Abstract
Rice blast caused by Magnaporthe oryzae poses a major threat to world food supply. Conditions conducive to the disease include insufficient light. Our previous studies found that light-dependent phosphorylation of light-harvesting complex II protein (LHCB5) mediates rice immunity; however, the underlying mechanisms, in particular, the upstream regulators of LHCB5 phosphorylation, remain unknown. Here, we show that OsLIKE1 functions as a lectin receptor-like kinase (LecRLK) to play a critical role in rice resistance against M. oryzae. OsLIKE1 interacts with and phosphorylates LHCB5. In addition, we find two phosphorylation sites in the kinase domain of OsLIKE1 important for its auto-phosphorylation and phosphorylation of LHCB5. Moreover, we find that variation in light-induced OsLIKE1 expression is driven by specific single-nucleotide polymorphisms (SNPs) within the promoter region. These SNPs promote the auto-phosphorylation of OsLIKE1 to phosphorylate LHCB5 and activate its immune function against the blast fungus. Collectively, these findings elucidate the mechanism by which light regulates OsLIKE1-dependent defense in rice and support SNP-based rice breeding programs that increase disease resistance.
Similar content being viewed by others
Introduction
Environmental conditions strongly influence pathogen–host interactions. Rice blast epidemics often occur under overcast and rainy conditions, not only due to the favorable environment for the pathogen but also because of insufficient light, which reduces host resistance1,2,3. Given the importance of breeding rice varieties resistant to blast disease, elucidating the mechanism of light-regulated rice immunity is a critical research goal. Previous studies have shown that light induces the phosphorylation of the light-harvesting complex II protein (LHCB5) upon infection by the rice blast fungus Magnaporthe oryzae. The phosphorylation of LHCB5 occurs in the cytoplasm, and the phosphorylation of the 24th threonine in its chloroplast transit peptide (cTP) can accelerate its transport from the cytoplasm to the chloroplast, thereby inducing the accumulation of reactive oxygen species (ROS) in the chloroplast and activating immunity2. Resistance governed by LHCB5 phosphorylation is linked to the progeny harboring the desirable haplotype promoter, establishing the genetic basis of LHCB5-regulated resistance mediated by phosphorylation2. However, the mechanism by which LHCB5 undergoes phosphorylation in response to light and pathogen infection remains unclear.
Receptor-like kinases (RLKs) play crucial roles in the growth, development, and immunity of plants. Plants deploy many RLKs as pattern recognition receptors that detect microbe- and host-derived molecular patterns, serving as the first layer of inducible immunity4,5. RLKs contain a variable ectodomain responsible for ligand binding, a single-pass transmembrane domain, an intracellular juxtamembrane domain, and a cytoplasmic kinase domain. The extracellular domains of RLKs are highly variable, allowing them to perceive a wide variety of signals4,6. Based on the structure of their ectodomains, RLKs are divided into different sub-groups, including lectin receptor-like kinases (LecRLKs), which can be further categorized into L-, G-, and C-types based on their extracellular lectin domains7,8. While many RLKs have been well characterized for their roles in PAMP recognition and plant defense, increasing evidence supports the crucial roles of LecRLKs in plant development and responses to abiotic and biotic stresses. For instance, LecRK-IX.2, a L-type LecRLK, acts as a positive regulator of PAMP-triggered immunity in Arabidopsis9. Similarly, LecRK-I.9 recognizes extracellular ATP signals and directly phosphorylates RBOHD, triggering immune response in Arabidopsis10. In rice, OsLecRK, a G-type LecRLK, was reported to confer plant defense against brown planthopper, blast disease, and leaf blight disease11. These studies demonstrated that LecRLKs contribute to plant defense against biotrophic pathogens.
Aside from biotic pathogen-induced stresses, plants often encounter various abiotic environmental challenges, such as salt, drought, heat, cold, and light. Several reports have demonstrated the involvement of LecRLKs in plant responses to salt stress, wounding, drought, and heat stresses12,13,14,15,16; however, little is known about how LecRLKs respond to light and their likely underlying molecular mechanisms. Here, we identified OsLIKE1 as a LecRLK that interacts with LHCB5 and found that OsLIKE1 undergoes autophosphorylation in response to light stimulation and pathogen infection, subsequently phosphorylating LHCB5 to enhance resistance to rice blast.
Results
OsLIKE1 interacts with LHCB5 on the plasma membrane
We have previously shown that light induces the phosphorylation of LHCB5 in rice upon M. oryzae infection2. To identify the likely kinases that phosphorylate LHCB5, we performed immunoprecipitation-mass spectrometry (IP-MS) using LHCB5 overexpressing rice plants (LHCB5-OE) inoculated with the wild-type M. oryzae strain Guy112. 121 RLK proteins containing the kinase domain were identified (Supplementary Data 1). We validated interactions between selected 40 RLKs and LHCB5 using the yeast two-hybrid (Y2H) assay. To minimize the effects of the extracellular and transmembrane domains on protein expression and interaction, we expressed only their kinase domains (KDs) in the yeast Saccharomyces cerevisiae. The Y2H assay showed that a LecRLK, Os03g60810, exhibits the most vital interaction with LHCB5 (Supplementary Fig. 1 and Fig. 1a). We named this protein OsLIKE1 (LHCB5 interacting kinase 1). OsLIKE1 encodes a protein of 701 amino acids with a signal peptide and a lectin domain at the N terminus, a KD at the C-terminal region, and a transmembrane domain (TM) situated between the lectinB and KD domains (Fig. 1b). To identify locations of the OsLIKE1-LHCB5 interaction, we first observed OsLIKE1 by transiently expressing the GFP-tagged OsLIKE1 in the rice protoplast. When stained with FM4-64 dye, green fluorescence colocalized with red fluorescence of FM4-64 (Fig. 1c). To substantiate this localization, we generated transgenic rice lines overexpression OsLIKE1 with a C-terminal GFP tag (OsLIKE1-OE) in the ZH11 background (Supplementary Fig. 2a), and isolated the plasma membrane, chloroplast, nuclear, and cytoplasmic proteins from OsLIKE1-OE rice plants. Immunoblot analysis using compartment-specific markers confirmed that OsLIKE1 is predominantly located in the plasma membrane fraction (Fig. 1d).
a Yeast two-hybrid analysis for the interaction between OsLIKE1 and LHCB5. Specific genes fused to pGADT7 and pGBKT7 were co-introduced into the yeast AH109 strain. Transformants were first plated on SD-Leu-Trp medium and then replicated on selective SD-Leu-Trp-His-Ade medium for 5 days. The plasmid pairs pGBKT7-Lam and pGADT7-RecT, as well as empty pGADT7 and pGBKT7 vectors, served as negative controls. P53 and RecT were used as a positive control. b A schematic Diagram of OsLIKE1 Domains. The domains include the signal peptide (SP: 1-35), Lectin_legB domain (LectinB: 38-276 aa), transmembrane domain (TM: 307-329), and protein kinase domain (KD: 335-641 aa). c OsLIKE1-GFP colocalizes with FM4-64 on the plasma membrane of rice protoplasts. Bar, 10 μm. d Subcellular fractionation and immunoblotting analysis of OsLIKE1-GFP protein in OsLIKE1-OE rice leaves. Immunoblotting analysis with GFP, H+ATPase (membrane marker), Actin (cytoplasmic marker), RbcL (chloroplast marker) and Histone H3 (nuclear marker) antibodies. e OsLIKE1 was co-immunoprecipitated with LHCB5 during M. oryzae inoculation. Complex proteins were purified from the transgenic rice lines overexpression OsLIKE1 with a C-terminal GFP tag (OsLIKE1-OE) under blast stress and immunoprecipitated with an anti-GFP antibody. The presence of LHCB5 in the complex was confirmed by immunoblotting with anti-LHCB5 antibody. f The interaction between OsLIKE1 and LHCB5 on the plasma membrane of rice protoplasts was confirmed using a BiFC assay. OsLIKE1-YFPC and LHCB5-YFPN, or an empty vector, were co-expressed in rice protoplasts. YFP and FM4-64 signals were observed under confocal microscopy. Bar, 10 μm. All experiments were repeated independently with similar results at least three times.
