Abstract
Live-cell fluorescence nanoscopy and single-molecule tracking offer profound insights into cellular biology, but their application is limited by fluorescent tags with low brightness, large size, and irreversible photobleaching. Here, we present ReTagX, a bright, regenerative, compact array tag system enabling rapid recovery of photobleached signals through reversible, fluorogenic labeling. Newly developed ReTag1 (~18 kDa) with highly fluorogenic MaP655-TMP achieves a signal-to-noise ratio up to 204-fold and allows fluorescence recovery within 1 minute in wash-free live-cell imaging. This tag extends STED imaging of mitochondrial TOM20 from fewer than 5 to over 300 frames, enabling long-term visualization of mitochondrial dynamics at 80 nm resolution for over 100 minutes. Multimeric ReTag1 arrays, such as ReTag8 (~156 kDa), increase brightness 5.5-fold and extend TNFR2 single-molecule tracking ~2.9-fold, preserving >80% of trajectories over 5 minutes. ReTagX provides a robust platform for high-resolution, long-term molecular visualization of cellular dynamics in living cells.
Similar content being viewed by others
Introduction
Advances in fluorescence imaging have profoundly transformed our understanding of biological systems. Super-resolution microscopy (SRM), or nanoscopy, surpasses the diffraction limit of light (~200 nm), enabling unparalleled visualization of cellular structures and protein dynamics at the nanometer scale1,2. Over the past three decades, a variety of nanoscopy techniques—such as Structured Illumination Microscopy (SIM)3, Stimulated Emission Depletion (STED)4, Single Molecule Localization Microscopy (SMLM)5,6,7, and Minimal Photon Fluxes (MINFLUX)8—have been developed to enable protein imaging at the nanoscale. Of these, live-cell super-resolution imaging is particularly valuable for monitoring target proteins in real-time during cellular processes, offering robust spatial and temporal resolution while avoiding artifacts introduced by cell fixation9,10. Furthermore, single-molecule tracking (SMT) in living cells enables the direct observation of individual protein dynamics and interactions, providing critical insights into cellular processes such as signal transduction, membrane organization, and intracellular trafficking11,12,13. However, despite significant advancements, time-lapse recording of protein dynamics at high resolution remains a key challenge due to limited brightness, large size, and irreversible photobleaching of conventional fluorescent protein tag system.
To optimize nanoscopy and SMT performance in living cells, small-molecule organic fluorophores are preferred for their superior brightness, photostability, and continuous potential for synthetic optimization. Recent advances by research groups including Johnsson14, Lavis15, Hell16, and ourselves have propelled the development of cell-permeable and fluorogenic dyes, such as JF and MaP dyes, which exist in a nonfluorescent spirocyclic form in the unbound state but transition to a fluorescent zwitterionic form upon binding to target proteins17. Meanwhile, Self-labeling protein (SLP) tags, including SNAP-tag18, CLIP-tag19, Halo-tag20, and TMP-tag21,22, have been extensively developed to enable selective protein labeling with these organic fluorophores (Fig. 1). The covalent interaction between SLP substrates and genetically fused tags allows precise labeling of intracellular proteins. This combination has enabled live-cell labeling and nanoscopy of various proteins, such as cytoskeletal components16,23, β-site APP-cleaving enzyme 1 (BACE1)24, and lysosome-associated membrane glycoprotein 1 (LAMP1)25. However, a major limitation of this approach is irreversible photobleaching, which hampers long-term tracking and imaging of dynamic cellular processes. The issue of photobleaching and irreversible signal loss is particularly acute in super-resolution techniques like STED microscopy26, where ultrahigh laser intensities (~370 mW) are routinely used, restricting imaging to only a few frames27.
In conventional self-labeling tags, fluorescent probes are covalently linked to the protein tag, leading to irreversible photobleaching under intense illumination. Array tags, which recruit multiple reporters, can exhibit a tenfold increase in brightness and extended tracking duration, but their large size (~1.4 mDa) may hinder protein diffusion and interactions. ReTagX (18–156 kDa) developed in this study consists of multiple repeats of a regenerative protein tag paired with highly fluorogenic rhodamines. These rhodamines exist in a non-fluorescent spirocyclic form when unbound, significantly reducing background signals and creating a pool of fresh probes. Upon binding to ReTagX, the rhodamines transition to their fluorescent zwitterionic states. Through reversible labeling with ReTag1, bleached dyes rapidly dissociate from the tag, allowing fresh probes to bind and enabling fluorescence self-recovery. In array tag ReTagX, even when one fluorophore was bleached or exchanged, other fluorophores on ReTagX continued to fluoresce, significantly enabling extended imaging duration. Created with BioRender.com.
Another classical protein labeling strategy is array tag that introduces tandem repeats of fluorescent reporters into a single target protein, producing significantly enhanced brightness and tracking duration (Fig. 1). For instance, in array tag systems like SunTag28and Array G29, ~24 peptides or small proteins (e.g., nanobodies) are fused to the target protein, recruiting multiple fluorescent proteins to significantly boost brightness and photobleaching resistance in single-molecule tracking. The increased brightness also allows for the use of lower laser power, further reducing photobleaching and extending the tracking time (Supplementary Table 1). However, the relatively large size (~1.4 mDa) of these fusion constructs can significantly interfere with the diffusion behavior and interaction of target proteins, which typically range from 10 to 160 kDa.
To overcome these limitations, regenerative tags that enable fluorescence self-recovery have emerged as promising tools for super-resolution and long-term single-protein tracking. Heilemann’s group pioneered a reversible labeling strategy using fluorophore-labeled oligonucleotides that bind to protein-specific antibodies, extending the number of STED images from 10 to 50 frames, corresponding to the fluorescence intensity decaying by half in fixed samples30. While effective in fixed cells, this strategy presents significant challenges in living cells. Efficient fluorescence self-recovery requires four key criteria: (1) Rapid and reversible labeling: Fluorogenic ligands must bind and release within seconds to minutes to efficiently exchange bleached ligands. (2) High fluorogenicity: The fluorescence of ligands upon binding should be significantly brighter than the background from unbound ligands, enabling a large “fresh probe pool” for efficient exchange and reducing phototoxicity inside cells. (3) High brightness: The bound ligand on protein tag must be sufficiently bright for accurate localization and tracking with high temporal resolution. (4) Small tag size: A compact tag minimizes interference with protein function, expression, and diffusion. Recent developments, such as reHaloTags31, exchangeable Halo-Tag Ligands (xHTLs)32,33, and srTAG34, have been introduced for fluorescence self-recovery, yet their large tag sizes, unsatisfied brightness, or slow binding kinetics constrain their effectiveness as regenerative tags or array-forming platforms for long-term live-cell nanoscopy and single-protein tracking (Supplementary Table 1).
To achieve for long-term live-cell superresolution imaging and tracking, we developed a bright, regenerative, and compact array tag system (ReTagX) coupled with highly fluorogenic ligands. By optimizing the spirocyclization of rhodamines and linker length, we engineered highly fluorogenic ligands that provide bright labeling with minimal background, achieving up to a 203.8-fold signal-to-noise ratio in wash-free live-cell imaging. Systematic mutations (L28Q, R44L, H45Q) generated a regenerative tag (ReTag) exhibiting a 105–106-fold increase in the dissociation rate constant (Koff), enabling reversible labeling within seconds to minutes. This remarkable fluorescence recovery from ReTag1 (~18 kDa) and a pool of fluorogenic ligands, extended STED imaging of mitochondrial TOM20 from fewer than five frames to over 300, achieving 80 nm resolution for 100 min. Moreover, multimeric ReTagX arrays (e.g., ReTag8, ~156 kDa) enhance brightness by 5.5-fold and enable >80% trajectories retention over 5 min, extending single-molecule tracking of membrane proteins such as TNFR2 by ~2.9-fold compared to HaloTag in live-cell imaging. This advancement paves the way for prolonged visualization of cellular dynamics with robust spatial and temporal resolution in living biological systems.
Results
Design of a compact, bright and regenerative array tag
Fluorescence recovery is most effectively achieved through the spontaneous replacement of bleached fluorophores on a protein tag with fresh fluorescent ligands (Fig. 1). Inspired by the labeling strategies of exchangeable Halo-Tags and Array G, we sought to develop a bright, regenerative, and compact array tag system (ReTagX, 18-156 kDa) featuring fluorogenic ligands spanning the green to red spectrum. This system incorporates multiple repeats of a regenerative protein tag, which has been systematically engineered to enable efficient self-recovery of photobleached signals through reversible labeling with fluorogenic ligands. Beginning with the TMP-Tag, derived from Escherichia coli dihydrofolate reductase (eDHFR) due to its relatively small size (18 kDa, around half the size of the Halo-Tag at 33 kDa)35, we initially focused on developing highly fluorogenic ligands for TMP-Tag and subsequently performed systematic mutations to optimize binding and release kinetics for developing a regenerative protein tag—ReTagX. Finally, we demonstrate the feasibility of the array tag system (ReTag1-8) and its fluorogenic ligands for long-term live-cell nanoscopy and single-protein tracking.
