Abstract
Smc5/6 is an essential genome maintenance complex that interacts with double-stranded (ds) DNA, single-stranded (ss) DNA, and ss-dsDNA junctions. DNA association underlies Smc5/6’s functions in managing intermediates generated during genome replication and repair. However, the mechanisms of this activity are not fully understood. Here, we report a single-molecule study examining Smc5/6 association with a dsDNA substrate containing a ssDNA gap with defined 3’ and 5’ junctions. We found that Smc5/6 associates with both 3’ and 5’ junctions but prefers the 3’ junction in the presence of the ssDNA-binding complex RPA. Further, Smc5/6’s junction association frequency and dwell time are regulated by two non-SMC subcomplexes and DNA binding residues of Smc6. Moreover, Smc5/6 prefers binding to junction sites free of the sliding clamp PCNA over those occupied with it. These results suggest that Smc5/6 utilizes its multiple structural modules to associate with junction sites in coordination with other genome maintenance factors.
Similar content being viewed by others
Introduction
The Structural Maintenance of Chromosomes (SMC) family of complexes is critical for genome management across species1,2. Among the three eukaryotic SMCs, namely cohesin, condensin, and the Smc5/6 complex (Smc5/6), Smc5/6 possesses a unique set of roles in regulating genome replication, DNA repair, and the DNA damage response3,4,5,6. Consistent with its diverse roles, Smc5/6 deficiencies can lead to multi-system genome instability syndromes in humans and chromosomal aberrations in yeast and other model organisms3,4,7,8,9. The cellular roles of Smc5/6 have been linked to its core biochemical activities, including its multimodal interactions with DNA10,11,12,13. Insights into its DNA interaction modes have begun to be revealed by biochemical and single-molecule studies. Recent reports on the yeast Smc5/6 have shown that it resembles other SMCs in interacting with dsDNA and extruding dsDNA loops, but can also bind to sites wherein dsDNA and ssDNA are adjacent to each other, forming ss-dsDNA junctions (junctions for simplicity)14,15,16. DNA junctions are frequently formed on intermediates generated during DNA replication, recombinational repair, and the DNA damage response, which are processes influenced by Smc5/617,18,19,20,21,22. As such, Smc5/6 association with junctions can have important functional implications in genome maintenance.
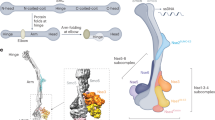
Smc5/6 compositions and functions are conserved among eukaryotes and have been best studied in budding yeast thus far. The budding yeast Smc5/6 holo-complex contains eight subunits, with the Smc5 and Smc6 subunits forming the backbone of the complex and dimerizing at their hinge regions3,4,5,6 (Fig. 1a, left). The SMC arm regions consist of long coiled-coil sequences connecting the hinge regions with the head regions, which contain DNA binding and ATPase sites. Among the six non-SMC elements (NSEs) that attach to the heterodimeric SMC backbone, the Nse1, Nse3, and Nse4 subunits form the Nse1-3-4 subcomplex (Nse1-3-4) that possesses DNA binding activity, while the Nse5 and Nse6 subunits form the Nse5-6 subcomplex (Nse5-6) that lacks direct DNA binding activity10,12,23,24,25,26,27,28,29. The Nse2 SUMO ligase subunit and Nse5-6 interact substantially with the SMC arm regions, whereas Nse1-3-4 primarily associates with the SMC head region10,12,23,24,25,26,27,28,29,30,31. Different subunits and subcomplexes of Smc5/6 can influence its DNA association. For example, ATP binding by Smc5 and Smc6 facilitates the complex to slide along dsDNA, while the inhibition of ATPase activity by Nse5-6 is linked to plasmid entrapment by Smc5/615,16,25,26,32.
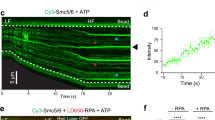
a (Left) A structural model of the budding yeast (S. cerevisiae) Smc5/6 holo-complex, which is composed of a Smc5-Smc6 heterodimer and the Nse1-6 subunits (PDB:7YQH)41. Five of the Nse subunits are organized into two subcomplexes: Nse1-3-4 and Nse5-6. The complex has a tripartite structure consisting of the hinge, arm, and head regions. (Right) A Coomassie blue stained SDS-PAGE gel picture of the Smc5/6 holo-complex after purification, with protein bands corresponding to subunits and molecular weight markers indicated. The band above Nse6 (*) contained chaperone proteins as determined by mass spectrometry. b (Left) A schematic of a ssDNA gap substrate containing a central region of ssDNA (5005 nt) flanked by two dsDNA arms of similar length (6452 bp and 6396 bp). An ATTO 647 N dye is attached to a thymidine located 177 bp from the 3’ junction site to mark the polarity of the tether. Utilization of fluorescently labeled trimeric RPA complex (composed of Rfa1-3 subunits) facilitates the visualization of the ssDNA gap region. (Right) An example kymograph showing the ssDNA gap substrate tethered with beads in optical traps. Fluorescently labeled RPA was withheld to highlight the dye position. White dashed lines indicate the border of the beads. c Representative kymograph of Smc5/6 binding behavior on the gapped substrate in the presence of A555-labeled Smc5/6 (green), LD655-labeled RPA (red), and ATP. The positions of the 3’ and 5’ ss-dsDNA junctions are marked by arrowheads. Another kymograph is included in Fig. S1d. d Dot plot showing junction dwell time distribution for Smc5/6 at 3’ (N = 315) and 5’ (N = 321) junctions. Center line marks median, box floor and height indicate 1st and 3rd quartiles, and minima/maxima of whiskers indicate the 5th and 95th percentiles, respectively. P values were determined from the two-tailed Mann–Whitney U test. Source data are provided in the Source data file. All subsequent plots of junction dwell time and junction encounter frequency, as well as P values, are presented in the same manner as shown here.
The interaction of Smc5/6 with DNA junctions was recently observed using dual-trap optical tweezers combined with multi-color scanning confocal microscopy15,16. These studies examined DNA substrates containing junction sites flanked by long ssDNA flaps, which were generated by stretching spontaneously nicked dsDNA tethers under high force (~60 piconewtons (pN), Supplementary Fig. 1a). In these substrates, free ssDNA flaps flanking the junction sites precluded the assessment of Smc5/6’s properties for binding to the junction alone and of possible preference for junctions with different polarities (i.e. with a 3’ vs. 5’ recessed end). To gain further insights into Smc5/6 association with DNA, we examined a ssDNA gap substrate that can be studied under physiological forces and contains junction sites without ssDNA flaps (Fig. 1b). This substrate resembles cellular ssDNA gaps that can be generated during DNA replication and repair processes. Further, it contains junction sites with different polarities that have distinct functional implications in cells, such as allowing DNA synthesis at 3’ ends versus boosting DNA damage checkpoint activation at 5’ ends33,34. By reporting Smc5/6’s behavior at distinct junction ends, this substrate could inform potential biological roles of Smc5/6 upon junction binding. This setup also allows the examination of how Smc5/6 reaches junction sites and how different subunits of the complex contribute to its junction association. Finally, this gap substrate enables assessment of how Smc5/6 coordinates with other genome maintenance factors for junction binding. Here, we focus on the replication protein A (RPA) complex that coats ssDNA and the proliferating cell nuclear antigen (PCNA) complex that supports processive DNA synthesis, as these are among the most abundant cellular complexes that associate with ssDNA gaps and DNA junctions, respectively, to influence many vital processes35,36.
Here, we examine a ssDNA gap substrate with defined junction sites and report Smc5/6 association with both 3’ and 5’ junctions, with a preference for 3’ junctions when RPA coats the ssDNA region. In addition, Smc5/6 frequently uses the one-dimensional search mode to reach junction sites, though direct binding can also be observed. Moreover, we examine three Smc5/6 variants containing specific mutations or lacking specific subcomplexes. The results suggest that previously mapped dsDNA binding residues on the Smc6 subunit are required for optimal DNA junction encounter frequency, that Nse5-6 promotes junction binding stability, and that Nse1-3-4 boosts both junction encounter frequency and binding stability of Smc5/6. Finally, we show that Smc5/6 and PCNA have a competitive relationship for junction targeting, suggesting that Smc5/6 mainly acts on PCNA-free junctions in genome maintenance processes.
Results
Experimental design to study Smc5/6 junction binding behaviors
We visualized Smc5/6 behavior on a ssDNA gap substrate that contained a 5-kb central ssDNA region flanked by an approximately 6-kb dsDNA arm on each side (Fig. 1b). This ssDNA gapped substrate was generated in two steps (Supplementary Fig. 1b). First, two optically trapped beads were manipulated to tether a 17.8-kb dsDNA molecule that contained two predefined nicks 5 kb apart in the central region. Second, the region between the nicks was melted under high force, and the resultant ssDNA fragment was flowed off from its complementary strand to produce the ssDNA gap substrate. The substrate contained an ATTO 647 N dye near the 3’ junction site, allowing the 3’ and 5’ junctions to be distinguished (Fig. 1b). Specifically, ATTO 647 N was conjugated to a thymidine located 177 nt from the 3’ junction site. Conjugation was mediated via a flexible C6 alkyl spacer to provide flexibility and reduce steric hindrance at the DNA backbone. As such, the dye was unlikely to affect large SMCs (~50 nm in length) sliding on dsDNA, as shown in previous studies14,37. The gap substrate was held at a force of 6 pN at which ssDNA and dsDNA have similar extensions and dsDNA adopts the B-form structure38.
