Abstract
Addressing the need for integrated molecular analysis in complex infectious conditions requires technologies that rapidly quantify both pathogens and host responses. This is particularly relevant in sepsis research, where existing tools typically measure either microbial or host biomarkers. Here, we present MIDAS (Multiplexed Intelligent Diffraction Analysis System), a proof-of-concept platform that integrates shape-encoded hydrogel particles with lens-free diffraction imaging and deep learning–based analysis to enable simultaneous quantification of bacterial RNA and inflammatory proteins in a single system optimized for potential point-of-care use. The assay completes multiplexed measurements in under 4 h, significantly faster than standard culture workflows ( ~ 20–40 h). Beyond spiked studies, MIDAS is evaluated using specimens from a clinically relevant porcine sepsis model, showing high concordance with culture, qPCR and ELISA. Although further clinical validation is required, this flexible platform may support the development of accessible host–pathogen profiling tools with broad applications in healthcare, agriculture, food safety, and beyond.
Similar content being viewed by others
Introduction
Sepsis—life-threatening organ dysfunction caused by infection1—affects more than 1.7 million Americans and kills ~270,000 annually, at a cost of USD$24 billion per year2,3. It is the leading cause of death among hospitalized patients4,5, and survivors of sepsis often face profound long-term health consequences (e.g., neurocognitive impairment)6,7,8. Prognosis for septic patients deteriorates hourly in the absence of appropriate treatment9,10; therefore, earlier identification of sepsis and focused treatment are keys to improve patient outcomes. Unfortunately, current diagnostic methods often fail to provide timely and produce actionable results in busy clinical settings. Clinicians currently rely on nonspecific clinical criteria (e.g., heart rate, blood pressure, respiration rate) to identify potentially septic patients, leading to high false positive rates11,12. Without rapid and specific diagnostic tests, patients with suspected sepsis are frequently subjected to unnecessary and excessive antibiotic administration, propelling the development of antibiotic resistance.
Sepsis pathophysiology is driven by complex interactions between the causal pathogen and the host response, making both sides essential for effective diagnosis1. However, the development of sepsis diagnostics to date has occurred along two parallel, unentwined tracks: (i) one attempting to identify pathogens and (ii) another seeking biomarkers of the host’s response to infection13,14. As such, existing approaches fail to capture the dynamic, two-way signaling between host and pathogens that is crucial for a comprehensive understanding of sepsis.
Current standard methods for pathogen detection and identification (culture-based methods) are slow (up to several days), difficult to apply at the point-of-care (POC), and limited in its ability to identify certain species. Meanwhile, host-response monitoring often relies on one-dimensional biomarker tests (i.e., lactate, procalcitonin, C-reactive protein, MR-proADM), which do not adequately capture the heterogeneity of sepsis15,16. Although advanced technologies, including genomic/transcriptomic technologies, have emerged for pathogen detection and host response monitoring (Supplementary Table 1)17,18,19,20,21,22, they are hampered by slow turnaround (hours to even days), high costs (too expensive for routine and repeated use, especially in low-income countries), high complexity (complex assay procedures and/or specialized bioinformatics expertise required for data interpretation), limited multiplexing capabilities, and a lack of flexibility to swap in other biomarkers for different diseases and conditions. Most importantly, no existing clinically accessible systems can simultaneously measure multiple disparate entities in a single system13,14. Thus, there is a critical need for generic and flexible diagnostic tools that are fast, accurate, practical, and capable of an integrated interrogation of both pathogens and the corresponding host responses—ideally at the POC and without requiring advanced computational tools or expertise, while delivering clear, actionable information to clinicians.
To fill the above gaps, we develop a generalizable strategy — MIDAS (Multiplexed Intelligent Diffraction Analysis System) — for the integrated analysis of multiple host proteins and bacterial genes in a single system. MIDAS synergistically integrates (i) a lens-free diffraction-based optical system for wide-field diffraction image acquisition, (ii) shape-encoded hydrogel sensor arrays for high multiplexing, and (iii) deep learning (DL) algorithms for fast, automated diffraction image analysis, all optimized for portability, simplicity, and POC use in busy clinical and/or low resource settings. MIDAS leverages lensless digital diffraction-based sensing, a technique that records raw diffraction images, defined as two-dimensional (2D) intensity maps of light scattered by particles. These are not direct spatial images of the particles themselves but optical signatures that encode structural and color features. This approach offers advantages over conventional optics (e.g., bright-field and fluorescence microscopy), particularly in terms of miniaturization, cost (<$500 per device), and the ability to achieve a wide field-of-view (FOV, ~30 mm2) with sufficiently high resolution ( ~ 3.3 µm). So far, we and other research groups have proven the utility of digital diffraction imaging in cellular23,24,25,26,27,28,29 and tissue-based30,31 diagnostics across various clinical contexts. To overcome the fundamental limitations of multiplexing and computational complexity that hinder digital diffraction imaging in high-dimensional molecular analysis, we incorporate shape-encoded hydrogel microparticles for single-color multiplexing. We reason that this approach would add a new dimension (shape) to the existing color encoding scheme of the diffraction-based optical system, significantly expanding its multiplexing power from two to virtually unlimited32,33,34,35,36. Additionally, we employ deep learning (DL) for rapid processing of diffraction images on local devices. Raw diffraction images are ideally suited for DL analysis, as they inherently yield large datasets and contain rich information (size, shape, amplitude, phase contrast) in the frequency domain that is hard to recognize by human vision and cognition.
This study presents MIDAS as a proof-of-concept platform that can quantify both proteins and nucleic acid markers in parallel. To the best of our knowledge, this is the first demonstration of a system capable of analyzing both pathogens and host responses simultaneously. We fabricate shape-encoded hydrogel particles using a custom-built microfluidic lithography system and functionalize them with capture probes (antibodies for proteins, oligonucleotides for RNAs) specific to target proteins and nucleic acids. We validate the platform by profiling two sets of markers spiked in human plasma—(i) five host inflammatory proteins and (ii) bacterial RNAs of the top five types of bacteria in positive culture cases—and benchmark MIDAS results against current reference standard methods (culture, qPCR, ELISA, and Luminex). In this work, we adopt the Sepsis-3 definition: life-threatening organ dysfunction caused by infection, irrespective of culture status or pathogen type. Within this framework, the present study focuses on the potential for bacterial sepsis, while the modular MIDAS platform can potentially be extended to other pathogens. Beyond spiked studies, we demonstrate multiplexed analysis of both protein and nucleic acid markers in clinically relevant porcine blood and plasma specimens in under 4 h. Although further validation using human patient samples is required to assess performance for sepsis diagnosis in clinical settings, these results highlight MIDAS’s potential to support future development of diagnostic tools, including possible point-of-care applications. With its modular design, MIDAS may also be adaptable for broader applications (e.g., healthcare, agriculture, food safety) in future work.
Results
Overall assay workflow and marker selection
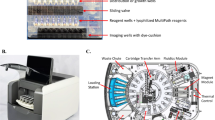
Figure 1 highlights the overall workflow of MIDAS analysis. First, a human biofluid sample (e.g., blood, urine, bronchoalveolar lavage fluid) is collected and split for parallel multiplexed assay of protein and RNA markers. The sample is then processed via ELISA-like colorimetric labeling protocol for protein analysis and hybridization technique for RNA analysis, using a cocktail of shape-encoded hydrogel particles bearing unique ‘gear’ codes to detect multiple target molecules. The stained particles are mounted in a designated area of a glass substrate and imaged by the MIDAS device. A custom-developed DL analysis algorithm is then applied for particle detection, classification, and marker expression readout. The MIDAS diagnostic procedure is completed within 3–4 h for both protein and RNA analyses, substantially faster than the standard culture (2–3 days).
Multiplexed Intelligent Diffraction Analysis System (MIDAS) is designed for the integrated analysis of both pathogens and host response in a single system under 4 h.
To identify or rule out sepsis, we used two sets of markers for the integrated analysis of the host–pathogen interface: (i) bacterial nucleic acids for pathogen detection and phenotyping, and (ii) inflammatory plasma proteins for assessing the host response. Bacteria were chosen as a model pathogen for this study because bacterial infections are the most common cause of sepsis. A recent study reported that bacteremia is present in 54.5% of patients with severe sepsis (n = 1167)37. For bacterial detection and phenotyping, we targeted 16S ribosomal RNA (rRNA) sequence due to their high abundance (103–105 strands in a single bacterium, as opposed to a single strand of genomic DNA), high sequence conservation across bacterial species (useful for general bacterial detection), and species-specific variable regions (useful for species-typing). Importantly, unlike cell-free microbial DNAs, RNAs require viable organisms, making it a better target for identifying patients with true infections. In this proof-of-concept study, we selected initial target markers, based on our preliminary data and literature search38,39,40,41,42,43,44: 1) bacterial 16S rRNA genes from the top five species in positive culture cases—Staphylococcus aureus, Escherichia coli, Enterococcus faecalis, Pseudomonas aeruginosa, and Klebsiella pneumoniae—as well as the universal target (i.e., a highly conserved region of 16S rRNA genes) for quantifying the total bacterial load in a sample, and 2) inflammatory plasma proteins, including IL-1β, IL-3, IL-6, IL-10, and TNF-ɑ, which are currently recognized as biomarkers for sepsis.
Particle design and synthesis for multiplexed assay
For the fabrication of shape-encoded particles, we used our custom-built microfluidic stop-flow lithography system to generate arrays of hydrogel microparticles (probe-particle pairs for target markers) in different shapes and sizes (Fig. 2a). This approach allows for high throughput ( ~ 10 particles/sec), high-resolution ( ~ 2 µm), highly flexible (shape/size), and low-cost manufacturing of hydrogel particles. For pilot testing, we used a simple gear-coding scheme to generate particles that can bear up to 2# of gear teeth unique codes divided by 2 for reflection and the number of gear teeth for rotation as there are no orientation indicators (Fig. 2b). We designed particles to be read with gear teeth used as means of encoding (0 = absent; 1 = present). In this study, we designed gear particles with eight teeth, so we have a total of 16 unique codes (28/(2・8) = 24). Of those, we selected five specific codes for the following protein and RNA analyses (Fig. 2c).
a Schematic of the stop-flow lithography system used for the high-throughput synthesis of shape-encoded hydrogel microparticles. b Shape encoding scheme allows generation of numerous unique codes and single-color multiplexed detection, achieving up to 16-plex with the gear design used in this study. c Comparison of particle imaging methods. Bright-field microscopy (BFM), raw diffraction (MIDAS), and numerically reconstructed phase (Recon) images. Raw diffraction images appear undecipherable. This experiment was independently repeated three times with similar results. d Schematic of the lensless diffraction-based optical sensor. The device is equipped with a high-resolution CMOS imager and an LED (λ = 625 nm) light source. The system body was fabricated in a photopolymer resin via 3D printing. e Field-of-view (FOV) of MIDAS ( ~ 30 mm2). Inset: MIDAS captures ~5,000 particles per acquisition, compared to ~820 particles using BFM. f Disposable sample cartridge for particle capture and imaging. g MIDAS signals strongly correlated with those measured by BFM (two-sided Pearson’s correlation, r = 0.98, P = 2.1 × 10-14). Comparison between BFM, raw diffraction (MIDAS), and numerically reconstructed transmittance (Recon) images of color-labeled particles. Data are shown as mean ± s.d. from technical replicates (n = 7 for e and 3 for g). Source data are provided as a Source Data file.
