Abstract
The final step of oocyte growth, which reorganizes chromatin from the non-surrounded nucleolus (NSN) to the surrounded nucleolus (SN) configuration is essential for embryonic development after meiotic maturation and fertilization. The underlying mechanisms remain unknown. We identify RNA polymerase II (RNAPII) degradation as the key driver of this process. Inhibitors that trigger RNAPII degradation, but not nucleoside-based transcription inhibitors, induce NSN-to-SN transition in oocytes. By establishing miniTrim-Away for nuclear proteins and using segregase and proteasome inhibitors, we demonstrate that RNAPII degradation is necessary and sufficient for NSN-to-SN transition. Further experiments reveal that RNAPII degradation results in a global collapsing force and a local attractive force required for the transition to SN configuration. Finally, embryos derived from NSN oocytes have aberrant RNAPII levels and localization, and are defective in maternal-to-zygotic transition. Our study elucidates the mechanistic framework of oocyte chromatin reorganization and presents a strategy for inducing fully grown oocyte nuclei.
Similar content being viewed by others
Introduction
Oogenesis is essential for sexual reproduction in mammals. Growing oocytes have a non-surrounded nucleolus (NSN) chromatin configuration1,2,3,4 and actively transcribe messenger RNAs (mRNAs)2,5, which are used to establish protein and mRNA stores for the next generation5,6,7,8. During the final stages of oogenesis, condensed chromatin forms a ring around the nucleolus in a surrounded nucleolus (SN) configuration1,2,3,4 and transcription ceases2,5,9,10,11,12,13,14,15. SN oocytes have a higher developmental competence than NSN oocytes3,16,17,18,19,20,21,22,23,24, and their nucleus primarily accounts for their differences in the success of embryonic development25. Therefore, the reorganization of chromatin from NSN to SN configuration is tied to the developmental potential of mammalian oocytes.
NSN-to-SN transition is accompanied by many characteristic changes in the nuclear architecture26. Compared to NSN nuclei, SN nuclei display a reduction in nucleolar components and ribosomal RNA (rRNA) synthesis27,28,29,30,31,32, clustering of centromeres and pericentromeric constitutive heterochromatin (chromocenters) around the nucleolus33,34,35, fewer but larger splicing-related nuclear speckles36,37,38,39,40 and altered histone modifications and DNA methylations35,41,42,43,44.
How the chromatin is reorganized during this key step of mammalian oocyte development is a long-standing question. Previous studies of knockout mice have reported several proteins that could alter the NSN/SN ratio of isolated oocytes31,45,46,47,48,49,50,51,52,53,54,55,56,57, but whether these proteins are involved in NSN-to-SN transition via directly driving this process or indirectly affecting oocyte growth is unclear. Furthermore, the physiological relevance of these proteins is not known: evidence that their expression or activity changes during NSN-to-SN transition in vivo is lacking. Here, we comprehensively mapped almost 30 nuclear components in NSN and SN oocytes to uncover the underlying molecular differences and to identify candidates that drive the physiological NSN-to-SN transition.
Results
Comprehensive analysis of NSN and SN nuclei
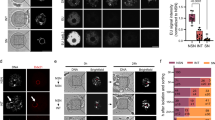
First, we established a workflow for studying NSN and SN oocytes by sorting freshly isolated germinal vesicle (GV) oocytes based on their chromatin configurations, followed by fixation, immunofluorescence staining, high-resolution microscopy, and machine learning-based 3D segmentation analyses (Fig. 1a). To avoid phototoxicity and DNA damage incurred by the use of UV-excitable Hoechst dyes3,58, we performed live staining of oocyte chromatin using the green-excitable SPY555-DNA or red-excitable 5-SiR-Hoechst dyes, which are routinely used for long-term live imaging of human oocytes59. NSN and SN oocytes were sorted with a high accuracy of 99.0% and 100%, respectively (Fig. 1b). 3D segmentation analysis confirmed that the chromatin volume of NSN and SN oocytes differed by two-fold (Fig. 1c), consistent with chromatin condensation60 and detachment from the nuclear envelope61 during NSN-to-SN transition.
a Workflow for live sorting of NSN and SN oocytes and 3D segmentation analyses of oocyte nuclei. b Manual scoring of chromatin configurations in sorted NSN and SN oocytes after fixation and staining with Hoechst. c Automated quantification of chromatin volume in NSN and SN oocytes. p < 0.0001. d Schematic representation of oocyte nucleus (scheme: chromocenters in green, nuclear bodies in magenta, and chromatin in blue). e Automated quantification of chromatin volume in NSN and SN oocytes treated with different small molecule inhibitors or overexpressing different nuclear components. P value for NSN treated with RNase A, ActD, BRM/BRG1i and SN treated with 1,6-hexanediol, CTCF OE, DZNep, HP1β OE is less than 0.0001; P(NSN-GSK-J4) = 0.001; P(NSN-CHX) = 0.0002; P(NSN-Quarfloxin) = 0.292; P(NSN-Bleomycin) = 0.5768; P(NSN-Hinokiflvone) = 0.9574; P(NSN-TSA) = 0.0888; P(NSN-Etoposide) = 0.0294; P(SN-TET mix OE) = 0.0791; P(SN-MATR3 OE) = 0.0558; P(SN-hnRNPU OE) = 0.1106; P(SN-A-485) = 0.4147; P(SN-Hinokiflavone) = 0.0077; P(SN-DNMT mix OE) = 0.0096. f, g Immunofluorescence images of NSN oocytes treated with triptolide or α-amanitin. h–k Automated quantification of total intensity of EU, chromatin volume, total intensity of pS2, and RPB1 in NSN oocytes treated with triptolide or α-amanitin. For total intensity of EU, pS2 RPB1, and chromatin volume between NSN oocytes treated with water and triptolide or α-amanitin, p < 0.0001. *p < 0.05. **p < 0.01. ***p < 0.001. ****p < 0.0001. N.S., not significant. All box plots show median (horizontal black line), mean (small black squares), 25th and 75th percentiles (boxes), 5th and 95th percentiles (whiskers), and 1st and 99th percentiles (crosses). Statistical significance is based on unpaired, two-tailed Student’s t test (c, e, h–k). The number of oocytes analyzed is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
To study the molecular differences between NSN and SN nuclei, we systematically evaluated the localization and staining intensity of 28 common nuclear components (Fig. 1d and Supplementary Fig. 1a–d). We found that 13 of them localized to chromatin, and the staining intensity for most of them was lower in SN nuclei. Seven nuclear components localized to chromocenters and their staining intensity were all lower in SN nuclei. Nucleolar proteins and the nuclear speckle marker SRSF2 also showed lower staining intensity in SN nuclei, in line with previous reports27,28,29,30,32,36,37,38,39,40. These data suggest that most nuclear components are downregulated during NSN-to-SN transition.
RNA polymerase II (RNAPII) inhibitors ectopically induce NSN-to-SN transition in mouse oocytes
NSN-to-SN transition did not spontaneously occur even after prolonged culture of NSN oocytes ex vivo (Supplementary Fig. 2a–c), in line with early studies3,62. In order to identify nuclear components potentially driving NSN-to-SN transition, we used small-molecule inhibition and overexpression approaches to transiently modulate the levels or activities of different nuclear components, thereby mimicking their differences in NSN and SN oocytes (Fig. 1d, Supplementary Fig. 1a and Supplementary Table 1). Many of the treatments that led to the desired changes in nuclear components altered the transcriptional activity and/or chromatin compaction in NSN and SN oocytes (Fig. 1e, Supplementary Fig. 2d, e and Supplementary Table 2). In particular, treating NSN oocytes with RNase A, ActD, BRM/BRG1i or GSK-J4 prominently silenced transcription and/or compacted chromatin, whereas CTCF overexpression or DZNep treatment of SN oocytes mildly activated transcription and/or decompacted chromatin. While no treatment was able to reverse NSN-to-SN transition, we found that ActD is able to induce NSN-to-SN transition as evidenced by the ring-like chromatin configuration around the nucleolus and the almost two-fold reduction in chromatin volume (Fig. 1e and Supplementary Fig. 2f, g).
We also considered the potential contributions of liquid-liquid phase separation (LLPS), which has recently been implicated in driving chromatin reorganization in sperm cells of flowering plants63. To test if LLPS drives NSN-to-SN transition, we treated SN oocytes with 1,6-hexanediol, which could disrupt LLPS via weakening hydrophobic interactions between proteins. Although 1,6-hexanediol drastically decompacted chromatin in SN oocytes (Fig. 1e), the morphology of chromatin did not resemble that of NSN oocytes, as characterized by the excessive fragmentation of chromocenters (Supplementary Fig. 2g).
ActD acts on both RNAPI and RNAPII64. The RNAPI inhibitor quarfloxin did not phenocopy ActD (Fig. 1e), implying that NSN-to-SN transition likely involves RNAPII. To test this, we treated NSN oocytes with two widely used RNAPII inhibitors, triptolide and α-amanitin, which covalently bind to the general transcription factor TFIIH to block transcription initiation65 or impair translocation to block transcription elongation64,66,67,68, respectively. As expected, both triptolide and α-amanitin almost completely abolished nascent transcription in NSN oocytes (Fig. 1f–h). Importantly, both treatments induced NSN-to-SN transition within 7 hours after treatment (Fig. 1f, g, i), as supported by live-cell imaging (Supplementary Fig. 3 and Supplementary Movies 1–4). Thus, RNAPII inhibitors can swiftly induce NSN-to-SN transition in mouse oocytes ex vivo.
Induced SN-like nuclei recapitulate epigenetic features, chromatin interactions and developmental potential in SN oocytes
To confirm that RNAPII inhibitors induce NSN-to-SN transition beyond morphological changes, we also examined changes in epigenetic features, chromatin interactions and developmental potential in induced SN-like nuclei. DNA damage marker γH2A.X and pan-histone H4 acetylation (H4ac) staining intensity were increased, whereas DNA methylation, heterochromatin protein HP1β, DNA hydroxymethylation and pan-histone dimethylation (KMe2) staining intensity were reduced in α-amanitin-treated NSN oocytes (Fig. 2a, b), similar to SN oocytes (Fig. 1d and Supplementary Fig. 1a–d). Low-input Hi-C profiling69 further revealed that the chromatin contact map from α-amanitin-treated NSN oocytes is more similar to SN oocytes as compared to NSN oocytes (Fig. 2c). Moreover, α-amanitin-treated NSN oocytes and SN oocytes had almost identical contact probability curves with significantly more long-range ( > 1 Mb) contacts than NSN oocytes (Fig. 2d), in line with previous reports60. Remarkably, in vitro fertilization (IVF) after meiotic maturation of α-amanitin-treated NSN oocytes yielded twice as many 2-cell embryos as compared to NSN oocytes (Fig. 2e). Thus, induced SN-like nuclei recapitulate epigenetic features, chromatin interactions and developmental potential in SN oocytes.
a Immunofluorescence images of NSN oocytes treated with α-amanitin. b Automated quantification of total intensity of different nuclear components in NSN oocytes treated with α-amanitin. Statistical significance is based on unpaired, two-tailed Student’s t test. P value for γH2A.X, 5hmC and KMe2 #2 is less than 0.0001; P(H4ac) = 0.0006; P(5mC) = 0.0071; P(HP1β) = 0.0133. c Chromatin contact maps of NSN, α-amanitin-treated NSN and SN oocytes. d Contact probability curves of NSN, α-amanitin-treated NSN and SN oocytes. e Manual scoring of developmental outcomes of NSN, α-amanitin-treated NSN and SN oocytes after meiotic maturation and IVF. (two-tailed Fisher’s exact test for 2-cell embryos from NSN and α-amanitin-treated NSN oocytes, p = 0.0106) *p < 0.05. **p < 0.01. ***p < 0.001. ****p < 0.0001. All box plots show median (horizontal black line), mean (small black squares), 25th and 75th percentiles (boxes), 5th and 95th percentiles (whiskers), and 1st and 99th percentiles (crosses). The number of oocytes analyzed is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
Silencing transcription is insufficient to ectopically induce NSN-to-SN transition
Apart from direct transcriptional inhibition, studies in other systems reported that triptolide and α-amanitin can also trigger RNAPII degradation70,71,72,73,74,75,76,77,78. We performed immunofluorescence staining with a monoclonal antibody against phospho-serine 2 (pS2) at the C-terminal repeat domain and with a monoclonal antibody against the N-terminal domain of the largest RNAPII subunit RPB1, and found that both phosphorylated and total RNAPII were indeed depleted in triptolide- and α-amanitin-treated NSN oocytes (Fig. 1f, g, j, k).
To determine if silencing transcription alone is sufficient to induce NSN-to-SN transition, we prematurely terminated transcription by incorporating modified adenosine analogues 8-aminoadenosine (8-AA) and tubercidin/7-deazaadenosine (7-DAA)79,80. Both 8-AA and 7-DAA suppressed nascent transcription to a level that was similar to triptolide and α-amanitin treatments (Supplementary Fig. 4a–c), but without degrading RNAPII (Supplementary Fig. 4a, b, d, e). Importantly, however, 8-AA and 7-DAA did not induce NSN-to-SN transition (Supplementary Fig. 4a, b, f). Thus, silencing transcription is insufficient to induce the NSN-to-SN transition ex vivo.