To further characterize the in vivo interaction between OsLIKE1 and LHCB5 during M. oryzae infection, we performed a time-course co-immunoprecipitation (co-IP) assay. Proteins precipitated with anti-GFP beads were detected with the anti-LHCB5 antibody only in OsLIKE1-GFP lines, while no interaction was observed in GFP-only controls. Notably, the OsLIKE1-LHCB5 interaction was dynamically regulated upon pathogen challenge, with the binding strength robustly increased at infectious stages (24 to 72 h) compared to non-infection (0 h). (Fig. 1e). Similar results were also obtained in Nicotiana benthamiana using either the KD domain-containing or full-length OsLIKE1 (Supplementary Fig. 3a and b). This in vivo interaction was further validated by the bimolecular fluorescence complementation (BiFC) assay in transient expression of rice protoplasts and N. benthamiana (Fig. 1e and Supplementary Fig. 3c). In this assay, OsLIKE1 tagged with N-terminal YFP (OsLIKE1-YFPN) and LHCB5 tagged with C-terminal YFP (LHCB5-YFPC) were co-expressed in host cells, and the results showed the occurrence of colocalization mainly at the plasma membrane (PM) (Fig. 1f).
OsLIKE1 phosphorylates LHCB5 to enhance resistance against rice blast
To examine whether LHCB5 is a substrate phosphorylated by OsLIKE1, we expressed and purified His-tagged OsLIKE1-KD and GST-tagged LHCB5 in E. coli. In vitro phosphorylation assays showed phosphorylation of LHCB5 by OsLIKE1-KD (Fig. 2a). Our previous study demonstrated that the LHCB5 T24 residue is the sole phosphorylation site, and that stimulation of this site (T24D) induced cell death in N. benthamiana2. Therefore, we transiently expressed full-length OsLIKE1, OsLIKE1-KD, LHCB5, LHCB5T24A and LHCB5T24D in N. benthamiana leaves. Cell death was observed in leaves co-expressed OsLIKE1 or OsLIKE1-KD with LHCB5 and LCHB5T24D, but not in those co-expressed OsLIKE1 or OsLIKE1-KD with LHCB5T24A (Supplementary Fig. 4). These results suggested that OsLIKE1 triggers the activation of LHCB5 phosphorylation and the immune response.
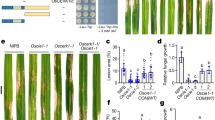
a OsLIKE1-KD phosphorylates LHCB5. A kinase assay was performed using OsLIKE1-KD-His as the kinase and GST-LHCB5 as the substrate, simultaneously detecting the autophosphorylation of OsLIKE1-KD and phosphorylation of LHCB5. Protein OsLIKE1-KD and LHCB5 were subjected to Mn2+-Phos-tag SDS-PAGE and normal SDS-PAGE followed by immunoblotting with anti-His and anti-GST antibodies. b Blast resistance of OsLIKE1-OE and OsLIKE1-KO plants following Guy11 spore spraying. Diseased leaves were photographed after 7 days. c The leaf lesion area of the leaves from (b) was assessed by Image J. The experiments were repeated twice with similar results. Values are the means of 3 replications, and error bars rep resent the SD (n = 3). Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures. d The relative fungal biomass of the leaves from (b) was detected by qRT-PCR. Values are the means of 3 replications, and error bars rep resent the SD (n = 3). Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures. e Guy11 infection is required for OsLIKE1 to phosphorylate LHCB5. Total proteins from OsLIKE1-OE and OsLIKE1-KO plants, sprayed with Guy11 spores and incubated for 2 days, were subjected to Mn2+-Phos-tag SDS-PAGE and regular SDS-PAGE, followed by immunoblotting with anti-GST and anti-LHCB5 antibodies. f OsLIKE1 is necessary for the phosphorylation of LHCB5. The LHCB5-HA fusion protein was expressed in rice protoplasts of ZH11, OsLIKE1-OE and OsLIKE1-KO, respectively. The LHCB5-HA fusion protein was subjected to Mn2+-Phos-tag SDS-PAGE and normal SDS-PAGE followed by immunoblotting with an anti-HA antibody. All experiments were repeated independently with similar results at least three times.
To further investigate whether OsLIKE1 contributes to blast resistance, we generated knock-out (OsLIKE1-KO) transgenic rice lines in the ZH11 background (Supplementary Fig. 2b). The OsLIKE1-KO lines #7 exhibited base deletion mutation, while lines #18 and #24 exhibited base insertion mutations (Supplementary Fig. 2b). In infection, OsLIKE1-OE rice lines showed strong resistance to M. oryzae, with significantly reduced lesion areas, whereas OsLIKE1-KO lines were more susceptible (Fig. 2b–d). No agronomically important traits differences were found among OsLIKE1-OE, OsLIKE1-KO, and the wild-type cultivar in the absence of M. oryzae infection (Supplementary Fig. 5). Moreover, compared to wild-type ZH11 plants, OsLIKE1-OE lines (#6, #18, and #20) maintained comparable plant height and yields upon M. oryzae infection. Strikingly, the OsLIKE1-KO lines (#7, #18, and #24) displayed more severe yield losses and stunted growth than infected ZH11, demonstrating that OsLIKE1 is essential for maintaining both productivity and plant architecture under blast stress (Supplementary Fig. 6).
Moreover, analyzed the phosphorylation of LHCB5 in ZH11, OsLIKE1-OE, and OsLIKE1-KO rice lines, both uninfected and infected by Guy11 using Mn2+-Phos-tag gel electrophoresis. We found that OsLIKE1 phosphorylation is induced by Guy11 infection in the OsLIKE1-OE compared to uninfected lines (Fig. 2e). Notably, LHCB5 phosphorylation was induced by Guy11 in the OsLIKE1-OE lines but not in ZH11 or OsLIKE1-KO lines (Fig. 2e). This result was also confirmed in the protoplasts of ZH11, OsLIKE1-OE, and OsLIKE1-KO rice lines treated with or without purified mycelial (PM) elicitors (Fig. 2f)2. Taken together, these findings demonstrated that OsLIKE1 enhances rice resistance to blast by phosphorylating LHCB5.
