Abstract
Designing proteins that bind with high affinity to hydrophilic protein target sites remains a challenging problem. Here we show that RFdiffusion can be conditioned to generate protein scaffolds that form geometrically matched extended β-sheets with target protein edge β-strands in which polar groups on the target are complemented with hydrogen bonding groups on the design. We use this approach to design binders against edge-strand target sites on KIT, PDGFRɑ, ALK-2, ALK-3, FCRL5, NRP1, and α-CTX, and obtain higher (pM to mid nM) affinities and success rates than unconditioned RFdiffusion. Despite sharing β-strand interactions, designs have high specificity, reflecting the precise customization of interacting β-strand geometry and additional designed binder-target interactions. A binder-KIT co-crystal structure is nearly identical to the design model, confirming the accuracy of the design approach. The ability to robustly generate binders to the hydrophilic interaction surfaces of exposed β-strands considerably increases the range of computational binder design.
Similar content being viewed by others
Introduction
There has been considerable recent progress in de novo protein binder design1,2,3,4,5,6. Both energy based1 and deep learning methods, such as RFdiffusion2, now enable design of protein binders given only the structure of the target of interest and (optionally) the specification of region of the target surface to bind3. Despite this progress, design of high affinity binders to hydrophilic regions of a target surface remains challenging since the exposed hydrogen bond donors and acceptors must be complemented with precisely positioned acceptors and donors on the designed binder to compensate for the loss of interactions with water. While RFdiffusion excels at generating backbones that are shape complementary to the targeted region of a protein surface, these solutions do not always provide detailed complementarity of hydrogen bonding donors and acceptors7. Many therapeutically relevant target proteins have β-sheets with unpaired and exposed β-strands; these often have non-canonical structures to reduce the tendency for aggregation8,9. β-strand targeted binders have been generated using pre-deep learning Rosetta methodology10, but this approach has limited ability to match the diversity of edge β-strand geometries. A generalizable deep learning method for designing β-strand pairing based binders to complement arbitrary target β-strand twists11, bends12, bulges13, and other irregular features14,15 could improve the design of binders to polar target surfaces.
Here, we demonstrate that RFdiffusion generation of binder backbone mediated hydrogen bonding interfaces with the target can yield designs with polar groups nearly perfectly complementing those on the target. We develop a general approach for guiding RFdiffusion denoising-diffusion trajectories towards such β-strand centric interfaces, and we experimentally validate this binder design approach.
β-strand interface conditioning improves binder designs
During training of RFdiffusion, a subset of training examples was provided with secondary structure and secondary structure block-adjacency (SS/ADJ) information; this was found to enable the conditioning of the model towards user defined protein monomer folds (as previously described by Watson et. al.)2. We explored whether providing interface conditioning information indicating a binder β-strand pairing with a target edge β-strand at the point of inference could yield designable strand pairing complexes. RFdiffusion takes as input desired residue secondary structures as well as an N by N block-adjacency matrix (N is the length of the designed protein), which specifies desired adjacencies between secondary structure blocks. These conditioning tensors can specify binder residues having desired secondary structure identities (helix, strand, loop, or masked) as well as the binder residue interaction identities (interacting, non-interacting, masked) with other binder residues (used for fold conditioning), or with target residues (used here for interface conditioning). Given the length of the designed interacting β-strand (L) and the identity of the target β-strand to be bound (T), RFdiffusion generates binders assigning a random set of L consecutive binder residues to form a β-strand pair with the target strand T, while the remainder of the binder output residues are not explicitly assigned a secondary structure or target interaction. We found that outputs from conditioned β-strand pairing runs indeed contained the user specified β-strand interfaces (Fig. 1a-b, Supplementary Fig. 1).
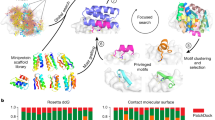
a Representation of β-strand interface conditioning information provided as a tensor to RFdiffusion to generate β-strand pairing binders. b Example RFdiffusion denoising trajectories without (top) and with (bottom) β-strand interface conditioning. Conditioning indicates that part of the binder scaffold should be a β-strand (gold) that contacts the indicated target edge strand (cyan). This information influences the denoising in very early trajectory timesteps (t), with the tertiary fold determined within 5 timesteps and the final output at t = 50. c Binder design success rates using β-strand interface conditioning or RFdiffusion default settings with hotspots indicating the target edge strand of interest. d Structural clustering of RFdiffusion output binder scaffolds using hotspot conditioning (left) or β-strand interface conditioning (right) t-distributed Stochastic Neighbor Embedding (t-SNE) transforms of all-by-all pairwise template modeling (TM) scores60 among binder scaffolds across all targets are plotted, with close proximity of points representing structural similarity. Output fold secondary structures are classified by color as indicated in the legend. Bold bordered data points indicate in silico successes (red circles, pAE interaction <10, pLDDT >85, ΔΔG < −30) and experimentally validated binders (black stars). Source data are provided as a Source Data file.
We supplied RFdiffusion with β-strand interface conditioning tensors for larger scale binder design campaigns against protein targets containing edge β-strands sufficiently exposed for binder access. To evaluate the generality of the approach, we selected targets that span a range of edge β-strand geometries. The seven selected targets also have considerable therapeutic relevance. Activin receptor-like kinases 2 and 3 (ALK-2 and ALK-3) are both Type I bone morphogenetic protein receptors that regulate growth of bone, vasculature, hair follicles, enamel as well as wound healing and tumor suppression in various soft tissues16,17,18,19,20. Platelet-derived growth factor receptor ɑ (PDGFRɑ) and Mast/stem-cell factor receptor (also known as KIT, SCFR, CD117) are both type III receptor tyrosine kinases that play roles in cardiomyocyte proliferation and heart tissue regeneration after myocardial infarction21,22,23,24,25,26,27,28,29. FCRL5 functions in critical signaling pathways for B-cell activation30. NRP1 is a coreceptor for various growth factor signaling pathways (TGF-1, EGF, VEGF, PI3K, HGF, and PDGF)31, a viral entry factor32, and plays a role in the of RAS/MAPK signaling in various cancers33,34. α-Cobratoxin (α-CTX), the lone non-receptor target, is a prominent toxin derived from elapid snakes35 that acts by blocking muscle and nerve acetylcholine signaling. All except α-CTX are therapeutic targets for different cancers36,37,38,39,40,41,42,43,44,45,46,47,48. Designed protein binders against these targets could be useful for antagonizing native signaling, targeting drug conjugates and other therapeutics to tumors, designing novel agonists49,50,51 and targeted receptor degradation52, inhibitors1, triggering cargo endocytosis52, or target therapeutics towards particular cell types53.
The set includes single β-sheet targets with highly exposed edge β-strands (α-CTX, FCRL5, NRP1) and somewhat occluded edge β-strands (ALK-2 and ALK-3), as well as immunoglobulin (Ig) fold β-sandwich domains (FCRL5, KIT, and PDGFRɑ). These were found to be challenging for binder design using standard hotspot-only conditioned RFdiffusion–either in silico design calculations had failed to generate designs predicted with high confidence to bind to the target, or experimental testing had failed to yield binders (ALK-3, KIT, see Data Availability). Because no experimentally determined structure exists for FCRL5, we designed binders against the AlphaFold 2 model54,55,56. To provide a stringent test of design specificity, the set includes some structurally related targets: ALK-2 and ALK-3 are members of the TGF-β superfamily57, and KIT and PDGFRɑ are type-III receptor-tyrosine kinases58 (Supplementary Fig. 2).
In a first in silico benchmark experiment, we compared the diversity and quality of binders designed using our interface conditioning method to designs generated using the standard RFdiffusion target site “hotspot” directed approach. Following the generation of 500 scaffolds using each diffusion approach, ten candidate binder sequences were generated for each scaffold by ProteinMPNN59 and each binder candidate complex was predicted with AlphaFold 254. Across all eight target sites across the seven target proteins (two sites were targeted for PDGFRɑ due to predicted structural variability), β-strand interface conditioning yielded design models with improved in silico binding metrics6 AlphaFold 2 interface predicted aligned error (pAE) and Rosetta ΔΔG (Fig. 1b), with over 9.2% of β-strand interface conditioned designs meeting reasonable quality metrics, in contrast to 0.98% success rates by RFdiffusion conditioned on target hotspots alone. β-strand interface conditioning yielded in silico design successes for all targets, whereas hotspot conditioning alone did not yield any in silico successes for two targets (ALK-3 and FCRL5). β-strand interface conditioning also yielded more globular binder designs with 88.7% of output scaffolds meeting radius of gyration criteria and only 25.5% of designs from the other methods meeting the same criteria (Supplementary Fig. 3). β-strand interface conditioning yielded outputs that covered a distinct protein structure space compared to other methods, as quantified by all-by-all pairwise template modeling (TM) scores60 between aligned binder scaffolds (Fig. 1c), with as expected a higher fraction of β-sheet containing binders and a decrease in ɑ-helical bundle outputs (Fig. 1d).
