Abstract
Ethylene coordinates numerous plant growth processes, particularly in cucurbit crops, yet its role in vegetative growth regulation remains largely unexplored. Here, we report the function of ethylene in controlling branch and tendril development in cucumber. We find that ethylene promotes branches formation but inhibits tendrils development in a dose-dependent manner. CRISPR-Cas9-generated gene-edited Csein2 and Csein3/Cseil1 mutants exhibit few branches and more tendrils. Exogenous ethylene can recover the branch/tendril defective phenotypes of the Csein3 and Cseil1 mutants but not those of the Csein2 mutant or the Csein3/Cseil1 double mutant. Transcriptomic and metabolic analyses reveal that CsCYP707A4 and CsTL are the key downstream targets of ethylene signaling. We show that CsEIN3 can bind to its promoters to activate the expression of CsCYP707A4 but inhibit the expression of CsTL, which leads to the opposite effect on branch and tendril development. The study sets the foundation for designing ideal plant architecture to increase production efficiency.
Similar content being viewed by others
Introduction
Ethylene is an important plant hormone that coordinates numerous plant growth and developmental processes, including seed germination, seedling growth, sex determination, fruit ripening, leaf and flower senescence, and the response to biotic and abiotic stress1,2,3,4,5,6,7. By examining ethylene-induced triple response and other typical ethylene responses in seedlings, researchers discovered numerous ethylene signaling mutants and elucidated the mechanisms underlying ethylene signal transduction8,9,10. Ethylene plays a crucial role in controlling sex differentiation in cucurbits and other crops. Ethylene synthesis-related genes ACS1, ACS7, ACS11, and ACO2 have been identified as key regulators for determining male or female flower development1,2,3,11. Similarly, the ethylene signaling-related genes CsEIN3/CsEIL1 and CsEIN2 have been revealed in the regulation of sex differentiation6,7,12,13. How ethylene regulates vegetative growth, such as branch development, remains unclear, despite its critical role in the determination of plant architecture, which consequently affects yield, product quality, and suitability for commercial production, has been investigated14,15,16.
Plant branches are key determinants of yield for many crops17,18,19. The branching structure directly determines the quantity of flowers and seeds in rice (Oryza sativa) and maize (Zea mays)20,21,22,23. Plants with more branches are considered to have ideal structures for field cultivation in the pickling industry17,24. However, redundant branches severely block sunlight and airflow, reduce yield and fruit quality, and increase the risk of disease and pest infestation25. To increase the yield of horticultural crops in greenhouse production, a two-stem or three-stem plant is preferred. Moreover, since high-density cultivation has become increasingly popular, plants with fewer branches or even non-branching cultivars are preferred. Tendrils are considered modified branches in Cucurbit crops such as cucumber. They are essential for climbing and wrapping around stick supports. Therefore, elucidating the regulatory mechanisms underlying branch and tendril development is crucial for increasing cucurbits' yields and reducing cultivation costs, particularly given the challenges associated with the increasing global population and environmental changes26.
Branch development is determined by the formation of axillary buds, which originate from axillary meristems. Accumulating evidence has shown that this process is influenced by environmental and hormonal signals17,22,27. Auxin (IAA) and strigolactone (SL) suppress branching, whereas cytokinin induces branch development19,23,28,29,30. Previous studies on branch development in Arabidopsis, rice, maize, and tomato have shown the cross-talk between plant hormones. For example, auxin can induce strigolactone activity but inhibit cytokinin activity during branch formation29. Gibberellin has been reported to inhibit branch formation in Arabidopsis, rice, and pea by functioning together with SL15,31. Further, brassinosteroids (BR) promote branch development in tomato, a process that is negatively regulated by gibberellin18. Abscisic acid can inhibit the growth of axillary buds in pea32, Arabidopsis13, tomato33, and cucumber34. Existing reports have shown the complex cross-talk of ethylene with auxin, cytokinin, and jasmonic acid35,36,37,38,39. But whether the interaction between ethylene and the other phytohormones functions in the regulation of cucumber branch and tendril development remains largely unexplored.
Recent reports showed that auxin and abscisic acid play key roles in controlling branch development in cucumber34,40. The expression of the auxin efflux carrier encoding gene CsPIN3 is repressed by CsBRC1, thereby increasing the auxin content in axillary buds and inhibiting branch formation40. The abscisic acid catabolism-related gene CsCYP707A4 has been demonstrated to regulate branch formation34. Alternatively, modified branch tendrils were reported to be regulated by gibberellin in cucurbit crops41. Tendril formation is controlled by three key genes, namely, CsTL, CsTEN, and CsUFO42,43,44. Therefore, different plant hormones and genes might be involved in the development of branches and tendrils. Identification of these factors could provide targets for breeding of cultivars suitable for high-efficiency production.
Here, we show that ethylene plays a key role in increasing the number of branches but decreasing the number of tendrils in cucumber in a dose-dependent manner. We reveal that the ethylene signaling components CsEIN2 and CsEIN3/CsEIL1 can regulate the expression of CsCYP707A4 and CsTL to promote branch development and inhibit tendril development, respectively. These results set the foundation for the development of cucumber cultivars suitable for highly efficient production.
Results
Ethylene oppositely regulates cucumber branch and tendril development
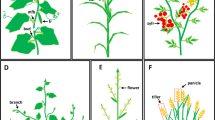
Exogenous ethylene was applied to cucumber plants 30 days after germination (DAG). A significant increasing of branch number was observed (Fig. 1a). Compared with the control, ethylene mainly promoted branching at nodes 2-6, with significant differences observed at nodes 2-4, where the number of branches increased by 1.6-fold (Fig. 1b). Intriguingly, tendril development was inhibited after ethylene treatment, with both the number and total length of tendrils at nodes 2–6 reduction of 32% and 26%, respectively, compared with those in the control (Fig. 1c, d). These results confirmed that the role of ethylene in inducing branching while delaying tendril development occurs throughout most time of the cucumber growth cycle, starting from the seedling stage (15 DAG) to maturity stage (30–45 DAG) (Supplementary Table 1).
a Mature plants (30 days after germination, DAG) were treated with exogenous ethylene for observation of branching and tendril development. The area in the dashed box is magnified on the right. Red arrows indicate the branching sites, while white triangles mark the positions of normally developed tendrils. Orange triangles highlight positions where tendrils are absent or exhibit delayed development. Scale bars= 5 cm. b Violin plots depicting the distribution of branch numbers at different nodes for ethylene-treated group and control group. Significant differences in the means (±SE, n = 15 independent plants), determined by one-way ANOVA followed by Tukey’s test, are indicated by asterisks (*P < 0.05, **P < 0.01). Violin plots showing the total number of tendrils (c) and total tendril length (d) at nodes 2-6 in mature plants of the ethylene-treated group and control group. Statistical significance in the means ( ± SE, n = 15 independent plants) was determined using a two-tailed Student’s t-test, and P values are provided. b–d The thick line within the plots indicates the median value, the X symbol indicates the mean value. e Phenotypes of branching and tendrils in cucumber seedlings (15 DAG) treated with different concentrations of exogenous ethylene (0.2 mM, 1 mM, and 5 mM ethephon with sterile water treatment serving as the control). Top to bottom: whole plant, branching details, tendril details. The white arrows indicate the branching sites, whereas the white triangles indicate the normal tendrils. Scale bars = 5 cm. Box plots showing the distribution of branch numbers (f) and tendril numbers (g) under different ethylene doses. The center line represents the median, the boundaries indicate the 25th and 75th percentiles. Whiskers extend to the largest and smallest values. Significant differences in the means (±SE, n = 15 independent plants) were determined using one-way ANOVA with Dunnett’s multiple comparisons test, and P values are indicated, and the X symbol indicates the mean value. (h) Correlation analysis of the average number of branches per node and the average number of tendrils per node using the two-tailed Pearson method. Source data are provided as a Source Data file.