MIDAS sensor for large field-of-view analysis
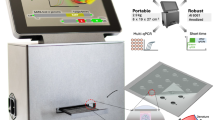
Building on our previous imager prototypes23,45, we designed the MIDAS sensing hardware to offer a compact, cost-effective system for wide-field diffraction image acquisition of shape-encoded hydrogel particles (Fig. 2d and Supplementary Fig. 1). Notably, conventional hydrogel particle analysis using one-way microfluidic channels has faced limitations in throughput, speed, and accuracy46. We reasoned that the diffraction-based optical sensor could address these challenges. The device consists of a light-emitting diode (LED), an aperture, and a complementary metal-oxide-semiconductor (CMOS) board camera. The interference patterns between the incident light and the scattered light from particles were directly recorded by the camera positioned underneath the sample. The sensing system was compact (7.5 (L) × 9.5 (W) × 15 (H) cm3) and cost-effective (~$330) with no intermediate optical components (e.g., lenses, filters) necessary (Supplementary Table 2).
We compared MIDAS performance with bright-field systems of comparable accessibility and cost: a conventional bright-field microscope (BFM), a digital handheld microscope, and a smartphone with a clip-on lens (Supplementary Fig. 2). Hydrogel particles (stained, unstained, or both) were imaged using all systems. The BFM (Nikon Eclipse TS100 with 4× objective; ~2.22 µm pixel size) provided a FOV of ~11 mm2 (~6.3 mm2 normalized to MIDAS resolution). The digital microscope (Andonstar V160; ~1.6 µm pixel size) yielded a FOV of ~4.9 mm2 (~5.3 mm2 normalized). The smartphone clip-on lens system (Samsung Galaxy S21 with Mobile Phone Lens; ~1.62 µm pixel size) offered a FOV of ~20 mm2, though with somewhat lower effective resolution for capturing finer particle features, likely due to the auto-focusing mode and limited numerical aperture of the clip-on lens system. By comparison, MIDAS provided a FOV of ~30 mm2—comparable to or slightly greater than the comparator systems—while offering sufficient resolution for feature-level analysis (Fig. 2e). In practice, this allowed detection of ~5000 particles in a single acquisition, compared to ~820 particles using the BFM (Fig. 2e, inset). When all systems were evaluated under the same monochromatic LED illumination used by MIDAS, comparator bright-field devices showed reduced contrast and feature visibility, consistent with their optimization for broadband white-light imaging, whereas MIDAS preserved particle feature visibility (Supplementary Fig. 2c). For applications requiring analysis of concentrated samples or parallelized compartments, custom PDMS microfluidic chips can be incorporated to further increase analysis capacity (Fig. 2f). We also tested whether diffraction patterns can be used to extract color information on particles. For conventional BFM, we obtained the same metrics by measuring blue levels in micrographs. The MIDAS color intensity profiles showed a high correlation with those from microscopy (Pearson’s correlation coefficient r = 0.98; Fig. 2g), confirming that diffraction-based sensing maintains color information on stained particles. Furthermore, the numerically reconstructed images also matched well with conventional BFM (Fig. 2g), providing additional validation of the system’s ability to maintain accurate color representation.
Deep learning (DL) algorithm for fast and accurate particle analysis
Conventional diffraction image analysis relies on computationally intensive reconstruction that typically require GPU-equipped servers or cloud-based processing. These requirements limit practicality in point-of-care or resource-limited settings, where reliable high-bandwidth connectivity is often unavailable. To address these limitations, we developed a DL-based workflow that directly interprets raw diffraction patterns, eliminating the need for image reconstruction. After centralized training, the resulting models can be deployed locally on standard computers (or even low-power microcontrollers) to enable fast, fully offline analysis.
A three-stage DL workflow was designed to automatically locate and classify barcoded particles in raw diffraction images: (1) particle recognition, (2) code classification, and (3) color readout. In the first stage, candidate particle regions were identified using a blob detection algorithm, followed by a convolutional neural network (CNN) trained to distinguish particles from background artifacts (Fig. 3a). The model was trained on a dataset of annotated diffraction images (930 particle and 88,357 non-particle), with data augmentation to improve performance across variations in stain level and orientation. Using a 70/10/20% train/validation/test split, the model achieved validation and test accuracies of ~100% and ~99.2%, respectively (Fig. 3b, and Supplementary Fig. 3). Bayesian optimization identified the optimal architecture within 47 trials, with an average 5-fold cross-validation accuracy of ~100%. The second stage of code classification used a separately trained CNN (Fig. 3c) to classify the code (i.e., gear design in this study) associated with each detected particle. The model was trained and evaluated using a 70/10/20% train/validation/test split. It achieved a best validation accuracy of ~100% and a test accuracy of ~94.1% (Fig. 3d, Supplementary Fig. 3). Bayesian optimization identified the final model architecture within 32 trials, with an average 5-fold cross-validation accuracy of ~98.0%. While our results demonstrate high learning curve accuracies, over 1000 epochs was needed to achieve accuracies > 90%. The accuracy trends remain stable yet volatile at ~4000 epochs onwards, reflecting the greater complexity of distinguishing between similar gear shapes compared to simple particle detection.
a Particle classifier. The input to this module was a particle-like raw diffraction image (128 × 128 pixel 2D image) proposed by a maximally stable extremal region blob detection algorithm. The image was processed through a trained convolutional neural network (NN) consisting of two convolutional layers (Conv2D), two pooling layers (Max pool), and two fully connected layers (FC). The NN labeled the image as a particle (1) or a non-particle (0) diffraction pattern. ReLU, rectified linear unit; Pd, dropout rate. b The particle detection NN was trained with 930 original particle and 88,357 non-particle diffraction patterns as well as augmented dataset. The held-out test set accuracy reached ~99.2% for particle detection. c Code classifier. Diffraction images classified as particles from (a) entered the next module for code classification. The code classification NN had eight Conv2D and three FC layers and produced the final output of shape classes or codes. PReLU, parametric ReLU. d The code classification network was trained with 930 original diffraction images and augmented dataset. The held-out test set accuracy was ~94.1%. Source data are provided as a Source Data file.
To further assess robustness, we evaluated the impact of z-position variability on classification accuracy, given that diffraction images can, in principle, be sensitive to particle height. MIDAS minimizes this variability in two ways: (i) particles fabricated by stop-flow lithography exhibit highly uniform thickness (19.9 ± 0.646 µm; Supplementary Fig. 4a)47,48, and (ii) particles sediment onto the top surface of a glass coverslip placed directly on the sensor, serving as the substrate, which ensures they lie in a common focal plane and maintain a constant particle-to-sensor distance. Our DL model, trained on raw diffraction images collected from multiple No. 1 coverslips as substrates (manufacturing tolerance ±5–20 µm), achieved consistently high performance in particle detection ( ~ 99%) and code classification ( ~ 94%), indicating robustness to modest height variations. Furthermore, when we compared coverslips of different thicknesses (No. 1, 150 ± 20 µm; No. 1.5, 170 ± 20 µm), we observed no measurable impact on DL accuracy (Supplementary Fig. 4b, c). Together, these results demonstrate that MIDAS is robust to the range of z-position variations expected under our assay conditions.
In the last stage of color readout, the algorithm simply computes the average pixel intensity inside a small area surrounding the detected particle. This intensity is then referenced against a calibration curve to determine the expression level of the corresponding target biomarker. The computed values showed strong correlation with colorimetric signals measured by bright-field microscopy (Fig. 2g), confirming that the diffraction-based method preserves staining information with high fidelity.
MIDAS assay characterization for protein analysis
The wide-field diffraction image acquisition capability and multiplexing power of MIDAS highlight the platform’s promise for comprehensive molecular profiling. First, we characterized the MIDAS assay for multiplexed protein analysis using a set of known and promising inflammatory biomarkers for sepsis. Figure 4a outlines the overall procedure of the MIDAS-protein assay. Briefly, the colorimetric labeling protocol involves: (i) conjugation of particles with capture antibodies, (ii) target extraction from plasma using a cocktail of antibody-conjugated particles, (iii) labeling particles with detector elements, and (iv) colorimetric signal amplification via alkaline phosphatase (AP) and BCIP/NBT substrate. The resulting precipitated indigo-blue color products accumulate within particles, enabling quantification of target biomarkers via color intensity (Fig. 4b).
a Assay schematic for target analyte detection. Target analytes (e.g., proteins, nucleic acids) in patient biofluid samples (e.g., blood, urine) are colorimetrically labeled with hydrogel microparticles. The enzyme-mediated color development method using an enzyme-substrate pair [alkaline phosphatase (AP) and BCIP/NBT] catalytically converts substrate molecules into insoluble color products that locally precipitate and amplify on the particles. b Representative bright-field and MIDAS raw diffraction images of hydrogel particles after being labeled with target analytes spiked in human plasma at different concentrations. c Comparison of IL-3 detection using MIDAS and conventional ELISA. d Titration curves of IL-1β, IL-3, IL-6, IL-10, and TNF-α in diluted human plasma. e MIDAS signals for differential detection of various cytokines. Note the high specificity and low background noise compared to other cytokines. f Heatmap of obtained MIDAS signals for detection specificity. Data represent mean ± s.d. of three independent experiments. a.u. arbitrary unit. Source data are provided as a Source Data file.
We first benchmarked MIDAS against a commercial ELISA kit for IL-3 detection in human plasma. MIDAS demonstrated a 28-fold improvement in sensitivity, with a limit of detection (LOD) of 4.09 pg/mL compared to 115 pg/mL for ELISA (Fig. 4c). The LOD was defined as the target concentration yielding a signal equal to three times the standard deviation of blank samples. We then extended the analysis to the five cytokines, including IL-3, IL-6, IL-10, IL-1β, and TNF-α (Fig. 4d) and observed that MIDAS consistently achieved higher sensitivities (i.e., lower LODs) across all targets compared to ELISA, while maintaining compatible or broader dynamic ranges (Supplementary Table 3).