The eight subunits of the budding yeast Smc5/6 complex were co-expressed in yeast, and the holo-complex (or 8-mer complex) was purified using a combination of affinity and gel filtration purification approaches, as done previously15 (Fig. 1a, right). The purified Smc5/6 exhibited ATPase activity as expected (Supplementary Fig. 1c). This activity was abolished upon mutating the ATP hydrolysis residues (Smc5-E1015Q and Smc6-E1048Q) as seen before, ruling out potential contamination from other ATPases15. Consistent with prior studies, the ATPase activity of the Smc5/6 8-mer complex was stimulated by DNA13,25,26,28 (Supplementary Fig. 1c). The Smc5/6 holo-complex was nonspecifically labeled with AlexaFluor555 (A555) through the N-terminal amines of its subunits. Smc5/6 labeled in this manner has been shown to behave similarly to the complex that is site-specifically labeled at the C-terminus of Smc515.
Smc5/6 associates with ss-dsDNA junction sites and has a preference for the 3’ junction
The ssDNA gap substrate described above was incubated with A555-labeled Smc5/6 and five-fold molar excess of LD655-labeled RPA in the imaging channel. The behaviors of Smc5/6 and RPA were monitored by scanning confocal microscopy. Successful generation of the ssDNA gap was evidenced by the LD655-RPA signals coating the expected central region of the DNA tether (Fig. 1c and Supplementary Fig. 1d). We included ATP in the reaction based on the consideration that ATP can influence Smc5/6 binding to dsDNA14,15,16. We followed the binding and movement of Smc5/6 along the entire length of the DNA tether for at least 200 s or until the tether broke for each kymograph.
We found that Smc5/6 did not bind within the ssDNA region when RPA was in excess under the low-force regime (~6 pN, Fig. 1c and Supplementary Fig. 1d). This finding is consistent with previous findings that Smc5/6 was excluded from RPA-coated ssDNA region under a high-force regime15. These data combined indicate that ssDNA fully coated by RPA is a poor substrate for Smc5/6 regardless of force applied to the DNA. In contrast, Smc5/6 was observed to bind the two dsDNA arms flanking the ssDNA gap and diffuse along dsDNA (Fig. 1c and Supplementary Fig. 1d). Its diffusion along dsDNA led to frequent encounters with ss-dsDNA junction sites, defined as sites separating the RPA-coated and RPA-free regions detected in our setup. Measuring the signal lifetime of encounter events at junction sites, a range of Smc5/6 dwell times at the 3’ and 5’ junction sites were observed (Fig. 1d). We described the dwell times by median and quartile range (Q1-Q3) and found that those at 3’ junctions with a median of 0.78 s (0.37–2.18 s) were significantly longer (P < 0.0001) than those at 5’ junctions with a median of 0.46 s (0.26–1.02 s) (Fig. 1d). These results reveal that Smc5/6 interacts with both 3’ and 5’ junction sites in the absence of ssDNA flaps; however, Smc5/6 prefers the 3’ junction in the presence of ATP and excess RPA.
Smc5/6’s preference for the 3’ junction requires RPA and is enhanced by ATP
As RPA is abundant in cells and impacts many DNA-processing reactions36, we asked if and how RPA influences Smc5/6 junction binding. To this end, we first omitted RPA in the experimental setup. Consistent with previous reports, Smc5/6 signal could be seen in RPA-free ssDNA regions (Fig. 2a and Supplementary Fig. 2a, b)15,16. These signals could reflect Smc5/6 binding to ssDNA or to internal junction sites that may form due to secondary structure generation in the absence of RPA. Strikingly, even though Smc5/6 remained present at both 3’ and 5’ junctions, the 3’ junction preference was lost when RPA was omitted, regardless of the ATP status (Fig. 2b). Without RPA, the median dwell times of Smc5/6 at 3’ and 5’ junctions were 0.74 s (0.37–1.92 s) and 0.70 s (0.46–1.47 s) in the presence of ATP, respectively, and were 1.1 s (0.52–2.7 s) and 1.2 s (0.55–2.9 s) in the absence of ATP, respectively (Fig. 2b). The longer junction dwell time in the absence of ATP compared with the addition of ATP suggests that without RPA, ATP may favor the complex to slide back onto dsDNA after a junction encounter. As the 3’ junction preference was lost without RPA despite the presence of ATTO 647 N dye in both RPA and no-RPA conditions, the dye was not the cause for the observed differential behaviors of Smc5/6. In conclusion, these data suggest that RPA is required for Smc5/6 to exhibit a preference for binding to 3’ DNA junctions.
a Representative kymograph of A555-labeled Smc5/6 (green) binding to the gapped DNA in the presence of ATP but no RPA. The 3’ junction is marked by an ATTO 647 N dye (red). The positions of the 3’ and 5’ ss-dsDNA junctions are indicated by arrowheads. Orange brackets indicate events with Smc5/6 dwell time longer than 50 s at 3’ and 5’ junctions. b Dot plot showing the residence time distribution for Smc5/6 at 3’ and 5’ junctions without RPA in the presence or absence of ATP. Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ junction +ATP (N = 470), 5’ junction +ATP (N = 421), 3’ junction -ATP (N = 317), 5’ junction -ATP (N = 443). c Representative kymograph of A555-labeled Smc5/6 (green) binding to the gapped DNA in the presence of ATP and equimolar LD655-labeled RPA (red). Pink arrowhead indicates a position inside the gapped region exhibiting Smc5/6 signals. d Dot plot showing the distributions of junction dwell times for Smc5/6 with equimolar RPA in the presence or absence of ATP. Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ junction +ATP (N = 301), 5’ junction +ATP (N = 513), 3’ junction -ATP (N = 407), 5’ junction -ATP (N = 494).
In alignment with this interpretation, Smc5/6’s 3’ junction binding preference was restored when RPA was added back to the reaction at an equimolar ratio to Smc5/6 (Fig. 2c, d and Supplementary Fig. 2c, d). Junction dwell times of Smc5/6 at 3’ and 5’ junctions differed from each other both in the presence of ATP (P < 0.0001) and in the absence of ATP (P < 0.0001) (Fig. 2d). The equimolar presence of RPA reduced, but did not abolish, Smc5/6 binding within the gapped region (Fig. 2c and Supplementary Fig. 2c, d), in contrast to the excess RPA condition wherein little Smc5/6 signal was observed on ssDNA (Fig. 1c and Supplementary Fig. 1d). In the presence of equimolar RPA, Smc5/6 associated with the ssDNA region at a few sites, rather than broadly distributed when RPA was omitted (Fig. 2c and Supplementary Fig. 2c, d). Taken together, these results show that RPA has a major effect on Smc5/6 association with both ssDNA and junction sites.
Under the condition in which Smc5/6 and RPA were added at an equimolar concentration, we found that ATP omission attenuated the differential binding behavior of Smc5/6 at 3’ versus 5’ junctions. While the median dwell time of Smc5/6 at 3’ and 5’ junction sites in the presence of ATP were 0.77 s (0.36–2.8 s) and 0.44 s (0.28–0.82 s), respectively, these became 0.54 s (0.32–1.1 s) and 0.41 s (0.26–0.64 s) in the absence of ATP, respectively (Fig. 2d). This effect was mainly due to a reduction in Smc5/6 dwell times at 3’ junction sites when ATP was absent compared to when ATP was present (P < 0.0001) (Fig. 2d). With this said, dwell times continued to be longer at the 3’ over 5’ junction without ATP (P < 0.0001). These results suggest that when ATP is omitted, Smc5/6 still shows preferential binding towards the 3’ junction in the presence of RPA, albeit to a lesser degree, which reflects a role for ATP in modulating but not determining Smc5/6 association with 3’ junctions.
Smc5/6 encounters junctions via 1D sliding or direct binding
We next examined the pathway by which Smc5/6 contacts a junction site. We focused on binding events lasting greater than 5 s, a threshold above which we considered as stable binding. Analyzing data collected in conditions with equimolar concentrations of Smc5/6 and RPA (5 nM each), we found that Smc5/6 utilized two modes to target junctions. These included Smc5/6 undergoing one-dimensional (1D) sliding on dsDNA before engaging with a junction, and Smc5/6 binding a junction directly from solution (referred to as direct binding) (Fig. 3a). The relative ratios between the 1D search versus direct binding modes are similar across the ATP status (present or absent) and the junction type (5’ or 3’) (Fig. 3b). In total, 1D search comprised two-thirds of Smc5/6 junction binding events (75/112 events) (Fig. 3b). These data suggest that Smc5/6 can undergo both 1D search or direct binding to engage with a junction site, with a preference for 1D search.
a Representative kymographs of the Smc5/6 complex (green) targeting a ss-dsDNA junction on the gapped substrate via one-dimensional sliding (Top) or direct binding from solution (Bottom) in the presence of LD655-RPA (red). White arrowheads indicate the beginning of Smc5/6 interaction with the junction. Black arrowheads indicate the position of the ss-dsDNA junction. Below each kymograph is a cartoon of the junction encounter modes (1D sliding vs. direct binding) exhibited by Smc5/6. b Pie charts showing the fractions of Smc5/6 junction association events preceded by 1D sliding (dark grey) versus direct binding (light grey) in the presence (Top) or absence (Bottom) of ATP. Binding events that lasted for more than 5 s were counted. Sample sizes are indicated inside the pie charts. Source data are provided in the Source data file.
Oligomerization state of Smc5/6 at junctions examined using a dye-free DNA substrate
Since A555 could be conjugated to the N-terminus of any subunit of Smc5/6, varied numbers of fluorophores attached to different complexes could cause heterogeneous signal intensities. Alternatively, varied levels of signal intensity could reflect different oligomerization states of Smc5/6. To distinguish between these two scenarios, we performed site-specific labeling of Smc5/6 by conjugating the Cy3 fluorophore to an S6-tag fused to Smc5 via the SFP enzyme (see Method). Our previous study has shown that Smc5/6 subjected to this labeling method exhibited similar dsDNA sliding behaviors as the N-terminal labeled Smc5/6, and both bind to ss-dsDNA junctions with ssDNA flaps15. We verified that the S6-tagged Smc5/6 8-mer complex showed comparable ATPase activity as the one without this tag, suggesting that both are enzymatically active (Supplementary Fig. 3a, b).