A second, much larger campaign was carried out for each target to generate designs for experimental testing. The sets were generated using β-strand targeted RFdiffusion and with standard hotspot conditioned RFdiffusion, followed by ProteinMPNN, and selection based on AlphaFold 2 and Rosetta metrics; partial diffusion scaffold optimization4,61 and additional sequence sampling was carried out to generate designs with improved metrics in several cases. β-strand pairing designs greatly outperformed other scaffold types in this in silico filtering stage for most targets with the exception of the PDGFRɑ target site, which was likely more amenable to helical bundle binder design success due to its flat, hydrophobic surface (Supplementary Fig. 2b). Structurally diverse designs with high AlphaFold 2 confidence (interface pAE <8 and pLDDT > 85) designs were selected for experimental screening by yeast surface display. While we ordered libraries including thousands of designs for some targets (KIT, PDGFRɑ, FCRL5, ALK-2, and ALK-3), binder success rates in our screen suggested that it would be possible to design strand pairing binders at smaller scales (Supplementary Table 1). Indeed, design sets for NRP1 and α-CTX yielded binders at 96 well scales. The α-CTX binder originally introduced by Vazques-Torres et. al., referred to in that work as LNG, will be referenced as α-CTX binding protein (α-CTXbp) in this work. α-CTXbp had been designed with the interface-conditioning method introduced here, but limited details on the method were revealed previously62. For PDGFRɑ, where both helical bundle designs and β-strand pairing designs passed in silico filters, significantly more binders were obtained with the strand directed approach than with standard RFdiffusion at this selection stage (Supplementary Fig. 4). The most enriched designs for binding affinity and expression from this assay were cloned, expressed in E. coli, and purified (Supplementary Fig. 5). SPR revealed mid- to sub-nanomolar affinity binders for our targets (Fig. 2a-c). In the following sections, we describe the active designs for each target in turn.
a Design models of binding complexes for each target. Left: Entire complex design models with binders indicated in green/gold and targets in teal/cyan. β-strand interface conditioned scaffold design yielded binders with interfacial β-strands (gold) forming strand-pairing hydrogen bonding interactions with target edge β-strands (cyan). Middle: Close up view of design model backbone hydrogen-bond interactions with putative hydrogen bonds shown in white. Right: schematic representation of strand pairing interactions showcase the diversity of sequence-independent β-strand pairing interactions. b Circular dichroism thermal melts. Full spectrum analyses (left) performed at 25 °C (cyan), 75 °C (green), and 95 °C (gold) assess the overall binder fold at these three temperatures, while single wavelength thermal melts (right) were measured at 217 nM to calculate binder Tm values. c SPR measurement (pink) of binding kinetics at 600pM, 4 nM, 30 nM, 200 nM, 1.5 μM, and 10 μM (association phases 1-6 on the X-axis). Fits for Kd determination (green) excluded the 10 μM data excluded due to signal aberrations at this high binder concentration. Binders were reproduced and similar SPR kinetics were fitted with n ≥ 2 for each binding protein. Source data are provided as a Source Data file.
TGF-β superfamily targets: ALK-2 and ALK-3
ALK-2 and ALK-3 are two of four known Type I Bone Morphogenetic Protein Receptors (BMPRI). Targeting BMPR therapeutics towards specific tissues and pathologies could have considerable utility, but achieving binding specificity for ALK-2 versus ALK-3 presents a difficult challenge due to their structural homology. The ALK-2 and ALK-3 ectodomains have 30% sequence identity and high structural similarity (Supplementary Fig. 2). On both targets, there is an edge β-strand with five consecutive residues with backbone atoms in very similar atomic positioning (RMSD 0.07), and each strand is partially occluded by an N-terminal coil. Prior to this study, multiple design campaigns—including campaigns utilizing RFdiffusion, ProteinMPNN, and AlphaFold 2—had failed to yield binders for ALK-3 (see Data Availability). The difficulty of designing binders to these targets is demonstrated in their lower success rates during our yeast surface display assay, with less than 1% of designs showing strong binding and enrichment (Supplementary Table 1).
We conditioned RFdiffusion on generating β-strand pairings to these five edge β-strand residues that were not occluded by the N-terminal coil. We identified binding proteins, ALK-2bp and ALK-3bp, with Kd values of 96 nM and 528 pM, respectively (Fig. 2c) by surface plasmon resonance (SPR). While both binders were intended to strand pair with the homologous target edge β-strand, the designed interaction footprints of their validated were quite distinct, with ALK-2bp forming a parallel β-strand interaction with the target β-strand and ALK-3bp pairing to the same strand in an antiparallel manner. ALK-3bp makes additional strand-pairing like contacts with the C-terminal coil of ALK-3 in a non-canonical hydrogen bonding pattern, with the binder β-strand curvature nearly perfectly complementing a bulge in the coil motif; these additional interactions may account for the four orders of magnitude greater binding affinity. This additional hydrogen bonding network with the C-terminal coil was not prespecified; RFdiffusion simply generated the secondary binder β-strand given the context of the initial β-strand motif specified by interface conditioning.
Single exposed edge β-strand: α-CTX, FCRL5, and NRP1
Three of the targets have more exposed edge β-strands. The acetylcholine receptor antagonist α-CTX is a three-finger toxin that consists of a single β-sheet with extended loops that bind a hydrophobic receptor pocket. We aimed to design a β-strand pairing binder that would sterically interfere with this interaction. FCRL5 weakly binds IgG to modulate B-cell activation. While the exact IgG binding site is unknown63, the receptor consists of several Ig domains. In the AlphaFold 2 model, the N-terminal Ig domain contains one three stranded sheet and one five stranded sheet, leaving an exposed β-strand available to target during design. Finally, we targeted the discoidin domain of NRP1, which contains a highly twisted β-sheet with a potential β-strand binding site where the edge β-strand twists about 90 degrees with exposed backbone polar atoms before entering into the protein core. While quite exposed, these target edge strands have distinctive structures that we hypothesized could allow for specific β-strand complementarity.
β-strand interface conditioned RFdiffusion resulted in a 5-fold improved in silico success rate for these targets compared to hotspot directed RFdiffusion which, even without conditioning, generated β-strand pairing binder designs with up to 20% of outputs (β-strand pairing designs were generated <5% of the time for the less exposed edge strand containing targets). Binders for α-CTX (originally introduced by Vazques-Torres, et. al.) and NRP1 (1.9 nM and 101 nM, respectively) were obtained by testing the 96 designs based solely on in silico metrics. The best binder to FCRL5, with a 82 nM Kd, was obtained from a yeast display library of 4841 designs. The FCLR5 and NRP1 target β-strands both had similar conformational twists, but FCLR5bp and NRP1bp utilize quite distinct antiparallel and parallel hydrogen bonding binding modes, respectively (Fig. 2a, right). Similar to FCLR5bp, α-CTXbp forms a mostly canonical antiparallel strand pair with a slight irregularity as the binder β-strand adapts to complement a small target strand bulge.
Type III receptor tyrosine kinases: KIT and PDGFRɑ
The type III RTK family receptors KIT and PDGFRɑ play roles in angiogenesis, tissue regeneration, and aberrant cancer signaling. Both receptor ectodomains are comprised of five Ig-like folds, with native ligands—stem cell factor and platelet derived growth factor—activating cellular signaling pathways by binding Ig domain 2 of KIT and PDGFRɑ, respectively, to induce receptor dimerization and intracellular cross-phosphorylation. Binders designed to occupy the ligand-binding pocket could act as antagonists to prevent aberrant signaling, and when oligomerized50, high-affinity binders could function as strong signaling agonists for tissue repair therapeutics. As such, we targeted the ligand binding sites on domain 2 of each receptor. As in the case of ALK-3, previous attempts to de novo design binders against KIT had failed (see Data Availability).
For KIT, a 65 nM binder (KITbp) was identified from a yeast surface library screening of 1298 designs. KITbp was designed to bind domain 3 strand on KIT which is part of the stem cell factor ligand binding site; the binding interface features an extensive canonical antiparallel hydrogen bond network, with 8 consecutive hydrogen bonds contributing a β-strand pair of 18 total binder and target residues (Fig. 2a, left). For PDGFRɑ, two binders were identified—PDGFRɑbp-7LBF (Kd = 137 pM) was designed to bind the cryo-EM structure (PDB accession code: 7LBF), while PDGFRɑbp-AF2model (Kd, 4 nM) was designed to bind the AlphaFold 2 model. They were enriched from 5427 and 189 member design libraries via yeast display, without any further experimental sequence optimization. Despite significant disagreement between the cryo-EM structure and the AlphaFold 2 model regarding the conformation of the target domain 2 Ig fold, design against both conformations yielded high-affinity binders. The binders may selectively induce the cognate PDGFRɑ conformation upon binding, as their design model conformations are not cross-compatible (Supplementary Fig. 7). PDGFRɑbp-7LBF forms a complex h-bond network in which the binder strand forms both parallel and antiparallel interactions with two different target strands of the target domain 2 Ig fold. This precisely complementary strand pairing h-bond network highlights the power of β-pairing conditioned RFdiffusion to design complex β-strand architectures to perfectly complement idiosyncratic target topologies.
Stability of binders with exposed edge β-strand interfaces
No obvious trends were observed between the size of the edge β-strand interface and aggregation propensity of the binders. All binders could be purified at high yields, and analysis of size exclusion chromatography curves reflects elution patterns consistent with monomeric binders being the most prominent purified species (Supplementary Fig. 5). Circular dichroism thermal melts (Fig. 2b) obtained for each binder indicate that the binder folds remain intact at high temperatures, even those with significant edge β-strand content. ALK-3bp was stable and monomeric up to 95 °C during a circular dichroism thermal melt experiment despite having four exposed edge β-strands. The binder with the second most β-strand content, ALK-2bp, was also thermostable with a measured TM of 81.2 °C. Binders with majority ɑ-helical support for their β-strand interfaces (KITbp, α-CTXbp, and PDGFRɑbp-AF2model) did not seem to be intrinsically more thermostable with measured TMs of 63.2 °C, 95 °C, and 95 °C, respectively. The binders with ɑ/β secondary structure content (2-3 β-strand sheets with 2 buttressing ɑ-helices; FCRL5bp, NRP1bp, and PDGFRɑbp-7LBF) had TMs of >95 °C, 77.2 °C, and 71.1 °C, respectively. The excellent solution behavior of the designs despite having edge β-strands clearly available for intermolecular interactions suggests that the same idiosyncratic features that enable high affinity and specificity (see below) target binding disfavor self-self interactions9.
