Abstract
The phase 2 KNOCKOUT (NCT05283135) study evaluates higher-than-approved doses of risankizumab, an interleukin-23 inhibitor, for treatment of moderate-to-severe plaque psoriasis. Patients received either 300 or 600 mg of risankizumab at Weeks 0, 4, and 16 and were monitored for 100 weeks without further dosing. Efficacy and safety were tracked throughout the study and lesional/non-lesional tissue was evaluated by RNASeq. The primary endpoint was the change in baseline in number and/or function of tissue resident memory T cells (TRM) at week 52. The secondary endpoints were Psoriasis Areas and Severity Index (PASI) 100 at weeks 28, 40, and 52 and safety events over 100 weeks. Nine patients per treatment group completed dosing. At Weeks 28 and 52, PASI 75/90/100 responses were: 94.4%/94.4%/83.3% and 77.8%/61.1%/44.4% of all patients, respectively, with no new safety signals. At Week 52, TRM cell numbers in lesional skin were markedly reduced, with numbers similar to non-lesional TRM numbers at Week 0. High skin clearance rates with higher-than-approved initial dosing of risankizumab with prolonged maintenance of skin clearance, despite the lack of continuous dosing, represent a potential alternative treatment strategy for patients with psoriasis.
Similar content being viewed by others
Introduction
Psoriasis is a common, chronic, immune-mediated inflammatory disease. Many highly efficacious medications now exist for treating patients with psoriasis1, but disease eventually recurs following withdrawal of these treatments2. Risankizumab, an anti-interleukin (IL)−23 biologic, is highly effective and safe for treating moderate-to-severe plaque psoriasis. Approximately 80% of patients achieve greater than 90% clearance of skin lesions, and approximately 60% reach complete skin clearance at 1 year, with few-to-no side effects3,4,5,6. In addition, the IMMhance study revealed that some patients undergoing risankizumab withdrawal demonstrate prolonged skin clearance long after their last dose; more specifically, approximately 10% of patients with completely clear skin after only three risankizumab 150 mg doses (at weeks 0, 4, and 16) maintained completely clear skin for up to 1 year before psoriasis recurred6. The phase 1 study for risankizumab in patients with psoriasis also showed long-term clearance of lesions after a single high dose of drug7. The combination of these prior clinical results suggests there is a need to scientifically explore the biologic basis of the duration of risankizumab’s therapeutic activity; specifically, there is a need to test whether high induction doses can result in more complete skin clearance and for longer periods of time.
Risankizumab is postulated to induce high levels of prolonged skin clearance due to an effect on resident memory T (TRM) cells8. TRM cells develop within tissues in response to pathogen exposure, are long-lived, and are responsible for tissue protection from re-exposure to the original pathogen9,10,11. Interestingly, TRM cells are also found in skin previously affected by active psoriasis, and are now believed to be the cells responsible for recurrences of psoriasis12,13,14. Recent evidence also suggests that psoriatic TRM cells are under the control of IL-2315.
Here, a pilot phase 2 study, KNOCKOUT, describes higher-than-approved induction doses of risankizumab, aiming to induce high levels of complete skin clearance and long-term remissions. Prolonged remission of psoriasis is also correlated with knockdown of TRM cells within tissue.
Results
Baseline demographics/disease characteristics and patient disposition
A total of 20 patients were enrolled in the study, with 10 randomized to receive 300 mg and 10 to receive 600 mg of risankizumab. Patient demographics and baseline characteristics are presented in Table 1. Of note, patients randomized to the 600 mg group, compared to the 300 mg group, had numerically higher weights (88.4 ± 20.7 kg vs. 86.7 ± 13.7 kg, mean ± SD), had numerically longer disease duration (26.0 ± 18.1 years vs. 16.9 ± 15.1 years, mean ± SD), and had numerically higher static Physician’s Global Assessment (sPGA) scores (6 of 10 with sPGA scores of 4, indicating severe psoriasis, vs. 1 of 10 with an sPGA score of 4).
Of the 20 enrolled patients, 9 patients in the 300 mg treatment group and 9 patients in the 600 mg treatment group completed all three risankizumab doses at weeks 0, 4, and 16. In the 300 mg treatment group, one patient received two doses but was lost to follow-up and was not considered in the efficacy analysis. Of the remaining 9 patients, 4 withdrew consent prior to week 52 to pursue other treatment options, 2 withdrew consent after week 52 to pursue other treatment options, and 3 completed the study. In the 600 mg treatment group, one patient received only 1 dose, was lost to follow-up, and was not considered in the efficacy analysis. Of the remaining 9 patients, 1 withdrew consent before week 52 due to relocation, 1 withdrew consent to seek other treatment options before week 52, 4 withdrew consent to pursue other treatment options after week 52, and 3 completed the study.
Of the 18 patients who completed dosing, 16 completed the week 52 visit. Thirteen of these 16 patients underwent week 52 lesional skin biopsies and had evaluable RNAseq data for analysis. No patients withdrew from the study due to an adverse event.
Efficacy
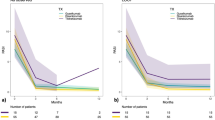
At week 16 (following two initial risankizumab doses at weeks 0 and 4), 100% (n = 18; 95% CI [81.5, 100.0]) of all patients achieved psoriasis area and severity index (PASI) 75, 94.4% (n = 17; 95% CI [72.7, 99.9]) achieved PASI 90, and 66.7% (n = 12; 95% CI [41.0, 86.7]) achieved PASI 100 (Fig. 1). At week 28 (following a third risankizumab dose at week 16), PASI 75/90/100 efficacy rates were 94.4% (n = 17, 95% CI [72.7, 99.9]), 94.4% (n = 17, 95% CI [72.7. 99.9]), and 83.3% (n = 17, 95% CI [58.6, 96.4]) (Fig. 1a–c). The 300 mg group generally had numerically slightly higher efficacy response rates when compared to the 600 mg dose group at all timepoints, which might be attributed to these patients having less severe baseline disease (Fig. 1c and Table 1). Treatment also had a profound effect on improving patient quality of life, as measured by the proportion of patients achieving a score 0 or 1 on the Dermatology Life Quality Index (DLQI); DLQI 0/1 was achieved by 88.9% (n = 16, 95% CI [65.3, 98.6]) of all patients at week 28 (Fig. 1d), indicating no skin disease effect on quality of life.