We went on to examine Cy3-labeled Smc5/6 on DNA in the presence of RPA and ATP. To rule out any potential effect of DNA dye on Smc5/6 binding behavior, we used a gapped DNA substrate containing different lengths for the dsDNA arms flanking the ssDNA region. In this case, 3’ versus 5’ junctions can be determined based on the asymmetry of the dsDNA arms (Supplementary Fig. 3c)39. Overall, a more uniform brightness of Smc5/6 signals on the DNA substrate was seen compared to the experiments using non-specifically labeled Smc5/6. Importantly, Smc5/6 dwell times at 3’ junctions were significantly longer than those at 5’ junctions (P < 0.0001, Supplementary Fig. 3d). This finding verifies our results using the non-specifically labeled Smc5/6 and DNA substrate with ATTO 647 N, with both experiments suggesting that Smc5/6 can bind to both junctions but prefers the 3’ junction. Quantification of the fluorescence intensities showed that about three quarters of Smc5/6 traces contained one fluorophore, while the remaining quarter contained two fluorophores (Supplementary Fig. 3e). This result implies that only one or two copies of the Smc5/6 complex bind the junction sites, and a higher oligomeric state is uncommon. Furthermore, measurements of Smc5/6 dwell time at either the 3’ or 5’ junctions showed no statistical difference between Smc5/6 traces containing one or two fluorophores (Supplementary Fig. 3f), suggesting that junction engagement is not impacted by potential Smc5/6 oligomerization.
Nse1-3-4 is required for optimal Smc5/6 binding to both dsDNA and junction sites
Next, we sought to test how non-SMC subcomplexes contribute to Smc5/6 junction binding. We first examined the role of the Nse1-3-4 subcomplex by studying a Smc5/6 complex lacking these three subunits, which is referred to here as the Smc5/6 5-mer (Fig. 4a). A cryo-EM structure of Smc5/6 encircling dsDNA has shown that the Nse1-3-4 subcomplex contains DNA binding sites on Nse3 and Nse47. Previous studies have also shown that the Smc5/6 5-mer is defective in binding dsDNA and that Nse1-3-4 alone is capable of dsDNA binding10,12,15,26,40. However, it was observed that the Smc5/6 5-mer complex can still associate with junction DNA flanked by long ssDNA flaps15.
a SDS-PAGE gel picture of purified Smc5/6 5-mer (lacking Nse1-3-4) and 6-mer (lacking Nse5-6) complexes. Coomassie blue stained bands correspond to the subunits as indicated. b Representative kymograph of A555-labeled Smc5/6 5-mer (green) binding behavior on the gapped substrate in the presence of LD655-labeled RPA (red) and ATP. Black arrowheads indicate the positions of the 3’ and 5’ ss-dsDNA junctions. c Dot plot showing the distributions of junction dwell times for Smc5/6 8-mer (black) or 5-mer (gold) at 3’ and 5’ junctions in the presence of RPA and ATP. Plots and statistical analyses are presented as in Fig. 1d. Sample sizes: 3’ 8-mer (N = 301), 3’ 5-mer (N = 156), 5’ 8-mer (N = 513), 5’ 5-mer (N = 141). d Representative kymograph of A555-labeled Smc5/6 6-mer (green) binding behavior on the gapped substrate in the presence of LD655-labeled RPA (red) and ATP. Black arrowheads indicate the positions of the 3’ and 5’ ss-dsDNA junctions. e Dot plot showing the numbers of Smc5/6 binding events at the 3’ or 5’ junctions per 100 s in the presence of ATP for the 8-mer Smc5/6 (black) versus the 6-mer Smc5/6 (magenta). Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ 8-mer (N = 37), 3’ 6-mer (N = 44), 5’ 8-mer (N = 37), 5’ 6-mer (N = 46). f Dot plot showing the distributions of junction dwell times for Smc5/6 8-mer (black) and 6-mer (magenta) in the presence of RPA and ATP. Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ 8-mer (N = 301), 3’ 6-mer (N = 1,443), 5’ 8-mer (N = 513), 5’ 6-mer (N = 1,468).
We purified the Smc5/6 5-mer and showed it has drastically reduced ATPase activity as reported previously15,41 (Supplementary Fig. 4a). We used the reaction condition containing ATP and equimolar Smc5/6 and RPA (5 nM each), at which the 8-mer Smc5/6 displayed a 3’ junction binding preference (Fig. 2c, d). Under the same reaction condition, Smc5/6 5-mer showed sparse signals on dsDNA (Fig. 4b and Supplementary Fig. 4b). This result is consistent with previous single-molecule data using DNA substrates tethered under high force and with biochemical data that Nse1-3-4 is important for dsDNA binding10,12,15,26,40. Further, the junction dwell times of the 5-mer complex were significantly reduced compared to those of the 8-mer for both 5’ and 3’ junctions, and the 3’ junction preference was also lost (Fig. 4c). At the 3’ junction, the 5-mer complex had a median dwell time of 0.30 s (0.18–0.63 s) compared to the 8-mer dwell time with a median of 0.77 s (0.36–2.8 s), and the difference was statistically significant (P < 0.0001). At the 5’ junction, the 8-mer median dwell time of 0.44 s (0.28–0.82 s) compared to the 5-mer median dwell time of 0.28 s (0.18–0.51 s) was also significantly different (P < 0.0001). However, no statistical difference was found between the dwell time distribution of the 5-mer at 3’ versus 5’ junctions (P = 0.37). These data suggest that Nse1-3-4 is required for optimal Smc5/6 association with junction sites in addition to its role in dsDNA binding. The retained, albeit diminished, ability of the Smc5/6 5-mer to bind junction sites suggests that additional DNA contact sites also contribute to junction association.
Nse5-6 enhances Smc5/6 stability at junctions but inhibits its dsDNA binding
The Nse5-6 subcomplex lacks DNA binding activity but can regulate DNA entrapment and looping as well as ATPase activities of Smc5/614,24,25,26,32,41. To interrogate whether and how this regulatory subcomplex affects Smc5/6 junction binding, we purified the Smc5/6 complex lacking Nse5-6 (termed here as the Smc5/6 6-mer) (Fig. 4a). As seen previously, the Smc5/6 6-mer exhibited an increased ATPase activity compared to the Smc5/6 8-mer (Supplementary Figs. 1c and 4a)25,26,41. Using the same single-molecule setup as described above, we observed that the 6-mer complex showed more robust dsDNA association compared with the 8-mer complex. This was evident even when the 6-mer was used at a reduced concentration (2 nM), a condition necessary to resolve individual 6-mer complexes in our setup (Fig. 4d and Supplementary Fig. 4c). This behavior suggests that Nse5-6 is a negative regulator of dsDNA binding by Smc5/6, which is consistent with previous reports14.
The enhanced dsDNA binding for the Smc5/6 6-mer compared with the 8-mer complex raised the possibility that it may target the junction sites more efficiently via 1D diffusion. We found that the Smc5/6 6-mer exhibited a three-to-five-fold increase in encounter frequency with 3’ and 5’ junctions compared to the 8-mer complex (Fig. 4e). Note that the encounter frequency is not only influenced by the diffusion coefficient of Smc5/6 on dsDNA, but also by its binding lifetime on DNA and its propensity to form long-lived interactions with the junction. Indeed, the 6-mer’s dwell times at both 5’ and 3’ junctions were reduced compared with the 8-mer, while the 3’ junction preference was maintained (Fig. 4f). At the 3’ junction, the 6-mer complex exhibited a median dwell time of 0.48 s (0.23–1.2 s) compared with the 8-mer dwell time of 0.77 s (0.36–2.8 s) and the difference was statistically significant (P < 0.0001). At the 5’ junction, the median dwell times for 8-mer and 6-mer were 0.44 s (0.28–0.82 s) and 0.37 s (0.19–0.77 s), respectively, and the difference was also statistically significant (P < 0.0001). Further, the dwell time distributions of the 6-mer complex at the 3’ versus 5’ junction were found to be significantly different (P < 0.0001).
These results suggest that while the Smc5/6 6-mer makes more frequent encounters with junctions than the 8-mer, these encounters are less conducive to stable associations. This behavior may be due to the 6-mer’s greater tendency to transition back to the dsDNA region. As such, the Nse5-6 subcomplex can regulate Smc5/6’s distribution between dsDNA and junctions, serving as a molecular toggle for different Smc5/6 functionalities. This effect differs from that of Nse1-3-4, which constitutively enhances Smc5/6 association with both dsDNA and junctions.
