Abstract
The bacterial flagellum, essential for motility and pathogenesis, requires the filament cap (FliD) to polymerize flagellin (FliC). However, the mechanisms governing the transition from the hook junction to the filament elongation remain elusive, obscured by stoichiometric mismatches and barely detectable interactions. To resolve this, we deploy solution NMR to characterize ultra-weak interactions, quantify affinities (KD ≈ 0.1 mM for junction protein FlgL; 1.65 mM for FliC). These data enable rational complex stabilization for cryo-EM structure determination of Salmonella FliD pentamers complexed with FlgL or FliC, revealing that both substrates engage an identical conserved surface in a 5:5 stoichiometry. Integrating these structures into native flagellar tip densities reveal a 5:11 FliD:FlgL/FliC architecture, where six additional subunits barely detected by NMR dock at secondary sites. Mutations that disrupt or enhance these interfaces impair motility and filament integrity, while disulfide-locked FliD pentamers confirm that cap rigidity is crucial for elongation. These findings support a rotary cap mechanism where ultra-weak binding and structural fidelity of the cap ensure efficient flagellin polymerization. Our study resolves the long-standing paradox of stoichiometric mismatch in flagellar filament biogenesis, providing a blueprint for the assembly of dynamic macromolecular machines.
Similar content being viewed by others
Introduction
The bacterial flagellum is a highly regulated macromolecular rotary nanomachine1,2,3. It functions as a motor equipped with a propeller, enabling bacterial mobility1,4,5 and playing a critical role in bacterial survival, reproduction, colonization, and pathogenicity6,7,8. This complex apparatus is comprised of more than 25 conserved proteins exported through a dedicated type III secretion system (T3SS)6,9, organized into three main regions: the basal body, the proximal hook, and the distal helical filament (Fig. 1)10,11. The basal body acts as a motor and protein export apparatus, driven by proton-motive force10,12,13,14,15. The hook, a universal joint in a curved structure, connects the basal body to the filament and transmits the torque from the stator to the filament tail1,11. Assembly of the ∼55 nm long hook16,17,18 needs ~120 copies of the hook protein FlgE19,20,21,22,23. The long helical filament24,25,26,27,28, composed mainly of polymerized flagellin (FliC), functions as a propeller to generate thrust and adapts to different environments through various super-helical geometries27,29,30.
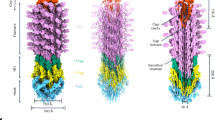
This schematic illustrates the hierarchical organization and spatial positioning of the major components of the bacterial flagellum, a highly complex nanomachine comprising two primary modules: the rotary motor and the helical propeller. The rotary motor drives the rotational movement of the flagellum, which is transmitted through the basal body and the hook (composed of FlgE) to the filament. Filament assembly initiates at the hook-filament junction tip (left panel), where the FlgL ring (colored dark yellow) serves as an anchoring point for the growing filament and the filament cap (colored dark magenta). The cap docks at this junction, playing a critical role in recognizing and positioning incoming FliC (flagellin) subunits (colored teal) as the filament elongates. The process continues until the filament is fully assembled (right panel), ensuring the structural integrity required for the flagellum’s function as an efficient helical propeller.
Assembly of the flagellum proceeds through a highly ordered process, beginning with the basal rod, followed by the hook, hook-filament junction, filament cap, and ultimately the filament8,10,11. The filament elongates starting from the interface between the hook-filament junction and the filament cap31,32(Fig. 1). The hook-filament junction consists of two layers of morning glory-like structures, built by the junction protein FlgK and junction protein FlgL11,33,34. In the junction, a ring-shaped FlgK oligomer is in contact with the hook and the FlgL ring19,24 (Fig. 1). After the assembly of the hook junction, the filament cap protein FliD is secreted to the distal end of the hook junction29,35. The secreted FliD docks at the FlgL ring at the tip of the hook and forms a cap-shaped oligomer that nucleates filament elongation36,37(Fig. 1). Despite these evidences implicating interactions between FliD and other flagellar components, direct biochemical characterization of these interactions in vitro remains scarce.
Beyond interaction features, a comprehensive understanding of filament assembly demands structural elucidation of flagellar components at the growing distal end6. Advances in cryo-electron microscopy (cryo-EM) and X-ray crystallography have resolved high-resolution structures of individual components24,38,39,40. FliD, the cap protein, forms a homo-tetramer to homo-hexamer stool-shaped complex40,41,42,43; FlgL, the hook-filament junction component directly involved in filament assembly, remains mysterious except for its truncated and monomeric structures till very recent39,44,45; FliC, the primary flagellin protein, polymerizes into a helical filament and its structure has been determined at near-atomic resolution, which contributes to the filament rigidity and motility46,47. Pioneering work during the last three decades, based on elegant electron microscopy, proposed the influential hypothesis of a rotating FliD cap acting as a dynamic polymerization chaperone and rotating incrementally to sequentially position incoming flagellin subunits for processive filament elongation31,37,41. This model provided a crucial conceptual framework, yet the underlying molecular mechanics—how the pentameric cap dynamically interfaces with the flagellum’s prevailing 11-fold symmetry—remained a structural and biophysical paradox. While a recent near-atomic structure provided key insights into the FliD cap atop the FlgL ring35, the fundamental principles governing the symmetry mismatch accommodation, the atomic details of subunit binding, and the functional necessity of precisely tuned interaction affinities have remained elusive. This lack of clarity has hindered a mechanistic understanding of how the flagellar cap discriminates the hook-filament junction from flagellin to mediate rapid filament assembly.
Here, we address these challenges through an integrated approach combining solution-state nuclear magnetic resonance (NMR) spectroscopy to characterize ultraweak FliD-substrate interactions, with cryo-EM structural analysis of transient complexes and native flagellar assemblies. Our structural and biochemical results reveal how finely tuned binding affinities and cap architecture coordinate filament assembly via a rotary mechanism, resolving the paradox of symmetry mismatch and ultraweak interactions in the assembly of this essential macromolecular machine.
Result
Interaction of FliD with FlgL and FliC
The flagellar filament cap protein FliD of Salmonella enterica serovar Typhimurium is composed of 467 residues and harbors four major domains: D0, D1, D2, and D3 (Fig. 2a, b). The D0 domain features an N-terminal signal peptide region (SPR, residue 1–44) and a C-terminal chaperone binding region (CBR, residues 411–467), which are crucial for protein secretion through the T3SS component. The D1 domain (residues 44–72, 268–411) facilitates flagellin targeting and filament extension, while the D2 (residues 72–103, 223–268) and D3 (residues 103–223) (Polymerization domain, PD) domains intrinsically oligomerize to form the cap structure of the flagellar filament (Fig. 2a, b).
a Domain organization of FliD protein from Salmonella enterica serovar Typhimurium. b Cartoon rendering of the FliD structure with a color gradient from blue (N-terminus) to red (C-terminus). c Representative 2D class averages of the full-length FliD and FliD d0 domain truncation construct (FliDd1d2d3, residues from 44 to 411) derived from cryo-EM single particle classification. d, e Cryo-EM structure of FliD homo-pentamer rendered as a cartoon mode, with Chain A colored according to a rainbow gradient scheme. f Expanded view of the FliD Homo-pentamer interface. Residues involved in intermolecular contact are shown as ball-and-sticks. g Sequence conservation analysis of the residues involved in FliD pentamerization. The equilibrium was assessed by analytical size-exclusion chromatography, with the monomer and pentamer peaks quantified by integrated peak area. h Effect of key amino acid substitution on the equilibrium of FliD monomer-pentamer in solution. i Solvent-exposed surface of FliD rendered with the YRB color scheme74 to highlight hydrophobicity and charge. Hydrophobic, negative-charged, and positive-charged residues were colored in yellow, red, and blue, respectively. Others were colored in grey. j The FlgL and FliC binding site on FliD determined by NMR titration is rendered as surface mode and colored in dark orange and cyan, respectively. Notably, all residues implicated in FliC binding fall within the FlgL-binding region; thus, the overlapping interface is represented by the cyan-colored surface. Unassigned residues located at the potential interface are colored in light grey. k Sequence conservation analysis of FliD, colored based on residue identity conservation scores determined by Consurf server75.
Our initial trial shows that spontaneous oligomerization of the full-length FliD would form high levels of aggregation rather than a well-defined cap-like oligomer (Fig. 2c and Supplementary Fig. 1). Subsequently, we designed and screened FliD truncations to form a homogeneous oligomer. The FliD D0 domain truncation successfully yielded a homopentamer (Fig. 2c), and its cryo-EM structure was solved at 3.06 Å resolution (Fig. 2d, e, and Supplementary Fig. 2 and Supplementary Table 1). This structure showed significant orientational differences compared to previously reported X-ray structures of the FliD D2-D3 domains (Supplementary Fig. 3a, b).
