Abstract
Aging is an inevitable process integrating chronological alterations of multiple organs. A growing aging population necessitates feasible anti-aging strategies to deal with age-associated health problems. We previously performed a proteomics analysis in a healthy-aging cohort, and revealed an age-related downregulation of ARMH4. Here we generate a whole-body Armh4-knockout mouse line, and investigate its impact on systemic aging. Under normal feeding conditions, Armh4 deficiency significantly lowers spontaneous mortality and extends maximum lifespan. In the female mice, Armh4 deficiency postpones sexual maturity for one week. At the organ level, the age-related pathologies of the heart, liver, kidney, and spleen are substantially alleviated by Armh4 deletion. Mechanistically, ARMH4 interacts with IGF1R/FGFR1 to sensitize the activation of PI3K-Akt-mTORC1 and Ras-MEK-ERK pathways, consequently promoting protein synthesis and inhibiting autophagy. Moreover, ARMH4 is required for the maintenance of IGF1R/FGFR1 expressions through regulating the transcription factor c-Myc. Therefore, ARMH4 maintains a positive-feedback growth signaling to promote aging.
Similar content being viewed by others
Introduction
With the rapid advancement of socio-economic conditions and healthcare services, global life expectancy has been steadily increasing. However, as the proportion of elderly people increases, the pressure on healthcare resources and social services is continuously intensifying. Healthy aging emphasizes the importance of extending healthspan by focusing on strategies that reduce the duration of organ dysfunction and disease, rather than merely prolonging lifespan1. The concept of healthy aging provides a valuable framework for addressing the multifaceted challenges associated with aging populations.
Aging is an inevitable process characterized by the gradual decline of organ function under the combined influence of genetic and environmental factors2,3. The characteristics of aging at the cellular and molecular levels encompass multiple dimensions, including protein homeostasis imbalance, decline in autophagic function, dysregulation of nutrient sensing, exacerbation of cellular senescence, persistent chronic inflammation, and so on4,5. Autophagy is a critical mechanism of cellular self-cleaning. This process is not only vital for maintaining protein homeostasis but also broadly influences the clearance and renewal of non-protein macromolecules and subcellular organelles6. Genetic inhibition of autophagy accelerates the aging process in model organisms, and strategies capable to enhance autophagic flux can extend healthspan and lifespan7,8.
The nutrient-sensing network represents a complex interplay between extracellular ligands and their specific receptors, triggering a series of precisely regulated intracellular signaling cascades, such as PI3K-Akt-mTOR and Ras-MEK-ERK pathways9,10. At the core of this nutrient-sensing network lies the mechanistic target of rapamycin (mTOR), whose activity is intricately regulated by nutritional status and environmental growth or stress signals11. Inhibition of mTOR extends lifespan in model organisms and confers protection against age-related pathologies12,13,14,15,16, while its activation has been associated with acceleration of aging and shortened lifespan17. However, application of mTOR inhibition in human remains challenging due to its pivotal role in cell survival.
Although significantly extending human lifespan remains impossible so far, cumulating evidence suggests that caloric restriction and exercise can delay aging18,19. Furthermore, the blood of young mice has been proven to restore the function of multiple organs in old mice in heterochronic parabiosis studies, indicating that it is feasible to reverse the pathophysiological process of aging by targeting blood factors20,21. We previously performed a proteomics study in a healthy aging cohort, and identified 9 gender-independent factors of which the expressions were sensitive to aging22,23. One of these proteins is armadillo like helical domain containing 4 (ARMH4), which was clustered together with the insulin like growth factor 1 (IGF1), a canonical factor involved in development and aging24. ARMH4 is a single-pass type I membrane protein with limited information about its biological functions. ARMH4 has been shown to be expressed in the early stages of mouse embryonic heart development, especially in the progenitor cells of the first and second heart fields25. Shuster et al.26 reported that both knockdown and overexpression of ARMH4 led to incomplete dendritic development of Purkinje cells, suggesting an involvement of ARMH4 in cell differentiation. Nevertheless, the molecular function of ARMH4 in aging remains unclear.
In this study, we developed a whole-body Armh4 gene knockout mouse line, and investigated its impact on systemic aging. We found that Armh4 deficiency protects multiple organs from senescence-associated disorders and extends lifespan. ARMH4 interacts with insulin like growth factor 1 receptor (IGF1R)/fibroblast growth factor receptor 1 (FGFR1) to sensitize the activation of downstream PI3K-Akt-mTOR and Ras-MEK-ERK pathways, consequently regulating protein synthesis and autophagy. Meanwhile, ARMH4 maintains normal IGF1R/FGFR1 expressions through regulating the transcription factor c-Myc. Our findings reveal an ARMH4-mediated positive-feedback growth signaling circuit that contributes to systemic aging. Targeting ARMH4 would be a promising strategy to achieve healthy aging.
Results
Armh4 deficiency promotes healthy aging and extends longevity in mice
We previously identified ARMH4 as an aging biomarker that was significantly downregulated in the elderly in both male and female (Fig. 1a)22,23. The data from The Temporal Expression during Development Database (TEDD) revealed that ARMH4 expression was generally reduced across multiple organs in the adult stage compared to the embryonic period (Supplementary Fig. 1a), suggesting that ARMH4 is a development-sensitive factor. Moreover, analysis of the public GTEx database revealed reduced ARMH4 expression in blood samples from individuals aged 70–79 years compared to those aged 20–29 years (Fig. 1b). To corroborate these findings across species, we collected plasma samples from young (8-week-old) and aged (25-month-old) mice, and examined ARMH4 expression using Western blot analysis. The results showed that ARMH4 expression was significantly reduced in the plasma of aged mice compared to young mice (Supplementary Fig. 1b). Consistently, the expression levels of ARMH4 in the brain, lung, heart, liver, kidney, and muscle were also significantly lower in the aged mice than those in the young mice (Fig. 1c, and Supplementary Fig. 1c, d). The conserved downregulation of ARMH4 during aging implicates an important role of ARMH4 in systemic aging.
a The level of circulating ARMH4 decreased in aged people compared with young adults detected by mass spectrometry23. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent subjects. b The analysis of the public GTEx database about ARMH4 expression in blood samples from individuals aged 70-79 years compared to those aged 20-29 years. Young group are presented as box plots showing the median (0.0123), the 25th (0.0080) and 75th (0.0199) percentiles, and the minimum (0.0000) and maximum (0.0749) values, n = 69 (20–29); Old group are presented as box plots showing the median (0.0096), the 25th (0.0000) and 75th (0.0138) percentiles, and the minimum (0.0000) and maximum (0.0266) values, n = 26 (70–79). c The protein levels of Armh4 in the brain, lung, heart, liver, kidney and muscle from young (2–month-old) and aged (25-month-old) mice measured by western blot. Representative blot of n = 3 independent mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent mice. d Schematic diagram illustrating the Armh4 gene knockout (KO) strategy in mouse. e Validation of Armh4 KO at the mRNA level in mouse tissues using qRT-PCR. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. f Mendelian inheritance of the breeding offspring from heterozygous Armh4-KO mice. g Representative images showing fur appearance of wild type (WT) and Armh4-KO mice at 2 months (young) or 25 months (aged) of age. Scale bar, 1 cm. h Weights of WT and Armh4-KO mice during natural aging. Data are presented as mean values +/- SEM, with sample sizes of n = 12/9, 11/9, and 8/9 (WT/KO) for the 6–15, 18–21, and 24-month groups, respectively. i Representative Masson’s trichrome staining images showing the infrastructure of skins from young/aged WT and Armh4-KO mice. n = 3. Scale bar, 200 μm. j Representative images showing the hair regeneration capacity after excising a patch of dorsal fur from 21-month-old WT or Armh4-KO mice. n = 3. Scale bar, 1 cm. k, l Survival curves of two panels of WT and Armh4-KO mice under natural feeding conditions. The mice from the first panel (k) were euthanized at 25 months of age for molecular studies, while mice from the second panel (l) were further raised until natural death. Statistical significance was determined by a two-tailed unpaired t-test, and the exact P values are reported directly on the figures. Source data are provided as a Source Data file.
To investigate the physiological function of ARMH4, we generated a whole-body Armh4 knockout (KO) mouse line by deleting the exon 2 using the CRISPR-cas9 technology, which led to a frame shift and early translation termination of the ARMH4 protein (Fig. 1d). The ablation of Armh4 was verified by genotyping (Supplementary Fig. 1e) and quantitative real-time polymerase chain reaction (qRT-PCR; Fig. 1e). Homozygous Armh4-deficient mice demonstrated normal reproductive capabilities and exhibited standard Mendelian inheritance patterns (Fig. 1f), suggesting that ARMH4 is dispensable for fertility. However, starting from 15 months of age, the fur of wild-type (WT) mice began to exhibit dryness and greying, whereas the fur of KO mice remained dark and lustrous (Fig. 1g, Supplementary Fig. 1f). The body weights of adult Armh4-deficient mice were 2-3 g lower than WT mice (Fig. 1h, Supplementary Fig. 1g). After 15 months, the weight gain of WT mice gradually slowed down, while the body weights of Armh4-deficient mice remained steadily growing until 24-month-old (Fig. 1h). Masson staining revealed that the loss of Armh4 significantly mitigated the age-related wrinkling of the epidermis and the structural disorganization of the dermis and hypodermis (Fig. 1i). We subsequently assessed hair regeneration capacity by excising a patch of dorsal fur in 21-month-old mice. The results indicated that the hair regrowth rate in KO mice was remarkably faster than that in WT mice (Fig. 1j).
Next, we assessed frailty index (FI) scores and evaluated motor function, including running endurance, rotarod performance, and grip strength, in both male and female mice at 15 months of age (Supplementary Fig. 1h–k). The FI scores of KO mice did not show significant difference from WT mice in female. However, in male, WT mice exhibited pronounced loss of hair pigmentation, whereas KO mice remained unaffected alongside aging (Supplementary Fig. 1f). In the tests assessing motor coordination and balance (running endurance and rotarod performance), male KO mice demonstrated significantly enhanced physical endurance and balance compared to their WT counterparts (Supplementary Fig. 1i-k). These results suggest that the deletion of Armh4 promotes healthy aging.
From 20 months on, WT mice began to exhibit instances of spontaneous mortality, while KO mice did not show any deaths in a cohort that was euthanized at 25 months for molecular assessments (Fig. 1k), suggesting that ARMH4 depletion lowers aging-associated mortality. In a separate cohort, we further housed the mice (5 WT and 4 KO) to natural death, and found that Armh4 knockout extended maximum lifespan (Fig. 1l). Collectively, these data suggest that ARMH4 depletion delays aging and extends lifespan in mice.
