Abstract
For the MHC, MR1 and CD1 systems, antigen recognition involves contact of the membrane distal surfaces of both the αβ T cell receptor (TCR) and the antigen-presenting molecule. Whether other antigen display mechanisms by antigen-presenting molecules operate remains unknown. Here, we report mass spectrometry analyses of endogenous lipids captured by CD1c when bound to an autoreactive αβ TCR. CD1c binds twenty-six lipid species with bulky headgroups that cannot fit within the tight TCR-CD1c interface. We determined the crystal structures of CD1c presenting several gangliosides, revealing a general mechanism whereby two lipids, rather than one, are bound in the CD1c cleft. Bulky lipids are oriented sideways so that their polar headgroups protrude laterally through a side portal of the CD1c molecule - an evolutionarily conserved structural feature. The sideways-presented ganglioside headgroups do not hinder TCR binding and so represent a mechanism that allows autoreactive TCR recognition of CD1c. In addition, ex vivo studies showed that the sideways-presented gangliosides can also represent TCR recognition determinants. These findings reveal that CD1c simultaneously presents two lipid antigens from the top and side of its cleft, a general mechanism that differs markedly from other antigen-presenting molecules.
Similar content being viewed by others
Introduction
αβ T cell activation relies on T cell receptor (TCR) contact with antigens bound to antigen-presenting molecules. While Major Histocompatibility Complex (MHC) molecules, the MHC class I related molecule (MR1), and CD1 molecules present peptides, metabolites, and lipids, respectively, the positioning of αβ TCRs relative to antigen-presenting molecules is generally conserved across these three systems1. The membrane distal surface of the αβ TCR heterodimer contacts the membrane distal surface of the antigen-presenting molecule, along with antigen protruding from the antigen-binding cleft2,3,4,5. Accordingly, conventional depictions of this aligned, end-to-end contact mechanism show the antigen-presenting molecule below the αβ TCR, and antigens protrude ‘upwards’ for direct αβ TCR contact. For over three decades, this central end-to-end binding concept has shaped the general understanding of what governs αβ TCR specificity for antigen and antigen-presenting molecules.
The other conserved feature of antigen display to T cells by MHC, MR1 and CD1 is 1:1 stoichiometry of antigen bound to antigen-presenting molecules6. Recently, an exception was identified, where each human CD1b protein binds two lipid ligands, one on top of the other, so that the lower lipid acts as an inert scaffold and the upper lipid protrudes upward to act as the antigen contacting TCRs sitting ‘atop’ of CD1b7,8. Among the four human CD1 antigen presenting molecules, CD1c likely has unique immunological functions, as it is expressed on subsets of dendritic cells (DC) in lymphoid tissues, marks marginal zone B cells, and can be found on malignant lymphocytes9,10,11. Yet among human CD1 antigen-presenting molecules, the least is known about the display mechanisms of CD1c12,13,14,15,16. To expand understanding of lipid display by CD1c to TCRs, here we analyse cellular lipids trapped between CD1c and an αβ TCR and discover that CD1c uses a distinct antigen-display mechanism. Wherein, CD1c binds two orthogonally positioned lipids with distinct structural motifs that protrude through distinct portals on the top and side of the cleft to control TCR binding and response. These findings suggest a previously unrecognised pathway for immune regulation via CD1c-lipid-TCR axis.
Results
Trapping ligands between CD1c and the 3C8 TCR
To understand self-lipid display by CD1c, we sought to trap and analyse endogenous lipids in CD1c-lipid-3C8 TCR complexes, taking advantage of a new lipidomics platform17. Secreted CD1c-lipid complexes from HEK293T cells and recombinant 3C8 TCR protein underwent size exclusion chromatography to separate CD1c-lipid complexes that do or do not bind the 3C8 TCR. After eluting lipids in organic solvents, time-of-flight mass spectrometry (TOF-MS) detected lipids that promote or block TCR binding18,19. Recent implementation of sensitive MS methods now allows lipidome-scale detection17, yielding complex patterns of lipid ligands with 630 unique ion chromatograms detected across 11 fractions, with each ‘molecular event’ corresponding to a lipid defined by m/z, retention time and intensity values (Fig. 1).
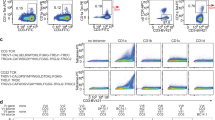
a Size-exclusion profiles of CD1c plus TCR show that complexes comigrating with CD1c monomers (fractions 37-39), and a biphasic curve with weakly (fractions 33-35) and strongly (fractions 26-31) excluded complexes. b, c Targeted MS analysis of lipids strongly and weakly excluded identified small monoacyl ligands, and a dual chain weak TCR ligator, sphingomyelin, respectively. d–f Whereas the single chain headless lipid, monoacylglycerol (MAG), binds inside CD1c to allow a tight CD1c-TCR interface that surrounds the F′ antigen exit portal (olive), lipidomics analysis of weak ligators of CD1c-TCR identified 26 dual chain lipids with large headgroups (red), whose mechanism of CD1c binding was unknown. g Unsupervised clustering analysis of ligands shows strongly excluded fractions are headless single chain lipids, weakly excluded fractions are dual chain lipids with large headgroups. Elutional lipidomics was performed twice with similar results.
Fractions 37–39 of CD1c-TCR mixtures contained proteins that comigrated with separately added CD1c monomers, so these TCR-unbound CD1c proteins likely contained non-antigenic lipids (Fig. 1a). Unexpectedly, early eluting CD1c-TCR complexes showed a biphasic response with strongly (fractions 26–31) and weakly (fractions 33–35) excluded complexes, which likely contained lipids that promoted durable versus intermittent CD1c binding to the TCR, respectively. To test this interpretation, we scanned fractions 26-31 for m/z values of two known antigens, C18:1 monoacylglycerol and C16:0 fatty acid15, finding both, which directly demonstrated antigen trapping by CD1c-TCR complexes (Fig. 1b). Although weakly excluded fractions 33–35 were expected to contain weak T cell agonists, targeted analysis instead detected C42:2 sphingomyelin (42:2 SM), which is a known TCR blocker for CD1a20 and CD1d21,22 (Fig. 1c). This result was unexpected because prior work showed that the 3C8 TCR docked flush to CD1c, fully surrounding the F’-portal15 (Fig. 1d), which predicts that the phosphocholine unit of 42:2 SM cannot fit between CD1c and the TCR, and it was known that shorter length SM (34:1) could act as a blocker of autoreactive CD1c-restricted TCRs15.
Lipidome-scale analysis
To address this apparent contradiction, unbiased analysis sought additional lipids from weakly excluded CD1c fractions that might identify ligand motifs. From 630 total CD1c-associated events, we censored alternate adducts and isotopes and focused on events that peaked in weakly excluded fractions that showed mass greater than that of C34:1 SM (molecular weight 702) (Fig. 1e), yielding 26 events (Supplementary Fig. 1) solved by collisional mass spectrometry (CID-MS) (Supplementary Fig. 2). After demonstrating a lack of non-specific binding to TCR alone and confirmation of peak capture in early excluded fractions (Fig. 1e and Supplementary Fig. 3), this analysis detected dual chain lipids with bulky headgroups, including three additional SMs and twenty-two other lipids in seven classes: ganglioside GM3 (GM3), ganglioside GM2 (GM2), hexosylceramide (hexCer), sulfatide, phosphatidylserine (PS), phosphatidylcholine (PC), and phosphatidylinositol (PI) (Fig. 1f).
All of these lipid headgroups are considered bulky in the sense that, unlike single chain headless antigens, they all contain one or more glycans or charged inorganic moieties and so were considered unlikely to pass the tight CD1c-3C8 interface. Yet, GM2 and GM3 were notable for particularly large headgroups comprised of tri- or tetra-saccharides. To quantitate each lipid species, we obtained standards for seven of the eight classes, and using GM3 as a surrogate for GM2, generated curves under electrospray conditions used to measure CD1c eluents (Supplementary Fig. 4a, b). While the standards did not replicate all of molecular variations in natural molecules, the measured class-specific relative ionisation efficiency allowed MS intensity signals to be converted into estimated molar ratios for 26 lipids in eight classes (Supplementary Fig. 4c, d), finding that GM3 was the most abundant lipid in these weakly excluded CD1c samples and that, combined, gangliosides accounted for 54 % of lipid among these 26 ligands detected. Thus, lipids with bulky headgroups were nearly universally present in these fractions and gangliosides were dominant, consistent with possible roles in regulating T cell response.