The structure shows that the FliD pentamer is stabilized by the intermolecular interactions between D2 and D3 domains, including several hydrophobic interactions involving residues I234 and I236 from D2 domain, V172, and V174 from D3 domain, as well as a salt bridge between E237 in D2 domain and K175 in D3 domain (Fig. 2f). Especially, the high conservation of these hydrophobic residues across species suggests their functional importance (Fig. 2g and Supplementary Fig. 3c, d). Substitution of individual residues at the interface destabilized the pentamer, as measured by a shift in the monomer-pentamer equilibrium using size-exclusion chromatography, highlighting the critical role of these interactions (Fig. 2h and Supplementary Fig. 3e, f). The FliD pentamer maintains a stable core, evidenced by 7100 Ų of buried hydrophobic interface (Supplementary Fig. 3d). This buried surface area (~7100 Ų) falls within the range of stable oligomeric interfaces—comparable to molecular chaperone SecB (Homo-tetramer, 8373 Ų)48,49, yet significantly larger than transient signaling complexes like EIIA-EIIB (702 Ų) in phosphotransferase system50, underscoring the inherent stability of the FliD pentameric core. However, its solvent-exposed surface lacks large hydrophobic or charged patches (Fig. 2i), indicating an absence of strong, specific binding sites for other flagellar subunits. This non-specific binding landscape suggests FliD engages FlgL and FliC through weak, transient interactions—a design essential for its role as a rigid platform that enables rapid subunit exchange during processive filament elongation.
To gain atomic interaction of FliD with other filament-associated proteins, we used the solution NMR spectroscopy. Given the large size of FliD, this protein was segmented into three constructs based on its domain boundaries using the “divide and conquer” strategy51 (Fig. 2a, b). The interaction between each segmented domain and FliC/FlgL was analyzed using NMR titration experiments. NMR analysis revealed that FliC and FlgL bind specifically to the D1 domain of FliD (Supplementary Fig. 4a–c). Significant line broadening observed during titration indicates chemical exchange on the microsecond-to-millisecond timescale, which is characteristic of weak, transient protein–protein interactions. While only minimal chemical shift perturbations were observed for the D2-D3 domains (Supplementary Fig. 4d–f), indicating substantially even weaker or nonspecific binding compared to the interactions at the D1 domain.
The 1H-15N correlated spectra of FliDd1 showed well-dispersed peaks, enabling near-complete resonance assignment (Supplementary Fig. 5a, b). Upon the addition of FlgL and FliC to labeled FliD-D1, significant line broadening was observed in the 1H-15N correlated spectra, indicating interactions between these proteins and the D1 domain. Further NMR analysis revealed that FlgL and FliC share an overlapping binding site on the D1 domain, located on the outer surface of the leg region within the stool-like cap structure (Fig. 2j and Supplementary Fig. 5c). The location of this binding site is consistent with its position during flagellar filament assembly and is situated in the most conserved region of FliD (Fig. 2k), underscoring the functional importance of this interface.
Structure of the flagellar filament cap FliD in complex with FlgL
FlgL, the hook-filament junction protein, represents the smallest flagellar propeller component52. Unlike multidomain FliD, FlgL comprises only D0 and D1 domains (Fig. 3a, Supplementary Fig. 6a). Since it was found that FliD-D1 could interact with FlgL, the affinity between these two proteins was quantified using NMR titration experiments (Supplementary Fig. 6b). The dissociation constant (Kd) between FliD-D1 and FlgL was determined to be ~100 μM (Fig. 3b), consistent with the established classification of an ultra-weak interaction (Kd in the high micromolar to millimolar range), which is distinct from conventional “weak” binding events50,53,54. This ultra-weak binding precluded stable complex formation for structural studies. Initial attempts to stabilize the native complex using GraFix (gradient fixation)52 were unsuccessful due to complex dissociation, prompting the use of interface-guided fusion constructs to elevate the complex ratio by mimicking proteins in infinite high concentrations55,56. We designed several fusion constructs linking FlgL to the N- or C-terminus of FliD with flexible linkers. The construct yielding a homogeneous, monodisperse complex suitable for high-resolution cryo-EM analysis was the one with the FlgL D1 (resi 40–280) domain fused to the N-terminus of full-length FliD (Supplementary Fig. 6c–f). This fusion construct allowed for successful structure determination. The cryo-EM structure of the FliD-FlgL complex was resolved at a resolution of 3.30 Å (Supplementary Table 1, and Supplementary Fig. 7).
a Domain organization of the FlgL protein from Salmonella. b Chemical shift perturbation fitting profile for 15N labeled FliDd1 upon titration with unlabeled FlgL. c, d Cryo-EM structure of FliD in complex with FlgL rendered in surface and cartoon, with FliD and FlgL colored in magenta and dark orange, respectively. e Intermolecular interaction details of the FliD-FlgL complex. Residues participating in intermolecular interaction are shown as ball-and-sticks. Hydrogen bonds and salt bridges are shown as broken lines. The expanded views of the three major interfaces were highlighted in the yellow dashed boxes. f Alternation of the key residues on the binding affinity of FliD-FlgL interaction. The effect is quantified as the percentage of complex formation in solution determined by three independent replicates of pull-down assays (Supplementary Fig. 9). The error bar represents standard deviation (SD) from n = 3 replicates. g Close-up views of the FlgL/FliD interface and selected residues for Intermolecular disulfide bond engineering. Residue pairs with Cβ–Cβ distances of 4–6 Å are as 1: FliD E300C/FlgL Q71C, 2: FliD P301C/FlgL Q71C, 3: FliD A298C/FlgL L75C,4: FliD Q348C/FlgL T89C. h Coomassie stained SDS-PAGE with nonreducing (-) vs. reducing samples (+) of wild-type and mutant FliD/FlgL complexes, validating crosslink formation. This experiment was conducted independently three times.
The overall structure of the FliD-FlgL complex reveals a 5:5 stoichiometry, where each FlgL monomer binds independently to a single FliD subunit (Fig. 3c, d). The single FliD-FlgL interface buries a total surface area of 955 Ų, with two major helices from FlgL interacting with a flat, elongated region on the D1 domain of FliD (Fig. 3e). This binding interface is in agreement with our NMR titration data (Fig. 1j) and provides structural validation for earlier hypothetical models that were based on low-resolution electron microscopy images31,37. Structural analysis of the interface reveals a highly hydrophilic and complementary surface, suggesting that polar interactions and van der Waals forces collectively contribute to stabilizing the complex (Supplementary Fig. 8).
Three major interaction motifs were identified at the interface (Fig. 3e). First, a salt bridge forms between FlgL E92 carboxyl group and FliD R326 side chain, complemented by a hydrogen bond with FliD T340. Second, a series of hydrophobic contacts occur between FliD Y296, A298, and L313, and FlgL L254 and L75. Third, another salt bridge forms between FliD R29 and FlgL D266, along with additional hydrophobic interactions between FliD P32 and FlgL L269 and L273. Notably, the structural data confirmed that there is no direct interaction between FlgL monomers. Mutagenesis studies, in which key residues involved in these interactions were substituted, resulted in decreased stability of the FliD-FlgL complex (Fig. 3f, and Supplementary Fig. 9), further confirming the importance of these specific interactions in this transient complex formation. To further validate our structure, we engineered disulfide bonds between four residue pairs at the interface (Cα distance <5 Å). Oxidation-induced crosslinking of these cysteine mutants confirmed the accuracy of our structure (Fig. 3g, h).
Structure of the flagellar filament cap FliD in complex with FliC
The flagellin FliC exhibits a domain architecture highly similar to that of full-length FliD (Fig. 4a, and Supplementary Fig.10a). Although there is extensive evidence for the important role of FliD in flagellin assembly, there is no publication reporting direct interaction between FliD and other filament components in vitro25,37,41,57. This suggests that these FliD-containing complexes are highly transient and dynamic. Benefiting from the solution NMR in characterizing ultra-weak and transient interactions50,58, we quantified the dissociation constant (Kd) of the FliD-FliC interaction at 1.65 mM with stoichiometry 1:1 (Fig. 4b, and Supplementary Fig. 10b), confirming the ultra-weak binding affinity. To stabilize this ultra-transient complex for structure analysis, we employed the fusion approach as FlgL-FliD complex stabilization. Since full-length FliC has a strong filamentation propensity, we first identified that D0 domain truncation yielded monodisperse protein. After screening multiple fusion constructs, the optimal construct for structural studies is covalently linked the N-terminus of FliC D1D2D3 (residues 45–455) to the C-terminus of full-length FliD (Supplementary Fig. 10c–f). Using this construct, we identified two distinct complexes and determined their cryo-EM structures - one with 5:5 stoichiometry at 3.30 Å resolution and another with 10:10 stoichiometry at 3.27 Å resolution (Supplementary Table 1, and Supplementary Fig. 11).