Armh4 deficiency postponed puberty in female mice
As evolutionary theories predict, lifespan is consistently correlated with the age of reproduction across mammalian species27. Furthermore, in inbred mouse strains, the timing of sexual maturity is also positively associated with lifespan, with IGF1 identified as a crucial mediator28. We then tested whether Armh4 deficiency influence the timing of puberty. In C57BL6J mice, sexual maturity typically occurs from 6 to 8 weeks of age. Vaginal smear test29 showed that, in female WT mice, a significant number of anucleate cornified cells appeared at 6 weeks of age, whereas this physiological change in KO female mice occurred only till 7 weeks of age (Supplementary Fig. 2a). Gross examination and hematoxylin and eosin (H&E) staining of the ovaries revealed that female KO mice had a greater number of pre-ovulatory follicles30 at 7 weeks of age, when female WT mice already exhibited a predominance of mature follicles (Supplementary Fig. 2b, c). Enzyme-linked immunosorbent assay (ELISA) showed that the ratio of luteinizing hormone (LH) to follicle-stimulating hormone (FSH), an index of ovulation31, was significantly lower in KO mice compared to WT mice (Supplementary Fig. 2d). These data suggest that Armh4 deficiency delays sexual maturation in female mice. Notably, no obvious effects of Armh4 knockout on male sexual maturation were observed in terms of testis weight, sperm count, the ratio of LH/FSH, or testosterone at 7 weeks of age (Supplementary Fig. 2e–h).
Armh4 deficiency prevents cellular senescence and metabolic dysfunction in the liver
We next investigated the impact of Armh4 deficiency on the aging process at the organ level. Although there was no significant difference in body weight between 25-month-old Armh4-KO and WT mice (Fig. 2a), heart and lung weights were significantly lower in aged Armh4-KO mice compared to WT mice (Fig. 2b). Liver weights of aged mice with Armh4 deficiency also exhibited a decrease trend, despite not approaching statistical significance due to large variations (P = 0.132; Fig. 2b, c). H&E staining of liver tissues showed that Armh4 KO significantly ameliorated immune cell infiltration in aged mice (Fig. 2d). Masson trichrome staining showed that aging-related hepatic fibrosis was significantly alleviated by Armh4 deletion (Fig. 2e), which correlated with a marked reduction in the expression of fibrosis marker genes, including Acta2, Col1a1, and Ki67 (Fig. 2f). Aged livers exhibited a robust increase in β-galactosidase (β-Gal) activity compared to young livers32, however, this feature was significantly reversed in Armh4-KO mice (Fig. 2g). We then examined the factors involved in senescence-associated secretory phenotype (SASP)32. qRT-PCR assay showed that Armh4 deficiency significantly suppressed aging-related inductions of SASP factors, including p16, p21, p53, Il1b, and Tnfa, except for Il6 (Fig. 2h). These data suggest that ARMH4 inhibition suppresses cellular senescence in the liver. Moreover, aging-associated morphological and molecular changes in spleen and kidney were substantially alleviated in Armh4-deficient mice compared to WT mice (Supplementary Fig. 3 and Supplementary Fig. 4), suggesting that ARMH4 functions as a systemic pro-aging factor in multiple organs.
a Body weight measurements of young (2-month-old) and aged (25-month-old) male WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 9 (WT young), 6 (KO young), 7 (WT aged), and 7 (KO aged) independent mice. b Organ weight measurements of young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 9 (WT young), 6 (KO young), 7 (WT aged), and 7 (KO aged) independent mice. c Representative liver images from young and aged WT and Armh4-KO mice. Scale bar, 5 mm. d H&E staining of the liver and statistical analysis of inflammatory cell infiltration in young and aged WT and Armh4-KO mice. Scale bar, 50 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 18 fields from 3 independent mice per group. e Liver Masson staining and statistical results in young and aged WT and Armh4-KO mice. Scale bar, 50 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 15 fields from 3 independent mice per group. f Liver fibrosis index qRT-PCR results in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. g Liver β-gal staining and statistical results in young and aged WT and Armh4-KO mice. Scale bar, 200 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 15 fields from 3 independent mice per group. h qRT-PCR results for liver aging and inflammation markers in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. i Blood glucose levels during GTT in 21-month-old mice. n = 10 (WT), 9 (KO). j qRT-PCR results for liver glucose metabolism markers in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. k qRT-PCR results for liver metabolic function markers in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. l qRT-PCR results for liver fatty acid metabolism markers in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. m–q ELISA quantification of serum metabolic profiles in young and aged WT versus Armh4-KO mice at 8-week (young) or 25-month (aged) old: (m) Triglycerides (TG), (n) total cholesterol (TCh), (o) high-density lipoprotein cholesterol (HDL-C), (p) low-density lipoprotein cholesterol (LDL-C), (q) AST/ALT. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 (young groups) and n = 6 (aged groups) independent mice. Statistical significance was determined by two-way ANOVA, and the exact P values are reported directly on the figures. Source data are provided as a Source Data file.
Given that most aging-related metabolic events occur in the liver33,34, we then investigated whether ARMH4 influences metabolic pathways. We performed the glucose tolerance test (GTT) in 21-month-old male mice, and found that Armh4 KO substantially compromised glucose clearance upon glucose challenge compared to WT controls (Fig. 2i). However, Armh4 deficiency did not affect the expression levels of genes involved in glucose handling pathways in the liver (Fig. 2j). GTT in 15-month-old mice showed that Armh4 deficiency slightly slowed glucose clearance in both males and females (Supplementary Fig. 5a, b). We next subjected the mice to a 24-h short-term fasting challenge followed by refeeding (Supplementary Fig. 5c). The results showed that Armh4-KO mice of both sexes consumed approximately 20% less food than their WT counterparts during refeeding (Supplementary Fig. 5c). Additionally, Armh4-KO mice exhibited significantly less weight loss after fasting, as well as less weight regain post-refeeding, than WT controls (Supplementary Figs. 5d, e). However, Armh4 deficiency had marginal effects on fasting/refeeding-associated changes in blood glucose (Supplementary Figs. 5f, g) and insulin (Supplementary Figs. 5h, i). These data suggest that ARMH4 regulates glucose homeostasis via a yet unidentified insulin-independent regulatory mechanism.
When looking into lipid metabolism, we found that the aged liver exhibited substantially increased expression of genes involved in cholesterol metabolism, including Hmgcr, Cyp7a1, Abcg1, and Ldlr, which were largely reversed by Armh4 deletion (Fig. 2k). While Armh4 deficiency did not affect fatty acid biosynthesis or oxidation, it significantly suppressed the aging-associated induction of Fatp1 and Cd36 (Fig. 2i), factors responsible for fatty acid transport35,36. In despite of the changes in gene reprogramming related to lipid metabolism, no significant differences were observed for blood triglycerides (TG), total cholesterol (TCh), high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), or the AST/ALT ratio (Fig. 2m–q). These data suggest that the function of ARMH4 in metabolism might be complex.
Armh4 deficiency decelerates cardiac aging
During aging, the heart undergoes myocardial remodeling similar to pathological cardiac hypertrophy37,38. While Armh4 KO did not affect cardiac contractile function throughout aging (Fig. 3a–c), echocardiographic assessment of the E/A ratio in 15-month-old mice revealed improved diastolic function of the hearts from aged Armh4-KO mice compared to those from WT controls (Fig. 3d). Furthermore, Armh4 deficiency substantially decelerated aging-associated enlargement of the left ventricular chamber and attenuated the increase in left ventricular posterior wall thickness (Supplementary Fig. 6a–d). Consistently, Armh4 KO significantly reduced the age-dependent increase in left ventricular mass as calculated from short-axis echocardiography (Fig. 3e).
a Representative echocardiographic images parasternal short-axis view (SAX) at the papillary muscle level from young and aged WT and Armh4-KO mice heart. b, c The impacts of Armh4 deficiency on ejection fraction (EF%) (b) and fractional shortening (FS%) (c). Data are presented as mean values +/- SEM. with sample sizes of n = 12/9, 11/9, and 8/9 (WT/KO) for the 6-15, 18-21, and 24-month groups, respectively. Statistical significance was determined by a two-tailed unpaired t-test. d Representative echocardiographic images on the apical four-chamber view (A4C) from WT and Armh4-KO mice (male and female) heart, and mitral E/A ratio was measured. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 16 (WT) and n = 10 (KO) independent mice at 15 months of age. Statistical significance was determined by a two-tailed unpaired t-test. e The impacts of Armh4 deficiency on left ventricle (LV) mass. Data are presented as mean values +/- SEM, with sample sizes of n = 12/9, 11/9, and 8/9 (WT/KO) for the 15, 18-21, and 24-month groups, respectively. Statistical significance was determined by a two-tailed unpaired t-test. f Representative heart images from young and aged WT and Armh4-KO mice. Scale bar, 5 mm. g H&E staining of the heart in young and aged WT and Armh4-KO mice. Scale bar, 50 μm. h WGA staining of the heart and statistical results of cardiomyocyte area in young and aged WT and Armh4-KO mice. Scale bar, 50 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 24 fields from 3 independent mice per group. Statistical significance was determined by two-way ANOVA. i Heart masson staining and statistical results in young and aged WT and Armh4-KO mice. Scale bar, 50 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 15 fields from 3 independent mice per group. Statistical significance was determined by two-way ANOVA. j qRT-PCR results of cardiac hypertrophy-related genes in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. Statistical significance was determined by two-way ANOVA. k qRT-PCR results of cardiac SASP-related genes in young and aged WT and Armh4-KO mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 5 independent mice. Statistical significance was calculated with two-way ANOVA. l, PCA analysis results of cardiac RNA-seq from young and aged WT and Armh4-KO mice, with sample sizes of n = 4 independent mice. m, Venn diagram analysis of differential genes from young and aged WT and Armh4-KO mice, with sample sizes of n = 4 independent mice. n, Heatmap analysis of differentially expressed genes across WT-young, WT-aged, and Armh4-KO groups, with sample sizes of n = 4 independent mice. o, Ericher analysis of differential expressed genes between WT and Armh4-KO aged mice, with sample sizes of n = 4 independent mice. The exact P values are reported directly on the figures. Source data are provided as a Source Data file.
Anatomy of the 25-month mice showed that, consistent with a decrease in heart weight (Fig. 2b), the heart size of aged Armh4-KO mice was smaller than that of WT mice (Fig. 3f). H&E staining and wheat germ agglutinin (WGA) staining revealed that the aging-related enlargement of cardiomyocyte cross-section area was largely blunted by Armh4 deficiency (Fig. 3g, h). Masson trichrome staining showed that Armh4 deficiency significantly reduced myocardial fibrosis occurred alongside aging (Fig. 3i). Consistently, Armh4 KO significantly blocked the age-related inductions of hypertrophy markers, including Nppa and Myh7, albeit that no statistical significances was achieved for Nppb, Myh6 or Acta1 (Fig. 3j). Moreover, the expressions of SASP factors were significantly lower in aged Armh4-KO mice than that in WT mice (Fig. 3k). These data suggest that ARMH4 inhibition delays cardiac aging.
To explore the underlying mechanisms, we performed an RNA sequencing analysis using the left ventricular tissues from young and aged mice with or without Armh4 KO. Principal component analysis (PCA) showed that the absence of Armh4 caused a global impact on transcriptome in young mice, and slightly shifted the gene expression pattern towards juvenescence in aged mice (Fig. 3l). Venn diagram showed that there were more overlapped items between the upregulated genes in the aged hearts and the downregulated genes after Armh4 deletion than the parallel comparison, and vice versa (Fig. 3m). Consistently, heatmap analysis revealed largely attenuated changes of the aging-sensitive genes at the transcriptome level (Fig. 3n), particularly those involved in aging and longevity, such as Zbp1, S100a9, B2m, and Hspa1b39,40,41,42. Functionally, these genes mainly covered immune-related pathways in the functional enrichment analysis (Fig. 3o). Gene set enrichment analysis (GSEA) also revealed an enrichment of inflammatory response pathways in the downregulated genes after Armh4 KO (Supplementary Fig. 6e–j). These data suggest that ARMH4 may contribute to the aging-related chronic inflammation in the heart.
