Abstract
A central challenge in cancer research is to identify the secreted factors that sustain tumor cell survival. This is best exemplified in Hodgkin lymphoma, where malignant cells constitute a minor fraction of the tumor and rely on signals from the microenvironment for survival. Using genome-wide transcriptional profiling with spatial and single-cell resolution, we show that the neighborhood around malignant cells forms a distinct niche of 31 non-malignant cell types, enriched in helper T cells and myeloid cells, but depleted of plasma cells. Moreover, our spatial analysis nominates IL13 as a candidate survival factor. Recombinant IL13 augments malignant cell growth in vitro, and genome-wide loss-of-function screens across >1000 human cancer cell lines identify IL4R and IL13RA1, heterodimeric components of the IL13 receptor, as uniquely essential in Hodgkin lymphoma. Importantly, blocking antibodies phenocopy genetic inactivation. Our findings provide a biological rationale for testing IL13-directed therapies, which are already FDA-approved, in Hodgkin lymphoma.
Similar content being viewed by others
Introduction
Sustained growth factor signaling is a fundamental hallmark of cancer1,2. Most prior studies have focused on the somatic cancer genome as a source of insight into the mechanisms explaining such aberrant signaling, as similarly comprehensive analyses of the tumor microenvironment have been technically infeasible. Yet not all cancers have somatic mutations that readily explain tumor survival, and the fact that human tumors often contain high proportions of non-malignant cells (fibroblasts, lymphoid cells, and myeloid cells) suggests that such microenvironmental cells may contribute to tumor cell survival. Consistent with this hypothesis, establishing cancer cell lines is often difficult, if not impossible, presumably because the tumor cells cannot survive when deprived of microenvironment-derived survival factors.
Survival signals are particularly important in B-cell lymphomas because normal B cells are highly dependent upon signals transduced through cytokine and antigen receptors within lymph node follicles3. Malignant B cells can overcome the limiting availability of these growth signals through somatic mutations in the B cell receptor or its downstream effectors4,5, and therapeutics targeting this pathway have proven effective in clinical trials of diffuse large B-cell lymphoma (DLBCL)6,7,8.
Whereas DLBCL typically grows as monotonous sheets of malignant B cells with a relatively minor population of non-malignant immune cells, other B-cell lymphomas are composed of a mixture of malignant B cells and abundant immune cells9,10,11,12, suggesting that a cell non-autonomous survival mechanism might be at play. The most extreme example of a microenvironment-associated cancer is classic Hodgkin lymphoma (cHL), where the rare malignant Hodgkin Reed Sternberg (HRS) cells that comprise only 1-5% of the tumor’s cellularity are embedded in an extensive inflammatory infiltrate13. Several lines of evidence suggest the importance of the Hodgkin lymphoma microenvironment for HRS cell survival (as opposed to simply being a reactive infiltrate). First, HRS cells are notoriously difficult to propagate in vitro or in immunodeficient mice, with the success rate in establishing HRS cell lines being <1%14,15,16. Second, when cHL tumors spread to non-lymphoid organs, the HRS cell microenvironment is reconstituted at those distant sites10,11. Third, somatic activating mutations that lead to constitutive activation of oncogenes are relatively uncommon in cHL17,18,19,20, suggesting that growth and survival signals may be derived from the HRS cell niche.
Importantly, HRS cells have been shown to have immune evasion mechanisms, including loss of function mutations in MHC class I and increased expression of PD-L1 through copy number gains21,22. Recent immunophenotyping studies using candidate markers have demonstrated that distinct subsets of PD-L1+ macrophages and CD4 + T cells (including regulatory CTLA-4+ and LAG3+ subsets) surround HRS cells23,24,25. Single-cell transcriptomic studies of cHL have been reported25,26,27, but those studies lacked the ability to capture certain myeloid cells (which can make up > 50% of tumor cellularity28) and stromal cells, and they lacked spatial resolution. Therefore, an unbiased, comprehensive analysis of the cHL microenvironment has been desperately needed by the field.
In this study, we use spatially resolved expression profiling to systematically define the molecular and cellular architecture in the intact cHL tumor microenvironment. We identify immune evasive and trophic signaling mechanisms and functionally validate a critical role for IL13 in promoting HRS cell survival, suggesting that targeting IL13 signaling may have therapeutic benefit for patients with Hodgkin lymphoma.
Results
Transcriptional programs and topology of HRS cells
We first generated a comprehensive single-nucleus and spatially resolved RNA sequencing dataset (Fig. 1a, supplementary data 1) using samples from newly diagnosed cHL patients (n = 12), including EBV-positive and EBV-negative cases, as well as from control reactive lymphoid tissues (RLN, n = 7). Histopathologic analysis, single-nucleus RNA sequencing (snRNAseq), and spatial profiling using slide-seqV2 were performed on serial sections from each specimen (≥2 replicates for each sample and each technology platform). Following quality control (see Methods, Supplementary Fig. 1a–c, 2a, Supplementary Data 2, 3), the resulting dataset yielded 1.9 million spatially resolved RNA profiles and 308,403 single-nucleus RNA profiles after quality control (Fig. 1b, c).
a Schematic of experimental design and analyses. Created in BioRender. Shanmugam, V. (2025) https://BioRender.com/nkuwxyx. b an overview of the single-nucleus RNA sequencing (snRNAseq) dataset. Uniform manifold approximation and projection (UMAP) embedding of snRNAseq profiles colored by cell type annotation (n = 308,403 nuclei from 15 subjects). c representative slide-seqV2 array of a Hodgkin lymphoma tumor with hematoxylin and eosin (H&E) stained section (top) and spatial plot of cell types colored by cell type (bottom), same color scheme as b. A total of 62 slideseqV2 arrays sampling different regions of samples from 19 subjects showed expected patterns of cell type distribution based on morphology. d identification of HRS cells by copy number variant (CNV) inference using inferCNV and EBV-specific read mapping. UMAP embedding showing CNV burden (quantified by CNV score) across cell types (n = 308,403 nuclei from 15 subjects). The top right inset shows the UMAP embedding of HRS cells identified by high CNV burden colored by EBV status determined by in situ hybridization (positive or negative). The lower right inset shows the same UMAP embedding with the expression levels of EBV-specific transcripts (n = 6720 nuclei from 10 patient samples). e normalized and scaled gene expression arranged by chromosome number showing CNVs in EBV- and EBV+ HRS cells (grouped by donor), relative to non-tumor cells (top) that show no significant evidence of CNVs. f Left: heatmap showing shared gene expression programs (normalized and scaled expression) in tumor cells across multiple donors identified by non-negative matrix factorization, grouped by EBV status and donors. Right: enrichment analysis results of curated gene sets (columns) in each program (rows). Pathway enrichment analysis was performed using a one-sided hypergeometric test and P values were adjusted for multiple comparisons using the Benjamini-Hochberg method. g spatial contour plot showing the clustered topology of HRS cells in tissue (representative array from donor H12-200). This analysis was repeated in 20 arrays with sufficient numbers of tumors cells and the majority of samples showed similar findings (Supplementary Data 4). For c and g, scale bars, 500 μm.
We assigned cell types based on the snRNAseq profiles using unbiased classifiers followed by manual curation (see Methods). This analysis captured the diversity of cell types expected based on the histopathology of cHL, including myeloid, lymphoid, and stromal cell types (Fig. 1b, c). Given their importance and low abundance, HRS cells were identified using multiple orthogonal strategies. First, we identified putative HRS cells using the well-established marker genes TNFRSF8 (CD30), CD274 (PD-L1), PAX5, and IRF4. Next, we performed copy number variant inference to determine if this putative HRS cell population harbored shared copy number variation seen in clonal tumor cells but not normal cells. Strikingly, the putative HRS cell population showed strong evidence of copy number variation, whereas the other cell types did not, indicating that we had appropriately identified HRS cells (Fig. 1d, e). We observed substantial inter-patient heterogeneity in HRS cell gene expression and copy number profiles (Fig. 1e, f), as shown in previous studies29,30,31,32. EBV-positive HRS cells (as determined by in situ hybridization using EBV transcript-specific probes) were associated with higher levels of EBV transcript detection by RNAseq, supporting the high quality of the dataset (Fig. 1d). Notably, mapping RNA reads from EBV-negative HRS cells to pathogen reference databases did not reveal a signal suggestive of other pathogens playing a role in cHL (see Methods). Intriguingly, EBV-negative HRS cells tended to express expression programs typically seen in neuroglial cells (Fig. 1f, Supplementary Fig. 3a, b). This finding was validated using previously reported gene expression datasets of microdissected primary HRS cells33,34 and cell lines35 (Supplementary Fig. 3c–k) and is consistent with observations in a recently published single-cell RNAseq dataset of HRS cells29. The mechanism explaining this distinct transcriptional program requires further investigation.
Using the spatial profiling data, we next performed probabilistic cell-type mapping using the RCTD method36 to define the topology of HRS cells in tissues, validating the results by spatial plotting of marker genes, copy number variant inference, and EBV read mapping (Supplementary Fig. 3l-o). Strikingly, in most tumors examined, the HRS cells showed a clustered spatial distribution (Fig. 1g, Supplementary Fig. 3p, Supplementary Data 4). This co-localization could reflect evidence of positive selection for paracrine HRS-HRS cell interactions.
Defining the immune microenvironment in cHL
We next focused on the immune and stromal cells, which comprise the vast majority of cells within cHL tumors. We identified the presence of 23 distinct immune cell types (Fig. 2a, Supplementary Figs. 4a, b, 5a, b, e, f, 6a, b) and used compositional analysis37 to compare their abundance in cHL to that observed in reactive lymphoid tissues. We observed a striking increase in the abundance of activated and differentiated T cell subsets, including regulatory T cells, activated/proliferating CD4+ T cells, exhausted CD8 + T cells, and a concomitant decrease in less differentiated naive T cells. Moreover, monocytes/macrophages were significantly expanded, particularly in EBV+ cHL (Fig. 2b, Supplementary Figs. 4c, 5c, d, h, i; 6c, d).
a UMAP embedding of immune cells colored by cell type annotation (n = 277,911 nuclei from 15 subjects). b compositional analysis of immune cells in Hodgkin lymphoma (HL) versus reactive lymphoid tissue (RLN) samples. c unsupervised consensus tensor factorization reveals a shared HL-associated myeloid cell program. d compositional analysis of myeloid subsets across HL and RLN samples. e UMAP embedding of stromal cells colored by cell type annotation (n = 23,772 nuclei from 15 subjects). f compositional analysis of stromal cells in HL versus RLN samples. g, consensus tensor factorization reveals a distinct activated pro-fibrotic, myofibroblast-like fibroblastic reticular cell program. For all these plots (including b, d, and f), n = 15 (10 HL and 5 RLN) samples from independent subjects (biological replicates). For the boxplots (b, d, f), boxes represent the interquartile range (25th − 75th percentiles) with the median shown as a central line; whiskers extend to the most extreme data points within 1.5 × IQR of the quartiles. All data points are shown in the boxplots. Statistical significance of the compositional analysis (b, d, f) was assessed using the scCODA framework, which estimates posterior inclusion probabilities for each covariate-cell type effect and applies a Bayesian FDR threshold to determine significance. Effects with posterior inclusion probabilities exceeding the cutoff corresponding to FDR < 0.20 are marked with *. Tests are two-sided, as the model estimates both enrichment and depletion.