For multiplexing, the MIDAS assay showed excellent sensitivity and specificity among the five targets with negligible cross-reactivity between targets (Fig. 4e, f and Supplementary Table 4). The assay also showed high accuracy, with recovery rates for all targets falling within the commercially acceptable range (70-130%)49 (Supplementary Table 4).
MIDAS assay characterization for nucleic acid analysis
Next, we evaluated MIDAS for multiplexed detection of bacterial nucleic acids. The MIDAS-nucleic acid assay uses a sandwich hybridization technique in which each hydrogel particle carries a capture oligonucleotide that selectively binds a target sequence generated from bacterial RNA. After hybridization, a second oligonucleotide labeled with biotin is introduced to tag the opposite region of the same amplicon, enabling enzymatic colorimetric readout through streptavidin-alkaline phosphatase (StAv-AP) and BCIP/NBT development (Fig. 5a). Target-positive particles accumulate an intense blue precipitate, whereas negative controls show minimal background, confirming the specificity of the hybridization conditions (Supplementary Fig. 5). Probes and primers specific to bacterial targets are adapted from literature50 and modified as needed (Supplementary Table 5).
Probes targeting hypervariable regions of bacterial 16S rRNA sequences were used to specifically detect various bacterial types. RNA was extracted from bacterial cultures, amplified by asymmetric RT-PCR (35 cycles) using specific primers for each bacterial type, and detected using the corresponding probe conjugates. a Sequences of probes specifically targeting a conserved region of bacterial 16S rRNA. Two probe sequences with a single base difference were used for capture and detection purposes. b Serial dilutions of synthetic DNA or bacteria-derived DNA were used as detection targets. Bacteria-derived DNA molecules were obtained via asymmetric RT-PCR of E. coli 16S rRNA (35 cycles). Synthetic DNA had the same sequence as bacteria-derived DNA. The detection limit was 3.01 pM. c Bacterial detection in blood. Serial dilutions of E. coli were spiked into human blood and processed by extracting the RNA using the same procedure as described above. d Universal detection of bacteria for quantifying the total bacterial load in a given sample. Universal probes detected five individual bacterial types (each at 100 CFU) as well as mixtures of different bacterial types. The dotted line represents the mean signal for a single bacterial species. e MIDAS signals for differential detection of various bacterial types. Note the high specific signals and low background noise against other bacteria. f Heat maps comparing the specificity of the MIDAS assay with that of qPCR. Data represent mean ± s.d. of three independent experiments. a.u. arbitrary unit. Source data are provided as a Source Data file.
To determine analytical sensitivity of the MIDAS-nucleic acid assay, we first examined Escherichia coli as a representative model organism. Serial dilutions of synthetic oligonucleotides and cDNA generated from bacterial RNA were analyzed, yielding a LOD of ~3.0 pM [DNA] (Fig. 5b), equivalent to ~150 amol of target in a 50-µL sample. Because clinically relevant bacteremia often involves extremely low organism burdens, we next spiked whole blood with defined numbers of E. coli. After sample processing and amplification, MIDAS reliably detected as few as ~2 bacteria per mL of blood (Fig. 5c). The number of PCR cycles affected the dynamic range: ~35 cycles yielded the broadest quantification window, whereas additional cycles increased sensitivity at the cost of a narrower linear range (Supplementary Fig. 6). Increased variability at the lowest concentrations was consistent with expected stochastic sampling during serial dilution.
The assay was then expanded to five clinically relevant pathogens associated with sepsis—Staphylococcus aureus, Escherichia coli, Enterococcus faecalis, Pseudomonas aeruginosa, and Klebsiella pneumoniae. Using species-specific probe sets (Supplementary Table 5), MIDAS achieved LODs ranging from 1.41 to 35.8 pM (Supplementary Fig. 7). The platform generated quantitative signals across several orders of magnitude of bacterial load, enabled in part by the inherently high abundance of 16S rRNA (103–105 copies per bacterium), which supports robust amplification and sensitive detection.
To support broad pathogen screening, we also implemented a “universal” detection strategy targeting a conserved region of the 16S rRNA gene across multiple species. Two capture sequences differing by one nucleotide were paired with a common biotinylated detection probe. This configuration produced consistent signals across different species and accurately reflected the total bacterial burden in mixed samples containing ~100 CFU of each organism (Fig. 5d), demonstrating its utility as a first-line screening tool for pathogen presence.
Finally, we applied MIDAS for species-level discrimination using probes directed to hypervariable regions of the 16S sequence. Using synthetic oligonucleotides and RNA-derived amplicons from each of the five species, the multiplexed assay showed distinct, non-overlapping signals for each species with minimal off-target binding (Fig. 5e, and Supplementary Table 6). Recovery rates (70–130%) were within accepted analytical ranges. Compared to qPCR, MIDAS exhibited improved specificity (Fig. 5f), likely due to its requirement for simultaneous recognition by both capture and detection probes. Hybridization conditions (e.g., temperature, ionic strength, incubation time) can be tuned to further minimize non-specific interactions. Thanks to its flexible and modular design, the platform may also be adapted to differentiate phenotypic traits such as antimicrobial resistance within closely related bacterial strains.
MIDAS validation
We evaluated the clinical utility of the MIDAS assay using clinically relevant specimens derived from a well-established swine model of sepsis51,52. This model offers several key advantages over human studies for early-stage diagnostic validation: (i) it provides definitive ground truth regarding infection etiology, (ii) it enables high-frequency serial sampling—including pre-sepsis baseline measurements—for internal comparisons, (iii) it permits collection of larger blood volumes needed for parallel testing with current standard assays (e.g., blood culture, qPCR, ELISA), and (iv) it avoids common confounding variables such as underlying comorbidities present in human cohorts.
To confirm that the swine infection model reflected sepsis rather than isolated bacteremia, we evaluated animals using the Sepsis-3 definition of sepsis: life-threatening organ dysfunction caused by infection1. Operationally, Sepsis-3 defines sepsis as an increase in sequential organ failure assessment (SOFA) score of ≥2 points due to infection. Using available data from the pigs and SOFA score definitions (Supplementary Table 7), we determined organ-specific SOFA scores (Supplementary Table 8) at multiple timepoints between baseline (pre-infection) and experiment termination (range 6-24 h). Following E. coli inoculation, all animals showed an increase of ≥2 points in at least one organ system SOFA score, thereby meeting Sepsis-3 criteria. These findings support that this swine model reproduces the clinical syndrome of sepsis as currently defined.
Whole blood and plasma samples were collected from swine at baseline and at multiple timepoints up to 24 h post-infection after injecting E. coli inoculum into the renal parenchyma. Samples were stored ( − 80 °C) until assay at which time they were analyzed using current reference standard diagnostic methods (culture, qPCR, ELISA/Luminex) alongside MIDAS assays, which completed both bacterial RNA and host protein profiling in under 4 h. For swine samples, we adapted the MIDAS protein assay by substituting porcine-specific antibodies and optimizing the assay conditions accordingly (Supplementary Fig. 8). Note that IL-3 was not included in the porcine panel because no commercial porcine IL-3 antibodies were available.
Figures 6–7 present representative results from the MIDAS assays, and Supplementary Table 9 provides a side-by-side comparison of assay performance between MIDAS and conventional methods. Cutoff thresholds for marker positivity were defined as 3× SD of control samples (Supplementary Table 10). Across the tested samples, MIDAS showed high concordance with culture and qPCR, accurately detecting bacterial presence and species in 30 out of 32 samples (93.8%). The two discordant cases (#4 T12 and #5 T4) had very low bacterial loads (1 CFU by culture), where non-detection may reflect sampling constraints. In parallel, cytokine measurements by MIDAS correlated strongly with reference methods (Luminex/ELISA), with a Pearson correlation coefficient (r = 0.87).
a, b Detection of bacterial pathogens using MIDAS with (a) universal and (b) specific probes for bacteria types. Total RNA was extracted from the blood samples (400-µL aliquot from each sample) and amplified via asymmetric RT-PCR (40 cycles) using both universal and specific primers for bacterial types. Amplicons were then detected by the MIDAS assay. In a all baseline samples (n = 8) tested negative and post-inoculation samples (n = 8) tested positive, consistent with standard culture and qPCR results. ND indicates no pathogen detected. In b MIDAS correctly identified the pathogen types in post-inoculation samples. Negative values within 3 × SD of the background signal were set to zero. c Heatmap summarizing MIDAS signal intensities for universal and specific detection across samples. Whole blood samples were collected from eight swine at baseline (denoted as BL) and post-inoculation (e.g., T2) with E. coli. TD indicates time of death. Data represent mean ± s.d. from two-three repeated tests. a.u. arbitrary unit. Source data are provided as a Source Data file.
a Heatmap comparing MIDAS results with the reference standard (Luminex/ELISA) across all samples. Serial plasma samples (n = 17) were collected from five swine at baseline (BL) and post-inoculation time points (e.g., T2) with E. coli. A 40-µL aliquot from each sample was used for the MIDAS assay to detect a panel of protein markers, including IL-1β, IL-6, IL-10, and TNF-α. b MIDAS results showed strong match with reference measurements (two-sided Pearson’s correlation, r = 0.87, P = 4.5 × 10-26). Each data point represents the mean value of two-three repeated tests. Data are plotted on log–log scales to better visualize low-concentration values. The dashed line indicates the best linear fit. Source data are provided as a Source Data file.
Although these initial findings are promising, further validation studies using fresh biological samples are needed. This will require increasing assay throughput, as well as larger-scale prospective swine studies and ultimately clinical validation using patient samples.
Discussion
In this study, we developed MIDAS, a multiplexed assay platform that integrates protein and nucleic acid analysis to address a key gap in sepsis diagnostics: the simultaneous detection of pathogens and host immune responses—an essential but often overlooked aspect of current diagnostic strategies. As summarized in Supplementary Table 1, several recently FDA-cleared platforms—including BioFire® BCID2, T2Dx®, IntelliSep™, TriVerity™, SeptiCyte RAPID™, Sepsis ImmunoScore™, and Abionic’s IVD CAPSULE PSP™—have made important advances in either pathogen identification or host-response profiling to improve diagnostic accuracy. However, these technologies typically focus on a single diagnostic axis and often rely on centralized laboratory infrastructure. In contrast, MIDAS is, to the best of our knowledge, the first system to enable molecular multiplexing of both pathogen and host biomarkers within a single platform, offering promising potential for future development toward sepsis diagnostics and research applications, pending further clinical validation.