Different ethylene dosage was also applied (refer to Methods) to validate the opposite induction pattern of tendril and branch development. The results showed that with increasing concentrations of ethylene, more branches were produced, and the number of branches increased to an average of 3.5 upon treatment with 5 mM ethylene (Fig. 1e, f). Moreover, higher concentrations of ethylene significantly reduced the number of tendrils (from 2 to 0.2) (Fig. 1e, g). Analysis of the ethylene-responsive marker gene CsERF1 revealed a significant dose-dependent expression pattern following treatment with varying concentrations of exogenous ethylene (Supplementary Fig. 1a). Pearson’s analysis revealed a significant negative correlation between the average numbers of branches and tendrils per node (Fig. 1h). Moreover, a positive correlation (r = 0.9321) was detected between the ethylene dosage and branch-to-tendril ratio (Supplementary Fig. 1b). The same treatment was conducted in different genotypes of cultivated cucumbers, including northern and southern types. The consistent results indicate that the effects of ethylene on branching and tendril development were genotype independent (Supplementary Fig. 2).
CsEIN2, CsEIN3, and CsEIL1 are involved in the regulation of branch and tendril development
To confirm the function of ethylene signaling in regulation of branch and tendril development in cucumber, we generated CRISPR/Cas9-edited lines of the core ethylene signaling genes CsEIN2, CsEIN3, and CsEIL1. Two representative T2 generation homozygous mutant lines were used for branch and tendril analysis. These include the Csein2-1/2 lines (with 4 bp and 8 bp deletions, respectively), the Csein3-1/2 lines (with 50 bp and 1 bp deletions, respectively), and the Cseil1-1/2 lines (with 4 bp and 5 bp deletions, respectively) (Fig. 2, Supplementary Fig. 3). All these mutants had frameshift mutations leading to premature termination. They displayed fewer branches and more tendrils than those of wild-type (Fig. 2). The branches at nodes 1–20 of each mutant grown in a greenhouse with natural light during the spring and autumn seasons were investigated. The results showed that the Csein2-1/2 lines exhibited few branches per node (by approximately 94% reduction) (Fig. 2c) with almost no lateral branches during the spring cultivation season (Supplementary Fig. 4), whereas the wild-type possessed an average of 0.26 lateral branches per node. Moreover, the lateral branches of the mutants Csein3 and Cseil1 decreased by approximately 34% and 40%, respectively, compared with those of the wild-type (Fig. 2c).
a Editing details of Csein2, Csein3 and Cseil1 homozygous mutants mediated by the CRISPR/Cas9 system. The target sequences and the PAM sites are marked with green lines. Red dashes represent deletions compared with the wild-type sequence. The deleted nucleotides are marked above the sequence. The red letters represent the additional sequences. The dashed boxes show the corresponding premature termination site of the protein. The green boxes on the right indicate regions consistent with the wild-type (WT) protein sequence, and the gray boxes indicate missense protein sequences resulting from frameshift mutations. b Observation of branch and tendril phenotypes of mature Csein2, Csein3, and Cseil1 mutants, the Csein3/Cseil1 double mutant, and WT plants grown in greenhouses. To better display the branches, the leaves within the dashed box were removed and are shown below, while the branches were left untouched. The arrows indicate the branches. Scale bars: 20 cm. c Box plots showing the distribution of the average branch number per node of mature Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants and WT plants. Twenty nodes were observed for each plant. (d) Tendril phenotype observation of the Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants and WT seedlings. e Box plots showing the distribution of the average number of tendrils per node of the Csein2, Csein3, and Cseil1 mutants, Csein3/Cseil1 double mutant, and WT seedlings. c and e The centerlines of the box plots represent the median values, and the boundaries indicate the 25th percentiles (lower) and the 75th percentiles (upper). Whiskers extend to the largest and smallest values, and the X mark indicates the mean value. Significant differences in the means (±SE, n = 10 independent plants) between various mutants and the WT were determined using one-way ANOVA followed by Tukey’s HSD test, with different letters denoting significant differences. P values for the mutants compared with the WT are indicated above the corresponding boxes. Source data are provided as a Source Data file.
To clarify the potential redundancy of CsEIN3 and CsEIL1 function in controlling branch development, the homozygous double mutant Csein3/Cseil1 was generated. The branch numbers of Csein3/Cseil1 were significantly lower than those of the wild-type and each of the Csein3 and Cseil1 single mutants (Fig. 2b, c). The results indicate that both CsEIL1 and CsEIN3 were involved in controlling branch development but they functioned redundantly.
Tendrils were increased in Csein2, Csein3, Cseil1 and Csein3/Cseil1 mutants compared to that of the wild-type (Fig. 2b). The first tendril appeared significantly earlier in these mutants (Fig. 2d). Statistical analysis showed that the wild-type cucumber seedlings possessed 0.25 tendrils per node on average, whereas the Csein2 and Cseil1/Csein3 mutants exhibited three times more tendrils than that of the wild-type seedings (Fig. 2e).
To further verify the function of ethylene in regulating branch and tendril development, we generated overexpression lines of the core ethylene signaling transcription factor CsEIN3 (Supplementary Fig. 5a–c). Two independent lines exhibited constitutively elevated CsEIN3 expression in the axillary region (Supplementary Fig. 5a). These lines also exhibited reduced tendril development (Supplementary Fig. 5d), which was consistent with the phenotype observed under exogenous ethylene treatment (Fig. 2). The average numbers of tendril per node were decreased by approximately 35% in 30 DAG plants. In contrast, the overexpression lines presented a pronounced increase in the number of branches (Supplementary Fig. 5f, g). Statistical analysis revealed that the average number of branches per node in the overexpression lines was nearly doubled relative to that of the wild-type (Supplementary Fig. 5h).
Taken together, these results suggest that ethylene signaling genes CsEIN2 and CsEIN3/CsEIL1 played a crucial role in promoting branch formation and inhibiting tendril development.
Transcriptome and metabolome analyses to explore targets of the ethylene pathway
To identify the target genes involved in modulating branch and tendril development downstream of the ethylene pathway, we generated transcriptome and metabolome data. Transcriptome analysis revealed that compared with the wild-type, Csein2, Csein3/Cseil1 mutants presented 1252 and 3719 differentially expressed genes (DEGs), respectively. There were 342 shared DEGs between these two groups (Csein2 vs. wild-type, Csein3/Cseil1 vs. wild-type). Additionally, compared with mock treatment, the exogenous ethylene treatment resulted in 1528 DEGs (Fig. 3a, Supplementary Data 1). There were 133 shared DEGs across all three comparison groups (Csein2 vs. wild-type, Csein3/Cseil1 vs. wild-type, and ET treatment vs. mock treatment; Fig. 3a). Furthermore, metabolomic analysis revealed that the endogenous concentrations of abscisic acid and auxin significantly increased by approximately 90% and 50%, respectively, in the Csein2 mutant and Csein3/Cseil1 double mutant. However, the endogenous abscisic acid and auxin decreased by 41% and 42%, respectively, after ethylene treatment (Fig. 3d). In addition, we didn’t observe significant difference on strigolactone and brassinosteroid accumulations between wild-type and the mutants or before and after ethylene treatment (Fig. 3d). These results demonstrate that abscisic acid and auxin could be involved in the signaling of ethylene in regulation of branch and tendril development.
a Venn diagram illustrating the number of differentially expressed genes (DEGs) in the axillary region of seedlings of the Csein2 mutant and Csein3/Cseil1 double mutant compared with the wild-type (WT) 15 days after germination (DAG); the DEGs in 15 DAG seedlings after ethylene treatment (1 mM) and control (mock) treatment; and the shared DEGs across these comparison groups. Relative expression levels of critical genes downstream of ethylene signaling that control branch (b) and tendril (c) development in the WT 24 h after ethylene treatment (1 mM), as well as in the Csein2 and Cseil1/Csein3 plants. The data are presented as the means ± SEs (n = 3 independent plants). Significant differences in means between various mutants and the wild-type (WT) were determined using one-way ANOVA followed by Dunnett’s multiple comparisons test and are indicated by asterisks (*P < 0.05, **P < 0.01). (d) Determination of the levels of hormones known to be involved in branching in the axillary region of 15 DAG WT seedlings 24 h after ethylene treatment (1 mM), as well as in the Csein2 and Cseil1/Csein3 plants. The data are presented as the means ± SEs (n = 6 biological replicates of the Csein2 mutant, Csein3/Cseil1 double mutant, and WT were used to quantify the levels of ABA and IAA, and n = 5 biological replicates of the Csein2, Csein3/Cseil1, and WT were used to quantify the levels of SL and BR). Each biological replicate consisted of pooled axillary regions from 5 independent plants. Significant differences in the means between various mutants and the WT were determined using one-way ANOVA followed by Dunnett’s multiple comparisons test, and P values are indicated above the corresponding boxes. Source data are provided as a Source Data file.
