Abstract
The worldwide rise of neurological and psychiatric conditions poses major challenges. However, current global research remains fragmented, dominated by limited cohorts and poorly integrated datasets that disconnect whole-body health, exposome, and brain health. Theories rarely unify brain measures with extracerebral factors or capture heterogeneity in individual trajectories. We introduce multimodal diversity, a non-linear, non-simplistic causal and ecological construct integrating data representation, whole-body and exposomic factors, and computational modeling to address this situated, embedded, and embodied complexity. This heuristic metamodel integrates global, multilevel data into personalized predictions fostering population inclusion, multimodal integration, diagnostic precision, and equitable, context-sensitive advances in brain health.
Similar content being viewed by others
Introduction
The global prevalence of neurological, neurodevelopmental, neurodegenerative, and psychiatric disorders has risen significantly1,2,3. Epidemiological, social, and economic estimates suggest that these diseases pose significant burdens and financial costs, impacting human health and societal welfare1,4. Together, they heighten the risk of comorbidity, chronic systemic diseases and mortality3. Brain disorders remain among the top ten leading causes of disability worldwide5,6. Neurological conditions were the most prevalent of the leading causes of disability in 2021, accounting for 443 million disability-adjusted life years (DALYs)1,5,6. The number of people living with dementia is projected to rise from 57.4 million in 2019 to 152.8 million by 2050, with the highest increase anticipated in low- and middle-income countries7. DALYs due to Alzheimer’s disease (AD) and Alzheimer’s disease and related dementias (ADRD) worldwide increased from 9.66 million to 25.28 million between 1990 and 20191. Psychiatric conditions accounted for 125.3 million DALYs4 in 2019. Between 1990 and 2019, DALYs rates for psychiatric conditions in adults increased from 803.8 to 833.2 per 100,0008. These stark estimates have spurred international calls to advance brain health through enhanced research, public health initiatives, policy development, targeted interventions, and improved treatments. The scientific fields require more diverse, complex, multilevel, accurate, global, robust, and multimodal approaches7,9,10,11,12,13.
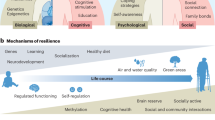
Multimodal diversity in brain health12 (Box 1, Fig. 1) refers here to a multilevel construct that combines three dimensions (Table 1): (i) data representation, encompassing heterogeneous sources ranging from omics to neuroimaging, behavioral, and clinical indicators; (ii) multilevel integration of brain and extracerebral influences, including whole-body health metrics (e.g., cardiometabolic status, immune function, gut microbiome, interoception) and environmental factors (physical and social determinants or exposome); and (iii) computational architectures, capable of handling the intrinsic multimodal complexity, heterogeneity, and probabilistic trajectories inherent to such diverse data. Unlike traditional approaches that consider single modalities or levels of analysis, multimodal diversity emphasizes synergistic modeling that integrates biological, systemic, and environmental measures to generate more accurate, and individualized predictions12,13,14,15,16. This definition transcends traditional notions of diversity (often confined to gender, ethnicity, race, or minority status) by focusing on complex model interactions that exhibit significant variation across individuals and populations due to stochastic and probabilistic dynamics (i.e., the random but structured processes driving heterogeneous outcomes across the lifespan16).
The figure illustrates the dimensions of multimodal diversity, showcasing data diversity (a) and its integration into personalized models through advanced modeling approaches (b). a depicts data diversity with the hierarchical data layers contributing to brain health determinants, spanning micro- to macro-levels between and within data types. The first level focuses on genetic and epigenetic factors, including genetic risks, mutations, DNA methylation, and social epigenomics. The second level represents multimodal brain models (MBM), encompassing brain microarchitecture, neurotransmission, structures, functions, and global networks. The third level integrates whole-body biology, combining multi-omics data such as transcriptomics, proteomics, metabolomics, and microbiomics with systemic health indicators like cardiometabolic health. The fourth level incorporates behavioral, cognitive, and clinical data (BCC). The fifth level includes exposome factors, such as social determinants (e.g., socioeconomic status, social interactions, and social identities) and physical exposures (e.g., green spaces, nanoplastics, air pollution, and heatwaves). b outlines the diversity in modeling approaches required to effectively integrate multimodal data diversity into brain health research. It emphasizes data diversity, the generation of low-dimensional brain representations, the incorporation of extracerebral influences such as whole-body health and exposome data, and the need to account for heterogeneity and individual trajectories. This framework highlights the potential of multimodal diversity to advance precision medicine by addressing complex brain health determinants and personalizing interventions. SES socioeconomic status.
Theoretically and methodologically, this concept emphasizes synergistic computational modeling and integration of multimodal data10,12,13,14,15,16 in neurology and psychiatry11,17,18,19,20,21,22,23,24. There are urgent calls to address methodological barriers, global collaboration, and improved computational models10,15,17,25,26,27,28,29,30,31. However, challenges related to underrepresentation, the reliance on single-level approaches in global settings (e.g., those limited to a single data type such as genetic variants, or neuroimaging), and the need for theoretical frameworks that integrate biological, behavioral, and environmental layers must be addressed jointly12.
This perspective begins by highlighting how the limited funding, insufficient samples representation, inadequate global data sharing, and the absence of unified theoretical frameworks undermines advancements in brain health. While our framework is focused mainly on brain health, we use AD/ADRD as an illustrative case, while also highlighting applications to psychiatric and neurodevelopmental disorders. We propose a heuristic framework that emphasizes integrating global and multimodal data and theoretical models to tackle the core challenges for computational modeling. We discuss the core ethical, social, and technical barriers of this initiative. Finally, we outline a future agenda and recommendations to enhance computational models, deep phenotyping, neuroecological links, transdisciplinary collaboration, and inclusive brain health research.
Why multimodal diversity is essential for precision brain health
Our perspective focuses on some of the main challenges of precision in brain health driven by funding constraints, methodological hurdles, and broader scientific infrastructures. A core barrier is the limited funding and consequent lack of substantial multimodal data in global settings. Current research approaches often fail to represent diverse populations and incorporate various data types32,33, leading to simplified predictions34,35 and restricted applicability11,36. By constraining the focus to single data types29,37 and excluding environmental-biology interactions, approaches can neglect important factors that impact brain health38,39,40,41.
First, limited representation of diverse populations17,18,22,23 leads to a lack of generalizability24, decreasing the effectiveness of diagnosis and treatments12,24,25. Genome-wide association studies (GWAS) and polygenic risk studies are predominantly based on European descent populations, biasing the identification of risk factors36,42,43,44,45,46 and hindering tailored prediction models and treatments29,37. Ancestry diversity affects the understanding of disease prevalence, genetic and biological mechanisms, and brain phenotypes47. Reduced population diversity limits specificity48,49,50,51, neglecting complex biological mechanisms52 and diminishing clinical insights50, risk identification53,54, cognitive evaluations53,55, and neuroimaging characterization56. These gaps have a negative impact on accurate diagnosis, biomarker development, and successful clinical trials36,49,57,58.
Multiple factors contribute to increased data variability in underrepresented populations from Latin America, Africa, Middle East and Asia. These populations often exhibit more significant genetic admixture51,59 than in the U.S. or European populations. Greater multimodal heterogeneity is also evident, particularly in whole-body and environmental data12,60. Populations in the Global South frequently experience higher burdens of cardiometabolic conditions61 and show stronger biological embedding of structural inequalities. Most predictive models are trained on large U.S. and European datasets. These problems are observed across Global South populations and minorities within countries such as the US62,63. When applied to underrepresented populations with different data distributions, these models often exhibit poor generalizability and higher misclassification rates34,56,62.
Second, most research in brain health concentrates on isolated genetic and brain data13,49,64, neglecting the influence of whole-body health and multi-omics60,65. Approaches usually focus on single-level explanations, such as genetics36,42,43,44,45,46 while overlooking biology-environment interactions14,15,64 or vice versa. Although some models (see below) emphasize how the body serves as a dynamic interface linking the environment and the brain, most existing research remains predominantly theoretical. The exposome13 refers to the totality of exposures an individual experiences throughout their lifetime encompassing physical (e.g., pollutants, heat waves) and social influences (socioeconomic status, political stress, loneliness, poverty, and cultural factors). The exposome has only recently begun to be applied to brain health9,13,58 with few examples combining it with genetics, whole-body health, and brain function10,66,67. These brain-body-environment interactions require new conceptual models9,68 for data integration69,70. How to systematically model extracerebral factors is poorly understood, limiting predictive accuracy69. Models that rely on averages frequently miss critical individual differences, leading to inaccurate predictions, especially in diverse populations34. A metamodel may enables a comprehensive understanding of brain health by integrating biology-environment interactions, and population heterogeneity15,71, informing global health policies and advancing precision medicine72,73.
Third, computational modeling often fails to integrate global data and account for heterogeneity, thereby hampering the development of tailored diagnostic tools and treatments34,35. The underrepresentation of non-White and diverse populations in global datasets compromises the generalizability, fairness, and clinical utility of artificial intelligence (AI) and machine learning models56,74,75. Models trained on demographically skewed data risk misclassification, poor prediction, and reinforcement of systemic inequities. Equitable modeling requires transparent reporting (clear documentation of data sources, methods, and limitations), interdisciplinary validation (cross-field review and replication of findings), and stakeholder engagement (involving community and policy representatives)76. Key needs include assessing data heterogeneity, incorporating multimodal inputs (i.e., systemic biology and exposomic layers9,28,77) and applying individualized probabilistic modeling to capture complex risk trajectories73. These strategies are essential for advancing truly equitable, globally relevant brain health research. A structured metamodel -a computational framework designed to handle data heterogeneity, integrate multimodal sources, and model individual trajectories and extracerebral influences-, is crucial for advancing brain health research29.
Multimodal diversity frameworks in data and models
We present the building blocks of a heuristic framework incorporating synergistic computational architectures10,11,12,13,14,15,16,53,77,78,79,80,81,82,83 and personalized models72,73,84. We first summarize emerging evidence on multimodal diversity. Then, we explore how integrating computational approaches can provide promising pathways for improved theorization and prediction.
Emerging evidence in brain health and disease
Evolving evidence and calls from the World Health Organization (WHO), the Alzheimer’s Association, and the National Institute on Aging highlight the importance of incorporating population diversity, loneliness, social determinants of health, and complex biological, neuropathological, and exposomic measures in brain health research36. Research requires addressing infrastructural, methodological, and funding barriers that hinder the development of diverse, biologically and contextually informed datasets. Despite advances from initiatives like UK Biobank85 and FINNGEN86, these datasets face challenges regarding population heterogeneity, rich ancestries, and global representation39,87,88. Bridging these gaps demands global collaboration to build inclusive, multilevel, lifespan-oriented cohorts. Key priorities include lifespan enrollment89,90,91,92 and the development of biorepositories to reduce disparities in genomic and biomarker data74,93,94,95,96.
Funding agencies supporting large-scale initiatives have started to promote increased data diversity and multimodal integration and comparisons across global datasets are now more feasible. Examples include the NIH All of Us program (with 77% of participants having underrepresented backgrounds), UK Biobank; the Innovative Medicines Initiative (IMI) – European Union; the Precision Medicine Initiative (PMI, NIH, USA); the NIH - Brain Research Through Advancing Innovative Neurotechnologies (BRAIN) Initiative (USA); the NIH - Research Domain Criteria (RDoC) Initiative (USA), the NIH ECHO program, as well as programs in the Global South10,26,27,28,29,31,62,97,98,99,100,101,102,103. Initiatives such as H3Africa104, FINGER105, BrainLat96,106 and ReDLat107 demonstrate that ethically governed, context-sensitive infrastructures are feasible in the Global South, though broader implementation is needed. These programs evidence feasible, collaborative pathways supporting large-scale inclusion across populations. Sustained progress will require coordinated international efforts, culturally grounded design, regional leadership, and structural reforms (federated data sharing, open-access platforms, and long-term ethical governance) to ensure equity and generate actionable insights for global brain health.