To further explore the functional impact of ARMH4, we employed an in vitro senescence model using human AC16 cardiomyocytes with H2O2 treatment (400 μM)43. Knockdown of Armh4 with specific siRNA significantly reversed the enhancement of senescence-associated β-galactosidase (SA-β-gal) activity after H2O2 treatment (Fig. 4a, b). On the other hand, the SA-β-gal signal was significantly aggravated by Armh4 overexpression (Fig. 4c, d). Consistently, in mouse embryonic fibroblasts (MEFs) isolated from Armh4+/+, Armh4+/- (heterozygotes), and Armh4-/- (homozygotes) embryos (E13.5), the H2O2-induced expressions of SASP factors (p16, p21, p53, Il1b, Il6, and Tnfa) were largely inhibited by Armh4 deficiency in a dose-dependent manner (Fig. 4e). Furthermore, we also observed a detrimental role of ARMH4 in cell senescence in human umbilical vein endothelial cells (HUVECs) treated with D-galactose (D-gal; 10 mg/mL) (Fig. 4f–i). These data suggest that ARMH4 is a general pro-senescence factor across species and cell types.
a Validation of siRNA-mediated Armh4 knockdown efficiency in AC16 cells. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by two-way ANOVA. b Quantification of SA-β-gal cells in H2O2 (400 μM)-induced senescence model of AC16 cells following Armh4 knockdown. Scale bar, 200 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for 12 fields from 3 biological replicates. Statistical significance was determined by two-way ANOVA. c Validation of Armh4 overexpression efficiency via plasmid transfection in AC16 cells. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by two-way ANOVA. d Quantification of SA-β-gal cells in H2O2 (400 μM)-induced senescence model of AC16 cells following Armh4 overexpression. Scale bar, 200 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for 12 fields from 3 biological replicates. Statistical significance was determined by two-way ANOVA. e qRT-PCR results of SASP-related genes and Armh4 in WT, HET and KO MEFs. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was calculated with one-way ANOVA. f Validation of siRNA-mediated Armh4 knockdown efficiency in HUVECs. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by two-way ANOVA. g Quantification of SA-β-gal cells in D-Gal (10 mg/mL)-induced senescence model of HUVECs following Armh4 knockdown. Scale bar, 200 μm. Graph bars represent the mean percentage of SA-β-gal-positive area in Armh4-knockdown cells treated with PBS or D-Gal, relative to their respective negative controls ± SEM. Black circles represent individual data points for 9 fields from 3 biological replicates. Statistical significance was determined by two-way ANOVA. h Validation of Armh4 overexpression efficiency via plasmid transfection in HUVECs. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by two-way ANOVA. i Quantification of SA-β-gal cells in D-Gal (10 mg/mL)-induced senescence model of HUVECs following Armh4 overexpression. Scale bar, 200 μm. Data are presented as mean values +/- SEM. Black circles represent individual data points for 9 fields from 3 biological replicates. Statistical significance was determined by two-way ANOVA. The exact P values are reported directly on the figures. Source data are provided as a Source Data file.
ARMH4 regulates protein homeostasis via mTOR in cardiomyocytes
Imbalance of protein homeostasis and decline in autophagic function are key hallmarks of cellular senescence4. Beyond the inflammatory pathways, ribosome was also identified as a prominent category among the downregulated genes following Armh4 KO in GSEA (Supplementary Fig. 7a), suggesting an impact of ARMH4 on translation activity. We then assessed translation activity using a puromycin incorporation assay in MEFs44. Compared to WT MEFs, Armh4-KO MEFs exhibited a significant reduction in protein synthesis rate (Fig. 5a). Consistently, in isolated neonatal rat ventricular myocytes (NRVMs), silencing Armh4 with a specific siRNA significantly suppressed protein synthesis rate, while adenovirus-mediated Armh4 overexpression significantly enhanced protein synthesis rate (Fig. 5b). Interestingly, administration of exogenous recombinant ARMH4 protein similarly promoted protein synthesis in NRVMs (Fig. 5c), verifying that the regulation of translation activity is derived from the circulating ARMH4.
a Puromycin incorporation assay of WT and Armh4-deficient MEFs to assess the effect on protein synthesis. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. b Puromycin incorporation assay of NRVMs with/without siRNA-mediated knockdown of Armh4 or adenoviral overexpression of Armh4. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. c Puromycin incorporation assay of NRVMs with/without exogenous administration of recombinant ARMH4. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. d Western blot analysis of autophagy-related markers in WT and Armh4-deficient MEFs. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. e Immunofluorescence analysis using GFP-LC3-RFP plasmid labeling in WT and Armh4-deficient MEFs to evaluate the impact of Armh4 knockout on autophagy. Scale bar, 20 μm. f, g Western blot analysis for autophagy-related markers p62 and LC3 in NRVMs with/without siRNA-mediated knockdown of Armh4 (f) or adenoviral overexpression of Armh4 (g). Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. h Western blot analysis for autophagy-related markers p62 and LC3 in NRVMs treated with/without recombinant ARMH4. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test, and the exact P values are reported directly on the figures. Source data are provided as a Source Data file.
We next examined the impact of ARMH4 on autophagy. Loss of Armh4 significantly reduced the expression of p62 and enhanced the LC3 II/I ratio in MEFs (Fig. 5d), suggesting an augment of autophagic activity after Armh4 deficiency. Moreover, we transfected the cells with a plasmid expressing a GFP-LC3-RFP fluorescent probe capable to indicate autophagic flux45. Fluorescence imaging showed that the autophagic flux was remarkably enhanced in Armh4-KO MEFs, whereas it could be completely blunted by ectopically overexpressing Armh4 (Fig. 5e). In NRVMs, whereas Armh4 knockdown caused a significant decrease in p62 and an enhancement of the LC3 II/I ratio (Fig. 6f), both genetic overexpression of Armh4 and administration of recombinant ARMH4 protein conversely suppressed the autophagic activity (Fig. 5g, h).
a Western blot analysis of phosphorylated p70S6K, Akt, ERK, and LC3 in the hearts of young and aged WT and KO mice. Representative blot of n = 3 independent mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent mice. Statistical significance was determined by two-way ANOVA. b Western blot analysis of phosphorylated, p70S6K, Akt, ERK, and LC3 in the livers of young and aged WT and KO mice. Representative blot of n = 3 independent mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent mice. Statistical significance was determined by two-way ANOVA. c Western blot analysis of phosphorylated mTOR, p70S6K, Akt, ERK in Armh4-deficient MEFs compared to the WT controls. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. d, e Western blot analysis of phosphorylated mTOR, p70S6K, Akt, and ERK in NRVMs with/without siRNA-mediated knockdown of Armh4 (d) and adenoviral overexpression of Armh4 (e). Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. f Western blot analysis of phosphorylated p70S6K, Akt, and ERK in NRVMs treated with recombinant ARMH4. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. The exact P values are reported directly on the figures. Source data are provided as a Source Data file.
Protein synthesis and autophagy are coordinated cell processes integrally controlled by the mTOR signaling pathway, which plays a central role in growth and aging14,46,47. We then investigated the impact of ARMH4 on the mTOR signaling pathway. Western blot analysis for proteins isolated from cardiac and hepatic tissues revealed that Armh4 deficiency substantially attenuated the phosphorylation levels of p70S6K and Akt (Fig. 6a, b), substrates of mTOR complex 1 and 2 (mTORC1/2), respectively48,49. Concurrently, the LC3-II/I ratio was significantly enhanced by Armh4 deletion (Fig. 6a, b). In MEFs, loss of Armh4 substantially suppressed the phosphorylation of mTOR as well as the phosphorylations of p70S6K and Akt compared to the WT control (Fig. 6c). Consistently, these phosphorylations were largely reduced by Armh4 knockdown, but enhanced by ectopically overexpression of Armh4 and administration of recombinant ARMH4 protein in NRVMs (Fig. 6d–f). These data suggest that ARMH4 may promote aging-related protein dyshomeostasis through activating both mTORC1 and mTORC2.
ARMH4 regulates nutrient-sensing signaling networks
Interestingly, manipulating the expression of ARMH4 also positively regulated the phosphorylation level of extracellular signal-regulated kinase (ERK) in addition to the mTOR signaling cascades (Fig. 6a–f), suggesting that the regulation by ARMH4 takes place in very upstream of these nutrient-sensing signaling pathways. Growth factor receptors, particularly IGF1R and FGFR1, regulates both mTOR and mitogen-activated protein kinase (MAPK) pathways (Fig. 7a)50,51. In the co-immunoprecipitation (co-IP) assays using HEK293 cells ectopically expressing FLAG-tagged ARMH4 and HA-tagged IGF1R/FGFR1, we found that ARMH4 could pull down both IGF1R and FGFR1 (Fig. 7b). Immunofluorescence assay showed that ARMH4 co-localized with both IGF1R and FGFR1 on cell membrane (Fig. 7c). Immunoprecipitation (IP) assay with antibodies targeting endogenous ARMH4, IGF1R, and FGFR1 further confirmed the interaction between ARMH4 and IGF1R/FGFR1 in heart tissues in vivo (Fig. 7d, e). Structural modeling using AlphaFold 3 ref. 52 revealed that presence of ARMH4 encompassed the dimerized receptors, and stabilized the interactions of these receptors with their corresponding ligands, IGF1 and fibroblast growth factor (FGF), respectively (Fig. 7f). Particularly, in the IGF1-IGF1R complex, ARMH4 exhibited a predictable filamentous α-helix structure rather than a promiscuous loop structure in the FGF-FGFR1 complex (Fig. 7f), suggesting structural flexibility of ARMH4 in its molecular function. To further explore potential interfaces underlying ARMH4-IGF1R interaction, we employed AlphaFold 3 for structural prediction (Fig. 7g), and identified five candidate domains in ARMH4 possibly responsible for its interaction with IGF1R (Fig. 7h). We then constructed Flag-tagged ARMH4 truncations deleting each of these domains, and performed co-IP assays together with HA-tagged IGF1R. The results showed that deletion of residues 611-621 (EGQEDEDEEDE), but not 189-199 or 541-546, largely blocked the interaction between ARMH4 and IGF1R, albeit that the function of 420-434 and 698-705 could not be determined due to their protein instability (Fig. 7i). These results suggest that the 611-621 segment rich in negatively charged amino acids in ARMH4 might be essential for its interaction with IGF1R.
a Schematic diagram of IGF1R and FGFR1 activation pathways. b, c Co-IP (b) and immunofluorescence (c) to detect the interaction between ARMH4 and IGF1R/FGFR1 in HEK293 cells co-transfected with pcDNA3.1-hArmh4-3FLAG and pcDNA3.1-hIgf1r-HA, as well as pcDNA3.1-hArmh4-3FLAG and pcDNA3.1-hFgfr1-HA, Scale bar, 10 μm. All experiments were performed twice independently with comparable results, and representative data from one experiment are shown. d, e Validation of ARMH4-IGF1R (d) and ARMH4-FGFR1 (e) interaction using endogenous IP in heart tissues. All experiments were performed twice independently with comparable results, and representative data from one experiment are shown. f AlphaFold 3 simulation of the effect of ARMH4 on ligand-receptor binding for IGF1/IGF1R and FGF2/FGFR1. g AlphaFold3 simulation of the specific binding region of ARMH4 within the IGF1/IGF1R ligand-receptor complex. h Schematic diagram of the designed plasmid construct targeting the predicted binding sequence between ARMH4 and IGF1R. i Co-IP analysis of the interaction between IGF1R and ARMH4, including its isoforms, in HEK293 cells. All experiments were performed twice independently with comparable results, and representative data from one experiment are shown. Source data are provided as a Source Data file.
