of photoactive yellow protein. Science 346, 1242–1246 (2014).
Barends, T. R. M. et al. Direct observation of ultrafast collective motions in CO myoglobin upon ligand dissociation. Science 350, 445–450 (2015).
Pande, K. et al. Femtosecond structural dynamics drives the trans/cis isomerization in photoactive yellow protein. Science 352, 725–729 (2016).
Schmidt, M. Mix and inject: reaction initiation by diffusion for time-resolved macromolecular crystallography. Adv. Condens. Matter Phys. 2013, 1–10 (2013).
Stagno, J. R. et al. Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography. Nature 541, 242–246 (2017).
Olmos et al. Enzyme intermediates captured “on the fly” by mix-and-inject serial crystallography. BMC Biol. 16, 59 (2018).
Dasgupta, M. et al. Mix-and-inject XFEL crystallography reveals gated conformational dynamics during enzyme catalysis. Proc. Natl. Acad. Sci. USA 116, 25634–25640 (2019).
Pandey, S. et al. Observation of substrate diffusion and ligand binding in enzyme crystals using high-repetition-rate mix-and-inject serial crystallography. IUCrJ 8, 878–895 (2021).
Takahashi, S. et al. Folding of cytochrome c initiated by submillisecond mixing. Nat. Struct. Biol. 4, 44–50 (1997).
Chiu, Y. C. et al. Kinetic and structural studies on the catalytic role of the aspartic acid residue conserved in copper amine oxidase. Biochemistry 45, 4105–4120 (2006).
Klema, V. J. & Wilmot, C. M. The role of protein crystallography in defining the mechanisms of biogenesis and catalysis in copper amine oxidase. Int. J. Mol. Sci. 13, 5375–5405 (2012).
Matsuzaki, R., Fukui, T., Sato, H., Ozaki, Y. & Tanizawa, K. Generation of the topa quinone cofactor in bacterial monoamine oxidase by cupric ion-dependent autooxidation of a specific tyrosyl residue. FEBS Lett. 351, 360–364 (1994).
Kim, M. et al. X-ray snapshots of quinone cofactor biogenesis in bacterial copper amine oxidase. Nat. Struct. Biol. 9, 591–596 (2002).
Kishishita, S. et al. Role of copper ion in bacterial copper amine oxidase: spectroscopic and crystallographic studies of metal-substituted enzymes. J. Am. Chem. Soc. 125, 1041–1055 (2003).
Murakawa, T. et al. Quantum mechanical hydrogen tunneling in bacterial copper amine oxidase reaction. Biochem. Biophys. Res. Commun. 342, 414–423 (2006).
Murakawa, T. et al. High-resolution crystal structure of copper amine oxidase from Arthrobacter globiformis: assignment of bound diatomic molecules as O2. Acta Crystallogr. D Biol. Crystallogr. 69, 2483–2494 (2013).
Murakawa, T. et al. Probing the catalytic mechanism of copper amine oxidase from Arthrobacter globiformis with halide ions. J. Biol. Chem. 290, 23094–23109 (2015).
Murakawa, T. et al. In crystallo thermodynamic analysis of conformational change of the topaquinone cofactor in bacterial copper amine oxidase. Proc. Natl. Acad. Sci. USA 116, 135–140 (2019).
Murakawa, T. et al. Neutron crystallography of copper amine oxidase reveals keto/enolate interconversion of the quinone cofactor and unusual proton sharing. Proc. Natl. Acad. Sci. USA 117, 10818–10824 (2020).
Murakawa, T. et al. Serial femtosecond X-ray crystallography of an anaerobically formed catalytic intermediate of copper amine oxidase. Acta Crystallogr. D. Struct. Biol. 78, 1428–1438 (2022).
Murakawa, T. et al. Neutron crystallography of a semiquinone radical intermediate of copper amine oxidase reveals a substrate-assisted conformational change of the peptidyl quinone cofactor. ACS Catal. 13, 12403–12413 (2023).
Murakawa, T. et al. Microcrystal preparation for serial femtosecond X-ray crystallography of bacterial copper amine oxidase. Acta Crystallogr. F. Struct. Biol. Commun. 77, 356–363 (2021).
Shimizu, E. et al. Purification and properties of phenylethylamine oxidase of Arthrobacter globiformis. Biosci. Biotechnol. Biochem. 61, 501–505 (1997).
DePonte, D. P. et al. Gas dynamic virtual nozzle for generation of microscopic droplet streams. J. Phys. D Appl. Phys. 41, 195505 (2008).
Tono, K. et al. Diverse application platform for hard X-ray diffraction in SACLA (DAPHNIS): application to serial protein crystallography using an X-ray free-electron laser. J. Synchrotron Radiat. 22, 532–537 (2015).
Kataoka, M. et al. Detection of the reaction intermediates catalyzed by a copper amine oxidase. J. Synchrotron Radiat. 18, 58–61 (2011).
Juers, D. H. & Ruffin, J. MAP_CHANNELS: a computation tool to aid in the visualization and characterization of solvent channels in macromolecular crystals. J. Appl. Crystallogr. 47, 2105–2108 (2014).