Genetic diversity and scientific discovery
The number of well-powered, large-scale genetic studies using GWAS and polygenic risk scores worldwide, particularly in the Global South, for psychiatric (i.e., schizophrenia108, bipolar109 or depression110), and neurological conditions (i.e., Parkinson’s disease111 and stroke112) is small but growing. Over 40 new genetic risk variants not previously reported in European populations have been identified. These findings can potentially generate discriminative models across different ancestries. Distinct linkage disequilibrium and haplotype patterns in Global South populations sharpen fine-mapping and causal inference. Polygenic risk scores (PRS) trained mainly on European datasets underscore the need for ancestry-aware discovery and validation113. Local-ancestry–aware methods such as Tractor enable robust GWAS in admixed populations, producing ancestry-specific effect estimates and recent schizophrenia GWAS across African, Latino, and East Asian cohorts confirm their value114. Large, multi-ancestry cohorts such as NeuroGAP-Psychosis in Africa and the Latin American Genomics Consortium are examples of this effort115,116. Asian GWAS have also identified new pleiotropic loci for depression and schizophrenia (e.g., 1q25.2)117, while large East Asian–European meta-analyses revealed 53 novel schizophrenia loci through trans-ancestry fine-mapping118. Rare CNV variants for major psychiatric disorders119 and specific genetic footprints associated with stroke120 have also been reported in Chinese populations.
AD/ADRD
Specific mutations not previously reported in Europe and the US, include the PSEN1 E280A mutation (the Paisa mutation), found in Medellin, Colombia, and responsible for the largest early-onset AD population in the world. The high prevalence of the PSEN1 E280A mutation in the Colombian population stems from a founder event, with ancestral origins predominantly traced to Spanish lineage121,122,123. APOE effects on AD/ADRD vary by haplotype and local ancestry in the Global South. Protective variants such as APOE ε3 Christchurch124 and RELN125, identified in LAC, renew genetics-informed trial prospects51. In African-ancestry cohorts, meta-analyses identified MPDZ and 11 additional AD-risk loci126 and showed attenuated APOE ε4 risk compared with Europeans43. APOE ε2 is generally more protective in Europeans than in other ancestries, while the ε2/ε3 genotype confers ~8-fold higher AD risk in African Americans yet reduced risk in Europeans127. These findings stress the need for diverse cohorts to expand discovery and enable ancestry-informed therapies.
Whole-body health
Whole-body health offers a systemic framework to elucidate how physiological networks across the cardiometabolic, inflammatory, and immune domains mediate the brain’s adaptation to environmental demands11,12,13,60,64,128,129,130,131,132,133. This approach allows to embed exposomic perturbations within multi-organ and multi-omics architectures to provide a mechanistic bridge linking external exposures to neurobiological trajectories of resilience and disease13,16. Multi-omics has emerged as a promising path in psychiatry65 and neurology41,65,134,135. Combining omics with clinical, neuropathological, and neuroimaging data improves greater prediction accuracy41,65,136,137. Inflammation processes have become relevant in psychiatry, neurology138, and neurodegeneration129,134,136,139. Pro-inflammatory cytokines, including mediators such as Interleukin-1β (IL-1β), IL-6, and the nucleotide-binding and oligomerization domain like receptor (NLR) family, are critical for synaptic dysfunction and allostatic load in brain disorders140. Corticosteroids and sex hormones (testosterone and estradiol) play crucial roles in modulating neurodevelopment, brain structure, and function141; and their dysfunction is associated with multiple conditions141.
Proteome-wide analyses of brain tissue and peripheral fluids have identified proteins and pathways linked to conditions such as schizophrenia and other mental disorders142. Integrating proteomic QTLs, transcriptomic QTLs, and GWAS signals enables prioritization of candidate genes beyond what genomics alone can achieve142. Proteome-wide association studies, Mendelian randomization, and imaging phenotypes have linked proteins such as EGFR and TMEM106B to both brain structure and risk for schizophrenia or neurodevelopmental disorders143. Transcriptomics–MRI integration further shows how gene expression aligns with structural and functional changes in mental illness144, while multimodal fusion uncovers hidden pathophysiological links145. Spatial multi-omics and mass spectrometry-based neuroproteomics has provided detailed maps of protein-level changes (post-translational modifications and protein–protein interactions) which are central to the pathogenesis of diseases146,147.
AD/ADRD
The AT(N) (A=amyloid-beta, T=tau, and N=neurodegeneration) framework reflects AD brain changes but most likely is insufficient for accurate diagnosis in global settings47,148,149,150. Combining this framework with multi-omics and neuroimaging can enhance predictions145,151, clinical characterization152 and the discovery of treatment targets152. RNA sequencing and proteomic screening evidence upregulation of transcription- and chromatin-related genes, including histone acetyltransferases for H3K27ac and H3K9ac153. One study154 identified 137 proteins with distinct trajectories in causative models of AD, including many that changed before traditional AD biomarkers. GWAS have identified AD-susceptible genes involved in amyloid plaque and neurofibrillary tangle formation, cholesterol metabolism, endocytosis, and the innate immune system155. AD/ADRD loci impact immune mechanisms in AD155. Among other multi-omics markers136,137, metabolomics has revealed brain insulin dysfunction. Transcriptomics reveal a significant impact of ancestry, particularly affecting immune pathways156. Postmortem brain multi-omics data and in vivo samples are very limited. Biophysical and semi-empirical modeling can help to incorporate these scarce data via prios36,157,158. Other more accessible modalities (i.e., plasma proteomics, cell transcriptomics) offer promising non-invasive proxies for brain health41,159,160 and multi-modal integration41,159,161. Brain-blood correspondence in animal models69,162 can also validate peripheral multi-omics biomarkers.
Epigenomics also evidence the impact of diversity163,164,165. Ancestral epigenetic profiles demonstrate differential DNA methylation signatures in imprinted gene regulatory zones within the inflammasome (e.g., NLRP1 and MEST/MESTIT1) across individuals with AD166. Early developmental alterations in DNA methylation may contribute to ancestry risk166. Differences in DNA methylation age acceleration (e.g., epigenetic clocks) have been linked to immune system aging and inflammation, influenced by environmental and lifestyle factors167. Organ clocks (the biological age measures and mechanisms within specific organs that regulate their function) utilize transcriptome, epigenome, and proteome to integrate the physiological rhythms and aging processes of heart, liver, kidneys, gut, and brain168. The combination of genomics, particularly polygenic risk scores, with organ clocks represents a powerful strategy for predicting mortality indices, disease pathology, resiliency, and healthy aging16,169. Whole-body health, including cardiometabolic factors are critical predictors of brain health and disorders60,61. Gut microbiome health is sensitive to exposome influences and is a crucial predictor of brain health66. Cardiovascular, metabolic, and gut microbiome are key predictors of behavior and clinical outcomes60,170.
Despite these advances, several fields of omics remain immature and underdeveloped. Whole-body modeling frameworks that integrate longitudinal multi-omics at the individual level are still in their infancy, particularly in underrepresented populations. The gut-brain axis, although promising, lacks standardized protocols and causal models in human studies. Postmortem brain multi-omics data, essential for validation, are scarce and geographically skewed, limiting global generalizability. Conversely, promising advances are emerging from the integration of epigenetic clocks, peripheral omics (e.g., plasma proteomics), and organ-specific biological age measures.
Biology–environment interactions
Understanding biology-environment interactions58,171 requires multimodal frameworks70 capturing the biological embedding of macrosocial factors (e.g., country-level inequality) on biology and brain health. The One-Health approach emphasizes the interconnectedness of human, animal, and environmental health and is critical for precision brain health13. Water and air pollution, plastics and nanoplastics, climate change, and heat waves are essential to the physical exposome9,13. Social inequality, loneliness, political polarization, and injustice are social exposomes9,48 highlighting the need to examine macro- and individual-level determinants9,13,48,54,172,173.
Environmental exposures strongly influence brain biology and increase the risk of psychiatric and neurological disorders. Long-term air pollution, particularly PM2.5 and NOₓ174 have also been linked to structural and functional brain alterations, as well as to depression and other mental disorders, through mechanisms involving inflammation, oxidative stress, and genetic susceptibility175,176. Socioeconomic factors such as lower family income, parental education, and neighborhood disadvantage predict reduced cortical surface area and a greater risk of neurodevelopmental conditions48,177,178,179,180. Furthermore, early adversities have been consistently associated with structural and functional brain changes and with the development of psychiatric disorders181,182.
External exposomes dysregulate internal biological processes13. Life adversity, unhealthy lifestyles, and loneliness are associated with impaired hypothalamic-pituitary-adrenal (HPA) axis activity, reducing glucocorticoid sensitivity, triggering insulin resistance, and causing endothelial dysfunction183. These adversities are linked to inflammation, elevated peripheral inflammatory cytokines184,185,186,187, and impacts on the microbiome, ultimately increasing the risk of non-communicable diseases, neuropsychiatric and neurological conditions184,186,187. The prediction of clinical outcomes increases when biological variables are integrated with exposomes10,29,79,80. Different factors, including socioeconomic status, environmental factors, perceived discrimination, and country-level inequality levels10,67,79,80,173,188 influence neuroendocrine, brain-gut microbiome66, and neurocognitive markers10. Social determinants of health48 predict healthy aging better than sex and age in diverse populations53,188,189.
Social epigenomics163,190,191 combines exposomes and epigenetics. Social determinants of health, discrimination, exclusion, and socioeconomic adversities influence DNA methylation191. Epigenome-wide association studies (EWAS) of socioeconomic status and social mobility have implicated genes involved in inflammation, immune function, stress response, metabolic processes, and neural development. Biological age is influenced by ancestry, lifestyles, and social factors190,192,193. Hispanic populations have lower intrinsic epigenetic age acceleration, indicating slower cellular aging (i.e., the Hispanic paradox, with slower aging processes despite having increased risk factors)167,194. Effects of social disparities on epigenetics can also be mitigated by lifestyle changes in some ethnic groups. Methylomics and transcriptomics analyses from the human postmortem brain and studies on genetic variation reveal a significant impact of genetic ancestry, particularly affecting genes involved in immune response pathways156.
AD/ADRD
Different exposomes impact the brain structure-functioning48,171, health53,189, and disease10. Ancestry and social exposome influence biomarkers for AD/ADRD195. Hispanic populations have increased adverse physical and social exposomes and higher AD biomarkers AT(N) such as Aβ40 and reduced neurofilament plasma levels compared to non-Hispanic White populations in the US196. Cardiometabolic diseases, unequally distributed worldwide and influenced by the internal and social exposomes, modulate AT(N) biomarkers195. Socioeconomic factors affect the relationship between brain markers and ancestry195. Cerebrospinal fluid Aβ42/40 levels are negatively correlated with total tau and p-tau181 in Caucasians, but the correlation is weaker in non-Hispanic Black individuals51. CSF Aβ42/40, total tau, and p-tau181 demonstrate weaker correlations with cognitive measures in Black than White participants197. Effects of ancestry and social disparity in AD/ADRD have also been reported in organ194,198, circadian198, and brain clocks10.
The dependence on proxies is an important limitation, as these are indirect indicators used when direct measures are unavailable. Unlike endophenotypes (biologically grounded intermediate traits) or surrogate outcomes (validated substitutes for clinical endpoints), proxies provide pragmatic but non-causal markers. Although much of the evidence is not causal, progress in the field is already evident1,2,3,4,5. A hierarchy of exposome factors5,6,7 can situate biological mechanisms alongside broader social, economic, and environmental factors. A new generation of methods is actively tackling this challenge8, including the NIH Gateway Exposome Coordinating Center, and the predictive exposome databases9 are bringing multiple innovations. High-resolution exposure assessment10, and evaluations of exposure-wide association studies (ExWAS) can improve specificity and granularity1,10,11,12,13. Compared to traditional self-report instruments, which are prone to recall and response bias, these emerging biomarker-based tools offer greater accuracy, temporal resolution, and translational value.
Modeling and theorization
Different computational and theoretical architectures are required to assess the flexibility and complexity needed to incorporate multimodal diversity (Box 2). Specific examples include dynamic causal modeling199, Bayesian architectures200, active inference models201, and biophysical modeling78. These methods have proven effective in integrating multimodal data while being generative, recursive, and continuously updated as new information becomes available15. Data-driven generative approaches enable the development of single-subject and personalized translational models65,71,84. This framework does not support the idea of extensive, universal, or one-size-fits-all models of brain health. Instead, it advocates for tailored models in which the relevance of predictors and outcomes varies according to population-specific data, contextual variables, and multimodal variation. The metamodel framework requires tailored procedures that can be instantiated using different techniques (Tables 2 and 3). We propose four main avenues to develop future computational architectures.
