Abstract
Since their discovery, CRISPR systems have been repurposed for programmable targeted genomic editing, leading to applications for gene disruption, single base editing, insertion, deletion, and manipulation of short genomic sequences. Pairing Cas9 nickase with reverse transcriptase allows applications for insertion, substitution, and deletion of short genomic sequences from an RNA template without generating double stranded breaks however this technology typically shows reduced efficacy in post mitotic cells, limiting its translatability in vivo. Here we present a novel, ligase-based method that addresses these limiations. We introduce edits through delivery and ligation of a synthetic DNA donor to genomic nicks created with Cas9 nickase and report editing activity in cell lines, primary cell cultures, and adult mice via nonviral delivery. With favorable on target outcomes compared to transcription-based editing in key cell types, good tolerability, and deliverability, ligation-mediated gene editing has the potential to further advance genomic medicine.
Similar content being viewed by others
Introduction
CRISPR/Cas9 revolutionized genome engineering due to its ease of retargeting, requiring only modifying a portion of the RNA sequence of CRISPR/Cas9 to match the genomic target1,2. Cas9 was suited to gene disruption as DNA cleavage resulted in double-strand breaks (DSBs). However, deleterious side effects including heterogenous on-target edit outcomes containing insertions or deletions (indels), potential off-targets, chromosomal rearrangements, and even chromosomal loss, discouraged its use for many genetic diseases3. Engineering of the CRISPR/Cas9 system led to tools with greater finesse, such as single-strand nicking (nCas9) or complete catalytic inactivation (dCas9)2. dCas9 coupled with deaminases enabled specific single base changes4,5 and combination of nCas9 with writing enzymes such as a Moloney murine lentiviral reverse transcriptase (MMLV-RT, RT) and PhiC29 DNA polymerase allowed writing of any small corrections6,7. However, transcription-based editing is limited by decreased efficiencies of long edits ( > 10 nucleotides (nt)) and poor performance in primary cells, likely due to decreased intracellular dNTP pools8,9. To circumvent these observations, we sought out an editing method independent of exogenous writing enzymes.
When nCas9 nicks the genome, it leaves behind a 3’ hydroxyl group6. This moiety can be extended by RT but is also a suitable substrate for DNA ligase, which in the presence of a heterologous DNA donor, can result in DNA insertion. Pre-synthesizing the donor removes the uncertainty of endogenous dNTP availability and simplifies writing to a single step. To bring together guide and donor, we devised a splinting DNA (splint) containing three hybridization domains that bind to the donor, the ligase-mediated guide RNA (lmgRNA), and the genomic target site. These three interactions serve two key functions: efficient delivery of the donor via nCas9 ribonucleoprotein (RNP) nuclear localization and alignment of the donor to the nicked genomic flap for ligation. The inclusion of a nicking guide (ngRNA) targeting the opposite strand approximately 50 – 100 base pairs (bp) away from the edit site improves efficiency and is employed for all method comparisons.
Through chemical and sequence optimization of each oligonucleotide component to improve affinity, stability, and activity, we report efficiencies of small corrections (1–3 nt) in therapeutically relevant loci up to 52.4% in HEK293T cells and 66.0% in primary human hepatocytes (PHH) with up to 100.0% fidelity (percentage of edits lacking any indels of total edits). Pairing two complexes together (paired L-PGI, pL-PGI), we observe up to 82.4% total efficiency and 62.7% fidelity for placement of a 38 bp Bxb1 integrase recognition site (attB) in PHH. Using pL-PGI to place attB leads to 14.4% on-target integration of a 30 kilobase (kb) transgene cargo when co-delivering integrase and helper-dependent adenovirus template (HDAd), representing 59.6% usage of available attBs. pL-PGI demonstrates good translatability across important model species in vitro, resulting in similar or significantly superior efficiency and fidelity compared to prime editing (PE). Lastly, we present mouse data showing 7.9% attB placement using pL-PGI compared to 2.5% by dual PE, demonstrating that the advantages of L-PGI seen in vitro translate in vivo. In summary, due to its good tolerability, high potency, favorable fidelity and compatibility with therapeutically relevant delivery modalities such as lipid nanoparticles (LNP), we believe L-PGI has the potential to expand the gene editing toolbox and enable the development of treatments for previously inaccessible genetically-driven diseases.
Results
L-PGI mechanism and optimization
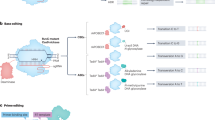
Ligase-mediated programmable genomic integration (L-PGI) capitalizes on nicking of the non-target strand of the genome by nCas9, leaving a free hydroxyl group on the 3’ DNA flap6. DNA ligase recognizes the free 3’ DNA end and can directly modify the genome using a donor DNA sequence containing a free 5’ phosphate. After ligation, incorporation of the free 3’ end of the donor into the genome is aided by a microhomology arm (10–20 nt) downstream of the edit. Using a complementary DNA splint that contains a donor binding site (DBS), genomic flap binding site (FBS) and guide binding site (GBS) to link the donor, genome and 3’ end of a single guide RNA, all the editing components can be colocalized to enable efficient gene editing (Fig. 1a). The newly ligated donor DNA then competes with the endogenous strand, with successful incorporation resulting in editing (Fig. 1b)6. Using a Cy5-labeled dsDNA substrate in a biochemical assay, we confirmed that all reaction components were necessary for editing (Lane 3, Fig. 1c).
A Schematic diagram illustrating the L-PGI editing complex and binding of all hybridization domains in oligonucleotides with star indicating ligation site. B Diagram of edit incorporation for an A to C transversion. The donor DNA (green) contains the desired edit with homologous sequence arms to the left and the right of the base edit. After ligation, the donor competes with the endogenous strand for genome incorporation via the mismatch repair (MMR) pathway. C Validation of edit strategy using recombinant protein and synthetic oligonucleotides in vitro visualized by SDS-PAGE. Lane 1: 50 nt and 200 nt DNA fragment positive controls representing the expected edit products. Lanes 2–5: L-PGI reactions containing (+) or excluding components (-) as labeled. Lane 3 (red) shows expected edit product only when all components are included. D Optimization of DBS/SBS length with statistical comparison between 24 vs 26 nt and 26 vs 32 nt. E Optimization of FBS length with statistical comparison of 9 vs 11 nt and 11 vs 13 nt. F Effect of splint GBS and lmgRNA SBS lengths. Red = locked nucleic acid (LNA), blue = 2’ O-methylation, purple = ribonucleic acid (RNA), black = DNA base, and * = phosphorothioate bond. Nucleotide sequences are shown as they are bound to each other and in some cases a part of the GBS or SBS functions as a single stranded linker. All splints include the same DBS and FBS (not shown) and lmgRNAs include identical scaffolds and spacers (not shown). G Testing linked and split mRNA architecture with varying placements of leucine zippers (LZ), P2A, or XTEN linkers on nCas9 and either T4, SplintR, or human ligase IV (hLig4) truncations. Untreated control represents untransfected cells. D–G utilize the HEK293T GFP reporter cell line assayed by flow cytometry and show results of biological replicates n = 3. Ordinary one-way ANOVA without matching for Gaussian distributions used to calculate all p values. H Summary of optimization results. All error bars represent standard deviation. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-3, 6.