Bulky sphingolipids are permissive to TCR binding
Thus, lipidomics outcomes supported two unexpected and general conclusions. First, strongly and weakly excluded complexes released lipids that clustered in two structural motifs (Fig. 1g): single chain headless molecules versus dual chain bulky lipids, which implies two modes of lipid antigen capture or display by CD1c. Second, the tight interface between CD1c-3C8 TCR15 (Fig. 1d) was inconsistent with bulky headgroup positioning at the membrane distal surface of CD1c as predicted by general CD1 presentation models23,24. Further functional testing highlighted this conundrum. To investigate the effect of bulky headgroup lipids on CD1c-TCR recognition ex vivo, the SCARB1-deficient HEK293T cells lacking the SR-B1 activity that interfered with CD1c tetramers were used25. We further selected GD3, along with its closely related GM3, to investigate how the presence of an additional sialic acid influenced T cell recognition of CD1c. The treatment of CD1c tetramers with bulky gangliosides GM3 or GD3 failed to suppress the autoreactive recognition of CD1c with 3C8 TCR transfected cells, unlike that of the antigen-specific CD1b-restricted TCR control24 (Fig. 2a). Also, surface plasmon resonance (SPR) found similar steady state affinities (KD) of 3C8 TCR towards CD1c with endogenous lipids (CD1c-endo), CD1c-MAG and CD1c-GD3, as though lipids with absent, small or large headgroups all allow CD1c to bind the 3C8 TCR similarly (Fig. 2b).
a FACS plots depicting HEK293T.SCARB1-/- cells with surface-expressing 3C8 TCR (upper) and CD1b-restricted GEM42 TCR (lower) stained with CD1c tetramers with no added ligand (CD1c-endo), the indicated ligand or streptavidin-PE (SAv-PE) showing mean fluorescence intensities (MFI). b Surface plasmon resonance (SPR) sensorgrams of the 3C8 TCR bound to the CD1c-lipid complexes show steady state affinities (KD). Error bars indicate mean ± S.D. of n = 4 or 5 independent experiments. c Left, Structure of CD1c presenting mycobacterial lipid PM. Right, surface representation demonstrates two major opening portals, F′ and G′. d–f Structure of CD1c presenting gangliosides GD3, GM3, and GM1, respectively. Left, Top-down view showing MAG (white) or MPM (light yellow) in the A′-pocket, with GD3 (yellow), GM3 (red), or GM1 (pink) in the F′-pocket. Centre, Chemical structure of lipid headgroup and side view showing the headgroups protruding out of the G′-portal of CD1c. Details of interaction between sugars in the headgroups and CD1c are indicated, with hydrogen bonds represented as green dashes. Right, Surface representation showing the shape complementarity of the protein with the headgroups, with CD1c amino acids contacting the ganglioside headgroups individually colour coded. Sugar abbreviations: Glc glucose, Gal galactose, Neu5Ac N-acetylneuraminic acid or sialic acid, GalNAc N-acetyl-galactosamine.
CD1c uses a sideways antigen-displaying mechanism
CD1c cleft architecture is defined by the A′ and F′-pockets that bind lipids and connect to the external solvent via D′/E′, G′-, and F′-portals13. The F′-portal allows antigen protrusion to the top of CD1c and other CD1 isoforms for TCR contact14,26. We crystallised foreign phosphomycoketide (PM) with CD1c (Fig. 2c, Supplementary Fig. 5a, and Supplementary Table 1), which resided in the A′-pocket in an upright position, similar to the seating of MAG in prior structures, but unlike MAG, PM did extend through the F′-portal of CD1c, analogous to that of the CD1c-mannosylphosphomycoketide (MPM) structure14. The F′-pocket also harboured one or two single chain spacer lipids, hinting that CD1c might normally bind more than one lipid.
To investigate how lipids with bulky headgroups are permissive for 3C8 TCR binding, we determined the structures of mock-treated CD1c, CD1c-GD3, CD1c-GM3, CD1c-MPM-GM1, and CD1c-MPM-GD3 (see methods and Supplementary Table 1 and Supplementary Fig. 5b). GD3 and GM3 differ by an additional sialic acid moiety in GD3 (Fig. 2d, e), whereas GM1 ganglioside matches the core structure of GM3, but contains two additional branched sugars (Fig. 2f). In the CD1c-GD3 and CD1c-GM3 structures, unbiased electron density corresponding to MAG was clearly visible in the A′-pocket, and was positioned similarly to that observed in previous 3C8 TCR-CD1c-endo ternary complex15, as well as MAG bound in the CD1c-mock structure (Supplementary Fig. 5b). MPM was built into the density in the A′-pocket of the CD1c-MPM-GM1 and CD1c-MPM-GD3 except for the flexible mannosyl headgroup as observed in other CD1c-mycoketide structures26.
Unlike other CD1 structures, where antigen headgroups protrude to solvent through the F′-portal27, no electron density was observed at the F′-portal in the CD1c-ganglioside structures. Instead, continuous unbiased electron density was observed that extended laterally across the F′-pocket and exited via a sideways G′-portal of CD1c (Fig. 2c–f, Supplementary Fig. 5c–f). Here, the two tails of the ceramide moiety (Fig. 2d–f) adopted the previously observed position occupied by spacer lipids13, and the G′-portal is shown here as an escape tunnel that allows large glycans to protrude to the surface of CD1c (Fig. 2b, Supplementary Fig. 5a, b). Namely, the two chains of extended laterally across the F′-pocket, where the amide linkage hydrogen bonded to Asp88 and Lys91 (Fig. 2d–f and Supplementary Fig. 5), which anchored the sphingosine chain on top of the fatty acyl chain in all structures (Fig. 2d–f, Supplementary Fig. 5, right column). The fatty acid was resolved up to 18 carbons in length. Therefore, with the C18 sphingosine chain, we modelled the ceramide tail length of 36:1.
Extending early work that defined the G′-portal13, the bulky headgroups of GD3 and GM3 protruded through the G′-portal (Fig. 2d, e), where the electron density for all sugars suggested a stable interaction with the outer surface of CD1c (Supplementary Fig. 5c, d, f). The branched sialic acid moiety of GM1 was not well resolved however, suggesting less stable contact with CD1c (Supplementary Fig. 5e). The bent headgroups of GD3 and GM3 wrapped around the extended loop (His84 – Phe94) of the α1-helix of CD1c and made a series of polar-mediated contacts with each sugar moiety (Fig. 2d, e). Here, the surface contour of the α1-helix exhibited high shape complementarity with the GD3 headgroup, where Gln87 and Tyr89 generate a concave curvature to position the galactose (Gal) and two neuraminic acids (Neu5Ac). GM3, which lacks the terminal Neu5Ac, formed a similar curvature, but without contacting Gln21 and Ser86 (Fig. 2d, e). The branched GM1 headgroup did not fully engage with CD1c and formed hydrogen bonds via N-acetyl-galactosamine (GalNAc) to Tyr89 (Fig. 2f). The sideways positioning of the gangliosides provided immediate structural insight into their permissive nature for 3C8 TCR binding, as they adopt an unprecedented display mechanism whereby the bulky headgroups protrude sideways from the G′-portal of CD1c, positioning them away from the 3C8 TCR binding site at the A′-roof of CD1c (Supplementary Fig. 6 and Supplementary Movie 1).
CD1c volume versus lipid length
After implementing criteria for demarcating G′-portal boundaries, new (Fig. 2) and prior CD1c structures13,14,15,26 provided data for volume estimates of the CD1c cleft8. The revised consensus estimate from all structures was 2050 ± 180 Å3, suggesting that CD1c accommodate ~ 50–54 CH2 units (C50-54). This value is ~ 1.4-fold larger than estimates for the CD1a and CD1d cleft (~C38-39)8, which normally bind one ligand with two alkyl chains ( ~ C38). Thus, since CD1a and CD1d bind one dual chain lipid8, the ~40 % larger cleft volume in CD1c is consistent with dual ligand binding in general. The measured volume approximately matches the overall length of one dual chain plus one single chain lipid, as observed in individual structures reported here (Fig. 2c–f).
