Abstract
Understanding the impact that subtle variations (missense mutation, environmental change, ion chelation, ligand binding, etc.) have on protein structure helps to reveal their biological effects, but remain extremely challenging due to the difficulty in measuring and locating the changes in protein structure. Herein, a method entitled MELO is therefore constructed, which enable a systematic measurement based on residues’ geometric characteristics & relative distance and a high-throughput location of structural change based on secondary structure variation & protein segment shift. Our method performs best in capturing the structure changes of various degrees of magnitude (some increases were >30%) and is capable of precisely locating the regions of alterations for critical case studies. Moreover, it identifies over 10,000 structural changes induced by subtle variation that existing methods fail to detect. An online server allows users to upload their structures for comparison, and all those structural changes identified in this study have also been made available for download.
Similar content being viewed by others
Introduction
The function of a protein is closely related to its three-dimensional structure1,2. Any changes in structure (caused by mutation3, environmental change4, metal ion chelation5 or ligand binding6, as shown in Supplementary Fig. S1) can significantly affect the protein functions7,8,9, which subsequently results in physiological variation10 or pathological response11. In other words, it is demanded to measure the structural variation12 and locate the altered region13, which are crucial for accelerating the cutting-edge research of protein function prediction, etiology analysis, drug discovery14, etc. Traditionally, researchers rely on structural alignment and visual inspection to discover structural variations15. However, such strategy is not only inefficient and incapable of quantifying results but also carries high degree of subjectivity, making it challenging to capture critical structural variations, especially when analyzing complex proteins16,17. Thus, developing a high-throughput scoring method, which realizes objective, accurate and reliable measurement of structural variation and location of altered region, is of significant importance18,19.
Till now, two types of scoring method have been available, which include superposition-based and superposition-free ones20. For the superposition-based methods (such as RMSD21 and TM-score22), they are valued for their ability to offer quantitative assessment of structural variation, allowing effective comparison across diverse protein structures23. Particularly, TM-score offers a standardized approach by providing a fixed scoring range, which is not dependent on the size of the protein, and introduces clear thresholds for assessing structural similarity24. This feature makes it useful for comparing proteins of different sizes25. For the superposition-free methods, they are designed to evaluate structural changes without needing to align the protein structures beforehand26. Taking LDDT27 as an example, it is known to be insensitive to structure rotation and translation, allowing it to disregard the errors caused by the absolute coordinates of amino acids, thereby reducing the bias introduced by inaccuracies in the alignment process28.
However, subtle variations (such as the mutation in sequence) are reported to be able to induce obvious structure changes29,30 (even disruptions3) that the existing methods are not always able to capture, nor can the methods locate the regions of alteration31. As shown in Supplementary Fig. S1, it is clear that existing methods are insufficient to identify structural changes arising from the subtle perturbations. Critical issues include: a) different methods result in inconsistent conclusions; b) use of different alignment methods lead to significant scoring variations; and c) different method versions result in divergent outcomes (as discussed in Supplementary Fig. S1). Moreover, existing methods typically focus on assessing difference between two structures using a simple score, and their validity in locating the structural change was yet to be assessed. These analyses above highlight the critical need in developing effective method to measure the structural variation between proteins and locate their corresponding region of alteration.
In this work, we develop a scoring method entitled MELO capable of measuring and locating the structural variation of proteins. This method: a) enables comprehensive measurement by collectively considering the changes in Geometric characteristics and Relative distances among amino acids, and b) realizes high-throughput location of structure alterations by simultaneously assessing the variations of secondary structures and the shifts among protein segments. Compared with available methods, MELO performs best in identifying the structure change of various degrees of magnitude (some increases were > 30%) and is capable of precisely measuring such structure change and locating altered region in critical case studies. Additional work identified >10,000 structural changes induced by subtle variation that existing methods fail to detect. Finally, we construct an online server to allow users to upload their own structures for enabling customized comparison, and all those structure changes ( > 10,000) we identify in this study had also been made available for download (which was reported to be urgently demanded by research community3). A standalone program and the source code of MELO is made available.
Results and discussion
Constructing the MELO for Measuring and Locating the Structure Variations
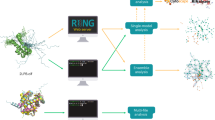
The MELO was developed in this study to accurately measure the structure changes induced by subtle variation (SCinSV) and precisely locate the regions of the changes, which functioned by assessing the degree of structure changes from two perspectives: a) it calculated the changes of residue’s geometric characteristics to measure the variations in secondary structure (VSSmeasur), and provided the location of those variations (PLOTVSS). Specific details were demonstrated in Fig. 1b) it normalized and weighted the relative distances among residues to assess the shift among protein segments (SPSmeasur), and offered the location of the shift (PLOTSPS). Specific details were described in Fig. 2. Moreover, this study applied a thresholding approach proposed by TM-score24 to analyze tens of millions of protein pairs collected from SCOP232 and CATH33, which helped to identify the thresholds customized for MELO to indicate structure change. Particularly, the protein pairs having variations in secondary structures or shifts among structure segments would be assessed by MELO as VSSmeasur ≥ 0.5 or SPSmeasur ≥ 0.5, respectively. The detailed process to identify the thresholds was shown in Supplementary Fig. S2. In the following sections, we compared the performance of MELO with that of the existing methods in assessing SCinSV, and several case-analyses were further provided to enable the in-depth comparison.
a preprocessing the structure preparation and representation. S-W- & US-align-based algorithms were used to align two protein sequences (SA and SB), establishing the residue-level correspondences and ensuring a precise mapping promoting downstream analyses. b computing geometric characteristics of studied amino acids. Key geometric characteristics of amino acids were calculated for both SA and SB, including: bend angles (KAPPA), dihedral angles (ALPHA), peptide backbone torsion angles (PHI and PSI), water exposed surface (ACC), etc. All characteristics were encoded into vectors (such as V1, V1’ for SA and V2, V2’ for SB). c depicting PLOTVSS based on geometric characteristics among residues. Variations in secondary structure were located using PLOTVSS. The darker the residue’s color was, the higher its ci value was, which reflected greater variation in its corresponding secondary structure. Moreover, the PLOTVSS showed the type of secondary structure in SA and SB, with the classification information described in Supplementary Fig. S25. d calculating VSSmeasur based on the geometric characteristics among residues. VSSmeasur scores were calculated to quantify overall secondary structure variation. Particularly, the closer the VSSmeasur is to 1, the more different the secondary structures of two studied proteins are.
a preprocessing structure preparation and representation. The S-W- & US-align-based algorithms were adopted to align sequences of two protein (SA and SB), establishing the residue-level correspondence and facilitating the precise pairwise distance calculation for detecting segmental shifts. b computing and normalizing the relative distances among amino acids. Pairwise distances between residues were computed for protein pairs after sequence alignment (such as dA1, dA2, dA3 for protein A and dB1, dB2, dB3 for protein B). Changes between these distances from two proteins were calculated and normalized based on weighting function using the maximum pairwise distances, resulting in SPSmeasur. c calculating SPSmeasur and generating PLOTSPS based on relative distance. The PLOTSPS depicted the protein segment shifts between two proteins. The vertical axis demonstrated the sequence of protein A (SA), and the horizontal axis provided the sequence of protein B (SB). The darker the color of residue pair was, the greater shift in its corresponding protein segment was. As shown, the area demarcated by the black dash line highlighted significant segment shift between domain 1 (colored in pink) and domain 2 (colored in blue), contrasting with minimal internal change within each domain.
Examination by the gradually deepening classification levels of SCOP2
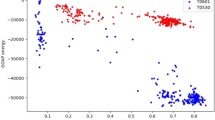
Four benchmarks were collected to represent varying degrees of structure changes based on the four classification levels of SCOP232 (class, fold, superfamily, and family), which contained 0.308, 0.409, 0.411, and 0.413 billion structure pairs, respectively. Since TM-score is the only existing method with a clear threshold for distinguishing protein structure changes (score < 0.5 was adopted to indicating structure change24 or unsuccessful prediction of structures34), the performance of MELO was first compared with this established method using four benchmarks. As shown in Fig. 3a together with a corresponding Supplementary Table S1 providing the comprehensive raw data, as the SCOP2 level deepen (from class, to fold, then to superfamily, and finally to family), the accuracy of TM-score in differentiating the structural change declined from 99.8% to 93.8%, while our MELO consistently maintained the accuracies of > 99.7% at all four levels. In other words, MELO and TM-score worked well for discovering the changes in “major secondary structure” and “arrangement of secondary structure elements” (changes at class and fold levels of SCOP2), while MELO provided greater precision than TM-score in capturing the change at superfamily and family levels (increase by about 5%).
a comparing the accuracies of two methods across four SCOP2 classification levels (Class, Fold/IUPR, Superfamily, and Family). MELO maintained a consistent accuracy >99.9% at all levels, providing greater precision than TM-score in identifying the change at the level of superfamily and family (increased by about 5%). b accuracy assessments between MELO and TM-score in identifying structural change induced by relatively small sequence alterations (the protein pairs with sequence identity >70%). MELO kept consistent accuracy above 99.9% at all levels, substantially outperforming TM-score. c the assignments of two million structure pairs (sequence identity >70%) into four Quadrants (Q1 to Q4) based on the results of VSSmeasur and SPSmeasur. The pie charts provided the level of agreement between MELO and TM-score in each Quadrant. In Q1, the MELO and TM-score mostly achieved the same conclusions with 0.01% disagreement. In Q2, Q3 and Q4, where MELO found structure changes, the findings of two methods were largely inconsistent, with 86.45%, 97.23% and 91.95% pairs having distinct conclusion, respectively. d a representative pair of proteins in Q2 (Borrelia burgdorferi OspA before and after the replacement of two β-hairpin sequences with Gly-Gly and shortening of one β-hairpin). Shifts among protein segments were captured by MELO (VSSmeasur = 0.13, SPSmeasur = 0.63), while both TM-score and LDDT failed to identify these structural changes. e a representative protein pair in Q3 (intestinal fatty acid-binding protein and its helix-less variant). Both the variations in secondary structures and shifts among protein segments were successfully identified by MELO (VSSmeasur = 0.66 & SPSmeasur = 0.65), but not found by TM-score and RMSD. f a representative protein pair in Q4 (change of some ordered secondary structures to disordered ones in SARS-CoV-2 spike protein induced by multi-point mutations). Such changes were successfully captured by MELO (VSSmeasur = 0.56 & SPSmeasur = 0.35), but did not discovered by other existing methods.
To further assess the accuracy of methods in identifying structural change induced by relatively small sequence alteration, the protein pairs with sequence identity > 70% were chosen from the above benchmarks. As provided in Fig. 3b, as the SCOP2 level deepen, the accuracy of TM-score in identifying structural changes declined from 96.5% to 62.4%, while that of our MELO remained the accuracies of > 99.9% at all four levels. Moreover, there is a dramatical decline in the accuracies ( ~ 30%) of TM-score between class/fold and superfamily/family, which indicated our MELO as well-performing in revealing structure changes of different degrees of magnitude (compared with the accuracies of TM-score in the SCOP2 level of superfamily and family, that of the MELO enhanced significantly by > 30%). All in all, MELO performed well in identifying the structural changes, particularly those induced by relatively small sequence alteration.
Assessment based on the Structural Changes Induced by Subtle Variations
To measure the ability of methods to discover the structure changes induced by subtle variation (SCinSV), all single proteins in the PDB35 were paired and filtered as described in ‘Collecting Benchmark Datasets for Assessing Methods’ Performances’, resulting in over two million structure pairs with sequence identity greater than 70%. Such kind of variation included the small sequence alteration (such as missense mutation3), environmental change4, ion chelation5, ligand binding6, and so on. Using MELO, those two million pairs could be first divided into four Quadrants (from Q1 to Q4, shown in Fig. 3c) by those two metrics (VSSmeasur and SPSmeasur) and thresholds. Particularly, Q1 indicated the pairs of similar structures (VSSmeasur <0.5, SPSmeasur < 0.5), Q2 presented the pairs having shifts among protein segments (VSSmeasur <0.5, SPSmeasur ≥ 0.5), Q4 denoted the pairs with variations in secondary structures (VSSmeasur ≥ 0.5, SPSmeasur < 0.5), and Q3 gave the pairs with both shifts among segments and variations in secondary structure (VSSmeasur ≥ 0.5, SPSmeasur ≥ 0.5).