β-strand pairing binder interfaces are target specific
We next investigated whether the identified binders were specific for their designed target and did not form off target high affinity β-strand pairing interactions. To test this, we performed an all-by-all SPR experiment where each binder was tested for binding affinity against each of the receptor targets in the test set. At 1.5 μM and 200 nM concentrations, all of the binders showed strong SPR response for their intended target receptor compared to off-target receptors (Fig. 3a). There was no evidence for off-target binding of the ALK-2, ALK-3, KIT, and PDGFRɑ binding proteins to related family members (e.g., ALK-2bp did not bind strongly to ALK-3). This high specificity may arise because the binders for each structurally similar target pair (i.e., ALK-2 and ALK-3, KIT and PDGFRɑ, FCRL5 and NRP1), were designed with different β-strand pairing hydrogen bonding arrays, and each binder makes additional non-strand-pairing contacts with the intended target. With the exception of ALK-3bp which forms exclusively β-strand contacts with ALK-3, all designed binders form additional ɑ-helical contacts that are complementary to neighboring target surface topologies. Overall, the interface side chain contacts resemble natural protein interfaces, with binder side chains making electrostatic and shape complementarity interactions with target side chains. The polar interactions made by strand-pairing lead to lower overall hydrophobicity, as measured by Spatial Aggregation Propensity (Supplementary Fig. 6b).
a Designed binder SPR response traces against all targets. For each binding protein, immobilized cognate target protein yielded the strongest SPR binding response at various binder concentrations (7-fold increasing binder concentrations ranging from 600pM to 1.5 μM, from left to right in each trace). Average response units for the (b) 1.5 μM and (c) 200 nM binder concentrations. Grid colors indicate response units relative to the maximum response for each binder. Source data are provided as a Source Data file.
KITbp:KIT complex structure confirms design model accuracy
To assess the accuracy of our design method, we solved the structure of KITbp in complex with domains 1–3 of KIT at 2.8 Å resolution (Fig. 4a, Supplementary Table 2). The crystal structure has near exact agreement with the computational design model with 1.9 Å all-atom RMSD between the design model and the structure. Over the designed binder alone, the RMSD over all atoms between the crystal and design is 2.0 Å, and over the backbone, 0.98 Å RMSD (Fig. 4b). The design model and crystal structure align with atomic level accuracy over nearly all interfacial side chain residue atoms. The KITbp binding site on KIT overlaps with that of the KIT native ligand stem cell factor (SCF), as all the binder designs targeted this site. Consistent with this, saturating concentrations of SCF (60μM) reduced KIT library binding sort counts by 99% in a yeast surface display binding assay (Supplementary Table 2).
a Crystal structure (PDB ID: 9H71) of KITbp (cyan) in complex with KIT (green) and 2Fo-Fc electron density contoured at +1.0 RMSD (gold). b The KITbp crystal structure superimposes on the binder design model (grey) with subangstrom backbone atom RMSD, and 2.0 Å all-atom RMSD. c Close-up views of the binder interface reveal high shape complementarity of the binder-target complex (top). The binder interface consists of hydrophobic and polar interactions between binder core-boundary residues and KIT receptor domain 2 core-boundary residues (middle). The binder forms an extensive hydrogen-bond β-pairing interaction with the targeted domain 2 edge strand (bottom). There are multiple side-chain interactions between binder and KIT tyrosine residues (Y38 and Y39 in the binder; Y259 and Y269 in KIT) in the core-boundary interaction, and between binder glutamate residues (E14, E28, E24, E35) and KIT hydrogen bond donors (Q256, K258, N260, W262, H263, R271). The binder also makes electrostatic interactions with both the opposite domain 2 edge strand and KIT domain 3. Source data are provided as a Source Data file.
The 29 residue binder solvent exposed surface area (SASA) of the binder interface (1164 Å2) consists of 8 strand-pairing binder residues (from Gly21 to Asp28, 291 Å2) and 21 additional interface residues (873 Å2). By heavy atom SASA, the polar atoms comprise 65% of the strand-pairing interface, a somewhat higher proportion compared to the rest of the interface surface, which was only 45% polar. There are 21 sidechain-sidechain and backbone-backbone hydrogen bonds between KITbp and KIT (calculated by HBPLUS64), many more than in previous designed binders (Supplementary Fig. 6) and in most native protein-protein interfaces with similar interface sizes65 (Fig. 4c). The eight strand pairing binder residues participate in nearly half (9) out of the total (21) hydrogen-bonds in the 29 residue binder interface. The majority of these hydrogen bonds were predicted accurately in our design model (16 out of 21). At the center of the interface, a network of four binder and KIT tyrosine residues (Y38, Y39, Y259, Y269) form a highly complementary interface between the binder interfacial helix and core-boundary of the KIT domain 2 Ig fold. Lining the perimeter of the interface are numerous polar interactions, including the designed β-pairing interface as well as several polar side-chain interactions.
KITbp contains an unpaired β-strand that pairs with the target, with three buttressing ɑ-helices that tether the β-strand in place. Foldseek66 alignment of KITbp against the PDB did not identify significant matches to known protein structures. We were unable to find examples of lone β-strands in natural proteins, except those found in protein-protein complexes where a loop in one partner forms a β-strand conformation that extends the β-sheet of the other partner67. A similar induction of β strand formation upon binding may occur with KITbp; Alphafold 2 prediction of the monomeric KITbp without KIT target places the β-strand pairing residues in a coil conformation that better shields the binder’s hydrophobic core residues and hydrogen bonding atoms of the β-strand interface (Supplementary Fig. 8); such KIT-dependent conformational switching may contribute to the observed binding specificity.
Biological functionality of designed binders
We next sought to assess the biological functionality of the designs. FCRL5 is internalized and transits through the endocytic degradation pathway upon binding to antibodies63, and we investigated whether FCRL5bp could be similarly internalized. We incubated cells expressing FCRL5 with FCRL5bp tagged with pHrodo DeepRed, a pH sensitive (~5 pKa) dye that emits 655 nm fluorescence at late endocytic vesicle pH, and observed binder internalization that correlated with FCRL5 expression levels, reaching a steady state in this expression system (Fig. 5a). We hypothesized that FCRL5bp could thus be useful as an Endotag52 for targeted protein degradation, as recently demonstrated for other designed proteins. We fused FCRL5bp to EGFRbp (EGFRn from Cao et al.) in both terminal orientations and compared the ability of this molecule to degrade EGFR by endocytic lysosomal trafficking induced by FCRL5 binding. After treating cells with 50 nM of either EGF, the binder fusions, or each binder alone, EGFR was robustly degraded in a manner dependent on doxycycline induced expression of FCRL5.
a HeLa cells engineered to express FCRL5 (HeLa-FCRL5) receptor in a Dox-inducible manner were treated with 50 nM neutravidin-labeled pHrodo DeepRed complexed FCRL5bp, and live cell imaging (top panels) was used to measure overlap of GFP (FCRL5-expressing cells) and pHrodo Red fluorescence (internalized FCRL5bp). 1000 ng/mL doxycycline results in strong GFP expression and intracellular pH activated pHrodo Red fluorescence. Internalization (measured arbitrary units calculated by overlap of red and green fluorescence divided by phase area) reached steady state in 18 hours (images are from this timepoint); values are presented as mean ± SD for n = 3 replicate wells. b, c EGFR is degraded HeLa-FCRL5 cells by de novo binders. Measuring EGFR degradation by immunostaining and Western blot (b) or flow cytometry (c); n = 3 independent biological replicates for western blots and flow cytometry. FCRL5bp fusion to EGFRbp resulted in comparable degradation to the native degradation mechanism (EGF signaling). Gating strategies for flow cytometry are provided in Supplementary Fig. 9. d, e Western blot analysis of PDGFRα inhibition in Chinese Hamster Ovary cells engineered to overexpress PDGFRα (repeated four times with similar results). Levels of phosphorylated PDGFRα, Erk, and Akt were measured by immunoblots with fluorescent antibodies. Signals were normalized by the fluorescent signal of an antibody against the constitutively expressed housekeeping proteins S6 or actin. Data are presented as mean ± SD of n = 3 independent biological replicates. Source data are provided as a Source Data file.
PDGFRɑbp-7LBF was designed to sterically occlude the binding site of the native ligand PDGF-AA, and indeed PDGFRɑbp-7LBF blocked activation of signaling by PDGF-AA through the Akt and Erk pathways with nanomolar inhibition of ERK phosphorylation (IC50 = 19.6 nM) and AKT phosphorylation (IC50 = 38.25 nM) in a CHO cell line engineered to stably express PDGFRɑ (Fig. 5d-e). ALK-2bp, ALK-3bp, and NRP1bp could not be tested for functional agonism in a straightforward manner, as the native ligands for ALK-2 and ALK-3 promiscuously activate signaling with other BMP receptors and NRP1 acts as a cofactor in signaling for various RTKs. In a companion study, α-CTXbp was found to potently neutralize α-CTX, protecting mice from a lethal neurotoxin challenge62.