The G′-portal is a unique feature of CD1c
Overall, CD1c possesses a distinctive binding pocket that exhibits a mismatch with lipid size, attributable to its flexible F′-pocket. Compared to other CD1 isoforms, and despite the similarity in the overall structure (Supplementary Fig. 7)12, the F′-pocket of CD1c is readily open, so that dual tail lipids can replace the two spacer lipids, and the large headgroup can protrude via the G′-portal (Figs. 2c and 3a). The G′-portal is formed by the arrangement of a structurally defined constellation of residues (Fig. 3a). The two α-helices of CD1c were remodelled upon accommodating these sideways ligands (Fig. 3b). Despite this, the dimensions of the G′-portal did not undergo large change, with its major axis (~16.0–17.2 Å) or minor axis (~10–10.6 Å) (Fig. 3c). Overall, CD1c structurally aligned closely with the other three human CD1 isoforms (Supplementary Fig. 7b). In contrast, while the smaller side chains of Ser90, Ser139, Ser143 and Leu147 make the solvent accessible G′-portal for CD1c, the corresponding positions in the other CD1 isoforms contain more bulky residues that block this G′-portal site (Fig. 3d). As such, comparing these structures to prior structures of other CD1 isoforms, the G′-portal is essentially an invariant and unique feature of CD1c. Amino acid sequence alignments of CD1C genes across 15 mammalian species (Supplementary Fig. 8 and Supplementary Table 2) showed ~ 98 % sequence identity among primates and 28–69 % identity in non-primates. Except for equine CD1c, conservation of the small residues that form the G′-portal were present in all species. Thus, this lateral portal is likely conserved throughout the evolution of placental mammals.
a Details at the G′-portal of CD1c showing the protrusion of the GD3 headgroup (yellow ball-and-stick). Amino acid side chains are depicted in space-filling format. The G′-portal is constituted by nine amino acids, including four large residues His84, Gln87, Tyr89, and Lys91, and five smaller residues that create the accessible entry point, i.e. Ser90, Leu140, Ser139, Ser143, and Leu147. b Structural alignment of CD1c-GD3 (teal) versus CD1c-mock (pink) showing the movements of the α-helices, with arrows depicting direction of motion from the CD1c-mock to CD1c-GD3 structures. Cα atoms of Asp86 and Leu147 are depicted in spheres with the corresponding colours. Lipids are depicted in ball-and-stick, with GD3 coloured in yellow and spacer lipids in CD1c-mock coloured in white. c Comparison of the G′-portal in the structures of CD1c-mock versus CD1c-GD3. The major axis is measured from Lys91 Cβ to Leu147 Cγ, and the minor axis is measured from His84 Cβ to Leu140 Cγ. d Side view to the G′-portal of CD1c in comparison with the CD1a (brown), CD1b (green) and CD1d (blue), represented as surfaces. The blocking residues of the corresponding G′-portal position in each CD1 isoform are individually coloured as footprint. e Top view to the F′-portal of all CD1 isoforms demonstrating the sideways presentation of CD1c while the same GD3 lipid (yellow spheres) is presented upright in the other isoforms. CD1d-GD3 was reported previously (PDB ID 3AU1)28. f Comparison of the CD1c G′-portal versus the open end of MHC II. CD1c and GD3 are in the same colour code as previous panels. The presented peptide in MHC II is depicted in green-cyan ball-and-sticks. Right, Equivalent position of the G′-portal in CD1c and the open end of MHC II. The structures are portrayed in surface representation. The structure of MHC II-peptide without superposition of CD1c-GD3 is placed at the right for reference. MHC II-peptide was reported previously (PDB ID 7T6I)52.
To ascertain if sideways antigen presentation occurs with other CD1 isoforms, we determined the structures of CD1a-GD3 and CD1b-GD3 (Supplementary Table 1) for comparison with the published CD1d-GD3 structure28 (Fig. 3e and Supplementary Fig. 9). For CD1a, CD1b and CD1d, the GD3 headgroup exited upwards through the F′-portal. Thus, the lateral protrusion mechanism is unique to CD1c, and can be compared to the open-ended cleft of MHC II that allows overhang by the ragged ends of peptides6. MHC II has a notch that allows peptides to protrude upward, whereas by comparison the ganglioside headgroups escape sideways through a fully formed portal in the side wall of CD1c (Fig. 3f). The MHC II notch and the CD1c G′-portal represent structural modifications that likely act as sizing mechanisms; the MHC II allows for escape of excess peptide, but glycans egressing from the G′-portal adhere to the outside of CD1c and approach the TCR recognition site, so might tune T cell responses.
Sideways presentation modulates recognition of upright lipids
The autoreactive 3C8 TCR interacted exclusively with CD1c, without co-contacting the headless MAG sequestered within the A′-pocket15. To establish whether the sideways-presented GD3 ganglioside influences TCR recognition of a headed lipid in the A′-pocket of CD1c, we focused on the DN6 TCR, which demonstrates specificity for CD1c presenting the mycobacterial PM, and is predicted to dock over the F′-portal of CD1c29,30. The crystal structure of CD1c-PM (Supplementary Table 1) confirmed the upward pointing nature of PM from the F′-portal (Fig. 2c). SPR analysis demonstrated that the DN6 TCR remained similarly reactive to CD1c complexes treated with PM and GD3, compared to PM alone, but the response units (Rmax) were two-fold diminished compared to CD1c presenting PM alone (Fig. 4a). Similarly, SPR analysis of another PM-specific TCR called 22.530 showed that, despite equivalent affinity for CD1c-PM and CD1c-PM-GD3, response units were ~ 3-fold lower with CD1c-PM-GD3 (Fig. 4a), suggesting that a sideways lipid impacted TCR binding in some manner. Similarly, after transducing the DN6 TCR into SKW-3 cells, SKW-3.DN6 bound CD1c-PM tetramers in the presence and absence of GD3, but not to CD1c-endo, and CD1c-GD3 (Fig. 4b, c), confirm the specificity of the T cell line for CD1c-PM and suggests that the G′-sideways-presented GD3 still allowed the TCR interaction with the A’-upright-presented PM. However, consistent with the SPR data, the CD1c-PM-GD3 exhibited reduced staining on SKW-3.DN6 TCR cells.
a Steady-state binding sensorgrams and equilibrium plots of DN6 (upper) and 22.5 (lower) TCRs against CD1c-PM with (blue) and without (red) the presence of GD3 using SPR. Error bars indicate mean ± S.D. of n = 3 independent experiments, with respective steady state affinities (KD) and Rmax values indicated in the equilibrium and sensorgram plots respectively. b Staining of DN6 TCR transduced SKW-3 cells against CD1c-endo, -PM, -PM-GD3, and -GD3 tetramers, with streptavidin-PE (SAv-PE) as negative control. Black-outlined gate shows tetramer-positive cells, with corresponding mean fluorescence intensities (MFI) indicated in each plot. c CD69 activation assay of DN6 transduced SKW-3 cells against plate-bound CD1c-lipids. Error bars indicate mean ± S.D. of n = 3 independent biological repeats. Two-sided t-test statistical analysis was performed between CD1c-PM and CD1c-PM-GD3, with the resulting p-value = 0.0012 (indicated with **). d Binding pocket comparison of CD1c-PM (copper-yellow, open conformation) and CD1c-MPM-GD3 (deep teal, closed conformation). e Structural alignment of CD1c-PM (open conformation) and CD1c-MPM-GD3 (closed conformation) showing the steadiness of the α2-helix and the movement of the α1 helix. Colour code is same as in (d). f Back view to the α1-helix showing the pivoting of the helix at Asn60 with and without the presence of GD3 ganglioside. Colour code is same as in d. Cα of Asn60 and Asp83 are depicted in spheres. g Details of the movement of Asp83 (spheres) in the presence of GD3 ganglioside which leading to the displacement of Phe72, Phe75, and Arg79 on the α1-helix.