Low-dimensional integration of multilevel brain measures
Multimodal diversity can be mapped to low-level biological mechanisms using methods from artificial intelligence202 and dynamic systems modeling15. On the one hand, non-linear deep learning dimensionality reduction techniques enable the generation of highly informative low-dimensional latent representations of brain spatiotemporal dynamics. Non-linear low-dimensionality reduction approaches can more accurately characterize different conditions than the traditional statistical approaches relying on assumptions of linearity and stationarity82,83,203. Machine learning can be complemented by theory-driven dynamical system measures, such as metastability (the brain’s ability to transition between globally integrated and segregated states204), irreversibility (linked to brain complexity, entropy and flexible brain transitions205), and criticality (a metric of proximity to a dynamic regime in-between order and chaos), which have proven effective in capturing and conveying the structural and functional complexity of the brain across levels in brain disorders205.
Generative biophysical whole-brain models help to investigate mechanisms underlying data-driven characterizations15. These models integrate multiple heterogeneous data sources, encompassing brain structure and function, connectomics, neurochemistry, synaptic processes, or cytoarchitecture206. Moreover, those models can track brain circuitry integrity and function at multiple spatial and temporal scales combining fMRI, electroencephalography (EEG), magnetoencephalography (MEG), or PET, enhancing model predictive power and multilevel consistency206.
Embodied and generative models for extracerebral influences
Combining different organ clocks with computational architectures helps to characterize brain health and disorders168,169,198. Accelerated aging in particular organs, such as the heart, is linked to increased risks of brain disorders168. For AD, such risk is independent of the blood-based biomarkers168. The global effects of such extracerebral influences can be modeled as descriptors of the global state of the system (order parameters), capturing and summarizing the nature of dynamic interactions within the brain15. Order parameters can include whole-body signals, such as autonomic responses, or exposome factors, thus addressing biology-environmental interactions15,.
From a data-driven approach, the effect of extracerebral influences on the brain can be determined using low-dimensional latent representations of spatiotemporal neural dynamics207. A concrete example are variational autoencoders, generative models combining deep learning with probabilistic inference, to track low-dimensional latent representations of multilevel brain states in health and disease82,203. This approach could be extended to incorporate integrated representations of the brain, bodily states, and exposome, providing a set of variables simultaneously encoding multiple dimensions15. These representations can be studied in response to changes in extracerebral factors, including dietary inputs, physical activity, stress exposure, and other exposomes9,82.
Incorporating heterogeneity and individual trajectories
Individual and personalized analyses are critical for global healht208. This requires single-subject data and estimates of how individual’s biology interacts with heterogeneous environmental factors. In biophysical generative models, country-level heterogeneities can be studied as modulators of parameters linked to neural activity, metabolism, structural atrophy, and pathological proteins82,206,208. Examples are found in the BigBrain project209, integrating tissue models and neuroanatomical individual data across scales and providing microstructural maps enriched with cellular and molecular levels merged with fMRI connectivity models, as well as simulations of diverse brain disorders. Other initiatives provide personalized transcriptomic210, proteomic211, and neurochemical maps212 to enrich biophysical generative models.
Perturbational approaches can be used for personalized medicine15, where the effect of changing variables (perturbations) on the observed phenotype can be systematically mapped213. This approach can also be used as a data augmentation technique to enhance deep learning frameworks, generating synthetic data that incorporates the characteristics of underrepresented groups, improving sensitivity and generalizability214. Another way to increase personalization is by Bayesian models, which provide a probabilistic framework that continuously updates predictions as individual data becomes available. This adaptability is essential for personalized brain health assessments that have facing scarce and heterogeneous data, enabling the integration of individual-specific data into models originally built from large population datasets.
Personalized models and individual trajectories can be complementary. Personalized models integrate prior information (genetic, clinical, or environmental data) to tailor predictions to individuals. These models can employ supervised learning and probabilistic frameworks that adapt group-level findings to individual profiles215. In contrast, models of individual trajectories focus on within-person changes over time, capturing temporal dynamics like disease progression or treatment response. Techniques such as Bayesian updating, hidden Markov models, and state-space modeling are commonly used to estimate these temporal trajectories216,217. Combining both strategies enables more accurate, dynamic, and context-sensitive assessments, advancing individualized approaches throughout the lifespan.
Health and disease are inherently variable and not suited to one-size-fits-all approaches218. Normative modeling generates population-based reference trajectories, quantifies individual deviations, enabling detection of atypical patterns in healthy and clinical populations219,220,221,222. Normative modeling uses continuous trait modeling (e.g., brain age deviation), thresholding to define clinical significance, and hybrid models222. Hierarchical Bayesian methods and data integration have enhanced their biological relevance222,223. Lifespan-informed designs demonstrate how these models improve representativeness and mechanistic insight220,221. Medical digital twins132 represent a natural evolution of these models, allowing real-time simulation of individualized biological systems. The combination of multimodal data spanning organ-level dynamics, omics, and exposomic inputs can help AI-driven digital twins to provide personalized aging clocks of health and disease.
A metamodeling heuristic framework
Current theoretical explanations do not systematically address the challenges posed by integrating brain measures with extracerebral information into individual trajectories15,34. New computational strategies can help to incorporate multimodal diversity in psychiatry224 and neurology185,225. The combination of multimodal data, brain-extracerebral integration, and personalized approaches provides bona fide examples, handling population heterogeneity, multilevel metrics, and neuroecological links10 with emerging applications77,78,81,82,208,214.
Future models must effectively integrate diverse data sources while accounting for individual differences and the specific needs of diverse populations. While current methods show promise, they require significant enhancements to better capture non-linear interactions and individual variability15. Achieving this will require integrating advanced tools, such as synthetic data generation and probabilistic modeling. Additionally, balancing simplification and precision is crucial in future developments to avoid oversimplifying complex interactions while ensuring practical and actionable outcomes.
There is an overreliance on causal models and explanations of brain health, especially in neuroscience research226. Causality, although bolstered by powerful technologies, is fundamentally limited15. Studying isolated neural components or perturbing circuits offers causal claims about necessity and sufficiency but fails to reveal the embedded principles that govern behavior, largely due to ecological constraints, brain degeneracy, multiple realizability, and the lack of a direct mapping between structure and function. Pure causal models often fall into the mereological fallacy (attributing organismal or environmental functions to parts like neurons or isolated factors), overlooking brain health as ecological phenomenon. They lack the capacity to explain why and how health and disease emerge beyond mechanical causation. Brain health must move beyond pure causal manipulation towards a pluralistic approach that prioritizes precise neurosyndemics as a foundational step15. Understanding this embedded and situated nature through complexity science can help uncover emergent properties triggered by exotype-endotype interactions9. Synergistic explanatory frameworks13,172,227,228,229 better capture the complexity of brain health through integration across levels, enhancing both explanatory power and translational relevance. Bona fide examples of such integrative models include brain and organ aging clocks168,169,230, allostatic load131,231,232,233,234,235, and the biological embedding of adversity236,237,238.
Dimensional approaches for psychiatry and neurology
Although our perspective is mainly focused on brain health, multimodal diversity in research supports a neurological and psychiatric dimensionality. The core heuristics apply equally to psychiatric disorders. Neuropsychiatric genomics continues to be underrepresented in populations with high genetic admixture, distinct environmental exposures, and healthcare disparities, limiting equity in precision psychiatry239,240. GWAS including African and Latino participants have identified novel variants for depression, schizophrenia, anxiety, and bipolar disorder—missed in European-only studies116,240. Studies from the Middle East have uncovered multiple novel gene–phenotype associations in psychiatric disorders. In autism spectrum disorder (ASD), numerous novel candidates include GLT8D1, HTATSF1, OR6C65, ITIH6, and DDX26B in Saudi families241, DPP4, TRHR, and MLF1 in Lebanese families242, and TRPC4 and SCFD2 in Qatari cohorts243. Copy number studies in Lebanon further implicated PTDSS1 and AREG as novel ASD susceptibility loci244. In schizophrenia, Israeli trio and founder-population studies revealed a burden of rare damaging de novo mutations245, while Iranian families carrying AGO1 truncating variants showed a spectrum from autism to schizophrenia246. For bipolar disorder, Turkish multiplex families yielded novel variants in TMTC1 and STARD9247. These findings underscore how Middle Eastern cohorts are expanding the catalog of rare and novel genetic contributors to psychiatric disease.
Environmental, cultural, and social determinants shape psychiatric conditions14,15,64,177,178. Global exposomic variation, combined with individual and societal-level risks, has been linked to general psychopathology248,249,250. In schizophrenia and bipolar disorder, factors such as urbanization, gross domestic product per capita, employment, and air pollution influence clinical expression251, underscoring the need for context-sensitive diagnosis and treatment250. Computational psychiatry has improved classification and prediction by integrating neuroimaging, genetics, behavior, and environmental data252,253,254. Investigating dimensional biological and exposome pathways in psychiatry and neurology14,64,255 is critical to advance the field. There is a high prevalence of neurodegenerative conditions among patients with anxiety, depression, schizophrenia, bipolar disorder, personality disorders, and stress-related conditions256, with overlapping genetic causes257, shared omics mechanisms256, and social risks5. Addressing these shared multifactorial interactions255 requires adopting transdiagnostic, dimensional, and integrative approaches.
Emerging models guided by the BePRECISE consortium20, emphasize integrating diverse datasets and transparent methodologies to ensure inclusivity, reproducibility, and actionable outcomes for psychiatry and neurology. These approaches align with the principles proposed here, incorporating genetic, biological, and exposome data to refine predictive models and improve clinical interventions.. In psychiatry, deep phenotyping following BePRECISE guidelines combines neuroimaging, genetic markers, and environmental exposures to enable the identification of depression subtypes associated with specific treatment responses47. These dimensional frameworks can help advance the fields of psychiatry and neurology and lay the groundwork for inclusive tackling of the real-world complexity of health and disease.
Conclusions: a future agenda
Recent promising advances in specific fields (Box 3) call for a more comprehensive science of precision brain health. Developing a neurosyndemic framework, incorporating diverse, multimodal models, and accounting for individual variability are essential to achieving a meaningful impact in real-world applications. We outline future strategies to provide more holistic, ecologically valid models that better reflect the complexities of human brain health (Table 1).
Critical steps are required to develop the advanced metamodeling frameworks (Fig. 2). These must simultaneously combine different strategies (spatiotemporal modeling via dimensionality reduction, whole-brain biophysical modeling, deep learning, and probabilistic structures) to effectively incorporate data representation and integration, extracerebral influences, inherent heterogeneity, and personalized modeling. Model perturbation strategies can be relevant to gain mechanistic insights15. Future metamodel frameworks should focus on human behavior and neural activity within real-world environments with more naturalistic and ecologically valid research paradigms11,258. Wearables, natural speech analysis, virtual reality, augmented reality, real-time neuroimaging and neurofeedback, citizen science, and ecological simulations may help better assess diversity11,258.
A framework for multimodal diversity that fosters precision medicine approaches needs to address the complexity of brain health and disorders while guiding personalized strategies. The a illustrates the concept of diversity in data and models for brain health, integrating inputs from epi/genetics (polygenic risk, genetic causation, epigenetic clocks), whole-body health (cardiometabolic measures, omics), neurocognition (MBM: multimodal brain measures; and BCC: behavior, cognitive and clinical levels), and the exposome (social and physical factors). These heterogeneous data streams converge into different combinations of computational approaches (lower part of the panel) addressing low-dimensional latent representations, multimodal brain integration, extracerebral measures, and individual heterogeneity, ultimately enabling tailored model types that enhance precise brain health. b shows multimodal diversity models integrating social factors (e.g., loneliness, economic hardship, political influences, and pollution) with data from genetic risks to exposome exposures. A nested modeling approach (hierarchical method in which a simpler model is embedded within a more complex one, allowing refinement across levels of complexity) supports the differentiation of risk profiles in individuals with brain disorders (neurological and psychiatric) and healthy controls (HC). c exemplifies the utility of this framework in brain health, highlighting how integrating multiple biological levels enhances the predictive power of biomarkers, clinical profiles, and neurocognitive data for brain health and disorders. d illustrates derived precision health applications in Alzheimer’s Disease (AD). This approach accounts for cases with differential genetic and probabilistic (environmental) burdens. Examples of future tailored interventions include gene therapy for individuals with high genetic burdens (e.g., inspired by the Christchurch protective gene for the PSEN1 mutation) and multicomponent interventions (e.g., enhanced social interactions, physical activity, healthy diets, and medical monitoring) tailored for those with probabilistic burdens. Created in BioRender. Migeot, J. (https://BioRender.com/l00oejq) (brain, body, and organs illustrations).