Targeted degradation of RNAPII is sufficient to induce NSN-to-SN transition in mouse and human oocytes
Our data so far suggest that RNAPII degradation might be critical for the NSN-to-SN transition induced by triptolide and α-amanitin beyond transcriptional inhibition. RNAPII is an essential protein81 and existing oocyte-specific driver lines all express Cre recombinase during early oocyte growth82, precluding conditional knockout analysis. We aimed to test this hypothesis by acutely degrading RNAPII in NSN oocytes using Trim-Away, an endogenous protein degradation technique based on Trim21-mediated proteasomal degradation of antibody-antigen complexes83,84. Trim-Away was developed for targeted degradation of cytoplasmic proteins, but RNAPII is a nuclear protein that does not shuttle in and out of the nucleus, so we sought to use the single-chain fragment variable (scFv) for Trim-Away in the nucleus. To this end, we initially tested a scFv developed against pS2 (clone 42B3)85 by expressing a GFP-fusion protein in oocytes. Although the 42B3-GFP construct did not localize to the nucleus (Supplementary Fig. 5a), a 42B3-split GFP construct was able to enter the nucleus and colocalize with pS2 on the chromatin in NSN oocytes (Supplementary Fig. 5b). These data show that the size of fusion tag could interfere with the nuclear localization of scFv. Hence, we fused 42B3 with the RING domain of Trim21 (t21R)86 rather than the Fc domain of IgG83 to develop a miniTrim-Away approach for targeted degradation of nuclear proteins (Fig. 3a).
a Schematic representation of the miniTrim-Away approach. b Immunofluorescence images of NSN oocytes expressing 42B3-t21R to deplete phosphorylated RNAPII for 3 or 16 h. c–f Automated quantification of total intensity of pS2, chromatin volume, total intensity of EU and RPB1 in NSN oocytes expressing 42B3-t21R to deplete phosphorylated RNAPII for 3 or 16 h. For the total intensity of EU, pS2, RPB1, and chromatin volume between the control group and NSN oocytes expressing 42B3-t21R for 3 h or 16 h, all p < 0.0001. g Immunofluorescence images of human oocyte expressing 42B3-t21R to deplete phosphorylated RNAPII for 16 h. h Automated quantification of chromatin surface area-to-volume ratio (SA/V) in human oocytes with partially compacted and fully compacted chromatin. p = 0.0003. i Still images from time-lapse movies of human oocytes with partially compacted chromatin expressing 42B3-t21R. Time is given as hours:minutes from the start of imaging immediately after injection. Yellow dotted circles highlight the nuclei. j Automated quantification of chromatin SA/V in human oocytes with partially compacted chromatin expressing 42B3-t21R over time. Data are presented as mean values +/- SD. ***p < 0.001. ****p < 0.0001. All box plots show median (horizontal black line), mean (small black squares), 25th and 75th percentiles (boxes), 5th and 95th percentiles (whiskers), and 1st and 99th percentiles (crosses). Statistical significance is based on unpaired, two-tailed Student’s t test (c–f, h). The number of oocytes analyzed is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
Expression of 42B3-t21R in NSN oocytes depleted not only pS2 (Fig. 3b, c), but also RNAPII phosphorylated at other residues at the C-terminal repeat domain (Supplementary Fig. 5c) and unphosphorylated RNAPII (Supplementary Fig. 5d). These data suggest that RNAPII dynamically cycles between unphosphorylated and different phosphorylated states87, consistent with active transcription in NSN oocytes. NSN oocytes expressing 42B3-t21R for 3 hours underwent a partial NSN-to-SN transition, as supported by the 1.5-fold reduction in chromatin volume (Fig. 3b, d). The presence of residual nascent transcription (Fig. 3b, e) hinted that the partial NSN-to-SN transition could be due to the residual RNAPII (Fig. 3b, f), so we extended the expression time from 3 to 16 hours. Remarkably, NSN oocytes expressing 42B3-t21R for 16 hours underwent a full NSN-to-SN transition, as confirmed by the 2-fold reduction in chromatin volume and live-cell imaging (Fig. 3b, d, Supplementary Fig. 5e–k and Supplementary Movies 5, 6).
We also tested whether RNAPII degradation drives NSN-to-SN transition in human oocytes. Because immature human GV oocytes leftover from fertility treatment rarely had fully NSN chromatin configuration (1/52), we utilized the ones that had partially compacted chromatin for experiments (Fig. 3g) and specifically used surface area-to-volume (SA/V) ratio instead of volume to quantify the less drastic changes in chromatin (Fig. 3h). Likewise, 42B3-t21R was able to fully deplete RNAPII in human oocytes (Fig. 3g), and we found that this treatment can trigger complete NSN-to-SN transition (Fig. 3i and Supplementary Movie 7,8), as supported by the further reduction in chromatin SA/V ratio (Fig. 3j). Thus, targeted degradation of RNAPII is sufficient to induce NSN-to-SN transition in mouse and human oocytes.
Segregase- and proteasome-mediated degradation of RNAPII is necessary for NSN-to-SN transition
Next, we asked whether RNAPII degradation is physiologically relevant to NSN-to-SN transition in vivo. Previous work reported that the levels of phosphorylated RNAPII are lower in SN oocytes as compared to NSN oocytes14,88,89,90,91,92,93,94,95,96, but whether this observation reflects simply dephosphorylation or downregulation at protein level is not clear. To this end, we compared the abundance of phosphorylated as well as of total RNAPII in NSN and SN oocytes by immunofluorescence staining and immunoblotting. We found that phosphorylated RNAPII is not converted to the unphosphorylated form but depleted at protein level (Supplementary Fig. 6a–d), accounting for the substantial reduction in total RNAPII levels in SN oocytes (Supplementary Fig. 6e–h). As orthogonal approaches, we imaged oocytes in follicles at different growth stages on mouse ovary sections and oocytes harvested from ex vivo follicle culture. Both approaches showed that phosphorylated RNAPII was progressively depleted during late stages of follicle growth (Fig. 4a–g), concomitant with NSN-to-SN transition (Fig. 4a, d, h). Thus, phosphorylated RNAPII was specifically depleted during NSN-to-SN transition in vivo.
a Immunofluorescence images oocytes from follicles at different growth stages on mouse ovary sections. b, c Automated quantification of total intensity of pS2 and RPB1 in oocytes from follicles at different growth stages on mouse ovary sections. Compared to primordial follicle, Padjusted(primary follicle-pS2)=0.60178; Padjusted(primary follicle-RPB1) = 0.97547; Padjusted(secondary follicle-pS2)=0.99954; Padjusted(secondary follicle-RPB1) = 0.00911; Padjusted(antral follicle-pS2)=0; Padjusted(antral follicle-RPB1) = 0. d Immunofluorescence images of oocytes harvested from mouse secondary follicles cultured for different days. e–h Automated quantification of total intensity of pS2, RPB1, chromatin-bound RPB1 and chromatin volume in growing oocytes harvested from mouse secondary follicles cultured for different days. Compared to D14 + 3, Padjusted(D14 + 5-pS2)=1.55805E-4; Padjusted(D14 + 5-RPB1) = 1.21251E-6; Padjusted(D14 + 5-chromatin-bound RPB1) = 9.15881E-8; Padjusted(D14 + 5-chromatin volume)=0.14783; Padjusted(D14 + 7-pS2)=0.86434; Padjusted(D14 + 7-RPB1) = 0.16675; Padjusted(D14 + 7-chromatin-bound RPB1) = 1.39079E-5; Padjusted(D14 + 7-chromatin volume)=0.20583; Padjusted(D14 + 9-pS2)=5.37076E-4; Padjusted(D14 + 9-RPB1) = 0.06188; Padjusted(D14 + 9-chromatin-bound RPB1) = 0.9796; Padjusted(D14 + 9-chromatin volume)=0. i Immunofluorescence images of oocytes harvested from mouse secondary follicles treated with CB-5083, MG132 and flavopiridol on D14 + 7 for 2 days. Yellow arrowheads highlight the nucleolus. j Automated quantification of total intensity of RPB1 in oocytes harvested from mouse secondary follicles treated with CB-5083, MG132 or flavopiridol(Flavo.) on D14 + 7 for 2 days. Compared to DMSO, Padjusted(CB-5083) = 0; Padjusted(MG132) = 1.07948E-6; Padjusted(flavopiridol)=0.77002. k Manual scoring of chromatin configurations in oocytes harvested from mouse secondary follicles treated with CB-5083 (two-tailed Fisher’s exact test, p = 0.0169), MG132 (two-tailed Fisher’s exact test, p = 0.0569) or flavopiridol. l Immunofluorescence images of NSN and SN oocytes from wildtype and Ubb cKO mice. m Manual scoring of chromatin configurations in oocytes from wildtype and Ubb cKO mice (two-tailed Fisher’s exact test, p = 0.1296). *p < 0.05. ****p < 0.0001. N.S., not significant. Scatter intervals are presented as mean values +/- S.D. Statistical significance is based on one-way ANOVA followed by post-hoc Tukey’s multiple comparison test (b, c, e–h, j). The number of oocytes analyzed is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
To decipher how RNAPII levels decrease during NSN-to-SN transition, we first examined the transcriptional regulation of Rpb1. Briefly, we performed low-input whole-genome bisulfite sequencing (WGBS)97, ATAC-seq98 and fractionated RNA-seq and found that DNA methylation levels, chromatin accessibility, actively transcribing and stored mRNA levels of Rpb1 were not different between NSN and SN oocytes (Supplementary Fig. 7a). Consistent with our multiomics analyses, RT-qPCR confirmed that Rpb1 mRNA levels were only marginally reduced in SN oocytes as compared to NSN oocytes (Supplementary Fig. 7b). We then examined the translational regulation of Rpb1 by Ribo-seq99, poly(A) tail-length assay, RIBOmap100 and CHX treatment. Rpb1 mRNA did not display a significant difference in ribosome occupancy nor poly(A) tail-length (Supplementary Fig. 7a, c). In addition, RIBOmap analysis and CHX treatment confirmed that Rpb1 mRNA was not actively translated in NSN or SN oocytes (Supplementary Fig. 7d–f). Thus, the synthesis of RNAPII protein does not change during NSN-to-SN transition.
To determine whether the depletion of phosphorylated RNAPII is mediated by direct protein degradation during NSN-to-SN transition, we acutely treated ex vivo cultured follicles with inhibitors against segregase and proteasome, two key players of proteolysis in cells. Both segregase inhibitor CB-5083 and proteasome inhibitor MG132 suppressed RNAPII degradation (Fig. 4i, j). Strikingly, they also blocked NSN-to-SN transition (Fig. 4i, k). Interestingly, however, inhibiting RNAPII phosphorylation with the pan-CDK inhibitor flavopiridol could not prevent RNAPII degradation (Fig. 4i, j), suggesting that phosphorylation marks the fraction of RNAPII to be degraded but does not mediate the degradation process.
To investigate whether the degradation of phosphorylated RNAPII is mediated by ubiquitination as previously reported in other systems71, we examined the chromatin configuration of GV oocytes from wildtype and conditional knockout (cKO) mice for Ubb, the predominant polyubiquitin gene that encodes 70% of ubiquitin protein and phenocopies proteasome inhibition upon knockout in oocytes101. While we do not exclude the possibility that the minor amount of ubiquitin protein remaining in Ubb cKO is sufficient for RNAPII degradation, quantifications revealed no significant difference in their NSN/SN ratio (Fig. 4l, m), suggesting that ubiquitination is unlikely to be involved in RNAPII degradation in oocytes. Taken together, RNAPII degradation is necessary for NSN-to-SN transition in vivo.
RNAPII degradation induces NSN-to-SN transition via stripping chromatin-bound RNAPII
To understand how RNAPII degradation induces NSN-to-SN transition, we first compared the levels of RNAPII in oocytes with that in somatic cells, which do not undergo major chromatin reorganization upon RNAPII degradation102,103,104,105,106,107. Immunoblotting revealed that RNAPII in NSN oocyte is around two- and 12-fold more abundant than that in SN oocyte and somatic cell, respectively, predominantly in the phosphorylated form (Fig. 5a). High-resolution imaging further revealed that phosphorylated RNAPII is found on chromatin in NSN oocytes, whereas unphosphorylated RNAPII is found in the nucleoplasm in both NSN and SN oocytes (Fig. 5b), implying that RNAPII degradation primarily removes phosphorylated RNAPII during NSN-to-SN transition.
a Immunoblots of NSN, SN oocytes and HeLa cells. Black arrows mark phosphorylated (IIO), unphosphorylated (IIA) and tailless (IIB) RPB1. b Immunofluorescence images of NSN and SN nuclei. c Pie chart showing the distribution of pS2 CUT&Tag peaks in NSN oocytes. d Metagene plot of pS2 enrichment for genes of different expression levels in NSN oocytes. e Metagene plot of chromatin accessibility for genes of different expression levels in NSN oocytes and in SN oocytes. f Metagene plot of transcription for genes of different expression levels in NSN oocytes. g Still images from time-lapse movies of NSN oocytes treated with water or α-amanitin with tracking of chromocenters. h Automated quantification of MSD of chromocenters in NSN oocytes treated with water, 42B3-t21R, α-amanitin(α-aman.) or triptolide. Padjusted(42B3-t21R)=4.3521E-6; Padjusted(α-amanitin)=5.97982E-6; Padjusted(triptolide)=2.11141E-4. i Still images from time-lapse movies of NSN oocytes injected with water or overexpressing H2B-mClover3 with tracking of chromocenters. j Automated quantification of MSD of chromocenters in NSN oocytes injected with water or overexpressing H2B-mClover3. p = 0.0766. k Immunofluorescence images of α-amanitin-treated NSN oocytes overexpressing H2B-mClover3 or expressing αHIST-αLMN. Yellow arrowheads highlight the nucleolus. l, m Automated quantification of chromatin volume and surface area in α-amanitin-treated NSN oocytes overexpressing H2B-mClover3 or expressing αHIST-αLMN. Padjusted(H2B OE-chromatin volume)=5.42698E-4; Padjusted(αHIST-αLMN-chromatin volume)=0.00546; Padjusted(H2B OE-surface area)=0.89125; Padjusted(αHIST-αLMN-surface area)=5.01899E-5. n Immunofluorescence images of oil-injected NSN oocytes treated with water or α-amanitin. Yellow dotted circles highlight the oil droplet. o Immunofluorescence images of EU-low GV oocytes from wildtype and Npm2 cKO mice. The yellow arrowhead highlights the absence of a nucleolus. p Automated quantification of chromatin volume in GV oocytes from wildtype and Npm2 cKO mice. P(EU-high)=0.0924; P(EU-low)=0.6998. *p < 0.05. ****p < 0.0001. N.S., not significant. Box plots show median(middle line), mean(small squares), 25th and 75th percentiles(boxes), 5th and 95th percentiles(whiskers), and 1st and 99th percentiles(crosses). Unpaired, two-tailed Student’s t test (j, p), and one-way ANOVA followed by post-hoc Tukey’s multiple comparison test (h, l, m). The number of oocytes analyzed or loaded is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
To profile the chromatin binding patterns of phosphorylated RNAPII prior degradation, we optimized low-input CUT&Tag108 (Supplementary Fig. 8a) and found widespread binding of phosphorylated RNAPII to different chromosomes in NSN oocytes (Supplementary Fig. 8b), particularly being enriched at the promoter, gene body and intergenic regions (Fig. 5c). Metagene analyses revealed that genes expressed at higher levels have more RNAPII accumulated at promoters and found within gene bodies (Fig. 5d), consistent with more release of paused RNAPII for actively transcribed genes109,110. Unexpectedly, different from somatic cells109,110 and fertilized eggs94,111, a substantial fraction of phosphorylated RNAPII also accumulated at the 3’ distal intergenic regions (Fig. 5d), coinciding with the unusual open chromatin (Fig. 5e and Supplementary Fig. 8c) and single-stranded DNA signals112 at the transcription termination site (TTS) that are absent in SN oocytes. Because such enrichment did not result in transcription read-through113 (Fig. 5f and Supplementary Fig. 8d), this was likely due to paused RNAPII during transcription termination114.