We leveraged a HEK293T reporter cell line containing a blue fluorescent protein (BFP) that converts to green fluorescent protein (GFP) upon successful editing of 3 nt at positions +2, +4, and +5 from the nick for further development (Supplementary Fig. 1a)10. Initial experiments using an unmodified DNA splint did not yield detectable GFP. However, substituting locked nucleic acids (LNAs) in the GBS and DBS regions of the splint led to signal, likely due to increased stability of the splint and its affinity for the donor and lmgRNA (Supplementary Fig. 1b)11. We then systematically varied the lengths of the DBS, FBS, and GBS and found that 22–26 nt, 11–13 nt and 19–20 nt resulted in the highest GFP conversion for the three sequence components, respectively (Fig. 1d-f). Optimization of LNA position and number led to a splint design with LNAs alternating throughout the DBS and in the 3’ end of the GBS (Supplementary Fig. 1c). Addition of LNAs in the FBS did not improve efficiency, perhaps due to undesirable spacer-FBS binding (Supplementary Fig. 1d). Moving forward, we selected a 19 nt GBS, 13 nt FBS, and 24 nt DBS, although locus-specific sequence optimization is expected to further improve the system for individual targets.
We next explored other avenues for improving the L-PGI system, including enzyme selection, mRNA architecture and in vitro transcription (IVT) optimization. Initial work employed the well-characterized, highly efficient T4 ligase12. Compared with other ligases, including human DNA ligase IV (hLig4) and PBCV-1 DNA ligase (SplintR), T4 ligase consistently led to the highest GFP signal (Fig. 1g)13. We then investigated colocalization and fusion of nCas9 and ligase, as these strategies had been successfully for other genome editing technologies6. However, linking nCas9 and T4 ligase through either a flexible peptide linker, self-cleaving peptide, or leucine zippers (LZ) had either no effect or a detrimental effect on editing efficiencies (Fig. 1g)14. When LZ-based linking was used for SplintR, we found that LZ placement strongly impacted efficiency in an orientation-dependent manner (Supplementary Fig. 1e). We therefore included LZ on the C-terminus of ligase and N-terminus of nCas9 for all designs, hypothesizing that the gains for the lower efficiency ligase revealed efficiency improvements obscured by the high activity of T4 ligase. Finally, we found that including N1-methylpseudouridine during mRNAs production led to an almost 2-fold increase in GFP compared to uridine (Supplementary Fig. 1f)15. Based on these findings, we used split LZ co-localized nCas9 and T4 ligase translated from N1-methylpseudouridine mRNAs for subsequent L-PGI target efficiency analysis.
In the absence of cellular machinery, editing by L-PGI is dependent on all the components of the system. In cells, some editing was achieved when T4 ligase or phosphorylation of the 5’ end of donor DNA were omitted, implying that endogenous DNA repair can aid in L-PGI (Supplementary Fig. 1g). We determined that these events were not due to the splint acting as a template for homology-directed repair (HDR) by comparing L-PGI oligonucleotides with a BFP to GFP HDR donor. Lastly, Cas9 co-delivered with splint and donor produced no significant editing, suggesting that L-PGI edit is independent of HDR (Supplementary Fig. 1h).
L-PGI for therapeutic targets
L-PGI can correct point mutations in disease-associated loci and introduce longer edits. We explored a set of disease-relevant mutations in HEK293T16,17,18, and after extensive optimization of chemical modifications, enzyme architecture, and transfection conditions, were able to reach precise editing efficiencies of 25.8% to 52.4% for three different loci with <1.5% indel generation for each edit (Supplementary Fig. 2a-d). To enhance translatability into in vivo models and therapeutic applications, we focused subsequent work on liver-associated genomic targets in non-dividing primary human hepatocytes (PHH) and devised an edit quality metric, fidelity, defined as the proportion of sequencing reads aligned to the edit outcome with no insertions or deletions. L-PGI oligonucleotide components were designed based on high efficiency nCas9 spacers to install disease-relevant mutations in wild-type cells, including H1069Q in ATP7B for Wilson’s disease and C282Y in HFE for hemochromatosis17,19. In PHH, L-PGI demonstrated a 10-fold advantage in editing efficiency over PEMax at ATP7B, reaching 19.9% efficiency with 97.9% fidelity (Fig. 2a)20. Finding low efficiencies with initial PE controls, we moved to comparisons with internally engineered nCas9-RT variants with higher editing efficiencies, PE-TB1-3 (Fig. 2b)21. Against PE-TB2, L-PGI maintained a significant lead performing the 2nt mutation at HFE (Fig. 2c).
Results shown for n = 3 biological replicates for all experiments unless otherwise indicated. A Efficiency and fidelity of 2 nt substitution of +5 G to T and +8 G to T to install H1069Q with silent mutation in ATP7B assayed by next generation sequencing (NGS). L-PGI was compared to PE2, PE3, and PEMax. Ordinary one-way ANOVA without matching with Gaussian distribution used to calculate p value for efficiency comparison between PEMax and L-PGI. B Comparison of engineered nCas9-RT variant controls with published mRNA sequences for placement of 38 bp Bxb1 attachment site (attB) in F9 intron 1 via twin PE mechanism using paired guides with 20 overlap in PHH assayed by digital droplet PCR (ddPCR). Unpaired T test and ordinary one-way ANOVA with Gaussian distribution performed, respectively. n = 2 for PE6 test condition only. C Efficiency and fidelity of 2 nt substitution of UGC to UAU to install C282Y with silent mutation in HFE assayed by NGS. L-PGI is compared to PE-TB2 (nCas9 fused to engineered RT). Ordinary one-way ANOVA performed for efficiency to calculate p values, ns indicates not significant. n = 2 for L-PGI test condition only. D Schematic map showing location of guide target window in a 900 bp range spanning the transcription start site (TSS) preceding exon 2 of APOA1. E Efficiency of 1–3 nt substitutions performed by either PE or L-PGI shown side by side using the same spacer sequences for either pegRNA or lmgRNA and identical ngRNA by Sanger Sequencing and inference of CRISPR edits (ICE) analysis against untreated control. Paired T test performed with estimation plot showing up to 54.3 percentage gain in efficiency with average gain of 13.1 across all tested spacers. F Edit quality comparison for subset of APOA1 spacers showing comparable fidelity across loci and methods. SP4275 PE-TB3, SP4286, and SP4275 L-PGI test conditions contain n = 2 biological replicates. All error bars represent standard deviation. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-4.
We then explored the application of precision gene editing for modification of target gene expression as a potential therapeutic strategy for non-monogenic disorders. We selected APOA1 as proof of concept to treat cardiovascular disease22 and screened spacers within the promoter region to induce 1 – 3 nt edits with either L-PGI or PE. These edits would result in sequences resembling transcription factor (TF) binding motifs, which might upregulate APOA1 expression, leading to therapeutic benefit (Fig. 2d). We found up to 66.0% efficiency with L-PGI compared to 48.0% with PE-TB3 at the same cut site and an average of 13.1-point gain across all sites with comparable edit quality (Fig. 2e, f).
Further, we explored the possibility of placing entire TF binding sites. By shifting the placement of the homology arm in the donor or RT template, we attempted 14 bp edits as either a complete insertion, shifting endogenous sequence downstream, or as a substitution by simultaneously excising endogenous sequence of the same length (Fig. 3a). We found up to 34.9% total editing and superior fidelity using L-PGI compared to 5.0% using RT-based (pegRNA) editing (Fig. 3b, Supplementary Fig. 3a-d). Insertion generally outperformed sequence substitution both by ligation and RT-based methods likely due to the favorability of strand displacement closer to the nick site. We found that L-PGI maintained a lower indel rate across all small insertion types tested (Fig. 3c). The additional errors from PE appear as a combination of substitutions and indels and are likely due to RT errors (Fig. 3d). Although substitution errors also occur in L-PGI, they are likely the result of errors during chemical synthesis, which could be minimized with higher purity donors. We then tested longer insertions, including a 38 bp Bxb1 integrase attachment site (attB), and found up to 11.8% correct editing with L-PGI compared to 0.5% with RT (Fig. 3e). Though the edit outcomes for attB insertion via L-PGI were still favorable compared to RT, we observed a reduction in efficiency consistent with results in the GFP reporter HEK293T line using longer donor lengths.