Discussion
We show through in silico modeling and high throughput binding assays that β-strand interface conditioned RFdiffusion outperforms alpha-helical focused methods for the design of binders against seven edge β-strand containing targets. The binders are thermostable and target-specific, and the crystal structure of the KITbp:KIT complex shows the method has high structural accuracy. The higher success rate of binder design against polar targets containing β-sheets using the β-pairing targeted approach than with primarily ɑ-helical designs likely reflects the difficulty for the latter of complementing the many polar NH and CO groups on edge β-strands with designed sidechain-backbone hydrogen bonds–using geometrically matched β-strands this polar interface can be achieved in less convoluted fashion.
As noted by the Richardsons9, edge β strands of native proteins often have irregular geometries that they speculated functioned to suppress off-target β strand mediated interactions with themselves and with other proteins. The very high binding specificity and the lack of significant aggregation of our designs despite the availability of edge β-strands for intermolecular interactions supports the conjecture that non-canonical β strand geometries suppress off target pairing, and considerably extends the pre-deep learning observations of Sahtoe et al. in demonstrating that the irregular edge β-strands of native proteins can be systematically targeted provided that the geometry of the binder strand is sculpted to match the target.
Our β-strand targeted RFdiffusion approach should enable facile binder design against many previously challenging protein surfaces, including the many Ig domains and other β-sheet containing domains frequent in the extracellular domains of cell surface receptors. As many of these are current therapeutic targets, the method could contribute to new therapeutic interventions. While we acknowledge that the binders presented here may not be immediately feasible as therapeutic molecules, we expect that further optimization with these binders as starting points may improve their affinity, stability, pharmacokinetics, and other desired characteristics (e.g., mouse/human cross-reactivity, oligomerization, or protease stability) for therapeutic applications.
Methods
In silico comparison of RF diffusion-generated scaffolds
Binder scaffolds were generated with RFdiffusion conditioned with hotspots alone or with β-strand interface conditioning against the target edge β strand sites indicated throughout this work (Fig. 2, Supplementary Fig. 2). Interface conditioning tensors for this were pre-made with the script (see Data Availability) to ask RFdiffusion to create binder interface strands of equal length to the target edge β-strand. Scaffolds were compared by calculating the TM-score60 of each binder scaffold against all other binder scaffolds against all targets in the set. This data was transformed into a 2D-array with t-SNE transformation function in scikit-learn (version 0.24.2) to produce the graph in Fig. 2. After generation of ten sequences per scaffold with ProteinMPNN, sequence qualities were assessed with AlphaFold 2 initial guess6 and Rosetta FastRelax1.
Production scale computational protein design workflow
Target structures used as inputs for binder design were obtained from the Protein Data Bank for KIT (PDB:2E9W)68, NRP1 (PDB:2QQI)69, and PDGFRɑ (PDB:7LBF)70. Publicly available AlphaFold 2 models were used for the design of binders against ALK-2 (Uniprot:Q04771, Accessed October 2022), ALK-3 (Uniprot:P36894, Accessed October 2023), FCRL5 (Uniprot:Q96RD9, Accessed February 2023), and PDGFRɑ (Uniprot:P16234 Accessed April 2023). Target surfaces were assessed for surface hydrophobics and edge strands. Five to ten thousand binder backbone scaffolds complementing these sites were generated by standard RFDiffusion design with hotspot conditioning as well as β-strand interface conditioning. To condition RFdiffusion towards generating β-strand interfaces, the script introduced in this work (Data Availability) was used to create conditioning tensors that guide RFdiffusion toward generating interface strands of equal length to the target edge β-strand. ProteinMPNN sequences were generated for the output backbones and subject to in silico screening based on AlphaFold 2 initial guess (pAE interaction and binder pLDDT)6, AlphaFold 2 monomer pLDDT54, Rosetta ΔΔG1, radius of gyration (ROG), and spatial aggregation propensity (SAP)71. The cutoff values for these metrics varied somewhat for each target protein, but obtained minimum values of pAE interaction <10, pLDDT >85, ΔΔG < −30, a variable ROG filter adjusted for binder size length/4.5 = ROG, and SAP < 45 (more details can be found in Supplementary Table 1). In some cases, in silico optimization was carried out by iterative partial RFdiffusion and ProteinMPNN cycling72. These filters were used for in silico selection of top binder designs from hundreds of thousands of sequences, resulting in libraries ranging in size from 96 to 5000 that were screened experimentally. In silico and experimental library sizes depended on the in silico filter success rate, and the perceived difficulty of designing a binder interface based on target surface concavity, target surface hydrophobicity measured by SAP, and known previous experimental challenges (as for ALK-2, ALK-3, KIT, unpublished data). To ensure our binder design libraries sampled diverse sequences and structures, we clustered designs by sequence similarity or RMSD and selected a subset of designs such that each cluster was equally represented in the final library for experimental characterization.
DNA library preparation
For ALK-2, ALK-3, KIT, FCRL5, and PDGFRɑ, 104 scale DNA libraries were generated by using DNAworks2.0 reverse translating designed amino acid sequences that optimally reflected the Saccharomyces cerevisiae codon frequency table. Additional 5‘ and 3‘ adapters were included to enable PCR amplification of libraries by single sets of primers. All libraries were amplified using Kapa HiFi polymerase (Kapa Biosystems) with a qPCR machine (Bio-Rad, CFX96). In detail, the libraries were first amplified in a 25 μl reaction, and the PCR reaction was terminated when the reaction reached half maximum yield to avoid overamplification. The PCR product was loaded onto a DNA agarose gel. The band with the expected size was cut out, and DNA fragments were extracted using QIAquick kits (Qiagen). Then, the DNA product was re-amplified as before to generate enough DNA for yeast transformation. The final PCR product was cleaned up with a QIAquick Clean up kit (Qiagen). For the yeast transformation step, 2–3 µg of linearized modified pETcon vector (pETcon3) and 6 µg of insert were transformed into the EBY100 yeast strain using the lithium-acetate/PEG protocol of ref. 73.
DNA libraries for deep sequencing were prepared using the same PCR protocol, except the first step started from yeast plasmid prepared from 5 × 107 to 1 × 108 cells by Zymoprep (Zymo Research). Illumina adapters and 6-bp pool-specific barcodes were added in the second qPCR step. Gel extraction was used to obtain the final DNA product for sequencing. All the different sorting pools were sequenced using Illumina NextSeq sequencing.
Target protein preparation
Biotinylated target protein was commercially available for KIT (Acro Biosystems, CD7H5255), FCRL5 (Acro Biosystems, FC5-H82E3), NRP1 (Acro Biosystems, NR1-H82E3), α-CTX (Latoxan, L8114), and PDGFRɑ (Sino Biological, 10556-H27H-B). For ALK-2, ALK-3 biotinylated receptor was expressed as avi-tagged ectodomain constructs in E. coli, purified, and biotinylated as described by Tao Huang and Andrew P. Hinck74.
Yeast display
Saccharomyces cerevisiae EBY100 strain cultures were grown in C-Trp-Ura medium supplemented with 2% (w/v) glucose. For induction of expression, yeast cells were centrifuged at 4500 × g for 5 min and resuspended in SGCAA medium supplemented with 0.2% (w/v) glucose at the cell density of 1 × 107 cells per ml and induced at 30 °C for 16–22 h. Cells were washed with PBSF (PBS with 1% (w/v) BSA) and labeled with biotinylated targets using two labelling methods: with-avidity and without-avidity labeling. For the with-avidity method, the cells were incubated with biotinylated target, together with anti-c-Myc fluorescein isothiocyanate (FITC, Miltenyi Biotech) and streptavidin–phycoerythrin (SAPE, ThermoFisher). The concentration of SAPE in the with-avidity method was used at one-quarter of the concentration of the biotinylated targets. For the without-avidity method, the cells were first incubated with biotinylated targets, washed and secondarily labeled with SAPE and FITC. All the original libraries of de novo designs were sorted using the with-avidity method for the first few rounds of screening to exclude weak binder candidates, followed by several without-avidity sorts with different concentrations of biotinylated targets. Binder affinities were estimated from this titration data using the software provided (Data Availability) to estimate SC50, originally introduced by Cao et. al.
Protein expression and purification
Protein binder designs were ordered as synthetic genes (eBlocks, Integrated DNA Technologies) and cloned via BsaI overhangs into the target cloning vector, LM062775 for Golden Gate assembly. Golden Gate reaction mixtures were transformed into a chemically competent expression strain (BL21(DE3)), and overnight outgrowth cultures were used to seed 500 mL protein expression cultures in auto-induction media (autoclaved TBII media supplemented with Kanamycin, 2 mM MgSO4, 1×5052) and to propagate expression plasmid for isolation (QIAprep Spin Miniprep Kit) and sanger sequencing services performed by GENEWIZ (Azenta Life Sciences). The following day (20-24 hrs later), cells were harvested and lysed by sonication (QSonica Q500 Sonicator), and clarified lysates were purified by immobilized metal affinity chromatography using Ni-NTA agarose resin (Qiagen). Samples were eluted in a Tris elution buffer containing 300 mM imidazole, sterile filtered with 0.22μm Polyvinylidene Fluoride syringe filter prior to size exclusion chromatography. Protein designs were then screened via SEC using an AKTA FPLC outfitted with an autosampler capable of running samples from a 96-well source plate. The protein binders were run on a Superdex75 Increase 5/150 GL column (Cytiva 29148722) at room temperature.