Beyond differences in cell type abundance, we observed substantial transcriptional changes within the immune cells. For example, cHL-associated germinal center B cells showed increased expression of granzyme B38 (GZMB; Supplementary Fig. 4d), and cHL-associated regulatory T cells showed increased expression of IFNg-stimulated genes (IL12RB2, CXCL10) and NRN (neuritin), a secreted protein that enhances the expansion and suppressive function of Treg cells39 (Supplementary Fig. 5g).
Unsupervised consensus tensor factorization analysis similarly revealed a distinct tumor-associated immunosuppressive myeloid cell program (Fig. 2c) with coordinated upregulation of inhibitory receptors (MARCO, LILRB4), metal ion transporters (SLC39A8, SLC30A4), metal-binding proteins (MT1, MT2), and IFN and IL4/IL13-stimulated genes (ALOX15, KYNU). Further annotation of the monocyte/macrophage population revealed enrichment of inflammatory monocytes40 and the recently described41 IL4I1+/IFN-primed and HES1+ macrophages (Fig. 2d, Supplementary Fig. 6e, f). The enrichment of IL4/IL13-stimulated genes in myeloid cells is particularly notable as it suggests ongoing IL4/IL13 signaling in the tumor microenvironment.
Eight distinct stromal cell types were also identified in cHL tumors (Fig. 2e, Supplementary Fig. 6a), including enrichment of fibroblastic reticular cells independent of EBV status (Fig. 2f, Supplementary Fig. 7b). These cells exhibited a pro-fibrotic program characterized by expression of FAP, POSTN, collagens, and the pro-fibrotic cytokines CCL11 and IL19 (Fig. 2g, Supplementary Fig. 7c) and likely mediate the extensive fibrosis that is a histopathologic hallmark of cHL tumors10.
Importantly, spatial mapping of the immune microenvironment (Supplementary Fig. 8a, b) revealed evidence of a distinct microenvironmental niche surrounding HRS cells. Specifically, we observed the enrichment of CD4+ T cells, follicular dendritic cells, and monocyte/macrophages near HRS cells. In contrast, fibroblasts, plasma cells, and plasmacytoid dendritic cells were depleted from HRS cell neighborhoods (Fig. 3a, b). Consistent with these observations, immune cells near HRS cells had a distinct gene expression program compared to those same cell types located further away, perhaps explained by HRS cell-derived cytokine gradients (Fig. 3c, Supplementary Fig. 8c–e, h). We confirmed the existence of an HRS cell niche by generating an independent spatial dataset consisting of 637,000 cells from 4 formalin-fixed paraffin-embedded cHL samples analyzed on the CosMx platform measuring 1000 transcripts (Fig. 3d, Supplementary Fig. 2b-d, 9a, b, supplementary data 5). These results argue in favor of the existence of a highly structured HRS cell niche as opposed to a random assortment of immune and stromal cell types (Fig. 3e).
a Mean enrichment ratios of immune cell types (colored by cell type) aggregated across the entire multi-sample, multi-replicate slideseqV2 dataset (n = 38, independent arrays sampling different tumor regions from 12 patients). Statistical significance was assessed by calculating z-scores for log₂(mean enrichment ratios) relative to zero and deriving two-sided P values from the standard normal distribution. The size of the points indicates the P-value; mean ± s.e.m. is depicted. b representative array showing spatial enrichment of CD4+ T cells and depletion of plasma cells around clustered HRS cells. The images are representative of 38 independent arrays sampling different tumor regions from 12 patients. Scale bar: 500 μm. c volcano plot showing cell-type specific differentially expressed genes in relation to HRS cell proximal/dense regions. A positive log2(fold change) (right of volcano plot) indicates genes that show higher expression in HRS cell proximal/dense regions. Fold changes and q-values are calculated using the C-SIDE framework (see Methods). q-values denote p-values (two-sided) adjusted for multiple comparisons using the Benjamini–Hochberg false discovery rate (FDR) method. Results are aggregated across the entire multi-sample, multi-replicate slide-seqV2 dataset (n = 38, independent arrays sampling different tumor regions from 12 patients). d CosMx analysis showing enrichment of IL10+ CD4+ T cells within HRS cell dense/proximal regions. This image is representative of multiple fields of view sampling 4 cHL samples. Scale bar, 100 μm. e schematic representation of the structured model where immune cells show a non-random spatial distribution around clustered tumor cells in contrast to the null hypothesis random spatial model where immune cells are randomly distributed. Created in BioRender. Shanmugam, V. (2025) https://BioRender.com/1uaubi2. f schematic overview of the approach for systematic discovery and prioritization of spatially-informed tumor-immune ligand-receptor interactions within the HRS cell niche. Created in BioRender. Shanmugam, V. (2025) https://BioRender.com/0hekpqv. g scatterplot of tumor-targeted ligand-receptor interactions based on spatial proximity (spatial association metric, see methods) and normalized ligand/receptor expression (sca-LRscore metric). Interactions that show high ligand/receptor expression and spatial proximity are called out and shown on the right as a bubble plot. h RNA fluorescence in-situ hybridization analysis of IL13 expression (yellow) in HRS cells and the microenvironment. The red TNFRSF8 (encoding CD30) probe highlights HRS cells. Scale bar; 20 µm. The data are representative of independent experiments on three different patient samples. i schematic diagram of systematically prioritized paracrine tumor-microenvironment interactions in Hodgkin lymphoma, including putative growth factors and immunosuppressive interactions.
Identification of ligand-receptor interactions in the HRS cell niche
We next sought to identify secreted or cell surface ligands expressed by microenvironmental cells in proximity to HRS cells. We further prioritized such survival factors by requiring their receptors to be expressed in HRS cells. This spatially-aware ligand-receptor analysis revealed multiple candidate immunoregulatory and growth factor interactions validated in the independent dataset and via in situ hybridization (Fig. 3f–i, Supplementary Figs. 8f, g, and 9c–n). This analysis recovered known interactions involving CD274 (PD-L1), CCL17, and CCL22 ligands, highlighting the effectiveness of the approach.
Experimental validation of candidate juxtacrine and paracrine growth factor interactions
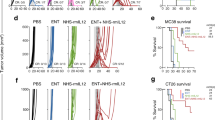
Having systematically identified candidate spatially associated ligand-receptor interactions that might support HRS cell growth or survival, we sought to test the importance of these associations experimentally. We first performed gain-of-function experiments, asking whether any of the 36 tested recombinant candidate ligands increased the abundance of the HRS cell lines L1236 and UHO1 when grown in either low serum or low-density conditions. 20/36 (55%) yielded significant increases in cell numbers, with the greatest effect seen with CD40 ligand (CD40LG) and IL13 (Fig. 4a and Supplementary Figs. 10a, b).
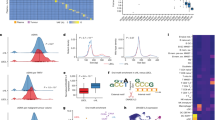
a Barplot showing the effect of recombinant factors on the relative viability (normalized to untreated control) of L1236 cells at three days under low serum conditions; mean is depicted (n = 3, independent wells from the same experiment). Bars in red denote factors that induced > 120% relative viability compared to control. b waterfall plots (ordered by increasing CRISPR gene effect score) showing genetic dependencies of 3 cHL cell lines (L1236, UHO1, L428). c, d scatterplot of IL4R and IL13RA1 (c) and IL4R and STAT6 (d) gene effect scores across the DepMap dataset. Each point (n = 1086) represents an independent cell line in the dataset. Boxplots show the median (center line), interquartile range (box), and the most extreme data points within 1.5 × the interquartile range (whiskers). Points beyond this range are plotted individually as outliers. e effect of CRISPR/Cas9-mediated knock out of IL13RA1 or IL4R on L1236 cell viability (viable cell counts). f rainplot of IL13 expression showing that the cHL cell lines (L1236, L428, and UHO1) show outlier expression of IL13. g effect of CRISPR/Cas9-mediated knock out of IL13 on L1236 cell viability (viable cell counts), with or without recombinant IL13 supplementation (rIL13). h dose-response analysis of tralokinumab, lebrikizumab, and dupilumab on L1236 cells. IC50 values are indicated on the right. The relative viability values represent the mean of two independent experimental replicates. The Cmax range of these antibodies reported in clinical trials is indicated in grey. i time course analysis of the effect of tralokinumab, lebrikizumab, dupilumab (0.5 µM each), and isotype control antibody on L1236 cell viability (n = 3, independent wells from the same experiment); mean values are depicted. j schematic representation of the proposed model of spatial organization and interactions that mediate tumor cell growth within the intact cHL tumor microenvironment. Created in BioRender. Shanmugam, V. (2025) https://BioRender.com/el2n84m. k schematic summary of genetic and pharmacologic perturbations of the IL13-IL4R-IL13RA1-STAT6 axis that induce HRS cell death. Created in BioRender. Shanmugam, V. (2025) https://BioRender.com/zwqm0x5. The data in (e, g, and i) are of three independent experiments. Source data are provided as a Source Data file.
To test whether IL13 and CD40LG signaling were essential for the survival of HRS cells, we first turned to the Cancer Dependency Map (DepMap)35, a compendium of 1086 human cancer cell lines subjected to genome-wide CRISPR/Cas9 knock-outs. Whereas CD40 was not a dependency in any of the DepMap models, knock-out of either component of the obligate heterodimeric receptor for IL13 (IL4R and IL13RA1) resulted in a profound loss of HRS cell viability. Furthermore, STAT6, the downstream transcriptional mediator of IL13 signaling, scored as an HRS cell dependency. Specifically, all three HRS cell lines in the DepMap required IL4R, IL13RA1, and STAT6 expression for survival, whereas none of the other cell lines in the database exhibited such dependency (Fig. 4b–d). This finding is unlikely to be seen by chance alone (IL13RA1, P = 4.8 x 10−9; IL4R, P = 5.1 x 10−6; STAT6, P = 4.7 x 10−9; Fisher’s Exact Test of Independence). We further validated this result using independent, individual gRNAs targeting IL4R and IL13RA1 and showed they result in loss of HRS cell viability (Fig. 4e). Analysis of these cells showed induction of cleaved caspase 3, consistent with cell death via apoptosis (Supplementary Fig. 10c). We hypothesize that these HRS cell lines underwent selection for their ability to express IL13, a cytokine that is not typically expressed at high levels by normal B cells42. Consistent with this hypothesis, the HRS cell lines showed outlier expression of IL13 (Fig. 4f), and knock-out of IL13 similarly resulted in loss of cell viability that was rescuable with recombinant IL13 (Fig. 4g, Supplementary Fig. 10d). Importantly, we observed minimal differences in IL13 and IL13 receptor expression in EBV+ and EBV- tumors, consistent with a previous study43 (Supplementary Fig. 10g).