A Illustration showing the 14 nt edit with and without excision of endogenous sequence representing end cases preserving either endogenous sequence or reading frame. The endogenous sequence downstream of the nick is shown in blue and red. The donor contains the 14 nt insertion followed by sequence homologous to the red for sequence replacement or to the blue for sequence insertion. B Highest total efficiencies and maximum fidelities observed using either sequence replacement or insertion observed using either PE-TB2 or L-PGI in PAH intron 1 obtained through component ratio optimization. Ordinary one-way ANOVA with Gaussian distribution performed for biological replicates n = 2 with p values shown. Efficiencies and fidelity vary depending on component ratio, for complete data set refer to Supplementary Fig. 3. C Comparison of lowest indel generation rates observed between PE-TB2 and L-PGI. Indel rate is calculated as the rate of total erroneous deletions and insertions occurring within the nick to nick edit window normalized to total edited reads. P values determined with Ordinary one-way ANOVA with Gaussian distribution for multiple comparisons. n = 3 for HFE 2nt edit PE-TB2 condition and 2 for all other test conditions. D Representative NGS paired-end alignment showing top 15 reads each of 14 nt insertions in PAH intron 1 of PHH using L-PGI (top) and PE-TB2 (bottom). Black bars indicate deletions, highlighted bases show substitutions, and underlined bases contain insertions. In L-PGI the dominant errors are point substitutions in the donor region (24 nt in red in reference alignment) and excisions between the end of the donor and the nicking site on the opposite strand (2 nt in red). PE results in additional insertions and deletions in the edit window. E Comparison of 38 nt Bxb1 attB placement efficiency in PAH intron 1 using L-PGI or PE-TB2 by NGS showing correct editing rate only. Ordinary one-way ANOVA with Gaussian distribution performed with n = 3. All error bars represent standard deviation. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-4.
Paired L-PGI
To combat the efficiency drop with longer edits, we paired two L-PGI complexes (paired L-PGI, pL-PGI) to simultaneously introduce edits on opposite strands of the genome (Fig. 4a). Donors can be designed either containing exogenous sequence for gene replacement, or complementary to the genome beyond the opposing nick for precise deletion (Fig. 4b). In both scenarios, the genomic region between the two nicks is excised by 5’ exonuclease degradation23, resulting in a size change detectable by gel electrophoresis following target amplification24,25. We used pL-PGI to either delete the C9ORF72 hexanucleotide repeat expansion in HEK293T cells (Supplementary Fig. 4a) or replace the 131 bp region with a 38 bp Bxb1 attB site (Supplementary Fig. 4b). We quantified the frequency of these and additional edits using NGS amplicon sequencing and found that pL-PGI could install a 33 bp Pa01 attB site at NOLC1 locus in HEK293T cells with ~1.5–2-fold efficiency improvement over RT-based editing and ~60% reduction in indels (Fig. 4c)26.
A Schematic diagram of paired L-PGI complexes targeting opposite strands to install paired reverse complementary donors. B Design strategies demonstrating replacement type edit (left) by encoding the desired edit in reverse complementary partial overlap donors or deletion type edit (right) by having donors homologous to the genome flanking the sequence to be excised. C Pa01 33 bp attB placement efficiencies using paired complexes PE2, PEMax, or pL-PGI in NOLC1 of HEK293T measured by NGS for biological replicates n = 3. Control defined as untransfected cells. Ordinary one-way ANOVA for multiple comparisons with Gaussian distribution performed for precise edit efficiencies by edit method with p values shown. D Efficiencies of Pa01 33 bp attB placement in NOLC1 of hematopoietic stem cells (HSCs), Bxb1 38 bp attB placement in B2M of induced pluripotent stem cells (iPSCs), and 42 bp edit containing Bxb1 38 bp attB with TAAT stop codon in CIITA of iPSCs, all results assayed by ddPCR. Control conditions received electroporation only with no transfection reagents. Each condition performed once n = 1. E Efficiencies of 175 bp excision at VEGFA of HEK293T, iPSC, and PHH by either dual PE2 or pL-PGI by NGS for biological replicates n = 3. All error bars represent standard deviation. Source data are provided in Source Data file. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-4.
Evaluation in additional cell types showed that pL-PGI is also active in hematopoietic stem cells (HSCs) and induced pluripotent stem cells (iPSCs), achieving up to 21.2% and 16.9% attB placement, respectively without selection (Fig. 4d). pL-PGI was also used to delete 175 bp at the VEGFA locus, leading to 3-fold higher editing in HEK293T, 1.6-fold higher editing in iPSCs, and 16-fold higher editing in PHH compared to RT editors (Fig. 4e). To test the limit of edit size, we attempted a 345 bp miniGFP insertion in HFE using pL-PGI with 55 nt overlap in PHH and detected up to 0.5% editing by ddPCR, likely the result of limited delivery of long donors (Supplementary Fig. 4c)27.
The use of DNA donors and splint molecules has the potential to activate innate immune responses in treated cells. To investigate this, we examined pL-PGI cytotoxicity in PHH using an ATP-based cell viability assay and found no significant decrease in cell health using optimized transfection conditions (Supplementary Fig. 5a, b). Assessment of potential innate immune response using an IFN-beta enzyme-linked immunoassay also yielded no significant response in PHH (Supplementary Fig. 5c).
pL-PGI for integrase mediated genomic insertion
To optimize pL-PGI for integration in PHH, we symmetrically walked the overlap around the central dinucleotide while maintaining full complementarity between the splint and donor and found that 10 bp showed both best efficiency and fidelity (Fig. 5a). We found that 20 bp overlap efficiencies could improve by decreasing just the splint DBS length, thereby minimizing unwanted LNA exacerbation of splint secondary structure (Supplementary Fig. 5d). We also explored placement of the Bxb1 attP attachment site due to a potentially favorable off-target cryptic site profile in the human genome compared to attB28. We varied the donor overlaps for the 52 bp attP sequence in a similar manner and saw that fidelity was correlated with donor length while efficiency was inversely related (Supplementary Fig. 5e). From our attB and attP insertion studies, we concluded that optimal editing can be achieved with 10 bp overlap attB.
All results show biological replicates of n = 3 unless otherwise stated. A Efficiency and fidelity of attB placement with symmetrically varying overlap for forward and reverse donors in F9. Donor and splint DBS were kept locked to maintain full donor splint hybridization (24, 29, 38 nt for 10, 20, and 38 overlap, respectively). Control is untreated cells. Each condition for efficiency testing performed n = 2 and for fidelity, n = 6. % edit quantified as precise editing only and % fidelity quantified as precise editing out of total editing for all groups. Ordinary one-way ANOVA for multiple comparisons with Gaussian distribution used to calculate p values. B Application of 10 overlap pL-PGI for attB placement in F9, Albumin (ALB), and PAH in PHH versus dual 20 overlap PE-TB2 showing frequency of precise editing only by NGS. C Schematic illustration of 2-step attB placement followed by Bxb1-mediated integration of viral cargo showing expected genomic products. ITR = inverted terminal repeats, attP = attachment site P, attL = post recombination left junction, attR = post recombination right junction. D Residual unconverted attB and total gene integration efficiency of helper-dependent adenovirus (HDAd) comparing attB placed by either dual PE or pL-PGI. Sum of both bars represents total edit efficiency for each attB placement method. Control is untreated cells. E Comparison of attB conversion rate (calculated as ratio of integration to integration + residual attB) between dual PE and pL-PGI across both left and right junctions. Ordinary one-way ANOVA for multiple comparisons used for calculation of p values. All error bars indicate standard deviation. Source data are provided in Source Data file. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-4, 7.