Circular dichroism
Far-ultraviolet circular dichroism measurements were carried out with a JASCO-1500 instrument equipped with a temperature-controlled multi-cell holder. Wavelength scans were measured from 260 to 190 nm at 25 and 95 °C. Temperature melts monitored the dichroism signal at 217 nm in steps of 2 °C/min with 30 s of equilibration time. Wavelength scans and temperature melts were performed using 0.3 mg/ml protein in PBS buffer (20 mM NaPO4, 150 mM NaCl, pH 7.4) with a 1 mm path-length cuvette. Melting temperatures were determined by fitting the data with a sigmoid curve equation. For designs retained more than half of the mean residue ellipticity values, Tm values are reported as greater than 95 °C. Tm values of the other designs were determined as the inflection point of the fitted function.
Surface plasmon resonance measurement of binding kinetics
Binding kinetics were analyzed via Surface Plasmon Resonance (SPR) on a Biacore 8 K (Cytiva) operated at 25 °C with a data collection rate of 10 Hz. Receptor binding was measured by capturing biotinylated receptor ectodomains on a streptavidin-coated CAPture chip using the Biotin CAPture Kit (Cytiva #28920234). Biotinylated receptors (0.125 µg/mL) were injected for 100 s at a flow rate of 10 µL/min in HBS-EP⁺ buffer (0.01 M HEPES pH 7.4, 0.15 M NaCl, 3 mM EDTA, 0.005% v/v Surfactant P20; Cytiva #BR100669) to achieve capture levels of approximately 250 RU. Analytes were prepared by serial 1:7 dilution from 10 µM stocks in HBS-EP⁺ and injected at a flow rate of 30 µL/min to monitor association, followed by dissociation in running buffer (HBS-EP⁺) at the same flow rate. Single-cycle kinetics were performed by injecting increasing analyte concentrations sequentially over the same captured ligand surface (six concentrations per cycle, 100 s association, 60 s dissociation between each cycle, and 750 s final dissociation). Regeneration was achieved by two 60 s pulses of regeneration buffer at 10 µL/min. Blank-injection (buffer-only) subtraction was applied for all analyte injections, double referencing (reference-surface + blank) was performed. Sensorgrams were processed and fit in the Biacore Insight Evaluation Software using a 1:1 Langmuir binding model. An exception was made to use a heterogenous kinetic fit model for α-CTXbp as this was found in ref. 62 to better model the partially biphasic binding properties for the complex. Global fit was applied for kₐ, kd, Rmax, and tc parameters with default initial values 10e5, 10–e, Ymax, 10e8 initial values. Drift was fitted locally (and is reported in the kinetic parameters in the Source Data file), and RI was set to 0 as a constant. Fitting drift was capped at a maximum contribution 15%, where the drift contribution over the 750 s fit could not exceed 15% of the Rmax parameter (capping only performed for KITbp and FCRL5bp for optimal fitting values χ2 and SE, while other binders were affected). 10 μM data was excluded from the fit due to non-ideal solution behaviour, observable in the overlaid fits in Fig. 2.
Surface plasmon resonance to measure target specificity
All by all binder-target interactions were measured according to SPR protocol described above. Identical binder titres and receptor loading protocols were used to maintain as much signal consistency as possible, with binder titre values ranging in seven fold dilutions from 10 μM to 600pM to capture nonspecific binding across a large concentration spectrum. Non-specific binding responses were evaluated by averaging response values across the association phase for a given titre, normalizing responses such that all values were positive, and taking each response as a fraction of the maximum observed response (for each binder the maximum response was observed against the target protein for which it was designed).
Recombinant production of KITD1-D3 for X-ray crystallography
KIT123, i.e., extracellular domains D1–3 of the tyrosine kinase receptor KIT, was recombinantly produced via transient expression of suspension-adapted HEK 293 cells. Cells were grown and maintained in a 1:1 mixture of the Freestyle (Gibco) and the Ex-Cell (Merck) medium. Before the transfection, cells at the density of 1.5× 106 cells.mL–1 were centrifuged at 250 × g for 6 min and resuspended in the pre-warmed FreeStyle medium only to reach the density of 3.0 × 106 cells.mL–1 and incubated at 37 °C, 130 rpm, 70 % humidity, and 8.0 % CO2 for 15 min. The cells were subsequently added 450 µg of the plasmid DNA carrying the target construct per 100 mL of the medium. After 5 min, the cells were added 900 µg of linear polyethylenimine 25 kDa (Polysciences) and 3 µmol of kifunensine per 100 mL of the medium and continued incubation. After additional 5 h, an equal volume of the Ex-Cell medium to the FreeStyle medium was added to the cultures to return back to the density of ~1.5 × 106 cells.mL–1. 24 h post-transfection, the cells were added D-glucose and valproic acid to the resulting concentration of 55 mM and 3.5 mM, respectively. 96 hours post-transfection, the cells were harvested by centrifugation at 500 × g and 4 °C for 10 min and the conditioned medium (supernatant), carrying the recombinantly produced protein of interest, was collected, added 10,000 U of Endo Hf (NEB) per 100 mL of the medium, and incubated for 4 h at room temperature to remove heterogenous N-linked glycans and facilitate the subsequent crystallization attempts. After filtering the incubated medium through a 0.22 µm filter, the clarified sample was loaded to a 1 mL Ni-NTA HisTrap HP column (Cytiva) equilibrated with HEPES-buffered saline (HBS; 20 mM HEPES, 150 mM NaCl, pH 7.4). The column was washed with 10 mM imidazole in HBS and the His-tagged protein of interest was eluted using 150 mM imidazole in HBS. The eluted protein fraction was subsequently loaded to a Superdex 75 Increase 10/300 GL column (Cytiva) to simultaneously remove aggregates and remaining impurities and to exchange buffer to HBS with no imidazole. The fractions corresponding to the protein of interest were pooled together, their purity was analyzed by SDS-PAGE (Bio-Rad), and the concentration was determined using the NanoDrop Spectrophotometer (Thermo Fisher Scientific).
Crystal structure of KIT123 in complex with KITmb
The KIT123–KITbp complex was formed by adding a 3-fold molar excess of the purified KITbp to the recombinantly produced, EndoH-treated KIT123 receptor (domains D1–3 of the ectodomain). The complex was isolated using size-exclusion chromatography equipped with a Superdex 75 Increase 10/300 GL column (Cytiva) equilibrated with HEPES-buffered saline (HBS; 20 mM HEPES, 150 mM NaCl, pH 7.4). Fractions corresponding to the KIT123–KITbp complex were pooled and concentrated by centrifugal ultrafiltration to the concentration of 6.1 mg.ml–1. Sparse-matrix crystallization screens were carried out in 96-well 3-drop plates (Molecular Dimensions) using the BCS-Screen (Molecular Dimensions) at 293 K and the sitting-drop method. The vapour-diffusion geometry was used to set up sitting drops consisting of 100 nL of a protein solution and 100 nL of each reservoir solution using a Mosquito nanolitre crystallization robot (SPT Labtech). The protein complex crystallized in the condition G11 (0.2 M sodium/potassium phosphate pH 7.5, 0.1 M HEPES pH 7.5, 22.5 % PEG Smear Medium, 10 % glycerol). Crystals were cryo-protected with mother liquor supplemented with 25% v/v glycerol and subsequently flash-cooled by direct plunging into liquid nitrogen. X-ray diffraction data of protein crystals were collected at the P13 beamline (PETRA III, EMBL Hamburg). Obtained data were processed using XDS76 and severe data pathologies, including strong anisotropy and translational noncrystallographic symmetry, were revealed, yielding similar characteristics as reported previously77. Based on these findings, the data were elliptically-truncated and corrected using the STARANISO78,79 server and accordingly treated during the following steps. Initial phases were determined by maximum-likelihood molecular replacement in Phaser80 using the domains D1–3 part of the KIT-SCF structure (PDB ID: 2E9W)68 as a search model. Model (re)building was performed in Coot81, and coordinate and ADP refinement was performed in PHENIX82. Model and map validation tools in Coot, the PHENIX suite, and the PDB_REDO server83 were used to validate the quality of crystallographic models. Atomic coordinates and structure factors of the protein-protein complex were deposited in the Protein Data Bank under the PDB code 9H71.
In silico binder interface characterization
SAP and SASA metrics were calculated using PyRosetta and PyMOL software using the calculate_sap and get_area functions, respectively. ΔSAP and ΔSASA were obtained by calculating each metric for the binder before and after removing target proteins from complex models. To calculate the polar interface surface, the ΔSASA was calculated for each binder atom. The sum of oxygen and nitrogen ΔSASA values yielded the polar interface values, while all carbon residues were considered to be non-polar. Using the DSSP algorithm, binder-target strand pairing interactions were identified by counting β-strand conformation binder and target residues with backbone interactions less than 3.5 Å in distance. Strand pairing orientation (parallel or antiparallel) was determined by calculating the average position of the N-terminal and C-terminal halves of the binder and target strand-pairing residues. If the distance between the binder N-terminal strand contacts and the target N-terminal strand contacts was less than that of the distance to the target C-terminal strand contacts, the strand-pair was determined to be parallel. Otherwise, it was determined to be antiparallel. The software used to determine these metrics is provided in the Data Availability section.