OsLIKE1 forms a homodimer that undergoes autophosphorylation to subsequently phosphorylate LHCB5
Receptor homodimerization or oligomerization is a common mechanism for ligand-mediated receptor activation and intracellular signal amplification17,18,19. Therefore, we tested OsLIKE1 self-interaction through GST pull-down and BiFC assays (Fig. 3a, b). Furthermore, we generated stable transgenic rice co-expressing OsLIKE1-GFP and OsLIKE1-Flag, with GFP and OsLIKE1-Flag co-expression serving as negative controls. Co-IP assays demonstrated that OsLIKE1-Flag was specifically detected in anti-GFP immunoprecipitates from OsLIKE1-GFP/OsLIKE1-Flag lines but not from control lines (Fig. 3c), confirming OsLIKE1 homodimerization in planta. Using native PAGE analysis, we found that KDs are mostly present in dimers (Fig. 3d). To examine which domains of OsLIKE1 might be involved in self-interaction, deletion mutants were made in the extracellular Lectin-binding (LectinB) domain and the intracellular KD. GST pull-down and Co-IP assays revealed that the deletion of KD, but not LectinB, nearly abolished self-interaction (Figs. 3e, f, and Supplementary Fig. 7).
a In vitro pull-down assay of GST-OsLIKE1 and OsLIKE1-His. Recombinant GST-OsLIKE1 or GST bound to glutathione Sepharose beads was incubated with E. coli BL21(DE3) cell lysates containing OsLIKE1-His. Eluted proteins were analyzed by immunoblot (IB) with monoclonal anti-GST and monoclonal anti-His antibodies. b Interaction of OsLIKE1 itself in rice protoplasts by the BiFC assay. OsLIKE1-YFPC and OsLIKE1-YFPN or empty vectors were co-expressed in rice protoplasts for detecting YFP signals under confocal microscopy. Bar, 10 μm. c Co-IP assay of OsLIKE1 homodimers. Total proteins were extracted from leaves of stable transgenic rice lines co-overexpression OsLIKE1-GFP/OsLIKE1-Flag and GFP/OsLIKE1-Flag in the ZH11 background. Proteins complexes were immunoprecipitated using GFP-trap beads, and the presence of OsLIKE1-GFP (detected with anti-GFP antibody) and OsLIKE1-Flag (detected with anti-Flag antibody) was confirmed by immunoblotting. d Immunoblot analysis of OsLIKE1-KD-His protein purified from E. coli BL21(ED3). The purified OsLIKE1-KD-His protein was subjected to native-PAGE followed by immunoblotting with a monoclonal anti-His antibody. e In vitro pull-down assay of KDs in OsLIKE1. Recombinant GST-OsLIKE1-KD or GST bound to glutathione Sepharose beads was incubated with E. coli BL21(DE3) cell lysate containing OsLIKE1-KD-His. The eluted protein was analyzed by immunoblot with monoclonal anti-GST and monoclonal anti-His antibodies. f Co-IP assay of KDs in OsLIKE1. Complex proteins were purified from N. benthamiana leaves expressing C-terminal GFP-tagged OsLIKE1-KD and immunoprecipitated with an anti-GFP antibody. The presence of OsLIKE1-KD-GFP and OsLIKE1-KD-HA in the complex was determined by immunoblot. OsLIKE1-KD-HA was immunoprecipitated with an anti-HA antibody. g OsLIKE1 phosphorylation sites identified in vitro by LC-MS/MS. The observed “y” and “b” ions are numbered. h The protein of full-length OsLIKE1 and phosphorylation sites mutations (T522 and T523) were purified with Ni-NTA agarose from E. coli BL21(DE3) and then treated with a kinase reaction buffer. The protein was purified and subjected to Mn2+-Phos-tag SDS-PAGE and normal SDS-PAGE, followed by immunoblotting with an anti-His antibody. i OsLIKE1 phosphorylates LHCB5 at 523 threonine but not at 522 threonine. The full-length OsLIKE1 and phosphorylation sites mutated proteins (T522 and T523) were used as kinases and GST-LHCB5 as the substrate is shown. Protein LHCB5 and OsLIKE1-KD with its mutant were subjected to Mn2+-Phos-tag SDS-PAGE and normal SDS-PAGE followed by immunoblotting with anti-GST and anti-His antibodies. All experiments were repeated independently with similar results at least three times.
To identify the amino acids involved in OsLIKE1 self-interaction, molecular modeling and docking analysis were carried out. The OsLIKE1 3D model was obtained from AlphaFold2 calculations, and most regions were confident with pLDDT> 90 (Supplementary Fig. 8a). ClusPro was used to predict that the interaction regions between KDs. Specifically, Loop2 (L2), Loop4 (L4), Loop6 (L6), Loop8 (L8), Loop10 (L10), Loop12 (L12), Loop14 (L14), and β-strand 12 (β12) and the amino acid residues involved were predicted to form respective hydrogen bonds (Supplementary Fig. 8b–d). Moreover, GST pull-down and co-IP with alanine substitution were carried out to verify this predicted model. The results showed that mutations in L6 (positions 429, 430, and 432) and L14 (positions 566, 567, 571, and 573) nearly abolish the interaction and homodimer formation (Supplementary Fig. 9a and b).
To determine whether OsLIKE1 forms homodimer to facilitate auto-phosphorylation, we expressed and purified OsLIKE1-KD and performed an in vitro phosphorylation assay using Mn2+-Phos-tag SDS-PAGE. We observed a slower migrating band of OsLIKE1-KD in the presence of ATP compared to its absence, indicating that phosphorylation occurs in OsLIKE1-KD (Fig. 3h). Mutations in L6 and L14 nearly abolish this auto-phosphorylation (Supplementary Fig. 9c). To further identify the phosphorylation sites of OsLIKE1-KD, we performed mass spectrometry analysis with the above purified OsLIKE1-KD, which showed two differentially phosphorylated residues in T522 and T523 (Fig. 3g and Supplementary Fig. 10). Mutation of both residues to alanine (OsLIKE1-KDT522A/T523A), rather than a single mutation, prevented OsLIKE1-KD phosphorylation (Fig. 2h). Furthermore, mutated the phosphorylated residues in OsLIKE1-KD, and found that T523A and T523A/T523A mutations abolished the phosphorylation of LHCB5 by OsLIKE1-KD, but not T522A mutation (Fig. 3i). These results suggested that OsLIKE1 can form homodimers through its KD to facilitate auto-phosphorylation and LHCB5 phosphorylation.
The light-inducible promoter drives the expression of OsLIKE1 and promotes its auto-phosphorylation
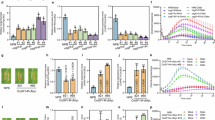
Our previous studies have demonstrated that LHCB5 is subject to light-induced phosphorylation during infection of M. oryzae2. To further investigate whether the auto-phosphorylation of OsLIKE1 and its associated resistance were light-dependent, we examined OsLIKE1 phosphorylation in OsLIKE1-OE rice lines inoculated with Guy11 at different light intensities. Interestingly, we found that OsLIKE1-OE rice lines show enhanced resistance at various light intensities compared to ZH11 (Fig. 4a). Moreover, both OsLIKE1 and LHCB5 can undergo phosphorylation in OsLIKE1-OE rice lines infected by Guy11, and this phosphorylation showed no difference at 50 and 200 μmol photons m−2s−1 (Fig. 4b).