We next compared optimized configurations of both pL-PGI and dual PE-TB2 at therapeutic loci F9, Albumin, and PAH29,30,31. We found 20.7–38.8% total editing efficiencies with pL-PGI, matching or outperforming RT-based editing at all loci tested (Fig. 5b). The fidelity of RT-based editing appears to be more locus-dependent than pL-PGI, while both methods see reduced fidelity compared to single complex editing due to the possibility of NHEJ repair prior to either edit installation or overlap hybridization.
Initial efforts using fully methylated donors for integrase-mediated gene insertion found low attB conversion after 2-step PGI (Fig. 5c)32,33. Hypothesizing that chemical modifications were recruiting methyl-CpG-binding proteins thereby blocking integrase access, we then screened additional chemical modifications for attB placement in PHH and found that removal of all methylation while retaining two phosphorothioate bonds resulted in preservation of donor stability, edit efficiency, and improved Bxb1 activity in cells (Supplementary Fig. 5f, g). Using this donor modification pattern, we were able to install 38 bp attB into intron 1 of PAH and in the presence of Bxb1 mRNA and helper-dependent adenoviral (HDAd) DNA cargo (Supplementary Fig. 5h), measured 14.4% whole gene insertion and 59.6% attB conversion by ddPCR (Fig. 5d, e, Supplementary Fig. 5i). We found higher attB conversion with pL-PGI than RT-based editing, consistent with the improvements in fidelity seen with pL-PGI.
Translating pL-PGI across species in vitro
Using appropriately designed lmgRNAs, donors and splints, we were able to efficiently install 38 bp attBs into intron 1 of the PAH gene in primary mouse hepatocytes (PMH) and primary cynomolgus hepatocytes (PCH), both important surrogate cell types for animal models (Fig. 6a). Comparing pL-PGI to optimized dual PE-TB2 attB insertion, we observed similar correct and total edit efficiency in both PCH and PHH. However, in PMH we observed almost no correct insertion using dual PE-TB2, while pL-PGI was able to achieve 10.0% placement of functional attBs. pL-PGI and dual PE-TB2 showed comparable fidelity in PCH but differed in PHH and PMH, with 1.5-fold and 10-100-fold higher fidelity, respectively (Fig. 6b). Additionally, pL-PGI and dual PE-TB2 lead to different imperfect editing outcomes (Supplementary Fig. 6a,b). RT-based insertion showed partially written sequence or incorrect sequence that is likely from incorrect reverse transcription or partial strand degradation. pL-PGI generally results in insertions containing the entire forward or reverse donor with blunt end joining to the second nick site, reflecting the benefit of pre-synthesized donors against de novo transcription in cells.
A Perfect edit and indel efficiencies at PAH intron 1 in PHH, primary cynomolgus hepatocytes (PCH), and primary mouse hepatocytes (PMH), respectively using guides targeting the identical nicking sites for both edit methods in each comparison. PHH and PMH show results of biological replicates n = 3 and PCH results of biological replicates n = 2. Ordinary one-way ANOVA with Gaussian distribution performed for all analyses. B Fidelity comparison between pL-PGI and dual PE-TB2 for the same edits in PHH, PCH, and PMH, respectively, calculated by taking the percent of perfect beacons out of total edits. PHH and PMH show results of biological replicates n = 3 and PCH results of biological replicates n = 2. C pL-PGI LNP formulation payload schematic depicting all in one (AIO) delivery and split delivery. AIO: single particle with all 8 components: nCas9 mRNA (1), ligase mRNA (2), forward lmgRNA (3), reverse lmgRNA (4), forward splint (5), reverse splint (6), forward donor (7), and reverse donor (8). Split: two particles each containing half of both mRNAs and either forward guide, forward splint, forward donor or reverse guide, reverse splint, and reverse donor. D Potency of dual PE-TB2 AIO versus AIO and split pL-PGI LNP targeting F9 in PHH assayed by ddPCR showing biological replicates n = 3 for each treatment dose. 4-parameter nonlinear curve calculated showing EC50 (ng/well) and R squared goodness of fit for each group. E Dual PE-TB3 vs pL-PGI for attB placement in mouse PAH assayed by ddPCR. Ordinary one-way ANOVA performed on biological replicates n = 5 for all groups. Control group treated with saline vehicle. F PE-TB3 vs L-PGI for insertion of a 14 nt edit in mouse PAH assayed by ddPCR showing 2-fold gain in efficiency. Ordinary one-way ANOVA performed on biological replicates n = 5. G Alanine transaminase (ALT) and aspartate transaminase (AST) levels in units/L (U/L) in mouse serum collected 24 h and 7 days after LNP injection. Sampled from individual animals n = 3. Ordinary one-way ANOVA performed for p values. All error bars indicate standard deviation. All one-way ANOVA tests assume Gaussian distribution with no matching. Source data are provided in Source Data file. Source data are provided in Source Data file. Nucleotide reagents provided in Supplementary Data 1-4, 7.
Using pL-PGI for therapeutic indications also involves moving to higher purity reagents and formulation of the components in a relevant delivery modality such as lipid nanoparticles (LNP). We tested HPLC-purified synthetic F9 lmgRNAs and found a 1.5-fold improvement over crude lmgRNAs which translated to an overall ~3-fold improvement in attB placement efficiency compared to RT-based attB placement with purified synthetic atgRNAs in PHH (Supplementary Fig. 6c). We also tested the effect of purification on donors and found improvement to 82.4% total editing with 62.7% fidelity at PAH in PHH (Supplementary Fig. 6d). Next, we compared all in one (AIO) LNP and split LNP packaging designs for attB placement in F9, theorizing that separate formulations may minimize unwanted cross-hybridization between oligonucleotide components (Fig. 6c). We found AIO and split pL-PGI performed similarly, and both were more potent than the PE LNP (Fig. 6d). AIO was selected for in vivo for simplicity. Lastly, we formulated mouse F9 reagents into LNPs and found 10-fold higher attB placement in PMH with pL-PGI than dual PE-TB2, consistent with earlier MessengerMAX results (Supplementary Fig. 6e).
L-PGI and pL-PGI in vivo
We selected the lead L-PGI 14 nt insertion and pL-PGI attB placement architecture and compared their efficiency in vivo against PE-TB3 and dual PE-TB3, respectively. High purity guide RNAs were synthesized in-house for pL-PGI and dual PE-TB3 and other synthetic oligonucleotide components were provided by an external vendor (IDT) without purification. mRNAs and oligonucleotide components were co-formulated and dosed to adult mice. Using ddPCR as a preliminary readout, we found up to 12.1% attB placement in one animal with an average of 7.9% by pL-PGI and an average of 2.5% by dual PE-TB3, representing a 3-fold advantage (Fig. 6e). For 14 nt insertion, we found an average of 3.8% editing by L-PGI and 1.9% by PE-TB3, a 2-fold advantage (Fig. 6f). Comparison with PMH in vitro suggests that these efficiency differences are primary fidelity driven, though future deep sequencing would be essential to further interpretation.
While earlier experiments detected no toxicity in PHH, in mice pL-PGI resulted in elevated alanine transaminase (ALT) and aspartate transaminase (AST) levels compared to control groups at 24 h that subsided by 7 days (Fig. 6g). While this did not translate to any adverse reaction in animals or prevent editing, toxicity remains a potential risk that may be mitigated by further optimization of the system (allowing for decreased doses) or addition of immune suppression cotreatments.
Discussion
Large precise insertions using CRISPR/Cas9 based systems have been challenging using the available gene editing systems1,32,33,34,35,36. Here, we describe L-PGI and pL-PGI, two technologies that capitalize on the biochemistry of CRISPR/Cas9 nicking to ligate edits using synthetic DNA donors without requiring exogenous writing enzymes.