a Domain architecture of Salmonella flagellin FliC. b NMR-derived binding isotherm of 15N-labeled FliDd1 titrated with unlabeled FliC, plotted as chemical shift perturbations (CSPs) and fitted with one-site binding mode. c, d Cryo-EM structures of FliD-FliC complexes in 5:5 and 10:10 stoichiometries (surface and cartoon representations; FliD, magenta; FliC, dark cyan). e Inter-pentamer interactions in the FliD decamer, highlighting key residues with ball-and-stick. f Size-exclusion chromatography (SEC) analysis of full-length FliD and interface mutants, quantifying aggregation propensity. g Molecular interface between FliD and FliC, with expanded views of binding sites (hydrogen bonds and salt bridges shown as dashed lines). h Binding affinity modulation by FliD-FliC interface mutations, quantified as percent complex formation via NMR titration (Supplementary Figs. 14 and 15). The statistics data are presented as mean peak intensity ratio values ± SD with n = 7 from the NMR titration spectra. i Intermolecular disulfide bond engineering of FliD-FliC complex. Residue pairs with Cβ–Cβ distances of 4–6 Å are as 1: FliD S352C/FliC N83C, 2: FliD T351C/FliC E84C, 3: FliD E349C/FliC N87C, 4: FliD Q348C/FliC Q90C, 5: FliD Q348C/FliC Q90C. j SDS-PAGE with nonreducing (-) vs. reducing samples (+) of wild-type and mutant FliD/FliC complexes, validating crosslink formation (Coomassie stain). This experiment was conducted in three independent duplicates.
The structures reveal that each FliC subunit binds independently to one FliD subunit, with FliD oligomerizing in a C5 or D5 symmetry (Fig. 4c, d). In the C5 symmetry structure, the oligomerization interface mirrors that of the FliD pentamer. However, in the D5 symmetry conformation, an additional interface was observed where the C-terminal of FliD forms an antiparallel helix bundle (Fig. 4d). This inter-pentamer interaction is mediated by hydrophobic contacts involving residues Y436, F440, L446, M446, and L450 (Fig. 4e, and Supplementary Fig. 12a). Mutating these hydrophobic residues into polar amino acids restores aggregation in full-length FliD (Fig. 4f, and Supplementary Fig. 12b, c). Although this interface is visualized and confirmed in our experiments, it is likely non-physiological, as previously suggested40. Instead, we propose that these hydrophobic residues contribute to chaperone-mediated substrate anti-aggregation59 and in recognizing newly translocated FliC subunits.
All intermolecular contacts within the FliD-FliC complex occur either between FliD-FliD, or FliD-FliC, with no direct interaction between FliC subunits. The FliD-FliC interface, although burying a surface area of 1296 Ų and showing shape complementarity (Supplementary Fig. 13), features only two key interactions that contribute to the ultra-weak binding affinity (Fig. 4g). One is a salt bridge between FliD K361 and FliC E94, while the other involves FliC I72 inserting into a hydrophobic cleft in FliD formed by residues Y296, A298, and L313. Unlike the FliD-FlgL complex, no significant third interface was observed (Fig. 4g). Notably, the overall structure of the FliD pentamer remains unchanged upon FliC binding, with a root-mean-square deviation of 1.109 Šfor the backbone atoms between the apo and FliC-bound states. Substitutions of amino acids at the FliD-FliC interface, particularly those disrupting key interactions, markedly reduced the binding affinity between FliD and FliC (Fig. 4h, and Supplementary Figs. 14 and 15). Disulfide bond crosslinking further validates the transient structure of the complex in solution (Fig. 4i, j).
Structural models of the hook-filament junction tip and filament tip
In the native flagellum, the filament and hook are formed by repeating arrays of 11 subunits of FliC or FlgE per turn24,47,60,61. Though without direct and precise evidence, the hook-filament junction is also expected to contain 11 copies of FlgK as well as FlgL34,62. This indicates the stoichiometry of the FliD-FliC or FliD-FlgL complex should be 5:11 rather than the 5:5 stoichiometry we reported here. To reconcile this apparent stoichiometric discrepancy, we sought to determine the structure of the native hook-filament junction tip and the filament tip directly from bacteria flagella.
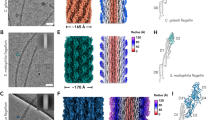
To obtain the native hook-filament junction tip (Fig. 5a), we knocked out the fliC gene to arrest flagellar assembly before filament elongation, preserving the flagellum in the pre-elongation state at the hook-filament junction. The native flagellar junction tip, as well as the filament tip as control, were mechanically shaved from the modified Salmonella strain and assessed by negative staining electron microscopy (Fig. 5b, c, and Supplementary Fig. 16a, b). The resulting cryo-EM density map revealed a clear structural arrangement, showing a FliD pentamer embedded within an 11-fold symmetric FlgL ring (Fig. 5d). Despite the moderate resolution of ~6.2 Å (Supplementary Fig. 16c), our determined FliD-FlgL structure at 5:5 stoichiometry, fits confidently within the density map. Additionally, six extra FlgL subunits could be docked onto the complex (Fig. 5d). The structure model of the hook-filament junction tip is depicted in Fig. 5e, f. 11 FlgL subunits are arranged in a pattern of five up-and-down pairs, with one extra subunit positioned at a downer step (Fig. 5e). The extra FlgL does not interact with or occupied by FliD (Fig. 5e), indicating the presence of an accessible site primed for the incorporation of the first flagellin subunit.
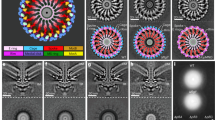
a Simplified schematic of the hook tip before flagellin elongation. b Representative 2D class averages of the native hook tip derived from Cryo-EM single particle 2D classification. c A representative mask slice of the hook tip was obtained from homogeneous refinement of the cryo-EM dataset with CryoSPARC76. d Model fitting of FliD and FlgL into the native Cryo-EM map of native flagellar hook tip, showing the determined 5:5 FliD-FlgL structure (colored as in previous figures) and six additional FlgL subunits (dark green). e, f Cartoon rendering of the model structure between 5 copies FliD in complex with 11 copies FlgL. FlgL positioning at “downer” conformation was highlighted with a green edge. g A simplified schematic of the hook tip before flagellin elongation. h A representative 2D class of the native filament tip derived from Cryo-EM single particle 2D classification. i A representative mask slice of the filament tip obtained from homogeneous refinement of the cryo-EM dataset with CryoSPARC. j Model fitting of FliD and FliC into the native Cryo-EM map of native flagellar filament tip. The determined FliD-FliC structure at 5:5 stoichiometry is colored in consistency. The extra 6 FlgL subunits were colored in light marine. The smeared density corresponding to the D3 domain of FliC in the “up” conformation results from a combination of limited particle numbers in this specific state and the intrinsic structural flexibility of the D3 domain itself. k, l Cartoon rendering of the model structure of FliD in complex with FliC at 5:11 stoichiometry. FliC positioning at “downer” conformation was also highlighted with a green edge.
Using a similar approach, we obtained a cryo-EM density map of the native filament tip (Fig. 5i). This map, resolved at 6.97 Å, revealed a comparable architecture for the FliD-FliC assembly at a 5:11 stoichiometry, akin to the FliD-FlgL complex (Fig. 5j, and Supplementary Fig. 16b, c). Identically, five additional FliC subunits bind to the FliD-FliC 5:5 complex at a secondary binding site, adopting a parallel down conformation (Fig. 5k, l). One additional FliC subunit is positioned in an even lower conformation adjacent to one of the five FliC subunits (Fig. 5k). This concave subunit exhibits no direct interaction with neighboring FliD molecules (Fig. 5k), leaving its binding surface exposed. This distinct configuration suggests that this subunit may serve as the site for newly translocated FliC during another round of flagellar elongation.
The in vitro complexes we characterized were observed at a 5:5 stoichiometry, rather than the 5:11 stoichiometry seen in native structures. We propose that the six additional subunits are stabilized through a combination of lateral interactions with adjacent FlgL/FliC subunits and weaker, dynamic contacts with secondary sites on FliD, collectively enabling the stable 5:11 architecture observed in native flagella. Despite attempts to stabilize this minor conformation through the design and screening of FliD-FlgL as well as FliD-FliC fusion constructs, we all failed likely due to the extremely transient nature of this interaction.