a Blast resistance of OsLIKE1-OE plants following Guy11 spore spraying at different light intensities (200 mmol photons m⁻²s⁻¹ and 50 mmol photons m⁻²s⁻¹). Diseased leaves were photographed after 6 days. b The phosphorylation of OsLIKE1 and LHCB5 is not affected by light intensity. Total proteins of OsLIKE1-OE rice leaves treated at different light intensities after 2 days were subjected to Mn2+-Phos-tag SDS-PAGE and normal SDS-PAGE, followed by immunoblotting with anti-GFP and anti-LHCB5 antibodies. c OsLIKE1-mediated resistance is associated with its SNP type 2 promoter. The 400 accessions of 3 K rice materials were cultivated in a natural disease nursery (Ganyu District, Jiangsu Province, China) and classified according to rice blast severity ratings (See Materials and “Methods” section), obtaining field resistance data for all 400 accessions against rice blast. These data were subjected to association analysis with the promoter variations of OsLIKE1 (detailed data are provided in Supplementary Table 3). Statistical analysis revealed that the SNP type 2 promoter variant showed a significant correlation with blast resistance (Spearman r = 0.42, P <0.001). Data are displayed as box and whisker plots with individual data points: median, first and third quantiles, maximum and minimum (n = 400). Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures.d The expression levels of OsLIKE1 genes in TP309 and YG456 inoculated with Guy11 at different light intensities were analyzed by qRT-PCR. Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures, Data are displayed as scatter plots with individual data points: median, maximum and minimum (n = 3). e The expression levels of OsLIKE1 genes in non-infected TP309 and YG456 at different light intensities were analyzed by qRT-PCR. Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures, Data are displayed as scatter plots with individual data points: median, maximum and minimum (n = 3). f The expression levels of OsLIKE1 genes in TP309 and YG456 treated with or without Guy11 infection were analyzed by qRT-PCR. Data are analyzed by two-sidedsided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures, Data are displayed as scatter plots with individual data points: median, maximum and minimum (n = 3). g Evaluation of OsLIKE1 expression driven by distinct promoters in transgenic rice lines. The activity of three promoter variants—pUbi (ubiquitin promoter), pSNPtype1, and pSNPtype2—driving OsLIKE1 expression in transgenic rice lines (pC1305-pUbi:OsLIKE1, pC1305-pSNPtype1:OsLIKE1, and pC1305-pSNPtype2:OsLIKE1) was assessed. After 12 h of exposure to different light intensities (200 μmol photons m⁻²s⁻¹ and 50 μmol photons m⁻²s⁻¹), total proteins were extracted and analyzed by SDS-PAGE followed by immunoblotting using an anti-GFP antibody. Protein loading is indicated with Ponceau staining (Ponceau S). h The expression of OsLIKE1 genes in sixteen rice varieties at different light intensities were analyzed by qRT-PCR. The data represent the mean ± standard error of three biological replicates (n = 3). Data are analyzed by two-sided ANOVA followed by Tukey’s test. Adjusted p values are shown in figures. i LHCB5 was phosphorylated in twelve rice varieties. Proteins from sixteen rice varieties leaves inoculated with Guy11 at 2 dpi, extracts treated with phosphatase inhibitors, were subjected to normal and Mn2+-Phos-tag SDS-PAGE followed by immunoblotting with the LHCB5 polyclonal antibody. All experiments were repeated independently with similar results at least three times.
We speculated that this phenomenon may be related to the overexpression of OsLIKE1 and, further, different OsLIKE1 expression levels may exist in natural populations often associated with single-nucleotide polymorphisms (SNPs) within the promoter region. Therefore, we investigated the variance of OsLIKE1 promoter sequences from 3000 sequenced rice genomes20. SNPs were identified and separated into 20 groups using O. sativa ssp japonica cv. TP309 (SNP type 1) as the control, and most SNPs were distributed within types 1 and 2 (Supplementary Data 2 and Supplementary Fig. 11a). Notably, using Spearman’s rank correlation analysis to assess the relationship between disease resistance/susceptibility and SNPs across > 400 rice accessions from the 3 K dataset, we discovered a significant positive correlation between SNP type 2 and enhanced resistance (r = 0.42, p < 0.001; Fig. 4c). This finding further substantiates the evidence that OsLIKE1 promoter variations may play a key role in regulating immune responses.
To investigate the potential association between SNP type 2 and light-dependent OsLIKE1 transcription, we examined the transcription levels of OsLIKE1 in TP309 and YG456 rice varieties inoculated with or without M. oryzae at different light intensities. The results showed that a light intensity of 200 μmol photons m−2s−1, compared to 50 μmol photons m−2s−1, significantly induced OsLIKE1 expression in YG456. However, light intensity had no effect on OsLIKE1 transcription in the TP309 variety (Fig. 4d, e). Additionally, M. oryzae infection did not significantly induce the transcription of OsLIKE1 in either TP309 or YG456 at a light intensity of 200 μmol photons m−2s−1 (Fig. 4f). To determine the transcriptional dynamics of OsLIKE1 and LHCB5 in response to M. oryzae infection under varying light intensities, we analyzed their expression in rice cultivars ZH11 and TP309. We found a striking contrast: the transcription of OsLIKE1 remained largely unchanged across all conditions, indicating it is not induced by either pathogen infection or light (Supplementary Fig. 12a and b). In contrast, LHCB5 expression was significantly upregulated by both light exposure and pathogen challenge, with a particularly pronounced induction observed at 48 and 72 hpi (Supplementary Fig. 12c and d), a pattern consistent with previous reports2.
We further amplified the promoter sequences of OsLIKE1 from YG456 and TP309 and analyzed SNP types. The results showed that the SNPs of OsLIKE1 promoter in YG456 belong to type 2 (Supplementary Fig. 11a and Supplementary Fig. 13). To validate the correlation between OsLIKE1 promoter SNPs and expression levels, we expressed the GFP reporter gene driven by different promoters (p35S:GFP, pOsLIKE1type 1:GFP and pOsLIKE1type 2:GFP) in rice protoplasts at different light intensities. The result showed that constructs p35S:GFP and pOsLIKE1type 1:GFP show no other levels of green fluorescence at 50 and 200 μmol photons m−2s−1, while pOsLIKE1type 2:GFP shows a significantly higher level of green fluorescence at 200 μmol photons m−2s−1 thanthat at 50 μmol photons m−2s−1 (Supplementary Fig. 11b). Western blot analysis also showed a higher GFP protein level under the control of pOsLIKE1type 2 promoter at 200 μmol photons m−2s−1 thanat 50 μmol photons m−2s−1, indicating that type 2 SNPs in the OsLIKE1 promoter are indeed light-inducible (Supplementary Fig. 11c).
To validate the stability of these findings in rice, we generated stably inherited transgenic rice lines expressing OsLIKE1 gene driven by three distinct promoters: the ubiquitin promoter (pUbi), the SNP type 1 promoter (pSNPtype1), and the SNP type 2 promoter (pSNPtype2). The constructs pC1305:pUbi:OsLIKE1 and pC1305:pSNPtype1:OsLIKE1 exhibited comparable levels of green fluorescence at both 50 and 200 μmol photons m−2s−1. In contrast, pC1305:pSNPtype2:OsLIKE1 showed significantly higher fluorescence intensity at 200 μmol photons m−2s−1 compared to 50 μmol photons m−2s−1 (Fig. 4g). Identical results were observed when OsLIKE1 was replaced with the GFP reporter gene in transgenic rice lines (Supplementary Fig. 11d), confirming the light-dependent regulatory role of the SNP type 2 promoter.