Potentially explaining this downward tuning of response, superposition of CD1c-PM and CD1c-MPM-GD3 demonstrated interchangeable ligand-dependent conformational states that represent the ‘open’ and ‘closed’ conformers. In the ‘open’ CD1c-PM conformation the CD1c cleft volume was 2140 Å3, while the ‘closed’ cleft volume of CD1c-MPM-GD3 was reduced to 1995 Å3 (Fig. 4d). While the α2-helix remained relatively fixed (Fig. 4e), the α1-helix in the closed form pivoted 4.9°, as the GD3 headgroup brought the C-terminus of the α1-helix downward by 2.9 Å (Fig. 4f). This effect, in turn, displaced C-terminal region of the α1-helix that is critical for the binding of DN6 and 22.5 TCRs30 (Fig. 4g and Supplementary Movie 2). Thus, expanding prior work showing that CD1c is, in general terms, flexible, these data indicate that CD1c exists in a dynamic equilibrium between open and closed conformers where the DN6 and 22.5 TCRs favour binding the open conformer, whereas the presence of the bound GD3 promoted the closed conformer. This effect might limit TCR recognition and account for approximately half of the population in solution.
CD1c-ganglioside mediated T cell responses
We next sought to investigate whether the sideways presented gangliosides represented TCR recognition determinants (Supplementary Fig. 10a). Because CD1c tetramer+ cells are rare in healthy blood25, and CD1c-endo tetramer+ cells were present at similar frequencies to CD1c-GD3 or CD1c-GM3 tetramer+ cells (Fig. 5a), we FACS sort-enriched these cells from nine donors using GD3-loaded CD1c tetramers and expanded them in vitro. We then evaluated their antigen specificity toward distinct lipids on the basis of staining intensity. These expanded polyclonal T cells were enriched with CD1c tetramer+ cells, including those from both the αβ and γδ T cell lineages. Consistently across three donors, most T cells stained with both unloaded CD1c-endo tetramers as well as ganglioside-loaded CD1c tetramers, with mycobacterial PM loaded CD1c as a control (Fig. 5b), suggesting that most CD1c-ganglioside-reactive clones exhibit direct CD1c autoreactivity, in line with the sideways-presented headgroup locating distally to the TCR docking platform. Nonetheless, staining profiles demonstrated that the staining intensity of some populations were indeed two to three-fold enhanced by CD1c-ganglioside relative to CD1c-endo (across three donors) or regained with CD1c-ganglioside relative to CD1c-PM (donors 2 and 3) (Fig. 5b), consistent with a tuning effect on CD1c-directed responses.
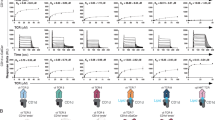
a (Left) Representative FACS plots showing CD1c tetramer staining of CD3+ T cells and CD3- lymphocytes. Cells were stained with CD1c tetramers loaded with PM, GD3, or GM3, or with unloaded CD1c tetramers (CD1c-endo). (Right) Scatter plot showing frequencies of CD1c tetramer+ T cells within the T cell populations from n = 20 PBMC samples. b Representative FACS plots showing in vitro expanded CD3+ T cells stained with CD1c tetramers loaded with the indicated lipids. The MFI of CD1c tetramers is shown next to each gate. c Representative contour plots showing CD1-lipid tetramer staining on HEK293T.SCARB1-/- cells transiently transfected to express CD1c restricted TCRs shown in Supplementary Fig. 8b, c or control TCRs, 22.5 (CD1c-PM reactive). MFI values of tetramer positive cells are shown in the top left of each plot, gated comparatively across all samples. The experiments were performed in two independent repeats. d Representative FACS plots showing F10 T cell clone stained with CD1c tetramers loaded with the indicated lipids. e CD69 expression and IFNγ production of F10 T cell clone after stimulation with bead-bound CD1c-lipid, which was pre-incubated with an anti-CD1c monoclonal antibody (0.5 μg/mL) or isotype control (0.5 μg/mL). Error bars indicate mean ± SEM of n = 3 independent biological repeats. CD69 expression data were normalised to relevant controls. Statistical analysis was performed using two-way ANOVA; *p < 0.05 (IFNγ: CD1c-GD3 with antibody vs isotype, 0.0190; CD1c-GM3 with antibody vs isotype, 0.0145; CD1c-endo isotype vs CD1c-GD3 isotype, 0.0238), **p < 0.01 (CD69: CD1c-endo isotype vs CD1c-GD3 isotype, 0.0045; IFNγ: CD1c-endo isotype vs CD1c-GM3 isotype, 0.0054), ***p < 0.001 (CD69: CD1c-GD3 with antibody vs isotype, 0.0005), and ****p < 0.0001 for the last two pairs.
TCR sequencing of these candidate T cell clones revealed a polyclonal TCR repertoire, enriched for TRBV4-1 for the αβ T cells and TRDV1 for the γδ T cells, confirming emergent TCR motifs for CD1c autoreactive TCRs (Supplementary Fig. 11a)25,31. TCRs from two such clones, one αβ (clone GL1) and one γδ (clone GL4) TCR, were sequenced and subsequently transfected for surface expression in SCARB1-deficient HEK293T cells and the cells subsequently stained with a panel of CD1c tetramers loaded with distinct lipids (Fig. 5c and Supplementary Fig. 11b–d). CD1c-endo reactivity of the GL1 TCR was largely unaffected by the presence of gangliosides GD3 or GM3, however, staining was reduced by PM. This staining was rescued by CD1c tetramers loaded with both PM and GD3, suggesting that the presence of the ganglioside is not the primary site of recognition but may fine tune PM-responsiveness. The GL4 TCR was largely unaffected when CD1c was loaded with distinct individual lipids, suggesting that the sideways-presented gangliosides are permissive to CD1c-autoreactivity. When tetramers were loaded with both PM and GD3, however, the staining intensity was approximately doubled relative to CD1c-endo, thereby suggesting that the co-presentation of PM and GD3 may represent an added molecular determinant for antigenicity. Overall, the sideways-presented lipids may have differing modulatory effects on CD1c-TCR interaction depending on the specific TCR involved, i.e. inhibitory effect as in DN6 and 22.5 TCRs (Fig. 4) or enhancing effect as in GL1 and GL4 TCRs (Fig. 5c).
We next examined CD1c-ganglioside tetramer reactivity in tonsils (Supplementary Fig. 10b), which are enriched in CD1c+ APCs compared to blood32,33. From three tonsil samples, dual CD1c-endo and CD1c-GD3 tetramer staining identified four αβ T cell lines with preferred reactivity towards CD1c-GD3 over CD1c-endo (Supplementary Fig. 11e). We sorted and expanded one CD4+ T cell clone (F10) (Supplementary Fig. 11f), which confirmed greater reactivity towards CD1c-GM3 and CD1c-GD3 over CD1c-endo, and then extended these analyses to a broader range of CD1c-restricted ligands (Fig. 5d). This confirmed lack of reactivity towards CD1c-endo and other ‘upward pointing’ antigens, including lysophosphatidylglycerol (LPG), lysophosphatidic acid (LPA) and PM. Yet specific reactivity towards GD3 and GM3 exists (Fig. 5d). As this F10 clone showed no reactivity towards GD1a, a branched ganglioside with two additional sugar headgroup moieties compared to GD3, this indicated that specificity determinants common to GD3/GM3 underlie this TCR-mediated recognition. The CD1c-ligand tetramer reactivity of the F10 T cell clone was also mirrored in activation assays, as measured by CD69 upregulation and IFN-γ production, which were blocked by an anti-CD1c mAb, thereby underscoring the CD1c-restricted nature of this T cell response (Fig. 5e). Accordingly, while the sideways presentation of gangliosides can lead to the formation of TCR recognition determinants, one role is simply lateral escape, which might lead to as yet discovered other lateral interactions, but, as observed here, commonly leads to not impeding the broader CD1c-restricted autoreactive TCR repertoire.