In addition to its clinical relevance, MIDAS represents a significant technological advancement. It builds upon two well-established strategies: (i) lens-free diffraction-based sensing and (ii) shape-encoded microparticle-based bioassays. Lens-free diffraction-based optical sensing/imaging has been widely recognized as a powerful, low-cost method for wide-field diffraction image acquisition, enabling large-area analysis without complex optics. Digital platforms, including our own, have demonstrated strong performance in cellular23,24,25,26,27,28,29 and tissue30,31 analysis across large fields of view using minimal optical hardware. However, their application in molecular diagnostics has been limited, largely due to reliance on low-plex color encoding schemes23. In parallel, shape-encoded hydrogel microparticles have been used in prior studies, including our own, for multiplexed detection of biomolecules (e.g., proteins, nucleic acids) using both fluorescence and colorimetric readouts32,33,34,35,36,53,54,55. While these studies established important groundwork, decoding in such systems is typically rule-based and often limited by particle orientation, image quality, and fabrication variability.
To address these limitations, MIDAS synergistically integrates shape-encoded hydrogel microparticles with lens-free digital sensing and DL-based decoding. Compared to traditional multiplexing platforms such as planar arrays and fluorescence-based microbeads, shape-encoded particles offer distinct advantages, including a high surface-to-volume ratio, rapid and sensitive reaction kinetics, high encoding capacity, and improved decoding robustness56,57. Despite these advantages, traditional decoding approaches, such as coulter counter58, flow cytometry59, and rule-based computer vision36, remain limited in throughput and accuracy. For example, flow cytometry and Coulter counters require unidirectional particle alignment through microfluidic channels, which restricts particle design and reduces encoding capacity58,59, while computer vision systems often fail when particles are flawed, occluded, or randomly oriented46. MIDAS overcomes these challenges by enabling robust, orientation-invariant decoding directly from raw diffraction images using custom DL models. Our workflows are optimized for multiplexed analysis of shape-encoded particles on low-power devices (e.g., microcontrollers, smartphones), eliminating the need for computationally intensive image reconstruction. The resulting platform is compact, low-cost, and well suited for future POC deployment. To the best of our knowledge, MIDAS is the first system to combine these three elements—shape-encoded particles, lens-free diffraction sensing, and DL-based decoding—into a single, self-contained platform for simultaneous, multiplexed analysis of large particle populations.
MIDAS offers complementary capabilities to conventional diagnostic methods such as ELISA, standard culture, and qPCR, particularly in detection sensitivity, multiplexing capability, assay time, cost, and it can also be easily applied to POC scenarios. In this proof-of-concept study, the MIDAS-protein assay demonstrated 5-plex protein detection with a detection sensitivity (LOD) of 4.09 pg/mL for IL-3, which is a > 25-fold improvement compared to the 115 pg/mL LOD of standard ELISA. Similarly, the MIDAS-nucleic acid assay showed high specificity and sensitivity in both detection and phenotyping a panel of 5 types of bacteria, achieving LOD as low as ~2 pM for bacterial pathogens using 35 PCR cycles, equivalent to ~3-4 CFU/mL in biofluid sample. Notably, MIDAS sensitivity could be further enhanced by increasing the PCR cycle number to 40, enabling detection of as few as 2 bacteria in 1 mL of blood. This level of sensitivity is crucial given that the microbial load in septic patient blood is typically low, ranging between 1 and 100 CFU/mL60,61. These results highlight the potential of MIDAS as a promising tool for both pathogen detection/identification and host-response monitoring in experimental settings. Further validation, including assessment of diagnostic accuracy in clinical samples, is required to determine its clinical applicability.
The current work was designed to prove the overall MIDAS concept. As such, extensive clinical testing was beyond the scope of this developmental study. However, the data from the first prototype suggest several areas for improvement:
First, although MIDAS is ultimately intended for use at the POC, the current system is an early-stage prototype. To realize a true sample-in, answer-out device suitable for urgent care, rural clinics, or emergency response environments, it will be essential to automate sample preparation and fluid handling. Integration with microfluidic cartridges62,63 and simplified few-step protocols64,65,66 could substantially reduce assay time, minimize user intervention, and enhance usability. Our demonstration of MIDAS operation in an outdoor, non-laboratory setting (Supplementary Fig. 9) underscores its portability and robustness, but further engineering advances are needed for clinical deployment.
Second, in this proof-of-concept, we demonstrated 5-plex detection using 16 available particle codes. While modest compared to high-end systems such as spectral flow or mass cytometry ( > 40-plex), those platforms often depend on costly instrumentation and centralized infrastructure. In contrast, MIDAS is designed to fill a different niche: a compact, low-cost platform capable of simultaneous RNA and protein detection at the point of need. Moreover, we have previously reported alternative particle designs46 with substantially higher multiplexing potential ( > 500-plex), underscoring the scalability of this approach for future applications.
Third, the lensless optical design of MIDAS provides a wide field of view with sufficient resolution to capture thousands of particles in a single frame, which enables rapid and efficient multiplexed analysis within a compact, low-cost format. While the current system is implemented in a batch-loading format, integration with continuous or stop-flow microfluidics, along with resolution-enhancing strategies (e.g., subpixel shifts, structured illumination, or DL-based super-resolution) could further increase both throughput (i.e., sample processing capacity) and resolution. In this proof-of-concept study, we prioritized a design that balances practicality, simplicity, and performance for potential POC implementation—intended to complement, rather than compete with, rapidly evolving optical systems and advanced high-throughput technologies.
Fourth, MIDAS achieves detection limits down to 2 CFU/mL of blood—meeting or exceeding conventional culture thresholds. However, detection of a single bacterium (e.g., ~1 CFU/mL in biofluid) remains inherently constrained by sampling, as the organism may simply be absent from the tested aliquot. This is a fundamental limitation common to all diagnostic platforms, regardless of sensitivity. To further mitigate this challenge, MIDAS could be coupled with additional signal amplification strategies such as hybridization chain reaction54, tyramide signal amplification67, and rolling circle amplification68—all of which we have previously validated in hydrogel particle systems—and potentially integrated with high-sensitivity fluorescence detection.
Fifth, our study addresses sepsis as defined by Sepsis-3: life-threatening organ dysfunction caused by infection, irrespective of culture status or pathogen type. While Sepsis-3 defines sepsis independent of culture results, culture negativity remains a major diagnostic challenge in clinical practice; it often reflects (i) bacterial infections missed due to prior antibiotics, low pathogen burden, or sampling limitations—fundamental limitations that affect all diagnostic platforms, including culture and qPCR—or (ii) non-bacterial etiologies (e.g., viral or fungal), which would require incorporating additional targets. Within this framework, the present implementation focuses on bacterial sepsis, detecting bacterial RNA and host cytokine markers to capture both pathogen- and host-side signatures on a compact, low-cost platform. Although demonstrated here as a proof-of-concept, its modular design could potentially be extended to include viral and fungal etiologies in future work. Future studies will evaluate performance in culture-negative bacterial cases and non-bacterial sepsis using clinically relevant samples.
Sixth, the present system supports species-level detection, which is consistent with conventional clinical practice using cultures, where species identification is generally sufficient to guide empiric antibiotic therapy while awaiting susceptibility results. Strain-level resolution and antimicrobial resistance profiling are not yet implemented in this study, but these capabilities could be added by incorporating primers or probes targeting resistance markers (e.g., mecA, vanA, PVL genes)50 or SNPs associated with drug resistance68. The modular design of MIDAS facilitates such adaptation, as illustrated by our preliminary tests with synthetic mecA and PVL targets, which showed highly specific detection (Supplementary Fig. 10) but have not yet been fully validated. Rather than a limitation, this is a clear and achievable next step toward expanding the clinical utility of MIDAS.
Seventh, our retrospective validation study demonstrated that MIDAS can detect both pathogen and host signals from clinically relevant swine specimens; however, a limitation is that the whole blood samples were frozen, which can compromise bacterial viability and lead to underrepresentation of true bacterial burden. Since both the MIDAS assay and the reference standard methods (culture and qPCR) were performed on the same set of frozen samples, the comparative assessment remains valid.
Lastly, although a full cost analysis for clinical implementation would need to account for labor, infrastructure, quality control, and regulatory compliance, our prototype-stage estimates place core component costs at ~$350, with reagent costs of ~$5 per 5-plex protein assay and ~$20 per 5-plex RNA assay (Supplementary Table 2), based on raw material costs under academic research conditions. Although bacterial cultures are often perceived as cheap, they indeed require substantial infrastructure and skilled personnel, limiting use in many global settings. By contrast, rapid molecular tests like MIDAS may be more scalable and affordable—similar to how GeneXpert has replaced culture-based tuberculosis testing in much of the world where lab access is limited. Our long-term goal is to make the system an affordable and accessible diagnostic solution across both high- and low-resource settings, which could significantly impact global health and address equity challenges. Building on the retrospective validation study using a swine model of sepsis presented here, we plan to pursue larger-scale prospective swine studies and ultimately clinical validation with patient samples. These efforts will be essential for confirming the robustness, accuracy, and real-world applicability of MIDAS across diverse clinical scenarios.
In summary, the MIDAS platform provides a modular, multiplexed approach for simultaneous analysis of pathogen and host biomarkers. By addressing key limitations of current diagnostic methods—in particular, slow turnaround times, high costs, limited multiplexing capabilities, and low flexibility in handling various biological entities—MIDAS could offer a promising foundation for future development toward improved sepsis diagnostics, pending further clinical validation, including assessment of diagnostic accuracy using clinical samples. Its versatile and modular design will allow for expansion beyond the initial sensor elements and marker panels selected in this study (e.g., to include additional sepsis etiologies and antimicrobial resistance genes). With further validation and development, possibly in conjunction with its capability for single-cell analysis23, tissue pathology31, and viral plaque assays69, we envision that this versatile approach could have broad implications across multiple domains.