To test if RNAPII degradation induces NSN-to-SN transition via clearing RNAPII from chromatin, we sought to reduce the levels of chromatin-bound RNAPII without degrading it. To this end, we treated NSN oocytes with THZ1 and DRB, inhibitors of CDK7 and CDK9 that sequentially phosphorylates RNAPII during active transcription cycle115, respectively. As expected, THZ1 was more effective than DRB in inhibiting transcription and reducing phosphorylated RNAPII levels without affecting total RNAPII levels (Supplementary Fig. 9a-d). We found that DRB only partially released chromatin-bound RNAPII concomitant with a partial NSN-to-SN transition, whereas THZ1 fully released chromatin-bound RNAPII concomitant with a full NSN-to-SN transition (Supplementary Fig. 9a, e, f). Importantly, pooling and plotting control, DRB- and THZ1-treated oocytes together revealed a linear relationship between the levels of chromatin-bound RNAPII and chromatin volume (Supplementary Fig. 9g). Taking advantage of DRB as a non-covalent inhibitor, we further investigated whether inducing NSN-to-SN transition alone is sufficient to improve the developmental potential of NSN oocytes. Indeed, significantly more embryos derived from NSN oocytes transiently treated with DRB before meiotic maturation and IVF progressed beyond 2- and 4-cell stages as compared to embryos derived from non-treated NSN oocytes (Supplementary Fig. 9h). Taken together, RNAPII degradation strips chromatin-bound RNAPII to induce NSN-to-SN transition.
Removal of chromatin-bound RNAPII induces a global collapsing force and a local attractive force required for NSN-to-SN transition to take place
Studies in other systems reported that chromatin-bound RNAPII could sterically constrain chromatin movement104,116. Indeed, tracking of chromatin structures such as chromocenters revealed an increase in mean square displacement (MSD) upon the induction of NSN-to-SN transition (Fig. 5g, h), suggesting that the transition to SN configuration involves an increase in chromatin mobility. To directly test the contributions of chromatin dynamics, we sought to reduce chromatin mobility by overloading histone subunit H2B (Fig. 5i, j). Interestingly, upon induction of NSN-to-SN transition, H2B overexpression markedly suppressed the collapse of peripheral chromatin, but without interfering with the formation of the chromatin ring around the nucleolus (Fig. 5k, l, Supplementary Fig. 10a, b and Supplementary Movies 9, 10). This result suggests that the increase in chromatin mobility only contributes in part to the transition to SN configuration, leading us to a model whereby NSN-to-SN transition involves both a collapsing force as a result of the increase in global chromatin dynamics and an attractive force as a result of the nucleolus being converted into an attractive surface for local chromatin attachment.
To demonstrate the presence of the global collapsing force, we designed a histone nanobody-lamin nanobody fusion construct (αHIST-αLMN) to artificially anchor chromatin to the periphery of the nucleus in order to counteract the collapse of chromatin during NSN-to-SN transition. Indeed, expression of αHIST-αLMN suppressed the collapse of peripheral chromatin without interfering with the formation of the chromatin ring around the nucleolus, similar to H2B overexpression (Fig. 5k–m, Supplementary Fig. 10a, c, d, and Supplementary Movies 11, 12). Notably, chromatin detachment from the nuclear envelope61 alone was insufficient to generate a collapsing force, as evidenced by the insignificant changes in chromatin volume upon acute depletion of lamin by miniTrim-Away (Supplementary Fig. 10e–h). Next, to demonstrate the presence of the local attractive force, we attempted to create an artificial nucleolus but with an inert surface by microinjecting a drop of silicone oil into the oocyte nucleus. Prior to inducing NSN-to-SN transition, chromatin surrounded neither the nucleolus nor the oil droplet (Fig. 5n). However, after inducing NSN-to-SN transition, chromatin surrounded the nucleolus but not the oil droplet, concomitant with the displacement of nucleolar proteins such as NCL and B23 from the surface of the nucleolus (Fig. 5n and Supplementary Fig. 10i–k).
To further show that these two forces are independent of each other, we compared the degree of chromatin compaction in EU low (transcriptionally inactive, SN-like) GV oocytes from wildtype and nucleolus-deficient Npm2 cKO mice117,118. Quantifications revealed no significant difference in their chromatin volume (Fig. 5o, p), confirming that the absence of nucleolus abolishes the chromatin ring without affecting the collapse of peripheral chromatin that accounts for the major changes in chromatin volume. Thus, removal of chromatin-bound RNAPII increases chromatin dynamics and triggers nucleolar surface remodeling, resulting in a global collapsing force and a local attractive force required for the transition to SN configuration (Supplementary Fig. 11).
Embryos derived from NSN oocytes have aberrant RNAPII levels and localization and are defective in maternal-to-zygotic transition
Most embryos derived from NSN oocytes had poor developmental competence and were arrested early at 1-cell stage (Fig. 2e), the time when maternal-to-zygotic transition takes place in mice with maternal mRNAs being degraded and replaced by newly transcribed mRNAs during ZGA119,120. We reasoned that the high levels of maternal RNAPII in NSN oocytes may result in aberrant RNAPII levels after meiotic maturation and fertilization121, later contributing to defects in ZGA, a process primarily executed by RNAPII and to a lesser extent RNAPI122. Indeed, immunoblotting confirmed that the undegraded RNAPII contributes to the higher levels of RNAPII in MII oocytes and 1-cell embryos derived from NSN oocytes, independent of de novo translation of RNAPII during meiotic maturation (Fig. 6a). Surprisingly, despite the higher levels of RNAPII, immunofluorescence staining revealed lower levels of total and phosphorylated RNAPII in the nuclei, concomitant with lower transcriptional activities in 1-cell embryos derived from NSN oocytes (Fig. 6b–e). In addition, we observed smaller chromatin volume and excessive nucleolar fragmentation (Fig. 6f–h), indicative of defects and possibly a delay in pronucleus formation in 1-cell embryos derived from NSN oocytes.
a Immunoblots of GV oocytes, MII oocytes and 1-cell embryos derived from NSN and SN oocytes. Black arrows mark phosphorylated (IIO) and unphosphorylated (IIA) RPB1. b Immunofluorescence images of 1-cell embryos derived from NSN and SN oocytes. c, d, e, f Automated quantification of total intensity of RPB1, pS2, EU, and chromatin volume in 1-cell embryos derived from NSN and SN oocytes. P(pS2)=0.0045; P(RPB1) < 0.0001; P(EU) = 0.0253; P(chromatin volume)=0.0007. g Immunofluorescence images of pronuclei of 1-cell embryos derived from NSN, SN, and in vivo ovulated (IVO) oocytes. h Manual scoring of the number of nucleolus in the pronuclei of 1-cell embryos derived from NSN, SN, and IVO oocytes. (two-tailed Fisher’s exact test for male pronuclei, p = 0.0001; two-tailed Fisher’s exact test for female pronuclei, p = 0.6201) i Volcano plot showing differently expressed transcripts in 1-cell embryos derived from NSN and SN oocytes. j Euler diagrams showing the overlap between downregulated genes and minor ZGA transcripts and overlap between upregulated genes and maternal transcripts. k GO analysis of downregulated and upregulated genes not overlapping with minor ZGA transcripts and maternal transcripts. l ATAC-see images of 1-cell embryos derived from NSN and SN oocytes. m Automated quantification of total intensity of ATAC-see signal in 1-cell embryos derived from NSN and SN oocytes. p = 0.1052. n Immunofluorescence images of TSA-treated 1-cell embryos derived from NSN oocytes. o Manual scoring of TSA-treated 1-cell embryos derived from NSN oocytes. (two-tailed Fisher’s exact test for DMSO- and TSA-treated 1-cell embryos derived from NSN oocytes, p = 0.3089). *p < 0.05. **p < 0.01. ***p < 0.001. ****p < 0.0001. N.S., not significant. Box plots show median(middle line), mean(small squares), 25th and 75th percentiles(boxes), 5th and 95th percentiles(whiskers), and 1st and 99th percentiles(crosses). Unpaired, two-tailed Student’s t test (c–f, l). The number of oocytes analyzed or loaded is specified in italics. Scale bars are 5 µm. Source data are provided as a Source Data file.
We further performed low-input RNA-seq to compare 1-cell embryos derived from NSN and SN oocytes (Fig. 6i). Notably, 42 out of 223 downregulated genes were minor ZGA transcripts, whereas 52 out of 168 upregulated genes were maternal transcripts, indicative of defects in minor ZGA and maternal mRNA degradation in 1-cell embryos derived from NSN oocytes (Fig. 6i, j). Gene ontology (GO) analysis further revealed that remaining downregulated genes are mostly related to RNA splicing and cell division, whereas remaining upregulated genes are mostly related to oocyte development, negative regulation of fertilization and catabolic processes (Fig. 6k), suggesting additional nuclear and cytoplasmic defects in 1-cell embryos derived from NSN oocytes.
To determine whether the reduced binding of RNAPII to chromatin and hence lower transcriptional activity could be a result of lower chromatin accessibility94, we performed ATAC-see123 but detected no significant difference in chromatin accessibility between 1-cell embryos derived from NSN and SN oocytes (Fig. 6l,m). Moreover, trichostatin A (TSA) treatment, which could ectopically enhance chromatin accessibility and increase transcriptional activity124, did not rescue the developmental arrest in 1-cell embryos derived from NSN oocytes (Fig. 6n, o). These data suggest defects in zygotic factors that promote RNAPII loading into the nucleus and/or binding to chromatin. Taken together, embryos derived from NSN oocytes are defective in maternal-to-zygotic transition, accounting for their arrest at as early as 1-cell stage and poor development beyond.
Discussion
Our data elucidate the previously unknown mechanism and significance of chromatin reorganization in growing mouse oocytes: RNAPII degradation drives the transition of oocyte chromatin from the NSN to SN configuration, which would otherwise lead to multiple defects after meiotic maturation and fertilization. NSN-to-SN transition and transcriptional silencing are two unique features of the final step of oocyte growth. It is generally believed that independent mechanisms drive these two events in mammalian oocytes89,118,125,126. We challenge this conventional view by showing that RNAPII degradation strips chromatin-bound RNAPII, simultaneously silencing transcription and inducing two forces required for chromatin reorganization to the SN configuration (Supplementary Fig. 11). This elegant coupling mechanism ensures that the silencing of maternal transcription is tightly coordinated with the preparation of maternal chromatin for subsequent embryonic development.
In this study, we systematically investigated many nuclear components and directly tested their potential contributions by mimicking their differences in NSN and SN oocytes using small-molecule inhibition and overexpression. Many of them are epigenetic marks and factors, which have been implicated in mammalian oocyte development. While modulating many of these factors could alter transcriptional activity and/or chromatin compaction in oocytes, they are not sufficient to induce or reverse NSN-to-SN transition. Nevertheless, we do not exclude the possibility that they help to ensure complete transcriptional silencing and/or chromatin compaction in fully grown oocytes50,96,126,127,128,129,130,131. Interestingly, previous studies knocking out a number of proteins such as MSY2/YBX245,90, MATER48, EPAB49, RPS2651, and LSM14B54,55,56,57, which are cytoplasmic factors that have no apparent interactions with RNAPII, could alter the NSN/SN ratio of isolated oocytes. These proteins may have exerted their effects via regulating the stability and/or translation of machineries in the upstream degradation pathway, thus indirectly affecting RNAPII degradation.
Degrading RNAPII using triptolide, α-amanitin or auxin-inducible degron results in little changes in gross chromatin morphology in other systems102,103,104,105,106,107,132, including early mammalian embryos133,134,135,136,137,138. Why does RNAPII degradation have more profound impacts on chromatin in mammalian oocytes than in other systems? Recent studies imply that chromatin-bound RNAPII levels increase with and serve as a limiting factor for cell size139,140. As growing oocytes expand in size and have increasing needs for transcription, they continue to load RNAPII into their nucleus and accumulate more phosphorylated RNAPII on chromatin. Indeed, RNAPII levels in NSN oocytes were around 12.5- and 2-fold higher than those in somatic cells or blastomeres, respectively. This may explain the more profound effects on the chromatin in mammalian oocytes upon the removal of chromatin-bound RNAPII.
Finally, we successfully used scFv for Trim-Away of a non-shuttling nuclear protein. Both fragment antigen-binding (Fab) and its genetically encoded form scFv have been extensively used for imaging endogenous proteins before141,142. scFv has advantages of its smaller size and not artificially clustering antigens over the bivalent IgGs83, and advantages of higher stability and wider availability over nanobodies86. The direct fusion of scFv to t21R also circumvents the need for co-delivering Trim21, opening unprecedented opportunities for using off-the-shelf Fabs or scFvs like 42B385 to study the functions of nuclear proteins in primary or nondividing cells without the need for any genetic manipulation. Although chemical tools like triptolide and α-amanitin are available, they have pleiotropic effects in various systems, and how they induce RNAPII degradation is still largely unclear. The addition of our 42B3-t21R construct to the current toolbox will be invaluable for studying the acute effects of depleting endogenous RNAPII in diverse mammalian species and cell types.
Methods
Data reporting
No statistical methods were used to predetermine the sample size for experiments. Experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. Methods of this research comply with all relevant ethical regulations approved by the National Institute of Biological Sciences, Beijing, Reproductive and Genetic Hospital of CITIC-XIANGYA, and Beijing Perfect Family Hospital.
Preparation and culture of mouse oocytes and follicles
All mice were maintained in a specific pathogen-free environment, including a 12-h light/dark schedule, regulated temperature (approximately 22 °C), and 50–60% relative humidity, at the Animal and Plant Center of the National Institute of Biological Sciences, Beijing, according to international animal welfare rules. Standard chow and water were available without restriction. All experiments with mice complied with the ethical guidelines of the Animal Ethics Committee of the National Institute of Biological Sciences, Beijing. Ubb cKO mice were generated by breeding C57BL/6 JGpt-Ubbem1Cflox/Gpt (GemPharmatech) with B6.Cg-Tg(Msx2-cre)5Rem/BortJ143 (The Jackson Laboratory), then backcrossing with C57BL/6JGpt-Ubbem1Cflox/Gpt. Npm2 cKO mice were generated by breeding C57BL/6JGpt-Npm2em1Cflox/Gpt (GemPharmatech) with B6.Cg-Tg(Msx2-cre)5Rem/BortJ143 (The Jackson Laboratory), then backcrossing with C57BL/6JGpt-Npm2em1Cflox/Gpt.
Oocytes of around 60-80 μm in diameter were isolated from ovaries of 8- to 10-week-old unprimed FVB/N or C57BL/6J (Charles River Laboratories) or Ubb cKO or Npm2 cKO female mice by puncturing preantral and antral follicles on the surface of ovaries with needles. Cumulus-enclosed GV oocytes were stripped by repeated mouth pipetting and arrested at prophase in homemade phenol red-free M2 supplemented with 250 μM dibutyryl cyclic AMP (dbcAMP) (Sigma-Aldrich or MedChemExpress) under paraffin oil (ACROS Organics) at 37 °C. Generally, around 50 oocytes can be isolated from one 8- to 10-week-old unprimed FVB/N, and around 30 oocytes can be isolated from one 8- to 10-week-old unprimed C57BL/6 J female mouse.