Development of multicolor highly fluorogenic ligands
In developing highly fluorogenic ligands for the TMP-Tag, we employed sulfonamide-modified carbo-rhodamine (MaP618m) as a fluorescent reporter due to its robust brightness, photostability, and ability to transition from a nonfluorescent spirocyclic state to a fluorescent zwitterion upon protein binding. Trimethoprim (TMP)36, a drug with picomolar affinity for TMP-Tag in the presence of NADPH37 and excellent cell permeability, was conjugated with MaP618m via alkane linkers of varying lengths (C2, C4, C6), yielding MaP618m-C2TMP, MaP618m-TMP, and MaP618m-C6TMP (Fig. 2a and Supplementary Fig. 1). A control probe, CPY-TMP, was synthesized using carbo-rhodamine with a C4 linker (Supplementary Fig. 1).
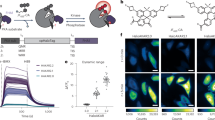
a Chemical structures of TMP-Tag ligands consisting of Trimethoprim tethered rhodamines with varying linker length. b Absorption and emission spectra of MaP618m-TMP (250 nM) in the presence (plain line) and absence (dashed line) of TMP-Tag (500 nM) after 1 h incubation. c Fluorescent response of TMP-Tag/Halo-Tag ligands (250 nM, gray dots) upon binding to TMP-Tag/Halo-Tag (500 nM, red boxes). d Binding modes of TMP ligands with TMP-Tag (PDB ID: 7NAE). e Wash-free live-cell confocal images of co-cultured untransfected U2OS cells and U2OS SNAP-TMP-NLS-expressing cells labeled with fluorescent ligands (200 nM or 10 μM) for 30 min. Untransfected U2OS cells are marked with white dashed lines. The calibration bar was indicated in the upper right corner. f Statistical charts of signal-over-background signals (S/B) and nuclear brightness of live U2OS imaged in (e). ** indicates statistical significance (p = 0.0015, p < 0.01), not significant (n.s.), significant: p ≤ 0.0001 (****), n = 20 cells (S/B) and 15 cells (brightness) from three individual experiments. Chemical structures of TMR (g) and SiR (l) derivatives coupled to TMP-Tag substrate. Absorption and emission spectra of TMR (h) and SiR (m) -derived TMP probes (1 μM) in the presence (plain line) and absence (dashed line) of TMP-Tag (2 μM) after 1 h incubation. i, n Wash-free live-cell confocal images of co-cultured untransfected U2OS cells and U2OS SNAP-TMP-NLS-expressing cells labeled with TMR-TMP, SiR-TMP, MaP555-TMP, and MaP655-TMP for 30 min. j, o Statistical charts of signal-over-background signals (S/B) and nuclear brightness of live U2OS imaged in (i, n). The cells were pre-labeled with 350 nM SiR-BG (j) or 200 nM R500-BG (o) for 2 h prior to the treatment with 200 nM TMP probes. Normalized fluorescence intensities of TMR-derived (k) or SiR-derived (p) ligands at different time points for nuclear protein labeling in live U2OS cells. In e, i, n, the untransfected cells are marked with white dashed lines. The white numbers indicate the average fluorescence intensity ratio between the nuclear signal (transfected cells) and the cytosol signal (normal cells) (Fnuc/Fcyt). The yellow numbers indicate the average fluorescence intensity ratio between the cytosol signal (normal cells) and the buffer signal (Fcyt/Fbuf). n = 20 cells from three individual experiments, mean values ± S.E.M. Significance was calculated using two-sided t-tests, not significant (n.s.), significant: p ≤ 0.0001 (****). In e, i, n, Scale bar: 10 μm. Box-and-whisker plots in (j, o): Center line represents the median; whiskers extend from minimum to maximum values; all individual data points are shown. Source data are provided as a Source Data file.
Spectroscopic characterization in absorbance and fluorescence was next examined for fluorescent ligands in the presence of TMP-Tag. MaP618m-TMP (C4 linker) demonstrated high fluorogenicity, exhibiting a 30.4-fold increase in absorbance at 616 nm and a 190.2-fold enhancement in fluorescence intensity at 636 nm (Fig. 2b, c and Supplementary Fig. 2), significantly surpassing the performance of the control probe CPY-TMP (Supplementary Fig. 3). Importantly, the interaction between the TMP-Tag and MaP618m-TMP produced greater brightness and fluorogenicity than that of Halo-Tag with ligands (MaP618m-Halo) using the same fluorogenic reporter MaP618m (Fig. 2c and Supplementary Fig. 3). Additionally, it reveals that linker length critically influenced fluorogenicity: MaP618m-C2TMP and MaP618m-C6TMP showed lower fluorescence increases of 73.0-fold and 36.7-fold, respectively (Fig. 2c), attributed to differential interactions with the TMP-Tag surface. Molecular docking studies (PDB: 7NAE) supported these findings, revealing that the C4 linker allowed optimal stabilization of the zwitterionic form on the TMP-Tag, while the shorter C2 linker restricted conformation within the binding pocket, and the longer C6 linker exposed the dye to solvent (Fig. 2d).
We then evaluated the performance of TMP ligands in live-cell imaging using U2OS cells expressing a nuclear-localized SNAP-TMP-NLS fusion protein, co-cultured with untransfected U2OS cells. Notably, U2OS cells incubated with 200 nM of the control probe CPY-TMP exhibited a high nucleus-to-cytosol signal ratio of 26.3-fold (Fig. 2e, f). However, increasing the probe concentration to 2 μM resulted in significant nonspecific staining and background signals, with a significantly reduced cytosol-to-medium signal ratio of 3.7-fold (Supplementary Fig. 4c). This inability to use high-concentration probes hinders the formation of an efficient pool for fluorescence self-recovery in live-cell imaging. In contrast, the background signal from untransfected U2OS cells incubated with MaP618m-TMP was negligible with no visible aggregation and indistinguishable from the medium, at both 200 nM, 2 μM and 10 μM (Fig. 2e and Supplementary Figs. 4, 5), a result not previously achieved in the literature. Importantly, MaP618m-TMP enables bright fluorescence labeling of nucleic TMP-NLS within 6 min (Supplementary Fig. 4f), yielding a remarkable nucleus-to-cytosol signal ratio of 110.4-fold and 134.0-fold at 200 nM and 10 μM MaP618m-TMP, respectively (Fig. 2e, f).
The performance of highly fluorogenic TMP ligands in intracellular protein labeling enables the development of fluorescent ligands across the green to red spectral range. Leveraging the comparable fluorogenicity of the TMP-Tag and HaloTag17,38, we synthesized fluorogenic TMP-Tag ligands (MaP512-TMP, MaP555-TMP, SiR-TMP, and MaP655-TMP) by replacing the Halo-Tag substrate (chloroalkane) with trimethoprim, linked via a C4 alkane chain (Supplementary Figs. 1 and 3). The fluorogenicity of newly developed fluorogenic ligands was first confirmed by the increased D50 and decreased Log KL-Z, representing the shifted equilibrium from zwitterionic states to spirocyclic form (Supplementary Figs. 6 and 7). Compared to the control probe R500-TMP, the green fluorogenic ligand MaP512-TMP demonstrated substantial improvements in both absorbance (11.8-fold) and fluorescence (31.5-fold), enabling rapid intracellular TMP-Tag labeling (<15 min) with a high signal-to-background ratio of 50.1-fold at 200 nM (Supplementary Fig. 8). However, at 1 μM, some background signal and nonspecific staining were observed, reducing the signal-to-background ratio to 8.9-fold (Supplementary Fig. 8). Notably, MaP555-TMP show moderate fluorogenicity (Fig. 2g, h) and efficiently labeled nuclear TMP-Tag, yielding an excellent nuclei-to-cytosol signal ratio (Fnuc/Fcyt = 36.5) under wash-free conditions within 5 min (Fig. 2i–k). Importantly, even at 1 μM, MaP555-TMP maintained a low background signal and bright labeling of nucleic protein with a high nuclei-to-cytosol signal ratio (Fnuc/Fcyt = 32.2) (Fig. 2i, j and Supplementary Fig. 9).