Smc6’s DNA binding sites regulate Smc5/6 junction binding
Our finding that Smc5/6 still interacts with junctions in the absence of Nse1-3-4, albeit at a reduced capacity (Fig. 4b, c), suggests that other DNA binding modules also contribute to Smc5/6 binding at DNA junctions. A recent cryo-EM and mutagenesis study mapped DNA binding sites on the head regions of Smc5 and Smc6, in addition to those on the Nse1-3-4 subcomplex10. DNA binding sites on the Smc6 subunit were shown to aid in Smc5/6 association with chromatin and support cell viability10. Thus, we examined a Smc5/6 6-mer variant that contained mutations of the Smc6 DNA binding sites (K129A, K140A, K177A, K200A, K201A, K202A) (Fig. 5a)10. The purified mutant 6-mer complex (Smc6DNAm) exhibited an ATPase activity comparable to the wild-type 6-mer complex (Fig. 5b and Supplementary Fig. 4a). However, compared to the wild-type complex, the Smc6DNAm complex exhibited fewer signals on the gapped DNA substrate and reduced encounter frequencies at the 3’ and 5’ junctions (Fig. 5c, d and Supplementary Fig. 4d). Junction dwell time analyses based on the Mann-Whitney U test indicated that the Smc6DNAm complex generally showed longer dwell time than the wild-type complex, but had much fewer events exceeding 10 seconds (Fig. 5e). At the 3’ junction, Smc6DNAm exhibited a median dwell time of 0.80 s (0.40–1.8 s) compared with the wild-type 6-mer’s median dwell time of 0.48 s (0.23–1.2 s), and the difference was statistically significant (P < 0.0001). At the 5’ junction, the median dwell times for Smc6DNAm and wild-type 6-mer complex were 0.50 s (0.30–1.4 s) and 0.37 s (0.19–0.77 s), respectively, and the difference was also statistically significant (P < 0.0001). Collectively, these data suggest that the examined DNA binding sites on the Smc6 head region contribute to DNA association, but other DNA binding sites, such as those located on Nse3, Nse4, and Smc5 subunits10,12,26,40, can also be important for junction association.
a Structural model depicting the DNA binding sites on Smc6. A cryo-EM structure of a portion of the Smc5/6 6-mer encircling a single dsDNA and six Smc6 DNA binding residues are shown (PDB: 7TVE)10. b SDS-PAGE gel picture of purified Smc5/6 6-mer complex containing mutated DNA binding residues within the Smc6 subunit (Smc6DNAm). Coomassie blue stained protein bands correspond to the subunits as indicated. c Representative kymograph of A555-labeled Smc6DNAm 6-mer (green) binding behavior on the gapped substrate in the presence of LD655-labeled RPA (red) and ATP. Black arrowheads indicate the positions of the 3’ and 5’ ss-dsDNA junctions. d Dot plot showing the numbers of Smc5/6 binding events at the 3’ or 5’ junctions per 100 s in the presence of ATP for the 6-mer wildtype (magenta) versus Smc6DNAm (purple). Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ WT (N = 44), 3’ Smc6DNAm (N = 14), 5’ WT (N = 46), 5’ Smc6DNAm (N = 14). e Dot plot showing the distributions of junction dwell times for Smc5/6 6-mer WT (magenta) or Smc6DNAm (purple) in the presence of 5 nM LD655-RPA and 2 mM ATP. Plots and statistical analyses are presented as in Fig. 1d. Sample sizes are: 3’ WT (N = 1,443), 3’ Smc6DNAm (N = 275), 5’ WT (N = 1,468), 5’ Smc6DNAm (N = 202).
PCNA limits Smc5/6’s junction association and dsDNA sliding
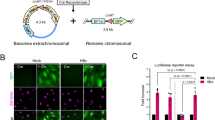
Thus far, our data have provided insights into how Smc5/6 achieves junction binding, its preference for junction polarity, the roles of its subcomplexes and DNA binding sites, and the influence of RPA. Next, we sought to explore how Smc5/6’s engagement with junction sites is influenced by another junction binding complex—proliferating cell nuclear antigen (PCNA). PCNA is a ring-shaped complex that binds DNA polymerases to enhance their processivity and is loaded onto DNA by the replication factor C (RFC) in an ATP-dependent manner35,42,43. Using dual-color imaging, we tracked the behaviors of A555-labeled Smc5/6 8-mer and LD655-labeled PCNA on the gapped DNA substrate in the presence of RFC, RPA, and ATP (Fig. 6a). As reported before, PCNA exhibited 1D diffusive behavior on dsDNA44. Interestingly, when Smc5/6 and PCNA were both observed to diffuse on dsDNA, they recoiled from each other upon encounter in most cases (“Short colocalization” in Fig. 6b, c). Only rarely did they stably associate and diffuse together (“Long colocalization” in Fig. 6b, c) or bypass each other (“Bypass” in Fig. 6b, c). Given that PCNA and Smc5/6 can both be topologically linked to dsDNA10,14,15,16,44, these observations suggest that the two do not form a stable super-complex on dsDNA or create topological openings wide enough for the other to pass through.
a Representative kymograph of A555-Smc5/6 (green) and LD655-PCNA (red) on a gapped substrate in the presence of RFC, RPA, and ATP. b Examples of categories of Smc5/6 and PCNA behaviors upon their encounter on dsDNA. (Top) The two complexes exhibited transient colocalization (arrowheads) followed by a recoil upon encounter. Arrow indicates the position of DNA junction. (Middle) The two stably colocalized and co-diffused on dsDNA. (Bottom) They bypassed each other. Arrowhead indicates a bypass event. c Plot showing the fraction of events (N = 1640) for each of outcomes of Smc5/6 and PCNA encounter on dsDNA as described in panel b. Bar heights indicate means; error bars represent SEM (N = 12 kymographs). Statistical analyses are presented as in Fig. 1d. d Plot showing the numbers of Smc5/6 binding events at the 3’ junctions per 100 s. The gapped substrate was incubated with LD655-PCNA, RFC, RPA, and ATP before transitioning to channel containing A555-Smc5/6 8-mer, RPA, and ATP. For “PCNA-” condition, the first incubation was omitted. Sample sizes are: PCNA- (N = 37), PCNA+ (N = 9). e Plot showing dwell time distributions for Smc5/6 at 3’ junctions under the condition where PCNA was omitted or included. Sample sizes are: PCNA- (N = 301), PCNA+ (N = 173). d, e Plots and statistical analyses are presented as in Fig. 1d. f A working model for Smc5/6 binding to junction DNA and its regulation by RPA and PCNA. Our findings suggest that Smc5/6 can arrive at junction sites via 1D diffusion or direct binding. Smc5/6 can then engage with 3’ or 5’ junction sites, with a preference for 3’ junctions when RPA coats ssDNA. RPA’s DNA binding domain (DBD)-A/B and DBD-C/D/E are shown as small and large ovals and bind to 3’ and 5’ of ssDNA, respectively. PCNA (ring) reduces Smc5/6 junction encounter frequency and dwell times. The Nse5-6 subcomplex reduces Smc5/6 junction encounter frequency but increases junction dwell times. Frequent junction encounter requires Smc6 DNA binding residues (star) and Nse1-3-4. Additionally, Nse1-3-4 increases junction dwell times.
The finding that PCNA and Smc5/6 remain as separate entities and are unable to bypass each other raised the possibility that PCNA could constrain Smc5/6’s movement on dsDNA and thus affect its access to junction sites. Indeed, we found that pre-loading PCNA onto the gapped substrate reduced the frequency of 3’ junction targeting by Smc5/6, suggesting that PCNA acts as an obstacle to Smc5/6’s junction access (Fig. 6d). Furthermore, the presence of PCNA reduced Smc5/6 dwell times at 3’ junctions (Fig. 6e), and Smc5/6 and PCNA rarely colocalized at junctions (Supplementary Fig. 5). These results indicate that PCNA and Smc5/6 antagonize each other’s junction binding, thus likely occupying different junction sites in the cell.
Discussion
Smc5/6 has important functions during several genome maintenance processes. During DNA replication, Smc5/6 stabilizes stalled replication forks and promotes replication termination13,17,18,45,46. In recombinational repair, Smc5/6 enables the processing of branched DNA repair intermediates19,20,21. A commonality of DNA substrates subjected to regulation by Smc5/6 is the presence of ssDNA gaps and ss-dsDNA junctions. Smc5/6 deficiencies in yeast consistently increase the level of DNA structures harboring ssDNA gaps47. Recent studies of budding yeast Smc5/6 revealed its preferential binding to junction sites flanking ssDNA gaps15,16, providing a plausible means to support its multiple roles in genome maintenance. Here, we dissected the mechanisms underlying Smc5/6 binding to junction sites using ssDNA gap substrates with defined 3’ and 5’ junction positions. We found that yeast Smc5/6 is capable of binding to both 3’ and 5’ junctions flanking ssDNA in the absence of flap DNA or the ssDNA-binding complex RPA. Moreover, RPA-coated ssDNA renders a 3’ junction preference for Smc5/6 in the presence of ATP. To reach junction sites, Smc5/6 frequently uses 1D diffusion, but it can also directly bind these sites from solution. Our examinations of Smc5/6 variants suggest that Smc5/6 uses multiple subunits to achieve optimal junction binding. While Nse1-3-4 generally promotes DNA binding, Nse5-6 regulates the balance of dsDNA association versus junction association by Smc5/6. Further, DNA binding sites in Smc6 are required for frequent encounter with junction sites. Finally, we showed that Smc5/6 prefers PCNA-free junctions and independently slides on dsDNA along with PCNA without passing each other. These findings, summarized in a working model (Fig. 6f), provide insights into how Smc5/6 associates with DNA junction sites through subunit collaboration and how this activity is influenced by two key genome maintenance complexes—RPA and PCNA.
We and others have previously reported that Smc5/6 binds to DNA junction sites next to extended ssDNA flaps under high force15,16. Here, we show that Smc5/6 has the intrinsic ability to bind “clean” ss-dsDNA junctions without ssDNA flaps under low force. In cells, clean junctions without flaps can flank ssDNA gaps produced due to failed replication, among other situations. Smc5/6 association with clean junctions observed in this work, or with a previously used replication fork containing a short ssDNA gap15, represents binding events that could facilitate Smc5/6-mediated repair of ssDNA gaps or stalled forks. Cells also contain ss-dsDNA junctions with extended ssDNA flaps, such as those generated by end resection or at degraded replication forks. Longer-lived Smc5/6 at these sites may underlie its function to protect and target DNA to specialized locations, such as nuclear envelope or the peripheries of heterochromatin48,49,50,51. Interestingly, Smc5/6 dwell times at flap-containing junctions are about an order of magnitude longer than those at clean junctions as measured in our experimental setup15. Thus, Smc5/6 binding to “clean” versus “flap” junctions could mediate differential cellular functions.