To verify the involvements of IGF1R and FGFR1 in ARMH4-mediated signal transduction, we examined the impacts of two inhibitors, picropodophyllin (PPP) and ferulic acid sodium (FA), specific for IGF1R and FGFR1, respectively. Treatments with either PPP (2.5 μM) or FA (2.5 μM) blocked the enhancement of ERK phosphorylation by Armh4 overexpression in NRVMs (Fig. 8a, b). Surprisingly, PPP treatment only blocked the ARMH4-mediated phosphorylation of Akt but not the phosphorylation of p70S6K; on the contrary, FA treatment only blocked the ARMH4-mediated phosphorylation of p70S6K but not the phosphorylation of Akt (Fig. 8a, b). These data suggest that, in despite of shared functions, IGF1R and FGFR1 differentially mediate the downstream activation of mTORC1 and mTORC2 by ARMH4. Furthermore, treatment with PPP or FA partially reversed the inhibition of autophagy (Fig. 8c, d) and the activation of translation (Supplementary Fig. 7b, c) induced by Amrh4 overexpression, suggesting that the impacts of ARMH4 on autophagy and translation activities are derived from combined effects of IGF1R and FGFR1 activations.
a, b Adenoviral overexpression of Armh4 with PPP (a) and FA (b) treatment in NRVMs followed by Western blot analysis of phosphorylated mTOR, p70S6K, Akt and ERK. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. c, d Adenoviral overexpression of Armh4 with PPP (c) and FA (d) treatment in NRVMs followed by western blot to detect the autophagy-related markers. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. e, f Adenoviral overexpression of Armh4 with siRNA-mediated knockdown of Igf1r (e) and Fgfr1 (f) in NRVMs followed by western blot analysis of phosphorylated p70S6K, Akt, ERK and LC3. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. g, h Adenoviral overexpression of Armh4 with IGF1 (g) and FGF2 (h) treatment in NRVMs followed by western blot analysis of phosphorylated p70S6K, Akt, ERK and LC3. n = 3. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by two-way ANOVA, and the exact P values are reported directly on the figures. Source data are provided as a Source Data file.
We next directly silenced Igf1r and Fgfr1 using specific siRNAs to further validate the downstream regulations. Unlike the effects of pharmaceutical inhibitors, knockdown of either receptor significantly abrogated the enhanced phosphorylations of p70S6K, Akt, and ERK by ARMH4 overexpression, while concurrently rescuing its suppressive effect on the LC3-II/I ratio (Fig. 8e, f). To determine preferential regulations of downstream effectors by IGF1R and FGFR1, we treated the cells with recombinant IGF1 and FGF ligands. The results showed that Armh4 knockdown largely abolished the enhanced phosphorylations of p70S6K, Akt, and ERK by IGF1 or FGF to diversified extent (Fig. 8g, h). Notably, FGF2 exhibited superior efficacy in activating ERK compared to IGF1 (Fig. 8g, h). These results suggest that ARMH4 sensitizes IGF1R and FGFR1 receptors to their corresponding ligands, and facilitates downstream activation of the nutrient-sensing signaling networks.
ARMH4 maintains a positive-feedback growth signaling circuit mediated by c-Myc
To our surprise, when investigating the molecular details about the activations of IGF1R and FGFR1 by ARMH4, we observed that the protein levels of both IGF1R and FGFR1 were significantly down-regulated in Armh4-deficient cardiac/hepatic tissues and MEFs compared to corresponding WT controls (Fig. 9a–c). These changes were obviously attributable to transcriptional regulations, since Armh4 KO significantly reduced their mRNA levels in MEFs (Supplementary Fig. 8a). Consistently, Armh4 knockdown in NRVMs significantly reduced both mRNA and protein levels of IGF1R/FGFR1 (Fig. 9d, and Supplementary Fig. 8b). On the other hand, Armh4 overexpression via adenoviral vectors or administration of recombinant ARMH4 protein conversely increased IGF1R/FGFR1 expression (Fig. 9e, f; Supplementary Fig. 8c–e). Furthermore, IGF1 administration in NRVMs induced an upregulation of IGF1R, and FGF2 administration induced an upregulation of FGFR1 both in ARMH4-dependent manners (Supplementary Fig. 8f–g). Nevertheless, neither IGF1 expression in the heart nor its circulating level in plasma was affected by Armh4 KO in mice (Supplementary Fig. 8h, i). These data suggest a transcriptional regulation of IGF1R/FGFR1 by ARMH4, thereby contributing as a positive feedback circuit of the growth factor signaling downstream of ARMH4.
a Western blot analysis of FGFR1, IGFR1 protein expression in the hearts of young and aged WT and KO mice. Representative blot of n = 3 independent mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent mice. Statistical significance was determined by two-way ANOVA. b Western blot analysis of FGFR1, IGFR1 protein expression in the livers of young and aged WT and KO mice. Representative blot of n = 3 independent mice. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 independent mice. Statistical significance was determined by two-way ANOVA. c Western blot analysis of FGFR1, IGFR1 protein expression in Armh4-deficient MEFs. Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. d, e Western blot analysis of FGFR1 and IGFR1 protein expression in NRVMs with/without siRNA-mediated knockdown of Armh4 (d) and adenoviral overexpression of Armh4 (e). Representative blot of n = 3 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. f Western blot analysis of FGFR1 and IGFR1 protein expression in NRVMs treated with recombinant ARMH4. n = 6. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. The exact P values are reported directly on the figures. Source data are provided as a Source Data file.
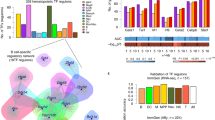
To explore the possible mediator of this positive feedback circuit, we predicted the transcription factors targeting the promoter regions of Igf1r and Fgfr1 using TF-Target Finder (TFTF)53. The results showed that the promoter regions of Igr1r and Fgfr1 shared binding motifs of a number of transcription factors, including c-Myc (Fig. 10a), which has been previously found to be a downstream target of mTOR54. Immunoblotting assay showed that the protein level of c-Myc was reduced by Armh4 knockdown but enhanced by its overexpression in NRVMs (Fig. 10b). Nevertheless, the mRNA level of c-Myc was not sensitive to ARMH4 manipulations (Fig. 10c, d), suggesting a post-transcriptional regulation of c-Myc by ARMH4. We then constructed a luciferase reporter containing the 5’-UTR region of c-Myc (Fig. 10e), and evaluated its impact on translation activity. The results showed that ARMH4 overexpression significantly augmented the translation efficiency of luciferase containing c-Myc 5’-UTR (Fig. 10e, f), indicating a regulation of c-Myc expression at the translational level. Bioinformatic analysis using the JASPAR database identified putative c-Myc binding motifs (CACGTG) within the promoter regions of both Igf1r and Fgfr1 (Fig. 10g, h). Silencing c-Myc in NRVMs significantly reduced the expressions of IGF1R and FGFR1 (Fig. 10i) and significantly blunted the upregulation of IGF1R and FGFR1 induced by Armh4 overexpression (Fig. 10j). Consistently, c-Myc knockdown substantially suppressed the ARMH4-mediated phosphorylations of p70S6K and Akt (Fig. 10k); and it also significantly reversed the ARMH4-mediated enhancement of protein synthesis rate and suppression of autophagy (Fig. 10l, m). These data suggest that c-Myc is involved in the positive feedback growth signaling circuit governed by ARMH4 during aging.
a TF-Target Finder predicted the transcription factors that co-regulated by IGF1R and FGFR1 screening diagram. b Western blot analysis of c-Myc protein expression in NRVMs upon Armh4 knockdown and overexpression. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. c, d qRT-PCR of Myc expression in NRVMs upon Armh4 knockdown (c) and overexpression (d). Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. e Schematic diagram of the dual-luciferase reporter constructs. f Dual-luciferase reporter assay in HEK293 cells. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 3 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. g Screening the c-Myc binding motifs by JASPAR. h Computational prediction of c-Myc binding motifs in IGF1R/FGFR1 regulatory regions using JASPAR. i Western blot analysis of c-Myc, IGF1R, FGFR1 expression in NRVMs upon Myc knockdown by siRNA. Representative blot of n = 6 biological replicates. Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 6 biological replicates. Statistical significance was determined by a two-tailed unpaired t-test. j–m Western blot analysis of IGF1R and FGFR1 expression (j), phosphorylated p70S6K, Akt, ERK (k), puromycin incorporation (l), the autophagy-related marker (m) in NRVMs with Myc knockdown followed by Armh4 overexpression. Representative blot of n = 4 biological replicates (j, k, m) and n = 3 biological replicates (l). Data are presented as mean values +/- SEM. Black circles represent individual data points for n = 4 biological replicates (j, k, m) and n = 3 biological replicates (l). Statistical significance was determined by two-way ANOVA. The exact P values are reported directly on the figures. Source data are provided as a Source Data file.
Discussion
The application of anti-aging strategies developed from animal studies to human remains a big challenge due to the genomic difference between species. Our previous study identified ARMH4 as an age-sensitive factor in a healthy-aging human cohort22,23. This study revealed that ARMH4 functions as a pro-aging factor by maintaining a positive-feedback circuit between IGF1R/FGFR1 signaling activity and their expressions. ARMH4 inhibition ameliorates cell senescence and decelerates systemic aging at multiple levels, suggesting that ARMH4 would be a promising target to promote healthy aging and extend lifespan (Supplementary Fig. 9a)55.
Aging is associated with changes in circulating levels of various molecules5. We previously identified ARMH4 in the blood that was down-regulated in a healthy-aging human cohort23. Both animal and cell experiments verified that ARMH4 functions as a pro-aging/pro-senescence factor by sensitizing growth factor signaling. Therefore, it is rationale to deduce that the downregulation of ARMH4 might be a compensatory effect due to enhanced cellular responses to growth signaling during aging. Nevertheless, the natural downregulation of ARMH4 expression might not be enough to significantly influence the aging process, given that genetic deletion of Armh4 offers prominent impacts of aging deceleration. These findings suggest that endogenous ARMH4 magnifies the nutrient-sensing network in cells, thereby functions as a driver of systemic aging.
ARMH4 was originally identified by Lee et al.56 in 2014, when it was firstly named as upstream of mTORC2 (UT2), given after its declared function interacting with and inhibiting RICTOR, an essential component of mTORC2, in hematopoietic progenitor cells. Beyond this, ARMH4 has also been shown negatively regulating the activation of STAT3 by interacting directly with GP130 and inhibiting the phosphorylation of STAT3 on tyrosine 705 ref. 57 De Luca et al.58 reported that overexpression of ARMH4 in primary renal epithelial cells reduced the expression of inflammatory mediators IL-1B and IL-8. In this study, we provided the first in vivo evidence for ARMH4’s intrinsic function using a whole-body knockout mouse model. Our data showed that genetic deletion of Armh4 substantially inhibited chronic inflammation in various organs during the aging process (Figs. 2h, 3k; and Supplementary Fig. 3d, 4d). Additionally, RNA-seq analysis revealed a globally suppressive effect of Armh4 knockout on immune responses (Fig. 3o; and Supplementary Fig. 6e), suggesting a pro-inflammatory role of endogenous ARMH4 during aging. This was further substantiated in cell-senescence models using human cell lines, such as AC16 and HUVECs (Figs. 4a–d, 4f–i). These findings shed lights on the molecular function of ARMH4 in immune activity, particularly in aging-associated inflammation.