Methods
Materials
The SU-8-coated Si wafer, patterned positively with a microchannel path, was purchased from MicroFit (Gtyeonggi-do, Korea). Polydimethylsiloxane elastomer (PDMS, Sylgard 184) was purchased from Dow Corning (Michigan, USA). Poly(ethylene glycol)diacrylate (Mn 700 Da), Poly(ethylene glycol) (Mn 600 Da), Poly(ethylene glycol) (Mn 200 Da), a 2-hydroxy-2-methypropiophenone (Darocur 1173) photoinitiator, Tris-EDTA buffer solution, Tween 20 and Human serum were purchased from Sigma-Aldrich (MO, USA). 100x Tris-EDTA buffer solution, sodium chloride, Streptavidin-alkaline phosphatase (StAv-AP) and nuclease-free water were purchased from Thermofisher scientific (MA, USA). Phosphated-buffer saline (PBS) was purchased from Welgene (Gyeongsangbuk-do, Korea). A BCIP/NBT alkaline phosphatase substrate kit was purchased from Vector Labs (CA, USA). 1 M Tris-HCL pH 9.0) was purchased from BioPrince (Gangwon-do, Korea). Antibody-antigen sets for sandwich immunoassay were purchased from R&D systems. All oligonucleotides used in this study were synthesized by Integrated DNA Technologies (IA, USA) and Bioneer (Daejeon, Korea). Human/porcine plasma and whole blood samples were obtained from Innovative Research as commercially sourced, fully de-identified materials.
Optical Systems
MIDAS sensing system
The light source had a 625-nm LED (M625D3, Thorlabs) paired with a 220-grit optical diffuser (DGUV10-220, Thorlabs) and a 100-μm aperture (Thorlabs). Optical components were aligned inside a custom-made holder. Diffraction images were recorded by a monochromatic 10.7-megapixel CMOS imager (DMM 27UJ003, The Imaging Source). The camera pixel size is 1.67 μm, and the field of view 6.5 × 4.6 mm2. We made a device body using a 3D printer (Prusa). A sample tray was designed for easy sample loading and alignment. The overall size of the device was 7.5 (L) × 9.5 (W) × 15.0 (H) cm3.
Benchmarking systems
For benchmarking, we compared four optical systems. MIDAS (described above) operated at ~50 lux. A bright-field microscope (BFM, Nikon Eclipse TS100) was equipped with a Plan Fluor 4 × /0.13 NA objective (Nikon) and a Basler acA1920-40uc camera (Edmund Optics), using the built-in white LED source ( ~ 105 lux). A digital handheld microscope (V160, Andonstar Technology) was equipped with a built-in lens with an integrated 8-LED ring light ( ~ 104 lux). Finally, a smartphone + clip-on lens system was tested using a Samsung Galaxy S21 paired with a universal 37-mm 0.45× wide-angle + macro clip-on lens (labeled ‘2-in-1 lens, 37 mm, 0.45× wide-angle + macro,’ purchased from AliExpress) under ambient laboratory lighting ( ~ 300-1000 lux).
Hydrogel particle synthesis and modification
PDMS microfluidic device fabrication
Microfluidic devices were initially designed using AutoCAD (Autodesk, CA, USA) and printed on photomask films (Han & All Technology, Gyeonggi-do, Korea). An SU-8 master mold was fabricated via conventional photolithography using silicon wafers coated with a negative photoresist, SU-8 25 with a thickness of 23 μm. PDMS mixed at a 10:1 base to curing agent ratio was poured on the SU-8 master mold and cured for 4 h at 70 °C. Following curing, the PDMS slab was detached from the SU-8 master and perforated using 1.0 mm and 10.0 mm biopsy punches (Miltex, PA, USA) to create inlet and outlet reservoirs, respectively. The PDMS mixture was coated onto glass slides and partially cured at 70 °C for 25 min. Subsequently, the prepared PDMS slab was affixed to the partially cured surface of the glass slide and allowed by curing overnight at 70 °C.
Hydrogel microparticle synthesis via Stop-flow lithography (SFL)
To execute SFL, a UV-pressure synchronization system controlled by a customized circuit board and a LabView (National Instruments, TX, USA) script was developed. With the microfluidic device mounted on an inverted microscope (Axiovert 200, Zeiss, Oberkochen, Germany), the prepolymer- loaded pipette tip was fitted into the microfluidic device. The prepolymer was injected into the device using air pressure regulated by a pressure regulator (ITV0031-3BL, SMC, Tokyo, Japan). A high-resolution photomask with various patterns (25 000 dpi) was embedded into the field-stop of the microscope. A LED lamp (Mounted high-power LED M365L2, Thorlabs, NJ, USA) served as the photo-polymerization source, and its intensity was controlled by an LED driver (T-Cube high power LED Driver, Thorlabs, NJ, USA). For this work, the UV intensity was kept constant at 3,000 mW/cm2.
Probe conjugation to hydrogel microparticles
For capture antibody conjugation, we used the ‘linker-free antibody conjugation method’ that combines the thiol group present in the antibody with the unreacted acrylate double bonds (UADBs) (C = C) present in the hydrogel microparticle70. This approach utilizes the unreacted acrylate double bonds (UADBs) remaining within the particles and facilitates the thiol coupling via a Michael addition reaction71. To initiate the process, 5 µL of 0.2 mM TCEP diluted in PBST (prepared by adding 0.05% v/v of Tween 20 in 1X PBS buffer) is combined with 5 uL of capture antibody diluted in PBST (12 µg/µL for IL-1 beta/IL-F2, 6 ug/µL for IL-3, 6 µg/µL for IL-6, 6 µg/µL for IL-10, 12 ug/uL for TNF-a), reducing the antibody for 1 h at room temperature. Next, 10 µL of the reduced capture antibody is incubated with 10 µL of particles ( ~ 2000 particles in PBST) at 25 °C for 48 h using a thermal shaker (Hangzhou Allsheng Instruments Co., Ltd., China) set at 15,000 rpm for agitation. Following conjugation, the capture antibody-conjugated hydrogel microparticles undergo five washes with PBST and are then stored at 4 °C. For MIDAS validation using porcine samples, the same procedure was followed using porcine capture antibodies at the following concentrations: 6 µg/µL for IL-1β, 2.4 µg/µL for IL-6, 12 µg/µL for IL-10, and 24 µg/µL for TNF-α).
For DNA capture probe conjugation, a solution containing 5 µL of 1 mM DNA probe diluted with 5 µL of 0.5 mM TCEP solution in PBST and TET, prepared by adding 0.05 % v/v of Tween 20 in 1X TE buffer, is used at room temperature. This solution is incubated for 1 h to reduce any disulfide bonds present in the thiol-modified DNA. Following reduction, 20 µL of the reduced DNA probe is added to 140 µL of particles ( ~ 2,000 particles in PBST) and allowed to react at 37 °C and 1500 rpm for 72 h. The hydrogel microparticles conjugated with the capture DNA probe undergo five washes with TET and are then stored at 4 °C.
MIDAS assay for protein profiling
The sandwich colorimetric assay was conducted in a 96-well filter plate using shape-encoded hydrogel microparticles (Supplementary Fig. 11). Supplementary Table 11 lists the antibodies used in this study. The assay uses 100 hydrogel particles per well (or test) suspended in PBST (PBS with 0.05% Tween 20) with 1% BSA as the blocking buffer. Each well receives 40 µL of human plasma sample. The target antigen was detected by incubating the particles at 1500 rpm and 25 °C for 2 h on a shaker, followed by three PBST rinses. Next, 40 µL of fresh PBST and 10 µL of detection antibody were added to the well. Detection antibodies are used at specified concentrations (18.75 ng/µL for IL-1β, 100 ng/µL for IL-3, 12.5 ng/µL for IL-6, 12.5 ng/µL for IL-10, 12.5 ng/µL for TNF-a). For MIDAS validation using porcine samples, the detection antibody concentrations were adjusted as follows: 3.75 ng/µL for IL-1β, 5 ng/µL for IL-6, 2.5 ng/µL for IL-10, 25 ng/µL for TNF-α. Antibody incubation was conducted at 25 °C and 1,500 rpm for 1 h, followed by three PBST rinses. StAv-AP was then applied at 25 °C and 1,500 rpm for 30 min, followed by five DIT (DI water with 0.05% Tween 20) rinses, ensuring complete removal of unattached reagents and PBS buffer. Finally, BCIP/NBT substrate solution was added to 50 µL of DIT, initiating a colorimetric reaction at 25 °C and 1,500 rpm for 7 min. The stained particles were mounted in a designated area on a glass substrate (typically a No. 1 coverslip), allowed to settle for ~1 min, and then detected using the MIDAS device.
MIDAS assay for RNA profiling
Bacteria culture and RNA extraction
All bacteria were purchased from the American Type Culture Collection (ATCC). Each species was expanded in suspension culture using the appropriate medium: S. aureus (#25923) in Staphylococcus broth (BD Biosciences); Enterococcus faecalis (#29212) in trypticase soy broth containing 5% defibrinated sheep blood (Hemostat Laboratories); E. coli (#25922) in Luria-Bertani (LB) media (BD Biosciences); Pseudomonas aeruginosa (#142), Klebsiella pneumoniae (#43816) in trypticase soy broth. For RNA extraction, cultures were pelleted at 6000 × g for 10 min, and total RNA was isolated by TRIzol lysis (Life Technologies) followed by purification with the Direct-zol RNA Microprep Kit (Zymo Research). RNA concentration and purity were verified by spectrophotometry (Nanodrop OneC, ThermoFisher). To model detection in clinically relevant samples, serial dilutions of cultured bacteria were spiked into human or porcine plasma or whole blood. For whole-blood processing, erythrocytes were lysed using DNA/RNA Shield with Proteinase K (Zymo Research) before TRIzol extraction. Subsequent RNA purification followed the same procedure used for bacterial cultures.
RT-PCR
Sequences for species-specific primers targeting 16S rRNA genes are listed in Supplementary Table 5. Complementary DNA (cDNA) was synthesized from total RNA using cDNA synthesis kit (Promega or Bioneer), incubating samples at 42 °C for 30–60 min in a programmable thermal cycler (Eppendorf Mastercycler Nexus Gradient). For asymmetric PCR, reactions were prepared using AccuPower 2X GreenStar Master Mix (Bioneer) with a 10:1 ratio of forward to reverse primer. The amplification protocol consisted of: 95 °C for 5 min (initial denaturation), 30–40 cycles of 94 °C for 15 s, 58 °C for 25 s, and 72 °C for 30 s, and 72 °C for 5 min (final extension). The number of cycles was selected based on the desired sensitivity and dynamic range.
MIDAS-DNA assay
Nucleic acid detection was performed using shape-encoded hydrogel particles suspended in TET buffer (Tris-EDTA with 0.05% Tween 20, containing 200 mM NaCl). Double-stranded PCR products were converted to single-stranded DNA by heating to 90 °C for 5 min, after which the denatured material was combined with particles for hybridization. Incubation proceeded for 60–100 min at 50 °C and 1,500 rpm. Particles were washed three times with TET (50 mM NaCl). Hybridization of the biotinylated detection probe (1 µM final concentration) was carried out for an additional 30–60 min under the same temperature and shaking conditions, followed by PBST rinses. StAv-AP labeling was performed at 37 °C for 30 min with shaking, and excess enzyme was removed by DIT washes (DI water containing 0.05% Tween 20). Colorimetric development was initiated by adding BCIP/NBT substrate to 50 µL of DIT and incubating at 42 °C for ~15 min. The stained particles were transferred to a glass substrate, allowed to settle for ~1 min, and then imaged using the MIDAS device.