We next asked whether the genetic dependency of IL13 signaling in HRS cells could be phenocopied pharmacologically. Specifically, we tested three monoclonal antibodies that block IL13 signaling through distinct mechanisms: dupilumab, which targets the alpha subunit of IL4 receptor (IL4R); tralokinumab, which blocks the binding of IL13 to IL13RA1 and lebrikizumab, which impairs the heterodimerization of IL4R and IL13RA1. An isotype control antibody served as a negative control. All three antibodies showed dose-responsive and time-dependent loss of viability of HRS cells and modest induction of apoptotic cell death, whereas the isotype control showed no effect (Fig. 4h, i, Supplementary Fig. 10e, f). These results are particularly notable because dupilumab, tralokinumab, and lebrikizumab are FDA-approved for use in patients with atopic dermatitis44,45,46,47. Importantly, we observed maximal growth inhibitory activity in HRS cells at concentrations that are readily achievable in patients treated at the clinically approved dose (Fig. 4h, Supplementary Fig. 10e).
Finally, we explored the possibility that IL4 upregulation might represent an adaptive resistance mechanism to IL13R blockade, given that IL13R forms an obligate heterodimer with IL4R. To test this hypothesis, we first assessed IL4 expression across 1,086 human cancer cell lines and found that the L1236 HRS cell line showed outlier levels of IL4 expression and also expresses IL4R and IL2RG (Supplementary Fig. 10h, i). However, these cells were sensitive to anti-IL13 and anti-IL4R antibodies (Fig. 4h), suggesting that autocrine IL4 secretion is not sufficient to mount resistance to anti-IL13 antibodies. Of course, it is possible that other mechanisms of resistance to anti-IL13 therapy could emerge under prolonged selective pressure.
Discussion
Anatomical pathologists have long recognized that tumors are complex admixtures of malignant cells and normal cell types. While hypothesis-driven research on the tumor microenvironment has yielded important insights, systematic genomic approaches have largely been restricted to the analysis of tumor cells. Researchers have typically gone to great lengths to purify cancer cells away from the microenvironment to enrich the signal coming from the malignant component of the tumor. More recently, single-cell transcriptomics has allowed for the analysis of all cells within a tumor, but the spatial information is lost with such approaches, thereby precluding the analysis of interactions between tumor cells and the normal cells immediately surrounding them.
Hodgkin lymphoma is one of the most enigmatic human tumors, with classic Hodgkin lymphoma (cHL) being composed of only rare tumor cells surrounded by an extensive array of infiltrating immune and stromal cells. While it is possible that these microenvironmental cells are unimportant for tumor cell survival, it has been well-documented that removing the malignant HRS cells from the microenvironment results in their failure to survive in vitro14,15,16. We, therefore, hypothesized that the microenvironment is a source of such survival and immune evasive factors.
To address this, we generated a comprehensive single-nucleus and spatial profiling dataset, composed of more than 2.2 million transcriptional profiles (all data are available here). Importantly, we believe this reference dataset includes all relevant cell types within cHL tumors, whereas standard single-cell profiling methods25,26,27,48 result in the loss of key cell types due to limitations of cell isolation methods. For example, macrophages do not survive isolation, yet they can comprise more than 50% of cells within cHL28. The quality of this reference dataset is supported by multiple lines of evidence, including the demonstration of clonality of rare HRS cells, the matching of cell types recognized by immunohistochemical analyses, and the confirmation of the recently described25,40 expansion of LAG3+ regulatory T cells and IL1B+ inflammatory monocytes that are thought to contribute to the ability of cHL to avoid immune surveillance.
Our study complements the growing body of literature supporting the presence of a distinct pro-tumorigenic cellular niche around HRS cells in cHL. Consistent with previous work, we observed the enrichment of CD4+ T cells, monocytes/macrophages, and myeloid dendritic cells and the depletion of plasmacytoid dendritic cells near HRS cells23,24,49,50,51. We additionally show that plasma cells are excluded from the immediate proximity of HRS cells, which may represent a potential mechanism of evading the humoral immune response. Previous hypothesis-driven studies23,24,25,49 have shown that specific immune checkpoint molecules are expressed in the cHL niche, including the myeloid markers (PD-L1, TIM3, IDO1) and CD4+ T cells (PD1, CTLA4, LAG3). Our genome-wide analysis recovers the recently reported52 interaction between CXCR5+ HRS cells and CXCL13-expressing macrophages and extends the observation to CXCL13-expressing CD4+ T cells, follicular dendritic cells, and fibroblasts in proximity to HRS cells. Our genome-wide study further expands our understanding of the niche by defining additional cell type-specific gene expression programs, including IL21+ and IL10+ CD4+ T cells and MMP12+ myeloid cells, that have not been previously described and require further investigation.
We experimentally validated one of the strongest findings of the cHL microenvironment, namely the nomination of IL13 as a potential survival factor for HRS cells. Gain- and loss-of-function studies supported the IL13 hypothesis. Analysis of the Cancer Dependency Map35 showed that IL4R and IL13RA1 were required for the survival of only three cell lines in the DepMap, and these are the only three cHL cell lines in the dataset. The only other gene with this pattern of selective dependency is STAT6, known to be regulated only by IL4 and IL13 signaling53,54,55. Of note, IL13 itself does not score in the DepMap dataset because of the pooled nature of those experiments; knock-out of IL13 would be rescued by nearby IL13-expressing cells. However, non-pooled experiments showed that IL13 is indeed required for the survival of cHL cell lines.
We note that CD4+ T cells in the cHL microenvironment express significant levels of IL13, consistent with IL13 serving as a key T cell cytokine. However, consistent with previous reports43,56,57,58,59, we also observed heterogeneous but robust IL13 expression in HRS cells themselves – a surprising phenomenon given that B cells typically do not express high levels of IL1342. The heterogeneous expression of IL13 in primary HRS cells suggests that HRS cells acquired IL13 expression over time, consistent with it representing a positive selection factor for survival. We speculate that surrounding T cells may establish IL13 signaling dependency in the early stages of cHL evolution, with subsequent selection for paracrine secretion of IL13 by HRS cells, either by epigenetic or somatic genome alterations. Consistent with this hypothesis, the Hodgkin lymphoma cell lines derived from patients with advanced disease with extranodal dissemination show strong and uniform expression of IL13. Furthermore, the recent description of somatic activating mutations in IL4R is consistent with this hypothesis, further supporting that HRS cells invoke multiple mechanisms to ensure IL13 signaling60. Moreover, the expression of IL13 and its receptor in HRS cells in EBV+ and EBV- disease suggests that this dependency may generalize to both disease subtypes43.
To explore the clinical translational potential of the role of IL13 signaling in cHL, we tested three IL13 receptor-blocking antibodies (Dupilumab, Lebrikizumab, and Tralokinumab) for their ability to recapitulate the effect of IL13, IL4R, or IL13RA1 genetic knock-out in cHL cell lines. In all three cases, antibody treatment at clinically achievable doses resulted in growth inhibition and apoptotic cell death. This finding builds upon observations from previous studies56,58,59,61 and is particularly noteworthy because these antibodies are already FDA-approved for treating atopic dermatitis44,45,46,47. We also note the recent report of using one of these antibodies, Dupilumab, to enhance responsiveness to PD1/PD-L1 checkpoint inhibitors in patients with lung cancer62. Taken together, these results suggest that IL13-blocking antibodies should be evaluated for their activity in patients with cHL.
A limitation of our study is the lack of in vivo validation of the IL13 hypothesis, due to the absence of in vivo models of Hodgkin lymphoma that faithfully recapitulate the microenvironment. Nevertheless, the availability of anti-IL13 antibodies that have been shown to be well-tolerated, coupled with in vitro activity at concentrations achievable with clinically-approved dosing, offers a clear path towards rapid testing of this hypothesis in a phase 1 clinical trial of patients with cHL.
Potential mechanisms of adaptive resistance will be important to explore in future studies. Our results suggest that autocrine IL4 expression may not be sufficient to mount resistance to anti-IL13 antibodies because cHL models that show high levels of IL4 expression show similar sensitivity to anti-IL4R (Dupilumab) and anti-IL13 (Lebrikizumab, Tralokinumab) antibodies. Our results also indicate that the cHL tumor microenvironment is a rich source of opportunities for therapeutic intervention, and the availability of a cHL spatial reference dataset should catalyze the evaluation of such targets. More broadly, our study describes a general approach for discovering cellular niches and microenvironment-derived therapeutic targets.
Methods
Tissue collection
After appropriate approval at institutional review boards at the Brigham and Women’s Hospital and Massachusetts General Hospital, and the Office of Research Subject Protection at the Broad Institute, excess excisional biopsy tissue specimens were collected from patients with newly diagnosed classic Hodgkin lymphoma. Additionally, reactive lymphoid tissue control samples were also collected. Demographic information for all study participants is provided in Supplementary Data 1. Written consent was obtained from participants whenever feasible under institutional umbrella research protocols. For cases where obtaining consent was not feasible, the institutional review board granted a waiver of informed consent for samples that were collected before 2015. Details of consent/IRB approval for each sample is provided in Supplementary Data 1. For the samples collected under an IRB-approved waiver of consent, we do not have explicit consent to share sex, age, and diagnosis, which in combination has the potential to identify patients. Therefore, age is expressed as ranges and individual sex information is redacted (summary statistics are reported). The specimens were embedded in OCT compound, snap-frozen, and stored at −80 °C. As a quality control step, tissue architecture was assessed by hematoxylin and eosin staining, and RNA integrity was determined using the Tapestation RNA ScreenTape system (RINe > 7.5).