Follicles were enzymatically isolated from ovaries of 14-day-old C57BL/6 J female mice in L-15 medium (Sigma-Aldrich) supplemented with 5% fetal bovine serum (FBS) (Sigma-Aldrich), 1x GlutaMAX (Thermo Fisher Scientific), and 0.1x penicillin G/streptomycin (Sigma-Aldrich) containing 1 mg/ml collagenase A (Roche) for 15 min at 37 °C. Intact follicles of around 100-120 μm in diameter with a centered oocyte were cultured in MEM alpha with GlutaMAX and nucleosides (Thermo Fisher Scientific) supplemented with 5% FBS, 0.03 μg/ml ovine follicle stimulating hormone (National Hormone and Peptide Program), 1x insulin/transferrin/sodium selenite (Sigma-Aldrich), and 0.1x penicillin G/streptomycin on 12-mm Transwell-COL collagen-coated 0.4-μm pore polytetrafluoroethylene (PTFE) membrane insert (Corning) at 37 °C and 5% CO2. Half of the medium surrounding the insert was replaced every 3 days. After 3 to 9 days of culture, in vitro grown oocytes were denuded in M2 with dbcAMP.
Preparation, culture and IVF of mouse MII oocytes
For in vitro fertilization of IVO MII oocytes, 4-week-old C57BL/6 J female mice were superovulated by sequential injection of 0.1 ml CARD HyperOva (Cosmo Bio) and 7.5 IU human chorionic gonadotropin (hCG) (ProSpec) after 48 h. Sperm was collected from caudae epididymides from >12-week-old FVB/N male mice and capacitated for 1 h in 400 μl of homemade phenol red-free HTF. MII oocytes were collected from oviducts 15 h after hCG. IVF was performed in 100 μl of HTF by adding 10 μl of 3.33×107/ml capacitated sperm for 3 h. Fertilized eggs were then washed in 1:1 HTF:EmbryoMax Advanced KSOM Medium (Sigma-Aldrich) for 30 min and further transferred to EmbryoMax Advanced KSOM Medium for long-term culture. MII oocytes, 1-cell embryos (2PN), and early and late 2-cell embryos were collected at 0, 7, 22, and 28h post fertilization (hpf), respectively.
For in vitro fertilization of MII oocytes matured from GV oocytes, FVB/N GV oocytes with a centered nucleus were first treated with water/100 μM α-amanitin (APExBIO) or DMSO/100 μM DRB (MedChemExpress) in M2 with dbcAMP for 7 h and then released into dbcAMP-free M2 for 13.5 h. Before the addition of sperm, a single hole was created in the zona pellucida of the MII oocytes using XYRCOS (Hamilton Thorne). IVF was performed as described above. After IVF, 1PN (parthenogenetic activation), 3PN (polyspermy), and abnormal embryos were excluded from further experiments.
Preparation and culture of human oocytes
The use of unfertilized human oocytes in this study was approved by the Institutional Review Board (IRB) at the National Institute of Biological Sciences, Beijing under the reference IRB22101001, Reproductive and Genetic Hospital of CITIC-XIANGYA under the reference LL-SC-2017-012 and Beijing Perfect Family Hospital under the reference 2020-09-08.
Oocytes were collected from patients aged 23–35 years who underwent ovarian stimulation for intracytoplasmic sperm injection (ICSI) as part of their assisted reproduction treatment owing to male factor-associated infertility at the Reproductive and Genetic Hospital of CITIC-XIANGYA and Beijing Perfect Family Hospital. Only oocytes that were GV at the time of ICSI and thus unsuitable for the procedure were used in this study. All patients gave informed consent for their surplus oocyte(s) to be used in this study. For oocytes collected at the Reproductive and Genetic Hospital of CITIC-XIANGYA, oocytes were vitrified using previously described homemade vitrification solutions to maximize their survival and polar body extrusion rate after thawing59. Briefly, oocytes were initially transferred to 20 μl of G-MOPS PLUS (Vitrolife). 20 μl of G-MOPS PLUS containing 7.5% DMSO and 7.5% ethylene glycol (Sigma-Aldrich) was then added and oocytes were incubated for 3 min at room temperature twice. 240 μl of G-MOPS PLUS containing 7.5% DMSO and 7.5% ethylene glycol were further added and oocytes were incubated for 6 to 10 min at room temperature. Before loading onto Cryotop, oocytes were washed in 300 μl of G-MOPS PLUS containing 15% DMSO, 15% ethylene glycol and 0.5 M D-(+)-trehalose (Sigma-Aldrich) twice. For thawing, Cryotop was inserted into 1 ml of prewarmed G-MOPS PLUS containing 1 M D-(+)-trehalose for 1 min at 37 °C. Oocytes were then transferred to 300 μl of G-MOPS PLUS containing 0.5 M D-(+)-trehalose for 3 min at room temperature, 300 μl of G-MOPS PLUS containing 0.25 M D-(+)-trehalose for 5 min at room temperature, and 300 μl of G-MOPS PLUS for 2 min at room temperature. Dead or morphologically abnormal oocytes after thawing were excluded from further experiments, and all other recovered oocytes were transferred to G-MOPS PLUS at 37 °C. For oocytes collected at Beijing Perfect Family Hospital, fresh oocytes were directly transferred to and cultured in G-MOPS PLUS at 37 °C within 3 to 5 h after retrieval from the ovaries. To select for less mature oocytes, only oocytes that were morphologically normal but did not undergo GV breakdown within 24 h after retrieval from the ovaries were used for experiments.
Drug treatment
All drugs except 1,6-hexanediol were prepared in water for embryo transfer or DMSO (Sigma-Aldrich). Oocytes were treated with 20 μM 8-AA (MedChemExpress), 20 μM A-485 (MedChemExpress), 100 μM α-amanitin, 100 nM ActD (MedChemExpress), 6 μM BRM/BRG1i (MedChemExpress), 10 μM bleomycin (MedChemExpress), 10 μg/ml CHX (Sigma-Aldrich), 100 μM DRB, 10 μM DZNep (MedChemExpress), 25 μg/ml etoposide (MedChemExpress), 100 μM GSK-J4 (MedChemExpress), 500 μM hinokiflavone (MedChemExpress), 10 μM quarfloxin (MedChemExpress), 2.5 μM THZ1 (MedChemExpress), 10 μM triptolide (MedChemExpress), 100 nM TSA (MedChemExpress) or 10 μM 7-DAA (MedChemExpress) in M2 with dbcAMP for 7 h. For 1,6-hexanediol treatment, oocytes were treated with 15% 1,6-hexanediol (Sigma-Aldrich) in M2 with dbcAMP for 10 min. For CHX treatment, oocytes were treated with 10 μg/ml CHX in M2 with dbcAMP for 24 h. For CB-5083, MG132, and flavopiridol treatments, cultured follicles were treated with 5μM CB-5083 (MedChemExpress), 25μM MG132 (MedChemExpress), or 5μM flavopiridol (MedChemExpress) on D14+7 for 48h. Embryos were treated with 5 nM TSA in EmbryoMax Advanced KSOM Medium for 16 h at 7 hpf.
Expression constructs and mRNA synthesis
To generate constructs for mRNA synthesis, we fused cDNAs with eGFP, mClover3, P2A, NLS(NP), t21R and/or V5 and subcloned them into pGEMHE to obtain 42B3-mClover3, 42B3-mClover3(11)-P2A-NLS-mClover3(1-10) (42B3-split mClover3), 42B3-t21R, mClover3-CTCF, DNMT1-V5, DNMT3A-V5, DNMT3B-V5, DNMT3L-V5, NLS-eGFP-hnRNPU, HP1β, αLMN-t21R, NLS-eGFP-MATR3, PAGAP-t21R, TET1-V5, TET2-V5 and TET3-V5. Histone-Chromobody and Lamin-Chromobody coding sequences (Chromotek) were fused and subcloned into pGEMHE to obtain αHIST-αLMN. pGEMHE-H2B-mClover3144 was also used. All mRNAs were synthesized and quantified as previously described84.
Microinjection
Mouse oocytes were microinjected with 3.5 to 14 pl of mRNAs as previously described84. 42B3-mClover3 mRNA was microinjected at a needle concentration (final concentration in the microinjection needle) of 560 ng/μl, 42B3-split mClover3 mRNA at 560 ng/μl, 42B3-t21R mRNA at 2965.9 ng/μl, mClover3-CTCF mRNA at 3040 ng/μl, DNMT1-V5 mRNA at 572.4 ng/μl, DNMT3A-V5 mRNA at 701 ng/μl, DNMT3B-V5 mRNA at 752.2 ng/μl, DNMT3L-V5 mRNA at 1057 ng/μl, NLS-eGFP-hnRNPU mRNA at 2251.2 ng/μl, H2B-mClover3 mRNA at 1145.8 ng/μl, αHIST-αLMN mRNA at 1584.2 ng/μl, HP1β mRNA at 952.5 ng/μl, αLMN-t21R at 2000 ng/μl, NLS-eGFP-MATR3 mRNA at 1148.1 ng/μl, PAGFP-t21R mRNA at 2000 ng/μl, TET1-V5 mRNA at 801.9 ng/μl, TET2-V5 mRNA at 528.9 ng/μl and TET3-V5 mRNA at 469.3 ng/μl. Oocytes were allowed to express the mRNAs for 3–4 h. Mouse oocytes were also microinjected with 7 pl of RNase A (Sigma) into the cytoplasm or 1 to 2 pl of dimethylsiloxane (Sigma-Aldrich) into the nucleus. Human oocytes were microinjected with 14 pl of 42B3-t21R mRNA at a needle concentration of 2965.9 ng/μl as previously described59.
Live staining of GV oocytes for visualizing chromatin configuration
SPY555-DNA (Spirochrome) and 5-SiR-Hoechst59 were reconstituted in DMSO. For visualizing chromatin configuration, oocytes were stained with SPY555-DNA or 5-SiR-Hoechst for 30 min at 1:500 dilution and 10 μM, respectively. For live imaging, SPY555-DNA or 5-SiR-Hoechst was present throughout imaging at 1:1000 dilution and 50 nM, respectively. For sorting GV oocytes with different chromatin configurations, individual oocytes were transferred to microdrops and manually categorized under an inverted widefield fluorescence microscope equipped with a fluorescent LED. GV oocytes exhibiting dispersed chromatin with a few bright punctate were categorized as NSN oocytes, whereas those displaying highly condensed chromatin with a complete perinucleolar ring were categorized as SN oocytes. Oocytes that could not be unambiguously assigned to either category were excluded from further experiments. NSN and SN oocytes were used for experiments immediately after sorting.
Immunofluorescence staining
Oocytes and embryos were fixed in 100 mM HEPES (pH 7.0, titrated with KOH), 50 mM EGTA (pH 7.0, titrated with KOH), 10 mM MgSO4, 2% methanol-free formaldehyde, and 0.5% triton X-100 for 15 to 60 min at 37 °C and washed in phosphate-buffered saline (PBS). For EU and L-AHA labeling, oocytes were pulsed with 1 mM EU (Thermo Fisher Scientific) or 100 μM L-AHA (Click Chemistry Tools) in M2 for 2 h at 37 °C before fixation and visualized using Click-iT RNA Alexa Fluor 488 Imaging Kit (Thermo Fisher Scientific) or AFDye 568 Azide (Click Chemistry Tools) or Click-iT HPG Alexa Fluor 488 Protein Synthesis Assay Kit (Thermo Fisher Scientific). For 5mC and 5hmC staining, fixed oocytes were treated with 4 M HCl for 10 min and neutralized with 100 mM Tris-HCl (pH 7.5) for 10 min. All antibody incubations and washings were performed in PBS with 0.5% triton X-100 (PBT) and 5% BSA (PBT-BSA) at 10 μg/ml overnight at 4 °C (for primary antibodies) and at 20 μg/ml for 1 h at room temperature (for secondary antibodies). Primary antibodies used were mouse anti-B23 (Santa Cruz Biotechnology #sc-56622), rabbit anti-CTCF (Sigma-Aldrich #07-729-25UL), mouse anti-5mC (Active Motif #39649), rabbit anti-5hmC (Active Motif #39792), rabbit anti-FBL (Abcam #ab166630), goat anti-GFP (Rockland Immunochemicals #600-101-215), mouse anti-γH2A.X (Sigma-Aldrich #05-636), rabbit anti-H3ac (Active Motif #39140), rabbit anti-H4ac (Sigma-Aldrich #06-598), rabbit anti-hnRNPU (Huabio #ET7107-10), rabbit anti-HP1α (Huabio #ET1602-8), mouse anti-HP1β (Sigma-Aldrich #MAB3448), rabbit anti-HP1γ (Huabio #ET1706-02), rabbit anti-H3K9me3 Abcam #ab8898), rabbit anti-KMe2 #1 (Abcam #ab7315), rabbit anti-KMe2 #2 (Abclonal #A5870), rabbit anti-KMe2 #3 (Abclonal #A18296), rabbit anti-KMe3 #1 (Thermo Fisher Scientific #PA5-116816), rabbit anti-KMe3 #2 (Abclonal #A20145), mouse anti-LMNAC (Sigma-Aldrich # SAB4200236), mouse anti-m3G/TMG (MBL #RN019M), rabbit anti-MATR3 (Huabio #ET7106-95), rabbit anti-NCL (Abclonal #A20910), rabbit anti-NuMA (Abcam #ab97585), rabbit anti-POLR1A (CST #24799), rabbit anti-POLR2B(Active motif #61558), rabbit anti-RPB1 (CST #149158S), mouse anti-CTD pY1 (Active Motif #91220), rat anti-CTD pS2 (Active Motif #61999), rabbit anti-CTD pT4 (Abclonal #AP1383), rat anti-CTD pS5 (Active Motif #67102), rat anti-CTD pS7 (Active Motif #61704), mouse anti-SRSF2 (Sigma-Aldrich #S4045), rabbit anti-SMARCA2 (Abclonal #A23291), rabbit anti-SMARCA4 (Abclonal #A19556), rabbit anti-SMC3 (Abcam #Ab128919) and rabbit anti-SMC4 (Sigma-Aldrich #HPA029449). Secondary antibodies used were Alexa Fluor 488-, 594-, or 647-conjugated AffiniPure Fab Fragment anti-goat IgG, anti-rabbit IgG, and anti-rat IgG (Jackson ImmunoResearch Europe), Alexa Fluor 568-conjugated Nano-Secondary anti-mouse IgG1, IgG2a and IgG2b (Proteintech). DNA was stained with Hoechst 33342 (Molecular Probes).