Deep phenotyping is essential for precision medicine, especially when it enables detailed clinical phenotypes (e.g., symptoms, disease progression) and endophenotypes across the whole body. Future efforts could lead to new disorder subtypes based on fine-grained clinical and biological correlates259. These advancements would support the creation of more global and extensive databases, tracking population representations and clinical subtypes. Developing human digital twins for brain health132 could integrate deep phenotyping data with temporal monitoring and prediction. These in silico replicas would enable scenario testing for future prevention and treatment, linking multi-organ and multi-omics to public health and clinical applications.
Research incorporating biological and environmental interactions requires further granularity. Regional public health strategies can benefit from integrating individual risk profiles and personalized interventions combining macro (i.e., aggregate-level exposome) and micro (i.e., individual-level brain health) factors16. Deep phenotyping in longitudinal cohorts can help to identify individual protective or risk factors that may influence broader geographical regions. In contrast, personalized medicine technologies in health and disease can help to improve healthcare accessibility, especially for marginalized communities. Implementing convergences between regional and individual models requires complexity modeling and big data32, while prioritizing data privacy and equity. Integrating population admixtures and multimodal data is essential for predicting clinical diagnoses characterizing healthy variation, clarifying frequent and infrequent phenotypes, and measuring individual trajectories. Such models help distinguish normative variability from clinically relevant deviations, a persistent challenge in neurology and psychiatry. For example, accounting for ancestry, lifestyle, social risks, and environmental exposures improves the identification of subclinical brain aging or stress-related vulnerabilities and resilience before disease onset16,192,193,218. Prediction models should support a broader goal: understanding how individuals respond to environmental risks and interventions across the life course16, rather than simply classifying disease states.
Future research should also bring more science of neurodevelopmental factors. Studying these factors can help minimize the risk of overemphasizing or misattributing the influence of factors from adulthood or older age in brain disorders260. Early-life adversities such as adversity, malnutrition, infections, and lack of cognitive stimulation are linked to disruptions in critical neurodevelopmental stages261 and contribute to the onset of psychiatric262 and neurological conditions263,264. This approach can explain mechanistically how such adversities affect biological pathways leading to brain disorders260. Despite the importance of early exposures in brain disorders89,90,91,256, yet most large-scale datasets lack of further assessment of neurodevelopmental measures to trace these trajectories. Expanding lifespan cohorts that capture neurodevelopmental milestones is crucial to understanding how early-life pathways contribute to midlife psychiatric and neurological disorders and their progression to late-life diseases220,221. Developmentally informed computational approaches, drawing on dynamic causal modeling, biophysical modeling, and normative modeling265 may enable more precise, equitable predictions by identifying deviations across lifespan that impact brain health outcomes.
Transdisciplinary collaboration is required to achieve multimodal diversity in clinical sciences, neuroscience, biology, epidemiology, computational modeling, and public health. Cutting-edge technologies (e.g., AI and biophysical modeling), knowledge sharing266, and open science practices and, are essential for future development. The brain capital267 and syndemics172,268,269 agendas can drive transdisciplinary innovation regarding critical challenges (climate change, pollution/plastics, conflict and stress, economic innovation) for brain and mental wealth13,267.
Towards addressing ethical, social, and technical barriers
Structural inequities shape data access, governance, and commercialization, reinforcing disparities in data ownership and use. Academia sets priorities, industry monetizes data, and policymakers regulate access—roles that can perpetuate or challenge injustice. Expanding datasets without safeguards risks eroding participant rights and centralizing control. Diversity must be valued not only for scientific validity but as a core ethical principle. To prevent harm, frameworks like WHO, STANDING Together, and FAIR36,39,103,218,270,271,272,273,274,275 can promote participatory governance, transparency, and fair benefit-sharing. AI bias on health data often arises from underrepresentation and uneven data quality100, compromising fairness, generalizability, and impact276,277. Restricted diversity can entrench health disparities100, but the solution must go beyond correcting algorithmic bias to embed ethical oversight, structural awareness, and stakeholder inclusion into equitable brain health modeling. Models must capture real-world heterogeneity, integrate multilevel data, and be co-developed with communities. Transparent governance276,278, global data-sharing frameworks103, and accountability are required prevent AI from reinforcing disparities. Other ethical risks linked to AI and big data, corporate misuse, circular predictions, confirmation bias, and automated decisions in low-interpretability models should not be underestimated276. Data protection, especially in the context of generative models capable of re-identifying individuals and challenging anonymization efforts need to be addressed32,113,279.
Citizen science280, community-based system dynamics, digital technologies, and participatory research approaches267,280 can help to democratize this framework. These strategies promote culturally competent recruitment and radical collaboration with community stakeholders, informing the equitable mapping of complex systems influencing health outcomes, including social determinants, healthcare access, and cultural factors281. Promoting equitable data sharing across jurisdictions, fostering public-private partnerships, building capacity in low-resource settings for active participation in discoveries, and reducing institutional and national barriers are crucial for advancing brain health interventions globally281. Such methods identify critical leverage points to improve data representation and accuracy, leading to practical, sustainable data capture tools and meaningful interventions for diverse populations267,280. We believe this framework will help to establish frank global brain health research agenda that benefits all populations, ensuring equity in knowledge and care across regions.
The proposed framework leverages advances in global multilevel data, theoretical and computational modeling, and transdisciplinary collaboration to build more comprehensive precision models of brain health. This approach may enhance our future understanding of complex brain health dynamics and intrinsically commit to equity, warranting scientific advancements benefit all people across the world.
References
Collaborators, G. B. D. F. Burden of disease scenarios for 204 countries and territories, 2022-2050: a forecasting analysis for the Global Burden of Disease Study 2021. Lancet 403, 2204–2256 (2024).
Harris, E. Neurological conditions are leading cause of disability worldwide. JAMA 331, 1440 (2024).
Momen, N. C. et al. Association between mental disorders and subsequent medical conditions. N. Engl. J. Med 382, 1721–1731 (2020).
Collaborators, G. B. D. M. D Global, regional, and national burden of 12 mental disorders in 204 countries and territories, 1990-2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet Psychiatry 9, 137–150 (2022).
Steinmetz, J. D. et al. Global, regional, and national burden of disorders affecting the nervous system, 1990–2021: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Neurol. 23, 344–381 (2024).
Feigin, V. L. et al. Global, regional, and national burden of stroke and its risk factors, 1990–2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet Neurol. 20, 795–820 (2021).
Wig, G. S. et al. Participant diversity is necessary to advance brain aging research. Trends Cogn. Sci. 28, 92–96 (2024).
McGorry, P. D. et al. The lancet psychiatry commission on youth mental health. Lancet Psychiatry 11, 731–774 (2024).
Hernandez, H. et al. The exposome of healthy and accelerated aging across 40 countries. Nat. Med 31, 3089–3100 (2025).
Moguilner, S. et al. Brain clocks capture diversity and disparities in aging and dementia across geographically diverse populations. Nat. Med 30, 3646–3657 (2024).
Ibanez, A. The mind’s golden cage and cognition in the wild. Trends Cogn. Sci. 26, 1031–1034 (2022).
Ibanez, A. Inspired by struggle: a personal journey to global precision brain health. iScience 28, 111918 (2025).
Ibanez, A. et al. Neuroecological links of the exposome and one health. Neuron 112, 1905–1910 (2024).
Ibanez, A. & Zimmer, E. R. Time to synergize mental health with brain health. Nat. Ment. Health 1, 441–443 (2023).
Ibanez, A., Kringelbach, M. L. & Deco, G. A synergetic turn in cognitive neuroscience of brain diseases. Trends Cogn. Sci. 28, 319–338 (2024).
Udeh-Momoh, C. T. et al. Resilience and brain health in global populations. Nat. Med 31, 2518–2531 (2025).
Lengnick-Hall, R. et al. Eight characteristics of rigorous multilevel implementation research: a step-by-step guide. Implement Sci. 18, 52 (2023).
Holmes, J. H. et al. Challenges for multilevel health disparities research in a transdisciplinary environment. Am. J. Prev. Med 35, S182–S192 (2008).
Lee, S. et al. Multimodal integration of neuroimaging and genetic data for the diagnosis of mood disorders based on computer vision models. J. Psychiatr. Res. 172, 144–155 (2024).
Morton, J. T. et al. Multi-level analysis of the gut-brain axis shows autism spectrum disorder-associated molecular and microbial profiles. Nat. Neurosci. 26, 1208–1217 (2023).
Ressler, K. J. & Williams, L. M. Big data in psychiatry: multiomics, neuroimaging, computational modeling, and digital phenotyping. Neuropsychopharmacology 46, 1–2 (2021).
Rohaut, B. et al. Multimodal assessment improves neuroprognosis performance in clinically unresponsive critical-care patients with brain injury. Nat. Med 30, 2349–2355 (2024).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
Duggan, M. R. et al. Proteomics identifies potential immunological drivers of postinfection brain atrophy and cognitive decline. Nat. Aging 4, 1263–1278 (2024).
Holmes, E. A. et al. Multidisciplinary research priorities for the COVID-19 pandemic: a call for action for mental health science. Lancet Psychiatry 7, 547–560 (2020).
Bjerke, I. E. et al. Data integration through brain atlasing: human brain project tools and strategies. Eur. Psychiatry 50, 70–76 (2018).
Doyle, A. E. et al. Advancing mental health research through strategic integration of transdiagnostic dimensions and genomics. Biol. Psychiatry 97, 450–460 (2025).
Liu, L., Feigin, V., Sacco, R. L. & Koroshetz, W. J. Promoting global collaboration for brain health research. BMJ 371, m3753 (2020).
Owolabi, M. O. et al. Global synergistic actions to improve brain health for human development. Nat. Rev. Neurol. 19, 371–383 (2023).
Haken, H. Synergetics of brain function. Int J. Psychophysiol. 60, 110–124 (2006).
Poline, J. B. et al. Data and tools integration in the Canadian Open Neuroscience Platform. Sci. Data 10, 189 (2023).
Matshabane, O. P. Promoting diversity and inclusion in neuroscience and neuroethics. EBioMedicine 67, 103359 (2021).
Hachinski, V. Integral brain health: Cerebral/mental/social provisional definitions. Alzheimers Dement 19, 3226–3230 (2023).
Greene, A. S. et al. Brain-phenotype models fail for individuals who defy sample stereotypes. Nature 609, 109–118 (2022).
Stephan, B. C., Kurth, T., Matthews, F. E., Brayne, C. & Dufouil, C. Dementia risk prediction in the population: are screening models accurate? Nat. Rev. Neurol. 6, 318–326 (2010).
Xue, D., Blue, E. E., Conomos, M. P. & Fohner, A. E. The power of representation: Statistical analysis of diversity in US Alzheimer’s disease genetics data. Alzheimers Dement (N. Y) 10, e12462 (2024).
Gorelick, P. B., Hainsworth, A. H. & Wallin, A. What will it take to achieve brain health globally? Cereb. Circ. Cogn. Behav. 6, 100209 (2024).
Lydon-Staley, D. M., Cornblath, E. J., Blevins, A. S. & Bassett, D. S. Modeling brain, symptom, and behavior in the winds of change. Neuropsychopharmacology 46, 20–32 (2021).
Matshabane, O. P., Whitted, C. G. & Koehly, L. M. Addressing diversity and inclusion challenges in global neuro-psychiatric and behavioral genomics research. Front Genet. 13, 1021649 (2022).