Single nucleus isolation and RNA sequencing
Fresh frozen tissues (embedded in OCT) were cryo-sectioned at 20-micron thickness on a Leica CM1950 Cryostat (Leica, US) at −20 °C. Pre-cooled 3 mm circular (3332 P/25, Integra) biopsy punches were used to isolate regions of interest from tissue sections. The punched tissue sections were placed in wells of a 6-well plate, and 1 mL of extraction buffer (82 mM Na2SO4, 30 mM K2SO4, 10 mM Glucose, 10 mM HEPES, 5 mM MgCl2, 1% Kollidon VA64, 1% Triton X100, 0.1% BSA, 667 units/mL RNase-inhibitor (Biosearch technologies, 30281-1)) was added to each well. The sample was then triturated slowly using a P1000 24 times over 2 minutes, repeated five times. After dissociation, the tissue was transferred to a Falcon tube, and the volume was diluted to 30 mL with a wash buffer (82 mM Na2SO4, 30 mM K2SO4, 10 mM Glucose, 10 mM HEPES, 5 mM MgCl2, 0.1% BSA, 67 units/mL RNase-inhibitor. Diluted samples were split into two 50 mL Falcon tubes and centrifuged at 600 g for 10 min. Most of the supernatant was removed, and the samples were repooled. Pooled samples were passed through a 40 μm strainer and collected in a pre-cooled 1.5 mL Eppendorf tube. In a pre-cooled centrifuge, samples were further concentrated for 10 min at 200 g. All but 50 μL of the supernatant was discarded, and the pellet resuspended. This solution was stained with DAPI (Thermo Fisher Scientific, 62248), and the nuclei were quantified using a C-Chip Fuchs-Rosenthal disposable hemocytometer (INCYTO, DHC-F01-5). The sample was then appropriately diluted to recover 10,000 cells according to the 10x Genomics 3’ gene expression (v3.1 chemistry) user guide (CG000204, Rev D). Partitioning oil and gel beads were added to the Chromium Next GEM Chip G with the single nuclei suspensions. Following successful GEM generation, reverse transcription was performed, and cDNA was recovered and cleaned up using Dynabeads MyOne SILANE according to the 10x Genomics protocol. The recovered cDNA was then amplified for 12 cycles, and the products were quantified using Agilent Bioanalyzer High Sensitivity DNA Chip. Once cDNA quality was confirmed to be within expectations, the libraries were prepared according to the Chromium Next GEM Single Cell 3’ Reagent Kits v3.1 steps. Following the 10x Genomics library preparation, the samples were qualified using the High Sensitivity DNA Chip and quantified using the Qubit 3.0 fluorometer. The libraries were normalized to 2 nM and pooled. Libraries were sequenced on an Illumina Nextseq 550/500 or a Novaseq 6000 S2 flow cell and demultiplexed for downstream analysis.
Spatial transcriptomics
Slide-seqV2
Alternate 10 μm thick tissue sections were thawed onto 3 mm Slide-seq arrays. The arrays were transferred immediately to a hybridization solution (6X sodium chloride sodium citrate (SSC) with 2 U/μL Lucigen NxGen RNAse inhibitor) for 30 minutes at room temperature. The intervening sections were used for hematoxylin and eosin, as well as immunohistochemical staining. Libraries were generated using the previously published Slide-seqV2 protocol63,64. The libraries were prepared using the standard Illumina protocol and sequenced on either Nextseq 550/500 or Novaseq 6000 S2/S4 flowcells at a depth of 100 million reads per array. Samples were pooled to a concentration of 4 nM and were sequenced with the previously described read structure64. The final dataset includes at least two biological replicates (multiple Slide-seq arrays) per patient sample.
Visium Spatial GeneExpression for FFPE V1
Two formalin-fixed paraffin-embedded samples of newly diagnosed classic Hodgkin lymphoma (from two unique patients) were selected based on the DV200 metric (>50% of RNA molecules greater than 200 base pairs in length) determined using the Tapestation RNA ScreenTape system. Two 5-micron sections from each tumor were placed on the four 6.5 x 6.5 mm capture areas of the Visium spatial gene expression slide. Deparaffinization, staining (Hematoxylin and Eosin), imaging, de-crosslinking, overnight probe hybridization, ligating, barcoding, and library construction were performed per manufacturer recommendations (protocol CG000404 Rev B). Quality control analysis of libraries was performed using the Tapestation system, and verified libraries were sequenced on a Nextseq 500 high-output flow cell at a depth of >10,000 reads per spot for sample A and >45,000 reads per spot for sample B, which approaches saturation ( > 80%) for both samples.
CosMx Spatial Molecular Imager
Formalin-fixed, paraffin-embedded (FFPE) tissue sections were prepared for CosMx SMI profiling as previously described65. NanoString® ISH probes were prepared by incubation at 95 °C for 2 min and placed on ice. The ISH probe mix (1 nM 980 plex ISH probe, 10 nM Attenuation probes, 1X Buffer R, 0.1 U/μL SUPERase•In™ [Thermofisher] in DEPC H2O) was pipetted into the hybridization chamber. The hybridization chamber was sealed to prevent evaporation, and hybridization was performed at 37 °C overnight. Tissue sections were rinsed of excess probes in 2X SSCT for 1 min and washed twice in 50% formamide (VWR) in 2X SSC at 37 °C for 25 min, then twice with 2X SSC for 2 min at room temperature and blocked with 100 mM NHS-acetate in the dark for 15 min. A custom-made flow cell was affixed to the slide in preparation for loading onto the CosMx SMI instrument.
RNA target readout on the CosMx SMI instrument was performed as described previously65. After RNA readout, the tissue samples were incubated with a 4-fluorophore-conjugated antibody cocktail against CD298/B2M (488 nm), PanCK (532 nm), CD45 (594 nm), and CD3 (647 nm) proteins and DAPI stain in the CosMx SMI instrument for 2 h. After unbound antibodies and DAPI stain were washed with Reporter Wash Buffer, Imaging Buffer was added to the flow cell, and eight Z-stack images for the 5 channels (4 antibodies and DAPI) were captured.
Single-molecule RNA fluorescence in situ hybridization
The RNAscope HiPlex V2 assay (ACDbio) was performed as previously described66. FFPE sections were rehydrated on ice for 3 hours before sectioning. Tissue was cut to 5 μm sections and placed on charged slides. Slides were air-dried at room temperature overnight. The following day, slides were incubated at 60 °C for 2 hours. Tissue sections were deparaffinized and rehydrated in xylene, followed by ethanol and water per Advanced Cell Diagnostics (ACD) recommendations. EBV-encoded RNA (EBER) in situ hybridization was performed on FFPE sections using the EBER probe on a Ventana BenchMark automated platform with iView Blue detection, following the manufacturer’s protocol and as previously described67.
Histopathology and immunohistochemistry
Hematoxylin and Eosin (H&E) staining
Frozen tissue sections on glass slides were dipped in xylene, hydrated with a graded ethanol wash series, and stained with hematoxylin. A weakly alkaline ammonium hydroxide solution was used as a nuclear “bluing” reagent. The sections were stained with eosin and dehydrated through a graded ethanol series, xylene, dehydrated, and coverslipped. Brightfield images were taken using the Leica Aperio VERSA Brightfield, Fluorescence & FISH Digital Pathology Scanner under a 10x objective.
Immunohistochemistry
Immunohistochemical staining of frozen and formalin-fixed paraffin-embedded tissue sections was performed according to clinical procedures in the Department of Pathology at the Brigham and Women’s Hospital, as previously described68. The following primary antibodies were used: CD30 (Ber-H2, Dako, lot 41775232, dilution 1:25), CD3 (LN10, Leica, lot 6082226, dilution 1:300), CD20 (L26, Dako, lot 41769962, dilution 1:250), PAX5 (BSAP, BD, lot 2301658, dilution 1:100), and PD-L1 (E1L3N, Cell Signaling Technology, lot 13, dilution 1:100).
Functional validation assays
Cell culture
Mammalian cell lines L428, L1236, and UHO1 were obtained from the Leibniz Institute DSMZ and cultured in media and conditions recommended by the distributor. After initial thawing and one month of passaging, cells were sent to LabCorp for STR Profiling and Mycoplasma testing. Cells were regularly tested for Mycoplasma contamination using the Mycoscope PCR Mycoplasma Detection Kit (BIO-CAT MY01050-GL).
Pharmacologic assays
We used relative viability measurements and total cell counts to determine the dose-dependent response and time-to-effect of three IL13-blocking antibodies. For these experiments, the following three antibodies were used: Tralokinumab (MedChemExpress HY-P99053, lot: 237441), Dupilumab (Selleck Chemicals A2038-1mg, lots 02-04), and Lebrikizumab (MedChemExpress HY-P99025, lot: 594161). Human IgG4 Lambda (Millipore Sigma I4765-1MG) and Kappa (MedChemExpress HY-P99003) were used as isotype controls.
Each therapeutic antibody or isotype control was serially diluted (500 nM to 5 pM, 10-fold) in 96-well plates. Then, 20,000 cells were added per well and cultured for nine days. On day nine, relative viability was assessed using the CellTiter-Glo (CtG) Luminescent Cell Viability Assay (Promega G7572) per the manufacturer’s instructions. Luminescence was quantified using an EnVision Multilabel Plate Reader, and each antibody-treated condition was normalized to its isotype control. For the time course experiments, all antibody and isotype control treatments were 500 nM. At regular intervals, viability measurements were made using CtG and viable cell counts (Vi-cell Blue). The culture media was changed every three days, and the antibody was replenished.
Cytokine supplementation assays
To investigate the effect of putative growth factors identified computationally, L1236 and UHO1 cells were supplemented with recombinant cytokines (concentrations and sources in Supplementary Data 6). Cytokines were reconstituted and aliquoted according to the manufacturer’s recommendations. The assays were performed in 96 well plates using 20,000 cells per well in triplicate. Cytokine was replaced during media changes every three days, and viability was measured using CtG.
CRISPR/Cas9 knockout
Kanamycin-resistant plasmids containing the top three CRISPR/Cas9 guides for IL13, IL13Ra1, and IL4R (guide sequences are in Supplementary Data 7) were obtained from the Genomic Perturbation Platform. After overnight culture of E. coli in LB medium (50 µg/mL Kanamycin), plasmid DNA was extracted with the QIAGEN Maxi Plasmid Purification kit (Cat. 12162). Lentiviral particles were produced in HEK293T cells by co-transfection of guide plasmids with psPAX2 (Addgene #12260) and pMD2.G (Addgene #12259), then concentrated using Lenti-X Concentrator (Takara PT4421 2). L1236 cells engineered to express Cas9 (with Blasticidin and Puromycin resistance) were maintained in standard conditions plus 4 µg/mL Blasticidin S HCl (Invitrogen A1113903) for two weeks to ensure Cas9 activity. For transduction, 10 million cells per condition (plus a non-transduced control) were seeded in 6-well plates. The following day, Polybrene (Millipore-Sigma TR-1003-G) was added to 8 µg/mL, and 300 µL of concentrated lentivirus was added to each well (or DMEM to non-transduced controls). Cells were incubated overnight, then media was replaced, and 2 µg/mL Puromycin Dihydrochloride (Invitrogen A1113803) was added to select transduced cells. Knockouts targeting IL13 were supplemented with 0.1 µg/mL of recombinant IL13 (PeproTech 200-13). Cells were counted regularly for 15 days on the Vi-cell Blue. Once the population neared the instrument’s lower detection limit, genomic DNA was extracted (QIAamp DNA Micro kit, QIAGEN 56304). Editing efficiency ( > 75% edited sequences) was confirmed by CRISPR sequencing (MGH CCIB DNA core; CRISPRESSOv2 analysis; primer and guide sequences in Supplementary Data 7 and editing efficiency in Supplementary Data 8), as no high-quality antibodies were available to verify protein knockout. Custom primers were obtained from IDT.