Open triangles below numbers represent doses of risankizumab. a PASI 75 responses. b PASI 90 responses. c PASI 100 responses. d DLQI 0/1 responses at week 28. Data are presented as mean ± 95% confidence interval.
Other measures of efficacy, including proportions of patients achieving sPGA scores of 0/1 (clear or almost clear skin) and 0 (clear skin), mean percent improvement in PASI, and mean absolute PASI, also demonstrated marked improvements in all patients (Fig. 2). At week 16, 94.4% (n = 17; 95% CI [72.7, 99.9]) of all patients achieved sPGA 0/1 (Fig. 2a) and 66.7% (n = 12; 95% CI [41.0, 86.7]) achieved sPGA 0 (Fig. 2b). Percent change in PASI improved by an average of 97.5% (94.8, 100) at week 16 and by 97.9% (94.3%, 100%) at week 28 (Fig. 2c). Mean absolute PASI dropped from 19.1 at baseline to 0.8 (0, 10) at week 16 and 0.8 (0, 14) at week 28 (Fig. 2d).
Open triangles below numbers represent doses of risankizumab. a PGA 0/1 responses. b PGA 0 responses. c Mean PASI improvement. d Mean PASI. Data are presented as mean ± 95% confidence interval.
Over the course of the second part of the study (weeks 28–100), when patients were only being assessed for efficacy and safety, they did not receive active treatment. High levels of efficacy were maintained by the majority of patients following drug withdrawal, 36 weeks following the last risankizumab dose. At week 52, 77.8% (52.4, 93.6) of patients achieved PASI 75, 61.1% (35.7, 82.7) achieved PASI 90, and 44.4% achieved PASI 100 (21.5, 69.2) (Fig. 3a, b). sPGA 0/1 and sPGA 0 responses were achieved by 61.1% (n = 11; 95% CI [35.7, 82.7]) and 44.4% (n = 8; [95% CI 21.5, 69.2]) of all patients, respectively, at week 52. Mean percent improvement from baseline PASI was 86.8% (Fig. 3c), and mean absolute PASI was 3.3 at this timepoint (as observed).
a PASI 90 responses. b PASI 100 responses. c PGA 0/1 responses. d PGA 0 responses. e Mean PASI improvement. f Mean PASI. Data are presented as mean ± 95% confidence interval, where error bars are reported.
Two of the six patients who completed the study (i.e., completed the week 100 visit) demonstrated completely clear skin (PASI 100 of 11.1%; n = 2; [95% CI 1.4, 34.7]) at week 100, 84 weeks following their last dose of risankizumab. One of these patients was a White man, had a disease duration of 11 months, weighed 103.3 kg, was naïve to biologic treatment, had a baseline PASI of 12.6, and was in the 300 mg dose group. The second patient was an African American man, had a disease duration of 39 years, weighed 103.0 kg, had received prior secukinumab treatment, had a baseline PASI of 21.2, and was in the 600 mg dose group. PASI 75 and 90 rates at week 100 were 27.8% and 22.2%, respectively, in the combined dose group. Mean percent improvement from baseline PASI was 62.6% ([47.8, 77.3]; Fig. 3c) and mean absolute PASI was 7.7 ([0, 21]; Fig. 3d) for the remaining 6 patients at week 100 (as observed). Patient numbers were too small, and baseline disease characteristics too unbalanced, to make definitive conclusions regarding remission times in the 300 mg group when compared to the 600 mg group; visualization of individual patient response curves provided some insight into this analysis (Supplementary Fig. 1).
Safety
High induction doses of risankizumab were well-tolerated, and no new safety signals were observed (Table 2). A total of 22 treatment-emergent adverse events (TEAEs) were reported through week 52, with none related to the study treatment according to the investigator (Table 2). One serious adverse event (SAE) occurred and was considered severe and unrelated to study treatment; this patient experienced a right tibial fracture and meniscal tear while playing soccer. No TEAEs lead to discontinuation, dose reduction, or interruptions in treatment (Table 2). Examination of routine laboratory parameters at weeks 4, 8, 22, and 46 revealed no significant abnormalities or trends. No major safety issues occurred between weeks 52 and 100.
High induction doses of risankizumab downregulate CD8+ TRM17 cell number and function
To investigate the mechanisms underlying long-term clinical remissions observed with high induction doses of risankizumab, longitudinal single-cell RNA-sequencing of full-thickness skin biopsy samples was performed at Baseline (lesional and non-lesional) and at week 52 (lesional) from both 300 and 600 mg dose groups. The dataset included >100,000 cells. First, total T cell populations were analyzed before and after treatment. Uniform Manifold Approximation Projection (UMAP) was used for T cell subtype visualization; T cells were classified into 11 distinct sub-clusters (Fig. 4a and Supplementary Fig. 2a). Baseline lesional skin had increased proportions of CD4+ naïve and cytotoxic T cells, central memory T cells, CD8+ cytotoxic T cells, and TRM cells, and decreased numbers of T effector memory cells in comparison to baseline nonlesional skin (Supplementary Fig. 2). At week 52, T cell populations in lesional skin had returned close to levels observed in baseline non-lesional skin (Supplementary Fig. 2).
a UMAP plot showing T cells colored by subtypes. b UMAP plots showing the expression levels of CD8A, CD4, ITGAE, CD69, IL17A, IL22, IFNG, and CCR6 in the T cell subtypes. c Plot showing the cell abundance in CD8 + TRM1 and CD8 + TRM17 following treatment with 300 or 600 mg of risankizumab. Every dot represents a patient. Lesional baseline (n = 13), non-lesional baseline (n = 12), 300 mg (n = 8) and 600 mg of risankizumab (n = 5) (one-way ANOVA). d Dot plot showing IL17A, IL17F, and IL22 expression comparing post-treatment (300 and 600 mg) to baseline in the CD8 + TRM17. e Violin plot showing the tissue-resident module score in CD8+ T cell subtypes. In the box plot, the centerlines denote median values (50th percentile); the bounds of the boxes represent the first and third quartiles (25th and 75th percentiles); the whiskers indicate the smallest and largest values within 1.5× the interquartile range. f Pseudotime trajectory colored by the pseudotime. g Pseudotime trajectory colored by three CD8+ T cell subtypes. h Expression of transcriptional signature plotted over pseudotime. i Pseudotime trajectory colored by three CD8+ T cell subtypes, split into different groups.
