Abstract
Fragile X-associated tremor/ataxia syndrome (FXTAS) is a neurodegenerative disorder caused by CGG repeat expansions in the FMR1 gene. While CGG repeat toxicity is established, the precise molecular mechanisms driving neurodegeneration remain unclear. Here, we show that a multi-omics strategy combined with TWAS reveals brain-region-specific molecular signatures and striking gene dysregulation in inhibitory neurons. Using conditional mouse models, we demonstrate that selective expression of expanded CGG repeats in GABAergic neurons is sufficient to recapitulate key pathologic hallmarks of FXTAS. We identify PRKCG as a genetic modifier of FXTAS, with cross-species evidence linking its overexpression to disease onset. Many dysregulated mRNAs in GABAergic neurons are targets of hnRNPA2/B1, an RNA-binding protein sequestered by CGG repeat RNA. Functional screening in Drosophila further establishes PRKCG as a potent modulator of CGG-associated neurotoxicity. These findings uncover a critical role of GABAergic neurons in FXTAS pathogenesis and position PRKCG as a promising therapeutic target.
Similar content being viewed by others
Introduction
The aberrant expansion of CGG repeats in the 5′ untranslated region (UTR) of the Fragile X messenger ribonucleoprotein 1 (FMR1) gene is implicated in Fragile X-associated disorders. Alleles harboring 55–200 repeats, falling within the premutation (PM) range, lead to the onset of Fragile X-related tremor/ataxia syndrome (FXTAS), a neurodegenerative disorder characterized by intention tremor, cerebellar gait ataxia, progressive cognitive decline, and Parkinsonism1. However, FXTAS shows incomplete penetrance, affecting only ~40% of male and 16% of female premutation carriers in late adulthood2, highlighting the need to better understand the molecular and cellular factors driving the onset of disease.
The “RNA toxicity” and “repeat-associated non-AUG (RAN) translation” are two major models proposed to explain FXTAS pathogenesis3,4,5,6,7. FXTAS carriers exhibited increased mRNA levels of FMR1 but displayed either no detectable or a slight reduction in FMRP protein levels, which is proportional to the length of the expanded CGG repeats8,9. In the RNA toxicity model, the upregulation of expanded CGG-repeat FMR1 mRNA triggers the formation of nuclear RNA aggregates, which can sequester various RNA-binding proteins, such as Pura3, heterogeneous nuclear ribonucleoprotein A2B1 (hnRNPA2B1)10, src-associated substrate during mitosis of 68 kDa (SAM68)11, DiGeorge syndrome critical region gene 8 (DGCR8)12, and TAR DNA binding protein of 43 kDa (TDP-43)13. This sequestration is believed to impede their normal biological function. In the RAN translation model, the FMR1 mRNA containing expanded CGG repeats initiates translation outside the traditional AUG start codon5. This leads to the generation of a polyglycine-containing protein, FMRpolyG, along with additional repeat-associated RAN translation products. FMRpolyG can be detected in brain tissues of FXTAS patients and mouse models and has been shown to be toxic in flies and cultured neurons, implicating its role in FXTAS pathogenesis5,7,14,15. Despite these insights, it remains unclear whether FXTAS is driven primarily by toxic RNA gain-of-function, RAN translation, or a combination of both mechanisms.
Animal models of FXTAS offer a valuable platform for advanced examination of how the CGG premutation induces alterations in gene activity across various tissues. A recent study suggested that expression of both FMRpolyG and expanded CGG repeat RNA (FMRpolyG+RNA) in the brain is pathogenic, recapitulating key FXTAS phenotypes, such as intranuclear inclusion formation and locomotor dysfunction7. As one of the key neuropathological hallmarks of FXTAS, intranuclear inclusions are found in neurons and astroglia throughout the brain that are immunoreactive for ubiquitin16,17,18,19, raising the question of how individual cell types are affected by the expression of expanded CGG repeats. However, previous studies provided only a limited understanding of the cell-type specificity associated with FXTAS pathogenesis. It remains unclear whether certain cell types have unique transcriptional signatures upon the expression of expanded CGG repeats. Additionally, it remains to be determined whether specific cell types are more vulnerable to the toxic effects of CGG repeats.
Our previous work using a robust Drosophila screening platform has uncovered genetic factors modulating CGG-associated toxicity, including the identification of PSMB5 as a suppressor of CGG-associated neurodegeneration in FXTAS3,20,21. Given the incomplete penetrance of FXTAS among fragile X premutation carriers, there remains a critical need to identify additional modifiers that could serve as genetic biomarkers and therapeutic targets. One promising strategy is transcriptome-wide association study (TWAS), which integrates eQTL data from relevant tissues/cell types with GWAS findings to pinpoint genes that drive disease risk through altered gene regulation. In several neurodegenerative diseases, such as Alzheimer’s disease (AD)22,23,24, Parkinson’s disease (PD)25,26,27, and amyotrophic lateral sclerosis (ALS)28,29,30,31, TWAS has provided insights into disease pathogenesis by uncovering the downstream effects of genetic risk loci. This approach holds significant promise for advancing our understanding of FXTAS and for prioritizing candidate genes for further study.
In this study, we employed a comprehensive multi-omics strategy in a FXTAS mouse model to map the molecular landscape in the brain. Our analyses revealed striking brain–region–specific signatures and profound gene dysregulation in inhibitory neurons induced by FMRpolyG and CGG-repeat RNA expression. By selectively expressing expanded CGG repeats in distinct neuronal populations using conditional mouse models, we demonstrated that the expression of expanded CGG repeats in GABAergic neurons alone is sufficient to recapitulate key FXTAS hallmarks while having minimal impact on Glutamatergic neurons. TRAP-seq further identified gene expression alterations specific to GABAergic neurons, many of which are bound by hnRNPA2B1, an RNA-binding protein sequestered by CGG repeat RNA. Moreover, a TWAS on premutation carriers identified 66 genes significantly linked to onset of FXTAS, with nine genes modulating CGG repeat–associated toxicity in an FXTAS fly model. Notably, cross-comparison of our datasets from various model systems revealed that upregulation of PRKCG, a known risk gene for SCA14, is associated with the onset of FXTAS. Finally, we showed that overexpression of human PRKCG and its fly ortholog, aPKC, induces neuronal toxicity in the FXTAS fly model. Together, our integrated analyses highlight the critical role of GABAergic neurons in driving FXTAS pathogenesis and identify PRKCG as a promising therapeutic target.
Results
Distinct transcriptional dysregulation in different brain regions of the FXTAS mouse model
Transgenic mice expressing both expanded CGG repeats and FMRpolyG in the brain exhibit FXTAS-like motor symptoms7, implicating the brain as the primary site of pathology. In this mouse model, the human FMR1 5′-UTR harboring 99 CGG repeats is inserted into the Rosa26 locus under a CAG promoter, with transcription silenced by a loxP-flanked polyadenylation cassette. Crossing these mice with Nestin-Cre transgenic animals, which express Cre recombinase in neuronal and glial progenitors, removes the cassette and activates expression of both CGG-repeat RNA and FMRpolyG protein throughout the brain (Fig. 1a). To elucidate the molecular mechanisms underlying FXTAS, we analyzed transcriptomic profiles from the cortex, cerebellum, and hippocampus at the onset of phenotype (3–4 m) and late progression stage(6–7 m) stages of FXTAS and control mice. Based on the number of differentially expressed genes (DEGs) detected across disease progression, the cerebellum was identified as the most affected brain region, with the cortex showing an intermediate number of DEGs relative to the cerebellum and the hippocampus (Supplementary Data 1). The hippocampus displayed the fewest DEGs. At the onset stage (3–4 months; Fig. 1b), we observed subtle transcriptomic changes in the bulk brain tissue of FXTAS mice compared to the control animals (nDEGs with P < 0.01, cortex n = 127; cerebellum n = 162; hippocampus n = 94; Supplementary Fig. 1a, b). Pairwise Fisher’s exact tests revealed significant differences in the numbers of DEGs among brain regions (all P < 0.05). Minimal overlap among brain regions and distinct KEGG pathway enrichments indicate strong region-specific signatures (Fig. 1c). Notably, some of these pathways have been implicated in FXTAS or ataxia32,33,34,35,36, encompassing the observation of Ras and MAPK signaling cascade in the cortex and the enrichment of mitophagy and Parkinsonism disease in the cerebellum. Hist1h1e, an established intellectual disability (ID)-associated gene37, was the only shared DEG identified across all three regions.