The transient complexes formation is required for flagellar assembly and bacteria motility
Structural dissection and comparison of the resolved complexes reveal key features of the dynamic, weak-affinity interactions governing flagellar filament components positioning by the filament cap FliD (Fig. 6a). FlgL and FliC occupy a shared binding site on FliD, revealing a conserved mechanism for substrate recognition by the filament cap. (Fig. 6a, b, and Supplementary Fig. 17a, b). Although the residues involved in FliD binding differ significantly between FlgL and FliC, one key interaction that dominantly contributes to the complex stability shows the same form and physicochemical property. Specifically, the hydrophobic side chains of FlgL L254 and FliC I72 interact similarly with hydrophobic groove forms by FliD Y296, A298, and L313 (Fig. 6b). The hydrophobic property of these residues is highly conserved in FliD, FlgL, and FliC (Fig. 6c). Noteworthy, polar and neutral residues distributed across the interface of both binary complexes form extended, complementary flat-strip interfaces that stabilize these ultra-weak interactions via van der Waals forces (Supplementary Fig. 8 and Fig. 13). In the native configurations, a second docking site was observed (Fig. 6b, and Supplementary Fig. 17a, b). Distinguished from the major docking site, no specific electrostatic or hydrophobic interactions were confirmed in the structural models, suggesting the significant contribution of shape complementarity and van der Waals interactions in mediating this interaction (Fig. 6b, and Supplementary Fig. 17c–e).
a Superposition of the FliD-FlgL and FliD-FliC complexes, with FliD rendered as a surface and FlgL (or FliC) as a cartoon. For clarity, only one subunit FlgL or FliC is depicted. b Cartoon representation of the overlaid FliD-FliC and FliD-FlgL complexes, highlighting the shared binding region of FliC and FlgL on FliD with magenta dash lines. The dark dashed lines indicate the overlapping region where the six additional subunits of FliC and FlgL interact with FliD. c Sequence conservation analysis of key interfacial residues. Panels from left to right show the conservation of residues in FliD (magenta), FlgL (orange), and FliC (teal) that are involved in the shared binding site. d Effect of amino acid alterations at the contact region on bacterial motility kinetics. Bar graphs represent the relative average colony diameter compared to wild-type strains, with individual data points representing relative diameters from each experiment. P-values were determined using a one-way ANOVA test (PRISM software, ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.1). Error bars represent standard deviations from at least 3 (FliD), 7 (FlgL), and 6 (FliC) independent replicates. Bars indicating mutations in FliD, FlgL, or FliC are colored magenta, dark yellow, and dark cyan, respectively. e Sequence conservation analysis of the second binding site on FliD for extra FlgL and FliC. f Effect of amino acid from second binding site substitutions on bacteria motility. Data are presented as mean motility diameter ratio values ± SD with at least 5 independent replicates. g Representative raw negative-staining electron microscope images of Salmonella with FliD, FlgL, FliC single point mutations. h Effect of single-point mutations in FliD, FlgL, or FliC on flagellar integrity, as assessed by statistical analysis of the percentage of normal length flagella (about 5 μm) from 10 randomly collected negative-stain images. All bacterial cells visible in each image were scored (total n > 15 cells per strain). Error bars indicate the standard deviation of flagella counts per cell.
To evaluate the functional significance of the observed interactions, we introduced key mutations that disrupt complex formation, as identified by our binding experiments (Figs. 3f and 4h), into corresponding Salmonella strains. In addition, residues located at the interface and conserved in bacteria species were also mutated (Fig. 6c, and Supplementary Figs. 18 and 19). The impact of these mutations on motility was quantified by measuring the colony diameter in motility assays. Mutations targeting key interaction residues, such as FliD Y296G, FliD L313G, FlgL L254G, and FliC I72G, exhibited statistically significant reductions in motility (Fig. 6d, and Supplementary Fig. 20a–c). Additionally, we assessed mutations at the second interface, where residues are relatively conserved (Fig. 6e, and Supplementary Fig. 18a). Mutations of these residues also led to a reduction in motility (Fig. 6f, and Supplementary Fig. 20d).
Conventional motility assays provide only a rough estimation of flagellar assembly. To directly and precisely visualize the effects of disrupted interaction on flagellum elongation, transmission electron microscopy with negative staining was used to image the flagella of wild-type and mutant Salmonella strains (Fig. 6g). Quantitative analysis revealed a significant reduction of fully assembled flagella in the mutant strains (Fig. 6h), highlighting the critical role of proper positioning of flagellar components by the flagellar cap in assembly and motility.
Structural plasticity governs FliD-mediated filament assembly
Our structural and biochemical analyzes establish that ultraweak interactions are essential for filament elongation and bacterial motility. To test whether modulating binding affinity affects assembly, we engineered structure-guided mutations intended to strengthen the FliD-FlgL and FliD-FliC interfaces (Fig. 7a, b). Quantitative pull-down assays confirmed up to a 3.5-fold enhancement in binding affinity for the most effective FliD-FlgL interface mutant (Fig. 7b, and Supplementary Fig. 21a). Beyond mutations designed to enhance hydrophobic or electrostatic interactions (e.g., V299I, D349R), we also identified key residues where substituting long side chains with glycine enhanced binding. This suggests these wild-type residues serve as stereospecific blockades that help maintain the weak interaction. Applying the same strategy to the FliD-FliC interface yielded affinity-enhancing mutants (Fig. 7c, d, and Supplementary Fig. 21b). Notably, all such mutants—whether targeting FliD-FlgL or FliD-FliC interactions—impaired bacterial motility to varying degrees (Fig. 7e, and Supplementary Fig. 22), demonstrating that finely tuned, low-affinity binding is critical for efficient filament assembly and function.
a FliD–FlgL interface (FliD, dark magenta; FlgL, dark orange) with key interface residues shown. b Relative binding affinities of FliD–FlgL interface mutants from pull-down assays and FliD–FliC mutants from NMR titrations (mean ± SD, n = 3–4). c FliD–FliC binding interface (FliD, magenta; FliC, cyan) highlighting residue FliC N269 and neighboring side‑chains. d Relative NMR peak‑intensity changes reflecting altered FliD–FliC binding affinity (mean ± SD, n = 5). e Motility impairment in Salmonella expressing affinity‑enhancing mutants of FliD (magenta), FlgL (orange), or FliC (cyan). Data show relative mean swim diameter ± SD from at least 6 (FliD), 9 (FlgL) and 8 (FliC) replicates; with ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.1 (one‑way ANOVA). f Top view of the FliD pentamer with interface residues A91 and A168 (Cα–Cα distance 4.3 Å). g, h SDS pages and images of EM negative stain for FliD2C mutant (A91C/A168C) in condition with or without reducing agent DTT. i Schematic of the disulfide‑stabilized FliD cap in the flagellar T3SS. j Negative‑stain EM images of Salmonella expressing FliD or FliD2C mutant. k Motility of strains expressing wild‑type FliD or FliD2C with or without inducer IPTG. l Stepwise model of cap‑mediated filament elongation. The pentameric FliD cap (magenta) docks onto the FlgL ring (yellow) (1). The first FliC subunit binds the exposed ‘down’ FlgL site (2), followed by sequential FliC incorporation into gaps between FliD subunits (3,4). After five subunits are inserted, competition with FlgL drives cap rotation (5), freeing new sites for further FliC addition (6–8). This cycle repeats, extending the filament.
Given the dynamic nature of the FliD-FliC/FlgL complex during assembly, disulfide bond crosslinking was used to examine the rigidity of the FliD pentamer cap in filament extension. Structural analysis revealed A91 and A168 at adjacent subunit interfaces (Cβ-Cβ distance: 4.3 Å; Fig. 7f). We generated a disulfide-locked FliD variant (FliD2C: A91C/A168C) that formed covalent pentamers upon oxidation (Fig. 7g, h). SDS-PAGE under non-reducing conditions confirmed >40% crosslinking efficiency (Fig. 7g), while negative-stain EM showed intact cap structures indistinguishable from wild-type (Fig. 7h, i). Critically, Salmonella expressing FliD2C exhibited no significant changes of flagellar ultrastructure and bacteria motility. These results demonstrate that the pentameric conformation remains static during elongation and that oligomer rigidity—not dynamic reorganization—enables filament in vivo assembly.
Discussion
The bacterial flagellum is a highly conserved and complex appendage essential for various physiological processes, including cell motility, pathogenesis, biofilm formation, and environmental adaptation. As a molecular nanomachine, the flagellum enables bacterial locomotion via a filamentous propeller driven by a membrane-embedded rotary motor. The assembly of this propeller, which consists of multiple filament subunits, is a highly dynamic process requiring precise molecular coordination. At the starting point of the axial assembly, flagellin FliC is transported through the central channel of the hook and integrated at the interface between the filament cap (FliD) and the hook-filament junction (FlgL). Although the significance of the filament cap in positioning and elongating flagellin has been previously recognized, the detailed structural and mechanistic understanding of this process has remained elusive. Herein, we present atomic-level insight into the structural features underlying the molecular mechanisms by which the flagellar filament cap orchestrates other filament components.