Although ARMH4 is a global protein expressed in multiple organs, it is highly enriched in the progenitor cells of the first and second heart fields during early stages of embryonic development, and may serve as a specific biomarker for myocardial derivatives of the first and second heart fields as a downstream responsive gene of the Wnt/β-catenin signaling pathway25. In kidney, the expression of ARMH4 increases with the differentiation of podocytes, governed by the transcription factor LMX1B58. ARMH4 can also be found in the nervous system of mouse embryos. Shuster et al.26 reported that ARMH4 was involved in the development of Purkinje cell dendrites, where both knockdown and overexpression of ARMH4 led to incomplete dendritic development. Despite the early evidence for the involvement of ARMH4 in development, its impact on systemic aging remains undetermined. This study fills the gap by identifying ARMH4 as a master pro-aging factor in multiple organs. In particular, RNA-seq analysis using heart tissues revealed that Armh4 ablation largely reversed the aging-associated transcriptome reprogramming, including previously reported senescence-associated genes, Zbp1, S100a9, B2m, and Hspa1b39,40,41,42. These findings suggest a global impact of ARMH4 on cardiac aging. However, genetic ablation of Armh4 does not affect fertility or embryonic development in mice, despite of a direct regulation of IGF/FGF signaling that has been tightly associated with development. These findings suggest that the function of ARMH4 might be context-dependent at different development stages.
ARMH4 has been previously identified as a single-transmembrane protein58. Our immunofluorescence assay verified that ARMH4 was mainly localized on cell membrane (Fig. 7c). Yet, we firstly identified ARMH4 in plasma samples from young and elderly individuals using mass spectrometry22,59, suggesting that ARMH4 is also a secretory protein. Furthermore, in Western blot analysis of plasma samples for ARMH4, the ARMH4 band was not detected at the expected 85 kDa but was observed between 55 ~ 70 kDa (Supplementary Fig. 1b). This observation suggests that ARMH4 may contain a proteolytic cleavage site allowing it to be released into the circulation. Supplementation of recombinant ARMH4 protein exerted similar effects on IGF1R and FGFR1 expressions as its overexpression with adenoviral vectors (Fig. 9e, f), suggesting that the secreted form of ARMH4 might be the functional one in regulating aging-related signaling. Interestingly, both our Western blot results and previous studies showed that the actual ARMH4 band is situated between 100 ~ 130 kDa, but not at predicted 85 kDa (Figs. 1c, 5d–h)58, implicating potential post-translational modification (PTM; e.g., glycosylation) events during its molecular maturation. Surprisingly, in the co-IP assay, the molecular size of ARMH4 varied with different interacting proteins; i.e., larger than 180 kDa when co-expressed with IGF1R, but lower than 180 kDa when co-expressed with FGFR1 (Fig. 7b), suggesting different kinds of PTMs may occur on ARMH4 after its binding to different target receptors.
In the program-based aging theory, aging has been considered as a natural process within the frame of development60. A strong argumentation of this theory is IGF1 signaling, which is not only essential for development and maturation, but also acts as a potent pro-aging factor61. Certain alterations or mutations of IGF1 signaling axis cause developmental or aging-related problems, such as diabetes, but others have been reported to extend lifespan and delay age-related decline46,47,62,63,64,65,66. This paradox highlights the trade-off between growth and survival: while moderate IGF-1 activity supports maintenance and repair, chronic overstimulation may drive the pathological aging progress. Therefore, finely tuned IGF1 signaling is crucial for the balance in its biological actions. Our findings demonstrated a unique modulatory function of ARMH4 on IGF1R signal transduction activity, providing a feasible molecular tool to manipulate IGF1 axis for dealing with aging-related issues. In addition to IGF1R, ARMH4 also directly interacts with FGFR1 to sensitize its activation following stimulation by FGFs. FGFs are pleiotropic mitogens that regulate cell migration, proliferation, differentiation, and survival67,68. The role of FGFR1 signaling in aging is complex: activation of FGFR1 by a subset of FGFs has been reported to promote cell senescence69,70, whereas its activation by FGF21 or FGF23 can counter oxidative stress and apoptosis, leading to delayed aging71,72. Here we demonstrated that FGFR1 also contributed to ARMH4-mediated activation of growth signaling that triggered the aging-related protein over-synthesis and autophagy impairment, suggesting a complex regulation of endogenous FGFR1 during natural aging.
As the hub of nutrient-sensing signaling, mTOR plays a central role in the regulation of aging and age-related diseases16. Interventions that inhibit mTORC1 have been reported to delay aging12. Compared with mTORC1, the role of mTORC2 in aging scenario has received much less attention. Nevertheless, genetic ablation of the mTORC2-specific subunit, RICTOR, has been shown to attenuate the aging-related pathologies73. We found that both mTORC1 and mTORC2 positively responded to ARMH4 expression, as evidenced by both in vitro (Fig. 6c–f) and in vivo (Fig. 6a, b) data. Interestingly, mTORC1 and mTORC2 activities seemed to be differentially regulated by the IGF1R and FGFR1 signaling (Fig. 8). The diverse regulations of these distinct complexes and their corresponding impacts on downstream outcomes during aging need to be further elucidated.
Feedback circuits are common signaling strategies adopted by cells to maintain metabolic homeostasis in response to variable conditions. For instance, telomere dysfunction activates ATM kinase, which triggers p53 activation; p53 inhibits the telomere component TRF2, further promoting ATM activation, ultimately driving cellular senescence or apoptosis74. Our findings reveal that ARMH4 interacts with IGF1R/FGFR1 to enhance downstream mTORC1/2 activation, which promotes the translation activity of c-Myc, and in turn increases Igf1r/Fgfr1 transcriptional expressions, thereby establishing a positive feedback circuit with crucial significance in the aging process. c-Myc is a transcription factor regulating cell proliferation, metabolism, and differentiation75. The mTORC1/S6K1 axis promotes c-Myc translation in an eIF4B-dependent manner76. Activation of mTORC1 is required for c-Myc-driven hepatocarcinogenesis77. These observations support bi-directionally positive regulations between c-Myc and mTOR. Therefore, targeting ARMH4 might be a promising avenue to press the brake on the positive-feedback circuit continuously delivering pro-senescent effects.
Taken together, our study demonstrates that ARMH4 defines a positive-feedback growth signaling circuit by both sensitizing IGF1R/FGFR1 signaling activity and enhancing their expressions via c-Myc. Targeting ARMH4 holds translational potential to promote healthy aging and extend lifespan.
Methods
Animal care
All experiment procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of Fuwai Hospital Chinese Academy of Medical Sciences (FW-2022-0011) and performed in accordance with the guide for the care and use of laboratory animals published by National Institutes of Health, USA (8th edition). All mice were raised in a specific pathogen free environment (room temperature, 24 ± 3 °C; room humidity, 55 ± 5%) with a 12 h light/12 h dark cycle and fed normal chow. Armh4 KO mice were generated using clustered regularly interspaced short palindromic repeats (CRISPR)-cas9 technology by GemPharmatech Inc. (Nanjing, China). Guide RNAs (gRNAs) were designed ahead of and after exon 2 for cleavage under the action of CRISPR/Cas9. Mice were normally fed during the entire natural aging process.
Cell culture
Neonatal rat ventricular myocytes (NRVMs) were isolated and cultured as previously described78. Briefly, the hearts were extracted from 1-3 d Sprague-Dawley (SD) neonatal rat. After removing the atrium and the large vessels attached to the surface of the heart, ventricles were fully cut into tissue debris. The fine-minced ventricular tissues were digested in an ADS buffer (NaCl 120 mM, HEPES (pH 7.4) 20 mM, NaH2PO4 8 mM, glucose 6 mM, KCl 5 mM, MgSO4 0.8 mM) containing 0.02% type II collagenase (Sigma, #1148090) and 0.06% Pancreatin (Sigma, #T2600000) at 37 °C for 20 min/time for 7 times. Cardiomyocytes were separated from non-myocardial mesenchymal cells by density centrifugation using Percoll (GE, #17-0891-09), and cultured in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone, #SH30243.01) containing 10% (v/v) heat-inactivated fetal bovine serum (FBS; Gibco, #10099141 C) and 1% penicillin and streptomycin (Gibico, #15140163) and 1‰ BrdU (Sigma, #B5002) for 24 h. The medium was then replaced with DMEM containing 1% insulin-transferrin-selenium (ITS; Cyagen Biosciences, USA) for another 24 h before transfection or treatment. The adenovirus overexpressing Armh4 was packaged by Vigene Biosciences Co., Ltd. (Shandong, China). Plasmids were transfected into NRVMs by Nanoparticle In Vivo Transfection Reagent (Altogen, #5031) as previously described78, and siRNAs were transfected by Lipofectamine RNAiMAX (ThermoFisher, #13778150).
The primary mouse fibroblasts (MEFs) were derived from 12.5-14.5-day-old mouse embryos. After removing the embryos, the head, internal organs, and limbs were removed. The trunk was digested with 0.06% Pancreatin (Sigma, #T2600000) at 37 °C for 20 min. The digestion was terminated by adding serum, followed by centrifugation at 1000 × g for 5 min. MEFs were cultured in DMEM containing 10% (v/v) FBS, 1% penicillin and streptomycin. Cultures were maintained at 37 °C in a humidified atmosphere containing 5% CO2 and expanded in cell culture dish. HEK293 (CELLCOOK, Cat# cc4002) cells were cultured in DMEM containing 10% (v/v) FBS, 1% penicillin and streptomycin. Reagents, including puromycin (ScienCell, #0543), Picropodophyllin (MedChemExpress, #15494), Ferulic acid sodium (MedChemExpress, #N0060A), IGF-1 (MedChemExpress, #HY-P7203), FGF-2 (MedChemExpress, # HY-P7091). Plasmids and siRNAs were transfected into MEFs and HEK293 by jetPRIMER (polyplus, #101000046). The siRNA sequences were listed in Supplementary Table 1.
Plasmids
The full-length cDNA sequence of human, mouse and rat Armh4 (hArmh4, mArmh4 and rArmh4), human Igf1r and Fgfr1 (hIgf1r, hFgfr1), mouse Myc (mMyc) were obtained by the National Center for Biotechnology Information (NCBI). The pcDNA3.1-hArmh4-3FLAG, pcDNA3.1-mArmh4-3FLAG and pcDNA3.1-rArmh4-3FLAG recombinant plasmids were constructed by cloning the coding region of hArmh4, mArmh4 and rArmh4 into the NheI (NEB, #R3131) and XhoI (NEB, #R0146) sites of the pcDNA3.1-3FLAG plasmid. The pcDNA3.1-hIgf1r-HA and pcDNA3.1-hFgfr1-HA recombinant plasmids were constructed by cloning the coding region of hIgf1r and hFgfr1 into the NheI and XbaI (NEB, #R0145) sites of the pcDNA3.1-HA plasmid. pLV3-CMV-Myc(mouse)-3FLAG-Fluc-Puro was purchased from MIAOLING BIOLOGY (#P58001). Massive plasmid replication was performed by transforming the plasmids into E.coli DH-5α (Vazyme, #C502). rArmh4 was encompassed into the adenovirus vector, the adenovirus was packaged by Vigene Biosciences Co., Ltd (Shandong, China).