We first showed that L-PGI and pL-PGI are capable of a broad range of edits from point mutations to insertion of hundreds of bases in cycling cells and primary hepatocytes. L-PGI has significantly higher efficiency and fidelity vs RT systems for medium sized ( ~ 14 bp) edits, a class essential for applications such as endogenous gene augmentation via installation of transcription factor binding sites. The potential for long insertions can also be applied to circumvent the PAM availability restriction in Cas9 targeting. While we have not fully explored the maximum size limit of the L-PGI system, the ability to simultaneously delete and insert sequences potentially opens the door to exon replacements, which could treat a multitude of genetic diseases caused by repeat expansions37.
We then demonstrated through a series of optimizations in primary mouse, human, and cynomolgus monkey hepatocytes that L-PGI and pL-PGI are efficient methods for enabling multi-kilobase gene insertion via writing of integrase landing sites. pL-PGI leads to more accurate editing outcomes than RT-based editing, resulting in significantly higher Bxb1 integration efficiency. Due to its superior capacity for longer insertions without compromising efficiency, pL-PGI can alternatively place 52 bp attP for Bxb1 integration. As integrases are not cargo size limited, this technology can enable precise and directed whole gene replacement that restores function under endogenous gene regulation32.
Lastly, we evaluated the impact of reagent purity on potency, the feasibility of formulating 8 separate RNA and DNA components in a single LNP, and the efficiencies of L-PGI and pL-PGI in adult mice. Compared to RT-based editing, L-PGI resulted in 2-fold higher 14 nt insertion and pL-PGI resulted in 3-fold higher attB placement in the mouse PAH gene, successfully reproducing the gains seen in PMH. Though elevated liver toxicity markers were observed post injection, these effects were transient and may be reduced through further reagent optimization and component engineering. The use of DNA donors and splint molecules has the potential to activate the innate immune response, which could lead to toxicity. At the doses used in this work, we did not observe decreased cell health or IFN-beta activation from L-PGI reagents in PHH cells. It is possible that the DNA response will be cell type dependent and a further investigation into this issue is warranted.
While we did not explore off-target editing, we consider it to be critical for further development of L-PGI for therapeutic use. Due to the dependence on nicks (vs DSBs), we expect minimal off-targets, similar to what has been seen in the prime editing field38. Any off-target edits are likely to be driven by spacer specificity and specificity can be benchmarked using guides with known off-target sites such as HEK46. Given the lack of on-target editing when splint or donor is omitted and previous reports showing un-detectable off-targets using ssDNA donor editing (HDR)39, we anticipate L-PGI to have very low off-target risk. Nonetheless, this hypothesis will require experimental verification using orthogonal approaches to provide sufficient sequencing depth and quantifying power.
In summary, the presented body of work demonstrates that L-PGI is an effective and versatile gene editing system for in vivo liver therapeutics with key advantages in the described use cases. Future directions include off-target profiling, toxicity reduction, expansion of editing capabilities, and further application for novel therapeutic strategies. While its synthetic format is suitable for transient delivery systems such as lipid, peptide, or polymer-based delivery, viral delivery systems such as AAV present challenges. To leverage the favorable characteristics of AAV, re-configuring the system for viral compatibility may be desirable for some applications. Due to its efficacy even in challenging species for RT-based editing such as mouse, L-PGI may have greater universal efficacy and could be coupled with delivery advancements for future therapies targeting important cell types implicated in CNS disorders. By offering solutions to fundamental shortcomings of RT-based editing, L-PGI may play an important role in the next generation of gene editing therapies.
Methods
Animal work ethics statement
All animal experiments were approved by the Explora BioLabs Institutional Animal Care and Use Committee (IACUC) at the Charles River Accelerator and Development Laboratory (CRADL), Watertown, MA (Protocol No. EB17-004-302), and were conducted in accordance with institutional and federal guidelines for the humane care and use of laboratory animals. All work complied with the Guide for the Care and Use of Laboratory Animals (8th Edition, NIH) and the AVMA Guidelines for the Euthanasia of Animals (2020).
Plasmid template generation
Coding sequences were derived from Addgene # 132774, 174820, 207854 and synthesized as gBlocks (Integrated DNA Technologies, IDT) to be cloned into an IVT vector that contains a CMV enhancer and promotor, a single copy of the 5’UTR and two copies of the 3’UTR from the human beta globin gene, and polyA tail using restriction digest and Gibson Assembly. Plasmids were transformed into 5-alpha Competent E. coli (NEB, C2987H) and sequence verified by Elim Biopharm and scaled up with QIAGEN Plasmid Plus Midi Kit (QIAGEN).
In vitro transcription of mRNA
All mRNAs were generated via in vitro transcription (IVT) reactions using the HiScribe T7 High Yield RNA Synthesis Kit (New England Biolabs). Plasmid DNA containing coding sequences were linearized using an XbaI restriction site located immediately downstream of the polyA tail. Linearized plasmids were then purified via phenol:chloroform extraction followed by ethanol precipitation. mRNAs were produced via IVT reactions that contain Uridine-Triphosphate or N1-Methylpseudouridine-5’-Triphosphate (TriLink BioTech), and capped co-transcriptionally with CleanCap Reagent AG (3’ OMe) (TriLink BioTech). IVT reactions were incubated at 37 °C for 2 h, followed by DNAse I digestion of the template DNA. mRNA products were purified using LiCl precipitation, quantified by Qubit Fluorometric Quantification (Thermo Fisher Scientific), and checked for integrity by denaturing gel electrophoresis.
Preparation of synthetic oligonucleotides
Splints and donor DNAs were ordered from IDT as 100 nmole DNA oligos in IDTE buffer. The ssODN used for HDR was ordered from IDT as a 4nmole Ultramer DNA Oligo. lmgRNAs, pegRNAs, and atgRNAs were ordered from IDT as Custom Alt-R gRNAs, 10 nmol with standard desalting. lmgRNAs and atgRNAs were purified by HPLC. Nicking gRNAs were ordered from Synthego as synthetic sgRNA, 5 nmol. Synthego sgRNAs included Synthego’s standard scaffold and chemical modification pattern, while IDT gRNAs were custom specified. Splints and donor DNAs were annealed together in NEBuffer 2 at 2 μM by heating to 95 °C for 2 min and ramping down to 25 °C over a period of 30 min.
In vitro biochemical assays
In vitro reactions were performed by first incubating lmgRNA (30 nM final) and annealed splint and donor DNA (30 nM final) with recombinant S. pyogenes nicking Cas9 (nCas9; IDT; 30 nM final) for 10 min at room temperature, followed by the addition of T4 ligase (NEB; 200U final), ATP (1 mM final), and 5’-Cy5-labeled dsDNA substrate (3 nM Final). Reactions were carried out in the presence of NEB Buffer 3.1 (1x final) at 37 °C for 1 h (final volume of 10ul). Reactions were terminated by the addition of 0.5% SDS and 100ug/ml Proteinase K and incubated at 37 °C for 30 min. Reaction products were then combined with 2x formamide gel loading buffer (90% formamide; 10% glycerol; 0.01% bromophenol blue), denatured at 95 °C for 10 min, and separated by denaturing urea PAGE gel (15% TBE-urea, 55 °C, 200 V). DNA products were visualized by Cy5 fluorescence signal using a LI-COR Odyssey CLx imager.