Three protein pairs being representative of three Quadrants Q2, Q3, and Q4 were then selected and shown in Fig. 3. As illustrated in Fig. 3d, a dramatic shift of a domain (from BLUE to PINK) in Borrelia burgdorferi OspA protein was reported to be induced by replacing two β-hairpin sequences with the Gly-Gly and shortening of a β-hairpin36. Such extensive shifts were successfully captured by MELO (VSSmeasur = 0.13 & SPSmeasur = 0.63; being in Q2 having shifts among protein segments), but not detected using TM-score. Moreover, as offered in Fig. 3e, a change of two α-helices & one β-sheet to a disordered region and a spatial displacement were shown between intestinal fatty acid-binding protein and its helix-less variant37. Both variations in secondary structures and shifts among protein segments were identified by MELO (VSSmeasur = 0.66 & SPSmeasur = 0.65; being in Q3), but not discovered by TM-score. In addition, as shown in Fig. 3f, a clear change of some ordered secondary structures to disordered ones in SARS-CoV-2 spike protein was reported to be induced by multi-point mutation38. Such changes in the secondary structure were also successfully identified by MELO (VSSmeasur = 0.56 & SPSmeasur = 0.35; being in the Q3 of Fig. 3c), but remained undiscovered using the TM-score.
Both RMSD21 and LDDT27 were widely used in protein structure comparisons. However, these two methods were absent of established threshold, which made their determination of structural changes relatively subjective20. In other words, a variety of ways of determination were used in existing studies39,40,41,42, and empirical rules could be generalized for RMSD (RMSD < 2 Å denoted highly similar structure, while RMSD > 5 Å indicated significant structural change) and LDDT (LDDT > 0.9 represented structure consistency, 0.7 < LDDT < 0.9 implied high local similarity with minor deviation, while LDDT < 0.5 suggested extensive structural discrepancy). Based on these empirical rules, an extra evaluation on those protein pairs being representative of Q2, Q3, and Q4 was conducted using RMSD and LDDT. As shown in Fig. 3d, e, f RMSD failed to identify two out of the three structural changes (as shown in Fig. 3e, f) and LDDT was not able to detect the structure changes in Fig. 3d, f. As a result, MELO was found effective in not only discovering SCinSV but also differentiating the types of structural change (protein segment shifts or secondary structure variations).
Moreover, based on the above analyses, it was reasonable to infer that many structural changes may not be detected if existing methods were used to conduct large-scale screening on millions of structure pairs. Therefore, MELO and TM-score, having established thresholds, were studied to compare their evaluating results in four Quadrants (from Q1 to Q4, illustrated in Fig. 3c). In Q1, the MELO and TM-score mostly reached the same conclusion with 0.01% disagreement, which led to a total of 205 disagreed pairs. Our in-depth analysis further identified that all these disagreed pairs had their aligned sequences less than 40 amino acids, and this is consistent with AlphaFold’s observation that TM-score was less effective for short sequence34. Specifically, as shown in Supplementary Fig. S3, all 205 disagreed protein pairs (discovered by TM-score, but missed by MELO; highlighted using black dots) were illustrated, and four typical exemplar pairs (selected from four corners of Q1) were offered in Supplementary Fig. S3. As shown, those four exemplar pairs were only identified by TM-score as the ones of significant structural change, but “missed” by all other methods (such as MELO, LDDT, and RMSD), and our visual inspections further confirmed that there was non-significant conformational change in all those four exemplar pairs. In Q2, Q3 and Q4, where MELO discovered structure change, the findings of two methods were significantly inconsistent, with 86.45%, 97.23% and 91.95% pairs having distinct conclusions, respectively. In other words, our MELO identified 12,562 SCinSV protein pairs from PDB, while TM-score identified 1,411. Such difference in numbers provided a large number of protein pairs requesting in-depth structure analysis.
Additionally, Supplementary Fig. S4 was further drawn to compare the assessing results of MELO and that of the software version of LDDT (LDDT-software, capable of performing high-throughput screening)27 when screening two million structure pairs. Till now, there had been no reported consensus on the LDDT cutoff for identifying the protein pairs of significant structural changes, and a threshold of LDDT < 0.6 (adopted by recently-published Foldseek42 and applied widely in previous studies43,44) was thus used in this study. As given in Supplementary Fig. S5 (an enlarged version of Q1 in Supplementary Fig. S4), a total of 147,663 structure pairs (identified by LDDT, but missed by MELO; highlighted using black dots) were discovered, and four typical exemplar pairs (selected from the corner of Q1) were further illustrated. As shown, those four exemplar pairs were only identified by the software version of LDDT as the ones of significant structural change, but “missed” by all other methods (such as MELO, TM-score and RMSD), and visual inspection confirmed that there was non-significant conformational change in all those four exemplar pairs. It is essential to emphasize that there is significant discrepancy between the evaluating outcomes of the software27 and server45 versions of LDDT. Particularly, as shown in Supplementary Fig. S5, there were dramatic differences between LDDT-server and LDDT-software, and the evaluating outcomes of LDDT-server were consistent with that of MELO, TM-score and RMSD (being completely opposite to that of LDDT-software assessed in the above section). However, LDDT-server cannot realize a high-throughput screening, making it extremely time-consuming to scan all two million structure pairs (our preliminary evaluation gave an estimation of 20,000 hours for completing whole scanning). In other words, in order to identify the cases where LDDT found structural changes but missed by MELO from millions of protein pairs, only LDDT-software could be adopted to enable a high-throughput screening. All in all, current versions of LDDT were either inaccurate in identifying protein pair of significant conformation change from massive amounts of data (LDDT-software) or incapable of realizing high-throughput screening due to its extremely time-consuming nature (LDDT-server).
Assessing the false positive rate of MELO in structure change discovery
To assess the false positive rate of our MELO in identifying the protein pairs of non-significant structural changes, a set of benchmark data was collected from PSCDB database46. Particularly, PSCDB categorized 839 protein pairs into 7 classes based on their conformation change before and after ligand binding, and a class titled ‘no significant motion’ comprised 311 pairs that give no significant movement upon binding (resolution <3.0 Å, sequence identity >95%) was found. This class of protein pair could be used as a suitable benchmark for assessing false positive rate of MELO and other existing tools in structure change discovery. As a result, our analysis found that MELO, TM‑score and RMSD achieved 100% accuracy (false positive rate equaled to zero) in identifying those 311 pairs as structurally unchanged. In contrast, the local version of LDDT misclassified 15 (false positive rate equaled to 4.8%) out of those 311 pairs as having structural change, and all 15 pairs were illustrated in Supplementary Fig. S6. As described, our visual inspections identified that non-significant structural change was found for 15 pairs, indicating a relatively higher false positive discovery rate of the local LDDT comparing with other methods (including MELO). Considering the good performance of MELO in discovering the SCinSV (as discussed in the above two sections), the false positive analyses above indicated that, compared with available methods, MELO performed better in discovering SCinSV without sacrificing the accuracy in discovering the structure pairs of non-significant changes (low false positive rate).
Comparing the methods’ abilities to locate the protein structure changes
To compare the performance of MELO with that of LDDT in locating the protein structure change, we first collected two benchmark datasets from Protein Structural Change Database (PSCDB)46. PSCDB database, to the best of our knowledge, provides location data for structural changes and further classifies all these changes into two major categories: ‘local motion’ and ‘domain motion’. As described by PSCDB, the ‘local motion’ is defined as the changes “occurring in a local protein segment” that are induced either upon or regardless of ligands binding, while the ‘domain motion’ is characterized by the changes “happening among protein domains” that are induced either upon or regardless of ligands binding. As stated in PSCDB publication46, “these structures represent a range of variations in the native structures that are associated with their molecular functions”. A total of 242 protein pairs with ‘local motion’ and 119 protein pairs with ‘domain motion’ are thus collected from PSCDB, which were utilized here as the testing data to compare the capacities of LDDT and SPSmeasur in locating the structural changes. The performances of LDDT and SPSmeasur in locating changes are then assessed using a well-established metric: maximum F1-score (Fmax), which had been widely used by many studies47,48,49. Performance comparisons between LDDT and SPSmeasur are enabled by computing the difference between their F1-scores (ΔF1). A positive ΔF1 indicates that SPSmeasur achieves better performance than LDDT, with the larger values reflecting greater improvements. Conversely, a negative ΔF1 suggests that LDDT performs better.
As shown in Supplementary Fig. S7a, the distribution of ΔF1 on the ‘local motion’ is overall right-skewed, indicating that most pairs have ΔF1 ≥ 0. Specifically, 94 pairs fell within the [0,0.1) range, 52 pairs within [0.1,0.2), 34 pairs within [0.2,0.3) & 19 pairs within [0.3,0.4). Cases where ΔF1 ≥ 0.9 are also observed, and the instances of ΔF1 < 0 are rare, showing only in [-0.4,0) range, with a few occurrences. These suggested that residue-wise SPSmeasur generally performed as well as or even better than LDDT in locating structural change for most local structural changes, with an observable improvement (ΔF1 ≥ 0) in 224 (92.6%) out of 242 protein pairs. For example, for the protein pair CL.74 (1KTG & 1KT9; ΔF1 is ~0.5), both the residue-wise SPSmeasur and LDDT-server can effectively capture the local structure changes indicated in PSCDB (as highlighted by a RED circle on the left side of Supplementary Fig. S7b and two GREEN circles on the right side of Supplementary Fig. S7b). However, as given by the purple circles and purple ribbons in the lower panel on the right side of Supplementary Fig. S7b, LDDT-server generated many false positive identifications of local structure changes, leading to lower specificity (89.2%) than that (100%) of SPSmeasur, and the F1 of LDDT-server (0.56) is therefore lower than that (1.00) of SPSmeasur. Supplementary Figs. S8-S12 provided detailed description of representative cases across the ΔF1 ranges from [0.9,1.0] to [0,0.1), where SPSmeasur’s structural change identification aligned more closely with the PSCDB-indicated regions. Moreover, as ΔF1 increased, SPSmeasur’s enhancements over LDDT-server in locating local structure change became more pronounced.
In addition to ‘local structure changes’, there are also structure changes between protein domains, which are equally critical for understanding protein function. Thus, we further tested the locating abilities of residue-wise SPSmeasur and LDDT-server on the dataset of ‘domain motion’ in PSCDB. The results identified that, residue-wise SPSmeasur outperformed LDDT-server in locating domain motion (Supplementary Fig. S13a). Particularly, there is no observation of protein pairs with ΔF1 < 0, indicating the good capability of our residue-wise SPSmeasur. Moreover, the peak of data distribution in Supplementary Fig. S13a was substantially shifted to the right side (ΔF1 > 0) compared with that in Supplementary Fig. S7a. There are 43 pairs in the [0.5,0.6) range and 24 pairs in the [0.6,0.7) range, which indicated that for protein pairs with domain motion, residue-wise SPSmeasur showed more improved locating performances over residue-wise LDDT-server in most of the studied protein pairs. For example, the protein pair ID.16 (1W9J-1FMV) gave a ΔF1 around 0.5, indicating a great inter-domain displacement. Residue-wise SPSmeasur provided better specificity (99.4%) and recall (85.5%) compared with those (85.7% and 39.1%, respectively) of residue-wise LDDT-server (as illustrated on the right side of Supplementary Fig. S13b). The Supplementary Figs. S14–S18 illustrated detailed case examples for protein pairs across ΔF1 range from [0.9,1.0] to [0,0.1), indicating that the discovery results of SPSmeasur aligned well with those indicated in PSCDB. Furthermore, as ΔF1 increased, SPSmeasur’s improvement over LDDT-server in locating the ‘domain motion’ became more pronounced.