Wilce, M. C. et al. Crystal structures of the copper-containing amine oxidase from Arthrobacter globiformis in the holo and apo forms: implications for the biogenesis of topaquinone. Biochemistry 36, 16116–16133 (1997).
Parsons, M. R. et al. Crystal structure of a quinoenzyme: copper amine oxidase of Escherichia coli at 2 A resolution. Structure 3, 1171–1184 (1995).
Shepard, E. M. & Dooley, D. M. Intramolecular electron transfer rate between active-site copper and TPQ in Arthrobacter globiformis amine oxidase. J. Biol. Inorg. Chem. 11, 1039–1048 (2006).
Smith, N. et al. Changes in an enzyme ensemble during catalysis observed by high-resolution XFEL crystallography. Sci. Adv. 10, eadk7201 (2024).
Butryn, A. et al. An on-demand, drop-on-drop method for studying enzyme catalysis by serial crystallography. Nat. Commun. 12, 4461 (2021).
Ishikawa, T. et al. A compact X-ray free-electron laser emitting in the sub-ångström region. Nat. Photon. 6, 540–544 (2012).
Wang, D., Weierstall, U., Pollack, L. & Spence, J. Double-focusing mixing jet for XFEL study of chemical kinetics. J. Synchrotron Radiat. 21, 1364–1366 (2014).
Kubo, M. et al. Nanosecond pump-probe device for time-resolved serial femtosecond crystallography developed at SACLA. J. Synchrotron Radiat. 24, 1086–1091 (2017).
Kameshima, T. et al. Development of an X-ray pixel detector with multi-port charge-coupled device for X-ray free-electron laser experiments. Rev. Sci. Instrum. 85, 033110 (2014).
Yumoto, H. et al. Focusing of X-ray free-electron laser pulses with reflective optics. Nat. Photon. 7, 43–47 (2013).
Nakane, T. et al. Data processing pipeline for serial femtosecond crystallography at SACLA. J. Appl. Crystallogr. 49, 1035–1041 (2016).
Joti, Y. et al. Data acquisition system for X-ray free-electron laser experiments at SACLA. J. Synchrotron Radiat. 22, 571–576 (2015).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014).
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Crystallogr. 45, 335–341 (2012).
Yefanov, O. et al. Accurate determination of segmented X-ray detector geometry. Opt. Express 23, 28459–28470 (2015).
Gevorkov, Y. et al. XGANDALF—extended gradient descent algorithm for lattice finding. Acta Crystallogr. A Found. Adv. 75, 694–704 (2019).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Beilsten-Edmands, J., Parkhurst, J. M., Winter, G. & Evans, G. Processing serial synchrotron crystallography diffraction data with DIALS. Methods Enzymol. 709, 207–244 (2024).
Acknowledgements
Diffraction experiments were performed using BL2 at SACLA and BL44XU at SPring-8 with the approval of the Japan Synchrotron Radiation Research Institute (proposal 2017B8038, 2020A6507, 2020A8013, 2021A6606, 2021A8013, 2021B6606, 2021B8029, 2022A6705, 2022A8020, 2022B6705, 2022B8013, 2023A6805, 2023A8009, 2023B6805, 2023B8013, 2024A8021, 2024B8016, and 2025B6504). The authors are grateful to the SACLA and SPring-8 staff for their assistance and guidance. This study was supported by the Platform Project for Supporting Drug Discovery and Life Science Research of the Japan Agency for Medical Research and Development under grant numbers JP21am0101070 (S.I.), JP22ama121001 (E.N.), by JSPS KAKENHI Grants 20H05448, 22H04757, 23K18117, and 24K01687 to Tak. M. from Japan Society for the Promotion of Science, and by the Cooperative Research Program of Network Joint Research Center for Materials and Devices (MEXT) under grant number 20251295 to Tak. M.
Author information
Authors and Affiliations
Contributions
Tak.M., M.Suz., M.Sug., E.N., K.Ta., and T.Ok. conceived the study; Tak.M., M.Suz., and T.Ok. designed the experiments; Tak.M. purified the protein; Tak.M. and T.A. prepared the microcrystals; Tak.M., M.Suz., K.F., T.Ma., E.M., I.M., I.K., Tai.M., H.M., Y.M., N.N., T.Oo., K.S., T.S., T.H., T.To., and T.Ok. collected the data at SACLA; Tak.M. and M.Suz. processed the data; Tak.M. conducted structural analysis; M.Suz. and T.Sun. performed the computations; T.K., K.To., T.Ta., M.Sug., L.F., J.K., R.T., S.I., and E.N. developed the SFX systems at SACLA; Tak.M., M.Suz., T.Y., K.Ta., and T.Ok. managed the project; Tak.M. wrote the original draft; Tak.M., K.Ta., and T.Ok. prepared the final version of the manuscript and figures with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Murakawa, T., Suzuki, M., Fukui, K. et al. Real-time capture of domain movements during copper amine oxidase catalysis by mix-and-inject serial crystallography. Nat Commun 16, 11149 (2025). https://doi.org/10.1038/s41467-025-67230-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67230-5