Among the 133 DEGs (Supplementary Data 1), there was one abscisic acid metabolome gene, CsCYP707A4 (CsaV3_4G034320), which was reported to control branching in cucumber34. Quantitative polymerase chain reaction (qPCR) analysis confirmed that its expression decreased by 37% in Csein2 and Csein3/Cseil1 mutants, and nearly doubled after ethylene treatment (Fig. 3b). These results indicate that CsCYP707A4 might play a key role in ethylene-regulated branch formation. However, we did not find any DEGs related to auxin biosynthesis and metabolism. In the meantime, we analyzed the expression of the previously reported branching-related genes. We found that expressions of CsGRF1, CsAGL16, and CsREV were not changed, but expressions of CsBRC1 and CsPIN3 were contrasted to ethylene-regulated branch formation (Supplementary Fig. 6).
Additionally, we observed that a CsTL was included on the list of the 133 DEGs, which was previously reported to control tendril development44. qPCR analysis confirmed that the CsTL expression was significantly increased in Csein2 and Csein3/Cseil1 mutants, and decreased by 40% following ethylene treatment (Fig. 3c). We also analyzed the other reported tendril-controlling genes, such as CsTEN and CsUFO, but did not detect any significant changes (Supplementary Fig. 6). We also evaluated the effects of exogenous ABA and IAA treatments on tendril development. Exogenous application of ABA or IAA did not lead to an increase in tendril number (Supplementary Fig. 7). These results suggest that CsTL functioned as a key regulator affecting tendril development downstream of the ethylene pathway.
Interestingly, the expression patterns of CsCYP707A4 and CsTL were opposite to each other among Csein2, Csein3, and Cseil1 mutants or after ethylene treatment (Fig. 3b, c), which implies that ethylene regulated branch and tendril development differentially.
CsEIN2 and CsEIN3/CsEIL1 induce CsCYP707A4 expression to promote branch development
To explore the possible mechanisms of the ethylene signaling genes CsEIN2, CsEIN3, and CsEIL1 in regulating branch formation, exogenous ethylene treatment was applied to Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants. We observed the branch number increasing in Csein3 and Cseil1 mutants. But the same effect was not observed in Csein2 and Csein3/Cseil1 mutants (Fig. 4a, b). Statistical analysis revealed that the branch numbers of Csein3 increased by approximately 3.3-fold after ethylene treatment, and that of Cseil1 increased by approximately 2-fold after ethylene treatment (Fig. 4b). Correspondingly, expression of CsCYP707A4 was significantly upregulated in Csein3 and Cseil1 mutants, but unchanged in Csein2 and Csein3/Cseil1 mutants (Fig. 4c). In addition, abscisic acid content was significantly decreased (Fig. 4c, d). To further clarify the relationship between CsCYP707A4 expression and ethylene-induced branch development, different concentrations of ethylene treatments (0.2 mM, 1 mM, and 5 mM ethephon) were conducted. Pearson’s statistical analysis revealed that the increased branch numbers in wild-type were correlated with the expression of CsCYP707A4, with an r value of 0.9432 (Fig. 4e). In contrast, this correlation was completely disrupted in Csein2 and Csein3/Cseil1 mutants (Fig. 4f, g). These results indicate that ethylene-induced branch development could depend on the expression of CsCYP707A4.
a Branch phenotypes of the Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants and wild-type (WT) mature plants after ethylene treatment. Images were captured 15 days post-treatment. The lower leaves were removed for clarity. The arrows indicate branch location. b Branch numbers in the above mutants and WT in the mature stage after ethylene treatment. Data are presented as means ± SE (n = 9 independent plants). Significant differences in the means were determined using a two-tailed Student’s t-test (P values are indicated). c Changes in the expression of CsCYP707A4 among the mutants and WT after 4, 24, 48 hours ethylene treatment. Data are presented as means ± SEs (n = 3 independent plants). d Changes in ABA content among the mutants and WT after 4, 24, 48 hours ethylene treatment. Data are presented as means ± SE (specific biological replicates numbers are indicated below each bar). c and d Significant differences between various mutants and the WT were determined using one-way ANOVA with Tukey’s HSD test. Exact P values are shown above bars for comparisons with significant differences (P < 0.05). Linear correlation between CsCYP707A4 expression and branch numbers per node in the WT (e), Csein2 (f), and. Csein3/Cseil1 (g) after ethylene treatment. Correlation analysis was performed using the two tailed Pearson method. h CsEIN3 binding sites identified by DAP-seq. (i) CsCYP707A4 promoter analysis showing potential CsEIN3 binding sites (p1–p4, orange triangles). Potential binding sequences are highlighted, with core elements colored; positions are indicated with the ATG-proximal base set to -1. j EMSAs validated the in vitro binding of CsEIN3 to p1 site on the CsCYP707A4 promoter. 3 independent experiments yielded identical results. Probe sequences are listed in Supplementary Data 2. k Yeast one-hybrid assays confirmed that CsEIN3 binding to p1 sites on the CsCYP707A4 promoter. Concentrations of aureobasidin A are indicated. l Dual-luciferase assays validated the transcriptional regulation of CsCYP707A4 by CsEIN3. The centerline represents the median. The statistical significance the differences between means (n = 15 biological replicates) was determined via two-tailed Student’s tests; P values are shown. Source data are provided as a Source Data file.
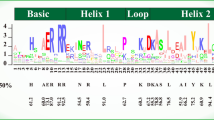
Next, we tried to illustrate the molecular mechanisms on how CsEIN3 regulates CsCYP707A4. Subcellular localization assay revealed that CsEIN3 was primarily located in the nucleus (Supplementary Fig. 8). The autoactivation assay demonstrated that CsEIN3 functions as a transcriptional activator (Supplementary Fig. 8). DAP-seq analysis was conducted using purified CsEIN3-BD protein containing its DNA-binding domain and the complying DNA library of wild-type. DAP-seq result defined that the core CsEIN3 binding motif was ‘A(G)TG(A)CATGC(A)AC(T)’. There were several potential CsEIN3 binding elements (p1-p4) in the promoter of CsCYP707A4 (Fig. 4h, i, p1). Electrophoretic mobility shift assay (EMSA) demonstrated that CsEIN3 could directly bind to the p1 element but not to the p2–p4 elements (Fig. 4j and Supplementary Fig. 9). Moreover, yeast one-hybrid (Y1H) assays verified that CsEIN3 could specifically bind to p1 (Fig. 4k and Supplementary Fig. 9). The dual-luciferase assay in a tobacco system showed that CsEIN3 could directly activate the transcription of CsCYP707A4 (Fig. 4l). In addition, expression of CsCYP707A4 was significantly upregulated in the auxiliary region of CsEIN3 overexpression lines compared to the wild-type (Supplementary Fig. 10). Therefore, CsEIN3 could bind to the promoter of CsCYP707A4 and activate its expression, thereby influencing the abscisic acid (ABA) content in the axillary region and promoting branch formation.