Discussion
Over four decades, structural studies on MHC-peptide, CD1-lipid and MR1-metabolite complexes provided foundational molecular insight into how antigens are displayed for T cell surveillance1. The general view is that antigens are anchored in 1:1 stoichiometry on antigen-presenting molecules and protrude upward toward TCR contact residues6. The dual upwards and sideways presentation by CD1c deviates fundamentally from this core paradigm. The 2:1 lipid:CD1c stoichiometry and sideways presentation is likely a general mechanism for CD1c, as large numbers of lipid ligands follow strong, weak or absent binding patterns; ganglioside loaded tetramers stain polyclonal T cells; simultaneous binding of one dual chain and one single chain lipid match the larger cleft volume of CD1c; and sequences that form the unique G′-portal are conserved across mammalian CD1C genes. These architectural features are not present in other CD1 isoforms, whereupon gangliosides are presented in the standard ‘upright’ mode and are generally considered to represent inert antigens or blockers CD1-restricted T cell responses12.
What are the implications of CD1c presenting gangliosides, or other lipid-based antigens in a sideways manner? Firstly, direct T cell reactivity towards CD1c is a common mode of action for this isoform15,34 and our results suggest that, in general, the CD1c-restricted autoreactive T cells will not be impeded by ganglioside binding presentation. Second, instead of being T cell recognition determinants, sideways presented gangliosides may perform a scaffolding role in stabilising dynamic CD1c plasticity and directing correct antigenic lipid presentation in the A’-pocket, similar to the role of scaffold lipids for CD1b8. Finally, the sideways presented ganglioside may represent a determinant for an innate receptor, occurring via a lateral approach in cis or trans from the antigen presenting cell or T cell, or an atypical alternate TCR binding site. These data do not predict simultaneous dual recognition by two TCRs, and steric considerations argue against this possibility. However, a more lateral TCR approach to antigen presenting molecules has been observed in αβ TCR binding to MHC II via a ‘tilted’ docking topology35. Furthermore, γδ TCRs are known to decorate MHC I like molecules in a range of docking modalities36,37, suggesting that sideways presented antigens might furnish specific ligands for γδ T cells.
Among the four human CD1 proteins with known antigen presenting function, CD1c is the least described in the literature. This situation does not imply lesser function, but more likely arises from two challenges in studying in vivo response to CD1c: lack of natural expression in mouse models and its intrinsic structural flexibility. The former issue is being addressed with better human and mouse transgenic models, and the latter, which impeded CD1c structural solution for a decade, is now overcome through the use of hybrid and wild type proteins14. Here, we show that structural flexibility is likely the physiological underpinning of its ligand-driven dual conformation mechanism of TCR recognition. Further, CD1c is not somehow less evolutionarily conserved in placental mammals38 or less frequently expressed in human tissues as compared to other isoforms. Instead, CD1c is a specific marker for DC and B cell subsets39, and is highly expressed in tonsils and other secondary immune tissues in health39 and autoimmune disease10. Moving past predictions of universal 1:1 binding stoichiometry8 but here we show how two antigens bind in orthogonal positions within CD1c. The CD1 ligand surveys and T cell responses establish the chemical identity of dozens of ligands, along with their antigenic classes defined as headless, headed or bulky, as well as the positioning, defined as ‘upright’ through the F’-portal or ‘sideways’ through the G’-portal as two simultaneously active modes of display. Now headless agonists (MAG, fatty acids), headed antigens (MPM, PM) and bulky laterally presented sphingolipids (GM3, GM1, GD3) and can be implemented in functional studies to establish whether CD1c-restricted T cell immunity involving self headless molecules or gangliosides plays a role in homoeostasis or disease, such as the nervous system or cancerous cells, where gangliosides are enriched40.
Methods
Protein expression and purification
Genes encoding human CD1c, CD1a, or hCD1d with human β2-microglobulin (β2m) were cloned into a single construct separated by a 2A self-cleavage peptide in the pHLSEC vector. The construct followed the sequence order of β2m-fos-2A-CD1c-jun, β2m-fos-2A-CD1a-jun, and β2m-fos-2A-hCD1d-jun with the fos/jun leucine zipper facilitating the post-translational association of the two proteins. Each gene was preceded by the mouse Igκ signal sequence to enable secretion of the heterodimer complex into the culture medium. The CD1 proteins were engineered with a hexa-histidine tag for affinity purification and an Avi-tag for biotin labelling. Two thrombin cleavage sites were introduced before the fos/jun sequences. The single-plasmid construct encoding CD1c, CD1a, or CD1d and β2m was transfected into Expi293F GntI(-) cells. For CD1b, a similar design was constructed for insect Trichoplusia ni High Five cell expression. The secreted CD1c-β2m, CD1a-β2m, CD1b-β2m and hCD1d-β2m heterodimer complexes (referred to hereafter as CD1c, CD1a, CD1b, and hCD1d) were harvested and subjected to a series of purification steps, including affinity chromatography, size-exclusion chromatography, and anion exchange chromatography, with purification validated by SDS-PAGE.
Genes encoding the α- and β-chains of DN6, 22.5, and 3C8 TCRs were cloned into the pET30a vector. For biotin labelling, the Avi-tag was added to the β-chain of DN6 TCR and the α-chain of 3C8 TCR. These constructs were transformed into BL21 (DE3) Escherichia coli cells. Protein expression was induced using 1 mM isopropyl β-D-1-thiogalactopyranoside (I2481C25, GoldBio), and expressed for four hours at 37 °C before harvesting by centrifugation. The inclusion bodies were collected and washed with a buffer containing 50 mM Tris, 100 mM NaCl, 1 % (v/v) Triton X-100, 2 mM dithiothreitol (DTT), 1 mM EDTA, and 0.5 mM phenylmethylsulfonyl fluoride (PMSF, P-470-50, GoldBio) at pH 8.0. The washed inclusion bodies were solubilized in a buffer comprising 50 mM Tris pH 8.0, 6 M guanidine hydrochloride, and 10 mM DTT. For refolding, 50 mg of each cognate α- and β-chain were gradually injected to a refolding buffer containing 50 mM Tris, 5 M urea, 440 mM L-arginine, 4 mM EDTA, 0.5 mM PMSF, 2 mM reduced glutathione, and 0.2 mM oxidised glutathione at pH 8.0. Refolding was carried out at 4 °C for 48 h. The refolded αβ heterodimers of each TCR were purified through sequential DEAE-cellulose anion exchange, strong anion exchange (HiTrapQ HP), size-exclusion chromatography, and hydrophobic interaction chromatography, with purification validated by SDS-PAGE. To improve the production yield of the 3C8 TCR, gene encoding for its α- and β-chains were separately re-constructed into pHLSEC vector for mammalian expression wherein the constant domains of both chains were further engineered with seven mutations described by Froning et al.41 and an extra mutation of Lys44Gln on the α-chain to assist the stabilised pairing between two chains. The purification of mammalian expressed 3C8 TCR followed the same method with that of CD1c protein.
TCR trapping of CD1c-lipid complexes
Recombinant 3C8 TCR (1.75 mg) was mixed with recombinant CD1c protein (1.75 mg), 3C8 TCR alone (100 µg) or CD1c protein alone (100 µg) in Tris buffered saline (TBS) pH 8.0 and run over Sephadex 200 16/600 size-exclusion column at 0.5 mL/min, using an AKTApurifier FPLC (GE). Approximately 2 µg of protein from protein containing fractions, based on UV elution profile, were analysed by SDS-PAGE for migration of TCR and CD1c species. In a separate experiment, 3C8 TCR alone (400 µg), CD1c protein alone (300 µg) or 3C8 TCR (6.5 mg) mixed with CD1c (4.5 mg) were run over two Sephadex 200 16/600 columns connected in series, at 0.5 mL/min. Fractions were collected and analysed by SDS-PAGE as per previous experiment. TCR trap experiments were completed twice with similar results.
Lipid elution and analysis
CD1c-lipid complexes were eluted of lipids with the Bligh and Dyer extraction42 that was adapted to small scale protein preparations17. After normalisation to 10 µM protein, fractions 26–28 and 29–30 were pooled due to low quantities within individual fractions. The injection volume was 10 µL for a reversed-phase HPLC column using an Agilent Poroshell EC-C18 column (1.9-micron, 3 × 50 mm) with a guard column (3 × 5 mm, 2.7 µm) using an Agilent 1260 HPLC system connected with an Agilent 6530 Accurate-Mass ESI-QToF mass spectrometer to acquire positive and negative ion mode data at a flow rate of 0.15 mL/min and the published gradient43. Intermediate eluting fractions (33-35) containing lipids allowing weak CD1c and 3C8 TCR interaction were compared to the late fractions (37-39) enriched for non-interacting CD1c and TCR monomers using Mass Hunter (Agilent) and the R package (version 3.4.2) XCMS44 for lipidomic peak analyses and in house designed software methods45. Lipid intensities are expressed as area-under-the-curve for identified lipids and were scaled and grouped by hierarchical clustering using Ward’s minimum variance method. Analysis and visualisation were performed in R.