Our structural analysis demonstrates that the transient complexes formation during flagellar filament assembly is predominantly stabilized by conserved hydrophobic and van der Waals interactions. Despite the relatively large interaction surfaces, direct pairwise interactions between specific residues are minimal, highlighting the dynamic nature of the assembly process. Key residues, such as FliD Y296 and L313, FlgL L256, and FliC I72, are critical for stabilizing these interactions and are highly conserved across bacterial species. In contrast, surrounding residues at the interface are smaller, hydrophilic, and less conserved, contributing to the conformational flexibility of the complexes. This flexibility suggests that multiple conformational states of the assembly intermediates coexist in solution, with the dominant conformation being the most stable and populated observed in our experiments. These states likely correspond to different stages of the transient FliD-FliC and FliD-FlgL complexes during filament growth. The ultra-transient nature of these interactions poses challenges for capturing the minor conformations at near-atomic resolution, which may require further exploration using native flagellar structures at high resolution in different assembly states.
In addition to the major conformations, our cryo-EM density maps of the native hook-filament junction and filament tip uncovered alternative binding configurations. No direct pairwise interaction was discovered in the alternative binding site, confirming an even weaker interaction than the major conformations, which reinforces the hypothesis that transient, ultra-weak interactions dominate the flagellar assembly process. Intervening this ultra-weak interaction did not significantly impair flagellar filament elongation or bacteria motility, suggesting that high-affinity binding is not essential for efficient filament assembly. Instead, these ultra-transient interactions may facilitate rapid flagellin incorporation, enhancing both the speed and flexibility of the assembly process. Similar ultra-weak interactions were also discovered in other protein nanomachines, such as bacteria pilus inter-subunits and the components of the pathogenic type three secretion system, where such weak interactions enable the assembly of the large nanomachines63,64.
A key finding of this study is the structural organization of the filament cap (FliD) and its role in coordinating the assembly of other filament components (FlgL and FliC) with a stoichiometry mismatch mechanism. FliD forms a pentameric, stool-like structure, with each “leg” interacting transiently with FlgL or FliC in an alternating up-down configuration. In the “up” state, FlgL or FliC docks onto FliD at a 5:5 stoichiometry, while in the “down” state, five FlgL subunits bind to a secondary site on FliD in a highly dynamic manner. Notably, an extra FlgL or FliC subunit adopts a distinct down configuration, remaining unbound to adjacent subunits, which provides an accessible site for subsequent flagellin incorporation.
These structural insights suggest a mechanism for initiating filament assembly at the hook-filament junction (Fig. 7i), providing structural and biophysical validation for the rotating cap model first proposed by Namba and coworkers31,37. At this critical juncture, the filament cap integrates with the FlgL ring, and the first flagellin (FliC) subunit binds specifically to the available FlgL subunit, guided by the hydrophobic interactions at the C-terminal region of FliD. This initial FliC binding event triggers a cascade of flagellin incorporation, where subsequent FliC subunits are sequentially positioned into the gaps between the legs of FliD in a left-handed spiral configuration. Once the five gaps are incorporated with FliC subunits, they compete with FlgL in “up” conformations for binding on FliD, driving the cap’s rotation and freeing additional gaps for further flagellin incorporation. This coordinated process continues with successive subunits competing for interactions with FliD, driving FliD’s rotation along the filament’s growth direction and extending the filament. This intricate, stepwise mechanism ensures the rapid and precise elongation of the filament, essential for bacterial motility.
While we characterize the interactions between monomeric FliD substrates as ultra-weak, it is important to consider the context of the native flagellum. The pentameric nature of the FliD cap likely creates a significant avidity effect, multiplying the overall binding strength for the undecameric FlgL or FliC ring. This avidity would provide the necessary stability for the cap to remain positioned at the growing tip, while the individual monomer-monomer interactions remain weak enough to allow rapid, low-energy subunit exchange during processive elongation.
In summary, our findings provide atomic-level insights into the molecular mechanisms underlying bacterial flagellar assembly, emphasizing the importance of transient, low-affinity interactions in driving filament subunits’ rapid and precise incorporation. This mechanism likely represents a general strategy employed by various bacterial nanomachines to achieve efficient assembly with minimal energy expenditure. These insights pave the way for future research into how such weak interactions can be exploited for therapeutic interventions targeting bacterial motility.
Methods
Bacteria strains, plasmids, and media
The bacterial strains utilized in this study are derived from Salmonella enterica serovar Typhimurium strain LT2 (also known as SGSC1412/ATCC 700720). Detailed information on the plasmids and primers used is provided in Supplementary Table 2. Bacteria were cultivated aerobically at 37 °C in liquid cultures as Luria-Bertani (LB) media, Terrific-Broth (TB) media, and the minimum media (M9). LB media contains Baco-Tryptone 10 g, yeast extract 5 g and NaCl 10 g per liter. One liter TB contains Baco-Tryptone 12 g, yeast extract 24 g, glycerol 4 mL, KH2PO4 2.3 g, and K2HPO4 12.5 g. M9 media contains the minimum necessary nine kinds of substances for bacteria growth, which are NH4Cl (or U-15N- NH4Cl) 1 g/L, Na2HPO4 6.78 g/L, KH2PO4 3.0 g, NaCl 0.5 g/L, MgSO4 0.24 g/L, CaCl2 0.01 g/L, glucose (or U-13C-glucose) 2 g/L, vitamin B 0.01 g/L and H2O 1 L. Antibiotic selection for plasmid maintenance, with ampicillin at 100 µg/mL, was applied to the cultures immediately before the bacterial transfer.
The λ-RED system was carried out to obtain scarless gene knockout from bacteria chromosomes through chromosomal recombination65. This process was conducted according to the methods reported in the literature. Mutant genes were introduced into bacteria stains with pBAD, or pTrc plasmid, and gene expression was induced with 0.1% arabinose or 50 µM IPTG, respectively.
Protein expression and purification
All the genes of the flagellar proteins were derived from Salmonella Typhimurium STM1913/700720 (ATCC). The DNA sequences encoding FlgL, FliD, FliC, as well as domains of these three proteins, were amplified with PCR and then inserted into pET16b vector with an N-terminal 8XHis-MBP-TEV (Tobacco etch virus protease site) tag or 8XHis-TEV tag. The fusion constructs for structure determination were prepared by fusing FlgL or FliC to the N-terminus of FliD with a highly flexible and disordered linker, GGS-linker. The amino acid sequence of the GGS-linker is GGSGSGSLVPRGGSGGSGGS. All mutants were made by site-directed single-point mutagenesis with PhusionR high-fidelity and efficiency DNA polymerase (NEB). Primers related to making all the plasmids are collected and listed in Supplementary Table 1. Non-isotopically labeled proteins were expressed with BL21(DE3) cells grown in Terrific-Broth (TB) or LB medium in the presence of ampicillin (100 mg/ml−1). Protein expression was induced at 16 °C with 0.5 mM isopropyl-beta-D-1-thiogalactopyranoside (IPTG) at OD600 ≈ 0.6 for about 36 h. Cells were collected at OD600 ≈ 1.8 by centrifugation at 6000 × g for 30 min at 4 °C and stored at −20 °C until purification.
For protein purification, cells were thawed and resuspended in lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, and 10 mM imidazole) and disrupted by a high-pressure homogenizer till the solvent became clear. The lysed cells were centrifuged at 20,000 × g for 1 h to remove the precipitant. The supernatant was passed through a 5 ml Ni Sepharose 6 Fast Flow column (GE Healthcare) pre-washed with lysis buffer. Non-specific bound proteins were removed by washing the Ni column with above 25 ml wash buffer (50 mM Tris-HCl pH 8.0, 500 mM NaCl, and 30 mM imidazole). The target proteins were eluted with about 20 ml elution buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, and 500 mM imidazole). The eluted proteins were subjected to TEV protease cleavage for tag removal by overnight dialysis with TEV cleavage buffer (10 mM Tris-HCl, pH 8.0, 5 mM beta-mercaptoethanol), followed by a second round of Ni column purification to remove the His-MBP or His tag from the supernatant. The flow-through was concentrated to less than 5 ml and injected into a HiLoad 26/600 Superdex 200 size exclusion column (GE Healthcare). The size exclusion buffer contains 20 mM Tris-HCl, pH 8.0, and 150 mM NaCl. The peak fraction was further purified by another round of size exclusion with analytical columns, including Superdex Increase 200 10/300 column (GE Healthcare) and SuperoseTM 6 column (GE Healthcare) with the same buffer. Purified proteins were assessed by SDS-PAGE analysis and concentrated in Amicon filters (Millipore). Protein concentration was spectrophotometrically determined with Lambert-Beer law using the UV absorbance at 280 nm and the corresponding extinction coefficient. The purified proteins were subject to Cryo-EM grid preparation or stored at −80 °C. All proteins were purified by the same methods.