Evaluating the dependency of methods on the protein sequence length
As reported, the measuring results of methods should not be significantly dependent on the size of analyzed proteins, a factor that can greatly impact the broad applicability of these methods50. In other words, if a scoring method shows significant fluctuation across the proteins of varying lengths, it may introduce biases in structure comparison, resulting in poor accuracy. In contrast, the highly-stable scoring methods can result in consistent evaluations across a range of protein sequence lengths, making them reliable for structure comparison20. Herein, the dependencies of four scoring methods (MELO, RMSD, TM-score and LDDT) on the sequence length of protein were analyzed based on the procedure described in ‘Calculating the Dependence of Methods on Protein Sequence Length’, and the evaluating results were illustrated in Supplementary Fig. S19. As provided in Supplementary Fig. S19a, with the increase of protein sequence length, the RMSD values showed an ascending trend in general by raising significantly from 1.5 to 5.7, which aligned with the observation of previous research22. In contrast, TM-score, LDDT, VSSmeasur and SPSmeasur remained relatively stable, indicating that RMSD was much more sensitive to the size of proteins compared with other three methods. To enable a direct comparison among TM-score, LDDT, VSSmeasur and SPSmeasur, they were further adjusted to a centered mean score value. As illustrated in Supplementary Fig. S19b, TM-score, LDDT, VSSmeasur, and SPSmeasur fluctuated around zero, maintaining their stability regardless of protein size. However, TM-score tended to be significantly lower when the protein sequence was short, which was consistent with the findings of previous study34. Meanwhile, LDDT gave a stronger oscillation around zero. As shown in Supplementary Fig. S19c, four violin plots of centered mean score values revealed the distribution of scoring results. The VSSmeasur and SPSmeasur gave tighter distributions around zero, indicating much higher consistency and lower dependency on protein size compared with existing methods, which made it reliable to measure SCinSV.
Among all methods, the RMSD is the only one without fixed score range. In this study, the raw measures of RMSD were further rescaled to a fixed score range of [0,1] using a transformation strategy identical to that of a previous report20. As depicted in Supplementary Fig. S20a, the rescaled RMSD (RMSDrescale, highlighted using BLACK line) indicated a clear dependency on protein size compared with MELO metrics, which gave the same conclusions as that reached in Supplementary Fig. S19a (great dependency of RMSD on the size of protein was observed). In contrast, as shown in Supplementary Fig. S20a and Supplementary Fig. S20b, the two metrics of MELO remained highly stable, indicating that our method was much less sensitive to the size of proteins compared with both RMSD and RMSDrescale.
Assessing the Reliance of Methods on Protein Pair’s Sequence Similarity
To evaluate methods’ reliance on the level of sequence similarity of studied structure pairs, two algorithms for sequence-based (Smith-Waterman51) and structure-based (US-align25) alignment were integrated into MELO. Based on these two algorithms, two million pairs of high sequence similarity ( > 70%) in Fig. 3c were assessed, which led to 3,682 pairs showing disagreements in MELO metrics. All results were provided in Supplementary Dataset. The visual inspections of all 3,682 pairs further identified that these disagreements arose primarily from an intentional staggering of the structurally divergent region by US-align for maximizing its global structural alignments25. Taking a typical structure pair (Mg²⁺-free vs Mg²⁺-bound KRAS; Supplementary Fig. S21) as example, the regions of βB & βC in KRAS were reported to have great structural shifts52. However, as shown in Supplementary Fig. S21a, those two regions (βB & βC) were staggered by the US-align (highlighted by BLUE dashed boxes), resulting in an underestimated value of MELO metric (SPSmeasur = 0.28; since massive amount of the values in the left triangle of the Supplementary Fig. S21c were completely missed for the regions of βB & βC).
When it comes to the application of the method based on Smith-Waterman alignment algorithm (S-W; given in Supplementary Fig. S21b), the regions of βB & βC were effectively matched (denoted by RED dashed boxes), resulting in a larger MELO metric (SPSmeasur = 0.61). In other words, the reported structure shift of βB & βC before and after Mg²⁺ binding (highlighted using PINK and BLUE ribbon, respectively, in Supplementary Fig. S21c) could be captured using S-W-based MELO, but the one based on US-align could not discover this shift. Other examples could be found in Supplementary Dataset. In conclusion, for structural pairs of high sequence similarity ( > 70%), structure-based alignment is more likely to underestimate structural changes compared with the sequence-based one when being integrated into MELO. This finding aligned with previous work53, stating: “when two sequences can be aligned in a statistically meaningful way, sequence-based structural superposition offers good measure of structural changes”.
Furthermore, the S-W-based algorithm was found capable of discovering relationships between proteins whose sequence identities are > 30%, but prone to underperforming in the protein pairs of low sequence similarity54. Thus, MELO’s performance on measuring the protein pairs of low sequence similarity was evaluated. As shown in Supplementary Fig. S22a, a pair of remote homologs (the B1 domain of human protein G & human protein B, sharing sequence similarity of only 17.3%) was reported to show highly similar structures55. The S-W-based and US-align-based metrics were computed and shown in Supplementary Fig. S22b and Supplementary Fig. S22c, respectively. Particularly, significant structure change was reported by S-W-based MELO metric (VSSmeasur = 0.58 & SPSmeasur = 0.94), while non-significant structure change was reported by the US-align-based one (VSSmeasur = 0.40 & SPSmeasur = 0.48). Such result indicated the greatly enhanced performance of the structure-based alignment method, compared with the sequence-based one, in measuring the pairs of low sequence similarity (identity ≤30%).
According to the analyses above, the MELO was designed to incorporate a hybrid strategy that stipulates: “when the sequence identity of a protein pair is >30%, S-W-based alignment will be used; when the identity is ≤30%, US-align-based one will be applied”. This hybrid strategy had been incorporated, as a default setting, into all three versions of MELO (online server, software tool, and command-line package). Meanwhile, the users could also customize their selection of algorithm (either S-W or US-align) that best aligned with their preferences or requirements.
Mitigating the Influence of Protein Flexibility on Structure Comparison
Proteins in solution, especially those linear ones, usually exhibited inherent flexibility56. When protein segments are far apart and lack direct interactions, protein flexibility can lead to natural displacement among those distant segments57. Such displacement was usually considered to be a natural manifestation of structure dynamics, but frequently misinterpreted by existing scoring methods as significant structure changes58. Taking the baboon theta-defensin-2 (BTD-2) at two timepoints in solution59 (offered in Supplementary Fig. S23a) as an example, the observed differences between two structures mainly stemmed from the intrinsic flexibility of the protein, particularly in the highly flexible β-strand region. Under such circumstance, if the difference of the relative distances among amino acids was directly used to measure the shifts among protein segments, the structure changes induced by those intrinsic flexibilities would be overestimated. Particularly, as shown in the heatmap based directly on the relative distance in Supplementary Fig. S23b, four pairs of protein segments (labeled as S1, S2, S3 & S4 by boxes of PURPLE, PINK, BLUE and GREEN, respectively) that were far apart in the structure showed significant changes in their pairwise distances. This outcome indicated that the flexibility-driven shift was identified as significant structure change. To address this issue, our MELO introduced weighted normalization of relative distances among pairwise residues, effectively mitigating the bias induced by protein’s flexibility. As provided in the PLOTSPS of MELO offered in Supplementary Fig. S23c, after weighted normalization, those flexibility-driven shifts (S1-S4) were not considered by MELO as great structural change, which minimized the misleading overestimation induced by distantly-separated residues.
Case study comparison of MELO performance with that of existing methods
To assess the ability of methods in discovering SCinSV, four typical types of structure changes were identified from literatures and evaluated by scoring methods. These four types of changes included those a) indicating different functional states (ATP-dependent transporters’ variations between inward- and outward-facing states), b) induced by mutations/ligand binding (structural changes of cellular retinoic acid-binding protein induced by missense mutations/the binding of retinoic acid), c) caused by ion chelation (changes in GTPase KRAS’s structure before and after magnesium ion chelation), and d) stimulated by environmental change (glucokinase’s structure changes under three different glucose concentrations). Moreover, the structure change of multi-chain protein complex was reported to drive inter-subunit interactions and allosteric regulation, enabling functional assembly and activity transition that are not achievable through monomeric protein structural changes alone60. All these changes discussed above were complicated, which made the traditional visual inspection very subjective and impractical for large-scale analysis15. Furthermore, different superposition methods could bring about distinct findings depending on the applied structure alignment algorithms25. To assess the ability of MELO in addressing these issues, its performances on these five cases were compared with that of existing methods.
Performance assessment using ABC transporters for substrates transportation
ABC transporters played crucial role in the uptake/expulsion of substance in cells61. Two types of ABC transporters were studied here, which included ABCB1 and ABCG262. As illustrated in Fig. 4a, b the structures of two transporters contained two chains (Chain A and Chain B), and each chain consisted of a transmembrane domain (TMD) and nucleotide-binding domain (NBD)61. For ABCB1, the 1st-3rd α-helices, 4th-5th α-helices, and 6th α-helix & NBD1 in Chain A were grouped into A1, A2, and A3, respectively, while the 7th-9th α-helices, 10th-11th α-helices, and 12th α-helix & NBD2 in Chain B were appointed into B1, B2, and B3, respectively (shown in Fig. 4a). For ABCG2, its Chain A and Chain B were also illustrated in Fig. 4b. Those transporters (ABCB1 and ABCG2) were reported to undergo a conformational transition between two states (inward-facing and outward-facing) during their substrates transportation50. Both conformational states were illustrated for ABCB1 (as provided in Fig. 4c) and ABCG2 (as provided in Fig. 4d), which revealed the substantial structure difference between ABCB1 and ABCG2. Particularly, the TMDs in the Chain A & Chain B of ABCB1 interwove with each other to form two V-shaped structures (as described in the schematic diagram of Fig. 4c, A1, B2, and A3 were placed on the left size of ABCB1, and B1, A2, and B3 were on the right side); in contrast, the TMDs in the Chain A & Chain B of ABCG2 were largely independent, with no interweaving between two chains (as provided in the schematic diagram of Fig. 4d, Chain A was placed on the left size of ABCG2, and Chain B was located on the right side).
a schematic representation of ABCB1’s structure. This structure contained two chains (Chain A and Chain B), and each chain consisted of a transmembrane domain (TMD) and a nucleotide-binding domain (NBD). As provided, the 1st-3rd α-helices, 4th-5th α-helices and 6th α-helix-NBD1 in the Chain A were grouped to A1, A2 and A3, respectively, and the 7th-9th α-helices, 10th-11th α-helices and 12th α-helix-NBD2 in the Chain B were assigned into B1, B2 and B3, respectively. b schematic illustration of ABCG2’s structure. This structure contained two chains (Chain A and Chain B), and each chain consisted of one TMD and one NBD. c schematic illustration of ABCB1’s inward-facing and outward-facing states. The TMDs in Chain A and Chain B of ABCB1 interwove with each other to form two V-shaped structures. d schematic representation of ABCG2’s inward-facing and outward-facing states. The TMDs in Chain A and Chain B of ABCG2 were largely independent, with no interweaving between two chains. e assessing the structure change between the inward-facing state (IF, PDB:6A6N) and the outward-facing one (OF, PDB:6A6M) of ABCB1 using different scoring methods. Substantial shifts among six structure segments (A1, A2, A3, B1, B2, and B3) were discovered by MELO (VSSmeasur = 0.27 & SPSmeasur = 0.58), and one out of three existing methods (RMSD) identified the structure change. The PLOTSPS of MELO was depicted, which identified nine pairs of protein segments of obvious structure shifts (such as A1 vs B3, A3 vs B3, and B2 vs B3; largely colored in RED). f assessing the structure change between the inward-facing state (IF, PDB:6VXH) and outward-facing one (OF, PDB:6HZM) of ABCG2 using different methods. Extensive shifts between Chain A and Chain B were discovered by MELO (VSSmeasur = 0.37 & SPSmeasur = 0.57), and two out of three methods (LDDT and RMSD) identified the structure change. The PLOTSPS was also depicted, which captured one segment pair of obvious structure shifts (Chain A vs Chain B; largely colored in RED).
To reveal the structure changes of ABCB1 and ABCG2 between conformational states (inward-facing and outward-facing), four scoring methods (MELO, TM-score, LDDT, and RMSD) were employed. As offered in Fig. 4e, substantial shifts among six protein segments (A1, A2, A3, B1, B2, and B3) were captured by MELO (VSSmeasur = 0.27 & SPSmeasur = 0.58; being in the Q2 of Fig. 3c), and one out of three existing methods (RMSD) identified the structural changes. Meanwhile, as described in Fig. 4f, a great shift between two chains (Chain A and Chain B) were discovered by MELO (VSSmeasur = 0.37 & SPSmeasur = 0.57; being in the Q2 of Fig. 3c), and one out of three methods (RMSD) captured the structure changes. Such results showed the good performance of MELO and RMSD in capturing structure changes of two transporters.