Baez, S., Alladi, S. & Ibanez, A. Global South research is critical for understanding brain health, ageing and dementia. Clin. Transl. Med 13, e1486 (2023).
Eteleeb, A. M. et al. Brain high-throughput multi-omics data reveal molecular heterogeneity in Alzheimer’s disease. PLoS Biol. 22, e3002607 (2024).
Step, K., Ndong Sima, C. A. A., Mata, I. & Bardien, S. Exploring the role of underrepresented populations in polygenic risk scores for neurodegenerative disease risk prediction. Front Neurosci. 18, 1380860 (2024).
Kunkle, B. W. et al. Novel Alzheimer disease risk loci and pathways in African American individuals using the African Genome Resources Panel: a meta-analysis. JAMA Neurol. 78, 102–113 (2021).
Rajabli, F. et al. Ancestral origin of ApoE epsilon4 Alzheimer disease risk in Puerto Rican and African American populations. PLoS Genet 14, e1007791 (2018).
Parra, M. A. et al. Dementia in Latin America: paving the way toward a regional action plan. Alzheimers Dement 17, 295–313 (2021).
Kandiah, N. et al. Current and future trends in biomarkers for the early detection of Alzheimer’s Disease in Asia: Expert Opinion. J. Alzheimers Dis. Rep. 6, 699–710 (2022).
Meeker, K. L. et al. Socioeconomic status mediates racial differences seen using the AT(N) framework. Ann. Neurol. 89, 254–265 (2021).
Migeot, J. et al. Social exposome and brain health outcomes of dementia across Latin America. Nat. Commun. 16, 8196 (2025).
Ibanez, A., Legaz, A. & Ruiz-Adame, M. Addressing the gaps between socioeconomic disparities and biological models of dementia. Brain 146, 3561–3564 (2023).
Stephan, B. C. M. et al. Prediction of dementia risk in low-income and middle-income countries (the 10/66 Study): an independent external validation of existing models. Lancet Glob. Health 8, e524–e535 (2020).
McGlinchey, E. et al. Biomarkers of neurodegeneration across the Global South. Lancet Healthy Longev. 5, 100616 (2024).
Taylor, D. J. et al. Sources of gene expression variation in a globally diverse human cohort. Nature 632, 122–130 (2024).
Santamaria-Garcia, H. et al. Factors associated with healthy aging in Latin American populations. Nat. Med 29, 2248–2258 (2023).
Mostert, C. M. et al. Broadening dementia risk models: building on the 2024 Lancet Commission report for a more inclusive global framework. EBioMedicine 120, 105950 (2025).
Fittipaldi, S., Migeot, J. & Ibanez, A. Socioeconomic disparities harm social cognition. Trends Cogn. Sci. 28, 386–387 (2024).
Li, J. et al. Cross-ethnicity/race generalization failure of behavioral prediction from resting-state functional connectivity. Sci. Adv. 8, eabj1812 (2022).
Raman, R. et al. Disparities by race and ethnicity among adults recruited for a preclinical alzheimer disease trial. JAMA Netw. Open 4, e2114364 (2021).
Liu, F. et al. Environmental neuroscience linking exposome to brain structure and function underlying cognition and behavior. Mol. Psychiatry 28, 17–27 (2023).
Bruxel, E. M. et al. Psychiatric genetics in the diverse landscape of Latin American populations. Nat. Genet. 57, 1074–1088 (2025).
Hazelton, J. L. et al. Cardiovascular risk factors and the allostatic interoceptive network in dementia. Cardiovasc Res, 121, cvaf185 (2025).
Miranda, J. J. et al. Understanding the rise of cardiometabolic diseases in low- and middle-income countries. Nat. Med 25, 1667–1679 (2019).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Liao, Y. et al. Surveillance of health status in minority communities-racial and ethnic approaches to community health across the US (REACH US) risk factor survey, United States, 2009. MMWR Surveill. Summ. 60, 1–44 (2011).
Santamaria-Garcia, H. et al. Allostatic interoceptive overload across psychiatric and neurological conditions. Biol. Psychiatry 97, 28–40 (2025).
Sathyanarayanan, A. et al. Multi-omics data integration methods and their applications in psychiatric disorders. Eur. Neuropsychopharmacol. 69, 26–46 (2023).
Zhang, X. et al. Discrimination exposure impacts unhealthy processing of food cues: crosstalk between the brain and gut. Nat. Ment. Health 1, 841–852 (2023).
Legaz, A. et al. Structural inequality linked to brain volume and network dynamics in aging and dementia across the Americas. Nat. Aging 5, 259–274 (2025).
Maitre, L. et al. Multi-omics signatures of the human early life exposome. Nat. Commun. 13, 7024 (2022).
Yoon, J. H. et al. Integrative approach of omics and imaging data to discover new insights for understanding brain diseases. Brain Commun. 6, fcae265 (2024).
Vieira, S. et al. Multivariate brain-behaviour associations in psychiatric disorders. Transl. Psychiatry 14, 231 (2024).
Teufel, C. & Fletcher, P. C. The promises and pitfalls of applying computational models to neurological and psychiatric disorders. Brain 139, 2600–2608 (2016).
Dhamala, E., Yeo, B. T. T. & Holmes, A. J. One size does not fit all: methodological considerations for brain-based predictive modeling in psychiatry. Biol. Psychiatry 93, 717–728 (2023).
Falcon, M. I., Jirsa, V. & Solodkin, A. A new neuroinformatics approach to personalized medicine in neurology: The Virtual Brain. Curr. Opin. Neurol. 29, 429–436 (2016).
Lee, Y. S., Garrido, N. L. B., Lord, G., Maggio, Z. A. & Khomtchouk, B. B. Ethical considerations for biobanks serving underrepresented populations. Bioethics 39, 240–249 (2025).
Schoeler, T. et al. Participation bias in the UK Biobank distorts genetic associations and downstream analyses. Nat. Hum. Behav. 7, 1216–1227 (2023).
Collin, C. B. et al. Computational models for clinical applications in personalized medicine-guidelines and recommendations for data integration and model validation. J. Pers. Med. 12, 166 (2022).
Moguilner, S. et al. Multi-feature computational framework for combined signatures of dementia in underrepresented settings. J. Neural Eng. 19, 4 (2022).
Moguilner, S. et al. Biophysical models applied to dementia patients reveal links between geographical origin, gender, disease duration, and loss of neural inhibition. Alzheimers Res. Ther. 16, 79 (2024).
Hernandez, H. et al. Brain health in diverse settings: How age, demographics and cognition shape brain function. Neuroimage 295, 120636 (2024).
Gonzalez-Gomez, R. et al. Educational disparities in brain health and dementia across Latin America and the United States. Alzheimers Dement 20, 5912–5925 (2024).
Herzog, R. et al. Genuine high-order interactions in brain networks and neurodegeneration. Neurobiol. Dis. 175, 105918 (2022).
Sanz Perl, Y. et al. Model-based whole-brain perturbational landscape of neurodegenerative diseases. Elife 12, e83970 (2023).
Coronel-Oliveros, C. et al. Viscous dynamics associated with hypoexcitation and structural disintegration in neurodegeneration via generative whole-brain modeling. Alzheimers Dement 20, 3228–3250 (2024).
Leonardsen, E. H. et al. Constructing personalized characterizations of structural brain aberrations in patients with dementia using explainable artificial intelligence. NPJ Digit Med 7, 110 (2024).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Gilmore-Bykovskyi, A. L. et al. Recruitment and retention of underrepresented populations in Alzheimer’s disease research: A systematic review. Alzheimers Dement (N. Y) 5, 751–770 (2019).
Parra, M. A. Overcoming barriers in cognitive assessment of Alzheimer’s disease. Dement Neuropsychol. 8, 95–98 (2014).
Bhutta, Z. A., Bhavnani, S., Betancourt, T. S., Tomlinson, M. & Patel, V. Adverse childhood experiences and lifelong health. Nat. Med 29, 1639–1648 (2023).
Soh, Y. et al. State-level indicators of childhood educational quality and incident dementia in older black and white adults. JAMA Neurol. 80, 352–359 (2023).
Farina, F. R. et al. Next generation brain health: transforming global research and public health to promote prevention of dementia and reduce its risk in young adult populations. Lancet Healthy Longev. 5, 100665 (2024).
Raju, R. M. & Tsai, L. H. Dismantling silos: the case for an integrated approach to address childhood determinants of lifelong brain health. Neuron 111, 3706–3709 (2023).
Bolleddula, J. et al. No person left behind: Mapping the health policy landscape for genomics research in the Caribbean. Lancet Reg. Health Am. 15, 100367 (2022).
Signore, I. A. et al. The Chilean COVID-19 genomics network biorepository: a resource for multi-omics studies of COVID-19 and long COVID in a Latin American Population. Genes (Basel) 15, 1352 (2024).
Sohail, M. et al. Mexican Biobank advances population and medical genomics of diverse ancestries. Nature 622, 775–783 (2023).
Prado, P. et al. The BrainLat project, a multimodal neuroimaging dataset of neurodegeneration from underrepresented backgrounds. Sci. Data 10, 889 (2023).
All of Us Research Program Genomics, I Genomic data in the all of us research program. Nature 627, 340–346 (2024).
Bethlehem, R. A. I. et al. Brain charts for the human lifespan. Nature 604, 525–533 (2022).
Devaney, S. A., Malerba, L. & Manson, S. M. The “All of Us” Program and Indigenous Peoples. N. Engl. J. Med 383, 1892 (2020).
Ibrahim, H., Liu, X., Zariffa, N., Morris, A. D. & Denniston, A. K. Health data poverty: an assailable barrier to equitable digital health care. Lancet Digit Health 3, e260–e265 (2021).
Ibanez, A., Parra, M. A., Butler, C. & Latin, A. & the Caribbean Consortium on, D. The Latin America and the caribbean consortium on dementia (LAC-CD): from networking to research to implementation science. J. Alzheimers Dis. 82, S379–S394 (2021).
Blaisdell, C. J. et al. The NIH ECHO program: investigating how early environmental influences affect child health. Pediatr. Res. 92, 1215–1216 (2022).
Bajcsy, P. et al. Enabling global image data sharing in the life sciences. Nat. Methods 22, 672–676 (2025).
Mulder, N. et al. H3Africa: current perspectives. Pharmgenomics Pers. Med 11, 59–66 (2018).
Rosenberg, A., Mangialasche, F., Ngandu, T., Solomon, A. & Kivipelto, M. Multidomain interventions to prevent cognitive impairment, alzheimer’s disease, and dementia: from FINGER to World-Wide FINGERS. J. Prev. Alzheimers Dis. 7, 29–36 (2020).
Duran-Aniotz, C. et al. The Latin American Brain Health Institute, a regional initiative to reduce the scale and impact of dementia. Alzheimers Dement 18, 1696–1698 (2022).
Ibanez, A. et al. The multi-partner consortium to expand dementia research in Latin America (ReDLat): Driving Multicentric Research and Implementation Science. Front Neurol. 12, 631722 (2021).
Bigdeli, T. B. et al. Contributions of common genetic variants to risk of schizophrenia among individuals of African and Latino ancestry. Mol. Psychiatry 25, 2455–2467 (2020).
Cuellar-Barboza, A. B. et al. Polygenic prediction of bipolar disorder in a Latin American sample. Am. J. Med Genet B Neuropsychiatr. Genet. 192, 139–146 (2023).
Meng, X. et al. Multi-ancestry genome-wide association study of major depression aids locus discovery, fine mapping, gene prioritization and causal inference. Nat. Genet. 56, 222–233 (2024).
Kim, J. J. et al. Multi-ancestry genome-wide association meta-analysis of Parkinson’s disease. Nat. Genet. 56, 27–36 (2024).
Keene, K. L. et al. Genome-wide association study meta-analysis of stroke in 22000 individuals of African descent identifies novel associations with stroke. Stroke 51, 2454–2463 (2020).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Atkinson, E. G. et al. Tractor uses local ancestry to enable the inclusion of admixed individuals in GWAS and to boost power. Nat. Genet. 53, 195–204 (2021).