a Mouse breeding and overview of the experimental workflow. Male mice were used for tissues dissection and bulk RNA-Seq. b The numbers of up-regulated and down-regulated genes in each brain region at the onset of phenotype stage. Brain tissues were collected from 4 FXTAS mice and 4 littermate controls, except for cortex at 6–7 months, where 3 FXTAS mice were analyzed. Venn diagrams show a cross comparison of differentially expressed genes (DEGs) between the cerebellum, hippocampus, and cortex. Two-sided Fisher’s exact tests revealed significant differences in the numbers of DEGs across brain regions (cerebellum vs. cortex, P = 0.0353; cerebellum vs. hippocampus, P = 4.5 × 10⁻⁵; cortex vs. hippocampus, P = 0.0331). c Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses for DEGs in each brain region by using two-sided Fisher’s exact test with Benjamini–Hochberg (BH) correction for multiple comparisons. d The numbers of up-regulated and down-regulated genes in each region at the late disease progression stage. e Cross comparison of up-regulated and down-regulated DEGs between the cerebellum, hippocampus, and cortex are shown in the Venn diagrams. Fisher’s exact tests confirmed that overlaps of both up- and downregulated DEGs were significant (Supplementary Data 6; all P < 1e−19), with downregulated overlaps showing higher odds ratios and smaller P values. f, g Dot plot of gene ontology (GO) biological processes enriched in up-regulated and down-regulated DEGs by brain region (P < 0.05 and gene counts >10).
At the late progression stage (6–7 months; Fig. 1d), we found profound gene dysregulation, aligning with an increased statistical power (nDEGs with FDR < 0.05, cortex n = 671; cerebellum n = 1142; hippocampus n = 205; Supplementary Fig. 1c) (Supplementary Data 1). Cross-region comparisons (Fig. 1e) showed that downregulated genes exhibited greater overlap than upregulated genes, likely reflecting conserved pathways involved in neuronal dysfunction or cell loss across regions38. Fisher’s exact tests confirmed that overlaps of both up- and downregulated DEGs were significant (all P < 1e − 19), with downregulated overlaps showing higher odds ratios and smaller P values. Heatmaps of DEGs clearly separated brain regions by their expression profiles (Supplementary Fig. 1d). Gene Ontology (GO) analysis illuminated both shared molecular pathways and brain region-specific signaling cascades altered in the cortex and cerebellum, particularly for those upregulated DEGs (Fig. 1f). In both brain regions, we found significant upregulation of genes involved in the chromatin organization and ubiquitin-dependent protein catabolic processes, suggesting that expression of expanded CGG repeats significantly impacts chromatin structure and protein stability in the brain. Moreover, the cerebellum exhibited notable upregulation of genes involved in mRNA processing and DNA damage response, suggesting an increased effort to maintain genomic integrity and eliminate abnormal mRNA molecules. In the cortex, some up-regulated genes are specifically associated with the protein folding process, revealing an increased emphasis on preventing the accumulation of misfolded or aggregated molecules, which could be toxic to the brain. KEGG analysis further highlighted brain-region-specific molecular cascades altered by the expression of expanded CGG repeats (Supplementary Fig. 1e, f). In comparison, downregulated DEGs displayed more convergent functional profiles across cortex and cerebellum (Fig. 1g), with consistent overrepresentation of lipid metabolism, ion transport, and regulating locomotor behavior. This observation could provide molecular insight into systemic features of the FXTAS mice, including motor dysfunction and obesity7.
Single-nuclei RNA-seq for the prefrontal cortex of FXTAS and control mice
The brain tissue’s profound cellular heterogeneity makes it challenging to study molecular alterations in particular cell types that contribute to disease pathogenesis. In FXTAS, cell type-specific gene expression changes are poorly understood, particularly at time points relevant to the onset of disease-related symptoms. To gain deeper insights into cellular perturbations associated with the pathogenesis of FXTAS, we applied snRNA-seq to the prefrontal cortex (PFC) of FXTAS animals and littermate controls at the onset of the phenotype stage (3–4 m) (Fig. 2a). After quality control, a total of 13,902 high-quality nuclei were analyzed, comprising 7025 nuclei from FXTAS animals and 6877 from controls. Unsupervised clustering identified 13 transcriptionally distinct clusters with consistent expression patterns across individuals (Fig. 2b–d). Based on the expression profile of well-established marker genes of mouse brain39,40, we annotated these clusters to seven major cell types, including excitatory neurons (Exc, 5329 nuclei, 38.3% of total), inhibitory neurons (Inh, 3365, 24.2%), astrocytes (Ast, 1382, 9.5%), oligodendrocytes (Olig, 1834, 13.2%), oligodendrocyte precursor cells (OPC, 512, 3.7%), microglia (Mic, 1004, 7.2%), and endothelial cells (End, 530, 3.8%).
a Schematic pipeline of snRNA-seq on PFC of 2 male FXTAS mice and 2 male littermate controls. b Uniform manifold approximation and projection (UMAP) embeddings of 7025 FXTAS and 6877 control single-nuclei transcriptomes. Cell clusters were color-coded and annotated post hoc based on their transcriptional profile identities (see details in “Methods”). c Bar plot showing the total number of detected nuclei per cell type. d Heatmap showing the expression patterns of well-established marker genes per cell type. e Bar plots showing the number and proportion of detected nuclei associated with each cell type in FXTAS and control mice (two-sided unpaired t-test, *P = 0.04). f Bar plot showing the proportion of detected nuclei associated with each cell cluster in FXTAS and control mice (two-sided unpaired t-test, *P = 0.002).
Uniform manifold approximation and projection (UMAP) visualization showed broadly similar clustering patterns between FXTAS and control samples (Supplementary Fig. 2b). We next compared the abundance of the annotated cell types between FXTAS and controls. Most cell types were present at comparable proportions; however, inhibitory neurons were significantly increased in FXTAS (1834 nuclei, 26%) compared with controls (1531 nuclei, 22%; Welch’s t-test, P < 0.05) (Fig. 2e). This result was independently validated by immunofluorescent staining with an anti-GABA antibody, which confirmed an increase in GABA⁺ neuronal density in FXTAS PFC relative to controls (Welch’s t-test, P < 0.05; Supplementary Fig. 3). In addition, a specific inhibitory subcluster (Inh_2) contained a higher proportion of nuclei in the FXTAS group compared to controls (Welch’s t-test, P < 0.05) (Fig. 2f). It is important to note that due to the sensitivity of different cell types to tissue dissociation, the estimated proportions may not accurately reflect their actual distribution in the mouse brain. Nevertheless, the changes in cell-type ratios between FXTAS mice and controls may reflect a disease-associated cellular change linked to the expression of expanded CGG repeats in the brain.