Expression and purification of rat-Armh4 (rArmh4)-GST fusion protein
The recombinant plasmid pGEX-6p-1-rArmh4-GST was constructed using standard molecular biology techniques and transformed into E.coli T7 (Vazyme, #C504) expression cells. Small-scale test expressions were performed to optimize induction conditions, screening IPTG concentration (0.1-1.0 mM), temperature (16 °C, 25 °C, 37 °C), and induction time (2-6 h). Following confirmation of optimal expression conditions, a 200 mL LB culture was inoculated at a 1:50 ratio with a fresh overnight pre-culture and grown at room temperature with shaking until mid-log phase (OD600 ≈ 0.6). Protein expression was induced by adding IPTG to a final concentration of 0.1 mM, followed by incubation at 37 °C with shaking for 4 h. Cells were harvested by centrifugation at 4500 × g for 15 min at 4 °C. The pellet was resuspended in 15 mL of ice-cold PBS and lysed on ice using a probe sonicator (70% amplitude, 10 seconds pulse ON / 10 seconds pulse OFF cycles) for 30 min, or until the lysate clarified. The lysate was clarified by centrifugation at 12,000 × g for 30 min at 4 °C. The supernatant was incubated with Glutathione-Sepharose beads overnight at 4 °C on a rotary shaker. The beads were collected by centrifugation at 5000 × g for 5 min and washed three times with PBS. Bound recombinant protein was eluted by competitive elution using GST Elution Buffer (125 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10 mM reduced glutathione) with incubation on a rotary shaker for 30 min at room temperature. The eluate was recovered following centrifugation at 5000 × g for 3 min. The purified rArmh4-GST fusion protein concentration was determined using the BCA assay.
Histology
After sacrifice of the mice, the liver, spleen, and kidneys were harvested and dipped in saline solution. A portion of the tissues was rapidly frozen in liquid nitrogen for preservation, and another portion, along with the hearts, was fixed in 4% paraformaldehyde for 24 h at room temperature. Subsequently, these organs were embedded in paraffin using standard histological procedures and sectioned at 5 μm. The sections were subjected to hematoxylin-eosin (H&E) staining for histopathology, Masson trichrome staining for fibrosis, β-gal staining for aging or wheat germ agglutinin (WGA; Invitrogen, #W11261) staining for measurement of the cardiomyocyte cross-sectional area. The stained sections were visualized by a confocal microscope (Olympus FV3000, Tokyo, Japan) and analyzed by Image-Pro Plus 6.0 software (Media Cybernetics, Bethesda, USA).
Senescence-associated β-galactosidase (SA-β-Gal) staining
Treated AC16 (CELLCOOK, Cat# cc4030) and HUVEC (CELLCOOK, Cat#CC4004) cells were washed twice with PBS and fixed in 4% paraformaldehyde for 15 min at room temperature. Fixed cells were incubated with SA-β-Gal staining solution (Vazyme, #AB101-01) (940 μL Buffer A, 10 μL Buffer B, and 50 μL X-gal) for 12 h at 37 °C under dark conditions. Images were examined and captured using a confocal microscope. The percentage of SA-β-Gal-positive cells was quantified from random fields using ImageJ software.
Female mouse vaginal cell smear
A fine cotton swab soaked in physiological saline was gently inserted into the vagina of 5-week-old, 6-week-old, and 7-week-old mice at 0.5 cm, and then slowly rotated and removed. The secretions on the cotton swab were evenly smeared on the slide. After the slide was naturally dried, it was placed in 10% paraformaldehyde and fixed for 10 min. Then, according to the manufacturer’s instructions (Servicebio, #G1005), the slide was H&E stained using a H&E staining kit. The image was then captured under 20-fold magnification of H&E staining using an Olympus microscope, and three random fields were taken for each slide. The time of sexual maturity of female mice was determined by identifying the time when the vaginal cells changed with age.
Male sperm smearing and counting
Eight-week-old male mice were used to collect cauda epididymis, which was rinsed with PBS and minced in M2 medium (Sigma, #M7167). The samples were incubated at 37 °C in a 5% CO2 incubator for 30 min to allow sufficient sperm release into the medium. Sperm count was determined using 0.15% eosin staining and a hemocytometer-Countess 3 (invitrogen). Some of the sperm were stained with eosin using an H&E staining kit. The stained sections were visualized under a confocal microscope.
Glucose tolerance test (GTT)
Mice of the same age and sex (minimum n = 5 per group) were fasted for 16 h prior to the test, beginning at 5:00 PM the previous day (with water provided ad libitum). At 9:00 AM, body weight was measured and baseline (0 min) blood glucose levels were determined from tail vein blood. Following a 30-min acclimatization period, a glucose solution (2 g/kg body weight) was administered via intraperitoneal injection. Injections for all mice were completed within 1 min of each other to ensure consistency. Blood glucose levels were subsequently measured at 15, 30, 60, 90, and 120 min post-injection. The glucose excursion curve was plotted (blood glucose vs. time) and results were analyzed using GraphPad Prism software. Area under the curve (AUC) was calculated to quantify glucose tolerance.
Fasting-refeeding metabolic assessment
Age- and sex-matched mice (n ≥ 5 per group) were subjected to a 24 h fasting protocol initiated at 09:00 on Day 0. Prior to feed withdrawal, baseline measurements including random blood glucose, body weight, and blood collection (via tail vein puncture into tubes) were recorded. Mice were fasted with water available ad libitum until 09:00 on next day, at which point post-fasting blood glucose and body weight were measured followed by repeat blood collection. All animals were then provided pre-weighed chow for a standardized 6-h refeeding period. At 15:00 (6 h post-refeeding), final measurements of blood glucose, body weight, and food intake (determined by weighing remaining chow) were obtained with terminal blood collection. Blood samples were centrifuged at 3000 × g for 15 min at 4 °C to isolate plasma. Plasma insulin concentrations at baseline, 24 h fast, and 6 h refeeding timepoints were quantified using a commercial Insulin ELISA Kit (Elabscience, #E-EL-M1382) per manufacturer’s protocol. Metabolic parameters including: blood glucose (24 h fast vs. baseline), body weight (24 h fast vs. baseline), blood glucose (6 h refeed vs. 24 h fast), and body weight (6 h refeed vs. 24 h fast) were calculated. Data analysis and visualization were performed using GraphPad Prism 8.0.1
Frailty index (FI) assessment
Frailty in mice was assessed using a clinical frailty index (FI), which quantifies the accumulation of health deficits. A maximum of 31 clinical parameters were evaluated, covering appearance, musculoskeletal integrity, sensory function, gastrointestinal and urogenital systems, respiratory condition, and general physical indicators such as body weight and temperature. Each parameter was scored as 0 (normal), 0.5 (mild deficit), or 1 (severe deficit). The FI was calculated as the total score divided by the number of parameters assessed, resulting in a continuous value ranging from 0 to 1. Higher FI values indicate greater frailty. All assessments were performed using non-invasive observation and gentle handling to minimize stress.
Treadmill exhaustion test
Age- and sex-matched mice (minimum n = 5 per group) underwent a 5-day acclimatization protocol prior to formal testing, consisting of daily running sessions at a constant speed of 10 m/min on a 10% incline treadmill with electrical stimulation (1.22 mA, 10-second duration upon contact with the stimulus grid79. On the test day, mice were subjected to an initial 5 min phase at 10 m/min with 10% incline, followed by a progressive phase where speed increased incrementally by 2 m/min while maintaining the incline. The test was terminated when mice started maintaining contact with the stimulus grid for > 10 consecutive seconds despite encouragement, or at 30 min if exhaustion criteria were not met. Time to exhaustion (s), total distance run (m), and number of electrical stimuli received were recorded. Data were analyzed using GraphPad Prism 8.0.1.
Rotarod fatigue test
Age- and sex-matched mice (minimum n = 5 per group) underwent acclimatization training on the rotarod apparatus for five consecutive days prior to formal testing. Training sessions consisted of 2-3 rounds of the following protocol per day: 1 min at 4 rpm, 3 min at 10 rpm, and 3 min at 30 rpm, with 15 min rest intervals between rounds. On the test day, mice were placed on the rotarod starting at an initial speed of 4 rpm. The rod speed was then accelerated linearly at a rate of 20 rpm per minute up to a maximum speed of 40 rpm. The latency to fall from the rod was recorded for each mouse. Data were analyzed and graphs were generated using GraphPad Prism software 8.0.1.
Mouse echocardiography
Animal Preparation: Mice were anesthetized using isoflurane inhalation (typically 1.5–2% for maintenance) delivered via a nose cone. Chest hair was carefully removed using depilatory cream or clippers. The anesthetized mouse was placed in the supine position on a warming pad (37 °C) to maintain body temperature. Ultrasound transmission gel was applied to the chest. 1) Parasternal Short-Axis (SAX) View at the Papillary Muscle Level: A high-frequency ultrasound probe (VINNO) was positioned on the left precordium. The standard left ventricular short-axis view at the papillary muscle level was obtained. Measurements were performed on at least three consecutive cardiac cycles and averaged for the following parameters: Left ventricular end-systolic and end-diastolic interventricular septum thickness (IVSs, IVSd). Left ventricular end-systolic and end-diastolic posterior wall thickness (LVPWs, LVPWd). Left ventricular end-systolic and end-diastolic internal diameter (LVIDs, LVIDd). Left ventricular end-systolic and end-diastolic volumes (LVESV, LVEDV) were calculated using Simpson’s method of disks (biplane or single-plane as applicable). Left ventricular ejection fraction (LVEF) was calculated as: [(LVEDV - LVESV) / LVEDV] × 100%. Left ventricular fractional shortening (LVFS) was calculated as: [(LVIDd - LVIDs) / LVIDd] × 100%. 2) Apical Four-Chamber (A4C) View: The probe was repositioned over the cardiac apex and angled approximately 30 degrees towards the right shoulder to obtain the standard apical four-chamber view. Pulsed-wave Doppler was used with the sample volume placed at the tips of the mitral valve leaflets. The ratio of early (E) to late (A) diastolic mitral inflow velocities (E/A ratio) was measured over at least three consecutive cardiac cycles and averaged.
Enzyme-linked immunosorbent assays (ELISA)
Plasma levels of follicle-stimulating hormone (FSH), luteinizing hormone (LH), testosterone, insulin (INS) and insulin-like growth factor 1 (IGF-1), were measured with a quantitative enzyme-linked immunosorbent assay (Quantikine ELISA) kit, respectively, according to the manufacturer’s instructions. The kits for mouse FSH (#M0511), mouse LH (#M3053), mouse testosterone (#M0003), mouse insulin (#M1382), mouse IGF-1 (#M3006) were purchased from Elabscience.