We further selected 15 rice samples in 3k rice varieties to analyze SNP types and promoter transcription levels. We found that 12 of these samples belong to SNP type 2, and in these, the transcription and protein levels of OsLIKE1 were significantly upregulated under high light induction, compared to TP309 and SNP type 3 rice varieties (Fig. 4h and Supplementary Fig. 14a). All 12 rice samples notably showed OsLIKE1 auto-phosphorylation and LHCB5 phosphorylation (Fig. 4i and Supplementary Fig. 14b). Through genetic crosses between the resistant donor YG456 and susceptible LTH varieties, we previously demonstrated the stably inherited LHCB5 phosphorylation2. To validate whether the OsLIKE1 SNP type 2 promoter is heritable, we analyzed the OsLIKE1 promoter in progeny derived from LTH × YG456 crosses, which revealed that the previously reported blast-resistant progeny exhibiting LHCB5 phosphorylation exclusively carried the OsLIKE1 type 2 promoter variant (Supplementary Data 3). This finding demonstrated co-segregation of the OsLIKE1 type 2 promoter with heritable LHCB5 phosphorylation in the offspring. Together, these findings indicated that SNP-mediated promoter sequence divergence leads to different expression levels of OsLIKE1 in a light-dependent manner, which promotes OsLIKE1 auto-phosphorylation and LHCB5 phosphorylation.
Discussion
Environmental conditions strongly influence pathogen–host plant interactions. Many studies have demonstrated that environmental conditions like light, temperature, and humidity can affect disease development and innate host immunity21,22,23,24,25. Compared to the extensive research on regulating plant diseases by temperature and humidity, studies on the impact of light on plant diseases are relatively limited. Our previous studies proposed a model where during the M. oryzae and rice interaction, the host monitors the progression of infection through the phosphorylation of LHCB5 in a light-dependent manner, resulting in host immune response activation2. However, the mechanism of LHCB5 phosphorylation has not been elucidated. Our current studies establish the relationships among light conditions, OsLIKE1 mediated LHCB5 phosphorylation and blast resistance.
Plants rely on membrane-bound RLKs to perceive biotic and abiotic signals. The LecRLKs, a plant-specific RLK subfamily, have recently been found to play crucial roles in responses to abiotic and biotic stresses4,16,26. Here, we identified a LecRLK OsLIKE1 that interacts with and phosphorylates LHCB5 in vitro and in vivo. OsLIKE1 relies on the kinase domain to form homodimers that activate auto-phosphorylation, like most RLKs17,19,27,28. RLK auto-phosphorylation is a key mechanism that activates and amplifies downstream signaling pathways. One well-studied RLK is CERK1 which auto-phosphorylates and interacts with other proteins to trigger defense responses upon binding chitin in Arabidopsis thaliana29. We found two phosphorylation sites in the KD of OsLIKE1 that determine its auto-phosphorylation. One site (T523) is necessary for LHCB5 phosphorylation, indicating OsLIKE1 may have additional substrates similar to other model systems30,31,32. Notably, OsLIKE1 auto-phosphorylation is required for infection, suggesting OsLIKE1 may perceive other signaling molecules released from M. oryzae.
In response to pathogen infection, rice immunity is initiated by pattern recognition receptor (PRR) complexes, such as chitin and peptidoglycan (PGN) receptor OsCERK1, which subsequently activates immunity via downstream cytoplasmic kinases33,34,35,36. These results pointed out several important areas of investigation on the mode of action and regulation of OsLIKE1 in rice immunity: 1. The identification of OsLIKE1 ligands. 2. The identification of OsLIKE1 co-receptor for defense signaling should ligand recognition is not known. 3. The determination of relationships between OsLIKE1 and other PRRs in rice.
The phosphorylation of OsLIKE1 and LHCB5 is required for light induced responses regardless of any diseases. Several RLKs require environmental factors to activate their phosphorylation and subsequent signaling pathways. For example, the activation of A. thaliana FLS2 phosphorylation is response to environmental factors such as drought and salt37. The phosphorylation of rice LecRLK SIT1 is rapidly activated in the presence of salt that subsequently activates the MPK3/6 cascade pathway38. However, light-induced phosphorylation of RLKs has yet to be documented.
In this study, we demonstrated that different types of SNPs are present in the promoter region of OsLIKE1, and the type 2 SNPs are light-inducible. Under higher light intensity, the expression levels of OsLIKE1 significantly are increased, enabling it to perceive pathogen infection, form dimers, and undergo auto-phosphorylation. This is similar to studies by Espinoza et al. that CERK1 expression was upregulated under salt stress and unraveled the molecular mechanism that contributes to cross-tolerance between chitin and salt in A. thaliana39. The expression of PnLecRLK1 was rapidly induced by simulated cold, salt, and drought stresses in Pohlia nutans40. However, it remains challenging to explore how the expression of these RLKs is induced under these stresses. Importantly, SNPs identified in the OsLIKE1 promoter region are valuable resources for identifying upstream regulatory elements that govern transcription in connection to light perception. For example, an A to G transition in the promoter region of the BSR-D1 gene in M. oryzae lead to reduced expression via repressive MYB transcription factor binding, resulting in elevated resistance41.
In summary, our results support a model that light influences transcription of OsLIKE1, and this regulation is affected by various SNPs, especially type 2 SNPs, in the native promoter of OsLIKE1. Light-induced transcription of OsLIKE1 also promotes its homodimer formation and auto-phosphorylation, resulting in LHCB5 phosphorylation, which activates host immunity against M. oryzae infection (Fig. 5). Notably, phytochromes are central players in mediating light-dependent transcriptional reprogramming in plants42,43,44. A compelling question arising from our work is whether the OsLIKE1 promoter possesses an intrinsic light-responsive module that acts autonomously, or whether its light inducibility is mediated indirectly through the phytochrome signaling pathway. Elucidating this distinction will be a critical next step in fully understanding the upstream regulatory framework that connects light perception to OsLIKE1-mediated immune responses.
In natural rice populations, the promoter of OsLIKE1 contains different types of SNPs (mainly type 1 and type 2). Unknown transcription factors (FTs) regulated by light in rice bind to the type 2 promoter of OsLIKE1. There are 3 states of OsLIKE1 during the M. oryzae-rice interaction under different light intensities. a When light is insufficient, the TFs may be repressed and unable to initiate OsLIKE1 transcription (type 1 or 2). Without enough protein accumulation, OsLIKE1 cannot be activated, failing to phosphorylate LHCB5 and induce chloroplast immunity, leading to rice susceptibility to M. oryzae. b If the promoter of OsLIKE1 is type 1, the transcription factor cannot bind to it when light is sufficient. As a result, OsLIKE1 is not activated and rice remains susceptible. c If the promoter of OsLIKE1 in rice is type 2, the TFs bind and initiate OsLIKE1 transcription when light is sufficient. OsLIKE1 forms a homodimer and activates auto-phosphorylation, which then phosphorylates LHCB5 and induces immunity, leading to resistance against M. oryzae.
Methods
Plant varieties
R4D1 rice varieties were obtained from Bo Zhou’s lab at the International Rice Research Institute. Yangeng456 (YG456) was obtained from the Saline-Alkali Land Utilization and Research Institute, Liaoning, China. Oryza sativa japonica cv. ZH11 was transformed with constructs pCAM2300:OsLIKE1 and CRISPR-Cas9:OsLIKE1 with Agrobacterium tumefaciens-mediated transformation at EDGENE BIOT (Wuhan, China). The plants were grown in the greenhouse or fields at Nanjing Agricultural University in Nanjing, China.
Fungal strains and cultures
M. oryzae Guy11 was used as the wild-type strain in this study. The strains were cultured in a complete medium (CM). Liquid CM was employed to cultivate mycelia for DNA and RNA extraction. For conidiation, strain blocks were maintained on a straw decoction and corn (SDC) agar medium at 28 °C for 7 days in the dark, followed by 3 days of continuous illumination under fluorescent light.