Mouse ovaries were fixed in 100 mM HEPES (pH 7.0, titrated with KOH), 50 mM EGTA (pH 7.0, titrated with KOH), 10 mM MgSO4 and 2% methanol-free formaldehyde for 30 minutes at room temperature. They were then washed in PBS, cryoprotected in 30% sucrose overnight at room temperature, and embedded in OCT. 30 μm-thick cryosections were used for immunostaining. After removing OCT with PBS, ovary sections were blocked with PBT-BSA for 1 hour. Slides were then incubated with primary antibodies at 4 °C overnight. Following three 15-minute washes with PBT, slides were incubated with secondary antibodies for 1 hour at room temperature. DNA was stained with Hoechst 33342, and slides were mounted with 50% glycerol.
polyA RNA-FISH
Oocytes were fixed in 100 mM HEPES, 50 mM EGTA, 10 mM MgSO4, 2% methanol-free formaldehyde, and 0.5% triton X-100 for 15 to 60 min at 37 °C and washed in PBS. Oocytes were then washed with Wash Buffer A (Biosearch Technologies) containing 10% formamide and incubated with 200 nM Alexa Fluor 647-conjugated oligo dT30 (IDT) in hybridization buffer (Biosearch Technologies) supplemented with 10% formamide overnight at 37 °C. After probe incubation, oocytes were washed with Wash Buffer A for 30 min at 37 °C and stained with Hoechst 33342 in Wash Buffer A for another 30 min at 37 °C. Oocytes were further washed with Wash Buffer A and Wash Buffer B (Biosearch Technologies) for 30 min at 37 °C and room temperature, respectively, before imaging.
RIBOmap
RIBOmap was performed as previously described100. Oocytes were fixed in 100 mM HEPES, 50 mM EGTA, 10 mM MgSO4 and 2% methanol-free formaldehyde for 15 to 60 min at 37 °C and washed in PBS. Oocytes were then washed three times with hybridization buffer consisting of 2x SSC, 10% formamide, 0.1% triton X-100, 0.1 mg/ml yeast tRNA (Thermo Fisher Scientific) and 0.2 U/μl RiboLock RNase Inhibitor (Thermo Fisher Scientific). Hybridization solution was prepared by diluting all probes (Genewiz), including 5 splints (5’-ACAAAATAGAACCGCGGTCCTATTCAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA TAT CTT TAG T*G*T* /3InvdT/-3’; 5’-CATCGTTTATGGTCGGAACTACGACAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA TAT CTT TAG T*G*T* /3InvdT/-3’; 5’-AGGTTTCCCGTGTTGAGTCAAATTAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA TAT CTT TAG T*G*T* /3InvdT/-3’; 5’-TGTTATTGCTCAATCTCGGGTGGCTAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA TAT CTT TAG T*G*T* /3InvdT/-3’; 5’-AGATAGTCAAGTTCGACCGTCTTCTAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA AAA TAT CTT TAG T*G*T* /3InvdT/-3’), 5 Rpb1-specific padlocks (5’-/5Phos/AAGATA AACATCGTAGACTA GCTACAAAAAGCCTGCGCC TCAGGTCAT ACACTA-3’; 5’-/5Phos/AAGATA AACATCGTAGACTA TGTCAATATTGAAGGGTGTGACAATCT TCAGGTCAT ACACTA-3’; 5’-/5Phos/AAGATA AACATCGTAGACTA TCTATTACATCTTGTTTAGCTTTCTTAATAGTGT TCAGGTCAT ACACTA-3’; 5’-/5Phos/AAGATA AACATCGTAGACTA GTGAAGATCTCCACAATATCATTGGACG TCAGGTCAT ACACTA-3’; 5’-/5Phos/AAGATA AACATCGTAGACTA TGGTGAGGGGATGTATGGGC TCAGGTCAT ACACTA-3’) and 4 Rpb1-specific primers (5’-CTGCGCAGGCGCAAACC GTGTCTACGATG-3’; 5’-CCCCTGCGTACTAATTCCTGAAG GTGTCTACGATG-3’; 5’-CTCATTGTTATGAGCCTTCTCAATGA GTGTCTACGATG-3’; 5’-CACAGCCTCAATGCCCAGTA GTGTCTACGATG-3’; 5’-AGCTGGGAGACATAGCACCA GTGTCTACGATG-3’), to a final concentration of 50 nM with hybridization buffer. Oocytes were incubated with hybridization solution for 4 to 6 h at 40 °C. After hybridization, oocytes were washed twice with PBS with 0.1% triton X-100 and 0.1 U/μl RiboLock RNase Inhibitor (PBSTR) and once with 4x SSC diluted with PBSTR. All washes were performed for 10 min at 37 °C. Oocytes were then rinsed with PBSTR and incubated for 1.5 h at room temperature with ligation solution consisting of 0.25 Weiss U/μl T4 DNA ligase (Thermo Fisher Scientific), 1x T4 DNA ligase buffer, 0.5 mg/ml BSA, and 0.4 U/μl RiboLock RNase Inhibitor. After two washes with PBSTR for 5 min each, oocytes were incubated with amplification-labeling mixture consisting of 0.5 U/μl phi29 DNA polymerase (NEB), 1x phi29 DNA polymerase reaction buffer, 250 μM dNTP (NEB), 0.5 mg/ml BSA, 0.4 U/μl RiboLock RNase Inhibitor, and 1 μM labeling probe (5’-/5Cy3/CATACACTAAAGATAACAT-3’) (Genewiz) for 2 h at 30 °C. Oocytes were subsequently washed twice with PBSTR with 0.05 mg/ml BSA for 5 min each at room temperature before imaging.
ATAC-see
ATAC-see was performed as previously described123 after minor modifications. TAMRA-labeled oligonucleotides (Tn5MErev: 5’-[phos]CTGTCTCTTATACACATCT-3’; Tn5ME-A-TAMRA: 5’-/TAMRA/TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG-3’; Tn5ME-B-TAMRA: 5’-/TAMRA/GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAG-3’) (Ruiboxingke Biotechnology) were resuspended in water to a final concentration of 100 μM, and equimolar of Tn5MErev/Tn5ME-A-TAMRA and Tn5MErev/Tn5ME-B-TAMRA were mixed, denatured for 5 min at 95 °C and annealed by slowly cooling down to room temperature. Tn5 transposome was assembled by mixing 0.602 μl of Tn5MErev/Tn5ME-A-TAMRA, 0.602 μl of Tn5MErev/Tn5ME-B-TAMRA, 0.577 μl of 2× dialysis buffer [100 mM HEPES–KOH (pH 7.2), 0.2 M NaCl, 0.2 mM EDTA, 2 mM DTT, 0.2% Triton X-100 and 20% glycerol] and 3.221 μl of 50 μM Tn5 (ActiveMotif #81286) and incubating for 1 h at room temperature.
Embryos were fixed in 100 mM HEPES, 50 mM EGTA, 10 mM MgSO4, 2% methanol-free formaldehyde, and 0.5% triton X-100 for 15 to 60 min at 37 °C and washed in PBS. Embryos were then incubated with 4 μl of Tn5 transposome diluted with 10 μl of 2x TD buffer and 6 μl of water for 1 h at 37°C. After the tagmentation reaction, embryos were washed with PBS containing 0.01% SDS and 50 mM EDTA for 15 min at 55 °C thrice. Embryos were further washed with PBT-BSA before staining with Hoechst 33342 for 1 h at room temperature.
Confocal and light-sheet microscopy
For confocal imaging, fixed oocytes and embryos were imaged in 2 μl of medium (for live imaging) or PBS with 1% polyvinylpyrrolidone (PVP) and 0.5 mg/ml BSA (for fixed imaging) under paraffin oil in a 35-mm dish with a no. 1.5 coverslip (iBidi). Images were acquired with LSM 900 (Zeiss), LSM 980 (Zeiss), and Stellaris 5 (Leica) confocal laser scanning microscopes equipped with a 40x LD C-Apochromat 1.1 NA water-immersion objective (Zeiss), 40x C-Apochromat 1.2 NA water-immersion objective (Zeiss), or a 63x Plan-Apochromat 1.4 NA oil-immersion objective (Leica). Hoechst was excited with a 405-nm laser line and detected at 410 to 483 nm. mClover3 and Alexa Fluor 488 were excited with a 488-nm laser line and detected at 493 to 556 nm. Alexa Fluor 568 and Alexa Fluor 594 were excited with a 561-nm laser line and detected at 566 to 635 nm. Alexa Fluor 647 and 5-SiR-Hoechst were excited with a 633- or 640-nm laser line and detected at 645 to 700 nm.
For light-sheet imaging, oocytes were imaged in 0.5 ml of medium in four different compartments of a multiwell sample holder (Viventis Microscopy Sàrl). Dual-view images were acquired with LS2 Live (Viventis Microscopy Sàrl) equipped with an environmental incubator box and two 25×1.1 NA water-dipping objectives (Nikon). SPY555 was excited with a 561-nm laser line and detected with a 523/20 nm – 610/25 nm dual band-pass filter. Dual-view images were processed with a fusion script (Viventis Microscopy Sàrl) after acquisition.
Images of the control and experimental groups were acquired under identical imaging conditions on the same microscope. For some images, shot noise was reduced with a Gaussian filter. Care was taken that the imaging conditions (laser power, pixel-dwell time, detector gain, and exposure time) did not cause phototoxicity (for live imaging), photobleaching, and saturation.
Image quantification
To reproducibly segment chromatin (labeled with Hoechst 33342 or SPY555-DNA) or other nuclear structures (labeled with antibodies against different nuclear components), the Machine Learning Segmenter was used for automatic segmentations in Arivis (Zeiss). For each experiment, 3 to 5 z-planes from 5 to 10 cells or timepoints were manually annotated for foreground and background pixels and subjected to model training under the Fluorescence Fast mode. Suitable smoothing and threshold values were selected to refine the segmentation results before batch applications. Tracking in Arivis was used for tracking segmented chromocenters. Specific parameters used were: Brownian Motion (Centroid) (for Motion type), Center of Geometry (for Centroid), 3 µm (for Max. distance), and 2 (for Max. time gap). Total fluorescence intensity, volume, surface area and MSD of the segmented objects were exported into Excel (Microsoft) and OriginPro (OriginLab) for further processing.
Immunoblotting
40 mouse oocytes or embryos (per lane) were extensively washed in protein-free medium and snap-frozen in 1 μl of protein-free medium in liquid nitrogen. Before thawing, 7 μl of water and 4 μl of 4x NuPAGE LDS sample buffer (Thermo Fisher Scientific) with 100 mM dithiothreitol (DTT) were added. Samples were then thawed at 37 °C and snap-frozen in liquid nitrogen twice more before being heated for 5 min at 99 °C. For λPP treatment, oocytes were lyzed in 8 μl of buffer (1X NEBuffer for protein metallophosphatase and 1 mM MnCl2) by snap-freezing in liquid nitrogen and thawing at 37 °C three times, and incubated for 30 min at 30 °C after adding 0.16 μl of 400 U/μl λPP. Samples were resolved on a 15-well NuPAGE 8 to 16% Tris-Glycine Plus WedgeWell Gel of 1.0 mm thickness (Thermo Fisher Scientific) with Tris-Glycine SDS Running Buffer (Thermo Fisher Scientific or Beyotime). Proteins were transferred onto a methanol-activated 0.45-μm polyvinylidene difluoride (PVDF) membrane (Thermo Fisher Scientific) with SDS-free Towbin buffer at 200 mA for 2 h on ice. Blots were stained with No-Stain Protein Labeling Reagent (Thermo Fisher Scientific) before blocking. Blocking and antibody incubations were performed in Tris-buffered saline (TBS) with 5% skim milk and 0.1% tween-20. Primary antibodies used were rabbit anti-RPB1 (CST #149158S), mouse anti-CTD pY1 (Active Motif #91220), rat anti-CTD pS2 (Active Motif #61699), rabbit anti-CTD pT4 (CST #26319S) and mouse anti-CTD pS5 (4H8) (Santa Cruz sc-47701). Secondary antibodies used were horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (CST #7074P2), goat anti-rat IgG (CST #7077S) and horse anti-mouse IgG (CST #7076P2). Blots were developed with SuperSignal West Femto Maximum Sensitivity Substrate (Thermo Fisher Scientific) and documented with Amersham ImageQuant 500 (Cytiva). Care was taken that the exposure time did not cause saturation.
RT-qPCR
40 mouse oocytes (per sample) were extensively washed in protein-free medium and snap-frozen in 1 μl of protein-free medium in liquid nitrogen. Total RNA was extracted with TRIzol LS Reagent (Thermo Fisher Scientific) in Phasemaker Tube (Thermo Fisher Scientific) and treated with TURBO DNA-free Kit (Thermo Fisher Scientific). Reverse transcription (RT) was performed using SuperScript IV First-Strand Synthesis System (Thermo Fisher Scientific), and quantitative PCR (qPCR) was performed using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad) on CFX Opus 96 Real-Time PCR System (Bio-Rad). Endogenous Rpl13a was used as an internal control to calculate the relative transcript level. For amplifying Rpl13a, the following primers were used: 5’-GAGGTCGGGTGGAAGTACCA-3’ and 5’-TGCATCTTGGCCTTTTCCTT-3’. For amplifying Rpb1, the following primers were used: 5’-ACATGTGCAGGAAACATGAC-3’ and 5’-CTTCGGGTTATTAGAATCTACAAGC-3’.
Poly(A) Tail-Length Assay
40 mouse oocytes (per sample) were extensively washed in protein-free medium and snap-frozen in 1 μl of protein-free medium in liquid nitrogen. Total RNA was extracted with TRIzol LS Reagent in Phasemaker Tube and treated with TURBO DNA-free Kit. G/I tailing, RT and end-point PCR were performed according to the manual of the Poly(A) Tail-Length Assay Kit (Thermo Fisher Scientific). The poly(A) start site (PAS) fragment was obtained by PCR using the Rpb1-specific forward primer (5’- TTGGTGCCTGCTCTGG-3’) and the Rpb1-specific reverse primer (5’-TGGTCAAAATTAGTAAACTTTATTTCAATTTCAAAAAATAACAAA-3’) located immediately upstream of the PAS. The tail fragment was obtained by PCR using the same Rpb1-speicfic forward primer and a 35 nt universal reversal primer provided in the kit. The PCR products were resolved on a 2.5% agarose gel, and the poly(A) tail length was calculated by (size of tail fragment) – (size of PAS fragment) – 35.