Silicon-rhodamine (SiR) is well-established for its fluorogenic properties in protein labeling14. SiR-TMP exhibited moderate fluorogenicity upon binding to TMP tags (Fig. 2l and Supplementary Fig. 3), with a 9.4-fold increase in fluorescence in vitro and a favorable nuclei-to-cytosol signal ratio (Fnuc/Fcyt = 73.9) at a low concentration of 200 nM (Fig. 2n, o). However, increasing the concentration to 2 μM led to substantial background signal and nonspecific staining, limiting its effectiveness in forming an efficient “probe pool” for fluorescence recovery (Supplementary Fig. 9). To overcome this, we incorporated 3,5-difluorobenzene-1-sulfonamide into SiR-TMP, resulting in the highly fluorogenic TMP ligand MaP655-TMP (Fig. 2l). Spectroscopic characterization confirmed its superior fluorogenicity, with substantial enhancements in both absorption (36.2-fold) and emission (179.8-fold) (Fig. 2m). In live-cell imaging, MaP655-TMP at 200 nM displayed intense nuclear fluorescence with a significant contrast ratio (Fnuc/Fcyt = 179.1) within 10 min of staining (Fig. 2n–p). Crucially, even at 10 μM, MaP655-TMP maintained negligible background (Fcyt/Fmed = 1.0) and a high contrast ratio (203.8-fold) (Fig. 2n). Such high-contrast imaging and fast labeling of live U2OS cells with highly fluorogenic MaP618m-TMP or MaP655-TMP, especially with a pool of high-concentration probes, holds great potential for fluorescence self-recovery, while maintaining good biocompatibility at high concentrations and showing no adverse effect on cell proliferation activity (Supplementary Fig. 10).
Development of regenerative tag for live-cell fluorescence self-recovery
We next explored the potential of the non-covalent TMP-Tag for reversible labeling. SDS-PAGE followed by in-gel fluorescence scanning confirmed the non-covalent nature of the interaction, as fluorescent signal from the TMP-Tag protein band removed in SDS-PAGE (Supplementary Fig. 11). Confocal imaging of live U2OS cells expressing H2B-SNAP-TMP and labeled with MaP555-TMP (with SiR-BG as control) revealed a bright nuclear signal that remained stable for over 4 h after probe washout, suggesting unexpectedly high TMP–ligand complex stability (Supplementary Fig. 12). It is probably due to the ultra-strong binding affinity and stable closed conformational states of TMP-Tag with NADPH and Trimethoprim derived probe37. This was also confirmed by Biofilm Layer Interference (BLI) analysis, which revealed that MaP555-TMP exhibited a rapid association with wild-type TMP-tag (Kon = 0.925 × 105 M−1 s−1) but a relatively low dissociation rate (Koff = 4.88 × 10−7 s−1) (Fig. 3a and Supplementary Fig. 13). Such stable binding prevents the release of bleached fluorophores and subsequent fluorescence recovery through reversible labeling. Consequently, we introduced multiple mutations into the TMP-Tag to establish a reversible labeling system with fast binding kinetics, while preserving high fluorogenicity by carefully balancing exchange rate, binding specificity, and signal contrast. The high fluorogenicity effectively suppresses nonspecific background fluorescence, enabling the use of high probe concentrations to promote probe exchange and maintain strong target labeling. To maintain fluorogenicity, we initially introduced R44L and H45Q mutations, known to decrease binding with trimethoprim and NADPH but avoiding affect fluorogenicity due to few interactions with the fluorogenic ligand39. Meanwhile, more mutations, including M20V40, D27E41, L28Q41, and W30R42, which contribute to trimethoprim resistance by expediting its release from eDHFR (TMP-Tag), were further introduced (Fig. 3b). Screening results showed that the mutations (L28Q, R44L and H45Q) reduced the affinity of the TMP-Tag and its ligands (KD: 0.33-2.8 μM) while greatly maintaining their fluorogenicity (11.8-161-fold) (Fig. 3c–e). Furthermore, BLI data indicated that the dissociation rate of MaP555-TMP increased from 4.88 × 10−7 s−1 of wild-type TMP-Tag to 0.13 s−1 of ReTag1 containing L28Q, R44L, and H45Q, a remarkable enhancement of 266,000-fold (Fig. 3d, e and Supplementary Fig. 14). Additionally, other fluorogenic probes, including MaP618m-TMP and MaP655-TMP, CPY-TMP, and SiR-TMP, also demonstrated significantly increased dissociation rates, enabling reversible binding within seconds to minutes (Fig. 3e and Supplementary Figs. 15–19).
a Binding and dissociation kinetic of MaP555-TMP to wild-type TMP-Tag measured in BLI analysis. The red dotted line indicated the beginning of the dissociation step. b Mutation sites (magenta sticks) of TMP-Tag (PDB ID: 7NAE). Yellow sticks represent the Trimethoprim (TMP). c Heatmap showing the fluorogenicity of TMP-Tag and its variants with different ligands. The numbers of F/F0 mean the ratios of fluorescent intensities of 500 nM TMP probes in the presence and absence of TMP-Tag or its variants (1 μM). d Binding and dissociation kinetics of MaP555-TMP to ReTag1. e Binding affinities, kinetic, and fluorogenicity of fluorescent ligands to TMP-Tag and ReTag1, respectively. Kon and Koff were measured by Biofilm Layer Interference (BLI), and KD was determined using the steady-state affinity model in the FortéBio Octet Data Analysis software. f Reversible labeling of nuclear ReTag1 with MaP555-TMP. Fixed U2OS cells expressing H2B-SNAP-ReTag1 (nuclear localization signal) were covalently pre-labeled with 350 nM SiR-BG (red) and iteratively stained with 400 nM MaP555-TMP (yellow). Cells were washed with an imaging medium (2 cycles). Scale bars: 20 μm. Images recorded every 1 min. g Fluorescence intensities of MaP555-TMP was normalized to fluorescence from SNAP-tag labeled with SiR-BG at the varying time points in (f). h Statistical charts of signal-over-background signals (S/B) and nuclear brightness of live U2OS cells expressing H2B-SNAP-ReTag1 (gray and purple violin diagram) and H2B-SNAP-TMP (pink violin diagram) labeled with 200 nM and 1 μM MaP555-TMP (n = 20 cells). Significance was calculated using two-sided t-tests, not significant (n.s.). i Fluorescence recovery after repetitive photobleaching of live U2OS cells expressing H2B-SNAP-ReTag1 or H2B-SNAP-TMP-Tag were labeled with MaP655-TMP, respectively. 30 s / frame, 5 min / a cycle, totally 4 cycles. Scale bars: 20 μm. j Plots illustrating the normalized fluorescence intensities at the varying time point in i. n = three individual experiments, shadow indicates the S.D. value. Data are presented as mean ± S.D. Source data are provided as a Source Data file.
We next examined the reversible binding performance in cell imaging. U2OS cells were transfected with Histone2B-SNAP-ReTag1 and stained with SiR-BG as the control probe for quantification. Consistent with in vitro results, fluorogenic ligands exhibited the rapid binding and releasing processes, enabling the reversible labeling within seconds to minutes in cellular imaging Fig. 3f, g and Supplementary Figs. 20, 21). Notably, the ReTag1 labeled with MaP555-TMP maintained its robust signal-to-noise ratio (47.8-folds) and comparably high brightness to the wild-type TMP-Tag (Fig. 3h).
We next investigated the fluorescence self-recovery of the ReTag1 with highly fluorogenic ligands in live-cell imaging. Fluorescence recovery after photobleaching (FRAP) experiments were conducted in live U2OS cells expressing Histone2B(H2B)-SNAP-ReTag1 considering slow diffusion of H2B. U2OS cells expressing wild-type TMP-Tag served as a control group, with 50% of the fluorescence signal photobleached in each cycle of a four-time repetitive bleaching process. As anticipated, more than 80% of the fluorescence signal of the wild-type TMP-Tag labeled with fluorogenic ligands was bleached, with minimal fluorescence recovery observed, confirming few interference from H2B diffusion. In contrast, ReTag1 with MaP555-TMP, MaP618m-TMP, MaP655-TMP, SiR-TMP and CPY-TMP demonstrated rapid fluorescence recovery through reversible labeling (Fig. 3i, j and Supplementary Figs. 22–24). Notably, ReTag1 labeled with the highly fluorogenic MaP618m-TMP and MaP655-TMP achieved significant fluorescence recovery within 1–2 min, retaining 68.7% and 72.9% of the fluorescence signal under identical bleaching conditions. In contrast, low fluorogenic ligands CPY-TMP and SiR-TMP showed considerably reduced recovery of 37.6% and 55.4%, respectively (Supplementary Figs. 22, 23). Similarly, an exchangeable Halo-Tag system labeled with SiR-Hy532 was also compared under the same condition, whereas 49.9% of the original signal retained (Supplementary Fig. 25). These results highlight the critical role of high fluorogenicity for efficient fluorescence recovery. The reduced performance is likely due to the high population of the zwitterionic form in unbound CPY-TMP, SiR-TMP, and SiR-Hy5, which can absorb excitation light and generate reactive oxygen species (ROS), leading to photodamage of both the protein tag and fluorophore.