Resistance test and infection assays
For resistance tests, conidia were suspended to a concentration of 5 × 104 spores per milliliter in a 0.2% (w/v) gelatin solution. Five milliliters of this suspension were sprayed on two-week-old rice seedlings. The inoculated plants were then kept in a growth chamber at 25 °C with 90% humidity, in the dark for the first 24 h, followed by a 16/8 h light/dark cycle. The disease severity was assessed at 7 days post-inoculation.
The disease severity of leaf blast under field conditions in the disease nursery was evaluated using the standard 0–9 scale45, with all assessments conducted in situ to reflect actual field infection patterns. The ten-level rating system was defined as follows: level 0 represented completely healthy leaves with no visible disease symptoms under field observation; level 1 showed initial infection signs with small brown spots less than 1 mm in diameter, typically appearing in field conditions; level 2 exhibited larger brown spots measuring 1–2 mm in diameter, as commonly seen in early field infections; level 3 presented characteristic circular to elliptical gray lesions 1–2 cm in diameter, representing typical field symptom development; level 4 demonstrated spindle-shaped lesions 1–2 cm in length confined between two veins, with infected area not exceeding 2% of the total leaf area under field assessment; level 5 displayed spindle-shaped lesions covering 2–10% of the leaf area, indicating moderate field infection; level 6 showed lesions affecting 11–25% of the leaf area, representing significant field damage; level 7 exhibited lesions occupying 26–50% of the leaf area, reflecting severe field infection; level 8 presented lesions covering 51–75% of the leaf area, indicating advanced field disease progression; and level 9 showed lesions affecting more than 75% of the total leaf area, representing maximum disease severity under field conditions.
RNA isolation and qRT-PCR
RNA isolation was performed using TRIzol (Invitrogen Life Technologies, Shanghai, China). cDNA synthesis was carried out with the PrimeScript™ RT Reagent Kit (TaKaRa). Quantitative RT-PCR was conducted using the ABI 7500 Fast Real-Time System, and transcript analysis was performed with the 7500 System SDS software. To compare the relative abundance of target gene transcripts in different rice varieties, the average threshold cycle (Ct) values were normalized to rice actin for each treated sample using the 2-∆Ct method, where ∆Ct = (Ct_target gene - Ct_actin). Fold changes in gene expression among different rice varieties were calculated using the 2-∆∆Ct method, where ∆∆Ct = (Ct_experiment - Ct_actin) - (Ct_control - Ct_actin).
Yeast two-hybrid assays
The OsLIKE1-KD cDNA was cloned into the pGBKT7 vector to create the bait construct. The prey construct was generated by cloning LHCB5 cDNA into the pGADT7 vector. Both constructs were confirmed by sequencing analysis and subsequently transformed into the yeast strain AH109 using the BD library construction & screening kit (Clontech). Tryptophan (Trp + ) and leucine (Leu + ) transformants were isolated and assayed for growth on SD-Trp-Leu-His (histidine)-Ade (adenine) medium.
In vivo co-IP assay
Co-IP using stable transgenic rice lines, rice protoplasts, and N. benthamiana cells was performed according to our previous study with minor modification2,46. Briefly, samples were extracted with IP buffer (50 mM Tris-HCl, pH 7.5; 150 mM NaCl; 0.5% NP-40; 5 mM DTT; and protease inhibitor cocktail). The mixtures were incubated at 4 °C with gentle shaking for 1 h. The IP complex was captured by adding 50 μl GFP-Trap® (ChromoTek, Germany, gta-20) or Anti-FLAG Antibody (Engibody, AT0022), followed by shaking at 4 °C for an additional hour. The beads were recovered by centrifugation at 2500 × g for 30 seconds and washed six times with cold TBS (50 mM Tris-HCl, pH 7.5; 150 mM NaCl). Then, 50 μl of glycine elution buffer (pH 2.5) was added to the beads. After boiling for 5 min, the samples were loaded onto SDS-PAGE gels for western blot analysis, and detected using anti-LHCB5 antibody (Agrisera, AS09407, 1:2000 dilution) and anti-GFP antibody (Abmart, M20004, 1:5000 dilution).
In vivo BiFC assay and subcellular localization
For subcellular localization in rice protoplasts, a cDNA fragment corresponding to the entire coding sequence of OsLIKE1 was cloned into the pCAM2300 vector to generate pACT1:OsLIKE1. OsLIKE1-YFPN and LHCB5-YFPC fusion constructs were generated and cloned into the pICH86988 vector; pICH86988-YFPN and pICH86988-YFPC vectors were used as controls. The fusion constructs were transformed into protoplasts prepared from TP309 seedlings, following a previously described method47. The recombinant plasmids were transferred to A. tumefaciens GV3101. The GV3101 strains carrying the indicated constructs were mixed at OD600 = 0.4 each and infiltrated into leaves of 4-week-old N. benthamiana plants. Infiltrated leaves were observed 4 days later using a confocal laser scanning microscope.
In vitro pull-down assay
To construct GST- or His-fused plasmids, OsLIKE1-LectinB and OsLIKE1-KD were inserted into the pGEX4T-2 (GE Healthcare Life Science) or pET-32a (Navogen) vectors. The pull-down assay was carried out using the ProFound pull-down GST protein-protein interaction kit (Pierce) according to the manufacturer’s instructions45. Briefly, the GST-fusion protein was expressed in E. coli strain BL21 (DE3). Soluble proteins were incubated with 50 μl glutathione agarose beads (Invitrogen, 16101) for 1 h at 4 °C. The beads were washed six times and then incubated with an equal amount of bacterial lysates containing His-fusion protein for another hour at 4 °C. After washing the beads six more times, the presence of His-fusion proteins was detected by immunoblotting using an anti-His antibody (Abmart, M20001, 1:5000 dilution).
Production of polyclonal antibodies against OsLIKE1
To generate high-specificity polyclonal antibodies against OsLIKE1, we commissioned Abmart to synthesize and purify the custom peptide antigen (sequence: AVDQFLFPGFSGDGVATSGAAAVTSTGLLQLTNETKEVFGHGFYPKPVSLRNASTGAAASFSTTFVFAIVPKYPDAHGHGLAFALAPSVAVPGAVAGKYLGLFNTSDSTGQIKNKIVAVELDTARDDEFADIDDNHVGIDDNSLKSVNSSPAGYHDAATGGKLASVNLIGGEPLQVWIEYDGDSAWLEVTVSPAG) and perform rabbit immunization. Briefly, the immunization protocol consisted of 6 subcutaneous injections administered at 2-week intervals, using 200 μg of purified antigen per immunization emulsified with Freund’s adjuvant (complete adjuvant for primary immunization, incomplete for boosts). Serum was collected 7 days after the final boost and antibody titer was determined by ELISA, demonstrating strong immunoreactivity at dilutions exceeding 1:100. The antibodies specifically recognized an ~80 kDa band corresponding to OsLIKE1 in western blot analysis of rice protein extracts.
Phosphorylation assay
In vivo phosphorylation analysis was performed following a previously described method (2). Briefly, rice leaves were ground into powder in liquid nitrogen and resuspended in 1 ml of extraction buffer (10 mM Tris-HCl [pH 7.5], 150 mM NaCl, 0.5 mM EDTA, 0.5% NP-40), supplemented with 1 mM PMSF, 10 μl of protease inhibitor cocktail (Sigma, P8340), and 10 μl of phosphatase inhibitor cocktail 3 (Sigma, 524627).