Given the role of tissue TRM cells in psoriasis recurrence12, IL-23 inhibition was hypothesized to affect TRM cells in psoriasis lesional skin. Of all T cell clusters, IFNγ+/CD8+/CD69+ T cells (defined as CD8 TRM1) and IL-17A+/CD8+/ ITGAE+ T cells (defined as CD8 TRM17) predominantly account for tissue-specific immune memory and thus were assessed (Fig. 4b). In association with durable clinical improvement of skin lesions, total number of CD8 TRM17 was remarkably reduced following high induction doses of risankizumab, particularly in the 600 mg dose group (mean of 3 cells in each sample), compared with lesional baseline group (mean of 22 cells in each sample) (p = 0.035) (Fig. 4c). Downregulation of IL-17 signaling genes (IL17A, IL17F, and IL22) was also observed in CD8 TRM17 after anti-IL-23 treatment (Fig. 4d). Decreased numbers of CD8 TRM1 cells were documented as well (more so in the 600 mg group, with a mean of 2 cells in each sample), although these decreases did not reach statistical significance (Fig. 4c and Supplementary Fig. 2b). Furthermore, in CD8 TRM1 cells, risankizumab treatment at week 52 was associated with lower expression of classic TRM1 markers, including IFNG, RUNX3, FOXO1, EOMES, SLAMF7, and ITGA116 (Supplementary Fig. 4b). Pathway enrichment analysis of differentially expressed genes also showed downregulation of T cell differentiation function in CD8 TRM1 cells (Supplementary Fig. 4c). Among related T cell subtypes, in line with data shown for CD8 TRM17 cells and CD8 TRM1 cells, high induction doses of risankizumab also substantially reduced total numbers and proportions of CCR6+/IL22+ T cells (annotated as T17 cells) (Supplementary Fig. 3).
To better understand the retention and differentiation process of CD8+ TRM cells, pseudotime analysis was performed on three subtypes using Monocle17, to create a continuous trajectory where CD8+ T cells were ordered from CD8+/GZMA+/CST7+ (defined as CD8 T cells) to CD8 TRM1/CD8 TRM17 cells (Fig. 4f, g). To further validate this development path, the pseudotime trajectory of cell lineages of global T cells was examined using Slingshot18. This analysis revealed four distinct lineages originating from the CD8_Tc subset, separately progressing toward the CD8 TRM1 (through lineage 1) or CD8 TRM17 states (through lineage 4) (Supplementary Fig. 4d). These results support the conclusion that CD8 TRM1/CD8 TRM17 represent distinct differentiated states derived from CD8+ T cells (Supplementary Fig. 4e). The TRM module score, composed of conserved genes associated with tissue-resident memory T cell features16,18, was consistently elevated in CD8_TRM1 and CD8_TRM17 cells (Fig. 4e), indicating a more differentiated state. This finding supports the validity of the pseudotime trajectory.
To identify potential cytokines and transcription factors that drive CD8+ TRM differentiation, branched expression analysis modeling (BEAM) was performed. The type 17 immune response signature (IL17A, IL22, and RORC) was highly enriched in branch 2 (i.e., in CD8 TRM17 cells). By contrast, the core transcriptional signature of type 1 immune responses (IFNG, TBX21) was prominently enriched at the end of branch 1 (i.e., in CD8 TRM1 cells). Expression of GZMH gradually decreased in both branches during cell differentiation, which is a marker for CD8+ TRM cells with relatively poor cytotoxicity seen in psoriasis lesions (Fig. 4h). Remarkably, this trajectory pattern mirrored clinical remissions observed in patients, where less CD8 TRM17 cells aggregated at the end of branch 2 in week 52 post-treatment tissue (Fig. 4i). Lastly, network analysis using CellChat19 revealed significantly reduced signals released by both CD8 TRM17 cells and CD8 TRM1 cells in both post-treatment groups (300 and 600 mg), but particularly in the 600 mg dose group (Supplementary Fig. 5a, b). Notably, greater crosstalk between Treg and CD8 TRM17 cells was documented in the 600 mg group (Supplementary Fig. 5c), indicating a potential mechanism by which IL-23 inhibition may dampen immune responses and delay psoriasis recurrences.
High induction doses of risankizumab normalize inflammatory gene expression in keratinocytes
CD8+ TRM17 and CD8+ TRM1 cells are abundant in human epidermis20,21. To better characterize the epidermal environment surrounding these cells, keratinocytes were sub-clustered and annotated by differentiation state. This analysis defined four distinct keratinocyte differentiation states, including basal (KRT5, KRT14), cycling (MKI67, TOP2A), differentiated (KRT1, KRT10), and terminally differentiated (IVL, KRT2) keratinocytes (Fig. 5a, b). Differential expression analysis was then performed between Baseline and week 52 samples. Expression of S100A7, S100A8, and S100A9 was markedly decreased following risankizumab treatment, especially in the basal keratinocytes (Fig. 5c); moreover, pathway enrichment on differentially expressed genes (DEGs) implicated several major inflammatory genes dominant in these cells (Fig. 5d). Interestingly, IL-23 blockade markedly reduced major histocompatibility complex-1 (MHC-I) antigen presentation genes, including HLA-A, HLA-B, HLA-C, and B2M, mostly localized to basal keratinocytes (Fig. 5c).
a UMAP plot showing keratinocytes colored by subtypes. b Dot plot showing top marker genes for each keratinocyte subtype. c Dot plot showing the top differentially expressed genes comparing L_P (lesional posttreatment, 300 and 600 mg) to L_B (lesional baseline) in the basal and differentiated keratinocytes. d Bar plot showing the top 20 KEGG pathways enriched for the down-regulated genes in basal keratinocytes comparing L_P to L_B. KEGG pathway enrichment was performed using a hypergeometric test, and p values were adjusted for multiple testing using the Benjamini–Hochberg method. e UMAP plot showing basal keratinocytes colored by sub-clusters. f UMAP plot showing the basal keratinocytes colored by different groups. g Bar plot showing the abundance composition across the three groups for each sub-cluster of basal keratinocytes. h Box plot showing the MHC-I antigen presentation module score in the basal keratinocyte sub-clusters. In the box plot, the centerlines denote median values (50th percentile); the bounds of the boxes represent the first and third quartiles (25th and 75th percentiles); the whiskers indicate the smallest and largest values within 1.5× the interquartile range. i Violin plot showing gene expression in the basal keratinocyte sub-clusters, each sub-cluster split by sample group. j Circle plot showing ligand receptor relationships among basal keratinocytes, differentiated keratinocytes, CD8 + TRM1, and CD8 + TRM17.