Stevenson, A. et al. Neuropsychiatric Genetics of African Populations-Psychosis (NeuroGAP-Psychosis): a case-control study protocol and GWAS in Ethiopia, Kenya, South Africa and Uganda. BMJ Open 9, e025469 (2019).
Bruxel, E. M. et al. Psychiatric genetics in the diverse landscape of Latin American populations. Nat. Genet. https://doi.org/10.1038/s41588-025-02127-z (2025).
Song, Y. et al. Multitrait genetic analysis identifies novel pleiotropic loci for depression and schizophrenia in East Asians. Schizophr. Bull. 51, 684–695 (2025).
Lam, M. et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat. Genet. 51, 1670–1678 (2019).
Yue, W., Yu, X. & Zhang, D. Progress in genome-wide association studies of schizophrenia in Han Chinese populations. NPJ Schizophr. 3, 24 (2017).
Han, Y. et al. Genetic risk and prognosis of the first incident stroke survivors: findings from China Kadoorie Biobank and UK Biobank. Neurology 105, e213832 (2025).
Acosta-Uribe, J. et al. A neurodegenerative disease landscape of rare mutations in Colombia due to founder effects. Genome Med 14, 27 (2022).
Acosta-Uribe, J. et al. Genetic contributions to alzheimer’s disease and frontotemporal dementia in admixed Latin American Populations. Preprint at medRxiv https://doi.org/10.1101/2024.10.29.24315197 (2024).
Zapata-Restrepo, L. M. et al. Case of early onset Alzheimer’s disease associated with a novel PSEN1 variant identified in Colombia. NPJ Dement 1, 31 (2025).
Quiroz, Y. T. et al. APOE3 Christchurch heterozygosity and autosomal dominant Alzheimer’s Disease. N. Engl. J. Med 390, 2156–2164 (2024).
Lopera, F. et al. Resilience to autosomal dominant Alzheimer’s disease in a Reelin-COLBOS heterozygous man. Nat. Med 29, 1243–1252 (2023).
Ray, N. R. et al. Extended genome-wide association study employing the African genome resources panel identifies novel susceptibility loci for Alzheimer’s disease in individuals of African ancestry. Alzheimers Dement 20, 5247–5261 (2024).
Naslavsky, M. S. et al. Global and local ancestry modulate APOE association with Alzheimer’s neuropathology and cognitive outcomes in an admixed sample. Mol. Psychiatry 27, 4800–4808 (2022).
Hazelton, J. L. et al. Neuroimaging meta-analyses reveal convergence of interoception, emotion, and social cognition across neurodegenerative diseases. Biol. Psychiatry 97, 1079–1090 (2025).
Migeot, J. A., Duran-Aniotz, C. A., Signorelli, C. M., Piguet, O. & Ibanez, A. A predictive coding framework of allostatic-interoceptive overload in frontotemporal dementia. Trends Neurosci. 45, 838–853 (2022).
Migeot, J. et al. Allostasis, health, and development in Latin America. Neurosci. Biobehav Rev. 162, 105697 (2024).
Hazelton, J. L. et al. Altered spatiotemporal brain dynamics of interoception in behavioural-variant frontotemporal dementia. EBioMedicine 113, 105614 (2025).
Wen, J. Towards a multi-organ, multi-omics medical digital twin. Nat. Biomed. Eng. 9, 1386–1389 (2025).
Birba, A. et al. Allostatic-interoceptive overload in frontotemporal dementia. Biol. Psychiatry 92, 54–67 (2022).
Liufu, C. et al. Integration of multi-omics summary data reveals the role of N6-methyladenosine in neuropsychiatric disorders. Mol. Psychiatry 29, 3141–3150 (2024).
Xu, J. et al. Neurobiological substrates underlying the effect of genomic risk for depression on the conversion of amnestic mild cognitive impairment. Brain 141, 3457–3471 (2018).
Bao, J. et al. Integrative analysis of multi-omics and imaging data with incorporation of biological information via structural Bayesian factor analysis. Brief Bioinform https://doi.org/10.1093/bib/bbad073 (2023).
Clark, C., Rabl, M., Dayon, L. & Popp, J. The promise of multi-omics approaches to discover biological alterations with clinical relevance in Alzheimer’s disease. Front Aging Neurosci. 14, 1065904 (2022).
Ravichandran, K. A. & Heneka, M. T. Inflammasomes in neurological disorders - mechanisms and therapeutic potential. Nat. Rev. Neurol. 20, 67–83 (2024).
Badhwar, A. et al. A multiomics approach to heterogeneity in Alzheimer’s disease: focused review and roadmap. Brain 143, 1315–1331 (2020).
Pape, K., Tamouza, R., Leboyer, M. & Zipp, F. Immunoneuropsychiatry - novel perspectives on brain disorders. Nat. Rev. Neurol. 15, 317–328 (2019).
Monari, S. et al. Blunted Glucocorticoid Responsiveness to Stress Causes Behavioral and Biological Alterations That Lead to Posttraumatic Stress Disorder Vulnerability. Biol. Psychiatry 95, 762–773 (2024).
Luo, J. et al. Genetic regulation of human brain proteome reveals proteins implicated in psychiatric disorders. Mol. Psychiatry 29, 3330–3343 (2024).
Ren, P. et al. Atlas of Proteomic signatures of brain structure and its links to brain disorders. Nat. Commun. 16, 5092 (2025).
Fan, J. W. et al. Transcriptomics and magnetic resonance imaging in major psychiatric disorders. Front Psychiatry 14, 1185471 (2023).
Wang, Y. et al. Multi-modal intermediate integrative methods in neuropsychiatric disorders: A review. Comput Struct. Biotechnol. J. 20, 6149–6162 (2022).
Mao, Q. et al. Spatial multiomics analysis in psychiatric disorders. EC Psychol. Psychiatr. 12, 1–5 (2023).
Zhang, H. et al. Mass spectrometry imaging for spatially resolved multi-omics molecular mapping. Npj Imaging 2, 20 (2024).
Dubois, B. et al. Alzheimer disease as a clinical-biological construct-an international working group recommendation. JAMA Neurol. 81, 1304–1311 (2024).
Gleason, C. E. et al. Alzheimer’s disease biomarkers in Black and non-Hispanic White cohorts: A contextualized review of the evidence. Alzheimers Dement 18, 1545–1564 (2022).
Parra, M. A. et al. Biomarkers for dementia in Latin American countries: Gaps and opportunities. Alzheimers Dement 19, 721–735 (2023).
Xu, J. et al. A causal association of ANKRD37 with human hippocampal volume. Mol. Psychiatry 27, 4432–4445 (2022).
Palmqvist, S. et al. Blood biomarkers to detect alzheimer disease in primary care and secondary care. JAMA 332, 1245–1257 (2024).
Nativio, R. et al. An integrated multi-omics approach identifies epigenetic alterations associated with Alzheimer’s disease. Nat. Genet. 52, 1024–1035 (2020).
Shen, Y. et al. CSF proteomics identifies early changes in autosomal dominant Alzheimer’s disease. Cell 187, 6309–6326 e6315 (2024).
Andrews, S. J. et al. The complex genetic architecture of Alzheimer’s disease: novel insights and future directions. EBioMedicine 90, 104511 (2023).
Benjamin, K. J. M. et al. Analysis of gene expression in the postmortem brain of neurotypical Black Americans reveals contributions of genetic ancestry. Nat. Neurosci. 27, 1064–1074 (2024).
Xue, C. et al. AI-based differential diagnosis of dementia etiologies on multimodal data. Nat. Med 30, 2977–2989 (2024).
Wang, M. et al. Transformative network modeling of multi-omics data reveals detailed circuits, key regulators, and potential therapeutics for alzheimer’s disease. Neuron 109, 257–272e214 (2021).
Heffel, M. G. et al. Temporally distinct 3D multi-omic dynamics in the developing human brain. Nature 635, 481–489 (2024).
Chen, Z. et al. Sampling inequalities affect generalization of neuroimaging-based diagnostic classifiers in psychiatry. BMC Med 21, 241 (2023).
Chen, C. et al. Applications of multi-omics analysis in human diseases. MedComm (2020) 4, e315 (2023).
Wu, Y. & Xie, L. AI-driven multi-omics integration for multi-scale predictive modeling of genotype-environment-phenotype relationships. Comput Struct. Biotechnol. J. 27, 265–277 (2025).
McEwen, C. A. Connecting the biology of stress, allostatic load and epigenetics to social structures and processes. Neurobiol. Stress 17, 100426 (2022).
Bonham, L. W. et al. DNA methylation age from peripheral blood predicts progression to Alzheimer’s disease, white matter disease burden, and cortical atrophy. NPJ Dement 1, 7 (2025).
Caviedes, A. et al. Epigenetics of dementia remains unraveled in Latin American and Caribbean populations: A call for collaborative efforts. Alzheimers Dement 20, 9076–9078 (2024).
Cevik, S. E. et al. DNA methylation of imprint control regions associated with Alzheimer’s disease in non-Hispanic Blacks and non-Hispanic Whites. Clin. Epigenetics 16, 58 (2024).
Horvath, S. et al. An epigenetic clock analysis of race/ethnicity, sex, and coronary heart disease. Genome Biol. 17, 171 (2016).
Oh, H. S. et al. Organ aging signatures in the plasma proteome track health and disease. Nature 624, 164–172 (2023).
Nie, C. et al. Distinct biological ages of organs and systems identified from a multi-omics study. Cell Rep. 38, 110459 (2022).
Mulder, D., Aarts, E., Arias Vasquez, A. & Bloemendaal, M. A systematic review exploring the association between the human gut microbiota and brain connectivity in health and disease. Mol. Psychiatry 28, 5037–5061 (2023).
Finch, C. E. & Kulminski, A. M. The alzheimer’s disease exposome. Alzheimers Dement 15, 1123–1132 (2019).
Mendenhall, E., Kohrt, B. A., Logie, C. H. & Tsai, A. C. Syndemics and clinical science. Nat. Med 28, 1359–1362 (2022).
Legaz, A., Baez, S. & Ibanez, A. Unequal burdens: How structural socioeconomic inequality shapes brain health in aging and dementia. Neuroscience 569, 245–247 (2025).
Best Rogowski, C. B. et al. Long-term air pollution exposure and incident dementia: a systematic review and meta-analysis. Lancet Planet Health 9, 101266 (2025).
Fan, H. et al. Linking ambient air pollution to mental health: evidence based on the two-sample Mendelian randomization and colocalization study. Transl. Psychiatry 14, 489 (2024).
Xu, J. et al. Effects of urban living environments on mental health in adults. Nat. Med 29, 1456–1467 (2023).
Noble, K. G. et al. Family income, parental education and brain structure in children and adolescents. Nat. Neurosci. 18, 773–778 (2015).
Rakesh, D., Zalesky, A. & Whittle, S. Assessment of Parent Income and Education, Neighborhood Disadvantage, and Child Brain Structure. JAMA Netw. Open 5, e2226208 (2022).
Ibanez, A. & Farombi, T. Biological imprint of education and rights-based brain capital. Alzheimers Dement 21, e70222 (2025).
Gonzalez-Gomez, R. et al. Qualitative and quantitative educational disparities and brain signatures in healthy aging and dementia across global settings. EClinicalMedicine 82, 103187 (2025).
Chan, S. Y. et al. The influence of early-life adversity on the coupling of structural and functional brain connectivity across childhood. Nat. Ment. Health 2, 52–62 (2024).
Holz, N. E. et al. Early social adversity, altered brain functional connectivity, and mental health. Biol. Psychiatry 93, 430–441 (2023).
Raymond, C. et al. Early childhood adversity and HPA axis activity in adulthood:The importance of considering minimal age at exposure. Psychoneuroendocrinology 124, 105042 (2021).
Furman, D. et al. Chronic inflammation in the etiology of disease across the life span. Nat. Med 25, 1822–1832 (2019).
De Felice, F. G., Goncalves, R. A. & Ferreira, S. T. Impaired insulin signalling and allostatic load in Alzheimer disease. Nat. Rev. Neurosci. 23, 215–230 (2022).