Our work showed that Smc5/6 frequently uses 1D diffusion along DNA to reach junction sites. In cells, this mode of junction binding may operate within a short distance given the presence of nucleosomes on chromatin. However, 1D diffusion may prove to be effective in situations where nucleosome occupancy is highly dynamic, such as during replication and repair processes where nucleosomes are removed before reassembly. We also observed that, for about one third of the time, Smc5/6 directly binds to junction sites, providing another means for Smc5/6 to target junctions in the nucleus that circumvents the need to remove or navigate through protein barriers on DNA. The 1D search mode of Smc5/6 relies on its interaction with dsDNA, supported by our finding that Nse1-3-4, important for the complex’s dsDNA binding10,12,15,26,40, is required for optimal junction targeting. Further, our data suggest that the DNA binding residues of Smc6 are important for frequent junction targeting likely because they are involved in dsDNA encirclement as seen in a prior cryo-EM structure10. However, mutation of these residues did not reduce the overall dwell times of Smc5/6 at junction sites. This suggests that other DNA binding sites within Smc5/6 subunits are used to engage junctions. These could include dsDNA-binding residues mapped to the Smc5 head region, Nse3, Nse4, and possibly ssDNA-binding residues mapped on the hinge regions of Smc5 and Smc610,11,52. The ssDNA and dsDNA binding sites within Smc5/6 may allow its simultaneous engagement with both the ssDNA and dsDNA portions of the junction sites, a notion worth exploring in the future.
In contrast to the DNA binding sites of Smc6 and Nse1-3-4, Nse5-6 lacks DNA binding ability, but inhibits the complex’s ATPase activity14,24,25,26,32,41. Consistent with a regulatory role, we found that the Smc5/6 6-mer lacking Nse5-6 showed elevated dsDNA binding and increased junction encounter frequency, but reduced dwell times at junction sites. These results suggest that Nse5-6 can operate as a molecular toggle for different Smc5/6 functions. It adds to previous findings that Nse5-6 favors trapping of plasmid DNA but disfavors DNA loop extrusion14,32. We speculate that the inclusion of Nse5-6 in the Smc5/6 complex provides more time for downstream processing activities at DNA junctions, while limiting Smc5/6’s sliding and loop-extruding activities on dsDNA. Given that the Nse5-6 counterparts in fission yeast and human cells have been suggested to be more dynamically associated with the other subunits of the complex compared with the budding yeast Smc5/653,54,55, it is plausible that different organisms may have differential needs for Nse5-6-mediated regulation.
Our work agrees with previous reports that Smc5/6 can interact with ssDNA regions on the gap DNA tether15,16. However, we found that increasing RPA concentrations gradually diminish this binding without disrupting Smc5/6-junction association. Therefore, RPA availability in the cell can tune the distribution of Smc5/6 on DNA. Smc5/6 association with RPA-free ssDNA regions may be mediated by its hinge region, which has been found to bind better to ssDNA than dsDNA11. Alternatively, as RPA can dismantle hairpin DNA structures56,57, the observed Smc5/6 signals within the gap region in the absence of RPA may correspond to binding events at ss-dsDNA junctions around these hairpins. Future studies will be required to gain further insights into RPA-regulated Smc5/6 binding to ssDNA.
Besides reducing Smc5/6’s presence within the ssDNA region, RPA also renders a 3’ preference for Smc5/6 junction binding. This suggests that RPA can make 3’ junctions more permissible for Smc5/6 to engage with compared to 5’ junctions. We speculate that this effect may be linked to RPA’s own binding polarity to ssDNA. While the three core DNA-binding domains of RPA engage with the 5’ ssDNA, its two more dynamic DNA-binding domains engage with the 3’ portion of ssDNA57,58,59,60. Thus, it is possible that the flexible binding of RPA at the 3’ portion of ssDNA allows DNA to be more accessible to Smc5/6. Our data using site-specifically labeled Smc5/6 suggest that a single complex can associate with junction sites with a 3’ junction preference, and that multiple complexes in proximity may also engage with junctions. More studies will be needed to better understand the mechanism and functional implications for the junction polarity preference of Smc5/6.
Finally, we observed that Smc5/6 prefers PCNA-free sites and the two often slide separately on dsDNA without forming a composite sliding unit or passing each other. This suggests that these junction-binding factors serve distinct roles in the cell. Recent studies reported that PCNA and its loader RFC can become limiting in replication stress, resulting in the formation of many ssDNA gaps61,62. When gap junctions are not occupied by PCNA, we envision that Smc5/6 can fulfill critical roles at PCNA-free junctions to protect them from nuclease degradation and to recruit downstream proteins for gap filling and repair. Future studies on the complementary functions of these junction-binding factors under different stress conditions can further our understanding of ssDNA gap management in cells.
Methods
Purification of Smc5/6 and its variants
Smc5/6 8-mer and 5-mer complexes
Subunits of the S. cerevisiae Smc5/6 8-mer or 5-mer complex with Smc5 tagged at its C-terminus with the CBP tag, or the tandem CBP and S6 tag, were overexpressed in yeast through the galactose-inducible promoters as outlined in previously reported protocols15,63. The Smc5/6 8-mer variant Smc5/6-EQ contains the ATP hydrolysis site mutations (Smc5-E1015Q and Smc6-E1048Q)10,15.
The complexes were purified according to previously reported protocols15,63. Briefly, cells were first grown at 30 °C in YP media supplemented with 2% glycerol and 2% lactic acid till reaching late log phase growth. Then galactose was added to reach 2% final concentration, and cells were grown for another 4 h to induce protein expression. Cell pellets were frozen dropwise in liquid nitrogen in buffer E (45 mM HEPES-KOH, pH 7.6, 10% glycerol, 0.02% NP40) supplemented with 100 mM NaCl, 1 mM DTT, 1x protease inhibitor cocktail (Sigma), and 1x cOmplete Ultra EDTA-free protease inhibitor (Roche). Cell breakage was done using a freezer mill (SPEX CertiPrep 6850 Freezer/Mill) for 7 cycles of 2 min at a rate of 15 impacts per second. Cell powder was then resuspended with 1 volume of buffer E supplemented with 300 mM NaCl and 1 mM DTT, and extracts were cleared by centrifugation for 30 min at 177,469 × g at 4 °C. Protein extract was supplemented with 2 mM CaCl2 and incubated with calmodulin affinity resin for 2 h at 4 °C to pull down CBP tagged Smc5 along with the other subunits of Smc5/6. The resin was washed by 10 CVs of buffer E supplemented with 300 mM NaCl, 2 mM CaCl2, 1 mM DTT, and bound proteins were eluted with the same buffer supplemented with 1 mM EDTA, 2 mM EGTA, and without CaCl2. Peak fractions were then applied to a Superose 6 Increase column in buffer containing 45 mM HEPES-KOH pH 7.6, 10% glycerol, 200 mM NaCl, 0.02% NP40, 1 mM DTT. Proteins in the peak fractions were concentrated and frozen for storage.
Smc5/6 6-mer complex
The pET expression plasmid contained the S. cerevisiae genes encoding Smc5, Smc6, and the Nse1-4 subunits, with Smc6 fused with the 3C-Twin-Strep tag10,25. The plasmid was used to generate the expression construct of the Smc6DNAm 6-mer complex wherein a synthesized DNA fragment containing the indicated mutations (see text) was exchanged with the wild-type Smc6 gene using the Gibson Assembly Cloning Kit (NEB E5510S). All plasmids were sequenced to confirm correct construction.
Expression and purification of both types of Smc5/6 6-mer complex were done according to a previously reported protocol10. Briefly, E. coli BL21 (DE3) cells containing the expression plasmids were grown in TB-medium at 37 °C before the addition of 0.4 mM IPTG to induce protein expression at 22 °C for 16 h. All following steps were carried out at 4 °C. Cells were harvested and resuspended in lysis buffer (50 mM Tris-HCl, pH 7.5, 300 mM NaCl, 5% glycerol, 25 mM imidazole) containing also 5 mM DTT, 1 mM PMSF, and 150 U Benzonase (EMD Millipore). Cell breakage was achieved by sonication at 40% amplitude for 15 min with pulsing of 1 s on and 2 s off cycle. Cell lysate was centrifuged at 40,000 × g for 40 min, and the protein extracts were filtered through a 0.45-µm filter. Cleared lysate was loaded to a 5-mL Strep-tactin HP (Cytiva) column, pre-equilibrated with 25 mL of lysis buffer supplemented with 2 mM DTT. After washing the column with 50 mL of lysis buffer, proteins were eluted with 20 mL of the same buffer supplemented with 2 mM DTT and 5 mM desthiobiotin. Eluates were added to a 5-mL HiTrap Heparin column (Cytiva) that was pre-equilibrated with 10 mL lysis buffer supplemented with 2 mM DTT. After washing the column with 25 mL of the same buffer, bound proteins were eluted with 20 mL of elution buffer (20 mM Tris, pH 7.5, 1 M NaCl, 2 mM DTT). Peak fractions were concentrated using Amicon Ultra centrifugal filter units (30 kDa cutoff) before loading to a 24-mL Superose6 10/300 GL size-exclusion chromatography column (Cytiva). Gel filtration and storage buffer contains 20 mM HEPES, pH 7.5, 250 mM NaCl, 1 mM TCEP. Peak fractions containing Smc5/6 were concentrated and snap-frozen with 15% glycerol in aliquots.