Swine samples for validation studies
This swine sepsis study was approved by the University of Michigan Institutional Animal Care and Use Committee (IACUC protocol #PRO00008551). All procedures were performed in an AAALAC-accredited BSL2 facility at the Unit for Laboratory Animal Medicine (ULAM). Male and female Yorkshire mix swine (40–50 kg, 14–16 weeks old) were used. Sex was not used as an experimental variable, because all animals received the same inoculum and sampling schedule, and the study was not powered to detect sex-dependent differences. To model gram-negative sepsis, animals were inoculated with ~3.5 × 1011 CFU live E. coli (strain CFT073) suspended in 5–6 mL PBS, administered via continuous infusion (0.33 mL/min) over 15 min directly into the kidney parenchyma using a syringe pump51,52. Venous blood specimens were collected at baseline (pre-inoculation) and every 2 h post-inoculation for up to 12 h. Sodium heparin preserved whole blood was aliquoted and flash frozen in liquid nitrogen. Blood for plasma was collected in EDTA containing tubes. All specimens were stored at -80 °C until assay. Plasma samples were subsequently used for a porcine Luminex assay for cytokine quantification. For the MIDAS work, frozen whole blood and plasma samples were thawed and analyzed in a blinded fashion for the MIDAS assays and reference standard clinical methods (culture and qPCR) as applicable. For RNA analysis, ~400 µL of whole blood was centrifuged to pellet cells and processed as described above. For culture, 200–400 µL of whole blood was used. Protein detection assays were conducted using 40 µL of plasma, ELISAs were performed using 100 µL of plasma per target.
Reference assays for benchmarking
ELISA
We used the same antibodies as used in MIDAS and followed the vendor-recommended protocol (R&D Systems) for ELISA. Capture antibody was diluted to a recommended concentration in PBS and added to the Maxisorp 96-well plate (Nunc) for overnight incubation at 4 °C. After being washed three times with PBS-T, 2% BSA in PBS blocking solution was added to the plate for 2 h incubation at room temperature. Subsequently, serially diluted standard or samples were added to each well for at least 2 h incubation at room temperature. After the samples were discarded and washed three times with PBS-T, biotinylated detection antibodies (recommended concentrations, diluted in 0.1% BSA solution) were added to each well and incubated at room temperature for 2 h. Unbound antibodies were washed with PBS-T three times. High sensitivity streptavidin−HRP molecules (1:5000 diluted in 0.1% BSA solution) were added to the each well for 30 min at room temperature. After being washed out with PBS, a TMB solution was added to each well and incubated for 20 min, an equal volume of stopping solution (2 M H2SO4) was added, and the optical density was read at 450 nm.
qPCR
cDNA was synthesized from each bacterial RNA sample using the cDNA synthesis kit (AccuPower Rochetscript Cycle RT PreMix, Bioneer) under thermal cycling conditions (15 °C for 20 sec, 50 °C for 4 min, 60 °C for 20 sec) for 10 cycles, followed by 95 °C for 5 min. For quantitative real-time PCR, the cDNA derived from bacteria samples were added with the reverse transcription system (AccuPower® Master Mix, Bioneer) and specific primers (Supplementary Table 5). The reaction mixtures were then applied to the Exicycler™ 96 Real-Time Quantitative Thermal Block and thermal cycling was carried out for 40 cycles at conditions of 94 °C for 5 sec, annealing at 58 °C for 25 sec, and extension at 72 °C for 30 sec. Ct values were obtained using the Exicycler3 software (provided by the manufacturer), and relative amounts of target nucleic acid were calculated based on the Ct values.
Deep learning algorithms
Computational methods
A three-stage workflow was designed to automatically locate and classify barcoded particles in hologram images. In the first stage, the Maximally Stable Extremal Region (MSER) blob detection algorithm is used to propose particle-like blobs. 128 × 128 pixel regions centered on these blobs are then extracted from the hologram image as candidates for classification72. In the second stage, a trained classification model is used to determine which regions contain a barcoded particle. In the third stage, a separately trained code classification model predicts the barcode associated with each detected particle. Lastly, the algorithm computes the average pixel intensity inside a small area surrounding the detected particle, and this intensity is referenced against a calibration curve to determine target analyte concentration within the particle.
Particle recognition
To propose candidate regions (i.e., blobs) that may contain barcoded particles, the MSER blob detection algorithm is applied to each hologram, then the minimum bounding rectangle surrounding each blob identified by MSER is computed. A blob is some region of an image that is approximately constant over some set of properties. In the case of MSER, blobs refer to extremal regions—a connected region of an image that remains stable over a range of thresholds. Because the minimum bounding rectangles for the particles of interest are typically close to square, we reject any MSER blobs whose minimum bounding rectangles differ in width (w) and height (h) beyond a predefined threshold. Specifically, we only accept blobs whose minimum bounding rectangle satisfies the condition 0.8 <w/h < 1.2. The non-max suppression algorithm is applied to remove detected regions with significant overlap73. The remaining bounding boxes of these blobs are returned by this stage as candidates to be classified as either containing a particle or not.
The MSER algorithm has 12 hyperparameters that govern the blobs it will propose. To find a robust set of hyperparameters, a dataset of 195 hologram images containing particles was collected, and the center of each individual particle was manually marked. We then applied a Bayesian hyperparameter optimization algorithm to find a set of MSER hyperparameters that minimized the error between the ground-truth particle locations and the locations of the remaining bounding boxes proposed by MSER on each image after the non-max suppression algorithm was applied to remove overlapping bounding boxes, allowing an overlap of up to 50% of the area of one of the bounding boxes.
Following the MSER blob detection, the identified blobs are sent downstream to the particle classifier, which classifies a blob as either a barcoded particle or not a barcoded particle. Blobs classified as barcoded particles are then sent to the code classifier, which classifies the unique code identifier associated with the barcode shape (i.e., labels 1 to 5). Both classifiers are convolutional neural networks (CNNs) followed by a series of fully connected layers (Fig. 3). The convolutional layers incorporate approximate translational invariance to the workflow74, allowing particle shapes to be identified irrespective of their placement in an image. The number of fully connected layers, their size, and all other hyperparameters discussed herein were chosen via Bayesian hyperparameter optimization using OpTuna75.
The optimized NN architecture for particle classification (particle vs. non-particle) is shown in Fig. 3a. Max pooling follows each convolutional feature map image to keep important feature information in the image data while facilitating dimensionality reduction76. Batch normalization (2D) follows each convolutional layer to improve model optimization during training77. Latent data exiting the convolutional layers feeds into a fully connected layer consisting of 256 nodes, followed by two fully connected layers consisting of 192 nodes. Each of these fully connected layers uses a PReLU activation function to capture regression fit non-linearity78, a dropout layer to prevent overfitting during training79, and batch normalization (1D) to improve parameter optimization during training. The final fully connected layer of the model uses a softmax layer to predict the probability that a blob proposed by the MSER algorithm contains a single barcoded particle80.
Code Classification
The optimized NN architecture for code classification (code 1-N) is shown in Fig. 3c. Like the particle classifier, max pooling and batch normalization (2D) follows the first two convolutional layers. Latent data exiting the convolutional layers feeds into a fully connected layer of 64 nodes. Data then feeds into five fully connected nodes consisting of 576 nodes. The output layer consists of six nodes—one node for each barcode label. A softplus layer is used as the choice of activation function for the output layer, whereas PReLU is used after the rest of the layers as illustrated. Softmax is applied at the output logits to obtain probability of class labels.
Training datasets
To create ground-truth training samples for classifier training, a total of 195 lab-collected hologram images of particle distributions were manually labeled with both their correct code label and particle coordinates for each particle. A 128 × 128 pixel area surrounding each manually marked particle was extracted from each hologram and assigned a label of 1 (containing a particle) and the code label corresponding to the particle in that region. To remove potentially ambiguous code labels, regions containing more than one particle in the ground-truth labels were assigned a label of 0 (not containing a particle). Next, every hologram was split into 630 non-overlapping equally spaced 128 × 128 regions covering the entire hologram. Of these, regions that intersected a ground-truth region surrounding a particle were rejected such that the only remaining 128 × 128 regions did not intersect or contain any particles. These regions were extracted and assigned a label of 0. This process provided us with a dataset of 930 regions manually labeled as 1 and 88,357 regions automatically labeled as 0. To alleviate this class imbalance, the final dataset used to train the particle classifier contained all 930 regions labeled as 1 and a uniformly random selection of 930 regions labeled as 0 from our data generation process.
Because barcoded particles may be identified across a range of stain levels and rotated positions that were not included in our lab-collected dataset, the training data was augmented to improve classification performance across all combinations of stain level and rotation. If a batch of images is augmented, the batch is randomly rotated, horizontally flipped, and/or vertically flipped to simulate generating new samples for the model during training. For each transformation, a batch is augmented if a probability P chosen uniformly at random from the interval [0, 1) satisfies P < 0.2.
Training Strategy for the Classifiers
Training and testing datasets were formed from each classifier dataset with a 70/10/20% train/validation/test split to assess the finalized model with the chosen hyperparameters. Within the each training dataset, 5-fold cross-validation was used to choose the best hyperparameters for the two classification tasks/models based on the average cross-validation accuracy81. A fixed random seed of 100 was used globally for more reproducible random number generation and model training. A dropout rate of ~0.378% for the particle classifier and 1.36% for the code classifier was applied after each CNN’s fully connected layers. Dropout was only applied during training. Early stopping based on the validation dataset loss was using during both the Bayesian hyperparameter search with cross-validation and the final model training on the train/validation/test datasets.
Image data entering the model was normalized between 0 and 1 to improve model parameter optimization. Since image data is fed as unsigned 16-bit matrices of pixel data, incoming image data was normalized by dividing each matrix element by the maximum value possible for this type of integer (i.e., 65,535). Both CNN models were trained with the Adam optimizer82 and L2 regularization83, the hyperparameters of which were guided by Bayesian optimization.
Training, validation, and test datasets for each classifier learning task are stratified to enforce consistent proportion of labels between datasets during model training and assessment. Samples are first stratified by label and then by stain level as represented by the light-normalized average pixel intensity of barcode particle centers within a 32 × 32 pixel window. The distribution of average pixel intensities is binned, and a unique integer is assigned to each bin to generate a stain label for each sample.