Profound gene expression dysregulation in inhibitory neurons of FXTAS mice
We next examined cell type-specific transcriptional alterations in FXTAS. Across all populations, we identified 1670 DEGs (adjusted p < 0.05, Supplementary Data 2), with inhibitory neurons accounting for the majority (79%, n = 1285) (Fig. 3a). Of these, 1216 DEGs also met a log fold-change threshold of >0.1 (Supplementary Fig. 2c). Inhibitory neurons showed a strong bias toward gene downregulation, indicative of a broad suppression of transcriptional programs compared to other cell types (Fig. 3a, b). In contrast, excitatory neurons displayed moderate change by harboring 351 DEGs, while non-neuronal cells displayed minimal alterations. A cross comparison of DEGs between cell types suggested cell type-specific gene expression patterns (Supplementary Fig. 2d). We then investigated the perturbed molecular signatures associated with the pathogenesis of FXTAS. Inhibitory neuron DEGs were linked to critical pathways implicated in neurodegenerative disorders, including endocytosis, axon guidance, and ubiquitin-mediated proteolysis (Fig. 3c). This observation suggested that the expression of expanded CGG repeats dramatically disrupts cellular homeostasis, compromises neuronal connectivity, and impairs protein quality control in inhibitory neurons. In contrast, excitatory neuron DEGs were enriched for pathways related to parathyroid hormone (PTH) secretion and action, phosphoinositide (PI), and cyclic adenosine monophosphate (cAMP) signaling (Fig. 3c).
a The number of DEGs in each cell type (FDR < 0.05). b Violin plot showing the gene expression fold changes in excitatory and inhibitory neurons. c KEGG pathway analyses for DEGs in excitatory and inhibitory neurons. d Correlations between logFCs of DEGs in bulk sequencing (cortex) and logFC of snRNA-seq data within cell types. Correlations were assessed using Spearman’s rank correlation (two-sided) (ρ = Spearman’s rank correlation coefficient). Red dots indicate DEGs detected in snRNAseq data (FDR < 0.05). e Heatmaps show the top 200 DEGs in inhibitory and excitatory neurons. f Dot plot of GO biological processes (P < 0.01) enriched in top 200 DEGs by cell types.
We then examined the concordance between snRNA-seq and bulk RNA-Seq on the cortex (3–4 m) from FXTAS mice by examining the correlation of log-fold changes of genes between the two datasets. Despite modest overall correlations, inhibitory and excitatory neurons exhibited the strongest correlations with bulk gene expression (Fig. 3d; P < 0.001), particularly for DEGs with substantial fold changes. To further clarify the functional components associated with the most significant gene expression changes, we performed GO analyses on the top 200 DEGs in each neuronal cell type, considering their statistical significance and expression fold changes (Fig. 3e). Although several GO terms were common to both cell types, inhibitory neurons uniquely exhibited enrichment in categories related to locomotor behavior and balance control (Fig. 3f), suggesting their specific contribution to FXTAS-associated motor impairments.
Finally, we tested whether these transcriptional alterations could be attributed to higher transgene expression in inhibitory neurons. We appended the GFP sequence (fused in-frame with the human FMR1 5′UTR transgene) to the mouse reference genome and reprocessed the snRNA-Seq data. Transgene expression was estimated by calculating the summed GFP counts normalized to the number of nuclei per cell type in the FXTAS samples. As shown in Supplementary Fig. 2e, GFP-positive nuclei were prominently detected in excitatory neurons, inhibitory neurons, astrocytes, and OPC, but were rare in microglia and oligodendrocytes and absent in endothelial cells (Supplementary Fig. 2e). Average GFP expression was broadly comparable across these major neuronal and glial types. These results demonstrate that transgene expression is broadly distributed across neurons and glial populations derived from Nestin-positive progenitors. Therefore, the pronounced transcriptional dysregulation observed in inhibitory neurons is unlikely to be explained simply by transgene dosage alone, but instead suggests an intrinsic susceptibility of this neuronal population to CGG-repeat toxicity.
Mice expressing FMRpolyG+RNA in GABAergic neurons display the full spectrum of the phenotype
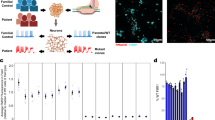
To further dissect the contribution of specific neuronal subtypes to FXTAS pathogenesis, we generated conditional mouse models by crossing Tagger mice41 with two Cre lines driven by Gad2 or CaMKIIα promoters. The Tagger mouse line bears a quad-cistronic transgene that stoichiometrically expresses four proteins to capture distinct populations of nucleic acids41. The resulting Gad2-Tagger and CaMKIIα-Tagger animals were then independently crossed with floxed FXTAS (FMRpolyG+RNA) transgenic mice to selectively express FMRpolyG protein and CGG repeat RNA in GABAergic and glutamatergic neurons, respectively (Fig. 4a). Quantitative RT-PCR revealed high expression of expanded CGG RNA in both cortex and cerebellum (Fig. 4b), validating the effective excision of the loxP cassette and successful transgene expression. Additionally, we found lower transgene expression in brain tissues of transgenic FMRpolyG+RNA; Gad2-Tagger mice compared to FMRpolyG+RNA; CaMKIIα-Tagger animals. This finding is consistent with the known abundance of neuronal types: ~70–85% of cortical neurons are excitatory and 15%-30% are inhibitory, confirming the selected promoters’ specificity.
a Schematic diagram for the breeding strategy of conditional FXTAS mouse models. b Quantitative RT-PCR analysis of transgene expression relative to the RplpO mRNA in cortex and cerebellum of control, FMRpolyG+RNA; Gad2-Tagger, or FMRpolyG+RNA; CaMKIIα-Tagger mice. Data are presented as mean ± SD; n = 3–4 biological replicates per group. Statistical significance was assessed using a two-sided unpaired t-test with Welch’s correction. c, d Time series rotarod test: the time before falling from a rotating rod of control, FMRpolyG + RNA; Gad2-Tagger, or FMRpolyG+RNA; CaMKIIα-Tagger mice. Both male and female mice were used. e, f Time series body weight recording. Both rotarod (c, d) and body weight (g, h) data were analyzed by two-way ANOVA with genotype and age as factors, followed by Šídák’s multiple correction. Data are presented as mean ± SD, and significance was defined as adjusted P < 0.05. We observed significant main effects of age and genotype-by-age interactions in rotarod performance and body weight of FMRpolyG+RNA; Gad2-Tagger mice. Both male and female mice were used. In FMRpolyG+RNA; CaMKIIα-Tagger mice, there is significant age effect on the body weight (F(4211) = 11.57, P < 0.0001), but no effect of genotype or interaction. g, h Immunohistochemistry against calbindin and FMRpolyG N terminus from control, FMRpolyG+RNA; Gad2-Tagger, and FMRpolyG+RNA; Nestin-Cre mice. Representative images are shown. Scale bar = 100 µm. Each experiment was independently repeated at least three times with similar results.
We then performed behavior assays and neuropathological staining to determine whether expanded CGG repeats in specific neurons could recapitulate the deficits observed in mice with brain-specific expression of CGG repeats7. Intriguingly, starting at 4 months of age, mice expressing expanded CGG repeats in GABAergic neurons displayed progressive locomotor deficiency (Fig. 4c) and significant weight gain compared to controls (Fig. 4e). Two-way ANOVA confirmed significant effects of genotype, age, and genotype × age interaction on rotarod performance (F(1,11) = 78.57, P < 0.0001; F(3,33) = 4.15, P = 0.0133; F(3,33) = 12.48, P < 0.0001), indicating progressive motor decline. Body weight analyses also revealed strong genotype, age, and interaction effects (F(1,25) = 15.83, P < 0.0001; F(1.74,43.56) = 308.8, P < 0.0001; F(4100) = 99.53, P < 0.0001), indicating genotype-associated weight gain over time. Of note, male mice displayed more pronounced motor deficits, with phenotypic changes observable as early as 3 months of age (Supplementary Fig. 4a, b). In contrast, females displayed delayed motor decline but earlier obesity (Supplementary Fig. 4c, d). This observation suggested a potential role of gender-specific factors in the progression of FXTAS. In contrast, mice expressing the transgene in glutamatergic neurons showed no evidence of weight gain or locomotor deficits (Fig. 4d, f). ANOVA tests revealed no significant genotype or interaction effects on rotarod performance. For body weight, only age effects were detected (F(4,211) = 11.57, P < 0.0001), consistent with normal growth. These results suggest that expression of expanded CGG repeats in GABAergic neurons alone is sufficient to drive FXTAS phenotypes.