Hi-C, data processing and analysis
Chromatin digestion was performed as previously described69 after minor modifications. 100 mouse oocytes (per sample) were fixed in PBS with 0.05% PVP and 1% methanol-free formaldehyde for 10 min at RT and 2.5 M glycine was added to a final concentration of 0.2 M to quench for 5 min at RT. Oocytes were transferred to 100 μl of PBS with 0.05% PVP and captured by adding 10 μl of washed ConA Beads Pro (Vazyme) for 10 min at RT. Oocytes were lysed in 12.5 μl of Hi-C lysis buffer [10 mM Trim-HCl (pH 8.0), 10 mM NaCl, 0.5 % NP-40] supplemented with cOmplete, Mini, EDTA-free Protease Inhibitor Cocktail (Roche) for 15 min on ice. Supernatants were discarded with 2.25 μl remaining, and 0.25 μl of 5% SDS was added before incubating for 10 min at 62 °C. 7.25 μl of nuclease-free water and 1.25 μl of 10% Triton X-100 were further added to quench for 15 min at 37 °C, and 1.25 μl of 10x NEBuffer 2 (NEB) and 5 U of MboI (NEB) were added to digest chromatin overnight at 37 °C. The next day, samples were incubated for 20 min at 62 °C to inactivate MboI and then cooled to RT. 2.5 μl of fill-in master mix [7.5 μl of 0.4 mM biotin-14-dATP (Active Motif), 0.3 μl of 10 mM dCTP, 0.3 μl of 10 mM dGTP, 0.3 μl of 10 mM dTTP and 1.6 μl of 5 U/μl of DNA polymerase I, Large (Klenow) Fragment (NEB)] was added to fill in restriction fragment overhangs and mark DNA ends with biotin, and samples were incubated for 90 min at 37 °C. Ligation master mix [33.2 μl of nuclease-free water, 11 μl of 10x T4 DNA Ligase buffer (NEB), 5 μl of 10% Triton X-100, 0.6 μl of 100x BSA and 0.5 μl of 2000 U/μl T4 DNA Ligase (NEB)] was further added, and samples were incubated for 4 h at RT. After removing supernatants, beads were resuspended with 15.5 μl of nuclease-free water, 1.75 μl of Lambda Exonuclease Buffer (NEB), 0.15 μl of Lambda Exonuclease (NEB) and 0.15 μl of Exonuclease I (NEB)] and incubated for 1 h at 37 °C to remove unligated ends. To release DNAs, 42.5 μl of nuclease-free water, 6 μl of 10% SDS and 2.5 μl of 20 mg/ml Proteinase K (Thermo Fisher Scientific) were added, and samples were incubated for 30 min at 55 °C. To reverse crosslinks, 6.5 μl of 5 M NaCl was added, and samples were incubated overnight at 68 °C. The next day DNAs were extracted with phenol-chloroform in Phasemaker Tube, precipitated with ethanol and resuspended in 5.5 μl of nuclease-free water. Library preparation was performed using the TruePrep DNA Library Prep Kit V2 for Illumina (Vazyme) after minor modifications. DNA fragmentation mix was scaled down to 10 μl. After tagmentation, the biotin pull-down process was begun by washing 12.5μl of 10mg/ml Sera-Mag SpeedBeads Neutravidin-Coated Magnetic Particles (Cytiva) or Dynabeads MyOne Streptavidin C1 (Thermo Fisher Scientific) with 40μl of 1x TWB [5mM Tris-HCl (pH 7.5), 1M NaCl, 0.5mM EDTA, and 0.05% Tween-20]. After resuspension in 25 μl of 2x BB [10 mM Tris-HCl (pH 7.5), 2 M NaCl, and 1 mM EDTA], beads were added to samples and incubated for 15 min at RT. Before PCR, beads were washed with 40 μl of 1x TWB twice and resuspended in 29 μl of nuclease-free water. The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit (Thermo Fisher Scientific) and qPCR, analyzed by a Fragment Analyzer System (Agilent) to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System (Illumina).
Reads were processed using HiC-Pro (v3.1.0) with default parameters in the HiC-Pro configuration file to obtain valid contacts. The “.bed” file of restriction fragments from MboI digestion was generated using the “digest_genome” Python script from HiC-Pro. Reads were initially mapped to mm10 (GRCm38.p6) reference genome, and unmapped reads were subsequently cut at restriction sites and re-mapped. After removing PCR duplicates and reads lacking restriction sites, contact matrix files were generated from the final set of valid contacts, which were then normalized by the ICE method. For visualization, the “hicpro2juicbox” shell script was employed to convert valid contacts into “.hic” files, which were then imported into Juicebox (v.2.20). For calculation of contact probability within chromosomes, valid contacts were converted into “.cool” files using Cooler (v0.9.3) and Cooltools (v0.6.1), and the resulting contact probability curves were smoothed and aggregated for plotting.
WGBS, data processing and analysis
30 mouse oocytes (per sample) were lyzed in 20 μl of lysis buffer consisting of 1x M-Digestion Buffer and 1 mg/ml Proteinase K (Zymo Research) for 20 min at 50 °C. Bisulfite conversion and library preparation were performed using the Pico Methyl-Seq Library Prep Kit (Zymo Research) after minor modifications. In the amplification with PrepAmp Primer step, PrepAmp primer was diluted from 40 μM to 20 μM. In the library amplification step, 10 cycles were used. The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit and qPCR, analyzed by a Fragment Analyzer System to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System.
Reads were processed using TrimGalore (v0.6.7) to remove adaptor and low-quality reads with parameters “-q 20, -e 0.1, --length 20, --stringency 1”, and then trimmed reads were mapped to mm10 (GRCm38.p6) reference genome using Bsmap (v2.90) with default parameters. Methylation level of each CpG site was estimated using the function “mcall” of Moabs (v1.3.9.6). Methylation level of each CpG site from multiple replicates was determined using total methylated read count across replicates versus total read count across replicates, ad CpG sites with less than 3 reads were discarded.
ATAC-seq, data processing and analysis
Library preparation was performed as previously described98 after minor modifications. 50 mouse oocytes (per sample) were treated with acidic Tyrode’s solution (Sigma-Aldrich) to remove the zona pellucida and transferred into 500 μl of ATAC-Resuspension Buffer [10 mM Tris-HCl (pH 7.4), 10 mM NaCl and 3 mM MgCl2] containing 0.1% NP-40, 0.1% tween-20 and 0.01% digitonin to lyze for nuclei using a glass capillary with <50 μm opening. The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit and qPCR, analyzed by a Fragment Analyzer System to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System.
Reads were processed using TrimGalore (v0.6,7) to remove adaptor and low-quality reads with parameters “-q 20, -e 0.1, --length 20, --stringency 1”, and then trimmed reads were mapped to mm10 (GRCm38.p6) reference genome using Bowtie2 (v2.4.2) with parameters “-L 25, -X 2000, --no-mixed --no-discordant”. Unmapped reads and low mapping quality reads (Q < 30) were discarded, and PCR duplicates were removed using the function “MarkDuplicates” from Picard tools (v2.27.2). ATAC-seq bigwig tracks were generated using the function “bamcoverage” from Deeptools (v3.5.1) with parameters “-binSize 50 -normSize 2494787188 -exactScaling -extendReads -normalizeUsing RPKM”. ATAC signal counts in each genomic region were computed using the function “computeMatrix” from Deeptools (v3.5.1) with parameters “-bs 200, --upstream 10000, --downstream 10000”.
RNA-seq, data processing and analysis
Karyoplast and cytoplast were separated by micromanipulation as previously described25. 5 μl of Dynabeads MyOne Streptavidin C1 was washed with 20 μl of Dynabead solution A (50 mM NaCl and 0.1 M NaOH) twice and 20 μl of Dynabead solution B (100 mM NaCl). Dynabeads were then resuspended in 5μl of 2x B&W buffer [10mM Tris-HCl (pH 7.5), 2M NaCl, and 1mM EDTA], and 5μl of 100μM biotinylated oligo-dT30VN (Sangon Biotech) was added for 20min at room temperature. Dynabeads were further washed with 20 μl of 1x B&W buffer four times and resuspended in 10 μl of nuclease-free water. 25 karyoplasts or 3 cytoplasts (per sample) were lyzed in 5 μl of lysis buffer [10 mM Tris-HCl (pH 7.5-8.0), 4% NP-40 and 0.2% SDS], and 0.24 μl of 5 M NaCl and 1 μl of Dynabeads were added for 20 min at room temperature. The supernatant was collected as the polyA-ve fraction, and 1 μl of 105-diluted ERCC RNA Spike-In Mix (Thermo Fisher Scientific) and 6.6 μl of nuclease-free water were added. Dynabeads were resuspended in 5 μl of lysis buffer as the polyA+ve fraction, and 1 μl of 105-diluted ERCC RNA Spike-In Mix and 7.6 μl of nuclease-free water were added. Library preparation was performed using the Ovation SoLo RNA-Seq System (Tecan) after minor modifications. In the primer annealing step, 1 μl of 50 μM random hexamer primer (Thermo Fisher Scientific) was added instead of 2.6 μl of First Strand Primer Mix. The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit and qPCR, analyzed by a Fragment Analyzer System to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System.
For RNA-seq of 2PN embryos, RNA libraries were constructed using the SMART-Seq Total RNA Pico Input Kit with UMI (Takara), following the manufacturer’s instructions with minor modifications. 3 embryos at 16 hpf were pooled as a sample. During the first PCR amplification step of library preparation, 5 cycles were performed, followed by 25 cycles in the second PCR amplification step. The resulting libraries were purified, quantified using the Qubit 1X dsDNA HS Assay Kit and qPCR, and assessed for quality and successful amplification using a Fragment Analyzer System. Libraries were then sequenced on the NovaSeq X Plus Sequencing System.
Reads were processed using TrimGalore (v0.6.7) to remove adaptor and low-quality reads with parameters “-q 20, -e 0.1, --length 10, --stringency 1”, and the reads with UMI barcode were trimmed with parameters “-q 20, -e 0.1, --length 10, --stringency 1 –clip_R2 14”. Then trimmed reads were mapped to mm10 (GRCm38.p6) reference genome plus ERCC.fasta using STAR (v2.7.10a) with default parameters. Multi-mapping reads were filtered using Samtools (v1.17), and PCR duplicates were removed using the function “MarkDuplicates” from Picard tools (v.2.27.2). Deduplicated reads were mapped to genes using HTseq (v2.0.3) as unstranded samples with default parameters. A gene/ERCC count was considered valid when present in at least five reads in at least two samples. Differentially expressed genes between groups of samples were identified using DESeq2 (v1.38.0). Gene counts of each sample were normalized by defining ERCC genes as the “controlGenes” when estimating the “sizeFactors” with DESeq2. For samples without ERCC, the gene counts of each sample were directly analyzed using the function ‘DEseq’ in DEseq2. The differentially expressed genes were identified based on an FDR cutoff of 0.05 and a log2FoldChange cutoff of 0.05.
For RNA-seq of 2PN embryos, differentially expressed genes were cross-referenced with minor ZGA transcripts (clusters IV, and V) or maternal transcripts (cluster II) from a previous study145 to calculate the overlap between minor ZGA transcripts and genes downregulated in NSN-derived 2PN embryos, as well as between maternal transcripts and genes upregulated in NSN-derived 2PN embryos. The non-overlapped genes further underwent GOBP enrichment analysis with Metascape146, and enrichment results were visualized as dot plots.
Ribo-seq, data processing and analysis
Library preparation was performed as previously described99 after minor modifications. 50 mouse oocytes (per sample) were lysed in 150μl of ice-cold lysis buffer [20mM Tris-HCl (pH 8.0), 150mM NaCl, 5mM MgCl2, 1% Triton X-100, and 1mM DTT] for 5min on ice, followed by snap-freezing in liquid nitrogen and thawing at 37°C three times. After spinning at 21300 xg for 5 min at 4 °C, the supernatant was digested with 0.25 μl of 2 U/μl TURBO DNase (Thermo Fisher Scientific) and 0.2 μl of 1:10-diluted 20 U/μl RNase T1/0.5 U/μl RNase A cocktail (Thermo Fisher Scientific) for 30 min at 37 °C. 5 μl of 20 U/μl SUPERaseIn RNase inhibitor (Thermo Fisher Scientific) and 95 μl of ice-cold lysis buffer were further added, and the mixture was extracted using TRIzol LS Reagent in a Phasemaker Tube. Purified RNAs were resuspended in 5 μl of PNK solution [5 U T4 PNK (NEB) in 1X T4 PNK reaction buffer] and incubated for 10 min at 37 °C. After pre-incubation, ATP was added to a final concentration of 10 mM, and the mixture was incubated for another 30 min at 37 °C before adding 5.5 μl of Gel Loading Buffer II (Thermo Fisher Scientific). Ribosome-protected fragments (RPFs) corresponding to the 25 to 35 nt band were separated using 6% TBE-Urea Gel (Thermo Fisher Scientific) and subjected to library preparation using NEXTFLEX Small RNA Seq Kit v3 (Perkin Elmer). The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit and qPCR, analyzed by a Fragment Analyzer System to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System.
Reads were processed using TrimGalore (v0.6.7) to remove adaptor and low-quality reads with parameters “-q 20, --phred33, --stringency 3, --length 20, --max_length 40”. As the NEXTFLEX Small RNA Seq Kit v3 randomly added 4 bases at the 5’ and 3’ end of each read, we clipped these random bases when trimming reads. Reads mapped to rRNA were removed using Bowtie (v1.3.1) with parameters “-v 1, -a, --best, --strata, -q, --phred33-quals”. Remaining reads were mapped to mm10 (GRCm38.p6) reference genome using STAR (v2.7.10a) with parameters “--outFilterType BySJout, -outFilterMismatchNmax 2, --outFilterMultimapNmax 1, -outFilterMatchNmin 16, -alignEndsType EndToEnd, -outSAMattrIHstart 0”. Differentially translated mRNAs were identified using DESeq2 (v1.38.0).
CUT&Tag, data processing and analysis
Library preparation was performed as previously described98 after minor modifications. 50 to 80 mouse oocytes (per sample) were pre-extracted with ice-cold extraction buffer [25 mM HEPES (pH 7.4), 50 mM NaCl, 3 mM MgCl2, 300 mM sucrose and 0.5% triton X-100] for 10 min on ice and washed three times through ice-cold extraction buffer without triton X-100. Pre-extracted oocytes were directly washed with antibody buffer and incubated with primary antibody without crosslinking. Primary antibodies used were mouse IgG (Beyotime #A7028), mouse anti-CTD pS5 (4H8) (Santa Cruz #sc-47701), rat IgG (Beyotime #A7031), rat anti-CTD pS2 (Active motif #61699). Secondary antibodies used were rabbit anti-mouse (Sigma-Aldrich #06-371) and rabbit anti-rat (Sigma-Aldrich #AP164). In the library amplification step, 16 to 21 cycles were used. The purified libraries were quantified by Qubit 1X dsDNA HS Assay Kit and qPCR, analyzed by a Fragment Analyzer System to confirm successful amplification and quality, and sequenced using Novaseq 6000 Sequencing System.
Reads were processed using TrimGalore (v0.6,7) to remove adaptor and low-quality reads with parameters “-q 20, -e 0.1, --length 20, --stringency 1”, and then trimmed reads were mapped to mm10 (GRCm38.p6) reference genome using Bowtie2 (v2.4.2) with parameters “-L 25, -X 2000, --no-mixed --no-discordant”. Unmapped reads and low mapping quality reads (Q < 30) were discarded, and PCR duplicates were removed using the function “MarkDuplicates” from Picard tools (v2.27.2). CUT&Tag bigwig tracks were generated using the function “bamcoverage” from Deeptools (v3.5.1) with parameters “-binSize 50 -normSize 2494787188 -exactScaling -extendReads -normalizeUsing RPKM”. CUT&Tag signals on genomic regions were computed using computeMatrix with parameters “-bs 200, --upstream 10000, --downstream 10000”.