CsEIN2 and CsEIN3/CsEIL1 suppress CsTL expression to inhibit tendril development
After ethylene treatment, the number of tendrils was significantly reduced in Csein3 and Cseil1 mutants as well as wild-type, but no significant change occurred in the Csein2 mutant or the Csein3/Cseil1 mutant (Fig. 5a, b). Statistical analysis revealed that the tendril numbers per node of wild-type decreased by 48% after ethylene treatment, and decreased by 33% in Csein3 and 39% in Cseil1 (Fig. 5b). Expression of CsTL was downregulated after ethylene treatment in Csein3 and Cseil1 as well as wild-type, but remained at the similar level in Csein2 and Csein3/Cseil1 mutants (Fig. 5c). Moreover, different concentrations of ethylene treatments (0.2 mM, 1 mM, and 5 mM ethephon) were conducted to assess the correlation between CsTL expression and the tendril development. Pearson’s statistical analysis revealed a significant correlation between the number of tendrils and CsTL expression in wild-type (Fig. 5d). In contrast, this correlation was absent in Csein2 and Csein3/Cseil1 (Fig. 5e, f). These results suggest that the inhibitory effect of ethylene on tendril development might depend on the regulation of CsTL expression by CsEIN2 and CsEIN3/CsEIL1.
a Tendril phenotypes of the Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants and wild-type (WT) in seedling stage after ethylene treatment (1 mM). Images were captured 15 days after the ethylene treatment. The three leaves closest to the apex are designated as L1, L2, and L3. Normally developed and delayed tendrils are marked by white and orange triangles, respectively. b Average number of tendrils per node in the mutants and WT in the seedling stage after ethylene or mock treatment. The data are presented as means ± SE (the number of independent plants is indicated below each bar). A two-tailed Student’s t-test was used to determine the significance of the differences (P values are indicated). c Relative CsTL expression levels in the mutants and WT at 4, 24, and 48 hours after ethylene treatment. Data are presented as means ± SE (n = 3 independent plants). Significant differences in the means between each ethylene treatment group at different time points and the untreated group (CK) were determined using one-way ANOVA with Tukey’s HSD test. Exact P values are shown above bars for significant differences (P < 0.05). Correlations between CsTL expression and the average number of tendrils per node under different ethylene concentrations in WT (d), Csein2 (e) and Csein3/Cseil (f). Correlation analysis was performed using the two-tailed Pearson method. Scale shows the colors corresponding to different concentrations of ethephon. g CsTL promoter analysis showing potential CsEIN3 binding sites (p1), indicated by green triangles. Core elements are colored and their positions annotated. h EMSAs validated the in vitro binding of CsEIN3 to the p1 site of the CsTL promoter. Three independent experiments yielded identical results. The probe sequences used are listed in Supplementary Data 2. i Yeast one-hybrid assays validated the binding of CsEIN3 to the p1 site of the CsTL promoter. The concentrations of aureobasidin A are indicated. j Investigation of the transcriptional regulation of CsEIN3 on CsTL using dual-luciferase assay. The statistical significance of the difference between means (±SE, n = 12 biological replicates) was determined via a two-tailed Student’s test. Source data are provided as a Source Data file.
Next, we tried to determine the possible regulation mechanism of CsEIN3 on CsTL. We identified a potential CsEIN3 binding element in 1191 bp upstream of the CsTL open reading frame (Fig. 5g). The binding site “ATGCATGA” perfectly matched with the DAP-seq result of CsEIN3 (Fig. 5g). Furthermore, we confirmed the binding of CsEIN3 to the CsTL promoter through EMSA (Fig. 5h) and Y1H assays (Fig. 5i). Subsequent dual-luciferase assays in a tobacco system also clearly revealed the repression of CsTL transcription by CsEIN3 (Fig. 5j). We also examined expression of CsTL in the two CsEIN3 overexpression lines. We observed that it was significantly downregulated in the axillary region compared with that in wild-type (Supplementary Fig. 10).
Discussion
Ethylene has been reported to be involved in biological processes ranging from seed germination to plant senescence1,45,46, but less attention has been given to its regulation of vegetative growth, such as branches and tendrils development in cucumber. In this study, we report that ethylene promoted branch development but inhibited tendril development. The gene-edited Csein2, Csein3, and Cseil1 mutants and Csein3/Cseil1 double mutant had reduced the number of branches and increased the number of tendrils (Fig. 2). Overexpression of CsEIN3 resulted in more branches but fewer tendrils (Supplementary Fig. 5). Additionally, exogenous ethylene treatment complemented the phenotypes of the Csein3 and Cseil1 mutants, but it could not restore the phenotypes of the Csein2 mutant and the Csein3/Cseil1 double mutant (Figs. 1, 4, and 5). Further, ethylene differentially regulated branch and tendril development in a dose-dependent manner, and higher ethylene levels led to more branches and fewer tendrils. Further studies demonstrated a significant correlation between CsCYP707A4 expression and branch numbers, as well as the expression of CsTL and tendril numbers, which were previously reported to control branches and tendrils development34,44. Biochemical and molecular analyses showed that transcription factor CsEIN3 could directly bind to the promoters of CsCYP707A4 and CsTL. Intriguingly, CsEIN3 activated the expression of CsCYP707A4 but inhibited the expression of CsTL (Figs. 4 and 5).
Hormone crosstalk plays key roles in plant growth and development. Branch development is likely the result of the interplay of multiple hormones with different targets17,22,27,29,30. Ethylene has been reported to participate in extensive cross-talk with other hormones, including auxin, gibberellin, cytokinin, and abscisic acid4,38,39, and these hormones have been shown to be involved in branch development15,19,28,33. In this study, we discovered that abscisic acid engaged in crosstalk with ethylene to control branch development. We observed that abscisic acid levels were decreased after ethylene treatment and increased in the Csein2 and Csein3/Cseil1 mutants (Figs. 3 and 4). These results are consistent with the previous report34. We also found that the level of auxin was increased in all the mutants, but decreased after ethylene treatment (Fig. 3). Due to the fact that auxin is known to inhibit branch development in cucumber40, we deduced that the crosstalk between ethylene and auxin might play key roles in branch regulation, although whether CsCYP707A4 affected auxin metabolism remained unknown. Further investigations are needed to reveal the mechanisms by which ethylene reduces auxin in the axillary region, and the cross-talk between ethylene and auxin or abscisic acid in controlling branch development.
Previous studies have revealed that CsBRC1 and CsCYP707A4 control branch development in cucumber, while CsTEN and CsTL regulate the F-box protein CsUFO, thereby controlling tendril development40,42,43,44. In this study, we revealed that ethylene-regulating branches were correlated with the expression level of CsCYP707A4, and this was independent of the expression of the CsBRC1 gene (Supplementary Fig. 6). There is a similar case reported in Arabidopsis that branch development was independent of CsBRC147. Additionally, tendrils are considered as modified lateral branches in cucumber42,43,44. Our study revealed that ethylene-controlled tendrils were dependent on CsTL, and that CsBRC1 was not involved in tendril development, which is consistent with previous research42,43. In addition, it was observed that the known tendril-controlling gene CsUFO was reduced at 96 hours after ethylene treatment; considering the delayed response of CsUFO, it is not supposed to be a direct downstream target gene of ethylene signaling in controlling tendrils (Supplementary Fig. 11).
On the basis of our findings, we proposed a model on how ethylene regulates branch and tendril development (Supplementary Fig. 12). High level of ethylene could inhibit expression of CsTL but promote expression of CsCYP707A4, which was accompanied by reducing abscisic acid and auxin content as well as fewer tendrils but more branches (Supplementary Fig. 12). In contrast, at low level of ethylene, CsTL expression was less inhibited, and at the same time, expression of CsCYP707A4 was less activated, leading to more tendrils but fewer branches. Therefore, the formation of branches and tendrils is oppositely regulated by ethylene in a dose-dependent manner (Supplementary Fig. 12). This study also demonstrated that CsEIN2 and CsEIN3/CsEIL1 are essential for ethylene-regulated branch and tendril development in cucumber. Exogenous ethylene did not restore the phenotypes of the Csein2 or Csein3/Cseil1 mutants, but complemented the Csein3 and Cseil1 mutants, respectively (Fig. 4, Fig. 5). In Arabidopsis, EIN3 and EIL1 are important ethylene signaling transcription factors downstream of EIN2, and they function redundantly in most cases, which resulted in distinct ethylene responses in the ein3 or eil1 single mutants compared to that of the ein3/eil1double mutant or the ein2 single mutant48,49,50,51. Ethylene response we observed in Csein2, Csein3, Cseil1, and Csein3/Cseil1 mutants is consistent with that reported in Arabidopsis, suggesting the conservation of the ethylene signaling pathway in these two plant species. Our data also indicated that CsEIN3 mediated the ethylene-induced activation of CsCYP707A4 expression and the suppression of CsTL. This dual regulatory function of EIN3 is consistent with previous findings52,53,54. To further elucidate the molecular model of ethylene-based regulation of branch and tendril development, high-order mutants are needed to establish the genetic relationships between CsEIN2 and CsEIN3/CsEIL1, and to define the relationships between these signaling components with CsCYP707A4 and CsTL.