Within basal keratinocytes, distinct separation of three groups suggested major shifts in keratinocyte function (Fig. 5f). There were four sub-clusters of basal keratinocytes primarily observed in Baseline lesion tissue: sub-cluster 5, 11, 12, and 13 (Fig. 5e, g). Notably, these four sub-clusters exhibited higher markers of MHC-I antigen presentation function (Fig. 5h, i). Given that the MHC-TCR combination triggers intracellular signaling cascades in T cells, resulting in effective immunity22, including in memory T cells22, we examined the ligand–receptor interaction among the two CD8+ TRM subtypes with basal keratinocytes. Indeed, basal keratinocytes were shown to interact with both CD8 TRM17 and CD8 TRM1 cells in psoriatic lesions through MHC/TCR/CD3 complexes (Fig. 5j). Conversely, the reduced expression of MHC-I molecules on basal keratinocytes was documented following risankizumab treatment, likely contributing to the diminished persistence of CD8+ TRM cells in the epidermis (Fig. 5i).
High induction doses of risankizumab disrupt persistence of CD8+ TRM cells in tissue through interference of IL-7 and IL-15 signaling
Given the observed shifts in CD8+ TRM cell and keratinocyte number/function within psoriatic tissue following risankizumab treatment, epidermal cell–cell communication changes were next evaluated. Ligand–receptor analyses on keratinocyte subtypes and CD8+ TRM1/17 cells were performed using CellChat. The number of ligand-receptor pairs significantly decreased following high-dose risankizumab treatment, particularly affecting crosstalk between keratinocytes and CD8+ TRM1/17 cells (Supplementary Fig. 6). CD8+ TRM are maintained long-term within tissue in a manner dependent on locally produced IL-7 and IL-1523. Notably, predicted cell–cell interactions between keratinocytes and CD8+ TRM cells, including IL-15 to IL-15R (in particular CD8+ TRM1), IL-7 to IL-7R, and TNFSF15 (TL1A) to TNFRSF25 (in particular CD8+ TRM17) were diminished, more so in the 600 mg dose group when compared to the 300 mg dose group (Fig. 6a–d) and in the high dose vs baseline (Fig. 6e, f). Of note, TL1A has been shown to play a critical role in the differentiation of Th17 cells and progression of autoimmune diseases24. Cell-cell crosstalk effects following risankizumab treatment were further validated through NicheNet (Supplementary Fig. 7), which showed similar results25.
a, b Ligand–receptor interaction probabilities among keratinocyte subtypes and CD8 + TRM1 or CD8 + TRM17 in 300 mg and 600 mg groups. c, d Ligand–receptor interaction probabilities among keratinocyte subtypes and CD8 + TRM1 or CD8 + TRM17 in baseline and 600 mg group. a–d The means of the average expression levels of two interacting molecules are indicated by the color heatmap (bottom panel), with blue to red representing low to high expression. P values were obtained via one-sided permutation tests and are indicated by the circle size. See Supplementary Data for exact P value. e, f Feature plots showing down-regulation of IL7, IL15, and TNFSF15 in keratinocytes.
Discussion
In the KNOCKOUT study reported here, patients with moderate-to-severe psoriasis treated with high induction doses of risankizumab (300 or 600 mg) at weeks 0, 4, and 16 showed rapid and high skin clearance rates at weeks 16 and 28. At week 28, 94.4% of all patients achieved PASI 75, 94.4% achieved PASI 90, and 83.3% achieved PASI 100. No new safety signals were observed in these patients despite the higher induction doses, consistent with the safety profile of patients with psoriasis treated with standard approved 150 mg doses of risankizumab. The majority of patients maintained efficacy at week 52, 36 weeks following their last drug dose. Remarkably, two patients (11.1%) demonstrated completely clear skin at week 100, 84 weeks following their last drug dose. Tissue analyses 36 weeks following the last risankizumab doses revealed significant reductions in tissue CD8+TRM cells, particularly for the 600 mg dose group, which are cells believed to be responsible for psoriasis recurrences following drug withdrawal. Significant reductions in cell–cell interactions between CD8+TRM cells and keratinocytes were also observed, along with significant down-regulations in IL-7, IL-15, and TL1A expression, providing plausible biologic mechanisms for the loss of CD8+TRM cells.
High induction doses of risankizumab produced higher initial rates of skin clearance compared to prior clinical studies with risankizumab. In the phase 3 UltIMMA 1 and 2 studies, 75.3% and 74.8% of patients receiving 150 mg of risankizumab, respectively, achieved PASI 90 at week 164. In the IMMhance study, 73.2% of patients receiving 150 mg risankizumab achieved PASI 90 at week 166. By contrast, all but one patient (94.4%) achieved PASI 90 at week 16 in the KNOCKOUT study reported here. Similar trends were observed for PASI 100 responses, with 35.9% of patients in UltIMMa 1, 50.7% of patients in UltIMMa 2, and 47.2% of patients in IMMhance achieving clear skin at week 164,6; 66.7% of patients in KNOCKOUT achieved PASI 100 at week 16. Remarkably, the PASI 100 response rate rose to 83.3% at week 28, following three doses of risankizumab. Thus, higher-than-approved initial doses of risankizumab lead to much higher short-term efficacy responses in patients with psoriasis. Furthermore, the KNOCKOUT complete clearance rate is approximately 20–30 percentage points higher than those reported for the best available treatments on the market to treat psoriasis, although any comparisons should be made with caution due to the small sample size of this pilot study1,26.