To better evaluate whether body weight as a confounding factor in rotarod deficits, we weight-matched subsets of FMRpolyG+RNA; Gad2-Tagger mice and controls at 3–4 months, when male Gad2 cohorts first exhibited motor decline. Even after weight matching, FMRpolyG+RNA; Gad2-Tagger mice exhibited significantly shorter rotarod latencies than wild-type controls at both ages (Supplementary Fig. 5), indicating that the observed onset of motor impairment is not attributable to differences in body weight. We further investigated whether the expression of expanded CGG repeats in GABAergic neurons could reproduce the phenotypic changes observed in FMRpolyG+RNA; Nestin-Cre animals. Remarkably, both transgenic mouse lines displayed similar levels of mobility impairment at 4 months of age (Supplementary Fig. 4e). However, by 6 months, FMRpolyG+RNA; Nestin-Cre mice had a higher body weight compared to FMRpolyG+RNA; Gad2-Tagger mice (Supplementary Fig. 4f), suggesting that in addition to inhibitory neurons, other cell types may also contribute to energy balance regulation. Finally, neuropathological examination showed a marked reduction of calbindin-labeled Purkinje cells and the accumulation of FMRpolyG-positive aggregates in FMRpolyG+RNA; Gad2-Cre mice, mirroring findings in FXTAS mice (Fig. 4g, h; Supplementary Fig. 6a, b). Taken together, these findings demonstrated that the expression of CGG repeats in GABAergic neurons alone was sufficient to induce FXTAS-like motor, metabolic, and neuropathological phenotypes, whereas the expression in glutamatergic neurons had minimal impact.
TRAP-Seq identifies age-dependent and brain region-specific dysregulation in GABAergic neurons of FXTAS
To further explore molecular signatures of GABAergic neurons during disease progression, we performed anti-HA translating ribosome affinity purification (TRAP) followed by RNA sequencing (TRAP-seq) on both cortex and cerebellum from FMRpolyG+RNA; Gad2-Tagger mice and their littermate controls (Fig. 5a). In these animals, an HA epitope is fused to the large ribosomal subunit protein 22 (Rpl22), allowing selective capture of ribosome-bound mRNAs from GABAergic neurons in response to expanded CGG repeats. Western blot analysis confirmed the expression of FMRpolyG in brain lysates of FMRpolyG+RNA; Gad2-Tagger individuals, but not in control animals (Supplementary Fig. 7a, b). Successful immunoprecipitation (IP) of HA-tagged ribosomes was further validated by anti-HA immunoblotting of brain lysates before and after IP (Supplementary Fig. 7c, d)
a Schematic diagram for the TRAP experiment by using mouse brain tissues. Male mice were used for tissues dissection. b Volcano plots of gene expression changes in the GABAergic neurons from cortex and cerebellum of FMRpolyG+RNA; Gad2-Tagger mice compared to control mice. In the cortex, TRA-Seq was performed on 4 FMRpolyG+RNA; Gad2-Tagger mice and 4 controls at 3–4 months, and on 3 FMRpolyG+RNA; Gad2-Tagger mice and 3 controls at 6–7 months. In the cerebellum, TRA-Seq was performed on 2 FMRpolyG+RNA; CaMKIIα-Tagger mice and 3 controls at 3–4 months, and on 3 mice per genotype at 6–7 months. c, e Venn diagrams showing the number of hnRNPA2/B1 targets by eCLIP-Seq in cortex or cerebellum, and the overlapping targets with dysregulated genes in GABAergic neurons. Two-sided Fisher’s exact tests confirmed that these overlaps were highly significant at both stages and in both brain regions (Supplementary Data 6). Heatmap showing the fold change direction of the shared targets at two stages. d, f Dot plots of GO biological processes (P < 0.05) enriched in dysregulated mRNA targets of hnRNPA2/B1 by stages.
TRAP-seq revealed substantial transcriptional dysregulation across both developmental stages, identifying 1485 DEGs in the cortex and 783 in the cerebellum (Fig. 5b) (Supplementary Data 3). Cross comparisons and KEGG pathway analyses indicated age-dependent and brain region-specific alterations (Supplementary Fig. 8a). For instance, DEGs at 3–4 months in GABAergic neurons were associated with key processes in axon development, such as the enrichment of key processes, including axon guidance, semaphorin-plexin signaling pathway, and axogenesis in both regions (Supplementary Fig. 8b, c). In the cerebellum, DEGs are associated with neuronal and motor-related processes, such as locomotor behavior (3–4 m) and skeletal muscle contraction (6–7 m), among a broader set of categories (Supplementary Fig. 8d). These molecular alterations indicate that pathways important for motor function are disrupted in GABAergic neurons, potentially contributing to the locomotor impairments observed in behavioral assays.
We next sought to leverage these cell type-specific gene expression profiles to the RNA toxicity mechanism contributing to FXTAS pathogenesis. HnRNPA2/B1 is one of the RNA-binding proteins that can become sequestered within the RNA foci formed by the accumulation of expanded CGG repeats RNA10. To link our current data to this model, we compared the TRAP-Seq DEGs with previously generated hnRNPA2/B1 eCLIP-seq datasets from adult mouse brain tissues (Supplementary Data 4). Surprisingly, nearly half of those TRAP-DEGs in GABAergic neurons overlapped with hnRNPA2/B1 targets, with the exception of cerebellar DEGs at the late stage (Fig. 5c, e). Fisher’s exact tests confirmed that these overlaps were highly significant at both stages and in both brain regions (Supplementary Data 6). Among them, 31 genes in the cortex and four genes in the cerebellum were consistently dysregulated at both early and late stages, including factors linked to neuronal development (Nfix, Nfia, Myt1l, Gdf11, Casz1) and protein folding (Unc45a, Hspb6). Other hnRNPA2/B1 targets were dysregulated in an age-dependent manner and exhibited distinct KEGG pathways (Fig. 5d, f), suggesting their dynamic, age-dependent roles in GABAergic neurons. Notably, we found that hnRNPA2/B1 targets mRNA of genes involved in axonal modulation and chromatin organization processes, particularly at the onset of the disease stage. These results could support a role for RNA-binding protein sequestration in driving GABAergic neuron dysfunction in FXTAS.
A combination of TWAS and Drosophila screen identifies 9 genes as genetic modifiers of FXTAS
To identify genetic modifiers of FXTAS, we performed a TWAS by integrating genetic and brain region-specific gene-expression panels from GTEx Consortium (cortex N = 184, cerebellum N = 189, and cerebellar hemisphere N = 158) with WGS summary statistic data from 159 FMR1 premutation carriers. Our analysis revealed 66 genes that reached a Bonferroni-corrected significant threshold of α = 1 × 10−4 across the three brain regions (Fig. 6a, b, c), with 31 genes meeting a more stringent threshold of 2.1 × 10−6 (Supplementary Data 5). These candidate genes play roles in RNA binding, protein ubiquitination, chromatin organization, phosphorylation, and DNA replication (Fig. 6d). Notably, nine genes are presented in at least two examined tissues, including LSM2, PSMB1, PRKCG, PPP6R1, RP11-289I10.2, DXO, SYNGAP1, VARS and CCDC106 (Supplementary Fig. 9a, b). Some candidates, such as PRKCG and RING1, have been implicated in ataxia42,43,44 or Parkinson’s disease45,46,47,48. Interestingly, PSMB1, which encodes a subunit of the 26S proteasome, was identified in this study, which aligns with our previous discovery of PSMB5 (encodes another subunit of the 26S proteasome) as a modifier of FXTAS through a WGS analysis21.
a–c Manhattan plot of association results from the FXTAS transcriptome-wide association study in cortex, cerebellum, and cerebellar hemisphere. Associations were tested using a logistic regression model that further adjusted for the covariates of FMR1 repeat length (both linear and quadratic terms) and the top 2 principal components of ancestry. The x-axis represents the genomic position of the corresponding gene, and the y-axis represents the -log10-transformed association combined P value derived from individual P values from single tissue model-based analyses. The red line shows P = 10-4. d A summary table showing the function of identified TWAS genes in FXTAS. e Drosophila screen identifies 9 genetic modifiers of CGG-associated neurotoxicity. Fly eye images demonstrate enhancement of CGG-associated neurotoxicity with RNAi knockdown of these 9 genes, including tsu (RBM8A), sli (SLIT3), Not3 (CNOT3), ValRS (VARS), Prosbeta6 (PSMB1), Zfrp8 (PDCD2), U2af50 (U2AF2), sff (BRSK1), and CG6013 (CCDC124).