In recent years, achieving an ideal plant architecture on the basis of branch development has become important for improving crop yield and quality in order to simplify cultivation management and reduce costs. In rice, maize, and tomatoes, ideal plant architectures have been investigated for high-yield production and different agricultural cultivation methods55,56. Dwarf and robust branches are suitable for urban horticulture, whereas few or no branches are more suitable for high-density cultivation in either the field or greenhouse conditions because of their capability to avoid the reducing energy loss and blocking of sunlight. This study provided an ideal plant architecture model to breed cucumber cultivars with more branches for urban cultivation or fewer branches for high-density production (Fig. 6). In addition, more tendrils are suitable for vertical production either in the field or in the greenhouse conditions since a high-efficiency mechanized harvest process could be applied by removing the trellises (Fig. 6; simplified cultivation type). Therefore, in practical urban agriculture production, enhanced ethylene signaling could be achieved through the application of several exogenous ethylene treatments or by using mutants with constitutively enhanced ethylene signaling. In contrast, ethylene inhibitors or ethylene signaling mutants (such as Cseil1 or Csein3) could be used to appropriately reduce ethylene signaling, which is useful for high-density planting.
Urban agriculture (left): Obtained by enhancing ethylene signaling, these plants present robust lateral branches, fewer tendrils, shorter internodes, and a dwarf and compact overall plant architecture. It is expected that 30,000 plants per hectare can be planted. Traditional cultivation (middle): Unmodified wild-type plants exhibit numerous lateral branches and a certain number of tendrils, resulting in a lower planting density. In protected cultivation, it is expected that 30,000 plants per hectare can be planted, whereas in traditional ground cultivation, it is expected that 20,000 plants per hectare can be planted. Simplified cultivation (right): Obtained by appropriately reducing ethylene signaling, these plants present fewer lateral branches and a moderate number of tendrils, making them suitable for high-density planting and mechanized harvesting. It is expected that 55,000 plants per hectare can be planted. Field planting diagrams for each type of cucumber are shown below, with plant spacing and row spacing indicated in the lower right corner of each diagram.
In conclusion, this study revealed the function of ethylene in promoting branch but inhibiting tendril development in cucumber, which is dependent on CsEIN2 and CsEIN3/CsEIL1. The results set the foundation for breeding ideal plant architecture for high-yield and high-efficiency cultivation through the modulation of ethylene signaling.
Methods
Plant materials and growth conditions
North China-type cucumber (Cucumis sativus) inbred lines 1830 and 9930, and those of other genetic backgrounds, such as HZ65 and HZ170, were used to evaluate the effects of ethylene on branch and tendril development. The inbred line 1830 was also used as the genetic transformation background material, and the resulting mutants were used for subsequent studies, such as transcriptome, hormone profiling, and correlation analyses.
For the plants grown in the climate chamber, the seeds were germinated in the dark at 28 °C, and the germinated seeds were transferred to a growth chamber at 26 °C/18 °C under a 16-hour light/8-hour dark cycle, with irrigation with quantitative Hoagland nutrient solution. For the plants grown in the standard solar greenhouse, the germinated seeds were transplanted and grown during the spring and autumn, when they received regular fertilization, watering, and pest management. Both the growth chamber and the standard solar greenhouse were housed at the National Engineering Research Center for Vegetables (NERCV) in Beijing (China).
Generation of ethylene signaling gene-edited and overexpression lines
To obtain the CRISPR/Cas9 gene-edited plants CsEIN2 (CsaV3_6G039650), CsEIN3 (CsaV3_1G042150), and CsEIL1 (CsaV3_6G004610), specific sgRNA target sites were designed and selected using the CRISPR-P 2.0 website (http://cbi.hzau.edu.cn/cgi-bin/CRISPR). Using the vector pCBC-DT1T2 as a template, a PCR fragment containing 2 sgRNA target sites was generated upon amplification of 4 partially overlapping primers and insertion into the modified vector pHSE401-aada-csvU6 with the pHSE401 backbone (Addgene plasmid # 62201 provided by Qi Jun Chen) using the Bsa I site and T4 ligase57. The resulting vectors were subsequently transformed into Agrobacterium strain EHA105, and genetic transformation of cucumber was carried out using the Agrobacterium-mediated cotyledon transformation method58. The cotyledon explants whose distal portions were removed were precultured for 2 days, infected with Agrobacterium strain EHA105, and cocultured for 2 days. The explants were subsequently transferred to shoot induction medium supplemented with 6-benzylaminopurine and phosphinothricin. After shoot formation, the regenerated shoots were transferred to rooting medium containing indole-3-acetic acid to develop into T0 plants. The primers used for constructing the gene editing vectors are listed in Supplementary Data 2.
T1 generation homozygous mutants of Csein3 and Cseil1 were self-pollinated to obtain T2 generation homozygous Csein3 and Cseil1 individual mutants. Homozygous T1 generation Csein3 and Cseil1 were crossed, and T2 generation homozygous Csein3/Cseil1 double mutants were selected. Heterozygous T1 generation Csein2 was self-pollinated and selected to obtain T2 generation homozygous Csein2 mutants. These T2 and subsequent homozygous lines were then used for further analysis. All primers used for gene-edited mutant identification are listed in Supplementary Data 2.
Exogenous ethylene treatment and sample collection
Exogenous ethylene treatment experiments (ethylene treatment, ET) were conducted on wild-type, Csein2, Csein3, and Cseil1 mutant, and Csein3/Cseil1 double mutant cucumber plants, which were grown in a climate chamber. Ethylene treatment was applied 15, 30, and 45 days after germination (DAG) by spraying the entire plant with ethephon at concentrations of 0 (mock/control treatment, sprayed with sterile water), 0.2 mM, 1 mM, and 5 mM. After being sprayed, the plants were placed in a sealed container for 4 hours and then removed and grown under the same conditions with sufficient ventilation. Each seedling was treated only once, followed by subsequent sampling and phenotypic analysis.
For 15 DAG plants (seedling treatment), at 4, 24, 48, and 96 hours after ethylene treatment, samples from the axillary region (including axillary meristems and already developed small axillary buds less than 0.5 cm) were collected for transcriptome, hormone metabolome, and qPCR analyses to examine the short-term and long-term effects of ethylene. For phenotypic observation and evaluation, the branching and tendril phenotypes of the 15-day-old ethylene-treated seedlings were recorded 15 days after treatment. The posttreatment phenotypes and expression levels of various factors were subsequently subjected to correlation analysis.
For 30 and 45 DAG plants (mature plant treatment), phenotypic evaluations were conducted 15 days after ethylene treatment. These data were used to confirm that ethylene maintains consistent biological functions in both 45-day-old mature plants and 15-day seedlings.
Observation and statistical collection of branch and tendril phenotypes
For plants grown in the climate chamber (wild-type and gene-edited lines with or without ethylene treatment), phenotypic observations and statistical analyses were conducted at 15, 30, 45, and 60 DAG.
For plants grown in the solar greenhouse, phenotypic observations (photographic) and statistical analyses were conducted at 60 DAG. Data from both the spring and autumn planting seasons were calculated separately to more accurately assess the effects of genetic modifications under actual production conditions.
To accurately assess branching and tendril phenotypes in seedlings at different ages, an axillary bud greater than 2 cm was recorded as a branch, and a tendril longer than 2 cm was recorded as a mature tendril. For each gene-edited mutant or wild-type plant with or without ethylene treatment, at least 5 individual plants were examined when they were grown in climate-controlled chambers, and at least 10 individual plants were examined in solar greenhouses. The data analysis method is detailed in “Correlation analysis of the numbers of branches and tendrils and the expression levels of various factors”.