In the IMMhance study, patients received 150 mg standard doses of risankizumab at weeks 0, 4, and 16, and then were withdrawn from drug6. At week 52, 52.4% and 30.2% of IMMhance patients achieved PASI 90 and PASI 100, respectively. It is intriguing to note that two KNOCKOUT patients remained completely clear of psoriasis at the week 100 timepoint, 84 weeks following their last risankizumab dose. This suggests that some patients may have psoriatic inflammation that is particularly sensitive to IL-23 inhibition. Indeed, certain patients have been reported as “super-responders” for their high levels of skin clearance following treatment with IL-23 inhibitors; notably, these patients had psoriasis disease duration of less than 2 years27,28. These prior reports, combined with the KNOCKOUT results reported here, introduce the question of whether psoriasis patients with short disease duration may be particularly responsive to high induction doses of risankizumab, i.e., the “hit hard, hit early” hypothesis29. This approach remains to be studied in psoriasis, since the KNOCKOUT cohort included only two patients with short disease duration. In summary, long-term KNOCKOUT data over 100 weeks indicate that larger initial start-up doses of risankizumab can lead to disease remission times that are longer than previously reported following drug withdrawal2, and that some patients may experience sustained disease modification.
No new safety signals were observed in this study. No TEAEs related to study treatment were reported, and no TEAEs led to study discontinuation, dose reductions, or interruption of study treatment. There were no deaths. One serious, severe TEAE was reported, a worsening right tibial fracture and meniscal tear, which was deemed unrelated by the investigator. Of note, high doses of risankizumab are FDA-approved for inflammatory bowel disease, where the drug is also considered generally safe30. Specifically, initial dosing of risankizumab is 600 mg intravenously at weeks 0, 4, and 8 for Crohn’s disease31,32 and 1200 mg intravenously at weeks 0, 4, and 8 for ulcerative colitis33, both much higher than the 150 mg dosing at weeks 0, 4, and 16 utilized for treating patients with moderate-to-severe psoriasis. In sum, the KNOCKOUT safety data reported here are consistent with the reported safety profile of risankizumab and other IL-23 inhibitors in psoriasis and other diseases3,34,35.
The importance of the IL-23/Th17 pathway in the development and maintenance of psoriasis has been well established36. While both classes of medications are highly effective for psoriasis, IL-23 inhibitors, compared to IL-17 inhibitors, demonstrate the longest disease-free intervals following drug withdrawal2. These differences are not solely due to the pharmacokinetics of these two classes of medications. Prolonged remissions following withdrawal of IL-23 inhibitors are believed, at least in part, to be due their effects on TRM cells8. With treatment of psoriasis in general, TRM cells remain in previous inflamed psoriatic tissue and are responsible for psoriasis recurrences in the same body locations following drug withdrawal12. Notably, guselkumab, an IL-23 inhibitor, but not secukinumab, an IL-17 inhibitor, decreased TRM cells 6 months following onset of treatment37. Tissue effects following treatment with tildrakizumab, another IL-23 inhibitor, were also recently studied; inflammatory fibroblasts were identified as early targets of treatment38. TRM cell survival has also been shown to be under the control of IL-2315. KNOCKOUT data confirm that risankizumab, an IL-23 inhibitor like guselkumab, also decreases TRM cells in psoriatic skin, and that this effect is pronounced and prolonged, being shown in tissue 36 weeks following the last dose of risankizumab.
Mechanisms that underlie prolonged clinical remissions following treatment with IL-23 inhibitors have not been previously studied in detail. Here, CD8+ TRM17 cells were shown to be significantly decreased in lesional tissue 36 weeks following the last dose of risankizumab. In addition, epidermal keratinocytes, comprising the environment surrounding CD8+ TRM17 cells, normalized pro-inflammatory gene expression following risankizumab dosing. Furthermore, predicted cell–cell interactions between keratinocytes and CD8+ TRM cells were diminished in week 52 lesional tissue, including IL-15 to IL15R, IL-7 to IL-7R, and TNFSF15 (TL1A) to TNFRSF25 interactions; these changes were more pronounced in the 600 mg dose group when compared to the 300 mg dose group. This is notable because CD8+ TRM are maintained long-term within tissue in a manner dependent on locally produced IL-7 and IL-1523. In summary, these tissue findings explain, at least in part, the biologic mechanisms underlying long-term psoriasis remissions following anti-IL-23 treatment.
A limitation of this study is the small sample size, and as such, this study is not powered to detect significant differences between treatment groups. Additionally, due to the small sample size, imbalances may be present in patient demographics and baseline characteristics. Further studies may increase the number of patients enrolled and explore additional initial doses, since little-to no efficacy differences were demonstrated between the 300 and 600 mg dose groups. The most plausible explanation for the lack of a dose response in efficacy is the imbalance in disease severity between the two dose groups, with the 300 mg group having much less severe disease compared to the 600 mg dose group. Additional studies and analyses may also further explore the changes in lesional skin at different timepoints other than week 52. Further understanding of the biologic changes in skin at a cellular level may lead to better understanding of the mechanisms that underlie high levels of skin clearance and high rates of disease remission over time.
In conclusion, treatment with high induction doses of risankizumab produced rapid and durable skin clearance in patients with psoriasis, with no new safety signals, while markedly reducing tissue TRM over time. This study demonstrated that high induction doses of risankizumab, labeled “knockout dosing,” was able to achieve rapid, long-lasting skin clearance by altering resident memory T cell populations. In the clinical setting, high induction doses could potentially provide fast relief for patients with higher skin burdens. Further research is needed to determine if knockout dosing may also relieve patient burden due to reduced treatment maintenance requirements and potentially reduce treatment costs.
Methods
Study design and patients
The KNOCKOUT (decreasing resident memory T cells while increasing clinical durability: higher induction doses of risankizumab for moderate-to-severe plaque psoriasis) study was a single-center, randomized, double-blinded, 100-week, interventional, phase 2 study. Twenty patients ≥18 years old from the Portland metropolitan area with chronic stable moderate-to-severe plaque psoriasis (sPGA 3/4, body surface area (BSA) ≥ 10, and PASI ≥ 12 for at ≥6 months) and in good general health were recruited. Exclusion criteria included treatment with topical psoriasis in the past 2 weeks, oral medication or phototherapy for psoriasis in the past 4 weeks, any biologic treatment for psoriasis in the past 4 months, or any prior treatment with risankizumab. Patients were randomized in a 1:1 ratio to receive either 300 mg (2× dosing) or 600 mg (4× dosing) of subcutaneous risankizumab at weeks 0, 4, and 16. A random number generator was used to generate the sequence for drug assignments: 1 or 2, with 1 = 300 mg and 2 = 600 mg. This was performed by a dedicated staff member at Oregon Medical Research Center who was not involved in patient efficacy or safety assessments during the trial. This person also prepared the drug for each patient at each visit and administered the subcutaneous injections. Patients were recruited from February 2022 to August 2022. This study was approved by the IRB, and informed consent was obtained for all patients.