Hyperparameters between the classifiers are chosen via Bayesian optimization using OpTuna. We optimize learning rate, batch size, fully connected layer size, number of fully connected layers, dropout rate per fully connected layer, and L2 regularization coefficient. The target function guiding optimization for each classifier is the average 5-fold cross-validation accuracy. The Tree Parzen Estimator (TPE) was the sampling algorithm to guide the optimization84. Each Bayesian trial requires training the model five times to obtain an accuracy measure for the target function. Because of this computational cost, a pruning algorithm was used to end unpromising trials early. The Hyperband pruning algorithm was chosen because of its demonstrated benchmark performance with the TPE algorithm85.
Color/Concentration Readout
The average pixel intensity of stained particles is proportional to the concentration of target biomarkers in the samples through a series of calibration curves (protein targets shown in Fig. 4). After a blob has been classified as containing a particle, we compute the average pixel intensity inside a 20 × 20 pixel area surrounding the center of the blob. Each pixel intensity is mapped back to the calibration curve to determine the protein concentration for each particle.
Statistics & Reproducibility
Statistical analyses were performed using GraphPad Prism version 10 (GraphPad Software Inc.). For comparisons between two groups, two-sided nonparametric t-tests were used. For correlation analyses, Pearson’s correlation coefficients (r) were calculated using two-sided tests, and linear least-squares fitting was performed with 95% confidence intervals. A p value < 0.05 was considered statistically significant. Details on data presentation and sample sizes are provided in the figure legends. All experiments were performed in triplicate unless otherwise noted, and 5–10 hydrogel particles were analyzed per replicate as technical measurements. In the study of swine samples, no statistical method was used to predetermine sample size, and experiments were not randomized. No data were excluded from analyses. Investigators performing MIDAS assays on swine samples were blinded to reference diagnostic results.
Ethical Statement
All research complied with relevant ethical regulations. The swine sepsis study was approved by the University of Michigan Institutional Animal Care and Use Committee (IACUC protocol #PRO00008551) and conducted in an AAALAC-accredited facility at the Unit for Laboratory Animal Medicine (ULAM). No human subjects were involved in this work.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data underlying all figures are provided with the paper. The deep-learning training datasets, processed inference outputs, model checkpoints, and associated intermediate data generated in this study have been deposited in Zenodo under the accession code 10.5281/zenodo.1762268286. Raw swine sample data include procedural metadata with institutional identifiers and cannot be made publicly available due to University of Michigan animal research policies; de-identified swine data are available from the corresponding author for research purposes only. Requests will require a data-use agreement and will be processed within ~2–4 weeks. Source data are provided with this paper.
Code availability
All code used in this study is publicly available on GitHub at (https://github.com/Cgruich/AI4SepsisMultiplexAssay.git). A corresponding Zenodo archive is available under https://doi.org/10.5281/zenodo.1761953887.
References
Singer, M. et al. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA 315, 801–810 (2016).
Lagu T., Rothberg M. B., Shieh M. S. Hospitalizations, costs, and outcomes of severe sepsis in the United States 2003 to 2007. Crit. Care Med. 40, 754 (2012).
Center for disease control and prevention: Sepsis clinical information. Available from https://www.cdc.gov/sepsis/clinicaltools/index.html. (2025, accessed 8 Dec 2025).
Rhee, C. et al. Prevalence, underlying causes, and preventability of sepsis-associated mortality in us acute care hospitals. JAMA Netw. Open 2, e187571 (2019).
Liu, V. et al. Hospital deaths in patients with sepsis from 2 independent cohorts. JAMA 312, 90–92 (2014).
Mira, J. C. et al. Sepsis pathophysiology, chronic critical illness, and persistent inflammation-immunosuppression and catabolism syndrome. Crit. Care Med 45, 253–262 (2017).
Iwashyna, T. J., Ely, E. W., Smith, D. M. & Langa, K. M. Long-term cognitive impairment and functional disability among survivors of severe sepsis. JAMA 304, 1787–1794 (2010).
Prescott, H. C., Langa, K. M. & Iwashyna, T. J. Readmission diagnoses after hospitalization for severe sepsis and other acute medical conditions. JAMA 313, 1055–1057 (2015).
Seymour, C. W. et al. Time to treatment and mortality during mandated emergency care for sepsis. N. Engl. J. Med 376, 2235–2244 (2017).
Evans, L. et al. Surviving sepsis campaign: International guidelines for management of sepsis and septic shock 2021. Crit. Care Med 49, e1063–e1143 (2021).
Williams, J. M. et al. Systemic inflammatory response syndrome, quick sequential organ function assessment, and organ dysfunction: Insights from a prospective database of ed patients with infection. Chest 151, 586–596 (2017).
Haydar, S., Spanier, M., Weems, P., Wood, S. & Strout, T. Comparison of qsofa score and sirs criteria as screening mechanisms for emergency department sepsis. Am. J. Emerg. Med 35, 1730–1733 (2017).
Reddy, B. et al. Point-of-care sensors for the management of sepsis. Nat. Biomed. Eng. 2, 640–648 (2018).
Chanderraj, R. & Dickson, R. P. Rethinking pneumonia: a paradigm shift with practical utility. Proc. Natl. Acad. Sci. 115, 13148–13150 (2018).
Pierrakos, C., Velissaris, D., Bisdorff, M., Marshall, J. C. & Vincent, J. L. Biomarkers of sepsis: Time for a reappraisal. Crit. Care 24, 287 (2020).
Schuetz, P. et al. Effect of procalcitonin-guided antibiotic treatment on mortality in acute respiratory infections: a patient level meta-analysis. Lancet Infect. Dis. 18, 95–107 (2018).
Murphy, C. N. et al. Multicenter evaluation of the BioFire filmarray pneumonia/pneumonia plus panel for detection and quantification of agents of lower respiratory tract infection. J. Clin. Microbiol 58, e00128-20 (2020).
Wilson, J. G. et al. Cytokine profile in plasma of severe covid-19 does not differ from ARDS and sepsis. JCI Insight 5, e140289 (2020).
Hassan, F. et al. Evaluation of bacterioscan 216dx in comparison to urinalysis as a screening tool for diagnosis of urinary tract infections in children. J. Clin. Microbiol 57, e00571-19 (2019).
Ismail, F., Mackay, W. G., Kerry, A., Staines, H. & Rooney, K. D. The accuracy and timeliness of a point of care lactate measurement in patients with sepsis. Scand. J. Trauma Resusc. Emerg. Med 23, 68 (2015).
Amundsen, E. K. et al. Prognostic value of nucleated RBCs for patients with suspected sepsis in the emergency department: A single-center prospective cohort study. Crit. Care Explor 3, e0490 (2021).
Horing, S., Massarani, A. S., Loffler, B. & Rodel, J. Rapid antibiotic susceptibility testing in blood culture diagnostics performed by direct inoculation using the Vitek(r)-2 and bd phoenix platforms. Eur. J. Clin. Microbiol Infect. Dis. 38, 471–478 (2019).
Min, J. et al. Computational optics enables breast cancer profiling in point-of-care settings. ACS Nano 12, 9081–9090 (2018).
Im, H. et al. Design and clinical validation of a point-of-care device for the diagnosis of lymphoma via contrast-enhanced microholography and machine learning. Nat. Biomed. Eng. 2, 666–674 (2018).
Benzerdjeb, N., Garbar, C., Camparo, P. & Sevestre, H. Digital holographic microscopy as screening tool for cervical cancer preliminary study. Cancer Cytopathol. 124, 573–580 (2016).
Im, H. et al. Digital diffraction analysis enables low-cost molecular diagnostics on a smartphone. Proc. Natl. Acad. Sci. 112, 5613–5618 (2015).
Merola, F. et al. Tomographic flow cytometry by digital holography. Light-Sci. Appl. 6, e16241 (2017).
Veli, M. & Ozcan, A. Computational sensing of staphylococcus aureus on contact lenses using 3d imaging of curved surfaces and machine learning. ACS Nano 12, 2554–2559 (2018).
Zhu, H. Y., Isikman, S. O., Mudanyali, O., Greenbaum, A. & Ozcan, A. Optical imaging techniques for point-of-care diagnostics (vol 13, pg 51, 2013). Lab Chip 13, 4890–4890 (2013).
Greenbaum, A. et al. Wide-field computational imaging of pathology slides using lens-free on-chip microscopy. Sci. Transl. Med 6, 267ra175 (2014).
Zhang, Y. et al. 3d imaging of optically cleared tissue using a simplified clarity method and on-chip microscopy. Sci. Adv. 3, e1700553 (2017).
Roh, Y. H. et al. Precipitation-based colorimetric multiplex immunoassay in hydrogel particles. Lab Chip 20, 2841–2850 (2020).
Lim, Y. J., Choi, J. H., Mun, S. J., Kim, J. & Bong, K. W. Real-time signal analysis with wider dynamic range and enhanced sensitivity in multiplex colorimetric immunoassays using encoded hydrogel microparticles. Anal. Chem. 96, 7204–7211 (2024).
Kim, J. Y., Mun, S. J., Roh, Y. H. & Bong, K. W. Rapid colorimetric analysis of multiple microRNAs using encoded hydrogel microparticles. Analyst 146, 5508–5516 (2021).
Lee, H., Lee, J., Lee, S. G. & Doyle, P. S. Hydrogel-based colorimetric assay for multiplexed microRNA detection in a microfluidic device. Anal. Chem. 92, 5750–5755 (2020).
Juthani, N. & Doyle, P. S. A platform for multiplexed colorimetric microRNA detection using shape-encoded hydrogel particles. Analyst 145, 5134–5140 (2020).
Komori, A. et al. Characteristics and outcomes of bacteremia among ICU-admitted patients with severe sepsis. Sci. Rep. 10, 2983 (2020).
Min, J. et al. Integrated biosensor for rapid and point-of-care sepsis diagnosis. ACS Nano 12, 3378–3384 (2018).
Weber, G. F. et al. Interleukin-3 amplifies acute inflammation and is a potential therapeutic target in sepsis. Science 347, 1260–1265 (2015).
Wacker, C., Prkno, A., Brunkhorst, F. M. & Schlattmann, P. Procalcitonin as a diagnostic marker for sepsis: A systematic review and meta-analysis. Lancet Infect. Dis. 13, 426–435 (2013).
Pan, B. et al. Cith3: A reliable blood biomarker for diagnosis and treatment of endotoxic shock. Sci. Rep. 7, 8972 (2017).