RNA-seq assay
For RNA-seq, axillary regions were collected from the second node of 15 DAG cucumber seedlings, including wild-type, Csein2 mutant, and Csein3/Cseil1 double mutant. Sampling was conducted 24 h after ethylene or mock treatment. At least three biological replicates were used for each genotype or treatment. Library preparation was followed by RNA sequencing on the DNBSEQ-T7 platform, generating 150 bp paired-end reads. The filtered clean reads were aligned to the cucumber 9930 reference genome (v3) using STAR (v2.5.3), and feature counting was performed with featureCount (v2.0.3). The fold change in expression between the treatment group (or mutant) and the control group (or wild-type) was calculated using fragments per kilobase million (FPKM). To ensure the accuracy of the results, differential gene screening was conducted on the basis of the following criteria to obtain a sufficient number of differentially expressed genes: (1) in all replicates of the comparison group, at least one replicate should have an FPKM ≥ 1; (2) for comparisons between the ethylene treatment and control groups, |fold change-1 | ≥ 0.5 and adjusted P value ≤ 0.05; (3) for comparisons between the mutants and wild-type, |fold change-1 | ≥ 0.35 and adjusted P value ≤ 0.05.
qPCR analysis of gene expression
The axillary regions of 15 DAG cucumber plants were collected, and total RNA was extracted using a Plant RNA Extraction Kit (Vazyme, RC411) and reverse transcribed into cDNA using a cDNA synthesis kit (Vazyme, R33). Quantitative reverse transcription PCR was performed with ChamQ SYBR qPCR Master Mix (Vazyme, Q311) on a Roche LightCycler 480II. The cucumber gene CsActin2 was used as an internal control. Relative mRNA expression was calculated using the 2−ΔΔCt method59. All the gene-specific primers used for qRT–PCR are listed in Supplementary Data 2.
Hormone profiling analysis
The axillary regions of 15-day-old Csein2-1, Csein3/Cseil1, and wild-type (WT) cucumber plants 24 h after 1 mM ET treatment were collected for the quantification of abscisic acid (ABA), auxin (IAA), strigolactone (SL), and brassinosteroid (BR). In addition, samples of untreated WT, Csein2-1, Csein3-1, Cseil1-1, and Csein3/Cseil1 plants and those 4, 24, 48, and 96 h after 1 mM ET treatment were subjected to ABA quantification. Each biological replicate consisted of a pool of axillary regions from 5 independent plants.
For the quantification of ABA and IAA, the procedure was as following: (1) Fifty milligrams of ground tissue was homogenized and extracted with 1 mL of methanol/water/formic acid (15:4:1, v/v/v) containing 10 μL of internal standard (100 ng/mL). The mixture was vortexed for 10 min and centrifuged at 10,000 × g for 5 min at 4 °C. The resulting supernatant was collected and evaporated to dryness. The residue was reconstituted in 100 μL of 80% methanol, passed through a 0.22 μm membrane, and analyzed by LC–MS/MS. (2) Chromatographic separation was achieved using an ExionLC™ AD UPLC system equipped with a Waters ACQUITY UPLC HSS T3 C18 column (100 mm × 2.1 mm, 1.8 μm). The mobile phase consisted of (A) water containing 0.04% acetic acid and (B) acetonitrile 0.04% acetic acid. Gradient elution was performed as follows: 5% B (0–1 min), linear increase to 95% B (1–8 min), maintain at 95% B (8–9 min), and return to 5% B (9.1–12 min). The flow rate was 0.35 mL/min, the column temperature was maintained at 40 °C, and the injection volume was 2 μL. (3) Mass spectrometric detection was performed on a QTRAP® 6500+ system equipped with an ESI Turbo Ion-Spray source operating in both positive and negative ionization modes. The key MS parameters were as follows: ion spray voltage, ±5500 V; source temperature, 550 °C; and curtain gas, 35 psi. (4) Data were acquired in scheduled multiple reaction monitoring (MRM) mode using Analyst 1.6.3 software. The declustering potential and collision energy were optimized for each MRM transition. Quantification was performed with MultiQuant 3.0.3 software.
For the quantification of SL and BR, the procedure was as follows: (1) One hundred milligrams of ground tissue was weighed into a precooled 2 mL microtube and extracted with 1 mL of ice-cold ACN: MeOH (1:1, v/v) containing internal standards. The extraction procedure included vertexing for 60 s, sonication in an ice water bath for 10 min, incubation at −40 °C for 2 h, and centrifugation at 4,000 × g for 15 min at 4 °C. A 900 μL aliquot of the supernatant was derivatized with 200 μL of 4-DMAPBA (1 mg/mL in ACN) at 75 °C for 1 h. The solution was then evaporated to dryness under a stream of N₂, reconstituted in 90 μL of 50% methanol, vortexed for 60 s, ultrasonicated for 120 s, and centrifuged twice at 10,000 × g for 10 min each at 4 °C. The final extract was filtered through a 0.22 μm membrane for UHPLC–MS/MS analysis. (2) Chromatographic separation was carried out using an ExionLC™ AD UHPLC system (SCIEX) equipped with a Kinetex C18 column (100 mm × 2.1 mm, 1.8 μm). The mobile phase consisted of (A) water containing 0.1% formic acid and (B) methanol containing 0.1% formic acid. Gradient elution was performed as follows: 55% B (0 min), 90% B (6 min), hold at 90% B (6–10 min), 55% B (10.1 min), and maintain for 12 min. The flow rate was 0.35 mL/min, the column temperature was 25 °C, and the injection volume was 2 μL. (3) Mass spectrometric detection was performed on a QTRAP® 6500+ system (SCIEX) with an ESI source. The key MS parameters were as follows: ion spray voltage, ±4500 V; ion source gas, 1/2, 50 psi; temperature, 450 °C (positive mode)/400 °C (negative mode); and curtain gas, 40/35 psi (positive/negative mode). (4) Data were acquired in scheduled MRM mode controlled by Analyst 1.7.3 software. Quantifier and qualifier transitions were optimized via flow injection analysis. Quantification was performed with MultiQuant 3.0.3 software.
Correlation analysis
Pearson’s correlation coefficients were used to assess the correlations between the branching phenotype and the expression of the CsCYP707A4 gene under different ethylene doses, as well as the correlation between the tendril phenotype and the expression of the CsTL gene. Additionally, the correlations between the tendril and branching phenotypes across different plant lines were analyzed.
A two-tailed P value of less than 0.05 was considered statistically significant, and a value of less than 0.01 was considered highly significant. Simple linear regression analysis was used to determine the strength and direction of the linear relationship between the different comparison combinations. XY scatter plots were generated using GraphPad Prism (version 10.1.2) to analyze trends between branch numbers, tendril numbers, and the expression levels of different factors during cucumber development.
Subcellular localization of CsEIN3
The full-length coding sequence of CsEIN3 and the truncated sequence (Supplementary Fig. 8) cloned from the 1830 line were inserted into the Xba I and Kpn I sites of the vector pSuper 130060, respectively. The resulting constructs were verified by sequencing and introduced into A. tumefaciens strain GV3101 through electroporation.
Agrobacterium was prepared for transient expression as follows. Bacteria containing each construct were grown in Luria–Bertani liquid medium supplemented with 0.1 mM kanamycin and 0.05 mM rifampin at 28 °C for 12 h and then pelleted by centrifugation (4000 × g for 10 min), diluted to OD600 = 1.0 in infiltration medium (sterile water supplemented with 10 mM 2-N-morpholino ethanesulfonic acid, 10 mM MgCl2, and 0.1 mM acetosyringone), and incubated in the dark at room temperature for 3 h. Bacteria with the construct and P19 were mixed at a 1:1 (v/v) ratio and infiltrated into the leaves of six-week-old N. benthamiana plants. The empty vector was used as a positive control. GFP signals with excitation at 488 nm were detected and visualized with an Olympus BX 51 fluorescence microscope (Olympus Corporation, Tokyo, Japan) 2 days after infiltration.