Protein isotope labeling for solution NMR characterization
15N labeled FliDd1, FliDd2d3 and FliDd1 mutants were expressed in BL21(DE3) cells and grown in minimal medium (M9) supplied with 1 g/L U-[15N]-NH4Cl. 15N-13C double labeled FliDd1 sample was prepared by supplementing the M9 medium with 1 g/L U-[15N]-NH4Cl and 2 g/L U-[13C]-glucose. Cells were induced at around OD600 = 0.9 with 0.5 mM IPTG at 16 °C for about 24 h. Typically, isotope-supplied cells were harvested at OD600 above 1.2 and stored at −20 °C till purification. Purification of isotope-labeled proteins was conducted the same as unlabeled protein samples except for storage buffer (50 mM NaPi, pH 6.8, 150 mM NaCl, 0.05% NaN3). U-[15N]-NH4Cl and U-[13C]-glucose were purchased from Cambridge Isotope Laboratories and Isotec.
NMR spectroscopy of triple resonance and NMR titration
The NMR triple resonance and NMR titration experiments were carried out on Bruker AVANCE III 600 MHz and Bruker AVANCE III 800 MHz instruments, respectively, equipped with cryogenic probes. All the NMR spectra were recorded at 25 °C and processed with NMRPipe and further analyzed with Sparky. The protein samples for NMR experiments were prepared in 50 mM NaPi, pH 6.8, 150 mM NaCl, 0.05% NaN3, and 10% deuterated D2O. Backbone resonance assignment of FliDd1 was achieved using a 15N-13C labeled sample at 0.8 mM with a series of triple resonance experiments, including HNCA, CBCACONH, and HNCACB all with the gpwg3d pulse program. Each three-dimensional (3D) experiment was performed with 64 scans with d1 = 1.5 s and 2D experiments were conducted with 12 scans with the same d1 time. The single-point NMR titration experiments were performed with 0.2 mM 15N labeled protein (FliDd1, FliDd2d3, or FliDd1 mutant) titrated with 0.2 mM unlabeled proteins (FlgL, FliC or FliC mutant). Residues located at the binding interface were screened and confirmed by comparing the peak intensities described by recording and analyzing the 1H-15N-HSQC spectra.
To determine the binding affinity of interaction between FliD and FlgL as well as FliC, we performed NMR titrations using 15N-labeled FliDd1 (0.2 mM) with increasing amounts of FlgL (0 mM, 0.05 mM, 0.10 mM, 0.15 mM, 0.2 mM, 0.3 mM, 0.4 mM, 0.6 mM) or FliC (0 mM, 0.1 mM, 0.3 mM, 0.5 mM, 1.0 mM, 1.5 mM, 2.0 mM). 2D-HSQC spectrum was applied to acquire the intensity value of each peak corresponding to substrate adding. The KD values for these titrations were obtained by fitting the data with a one-site-specific binding equation. Origin 8.5 was used for data fitting and figure presentation.
Negative stain electron microscopy
Negative staining was used to check protein sample homogeneity and visualize bacteria and flagellum in situ. Protein samples were diluted to 10 μg/ml and loaded onto glow-discharged carbon-film 400 mesh copper grids. In comparison, bacteria samples were diluted to OD600 (optical density at 600 nm wavelength) ≈ 0.05. After 90 s of incubation at room temperature, the extra solution on the grid surface was dried with filter paper. The sample-loaded grid was then stained and dried gradually with 5 μL saturated uranyl acetate for 10 s, 10 s, and 30 s. Finally, the grids were air-dried for 10 min and loaded to a 120 keV transmission electron microscope (FEI Talos 120).
Cryo-EM sample preparation, data collection, and process
Proteins for cryo-EM characterization were purified and concentrated to above 8 mg/mL. Before making the Cryo-EM grids, protein samples were diluted to 1 mg/ml, 2 mg/ml, 4 mg/ml, and 8 mg/ml. Grid screening was performed by making four Cryo-EM grids corresponding to the different protein concentrations. Cryo-EM grids were prepared using a Vitrobot Mark Ⅳ system (FEI), by adding a 3 μL protein sample to the glow-discharged Quantifoil R1.2/1.3 holey carbon gold grids. Grid freezing was conducted with a blot time of 1.5 s, a blot force of -5, and a wait time of 15 s under 95% humidity at 4 °C.
Cryo-EM data was collected using a 300 keV Titan Krios electron microscope (FEI) equipped with a K3 direct electron detector (Gatan). Cryo-EM images were recorded in super-resolution mode with EPU66 at a nominal magnification of 105,000× in a physical pixel size of 0.85 Å/pixel and a defocus range from −1.0 μm to −2.5 μm. A dose rate of about 30 e/Å2·s and an exposure time of 2.5 s were applied with 40 movie frames. A total of 3000, 600, and 900 movie stacks were collected for FliD, FlgL-FliD, FliC-FliD, respectively. The datasets were processed in CryoSPARC (version 4.2.1)67,68. Briefly, the raw movie stacks were imported to CryoSPARC, aligned with Patch Motion Correction, followed by contrast transfer function (CTF) estimation using Patch CTF Estimation and Exposure Curation to remove bad movies. Detailed overflow of the cryo-EM data processing for each sample was shown in Supplementary Figs. 2, 7 and 11 and summarized in Supplementary Table 1.
Model building and refinement
The initial models of salmonella FliD, FlgL, and FliC were obtained from the Alphafold webserver with the ID: AF-P16328-F1 [https://alphafold.ebi.ac.uk/entry/P16328], AF-P16326-F1 [https://alphafold.ebi.ac.uk/entry/P16326] and AF-P06179-F1 [https://alphafold.ebi.ac.uk/entry/P06179], respectively. Buried surface areas were calculated using the PISA server (https://www.ebi.ac.uk/msd-srv/prot_int/cgi-bin/piserver). UCSF ChimeraX69 was used to dock the initial models into the cryo-EM maps. From this starting point, manual rebuilding was conducted in Coot70. Further refinements were performed with several cycles of real space refinement in Phenix71 and Coot iteratively. Final structures were uploaded to Molprobility72 server for quality confirmation. Figures for structure presentation were prepared by PyMOL, UCSF Chimera73, and UCSF ChimeraX69. Statistical data of structural refinement was summarized in Supplementary Table 2.
Pull-down assay
To determine the effect of mutations on FlgL-FliD stability, we performed pull-down assays to test the relative binding affinity change between wild-type proteins and their mutants. Protein samples used for this experiment were desalted into the binding buffer (25 mM Tris-HCl and 150 mM NaCl at pH 8.0). Briefly, His-FlgL or its mutants was incubated with untagged FliD or its mutants at a 1:1 concentration ratio (0.03 mM, 4 mL) for 0.5 h at 4 °C. The mixtures were injected into a 1 mL His Trap HP column (Cytiva) pre-washed with binding buffer. Then the column was washed with wash buffer containing 50 mM Tris-HCl and 150 mM NaCl at pH 7.4 for 10 times column volume. Finally, the column was eluted with 1 mL elution buffer (50 mM Tris-HCl, 150 mM NaCl, 500 mM imidazole, pH 7.4). 5 μL of the elution mixed with loading buffer were loaded into a 15% SDS-PAGE gel. All the pulldown assays were repeated three parallel times. Quantitative analysis of the pulldown assay was performed by ImageJ and the figures were presented with Prism 10.0.
Preparation of the flagellar Hook-Junction-Cap and Filament-Cap
Salmonella △ flic and wild-type strains were used to prepare flagellar Hook-Junction-Cap and Filament-Cap, respectively. Bacterial cells were grown in 5 L of M9 media at 30 °C with shaking at 30 × g overnight until the OD600 reached around 1.0. Cells were harvested and collected by centrifugation at 3000 × g for 30 min at 4 °C. For Hook-Junction-Cap preparation, the cell pellet was suspended with 50–100 ml lysis buffer (50 mM Tris-HCl,150 mM NaCl, pH 7.4) and then loaded into a homogenizer to lysate the bacteria. The supernatant was collected by centrifugation at 20,000 × g for 30 min at 4 °C. To purify the flagellar hook-junction-cap component, the supernatant was desalted into the low salt buffer (50 mM Tris-HCl, pH 8.0) and injected into a DEAE Sepharose fast flow anion exchange column (GE Healthcare). A gradient from 0% to 100% elution buffer (50 mM Tris-HCl, 1 M NaCl, pH 8.0) was applied to separate impurities and collected elution for 2 ml per tube. Negative staining was conducted to check the samples tube by tube. Further purification was performed by using cation exchange with a column packed with sourceS resin (GE Healthcare), followed by negative staining to check the samples tube by tube. The final step was to concentrate the target sample and load it into a Superose 6 Increase size exclusion column pre-washed with buffer containing 20 mM Tris-HCl, 150 mM NaCl, pH 7.4. The sample purity of each fraction under visible peaks was checked by negative staining. The purified Hook-Junciton-Cap sample was concentrated to around 8 mg/ml and prepared to make Cryo-EM grids. For Filament-Cap preparation, collected cell pellets were gently suspended with 50–100 ml fresh M9 culture and stirred overnight at 16 °C to cut the flagella tip from the bacteria surface. The bacteria cells were removed by centrifugation at 6000 × g for 30 min. 50% (NH4)2SO4 was added to the supernatant to precipitate the flagellar filament from soluble impurities. The precipitant was collected by another round of centrifugation at 20,000 × g for 30 min and resuspended with buffer containing 20 mM Tris-HCl, 150 mM NaCl, pH 7.4. The final step of purification was the same as the Hook-Junction-Cap sample. Cryo-EM grids of the Filament-Cap sample were prepared as others and stored in liquid nitrogen till use.