Dual-luciferase reporter assay
The 5’ untranslated region (5’UTR) of the Myc gene was cloned upstream of the firefly luciferase (Fluc) coding sequence in the pGL3-Fluc reporter vector. Concurrently, an Armh4 overexpression construct was generated by subcloning full-length Armh4 cDNA into the pcDNA3.1 vector with a C-terminal 3FLAG tag. For transfection assays, HEK293 cells were co-transfected with: (1) the Renilla luciferase (Rluc) control vector (pRL-TK), (2) the Myc-5’UTR-pGL3-Fluc reporter construct, and (3) either pcDNA3.1-Armh4-3FLAG or empty vector control. At 48 h post-transfection, dual-luciferase activity was measured using the Beyotime Dual-Luciferase Reporter Assay Kit (#RG029S) according to manufacturer’s protocols. Firefly luciferase signals were normalized to Renilla luciferase activity for each sample. The normalized Fluc/Rluc ratio in Armh4-expressing cells was compared to vector controls to assess ARMH4-mediated translational regulation of c-Myc expression.
Western blot
Total proteins were extracted from cells or heart tissues with Radio Immunoprecipitation Assay (RIPA) lysis buffer (Beyotime, #P0013B). Cell lysate products are centrifuged at 4 °C for 15 min at 12,000 × g. Protein concentration is determined using the BCA protein assay kit, and equal amounts of protein extracts are subjected to gel electrophoresis (SDS-PAGE) for protein separation, followed by transfer onto a PVDF membrane (MERCK, #IPVH00010). The PVDF membrane is blocked with 5% skim milk at room temperature for 1 h, then incubated with specific primary antibody overnight at 4 °C. The next day, after washing three times with TBST, the membrane is incubated with secondary antibody at room temperature for 1 h, followed by another three washes with TBST, and visualized by ECL Western Blotting Detection Kit (Thermo Fisher, #32109). Specific primary antibodies included: anti-FGFR1 (1:1000, CST, #9740), anti-IGF1R (1:1000, CST, #9750), anti-GAPDH (1:1000, CST, #2118), anti-FLAG (1:1000, CST, #8146), anti-HA (1:1000, CST, #3724), anti-Puromycin (1:5000, Merck-Millipore, #MABE343), anti-p-mTOR (1:1000, CST, #5536), anti-mTOR (1:1000, CST, #2983), anti-p-p70S6K (1:1000, CST, #9234), anti-p70S6K (1:1000, CST, #2708), anti-p-Akt (1:1000, CST, #4060), anti-Akt (1:1000, CST, #9272), anti-p-ERK (1:1000, CST, #4370), anti-ERK (1:1000, CST, #9102), anti-p-4E-BP1 (1:1000, CST, #2855), anti-4E-BP1 (1:1000, CST, #9644), anti-p62 (1:1000, CST, #8025), anti-LC3B (1:1000, CST, #3868), anti-c-Myc (1:1000, Abcam, ab32072), and anti-ARMH4 (1:500, ATLAS, #hpa001580).
co-immunoprecipitation (co-IP)
HEK293 cells transfected with corresponding plasmids were homogenized using the precooled lysis buffer (50 mM Tris at pH = 7.5, 150 mM NaCl, 2 mM EDTA at pH = 8.0, 1% NP-40, 5% Glycerol), thoroughly mixed by pipetting, and lysed at 4 °C with rotation for 30 min. The lysates were then centrifuged at 4 °C at 12,000 × g for 15 min. A small portion of the cell lysate was collected as the input, and the remaining lysate was incubated with protein A beads (Thermo fisher, #20333) for 30 min for pre-clearing. The pre-cleared lysate was then incubated overnight at 4 °C with anti-FLAG M2 Affinity Gel (Sigma, #A2220) on a rotator. On the second day, the FLAG-beads were washed with the prepared wash buffer (50 mM Tris at pH = 7.5, 150 mM NaCl, 2 mM EDTA at pH = 8.0, 0.1% NP-40), and the protein was collected with 50 μL of wash buffer. After boiling at 100 °C for 10 min, the samples were detected using Western blot.
Immunoprecipitation (IP)
Heart tissues (10 mg) from WT mice were homogenized in 300 μL of lysis buffer using a tissue grinder, followed by incubation at 4 °C with rotation for 30 min. The lysates were then centrifuged at 4 °C at 12,000 × g for 15 min. A small portion of the supernatant was collected as the input, and the remaining lysate was incubated with protein A beads for 30 min for pre-clearing. The pre-cleared lysate was then incubated overnight at 4 °C on a rotator with anti-ARMH4 (1:50, ATLAS, #HPA001580), anti-IGF1R (1:200, CST, #9750), or anti-FGFR1 (1:200, CST, #9750) antibody. On the following day, protein A beads were added and incubated at 4 °C with rotation for 4 h. The beads were then washed with washing buffer, and bound proteins were eluted with 50 μL of washing buffer. After boiling at 100 °C for 10 min, the samples were subjected to Western blot analysis.
Immunofluorescence
After transfection, HEK293, primary MEF cells, mouse heart sections were fixed in 4% paraformaldehyde at room temperature for 10 min, rinsed with PBS three times for 3 min each, then incubated in permeabilization buffer (0.2% Triton X-100) for 10 min, and rinsed with PBS three times for 3 min each. After blocking with a 5% BSA solution at room temperature for 1 h, the cells were incubated with anti-FLAG (1:200, CST, #8146) or anti-HA antibodies (1:200, CST, #3724) at 4 °C overnight. On the following day, the cells were washed with PBST (0.1% Tween-20 in PBS) three times, and then incubated with Alexa Fluor 488 antibody (1:1000, CST, #4408) and Alexa Fluor 594 antibody (1:1000, CST, #8889) at room temperature for 1 h. Afterward, the cells were rinsed with PBST three times for 3 min each. Finally, the cell nuclei were stained with DAPI (Thermo, catalog number, #62248) for 10 min, and washed with PBST three times for 5 min each. Images were observed and captured under a confocal microscope.
Puromycin incorporation assay
Puromycin mimics the aminoacyl-end structure of the amino-acid-tRNA, and can be readily incorporated into nascent polypeptides, thus reflecting a snapshot of ongoing translation events78. Before harvesting, NRVMs were treated with puromycin (1 μM) for 30 min, primary MEFs were treated with puromycin (1 μM) for 15 min, and quickly collected to extract total proteins, followed by immunoblotting with the anti-Puromycin antibody. Protein synthesis rate was calculated by the histograms of Puromycin signal against Coomassie blue staining.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Total RNAs from heart, liver, spleen, and kidney tissues were extracted using the TRIzol reagent (Takara, Kusatsu, Japan) follow the manufacturer’s instructions. The RNAs were quantified using NanoDrop (Thermo Fisher Scientific). The cDNAs were synthesized from 1 μg RNA using a RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher, #K1622). qRT-PCR was performed using ChamQ SYBR qPCR Master Mix (Vazyme, #Q311) on QuantStudio 7 Pro Real-Time PCR Detection System (Thermo Fisher Scientific). The primer sequences used in this study were listed in Supplementary Table 2.
RNA sequencing (RNA-seq) analysis
The heart tissues were harvested from WT-Young, KO-Young, WT-Aged, and KO-Aged groups to extract the total RNA using the TRIzol reagent. RNA-seq analysis was performed by the BGISEQ platform (Beijing Genomics Institution, BGI) as previously described23. Four independent biological replicates were sequenced for each group. The differential expression genes (DEGs) between groups were calculated using the linear models for microarray data (limma) package in R with a threshold of |Fold change | > 1.5 and adjusted P value < 0.05. Gene ontology (GO) analysis of DEGs was performed by Metascape online database with default parameters. Gene set enrichment analysis (GSEA) was used to screen significantly enriched signaling pathways. Raw data have been uploaded to the Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo) database with the access number GSE277042.
Statistics & reproducibility
Statistical analyses were performed using GraphPad Prism 8.0.1 Software. All experimental data were presented as mean +/− SEM of at least three independent experiments, unless denoted elsewhere. Statistical significance for multiple comparisons was determined by one-way ANOVA or two-way ANOVA followed by Turkey’s test. Bonferroni adjustment was used for post hoc analysis. Student’s t test was used for comparisons between two groups. P < 0.05 was considered statistically significant. Schematics figures (Figs. 6a, b; 9a, b; 10e, h; and Supplementary fig. 9) were created with BioGDP55. G-power software was used to predict the effect size for animal experiments. No data were excluded from the analyses. Samples were randomly allocated into experimental groups using generated randomization. The investigations were blinded to group allocation during data collection and analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data have been uploaded to the GEO database (accession codes: GSE277042). All other raw data are included in the source data files submitted with the manuscript and are fully accessible. Source data are provided with this paper.
Code availability
Codes used for the RNA-seq analysis are publicly available from R.
References
Beard, J. R. et al. The World report on ageing and health: a policy framework for healthy ageing. Lancet 387, 2145–2154 (2016).
Moqri, M. et al. Biomarkers of aging for the identification and evaluation of longevity interventions. Cell 186, 3758–3775 (2023).
Moqri, M. et al. Validation of biomarkers of aging. Nat. Med 30, 360–372 (2024).
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. Hallmarks of aging: an expanding universe. Cell 186, 243–278 (2023).
Singh, P. et al. Taurine deficiency as a driver of aging. Science 380, eabn9257 (2023).
Levine, B. & Kroemer, G. Biological functions of autophagy genes: a disease perspective. Cell 176, 11–42 (2019).
Ulgherait, M. et al. Circadian autophagy drives iTRF-mediated longevity. Nature 598, 353–358 (2021).
Zhou, B. et al. Mitochondrial permeability uncouples elevated autophagy and lifespan extension. Cell 177, 299–314 e216 (2019).
De Luca, A., Maiello, M. R., D’Alessio, A., Pergameno, M. & Normanno, N. The RAS/RAF/MEK/ERK and the PI3K/AKT signalling pathways: role in cancer pathogenesis and implications for therapeutic approaches. Expert Opin. Ther. Targets 16, S17–S27 (2012).
Lavoie, H., Gagnon, J. & Therrien, M. ERK signalling: a master regulator of cell behaviour, life and fate. Nat. Rev. Mol. Cell Biol. 21, 607–632 (2020).
Widjaja, A. A. et al. Inhibition of IL-11 signalling extends mammalian healthspan and lifespan. Nature 632, 157–165 (2024).
Mannick, J. B. & Lamming, D. W. Targeting the biology of aging with mTOR inhibitors. Nat. Aging 3, 642–660 (2023).
Papadopoli, D. et al. mTOR as a central regulator of lifespan and aging. F1000Res 8, F1000 (2019).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Igarashi, M. & Guarente, L. mTORC1 and SIRT1 Cooperate to foster expansion of gut adult stem cells during calorie restriction. Cell 166, 436–450 (2016).
Johnson, S. C., Rabinovitch, P. S. & Kaeberlein, M. mTOR is a key modulator of ageing and age-related disease. Nature 493, 338–345 (2013).
Ortega-Molina, A. et al. A mild increase in nutrient signaling to mTORC1 in mice leads to parenchymal damage, myeloid inflammation and shortened lifespan. Nat. Aging 4, 1102–1120 (2024).
Barger, J. L. et al. A low dose of dietary resveratrol partially mimics caloric restriction and retards aging parameters in mice. PLoS One 3, e2264 (2008).
Stensvold, D. et al. Effect of exercise training for five years on all cause mortality in older adults-the Generation 100 study: randomised controlled trial. BMJ 371, m3485 (2020).