For in vitro phosphorylation analysis, His-OsLIKE1 and GST-LHCB5 were expressed in E. coli strain BL21 (DE3) (Sigma, CMC0014) and purified as previously described48. The reaction mixture was assembled by adding the following to microcentrifuge tubes: 0.5 μg of purified recombinant kinase OsLIKE1, 10 μg of potential substrate LHCB5, 3 μl of 10x reaction buffer (250 mM Tris, pH 7.5, 100 mM MgCl2, and 10 mM DTT), and water to a total volume of 27.5 μl. The reaction was initiated by adding 2.5 μl of 0.5 mM ATP (Sigma-Aldrich, A2383), mixed well, and incubated at 22 °C for 1.5 h. To stop the reaction, 10 volumes of cold acetone were added.
Samples for phosphorylation analyses were resolved on 10% SDS-polyacrylamide gels prepared with 50 μM acrylamide-pendant Phos-tag ligand and 100 μM MnCl2, according to the instructions provided by the Phos-tag Consortium (Wako, AAL-107). The gels were electrophoresed at 80 V/gel for 3-6 h. Prior to transfer, gels were equilibrated in transfer buffer containing 5 mM EDTA for 20 min, twice, followed by equilibration in transfer buffer without EDTA for 10 min. Protein transfer from the Mn2+-Phos-tag acrylamide gel to the PVDF membrane was performed overnight at 80 V at 4 °C, and the membrane was subsequently analyzed by Western blotting.
Subcellular fractionation
Isolate rice protoplasts from transgenic rice plant materials. For isolating protoplasts, a razor blade was used to cut the stems and leaves into 0.5 mm strips. They were then placed into a petri dish containing 10 mL enzyme solution (1.5% cellulase and 0.3% macerozyme). The vacuum was applied for 1 h for the infiltration of the enzyme solution. Plants were incubated for about 4 h in the dark with gentle shaking (40 × g) at room temperature and the solution was removed with a glass pipet. 10 mL W5 medium was added and allowed gently swirl (80 × g) for 1 h to release the protoplast. W5 Protoplasts were filtered through a 35 μm nylon mesh, transferred into an 8 mL glass vial, precipitated at 220 × g for 4 min. These protoplasts will be used for subcellular fractionation. Membrane protein extraction: Using the Membrane and Cytosol Protein Extraction Kit (Beyotime, P0033), add Membrane Protein Extraction Reagent A to the rice protoplasts and homogenize on ice with 30-50 strokes to disrupt the cells. Centrifuge at 4 °C and 14000 × g for 30 min to collect the membrane fragments. Then add Membrane Protein Extraction Reagent B to fully extract the membrane proteins. Subsequently, centrifuge at 4°C and 14000 × g for 5 min, and collect the supernatant as the membrane protein solution.The isolated membrane protein was used for western blot analysis with the H+ATPase antibody (Agrisera, AS07260). Nuclear and Cytoplasmic Protein Extraction: Rice protoplasts were resuspended in pre-cooled cell lysis buffer (10 mM pH7.9 HEPES, 10 mM KCl, 1.5 mM MgCl2, 0.5% NP-40, 1 mM DTT), incubated on ice for 15 min, followed by the addition of NP-40 and another 10 min incubation on ice. The mixture was centrifuged at 300 g for 5 min to obtain the precipitate (nuclei), retaining both the supernatant and the precipitate. The precipitate was washed twice with nuclear wash buffer (10 mM pH7.9 HEPES, 10 mM KCl, 1.5 mM MgCl2) and centrifuged at 800 × g for 10 min to obtain purified nuclei. The isolated nuclear Protein was used for western blot analysis with the Histone H3 antibody (Abmart, P30266M).The supernatant was subjected to ultracentrifugation at 100000 × g for 3 h, and the resulting supernatant was collected as the cytoplasmic protein fraction.The isolated cytoplasmic protein was used for western blot analysis with the Actin antibody (Abmart, T40104S).Chloroplast protein isolation: Rice leaves was soaked in 0.5% sarkosyl solution for 3-5 min, rinsed thoroughly, and ground using a mortar and pestle in 1.25 M NaCl, 40 mM HEPES (pH 7.6), 5 mM EDTA (pH8), 0.1% bovine serum albumin, 0.1% β-mercaptoethanol following the high salt protocol. The homogenate was filtered through Miracloth and centrifuged in a microcentrifuge at 12000 × g for 20 s. With large volumes, the homogenate was centrifuged at 3000 × g for 5 min. The pellet was resuspended and washed twice in sorbitol dilution buffer (SDB: 0.33 M sorbitol, 20 mM HEPES, 2 mM EDTA, 1 mM MgCl2, 0.1% bovine serum albumin). The chloroplasts were loaded onto step gradients (30%:70% Percoll in SDB) and centrifuged for 10 min at 4 °C in a microcentrifuge. For large volumes, the step gradients were centrifuged for 30 min at 4 °C at 1500 × g. The chloroplasts were taken from the 30%:70% Percoll interface, whereas the chloroplast excluded fraction remains on the top. Chloroplasts were shock-frozen in liquid nitrogen, resuspended in 50 μL of ice-cold protein extraction buffer, and centrifuged at 15000 × g at 4 °C for 10 min. The isolated chloroplast was used for Western blot analysis with the RbcL (Beijing Protein Innovation, AbP80037-A-SE) antibody.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting the findings of this study are available within the article, Supplementary Information, Supplementary Data, and Source Data. Source data are provided with this paper.
References
Zhang, H., Zheng, X. & Zhang, Z. The Magnaporthe grisea species complex and plant pathogenesis. Mol. Plant Pathol. 17, 796–804 (2016).
Liu, M. et al. Phosphorylation-guarded light-harvesting complex II contributes to broad-spectrum blast resistance in rice. Proc. Natl. Acad. Sci. USA 116, 17572–17577 (2019).
Ren, Y., Gao, P. & Zhu, F. Meteorological grade prediction of occurrence degree of rice blast in Jiangsu. Jiangsu Agricul Sci. 44, 151–161 (2016).
Tang, D., Wang, G. & Zhou, J. M. Receptor kinases in plant-pathogen interactions: more than pattern recognition. Plant Cell 29, 618–637 (2017).
Wu, Y. & Zhou, J. M. Receptor-like kinases in plant innate immunity. J. Integr. Plant Biol. 55, 1271–1286 (2013).
Shiu, S. H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004).
Shiu, S. H. & Bleecker, A. B. Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis. Plant Physiol. 132, 530–543 (2003).
Wang, J. et al. Investigation of evolutionary and expressional relationships in the function of the leucine-rich repeat receptor-like protein kinase gene family (LRR-RLK) in the radish (Raphanus sativus L. Sci. Rep. 9, 6937 (2019).
Luo, X. et al. A Lectin Receptor-like kinase mediates pattern-triggered salicylic acid signaling. Plant Physiol. 174, 2501–2514 (2017).
Wang, Y., Nsibo, D. L., Juhar, H. M., Govers, F. & Bouwmeester, K. Ectopic expression of Arabidopsis L-type lectin receptor kinase genes LecRK-I.9 and LecRK-IX.1 in Nicotiana benthamiana confers Phytophthora resistance. Plant Cell Rep. 35, 845–855 (2016).
Cheng, X. et al. A rice lectin receptor-like kinase that is involved in innate immune responses also contributes to seed germination. Plant J. 76, 687–698 (2013).
Sun, M. et al. Ectopic expression of GsSRK in Medicago sativa reveals its involvement in plant architecture and salt stress responses. Front Plant Sci. 9, 226 (2018).