Transcription factor autoactivation assay
The full-length coding sequence of CsEIN3 and its various truncated fragments (R1-R3; the specific locations are shown in Supplementary Fig. 8) were amplified and inserted into the pGBKT7 plasmid (Clontech, 630489). The recombinant pGBKT7-CsEIN3, pGBKT7-CsEIN3-R1, pGBKT7-CsEIN3-R2, pGBKT7-CsEIN3-R3, and pGADT7 plasmids were subsequently transformed into Y2H Gold yeast competent cells. The samples were subsequently placed on SD/-Trp plates and incubated upside down at 28 °C for 2 to 3 days. Single colonies were selected for shaking culture, and colony PCR identification was performed. Yeast cultures identified as positive were added to 2 mL of SD/-Trp liquid media and cultured at 28 °C for 12 hours. Afterward, the cells were centrifuged at 5500 × g for 1 minute to collect the cells, the supernatant was discarded, and the cells were resuspended in 200 μL of 0.9% NaCl. The cell suspension was diluted, and 10 μL was spotted onto SD/-Leu-Trp and SD-Ade/-His/-Leu-Trp plates for analysis. Finally, the samples were incubated upside down in a 28 °C incubator for 2 to 3 days, after which growth was observed to determine the self-activation of CsEIN3.
DNA affinity purification sequencing of CsEIN3
The DNA affinity purification sequencing (DAP-seq) procedure for CsEIN3 follows the protocol outlined by Bartlett61. Portions of the CsEIN3 coding sequences that contain the DNA-binding domain (CsEIN3-BD) were cloned and inserted into the pLic-MBP vector62 through homologous recombination. The fusion protein His-CsEIN3-BD-MBP and the control His-MBP were purified using Profinity IMAC Resin (Bio-Rad, 1560135). The DNA employed for the preparation of DNA libraries was extracted from the 1830 line, which represents the wild-type. The library preparation process was executed following the manufacturer’s protocol for the NEBNext® Ultra™ II DNA Library Prep Kit for Illumina. The final enriched DNA library was sequenced on a NextSeq500 platform. The sequencing data were subjected to quality control, filtering, and trimming using Trimmomatic, followed by alignment to the 9930 genome (v3) utilizing Bowtie (v.2.2.6). Duplicate reads were identified and eliminated using SAMtools. Peak calling was performed with MACS2 (v.2.2.7.1), and the identified peaks were normalized against an empty vector control. The primers used for the CsEIN3 DAP-seq are listed in Supplementary Data 2.
EMSA
The 30–40 bp promoter fragments with potential CsEIN3 binding sites for CsCYP707A4 (p1-p4) and p1 for CsTL were synthesized with or without a biotin tag (Sangon Biotech, China). The sequences of binding probes and mutant probes are listed in Supplementary Data 2. The DNA binding domain of CsEIN3 was constructed into the pET28b vector, which included a His tag for protein purification. The recombinant vector was subsequently transformed into Escherichia coli (BL21) for protein purification. Electrophoretic mobility shift assays were then performed using a chemiluminescent EMSA kit (Beyotime Biotechnology, China) according to the manufacturer’s instructions.
Yeast one-hybrid assay
Primers were designed for the four binding sites of CsEIN3 in CsCYP707A4 promoter (p1-p4) and p1 in CsTL promoter, and to obtain PCR fragments, which were then inserted into the pAbAi vector (Clontech, 630491). The recombinant pAbAi-proCsCYP707A4 p1-p4 vector was linearized and transformed into the Y1H Gold yeast strain. The full-length coding sequence of CsEIN3 was cloned from the 1830 line and inserted into the pGADT7 vector. The pGADT7-CsEIN3 and pGADT7 empty vector were then cotransformed with pAbAi-proCsCYP707A4 p1-p4 and proCsTL p1, respectively. The resulting strains were cultured as described in the “Analysis of activation in the yeast system” and subsequently grown on plates lacking leucine with different concentrations of aureobasidin A (AbAr) to confirm their interaction.
Dual-luciferase reporter assays
The CDS of CsEIN3 was cloned and inserted into the pGreenII 62-SK vector to generate effector constructs. A 1000-bp promoter sequence (−2054 bp to −1055 bp) containing the CsEIN3 binding site p1 of CsCYP707A4 and a 1361-bp promoter sequence (−1511 bp to −151 bp) containing the CsEIN3 binding site p1 of CsTL were cloned and inserted into the pGreenII 0800-LUC vector to generate reporter constructs63. The reporter and effector constructs were subsequently transformed into Agrobacterium strain GV3101. For Agrobacterium preparation, refer to the “Subcellular localization of CsEIN3” section. Afterward, equal concentrations of Agrobacterium strains containing the reporter and the effector were mixed in specific ratios to ensure a consistent proportion of the reporter gene across different treatments. The mixed Agrobacterium was injected into the leaves of 4-week-old tobacco plants, which were then incubated for 72 hours. The tobacco leaves were ground, and homogenized samples were treated with substrates from the Dual-Luciferase Assay Kit (Vazyme, China). The LUC (firefly luciferase) and REN (Renilla luciferase) signals were measured using a microplate reader (Berthold LB942). The LUC/REN ratio was calculated to determine the transcriptional activation or repression of CsEIN3 on the target promoters.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data and CsEIN3 DAP-seq data generated in this study have been deposited in the Genome Sequence Archive (GSA) under accession CRA022797 and CRA022795, respectively. The LC-MS data for hormone quantification have been deposited in the Open Archive for Miscellaneous Data (OMIX) under accession number OMIX012503. Source data are provided with this paper.
References
Boualem, A. et al. A conserved mutation in an ethylene biosynthesis enzyme leads to andromonoecy in melons. Science 321, 836–838 (2008).
Li, Z. et al. Molecular isolation of the M gene suggests that a conserved-residue conversion induces the formation of bisexual flowers in cucumber plants. Genetics 182, 1381–1385 (2009).
Boualem, A. et al. A cucurbit androecy gene reveals how unisexual flowers develop and dioecy emerges. Science 350, 688–691 (2015).
Binder, B. M. Ethylene signaling in plants. J. Biol. Chem. 295, 7710–7725 (2020).
Zhang, H. et al. Gain-of-function of the 1-aminocyclopropane−1-carboxylate synthase gene ACS1G induces female flower development in cucumber gynoecy. Plant Cell 33, 306–321 (2021).
Rashid, D. et al. Ethylene produced in carpel primordia controls CmHB40 expression to inhibit stamen development. Nat. Plants 9, 1675–1687 (2023).
Huang, H. et al. Harbinger transposon insertion in ethylene signaling gene leads to emergence of new sexual forms in cucurbits. Nat. Commun. 15, 4877 (2024).
Guo, H. & Ecker, J. R. Plant responses to ethylene gas are mediated by SCFEBF1/EBF2-dependent proteolysis of EIN3 transcription factor. Cell 115, 667–677 (2003).
An, F. et al. Coordinated regulation of apical hook development by gibberellins and ethylene in etiolated Arabidopsis seedlings. Cell Res 22, 915–927 (2012).
Wen, X. et al. Activation of ethylene signaling is mediated by nuclear translocation of the cleaved EIN2 carboxyl terminus. Cell Res 22, 1613–1616 (2012).
Chen, H. et al. An ACC oxidase gene essential for cucumber carpel development. Mol. Plant 9, 1315–1327 (2016).
Zhang, S. et al. The control of carpel determinacy pathway leads to sex determination in cucurbits. Science 378, 543–549 (2022).
Chatfield, S. P., Stirnberg, P., Forde, B. G. & Leyser, O. The hormonal regulation of axillary bud growth in Arabidopsis. Plant J. 24, 159–169 (2000).
McSteen, P. & Leyser, O. Shoot branching. Annu. Rev. Plant Biol. 56, 353–374 (2005).
Rameau, C. et al. Multiple pathways regulate shoot branching. Front. Plant Sci. 5, 741 (2015).
Li, J., Yao, X., Lai, H., Zhang, X. & Zhong, J. The diversification of the shoot branching system: A quantitative and comparative perspective in meristem determinacy. Curr. Opin. Plant Biol. 81, 102574 (2024).
Liu, X., Chen, J. & Zhang, X. Genetic regulation of shoot architecture in cucumber. Hortic. Res. 8, 143 (2021).
Xia, X. et al. Brassinosteroid signaling integrates multiple pathways to release apical dominance in tomato. Proc. Natl. Acad. Sci. USA. 118, e2004384118 (2021).
Yang, H. et al. The tomato WRKY-B transcription factor modulates lateral branching by targeting BLIND, PIN4, and IAA15. Hortic. Res. 11, uhae193 (2024).