Due to the limited statistical power of current data, instead of taking a statistical approach for prioritizing candidate genes, we prioritized 22 candidate genes for functional screening in our FXTAS Drosophila model by considering the ranking scores of their fly homologs. Knockdown of these genes using the UAS-GAL4 system in the fly eye revealed that 9 genes exhibited genetic modulation of CGG toxicity, as evidenced by a notable enhancement of disrupted eye morphology or the induction of lethality (Fig. 6e; Supplementary Fig. 11a). These results provide a set of candidate genes for further functional investigation.
PRKCG modulates the CGG repeat toxicity associated with FXTAS
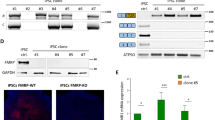
The estimated coefficients (β) of TWAS candidate genes revealed distinct patterns of their enrichment of gene expression levels between FXTAS individuals and controls (Supplementary Data 5). A positive β-value indicates predicted upregulation in FXTAS, while a negative value suggests downregulation. To determine whether these TWAS candidate genes are dysregulated in response to the expression of expanded CGG repeats in mammalian systems, we compared TWAS genes with expression profiles from bulk RNA-seq of FXTAS mouse brains. Five TWAS genes showed significant differential expression in cortex or cerebellum (Fig. 7a). Notably, Prkcg expression was elevated in cortex during the development of the disease, while others were primarily dysregulated at the late progression stage (6–7 m). Western blot analysis confirmed increased PKCγ protein in cortical tissue at 3–4 months but not at 6–7 months (Supplementary Fig. 10), might suggest a transient early upregulation that normalizes at later stages. Integration with TRAP-Seq data from the conditional FXTAS mouse model revealed 7 TWAS genes with altered expression in GABAergic neurons (Fig. 7b). Fisher’s exact test demonstrated significant overlap between TWAS candidates and TRAP-seq DEGs in cerebellar GABAergic neurons at 6–7 months (P = 0.018; supplementary Data 6). Among these, Prkcg was specifically upregulated in cerebellar GABAergic neurons at disease onset. TWAS analysis further indicated consistent positive enrichment of PRKCG across all three regions (Supplementary Fig. 9d), implying its upregulation in premutation carriers. Validation by using RT-qPCR in both mouse cortex and patient-derived neural progenitor cells (NPCs) confirmed significantly elevated PRKCG expression compared with controls (Fig. 7c, d). Additionally, we found that hnRNPA2/B1 binds to Prkcg mRNA. Together, these findings implicate increased PRKCG expression in FXTAS pathogenesis.
a Venn diagrams showing the number of TWAS candidate genes overlapping with DEGs from bulk RNA-Seq on the cortex and cerebellum of the FXTAS mouse model. Bar plot showing the normalized read counts of shared genes in FXTAS and control mice from bulk RNA-Seq. DEGs from 3 to 4 months were selected with P < 0.01; DEGs from 6 to 7 months were selected with FDR < 0.05. Data are presented as mean ± SD; n = 3–4 biological replicates per group. b Venn diagrams showing the number of TWAS candidate genes overlapping with DEGs from TRAP-Seq on GABAergic neurons from cortex and cerebellum of conditional FXTAS model. Bar plot showing the normalized read counts of shared genes in FXTAS and control mice from TRAP-Seq. DEGs from 3 to 4 months were selected with P < 0.01. Data are presented as mean ± SD; n = 2-3 biological replicates per group. Two-sided Fisher’s exact test demonstrated significant overlap between TWAS candidates and TRAP-seq DEGs in cerebellar GABAergic neurons at 6–7 months (P = 0.018; Supplementary Data 6). c Quantitative RT-PCR analysis of Prkcg expression relative to Gapdh in the cortex at two stages. Three biological replicates were used in each group. Data are presented as mean ± SD; n = 3 biological replicates per condition. Statistical significance was assessed using a two-sided unpaired t-test with Welch’s correction. Significance is defined as P < 0.05. d Quantitative RT-PCR analysis of PRKCG expression relative to GAPDH in NPCs from FXTAS patients and controls. Data are presented as mean ± SD; n = 3 biological replicates per condition. Statistical significance was assessed using a one-sided unpaired t-test. Significance is defined as P < 0.05. e Scheme of the overexpression of human PRKCG in the FXTAS fly model. f Overexpression of hPRKCG and aPKC resulted in enhancement of neurotoxicity in the FXTAS fly model. g Cell viability assays following shRNA-mediated knockdown of Prkcg (left) and hPRKCG overexpression (right) in N2a cells expressing expanded CGG repeats. Data are presented as mean ± SD and statistical significance was assessed using a two-sided unpaired t-test with Welch’s correction. Significance is defined as P < 0.05.
PRKCG, encodes protein kinase C gamma (PKCγ), a neuron-enriched isoform of the protein kinase C (PKC) family49. To further investigate its functional role in FXTAS, we overexpressed the Drosophila ortholog of PKC (aPKC) in the eye of the FXTAS fly model, resulting in the enhancement of the rough eye phenotype (Fig. 7f, the middle panel). Given the absence of a specific PRKCG ortholog in fly, we cloned the full-length cDNA of human PRKCG (hPRKCG) into a pUAST Drosophila vector and generated transgenic UAS lines that overexpressed hPRKCG under the control of a GAL4 driver (Fig. 7e). Upon crossing these transgenic lines with the FXTAS fly model, we observed that overexpression of hPRKCG induced the rough eye phenotype in the progenies (Fig. 7f, the right panel; Supplementary Fig. 11b), mirroring the effects of aPKC and demonstrating that PRKCG overexpression exacerbates CGG repeat-induced neurotoxicity.
To further validate this mechanism in mammalian cells, we manipulated PRKCG expression in Neuro2A (N2A) cells expressing CGG-repeat RNA by using knockdown and overexpression approaches. Short hairpin RNA (shRNA)-mediated knockdown of Prkcg significantly improved cell viability measured at 96 h (Welch’s t-test, P < 0.001) (Fig. 7g). Conversely, overexpression of hPRKCG further reduced cell viability compared with CGG repeats RNA expression alone, consistent with our fly genetic results. Taken together, the results from human genetic analyses, mouse models, fly screening, and cell-based toxicity assays demonstrate a critical role of PRKCG in FXTAS. These findings establish PRKCG as a biologically relevant modifier of CGG repeat–associated neuronal toxicity and support its prioritization as a candidate therapeutic target.
Discussion
Neurodegenerative diseases often exhibit region-specific pathologies and selective neuronal vulnerabilities. Prior FXTAS studies have primarily focused on bulk gene expression changes in selected brain regions of animal models50,51 and on proteomic analyses of human postmortem brain tissue52, offering limited insight into the complex molecular alterations underlying FXTAS. Moreover, the cellular heterogeneity of the central nervous system may mask toxic levels of expanded CGG repeat RNAs. Our study addresses these gaps by employing a multi-omics strategy in a FXTAS mouse model to systematically investigate the distinct brain regions, cell types, and pathways affected by expanded CGG repeats, thereby providing a deeper understanding of FXTAS molecular pathologies.
Our bulk RNA-seq analysis revealed distinct transcriptional signatures across the cortex, cerebellum, and hippocampus of the FXTAS mouse model. The cerebellum and cortex, both crucial for motor function, exhibited the most profound changes, whereas the hippocampus showed more modest dysregulation despite harboring FMRpolyG-positive aggregate7. GO analysis further suggested that the hippocampus engages distinct signaling pathways compared to motor-related regions, whereas the cerebellum and cortex exhibit alterations in chromatin organization, protein homeostasis, and RNA processing. This regional specificity may help explain why FXTAS patients predominantly exhibit motor deficits despite the widespread expression of expanded CGG repeats in the brain.