Importantly, the ultra-low background signal of highly fluorogenic ligands allowed for a substantial increase in concentration, forming a larger ligand pool to enhance the efficiency of replenishing bleached fluorophores. The bleached fluorescence of the regenerative ReTag1 could recover from 72.9% to 89.9% when the concentration of MaP655-TMP was increased from 200 nM to 2 µM (Fig. 3i, j). Similar enhanced fluorescence self-recovery in higher concentration of probe was also observed in MaP618m-TMP (Supplementary Fig. 22). We thus believe that both ligand reversibility and high fluorogenicity are crucial for achieving efficient fluorescence regeneration in live-cell imaging.
Long-term live-cell nanoscopy with ReTag1 and highly fluorogenic ligands
STED nanoscopy creates fluorescent images with up to 40 nm resolution in living cells2, but requires an intense depletion laser that causes severe photobleaching, typically limiting the number of image frames to fewer than five. This has been a persistent bottleneck in applying STED nanoscopy for tracking cellular dynamics43. We thus examined the performance of ReTag1 and highly fluorogenic ligands in live-cell STED nanoscopy.
STED imaging of live U2OS cells expressing Vimentin-ReTag1-Halo stained with MaP655-TMP enabled the capture of individual filament structures with a full-width at half maximum (FWHM) of 43 ± 5 nm, approximately five times lower than that from confocal imaging (Fig. 4a and Supplementary Fig. 27a). Moreover, the saturated stimulated emission power of MaP655-TMP labeled ReTag1 was determined to be Psat = 2.28 ± 0.03 mW by measuring fluorescence decay under varying STED laser powers at 775 nm (Fig. S27b)44,45. Importantly, over 15 frames of STED images of Vimentin were obtained using ReTag1 and MaP655-TMP, whereas less than 5 frames could be acquired with conventional Halo-Tag under the same conditions (Supplementary Fig. 26).
a Confocal and STED images of live U2OS vimentin-ReTag1-Halo-expressing cells labeled by 200 nM MaP655-TMP (219 mW STED laser power at 775 nm). The magnified regions (white boxes) and the plot of the fluorescence signal along the line profile in (a) are shown on the right. Scale bars: 2 μm (overview) or 500 nm (magnified region). b Time-lapse STED imaging of live U2OS TOM20-Halo-ReTag1-expressing cells labeled with 200 nM or 2 μM MaP655-TMP and TOM20-Halo-ReTag1-expressing cells labeled with 200 nM SiR-HTL or SiR-Hy5. Frame numbers indicated in the top right corner. Scale bar: 500 nm. c Normalized fluorescence intensities of mitochondria plotted as a function of frame numbers in STED images in b. n = three individual experiments, shadow indicates the S.D. value. d Diagram of biosynthesis, elongation, and fission process of mitochondria. Created with BioRender.com. e Time-lapse STED images of live U2OS cells expressing TOM20-Halo-ReTag1 after labeling with 2 μM MaP655-TMP. The number in the top-left corner indicates the image frame. Yellow arrow indicated the elongation process. White arrow indicated the ring shape of mitochondria. Purple arrow indicated the spherical mitochondria movement along the elongated mitochondria. Scale bar: 500 nm. Source data are provided as a Source Data file.
We further investigated the 3D STED microscopy of translocase of the outer membrane (TOM) 20 in the outer mitochondria membrane with ReTag1 and explored its potential in time-lapse live-cell STED microscopy (Fig. 4b and Supplementary Fig. 27c, d). The fluorescence of Halo-Tag with covalently bound SiR-HTL was severely bleached under the depletion laser of ~70 mW at 775 nm, limiting imaging to fewer than five frames. In contrast, the fluorescence intensity of MaP655-TMP remained at approximately 65% after 30 frames, surpassing the 50% retention observed with the reported Halo-Tag coupled with exchangeable ligand SiR-Hy5 (Fig. 4b, c). Remarkably, consistent with the findings from the FRAP experiment, increasing the concentration of MaP655-TMP to 2 µM maintained high contrast imaging and significantly enhanced photostability, achieving approximately 80% recovery after 30 frames. This resistance to photobleaching even allowed us to track mitochondria for more than 300 frames, revealing processes of fission, fusion and elongation of mitochondria with 80 nm resolution within 100 min (Fig. 4d, e and Supplementary Fig. 28; Supplementary Movie 1). To the best of our knowledge, this represents a distinctive achievement in the field.
To assess STED induced photodamage, we used a calcium sensitive fluorescent probe and observed no significant differences in intracellular calcium levels or cell morphology between STED irradiated cells and non-irradiated controls, indicating minimal acute cytotoxicity (Supplementary Fig. 29). Interestingly, we found a clear increase in ROS accumulation during prolonged STED imaging, particularly in cells labeled with fluorescent tags, compared to non-irradiated controls (Supplementary Fig. 30), probably causing mitochondrial morphological change (Fig. 4e). To support longer term STED imaging with reduced photodamage, the inclusion of ROS scavengers may help protect live cell structures from oxidative stress26.
We also investigate the performance of ReTag1 in SIM imaging, which provides super-resolution imaging (<100 nm) with ultrahigh speed (~ms) to track cellular dynamics46,47. The wash-free imaging of live U2OS cells expressing Vimentin-ReTag1-Halo labeled with MaP555-TMP, MaP618m-TMP, and MaP655-TMP revealed linear vimentin structures and dynamic movement at 91 ± 14 nm FWHM, significantly higher than wide-field imaging (512 ± 19 nm) (Supplementary Fig. 31 and Supplementary Movie 2). Additionally, the ReTag1 performs well with reported fluorogenic probes, enabling wash-free multi-color confocal and SIM imaging of various organelles, such as nucleus, F-actins, mitochondria, and vimentin simultaneously (Fig. 5a and Supplementary Figs. 32, 33).
a Wash-free three-color SIM image of live U2OS expressing Vimentin-Halo-ReTag1 stained with 200 nM of MaP655-TMP (cyan hot), 500 nM of MaP555-Actin (orange hot), and 2 µg/mL of Hoechst33342 (red) for 1 h. Scale bar: 10 μm. b Normalized fluorescence intensities of bleaching curves fitting from SIM images of Vimentin labeled with 400 nM MaP618m-TMP (red line, shadow indicates the S.D. value) and 400 nM MaP618m-Halo (blue line, shadow indicates the S.D. value). Data are presented as mean ± S.D., n= three individual experiments. c Wash-free wide field and SIM images of live U2OS cells expressing TOM20-Halo-ReTag1 stained with 200 nM MaP655-TMP. Scale bar: 10 μm. The magnified regions (white boxes) and the plot of the fluorescence signal along the line profile in (a) are shown on the right. Scale bar: 1 μm. d Time-lapse SIM imaging of live U2OS cells expressing TOM20-Halo-ReTag1 labeling with 200 nM MaP618m-TMP, scale bar: 10 μm. The biosynthesis (yellow arrow), elongation (red arrow) and fission (green arrow) of the mitochondria in the magnified regions (white box) was indicated with different time scale. Scale bar: 0.5 μm. e 2D-MINFLUX microscopy image of fixed U2OS cells expressing Vimentin-ReTag1 labeled with MaP655-TMP (2.5 nM). Confocal laser-scanning microscopy (CLSM) imaging was used as a reference. Magnification reveals Vimentin-ReTag1 with a localization precision of ∼5.0 nm. Scale bars: 0.1 μm. Source data are provided as a Source Data file.
We further confirmed the ReTag1’s utility in time-lapse SIM imaging by tracking mitochondrial translocase of the outer membrane (TOM20). The reversible and fluorogenic labeling of the ReTag1 with MaP618m-TMP provided better photobleaching resistance than Halo-Tag with covalently bound MaP618m-Halo (Fig. 5b). SIM imaging of live U2OS cells expressing TOM20-Halo-ReTag1 labeled with MaP655-TMP was successfully obtained with the FWHM of 118 ± 17 nm (Fig. 5c). Labeling with MaP618m-TMP not only revealed the specific distribution of TOM20 on the outer mitochondrial membrane with the FWHM of 123 ± 12 nm, but also showed mitochondrial dynamics such as fission, fusion, and elongation (Fig. 5d and Supplementary Fig. 34).
Notably, the reversible and fluorogenic labeling of the ReTag1 with highly fluorogenic ligands could be employed in MINFLUX, a microscopy method capable of achieving nanoscale localization based on the fluorescence “on-off” switching of fluorophores48. MINFLUX imaging of fixed U2OS cells expressing vimentin-ReTag1 with MaP655-TMP reveals individual vimentin filaments with an 5.0 nm localization precision (Fig. 5e and Supplementary Fig. 35).