Boyd, R. J., Avramopoulos, D., Jantzie, L. L. & McCallion, A. S. Neuroinflammation represents a common theme amongst genetic and environmental risk factors for Alzheimer and Parkinson diseases. J. Neuroinflammation 19, 223 (2022).
Muscatell, K. A., Brosso, S. N. & Humphreys, K. L. Socioeconomic status and inflammation: a meta-analysis. Mol. Psychiatry 25, 2189–2199 (2020).
Baez, S. et al. Structural inequality and temporal brain dynamics across diverse samples. Clin. Transl. Med 14, e70032 (2024).
Ibanez, A. et al. Healthy aging meta-analyses and scoping review of risk factors across Latin America reveal large heterogeneity and weak predictive models. Nat. Aging 4, 1153–1165 (2024).
Noren Hooten, N., Pacheco, N. L., Smith, J. T. & Evans, M. K. The accelerated aging phenotype: the role of race and social determinants of health on aging. Ageing Res. Rev. 73, 101536 (2022).
Martin, C. L., Ghastine, L., Lodge, E. K., Dhingra, R. & Ward-Caviness, C. K. Understanding health inequalities through the lens of social epigenetics. Annu Rev. Public Health 43, 235–254 (2022).
Amoruso, L. et al. Multilingualism protects against accelerated aging in cross-sectional and longitudinal analyses of 27 European countries. Nat. Aging 5, 2340–2354 (2025).
Coronel-Oliveros, C. et al. Creative experiences and brain clocks. Nat. Commun. 16, 8336 (2025).
Bobba-Alves, N. et al. Cellular allostatic load is linked to increased energy expenditure and accelerated biological aging. Psychoneuroendocrinology 155, 106322 (2023).
Meeker, K. L. et al. Effects of structural and social determinants of health and comorbidities on ethno-racial differences in ATN plasma biomarkers of Alzheimer’s disease: A HABS-HD study. Alzheimer’s. Dement. 19, e078605 (2023).
Asken, B. M. et al. Plasma Alzheimer’s biomarkers and brain amyloid in Hispanic and non-Hispanic older adults. Alzheimers Dement 20, 437–446 (2024).
Bonomi, S. et al. Relationships of cognitive measures with cerebrospinal fluid but not imaging biomarkers of alzheimer disease vary between black and white individuals. Ann. Neurol. 95, 495–506 (2024).
Hoyt, K. R. & Obrietan, K. Circadian clocks, cognition, and Alzheimer’s disease: synaptic mechanisms, signaling effectors, and chronotherapeutics. Mol. Neurodegener. 17, 35 (2022).
Stephan, K. E., Iglesias, S., Heinzle, J. & Diaconescu, A. O. Translational perspectives for computational neuroimaging. Neuron 87, 716–732 (2015).
Lope, D. J., Demirhan, H. & Dolgun, A. Bayesian estimation of the effect of health inequality in disease detection. Int J. Equity Health 21, 118 (2022).
Friston, K. The free-energy principle: a unified brain theory? Nat. Rev. Neurosci. 11, 127–138 (2010).
Ibanez, A. Intellectual cyborgs and the future of science. Trends Cogn. Sci. 27, 785–787 (2023).
Perl, Y. S. et al. Low-dimensional organization of global brain states of reduced consciousness. Cell Rep. 42, 112491 (2023).
Naik, S., Banerjee, A., Bapi, R. S., Deco, G. & Roy, D. Metastability in senescence. Trends Cogn. Sci. 21, 509–521 (2017).
Cruzat, J. et al. Temporal irreversibility of large-scale brain dynamics in alzheimer’s disease. J. Neurosci. 43, 1643–1656 (2023).
Ranasinghe, K. G. et al. Altered excitatory and inhibitory neuronal subpopulation parameters are distinctly associated with tau and amyloid in Alzheimer’s disease. Elife 11, e77850 (2022).
Patow, G., Martin, I., Sanz Perl, Y., Kringelbach, M. L. & Deco, G. Whole-brain modelling: an essential tool for understanding brain dynamics. Nat. Rev. Methods Prim. 4, 53 (2024).
Perl, Y. S. et al. The impact of regional heterogeneity in whole-brain dynamics in the presence of oscillations. Netw. Neurosci. 7, 632–660 (2023).
Amunts, K. et al. BigBrain: an ultrahigh-resolution 3D human brain model. Science 340, 1472–1475 (2013).
Akarca, D., Vertes, P. E., Bullmore, E. T., Team, C. & Astle, D. E. A generative network model of neurodevelopmental diversity in structural brain organization. Nat. Commun. 12, 4216 (2021).
Lam, K. H. B., Faust, K., Yin, R., Fiala, C. & Diamandis, P. The brain protein atlas: a conglomerate of proteomics datasets of human neural tissue. Proteomics 22, e2200127 (2022).
Markello, R. D. et al. Neuromaps: structural and functional interpretation of brain maps. Nat. Methods 19, 1472–1479 (2022).
He, C., Liu, J., Zhu, Y. & Du, W. Data augmentation for deep neural networks model in EEG classification task: a review. Front Hum. Neurosci. 15, 765525 (2021).
Moguilner, S. et al. Visual deep learning of unprocessed neuroimaging characterises dementia subtypes and generalises across non-stereotypic samples. EBioMedicine 90, 104540 (2023).
Kraus, B. et al. Insights from personalized models of brain and behavior for identifying biomarkers in psychiatry. Neurosci. Biobehav Rev. 152, 105259 (2023).
Roe, J. M. et al. Brain change trajectories in healthy adults correlate with Alzheimer’s related genetic variation and memory decline across life. Nat. Commun. 15, 10651 (2024).
Williams, J. P. et al. A Bayesian approach to multistate hidden Markov models: application to dementia progression. J. Am. Stat. Assoc. 115, 16–31 (2020).
Armstrong, T. The myth of the normal brain: embracing neurodiversity. AMA J. Ethics 17, 348–352 (2015).
Bethlehem, R. A. et al. Brain charts for the human lifespan. Nature 604, 525–533 (2022).
Schor, N. F. & Bianchi, D. W. Neurodevelopmental clues to neurodegeneration. Pediatr. Neurol. 123, 67–76 (2021).
Shabani, K. & Hassan, B. A. The brain on time: links between development and neurodegeneration. Development 150, dev200397 (2023).
Tozzi, L. et al. Personalized brain circuit scores identify clinically distinct biotypes in depression and anxiety. Nat. Med. 30, 2076–2087 (2024).
Rutherford, S. et al. The normative modeling framework for computational psychiatry. Nat. Protoc. 17, 1711–1734 (2022).
Scheffer, M. et al. A dynamical systems view of psychiatric disorders-theory: a review. JAMA Psychiatry 81, 618–623 (2024).
Jones, D. et al. A computational model of neurodegeneration in Alzheimer’s disease. Nat. Commun. 13, 1643 (2022).
Krakauer, J. W., Ghazanfar, A. A., Gomez-Marin, A., MacIver, M. A. & Poeppel, D. Neuroscience Needs Behavior: Correcting a Reductionist Bias. Neuron 93, 480–490 (2017).
Luppi, A. I. et al. A synergistic core for human brain evolution and cognition. Nat. Neurosci. 25, 771–782 (2022).
Luppi, A. I. et al. Dynamical models to evaluate structure-function relationships in network neuroscience. Nat. Rev. Neurosci. 23, 767–768 (2022).
Vohryzek, J., Cabral, J., Vuust, P., Deco, G. & Kringelbach, M. L. Understanding brain states across spacetime informed by whole-brain modelling. Philos. Trans. A Math. Phys. Eng. Sci. 380, 20210247 (2022).
Prattichizzo, F. et al. Organ-specific biological clocks: ageotyping for personalized anti-aging medicine. Ageing Res Rev. 96, 102253 (2024).
O’Connor, D. B., Thayer, J. F. & Vedhara, K. Stress and health: a review of psychobiological processes. Annu Rev. Psychol. 72, 663–688 (2021).
Kuhn, M. et al. Mismatch or allostatic load? Timing of life adversity differentially shapes gray matter volume and anxious temperament. Soc. Cogn. Affect Neurosci. 11, 537–547 (2016).
Rodriquez, E. J., Kim, E. N., Sumner, A. E., Napoles, A. M. & Perez-Stable, E. J. Allostatic load: importance, markers, and score determination in minority and disparity populations. J. Urban Health 96, 3–11 (2019).
Juster, R. P., McEwen, B. S. & Lupien, S. J. Allostatic load biomarkers of chronic stress and impact on health and cognition. Neurosci. Biobehav Rev. 35, 2–16 (2010).
Thomson, E. M. et al. Air pollution, stress, and allostatic load: linking systemic and central nervous system impacts. J. Alzheimers Dis. 69, 597–614 (2019).
Hastings, P., Johnson, L., Ugarte, E., Robins, R. & Guyer, A. The biological embedding of adversity and inequity: Examining the physiological impacts of discrimination and poverty on ethnically and economically minoritized youths. Psychoneuroendocrinology 153, 106261 (2023).
Kumsta, R. et al. The role of stress in the biological embedding of experience. Psychoneuroendocrinology 156, 106364 (2023).
Erickson, A. C. & Sbihi, H. Biological embedding, the air we breathe, and carcinogenesis. Lancet Planet Health 2, e149–e150 (2018).
Eckstrand, K. L., Singh, M. K. & Ajilore, O. Diversity, equity, and inclusivity in biological psychiatry research. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 7, 1195–1197 (2022).
Zurn, P., Stramondo, J., Reynolds, J. M. & Bassett, D. S. Expanding diversity, equity, and inclusion to disability: opportunities for biological psychiatry. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 7, 1280–1288 (2022).
Al-Mubarak, B. et al. Whole exome sequencing reveals inherited and de novo variants in autism spectrum disorder: a trio study from Saudi families. Sci. Rep. 7, 5679 (2017).
Kourtian, S. et al. Candidate genes for inherited autism susceptibility in the Lebanese population. Sci. Rep. 7, 45336 (2017).
Gupta, V. et al. Genetic Variant Analyses Identify Novel Candidate Autism Risk Genes from a Highly Consanguineous Cohort of 104 Families from Oman. Int J. Mol. Sci. 25, 13700 (2024).
Soueid, J. et al. RYR2, PTDSS1 and AREG genes are implicated in a Lebanese population-based study of copy number variation in autism. Sci. Rep. 6, 19088 (2016).
Lencz, T. et al. Novel ultra-rare exonic variants identified in a founder population implicate cadherins in schizophrenia. Neuron 109, 1465–1478e1464 (2021).
Mir, A., Khorram, E., Song, Y., Lee, H. & Tabatabaiefar, M. A. A novel heterozygous truncating variant in the AGO1 gene in an Iranian family with schizophrenia as an unreported symptom. Ann. Hum. Genet 87, 295–301 (2023).
Colak-Genis, E. et al. Investigation of genetic changes in three families with bipolar disease. Mol. Syndromol. 15, 464–473 (2024).
Fullerton, J. M. & Tesfaye, M. Global diversity in bipolar disorder: the role of cultural and social differences with a view to genomics. Biol. Psychiatry 98, 354–364 (2025).
Pries, L. K. et al. Predictive performance of exposome score for schizophrenia in the general population. Schizophr. Bull. 47, 277–283 (2021).
Chiao, J. Y. & Blizinsky, K. D. Population disparities in mental health: insights from cultural neuroscience. Am. J. Public Health 103, S122–S132 (2013).
Li, X. et al. The global burden of schizophrenia and the impact of urbanization during 1990-2019: An analysis of the global burden of disease study 2019. Environ. Res 232, 116305 (2023).
McFadyen, J. & Dolan, R. J. Spatiotemporal precision of neuroimaging in psychiatry. Biol. Psychiatry 93, 671–680 (2023).
Hauser, T. U., Skvortsova, V., De Choudhury, M. & Koutsouleris, N. The promise of a model-based psychiatry: building computational models of mental ill health. Lancet Digit Health 4, e816–e828 (2022).
Halassa, M. M. et al. Developing algorithmic psychiatry via multi-level spanning computational models. Cell Rep. Med 6, 102094 (2025).
Bruckl, T. M. et al. The biological classification of mental disorders (BeCOME) study: a protocol for an observational deep-phenotyping study for the identification of biological subtypes. BMC Psychiatry 20, 213 (2020).