Apoptosis assays
Caspase 3/7 activity was measured 48 hours after antibody treatment or transduction/selection using the CellEvent green fluorescence assay per manufacturer recommendations. Hoechst 33342 was used for nuclear counterstaining. The assay was performed with at least three technical replicates. Imaging was performed using the EVOS M7000 imaging system using the DAPI and GFP channels. The mean nuclear intensity of CellEvent green fluorescence was calculated per replicate using CellProfiler and normalized to the baseline level of fluorescence observed with the intergenic or isotype control. Staurosporine (1μM) was used as positive control.
Dependency Map data analysis
Gene effect scores derived from CRISPR knockout screens published by Broad’s Achilles35 and Sanger’s SCORE69 projects were downloaded from the internal DepMap portal (public 22Q2). Gene effect scores were inferred by Chronos70, with negative scores implying a fitness defect following gene knockout. As previously described, the scores are normalized such that non-essential and common essential genes have a median score of 0 and −1, respectively. A cell line was considered dependent if the probability of dependency was > 0.5. The probability of dependency is calculated for each gene effect score in a cell line as the probability that the score arises from a distribution of essential gene scores rather than non-essential gene scores71. To test whether any of our predicted ligand-receptor interactions were dependencies in Hodgkin lymphoma cell lines, we focused our analysis on our prioritized list of ligands and receptors and excluded common essential genes. We performed a Fisher’s exact test of independence to determine if there was any association between dependency on these genes (IL13RA1, IL4R, STAT6) and cancer type (Hodgkin lymphoma versus other cell lines).
Single nucleus RNA sequencing data analysis
Count matrix generation
Raw BCL files were demultiplexed and converted to FASTQ, which were then aligned to the GRCh38 human genome reference (cellranger reference 3.0.0, Ensembl v93 gene annotation, introns included) using the Cellranger count and mkfastq commands that are implemented in the Cumulus72 Cellranger workflow (v2.3.0) on Terra.
Ambient RNA correction and excluding empty droplets
We used the ‘remove-background’ function of CellBender v0.2.073 as implemented in the Cumulus CellBender workflow (v2.1.1) on Terra. The parameter ‘expected-cells’ was obtained from the Cell Ranger metric ‘Estimated Number of Cells’.
Doublet prediction
The gene-barcode expression count matrices were processed individually in R using Seurat v4. Initial filters were applied to exclude genes expressed in fewer than 3 cells and cells with 200 or fewer genes. ScDblFnder74 was then used to estimate doublets for each sample individually, with the expected number of doublets set as 1% per 1000 cells captured. The uncorrected expression count matrix (before ambient RNA correction using CellBender) was used as the input for doublet prediction.
Quality control, filtering, and dimensionality reduction
All the individual sample datasets were merged into one combined dataset. Filters were applied to exclude all estimated doublets, cells with 200 or fewer genes, 400 or fewer transcripts, and % mitochondrial reads > 5%. The combined, filtered count matrix was log-transformed and normalized using the NormalizeData function and the LogNormalize method. The top 2000 highly variable genes were selected for downstream analysis using the ‘vst’ method in the FindVariableFeatures function. The expression matrix was then scaled and centered using the ScaleData function. We then performed dimensionality reduction using the principal component analysis (PCA) and further performed uniform manifold projection and approximation (UMAP) using the first 50 principal components. The UMAP embeddings were used only for visualization purposes.
Data Integration
To facilitate cell type annotation, we used Harmony v0.175 to construct a shared embedding in which cells group by shared cell types across multiple samples while minimizing technical sources of variation. We performed integration using multiple covariates that reflect multiple partially related sources of biological and technical variation in the data (sequencing batch, library preparation batch, sequencing technology, donor, and GEM lane); see supplementary data 1 for complete details of batch variables. Convergence was achieved in 10 iterations, and the first 50 Harmony-corrected PCs were used to compute UMAP embeddings and a shared nearest-neighbor graph for downstream analysis and visualization.
Cell type annotation
Preliminary cell type labels were obtained by applying the SingleR76 algorithm using multiple immune cell reference datasets (HumanPrimaryCellAtlasData, BlueprintEncodeData, DatabaseImmuneCellExpressionData, MonacoImmuneData). The CellTypist77 algorithm was also used to assign preliminary cell type identities using the Immune_All_Low and Immune_All_High references. Initial broad cell type labels were assigned manually using a combination of automated annotation results and known cell type markers/signatures of immune and stromal cell types. This approach led to the identification of tumor (HRS) cells, B/plasma cells, T/NK cells, myeloid cells, and stromal cells (fibroblasts and endothelial cells). Next, we split the Seurat object into these major cell types and re-ran the scaling, dimensionality reduction, and clustering procedure described above. The resulting clusters were annotated using the same approach, and the cell type labels were added to the main integrated object. The myeloid cells were classified into finer cell types based on label transfer analysis (as implemented in Seurat78) from the MoMac-verse reference dataset41. Briefly, cell type label transfer was performed by projecting the PCA structure of the reference dataset onto our query dataset. First, common anchors were identified between our dataset and the MoMac-verse dataset using FindTransferAnchors, and then TransferData was used to classify the cells in our dataset based on the cell type labels of the MoMac-verse dataset. Principal component analysis was used as the dimensionality reduction method, and the first 50 principal components were used for this analysis.
CNV inference
We use inferCNV v.1.7.179 to identify evidence for somatic large-scale chromosomal copy number alterations in HRS cells by comparing gene expression intensity across positions of tumor genome in comparison to a set of reference ‘normal’ cells in the single nuclei dataset. For this analysis, we subset the data to include samples from cHL patients only. We use the cell-type annotation described previously to select HRS cells as the malignant set and all other cell types as the ‘normal’ reference. Due to performance constraints, we downsampled the reference cells so that there were a maximum of 7000 cells from each reference cell type. We use the “denoise,” “HMM,” and “cluster_by_groups” settings to enable the denoising procedure, CNV predictions via HMM, and separate clustering of cells from each patient, respectively. We set the cutoff value to 0.3 and the sd_amplifier to 2. All other settings were set to the default values. Gene expression levels were represented in a heatmap where genes were sorted by genomic position and ordered within each chromosome.
Cell-type compositional analysis
We use the single-cell compositional data analysis framework (scCODA)37 to compute the statistical significance of differences in cell-type composition between sample groups in the single-nuclei data. We specifically test for effects of the binary covariates disease_status (10 Hodgkin Lymphoma versus 5 Healthy samples) and ebv_status (5 EBV positive versus 5 EBV negative samples) in cell-type proportion obtained from level 4 annotations. To run scCODA, we first compute the proportion of each of the cell types for each donor. We then use scCODA’s automatic reference selection to select a reference cell type with the least relative abundance dispersion over all samples while being present in at least 95% of the samples. The reference cell type selected for the single-nuclei data using this criterion was ILC3. We run scCODA’s parameter inference routine using the Hamiltonian Monte Carlo sampling method with the default chain length of 20,000. Following the guidelines in the scCODA package, we set an FDR of 0.2 for the disease_status analysis and an FDR of 0.4 for the ebv_status analysis. The main figures report the cell types for which the given covariate has a statistically credible effect.
Cell type-specific differential expression analysis
To identify genes differentially expressed between different sample groups within specific cell types in the single-nuclei data, we perform pseudo-bulk differential gene expression analysis using muscat80 and DESeq2. We first obtain pseudo-bulk expression profiles by summing the raw UMI counts for each gene, sample, and cell type using the aggregateData method in muscat. We filter out cell-type-sample pairs with less than 10 cells to ensure sufficient statistical power. All of muscat’s internal sample and gene filtering steps were also enabled. We then test differential expression using the DESeq2 method incorporated in muscat’s pbDS function. A gene was considered differentially expressed in a given cell type if the Benjamini-Hochberg adjusted local (within cell-type) p-value was <0.05 and absolute fold change was > 2.
EBV reference mapping
For both 10x genomics single-nuclei and slide-seq spatial transcriptomics libraries, we used host genome (hg38) mapped BAM files as our starting point and extracted unmapped reads (flag 4 set) using samtools, which contain potential viral or otherwise non-host derived transcripts. To find EBV transcripts, we used the NCBI reference sequence NC_007605.1. Several low-complexity regions of this genome would give rise to false-positive alignments of low-quality reads. As such, we used USEARCH to mask these regions using the fastx_mask function. We mapped the unmapped reads using the STAR aligner to this filtered genome. For 10x libraries sequenced with 90 cycles on read2, we used a stringent alignment score (tag ‘AS’) of 70 and a number of mismatches (tag ‘nM’) of at most 4 to assign confident EBV calls. For slide-seq libraries sequenced with 40 cycles, we used an alignment score of 30 and a number of mismatches of at most 2. Barcodes and UMIs associated with each confidently mapped read were then extracted from the original BAM file and deduplicated to find the number of EBV-mapped transcripts in each 10x droplet or slide-seq spatial spot.
Disease-associated immune cell signatures analysis
We use tensor factorization framework Consensus Zero Inflated Poisson Tensor Factorization (C-ZIPTF)81 to identify significant gene expression programs that can be associated directly with biological processes within specific cell types and donor contexts from the single-nuclei data in an unsupervised manner. We first create a pseudo-bulk expression tensor by summing the raw UMI counts for each gene, donor, and cell type. The pseudo-bulk data is represented as a 3-way tensor \(\chi\) with dimensions \({N}_{D}\times {N}_{C}\times {N}_{G}\), where \({N}_{D}\), \({N}_{C}\), and \({N}_{D}\) represent the number of donors, cell types, and genes, respectively. \({\chi }_{i,j,k}\) denotes the aggregate counts of gene \(k\) across cell type \(j\) for donor \(i\). To facilitate the biological interpretability of factors and reduce noise in the tensor formed, we removed genes that are either not provided with HGNC symbols or not labeled as protein-coding in the HGNC database82. We also filtered out genes with a total count of less than 50 across all cells. The resulting tensor was total-count normalized, so each sample-cell type pair has \(1{0}^{6}\) counts. We then apply C-ZIPTF on the tensor formed.