Lipid identification
CID-MS analysis was performed on an Agilent 6546 Accurate Mass Q-TOF instrument and carried out with a collision energy of 15–80 V with the isolation width set to 1.3 m/z and comparing signals to recently generated lipidomic maps or to synthetic standards when necessary.
In vitro lipid loading
Monoacylglycerol 17:0 (MAG) was obtained from Sigma-Aldrich (SMB00506), while other lipids including gangliosides GD3, GM3, GM1, GD1a, and lysophosphatidylglycerol (LPG) and lysophosphatidic acid (LPA) were sourced from Avanti (860060, 860058, 860065, 86055, and 858125 and 857230, respectively). Phosphomycoketide (PM) and mannosyl β1-phosphomycoketide (MPM) were generously provided by Adriaan Minnaard (Groningen University). Lipids were solubilized via sonication in a buffer containing 10 mM HEPES, 150 mM NaCl, pH 7.5, and 0.5 % tyloxapol. For direct loading, CD1c protein containing endogenous lipids (CD1c-endo) was incubated with 0.5 % (w/v) tyloxapol (Sigma) in a buffer containing 10 mM HEPES and 150 mM NaCl, pH 7.5, at room temperature for 16 h. The tyloxapol-treated CD1c protein (CD1c-mock) was subsequently incubated with PM or MPM at a molar ratio of 1:10. For crystallisation, CD1c-endo was incubated with MAG at a molar ratio of 1:5 in the same buffer with 0.5 % tyloxapol at room temperature for 16 h. The CD1c-MAG complex was then further incubated with the desired lipids (PM, GD3, GM3, GM1, GD1a) at a molar ratio of 1:10 under the same conditions as the MAG displacement step. For optimisation of GM3 loading, GM3 was mixed with the GD3-loaded CD1c at a molar ratio of 10:1. For GM1 loading, yields were insufficient for crystallisation, so MPM was then used as a displacement medium before GM1 treatment. As a control, GD3-loaded CD1c was also mixed with MPM, with lipid-loaded CD1c samples validated using analytical MonoQ anion exchange chromatography and Phastgel iso-electric focusing (Cytiva). For CD1a and CD1b, GD3 loading used methods for CD1a-sphingomyelin 42:220 and CD1b-GMM46 complex formation, respectively. For tetramer staining that strictly required uniform biotinylation, biotin-labelled CD1c-endo was aliquoted into 5 μg vials at 20 μM. Subsequently, the desired lipid stocks (2000 μM) were added (200 μM), and the mixture was incubated at room temperature for 16 h and validated using iso-electric focusing.
Structural determination of CD1c-lipid complexes by X-ray crystallography
To facilitate crystallisation, the CD1c construct was engineered by replacing its α3 domain with that of CD1b, a method previously reported14 and utilised in our recent study26. The resulting chimeric construct was cloned into the pHLSEC vector, maintaining a design similar to the wild-type CD1c construct to enable comparable expression, purification, and lipid loading protocols as described above. After loading the desired lipids, the leucine zipper was cleaved from the CD1cchimeric-β2m heterodimer complex by thrombin (T6634; Sigma-Aldrich). Final size-exclusion chromatography was performed in a buffer containing 10 mM Tris and 150 mM NaCl at pH 8.0. The purified proteins were concentrated to 5 mg/ml and subjected to crystallisation using the hanging drop vapour diffusion method at 20 °C. The mother liquor contained 100 mM N-cyclohexyl-2-aminoethanesulfonic acid (CHES) at pH 9.4, 1.05 M trisodium citrate, and 25 mM triglycine. Crystals of CD1c-lipid complexes were cryoprotected using 50 % (w/v) sodium malonate and flash-frozen in liquid nitrogen. Crystals of CD1a-GD3 and CD1b-GD3 were obtained using the same hanging drop vapour diffusion method at 20 °C as previously described20,46. In brief, CD1a crystals were grown in a condition containing 21 % (w/v) PEG 1500, 20 mM DL-malic acid, 40 mM MES, 40 mM Tris, pH 5.5, whereas CD1b crystals were grown in a condition containing 22 % (w/v) PEG 3350 and 100 mM sodium iodide.
X-ray diffraction data were collected at the MX2 beamline of the Australian Synchrotron, part of ANSTO, and made use of the Australian Cancer Research Foundation (ACRF) detector. Data were processed using XDS47 for indexing, integration, and symmetry assignment, followed by scaling with Aimless48. Molecular replacement was performed using Phaser-MR49 with the high-resolution structure of CD1c-MPM1 (PDB ID: 7MX4) as the search model for CD1c data sets, CD1a-SM 42:2 (PDB ID: 7KP0) for CD1a-GD3 dataset, and CD1b-PC (PDB ID: 6D64) for CD1b-GD3. Structures were refined using Coot50 and phenix.refine51. The three-dimensional structures of CD1 with different lipid ligands were visualised in PyMOL, with the β-sheet of CD1c serving as a reference for structural alignment.
Surface plasmon resonance
Steady-state equilibrium affinity measurements for CD1c-lipids and TCRs were conducted at 25 °C using a BIAcore T200 instrument (Cytiva). The experiments utilized a running buffer containing 10 mM Tris and 150 mM NaCl at pH 8.0. Biotinylated CD1c-lipids were immobilised on streptavidin sensor chips to achieve approximately 2000 response units (RU) per flow cell, with hCD1d-endo serving as the reference control. TCRs, serving as analytes, were flowed over the sensor chip at 5 µL/min for 60 s, with the final response normalised by subtracting the response of the reference control. Serial dilutions were injected, with maximal concentrations of 100, 150, and 200 µM for DN6, 3C8, and 22.5 TCR, respectively. Affinity values and sensorgram plots were analysed and generated using BIAevaluation and GraphPad Prism software.
Human sample collection
Peripheral blood mononuclear cells (PBMC) were obtained from healthy human donors from the Australian Red Cross Blood Service after approval from the University of Melbourne Human Ethics Committee (1035100). Tonsils were collected from recurrent tonsillitis patients who consented to the use of their tissue for research at the John Radcliffe Hospital, Oxford, United Kingdom. Our study was reviewed and approved by the Oxford Radcliffe Biobank (ORB) Tissue Access Committee to obtain pseudonymised tissue samples and associated clinical data from patients recruited under ORB. All procedures were performed according to the Declaration of Helsinki guidelines, with all recruited volunteers provided written informed consent.