Clinical assessments and safety
Clinical assessments for psoriasis included PASI, sPGA, and BSA. These were assessed at weeks 0, 2, 4, 8, 12, 16 and then every 6 weeks until week 100. Collection of safety events was performed at each of these visits as well. Quality of life was assessed by the DLQI at weeks 0, 28, and 52. Routine laboratory tests, including complete blood count and serum chemistries, were assessed at weeks 4, 8, 22, and 46. All patients underwent baseline lesional and non-lesional biopsies at the Oregon Medical Research Center. The primary endpoint was the change from baseline in the number and/or effector function of epidermal CD8 + CD103 + TRM cells at week 52 in patients receiving either 300 or 600 mg of risankizumab. Secondary endpoints were PASI 100 at weeks 24, 40, 52, and 100 and safety events over 100 weeks in patients receiving 300 or 600 mg of risankizumab.
Statistical analyses
This study was exploratory in nature and thus was not statistically powered to detect differences between the treatment arms. All analyses were performed on the full analysis set, which included all randomized subjects who received at least 1 dose of risankizumab. Efficacy was reported using modified non-responder imputation, where patients who did not receive all three induction doses due to reasons other than lack of efficacy or intolerability were excluded. Continuous outcomes were summarized with mean, standard deviation, and 95% confidence interval, and compared between the two treatment arms with a t-test. For binary outcomes, proportions are summarized with percentages and 95% confidence intervals and compared between the two treatment arms with Fisher’s exact test. Analyses were performed with SAS software, version 9.4 (SAS Institute).
Generation of single-cell suspensions for scRNA-seq
Each biopsy was bisected vertically from the epidermis down with half of a 6 mm biopsy was cryopreserved in CryoStor CS10 media (BioLife Solutions). Samples were thawed on ice, washed briefly in cold Hank’s Balanced Saline Solution (Gibco, Thermo Fisher Scientific) to remove residual CryoStor CS10 media, and bi-sected before being enzymatically digested in either 0.25% Trypsin-EDTA (Gibco, Thermo Fisher Scientific) with 10 U/mL DNase I (Thermo Fisher Scientific) for 1 h at 37 °C quenched with FBS (Atlanta Biologicals) or 0.2% Collagenase II (Life Technologies) and 0.2% Collagenase V (Sigma) with 10 U/mL DNase I in plain medium for 1.5 h at 37 °C with rotation. The resulting cell suspensions were filtered through 70-μm cell strainers twice and re-suspended in PBS containing 0.04% BSA. Dermal and epidermal cells were combined in a 2:1 ratio to enrich for dermal cell populations, and libraries were constructed by the University of Michigan Advanced Genomics Core on the 10X Genomics Chromium system with chemistry v2 and v3 and sequenced on the Illumina NextSeq to generate 150 bp paired-end reads targeting 10,000 cells per sample at 20,000 reads/cell. Sequence data have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE305818. [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE305818]
Cell clustering and cell type annotation
The Seurat R package (v5.0.1) was employed to perform cell clustering on the combined matrix39. Cells with fewer than 200 features or exhibiting over 10% mitochondrial expression were excluded as low-quality cells. Expression levels were normalized for each cell using the NormalizeData function with default settings. Variable genes were identified using FindVariableFeatures, also with default settings. The ScaleData function was applied to scale and center the dataset. Principal component analysis (PCA) was conducted on the variable genes, with the first 20 principal components used for cell clustering and UMAP dimensional reduction. Clusters were determined by the FindNeighbors and FindClusters functions, with a resolution of 0.5. Cluster marker genes were identified through the FindAllMarkers function. Cell type annotations were assigned by matching cluster markers to canonical cell type signature genes. To calculate the disease composition for each cell type, the number of cells from each disease condition was counted per cell type, divided by the total number of cells for each condition, and scaled to 100 percent for each cell type.
Pseudotime trajectory construction
Pseudotime trajectory analysis was performed using the Monocle package (v2.28.0)17. Raw cell counts obtained from the Seurat analysis were normalized using the estimateSizeFactors and estimateDispersions functions with default settings. Genes with an average expression greater than 0.5 and present in more than 10 cells were retained for subsequent analysis. Variable genes were identified using the FindVariableFeatures function. Cell ordering was determined by the orderCells function, and the trajectory was constructed using reduceDimension, both with default parameters.
Differential expression analysis
Differentially expressed genes between lesional baseline and lesional post-treatment (week 52) samples were identified using the FindMarkers function. Genes present in fewer than 20% of cells in either sample group were excluded from the analysis.
Cell–cell interaction inference
The ligand-receptor (LR) interaction analysis was first performed through CellChat (v1.6.1)20. The main analysis was based on T cell subtypes or with keratinocyte subtypes. A separate run was performed for each sample group, and the number of significant interactions was calculated for each cell type pair. Cellular interactions were also confirmed by using nichenetr (v2.0.6) to analyze the different LR pairs between the lesional baseline and lesional post-treatment (week 52) sample data26. Briefly, keratinocytes were defined as “sender” cells, and CD8 + TRM17 cells were defined as “receiver” cells. Moreover, interactions among CD8 + TRM1/17 and keratinocyte subtypes were further detected using iTALK (v0.1.0) based on known LR pairs, including checkpoint, cytokine, and growth factor40.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
AbbVie is committed to responsible data sharing regarding the clinical trials we sponsor. This includes access to anonymized, individual, and trial-level data (analysis data sets) as well as other information (e.g., protocols, clinical study reports, or analysis plans), as long as the trials are not part of an ongoing or planned regulatory submission. This includes requests for clinical trial data for unlicensed products and indications. These clinical trial data can be requested by any qualified researchers who engage in rigorous, independent, scientific research, and will be provided following review and approval of a research proposal, Statistical Analysis Plan (SAP), and execution of a Data Sharing Agreement (DSA). Data requests can be submitted at any time after approval in the US and Europe and after acceptance of this manuscript for publication. The data will be accessible for 12 months, with possible extensions considered. For more information on the process or to submit a request, visit the following link: https://vivli.org/ourmember/abbvie/ then select “Home”. Available source data is provided with this paper. The sequencing data generated in this study have been deposited in the Gene Expression Omnibus under accession number GSE305818. [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE305818] Source data are provided with this paper.