Motility assay
Motility assays were conducted following established protocols with some modifications. In brief, Salmonella strains with specific gene knockout (flid-, flgl-, flic-) were transformed with pBAD plasmids carrying wild-type genes or their mutants. Fresh overnight cultures (OD600 = 0.5, 1 μL) were stabbed in 0.2% LB-agar plates (0.3% agar, 1%Tryptone, 0.5% Nacl, ampicillin 100 μg/mL, 0.125% arabinose and 0.6% glucose). The plates were incubated for 6 h at 37 °C. After incubation, images of the plates were captured using a LAS 4000 imaging system (GE Healthcare), and the colony diameters were measured with ImageJ software. Each strain was tested in at least three independent replicates. The data were analyzed and presented graphically using Prism 10.
Flagella length measurements
Salmonella strains containing mutant plasmids were grown overnight in LB media containing the appropriate anti-biotics and 0.125% arabinose and shaken at 30 rpm, at 37 °C. Cells were collected by centrifugation at 1000 × g for 10 min, and the pellet was resuspended in 500 µl of M9 medium. Next, 5 µl of bacterial suspension were placed on a freshly glow-discharged carbon-covered electron microscopic support grid for 1 min. The grid was washed twice with PBS buffer and stained for 1 min with an aqueous solution of 0.75% uranyl acetate. The grids were operated on a 120 keV FEI Tecnai 20 electron microscope (Thermo Fisher Scientific). Flagella were counted for 20–40 cells in each sample as indicated in the Figure legends. Use image software to count the number and length of flagella in the sample. Each analysis was repeated a minimum of three times.
Statistics and reproducibility
Pearson’s correlation coefficient (P value) of statistical analysis was conducted with GraphPad Prism 10.0 software. The P-values were determined with one-way ANOVA followed by Dunnett’s multiple comparisons test using wild-type protein or strains as control. Experiments related to statistics data were repeated in at least three independent preparations. Error bars indicate the standard error of the mean (n = 3, **P < 0.001, ***P < 0.0001). Source data are provided as a Source Data file. Each statistical analysis was performed on data obtained from at least three independent biological replicates. For representative electron microscopy images, more than five randomly selected micrographs were acquired per condition from each independent replicate.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Atomic coordinates for the Cryo-EM structures and correlated density maps have been deposited in the Protein Data Bank and Electron Microscopy Data Base, respectively, under accession number as: PDB:9M6I and EMDB: 63663; PDB:9M68 and EMDB: 63661; PDB:9M67 and EMDB: 63660; PDB:9M6H and EMDB:63662. NMR data have been deposited to Biological Magnetic Resonance Bank under accession number 53375. Source data for the pull-down assay, motility assay, and flagellar filament statistics is included in Source Data files. Raw images of the SDS-PAGE gels are provided in Supplementary Fig. 23. Source data are provided as a Source Data file. Source data are provided with this paper.
References
Wadhwa, N. & Berg, H. C. Bacterial motility: machinery and mechanisms. Nat. Rev. Microbiol. 20, 161–173 (2022).
Chang, Y. J., Carroll, B. L. & Liu, J. Structural basis of bacterial flagellar motor rotation and switching. Trends Microbiol. 29, 1024–1033 (2021).
Hu, H. D. et al. Structural basis of torque generation in the bi-directional bacterial flagellar motor. Trends Biochem. Sci. 47, 160–172 (2022).
Chen, S. Y. et al. Structural diversity of bacterial flagellar motors. EMBO J. 30, 2972–2981 (2011).
Xie, L., Altindal, T., Chattopadhyay, S. & Wu, X. L. From the cover: Bacterial flagellum as a propeller and as a rudder for efficient chemotaxis. Proc. Natl. Acad. Sci. USA 108, 2246–2251 (2011).
Worrall, L. J., Majewski, D. D. & Strynadka, N. C. J. Structural insights into type III secretion systems of the bacterial flagellum and injectisome. Annu. Rev. Microbiol. 77, 669–698 (2023).
Chaban, B., Hughes, H. V. & Beeby, M. The flagellum in bacterial pathogens: for motility and a whole lot more. Semin. Cell Dev. Biol. 46, 91–103 (2015).
Minamino, T. & Kinoshita, M. Structure, assembly, and function of flagella responsible for bacterial locomotion. EcoSal 11, eesp00112023 (2023).
Macnab, R. M. Type III flagellar protein export and flagellar assembly. Biochim. Biophys. Acta 1694, 207–217 (2004).
Chevance, F. F. & Hughes, K. T. Coordinating assembly of a bacterial macromolecular machine. Nat. Rev. Microbiol. 6, 455–465 (2008).
Macnab, R. M. How bacteria assemble flagella. Annu. Rev. Microbiol. 57, 77–100 (2003).
Deng, W. et al. Assembly, structure, function and regulation of type III secretion systems. Nat. Rev. Microbiol. 15, 323–337 (2017).
Costa, T. R. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359 (2015).
Minamino, T. & Namba, K. Distinct roles of the FliI ATPase and proton motive force in bacterial flagellar protein export. Nature 451, 485–488 (2008).
Erhardt, M., Mertens, M. E., Fabiani, F. D. & Hughes, K. T. ATPase-independent type-III protein secretion in Salmonella enterica. PLoS Genet. 10, e1004800 (2014).
Hirano, T., Yamaguchi, S., Oosawa, K. & Aizawa, S. Roles of FliK and FlhB in determination of flagellar hook length in Salmonella typhimurium. J. Bacteriol. 176, 5439–5449 (1994).
Erhardt, M., Singer, H. M., Wee, D. H., Keener, J. P. & Hughes, K. T. An infrequent molecular ruler controls flagellar hook length in Salmonella enterica. EMBO J. 30, 2948–2961 (2011).
Shibata, S. et al. FliK regulates flagellar hook length as an internal ruler. Mol. Microbiol. 64, 1404–1415 (2007).
Samatey, F. A. et al. Structure of the bacterial flagellar hook and implication for the molecular universal joint mechanism. Nature 431, 1062–1068 (2004).
Matsunami, H., Barker, C. S., Yoon, Y. H., Wolf, M. & Samatey, F. A. Complete structure of the bacterial flagellar hook reveals extensive set of stabilizing interactions. Nat. Commun. 7, 13425 (2016).
Kato, T., Makino, F., Miyata, T., Horvath, P. & Namba, K. Structure of the native supercoiled flagellar hook as a universal joint. Nat. Commun. 10, 5295 (2019).
Makishima, S., Komoriya, K., Yamaguchi, S. & Aizawa, S. I. Length of the flagellar hook and the capacity of the type III export apparatus. Science 291, 2411–2413 (2001).
Shaikh, T. R. et al. A partial atomic structure for the flagellar hook of Salmonella typhimurium. Proc. Natl. Acad. Sci. USA 102, 1023–1028 (2005).
Bulieris, P. V., Shaikh, N. H., Freddolino, P. L. & Samatey, F. A. Structure of FlgK reveals the divergence of the bacterial Hook-Filament Junction of Campylobacter. Sci. Rep. 7, 15743 (2017).
Evans, L. D., Poulter, S., Terentjev, E. M., Hughes, C. & Fraser, G. M. A chain mechanism for flagellum growth. Nature 504, 287–290 (2013).
Zhao, Z. et al. Frequent pauses in Escherichia coli flagella elongation revealed by single cell real-time fluorescence imaging. Nat. Commun. 9, 1885 (2018).
Kreutzberger, M. A. B. et al. Convergent evolution in the supercoiling of prokaryotic flagellar filaments. Cell 185, 3487–3500.e3414 (2022).
Fujii, T., Iwane, A. H., Yanagida, T. & Namba, K. Direct visualization of secondary structures of F-actin by electron cryomicroscopy. Nature 467, 724–728 (2010).
Evans, L. D., Hughes, C. & Fraser, G. M. Building a flagellum outside the bacterial cell. Trends Microbiol. 22, 566–572 (2014).
Calladine, C. R., Luisi, B. F. & Pratap, J. V. A “mechanistic” explanation of the multiple helical forms adopted by bacterial flagellar filaments. J. Mol. Biol. 425, 914–928 (2013).