Tetramer staining
Biotinylated CD1c monomers were co-expressed with biotin ligase in Expi293F GntI(-) cells and purified, and the desired lipids were loaded using the direct loading method described above. For control (CD1c-mock), an equivalent volume of 0.5 % tyloxapol was added and incubated overnight at RT. The next day, monomers were tetramerized with streptavidin-PE (BD), and cells were typically stained with 1 μg/ml tetramers. PBMC or tonsil mononuclear cell (MNC) samples were first incubated with anti-CD36 antibody (5-271; BioLegend; 10 μg/mL, 15 min, RT), followed by incubation with tetramers for 30 min at RT. Subsequent staining with an antibody cocktail was carried out at 4 °C prior to acquisition on an LSRFortessa (BD) and analysis using FlowJo software. For transient TCR expression, full TCR sequences were synthesised and subcloned into the pMIG-II plasmid. Because HEK293T cells readily express SR-B1 protein, encoded by SCARB1 gene – a member of CD36 protein family that interferes with the CD1c tetramer and TCR interactions – we used SCARB1-defficient HEK293T cells for this transient TCR transfection purpose. SCARB1-deficient HEK293T cells were co-transfected with the TCR and CD3 expression plasmids using FuGene HD. At three days post-transfection, cells – as well as SKW-3 stable cell lines – were stained with the indicated antibodies and CD1 tetramers loaded with specific antigens. For functional assessment, CD1c tetramer-positive T cells from PBMCs were sorted into U-bottom 96-well plates pre-coated with anti-CD3 (UCHT1; 10 μg/mL; BioLegend) and anti-CD28 (Clone 28.2; 2 μg/mL; BD Biosciences) and cultured in the presence of rHuIL-2 (200 U/mL), rHuIL-7 (50 ng/mL), and rHuIL-15 (5 ng/mL). Cells are cultured in the complete medium consisting of RPMI-1640 (21870-076, Gibco) supplemented with 10 % foetal bovine serum (FBS) (JRH Biosciences), penicillin (100 U/mL) and streptomycin (100 μg/mL) (11074440001; Roche), GlutaMAX (35050-061; Gibco; 2 mM), sodium pyruvate (11360-070; Gibco; 1 mM), nonessential amino acids (11140-050; Gibco; 0.1 mM), HEPES buffer (15630-080; Gibco; 15 mM) (pH 7.2 to 7.5), and 2-mercaptoethanol (50 μM; Sigma-Aldrich). After two days, cells were transferred to new wells and co-incubated with 75 K irradiated CD1c K562 cells, allowing T cell proliferation for 10–14 days in complete RPMI medium supplemented with cytokines. CD1c tetramer-positive T cells from tonsil MNCs were sorted into U-bottom 96-well plates and cocultured with irradiated PBMCs, allowing T cell proliferation for 14-21 days in complete RPMI medium supplemented with 10% heat-inactivated human serum and rHuIL-2 (200 U/mL; BioLegend).
Bead-based activation assay
Biotinylated CD1c monomers loaded with the desired lipids were incubated with coreceptor antibodies (anti-human CD3 antibody, 130-093-377; anti-human CD28 antibody, 130-093-386; anti-human CD2 antibody, 130-093-376; Miltenyi Biotec) and magnetic beads (14011; ThermoFisher) overnight at room temperature. The next day, coated beads were incubated with 50 K expanded T cells in the complete RPMI medium supplemented with rHuIL-2 (25 U/mL; BioLegend) and anti-CD11a antibody (2.5 μg/mL; HI111; BioLegend) for at least 16 h. For CD1c blockade experiments, the beads were incubated in complete RPMI medium in the presence of anti-CD1c antibody (L161; BioLegend) or isotype control (Mouse IgG1, κ; BioLegend) at 0.5 μg/mL at room temperature for 1 h. T cells were stained with the indicated antibodies at 4 °C. The supernatant was harvested and the concentration of IFNγ was measured by ELISA (430116; BioLegend).
ELISA
IFNγ ELISA (430116; BioLegend) used capture antibodies diluted in coating buffer, which were incubated at room temperature for 2 h. The plates were then washed with PBS containing 0.1 % Tween-20 and blocked with assay diluent at room temperature for 1 h. Next, diluted supernatant samples and cytokine standards were added for 2 h. Following wash with PBS-Tween-20, detection antibodies were added and incubated for 1 h. Avidin conjugated with HRP was incubated for 30 min. The reaction was developed using TMB substrate solution (34022; ThermoFisher), and the reaction was stopped by adding the stop solution (S5814; Merck). Absorbance was measured at 450 nm using a ClARIOstar Plus reader (BMG Labtech).
Plate-bound CD69 activation assay
Purified CD1c-lipids were coated on flat bottom 96-well polystyrene plates (Corning) at 12.5 μg/mL in phosphate-buffered saline pH 7.4 (PBS) at 4 °C overnight. Wells were washed once using PBS and PM lipid was added to the final concentration of 2 μM followed by an incubation at 37 °C for 4 h. Approximately 100 K SKW-3.DN6 cells were cultured in the coated plate for 16 h before surface staining. Staining with an antibody cocktail was carried out at 4 °C before acquisition on an LSRFortessa (BD) and analysis using FlowJo software, in three independent biological replicates.
Single-cell TCR sequencing
In vitro expanded T cells were stained with a panel of CD1c-lipid tetramers to assess their antigen preference prior to single-cell sorting. T cell populations were selected for sequencing based on their higher staining with ganglioside-loaded CD1c tetramers compared to unloaded CD1c controls. TCR sequences were obtained in Fig. 5b as previously described25. Single T cells were then sorted individually into 96-well polymerase chain reaction (PCR) plates. Complementary DNA (cDNA) was synthesised using SuperScript VILO cDNA synthesis kit (Thermo Fisher Scientific). Each TCR chain fragments was then amplified through two rounds of semi-nested PCR using multiplexed primers. TCR amplicons were subsequently purified and subjected to Sanger sequencing. The resulting sequences were analysed with the IMGT/V-QUEST online tool.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The crystallographic datasets generated and analysed within the current study were deposited to the Protein Data Bank (PDB) under codes 9OHT (https://doi.org/10.2210/pdb9OHT/pdb, CD1c presenting GD3 ganglioside), 9OHU (https://doi.org/10.2210/pdb9OHU/pdb, CD1c presenting GM3 ganglioside), 9OHV (https://doi.org/10.2210/pdb9OHV/pdb, CD1c presenting dual lipids MPM and GD3 ganglioside), 9OHW (https://doi.org/10.2210/pdb9OHW/pdb, CD1c presenting GM1 ganglioside), 9OHX (https://doi.org/10.2210/pdb9OHX/pdb, CD1c presenting endogenous lipids), 9OHY (https://doi.org/10.2210/pdb9OHY/pdb, CD1c presenting phosphomycoketide in its open conformation), 9OHZ (https://doi.org/10.2210/pdb9OHZ/pdb, Crystal structure of CD1a presenting ganglioside GD3), and 9OI0 (https://doi.org/10.2210/pdb9OI0/pdb, Crystal structure of CD1b presenting ganglioside GD3). All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu Rev. Immunol. 33, 169–200 (2015).
Borg, N. A. et al. CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor. Nature 448, 44–49 (2007).
Garboczi, D. N. et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Garcia, K. C. et al. An alphabeta T cell receptor structure at 2.5A and its orientation in the TCR-MHC complex. Science 274, 209–219 (1996).
Patel, O. et al. Recognition of vitamin B metabolites by mucosal-associated invariant T cells. Nat. Commun. 4, 2142 (2013).
Adams, E. J. & Luoma, A. M. The adaptable major histocompatibility complex (MHC) fold: structure and function of nonclassical and MHC Class I–like molecules. Annu. Rev. Immunol. 31, 529–561 (2013).
Gadola, S. D. et al. Structure of human CD1b with bound ligands at 2.3A, a maze for alkyl chains. Nat. Immunol. 3, 721–726 (2002).
Huang, S. et al. CD1 lipidomes reveal lipid-binding motifs and size-based antigen-display mechanisms. Cell 186, 4583–4596 (2023).
Lepore, M. et al. A novel self-lipid antigen targets human T cells against CD1c+ leukemias. J. Exp. Med. 211, 1363–1377 (2014).
Roura-Mir, C. et al. CD1a and CD1c activate intrathyroidal T cells during graves’ disease and hashimoto’s thyroiditis. J. Immunol. 174, 3773–3780 (2005).
Villani, A. C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Cao, T. P. et al. A structural perspective of how T cell receptors recognize the CD1 family of lipid antigen-presenting molecules. J. Biol. Chem. 300, 107511 (2024).
Mansour, S. et al. Cholesteryl esters stabilize human CD1c conformations for recognition by self-reactive T cells. Proc. Natl. Acad. Sci. USA 113, E1266–E1275 (2016).
Scharf, L. et al. The 2.5Å structure of CD1c in complex with a mycobacterial lipid reveals an open groove ideally suited for diverse antigen presentation. Immunity 33, 853–862 (2010).
Wun, K. S. et al. T cell autoreactivity directed toward CD1c itself rather than toward carried self lipids. Nat. Immunol. 19, 397–406 (2018).
Szoke-Kovacs, R. et al. A CD1c lipid agnostic T cell receptor bispecific engager redirects T cells against CD1c(+) cells. Front Immunol. 16, 1614610 (2025).