Briefly, C-ZIPTF employs a Bayesian approach to factorize tensor \(\chi\) into a set of factors corresponding to donor, cell type, and gene modes by modeling the multidimensional count data using a zero-inflated Poisson distribution. Pseudo-bulk counts \(\chi\) are modeled as draws from a Zero-inflated Poisson distribution, i.e., for every \({ijk},1\le i\le {N}_{D},1\le j\le {N}_{C},1\le k\le {N}_{G}\):
where \({x}_{{ijk}}\ge 0,{\lambda }_{{ijk}}\) is the expected Poisson count, and \({p}_{{ijk}}\) is the probability of extra zeros. As an abbreviation, it is written as \({\chi }_{{ijk}}\approx {ZIP}({\lambda }_{{ijk}},{p}_{{ijk}}).\) C-ZIPTF was applied to the pseudo-bulk tensor with a rank of 24. The factorization achieved a high explained variance of 0.916 and a strong cophenetic correlation of 0.931.
The gene latent factors \({g}^{r}\), 1 ≤ r ≤ R, represent ranked lists of genes with associated weights. They reveal the expression pattern by capturing co-regulation and co-expression patterns among groups of genes. To identify the top genes defining each factor, we first normalized the gene mode factor matrix such that the L1 norm of each factor equaled 1. We then filtered out genes based on two criteria: (1) genes whose maximum loading across all factors was less than the median of all gene loadings and (2) genes whose entropy was less than the median entropy plus twice the median absolute deviation. Additionally, we conducted gene set enrichment analysis of these factors using GSEApy83 in Python, offering a broader perspective on the associated biological functions and pathways. We assess the loadings in \({d}^{r}\) to determine the activity level of the GEP in individual donors. Similarly, the cell type loadings in \({c}^{r}\) indicate the activity level of the GEP across different cell types.
Intratumor heterogeneity analysis
To define the core gene expression programs in HRS cells, we conduct matrix factorization using Zero-Inflated Poisson Tensor Factorization. We subset the data to include only HRS cells and obtain a two-dimensional matrix of cells × genes. Here again, we remove genes that are either not provided with HGNC symbols or not labeled as protein-coding in the HGNC database82 and filter out genes with a total count of less than 50 across all cells. This matrix was then total-count normalized so that each cell had 106 counts. We then apply ZIPTF to this matrix, as matrices are essentially two-way tensors. To identify the top genes for each factor, we normalize the gene mode factor matrix such that the L1 norm of each factor equals 1. We then filter out genes based on two criteria: genes whose maximum loading across all factors is less than the median of all gene loadings and genes whose entropy is less than the median entropy plus twice the median absolute deviation. The programs are then annotated manually using gene set enrichment analysis and prior knowledge.
Ligand receptor analysis
Ligand receptor interaction inference was performed using the LIANA84 tool on the single-nucleus RNAseq dataset using the OmniPath85 Consensus ligand-receptor interaction database at intermediate cell type annotation hierarchy (“cell_types_level_3”). Ligands or receptors expressed in less than 5% of cells per cluster were excluded. For multimeric receptor complexes, the expression of individual subunits was considered, i.e., receptor complex annotations were not used to reduce the stringency of this analysis due to the sparsity of this annotation and expression levels of receptor complex members. Interactions were prioritized using expression (SingleCellSignalR LRscore86, > 0.4), cell type specificity (NATMI edge specificity87, > 0.01), and ligand-receptor colocalization metrics (described later). LRscore is a non-negative regularized score comparable between datasets (ranges from 0-1) and reflects the expression magnitude of ligand and receptor. The NATMI edge specificity score ranges from 0 to 1, where 1 means the ligand and receptor are uniquely expressed in each pair of cell types.
Spatial transcriptomics data analysis
Slide-seqV2 alignment and quality control
Demultiplexing, genome alignment, and spatial barcode matching were performed using Slide-seq tools v0.2 pipeline64. The resulting count and spatial barcode matrices were loaded into Seurat for downstream analysis. The “stray” barcodes not present in the densely packed center of the array were cropped out and excluded from analysis by manual inspection of each array.
Visium data processing
The demultiplexed FASTQ files were aligned against GRCh38-2020-A human transcriptome reference, and expression count matrices were generated using Space Ranger (v1.3.1). Space Ranger was also used to align the acquired images with the spatially resolved sequencing data.
Cell type deconvolution
RCTD v2.0.0, part of the spacexr v2.0.2 package88,89 was used for cell type deconvolution. For the slide-seqV2 dataset, RCTD was run in doublet mode for each sample individually (replicate mode) with the appropriate snRNAseq dataset (lymph node or Hodgkin lymphoma) with corresponding broad cell type labels (level 3) that were assigned as described in “cell type annotation” above. Importantly, beads that were confidently classified as a singlet or doublet were retained for any subsequent analysis. The beads that were not confidently classified (doublet_uncertain and reject classes) were excluded. This was facilitated by the high spatial resolution (10 microns) of our method, SlideseqV2. Therefore, we believe that this analysis closely approximates cell count rather than beads. Some specific cell types were grouped into more coarse cell type classes to allow the model to fall back to the cell class if classification at the level of the granular cell type is not possible. CD4+ and CD8+ T cells were grouped under the T cell class, fibroblasts and follicular dendritic cells were grouped under the stromal cell class, monocytes/macrophages and myeloid dendritic cells were grouped under the “myeloid cell” class, and the lymphatic and vascular endothelial cells were grouped under the “vascular” cell type class. For the Visium dataset, RCTD was run in full mode.
CosMx image processing and feature extraction
Raw image processing and feature extraction were performed using the SMI data processing pipeline65 which includes registration, feature detection, and localization. 3D rigid image registration was made using fiducials embedded in the samples matched with the fixed image reference established at the beginning of the SMI run to correct for any shift. Secondly, the RNA image analysis algorithm was used to identify reporter signature locations in the X, Y, and Z axes along with the assigned confidence. The reporter signature locations and the associated features were collated into a single list. Lastly, the XYZ location information of individual target transcript was extracted and recorded in a table by secondary analysis algorithm, as previously described65.
CosMx cell segmentation
The Z-stack images of immunostaining + DAPI were used to draw cell boundaries on the samples. A cell segmentation pipeline using the Cellpose machine learning algorithm90,91 was used to accurately assign transcripts to cell locations and subcellular compartments. The transcript profile of individual cells was generated by combining target transcript location and cell segmentation boundaries. Cells with fewer than 20 total transcripts assigned were omitted from the analysis.
Spatial clustering analysis
To determine whether the tumor cells exhibited a random, dispersed, or clustered distribution, we computed Ripley’s statistics using Ripley’s L function (variance-normalized version of Ripley’s K statistic) as implemented in the Squidpy package92. Arrays with insufficient numbers of tumor cells were excluded from this analysis. The Ripley’s L function was computed for each array with sufficient numbers of tumor cells ( > 100). The results were visualized and interpreted to assess the spatial organization of tumor cells. For interpretation, we use confidence envelopes: if Ripley’s L at distance d, L(d), exceeds the upper confidence envelope, it indicates statistically significant clustering at that distance; if L(d) falls between the upper and lower confidence envelopes, the distribution is not significantly different from random; and if L(d) falls below the lower confidence envelope, it suggests statistically significant dispersion or regular spacing at that distance.
Spatial cell type enrichment
We explore the significant enrichment of cell types surrounding the HRS cells using spatial transcriptomics data, including techniques such as Slide-seqV2 and the CosMx Spatial Molecular Imager (SMI). For Slide-seqV2, cell type deconvolution was performed using RCTD v2.0.0 as discussed above in “Cell type deconvolution”. For CosMx, cell type enrichment analysis utilizes the single-cell resolution data generated through the segmentation pipeline described earlier in “CosMx cell segmentation”. This approach allows direct assignment of transcripts to individual cells and enables enrichment analysis without the need for further deconvolution.
We construct a neighborhood graph by considering the spatial locations of the beads (voxels). We count the number of beads classified as cell type \(i\) within \(r\ge 0\) microns to HRS cells, denote it as \({{N}_{r}}^{0}({C}_{i})\). To assess the statistical significance of enrichment or depletion of each cell type, we generate a null distribution by randomly permuting the cell type labels of non-HRS cells only, while keeping HRS cell labels and their spatial positions fixed. This approach preserves the co-localizing nature of HRS cells. For each permutation iteration j, the count of cel type i within radius \(r\) microns of HRS cells is recorded as \({{N}_{r}}^{j}({C}_{i})\). We estimate the mean and the variance (\({\mu }_{r}({C}_{i})\), \({\sigma }_{r}({C}_{i})\)) of the counts across permutations and calculate the enrichment score as following
We also perform a one-tailed p-value test to assess the enrichment (or depletion) of cell types. Let \(H\left(.\right)\) be the indicator function such that \(H(x)=1\) if \(x > 0,\) and \(0\) otherwise, then we
We individually perform this test for each tissue and calculate the median of the enrichment scores across all tissues. The resulting p-values from each tissue are combined using Fisher’s combined probability test by utilizing the scipy.stats.combine_pvalues function from the SciPy library. To control the expected proportion of false discoveries among the rejected hypotheses, we apply the Benjamini-Hochberg procedure for false discoveryrate (FDR) correction. This adjustment ensures a more accurate control of the overall false positive rate in our analysis.
Cell-type specific spatial differential expression
After cell type deconvolution using RCTD, we performed cell type specific differential expression analysis across the entire multi-sample multi-replicate slideseqV2 dataset using C-SIDE89. The samples represent tumors from different patients, and replicates refer to other sections/arrays from the same tumor. We hypothesized that the local density and proximity of tumor cells are plausible predictors of gene expression in non-tumor cells. Therefore, we first calculated the normalized (min-max normalization per array) density of HRS cells for each pixel within a 200 µm radius. Second, for each pixel, we calculated the inverse of the distance to the nearest tumor cell (if distance = 0, proximity = 1). We then ran CSIDE in replicate mode (run.CSIDE.replicates) using these two orthogonal continuous explanatory variables to test for differentially expressed genes in non-tumor cells as a function of local proximity or density of tumor cells. Default parameters were used except for the normalized expression threshold for gene selection (gene_threshold = 1e-06) and the minimum number of cells of each cell type to be used (cell_type_threshold = 0). As implemented in CSIDE (CSIDE.population.inference), population-level differential expression analysis was performed across all samples to identify genes that consistently show a similar pattern of differential expression across multiple samples. These tests are two-sided. Default parameters were used except for the minimum number of groups (if use.groups) for which a gene must converge (MIN.CONV.GROUPS = 3). Genes with a log2 fold change value of > 0.5 and a q value of <0.05 were considered significantly differentially expressed. Cell type-specific genes were prioritized to minimize the potential of false-positive genes that may result from cell type misclassification and/or diffusion. Cell-type-specific genes were enriched by setting the ct_prop (minimum ratio of expression in a given cell type compared to all other cell types) threshold at a stringently high level (ct_prop > 0.5).