Moreover, the mechanisms underlying the structure change of ABCB1 & ABCG2 were distinct from each other50,63. Particularly, the transportation of substrates by ABCG2 was reported to be mainly induced by an inter-chain conformation change63 between Chain A and Chain B (shown in Fig. 3d), while substrate transportation by ABCB1 was found closely associated with two types of mechanism50 (shown in Fig. 3c), including an inter-chain change between its Chain A and Chain B and the inner-chain changes (between the segments B2 and B1-B3; between the segments A2 and A1-A3). In order to locate these sophisticated changes of both inter-chain and inter-segment types, the residue-wise LDDT, RMSD, and SPSmeassur were computed to evaluate whether the change could be captured. Particularly, for LDDT, both the alignment and residue-wise calculation were realized by SWISS-MODEL45; for RMSD, the align function in PyMOL was adopted to superimpose structures and compute residue-wise measurements; for SPSmeassur, after alignments, a residue-wise average value of the normalized difference of relative distance (Eq. 3) for each amino acid was calculated. Then, a visualization of residue-wise LDDT, RMSD, and SPSmeassur was implemented (the details of these implementations and all necessary source codes were made publicly available on GitHub at https://github.com/idrblab/MELO). Finally, the above calculations were applied to specific protein pairs to realize the computation of and comparison among residue-wise LDDT, RMSD, and SPSmeassur. Taking the structure pair (inward-facing and outward-facing) of ABCB1 offered in Fig. 4 as example, its residue-wise LDDT, RMSD, and SPSmeassur were calculated and demonstrated in Fig. 5a. As offered, clear structure changes in the substrate entrance region (at the bottom of ABCB1 in Fig. 5a) & the transmembrane region were identified by both residue-wise SPSmeassur and residue-wise RMSD (highlighted by gradual RED shading; the redder it is, the greater the change); the residue-wise LDDT could also find certain level of changes in the substrate entrance region, but highlighted obvious change at the top of ABCB1 instead of its transmembrane region. Those changes in the protein segment A2 of ABCB1 colored by residue-wise LDDT, RMSD and SPSmeassur were also illustrated in Fig. 5b. Taking Fig. 5a and Fig. 5b together, although there is noticeable divergence among methods’ findings, they are all able to indicate local structural changes.
a the structure alterations captured by residue-wise LDDT, RMSD, and SPSmeasur. Clear structure changes in the substrate entrance region (at the bottom of ABCB1) & the transmembrane regions were discovered by residue-wise SPSmeasur and residue-wise RMSD (highlighted by gradual RED shading; the redder it is, the greater the change); the residue-wise LDDT could also find certain level of changes in the substrate entrance region, but highlighted changes at the top of ABCB1 instead of its transmembrane regions. b the structure alterations in the segment A2 of ABCB1 discovered by residue-wise LDDT, RMSD, and SPSmeasur. c the PLOTSPS in MELO that captures the inter-chain & inter-segment changes. The structure change of segment A2 relative to five other segments (A1, A3, B1, B2 and B3; RED dashed box) were visualized, the conformation alterations of which were further depicted in (d) A2 relative to A1, e A2 relative to A3, f A2 relative to B1, g A2 relative to B2 and (h) A2 relative to B3. One inter-chain change (A2 relative to B2), two inner-chain changes (A2 relative to A1; A2 relative to A3), and two non-significant changes (A2 relative to B1; A2 relative to B1) were identified.
Furthermore, as demonstrated in Fig. 4e, f the PLOTSPS of MELO was depicted to further discover those changes of inter-chain & inter-segment types, because it offered the in-depth information describing the inter-residue changes of distance. Taking the ABCB1 demonstrated in Fig. 4e as an example, the PLOTSPS of MELO discovered a total of nine pairs of protein segments with noticeable structure shifts (such as A1 vs B3, A3 vs B3, and B2 vs B3; largely colored in RED). If all nine identified segment pairs were collectively considered, an inter-chain change between Chain A and Chain B together with the inner-chain changes (between segments B2 and B1-B3; between the segments A2 and A1-A3) could be captured by MELO, which were consistent with previous study50 and the schematic diagram shown in Fig. 4c. Similarly, when it came to the ABCG2, the PLOTSPS was also illustrated in Fig. 4f, which captured the inter-chain changes between Chain A vs Chain B (mostly colored in RED) that were described in Fig. 4d.
The ability of MELO’s PLOTSPS in capturing the inter-chain & inter-segment changes was also described by Fig. 5c, since it could be applied to analyze the relative changes among protein segments. As shown, the structure change of protein segment A2 relative to five other segments (A1, A3, B1, B2 and B3; RED dashed boxes) were calculated and then visualized in the Fig. 5d (A2 relative to A1), Fig. 5e (A2 relative to A3), Fig. 5f (A2 relative to B1), Fig. 5g (A2 relative to B2) and Fig. 5h (A2 relative to B3). As demonstrated, one inter-chain change (between A2 and B2) and two inner-chain changes (between A2 and A1 & between A2 and A3) were identified, which successfully reproduced the observation in previous report50. Moreover, the non-signification structure change of A2 relative to B1 (Fig. 5f) and B3 (Fig. 5h) was discovered by MELO, which also aligned well with the schematic diagrams in Fig. 4c. All in all, the PLOTSPS realized a highly flexible recognition and visualization of the relative structure changes among protein segments, which had been integrated into the online software tool.
Performance Assessment Using CRABPⅡ during Mutation and Retinoic Binding
The protein cellular retinoic acid-binding protein 2 (CRABPⅡ) was found to play a crucial role in intracellular transport and retinoic acid regulation, which was critical for maintaining normal cellular function64. Apart from the most common conformations (a total of 39 conformations of identical sequences deposited by different research laboratories), holo-CRABPⅡ (HC), binding retinoic acid65, two additional conformations were collected from PDB, such as: apo-CRABPⅡ (AC) without retinoic acid66, and mutant CRABPⅡ (MC) with the R35D and K36D mutation67. To reveal the structure changes induced by mutation (between HC and MC) and ligand binding (between HC and AC), four methods (MELO, TM-score, LDDT, and RMSD) were adopted. As provided in Fig. 6a, extensive shifts among the protein segments between MC and HC were successfully revealed by MELO (0.21 <VSSmeasur < 0.30 & 0.58 <SPSmeasur < 0.61; being in the Q2 of Fig. 3c), and none of those available methods discovered the structure changes. In the meantime, as shown in Fig. 6a, both the variations in secondary structures and shifts among protein segments between AC and HC were found by MELO (0.53 <VSSmeasur < 0.58 & 0.55 < SPSmeasur < 0.59; being in the Q3 of Fig. 3c), and two methods (LDDT & RMSD) found the structural changes. Furthermore, Fig. 6a revealed that no significant change in structure was observed among 39 conformations of HC by MELO (0.01 <VSSmeasur < 0.21 & 0.03 <SPSmeasur < 0.26; being in the Q1 of Fig. 3c), which highlighted the stability and accuracy of MELO in discriminating changed structure from the unchanged one. Since VSSmeasur and SPSmeasur ranged from 0 to 1 (like TM-score and LDDT), Fig. 6b employed violin plot to compare these three scores by analyzing their abilities to discern the structure change among three conformations of CRABPⅡ. Particularly, our MELO was able to identify the great shifts among protein segments between MC and HC, and discover both the variations in secondary structure and shifts among protein segments between AC and HC. However, TM-score could not identify structure change not only between MC and HC but also between AC and HC, and LDDT was also not functional enough to effectively identify the corresponding structure changes.
Apart from the most common conformations (39 conformations of identical sequences deposited by different research groups), holo-CRABPⅡ (HC), which bound retinoic acids, two conformations were collected from PDB: apo-CRABPⅡ (AC) without retinoic acid and mutant CRABPⅡ (MC) with the R35D and K36D mutation. a evaluating outcome for three groups of structure pairs (HC vs HC, MC vs HC, and AC vs HC) based on VSSmeasur (x-axis) and SPSmeasur (y-axis) of MELO. Great shifts among protein segments between MC and HC were captured by MELO (0.21 <VSSmeasur < 0.30 & 0.58 <SPSmeasur < 0.61). Both the variations in secondary structures and shifts among protein segments between AC and HC were discovered by MELO (0.50 <VSSmeasur < 0.53 & 0.55 <SPSmeasur < 0.59). No significant change in structure was observed among 39 conformations of HC by MELO (0.01 <VSSmeasur < 0.21 & 0.03 <SPSmeasur < 0.26), which illustrated the stability and accuracy of MELO in differentiating changed structure and the unchanged one. b violin plots for comparing three methods by analyzing their abilities to discern the structure change among three conformations of CRABPⅡ. MELO was able to identify the extensive shift among protein segments between MC and HC, and reveal the variations in secondary structure and shifts among protein segments between AC and HC. TM-score could not identify structure change not only between MC and HC but also between AC and HC, and LDDT was not useful enough to identify the corresponding structure changes. c assessing structure change between mutant CRABPⅡ (MC, PDB:7OXW) and holo-CRABPⅡ (HC, PDB:6HKR) based on different methods. Only MELO could capture the shifts among segments (VSSmeasur = 0.25 & SPSmeasur = 0.59). d evaluating the structure change between apo-CRABPⅡ (AC, PDB:1BLR) and holo-CRABPⅡ (HC, PDB:6HKR) using different methods. Only MELO could detect the variations in secondary structure and shifts among protein segments (VSSmeasur = 0.53 & SPSmeasur = 0.58.
According to the reported structures67, the details of structural changes between MC (colored in RED) and HC (colored in BLUE) were described in Fig. 6c, indicating that the MELO could identify the shifts among segments (VSSmeasur = 0.25 & SPSmeasur = 0.59; being in Q2 of Fig. 3c). Moreover, the mutations of R35D and K36D could further induce the disappearance of the binding pocket of retinoic acid67, which was identified by the PLOTSPS of MELO (as offered in Supplementary Fig. S24a). As illustrated, significant shifts were discovered for the second helix (depicted in Supplementary Fig. S24b), the βC-βD (given in Supplementary Fig. S24c) & the βE-βF (offered in Supplementary Fig. S24d), which made the binding pockets of retinoic acid disappeared (from BLUE to RED). Similarly, the structure change between AC (colored in PINK) and HC (colored in BLUE) was presented in Fig. 6d, illustrating that only MELO could detect the variations in secondary structure and the shifts among protein segments (VSSmeasur = 0.53 & SPSmeasur = 0.58; being in the Q3 of Fig. 3c). Meanwhile, the absence of retinoic acid was reported to make binding pocket expand significantly and secondary structure change extensively66, which was captured by the PLOTVSS and PLOTSPS of MELO (as provided in Fig. 7a, b). As described, extensive variations were discovered for the second helix, βD and βJ (as described in Fig. 7a, c). Meanwhile, significant shifts were discovered for the second helix (as illustrated in Fig. 7d), and βE-βF (as described in Fig. 7e), which made the binding pocket expanded extensively (from BLUE to PINK). In summary, based on the above analyses, MELO was found capable of identifying not only the variations in secondary structures but also the shifts among protein segments in the protein CRABPⅡ.
a the PLOTVSS generated by MELO for locating the variations in secondary structure. Three regions of obvious alterations were highlighted, which included the second helix (purple box), βD loop (brown box), and βJ loop (umber box). T: β-turn; E: β-strand; B: β-bridge; H: α-helix; I: Π-helix; G: 310-helix; P: κ-helix. b the PLOTSPS generated by MELO for locating the shifts among protein segments with darker color showing regions of significant displacements. The second helix (purple box) and the βE-βF loop (umber box) were captured by MELO as key regions of segmental shifts. c structural superimpositions of apo-CRABPII (AC, PDB:1BLR) and holo-CRABPⅡ (HC, PDB:6HKR), providing main variations in secondary structures (such as the second helix (purple box), the βD loop (brown box) & the βJ loop (umber box). d shift was discovered for the second helix. e shift was identified for the βE-βF. All in all, the MELO was found capable of identifying not only the variation in secondary structure but also the shift among protein segments in the studied protein of cellular retinoic acid-binding protein 2.