Purification of PCNA, RFC, and RPA complexes
PCNA
S. cerevisiae S6-tagged PCNA (Pol30) was expressed and purified following a previously described protocol42. Briefly, Pol30 containing a C-terminal S6-tag was expressed from a modified pET16 plasmid to allow the addition of an N-terminal cAMP-dependent protein kinase A recognition sequence (LRRASV) followed by a His-tag64. 48 L of E. coli BLR(DE3) cells (Novagen) were grown in LB media containing ampicillin at 30 °C to OD600 = 0.6. Protein expression was induced upon the addition of 1 mM IPTG for ~18 h at 15 °C. Cell pellets resuspended with an equal weight of Buffer 1 (20 mM Tris-HCl, pH 7.9, and 500 mM NaCl) containing 5 mM imidazole were disrupted by French press and centrifugation at 23,700 × g for 1 h at 4 °C. All following steps were carried out at 4 °C. Clarified supernatant was applied to a 150-mL HiTrap Nickel-chelating resin (Cytiva), pre-equilibrated with Buffer 1 containing 60 mM imidazole. The resin was washed with 300 mL Buffer 1 containing 60 mM imidazole before bound protein was eluted with a 1.6-L gradient of Buffer 1 with 60 mM imidazole to Buffer 1 with 500 mM imidazole. 25 mL fractions were collected, and the ones containing PCNA were pooled and dialyzed overnight against Buffer A (20 mM Tris HCL, pH 7.5, 5 mM DTT, 0.1 mM EDTA, 10% glycerol). Fractions were applied to a 200-mL DEAE Sepharose column, pre-equilibrated with Buffer A. The column was washed with 500 mL Buffer A, and bound proteins were eluted with a gradient of Buffer A without salt to Buffer A with 500 mM NaCl. Peak fractions containing PCNA were pooled and dialyzed overnight against Buffer A, and the protein was then aliquoted and stored in −80 °C.
RFC
S. cerevisiae RFC complex composed of full-length Rfc1-5 subunits was expressed using a dual vector strategy65. The genes of Rfc1 and Rfc5 subunits were cloned into a pLANT-2/RIL plasmid, whereas the genes of Rfc2-4 were cloned into a pET(11a) plasmid. Briefly, 48 L of E. coli BLR(DE3) cells (Novagen) grown the presence of ampicillin and kanamycin were allowed to each OD600 ~ 0.6 at 30 °C. Protein expression was induced by the addition of 1 mM IPTG for ~18 h at 15 °C. Cell pellets were resuspended in an equal weight of Tris-Sucrose (50 mM Tris-HCl, pH 7.5, 10% sucrose), followed by disruption by cold French press and cleared by centrifugation at 23,700 × g for 1 h at 4 °C. The supernatant was supplemented with Buffer A and 150 mM NaCl before being loaded onto a 30-mL SP-Sepharose Fast Flow column (Cytiva), pre-equilibrated with Buffer A and 150 mM NaCl. Bound protein was eluted with a 300-mL gradient of Buffer A with 150 mM NaCl to Buffer A with 600 mM NaCl. Peak fractions (eluted at ~365 mM NaCl) were pooled and diluted with Buffer A to achieve ~150 mM NaCl. The pooled fractions were applied to a 40-mL Q-Sepharose Fast Flow column (Cytiva), pre-equilibrated with Buffer A containing 150 mM NaCl. Bound proteins were eluted with a 400-mL gradient of Buffer A with 150 mM NaCl to Buffer A with a 600 mM NaCl. Peak fractions (eluted at ~300 mM NaCl) were pooled, aliquoted, and stored in –80 °C.
RPA
S. cerevisiae RPA was cloned, grown, and purified according to a previous protocol66. Briefly, plasmids encoding Rfa1, Rfa2, and Rfa3 under control of the IPTG inducible promotor were transformed into E. coli BL21-DE3 cells and induced with IPTG for 16 h at 15 °C. Protein was released from cells by French Press, then clarified by centrifugation and loaded onto a column, washed with 0.8 M KCl and 1.5 M Sodium thiocyanate (NaSCN), and then desalted using a hydroxyapatite column in 80 mM potassium phosphate. Protein complex was further purified on a MonoQ column in 200 mM KCl and eluted with a gradient to 1 M KCl. Peak fractions containing the RPA were pooled, aliquoted, and stored in −80 °C.
Protein labeling
Labeling Smc5/6 with AlexaFluor555 and RPA with LD655
We used AlexaFluor555 (A555) NHS ester (Thermo Fisher; Catalog # A20009) to non-specifically label the primary amines of Smc5/6 subunits15. Preferential N-terminal labeling was achieved by labeling at low pH (7.0) for an NHS ester reaction. Labeling buffer contained 45 mM HEPES-KOH pH 7.0, 200 mM NaCl, 1 mM DTT, and 0.25 mM EDTA pH 8.0. The Smc5/6 sample was incubated with A555 dye at a 1:5 molar ratio for 1 h at room temperature in the dark. The reaction was quenched with 30 mM Tris-HCl, pH 7.0, for 5 min. Excess dye was removed by buffer exchange with a Micro Bio-Spin P-30 Gel Column (Bio-Rad) with a storage buffer containing 45 mM HEPES-KOH, pH 7.6, 200 mM NaCl, 10% glycerol, and 1 mM DTT. The final labeling ratio was estimated to be >80% for all Smc5/6 samples. The labeled Smc5/6 complex was aliquoted, flash frozen, and stored at −80 °C. LD655-RPA was labeled using a similar protocol, but with a labeling buffer containing 50 mM HEPES-KOH pH 7.0, 150 mM NaCl, 1 mM DTT, and 0.25 mM EDTA, and a storage buffer containing 20% glycerol, 30 mM HEPES pH 7.9, 150 mM NaCl, 1 mM DTT, and 0.25 mM EDTA as previously described15.
Site-specific labeling of PCNA and Smc5/6
PCNA underwent labeling with SFP synthase (4’-phosphopantetheinyl transferase). The SFP synthase recognizes the short peptide S6-tag and catalyzes the covalent transfer of CoA-functionalized moieties to a serine residue within the tag via a phosphopantetheinyl linker. PCNA-S6, the SFP enzyme, and LD655-CoA were incubated at a 1:2:5 molar ratio in buffer containing 10 mM MgCl2 for 1 h at room temperature. The reaction was washed by 7 spins through Amicon Ultra 50 K filters in final storage buffer containing 20 mM Tris-HCl, pH 7.5, 150 mM KCl, 2 mM MgCl2, 1 mM DTT, 10% glycerol. The final labeling ratio was estimated to be 70% for each monomer.
A similar method was used to label S6-tagged Smc5/6 8-mer complex with the Cy3 dye and has been utilized previously15. Briefly, Smc5/6-S6, the SFP enzyme, and Cy3-CoA were incubated at a 1:3:5 molar ratio in buffer containing 10 mM MgCl2 at 4 °C overnight. Excess dye and the SFP enzyme were removed through buffer exchange using a Micro Bio-Spin™ P-30 Gel Column (BioRad) into a storage buffer containing 45 mM HEPES KOH, pH 7.6, 200 mM NaCl, 10% glycerol, 1 mM DTT, 0.02% IGEPAL, 0.01% Tween 20. Labeled Smc5/6-S6 was flash frozen and stored at −80 °C. The labeling ratio was estimated to be 30%, which is within the normal range of SFP-based labeling yield for Smc5/615. Protein concentration was determined through Bradford assay, and dye concentration was determined through Beer’s Law (A = εlc) using a Cy3 ε value of 150,000 M−1 cm−1 at 554 nm.
Single-molecule data acquisition
Single-molecule experiments were performed at room temperature on a LUMICKS C-Trap instrument combining three-color confocal fluorescence microscopy with dual-trap optical tweezers. A computer-controlled stage enabled movement of the optical traps within a microfluidic flow cell containing five channels. Laminar flow separated channels 1–3, which were used to form DNA tethers between 4.35-μm streptavidin-coated polystyrene beads (Spherotech). Under constant flow in channel 1, a single bead was caught in each optical trap. The traps were then moved to channel 2, containing the biotinylated DNA substrate of choice. By moving one trap back and forth in an axis parallel to the flow direction, a DNA substrate could be tethered between the two beads and detected via a change in the force-extension curve. The traps were then moved to channel 3, and the presence of a single DNA tether was verified by the force-extension curve. Orthogonal channels 4 and 5 served as the experimental and imaging channels that contained proteins of interest. The standard buffer used in these protein and imaging channels contained 50 mM potassium glutamate, 50 mM Tris-HCl pH 7.5, 2.5 mM MgCl2, and 1 mM DTT unless otherwise noted. Flow was turned off during data acquisition. The A555 and Cy3 fluorophores were excited by a 532-nm laser line, and the LD655 fluorophore was excited by a 638-nm laser line. The 532-nm laser operated at ~2.5 μW (10%) and the 638-nm laser at ~3.9 μW (10%) or ~2.0 μW (5%) measured at the objective. Kymographs were generated via a confocal line scan through the center of the two beads, with a line time of 45–75 ms and a pixel time of 0.1 ms.