Developing ReTagX for single-protein tracking
Single-protein studies are invaluable for understanding the operating principles of cellular machines but are hindered in living cells by limited brightness and tracking time due to photobleaching, even using small organic fluorophores49,50. Tandem repeat fluorescent reporters, such as SunTag28 and ArrayG29, have greatly advanced single-molecule tracking, but their large size (~1.3 MDa) may affect diffusion behavior and protein interaction. Inspired by Vale’s and Liphardt’s work28,29, we introduced a multiple-repeat ReTag1 fusion (ReTagX) with the highly fluorogenic MaP655 to enhance imaging contrast and tracking performance while retaining small tag size. We constructed ReTag4 (~78 kDa) by fusing four ReTag1 repeats with a (GGGGS)3 linker (Fig. 6a), and its incubation with MaP655-TMP resulted in a 55.5-fold increase in absorbance and a 170-fold increase in emission intensity, about four times higher than the original ReTag1(Fig. 6b). The increased brightness was confirmed by live-cell U2OS imaging of H2B-GFP-ReTagX-expressing cells (Fig. 6c, d), where low laser power was sufficient for high-contrast imaging and improved long-term SIM imaging (Fig. 6e, f).
a Illustration of ReTagX fused to the target protein. Created with BioRender.com. b Absorption and emission spectra of MaP655-TMP (10 μM) in the absence (orange line) and presence of ReTag1 (100 nM, pink line), ReTag2 (100 nM, blue line) and ReTag4 (100 nM, red line) after 1 h incubation. c Wash-free live-cell confocal images of co-cultured untransfected U2OS cells and U2OS H2B-GFP-ReTagX-expressing cells labeled with MaP655-TMP (200 nM) for 30 min. The untransfected cells are marked with white dashed lines. The white numbers indicate the average fluorescence intensity ratio between the nuclear signal (transfected cells) and the cytosol signal (normal cells) (Fnuc/Fcyt). d Statistical charts of nuclear brightness of TMP-probe in (c) (right, n = 23, 17, 19 cells, mean values ± S.E.M.). Not significant (n.s.), significant: p ≤ 0.0001 (****). e Wash-free wide field and SIM images of live U2OS cells expressing MitoNeet-ReTag4 stained with 200 nM MaP655-TMP. Scale bar: 5 μm. f Normalized fluorescence intensities of mitochondria plotted as a function of frame numbers in SIM images in U2OS cells expressing MitoNeet-ReTag1 and MitoNeet-ReTag4. n = three individual experiments, shadow indicates the SEM value. g Raw images of single ReTagX with different concentrations of MaP655-TMP (5 Hz). Scale bar: 10 μm. h Statistical charts of fluorescence intensity with background deduction for ReTagX (5 Hz, N = 100). Significant: p ≤ 0.0001 (****). i Representative fluorescence attenuation images of single ReTag1 (100 nM, orange line) and ReTag4 (100 nM, blue line, 500 nM, red line) (5 Hz). The dotted lines indicate the number of fluorophores on individual proteins. j Histogram showing the track-frame distribution of ReTag4 (8476 trajectory segments) and ReTag1(2786 trajectory segments) after fixed in coverslips. k Illustration of TNFR2 fused with either Halo-Tag or ReTag8. Created with BioRender.com. l Normalized single-molecule fluorescence intensities of TNFR2-Halo-Tag labeled with SiR-HTL (1 nM) and TNFR2-ReTag8 labeled with MaP655-TMP (200 nM) in live cells. Significant: p ≤ 0.0001 (****), n = 1495 and 1483 points from three independent experiments. m Representative trajectories of TNFR2-Halo-Tag labeled with SiR-HTL (1 nM) and TNFR2-ReTag8 with MaP655-TMP (200 nM) after 10 s and 50 s imaging, color coded by time. n Plot of the number of trajectories per 10 s interval for TNFR2 labeled with ReTag8, Halo-Tag, and TMP-Tag8. n = three individual experiments, error bar indicates the S.D. value. o Histogram showing the track-frame distribution of TNFR2-ReTag8 (2276 trajectories) and TNFR2-Halo-Tag (2222 trajectories) in live-cell imaging (20 Hz). Source data are provided as a Source Data file.
We further evaluated ReTag4’s performance in wash-free single-molecule imaging by purifying the protein and fixing it on a glass surface. The average brightness of single ReTag4 is significantly higher than ReTag1, especially at 500 nM, where it showed 2.9-fold higher brightness (Fig. 6g, h). Different from fluorescent imaging of bulk H2B–ReTag1, individual fluorophores bound to ReTag1 are visible for only tens of frames, after which the signal is lost due to dissociation or photobleaching, limiting its utility for long-term tracking due to signal loss (Fig. 6i). In contrast, ReTagX supports labeling with multiple fluorophores, showing a linear increase in fluorescence intensity with repeat number (Supplementary Fig. 36). ReTag4 exhibited high-frequency reversible labeling, particularly at 500 nM MaP655-TMP, enabling efficient multi-fluorophore exchange (Fig. 6i). Even when one fluorophore was bleached or exchanged, other fluorophores on ReTag4 continued to fluoresce, enabling extended single molecule tracking of over 100 frames in a streaming acquisition (Fig. 6j).
We next evaluated the performance of ReTag8 (~156 kDa), comprising eight ReTag1 repeats, for single-molecule tracking in live cells. To target the plasma membrane, ReTag8 was fused to a CAAX motif derived from the C-terminus of H-Ras (Supplementary Fig. 37a). In single-molecule imaging, the combination of ReTag8 with fluorogenic MaP655-TMP yielded a 5.2-fold increase in brightness compared to monomeric constructs (Supplementary Figs. 37b, c, and 38), enabling continuous maximum tracking of membrane proteins from approximately 50 frames to over 200 frames in living cells (Supplementary Fig. 37b–f).
We further fused ReTag8 to a membrane protein Tumor Necrosis Factor Receptor 2 (TNFR2) and conducted single-molecule tracking in live U2OS cells, using Halo-Tag and TMP-Tag8 (8xTMP-Tag repeats) as controls (Fig. 6k). Under identical imaging conditions (60% 640 nm laser, 60.4 W/cm², 50 ms exposure), TNFR2-ReTag8 exhibited a 5.5-fold increase in brightness relative to TNFR2-HaloTag (Fig. 6i and Supplementary Fig. 39). The measured diffusion coefficient of TNFR2–ReTag8 (0.28 ± 0.019 µm²/s) closely matched that of TNFR2-HaloTag (0.30 ± 0.028 µm²/s), indicating minimal perturbation to native mobility (Figs. 6m and S40). Moreover, ReTag8’s reversible fluorogenic labeling maintained a constant number of trajectories (>80%) for over 5 min observation, whereas fewer than 20% in Halo-Tag and TMP-Tag8 labeled proteins remained visible beyond 1 min under the same conditions (Fig. 6n and Supplementary Fig. 40; Supplementary Movie 3). The increased brightness and exchangeability of ReTag8 also enabled ~2.9-fold longer tracking durations compared to Halo-Tag (Fig. 6o and Supplementary Movie 4), thereby facilitating high-precision, long-term single-molecule imaging of membrane proteins in live cells.
Discussion
Irreversible photobleaching has long posed a significant challenge for time-lapse tracking cellular activities and protein dynamics in living cells, particularly in nanoscopy and SMT. Fluorescence regenerative tags present a promising solution to this issue, but it demands more stringent requirements in signal contrast and labeling kinetics. In this study, we introduce a template for developing an innovative bright and regenerative array tag through systematic engineering of fluorogenicity and reversible binding.
Highly fluorogenic ligands with ultra-low background are essential, as higher concentrations can maximize the replenishment of bleached fluorophores for signal recovery. Optimization of the spirocyclization of rhodamines and linker length has produced highly fluorogenic MaP618m-TMP and MaP655-TMP, enabling incubation concentrations to increase 50-fold up to 10 μM. The high concentration of unbound probes indeed provides a fresh pool to efficiently replenish bleached probes for fluorescence recovery in STED nanoscopy. Another crucial step for fluorescence regeneration is the reversible labeling with rapid binding and releasing kinetics. Mutations (L28Q, R44L, and H45Q) in the wild-type TMP-Tag result in the ReTag1, increasing the release rate by 266,000-fold and enables reversible and fluorogenic labeling within seconds to minutes.
The remarkable fluorescence self-recovery facilitated by ReTag1 and highly fluorogenic ligands enables time-lapse STED imaging of over 300 frames, a substantial improvement over the fewer than 5 frames achievable with the widely used HaloTag in conjunction with Silicon-rhodamine. This advancement holds significant promise for the long-term tracking of cellular dynamics at nanoscale resolution. However, phototoxicity remains a concern, potentially causing morphological alterations in organelles due to reactive oxygen species (ROS) generated by intense excitation and depletion laser irradiation. To mitigate this issue, further chemical modifications of ligands, such as the incorporation of COT moieties into fluorogenic probes51,52,53, and addition of ROS scavengers, could be explored to reduce ROS generation and further enhance the longevity and effectiveness of live-cell nanoscopy.