A key question in FXTAS is whether the pathogenic effects of expanded CGG repeats are driven by RNA gain-of-function, RAN translation, or both. Additional mechanisms, including DNA damage from R-loop formation, mitochondrial dysfunction, and 5hmC-mediated epigenetic modulation, have also been proposed53,54. Our findings support a combined mechanism. The marked upregulation of genes involved in chromatin organization, protein folding and ubiquitination, RNA processing, and DNA damage response in the cerebellum and cortex points to interconnected pathogenic processes. Accumulation of RNA foci disrupts mRNA processing, including alternative splicing, transport, and translation, ultimately resulting in broad dysregulation of gene expression4. FMRpolyG aggregates could further exacerbate these abnormalities by interfering with the function of RNA-binding proteins and ribosomal machinery55,56. Moreover, the presence of γH2AX (a sensitive DNA damage marker) in FXTAS inclusions underscores the role of genomic instability in disease progression57,58. Our integrated TRAP-Seq and eCLIP-Seq analyses identified numerous dysregulated mRNAs in GABAergic neurons that are bound by hnRNPA2B1, many of which are involved in axonal modulation and chromatin organization, implicating the RNA-binding protein sequestration as a key pathogenic mechanism.
We profiled the PFC by snRNA-seq, given its cellular diversity and its relevance to executive, behavioral, and preparatory motor functions59. Although cerebellar pathology is central to motor coordination deficits in FXTAS, the dominance of granule cells complicates the resolution of vulnerable subtypes such as Purkinje cells. As a neuropathological hallmark of FXTAS, intranuclear inclusions have been documented in both neurons and glial cells in Fragile X premutation mouse models as well as in the brains of FXTAS patients, suggesting that multiple cell types may be vulnerable to the toxic effects of expanded CGG repeat expression. It has also even been suggested that in early disease states, the inclusions may be protecting the cell by sequestering FMRpolyG1,17. Intriguingly, our single-nucleus data revealed inhibitory neurons as the most profoundly dysregulated population in FXTAS at early stages, exhibiting broad transcriptional suppression and enrichment for locomotor and balance-related pathways. Compared with the controls, FXTAS mice also showed increased proportion of inhibitory neurons in the PFC, suggesting excessive inhibition that could dampen neural circuitry and normal excitation. In contrast, a recent snRNA study highlighted the widespread cellular impact of expanded CGG repeats, suggesting the glial dysregulation in premutation pathophysiology at the end-of-life stage60. This points to a complex interplay among genetic factors (such as CGG repeat length), aging, and neurodegeneration in the development of FXTAS. For example, studies have shown a significant reduction in the abundance of inhibitory neurons in the frontal cortex during aging61. Such changes may mask the subtle effects of expanded CGG repeats on neuronal subtypes. Although we did not detect altered cell abundance or gene expression in astrocytes, this likely reflects dynamic shifts in cellular composition over the disease course. Further investigations are warranted to elucidate the stage-specific molecular signatures in both neuronal and glial populations and their contributions to FXTAS disease progression. It is also possible that some disease-associated mRNA effects are offset by homeostatic translational control, justifying future translational profiling and proteomic studies of the FXTAS brain.
Inhibitory neurons are essential for maintaining excitatory-inhibitory balance, which is crucial for motor function. Previous research provided some insights into the involvement of GABA system in the pathogenesis of FXTAS. For instance, elevated expression of several GABAA receptor subunits and proteins involved in GABAergic metabolism have been found in the cerebellum of an animal model of FXTAS62. Subsequent studies revealed altered GABAergic metabolism in premutation carriers of FXTAS63,64,65. However, these investigations focused on a limited set of genes, leaving gaps in our understanding of how alterations in GABAergic neurons contribute to the behavioral and neurological phenotype associated with FXTAS. Our work expands this view by showing widespread transcriptional dysregulation in PFC inhibitory neurons of the FXTAS mouse model, with GO categories enriched for locomotor behavior and balance control. Importantly, selective expression of expanded CGG repeats in GABAergic neurons alone was sufficient to induce motor decline, Purkinje cell loss, and intranuclear inclusions, thereby recapitulating phenotypes observed in pan-neuronal models7. Moreover, we observed sex-specific differences in disease onset and progression: male FXTAS mice exhibited earlier locomotor deficits, whereas female mice showed delayed but progressive changes, including earlier onset of weight gain. These findings parallel clinical observations in FXTAS patients, where penetrance, age of onset, and severity are generally higher in males than females1. In humans, this sex difference is often attributed to the X-linked nature of the FMR1 locus, with protective effects from the second X chromosome in females. However, our autosomal integrated model suggests that the sex-specific factors beyond X-linked dosage, such as hormonal influences, metabolic differences, or intrinsic neuronal vulnerabilities may contribute to FXTAS phenotypes.
Intriguingly, our integrative analyses also identified potential genetic biomarkers and therapeutic targets for FXTAS. Across bulk RNA-seq, TRAP-seq, and qRT-PCR, Prkcg expression was consistently elevated in FXTAS mouse tissues (Fig. 7), and validation in human premutation NPCs confirmed PRKCG upregulation. Functional studies demonstrated that PRKCG overexpression exacerbated CGG-repeat-induced neurotoxicity in Drosophila and mammalian cells, whereas shRNA-mediated knockdown of Prkcg mitigated toxicity. Mutations in PRKCG have been implicated in spinocerebellar ataxia-14 (SCA14)42,43,44 and Parkinson’s disease66,67. PKC activation is known to drive Purkinje cell degeneration68,69, and increased PKCγ activity leads to neurodegeneration via protein aggregation in SCA14. Both wild-type and mutant PKCγ overexpression has been found to form amyloid-like fibrils and aggregates in cultured cells70. Endogenous expression of two SCA14 mutations in the C1 domain of PKCγ led to cytoplasmic mis-localization and aggregation in both patient iPSCs and cerebellum42. However, the interplay between altered PKCγ activity and aggregation in contributing to SCA14 pathogenesis has yet to be elucidated. Here, our findings link elevated PRKCG expression to FXTAS pathogenesis, with hnRNPA2/B1 binding to Prkcg mRNA providing a mechanistic connection to CGG repeats RNA-mediated toxicity. These results nominate PRKCG as a promising biomarker and druggable target for FXTAS. However, how PRKCG contributes to the neuropathology of FXTAS needs further investigation.
A potential limitation of our study is that differences in disease phenotypes between FMRpolyG+RNA; Gad2-Tagger and FMRpolyG+RNA; CaMKIIα-Tagger mice could, in part, could be caused by differences in transgene expression levels. qRT-PCR results of cortex and cerebellum indicated lower transgene levels in Gad2-driven mice compared with CaMKIIα-driven mice (Fig. 4b), consistent with the relative abundance of inhibitory versus excitatory neurons in the cortex. Importantly, the transgene in these conditional mouse models is driven by the strong CAG promoter, which robustly induces neural expression71,72. The expected phenotypes, such as Purkinje cell loss in Gad2-Cre and Nestin-Cre but not CaMKIIα cohorts, further support effective induction in both lines. GFP-based snRNA-seq analysis further demonstrated broad transgene expression across excitatory, inhibitory, and glial lineages (Supplementary Fig. 2e). These findings argue against dosage alone explaining the strong inhibitory vulnerability, though precise quantification remains technically challenging. Future studies using spatial RNA imaging or lineage-specific quantification will be required to resolve this important question. Another limitation is that our model simultaneously produces expanded-repeat RNA and RAN-translated FMRpolyG3,4,5,6,7, making it difficult to dissect the relative contributions of RNA- versus protein-mediated toxicity in specific neuronal populations. Future work that can selectively block RAN translation or CGG-repeats RNA accumulation will be essential to define their respective roles in FXTAS pathogenesis.