Jeon, O. H. et al. Systemic induction of senescence in young mice after single heterochronic blood exchange. Nat. Metab. 4, 995–1006 (2022).
Loffredo, F. S. et al. Growth differentiation factor 11 is a circulating factor that reverses age-related cardiac hypertrophy. Cell 153, 828–839 (2013).
Li, L. et al. Tai Chi exercise improves age-associated decline in cerebrovascular function: a cross-sectional study. BMC Geriatr. 21, 293 (2021).
Lv, J. et al. Targeting FABP4 in elderly mice rejuvenates liver metabolism and ameliorates aging-associated metabolic disorders. Metabolism 142, 155528 (2023).
Kurosu, H. et al. Suppression of aging in mice by the hormone Klotho. Science 309, 1829–1833 (2005).
Conway, S. J., McConnell, R., Simmons, O. & Snider, P. L. Armadillo-like helical domain containing-4 is dynamically expressed in both the first and second heart fields. Gene Expr. Patterns 34, 119077 (2019).
Shuster, S. A. et al. In situ cell-type-specific cell-surface proteomic profiling in mice. Neuron 110, 3882–3896 e3889 (2022).
Yuan, R. et al. Genetic differences and longevity-related phenotypes influence lifespan and lifespan variation in a sex-specific manner in mice. Aging Cell 19, e13263 (2020).
Yuan, R. et al. Genetic coregulation of age of female sexual maturation and lifespan through circulating IGF1 among inbred mouse strains. Proc. Natl. Acad. Sci. USA 109, 8224–8229 (2012).
Quignon, C. Collection and analysis of vaginal smears to assess reproductive stage in mice. Curr. Protoc. 3, e887 (2023).
Myers, M., Britt, K. L., Wreford, N. G., Ebling, F. J. & Kerr, J. B. Methods for quantifying follicular numbers within the mouse ovary. Reproduction 127, 569–580 (2004).
Gotz, V. et al. Ovulation is triggered by a cyclical modulation of gonadotropes into a hyperexcitable state. Cell Rep. 42, 112543 (2023).
Islam, M. T. et al. Senolytic drugs, dasatinib and quercetin, attenuate adipose tissue inflammation, and ameliorate metabolic function in old age. Aging Cell 22, e13767 (2023).
Du, K. et al. Aging promotes metabolic dysfunction-associated steatotic liver disease by inducing ferroptotic stress. Nat. Aging 4, 949–968 (2024).
Yang, S. et al. A single-nucleus transcriptomic atlas of primate liver aging uncovers the pro-senescence role of SREBP2 in hepatocytes. Protein Cell 15, 98–120 (2024).
Lynes, M. D. et al. The cold-induced lipokine 12,13-diHOME promotes fatty acid transport into brown adipose tissue. Nat. Med 23, 631–637 (2017).
You, M. et al. Selenoprotein K contributes to CD36 subcellular trafficking in hepatocytes by accelerating nascent COPII vesicle formation and aggravates hepatic steatosis. Redox Biol. 57, 102500 (2022).
Czibik, G. et al. Dysregulated phenylalanine catabolism plays a key role in the trajectory of cardiac aging. Circulation 144, 559–574 (2021).
Moturi, S., Ghosh-Choudhary, S. K. & Finkel, T. Cardiovascular disease and the biology of aging. J. Mol. Cell Cardiol. 167, 109–117 (2022).
Radak, M. & Fallahi, H. Zbp1 gene: a modulator of multiple aging hallmarks as potential therapeutic target for age-related diseases. Biogerontology 24, 831–844 (2023).
Wang, C. et al. S100A9-driven amyloid-neuroinflammatory cascade in traumatic brain injury as a precursor state for alzheimer’s disease. Sci. Rep. 8, 12836 (2018).
Smith, L. K. et al. beta2-microglobulin is a systemic pro-aging factor that impairs cognitive function and neurogenesis. Nat. Med 21, 932–937 (2015).
Soti, C. & Csermely, P. Molecular chaperones and the aging process. Biogerontology 1, 225–233 (2000).
Davidson, M. M. et al. Novel cell lines derived from adult human ventricular cardiomyocytes. J. Mol. Cell Cardiol. 39, 133–147 (2005).
Li, C. et al. IRE1alpha Mediates the Hypertrophic Growth of Cardiomyocytes Through Facilitating the Formation of Initiation Complex to Promote the Translation of TOP-Motif Transcripts. Circulation 150, 1010–1029 (2024).
Kaizuka, T. et al. An autophagic flux probe that releases an internal control. Mol. Cell 64, 835–849 (2016).
Dorman, J. B., Albinder, B., Shroyer, T. & Kenyon, C. The age-1 and daf-2 genes function in a common pathway to control the lifespan of Caenorhabditis elegans. Genetics 141, 1399–1406 (1995).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A. C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993).
Liu, H. et al. Functional crosstalk between mTORC1/p70S6K pathway and heterochromatin organization in stress-induced senescence of MSCs. Stem Cell Res Ther. 11, 279 (2020).
Wang, W. et al. Cytoplasmic endonuclease G promotes nonalcoholic fatty liver disease via mTORC2-AKT-ACLY and endoplasmic reticulum stress. Nat. Commun. 14, 6201 (2023).
Pei, F. et al. Sensory nerve niche regulates mesenchymal stem cell homeostasis via FGF/mTOR/autophagy axis. Nat. Commun. 14, 344 (2023).
Stefani, C. et al. Growth factors, PI3K/AKT/mTOR and MAPK signaling pathways in colorectal cancer pathogenesis: where are we now? Int. J. Mol. Sci. 22, 10260 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Wang, J. et al. TFTF: an R-based integrative tool for decoding human transcription factor-target interactions. Biomolecules 14, 749 (2024).
Ma, J. et al. CD226 maintains regulatory T cell phenotype stability and metabolism by the mTOR/Myc pathway under inflammatory conditions. Cell Rep. 42, 113306 (2023).
Jiang, S. et al. Generic Diagramming Platform (GDP): a comprehensive database of high-quality biomedical graphics. Nucleic Acids Res 53, D1670–D1676 (2025).
Lee, D. et al. Transmembrane Inhibitor of RICTOR/mTORC2 in Hematopoietic Progenitors. Stem Cell Rep. 3, 832–840 (2014).
Lee, D. et al. Endogenous transmembrane protein UT2 inhibits pSTAT3 and suppresses hematological malignancy. J. Clin. Invest 126, 1300–1310 (2016).
De Luca, F., Kha, M., Sward, K. & Johansson, M. E. Identification of ARMH4 and WIPF3 as human podocyte proteins with potential roles in immunomodulation and cytoskeletal dynamics. PLoS One 18, e0280270 (2023).
Tang, L. et al. Bigu-Style Fasting Affects Metabolic Health by Modulating Taurine, Glucose, and Cholesterol Homeostasis in Healthy Young Adults. J. Nutr. 151, 2175–2187 (2021).
de Magalhaes, J. P. An overview of contemporary theories of ageing. Nat. Cell Biol. 27, 1074–1082 (2025).
Johnson, S. C. Nutrient Sensing, Signaling and Ageing: The Role of IGF-1 and mTOR in Ageing and Age-Related Disease. Subcell. Biochem 90, 49–97 (2018).
Ali, A. et al. Identification of functional rare coding variants in IGF-1 gene in humans with exceptional longevity. Sci. Rep. 15, 10199 (2025).
Wang, Y. et al. Targeting IGF1-Induced Cellular Senescence to Rejuvenate Hair Follicle Aging. Aging Cell, e70053 (2025).
Junnila, R. K., List, E. O., Berryman, D. E., Murrey, J. W. & Kopchick, J. J. The GH/IGF-1 axis in ageing and longevity. Nat. Rev. Endocrinol. 9, 366–376 (2013).
Mao, K. et al. Late-life targeting of the IGF-1 receptor improves healthspan and lifespan in female mice. Nat. Commun. 9, 2394 (2018).
Abdellatif, M. et al. Fine-tuning cardiac insulin-like growth factor 1 receptor signaling to promote health and longevity. Circulation 145, 1853–1866 (2022).
Xie, Y. et al. FGF/FGFR signaling in health and disease. Signal Transduct. Target Ther. 5, 181 (2020).
Coutu, D. L. & Galipeau, J. Roles of FGF signaling in stem cell self-renewal, senescence and aging. Aging 3, 920–933 (2011).
Zlinska, V. et al. Specific inhibition of fibroblast growth factor receptor 1 signaling by a DNA aptamer. Mol. Ther. Nucleic Acids 36, 102405 (2025).
Easter, M. et al. FGF receptors mediate cellular senescence in the cystic fibrosis airway epithelium. JCI Insight 9, e174888 (2024).
Wang, T. et al. Time-restricted feeding mitigates HFD-induced sarcopenic obesity in aging mice through improving the sensitivity of FGF21. J. Nutr. Biochem 140, 109893 (2025).
Chen, G. et al. alpha-Klotho is a non-enzymatic molecular scaffold for FGF23 hormone signalling. Nature 553, 461–466 (2018).
Xu, W., Chen, H. & Xiao, H. mTORC2: a neglected player in aging regulation. J. Cell Physiol. 239, e31363 (2024).
Horikawa, I., Fujita, K. & Harris, C. C. p53 governs telomere regulation feedback too, via TRF2. Aging (Albany NY) 3, 26–32 (2011).
Dang, C. V. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol. Cell Biol. 19, 1–11 (1999).
Dong, Y., Tu, R., Liu, H. & Qing, G. Regulation of cancer cell metabolism: oncogenic MYC in the driver’s seat. Signal Transduct. Target Ther. 5, 124 (2020).
Liu, P. et al. A functional mammalian target of rapamycin complex 1 signaling is indispensable for c-Myc-driven hepatocarcinogenesis. Hepatology 66, 167–181 (2017).
Wang, Z. et al. The long noncoding RNA Chaer defines an epigenetic checkpoint in cardiac hypertrophy. Nat. Med 22, 1131–1139 (2016).
Castro, B. & Kuang, S. Evaluation of muscle performance in mice by treadmill exhaustion test and whole-limb grip strength assay. Bio Protoc 7, e2237 (2017).
Acknowledgements
This study was supported by funds from Shenzhen Medical Research Fund (B2302026, ZW), National Key R&D Program of China (2022YFA1104500, ZW), National Natural Science Foundation of China (82370392, ZW), Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2025-I2M-XHJC-010, ZW; 2023-I2M-1-003, ZW; 2023-I2M-2-003, ZW), National High Level Hospital Clinical Research Funding (2025-GSP-GG-34, ZW), and Shenzhen Science and Technology Innovation Commission (RCJC20210706091947009, ZW).
Author information
Authors and Affiliations
Contributions
Z.W., H.L., and H.T. designed and supervised the study. Y.F. and B.W. performed most of the experiments with inputs from N.G., Y.B., A.X., and X.M. Q.G., J.Li., and J.W. participated in the animal experiments. J.Lv. performed the bioinformatic analysis. YF drafted the manuscript. All authors have read and approved the publication of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hua Wang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, Y., Wang, B., Guo, Q. et al. ARMH4 accelerates aging by maintaining a positive-feedback growth signaling circuit. Nat Commun 17, 812 (2026). https://doi.org/10.1038/s41467-025-67505-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67505-x