The compact size of ReTag1 (~18 kDa) allows the fusion of tandem repeats to construct regenerative ReTagX, which significantly enhances brightness and enables multi-labeling with reversible fluorophore exchange. ReTag8 demonstrates a 5.5-fold increase in brightness and a 2.9-fold longer single-protein tracking duration and an extended observation window of over 5 min, significantly outperforming the widely used Halo-Tag (<1 min). Future incorporation of additional ReTag1 repeats provides a powerful tool to enhance single-molecule tracking with minimal interference, though not suitable for stable-labeling applications like SunTag in translation studies. However, it is essential to carefully optimize probe concentration for signal contrast, account for potential homo-FRET quenching, and balance the tag size to avoid potential impacts on protein function, such as alterations in protein dynamics or the risk of aggregation.
In conclusion, we have developed a bright and compact regenerative array tag through systematic optimization of ligand fluorogenicity and reversible binding kinetics. This remarkable fluorescence self-recovery in ReTagX not only facilitates STED imaging with a resolution of 80 nm over 100 min, but also supports the long-term observation and tracking of membrane proteins at single-molecule resolution. This advancement in regenerative tagging sets the stage for long-term visualization of cellular dynamics and offers new insights into the roles of proteins within living biological systems.
Methods
The detailed procedures for the synthesis and characterization of all compounds, as well as their in vitro characterization and imaging experiments, are detailed in the Supplementary notes.
General experimental information for synthesis
All chemical reagents and dry solvents were purchased from commercial suppliers (Sigma-Aldrich, Bide Pharmatech, Adamas) and were used without further purification or distillation. Reactions were monitored by thin-layer chromatography (TLC) performed on TLC-aluminum sheets (Silica gel 60 F254, Merck) or liquid chromatograph-mass spectrometry (LC-MS 1260 Infinity II-6125, Agilent). Preparative RP-HPLC was performed on a 1260 Infinity II system (Agilent) on a C18 5 µm, 21.2 × 250 mm column (Supelco, flow rate 6 mL/min), solvent A: 0.1% v/v TFA in H2O, solvent B: acetonitrile. Typical gradient was from 10% to 95% B within 60 min. 1H NMR and high-resolution mass spectrometry (HRMS) were used to characterize all isolated compounds. Nuclear magnetic resonance spectra were recorded on a Bruker 400 MHz and 600 MHz instrument. All 1H NMR experiments are reported in units, parts per million (ppm), and were measured relative to the signals for residual dimethyl sulfoxide (2.51 ppm) in the deuterated solvent, unless otherwise stated. Multiplicities are shown as follows: s (singlet), d (doublet), t (triplet), q (quartet), dd (doublet of doublets), dt (doublet of triplets), and m (multiplets). HRMS were measured using ESI-QTOF mass spectrometer.
Protein-ligand docking simulation
The binding mode of the ligand to the protein was simulated using the CDDOCK module of Discovery Studio 4.0. The protein and ligand were assigned the CHARMM force field for the docking procedure. The crystal structure of eDHFR (PDB ID: 7NAE) was obtained from the Protein Data Bank (PDB), and the protein was prepared by removing water molecules and adding polar hydrogens. The protonation states of residues were carefully considered, with His residues protonated to HID, and Asp, Arg, Glu, and Lys residues set to their deprotonated states. The binding pocket was defined as the amino acid residues within 10 Å of the ligand. The three-dimensional structure of the ligand was generated using ChemBio3D Ultra software and energetically minimized with MMFF94, applying 5000 iterations and an RMS gradient of 0.05. Random ligand conformations were generated via high-temperature molecular dynamics simulations and random rotations. Following docking, the top 10 ranked protein-ligand conformations were selected, and the binding modes of the docked poses were visualized using PyMOL 1.1. All other docking parameters were set to default values.
General spectroscopic methods
Fluorescent probes for spectroscopy were prepared as stock solutions in DMSO and diluted such that the DMSO concentration did not exceed 1% (v/v). The measurements were taken in HEPES buffer (50 mM HEPES, 50 mM NaCl, pH 7.3) at ambient temperature (25 ± 2 °C) unless otherwise noted. Absorption spectra and emission spectra were measured on BioTek Cytation 5 Cell Imaging Multimode Reader using 384-well plate (Greiner, #781090) with optical bottom.
Plasmids and transfection
The plasmid pET51b (+) eDHFR(wt)-P30-Halo, pET51b (+) Halo-GGSX4-eDHFR(mt), pET51b (+) EGFP-ReTag1 and pET51b (+) EGFP-ReTag4 were constructed for protein expression and purification in vitro experiment. Plasmid pcDNA5/FRT/TO-SNAP-GGSX4-TMP -NLS and plasmid pcDNA5/FRT/TO-H2B-SNAP-ReTag1/pcDNA5/FRT/TO-H2B-SNAP-TMP host a C-terminal SNAP-tagged and eDHFR-tagged NLS and a N-terminal SNAP-tagged and eDHFR-tagged human H2B, respectively. Plasmid pLVX-Vimentin-ReTag1-Halo carries N-terminal ReTag1-tagged and Halo-tagged human vimentin, driven by the CMV promoter. Plasmid pcDNA5/FRT/TO-TOM20-HaloTag7-ReTag1 hosts a N-terminal Halo-tagged and human TOM20. Plasmid pcDNA5/FRT/TO-EGFP-TMP-CAAX/ pcDNA5/FRT/TO-EGFP-ReTag8-CAAX host a N-terminal ReTagX-tagged CAAX.
The appropriate fragments were inserted into the plasmid according to the protocol of Gibson. The synthesis of all primers and the subsequent sequencing of plasmids were executed by Sangon Biotech (Shanghai, China) following the standard protocol. The transfections were utilized the Lipofectamine 3000 transfection reagent (#L3000008). U2OS cells were seeded on the confocal plate (#781092) at an amount of 5 × 105 cells per well. After culturing for 24 h, plasmids were transfected into the cells. After 6 h, transfected cells were washed two times with 1X PBS and were maintained at fresh phenol red DMEM media with 10% FBS at least 12 h for the imaging. In single-molecule tracking, U2OS cells were imaged ~3 h post-transfection to maintain low protein expression levels. Specific plasmid sequences and construction methods are described in Supplementary notes.
General cell culture
U2OS cells (ATCC, BFN60700128) were cultured in Dulbecco’s Modified Eagle Medium (Gibco, #11960069) supplemented with 10% (v/v) FBS (Gibco #10099141 C). The cells were incubated at 37 °C and 5% CO2. Cell lines were split every 3–4 days or at confluence and underwent regular mycoplasma testing. The cells were incubated for 1–2 days before imaging. Imaging medium contains phenol red-free Dulbecco’s Modified Eagle Medium (DMEM) media with 10% FBS.
General no-wash live cell confocal microscopy
After transfection, the U2OS cells were incubated with 200-500 nM probes for 0.5-2 h at 37 °C in a 5% CO2 atmosphere and directly imaged using confocal microscopy without washing steps. A Leica SP8 confocal scanning microscope equipped with a white light laser, HC PL APO CS2 ×20.0 objective lens and spectral HyD detector was used. Microscopy conditions: R500 derivatives, λex 505 nm, detection range 515–600 nm; TMR derivatives, λex 555 nm, detection range 565–650 nm; CPY derivatives, λex 615 nm, detection range 625–700 nm; SiR derivatives, λex 640 nm, detection range 650–750 nm. U2OS cells were seeded on the confocal plate and then transfected with pcDNA5/FRT/TO-SNAP-GGSX4-TMP-NLS, pcDNA5/FRT/TO-H2B-SNAP-ReTag1, pcDNA5/FRT/TO-TOM20-HaloTag7-ReTag1, pLVX-Vimentin-ReTag1-Halo, and pcDNA5/FRT/TO-H2B-GFP-ReTagX.
Live cell STED microscopy
Live cells STED nanoscopy was performed using an Abberior STEDYCON 775/RESOLFT QUAD scanning microscope (excitation lines: 405 nm, 485 nm, 561 nm, 640 nm; STED lines: 775 nm) equipped with a UPlanSApo 100x/1.4 oil immersion objective lens (Abberior Instruments, Germany). Imaging was performed with avalanche photodiodes (APD) and spectral detection. Fluorophores, microscope and imaging parameters are described in Supplementary notes. For STED imaging, “Red Hot” LUTs were applied for data representation. STED performance was assessed by measuring the Full Width at Half Maximum (FWHM) of single intermediate filament fibers revealed by Vimentin-ReTag1-Halo and TOM20-Halo-ReTag1. STED images were acquired with a pixel dwell time of 10 µs and a pixel size of 25 nm with 5 average line scans. Image analysis was performed with ImageJ by extracting fluorescence intensity profiles perpendicular to vimentin filaments and mitochondria outer membrane. Mean diameters were calculated from at least three individual images (n ≥ 3) by fitting a Gaussian function, yielding FWHM.