Statistical analysis
Average (mean) and SD were calculated in Excel. Statistical significance is based on unpaired, two-tailed Student’s t test (for absolute values between two groups), one-way ANOVA followed by post-hoc Tukey’s multiple comparison test (for absolute values for multiple groups) and two-tailed Fisher’s exact test (for categorical values) calculated in OriginPro or Prism (GraphPad), assuming normal distribution and similar variance. All box plots show median (horizontal black line), mean (small black squares), 25th and 75th percentiles (boxes), 5th and 95th percentiles (whiskers), and 1st and 99th percentiles (crosses). All data were from at least two independent experiments with at least three biological replicates per experiments. P values are designated as *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. Nonsignificant values are indicated as N.S.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq, HiC, WGBS, ATAC-seq, Ribo-seq, Cut&Tag-seq data are deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2025) in National Genomics Data Center (Nucleic Acids Res 2025), China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA031449) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Materials will be provided with material transfer agreements as appropriate. Source data are provided with this paper.
References
Chouinard, L. A. A light- and electron-microscope study of the oocyte nucleus during development of the antral follicle in the prepubertal mouse. J. Cell Sci. 17, 589–615 (1975).
Mattson, B. A. & Albertini, D. F. Oogenesis: chromatin and microtubule dynamics during meiotic prophase. Mol. Reprod. Dev. 25, 374–383 (1990).
Debey, P. et al. Competent mouse oocytes isolated from antral follicles exhibit different chromatin organization and follow different maturation dynamics. Mol. Reprod. Dev. 36, 59–74 (1993).
Zuccotti, M., Piccinelli, A., Giorgi Rossi, P., Garagna, S. & Redi, C. A. Chromatin organization during mouse oocyte growth. Mol. Reprod. Dev. 41, 479–485 (1995).
Oakberg, E. F. Relationship between stage of follicular development and RNA synthesis in the mouse oocyte, Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis. Mutat. Res./Fundamental Mol. Mechanisms Mutagenesis 6, 155–165 (1968).
Moore, G. P. & Lintern-Moore, S. A correlation between growth and RNA synthesis in the mouse oocyte. J. Reprod. Fertil. 39, 163–166 (1974).
Fourcroy, J. L. RNA synthesis in immature mouse oocyte development. J. Exp. Zool. 219, 257–266 (1982).
Christians, E. et al. Gene expression and chromatin organization during mouse oocyte growth. Dev. Biol. 207, 76–85 (1999).
Bloom, A. M. & Mukherjee, B. B. RNA synthesis in maturing mouse oocytes. Exp. Cell Res 74, 577–582 (1972).
Moore, G. P., Lintern-Moore, S., Peters, H. & Faber, M. RNA synthesis in the mouse oocyte. J. Cell Biol. 60, 416–422 (1974).
Rodman, T. C. & Bachvarova, R. RNA synthesis in preovulatory mouse oocytes. J. Cell Biol. 70, 251–257 (1976).
Wassarman, P. M. & Letourneau, G. E. RNA synthesis in fully-grown mouse oocytes. Nature 261, 73–74 (1976).
Bouniol-Baly, C. et al. Differential transcriptional activity associated with chromatin configuration in fully grown mouse germinal vesicle oocytes. Biol. Reprod. 60, 580–587 (1999).
Miyara, F. et al. Chromatin configuration and transcriptional control in human and mouse oocytes. Mol. Reprod. Dev. 64, 458–470 (2003).
Zuccotti, M., Bellone, M., Longo, F., Redi, C. A. & Garagna, S. Fully-mature antral mouse oocytes are transcriptionally silent but their heterochromatin maintains a transcriptional permissive histone acetylation profile. J. Assist Reprod. Genet 28, 1193–1196 (2011).
Wickramasinghe, D., Ebert, K. M. & Albertini, D. F. Meiotic competence acquisition is associated with the appearance of M-phase characteristics in growing mouse oocytes. Dev. Biol. 143, 162–172 (1991).
Wickramasinghe, D. & Albertini, D. F. Centrosome phosphorylation and the developmental expression of meiotic competence in mouse oocytes. Dev. Biol. 152, 62–74 (1992).
Zuccotti, M. et al. Meiotic and developmental competence of mouse antral oocytes. Biol. Reprod. 58, 700–704 (1998).
Carabatsos, M. J., Elvin, J., Matzuk, M. M. & Albertini, D. F. Characterization of oocyte and follicle development in growth differentiation factor-9-deficient mice. Dev. Biol. 204, 373–384 (1998).
Zuccotti, M. et al. The analysis of chromatin organisation allows selection of mouse antral oocytes competent for development to blastocyst. Zygote 10, 73–78 (2002).
Liu, H. & Aoki, F. Transcriptional activity associated with meiotic competence in fully grown mouse GV oocytes. Zygote 10, 327–332 (2002).
Bellone, M., Zuccotti, M., Redi, C. A. & Garagna, S. The position of the germinal vesicle and the chromatin organization together provide a marker of the developmental competence of mouse antral oocytes. Reproduction 138, 639–643 (2009).
Fluks, M., Szczepanska, K., Ishikawa, T. & Ajduk, A. Transcriptional status of mouse oocytes corresponds with their ability to generate Ca2+ release. Reproduction 157, 465–474 (2019).
Zhu, S. et al. The chromatin accessibility landscape of mouse oocytes during configuration transition. Cell Prolif. 58, e13733 (2025).
Inoue, A., Nakajima, R., Nagata, M. & Aoki, F. Contribution of the oocyte nucleus and cytoplasm to the determination of meiotic and developmental competence in mice. Hum. Reprod. 23, 1377–1384 (2008).
Zuccotti, M., Garagna, S., Merico, V., Monti, M. & Alberto Redi, C. Chromatin organisation and nuclear architecture in growing mouse oocytes. Mol. Cell Endocrinol. 234, 11–17 (2005).
Zatsepina, O. V., Bouniol-Baly, C., Amirand, C. & Debey, P. Functional and molecular reorganization of the nucleolar apparatus in maturing mouse oocytes. Dev. Biol. 223, 354–370 (2000).
Shishova, K. V., Lavrentyeva, E. A., Dobrucki, J. W. & Zatsepina, O. V. Nucleolus-like bodies of fully-grown mouse oocytes contain key nucleolar proteins but are impoverished for rRNA. Dev. Biol. 397, 267–281 (2015).
Shishova, K. V., Khodarovich, Y. M., Lavrentyeva, E. A. & Zatsepina, O. V. High-resolution microscopy of active ribosomal genes and key members of the rRNA processing machinery inside nucleolus-like bodies of fully-grown mouse oocytes. Exp. Cell Res 337, 208–218 (2015).
Shishova, K. V., Khodarovich, Y. M., Lavrentyeva, E. A. & Zatsepina, O. V. Data on morphology, large-scale chromatin configuration and the occurrence of proteins and rRNA in nucleolus-like bodies of fully-grown mouse oocytes in different fixatives. Data Brief. 7, 1179–1184 (2016).
Zhang, J. et al. Mammalian nucleolar protein DCAF13 is essential for ovarian follicle maintenance and oocyte growth by mediating rRNA processing. Cell Death Differ. 26, 1251–1266 (2019).
Wang, T. & Na, J. Fibrillarin-GFP Facilitates the Identification of Meiotic Competent Oocytes. Front Cell Dev. Biol. 9, 648331 (2021).
Longo, F. et al. Nuclear localization of NORs and centromeres in mouse oocytes during folliculogenesis. Mol. Reprod. Dev. 66, 279–290 (2003).
Garagna, S. et al. Three-dimensional localization and dynamics of centromeres in mouse oocytes during folliculogenesis. J. Mol. Histol. 35, 631–638 (2004).
Bonnet-Garnier, A. et al. Genome organization and epigenetic marks in mouse germinal vesicle oocytes. Int J. Dev. Biol. 56, 877–887 (2012).
Vautier, D., Besombes, D., Chassoux, D., Aubry, F. & Debey, P. Redistribution of nuclear antigens linked to cell proliferation and RNA processing in mouse oocytes and early embryos. Mol. Reprod. Dev. 38, 119–130 (1994).
Borsuk, E., Vautier, D., Szollosi, M. S., Besombes, D. & Debey, P. Development-dependent localization of nuclear antigens in growing mouse oocytes. Mol. Reprod. Dev. 43, 376–386 (1996).
Ihara, M., Stein, P. & Schultz, R. M. UBE2I (UBC9), a SUMO-conjugating enzyme, localizes to nuclear speckles and stimulates transcription in mouse oocytes. Biol. Reprod. 79, 906–913 (2008).
Al Jord, A. et al. Cytoplasmic forces functionally reorganize nuclear condensates in oocytes. Nat. Commun. 13, 5070 (2022).
Dai, X. X. et al. PABPN1 functions as a hub in the assembly of nuclear poly(A) domains that are essential for mouse oocyte development. Sci. Adv. 8, eabn9016 (2022).
Kageyama, S. et al. Alterations in epigenetic modifications during oocyte growth in mice. Reproduction 133, 85–94 (2007).
Castillo-Fernandez, J. et al. Increased transcriptome variation and localised DNA methylation changes in oocytes from aged mice revealed by parallel single-cell analysis. Aging Cell 19, e13278 (2020).
Wasserzug Pash, P. et al. Pre-pubertal oocytes harbor altered histone modifications and chromatin configuration. Front Cell Dev. Biol. 10, 1060440 (2022).
Eleftheriou, K. et al. A transition phase in late mouse oogenesis impacts DNA methylation of the early embryo. Commun. Biol. 5, 1047 (2022).
Yu, J. et al. Transgenic RNAi-mediated reduction of MSY2 in mouse oocytes results in reduced fertility. Dev. Biol. 268, 195–206 (2004).
Gazdag, E. et al. TBP2 is essential for germ cell development by regulating transcription and chromatin condensation in the oocyte. Genes Dev. 23, 2210–2223 (2009).
Liu, Y. J., Nakamura, T. & Nakano, T. Essential role of DPPA3 for chromatin condensation in mouse oocytogenesis. Biol. Reprod. 86, 40 (2012).
Monti, M. et al. Developmental arrest and mouse antral not-surrounded nucleolus oocytes. Biol. Reprod. 88, 2 (2013).
Lowther, K. M. & Mehlmann, L. M. Embryonic Poly(A)-binding protein is required during early stages of mouse oocyte development for chromatin organization, transcriptional silencing, and meiotic competence. Biol. Reprod. 93, 43 (2015).
Xu, K. et al. Maternal Sall4 Is Indispensable for Epigenetic Maturation of Mouse Oocytes. J. Biol. Chem. 292, 1798–1807 (2017).
Liu, X. M. et al. Loss of oocyte Rps26 in mice arrests oocyte growth and causes premature ovarian failure. Cell Death Dis. 9, 1144 (2018).
Au Yeung, W. K. et al. Histone H3K9 Methyltransferase G9a in Oocytes Is Essential for Preimplantation Development but Dispensable for CG Methylation Protection. Cell Rep. 27, 282–293 e284 (2019).
Briley, S. M. et al. Global SUMOylation in mouse oocytes maintains oocyte identity and regulates chromatin remodeling and transcriptional silencing at the end of folliculogenesis. Development 150 https://doi.org/10.1242/dev.201535 (2023).
Li, H. et al. LSM14B is an Oocyte-Specific RNA-Binding Protein Indispensable for Maternal mRNA Metabolism and Oocyte Development in Mice. Adv. Sci. (Weinh.) 10, e2300043 (2023).
Shan, L. Y. et al. LSM14B controls oocyte mRNA storage and stability to ensure female fertility. Cell Mol. Life Sci. 80, 247 (2023).
Wan, Y. et al. LSM14B is essential for oocyte meiotic maturation by regulating maternal mRNA storage and clearance. Nucleic Acids Res 51, 11652–11667 (2023).
Zhang, H. et al. LSM14B coordinates protein component expression in the P-body and controls oocyte maturation. J. Genet Genomics 51, 48–60 (2024).
Purschke, M., Rubio, N., Held, K. D. & Redmond, R. W. Phototoxicity of Hoechst 33342 in time-lapse fluorescence microscopy. Photochem Photobio. Sci. 9, 1634–1639 (2010).
So, C. et al. Mechanism of spindle pole organization and instability in human oocytes. Science 375, eabj3944 (2022).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Borsos, M. et al. Genome-lamina interactions are established de novo in the early mouse embryo. Nature 569, 729–733 (2019).
De La Fuente, R. & Eppig, J. J. Transcriptional activity of the mouse oocyte genome: companion granulosa cells modulate transcription and chromatin remodeling. Dev. Biol. 229, 224–236 (2001).
Buttress, T. et al. Histone H2B.8 compacts flowering plant sperm through chromatin phase separation. Nature 611, 614–622 (2022).
Bensaude, O. Inhibiting eukaryotic transcription: Which compound to choose? How to evaluate its activity? Transcription 2, 103–108 (2011).
Titov, D. V. et al. XPB, a subunit of TFIIH, is a target of the natural product triptolide. Nat. Chem. Biol. 7, 182–188 (2011).
Bushnell, D. A., Cramer, P. & Kornberg, R. D. Structural basis of transcription: alpha-amanitin-RNA polymerase II cocrystal at 2.8 A resolution. Proc. Natl. Acad. Sci. USA 99, 1218–1222 (2002).
Gong, X. Q., Nedialkov, Y. A. & Burton, Z. F. Alpha-amanitin blocks translocation by human RNA polymerase II. J. Biol. Chem. 279, 27422–27427 (2004).
Brueckner, F. & Cramer, P. Structural basis of transcription inhibition by alpha-amanitin and implications for RNA polymerase II translocation. Nat. Struct. Mol. Biol. 15, 811–818 (2008).
Zhang, C. et al. tagHi-C Reveals 3D Chromatin Architecture Dynamics during Mouse Hematopoiesis. Cell Rep. 32, 108206 (2020).
Nguyen, V. T. et al. In vivo degradation of RNA polymerase II largest subunit triggered by alpha-amanitin. Nucleic Acids Res 24, 2924–2929 (1996).
Mitsui, A. & Sharp, P. A. Ubiquitination of RNA polymerase II large subunit signaled by phosphorylation of carboxyl-terminal domain. Proc. Natl. Acad. Sci. USA 96, 6054–6059 (1999).
Lee, K. B., Wang, D., Lippard, S. J. & Sharp, P. A. Transcription-coupled and DNA damage-dependent ubiquitination of RNA polymerase II in vitro. Proc. Natl. Acad. Sci. USA 99, 4239–4244 (2002).
Xie, S. Q., Martin, S., Guillot, P. V., Bentley, D. L. & Pombo, A. Splicing speckles are not reservoirs of RNA polymerase II, but contain an inactive form, phosphorylated on serine2 residues of the C-terminal domain. Mol. Biol. Cell 17, 1723–1733 (2006).
Wang, Y., Lu, J. J., He, L. & Yu, Q. Triptolide (TPL) inhibits global transcription by inducing proteasome-dependent degradation of RNA polymerase II (Pol II). PLoS One 6, e23993 (2011).