Wingo, T. S. et al. Shared mechanisms across the major psychiatric and neurodegenerative diseases. Nat. Commun. 13, 4314 (2022).
Smeland, O. B. et al. Genome-wide association analysis of parkinson’s disease and schizophrenia reveals shared genetic architecture and identifies novel risk loci. Biol. Psychiatry 89, 227–235 (2021).
Favela, L. H. & Amon, M. J. Reframing cognitive science as a complexity science. Cogn. Sci. 47, e13280 (2023).
Hampel, H. et al. The foundation and architecture of precision medicine in neurology and psychiatry. Trends Neurosci. 46, 176–198 (2023).
Walhovd, K. B., Lovden, M. & Fjell, A. M. Timing of lifespan influences on brain and cognition. Trends Cogn. Sci. 27, 901–915 (2023).
Xu, J. et al. Global urbanicity is associated with brain and behaviour in young people. Nat. Hum. Behav. 6, 279–293 (2022).
Danielsdottir, H. B. et al. Adverse childhood experiences and adult mental health outcomes. JAMA Psychiatry 81, 586–594 (2024).
Short, A. K. & Baram, T. Z. Early-life adversity and neurological disease: age-old questions and novel answers. Nat. Rev. Neurol. 15, 657–669 (2019).
Singhaarachchi, P. H. et al. Aging, sex, metabolic and life experience factors: contributions to neuro-inflammaging in Alzheimer’s disease research. Neurosci. Biobehav Rev. 162, 105724 (2024).
Kumar, S., Payne, P. R. O. & Sotiras, A. Normative Modeling using Multimodal Variational Autoencoders to Identify Abnormal Brain Volume Deviations in Alzheimer’s Disease. Proc SPIE Int Soc Opt Eng 12465, 1246503 (2023).
Zsadanyi, S. E. et al. Democratizing dementia research through global online conferences. Nat. Neurosci. 28, 1109–1112 (2025).
Ibanez, A. & Eyre, H. Brain capital, ecological development and sustainable environments. BMJ Ment. Health 26, e300803 (2023).
Cardona, J. F., Trujillo-Llano, C., Miller, J. & Ibanez, A. Peace in Colombia requires healing the scars of conflict on the mind and brain. Nat. Med 31, 2845–2849 (2025).
Cardona, J. F., Trujillo-Llano, C., Miller, J. & Ibanez, A. Address Colombia’s brain-health crisis. Nature 643, 336 (2025).
Alderman, J. E. et al. Tackling algorithmic bias and promoting transparency in health datasets: the STANDING Together consensus recommendations. Lancet Digit Health 7, e64–e88 (2025).
Lim, S. S. et al. Reporting guidelines for precision medicine research of clinical relevance: the BePRECISE checklist. Nat. Med 30, 1874–1881 (2024).
Maestre, G. et al. Promoting diverse perspectives: Addressing health disparities related to Alzheimer’s and all dementias. Alzheimers Dement 20, 3099–3107 (2024).
Rebbeck, T. R. et al. A Framework for Promoting Diversity, Equity, and Inclusion in Genetics and Genomics Research. JAMA Health Forum 3, e220603–e220603 (2022).
Diversity, equity and inclusion in neurology. Nat Rev Neurol 20, 199 (2024).
Fonseca, L. et al. Diversity matters: opportunities in the study of the genetics of psychotic disorders in low- and middle-income countries in Latin America. Braz. J. Psychiatry 43, 631–637 (2021).
Celi, L. A. et al. Sources of bias in artificial intelligence that perpetuate healthcare disparities-A global review. PLOS Digit Health 1, e0000022 (2022).
Yang, Y., Zhang, H., Gichoya, J. W., Katabi, D. & Ghassemi, M. The limits of fair medical imaging AI in real-world generalization. Nat. Med 30, 2838–2848 (2024).
The Lancet Digital, H A long STANDING commitment to improving health care. Lancet Digit Health 7, e1 (2025).
Stites, S. D. & Coe, N. B. Let’s not repeat history’s mistakes: two cautions to scientists on the use of race in alzheimer’s disease and alzheimer’s disease related dementias research. J. Alzheimers Dis. 92, 729–740 (2023).
Slade, M., Rennick-Egglestone, S. & Todowede, O. The transformative potential of citizen science for mental health. Lancet Psychiatry 11, 246–248 (2024).
Haklay, M. et al. Contours of citizen science: a vignette study. R. Soc. Open Sci. 8, 202108 (2021).
Schumann, G. et al. AddressIng Global Environmental Challenges To Mental Health Using Population Neuroscience: A Review. JAMA Psychiatry 80, 1066–1074 (2023).
Coronel-Oliveros, C. et al. Diversity-sensitive brain clocks linked to biophysical mechanisms in aging and dementia. Nat. Ment. Health 3, 1214–1229 (2025).
Eo, J., Kang, J., Youn, T. & Park, H. J. Neuropharmacological computational analysis of longitudinal electroencephalograms in clozapine-treated patients with schizophrenia using hierarchical dynamic causal modeling. Neuroimage 275, 120161 (2023).
Petzschner, F. H., Garfinkel, S. N., Paulus, M. P., Koch, C. & Khalsa, S. S. Computational Models of Interoception and Body Regulation. Trends Neurosci. 44, 63–76 (2021).
Guo, W., Wang, J. & Wang, S. Deep multimodal representation learning: A survey. Ieee Access 7, 63373–63394 (2019).
Septier, F. & Peters, G. W. An Overview Of Recent Advances In Monte-carlo Methods For Bayesian Filtering In High-dimensional Spaces. Theoretical aspects of spatial-temporal modeling, 31-61 (2015).
Hipolito, I. et al. Markov blankets in the brain. Neurosci. Biobehav Rev. 125, 88–97 (2021).
Langdon, C., Genkin, M. & Engel, T. A. A unifying perspective on neural manifolds and circuits for cognition. Nat. Rev. Neurosci. 24, 363–377 (2023).
D’Angelo, E. & Jirsa, V. The quest for multiscale brain modeling. Trends Neurosci. 45, 777–790 (2022).
Wilson, D. M. et al. Hallmarks of neurodegenerative diseases. Cell 186, 693–714 (2023).
Frisoni, G. B. et al. The probabilistic model of Alzheimer disease: the amyloid hypothesis revised. Nat. Rev. Neurosci. 23, 53–66 (2022).
Acknowledgements
A.I. is funded by ANID/FONDECYT Regular (1250091, 1210176, 1220995); ANID/FONDAP 15150012; ANID/PIA/ANILLOS ACT210096; ANID/FONDAP 15150012; Takeda CW2680521 and the Multi-Partner Consortium To Expand Dementia Research In Latin America [ReDLat, supported by Fogarty International Center (FIC) and National Institutes of Health, National Institutes of Aging (R01 AG057234, R01 AG075775, R01 AG21051, R01 AG083799, CARDS-NIH), Alzheimer’s Association (SG-20-725707), Rainwater Charitable foundation – Tau Consortium, the Bluefield Project to Cure Frontotemporal Dementia, and Global Brain Health Institute)]. CD-A is supported by ANID/FONDECYT Regular 1210622, ANID/PIA/ANILLOS ACT210096, Alzheimer’s Association (AARGD-24- 1310017) and ANID/FOVI240065. C.Y. is funded by the National Natural Science Foundation of China (Grant No. 82430063, 82030053). J.X is funded by the National Natural Science Foundation of China (Grant No. 82371924), the National Key Project of “Inter-governmental International Scientific and Technological Innovation Cooperation” (Grant No. 2023YFE0199700), the Tianjin Young Talents in Science and Technology (Grant No. QN20230336), the Tianjin Applied Basic Research Diversified Investment Foundation (Grant No. 21JCYBJC01360), and the Tianjin Medical University “Clinical Talent Training 123 Climbing Plan”. K.F. is funded by the Wellcome Trust (Ref: 205103/Z/16/Z). J.L.M.O. is supported by NIDA via R21DA050160 and DP1DA058737, the U.S. Department of Veterans Affairs via the National Center for Posttraumatic Stress Disorder and 1IK2CX002095-01A1, and the Yale Center for Brain and Mind Health. C.U receives funding support from The UKRI Medical Research Council (MR/Y019822/1), the Alzheimer’s Association (SAGA23-1141999), the Wellcome Leap and Temasek Trust Dynamic Resilience Award, the National Institute of Health (NIH) (RO1-AG074562), the Office for Veterans’ Affairs UK Defense and Security Accelerator (DASA) Fund (G2-SCH-2022-11-12245), the Global Brain Health Institute (UFRA-424 | CA-0241758), the Davos Alzheimer’s Collaborative Global Cohorts Fund, and the RoseTrees Foundation (Seedcorn2021\100220). C.S. is supported by the Swiss National Science Foundation (Grant No. 185897) and Synapsy. S.F. is funded by ZonMW (#73305095007 and #10510032120004), Health Holland, Topsector Life Sciences & Health (the Netherlands) (PPP-allowance; #LSHM20106). H.Z. is supported by the Swedish Research Council (#2023-00356, #2022-01018, and #2019-02397), the European Union’s Horizon Europe research and innovation program (Grant No. 101053962), Swedish State Support for Clinical Research (#ALFGBG-71320), and is a Wallenberg Scholar and Distinguished Professor at the Swedish Research Council. LTG is partially funded by K24AG053435. C.U. is additionally supported by The Wellcome Leap Dynamic Resilience Program (co-funded by Temasek Trust). The Geroscience Center for Brain Health and Metabolism is funded by FONDAP-15150012, FONDECYT-1150766, Michael J. Fox Foundation for Parkinson’s Research (17303), and FB210008. H.S-G is funded by NIH D43 Fogarty (Research Training for Equity in Alzheimer’s Disease and Brain Health in Colombia, number: 1D43TW012455-01A1), NIH R01 (Social epigenetics of Alzheimer’s disease and related dementias in Latin American countries, number: 1R01AG082056-01A1), Global Brain Health Institute and Alzheimer Association (“Brain health in individuals with exposition to high violence in Colombia”, number: GBHI ALZ UK-23-971135”). The contents of this publication are solely the responsibility of the authors and do not represent the official views of these institutions. The funders had no role in study design, data collection, analysis, publication decision, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
AI conceived the proposal. AI and HSG wrote the first draft and supervised and coordinated the work. J.M. prepared the figures under the supervision of AI and HSG. S.B., S.F., C.C.O., H.A.E., B.L., C.Y., P.S.S., V.H., K.F., and E.T. coordinated specific manuscript sections. A.I., C.D.A., J.M., S.B., S.F., C.C.O., H.A.E., C.U.M., H.Z., S.A., C.S., I.H.R., S.Fz., T.F., J.L.M.O., S.S., F.C., P.V.S., J.X., C.Y., L.G., B.L., P.S.S., K.Y., V.H., K.F., E.T., and H.S.G. reviewed, edited, provided critical comments, and approved the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
HZ has served at scientific advisory boards and/or as a consultant for Abbvie, Acumen, Alector, Alzinova, ALZpath, Amylyx, Annexon, Apellis, Artery Therapeutics, AZTherapies, Cognito Therapeutics, CogRx, Denali, Eisai, LabCorp, Merry Life, Nervgen, Novo Nordisk, Optoceutics, Passage Bio, Pinteon Therapeutics, Prothena, Quanterix, Red Abbey Labs, reMYND, Roche, Samumed, Siemens Healthineers, Triplet Therapeutics, and Wave, has given lectures sponsored by Alzecure, BioArctic, Biogen, Cellectricon, Fujirebio, Lilly, Novo Nordisk, Roche, and WebMD, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program. CS is a member of the scientific advisory board of Amazentis and Vandria and has given lectures sponsored by Schwabe Pharma. SF has received consulting fees from Biogen (2022). The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Muralidharan Sargurupremraj and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ibáñez, A., Duran-Aniotz, C., Migeot, J. et al. Computational whole-body-exposome models for global precision brain health. Nat Commun 16, 11078 (2025). https://doi.org/10.1038/s41467-025-67448-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67448-3
This article is cited by
-
Randomized controlled trial of resistance exercise and brain aging clocks
GeroScience (2026)