Live cell SIM microscopy
Live-cell imaging was performed using a super-resolution structured illumination microscopy (SIM) system (Multi-SIM from Beijing NanoInsights-tech Co., Ltd.). Cells were imaged in a temperature-controlled chamber (37 °C) with a CO₂ concentration maintained at 5%. The SIM system utilized a laser-based light source with multiple excitation wavelengths appropriate for the fluorophores used (488 nm, 561 nm, 594 nm and 640 nm). Super-resolution imaging was achieved using structured illumination in three dimensions (xy, xz, and yz planes). A total of 9 phases of structured illumination patterns were projected onto the sample, and each image stack was acquired sequentially at 100 nm lateral resolution. For z-stack imaging, optical sectioning was performed at 200–300 nm increments to obtain high-resolution 3D reconstructions. The SIM system used a high numerical aperture (NA) objective lens, typically 100xoil immersion (NA = 1.4). SIM images were reconstructed using HVL1 constrained deconvolution algorithm or DFCAN and DFGAN deep learning algorithms. The final reconstructed images were processed with a resolution of approximately 100 nm in the lateral direction and 300–400 nm in the axial direction. For analysis, images were further processed using ImageJ, and customized plugins were applied to enhance signal-to-noise ratio and correct for drift. Image analysis was performed with ImageJ by extracting fluorescence intensity profiles perpendicular to vimentin filaments and mitochondria outer membrane. Mean diameters were calculated from at least three individual images (n ≥ 3) by fitting a Gaussian function, yielding FWHM. More detailed experimental procedures are described in Supplementary notes.
MINFLUX microscopy
For MINFLUX, U2OS Vimentin-ReTag1-expressing cells were seeded on 18 mm #1.0 coverslips (41001118, Deckglaser), grown until a confluence of 60% was reached and chemically fixed by 2.4% paraformaldehyde (157-8, Electron Microscopy Sciences) and 0.1% glutaraldehyde (16220, Electron Microscopy Sciences) for 10 min. The cells were rinsed three times with PBS, followed by an additional brief wash in PBS and immersion in a 10 mM sodium borohydride (71320, Sigma) solution in PBS for 5 min. Cells were afterwards incubated with 2.5 nM MaP655-TMP in PBS with 100 μM NADPH. MINFLUX imaging was performed on an Abberior MINFLUX (Abberior Instruments GmbH, Göttingen, Germany) built on a motorized inverted microscope IX83 (Olympus, Tokyo, Japan) and equipped with 642 nm and 560 nm MINFLUX laser lines, 642 nm, 560 nm, 488 nm and 405 nm confocal lines. Imaging was performed with APDs in the spectral windows 650–685 nm and 685-720. Images were acquired using the default 2D imaging sequence, with an L in the last iteration step of 40 nm, and photon limit of 150 photons. Pinhole was set to 0.78 A.U., and excitation power at the periscope for the first iteration was ~ 96 µW. Four iterations were applied in the post-processing step to avoid false single-molecule emission events. To exclude detections originated from transient background, we set the maximum thresholds of 0.8 for the center frequency ratio (CFR) test and1 kHz for the fluorescence imaging. 39 Localizations from the same emission trace, i.e., with same TiD, further than three standard deviations with respect to the mean trace position were considered outliers and excluded from the trace. Only the traces containing at least 3 localizations within the first 180 min of the measurement were considered to calculate the localization precision. The experimental localization precision was estimated by co-aligning the mean values of all localizations obtained from individual emission traces fitted with a Gaussian function to estimate the overall standard deviation. For MINFLUX imaging, “Hot” LUTs were applied for data representation.
Sample preparation for single-molecule imaging
Clean coverslips were incubated with a mixture of PLL-(g)-PEG-biotin (50%, Susos) and PLL-(g)-PEG at 1:100,000 ratio for at least 2 h, followed by incubation with 1 mg/mL NeutrAvidin (Thermo Fisher Scientific) for 30 min to bind the biotin. For in vitro single-molecule imaging, the surfaces were then blocked with 1% BSA for 2 h, and finally, biotinylated purified protein was added and incubated for 15 min. In live-cell single-molecule imaging, the surfaces were incubated with 1 mg/mL RGD (pepnet) for 30 min, followed by blocking with 1% BSA for 2 h (cite https://doi.org/10.7554/eLife.67379). After each incubation step, the coverslips were washed three times with 1X TBS.
Cells were plated in 35 mm dishes and, upon reaching ~70% confluence, were transfected with 1 µg of plasmid DNA per dish for 5 h. The cells were then detached with trypsin and transferred onto pre-treated coverslips at a 1:3 split ratio. After 12 h of incubation in DMEM supplemented with 10% FBS at 37 °C and 5% CO₂, live-cell imaging was performed.
For covalent Halo-Tag labeling, cells were stained with 1 nM SiR-HTL for 30 min at 37 °C and 5% CO₂. Excess dye was removed by three washes with PBS, and the cultures were transferred to phenol-red–free DMEM supplemented with 10% FBS before imaging. For non-covalent TMP-Tag and ReTag labeling, Cells were washed three times with PBS and then incubated in phenol-red–free DMEM supplemented with 10% FBS and 200 nM MaP655-TMP for 30 min at 37 °C and 5% CO₂ prior to experiments.
Single-molecule super-resolution microscopy and analysis
For in vitro single-molecule imaging, purified proteins ReTag1 and ReTag4 immobilized on the substrate were tracked using an Olympus IX86 TIRF microscope. The imaging was conducted at 30% 640 nm laser intensity (30.4 W/cm2) and 200 ms exposure time to monitor their brightness after binding with 200 nM MaP655-TMP. The average of the 10 minimum values from each trajectory was used as the background, which was then subtracted from the signal.
Live-cell single-molecule imaging was performed on the same TIRF setup with a 640 nm excitation laser. For ReTag8-CAXX tracking and TMP-CAXX, 10% laser intensity (10.3 W/cm2) and 50 ms exposure time were used after binding with 200 nM MaP655-TMP. For TNFR2 imaging and tracking, acquisition was carried out at 60% laser intensity (60.4 W/cm2) with 50 ms exposure time. For the MSD experiment the imaging parameters the exposure time was shortened to 30 ms (still at 60% laser power) to improve temporal resolution. To quantify the fluorescence intensities of individual foci, a circular region of interest (ROI) with a diameter of 0.5 μm was selected. The average background intensity, measured in ten distinct areas of the cell that lacked fluorescent foci, was then subtracted from the signal. In signal processing, the following trajectories were excluded: those with fewer than 5 frames or maximal pairwise displacement <0.6 µm, as they were considered to represent non-specific dye binding signals.
Statistics and reproducibility
Sample sizes were chosen as indicated in the figure legends. No data were excluded from the analyses. The heatmap, bar and line charts in the figures were generated using GraphPad Prism 9.4.1 software. Figures 2f, j, o and 3h were analyzed by the two-tailed Student’s t-test. Significance was defined as a not significant (n.s.), **p < 0.01 and ****p < 0.0001. Figure 6d, h, l were analyzed by One-way ANOVA test. At least two independently replicates were performed for all experiments with similar results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. Source Data for the figures in the main text and in the Supplementary Information are provided in the Source Data file. Additional information and files are also available from the corresponding author. Source data are provided with this paper.
Change history
23 February 2026
In this article the affiliation details for Dongjuan Si, Zhihao Dong, Gang Xing, Hanqing Zhao, Cong Li and Lu Wang were incorrectly given as 'MOE Key Laboratory of Smart Drug Delivery, MOE Innovative Center for New Drug Development of Immune Inflammatory Diseases, School of Pharmacy, Endoscopy Center and Endoscopy Research Institute, Zhongshan Hospital, Fudan University, Shanghai, China' but should have been 'School of Pharmaceutical Sciences, MOE Key Laboratory of Smart Drug Delivery, MOE Innovative Center for New Drug Development of Immune Inflammatory Diseases, Endoscopy Center and Endoscopy Research Institute, Zhongshan Hospital, Fudan University, 201203, Shanghai, China'. The original article has been updated.
References
Sigal, Y. M., Zhou, R. & Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 361, 880–887 (2018).
Liu, Z., Lavis, L. D. & Betzig, E. Imaging live-cell dynamics and structure at the single-molecule level. Mol. Cell 58, 644–659 (2015).
Gustafsson, M. G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl. Acad. Sci. USA 102, 13081–13086 (2005).
Klar, T. A., Jakobs, S., Dyba, M., Egner, A. & Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl. Acad. Sci. USA 97, 8206–8210 (2000).
Schnitzbauer, J., Strauss, M. T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science. 355, 606–612 (2017).