In summary, our study provides a comprehensive exploration of the transcriptional alterations in the brain of the FXTAS animal model and reveals profound transcriptome dysregulation in inhibitory neurons. Using animal models, we demonstrated that expression of expanded CGG repeats in GABAergic neurons alone was sufficient to induce hallmark features of FXTAS, including intranuclear inclusions, Purkinje cell loss, and motor deficits. Our findings also identify a set of hnRNPA2B1-bound mRNAs and additional genetic modifiers through TWAS and fly screening, with PRKCG emerging as a particularly promising target. Taken together, our data highlight the critical role of GABAergic neuronal dysfunction in FXTAS and offer a potential therapeutic target for FXTAS.
Methods
Animals
Mice were housed in a pathogen-free facility at the Emory University Division of Animal Resources under standard conditions. Animals were maintained on a 12-h light/12-h dark cycle (lights on at 7:00 AM) with ad libitum access to food and water. The ambient temperature was maintained at 20–22 °C with a relative humidity of 40–60%. All animals were group-housed unless single housing was required for a specific procedure. Nestin-Cre transgenic (strain #003771; with C57BL/6 genetic background), Camk2a-Cre transgenic mice (strain #005359; with C57BL/6 genetic background), and Gad2-IRES-Cre knock-in mice (strain #010802; with C57BL/6 genetic background) were obtained from The Jackson Laboratory. Transgenic mice (with C57BL/6 genetic background) containing the entire human 5′ UTR of FMR1 with expanded 99 CGG repeats expressing CGG RNA and FMRpolyG protein were obtained from Baylor College of Medicine. Transgene expression was controlled by a loxP-flanked polyadenylation cassette and activated by Cre-mediated recombination.
The Tagger mouse line (with C57BL/6 genetic background) was obtained from Dr. Walker Jackson41. Under the control of CAG promoter and two transcription-terminating STOP cassettes (Frt-NeoR-Frt and LoxP-STOP-LoxP), Tagger mice can stoichiometrically express four proteins for the capture of distinct populations of nucleic acids, including HA-tagged ribosomal protein L22 (Rpl22-HA, for Ribo-Tag), Uracil phosphoribosyltransferase (TgUPRT, for TU-Tag), red fluorescent protein with a triple NLS (RFP-NLS, for Nuc-Tag), and FLAG-V5-tagged Argonaute2 (FLAG-V5-Ago2, for Ago-Tag). The mice were first crossed with ROSA26-Flpo mice (stock 007844, The Jackson Laboratory, Bar Harbor ME) to remove the FNF cassettes, creating LSL-Tagger lines. Then, Gad2-Cre and Gad2-IRES-Cre mouse lines were used to activate cell-type specific expression of Tagger transgene. The resulting animals were further crossed with FXTAS mice to obtain progenies with cell type-specific expression of both FMRpolyG, CGG RNA and Tagger transgene.
Male mice were used for all tissue dissection and all sequencing experiments including bulk RNA-seq, snRNA-seq, and TRAP-seq, to reduce molecular variability associated with sex-specific gene expression, particularly variability introduced by the female estrous cycle73. Both male and female mice were used for behavioral assays to ensure that the phenotypic consequences of expanded CGG repeats are assessed across sexes, thereby increasing the generalizability and translational relevance of the findings. Age- and sex-matched littermates were randomly allocated to experimental groups. The age and exact number of animals used in each experiment are provided in the corresponding figure legends and the Reporting Summary.
Time-series weight measurement and rotarod test
Both male and female mice were used for body weight measurements and rotarod tests. Body weights of mice with correct genotypes are recorded from 2 to 6 months old. Rotarod tests were conducted from 3 to 6 months old each month. The rod was cleaned and set to a fixed speed of 4.0 RPM before the test. Three testing trails were performed, during which the rotation speed accelerated from 4 to 40 rpm in 5 min. As the mice fall from the rod, each lane’s trip plate will stop the timer and display the latency in seconds. Trials were separated by 5–10 min intervals. The mean of latency was calculated and used as an indicator of locomotor performance.
Isolation of RNA from mouse brain tissue
For bulk RNA-seq, male mice were used, with a minimum of three biological replicates included for each genotype. Cortex, cerebellum and hippocampus were dissected from each mouse and homogenized in 1 mL of TRIzol (Invitrogen, 15596026) followed by standard RNA isolation protocol. Briefly, samples were incubated in TRIzol at RT for 5 min. Following incubation, 200 μL chloroform was added and samples were vigorously shaken for 15 s. The mixture was then incubated at RT for 15 min and centrifuged at 20,000 × g for 15 min at 4 °C. The upper aqueous layer, containing RNA, was carefully transferred to a new tube. To precipitate the RNA, 600 μL of isopropyl alcohol was added, and the samples were incubated at RT for 10 min before centrifugation at 20,000 × g for 10 min at 4 °C. The resulting RNA pellet was washed with 75% ethanol, air-dried for 10 min, and dissolved in RNase-free water. The quality and concentration of the RNA were assessed using the Qubit 2.0 RNA HS Assay (Thermo Fisher Scientific, Q32852).
Translating ribosome affinity purification (TRAP) and RNA isolation
The TRAP procedure41 were performed on cortex and cerebellum tissues from male mice. Brain tissues were dissected in pre-cold PBS buffer and homogenized in PBS containing 50 mM Tris (pH 7.5), 100 mM KCl, 12 mM MgCl2, and 1% Nonidet P-40, one mM DTT, 100 U/mL Ribolock RNase inhibitor, 100 μg/mL cycloheximide, and 1 tab/5 mL SigmaFast protease inhibitor cocktail (with EDTA, Sigma-Aldrich, S8830-2TAB). The homogenates were kept on ice for over 15 min and centrifuged at 20,000 x g for 10 min at 4 °C to remove nuclei and cell debris. Approximately 50 μL of S1 supernatant was kept for input RNA isolation, and the remaining supernatant was pre-cleared with Protein G Dynabeads (Thermo Fisher Scientific, 10003D) at 4 °C on a rotator for 30 min. The S2 supernatant was incubated with 10ul HA antibody (Roche, Anti-HA, 12CA5) on a rotator at 4 °C for 2 h, followed by incubating with protein G beads for an additional 2 h at the same condition. Afterward, the beads were collected and washed 3 times with High Salt Buffer (HSB: 50 mM Tris (pH 7.5), 300 mM KCl, 12 mM MgCl2 and 1% Nonidet P-40, one mM DTT, 50 U/mL Ribolock RNase inhibitor (Thermo Fisher Scientific, EO0382), and additional 3X wash by using Extra High Salt Buffer (EHSB: identical to HSB but containing additional 300 mM NaCl). Per each wash, beads were rotated gently at 4 °C for 2 min. During the last wash, 10% of the beads were saved for western blot, and the remaining beads were used for RNA isolation by TRIzol Reagent. The quality and concentration of the RNA were assessed using the Qubit 2.0 RNA HS Assay.
RNA-Seq and data analysis
The isolated mRNAs were prepared for the library according to the manufacturer’s protocols. Paired-end sequencing (2 × 150) was performed on an Illumina NovaSeq X-plus instrument. Sequencing reads were aligned to the mm10 mouse genome using TopHat v2.1.1. The number of reads aligned to each gene was tabulated using FeatureCount v1.5.3. All reads mapped to gene bodies were used for differential gene expression analysis using R package DEseq2 v1.30.1. DEGs were selected as those with a P < 0.05. DAVID web server was employed to perform GO analyses. Heatmaps were generated using R package pheatmap v1.0.13 with hierarchical clustering applied to rows.