Ligand receptor colocalization
To test the significance of colocalization of ligand-receptor pairs as a measure of interaction between cell types expressing those, we set up a series of permutation tests. We used count matrices of slide-seq libraries from lymphoma samples and ligand-receptor interacting pairs inferred from single-nuclei data using LIANA as our starting point. To remove potential bias due to empty regions of the arrays, which contain very few counts, we removed beads with the maximum number of detected genes below a certain threshold. This threshold was determined by finding the maximum between 150 (heuristically determined to reflect empty tissue sections in most arrays) or the value of the lower quartile of gene counts per bead (a higher value on a few arrays that had unusually high capture of transcripts). Furthermore, for each array, we did not analyze pairs where either ligand or receptor had less than 20 representative beads on that array. For specific highly expressed ligand or receptor genes such as B2M or CD3E, we observed that most of the beads on the arrays had nonzero expression. To binarize the expression of such genes into positive and negative categories, we considered beads that had expression below the 75th percentile of counts for that gene as negative. This value was selected heuristically so that only a few highly expressed genes would be affected. This thresholding mechanism would not affect the vast majority of genes that have mostly either counts of 1 or 0 in all beads. Subsequently, we counted the number of receptor-positive beads in an area around each ligand-positive bead as the observed value. We selected 1000 random sets of points equal to the number of ligand-positive beads from the set of beads deemed to contain tissue and counted the receptor-positive beads around each random ligand position. We calculated the significance of this permutation test as (1+n)/(N + 1), where n is the number of times the random permutation yielded a higher count than the observed value. We also calculated an effect size equal to the ratio of observed value and mean of values obtained from random permutations. We repeated this procedure across all arrays and over multiple radii. We combined the p-values for each radius using Fisher’s method and corrected them across the entire set of ligand-receptor pairs using the Benjamini-Hochberg method. The combined effect size for each pair and radius was calculated as the average of this value across all arrays. The product of the p-value and the effect size were used as the combined spatial association metric to reflect consistency (across replicates) and magnitude of colocalization (shown in Fig. 3). We show that for the top interactions inferred, the p values and effect size estimates are robust across multiple length scales (30, 100, 300 pixels, Supplementary Fig. 11) suggesting that a substantial number of top interactions are reproducibly identified across different distance thresholds, providing confidence that our main findings are credible are not artifacts of a specific spatial tuning parameter.
Statistics and Reproducibility
For sample size estimation, we aimed to have sufficient representation of samples (n > 3) with respect to EBV status (positive or negative), age (>50, <50) and histologic subtype (nodular sclerosis and mixed cellularity). These sample sizes are sufficient to support the claims made in the manuscript because several inferences were validated with an independent dataset or experimentally. No specific sample size calculation was used. No data were excluded from the analyses. Multiple orthogonal spatial assays (SlideseqV2, Visium, CosMx, RNAscope) were used to replicate and validate the key findings. All in vitro experiments were repeated at least twice on different days, reagent lots and experimentalist to ensure reproducibility. All attempts at replication were successful. The validation in vitro experiments were randomized i.e., cell line samples were randomly assigned to control (intergenic guides, isotype control) or experimental treatment conditions (cutting guides, antibodies). Whenever possible, investigators were blinded to group allocation for experiments. All the single-cell and spatial data collection and analysis were conducted with blinding of group allocation. Analysis of the in vitro viability and imaging data was conducted in a blinded fashion to minimize bias.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the processed single-nucleus and spatial transcriptomics datasets are freely accessible on figshare (https://doi.org/10.6084/m9.figshare.28220345). Because the data derive from human samples and contain potentially identifiable genetic information, the raw sequencing files have been deposited in the Database of Genotypes and Phenotypes (dbGaP), where they are available under controlled access under accession code phs004387. Access will be granted within three business days to qualified researchers for the sole purpose of health-related research, subject to final approval by the dbGaP Data Access Committee. Data will remain available through dbGaP for a minimum of 10 years after publication. Gene effect scores derived from CRISPR knockout screens and bulk RNA sequencing data of human cancer cell lines (including Hodgkin lymphoma cell lines) used in this study were published by the Broad’s Achilles35 and Sanger’s SCORE69 projects and are available in figshare93 [link] via the DepMap portal (22Q2 public release). The publicly available gene expression datasets of microdissected primary HRS cells used in this study are available in the GEO database under accession codes GSE1245334 and GSE3913333. The ‘MoMac-VERSE’ myeloid reference dataset41 used in this study for label transfer analysis of myeloid cells is available in the GEO database under accession code GSE178209 and https://gustaveroussy.github.io/FG-Lab/. Source Data for the in vitro studies are provided with this paper. Source data are provided with this paper.
Code availability
Code to reproduce all the analyses in this manuscript is available on Github: https://github.com/klarman-cell-observatory/chl_analysis and Zenodo: https://doi.org/10.5281/zenodo.1709452494. Code for the ligand-receptor colocalization analysis and quantification of unmapped (viral/microbial) reads are also available on GitHub (https://github.com/mehdiborji/LR_colocalization, https://github.com/mehdiborji/align_unmapped).
References
Hanahan, D. & Weinberg, R. A. The Hallmarks of Cancer. Cell 100, 57–70 (2000).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, 646–674 (2011).
Khaled, A. R. & Durum, S. K. Lymphocide: cytokines and the control of lymphoid homeostasis. Nat. Rev. Immunol. 2, 817–830 (2002).
Schmitz, R. et al. Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N. Engl. J. Med. 378, 1396–1407 (2018).
Chapuy, B. et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 24, 679–690 (2018).
Wen, T., Wang, J., Shi, Y., Qian, H. & Liu, P. Inhibitors targeting Bruton’s tyrosine kinase in cancers: drug development advances. Leukemia 35, 312–332 (2021).
Wilson, W. H. et al. Effect of ibrutinib with R-CHOP chemotherapy in genetic subtypes of DLBCL. Cancer Cell 39, 1643–1653.e3 (2021).
Younes, A. et al. PHOENIX investigators. Randomized phase III trial of ibrutinib and rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone in non-germinal center B-cell diffuse large B-cell lymphoma. J. Clin. Oncol. 37, 1285–1295 (2019).
Scott, D. W. & Gascoyne, R. D. The tumour microenvironment in B cell lymphomas. Nat. Rev. Cancer 14, 517–534 (2014).
Alaggio, R. et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 36, 1720–1748 (2022).
Campo, E. et al. The International Consensus Classification of Mature Lymphoid Neoplasms: a report from the Clinical Advisory Committee. Blood 140, 1229–1253 (2022).
Steen, C. B. et al. The landscape of tumor cell states and ecosystems in diffuse large B cell lymphoma. Cancer Cell 39, 1422–1437.e10 (2021).
Connors, J. M. et al. Hodgkin lymphoma. Nat. Rev. Dis. Prim. 6, 61 (2020).
Stein, H. & Diehl, V. First Hodgkin Cell Line L428 and the CD30 Antigen: Their Role for Diagnostic and Treatment of CD30-positive Neoplasms. Hematol. Oncol. Clin. North Am. 28, 1–11 (2014).
Zech, L., Haglund, U., Nilsson, K. & Klein, G. Characteristic chromosomal abnormalities in biopsies and lymphoid-cell lines from patients with Burkitt and non-Burkitt lymphomas. Int. J. Cancer 17, 47–56 (1976).
Kaplan, H. S. & Gartner, S. “Sternberg-reed” giant cells of Hodgkin’s Disease: cultivation in vitro, heterotransplantation, and characterization as neoplastic macrophages. Int. J. Cancer 19, 511–525 (1977).
Wienand, K. et al. Genomic analyses of flow-sorted Hodgkin Reed-Sternberg cells reveal complementary mechanisms of immune evasion. Blood Adv. 3, 4065–4080 (2019).
Reichel, J. et al. Flow sorting and exome sequencing reveal the oncogenome of primary Hodgkin and Reed-Sternberg cells. Blood 125, 1061–1072 (2015).
Maura, F. et al. Molecular evolution of classic Hodgkin lymphoma revealed through whole-genome sequencing of Hodgkin and Reed Sternberg cells. Blood Cancer Discov. 4, 208–227 (2023).
Gomez, F. et al. Ultra-deep sequencing reveals the mutational landscape of classical Hodgkin lymphoma. Cancer Res. Commun. 3, 2312–2330 (2023).
Roemer, M. G. M. et al. PD-L1 and PD-L2 Genetic Alterations Define Classical Hodgkin Lymphoma and Predict Outcome. J. Clin. Oncol. 34, 2690–2697 (2016).
Green, M. R. et al. Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 116, 3268–3277 (2010).
Carey, C. D. et al. Topological analysis reveals a PD-L1-associated microenvironmental niche for Reed-Sternberg cells in Hodgkin lymphoma. Blood 130, 2420–2430 (2017).
Patel, S. S. et al. The microenvironmental niche in classic Hodgkin lymphoma is enriched for CTLA-4–positive T cells that are PD-1–negative. Blood 134, 2059–2069 (2019).
Aoki, T. et al. Single-cell transcriptome analysis reveals disease-defining t-cell subsets in the tumor microenvironment of classic hodgkin lymphoma. Cancer Discov. 10, 406–421 (2020).
Chen, X. et al. T cell states, repertoire and function in classical Hodgkin lymphoma revealed through single-cell analyses. Cancer Immunol. Res. https://doi.org/10.1158/2326-6066.CIR-23-0547 (2024).
Aoki, T. et al. Single-cell profiling reveals the importance of CXCL13/CXCR5 axis biology in lymphocyte-rich classic Hodgkin lymphoma. Proc. Natl. Acad. Sci. USA 118, e2105822118 (2021).
Steidl, C. et al. Tumor-Associated Macrophages and Survival in Classic Hodgkin’s Lymphoma. N. Engl. J. Med. 362, 875–885 (2010).
de Kanter, J. K. et al. Single-cell RNA sequencing of pediatric Hodgkin lymphoma to study the inhibition of T cell subtypes. HemaSphere 8, e149 (2024).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Neftel, C. et al. An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma. Cell 178, 835–849.e21 (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Steidl, C. et al. Gene expression profiling of microdissected Hodgkin Reed-Sternberg cells correlates with treatment outcome in classical Hodgkin lymphoma. Blood 120, 3530–3540 (2012).
Weniger, M. A. et al. Human CD30+ B cells represent a unique subset related to Hodgkin lymphoma cells. J. Clin. Invest 128,, 2996–3007 (2018).
Tsherniak, A. et al. Defining a Cancer Dependency Map. Cell 170, 564–576.e16 (2017).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, (2021).
Chesneau, M. et al. Efficient expansion of human granzyme B-expressing B cells with potent regulatory properties. J. Immunol. 205, 2391–2401 (2020).
Gonzalez-Figueroa, P. et al. Follicular regulatory T cells produce neuritin to regulate B cells. Cell 184, 1775–1789.e19 (2021).
Paczkowska, J. et al. Cancer-specific innate and adaptive immune rewiring drives resistance to PD-1 blockade in classic Hodgkin lymphoma. Nat. Commun. 15, 10740 (2024).