References
Sbidian, E. et al. Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis. Cochrane Database Syst. Rev. 5, Cd011535 (2022).
Masson Regnault, M., Shourick, J., Jendoubi, F., Tauber, M. & Paul, C. Time to relapse after discontinuing systemic treatment for psoriasis: a systematic review. Am. J. Clin. Dermatol. 23, 433–447 (2022).
Gordon, K. B. et al. Long-term safety of risankizumab from 17 clinical trials in patients with moderate-to-severe plaque psoriasis. Br. J. Dermatol. 186, 466–475 (2022).
Gordon, K. B. et al. Efficacy and safety of risankizumab in moderate-to-severe plaque psoriasis (UltIMMa-1 and UltIMMa-2): results from two double-blind, randomised, placebo-controlled and ustekinumab-controlled phase 3 trials. Lancet 392, 650–661 (2018).
Warren, R. B. et al. Efficacy and safety of risankizumab vs. secukinumab in patients with moderate-to-severe plaque psoriasis (IMMerge): results from a phase III, randomized, open-label, efficacy-assessor-blinded clinical trial. Br. J. Dermatol. 184, 50–59 (2021).
Blauvelt, A. et al. Efficacy and safety of continuous risankizumab therapy vs treatment withdrawal in patients with moderate to severe plaque psoriasis: a phase 3 randomized clinical trial. JAMA Dermatol. 156, 649–658 (2020).
Krueger, J. G. et al. Anti-IL-23A mAb BI 655066 for treatment of moderate-to-severe psoriasis: safety, efficacy, pharmacokinetics, and biomarker results of a single-rising-dose, randomized, double-blind, placebo-controlled trial. J. Allergy Clin. Immunol. 136, 116–124.e117 (2015).
Blauvelt, A. Resident memory T cells in psoriasis: key to a cure? J. Psoriasis Psoriatic Arthritis 7, 157–159 (2022).
Mackay, L. K. et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Mueller, S. N., Zaid, A. & Carbone, F. R. Tissue-resident T cells: dynamic players in skin immunity. Front. Immunol. 5, 332 (2014).
Clark, R. A. Resident memory T cells in human health and disease. Sci. Transl. Med. 7, 269rv261 (2015).
Cheuk, S. et al. Epidermal Th22 and Tc17 cells form a localized disease memory in clinically healed psoriasis. J. Immunol. 192, 3111–3120 (2014).
Gallais Sérézal, I. et al. Resident T cells in resolved psoriasis steer tissue responses that stratify clinical outcome. J. Investig. Dermatol. 138, 1754–1763 (2018).
Matos, T. R. et al. Clinically resolved psoriatic lesions contain psoriasis-specific IL-17-producing αβ T cell clones. J. Clin. Investig. 127, 4031–4041 (2017).
Whitley, S. K. et al. Local IL-23 is required for proliferation and retention of skin-resident memory T(H)17 cells. Sci. Immunol. 7, eabq3254 (2022).
Park, S. L. et al. Divergent molecular networks program functionally distinct CD8(+) skin-resident memory T cells. Science 382, 1073–1079 (2023).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Abdelbary, M., Hobbs, S. J., Gibbs, J. S., Yewdell, J. W. & Nolz, J. C. T cell receptor signaling strength establishes the chemotactic properties of effector CD8(+) T cells that control tissue-residency. Nat. Commun. 14, 3928 (2023).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Mohammed, J. et al. Stromal cells control the epithelial residence of DCs and memory T cells by regulated activation of TGF-β. Nat. Immunol. 17, 414–421 (2016).
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol. 33, 169–200 (2015).
Falcão, A. M. et al. Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat. Med. 24, 1837–1844 (2018).
Adachi, T. et al. Hair follicle-derived IL-7 and IL-15 mediate skin-resident memory T cell homeostasis and lymphoma. Nat. Med. 21, 1272–1279 (2015).
Pappu, B. P. et al. TL1A-DR3 interaction regulates Th17 cell function and Th17-mediated autoimmune disease. J. Exp. Med. 205, 1049–1062 (2008).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Blauvelt, A. & Noe, M. H. The best psoriasis medications emerge. JAMA Dermatol. 160, 99–100 (2024).
Gargiulo, L. et al. A risankizumab super responder profile identified by long-term real-life observation-IL PSO (ITALIAN LANDSCAPE PSORIASIS). J. Eur. Acad. Dermatol. Venereol. 38, e113–e116 (2024).
Schäkel, K. et al. Early disease intervention with guselkumab in psoriasis leads to a higher rate of stable complete skin clearance (“clinical super response”): week 28 results from the ongoing phase IIIb randomized, double-blind, parallel-group, GUIDE study. J. Eur. Acad. Dermatol. Venereol. 37, 2016–2027 (2023).
Iversen, L. et al. Secukinumab treatment in new-onset psoriasis: aiming to understand the potential for disease modification-rationale and design of the randomized, multicenter STEPIn study. J. Eur. Acad. Dermatol. Venereol. 32, 1930–1939 (2018).
Ferrante, M. et al. Long-term safety and efficacy of risankizumab treatment in patients with Crohn’s disease: results from the phase 2 open-label extension study. J. Crohns Colitis 15, 2001–2010 (2021).
D’Haens, G. et al. Risankizumab as induction therapy for Crohn’s disease: results from the phase 3 ADVANCE and MOTIVATE induction trials. Lancet 399, 2015–2030 (2022).
Ferrante, M. et al. Risankizumab as maintenance therapy for moderately to severely active Crohn’s disease: results from the multicentre, randomised, double-blind, placebo-controlled, withdrawal phase 3 FORTIFY maintenance trial. Lancet 399, 2031–2046 (2022).
Louis, E. et al. Risankizumab for ulcerative colitis: two randomized clinical trials. JAMA 332, 881–897 (2024).
Papp, K. A. et al. Long-term safety and efficacy of risankizumab for the treatment of moderate-to-severe plaque psoriasis: Interim analysis of the LIMMitless open-label extension trial up to 5 years of follow-up. J. Am. Acad. Dermatol. 89, 1149–1158 (2023).
Blauvelt, A., Chiricozzi, A., Ehst, B. D. & Lebwohl, M. G. Safety of IL-23 p19 inhibitors for the treatment of patients with moderate-to-severe plaque psoriasis: a narrative review. Adv. Ther. 40, 3410–3433 (2023).