The ss-dsDNA gapped substrate (17,853 nt) was commercially purchased from LUMICKS (SKU: 00027). The substrate was stored as nicked dsDNA. To form the gap region, a tethered nicked dsDNA substrate was moved to channel 3, that only contained Tris-HCl pH 8.0. The absence of additional salt in the buffer aids in the denaturing of duplex DNA. The inter-bead distance was then gradually moved to 7.8 μm to melt the DNA duplex, at which a flow was introduced (~0.6 bar) for 10 s to facilitate the removal of the central DNA strand. The DNA tether was then relaxed and tested to determine if the force-extension curve had transitioned to an intermediate between a full dsDNA tether and a full ssDNA tether. If the tether remained a full dsDNA tether, the cycle of pulling and flowing was repeated until the gap substrate was formed or the tether snapped. When the gap substrate was successfully formed, the tether was held at ~6 pN in channel 3 and moved to channel 4 and/or channel 5 for experiments. For experiments involving a red dye (e.g., LD655-RPA), a kymograph was collected prior to the experiment in order to note the location/intensity of the ATTO 647 N dye and/or to photobleach the ATTO 647 N dye.
To visualize DNA binding by Smc5/6, the gapped tether was moved to channel 4, which contained A555-Smc5/6 (2–10 nM) ± ATP (2 mM) ± labeled or unlabeled RPA (5–25 nM) in standard buffer of 50 mM potassium glutamate, 50 mM Tris-HCl pH 7.5, 2.5 mM MgCl2, and 1 mM DTT. Kymographs lasted 200–250 s or until the tether broke. These protocols were also used for the experiments involving the wild-type Smc5/6 8-mer, Smc5/6 5-mer, 6-mer, and the Smc6DNAm variants. For the experiments where Smc5/6 and PCNA were allowed to bind DNA together, the gapped substrate was moved into channel 4 with 5 nM A555-Smc5/6, 2 nM LD655-PCNA, 2 nM RFC, 5 nM RPA, and 2 mM ATP in standard buffer. For the experiments where PCNA was allowed to bind DNA prior to Smc5/6, the gapped tether was incubated with 5 nM LD655-PCNA, 5 nM RFC, 5 nM RPA, and 2 mM ATP for 30 s and then moved to the experimental channel with 5 nM A555-Smc5/6 8-mer, 5 nM RPA, and 2 mM ATP. The 638-nm laser power was reduced to ~2.0 μW (5%) to lengthen the fluorescence lifetime of LD655.
To visualize Cy3-labeled Smc5/6 (5 nM) in the presence unlabeled RPA (5 nM) and ATP (2 mM), the gapped DNA substrate was constructed with an existing protocol39. The 19-kb substrate contained two pre-defined nicks to form a gap region flanked by 10-kb and 6-kb dsDNA arms. The asymmetry of the dsDNA arms was used to differentiate the 3’ and 5’ junctions.
Single-molecule data analysis
Kymographs were processed and visualized using custom software written by John Watters (“CTrapVis”, https://github.com/lumicks/harbor/tree/main/Visualization/C-Trap%20.h5%20File%20Visualization%20GUI), which incorporates tools from the lumicks.pylake Python library and other Python modules (Numpy, Matplotlib, Pandas). The start and end times of junction binding events for Smc5/6 were recorded via the photon profile for each kymograph. Frequency of junction binding events was calculated by dividing the number of recorded interactions for a junction on a kymograph by the total time of the kymograph and presented as number of events per 100 s. For analysis of junction targeting modes for Smc5/6, binding events that lasted more than 5 s were analyzed and categorized into whether Smc5/6 had been diffusing on DNA immediately prior to junction binding (1D search) or if it bound the junction with no prior 1D diffusion (direct binding). Events were excluded from analysis if Smc5/6 was already bound at the beginning of the kymograph. Graphs and plots were created using Prism. The number of Cy3-labeled Smc5/6 complexes per trajectory was estimated by dividing the averaged photon count for each trajectory by the photon count for a single Smc5/6 under the same imaging condition, following a similar procedure in a previous study67.
ATPase activity assay
Each examined form of the Smc5/6 complex, including 8-mer (wild-type or EQ), 6-mer, 5-mer, and Smc6DNAm, was incubated prior to labeling at 250 nM with 1 mM ATP and 80 nCi [α-32P]-ATP in 20 μL buffer containing 25 mM Tris-HCl pH 7.5, 2 mM MgCl2, 50 mM NaCl, 2% glycerol, and 1 mM DTT. For reactions including DNA, 40-bp dsDNA (2 µM) was added. Reactions were conducted at 30 °C for 2.5, 5, 7.5, and 10 min for 8-mer (wild-type), 6-mer, and Smc6DNAm or for 20, 40, and 60 min for 8-mer (EQ) and 5-mer. Reactions were terminated by the addition of 5 M formic acid to reach a final concentration of 0.83 M. Samples were then analyzed by thin layer chromatography, followed by phosphorimaging and quantification of ADP versus ATP levels46.
Statistics and reproducibility
Statistical tests and numbers of molecules or events analyzed are reported in the figure legends and in the text. P values were determined by the non-parametric, two-tailed Mann–Whitney U test, except noted otherwise, and were denoted in figure panels when possible. All protein gels were repeated and provided similar results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Kymographs used for analysis are available via Zenodo at https://zenodo.org/records/1779265868. Source data are provided with this paper.
Code availability
All specified scripts used to process and analyze C-trap data can be accessed on the LUMICKS Harbor site (“CTrapVis”, https://github.com/lumicks/harbor/tree/main/Visualization/C-Trap%20.h5%20File%20Visualization%20GUI)69.
References
Uhlmann, F. SMC complexes: from DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 17, 399–412 (2016).
Burmann, F. & Lowe, J. Structural biology of SMC complexes across the tree of life. Curr. Opin. Struct. Biol. 80, 102598 (2023).
Roy, S., Adhikary, H. & D’Amours, D. The SMC5/6 complex: folding chromosomes back into shape when genomes take a break. Nucleic Acids Res. 52, 2112–2129 (2024).
Peng, X. P. & Zhao, X. The multi-functional Smc5/6 complex in genome protection and disease. Nat. Struct. Mol. Biol. 30, 724–734 (2023).
Palecek, J. J. SMC5/6: multifunctional player in replication. Genes 10, 7 (2019).
Aragón, L. The Smc5/6 complex: new and old functions of the enigmatic long-distance relative. Annu. Rev. Genet. 52, 89–107 (2018).
Grange, L. J. et al. Pathogenic variants in SLF2 and SMC5 cause segmented chromosomes and mosaic variegated hyperploidy. Nat. Commun. 13, 6664 (2022).
van der Crabben, S. N. et al. Destabilized SMC5/6 complex leads to chromosome breakage syndrome with severe lung disease. J. Clin. Investig. 126, 2881–92 (2016).
Payne, F. et al. Hypomorphism in human NSMCE2 linked to primordial dwarfism and insulin resistance. J. Clin. Investig. 124, 4028–38 (2014).
Yu, Y. et al. Cryo-EM structure of DNA-bound Smc5/6 reveals DNA clamping enabled by multi-subunit conformational changes. Proc. Natl. Acad. Sci. USA 119, e2202799119 (2022).
Alt, A. et al. Specialized interfaces of Smc5/6 control hinge stability and DNA association. Nat. Commun. 8, 14011 (2017).
Zabrady, K. et al. Chromatin association of the SMC5/6 complex is dependent on binding of its NSE3 subunit to DNA. Nucleic Acids Res. 44, 1064–1079 (2016).
Kanno, T., Berta, D. G. & Sjogren, C. The Smc5/6 complex Is an ATP-dependent intermolecular DNA linker. Cell Rep. 12, 1471–82 (2015).
Pradhan, B. et al. The Smc5/6 complex is a DNA loop-extruding motor. Nature 616, 843–848 (2023).
Chang, J. T. H. et al. Smc5/6’s multifaceted DNA binding capacities stabilize branched DNA structures. Nat. Commun. 13, 7179 (2022).
Tanasie, N. L., Gutiérrez-Escribano, P., Jaklin, S., Aragon, L. & Stigler, J. Stabilization of DNA fork junctions by Smc5/6 complexes revealed by single-molecule imaging. Cell Rep. 41, 111778 (2022).
Ampatzidou, E., Irmisch, A., O’Connell, M. J. & Murray, J. M. Smc5/6 is required for repair at collapsed replication forks. Mol. Cell Biol. 26, 9387–9401 (2006).
Menolfi, D., Delamarre, A., Lengronne, A., Pasero, P. & Branzei, D. Essential roles of the Smc5/6 complex in replication through natural pausing sites and endogenous DNA damage tolerance. Mol. Cell 60, 835–846 (2015).
Branzei, D. et al. Ubc9- and Mms21-mediated sumoylation counteracts recombinogenic events at damaged replication forks. Cell 127, 509–522 (2006).
Bermúdez-López, M. et al. Sgs1’s roles in DNA end resection, HJ dissolution, and crossover suppression require a two-step SUMO regulation dependent on Smc5/6. Genes Dev. 30, 1339–56 (2016).
Bonner, J. N. et al. Smc5/6 mediated sumoylation of the Sgs1-Top3-Rmi1 complex promotes removal of recombination intermediates. Cell Rep. 16, 368–378 (2016).
Pond, K. W., de Renty, C., Yagle, M. K. & Ellis, N. A. Rescue of collapsed replication forks is dependent on NSMCE2 to prevent mitotic DNA damage. PLoS Genet. 15, e1007942 (2019).
Duan, X. et al. Architecture of the Smc5/6 Complex of Saccharomyces cerevisiae Reveals a Unique Interaction between the Nse5-6 Subcomplex and the Hinge Regions of Smc5 and Smc6. J. Biol. Chem. 284, 8507–15 (2009).
Yu, Y. et al. Integrative analysis reveals unique structural and functional features of the Smc5/6 complex. Proc. Natl. Acad. Sci. USA 118, e2026844118 (2021).
Taschner, M. et al. Nse5/6 inhibits the Smc5/6 ATPase and modulates DNA substrate binding. EMBO J. 40, e107807 (2021).