Performance assessment using GTPase KRAS before and after Mg2+ Chelation
The KRAS protein was a membrane-bound small GTPase that played a key role in cell growth, differentiation, and survival68. As offered in Supplementary Fig. S25a, the magnesium ion (Mg2+) acted as a critical cofactor, which facilitated the folding of KRAS structure to its active conformation69. To reveal the conformation alteration before and after Mg2+ chelation (between Mg2+-free and Mg2+-bound), three existing methods (MELO, TM-score, and LDDT) were used. As given in Supplementary Fig. S25b, great shifts among the protein segments between the Mg2+-free KRAS and Mg2+-bound one were accurately identified by MELO (0.16 <VSSmeasur < 0.25 & 0.60 <SPSmeasur < 0.64; being in Q2 of Fig. 3c), and the other two could not capture such change. Furthermore, Supplementary Fig. S25b revealed that no significant change in structure was discovered among those 42 conformations of Mg2+-bound KRAS by MELO (0.01 <VSSmeasur < 0.22 & 0.03 <SPSmeasur < 0.29; being in the Q1 of Fig. 3c), which denoted the stability and accuracy of MELO in discriminating changed structures from the unchanged ones. Since VSSmeasur and SPSmeasur ranged from 0 to 1, Supplementary Fig. S25c applied a violin plot to further compare three methods by analyzing their capabilities of discerning the structure change between Mg2+-free KRAS and Mg2+-bound one. All in all, our MELO could identify the critical conformation alteration in KRAS before and after Mg2+ chelation.
Based on the reported structure52, the details of structural change between Mg2+-free KRAS (in PINK) and Mg2+-bound one (in BLUE) were demonstrated in Supplementary Fig. S25d and Supplementary Fig. S25e. Upon the loss of Mg²⁺, the βB (given in Supplementary Fig. S25d) shifts significantly from the lower to upper part of the protein52, which was successfully captured by the PLOTSPS of MELO. Simultaneously, as given in Supplementary Fig. S25e, the βC and βD moved away from the pocket, which was also discovered by the PLOTSPS of our MELO. Furthermore, compared with those grids within the area demarcated by the brown lines of Supplementary Fig. S25e, those colored by purple lines in Supplementary Fig. S25d were much darker in red color, which indicated a more dramatic shift in βB compared with that in βC and βD. All in all, our MELO could not only locate the conformation alteration of KRAS induced by Mg2+ chelation but also capture the varying degrees of segmental changes.
Performance assessment using glucokinase in various glucose concentrations
The glucokinase (GCK) was key for the regulation of blood glucose homeostasis70. Changes in glucose concentration could lead to a transition of GCK among three different conformations71. As provided in Supplementary Fig. S26a, GCK transitioned from super-open conformation (SC) to intermediate-open one (OC), and then to closed one (CC) with the elevation of glucose concentration72. Such transition mainly involved a reduction in the angle between the large and small domains (from 100° to 65°, and then to 40°). To reveal these transitions (among SC, OC, and CC), three scoring methods (MELO, TM-score, and LDDT) were applied. As illustrated in Supplementary Fig. S26a, extensive shift among the protein segments between SC and CC was successfully identified by MELO (0.27 < VSSmeasur < 0.31 & 0.60 < SPSmeasur < 0.62; being in the Q2 of Fig. 3c), and none of those available methods discovered such structure change. Meanwhile, the structure changes between OC and CC could not be detected by MELO (0.18 <VSSmeasur < 0.23 & 0.41 < SPSmeasur < 0.48; being in the Q1 of Fig. 3c), but there was a great variation between the distribution of OC vs CC (colored in LIGHT RED) and that of CC vs CC (colored in GREEN) offered in Supplementary Fig. S26a. Moreover, no detectable change in structure was observed among 20 conformations of CC by MELO (0.01 < VSSmeasur < 0.16 & 0.01 < SPSmeasur < 0.15; being in Q1 of Fig. 3c), which denoted the stability and accuracy of MELO in discriminating changed structures from the unchanged ones. Violin plots were further employed in Supplementary Fig. S26b to compare the abilities of three methods to capture the transition of GCK among three conformations. Particularly, our MELO was able to identify the shifts among protein segments between SC and CC, but the TM-score and LDDT could not effectively identify the transition of GCK among three structural conformations.
Based on reported structures73, the transition between SC (colored in RED) and CC (colored in BLUE) were demonstrated in Supplementary Fig. S26c. Extensive shift between the ‘small domain’ and ‘large domain’ was reported, and a complicated internal shift within small domain was also observed73, which were successfully captured by the PLOTSPS of MELO (as offered in Supplementary Fig. S26c). Particularly, the darker RED color in the regions demarcated by brown lines highlighted the shift between small domain and large domain, and that demarcated by the blue lines indicated the internal shift within small domain. In the meantime, as provided in Supplementary Fig. S26d, compared with the structure change between CC and SC, that between OC (colored in PINK) and CC (colored in BLUE) were similar, but the degree of shift between small and large domain was much smaller, which was also effectively captured by the PLOTSPS of MELO. All in all, our MELO could not only locate the regions of structure changes during GCK’s transition but also capture the different degrees of such changes.
Performance assessment using heterotetramer adaptor protein complex AP2
The adaptor protein complex AP2 is a heterotetramer (composed of four chains of A, B, M, and S), which functions in protein transport via clathrin-coated vesicles in various membrane traffic pathways74. As demonstrated in Supplementary Fig. S27a, the AP2 undergoes a substantial structural transition from the closed conformation (CC) to an opened one (OC), which allows 2 critical cargo-binding motifs ([ED]xxxL[LI] and YXXΦ) to be exposed75. Such conformational changes involved two key components75: c1) a screw rotation of C-μ2 (one part of chain M) by ~127° about its long axis, with a 39 Å displacement, relative to N-μ2 (another part of chain M); c2) an inward collapse of the AP2 bowl, which is composed of chain A (α subunit) and chain B (β2 subunit). Based on two PDB structures (2VGL & 2XA7) of AP2 in different conformations (CC & OC), an analysis on the CC-OC transition was conducted based on MELO and available methods, and their performances in identifying such transition were carefully compared.
First of all, as shown in Supplementary Fig. S27b, substantial segmental shift between CC (PINK color) and OC (BLUE color) was captured using MELO (VSSmeasur = 0.31 & SPSmeasur = 0.72; being in the Q2 of Fig. 3c), whereas this transition could not be successfully identified by other existing methods (such as TM-score & LDDT) except for RMSD (9.258). Particularly, as offered by the RED color regions in MELO’s PLOTSPS of Supplementary Fig. S27b, the most prominent change is a movement of C-μ2 (one part of chain M), relative to N-μ2 (another part of chain M) and other chains of A, B and S (39 Å displacement between 2VGL and 2XA7). Thus, chain M, composed of C-μ2 & N-μ2, was further assessed and shown in Supplementary Fig. S27c. As shown, protein segment shift between C-μ2 and N-μ2 (depicted by component c175 above) was effectively found by both MELO (VSSmeasur = 0.384 & SPSmeasur = 0.732; being in Q2 of Fig. 3c) and RMSD, and the RED regions in MELO’s PLOTSPS of Supplementary Fig. S28c further highlighted the relative displacement of 39 Å for C-μ2 reported in previous study75. Second, the AP2 bowl (composed of chain A and chain B) was analyzed and illustrated in Supplementary Fig. S28d. As described, the inward collapse of AP2 bowl (demonstrated by component c275 above) was successfully identified by MELO (VSSmeasur = 0.279 & SPSmeasur = 0.549; being in the Q2 of Fig. 3c) and RMSD (9.77), and the noticeable RED color region in MELO’s PLOTSPS provided in Supplementary Fig. S28d further highlighted the collapse of the AP2 bowl during CC-OC transition that was discovered by previous publication75.
Meanwhile, it was reported that the chain A of AP2 remained “largely unaltered” in the CC-OC transition75, which inspires us to perform additional study on chain A using MELO and existing methods. As shown in Supplementary Fig. S28, MELO (VSSmeasur = 0.21; SPSmeasur = 0.36; being in Q1 of Fig. 3c), TM-score (0.74) and LDDT (0.79) did not identify a conformational change. In contrast, the RMSD (6.24) falsely suggested a significant structure change, which is in good agreement with a previous study76 reporting that the RMSD is prone to generating false positive discovery. Taking all the findings above together, our MELO was found able to capture both of the key components (c1 and c2) in the conformational transition of AP2, and capable of successfully avoiding the false discovery of the structurally-unchanged chain A.
Locating the subtle but critical protein structure variation using the MELO
In living organisms, protein structural changes are sometimes extremely small, yet these minor changes can result in significant functional alterations77. Although MELO gave good sensitivity when detected SCinSV, it might not be capable of detecting such extremely small change at the global level simply based on the metrics of VSSmeasur and SPSmeasur. Therefore, the PLOTVSS and PLOTSPS were offered by MELO to meet the crucial demands on highlighting the local changes of functional significances. Particularly, the PLOTVSS and PLOTSPS could provide the variations of secondary structure and the shifts among protein segments, respectively. Taking the myocyte enhancer factor 2B (MEF2B) as an example, a D83V mutation could greatly affect its binding to DNA and the subsequent gene activations78. As offered in Supplementary Fig. S29a, the D83V mutation disrupted the interaction forces stabilizing α-helix3 (both A and B), weakening the α-helix structure and transforming it to a disordered region79. Such changes were extremely subtle (as offered in Supplementary Fig. S29b), which led to the incapability of all existing methods (including MELO) of successful discovery at the global levels. However, the PLOTVSS (described in Supplementary Fig. S29c) and PLOTSPS (provided in Supplementary Fig. S29d) of MELO could not only indicate the transformation of the α-helix3 to disordered region (indicated by purple box) but also identified the corresponding shifts (denoted by the darker red color in the area demarcated by purple lines). All in all, although it was challenging to discover the extremely small structural change at the global level, our MELO could provide critical local information facilitating functional study, demonstrating its capability.
Developing web-server providing SCinSV data and identifying SCinSV
Herein, a unified metric titled ‘MELO-score’ was therefore proposed to assess whether there was structural changes between proteins. MELO-score was defined as the maximum of VSSmeasur and SPSmeasur. If MELO-score ≥ 0.5 (VSSmeasur ≥ 0.5 or SPSmeasur ≥ 0.5), a structural change was found. To show the functionality of MELO in measuring and locating the structure change of protein, a web-server (https://idrblab.org/melo/) was constructed, which offers the following capabilities: a) realizing customized assessments by allowing users to upload their own structures, b) offering downloadable version of MELO for enabling high-throughput analysis & protecting data security, c) providing > 10,000 structure changes identified by MELO, and d) offering in-depth description of both measurement and location for all those structural changes. To ensure the user-friendliness of this online server, two actions were further made: [a1] a result notification function that allows users to receive the calculation outcome through email after task submission is incorporated; [a2] a local software, enabling a high-throughput comparison of structures of any size, was developed. These can either avoid keeping user waiting for too long or safeguard user data privacy.
Moreover, all those two million protein pairs (sequence identity > 70%) and those 12,562 SCinSV structure pairs were made available for download from our server (https://idrblab.org/melo/). We provided these data publicly for two reasons. First, as described by one important previous publication3, “establishing a database for storing structure-disrupting mutations may allow for future renditions of AlphaFold2 or other artificial intelligence programs to include this information in their protein-folding predictions”, the data made publicly-available can partially meet the important need in this regard. Second, those data can also serve as a testing dataset for evaluating method. Especially, when a method is developed, the data can be used to assess its performance by comparing with that of the existing ones (including the MELO). Two functions were also incorporated to our online server to facilitate the interpretations of those data. (a) the data were linked to their corresponding subtle variations (such as mutation, ligand binding, & environmental change). For instance, structure change between two KRAS proteins was found by MELO, and their corresponding variations in ion binding (with/without Mg²⁺) was also described. In other words, those data may be useful for the actual structure analysis by considering the internal/external factor that probably induces structure change. (b) the data were also linked to their corresponding function annotation. Our analysis of those >10,000 structural changes discovered by MELO found that 22.8% of those protein pairs had their proteins different in functions. Taking the E. coli transcription factor RfaH as example (as demonstrated in Supplementary Fig. S30), two conformations were provided (2OUG & 2LCL) in our server, and their corresponding protein functions were also collected via matching to InterPro80. In particular, 2OUG was matched to IPR010215, indicating its inhibition of RNA polymerases; 2LCL was matched to IPR014722, highlighting its promotions of ribosome recruitment. In other words, our server helped to establish possible relation between SCinSV and its corresponding function alteration for 22.8% pairs. Furthermore, for the remaining 77.2% pairs annotated with identical functions, there may be undiscovered functional change awaiting further investigations, and these unexplored cases may offer a starting point for structure biologists when conducting analysis on revealing structure-function relation. Finally, we would like to emphasize that the main innovation and contribution of this work lie in the MELO methodology, not in those data that have been made publicly available only as supportive data source.