Hawkes, J. E., Yan, B. Y., Chan, T. C. & Krueger, J. G. Discovery of the IL-23/IL-17 signaling pathway and the treatment of psoriasis. J. Immunol. 201, 1605–1613 (2018).
Mehta, H. et al. Differential changes in inflammatory mononuclear phagocyte and T-cell profiles within psoriatic skin during treatment with guselkumab vs. secukinumab. J. Investig. Dermatol. 141, 1707–1718.e1709 (2021).
Francis, L. et al. Single-cell analysis of psoriasis resolution demonstrates an inflammatory fibroblast state targeted by IL-23 blockade. Nat. Commun. 15, 913 (2024).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Wang, Y. et al. iTALK: an R package to characterize and illustrate intercellular communication. Preprint at bioRxiv https://doi.org/10.1101/507871 (2019).
Acknowledgements
AbbVie and the authors thank the participants, study sites, and investigators who participated in this clinical trial. Medical writing support was provided by Trisha Rettig, PhD, of AbbVie. AbbVie funded this trial and participated in the trial design, research, analysis, data collection, interpretation of data, and the review and approval of the publication. All authors had access to relevant data and participated in the drafting, review, and approval of this publication. No honoraria or payments were made for authorship.
Author information
Authors and Affiliations
Contributions
A.B., R.J., H.P., J.E.G., and B.D.E. contributed to the design of the study. Experiments were conducted by R.J., L.C.T., R.B., J.F., M.G.K., and A.C.B. L.S. provided statistical analysis for efficacy and safety data. All authors contributed to the analysis and review of data and writing of the manuscript. The author order was assigned on the relative level of contribution each individual made to the research project.
Corresponding author
Ethics declarations
Competing interests
A.B. has served as a speaker (received honoraria) for Almirall, Eli Lilly, Sanofi, and UCB, has served as a scientific adviser (received honoraria) for AbbVie, Almirall, Alumis, Amgen, Anaptysbio, Apogee, Arcutis, Astria, Boehringer Ingelheim, Bristol Myers Squibb, Celltrion, Corvus, Dermavant, Eli Lilly, Galderma, GlaxoSmithKline, Immunovant, Incyte, IQVIA, Janssen, Leo, Lipidio, Merck, Novartis, Oruka, Paragon, Pfizer, Rani Therapeutics, Regeneron, Sanofi, Spherix Global Insights, Sun Pharma, Sycona, Takeda, UCB Union, and Zai Lab, and has acted as a clinical study investigator (institution has received clinical study funds) for AbbVie, Acelyrin, Almirall, Alumis, Amgen, Arcutis, Boehringer Ingelheim, Bristol-Myers Squibb, Dermavant, Eli Lilly, Galderma, Incyte, Janssen, Leo, Merck, Novartis, Pfizer, Regeneron, Sanofi, Sun Pharma, Takeda, and UCB, and owns stock in Lipidio and Oruka. R.J., L.T., R.B., J.F., M.G.K., A.B., and R.M. have no conflicts of interest to disclose. L.S. and H.P. are employees of AbbVie Inc. and may hold AbbVie stock, stock options, and/or patents. J.E.G. is supported by the A. Alfred Taubman Medical Research Institute as the Kenneth and Frances Eisenberg Emerging Scholar Award and a Doris Duke Charitable Foundation Grant. R.M. has served as a clinical investigator for AbbVie, Actavis, Advanced Tissue Sciences, Allergan, Amgen, Anacor, Astellas, Atrix, Barrier Therapeutics, Bayer, Berlex, Boehringer Ingelheim, Bristol Myers Squibb, Celgene, Cellgate, Celtic, Centocor, CombinatoRx, Connetics, Corixa, Cu Tech, Dermira, Dermik, Dow, DUSA, Eli Lilly, Ferndale, Fujisawa, Galderma, Genentech, Genetics Institute, GlaxoSmithKline, Graceway, Halcygen, HanAll Biopharma, HealthPoint, Human Genome Sciences, IDEC, Immunex, Intendis, ISIS, Isolagen, Janssen, Johnson & Johnson, Leo, Matrix, Medicis, MediGene, MedImmune, Merck, NanoBio, Novartis, Novum, Nycomed, Ortho McNeil, Paddock, Park Davis, Peplin, Perrigo, Pfizer, Promius, Regeneron, Research Testing Labs, Rhone-Poulenc, Roche, Rorer, Samumed, Sandoz, Schering Plough, Serono, Shionogi, Sirtris, Stiefel, Synta, Talima, Taro, Teva, Tolerx, Tolmar, Topica, Vertex, Viamet, Vicept, Wyeth, and XOMA. B.D.E. has received fees/honoraria/royalties as an advisory board member, contributor, and/or consultant for AbbVie, Acelyrin/Affibody, Amgen, AnaptysBio, Arcutis, Bristol Myers Squibb, Dermavant/Organon, Evelo, Evommune, Incyte, Janssen/Johnson & Johnson, LEO Pharma, Lilly, Navigator Medicines, Novartis, Ortho/Bausch Health, Priovant, Regeneron, Sanofi, up-to-date, and UCB; received speaking fees from AbbVie, Dermavant/Organon, Incyte, LEO Pharma, Lilly, Novartis, the National Psoriasis Foundation, Regeneron, Ortho/Bausch Health, and Sanofi; and received institutional funding as an investigator for AbbVie, Acelyrin/Affibody, Aclaris, Allakos/Concentra, Almirall, Alumis, Amgen, AnaptysBio, Apogee, Arcutis, Bristol Myers Squibb, Celldex, Celgene/Bristol Myers Squibb, Concert/Sun Pharma, Dermavant/Organon, DermBiont, Dermira/Lilly, Evelo, Incyte, Janssen/Johnson & Johnson, Kymab/Sanofi, LEO Pharma, Lilly, Novan/Ligand, Novartis, Pfizer, Regeneron, Sanofi, Sun Pharma, UCB, and Ventyx.
Peer review
Peer review information
Nature Communications thanks Marco Galluzzo, Luigi Gargiulo and Lajos Kemény for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Blauvelt, A., Jiang, R., Shi, L. et al. A randomized phase 2 clinical trial to treat moderate-to-severe plaque psoriasis patients with high-induction dosing of risankizumab. Nat Commun 17, 733 (2026). https://doi.org/10.1038/s41467-025-67475-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67475-0