Mulder, K. et al. Cross-tissue single-cell landscape of human monocytes and macrophages in health and disease. Immunity 54, 1883–1900.e5 (2021).
Punnonen, J. et al. Interleukin 13 induces interleukin 4-independent IgG4 and IgE synthesis and CD23 expression by human B cells. Proc. Natl. Acad. Sci. USA 90, 3730–3734 (1993).
Skinnider, B. F. et al. Interleukin 13 and interleukin 13 receptor are frequently expressed by Hodgkin and Reed-Sternberg cells of Hodgkin lymphoma. Blood, J. Am. Soc. Hematol. 97, 250–255 (2001).
Paller, A. S. et al. Efficacy and Safety of Tralokinumab in Adolescents With Moderate to Severe Atopic Dermatitis: The Phase 3 ECZTRA 6 Randomized Clinical Trial. JAMA Dermatol 159, 596–605 (2023).
Silverberg, J. I. et al. Two Phase 3 Trials of Lebrikizumab for Moderate-to-Severe Atopic Dermatitis. N. Engl. J. Med. 388, 1080–1091 (2023).
Guttman-Yassky, E. et al. Efficacy and Safety of Lebrikizumab, a High-Affinity Interleukin 13 Inhibitor, in Adults With Moderate to Severe Atopic Dermatitis: A Phase 2b Randomized Clinical Trial. JAMA Dermatol 156, 411–420 (2020).
Simpson, E. L. et al. SOLO 1 and SOLO 2 Investigators. Two phase 3 trials of dupilumab versus placebo in atopic dermatitis. N. Engl. J. Med. 375, 2335–2348 (2016).
Cader, F. Z. et al. Mass cytometry of Hodgkin lymphoma reveals a CD41 regulatory T-cell–rich and exhausted T-effector microenvironment. Blood 132, 825–836 (2018).
Stewart, B. J. et al. Spatial and molecular profiling of the mononuclear phagocyte network in classic Hodgkin lymphoma. Blood 141, 2343–2358 (2023).
Menéndez, V. et al. Mapping the spatial dynamics of the CD4+ T cell spectrum in classical Hodgkin lymphoma. Mod. Pathol. 37, 100551 (2024).
Veldman, J. et al. Rosetting T cells in Hodgkin lymphoma are activated by immunological synapse components HLA class II and CD58. Blood 136, 2437–2441 (2020).
Aoki, T. et al Spatially Resolved Tumor Microenvironment Predicts Treatment Outcomes in Relapsed/Refractory Hodgkin Lymphoma. J. Clin. Oncol. JCO2301115 https://doi.org/10.1200/JCO.23.01115 (2023).
Quelle, F. W. et al. Cloning of murine Stat6 and human Stat6, Stat proteins that are tyrosine phosphorylated in responses to IL-4 and IL-3 but are not required for mitogenesis. Mol. Cell. Biol. 15, 3336–3343 (1995).
Shimoda, K. et al. Lack of IL-4-induced Th2 response and IgE class switching in mice with disrupted Stat6 gene. Nature 380, 630–633 (1996).
Hou, J. et al. An interleukin-4-induced transcription factor: IL-4 Stat. Science 265, 1701–1706 (1994).
Kapp, U. et al. Interleukin 13 is secreted by and stimulates the growth of Hodgkin and Reed-Sternberg cells. J. Exp. Med. 189, 1939–1946 (1999).
Nakayama, S. et al. Role of mast cells in fibrosis of classical Hodgkin lymphoma. Int. J. Immunopathol. Pharmacol. 29, 603–611 (2016).
Natoli, A. et al. Targeting the IL-4/IL-13 signaling pathway sensitizes Hodgkin lymphoma cells to chemotherapeutic drugs: IL4/STAT6 regulate survival of Hodgkin lymphoma. Int. J. Cancer 133, 1945–1954 (2013).
Stanelle, J., Döring, C., Hansmann, M.-L. & Küppers, R. Mechanisms of aberrant GATA3 expression in classical Hodgkin lymphoma and its consequences for the cytokine profile of Hodgkin and Reed/Sternberg cells. Blood 116, 4202–4211 (2010).
Alig, S. K. et al. Distinct Hodgkin lymphoma subtypes defined by noninvasive genomic profiling. Nature 1–3 https://doi.org/10.1038/s41586-023-06903-x (2023).
Raia, V. et al. Dynamic mathematical modeling of IL13-induced signaling in Hodgkin and primary mediastinal B-cell lymphoma allows prediction of therapeutic targets. Cancer Res 71, 693–704 (2011).
LaMarche, N. M. et al. An IL-4 signalling axis in bone marrow drives pro-tumorigenic myelopoiesis. Nature 625, 166–174 (2024).
Stickels, R. et al. Library Generation using Slide-seqV2. (2021).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 1–7 https://doi.org/10.1038/s41587-020-0739-1.
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Song, J., Kim, J. Y., Kim, S. & Park, Y. Utility of Epstein-Barr viral load in blood for diagnosing and predicting prognosis of lymphoma: A comparison with Epstein-Barr virus-encoded RNA in situ hybridization. J. Mol. Diagn. 24, 977–991 (2022).
Shanmugam, V. et al Identification of diverse activating mutations of the RAS-MAPK pathway in histiocytic sarcoma. Mod. Pathol. 1 https://doi.org/10.1038/s41379-018-0200-x (2019).
Dwane, L. et al. Project Score database: a resource for investigating cancer cell dependencies and prioritizing therapeutic targets. Nucleic Acids Res 49, D1365–D1372 (2021).
Dempster, J. M. et al. Chronos: a cell population dynamics model of CRISPR experiments that improves inference of gene fitness effects. Genome Biol. 22, 343 (2021).
Dempster, J. M. et al. Extracting biological insights from the project Achilles genome-scale CRISPR screens in cancer cell lines. bioRxiv 720243 https://doi.org/10.1101/720243 (2019).
Li, B. et al. Cumulus provides cloud-based data analysis for large-scale single-cell and single-nucleus RNA-seq. Nat. Methods 17, 793–798 (2020).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Germain, P.-L., Lun, A., Garcia Meixide, C., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res 10, 979 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Domínguez Conde, C. et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, eabl5197 (2022).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Tickle, T., Georgescu, C., Brown, M. & Haas, B. inferCNV of the Trinity CTAT Project. 2019. Klarman Cell Observatory, Broad Institute of MIT.
Crowell, H. L. et al. Muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data. Nat. Commun. 11, 6077 (2020).
Chafamo, D., Shanmugam, V. & Tokcan, N. C-ziptf: stable tensor factorization for zero-inflated multi-dimensional genomics data. BMC Bioinforma. 25, 323 (2024).
Tweedie, S. et al. Genenames.org: The HGNC and VGNC resources in 2021. Nucleic Acids Res. 49, D939–D946 (2021).
Fang, Z., Liu, X. & Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 39, btac757 (2023).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-Seq data. Nat. Commun. 13, 3224 (2022).
Türei, D., Korcsmáros, T. & Saez-Rodriguez, J. OmniPath: guidelines and gateway for literature-curated signaling pathway resources. Nat. Methods 13, 966–967 (2016).
Cabello-Aguilar, S. et al. SingleCellSignalR: inference of intercellular networks from single-cell transcriptomics. Nucleic Acids Res. 48, e55 (2020).
Hou, R., Denisenko, E., Ong, H. T., Ramilowski, J. A. & Forrest, A. R. R. Predicting cell-to-cell communication networks using NATMI. Nat. Commun. 11, 5011 (2020).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. bioRxiv 2020.05.07.082750 https://doi.org/10.1101/2020.05.07.082750 (2020).
Cable, D. M. et al. Cell type-specific inference of differential expression in spatial transcriptomics. bioRxiv 2021.12.26.474183 https://doi.org/10.1101/2021.12.26.474183 (2022).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
DepMap, B. DepMap 22Q2 Public. https://doi.org/10.6084/M9.FIGSHARE.19700056.V2 (2022).
Chafamo, D., Shanmugam, V. & Tokcan, N. Genome-scale spatial mapping of the Hodgkin lymphoma microenvironment identifies tumor cell survival factors vigneshshanmug/chl_analysis: CHL manuscript analysis. (Zenodo, 2025). https://doi.org/10.5281/ZENODO.17094525.
Acknowledgements
This work was supported by funding from the Damon Runyon Cancer Research Foundation (PST: 46-24), the Lymphoma Research Foundation (V.S.), the Eric and Wendy Schmidt Center at the Broad Institute of MIT and Harvard (N.T.), and the National Cancer Institute R35CA242457 (T.R.G.). V.S. also thanks Dr. Jon Aster and Dr. Kathleen Burns for their mentorship and the Brigham and Women’s Hospital Department of Pathology and the Dana-Farber Cancer Institute Department of Oncologic Pathology for generously supporting his research training. We would also like to thank Catherine Wu and J. Bryan Iorgulescu for valuable discussions and conceptual and analytical input on spatial mapping of EBV-specific transcripts in SlideseqV2 datasets. We also thank the Nanostring Spatial Molecular Imager Technology Access Program team, particularly Emily Killingbeck and Youngmi Kim, for help with generating the CosMx dataset.
Author information
Authors and Affiliations
Contributions
V.S., F.C., and T.R.G. conceived the study with input from A.L., M.S., and S.R. V.S. performed experiments with help from S.S., S.H., I.B., J.W., H.M., N.N., E.M., and G.N. V.S., D.C., and N.T. performed analyses with help from M.B., C.U., D.C., J.W., and O.A. G.P. performed key immunohistochemical studies. M.S. and A.L. contributed key samples. V.S. and T.R.G. wrote the paper with contributions from all the authors.
Corresponding author
Ethics declarations
Competing interests
S.J.R. receives research support from Bristol Myers Squibb, Coherus, and KITE/Gilead. He is a member of the Scientific Advisory Board of Immunitas Therapeutics. F.C. is an academic founder of Curio Biosciences and Doppler Biosciences and a scientific advisor for Amber Bio. F.C.‘s interests were reviewed and managed by the Broad Institute per its conflict-of-interest policies. E.Z.M. is a founder of Curio Bioscience, Inc. M.A.S. has received research funding from Bristol Myers Squibb, AstraZeneca, Bayer, AbbVie, Genentech, and Novartis, and has served on advisory boards for Bristol Myers Squibb and AstraZeneca. T.R.G. is a paid advisor and/or equity holder in Dewpoint Therapeutics, Sherlock Biosciences, Amplifyer Bio, and Braidwell, Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zongming Ma and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shanmugam, V., Tokcan, N., Chafamo, D. et al. Genome-scale spatial mapping of the Hodgkin lymphoma microenvironment identifies tumor cell survival factors. Nat Commun 17, 838 (2026). https://doi.org/10.1038/s41467-025-67539-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67539-1