Manzo, S. G. et al. Natural product triptolide mediates cancer cell death by triggering CDK7-dependent degradation of RNA polymerase II. Cancer Res 72, 5363–5373 (2012).
Liao, R. & Mizzen, C. A. Site-specific regulation of histone H1 phosphorylation in pluripotent cell differentiation. Epigenetics Chromatin 10, 29 (2017).
Novais-Cruz, M. et al. Mitotic progression, arrest, exit or death relies on centromere structural integrity, rather than de novo transcription. Elife 7 https://doi.org/10.7554/eLife.36898 (2018).
Steurer, B. et al. Live-cell analysis of endogenous GFP-RPB1 uncovers rapid turnover of initiating and promoter-paused RNA Polymerase II. Proc. Natl. Acad. Sci. USA 115, E4368–E4376 (2018).
Stellrecht, C. M. & Chen, L. S. Transcription inhibition as a therapeutic target for cancer. Cancers (Basel) 3, 4170–4190 (2011).
Laham-Karam, N., Pinto, G. P., Poso, A. & Kokkonen, P. Transcription and Translation Inhibitors in Cancer Treatment. Front Chem. 8, 276 (2020).
Litingtung, Y. et al. Growth retardation and neonatal lethality in mice with a homozygous deletion in the C-terminal domain of RNA polymerase II. Mol. Gen. Genet 261, 100–105 (1999).
Lan, Z. J., Xu, X. & Cooney, A. J. Differential oocyte-specific expression of Cre recombinase activity in GDF-9-iCre, Zp3cre, and Msx2Cre transgenic mice. Biol. Reprod. 71, 1469–1474 (2004).
Clift, D. et al. A Method for the acute and rapid degradation of endogenous proteins. Cell 171, 1692–1706 e1618 (2017).
Clift, D., So, C., McEwan, W. A., James, L. C. & Schuh, M. Acute and rapid degradation of endogenous proteins by Trim-Away. Nat. Protoc. 13, 2149–2175 (2018).
Uchino, S. et al. Live imaging of transcription sites using an elongating RNA polymerase II-specific probe. J. Cell Biol. 221 https://doi.org/10.1083/jcb.202104134 (2022).
Zeng, J. et al. Target-induced clustering activates Trim-Away of pathogens and proteins. Nat. Struct. Mol. Biol. 28, 278–289 (2021).
Egloff, S. & Murphy, S. Cracking the RNA polymerase II CTD code. Trends Genet 24, 280–288 (2008).
Fulka, H., Novakova, Z., Mosko, T. & Fulka, J. Jr The inability of fully grown germinal vesicle stage oocyte cytoplasm to transcriptionally silence transferred transcribing nuclei. Histochem Cell Biol. 132, 457–468 (2009).
Abe, K., Inoue, A., Suzuki, M. G. & Aoki, F. Global gene silencing is caused by the dissociation of RNA polymerase II from DNA in mouse oocytes. J. Reprod. Dev. 56, 502–507 (2010).
Medvedev, S., Pan, H. & Schultz, R. M. Absence of MSY2 in mouse oocytes perturbs oocyte growth and maturation, RNA stability, and the transcriptome. Biol. Reprod. 85, 575–583 (2011).
Barnetova, I. et al. RNA polymerase II transcriptional silencing in growing and fully grown germinal vesicle oocytes isolated from gonadotropin-stimulated and non-stimulated gilts. Mol. Reprod. Dev. 79, 697–708 (2012).
Wei, H. et al. Unique subcellular distribution of RPB1 with a phosphorylated C-terminal domain (CTD) in mouse oocytes during meiotic division and its relationship with chromosome separation. J. Reprod. Dev. 61, 541–548 (2015).
Oqani, R. K. et al. P-TEFb kinase activity is essential for global transcription, resumption of meiosis and embryonic genome activation in Pig. PLoS One 11, e0152254 (2016).
Liu, B. et al. The landscape of RNA Pol II binding reveals a stepwise transition during ZGA. Nature 587, 139–144 (2020).
Sun, X. et al. Comprehensive analysis of nonsurrounded nucleolus and surrounded nucleolus oocytes on chromatin accessibility using ATAC-seq. Mol. Reprod. Dev. 90, 87–97 (2023).
Wu, D. & Dean, J. Reduced female fertility due to sequestration of RNA Pol II by pervasive transcription in exosome RNase-depleted oocytes. Cell Rep. 42, 113247 (2023).
Wang, C. et al. Reprogramming of H3K9me3-dependent heterochromatin during mammalian embryo development. Nat. Cell Biol. 20, 620–631 (2018).
Gassler, J. et al. Zygotic genome activation by the totipotency pioneer factor Nr5a2. Science 378, 1305–1315 (2022).
Hu, W. et al. Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation. Nat. Commun. 13, 5114 (2022).
Cheng, S. & Schuh, M. Two mechanisms repress cyclin B1 translation to maintain prophase arrest in mouse oocytes. Nat. Commun. 15, 10044 (2024).
Ryu, K. Y. et al. The mouse polyubiquitin gene Ubb is essential for meiotic progression. Mol. Cell Biol. 28, 1136–1146 (2008).
Haaf, T. & Ward, D. C. Inhibition of RNA polymerase II transcription causes chromatin decondensation, loss of nucleolar structure, and dispersion of chromosomal domains. Exp. Cell Res 224, 163–173 (1996).
Buchwalter, A. L., Liang, Y. & Hetzer, M. W. Nup50 is required for cell differentiation and exhibits transcription-dependent dynamics. Mol. Biol. Cell 25, 2472–2484 (2014).
Nagashima, R. et al. Single nucleosome imaging reveals loose genome chromatin networks via active RNA polymerase II. J. Cell Biol. 218, 1511–1530 (2019).
Caragine, C. M., Haley, S. C. & Zidovska, A. Nucleolar dynamics and interactions with nucleoplasm in living cells. Elife 8 https://doi.org/10.7554/eLife.47533 (2019).
Jiang, Y. et al. Genome-wide analyses of chromatin interactions after the loss of Pol I, Pol II, and Pol III. Genome Biol. 21, 158 (2020).
Hilbert, L. et al. Transcription organizes euchromatin via microphase separation. Nat. Commun. 12, 1360 (2021).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Wang, Z. et al. The phosphatase PP1 sustains global transcription by promoting RNA polymerase II pause release. Mol. Cell https://doi.org/10.1016/j.molcel.2024.10.046 (2024).
Kelley, J. R. et al. The PNUTS phosphatase complex controls transcription pause release. Mol. Cell https://doi.org/10.1016/j.molcel.2024.10.045 (2024).
Abe, K., Schauer, T. & Torres-Padilla, M. E. Distinct patterns of RNA polymerase II and transcriptional elongation characterize mammalian genome activation. Cell Rep. 41, 111865 (2022).
An, H. et al. KAS-seq profiling captures transcription dynamics during oocyte maturation. J. Ovarian Res 17, 23 (2024).
Grosso, A. R. et al. Pervasive transcription read-through promotes aberrant expression of oncogenes and RNA chimeras in renal carcinoma. Elife 4 https://doi.org/10.7554/eLife.09214 (2015).
Grosso, A. R., de Almeida, S. F., Braga, J. & Carmo-Fonseca, M. Dynamic transitions in RNA polymerase II density profiles during transcription termination. Genome Res 22, 1447–1456 (2012).
Parua, P. K. & Fisher, R. P. Dissecting the Pol II transcription cycle and derailing cancer with CDK inhibitors. Nat. Chem. Biol. 16, 716–724 (2020).
Ku, H. et al. Effects of Transcription-Dependent Physical Perturbations on the Chromosome Dynamics in Living Cells. Front Cell Dev. Biol. 10, 822026 (2022).
Burns, K. H. et al. Roles of NPM2 in chromatin and nucleolar organization in oocytes and embryos. Science 300, 633–636 (2003).
De La Fuente, R. et al. Major chromatin remodeling in the germinal vesicle (GV) of mammalian oocytes is dispensable for global transcriptional silencing but required for centromeric heterochromatin function. Dev. Biol. 275, 447–458 (2004).
Aoki, F., Worrad, D. M. & Schultz, R. M. Regulation of transcriptional activity during the first and second cell cycles in the preimplantation mouse embryo. Dev. Biol. 181, 296–307 (1997).
Yu, C. et al. BTG4 is a meiotic cell cycle-coupled maternal-zygotic-transition licensing factor in oocytes. Nat. Struct. Mol. Biol. 23, 387–394 (2016).
Bellier, S. et al. Nuclear translocation and carboxyl-terminal domain phosphorylation of RNA polymerase II delineate the two phases of zygotic gene activation in mammalian embryos. EMBO J. 16, 6250–6262 (1997).
Lin, C. J., Koh, F. M., Wong, P., Conti, M. & Ramalho-Santos, M. Hira-mediated H3.3 incorporation is required for DNA replication and ribosomal RNA transcription in the mouse zygote. Dev. Cell 30, 268–279 (2014).
Chen, X. et al. ATAC-see reveals the accessible genome by transposase-mediated imaging and sequencing. Nat. Methods 13, 1013–1020 (2016).
Wang, M., Chen, Z. & Zhang, Y. CBP/p300 and HDAC activities regulate H3K27 acetylation dynamics and zygotic genome activation in mouse preimplantation embryos. EMBO J. 41, e112012 (2022).
Xia, M. et al. PCBP1 is required for maintenance of the transcriptionally silent state in fully grown mouse oocytes. Cell Cycle 11, 2833–2842 (2012).
Chousal, J. et al. Chromatin Modification and Global Transcriptional Silencing in the Oocyte Mediated by the mRNA Decay Activator ZFP36L2. Dev. Cell 44, 392–402 e397 (2018).
Andreu-Vieyra, C. V. et al. MLL2 is required in oocytes for bulk histone 3 lysine 4 trimethylation and transcriptional silencing. PLoS Biol. 8 https://doi.org/10.1371/journal.pbio.1000453 (2010).
Ma, P., Pan, H., Montgomery, R. L., Olson, E. N. & Schultz, R. M. Compensatory functions of histone deacetylase 1 (HDAC1) and HDAC2 regulate transcription and apoptosis during mouse oocyte development. Proc. Natl. Acad. Sci. USA 109, E481–E489 (2012).
Nashun, B. et al. Continuous histone replacement by hira is essential for normal transcriptional regulation and de novo dna methylation during mouse oogenesis. Mol. Cell 60, 611–625 (2015).
Yu, C. et al. CFP1 Regulates histone H3K4 trimethylation and developmental potential in mouse oocytes. Cell Rep. 20, 1161–1172 (2017).
Demond, H. et al. Multi-omics analyses demonstrate a critical role for EHMT1 methyltransferase in transcriptional repression during oogenesis. Genome Res 33, 18–31 (2023).
Pancholi, A. et al. RNA polymerase II clusters form in line with surface condensation on regulatory chromatin. Mol. Syst. Biol. 17, e10272 (2021).
Jedrusik, A. et al. Maternally and zygotically provided Cdx2 have novel and critical roles for early development of the mouse embryo. Dev. Biol. 344, 66–78 (2010).
Fadloun, A. et al. Chromatin signatures and retrotransposon profiling in mouse embryos reveal regulation of LINE-1 by RNA. Nat. Struct. Mol. Biol. 20, 332–338 (2013).
Zhu, M. et al. Developmental clock and mechanism of de novo polarization of the mouse embryo. Science 370 https://doi.org/10.1126/science.abd2703 (2020).
Palmerola, K. L. et al. Replication stress impairs chromosome segregation and preimplantation development in human embryos. Cell 185, 2988–3007 e2920 (2022).
Pal, M., Altamirano-Pacheco, L., Schauer, T. & Torres-Padilla, M. E. Reorganization of lamina-associated domains in early mouse embryos is regulated by RNA polymerase II activity. Genes Dev. 37, 901–912 (2023).
Nakatani, T. et al. Emergence of replication timing during early mammalian development. Nature 625, 401–409 (2024).
Sun, X. M. et al. Size-Dependent Increase in RNA Polymerase II Initiation Rates Mediates Gene Expression Scaling with Cell Size. Curr. Biol. 30, 1217–1230.e1217 (2020).
Basier, C. & Nurse, P. The cell cycle and cell size influence the rates of global cellular translation and transcription in fission yeast. EMBO J. 42, e113333 (2023).
Kimura, H., Hayashi-Takanaka, Y., Stasevich, T. J. & Sato, Y. Visualizing posttranslational and epigenetic modifications of endogenous proteins in vivo. Histochem Cell Biol. 144, 101–109 (2015).
Conic, S. et al. Imaging of native transcription factors and histone phosphorylation at high resolution in live cells. J. Cell Biol. 217, 1537–1552 (2018).
Sun, X. et al. Conditional inactivation of Fgf4 reveals complexity of signalling during limb bud development. Nat. Genet 25, 83–86 (2000).
Cavazza, T. et al. Parental genome unification is highly error-prone in mammalian embryos. Cell 184, 2860–2877.e2822 (2021).
Abe, K. I. et al. Minor zygotic gene activation is essential for mouse preimplantation development. Proc. Natl. Acad. Sci. USA 115, E6780–E6788 (2018).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
We are grateful to the patients who participated in this study. We thank the staff from the Transgenic Animal Center & Animal and Plant Center, Imaging Facility and Bioinformatics Center at the National Institute of Biological Sciences, Beijing for technical assistance; the clinicians, nursing team, and embryology team at the clinics for their support of this study; Z. Han and Z. Li for help with human oocytes and mouse embryos; A. Andersen, Q. Liu, and X. Wang for critical comments on the manuscript; M. Schuh for cDNAs and constructs. The research leading to these results was funded by the Ministry of Science and Technology of the People’s Republic of China under a National Key Research and Development Program of China (2023YFA1800300), the National Institute of Biological Sciences, Beijing under internal grant to C.S. and the National Natural Science Foundation of China (NSFC) under a grant to J.G. (82371667). C.S is also a recipient of the Excellent Young Scientists Fund Program from the NSFC and the DAMO Academy Young Fellowship from the Alibaba DAMO Academy.
Author information
Authors and Affiliations
Contributions
J.W. and C.S. conceived the study, designed experiments and methods for data analysis; J.W. and C.S. performed all experiments and analyzed the data with the following exceptions: W.L. performed bioinformatics analysis of all next-generation sequencing data. J.W. and C.S. wrote the manuscript and prepared the figures with input from all authors; J.G. and G.L. supervised the collection and vitrification of human oocytes at the Reproductive and Genetic Hospital of CITIC-XIANGYA; X.X. supervised the collection and vitrification of human oocytes at the Beijing Perfect Family Hospital; B.L. supervised the bioinformatics analyses; and C.S. supervised the entire study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks David Albertini, Takashi Ishiuchi, Xin Wu, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Li, W., Guo, J. et al. Natural degradation of RNA polymerase II is essential for oocyte chromatin reorganization and maternal-to-zygotic transition. Nat Commun 17, 720 (2026). https://doi.org/10.1038/s41467-025-67476-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67476-z