Methods
Collecting benchmark datasets for assessing methods’ performances
A total of 72,544 structural domains were directly collected from the SCOP2 database32, which was known to group the protein structures into a hierarchical system of four levels (from class, to fold, then to superfamily, and finally to family; indicating the progressively finer distinctions in protein architectures32). The levels of ‘class’ and ‘fold’ indicated substantial variations in the ‘major secondary structures’ and ‘arrangements of secondary structural elements’, respectively, and the levels of ‘superfamily’ and ‘family’ described those structural differences distinguishing the ‘evolutionary origin’ and ‘homological similarity’ among proteins, respectively32. To assess the ability of methods in differentiating the structure changes of varied magnitudes (from class, to fold, to superfamily, to family; which denoted clear descending magnitude), a comprehensive set of protein pairs of structural variation was prepared for different levels of SCOP2 hierarchy. Taking the family level as an example, all structures were first collected from each of the 5,936 families, and all structures in one family were then paired with that in others. Similar procedure was repeated for the remaining three levels, which led to 0.308, 0.409, 0.411, and 0.413 billion pairs of structural variation for the level of class, fold, superfamily, and family, respectively. To further evaluate the accuracy of methods in identifying structural changes induced by relatively small sequence alteration, the protein pairs with sequence identity >70% were selected.
Moreover, a total of 38,128 structures of single proteins (which were unaffected by interactions among proteins in a complex) were collected from PDB35 to assess methods’ ability to discover the structure change induced by subtle variation (SCinSV). Such kind of variation included the small sequence alteration (such as missense mutation3), environmental change4, ion chelation5, ligand binding6, etc. To retrieve those critical data of SCinSV, a large-scale sequence similarity comparison between any two of the 38,128 proteins collected above were performed, and those pairs of >70%81 sequence similarity score (known as ‘identity’ in BLAST) were then collected, which resulted in more than two million structure pairs. Meanwhile, a variety of representative protein pairs indicating different types of SCinSV were further identified based on the resulting pairs, and these representative pairs were used to support five case-analyses in this research.
Preprocessing based on the structure preparation and representation
Before measuring protein structure changes, the MELO implemented a preprocessing workflow to guarantee the robustness and accuracy of the subsequent assessment, which consisted of two essential steps: structure preparation and representation (as illustrated in Figs. 1a and 2a). For the first step, experimentally-induced missing amino acids and incomplete side chains were repaired using binary search method82 and PDBFixer83, respectively, to produce complete input structure, which was key for avoiding alignment error and ensuring accurate comparison. For the second step, a hybrid strategy was applied, stipulating “when the sequence identity of a protein pair is > 30%, S-W-based alignment will be used; when the identity is ≤30%, US-align-based one will be applied”. This hybrid strategy was adopted here to establish the residue-level correspondences between two structures, which enabled the calculation of pairwise amino acid geometric characteristics and relative distances for measuring the structure changes.
Measuring secondary structure variation and protein segment shift
To comprehensively assess structure change, secondary structure variation and protein segment shift were measured by collectively considering the alterations in geometric characteristics and relative distances among residues. As a result, two numerical values indicating the variations of secondary structures (VSSmeasur) and shifts of protein segments (SPSmeasur) were calculated.
Calculating VSSmeasur based on the geometric characteristics among residues
Protein folding was governed by the complex interplay of forces, including hydrophobic effect, hydrogen bond, and other interactions84. The secondary structure of protein was determined by the Geometric characteristics among residues together with their interactions with surrounding environment85. In order to assess the variations of secondary structures, a total of ten geometric characteristics86 were first calculated for each residue i (as clearly described in Fig. 1b), and a 10-dimentional vector was therefore generated (as discussed in Supplementary Method S1). Second, a pairwise cosine similarity (\({{{\rm{c}}}}_{{{\rm{i}}}}\)) between two studied protein A and B for residue i was computed using two vectors (\({{{{\rm{v}}}}^{ \rightharpoonup }}_{{{\rm{i}}}}^{{{\rm{A}}}}\) and \({{{{\rm{v}}}}^{ \rightharpoonup }}_{{{\rm{i}}}}^{{{\rm{B}}}}\)) from two proteins (as described in Fig. 1c and the equation below), which resulted in the value of \({{{\rm{c}}}}_{{{\rm{i}}}}\) being fixed within the range of [0,1].
Third, the average value (\({{{\rm{c}}}}_{{{\rm{ave}}}}\)) of cosine similarities for all amino acids in two aligned proteins (of N sequence length after alignment) gave the measurement of the overall secondary structure variations between those two proteins. The \({{{\rm{c}}}}_{{{\rm{ave}}}}\) were then calculated for all those two million structure pairs collected from PDB, and the distribution frequencies of all pairs according to \({{{\rm{c}}}}_{{{\rm{ave}}}}\) values were shown in Supplementary Fig. S31. As given, the majority of the \({{{\rm{c}}}}_{{{\rm{ave}}}}\) values were concentrated in the range between 0 and 0.1. To enhance the resolution of \({{{\rm{c}}}}_{{{\rm{ave}}}}\) value in differentiating protein structure changes, a convex function was finally used to transform \({{{\rm{c}}}}_{{{\rm{ave}}}}\) to a new VSSmeasur.
This transformation substantially stretched the distribution frequencies for all those two million structure pairs (as shown in Supplementary Fig. S32), but the value of this metric remained within the range from 0 to 1. Particularly, the closer the VSSmeasur is to 1, the more different the secondary structures of two studied proteins are. All in all, this transformation ensured a robust and interpretable metric for comparing protein secondary structures (shown in Fig. 1d).
Calculating SPSmeasur based on the relative distances among amino acids
To systematically assess the shift among protein segments, the relative distance among residues of studied proteins should be calculated. Till now, two established tools (dRMSD87 & LDDT27) had been available to fulfill this kind of calculation. The dRMSD was a metric that was popular in comparing the pairwise distance between the residues in two proteins, which was reported to be limited by the lack of a fixed range (from zero to infinity, making the measuring results very difficult to standardize)22. Moreover, the dRMSD usually overestimated the changes of relative distances between distant amino acids among proteins58. In contrast, LDDT attempted to ignore the changes of relative distance between distant residues by focusing only on the adjacent ones. However, such ignorance of distant residues made LDDT highly dependent on the user-defined distance cutoffs, the poor selection of which could either exaggerate or understate the structural difference20. Thus, it is extremely necessary to introduce metric that could overcome the limitations from both scoring tools (dRMSD and LDDT).
Here, a method calculating the relative distance among residues was thus proposed, which was given as follows. First, the difference of the relative distance (between residue i and j) between protein A and B was computed (as illustrated in Fig. 2b), which was further normalized with weight by eliminating the influence of structure size (the performances of our proposed method were offered in Supplementary Fig. S19) and inherent distance magnitude (the performance of this proposed method was described in Supplementary Fig. S23). Taking together, the normalized difference of relative distances (between residue i and j) between protein A and B could be represented using the following equation.
where \({{{\rm{d}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{A}}}}\) and \({{{\rm{d}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{B}}}}\) represented the pairwise distance between residue i and j in the protein A and protein B, respectively; \({{{\rm{d}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{A}}}}-{{{\rm{d}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{B}}}}\) described the difference of relative distances (between residue i and j); \({{{\rm{D}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{norm}}}}\) denoted the normalized difference of relative distances, which resulted in the value of \({{{\rm{D}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{norm}}}}\) being fixed within the range of [0,1].
Second, the average value (\({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\)) for all residue pairs in two aligned proteins provided a good measurement of the protein segment shift between proteins. The \({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\) was then assessed for all two million protein pairs collected from PDB, and the distribution frequencies of all structure pairs according to \({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\) were described in Supplementary Fig. S33. As provided, the majority of the \({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\) values were concentrated between 0 and 0.1. To enhance the resolution of \({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\) in differentiating protein structure changes, a convex function was finally used to transform \({{{\rm{D}}}}_{{{\rm{ave}}}}^{{{\rm{norm}}}}\) to a new SPSmeasur.
Such transformation substantially extended the distribution frequencies of all those two million structure pairs (as shown in Supplementary Fig. S34), but the value of this metric remained within the range from 0 to 1. Particularly, the closer the SPSmeasur is to 1, the more different the protein segments of two studied proteins are. In summary, this transformation ensured a robust and interpretable metric for measuring the shifts of protein segments (shown in Fig. 2c).
Searching effective MELO threshold indicating structure changes
In order to assess the posterior probability of structure changes for all studied protein pairs, two groups of protein pair data (of changed structures vs of similar structures) were collected based on the strategy identical to that applied in previous work24. Particularly, structure domains were comprehensively collected from both SCOP232 (2022-06-29) and CATH (v4.3.0)33 in the first place, and were filtered by two criteria (the sequence of the collected proteins should be contiguous & the length of the collected proteins should exceed 80 amino acids; as described in Supplementary Fig. S2a). Second, the equivalent structure level (‘family’ in SCOP2 and ‘topology’ in CATH) was used in this study to identify those two groups of protein pairs. As reported in previous work24, any two proteins from the same family/topology were considered as ‘similar’ in their structures, and two proteins from different families/topologies were regarded as ‘changed’ in structure (as provided in Supplementary Fig. S2b). Third, the distribution of the two groups of structure pairs was measured using VSSmeasur (Supplementary Fig. S2c) and SPSmeasur (Supplementary Fig. S2d). As shown, those two groups (of changed structures vs of similar structures) of pairs were largely differentiated by the value of 0.5 for both VSSmeasur and SPSmeasur. Finally, the posterior probabilities of the pairs with structure changed were assessed by the following equation.
Taking the metric VSSmeasur measuring the variations of secondary structures as an example:
where \({{\rm{P}}}({{{\rm{X}}}}_{{{\rm{changed}}}})\) denoted the prior probability of the protein pairs showing changes in their secondary structures; \({{\rm{P}}}({{{\rm{VSS}}}}_{{{\rm{measur}}}})\) indicated the predictor prior probability of the secondary structure change equaling to VSSmeasur; \({{\rm{P}}}\left({{{\rm{VSS}}}}_{{{\rm{measur}}}}|{{{\rm{X}}}}_{{{\rm{changed}}}}\right)\) provided the likelihood of the secondary structure change equaling to VSSmeasur given that the protein pairs showed changes in their secondary structure; \({{\rm{P}}}({{{\rm{X}}}}_{{{\rm{changed}}}}|{{{\rm{VSS}}}}_{{{\rm{measur}}}})\) was the posterior probability of the protein pairs showing changes in their secondary structure given that the secondary structure variations equaled to VSSmeasur. In the meantime, the posterior probability of the protein pairs with similar structures could be further calculated using the equation as follows.
Taking the metric SPSmeasur measuring the shifts of protein segments as an example:
where \({{\rm{P}}}({{{\rm{X}}}}_{{{\rm{similar}}}})\) denoted the prior probability of the protein pairs having no shift of protein segments; \({{\rm{P}}}({{{\rm{SPS}}}}_{{{\rm{measur}}}})\) indicated the predictor prior probability of the protein segment shift equaling to SPSmeasur; \({{\rm{P}}}({{{\rm{SPS}}}}_{{{\rm{measur}}}}|{{{\rm{X}}}}_{{{\rm{similar}}}})\) described the likelihood of the protein segment shift equaling to SPSmeasur given that the protein pairs showed no shift of their protein segment; \({{\rm{P}}}({{{\rm{X}}}}_{{{\rm{similar}}}}|{{{\rm{SPS}}}}_{{{\rm{measur}}}})\) was the posterior probability of the protein pairs having no shift in the protein segment given that the protein segment shift equaled to SPSmeasur. Furthermore, in order to determining the effective threshold that denoted structure changes, the posterior probabilities of VSSmeasur (Supplementary Fig. S2e) & SPSmeasur (Supplementary Fig. S2f) were then searched, and the posterior probabilities for the pairs of changed structure and similar structure were colored in YELLOW and BLUE, respectively. Clearly, the lines of two series of posterior probability for changed structures (YELLOW) and similar structures (BLUE) intersected at the value of 0.5 for both VSSmeasur and SPSmeasur. These findings helped us to determine VSSmeasur = 0.5 and SPSmeasur = 0.5 as two effective thresholds that could denote the variations of secondary structures and the shifts of protein segments, respectively.