PDGFRɑ antagonism assay
Heparan-deficient Chinese hamster ovary cells stably overexpressing PDGFRα (CHO-PDGFRα) were grown to 70-90% confluency in CHO growth medium (Kaighn’s Modification of Ham’s F12 (F12K) medium (ATCC# 30-2004) + 10% Fetal Bovine Serum (FBS) (Biowest, #S1620) + 4% Penicillin-Streptomycin (P/S) (Gibco, #15140122) with 10 μg/mL puromycin (Gibco, #A11138-03). The cells were starved for 4 h in serum free F12K media and treated with synthetic and/or native ligand for 15 min at 37 degrees Celsius. Cells were subsequently washed with PBS and lysed with buffer containing 20 mM Tris–HCl (Sigma-Aldrich, 1185-53-1) (pH 7.5), 150 mM NaCl, 15% glycerol (Sigma-Aldrich, G5516), 1% triton (Sigma-Aldrich, 9002-93-1), 3% SDS (Sigma-Aldrich, 151-21-3), 25 mM β-glycerophosphate (Sigma-Aldrich, 50020-100 G), 50 mM NaF (Sigma-Aldrich, 7681-49-4), 10 mM sodium pyrophosphate (Sigma-Aldrich, 13472-36-1), 0.5% orthovanadate (Sigma-Aldrich, 13721-39-6), 1% PMSF (Roche Life Sciences, 329-98-6), 25 U benzonase nuclease (EMD, 70664-10KUN), protease inhibitor cocktail (PierceTM Protease Inhibitor Mini Tablets, Thermo Scientific, A32963), and phosphatase inhibitor cocktail 2 (Sigma-Aldrich, P5726). Lysates were collected, mixed with 4x Laemmli buffer (BioRad, #161-0747), and boiled at 95 Celsius for 10 min before 10 uL were loaded on to 4–10% SDS-PAGE gels and run for 30 min at 250 Volts.
Western blotting
The PDGFRɑ antagonism assay was analyzed using two different western blot techniques. One repeat of each was analyzed via traditional techniques which is as follows: after separation, proteins were transferred onto a nitrocellulose membrane (12 min, semi-dry transfer) and blocked for one hour in 5% bovine serum albumin. Membranes were probed with the following primary antibodies: phospho-PDGFRα (Tyr762) (Cell Signaling Technology, #24188) 1:1000 dilution, Phospho-Akt (Ser473) (Cell Signaling Technology, #9271) 1:1000 dilution, Phospho-p44/42 MAPK (Erk1/2) (Thr202/Tyr204) (Cell Signaling Technology, #9101) 1:10,000 dilution, and either S6 (Cell Signaling Technology, #2117) at 1:1000 dilution or H3 (Cell Signaling Technology, #9715) at 1:5000 dilution as a loading control. After overnight incubation on a rocker, the membrane was probed with HRP-conjugated secondary antibody, washed 3 times, and imaged with a Bio-Rad ChemiDoc Imager. The blot image was quantified using ImageJ peak band intensity. In addition, samples were analyzed using a Biotechne Jess Automated Western Blot Machine (Biotechne, #004-650). Samples were diluted 1:3 before being prepared as per the kit instructions for 12-230 kDA separation modules (Biotechne, #SM-W001). Samples were probed using the same primary antibodies as above via a RePlex assay (Biotechne, #RP-001) that was run using the default settings for anti-rabbit chemiluminescence (Biotechne, #DM-001). Signal was quantified using area under the curve for each protein of interest. All data points from each assay were normalized to a housekeeping gene in the same lane and then normalized to the 1 nM PDGF-AA condition within each experiment.
Vectors and constructs/lentiviral generation and infection
Lentiviral particles of the pHAGE-PDGFRɑ plasmid (Addgene, 116769)84 were generated by transfecting 60-80% confluent HEK293FT cells, maintained in 10 mL HEK growth medium (Gibco™ DMEM, high glucose (Gibco, 11965092) + 10% FBS + 1% P/S) on a 100 mm TC-treated culture dish, with 20 µg pHAGE-PDGFRɑ, 15 µg psPAX2 (Addgene, 12260), and 5 µg pMD2.g (Addgene, 12259) combined with 1.8 mL Opti-MEM medium (Gibco, 31985070) and 60 µg linear polyethylenimine. Transfected HEK293FT cells were replenished with fresh HEK growth medium 24 h after the transfection. The supernatant containing lentivirus was collected 48 and 72 h post-transfection. The collected supernatant was filtered through a 0.45 µM PES syringe filter. CHO-PDGFRα cells were generated by infecting 60–80% confluent heparan-deficient Chinese Hamster Ovary cells (CHO) cells (pgsD-677 cells) (# CRL-2244) with the filtered lentivirus-containing supernatant. Infected CHO cells were supplemented with fresh CHO growth medium 24 h after the lentiviral infection. 48 h after the infection, CHO cells were selected in the CHO selection medium (CHO growth medium containing 10 µg/mL puromycin). CHO selection medium was replaced every 48 h for 7 days. CHO-PDGFRα cells post-selection were maintained in the selection medium.
FCRL5 cell line preparation
Doxycycline-inducible expression of Flag-FCRL5-P2A-T2A-EGFP was generated by first seeding 1 ×106 WT HeLa cells in 10 cm dish (Genesee 25-202) in Dulbecco’s Modified Eagle Medium (DMEM, Gibco 11995073) supplemented with 10% heat-inactivated fetal bovine serum (HI FBS, Gibco A5256801) and 1% penicillin-streptomycin (PS, Gibco 15140122). The next day, cells were transfected using TransIT-LT1 Transfection reagent (Mirus Bio MIR2300) according to manufacturer protocol with 2:1 donor:sleeping beauty transposase plasmids (Addgene Plasmid #34879)85. After 72 h, cells were covered with selection media comprised of DMEM + 10 % HI FBS + 1% PS and 2 μg/mL puromycin (Invivogen 58-58-2). Selection media was replaced every 48 h until control WT cells were completely dead. Cells were maintained in selection media.
FCRL5bp internalization assay
Flag-FCRL5 HeLa cells were counted and seeded at 50,000 per well in a 24-well plate (Genesee 25-107) in DMEM + 10% HI FBS + 1% PS. The next day, cells were treated with either 0, 100, or 1,000 ng/mL doxycycline (Fisher Scientific BP26531), and incubated for 48 h at 37 °C 5% CO2. Cells were then lifted, counted using TrypanBlue, and seeded at 12,500 cells per well in a 96-well plate (Corning 3595) in phenol red-free DMEM (Gibco 31053028) supplemented with 10 % HI FBS and 0, 100, or 1000 ng/mL. Cells were incubated at 37 °C 5% CO2 for 8 h. Then, AviTagged-FCRL5bp was diluted in phenol red-free DMEM + 10% HI FBS and respective doxycycline concentrations. Then, FCRL5bp was complexed 1:1 with TFP ester-pHrodoDeepRed (Invitrogen P35358) labeled NeutrAvidin (Thermo Scientific 31000) for 15 min at 37 °C covered from light. Media was then replaced with respective treatments, and cells were monitored with a live-cell imaging incubator (Sartorius Incucyte S3) and internalization was quantified as the overlap of red and green fluorescent area divided by the total phase area in a well.
FCRL5bp-EGFRn bifunctional degradation assay
Flag-FCRL5 HeLa cells were counted and seeded at 300,000 per well in a 6-well plate (Genesee 25-105) in DMEM + 10% HI FBS + 1% PS and incubated overnight at 37 °C, 5% CO2. Then, cells were treated with either 0 or 1000 ng/mL doxycycline (Fisher Scientific BP26531) and incubated for 48 h at 37 °C, 5% CO2. Media was then replaced with treatments of 100 ng/mL EGF (Gibco AF10015) or 50 nM of respective FCRL5bp, EGFRn, or bifunctional treatment were prepared in media with and without 1,000 ng/mL doxycycline. Cells were incubated with treatments for 48 h at 37 °C, 5% CO2, then cells were lifted in 600 uL total volume. One-sixth of the total volume of cells was reserved for live-cell immunostaining and flow cytometry. Cells were washed three times by centrifuging for 5 min at 4 °C, 500 g, discarding supernatants, and resuspending in ice-cold PBS + 1% BSA (Sigma-Aldrich A3608). After the final wash, cells were resuspended in PBS + 1% BSA + 4 ug/mL anti-EGFR (Thermo Scientific MA513319) and incubated on ice for 30 min before repeating washing three times in PBS + 1% BSA. Cells were then resuspended in PBS + 1% BSA + 4 ug/mL Alexa Fluor 647 AffiniPure Goat Anti- Mouse IgG (H + L) (Jackson ImmunoResearch Laboratories 115-605-003) and incubated on ice for 30 min covered from light. Cells were then washed three times and quantified on BD Accuri C6 Plus flow cytometer, and data were processed using FlowJo v10 Software. The remaining cells were washed three times in ice-cold PBS before resuspending in RIPA lysis buffer (Thermo Scientific 89900) + protease inhibitor cocktail tablet (Thermo Scientific A32955) and lysing cells on ice for 15 min. Lysates were then centrifuged at 4 °C, 18,000 g for 15 min. The protein concentrations were quantified with a bicinchoninic acid assay kit (Thermo Scientific 23227) according to manufacturer’s protocols. Samples were prepared at equal protein with NuPage 4x lithium dodecyl sulfate buffer (Invitrogen NP0007) + 0.1 M DL- dithiothreitol (Thermo Scientific R0861), and boiled at 95 °C for 10 min before spinning down and loading on a 4–12% Bis-Tris protein gel (Bio-Rad 3450124). The gel was loaded and ran at 200 V for 1 h in 1x XT MES running buffer (Bio-Rad 1610789). The gel was then released from its cassette and transferred to a nitrocellulose membrane using a Trans-Blot Turbo Transfer Kit (Bio-Rad1704271) for 15 min at 25 A and 2.5 V. The membrane was then trimmed and blocked for 1 h at RT in Intercept (PBS) blocking buffer (LI-COR 927-70001). Then, the membrane was covered with blocking buffer + D38B1 anti-EGFR antibody (Cell Signaling Technology 4267) and incubated at 4 °C overnight on a rocker. The membrane was then washed three times for 5 min in PBS + 0.1 % Tween-20 (Thermo Scientific J20605.AP), and incubated for 1 h rocking at RT in goat anti-rabbit IgG (H + L) antibody IRDye 800CW (LI-COR 926-32211) in blocking buffer. The membrane washing procedure was repeated, then rinsed three times in PBS before imaging on an Odyssey CLx Imaging System. The membrane was then incubated overnight at 4 °C in blocking buffer + 7F9 anti-Vinculin antibody (Santa Cruz Biotech sc-73614). The washing procedure was repeated and the membrane was incubated for 1 h at RT in goat anti-mouse IgG (H + L) antibody IRDye 680RD (LI-COR 926-68070) in blocking buffer. The washing, rinsing, and imaging procedure was repeated. EGFR staining was quantified relative to vinculin staining using Image Studio software, and reported as a percentage relative to respective no doxycycline or +doxycycline control.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper, including all experimental data presented in this manuscript. Namely, In silico datasets, SPR traces, parameters for kinetic fits, and functional binder experimental data, and the data presented in the supplemental information file are included. Crystallographic data for the KITbp–KIT complex have been deposited in the Protein Data Bank under accession code PDB 9H71. Data corresponding to the α-CTX binder (α-CTXbp) were previously reported by Vázquez Torres in ref. 62. The Crystallographic data used here are available under PDB accession code 9BK6. No restrictions apply to data access. Larger files (PDB files, sequences, and analysis scripts for our in silico experiments, FACS data, design methods scripts and instructions, PDB files that were experimentally screened, and previously unpublished sequences that had been tested) can be found at: https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/in_silico_benchmark_outputs.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/anaysis_scripts.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/facs_data.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/target_pdbs.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/interface_tensors.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/experimentally_tested_design_pdbs.zip, https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/alk3_kit_failed_libraries.zip. Source data are provided with this paper.