Fuhrmann, M. et al. Super-resolution microscopy opens new doors to life at the nanoscale. J. Neurosci. 42, 8488–8497 (2022).
Schnell, U., Dijk, F., Sjollema, K. A. & Giepmans, B. N. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Chen, J. et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell 156, 1274–1285 (2014).
Brazda, P. et al. Ligand binding shifts highly mobile retinoid X receptor to the chromatin-bound state in a coactivator-dependent manner, as revealed by single-cell imaging. Mol. Cell Biol. 34, 1234–1245 (2014).
Nair, D. et al. Super-resolution imaging reveals that AMPA receptors inside synapses are dynamically organized in nanodomains regulated by PSD95. J. Neurosci. 33, 13204–13224 (2013).
Lukinavicius, G. et al. A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins. Nat. Chem. 5, 132–139 (2013).
Grimm, J. B. et al. A general method to fine-tune fluorophores for live-cell and in vivo imaging. Nat. Methods 14, 987–994 (2017).
Butkevich, A. N. et al. Fluorescent rhodamines and fluorogenic carbopyronines for super-resolution STED microscopy in living cells. Angew. Chem. Int Ed. Engl. 55, 3290–3294 (2016).
Wang, L. et al. A general strategy to develop cell-permeable and fluorogenic probes for multicolour nanoscopy. Nat. Chem. 12, 165–172 (2020).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Wang, T. Y. et al. The covalent trimethoprim chemical tag facilitates single-molecule imaging with organic fluorophores. Biophys. J. 106, 272–278 (2014).
Mo, J. et al. Third-generation covalent tmp-tag for fast labeling and multiplexed imaging of cellular proteins. Angew. Chem. 61, e202207905 (2022).
Lukinavicius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Karch, S. et al. A new fluorogenic small-molecule labeling tool for surface diffusion analysis and advanced fluorescence imaging of beta-site amyloid precursor protein-cleaving enzyme 1 based on silicone rhodamine: SiR-BACE1. J. Med Chem. 61, 6121–6139 (2018).
Lukinavicius, G. et al. Fluorogenic probes for multicolor imaging in living cells. J. Am. Chem. Soc. 138, 9365–9368 (2016).
Kilian, N. et al. Assessing photodamage in live-cell STED microscopy. Nat. Methods 15, 755–756 (2018).
Vicidomini, G., Bianchini, P. & Diaspro, A. STED super-resolved microscopy. Nat. Methods 15, 173–182 (2018).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Ghosh, R. P. et al. A fluorogenic array for temporally unlimited single-molecule tracking. Nat. Chem. Biol. 15, 401–409 (2019).
Spahn, C. et al. Protein-specific, multicolor and 3d sted imaging in cells with DNA-labeled antibodies. Angew. Chem. Int. Ed. Engl. 58, 18835–18838 (2019).
Holtmannspotter, M. et al. Reversible live-cell labeling with retro-engineered halotags enables long-term high- and super-resolution imaging. Angew. Chem. Int. Ed. Engl. 62, e202219050 (2023).
Kompa, J. et al. Exchangeable halotag ligands for super-resolution fluorescence microscopy. J. Am. Chem. Soc. 145, 3075–3083 (2023).
Catapano, C. et al. Long-term single-molecule tracking in living cells using weak-affinity protein labeling. Angew. Chem. Int. Ed. Engl. 64, e202413117 (2025).
Du, Z. et al. Self-renewable tag for photostable fluorescence imaging of proteins. J. Am. Chem. Soc. 145, 18968–18976 (2023).
Miller, L. W., Cai, Y., Sheetz, M. P. & Cornish, V. W. In vivo protein labeling with trimethoprim conjugates: a flexible chemical tag. Nat. Methods 2, 255–257 (2005).
Miller, G. P. & Benkovic, S. J. Strength of an interloop hydrogen bond determines the kinetic pathway in catalysis by Escherichia coli dihydrofolate reductase. Biochemistry 37, 6336–6342 (1998).
Baccanari, D. P., Daluge, S. & King, R. W. Inhibition of dihydrofolate reductase: effect of reduced nicotinamide adenine dinucleotide phosphate on the selectivity and affinity of diaminobenzylpyrimidines. Biochemistry 21, 5068–5075 (1982).
Lardon, N. et al. Systematic tuning of rhodamine spirocyclization for super-resolution microscopy. J. Am. Chem. Soc. 143, 14592–14600 (2021).
Yu, Q. et al. Semisynthetic sensor proteins enable metabolic assays at the point of care. Science 361, 1122–1126 (2018).
Watson, M., Liu, J. W. & Ollis, D. Directed evolution of trimethoprim resistance in Escherichia coli. FEBS J. 274, 2661–2671 (2007).
Krucinska, J. et al. Structure-guided functional studies of plasmid-encoded dihydrofolate reductases reveal a common mechanism of trimethoprim resistance in Gram-negative pathogens. Commun. Biol. 5, 459 (2022).
Cammarata, M. et al. Characterization of trimethoprim-resistant E. coli dihydrofolate reductase mutants by mass spectrometry and inhibition by propargyl-linked antifolates. Chem. Sci. 8, 4062–4072 (2017).
Lukinavicius, G. et al. Fluorescent dyes and probes for super-resolution microscopy of microtubules and tracheoles in living cells and tissues. Chem. Sci. 9, 3324–3334 (2018).
Gao, X. et al. Rhodamine-based fluorescent probe for dynamic STED imaging of mitochondria. Biomed. Opt. Express 15, 1595–1604 (2024).
Yang, X. et al. Mitochondrial dynamics quantitatively revealed by STED nanoscopy with an enhanced squaraine variant probe. Nat. Commun. 11, 3699 (2020).
Qiao, C. et al. Rationalized deep learning super-resolution microscopy for sustained live imaging of rapid subcellular processes. Nat. Biotechnol. 41, 367–377 (2023).
Li, D. et al. ADVANCED IMAGING. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Gwosch, K. C. et al. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nat. Methods 17, 217–224 (2020).
Moores, A. N. & Uphoff, S. Exceptional quantification of live-cell single-molecule tracking data for fluorophores with different photophysical properties. J. Phys. Chem. B 128, 7291–7303 (2024).
Zhang, L. et al. Advancing single-molecule biophysics: next-generation organic fluorophores with tailored labeling strategies. Chem. Biomed. Imaging 3, 572–598 (2025).
Liu, T. et al. Gentle rhodamines for live-cell fluorescence microscopy. ACS Cent. Sci. 10, 1933–1944 (2024).
Altman, R. B. et al. Cyanine fluorophore derivatives with enhanced photostability. Nat. Methods 9, 68–71 (2011).
Grenier, V. et al. Molecular prosthetics for long-term functional imaging with fluorescent reporters. ACS Cent. Sci. 8, 118–121 (2022).
Acknowledgements
This work was supported by the National Key R&D Program of China (2024YFA1307400, 2022YFF0708700), National Natural Science Foundation of China (22477016, 32171360, 32101213 to Z.C), International Science and Technology Cooperation Program under the 2023 Shanghai Action Plan for Science, Technology and Innovation (23430711800), Shanghai Pilot Program for Basic Research—Fudan University 21TQ1400100 (22TQ020), Natural Science Foundation of Shanghai (22ZR1404300); Shanghai Science and Technology Innovation Action Plan (22S31905500); Fudan University–CIOMP Joint Fund (FC2020-002); Medical Engineering Fund of Fudan University (yg2021-032, yg2022-2); National Key Research and Development Program of China (2023YFA1801200); National Science Fund for Distinguished Young Scholars (82025019); the Basic Research Pioneer Project by the Science and Technology Commission of Shanghai Municipality (STCSM). We thank Beijing NanoInsights-tech and Abberior for providing instrumentation for superresolution imaging.
Author information
Authors and Affiliations
Contributions
L.W. created the original concept and conducted the work; C.L., D.S., Z.D., and G.X. designed the experiments and performed the imaging experiments; D.S. and G.X. synthesized the probes; D.S., Z.D., and J.L. analyzed the data; L.Y. did the MINFLUX experiments; J.L., M.M., Y.Y., H.Z., B.D., and L.W. discussed the results; L.W., Z.C., L.Y., C.L., and D.S. wrote the paper with input from all the authors. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing financial interest(s): L.W. is the inventor of the patent “Cell-permeable fluorogenic fluorophores”. Lu Wang and Dongjuan Si are inventors on a patent application (Application No. 202510760855.2, titled “Preparation and Application of a Novel Reversible Protein Labeling Tag and Sulfonamide-Based Fluorescent Probes”) filed by [CNIPA]. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Michael Holtmannspötter and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Si, D., Dong, Z., Xing, G. et al. A bright and regenerative array tag with fluorogenic ligands for long-term live-cell nanoscopy and single-molecule tracking. Nat Commun 17, 576 (2026). https://doi.org/10.1038/s41467-025-67273-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67273-8