Correa, T. D. et al. Angiotensin ii in septic shock: Effects on tissue perfusion, organ function, and mitochondrial respiration in a porcine model of fecal peritonitis. Crit. Care Med 42, e550–e559 (2014).
Szederjesi, J., Almasy, E., Lazar, A., Hutanu, A. & Georgescu, A. The role of angiopoietine-2 in the diagnosis and prognosis of sepsis. J. Crit. Care Med. (Targu Mures) 1, 18–23 (2015).
Chousterman, B. G., Swirski, F. K. & Weber, G. F. Cytokine storm and sepsis disease pathogenesis. Semin Immunopathol. 39, 517–528 (2017).
Min, J. et al. Cytopan-portable cellular analyses for rapid point-of-care cancer diagnosis. Sci. Transl. Med. 12, eaaz9746 (2020).
Choi, J. H., Jang, W., Lim, Y. J., Mun, S. J. & Bong, K. W. Highly flexible deep-learning-based automatic analysis for graphically encoded hydrogel microparticles. Acs Sensors. 8, 3158–3166 (2023).
Dendukuri, D., Pregibon, D. C., Collins, J., Hatton, T. A. & Doyle, P. S. Continuous-flow lithography for high-throughput microparticle synthesis. Nat. Mater. 5, 365–369 (2006).
Dendukuri, D., Gu, S. S., Pregibon, D. C., Hatton, T. A. & Doyle, P. S. Stop-flow lithography in a microfluidic device. Lab Chip 7, 818–828 (2007).
Djoba et al. An evaluation of commercial fluorescent bead-based Luminex cytokine assays. PloS one 3, e2535 (2008).
Chung, H. J., Castro, C. M., Im, H., Lee, H. & Weissleder, R. A magneto-DNA nanoparticle system for rapid detection and phenotyping of bacteria. Nat. Nanotechnol. 8, 369–375 (2013).
Tiba, M. H. et al. A comprehensive assessment of multi-system responses to a renal inoculation of uropathogenic E.coli in swine. PLoS One 15, e0243577 (2020).
Tiba, M. H. et al. A novel swine model of the acute respiratory distress syndrome using clinically relevant injury exposures. Physiol. Rep. 9, e14871 (2021).
Appleyard, D. C., Chapin, S. C. & Doyle, P. S. Multiplexed protein quantification with barcoded hydrogel microparticles. Anal. Chem. 83, 193–199 (2011).
Kim, J. et al. Hydrogel-based hybridization chain reaction (hcr) for detection of urinary exosomal miRNAs as a diagnostic tool of prostate cancer. Biosens. Bioelectron. 192, 113504 (2021).
Lee, H., Kim, J., Kim, H., Kim, J. & Kwon, S. Colour-barcoded magnetic microparticles for multiplexed bioassays. Nat. Mater. 9, 745–749 (2010).
Le Goff, G. C., Srinivas, R. L., Hill, W. A. & Doyle, P. S. Hydrogel microparticles for biosensing. Eur. Polym. J. 72, 386–412 (2015).
Roh, Y. H., Lee, H. J. & Bong, K. W. Microfluidic fabrication of encoded hydrogel microparticles for application in multiplex immunoassay. Biochip J. 13, 64–81 (2019).
Song, C. W., Bae, S. H., Bong, K. W. & Han, C. S. Label-free multiplex detection of miRNA-assayed hydrogel barcode using a low-aspect-ratio micropore sensor. Sens. Actuators B-Chem. 380, 133376 (2023).
Chapin, S. C., Appleyard, D. C., Pregibon, D. C. & Doyle, P. S. Rapid microrna profiling on encoded gel microparticles. Angew. Chem. Int Ed. 50, 2289–2293 (2011).
Yagupsky, P. & Nolte, F. S. Quantitative aspects of septicemia. Clin. Microbiol. Rev. 3, 269–279 (1990).
Lamy, B., Dargère, S., Arendrup, M. C., Parienti, J. J. & Tattevin, P. How to optimize the use of blood cultures for the diagnosis of bloodstream infections? A state-of-the art. Front Microbiol 7, 697 (2016).
Hemmig, E., Temiz, Y., Gökçe, O., Lovchik, R. D. & Delamarche, E. Transposing lateral flow immunoassays to capillary-driven microfluidics using self-coalescence modules and capillary-assembled receptor carriers. Anal. Chem. 92, 940–946 (2020).
Sun, A. T. et al. An integrated microfluidic platform for nucleic acid testing. Microsyst. Nanoeng. 10, 66 (2024).
Najjar, D. et al. A lab-on-a-chip for the concurrent electrochemical detection of sars-cov-2 rna and anti-sars-cov-2 antibodies in saliva and plasma. Nat. Biomed. Eng. 6, 968–978 (2022).
Yeh, E. C. et al. Self-powered integrated microfluidic point-of-care low-cost enabling (simple) chip. Sci. Adv. 3, e1501645 (2017).
Jang, W., Kim, J., Mun, S. J., Kim, S. M. & Bong, K. W. Highly magnetized encoded hydrogel microparticles with enhanced rinsing capabilities for efficient microRNA detection. Biomedicines 9, 848 (2021).
Choi, J. H. et al. Tyramide signal amplification for a highly sensitive multiplex immunoassay based on encoded hydrogel microparticles. Analyst 150, 2118–2127 (2025).
Mun, S. J. et al. Multiplex genotyping of SNPs in genomic DNA via hydrogel-based assay mediated with muts and polyethylene glycol. Biosens. Bioelectron. 241, 115670 (2023).
Liu, T. R. et al. Rapid and stain-free quantification of viral plaque via lens-free holography and deep learning. Nat. Biomed. Eng. 7, 1040–1052 (2023).
Lee, H. J., Kim, J. Y., Roh, Y. H., Kim, S. M. & Bong, K. W. Linker-free antibody conjugation for sensitive hydrogel microparticle-based multiplex immunoassay. Analyst 144, 6712–6720 (2019).
Moon, H. J. et al. Elimination of unreacted acrylate double bonds in the polymer networks of microparticles synthesized via flow lithography. Langmuir 36, 2271–2277 (2020).
Matas, J., Chum, O., Urban, M. & Pajdla, T. Robust wide-baseline stereo from maximally stable extremal regions. Image Vis. Comput. 22, 761–767 (2004).
Felzenszwalb, P. F., Girshick, R. B., McAllester, D. & Ramanan, D. Object detection with discriminatively trained part-based models. Ieee Trans. Pattern Anal. Mach. Intell. 32, 1627–1645 (2010).
Fukushima, K. Neocognitron - a self-organizing neural network model for a mechanism of pattern-recognition unaffected by shift in position. Biol. Cybern. 36, 193–202 (1980).
Akiba T., Sano S., Yanase T., Ohta T., Koyama M. Optuna: A next-generation hyperparameter optimization framework. Kdd'19: Proceedings of the 25th Acm Sigkdd International Conferencce on Knowledge Discovery and Data Mining. 2623–2631 (2019).
Ciresan D., Meier U., Schmidhuber J. Multi-column deep neural networks for image classification. Ieee Conference on Computer Vision and Pattern Recognition (Cvpr). 2012, 3642–3649 (2012).
Ioffe, S. & Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. Int. Conf. Mach. Learn. 37, 448–456 (2015).
He, K. M., Zhang, X. Y., Ren, S. Q. & Sun, J. Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. IEEE Int. Conf. Comput Vis. (Iccv) 2015, 1026–1034 (2015).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Bridle, J. editor. Training Stochastic Model Recognition Algorithms As Networks Can Lead To Maximum Mutual Information Estimation Of Parameters. Advances in Neural Information Processing Systems (NIPS), (1989).
Raschka, S. Model evaluation, model selection, and algorithm selection in machine learning. Preprint at https://arxiv.org/abs/1811.12808. (2018).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. Proceedings of the 3rd International Conference on Learning Representations (ICLR, 2015).
Ng A. Y., editor. Feature Selection, L1 Vs. L2 Regularization, And Rotational Invariance. Proceedings of the 21st International Conference on Machine Learning; (2004).
Bergstra, J. S., Bardenet, R., Bengio, Y. & Kégl, B. Algorithms For Hyperparameter Optimization. Advances in Neural Information Processing Systems (NIPS, 2011).
Li, L. S., Jamieson, K., DeSalvo, G., Rostamizadeh, A. & Talwalkar, A. Hyperband: a novel bandit-based approach to hyperparameter optimization. J. Mach. Learn. Res. 18, 1–52 (2018).
Lim, Y. et al. Min J. Midas—rapid, multiplexed molecular profiling for integrated host–pathogen analysis (deep learning data/models/results): Zenodo; https://doi.org/10.5281/zenodo.17619538 (2025).
Gruich CA, M.; Goldsmith, B. Midas—rapid, multiplexed molecular profiling for integrated host–pathogen analysis (version 1.0.3): Zenodo; https://doi.org/10.5281/zenodo.17619538 (2025).
Acknowledgements
We would like to thank Thomas Flott for coordinating swine sample collection. We also acknowledge the following grant support: V Scholar Award (V2023-002) funded by the V Foundation to J.M., Lefkofsky Scholar Award funded by Lefkofsky Family Foundation to J.M., NIH R35GM157070 supporting J.M.’s effort in part, the Technology Innovation Program (20018111) funded by the Ministry of Trade, Industry and Energy (MOTIE, Korea) to K.B., and the Engineering Research Center of Excellence Program through the National Research Foundation of Korea (NRF) Grant funded by the Korea Government (MSIT) (RS-2025-02263336, RS-2023-00207833, RS-2025-16066885) to K.B. The swine studies were and KAS’s effort is, supported, in part, by a grant from the NIH R35GM136312.
Author information
Authors and Affiliations
Contributions
J.M. and K.B. conceived the study. Y.L., M.A.T., J.C., C.K., and N.F. performed the experiments. M.A., C.G., and B.G. developed the DL algorithms. K.A.S. provided the swine samples. Y.L., M.A., C.G., B.G., K.A.S., R.P.D., K.B., and J.M. analyzed the data. Y.L., M.A., C.G., B.G., R.P.D., K.B., and J.M. wrote the manuscript, which was edited by all authors.
Corresponding authors
Ethics declarations
Competing interests
J.M., K.B., B.G., and Y.L. are inventors on a provisional patent application related to this work, which has been assigned to and is being managed by the University of Michigan. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks James McLaren, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lim, Y.J., Asadi Tokmedash, M., Allen, M. et al. MIDAS: rapid, multiplexed molecular profiling for integrated host–pathogen analysis. Nat Commun 17, 349 (2026). https://doi.org/10.1038/s41467-025-67391-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67391-3