Yonekura, K. et al. The bacterial flagellar cap as the rotary promoter of flagellin self-assembly. Science 290, 2148–2152 (2000).
Yonekura, K., Maki-Yonekura, S. & Namba, K. Structure analysis of the flagellar cap-filament complex by electron cryomicroscopy and single-particle image analysis. J. Struct. Biol. 133, 246–253 (2001).
Green, A. G. et al. Large-scale discovery of protein interactions at residue resolution using co-evolution calculated from genomic sequences. Nat. Commun. 12, 1396 (2021).
Ikeda, T., Homma, M., Iino, T., Asakura, S. & Kamiya, R. Localization and stoichiometry of hook-associated proteins within Salmonella typhimurium flagella. J. Bacteriol. 169, 1168–1173 (1987).
Homma, M. & Iino, T. Locations of hook-associated proteins in flagellar structures of Salmonella typhimurium. J. Bacteriol. 162, 183–189 (1985).
Homma, M., DeRosier, D. J. & Macnab, R. M. Flagellar hook and hook-associated proteins of Salmonella typhimurium and their relationship to other axial components of the flagellum. J. Mol. Biol. 213, 819–832 (1990).
Maki, S., Vonderviszt, F., Furukawa, Y., Imada, K. & Namba, K. Plugging interactions of HAP2 pentamer into the distal end of flagellar filament revealed by electron microscopy. J. Mol. Biol. 277, 771–777 (1998).
Maki-Yonekura, S., Yonekura, K. & Namba, K. Conformational change of flagellin for polymorphic supercoiling of the flagellar filament. Nat. Struct. Mol. Biol. 17, 417–422 (2010).
Hong, H. J. et al. Crystal structure of FlgL and its implications for flagellar assembly. Sci. Rep. 8, 14307 (2018).
Al-Otaibi, N. S. et al. The cryo-EM structure of the bacterial flagellum cap complex suggests a molecular mechanism for filament elongation. Nat. Commun. 11 https://doi.org/10.1038/s41467-020-16981-4 (2020).
Song, W. S., Cho, S. Y., Hong, H. J., Park, S. C. & Yoon, S. I. Self-oligomerizing structure of the flagellar cap protein FliD and its implication in filament assembly. J. Mol. Biol. 429, 847–857 (2017).
Cho, S. Y. et al. Tetrameric structure of the flagellar cap protein FliD from Serratia marcescens. Biochem. Biophys. Res. Commun. 489, 63–69 (2017).
Cho, S. Y., Song, W. S. & Yoon, S. I. Crystal structure of the flagellar cap protein FliD from Bdellovibrio bacteriovorus. Biochem. Biophys. Res. Commun. 519, 652–658 (2019).
Song, W. S., Hong, H. J. & Yoon, S. I. Structural study of the flagellar junction protein FlgL from Legionella pneumophila. Biochem. Biophys. Res. Commun. 529, 513–518 (2020).
Einenkel, R. et al. The structure of the complete extracellular bacterial flagellum reveals the mechanism of flagellin incorporation. Nat. Microbiol. 10, 1741–1757 (2025).
Wang, F. et al. A structural model of flagellar filament switching across multiple bacterial species. Nat. Commun. 8, 960 (2017).
Kreutzberger, M. A. B. et al. Flagellin outer domain dimerization modulates motility in pathogenic and soil bacteria from viscous environments. Nat. Commun. 13, 1422 (2022).
Xu, Z., Knafels, J. D. & Yoshino, K. Crystal structure of the bacterial protein export chaperone secB. Nat. Struct. Biol. 7, 1172–1177 (2000).
Huang, C., Rossi, P., Saio, T. & Kalodimos, C. G. Structural basis for the antifolding activity of a molecular chaperone. Nature 537, 202–206 (2016).
Xing, Q. et al. Visualizing an ultra-weak protein-protein interaction in phosphorylation signaling. Angew. Chem. Int. Ed. Engl. 53, 11501–11505 (2014).
Huang, C. & Kalodimos, C. G. Structures of large protein complexes determined by nuclear magnetic resonance spectroscopy. Annu. Rev. Biophys. 46, 317–336 (2017).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Rowe, A. J. Ultra-weak reversible protein-protein interactions. Methods 54, 157–166 (2011).
Johansson, H. et al. Specific and nonspecific interactions in ultraweak protein-protein associations revealed by solvent paramagnetic relaxation enhancements. J. Am. Chem. Soc. 136, 10277–10286 (2014).
Jiang, Y., Rossi, P. & Kalodimos, C. G. Structural basis for client recognition and activity of Hsp40 chaperones. Science 365, 1313–1319 (2019).
Hu, J. et al. Structural basis of bacterial effector protein azurin targeting tumor suppressor p53 and inhibiting its ubiquitination. Commun. Biol. 6, 59 (2023).
Ikeda, T., Yamaguchi, S. & Hotani, H. Flagellar growth in a filament-less Salmonella fliD mutant supplemented with purified hook-associated protein 2. J. Biochem. 114, 39–44 (1993).
Lian, L. Y. NMR studies of weak protein-protein interactions. Prog. Nucl. Magn. Reson Spectrosc. 71, 59–72 (2013).
Xing, Q. et al. Structures of chaperone-substrate complexes docked onto the export gate in a type III secretion system. Nat. Commun. 9, 1773 (2018).
Shibata, S., Matsunami, H., Aizawa, S. I. & Wolf, M. Torque transmission mechanism of the curved bacterial flagellar hook revealed by cryo-EM. Nat. Struct. Mol. Biol. 26, 941–945 (2019).
Tan, J. et al. Structural basis of assembly and torque transmission of the bacterial flagellar motor. Cell 184, 2665–2679 e2619 (2021).
Jones, C. J., Macnab, R. M., Okino, H. & Aizawa, S. Stoichiometric analysis of the flagellar hook-(basal-body) complex of Salmonella typhimurium. J. Mol. Biol. 212, 377–387 (1990).
Gossert, A. D. et al. NMR structure of the Escherichia coli type 1 pilus subunit FimF and its interactions with other pilus subunits. J. Mol. Biol. 375, 752–763 (2008).
Rathinavelan, T. et al. NMR model of PrgI-SipD interaction and its implications in the needle-tip assembly of the Salmonella type III secretion system. J. Mol. Biol. 426, 2958–2969 (2014).
Blank, K., Hensel, M. & Gerlach, R. G. Rapid and highly efficient method for scarless mutagenesis within the Salmonella enterica chromosome. PLoS ONE 6, e15763 (2011).
Thompson, R. F., Iadanza, M. G., Hesketh, E. L., Rawson, S. & Ranson, N. A. Collection, pre-processing and on-the-fly analysis of data for high-resolution, single-particle cryo-electron microscopy. Nat. Protoc. 14, 100 (2019).
DiIorio, M. C. & Kulczyk, A. W. A robust single-particle cryo-electron microscopy (cryo-EM) processing workflow with cryoSPARC, RELION, and Scipion. J. Vis. Exp. https://doi.org/10.3791/63387 (2022).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in. Acta Crystallogr. D 75, 861–877 (2019).
Chen, V. B. et al. All-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Yang, Z. et al. UCSF Chimera, MODELLER, and IMP: an integrated modeling system. J. Struct. Biol. 179, 269–278 (2012).
Hagemans, D., van Belzen, I. A., Moran Luengo, T. & Rudiger, S. G. A script to highlight hydrophobicity and charge on protein surfaces. Front. Mol. Biosci. 2, 56 (2015).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (Grant NO.32371277 to Qiong Xing, Grant NO.32301028 to Wenxue Jiang), the Project of Technological Innovation Plan in Hubei Province (2024BCA001 to Qiong Xing) and Hubei Province Science and Technology Plan Project (2021CSA066 to Qiong Xing). The authors acknowledge all the staff, especially Dr Qianjun Chen and Hong yang, in cryo-EM center and computational center of Hubei University. NMR spectra were acquired at the National Center for Magnetic Resonance in Wuhan.
Author information
Authors and Affiliations
Contributions
Q.X. conceived the project and designed all the experiments. L.X.C., F.F.Z., and X.W. performed the experiments including sample preparation and all the assays. X.Q.C. and W.X.J. prepared the cryo-EM sample and performed the grid screening and data collection. X.D. and W.X.J. performed the NMR experiments and analysis the data. Q.X. and W.X.J. carried out the cryo-EM data process, model building and refinement. Q.X., L.X.C., F.F.Z., and X.D. wrote the paper. L.X.M. contributed to critically reading and helpful discussion. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks James Garnett and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, L., Cheng, X., Zhang, F. et al. Ultraweak interactions drive cap-mediated positioning and elongation of the bacterial flagellar filament. Nat Commun 17, 1127 (2026). https://doi.org/10.1038/s41467-025-67887-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67887-y