General cell culture conditions
HEK293T cells were purchased from ATCC and cultured in DMEM (11965092, Gibco) with 10% FBS (A3160501, Gibco). HEK293T GFP reporter cell line was provided by the Corn Lab10 and maintained similarly. Cells were dissociated with 0.25% Trypsin-EDTA (15400054, Gibco) and seeded in 96-well PDL-coated tissue culture plates (354210, Corning) at a density of 25k cells per well. Cryopreserved primary human hepatocytes (HMCPMS Hu8403, HMCPTS Hu8449, Hu8450, Gibco) were recovered in Cryopreserved Hepatocyte Recovery Media (CM7000, Gibco) and plated at 42k cells per well in Hepatocyte Plating Media (A1217601, CM300, Gibco) on Collagen I coated 96 well plates (354407, Corning). Cryopreserved primary cynomolgus monkey hepatocytes (MKCP10 CY427, Gibco) were recovered in Hepatocyte Plating Media and plated to 48k cells per well. Cryopreserved primary mouse hepatocytes (MSCP10 MC945, Gibco) were recovered in Hepatocyte Plating Media and plated to 20k cells per well. 8 hrs post recovery primary hepatocytes were washed and cultured in maintenance media with fresh media given again at 24 h (CM400, A1217601, A2737501, Gibco). Cell lines were not authenticated or tested for mycoplasma contamination.
Adenovirus-associated virus (AAV) production
Cargo was designed as self-complementary AAV containing Bxb1 attP attachment site, splice acceptor, and codon optimized human PAH exon 2-13 coding sequence with HiBit reporter tag (Promega) and cloned into plasmid backbone with Gibson assembly. AAV was produced by triple transfection with LK03 rep/cap and helper plasmid (Aldevron SF058826) in 293AAV Cell line (Cell Biolabs AAV-100). Virus was treated with PEG and purified with Iodixanol gradient centrifugation. Yield was determined by ITR assay titer using ddPCR.
Helper-dependent adenovirus (HDAd) production
Cargo was designed containing Bxb1 attP attachment site, splice acceptor, and codon optimized human F9 exon 2-8 coding sequence and assembled into in Ad5 gutless plasmid backbone. Cargo was linearized and transfected in HEK293 116 Cre+ cells (Baylor University), followed by helper virus transduction. HDAd was propagated by serial coinfection and purified by CsCl ultracentrifugation. Yield was determined by GFP reporter assay titer using ddPCR.
HEK293T transfection
After 16–24 h, the medium was changed to OptiMEM (31985070, Thermo Fisher Scientific) and the cells were transfected at approximately 70% confluency. A transfection mix for a single well included 0.4 uL of Lipofectamine 2000 (11668019, Thermo Fisher Scientific), 67 ng total mRNA, 2.1 pmol total gRNA, and 0.38 pmol total annealed splint and donor DNA. After 16 h, medium was changed back to DMEM with 10% FBS and cells were cultured until 2 days post transfection. For BFP to GFP conversion experiments, cells were dissociated with 0.25% trypsin, mixed with culture medium for trypsin inactivation, and analyzed with Attune NxT Flow Cytometer (Thermo Fisher Scientific). When multiple mRNAs, lmgRNAs, pegRNAs, or splints and donor DNAs were used in a single transfection, each was added in an equal amount to reach the total dosage described. When nicking gRNAs were used, they were added at half the dosage of the lmgRNA or pegRNA to reach the total dosage described.
Induced pluripotent stem cell (iPSC) electroporation
Human cryopreserved iPSCs were purchased from Fujifilm Cellular Dynamics and cultured in Supplemented Essential 8 Basal Media (A15169-01, A15171-01, Thermo Fisher Scientific) until 50 – 70% confluent. Cells were dissociated using Accutase (AT-104, Innovative Cell Technologies), counted, and per reaction 1e5 cells were electroporated with Neon NxT Transfection System (Thermo Fisher Scientific) using conditions V = 1500 V, Width = 20 ms, Pulses = 1 pulse. For each reaction the transfection mix contained 3 ug of each mRNA, 50 pmol of each guide, and 25 pmol of each splint and donor. Cells were then transferred to 48 well plates precoated with Vitronectin (A31804, Gibco) containing basal media with Clone R2 (100-0691, Stem Cell Technologies). Cells were given daily media changes until collection for gDNA and ddPCR analysis 7 days post transfection.
Hemopoietic stem cell (HSC) electroporation
Human cryopreserved mobilized peripheral blood CD34+ stem cells (MLEG34005C) were purchased from CGT Global and thawed in 1% human serum albumin 25% solution (MSPP-800120, Gemini) in DPBS (37350, StemCell). Cells were resuspended in CD34+ cytokine cocktail supplemented SFEM II media and seeded at 5e5 cells/mL for 2 days. 1e6 cells per condition were electroporated with Neon NxT Transfection System using conditions V = 1500 V, Width = 20 ms, Pulses = 1 pulse. Each reaction contained 10 ug of each mRNA, 270 pmol of each lmgRNA, 150 ng each splint, and 300 ng each donor. Cells were recovered in 6 well plate containing 2 mL of SFEM II media with 1x SFT cytokines and collected 3 days post transfection for gDNA purification and ddPCR analysis.
Primary hepatocyte transfection
PHH, PCH, and PMH were transfected with 0.3 μL Lipofectamine MessengerMAX (LMRNA015, Thermo Fisher Scientific) for co-delivery of all RNA and DNA components. Per reaction, the transfection mix contained 150 ng nCas9 mRNA, 150 ng ligase mRNA, 100 ng lmgRNA, 50 ng ngRNA, 5 ng splint, and 10 ng donor unless otherwise stated for optimization experiments. For pL-PGI transfections, the amount of each splint and donor was halved to result in the same total dosage. PE control conditions contained 300 ng nCas9-RT fusion and 100 ng pegRNA, 50 ng ngRNA, or 100 ng each dual atgRNAs. Cells were taken down 3 days post transfection for analysis. For integration experiments, cells were co-transfected with 200 ng Bxb1 mRNA and 1e6 multiplicity of infection (moi) for AAV or 1e3 moi for HDAd 2-3 days post attB placement according to previously described procedures and taken down after an additional 5 days of culture.
Genomic DNA extraction
For HEK293T cells, genomic DNA (gDNA) was extracted by removing medium, resuspending cells in 30 uL QuickExtract (QE0905T, LGC Biosearch Technologies), and incubating at 65 °C for 15 min followed by 98 °C for 10 min. Primary hepatocytes, iPSCs, and HSCs were lysed in a similar manner and gDNA extraction was followed by magnetic bead cleanup (A63882 AMPure XP, Beckman Coulter).
Target amplification and next-generation amplicon sequencing (NGS)
Target regions were amplified with 12.5 μL Q5 Hot Start High-Fidelity Master Mix (M0494X, NEB), 2 μL purified genomic DNA, and 2.5 μL of each 10 μM primer stock with water in a final volume of 25 μL for 25 cycles using annealing temperatures for the gene-specific part of the primers calculated by NEB’s online tool (https://tmcalculator.neb.com/). Complete list of primers provided in Supplementary data file. Amplified targets were imaged with gel electrophoresis in a 2% agarose gel run at 300 V for 30 min and imaged on a ChemiDoc (Bio-Rad) or used as a template for the Illumina barcoding PCR 2, run using 2 μL PCR1 product, 2.5 μL premixed barcoding primers, 12.5 μL Q5 master mix and water to final volume 25 μL for 12 cycles. PCR2 products were bead purified, pooled with 10% PhiX, denatured with NaOH, and loaded on Illumina MiSeq Micro Kit v2 with paired end sequencing. R1 and R2 Amplicon sequencing data were combined and analyzed using CRISPResso2. Input reads were trimmed to remove any Illumina adapters, if present. Sequences were then mapped to the amplicon references, which included both the wildtype allele and a prebuilt edited allele. Editing efficiency was quantified as the number of reads aligning to the edited allele divided by the total number of aligned reads. Insertions and deletions (indels) within the designated quantification window were assessed using a customized script that parsed the detailed mapping information generated by CRISPResso2. For beacon placement, the quantification window encompassed the entire 38 bp beacon region, while for prime editing, the quantification window was defined as the reverse transcription (RT) template region. Editing fidelity was calculated as the proportion of reads aligned to the edited allele without indels, divided by the total number of reads aligned to the edited allele.