Hallett, S. T. et al. Nse5/6 is a negative regulator of the ATPase activity of the Smc5/6 complex. Nucleic Acids Res. 49, 4534–4549 (2021).
Hallett, S. T. et al. Cryo-EM structure of the Smc5/6 holo-complex. Nucleic Acids Res. 50, 9505–9520 (2022).
Gutierrez-Escribano, P. et al. Purified Smc5/6 complex exhibits DNA substrate recognition and compaction. Mol. Cell 80, 1039–1054 e6 (2020).
Serrano, D. et al. The Smc5/6 core complex is a structure-specific DNA binding and compacting machine. Mol. Cell 80, 1025–1038 e5 (2020).
Zhao, X. & Blobel, G. A SUMO ligase is part of a nuclear multiprotein complex that affects DNA repair and chromosomal organization. Proc. Natl. Acad. Sci. USA 102, 4777–82 (2005).
Duan, X. et al. Structural and functional insights into the roles of the Mms21 subunit of the Smc5/6 complex. Mol. Cell 35, 657–68 (2009).
Taschner, M. & Gruber, S. DNA segment capture by Smc5/6 holocomplexes. Nat. Struct. Mol. Biol. 30, 619–628 (2023).
Johansson, E. & Dixon, N. Replicative DNA polymerases. Cold Spring Harb. Perspect. Biol. 5, a012799 (2013).
Waterman, D. P., Haber, J. E. & Smolka, M. B. Checkpoint responses to DNA double-strand breaks. Annu. Rev. Biochem. 89, 103–133 (2020).
Choe, K. N. & Moldovan, G. L. Forging Ahead through darkness: PCNA, still the principal conductor at the replication fork. Mol. Cell 65, 380–392 (2017).
Caldwell, C. C. & Spies, M. Dynamic elements of replication protein A at the crossroads of DNA replication, recombination, and repair. Crit. Rev. Biochem. Mol. Biol. 55, 482–507 (2020).
Barth, R. et al. SMC motor proteins extrude DNA asymmetrically and can switch directions. Cell 188, 749–763.e21 (2025).
Wasserman, M. R. & Liu, S. A tour de force on the double helix: exploiting DNA mechanics to study DNA-based molecular machines. Biochemistry 58, 4667–4676 (2019).
Chua, G. N. L. et al. A non-catalytic role for RFC in PCNA-mediated processive DNA synthesis. Cell (in press) https://doi.org/10.1016/S0092-8674(25)01478-3 (2025).
Jo, A., Li, S., Shin, J. W., Zhao, X. & Cho, Y. Structure basis for shaping the Nse4 protein by the Nse1 and Nse3 dimer within the Smc5/6 complex. J. Mol. Biol. 433, 166910 (2021).
Li, Q. et al. Cryo-EM structures of Smc5/6 in multiple states reveal its assembly and functional mechanisms. Nat. Struct. Mol. Biol. 31, 1532–1542 (2024).
Zheng, F., Georgescu, R., Yao, N. Y., Li, H. & O’Donnell, M. E. Cryo-EM structures reveal that RFC recognizes both the 3′- and 5′-DNA ends to load PCNA onto gaps for DNA repair. eLife 11, e77469 (2022).
Yao, N. et al. Replication factor C clamp loader subunit arrangement within the circular pentamer and its attachment points to proliferating cell nuclear antigen. J. Biol. Chem. 278, 50744–53 (2003).
Kochaniak, A. B. et al. Proliferating cell nuclear antigen uses two distinct modes to move along DNA. J. Biol. Chem. 284, 17700–10 (2009).
Peng, X. P. et al. Acute Smc5/6 depletion reveals its primary role in rDNA replication by restraining recombination at fork pausing sites. PLoS Genet. 14, e1007129 (2018).
Xue, X. et al. Restriction of replication fork regression activities by a conserved SMC complex. Mol. Cell 56, 436–45 (2014).
Agashe, S. et al. Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites. Nat. Commun. 12, 2111 (2021).
Mangione, R. M. et al. DNA lesions can frequently precede DNA:RNA hybrid accumulation. Nat. Commun. 16, 2401 (2025).
Whalen, J. M., Dhingra, N., Wei, L., Zhao, X. & Freudenreich, C. H. Relocation of collapsed forks to the nuclear pore complex depends on sumoylation of DNA repair proteins and permits Rad51 association. Cell Rep. 31, 107635 (2020).
Horigome, C. et al. PolySUMOylation by Siz2 and Mms21 triggers relocation of DNA breaks to nuclear pores through the Slx5/Slx8 STUbL. Genes Dev. 30, 931–45 (2016).
Ryu, T. et al. Heterochromatic breaks move to the nuclear periphery to continue recombinational repair. Nat. Cell Biol. 17, 1401–11 (2015).
Liu, H. W. et al. The SMC hinge is a selective gate for obstacle bypass. Nat. Commun. 16, 10457 (2025).
Oravcova, M. et al. The Nse5/6-like SIMC1-SLF2 complex localizes SMC5/6 to viral replication centers. Elife 11, e79676 (2022).
Pebernard, S., Wohlschlegel, J., McDonald, W. H., Yates, J. R. 3rd & Boddy, M. N. The Nse5-Nse6 dimer mediates DNA repair roles of the Smc5-Smc6 complex. Mol. Cell Biol. 26, 1617–30 (2006).
Raschle, M. et al. DNA repair. Proteomics reveals dynamic assembly of repair complexes during bypass of DNA cross-links. Science 348, 1253671 (2015).
Chen, H., Lisby, M. & Symington, L. S. RPA coordinates DNA end resection and prevents formation of DNA hairpins. Mol. Cell 50, 589–600 (2013).
Nguyen, B. et al. Diffusion of human replication protein A along single-stranded DNA. J. Mol. Biol. 426, 3246–3261 (2014).
de Laat, W. L. et al. DNA-binding polarity of human replication protein A positions nucleases in nucleotide excision repair. Genes Dev. 12, 2598–609 (1998).
Kolpashchikov, D. M. et al. Polarity of human replication protein A binding to DNA. Nucleic Acids Res. 29, 373–9 (2001).
Iftode, C. & Borowiec, J. A. 5’ −> 3’ molecular polarity of human replication protein A (hRPA) binding to pseudo-origin DNA substrates. Biochemistry 39, 11970–81 (2000).
Canal, B. et al. The DNA replication checkpoint limits Okazaki fragment accumulation to protect and restart stalled forks. Mol. Cell 85, 2462–2473.e6 (2025).
Chhetri, G. et al. PAF15-PCNA assembly exhaustion governs lagging strand replication and replisome integrity. bioRxiv https://doi.org/10.1101/2025.03.15.641049 (2025).
Li, S. et al. Molecular basis for Nse5-6 mediated regulation of Smc5/6 functions. Proc. Natl. Acad. Sci. USA 120, e2310924120 (2023).
Kelman, Z. & O’Donnell, M. Structural and functional similarities of prokaryotic and eukaryotic DNA polymerase sliding clamps. Nucleic Acids Res. 23, 3613–20 (1995).
Finkelstein, J., Antony, E., Hingorani, M. M. & O’Donnell, M. Overproduction and analysis of eukaryotic multiprotein complexes in Escherichia coli using a dual-vector strategy. Anal. Biochem. 319, 78–87 (2003).
Henricksen, L. A., Umbricht, C. B. & Wold, M. S. Recombinant replication protein A: expression, complex formation, and functional characterization. J. Biol. Chem. 269, 11121–32 (1994).
Chua, G. N. L. et al. Differential dynamics specify MeCP2 function at nucleosomes and methylated DNA. Nat. Struct. Mol. Biol. 31, 1789–1797 (2024).
Chang, J. T. H. Kymographs of Smc5/6 binding. Zenodo https://zenodo.org/records/17792658 (2025).
Watters, J. Scripts for single-molecule C-Trap data analysis. Zenodo https://zenodo.org/records/7618698 (2023).
Acknowledgements
We thank John Watters in the Shixin Liu Laboratory for technical support, Sophia Park in the Xiaolan Zhao Laboratory for helpful comments. J.T.C. was supported by an NCI F30 fellowship (F30CA275379). J.T.C. and B.J.K. were supported by a Medical Scientist Training Program grant to the Weill Cornell/Rockefeller/Sloan Kettering Tri-Institutional MD-PhD Program (T32GM152349) from the National Institutes of Health. V.M.B. and J.Z. acknowledge support from a training grant from NIGMS awarded to the Molecular Biophysics Program at Weill Cornell Graduate School (T32GM132081). G.N.L.C. was supported by an NIH F31 fellowship (F31MH132306). M.E.O. was supported by NIH (R01GM115809) and Howard Hughes Medical Institute. X.Z. was supported by NIH (R35GM145260) and Memorial Sloan-Kettering Cancer Center Core Grant P30 CA 008748. S. Liu was supported by the Alfred P. Sloan Foundation, the Marlene Hess Center for Research in Women’s Health and Biomedicine at Rockefeller University, and NIH (R01GM149862).
Author information
Authors and Affiliations
Contributions
X.Z. and S.Liu oversaw the project. J.T.C., V.M.B., and G.N.C. performed single-molecule experiments and analyzed the data. S.Li, V.M.B., and J.Z. prepared protein constructs and performed biochemical experiments. B.J.K. assisted with single-molecule experiments. M.E.O. and E.C.B. provided key reagents. J.T.C., V.M.B., X.Z., and S.Liu wrote the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maxim Molodtsov, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chang, J.TH., Miller-Browne, V., Chua, G.N.L. et al. Molecular determinants of Smc5/6 association with DNA junctions. Nat Commun 17, 1239 (2026). https://doi.org/10.1038/s41467-025-67999-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67999-5