Cheng, T. Y. et al. Lipidomic scanning of self-lipids identifies headless antigens for natural killer T cells. Proc. Natl. Acad. Sci. USA 121, e2321686121 (2024).
Brennan, P. J. et al. Structural determination of lipid antigens captured at the CD1d-T-cell receptor interface. Proc. Natl. Acad. Sci. USA 114, 8348–8353 (2017).
de Jong, A. et al. CD1a-autoreactive T cells recognize natural skin oils that function as headless antigens. Nat. Immunol. 15, 177–185 (2014).
Cotton, R. N. et al. CD1a selectively captures endogenous cellular lipids that broadly block T cell response. J. Exp. Med. 218, e20202699 (2021).
Melum, E. et al. Control of CD1d-restricted antigen presentation and inflammation by sphingomyelin. Nat. Immunol. 20, 1644–1655 (2019).
Rudolph, M. et al. Sortase A-cleavable CD1d identifies sphingomyelins as major class of CD1d-associated lipids. Front Immunol. 13, 897873 (2022).
Birkinshaw, R. W. et al. [alpha][beta] T cell antigen receptor recognition of CD1a presenting self lipid ligands. Nat. Immunol. 16, 258–266 (2015).
Van Rhijn, I. et al. A conserved human T cell population targets mycobacterial antigens presented by CD1b. Nat. Immunol. 14, 706–713 (2013).
Gherardin, N. A. et al. CD36 family members are TCR-independent ligands for CD1 antigen-presenting molecules. Sci. Immunol. 6, eabg4176 (2021).
Reijneveld, J. F. et al. Rational design of a hydrolysis-resistant mycobacterial phosphoglycolipid antigen presented by CD1c to T cells. J. Biol. Chem. 297, 101197 (2021).
Van Rhijn, I. & Moody, D. B. CD1 and mycobacterial lipids activate human T cells. Immunol. Rev. 264, 138–153 (2015).
Mallevaey, T. et al. A molecular basis for NKT cell recognition of CD1d-self-antigen. Immunity 34, 315–326 (2011).
Ly, D. et al. CD1c tetramers detect ex vivo T cell responses to processed phosphomycoketide antigens. J. Exp. Med. 210, 729–741 (2013).
Roy, S. et al. Molecular basis of mycobacterial lipid antigen presentation by CD1c and its recognition by αβ T cells. Proc. Natl. Acad. Sci. 111, E4648–E4657 (2014).
Guo, T. et al. A subset of human autoreactive CD1c-restricted T cells preferentially expresses TRBV4-1+TCRs. J. Immunol. 200, 500–511 (2017).
De Monte, A. et al. CD1c-related DCs that express CD207/langerin, but are distinguishable from langerhans cells, are consistently present in human tonsils. Front. Immunol. ume 7, 197 (2016).
Smith, M. E. F., Thomas, J. A. & Bodmer, W. F. CD1c antigens are present in normal and neoplastic B-cells. J. Pathol. 156, 169–177 (1988).
Gherardin, N. A. et al. Diversity of T cells restricted by the MHC class I-related molecule MR1 facilitates differential antigen recognition. Immunity 44, 32–45 (2016).
Hahn, M., Nicholson, M. J., Pyrdol, J. & Wucherpfennig, K. W. Unconventional topology of self peptide-major histocompatibility complex binding by a human autoimmune T cell receptor. Nat. Immunol. 6, 490–496 (2005).
Le Nours, J. et al. A class of γδ T cell receptors recognize the underside of the antigen-presenting molecule MR1. Science 366, 1522–1527 (2019).
Wegrecki, M. et al. Atypical sideways recognition of CD1a by autoreactive γδ T cell receptors. Nat. Commun. 13, 3872 (2022).
Van Rhijn, I., Godfrey, D. I., Rossjohn, J. & Moody, D. B. Lipid and small-molecule display by CD1 and MR1. Nat. Rev. Immunol. 15, 643–654 (2015).
Dougan SK, Kaser A, Blumberg RS. CD1 Expression on Antigen-Presenting Cells. In: T Cell Activation by CD1 and Lipid Antigens (ed Moody DB). (Springer, 2007).
Sarkar, A., Banerjee, S. & Biswas, K. Multi-dimensional role of gangliosides in modulating cancer hallmarks and their prospects in targeted cancer therapy. Front. Pharmacol. ume 14, 1282572 (2023).
Froning, K. et al. Computational stabilization of T cell receptors allows pairing with antibodies to form bispecifics. Nat. Commun. 11, 2330 (2020).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem Physiol. 37, 911–917 (1959).
van ‘t Klooster, J. S. et al. Periprotein lipidomes of Saccharomyces cerevisiae provide a flexible environment for conformational changes of membrane proteins. Elife 9, e57003 (2020).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Layre, E. et al. A comparative lipidomics platform for chemotaxonomic analysis of Mycobacterium tuberculosis. Chem. Biol. 18, 1537–1549 (2011).
Gras, S. et al. T cell receptor recognition of CD1b presenting a mycobacterial glycolipid. Nat. Commun. 7, 13257 (2016).
Kabsch, W. XDS. Acta Crystallogr. Sect. D. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution?. Acta Crystallogr. Sect. D. 69, 1204–1214 (2013).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. Sect. D. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. Sect. D. 68, 352–367 (2012).
Klobuch, S. et al. Human T cells recognize HLA-DP–bound peptides in two orientations. Proc. Natl. Acad. Sci. USA 119, e2214331119 (2022).
Acknowledgements
We thank the staff at the Australian Synchrotron MX beamline for assistance with data collection, and the Monash Macromolecular Crystallisation Platform. We thank Shin Yi Tin for technical assistance and Adriaan Minnaard for the PM and MPM reagents. We acknowledge the contribution to this study made by Christine Jesus, Eve Warner, David Maldonado-Perez and the rest of the team at the Oxford Centre for Histopathology Research and the Oxford Radcliffe Biobank, which are funded by the University of Oxford, the Oxford CRUK Cancer centre, the NIHR Oxford Biomedical Research Centre (BRC) (Molecular Diagnostics Theme/Multimodal Pathology Subtheme and the NIHR CRN Thames Valley network. This work was supported by NIH AR048632, AI 049313, AI 162584 (D.B.M.), the Wellcome Trust Discovery Award (302585/Z/23/Z, D.B.M., G.S.O., J.R.), the Australian Research Council DP210103064 (D.I.G., N.A.G.) and DE210101031 (A.S.), an NHMRC programme grant (1113293), NHMRC Emerging Leadership Award 2027104 (A.S.) and 2027058 (N.A.G.), NHMRC Investigator Awards to 2008981 (J.R.) 2008913 (D.I.G.), the UK Medical Research Council (G.S.O., Y.-L.C) and the Chinese Academy of Medical Sciences (CAMS) Innovation Fund for Medical Science (CIFMS), China (2024-I2M-2-001-1, G.S.O., Y.-L.C).
Author information
Authors and Affiliations
Contributions
T.-P.C. and G.-R.L. contributed equally, experimental design and primary data acquisition, performed analyses, solved structures and CD1c tetramer studies; T.-Y.C. and J.A.M. undertook mass spectrometry and bioinformatic analyses. Y.C., L.C., and R.F. solved CD1 binary structures, J.K. and E.Z.Q.N. assisted with research. T.S.F. and A.P.U. undertook TCR trap protein chemistry experiments. G.S.O., D.I.G., N.A.G., and Y.-L.C. supervised cellular immunology based experiments, provided reagents and funding. D.B.M., A.S., and J.R. are joint senior and corresponding authors, conceptualised alongside D.I.G., and supervised project, provided funding and resources, co-wrote the paper alongside T.-P.C. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have intellectual property though the Massachusetts General Brigham (D.B.M., G.S.O., J.R.) and the University of Oxford (G.S.O., Y.-L.C). G.S.O. and Y.-L.C. have relevant research collaborations with J&J Innovation. D.I.G. and N.A.G. hold PCT patents on therapeutic manipulation of CD1 family members. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Michael Birnbaum, Lawrence Stern, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, TP., Liao, GR., Cheng, TY. et al. Sideways lipid presentation by the antigen-presenting molecule CD1c. Nat Commun 17, 998 (2026). https://doi.org/10.1038/s41467-025-67732-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67732-2