Sanger sequencing
Primers were designed to amplify a 500–800 bp region surrounding the edit site and target amplification was performed using Platinum Superfi II Master Mix (12368010, Invitrogen) with 2 μL purified gDNA, 1.25 μL 10 μM of each primer, 12.5 μL master mix, and water up to 25 μL for 25 cycles of 10 s at 98 °C, 10 s at 59 °C, 30 s 72 °C, and final extension of 5 min after initial denaturation. Samples were sequenced by Genewiz (Azenta) and analyzed using Synthego inference of CRISPR edits (ICE v3) tool (https://ice.synthego.com/#/).
Cell viability assay
Toxicity of transfection conditions in cells was measured by CellTiter-Glo 2.0 (G9241, Promega) following manufacturer’s methods using 100 μL of premixed reagent per well in 96 well white-walled plates (165306, Thermo Fisher Scientific). Luminescence was detected on GloMax Discover Microplate Reader (GM3000, Promega).
Cell-based immune response assay
PHH were transfected with paired forward and reverse splints, donors, or lmgRNAs with varying chemical modifications at 12.5 nM final concentration in cell culture media using MessengerMAX delivery. Positive control AAV was dosed at 1e6 particles per cell and negative control condition was given MessengerMAX only. Cell culture media was harvested 6 hrs post transfection and assayed for IFN-beta using Simple Plex Human IFN-beta Cartridge (SPCKB-PS-000934, ProteinSimple) on Ella - Automated Immunoassay Platform (600-100, ProteinSimple) according to the manufacturer protocol for cell culture supernatants and analyzed with Simple Plex v3.2 software.
Droplet digital polymerase chain reaction (ddPCR)
Custom primers and probes were designed to measure editing in all referenced loci. Results were normalized to custom reference assays targeting unedited regions of the same genes in the respective species. Probes were dual labelled with 3′- 3IABkFQ and either 5′-carboxyfluorescein (FAM) for edit targets or 5′-hexachloro-fluorescein phosphoramidite (HEX) for reference. Assays were validated using gBlocks representing edit outcomes to test for both specificity and linearity. All primers, probes, and gBlocks were synthesized by IDT. Each reaction contained 12 µL of 2x ddPCR Supermix for probes (No dUTP) (1863025 Bio-Rad), 1.2 µL of each primer and probe mix to final concentration of 0.5 uM for each primer and 0.25 uM for each probe, 0.12 µL each of HindIII and Eco91I (FD0505 and FD0394, Thermo Fisher Scientific), 10–20 ng of DNA and water to a final volume of 24 µL. Droplets were generated on the AutoDG Instrument for automated droplet generation (186410, Bio-Rad). PCR amplification was performed with the following cycling parameters: initial denaturation at 95 °C for 10 min, followed by 40 cycles of denaturation at 94 °C for 30 s and combined annealing/extension step at 58 °C for 1 min, and a final step at 98 °C for 10 min. Data acquisition and analysis were performed on the QX200 Droplet Reader with QX Manager Software Standard Edition v2.0.
In vivo guide synthesis
Guide RNAs were synthesized on an AKTA Oligosynt synthesizer (Cytiva). Base-loaded 2000 Å CPG was packed into a 6.3 mL stainless steel column for synthesis scale of ~50 umol. After solid phase synthesis, the oligonucleotide on support was treated with 15 mL of AMA and incubated for 4 h at 25 °C. The solution was filtered, cooled on dry ice, and 15 mL of triethylamine trihydrofluoride was added dropwise. The reaction was heated for 4 h at 45 °C, then cooled on ice, quenched with ~10 volumes of water, and neutralized. The crude product was purified on an AKTA Avant system (Cytiva) using a PLRP-S 300 Å, 15–20 μm column with mobile phases 25 mM DBAA in H2O and 50% ACN at 60 °C. Sodium salt exchange and desalting of the final product were done by TFF on a Sartoflow Smart system, using a 0.14 m2 regenerated cellulose membrane with 10 kDa molecular weight cutoff.
Lipid nanoparticle (LNP) formulation
Lipid nanoparticles were formulated using the Precision Nanosystems Ignite. Nucleic acid payloads were diluted in a 50 mM pH 4.5 acetate buffer. All lipids were purchased from commercial vendors: ALC-0315 (Broadpharm), DSPC (NOF America), cholesterol (Avanti), and DMG-PEG2000 (NOF America). Lipids were diluted in ethanol and combined to create a final stock solution according to the desired molar ratio 46.3/9.4/42.7/1.6 of ALC/DSPC/Cholesterol/PEG-lipid, respectively. Mixing was performed to achieve a final LNP composition with an N/P ratio = 6. LNPs were then diluted and dialyzed overnight. LNPs were concentrated using ultracentrifugation and sterile filtered prior to dosing. All LNPs were analyzed using dynamic light scattering (DLS) on the Malvern Zetasizer and Quant-it Ribogreen (ThermoFisher R11490) to assess size, polydispersity, RNA concentration, and encapsulation efficiency.
Mouse work
Female CD-1 mice (Mus musculus, outbred strain, 6–8 weeks old) were obtained from Charles River Laboratories. Animals were housed in individually ventilated cages with autoclaved bedding and provided ad libitum access to standard chow and water. Rooms were maintained at 20–26 °C with 30–70% relative humidity and a 12 h light/dark cycle, as monitored daily by facility staff in accordance with Explora BioLabs Animal Care Guidelines (I-OG EB02.01). LNP formulations were administered intravenously via the tail vein at a dose of 6 mg/kg RNA, using five mice per treatment group. On day 7 post-injection, animals were euthanized and median-lobe liver tissue was collected for genomic DNA extraction (Quick-DNA/RNA Zymo Kit #R2131). ddPCR was performed as described for cell experiments, with efficiencies normalized to the Tfrc reference assay (Applied Biosystems 4458366). For serum chemistry, blood was collected from three animals per group at 24 h or 7 days after dosing and analyzed by IDEXX BioAnalytics for ALT and AST levels.
Statistics and reproducibility
Biological replicates in cell studies defined as replicates performed on randomly assigned populations of cells derived from a single cell line or primary cell isolation. Biological replicate in animal studies defined as individual animals. Sample size n defined for each experiment in figure captions. All error bars in figures represent standard deviation. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. In vitro reaction and blot in Fig. 1C were performed once.
Data analysis
NGS analysis was performed with CRISPRESSO v2 on a custom pipeline (see Code availability). ddPCR analysis was performed on QX Manager Standard v2.0. Flow gating and analysis were performed on Attune NxT Flow Cytometer. IFN-beta analysis was performed on Simple Plex v3.2 software. Graphs and data analysis were performed with GraphPad Prism 10.6.0 (796). All results are reported on biological triplicates, each assayed once unless otherwise indicated for in vitro experiments. All results are reported as mean and standard deviation showing individual results from biological replicates each assayed once for in vivo experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for main and supplemental figures are provided with this paper. The amplicon sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive database under accession code PRJNA135488040. The processed NGS and ddPCR data are available at figshare [https://doi.org/10.6084/m9.figshare.30494813]41. All data generated for figures are provided in the Source Data file. Source data are provided with this paper.
Code availability
The code for custom amplicon sequencing analysis is available without restriction under the MIT license at https://github.com/jessie-wangjie/tbAmpseq42.
References
Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 9, 1911 (2018).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Papathanasiou, S. et al. Whole chromosome loss and genomic instability in mouse embryos after CRISPR-Cas9 genome editing. Nat. Commun. 12, 5855 (2021).