Li, X. Y. et al. Control of tillering in rice. Nature 422, 618–621 (2003).
Mathan, J., Bhattacharya, J. & Ranjan, A. Enhancing crop yield by optimizing plant developmental features. Development 143, 3283–3294 (2016).
Barbier, F. F., Dun, E. A., Kerr, S. C., Chabikwa, T. G. & Beveridge, C. A. An update on the signals controlling shoot branching. Trends Plant Sci. 24, 220–236 (2019).
Du, Y., Wu, B., Xing, Y. & Zhang, Z. Conservation and divergence: Regulatory networks underlying reproductive branching in rice and maize. J. Adv. Res. 41, 179–190 (2022).
Soyk, S. et al. Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat. Genet. 49, 162–168 (2017).
Howe, V. The quest to find TLC: Tendril-less cucumbers, that is. Plant Cell 36, 2751–2752 (2024).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Yan, Y., Zhao, N., Tang, H., Gong, B. & Shi, Q. Shoot branching regulation and signaling. Plant Growth Regul. 92, 131–140 (2020).
Umehara, M. et al. Inhibition of shoot branching by new terpenoid plant hormones. Nature 455, 195–200 (2008).
Domagalska, M. A. & Leyser, O. Signal integration in the control of shoot branching. Nat. Rev. Mol. Cell Biol. 12, 211–221 (2011).
Chen, Y., Fan, X., Song, W., Zhang, Y. & Xu, G. Over-expression of OsPIN2 leads to increased tiller numbers, angle and shorter plant height through suppression of OsLAZY1. Plant Biotechnol. J. 10, 139–149 (2012).
Davière, J. & Achard, P. Gibberellin signaling in plants. Development 140, 1147–1151 (2013).
Arney, S. E. & Mitchell, D. L. The effect of abscisic acid on stem elongation and correlative inhibition. N. Phytol. 68, 1001–1015 (1969).
Cline, M. G. & Oh, C. A reappraisal of the role of abscisic acid and its interaction with auxin in apical dominance. Ann. Bot. 98, 891–897 (2006).
Chen, J. et al. The AGAMOUS-LIKE 16–GENERAL REGULATORY FACTOR 1 module regulates axillary bud outgrowth via catabolism of abscisic acid in cucumber. Plant Cell 36, 2689–2708 (2024).
Lehman, A., Black, R. & Ecker, J. R. HOOKLESS1, an ethylene response gene, is required for differential cell elongation in the Arabidopsis hypocotyl. Cell 85, 183–194 (1996).
Song, S. et al. Interaction between MYC2 and ETHYLENE INSENSITIVE3 modulates antagonism between jasmonate and ethylene signaling in Arabidopsis. Plant Cell 26, 263–279 (2014).
Yan, Z. et al. Type B response regulators act as central integrators in transcriptional control of the auxin biosynthesis enzyme TAA1. Plant Physiol. 175, 1438–1454 (2017).
Zemlyanskaya, E. V., Omelyanchuk, N. A., Ubogoeva, E. V. & Mironova, V. V. Deciphering Auxin-Ethylene Crosstalk at a Systems Level. Int. J. Mol. Sci. 19, 4060 (2018).
Dolgikh, V. A., Pukhovaya, E. M. & Zemlyanskaya, E. V. Shaping Ethylene Response: The Role of EIN3/EIL1 Transcription Factors. Front. Plant Sci. 10, 1030 (2019).
Shen, J. et al. CsBRC1 inhibits axillary bud outgrowth by directly repressing the auxin efflux carrier CsPIN3 in cucumber. Proc. Natl. Acad. Sci. Usa. 116, 17105–17114 (2019).
Fan, Z. et al. Tendril length is determined by gibberellin deactivation during thigmo response in cucumber. Plant J. 120, 901–909 (2024).
Wang, S. et al. A rare SNP identified a TCP transcription factor essential for tendril development in cucumber. Mol. Plant 8, 1795–1808 (2015).
Chen, Y. et al. CsUFO is involved in the formation of flowers and tendrils in cucumber. Theor. Appl. Genet. 134, 2141–2150 (2021).
Shen, J. et al. The GRAS transcription factor CsTL regulates tendril formation in cucumber. Plant Cell 36, 2818–2833 (2024).
Iqbal, N. et al. Ethylene role in plant growth, development and senescence: interaction with other phytohormones. Front. Plant Sci. 8, 475 (2017).
Huang, W. et al. A molecular framework of ethylene-mediated fruit growth and ripening processes in tomato. Plant Cell 34, 3280–3300 (2022).
Seale, M., Bennett, T. & Leyser, O. BRC1 expression regulates bud activation potential but is not necessary or sufficient for bud growth inhibition in Arabidopsis. Development 144, 1661–1673 (2017).
Alonso, J. M., Hirayama, T., Roman, G., Nourizadeh, S. & Ecker, J. R. EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 284, 2148–2152 (1999).
Ju, C. & Chang, C. Mechanistic insights in ethylene perception and signal transduction. Plant Physiol. 169, 85–95 (2015).
Harkey, A. F. et al. Identification of transcriptional and receptor networks that control root responses to ethylene. Plant Physiol. 176, 2095–2118 (2018).
Hao, D., Li, W. & Guo, H. Ethylene signaling in Arabidopsis: a journey from historical discoveries to modern insights. Plant Hormones. 1, e014 (2025).
Deng, L. et al. Tomato MED25 regulates fruit ripening by interacting with EIN3-like transcription factors. Plant Cell 35, 1038–1057 (2023).
Xu, M. et al. ETHYLENE INSENSITIVE3/EIN3-LIKE1 modulate FLOWERING LOCUS C expression via histone demethylase interaction. Plant Physiol. 192, 2290–2300 (2023).
Mahendrawada, L., Warfield, L., Donczew, R. & Hahn, S. Low overlap of transcription factor DNA binding and regulatory targets. Nature 642, 796–804 (2025).
Carbajal-Friedrich, A. A. & Burgess, A. J. The role of the ideotype in future agricultural production. Front. Plant Physiol. 2, 1341617 (2024).
Laurans, M. et al. Why incorporate plant architecture into trait-based ecology?. Trends Ecol. Evol. 39, 524–536 (2024).
Hooghvorst, I., López-Cristoffanini, C. & Nogués, S. Efficient knockout of phytoene desaturase gene using CRISPR/Cas9 in melon. Sci. Rep. 9, 17077 (2019).
Hu, B. et al. Engineering Non-transgenic Gynoecious Cucumber Using an Improved Transformation Protocol and Optimized CRISPR/Cas9 System. Mol. Plant. 10, 1575–1578 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25, 402–408 (2001).
Su, T. et al. WRKY42 Modulates Phosphate Homeostasis through Regulating Phosphate Translocation and Acquisition in Arabidopsis. Plant Physiol. 167, 1579–1591 (2015).
Bartlett, A. et al. Mapping genome-wide transcription-factor binding sites using DAP-seq. Nat. Protoc. 12, 1659–1672 (2017).
Yu, Y. et al. The key clock component ZEITLUPE (ZTL) negatively regulates ABA signaling by degradation of CHLH in Arabidopsis. Front. Plant Sci. 13, 995907 (2022).
Hellens, R. P. et al. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1, 13 (2005).
Acknowledgements
This work was supported by the National Key Research and Development Program (2022YFD1200501 and 2023YFF1000101), the National Natural Science Foundation of China (32202517), Beijing Academy of Agricultural and Forestry Sciences (JKZX202207), Young Scholars Program in Beijing, and National High-Level Talents Special Support Program for Young Top-Notch Talents.
Author information
Authors and Affiliations
Contributions
X.Z. and C.W. designed the project; T.Z., W.Z., Y.Z., and C.X. performed the experiments; J.Z., J.Y., H.Z., Y.Y., and C.W. analyzed the data; X.Z. and C.W. wrote the paper with input from the other coauthors; and C.W. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hongwei Guo and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Zhu, T., Zhang, W. et al. Ethylene promotes branch formation but inhibits tendril development in cucumber. Nat Commun 17, 745 (2026). https://doi.org/10.1038/s41467-025-67444-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67444-7