Locating secondary structure variation and protein segment shift
The MELO realized the discovery & illustration of the structurally-changed regions base on the pairwise values calculated using Eq. 1 and Eq. 3, which resulted in two visualized plots (PLOTVSS and PLOTSPS) indicating the locations of secondary structure variations and the locations of protein segment shifts, respectively. Moreover, the way to assess and then generate these two plots were also explicitly described in Fig. 1c and Fig. 2c, respectively.
Generating PLOTVSS based on the geometric characteristics among residues
A total of ten geometric characteristics86 were first calculated for each residue in a protein, and the secondary structures of this protein were determined by the calculated characteristics together with the residues’ interactions with surrounding environments85 (details were provided in both Supplementary Method S2 and Supplementary Fig. S35). Then, the PLOTVSS was generated according to the process described in Fig. 1c. As shown, the top and bottom rows displayed the sequence SA and SB of the protein A and B, and mutated residues were highlighted in RED font. The secondary structure symbols showing in the second and the penult rows were adopted directly from Supplementary Fig. S35, and the middle row provided the \({{{\rm{c}}}}_{{{\rm{i}}}}\) values calculated using Eq. 1 for the residue i. The darker the residue’s color was, the higher its \({{{\rm{c}}}}_{{{\rm{i}}}}\) value was, which reflected greater variation in its corresponding secondary structure.
Generating PLOTSPS based on the relative distances among amino acids
The PLOTSPS was generated according to the process shown in Fig. 2c. First, the differences of the relative distances (between residue i and j) between protein A and B were calculated, and further normalized using Eq. 3 for getting rid of the impact of structure size and inherent distance magnitude. As shown in Fig. 2c, the PLOTSPS was then depicted using vertical axis corresponding to sequence SA of protein A and horizontal axis corresponding to sequence SB of protein B, the color of each grid offered the pairwise \({{{\rm{D}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{norm}}}}\) values calculated using Eq. 3 for the residue pair of i and j. The darker the color of residue pair was, the higher the \({{{\rm{D}}}}_{{{\rm{i}}},{{\rm{j}}}}^{{{\rm{norm}}}}\) value was, which reflected greater shift in its corresponding protein segments. For example, the M1 was highlighted in dark RED color in Fig. 2c, which denoted that the Domain 2 (BLUE) corresponding to the residues 116-118 of protein A was significantly shifted from the Domain 1 (PINK) corresponding to the residues 37-39 of protein B. In the meantime, the color of M3 was very light, which described that no obvious shift was identified between the Domain 2 (BLUE) corresponding to the residues 114-116 of protein B and the Domain 2 (BLUE) corresponding to the residues 144-146 of protein A. In sum, both PLOTVSS and PLOTSPS generated by our MELO highlighted its ability to locating secondary structure variation and protein segment shift.
Calculating the dependence of methods on protein sequence length
To evaluate the consistencies of MELO and those available scoring methods (TM-score, LDDT and RMSD) across the proteins of different sequence lengths, all the two million structure pairs from PDB were first classified into 50 windows based on their aligned sequence lengths (window size = 20 residues), which resulted in (0, 20], (20, 40], …, and (980, 1,000]. Second, the structure variations of all pairs in each window (\({{{\rm{W}}}}_{{{\rm{i}}}}\)) were calculated, and a total of 50 mean scores (\({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}\), equaling to the average value of all calculated variations in a window) were generated. Taking RMSD as an example, its \({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}\) rose extensively with the increase of protein length (offered in Supplementary Fig. S19a), which was highly different from the trends of the remaining three methods, denoting that the RMSD was more sensitive to sequence length than the other three methods. Third, centered mean score (\({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}^{{{\rm{centered}}}}\)) was further calculated for each window (\({{{\rm{W}}}}_{{{\rm{i}}}}\)) using the following equation.
The \({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}^{{{\rm{centered}}}}\) helped to center the variations measured by different methods to the value of zero, which realized the objective evaluations of the dependence of methods on the length of sequence. In other words, by standardizing \({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}\), the \({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}^{{{\rm{centered}}}}\) were converted to the value of zero and scaled by their deviation, thus allowing for direct and consistent comparison. As shown in Supplementary Fig. S19b and Supplementary Fig. S19c, the \({{{\rm{Mean}}}}_{{{{\rm{W}}}}_{{{\rm{i}}}}}^{{{\rm{centered}}}}\) revealed that our MELO gave a more consistent measurement compared with other methods.
Evaluating the ability of methods on locating protein structure change
Two benchmark subsets (a local motion subset and a domain motion subset) were derived from PSCDB. First, for each protein pair, the “moving segments” indicated by PSCDB constituted the ground-truth set of changed residue pairs (true positives, TP, label 1), and all other aligned residues were treated as true negatives (TN, label 0). Residues that were unaligned, lack coordinates, or have ambiguous identifiers were removed prior to evaluation. Second, to obtain residue-wise scores of the MELO and LDDT, residue-wise SPSmeasur generated by MELO and residue-wise LDDT obtained from SWISS-MODEL were denoted \({{{\rm{s}}}}_{{{\rm{r}}}}^{{{\rm{MELO}}}}\) and \({{{\rm{s}}}}_{{{\rm{r}}}}^{{{\rm{LDDT}}}}\)。
Third, to prevent threshold-selection bias in the comparison, we applied a widely adopted approach, defined as the maximum F1-score across all thresholds (Fmax)47,48,49. Specifically, for the residue-wise SPSmeasur, a residue in a protein pair was labeled as “structurally changed” if its score \({{{\rm{s}}}}_{{{\rm{r}}}}^{{{\rm{MELO}}}}\) meets or exceeds the threshold \({{{\rm{\tau }}}}_{{{\rm{MELO}}}}\). The set of such residues was denoted by \({{{\rm{P}}}}_{{{\rm{MELO}}}}\left({{{\rm{\tau }}}}_{{{\rm{MELO}}}}\right)\):
Similarly, for the residue-wise LDDT, a residue in a protein pair was labeled as “structurally changed” if its score \({{{\rm{s}}}}_{{{\rm{r}}}}^{{{\rm{LDDT}}}}\) meets or exceeds the threshold \({{{\rm{\tau }}}}_{{{\rm{LDDT}}}}\). The set of such residues was \({{{\rm{P}}}}_{{{\rm{LDDT}}}}\left({{{\rm{\tau }}}}_{{{\rm{LDDT}}}}\right)\):
After obtaining the identified “changed-residue” sets \({{{\rm{P}}}}_{{{\rm{MELO}}}}\left({{{\rm{\tau }}}}_{{{\rm{MELO}}}}\right)\) and \({{{\rm{P}}}}_{{{\rm{LDDT}}}}\left({{{\rm{\tau }}}}_{{{\rm{LDDT}}}}\right)\), we used the PSCDB-indicated set of changed residues TP as the ground-truth positives. And the per-method location performance was summarized by the F1 score:
Within the threshold interval [0, 1.0] with a step size of 0.01, the threshold \({{{\rm{\tau }}}}_{{{\rm{m}}}}\) was exhaustively searched for each method to maximize the macro-average of the F1 across all protein pairs (F1 was first calculated for each pair, then averaged). The thresholds \({{{\rm{\tau }}}}_{{{\rm{MELO}}}}\) and \({{{\rm{\tau }}}}_{{{\rm{LDDT}}}}\) yielding the highest pair-wise F1 was fixed as the global threshold for residue-wise SPSmeasur and residue-wise LDDT. Finally, the relative advantage between residue-wise SPSmeasur and residue-wise LDDT was quantified as:
A positive ΔF1 indicated that SPSmeasur achieves superior performance in locating protein structure changes compared with LDDT-server, with larger values reflecting greater advantages. Conversely, a negative ΔF1 suggested inferior performance, with smaller values reflecting more pronounced disadvantages.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are permanently archived to the online repository (https://idrblab.org/melo/data) under CC BY 4.0 license. User can deploy MELO on multiple optimized platforms: an interactive web-service (https://idrblab.org/melo/), a local software tool (https://idrblab.org/melo/MELO.rar), a PyPI pip-installable tool (https://github.com/idrblab/MELO/) and (https://doi.org/10.5281/zenodo.17360644) for command-line integrations. Benchmark analyses demonstrated MELO scalability: MELO processes two million protein pairs in 109 hours, outperforming the TM-score (199 hours) & LDDT (1,044 hours); MELO offers the functions of discovering secondary structure variations and protein segment shifts, which were absent in available tools. For ultra-long sequences ( > 8,000 residues), MELO achieved rapid execution: 30 seconds (PyPI workstation), 90 seconds (local GUI) and 500 seconds (web-server). We show experimental structures from the PDB under accession numbers 1A57, 1BLR, 1ICN, 1KH0, 1PGB, 1V4T, 2LYE, 2VGL, 2XA7, 3QIC, 4DCH, 4EPV, 6A6M, 6A6N, 6BYY, 6BZ1, 6HKR, 6HZM, 6KWJ, 6KWU, 6M9W, 6VXH, 7CWO, 7OXW and 7XIZ. Source data are provided with this paper.
References
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Kuhlman, B. & Bradley, P. Advances in protein structure prediction and design. Nat. Rev. Mol. Cell Biol. 20, 681–697 (2019).
Buel, G. & Walters, K. Can AlphaFold2 predict the impact of missense mutations on structure?. Nat. Struct. Mol. Biol. 29, 1–2 (2022).
Levental, I. & Lyman, E. Regulation of membrane protein structure and function by their lipid nano-environment. Nat. Rev. Mol. Cell Biol. 24, 107–122 (2023).
Yang, M. & Song, W. Diverse protein assembly driven by metal and chelating amino acids with selectivity and tunability. Nat. Commun. 10, 5545 (2019).
Stenström, O., Diehl, C., Modig, K. & Akke, M. Ligand-induced protein transition state stabilization switches the binding pathway from conformational selection to induced fit. Proc. Natl. Acad. Sci. USA 121, e2317747121 (2024).
Chang, Y. G. et al. Circadian rhythms, a protein fold switch joins the circadian oscillator to clock output in cyanobacteria. Science 349, 324–328 (2015).
Streit, J. O. et al. The ribosome lowers the entropic penalty of protein folding. Nature 633, 232–239 (2024).
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease. Annu Rev. Biochem 75, 333–366 (2006).
Chao, Y. C., Merritt, M., Schaefferkoetter, D. & Evans, T. G. High-throughput quantification of protein structural change reveals potential mechanisms of temperature adaptation in Mytilus mussels. BMC Evol. Biol. 20, 28 (2020).
Tsuboyama, K. et al. Mega-scale experimental analysis of protein folding stability in biology and design. Nature 620, 434–444 (2023).
Ho, S. S., Urban, A. E. & Mills, R. E. Structural variation in the sequencing era. Nat. Rev. Genet 21, 171–189 (2020).
Wang, J. et al. Mapping allosteric communications within individual proteins. Nat. Commun. 11, 3862 (2020).
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Mifsud, J. C. O. et al. Mapping glycoprotein structure reveals Flaviviridae evolutionary history. Nature 633, 695–703 (2024).
Kondra, S., Sarkar, T., Raghavan, V. & Xu, W. Development of a TSR-based method for protein 3-D structural comparison with its applications to protein classification and motif discovery. Front Chem. 8, 602291 (2020).
Lema, M. & Echave, J. Assessing local structural perturbations in proteins. BMC Bioinforma. 6, 226 (2005).
Margelevicius, M. GTalign: spatial index-driven protein structure alignment, superposition, and search. Nat. Commun. 15, 7305 (2024).