Komor, A. C. et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Liu, B. et al. Targeted genome editing with a DNA-dependent DNA polymerase and exogenous DNA-containing templates. Nat. Biotechnol. 42, 1039–1045 (2024).
Ponnienselvan, K. et al. Addressing the dNTP bottleneck restricting prime editing activity. bioRxiv: The Preprint Server for Biology, 2023: p. 2023.10.21.563443.
Dunyak, M. T. et al. Low RT-based Genome Editing Fidelity in Mouse Hepatocytes: Challenges and Solutions. bioRxiv Preprint. https://doi.org/10.1101/2024.10.31.621282 (2024).
Richardson, C. D. et al. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34, 339–344 (2016).
Hagedorn, P. H. et al. Locked nucleic acid: modality, diversity, and drug discovery. Drug Discov. Today 23, 101–114 (2018).
Shi, K. et al. T4 DNA ligase structure reveals a prototypical ATP-dependent ligase with a unique mode of sliding clamp interaction. Nucleic Acids Res. 46, 10474–10488 (2018).
Ellenberger, T. & Tomkinson, A. E. Eukaryotic DNA ligases: structural and functional insights. Annu. Rev. Biochem. 77, 313–338 (2008).
Moll, J. R. et al. Designed heterodimerizing leucine zippers with a ranger of pIs and stabilities up to 10−15 M. Protein Sci. A Publ. Protein Soc. 10, 649–655 (2001).
Svitkin, Y. V. et al. N1-methyl-pseudouridine in mRNA enhances translation through eIF2α-dependent and independent mechanisms by increasing ribosome density. Nucleic Acids Res. 45, 6023–6036 (2017).
Hoban, M. D., Orkin, S. H. & Bauer, D. E. Genetic treatment of a molecular disorder: gene therapy approaches to sickle cell disease. Blood 127, 839–848 (2016).
Parisi, S. et al. Characterization of the most frequent ATP7B mutation causing Wilson disease in hepatocytes from patient induced pluripotent stem cells. Sci. Rep. 8, 6247 (2018).
Chen, Q., Shen, Y. & Zheng, J. A review of cystic fibrosis: Basic and clinical aspects. Anim. Models Exp. Med. 4, 220–232 (2021).
Waheed, A. et al. Hereditary hemochromatosis: effects of C282Y and H63D mutations on association with beta2-microglobulin, intracellular processing, and cell surface expression of the HFE protein in COS-7 cells. Proc. Natl. Acad. Sci. USA 94, 12384–12389 (1997).
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652.e29 (2021).
Doman, J. L. et al. Phage-assisted evolution and protein engineering yield compact, efficient prime editors. Cell 186, 3983–4002.e26 (2023).
Bhale, A. S. & Venkataraman, K. Leveraging knowledge of HDLs major protein ApoA1: Structure, function, mutations, and potential therapeutics. Biomed. Pharmacother. 154, 113634 (2022).
Liu, Y., Kao, H.-I. & Bambara, R. A. Flap endonuclease 1: a central component of DNA metabolism. Annu. Rev. Biochem. 73, 589–615 (2004).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Durrant, M. G. et al. Systematic discovery of recombinases for efficient integration of large DNA sequences into the human genome. Nat. Biotechnol. 41, 488–499 (2023).
Liang, G. T. et al. Enhanced small green fluorescent proteins as a multisensing platform for biosensor development. Front. Bioeng. Biotechnol. 10, 1039317 (2022).
Hazelbaker, D. et al. Large serine integrase off-target discovery and validation for therapeutic genome editing. bioRxiv 2024.08.23.609471;
Sharma, R. et al. In vivo genome editing of the albumin locus as a platform for protein replacement therapy. Blood 126, 1777 (2015).
Singh, K. et al. Efficient in vivo liver-directed gene editing using CRISPR/Cas9. Mol. Ther. 26, 1241–1254 (2018).
Richards, D. Y. et al. AAV-mediated CRISPR/Cas9 gene editing in murine phenylketonuria. Mol. Ther. Methods Clin. Dev. 17, 234–245 (2020).
Yarnall, M. T. N. et al. Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. Nat. Biotechnol. 41, 500–512 (2023).
Pandey, S. et al. Efficient site-specific integration of large genes in mammalian cells via continuously evolved recombinases and prime editing. Nat. Biomed. Eng., 1-18 (2024).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Zhao, Z. et al. Prime editing: advances and therapeutic applications. Trends Biotechnol. 41, 1000–1012 (2023).
Depienne, C. & Mandel, J. L. 30 years of repeat expansion disorders: What have we learned and what are the remaining challenges? Am. J. Hum. Genet. 108, 764–785 (2021).
Liang, S. Q. et al. Genome-wide profiling of prime editor off-target sites in vitro and in vivo using PE-tag. Nat. Methods 20, 898–907 (2023).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Nan, A. X. Ligase mediated programmable genomic integration (L-PGI). NCBI SRA, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1354880 (2025).
Nan, A. X. Ligase mediated programmable genomic integration (L-PGI). Figshare, https://doi.org/10.6084/m9.figshare.30494813 (2025).
Wang, J. Jessie-wangjie/tbAmpseq. GitHub https://github.com/jessie-wangjie/tbAmpseq# (2025).
Acknowledgements
We thank S. Malhotra, P. Liu, and P. Grewal for helpful discussions and support of the project; L. Kelley, B. Castro de Oliveira, M. Madera-Gustavo, and X. Shi for NGS support; D. Harrison for plasmids and V. Choudhary and M. Spicer for mRNAs; K. Zheng, J.Y. Zhang, and N. Pilla for virus engineering; N. Gandhi and N. Lessard for sample preparation and analysis; M. Cofone and D. Asante for oligonucleotide synthesis support. J.C. Garcia-Prieto and J.M. Sutton for analytical support; M. Bakalar, S. Svenson, M. Packer, and other members of Tome Biosciences for support and insight; Work was funded by Replace Therapeutics, a direct and wholly owned subsidiary of Tome Biosciences, and Tome Biosciences.
Author information
Authors and Affiliations
Contributions
A.X.N. and M.C. designed the research, performed experiments, analyzed data, generated figures, and wrote the manuscript under guidance from J.X. L.C. and A.S. performed in vitro experiments, cloned plasmids, and generated mRNA. B.E. designed AAVs, performed in vitro experiments, and analyzed data. C.B., N.S., D.L., K.M., R.A., and M.F. performed in vitro experiments and analyzed data. D.N. designed bioinformatic methods for Cas9 protospacer design and selection. J.W. designed bioinformatic pipeline for NGS analysis and analyzed sequencing data. C.B. and J.G. synthesized components for in vivo studies. W.L. and J.A. formulated LNPs. J.V.S. performed in vivo experiments with supervision by M.T.D. J.C.C. designed mRNA and contributed to the manuscript with guidance from S.K. S.H. conceived of initial concepts and supervised the research alongside J.D.F.
Corresponding authors
Ethics declarations
Competing interests
A.X.N., C.B., N.S., D.L., B.E., J.V.S., W.L., J.A., K.M., R.A., M.F., C.B., J.W., D.N., J.C.C., J.G., M.T.D., S.K., J.D.F., and J.X. are former employees of Tome Biosciences, Inc. M.C., L.C., A.S, and S.H. are former employees of Replace Therapeutics, Inc. M.C., L.C., and S.H. are inventors on patent application WO 2023/086834 A1 filed by Replace Therapeutics, Inc. that describes Replacer editing.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nan, A.X., Chickering, M., Bartolome, C.L. et al. Ligase-mediated programmable genomic integration (L-PGI). Nat Commun 17, 563 (2026). https://doi.org/10.1038/s41467-025-67255-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67255-w