Code availability
Code for running RFdiffusion has been released on GitHub, free for academic, personal and commercial use at https://github.com/RosettaCommons/RFdiffusion. An interface conditioning script that can be used to replicate the approach here is provided at https://files.ipd.uw.edu/pub/strand_pairing_binders_2025/interface_tensors.zip, along with scripts used to calculate SC50 values from yeast surface display titrations (as originally reported by Cao et. al.), scripts for calculating TMscores and for deriving and plotting t-SNE transformations of this data. The code is distributed under the MIT License and can be freely reused and modified. A script to run AF2 with an initial guess as was used in this work is provided at this Zenodo link: https://doi.org/10.5281/zenodo.7730843 ref. 6.
References
Cao, L. et al. Design of protein-binding proteins from the target structure alone. Nature 605, 551–560 (2022).
Watson, J. L. et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023).
Gainza, P. et al. De novo design of protein interactions with learned surface fingerprints. Nature 617, 176–184 (2023).
Vázquez Torres, S. et al. De novo design of high-affinity binders of bioactive helical peptides. Nature 626, 435–442 (2024).
Cao, L. et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science 370, 426–431 (2020).
Bennett, N. R. et al. Improving de novo protein binder design with deep learning. Nat. Commun. 14, 2625 (2023).
Stranges, P. B. & Kuhlman, B. A comparison of successful and failed protein interface designs highlights the challenges of designing buried hydrogen bonds: Successful Interface Designs Avoid Polar Interactions. Protein Sci. 22, 74–82 (2013).
Watkins, A. M. & Arora, P. S. Anatomy of β-strands at protein-protein interfaces. ACS Chem. Biol. 9, 1747–1754 (2014).
Richardson, J. S. & Richardson, D. C. Natural beta-sheet proteins use negative design to avoid edge-to-edge aggregation. Proc. Natl. Acad. Sci. USA 99, 2754–2759 (2002).
Sahtoe, D. D. et al. Transferrin receptor targeting by de novo sheet extension. Proc. Natl. Acad. Sci. USA 118, e2021569118 (2021).
Fujiwara, K., Ebisawa, S., Watanabe, Y., Toda, H. & Ikeguchi, M. Local sequence of protein β-strands influences twist and bend angles: Side Chain Influences β-Strand Twisting. Proteins 82, 1484–1493 (2014).
Fujiwara, K., Ebisawa, S., Watanabe, Y., Fujiwara, H. & Ikeguchi, M. The origin of β-strand bending in globular proteins. BMC Struct. Biol. 15, 21 (2015).
Richardson, J. S., Getzoff, E. D. & Richardson, D. C. The beta bulge: a common small unit of nonrepetitive protein structure. Proc. Natl. Acad. Sci. USA 75, 2574–2578 (1978).
Fox, N. K., Brenner, S. E. & Chandonia, J.-M. SCOPe: structural classification of proteins-extended, integrating SCOP and ASTRAL data and classification of new structures. Nucleic Acids Res. 42, D304–D309 (2014).
Chandonia, J.-M. et al. SCOPe: improvements to the structural classification of proteins - extended database to facilitate variant interpretation and machine learning. Nucleic Acids Res. 50, D553–D559 (2022).
Bragdon, B. et al. Bone morphogenetic proteins: a critical review. Cell. Signal. 23, 609–620 (2011).
Sorkin, M. et al. Hair follicle specific ACVR1/ALK2 critically affects skin morphogenesis and attenuates wound healing. Wound Repair Regen. 25, 521–525 (2017).
Liu, M., Goldman, G., MacDougall, M. & Chen, S. BMP signaling pathway in dentin development and diseases. Cells 11, 2216 (2022).
Sugimoto, H. et al. Activin-like kinase 3 is important for kidney regeneration and reversal of fibrosis. Nat. Med. 18, 396–404 (2012).
Ruan, X. et al. Activin receptor-like kinase 3: a critical modulator of development and function of mineralized tissues. Front. Cell Dev. Biol. 11, 1209817 (2023).
Marino, F. et al. Role of c-kit in myocardial regeneration and aging. Front. Endocrinol. 10, 371 (2019).
Di Siena, S. et al. Activated c-Kit receptor in the heart promotes cardiac repair and regeneration after injury. Cell Death Dis. 7, e2317 (2016).
Cimini, M. et al. C-kit dysfunction impairs myocardial healing after infarction. Circulation 116, I77–I82 (2007).
Yaniz-Galende, E. et al. Stem cell factor gene transfer promotes cardiac repair after myocardial infarction via in situ recruitment and expansion of c-kit+ cells. Circ. Res. 111, 1434–1445 (2012).
Lutz, M. et al. Local injection of stem cell factor (SCF) improves myocardial homing of systemically delivered c-kit + bone marrow-derived stem cells. Cardiovasc. Res. 77, 143–150 (2008).
Takematsu, E. et al. Transmembrane stem cell factor protein therapeutics enhance revascularization in ischemia without mast cell activation. Nat. Commun. 13, 2497 (2022).
Kim, B.-J. et al. Platelet-derived growth factor receptor-alpha positive cardiac progenitor cells derived from multipotent germline stem cells are capable of cardiomyogenesisin vitroandin vivo. Oncotarget 8, 29643–29656 (2017).
Kalra, K., Eberhard, J., Farbehi, N., Chong, J. J. & Xaymardan, M. Role of PDGF-A/B ligands in cardiac repair after myocardial infarction. Front. Cell Dev. Biol. 9, 669188 (2021).
Horikawa, S. et al. PDGFRα plays a crucial role in connective tissue remodeling. Sci. Rep. 5, 17948 (2015).
Ono, C. et al. Upregulated Fcrl5 disrupts B cell anergy and causes autoimmune disease. Front. Immunol. 14, 1276014 (2023).
Pandey, P. et al. New insights about the PDGF/PDGFR signaling pathway as a promising target to develop cancer therapeutic strategies. Biomed. Pharmacother. 161, 114491 (2023).
Saiz, M. L. et al. Epigenetic targeting of the ACE2 and NRP1 viral receptors limits SARS-CoV-2 infectivity. Clin. Epigenet. 13, 187 (2021).
Rizzolio, S. et al. Neuropilin-1 upregulation elicits adaptive resistance to oncogene-targeted therapies. J. Clin. Invest. 128, 3976–3990 (2018).
Tang, Y. H. et al. Neuropilin-1 is over-expressed in claudin-low breast cancer and promotes tumor progression through acquisition of stem cell characteristics and RAS/MAPK pathway activation. Breast Cancer Res. 24, 8 (2022).
de la Rosa, G., Corrales-García, L. L., Rodriguez-Ruiz, X., López-Vera, E. & Corzo, G. Short-chain consensus alpha-neurotoxin: a synthetic 60-mer peptide with generic traits and enhanced immunogenic properties. Amino Acids 50, 885–895 (2018).
Ensan, D. et al. Targeting ALK2: An open science approach to developing therapeutics for the treatment of diffuse intrinsic pontine glioma. J. Med. Chem. 63, 4978–4996 (2020).
Carvalho, D. et al. ALK2 inhibitors display beneficial effects in preclinical models of ACVR1 mutant diffuse intrinsic pontine glioma. Commun. Biol. 2, 156 (2019).
Zhou, X. P. et al. Germline mutations in BMPR1A/ALK3 cause a subset of cases of juvenile polyposis syndrome and of Cowden and Bannayan-Riley-Ruvalcaba syndromes. Am. J. Hum. Genet. 69, 704–711 (2001).