Schneider, T. R. Objective comparison of protein structures: error-scaled difference distance matrices. Acta Crystallogr D. Biol. Crystallogr 56, 714–721 (2000).
Olechnovič, K., Monastyrskyy, B., Kryshtafovych, A. & Venclovas, Č. Comparative analysis of methods for evaluation of protein models against native structures. Bioinformatics 35, 937–944 (2019).
Kabsch, W. A solution for the best rotation to relate two sets of vectors. Acta Crystallogr D. Struct. Biol. 32, 922–923 (1976).
Zhang, Y. & Skolnick, J. Scoring function for automated assessment of protein structure template quality. Proteins 57, 702–710 (2004).
Hamamsy, T. et al. Protein remote homology detection and structural alignment using deep learning. Nat. Biotechnol. 42, 975–985 (2024).
Xu, J. & Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5?. Bioinformatics 26, 889–895 (2010).
Zhang, C., Shine, M., Pyle, A. & Zhang, Y. US-align: universal structure alignments of proteins, nucleic acids, and macromolecular complexes. Nat. Methods 19, 1109–1115 (2022).
Ives, C. M. et al. Restoring protein glycosylation with GlycoShape. Nat. Methods 21, 2117–2127 (2024).
Mariani, V., Biasini, M., Barbato, A. & Schwede, T. lDDT: a local superposition-free score for comparing protein structures and models using distance difference tests. Bioinformatics 29, 2722–2728 (2013).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Gerasimavicius, L., Livesey, B. J. & Marsh, J. A. Loss-of-function, gain-of-function and dominant-negative mutations have profoundly different effects on protein structure. Nat. Commun. 13, 3895 (2022).
Ayaz, P. et al. Structural mechanism of a drug-binding process involving a large conformational change of the protein target. Nat. Commun. 14, 1885 (2023).
Bongirwar, V. & Mokhade, A. S. Different methods, techniques and their limitations in protein structure prediction: a review. Prog. Biophys. Mol. Biol. 173, 72–82 (2022).
Andreeva, A., Howorth, D., Chothia, C., Kulesha, E. & Murzin, A. G. SCOP2 prototype: a new approach to protein structure mining. Nucleic Acids Res 42, D310–D314 (2014).
Sillitoe, I. et al. CATH: increased structural coverage of functional space. Nucleic Acids Res 49, D266–D273 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Burley, S. K. et al. RCSB Protein Data Bank (RCSB.org): delivery of experimentally-determined PDB structures alongside one million computed structure models of proteins from artificial intelligence/machine learning. Nucleic Acids Res 51, D488–D508 (2023).
Kiya, M., Shiga, S., Ding, P., Koide, S. & Makabe, K. β-Strand-mediated domain-swapping in the absence of hydrophobic core repacking. J. Mol. Biol. 436, 168405 (2024).
Steele, R. A. et al. The three-dimensional structure of a helix-less variant of intestinal fatty acid-binding protein. Protein Sci. 7, 1332–1339 (1998).
Cao, Y. et al. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. Nature 608, 593–602 (2022).
Bryant, P., Kelkar, A., Guljas, A., Clementi, C. & Noe, F. Structure prediction of protein-ligand complexes from sequence information with Umol. Nat. Commun. 15, 4536 (2024).
Chen, J. N., Jiang, F. & Wu, Y. D. Accurate prediction for protein-peptide binding based on high-temperature molecular dynamics simulations. J. Chem. Theory Comput 18, 6386–6395 (2022).
Wissel, J., Muller, J., Ebersbach, G. & Poewe, W. Trick maneuvers in cervical dystonia: investigation of movement- and touch-related changes in polymyographic activity. Mov. Disord. 14, 994–999 (1999).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Robin, X. et al. Continuous Automated Model EvaluatiOn (CAMEO)-Perspectives on the future of fully automated evaluation of structure prediction methods. Proteins 89, 1977–1986 (2021).
Studer, G., Biasini, M. & Schwede, T. Assessing the local structural quality of transmembrane protein models using statistical potentials (QMEANBrane). Bioinformatics 30, 505–511 (2014).
Waterhouse, A. M. et al. The structure assessment web server: for proteins, complexes and more. Nucleic Acids Res 52, W318–W323 (2024).
Amemiya, T., Koike, R., Kidera, A. & Ota, M. PSCDB: a database for protein structural change upon ligand binding. Nucleic Acids Res 40, D554–D558 (2012).
Necci, M., Piovesan, D., Predictors, C., DisProt, C. & Tosatto, S. C. E. Critical assessment of protein intrinsic disorder prediction. Nat. Methods 18, 472–481 (2021).
Turner, N. L. et al. Reconstruction of neocortex: organelles, compartments, cells, circuits, and activity. Cell 185, 1082–1100 e1024 (2022).
Yang, Y., Zhang, H., Gichoya, J. W., Katabi, D. & Ghassemi, M. The limits of fair medical imaging AI in real-world generalization. Nat. Med 30, 2838–2848 (2024).
Hofmann, S. et al. Conformation space of a heterodimeric ABC exporter under turnover conditions. Nature 571, 580–583 (2019).
McWhite, C. D., Armour-Garb, I. & Singh, M. Leveraging protein language models for accurate multiple sequence alignments. Genome Res 33, 1145–1153 (2023).
Dharmaiah, S. et al. Structures of N-terminally processed KRAS provide insight into the role of N-acetylation. Sci. Rep. 9, 10512 (2019).
Kosloff, M. & Kolodny, R. Sequence-similar, structure-dissimilar protein pairs in the PDB. Proteins 71, 891–902 (2008).
Brenner, S. E., Chothia, C. & Hubbard, T. J. Assessing sequence comparison methods with reliable structurally identified distant evolutionary relationships. Proc. Natl. Acad. Sci. USA 95, 6073–6078 (1998).
Cheng, Q., Joung, I., Lee, J., Kuwajima, K. & Lee, J. Exploring the folding mechanism of small proteins GB1 and LB1. J. Chem. Theory Comput 15, 3432–3449 (2019).
Garcia de la Torre, J. & Hernandez Cifre, J. G. Hydrodynamic properties of biomacromolecules and macromolecular complexes: concepts and methods. J. Mol. Biol. 432, 2930–2948 (2020).
Salvi, N., Abyzov, A. & Blackledge, M. Solvent-dependent segmental dynamics in intrinsically disordered proteins. Sci. Adv. 5, eaax2348 (2019).
Neveu, E. et al. RapidRMSD: rapid determination of RMSDs corresponding to motions of flexible molecules. Bioinformatics 34, 2757–2765 (2018).
Conibear, A. C., Rosengren, K. J., Harvey, P. J. & Craik, D. J. Structural characterization of the cyclic cystine ladder motif of theta-defensins. Biochemistry 51, 9718–9726 (2012).
Shor, B. & Schneidman-Duhovny, D. CombFold: predicting structures of large protein assemblies using a combinatorial assembly algorithm and AlphaFold2. Nat. Methods 21, 477–487 (2024).
Robey, R. W. et al. Revisiting the role of ABC transporters in multidrug-resistant cancer. Nat. Rev. Cancer 18, 452–464 (2018).
Nielsen, J. et al. Structure of the human dopamine transporter in complex with cocaine. Nature 632, 678–685 (2024).
Yu, Q. et al. Structures of ABCG2 under turnover conditions reveal a key step in the drug transport mechanism. Nat. Commun. 12, 4376 (2021).
Feng, X. et al. CRABP2 regulates invasion and metastasis of breast cancer through hippo pathway dependent on ER status. J. Exp. Clin. Cancer Res 38, 361 (2019).
Tomlinson, C. W. E., Cornish, K. A. S., Whiting, A. & Pohl, E. Structure-functional relationship of cellular retinoic acid-binding proteins I and II interacting with natural and synthetic ligands. Acta Crystallogr D. Struct. Biol. 77, 164–175 (2021).
Wang, L., Li, Y., Abildgaard, F., Markley, J. L. & Yan, H. NMR solution structure of type II human cellular retinoic acid binding protein: implications for ligand binding. Biochemistry 37, 12727–11236 (1998).
Pastok, M. W. et al. Structural requirements for the specific binding of CRABP2 to cyclin D3. Structure 32, 1–15 (2024).
Zhang, Y. et al. Podocyte apoptosis in diabetic nephropathy by BASP1 activation of the p53 pathway via WT1. Acta Physiol. 232, e13634 (2021).
Poulin, E. J. et al. Tissue-specific oncogenic activity of KRAS(A146T). Cancer Discov. 9, 738–755 (2019).
Matschinsky, F. M. et al. Glucokinase activators for diabetes therapy: 2010 status report. Diab. Care 34, S236–S243 (2011).
Toulis, K. A., Nirantharakumar, K., Pourzitaki, C., Barnett, A. H. & Tahrani, A. A. glucokinase activators for type 2 diabetes: challenges and future developments. Drugs 80, 467–475 (2020).
Zak, K. M. et al. Crystal structure of Kluyveromyces lactis glucokinase (KlGlk1). Int J. Mol. Sci. 20, 4821 (2019).
Kamata, K., Mitsuya, M., Nishimura, T., Eiki, J. & Nagata, Y. Structural basis for allosteric regulation of the monomeric allosteric enzyme human glucokinase. Structure 12, 429–438 (2004).
Kelly, B. T. et al. A structural explanation for the binding of endocytic dileucine motifs by the AP2 complex. Nature 456, 976–979 (2008).
Jackson, L. P. et al. A large-scale conformational change couples membrane recruitment to cargo binding in the AP2 clathrin adaptor complex. Cell 141, 1220–1229 (2010).
Sargsyan, K., Grauffel, C. & Lim, C. How molecular size impacts RMSD applications in molecular dynamics simulations. J. Chem. Theory Comput 13, 1518–1524 (2017).
Cappelletti, V. et al. Dynamic 3D proteomes reveal protein functional alterations at high resolution in situ. Cell 184, 545–559 (2021).
Brescia, P. et al. MEF2B instructs germinal center development and acts as an oncogene in B cell lymphomagenesis. Cancer Cell 34, 453–465 (2018).
Lei, X. et al. The cancer mutation D83V induces an α-Helix to β-strand conformation switch in MEF2B. J. Mol. Biol. 430, 1157–1172 (2018).
Blum, M. et al. InterPro: the protein sequence classification resource in 2025. Nucleic Acids Res 53, D444–D456 (2025).
Cantarel, B. L., Morrison, H. G. & Pearson, W. Exploring the relationship between sequence similarity and accurate phylogenetic trees. Mol. Biol. Evol. 23, 2090–2100 (2006).
Cock, P. J. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Eastman, P. et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLoS Comput Biol. 13, e1005659 (2017).
Newberry, R. W. & Raines, R. T. Secondary forces in protein folding. ACS Chem. Biol. 14, 1677–1686 (2019).
Al Mughram, M. H., Herrington, N. B., Catalano, C. & Kellogg, G. E. Systematized analysis of secondary structure dependence of key structural features of residues in soluble and membrane-bound proteins. J. Struct. Biol. X 5, 100055 (2021).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Chen, H., Huang, Y. & Xiao, Y. A simple method of identifying symmetric substructures of proteins. Comput Biol. Chem. 33, 100–107 (2009).
Acknowledgements
The Natural Science Foundation of Zhejiang (RG25H300001); The National Key R&D Program of China (2024YFA1307503); The National Natural Science Foundation of China (22220102001, 82373790, U1909208 and 81872798); The Double Top-Class Universities (181201*194232101); The Fundamental Research Funds for Central University (2018QNA7023); Westlake Laboratory of Life Science & Biomedicine; Information Technology Center of Zhejiang University.
Author information
Authors and Affiliations
Contributions
F.Z. conceived the idea and designed the entire research; L.Y.Z. proposed MELO’s algorithms; Y.L. and Y.T.Z. built the website; M.X.L., L.Y.Z., Y.L., T.T.F., S.Y.S., C.B.G., H.C.S., X.N.S, M.K.L., M.J.M., and H.B.D. collected relevant datasets and provided biological supports; F.Z. wrote the manuscript. All authors reviewed and approved the latest version of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Richard Baker, Jiangning Song, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, L., Liao, Y., Zhang, Y. et al. Measuring and locating the changes in protein structure using MELO. Nat Commun 17, 1360 (2026). https://doi.org/10.1038/s41467-025-68110-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68110-8