Passricha, N., Saifi, S. K., Kharb, P. & Tuteja, N. Marker-free transgenic rice plant overexpressing pea LecRLK imparts salinity tolerance by inhibiting sodium accumulation. Plant Mol. Biol. 99, 265–281 (2019).
Zhao, J. L. et al. Mutual regulation of receptor-like kinase SIT1 and B’kappa-PP2A shapes the early response of rice to salt stress. Plant Cell 31, 2131–2151 (2019).
Yu, T. F. et al. Genome-wide analysis of CDPK family in Foxtail Millet and determination of SiCDPK24 functions in drought stress. Front Plant Sci. 9, 651 (2018).
Sun, Y., Qiao, Z., Muchero, W. & JG, C. hen Lectin receptor-like kinases: the sensor and mediator at the plant cell surface. Front Plant Sci. 11, 596301 (2020).
Pang X. & Zhou H. X. Activation of signaling receptors: do ligands bind to receptor monomer, dimer, or both? BMC Biophys 6, 7 (2013).
Stock, J. Receptor signaling: dimerization and beyond. Curr. Biol. 6, 825–827 (1996).
Cao Y. et al. The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. Elife 3, e03766 (2014).
Li, J. Y., Wang, J. & Zeigler, R. S. The 3000 rice genomes project: new opportunities and challenges for future rice research. GigaScience 3, 8 (2014).
Cheng, Y. T., Zhang, L. & He, S. Y. Plant-microbe interactions facing environmental challenge. Cell Host Microbe 26, 183–192 (2019).
Fu, D. et al. A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323, 1357–1360 (2009).
Cheng, C. et al. Plant immune response to pathogens differs with changing temperatures. Nat. Commun. 4, 2530 (2013).
Xin, X. F. et al. Bacteria establish an aqueous living space in plants crucial for virulence. Nature 539, 524–529 (2016).
Nomura, K. et al. Bacterial pathogens deliver water- and solute-permeable channels to plant cells. Nature 621, 586–591 (2023).
Gandhi A. & Oelmuller R. Emerging roles of receptor-like protein kinases in plant response to abiotic stresses. Int. J. Mol. Sci. 24, 14762 (2023).
Shimizu, T. et al. Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice. Plant J. 64, 204–214 (2010).
Albrecht, C., Russinova, E., Hecht, V., Baaijens, E. & de Vries, S. The Arabidopsis thaliana somatic embryogenesis receptor-like kinases1 and 2 control male sporogenesis. Plant Cell 17, 3337–3349 (2005).
Petutschnig, E. K., Jones, A. M., Serazetdinova, L., Lipka, U. & Lipka, V. The lysin motif receptor-like kinase (LysM-RLK) CERK1 is a major chitin-binding protein in Arabidopsis thaliana and subject to chitin-induced phosphorylation. J. Biol. Chem. 285, 28902–28911 (2010).
Ao, Y. et al. OsCERK1 and OsRLCK176 play important roles in peptidoglycan and chitin signaling in rice innate immunity. Plant J. 80, 1072–1084 (2014).
Li, Z. et al. OsRLCK 57, OsRLCK107 and OsRLCK118 positively regulate chitin- and pgn-induced immunity in rice. Rice (N. Y) 10, 6 (2017).
Huang, W. R. H. et al. Receptor-like cytoplasmic kinases of different subfamilies differentially regulate SOBIR1/BAK1-mediated immune responses in Nicotiana benthamiana. Nat. Commun. 15, 4339 (2024).
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 16, 537–552 (2016).
Akamatsu, A. et al. An OsCEBiP/OsCERK1-OsRacGEF1-OsRac1 module is an essential early component of chitin-induced rice immunity. Cell Host Microbe 13, 465–476 (2013).
Wang, C. et al. OsCERK1-mediated chitin perception and immune signaling requires receptor-like cytoplasmic kinase 185 to activate an MAPK cascade in rice. Mol. Plant 10, 619–633 (2017).
Yamada, K. et al. The Arabidopsis CERK1-associated kinase PBL27 connects chitin perception to MAPK activation. Embo J. 35, 2468–2483 (2016).
Yu, X. et al. FLS2-RBOHD module regulates changes in the metabolome of Arabidopsis in response to abiotic stress. Plant Environ. Interact. 4, 36–54 (2023).
Li, C. H. et al. The receptor-like kinase sit1 mediates salt sensitivity by activating mapk3/6 and regulating ethylene homeostasis in rice. Plant Cell 26, 2538–2553 (2014).
Espinoza, C., Liang, Y. & Stacey, G. Chitin receptor CERK1 links salt stress and chitin-triggered innate immunity in Arabidopsis. Plant J. 89, 984–995 (2017).
Liu, S. H., Wang, J., Chen, K. S., Zhang, Z. H. & Zhang, P. Y. The L-type lectin receptor-like kinase (PnLecRLK1) from the Antarctic moss enhances chilling-stress tolerance and abscisic acid sensitivity in. Plant Growth Regul. 81, 409–418 (2017).
Li, W. et al. A aatural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126 e115 (2017).
Kim, J. H. et al. Increasing the resilience of plant immunity to a warming climate. Nature 607, 339–344 (2022).
Leivar, P. ablo & Quail, P. eterH. PIFs: pivotal components in a cellular signaling hub. Trends Plant Sci. 16, 19–28 (2010).
Deeken, R. & Kaldenhoff, R. Light-repressible receptor protein kinase: a novel photo-regulated gene from Arabidopsis thaliana. Planta 202, 479–486 (1997).
Qi, Z. Q. et al. Molecular detection and analysis of blast resistance genes in rice main varieties in Jiangsu province, China. Agronomy 13, 157 (2025).
Liu, M. X. et al. The multifunctional ascorbate peroxidase MoApx1 secreted by Magnaporthe oryzae mediates the suppression of rice immunity. Plant Cell 37, koaf146 (2025).
Liu, M. X. et al. Targeting Magnaporthe oryzae effector MoErs1 and host papain-like protease OsRD21 interaction to combat rice blast. Nat. Plants 10, 618–632 (2024).
Liu, M. X. et al. Auxilin-like protein MoSwa2 promotes effector secretion and virulence as a clathrin uncoating factor in the rice blast fungus Magnaporthe oryzae. N. Phytol. 230, 720–736 (2021).
Acknowledgements
This research was supported by the key program of the Natural Science Foundation of China (NSFC) (32030091, Z.Z.), the Jiangsu Provincial Excellent Youth Science Foundation (BK20230100, M.L.), the NSFC Program (32172377, M.L.), the Innovational Fund for Scientific and Technological Personnel of Hainan Province (324CXTD428, Z.Z.), the Jiangsu Agriculture Science & technology Innovation Fund Program [CX(23)1032, Z.Z.], and the Biological Breeding-National Science and Technology Major Projects (2023ZD04070, L.Y. and Yuqiang Liu).
Author information
Authors and Affiliations
Contributions
J.L., M.L. and Z.Z. conceived and designed the study. J.L., L.S. and W.W. performed experiments with phenotypic and biochemical assays. J.W., W.C., Z.Y., L.Y., X.L. and G.L. contributed reagents, plant and fungal materials. J.L., H.Z., Z.Q., Yongfeng Liu, and Yuqiang Liu collected the data. J.L., P.W., M.L. and Z.Z. analysed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, J., Su, L., Wang, W. et al. Light-induced OsLIKE1 phosphorylation enhances rice resistance against blast disease. Nat Commun 17, 810 (2026). https://doi.org/10.1038/s41467-025-67504-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67504-y