Ehata, S. & Miyazono, K. Bone morphogenetic protein signaling in cancer; Some topics in the recent 10 years. Front. Cell Dev. Biol. 10, 883523 (2022).
Corless, C. L. et al. PDGFRA mutations in gastrointestinal stromal tumors: frequency, spectrum and in vitro sensitivity to imatinib. J. Clin. Oncol. 23, 5357–5364 (2005).
Simpson, J. E. et al. Autophagy supports PDGFRA-dependent brain tumor development by enhancing oncogenic signaling. Dev. Cell 59, 228–243.e7 (2024).
Jansson, S. et al. The PDGF pathway in breast cancer is linked to tumour aggressiveness, triple-negative subtype and early recurrence. Breast Cancer Res. Treat. 169, 231–241 (2018).
Evans, E. K. et al. A precision therapy against cancers driven by KIT/PDGFRA mutations. Sci. Transl. Med. 9, (2017).
Curtin, J. A., Busam, K., Pinkel, D. & Bastian, B. C. Somatic activation of KIT in distinct subtypes of melanoma. J. Clin. Oncol. 24, 4340–4346 (2006).
Küçükköse, E. et al. KIT promotes tumor stroma formation and counteracts tumor-suppressive TGFβ signaling in colorectal cancer. Cell Death Dis. 13, 617 (2022).
Rubin, B. P. et al. KIT activation is a ubiquitous feature of gastrointestinal stromal tumors. Cancer Res. 61, 8118–8121 (2001).
Sakuma, Y., Sakurai, S., Oguni, S., Hironaka, M. & Saito, K. Alterations of the c-kit gene in testicular germ cell tumors. Cancer Sci. 94, 486–491 (2003).
Hirota, S. et al. Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 279, 577–580 (1998).
Bryan, C. M. et al. Computational design of a synthetic PD-1 agonist. Proc. Natl. Acad. Sci. USA 118, e2102164118 (2021).
Edman, N. I. et al. Modulation of FGF pathway signaling and vascular differentiation using designed oligomeric assemblies. Cell 187, 3726–3740.e43 (2024).
Silva, D.-A. et al. De novo design of potent and selective mimics of IL-2 and IL-15. Nature 565, 186–191 (2019).
Huang, B. et al. Designed endocytosis-inducing proteins degrade targets and amplify signals. Nature 638, 796–804 (2025).
Chevalier, A. et al. Massively parallel de novo protein design for targeted therapeutics. Nature 550, 74–79 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Varadi, M. et al. AlphaFold Protein Structure Database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Hinck, A. P. Structural studies of the TGF-βs and their receptors - insights into evolution of the TGF-β superfamily. FEBS Lett. 586, 1860–1870 (2012).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Dauparas, J. et al. Robust deep learning-based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Gloegl, M. et al. Target-conditioned diffusion generates potent TNFR superfamily antagonists and agonists. bioRxiv https://doi.org/10.1101/2024.09.13.612773 (2024).
Vázquez Torres, S. et al. De novo designed proteins neutralize lethal snake venom toxins. Nature 639, 225–231 (2025).
Franco, A. et al. Human Fc receptor-like 5 binds intact IgG via mechanisms distinct from those of Fc receptors. J. Immunol. 190, 5739–5746 (2013).
McDonald, I. K. & Thornton, J. M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994).
Lo Conte, L., Chothia, C. & Janin, J. The atomic structure of protein-protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Cheng, P.-N., Pham, J. D. & Nowick, J. S. The supramolecular chemistry of β-sheets. J. Am. Chem. Soc. 135, 5477–5492 (2013).
Yuzawa, S. et al. Structural basis for activation of the receptor tyrosine kinase KIT by stem cell factor. Cell 130, 323–334 (2007).
Appleton, B. A. et al. Structural studies of neuropilin/antibody complexes provide insights into semaphorin and VEGF binding. EMBO J. 26, 4902–4912 (2007).
Kschonsak, M. et al. Structures of HCMV Trimer reveal the basis for receptor recognition and cell entry. Cell 184, 1232–1244.e16 (2021).
Chennamsetty, N., Voynov, V., Kayser, V., Helk, B. & Trout, B. L. Design of therapeutic proteins with enhanced stability. Proc. Natl. Acad. Sci. USA 106, 11937–11942 (2009).
Muratspahić, E. et al. De novo design of miniprotein agonists and antagonists targeting G protein-coupled receptors. bioRxivorg 2025 03, .644666 (2025). 23.
Benatuil, L., Perez, J. M., Belk, J. & Hsieh, C.-M. An improved yeast transformation method for the generation of very large human antibody libraries. Protein Eng. Des. Sel. 23, 155–159 (2010).
Huang, T. & Hinck, A. P. Production, isolation, and structural analysis of ligands and receptors of the TGF-β superfamily. Methods Mol. Biol. 1344, 63–92 (2016).
Wicky, B. I. M. et al. Hallucinating symmetric protein assemblies. Science 378, 56–61 (2022).
Kabsch, W. X. D. S. Acta Crystallogr. D. Biol. Crystallogr. 66, 125–132 (2010).
Jamshidiha, M. et al. Coping with strong translational noncrystallographic symmetry and extreme anisotropy in molecular replacement with Phaser: human Rab27a. Acta Crystallogr. D. Struct. Biol. 75, 342–353 (2019).
Tickle, I. STARANISO: use of a WebGL-based 3D interactive graphical display to represent and visualise data quality metrics for anisotropic macromolecular diffraction data. Acta Crystallogr. A Found. Adv. 75, e162 (2019).
Vonrhein, C. et al. Advances in automated data analysis and processing within autoPROC, combined with improved characterisation, mitigation and visualisation of the anisotropy of diffraction limits using STARANISO. Acta Crystallogr. A Found. Adv. 74, a360 (2018).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220 (2014).
Ng, P. K.-S. et al. Systematic functional annotation of somatic mutations in cancer. Cancer Cell 33, 450–462.e10 (2018).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Acknowledgements
This research was supported by The Audacious Project at the Institute for Protein Design (I.S., D.L. S.V.T., J.N.S., X.W., D.B.), NSF-GRFP program (S.A.R.), Howard Hughes Medical Institute (D.B., I.G., B.C., M.G.S, N.R., D.V., M.G., J.N.S.), The Nordstrom Barrier Institute for Protein Design Directors Fund (B.H.), The Open Philanthropy Project Improving Protein Design Fund (S.V.T. and B.C.), and AMGEN Donation to the Institute for Protein Design (X.W.). The project or effort depicted was or is sponsored by the Department of the Defense, Defense Threat Reduction Agency grant HDTRA1-21-1-0007 (M.G., X.W.). S.N.S. is supported by the Flanders Institute for Biotechnology (VIB), Belgium (grant no. C0101), Research Foundation Flanders (FWO), Belgium (grant nr. S000722N), and an FWO EOS research grant (grant no. G0H1222N). M.T. has received funding from the European Union’s Horizon Europe research and innovation programme under the Marie Skłodowska-Curie grant agreement No. 101155448 (HeartRepairKIT). T.P.J. acknowledges support from the Alliance programme under the EuroTech Universities agreement. M.B.V. has received funding from the European Union’s Horizon 2020 research and innovation programme under the MarieSkłodowska-Curie grant agreement No 899987. Biotinylated ALK-2 and ALK-3 were kindly provided by Andrew Hinck. We thank the staff of the beamline P13 (PETRA III, EMBL Hamburg) for technical support and beamtime allocation. We also acknowledge Preetham Venkatesh and Fiona Wang for support in testing the design methodology described in this study, Xinting Li for help with mass spectrometry analysis of proteins, Rohith Krishna for logistics support, and Ashish Phal for identifying targets of interest.
Author information
Authors and Affiliations
Contributions
I.S. and D.B. designed the research plan and goals. I.S. and J.L.W. conceived of the computational design strategy, and I.S. wrote the scripts for implementation into previously published RFdiffusion workflows for protein binder design. I.S., M.T., D.S.L., S.A.R., C.M., B.H., C.J.K., S.V.T., C.S.H., M.B.V., G.A., X.W., A.P.H., T.P.J., H.R.B., S.M.B., S.N.S., and D.B. identified an appropriate target set for applying the design method. I.S. designed all the reported binder sequences, with the exception of ALK-2bp (designed by D.S.L.) and NRP1bp (designed by C.K.). I.S. and B.C. designed sequences that were in the initial yeast surface display library screened for PDGFRɑ binders. I.S., D.S.L., B.H., I.G., D.V., M.G.S., and N.R. performed the yeast surface display assays for ALK-2, ALK-3, FCRL5, KIT, and PDGFRɑ. I.S., D.L., M.G., C.J.K., S.V.T., and J.N.S. contributed cloning, expression, purification, and binding characterization for the identified binders. M.T., K.H.G.V., K.V., and S.N.S. crystallized and solved the structure of KITbp in complex with the KIT receptor. Functional assays were performed by S.A.R. and S.M.B. (for FCRL5bp) and C.M., T.T.C., N.B., and H.R.B. (for PDGFRɑ).
Corresponding author
Ethics declarations
Competing interests
J.L.W. and B.H. are employed at Xaira Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jake Kerrison and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sappington, I., Toul, M., Lee, D.S. et al. Improved protein binder design using β-pairing targeted RFdiffusion. Nat Commun 17, 1101 (2026). https://doi.org/10.1038/s41467-025-67866-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67866-3