Isolation of nuclei from mouse cortex
All steps were performed on ice or at 4 °C. Frontal cortex from two male FXTAS mice and two male littermate controls was rapidly dissected in ice-cold homogenization buffer (HB: 260 mM sucrose, 30 mM KCl, 10 mM NaCl, 20 mM Tricine-KOH pH 7.8, 1 mM DTT, 0.5 mM Spermidine, 0.2 mM Spermine, 0.3% NP40, cOmplete Protease inhibitor (Sigma-Aldrich, 11697498001), and Ribolock RNase inhibitor (Thermo Fisher Scientific, EO0382) and immediately transferred to a Dounce homogenizer filled with 1 mL HB. Tissue was homogenized by 20 strokes with each pestle A and pestle B. The homogenate was then filtered through a 70um strainer followed by centrifugation for 10 min at 300 × g at 4 °C. The pallets were then resuspended in a homogenization buffer and mixed with 1 volume of 50% Iodixanol (Sigma-Aldrich, D1556) to make a final concentration of 25% iodixanol. Without disturbing the mixture, 30% Iodixanol was slowly layered underneath the 25% iodixanol mixture, and then 40% Iodixanol was gently added under the 30% mixture to generate a density gradient. A clearly defined interface should be visible between all three layers. The nuclei were centrifuged for 20 min at 1500 × g and 4 °C in a swinging bucket centrifuge with the brake off. After the centrifugation, the nuclei band was transferred from the interface between 30% and 40% iodixanol solutions to a pre-chilled tube. The nuclei were washed with wash buffer at 4 °C and counted after adding trypan blue.
Library preparation for snRNA-Seq
Libraries were generated with the 10x Genomics Chromium Single Cell Gene Expression reagent kit following the manufacturer’s instructions with minor modifications. Briefly, per sample, 16,100 nuclei were resuspended in 1x diluted nuclei buffer (10x Genomics) with 2% BSA, and 10,000 nuclei were captured. For 10,000 nuclei capture targeted, single nuclei expression libraries were sequenced to 20,000 read pairs per nuclei on an Illumina NovaSeq 6000 instrument at the Florida State University Translational Science Laboratory.
Data pre-processing and analyses of snRNA-seq
Raw reads were obtained with 10x Genomics pipeline CellRanger (v5.0.0) “mkfastq” with default parameters. Reads from each sample were then aligned to the mouse genome (mm10) with reference data “refdata-cellranger-mm10-2020-A-2.0.0”, and the gene-by-cell count matrices were generated. R package Seurat (V4.3.0.1) was performed for quality control (QC) analyses and downstream analyses, including data normalization, clustering, highly variable gene selection, etc. We created a Seurat object containing all four samples (two wildtypes and two FXTAS) and performed QC on the percentage of reads mapped to the mitochondrial (MT) genome. The MT percentage ranges from 0 to 3.37% with a mean of 0.01%, presenting high data quality; therefore, we did not filter any cells based on MT percentage. Next, according to the distribution of number of genes expressed in cells, we kept cells with at least 200 and at most 4000 genes expressed, resulting in 17,794 cells with 32,285 genes measured. In the downstream analyses, we first performed data normalization using “NormalizeData” with scale factor set as 10,000 and log-transformation. Then, “FindVariableFeatures” was used to select the top 2000 genes that exhibit high cell-to-cell variability with variance-stabilizing transformation. Data was then center scaled with “ScaleData,” and dimension reduction was performed using “RunPCA” with 30 principal components (PCs). To mitigate batch effect between conditions, we used harmony (v0.1.1) to perform batch correction. Next, “RunUMAP” was performed on the top 20 embeddings from harmony to obtain UMAP embeddings for visualization purposes. Based on the top 20 embeddings from harmony, we performed “FindNeighbors” to construct a shared nearest neighbors (SNN) graph and “FindClusters” with a resolution of 0.6 to obtain clusters using the Leiden algorithm.
Cell type annotation and differential gene expression analysis
Both unsupervised clustering and supervised classification methods were applied to identify seven major cell types. For unsupervised clustering, we used “FeaturePlot,” “DotPlot,” and “DoHeatmap” to visualize known marker genes (Slc17a7, Gad1, Gad2, Neurod2, Nrn1, Dlx1, Mobp, Mog, Acsbg1, Aqp4, Csf1r, Ctss, Flt1, Pdgfra) in each cluster for cell type identification purposes. In the meantime, two supervised classification methods (Cellcano74 and Seurat V4.3.0.1) were performed to double-confirm the identified cell types using two well-annotated prefrontal cortex datasets (GSE124952, GSE116470) as reference datasets. After careful curation, we removed clusters and cells that could not be identified and obtained 13,902 cells. Based on identified cell types, we performed differential gene expression analysis with “FindMarkers” in Seurat between conditions. Average log2FC (avg_log2FC) larger than 0.1 and adjusted p value less than 0.05 were used as cutoffs to identify DEGs per cell type.
Immunohistochemistry and immunofluorescence
Transcardiac perfusion was performed with saline followed by 4% paraformaldehyde (PFA) to clear blood and brain tissue. Brain tissue was post-fixed in 4% PFA overnight at 4 °C, and the Neuropathology Core of Emory Center for Neurodegenerative Disease prepared paraffin-embedded brain sections. Sections were deparaffinized at 58 °C for 40 min, immersed in xylene for 10 min, rehydrated through graded ethanol (100% and 95%), rinsed three times in water and three times in PBS. Antigen retrieval was performed in sodium citrate buffer (pH 6.0) at 95 °C for 30 min, followed by rinsing in PBS. To quench endogenous peroxidase activity, sections were incubated in 0.3% H₂O₂ in PBS for 10 min at room temperature, then permeabilized in 0.3% Triton X-100/PBS for 1 h and blocked in 2% normal goat serum (NGS) and 3% BSA for 2 h at room temperature. Sections were incubated overnight at 4 °C overnight with primary antibody against FMRpolyG (Sigma-Aldrich, MABN2280, clone 8FM-2F7) or Calbindin-D28k (ProteinTech, 66394-1-PBS) diluted in PBS containing 3% BSA, 2% NGS, and 0.1% Triton X-100. After three washes in PBS, sections were incubated for 1 h with biotin-conjugated goat anti-mouse secondary antibody (Jackson ImmunoResearch, 115-065-166; 1:500 in PBS with 3% BSA and 2% NGS), followed by three PBS washes. Detection was performed using the VECTASTAIN® Elite® ABC Kit (PK-6100) for 1 h, with three PBS washes, and visualized with 1× DAB substrate. Sections were washed three times in water and counterstained with hematoxylin. Finally, slides were dehydrated, covered, and imaged using a ZEISS Axio Imager.A2 microscope. For immunofluorescent, sections were co-stained overnight at 4 °C with primary antibodies against GABA (Thermo Fisher, PA5-32241) and NeurN (Sigma-Aldrich, MAB377), diluted in PBS containing 3% BSA, 2% NGS, and 0.1% Triton X-100. After three washes in PBS, sections were incubated for 1 h with secondary antibodies (Invitrogen: Alexa FluorTM 568 goat anti-mouse IgG (H + L), and Alexa FluorTM 647 goat anti-rabbit IgG (H + L)), followed by three PBS washes. Nuclear staining was performed with DAPI for 3 min at room temperature, after which sections were washed with PBS and mounted with antifade mounting medium. Slides were cured in the dark at room temperature for 1 h. Images were acquired using a Leica DMi8 microscope and analyzed with ImageJ.
Western blot analysis of mouse brain lysate
The brain tissue was homogenized using a Dounce homogenizer in lysis buffer containing 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% Triton X-100, and cOmplete Protease inhibitor (Sigma-Aldrich, 11697498001). Homogenates were incubated on ice for 30 min and centrifuged at 1200 × g for 10 min at 4 °C. The supernatant was transferred to a new tube and centrifuged again at 15,000 × g for 20 min at 4 °C. The final supernatant was collected on ice, and protein concentrations were quantified using a NanoDrop spectrophotometer according to the manufacturer’s instructions. Proteins were mixed with 4× Laemmli sample buffer (Bio-Rad, 34229167) containing 10% β-mercaptoethanol (Fisher Scientific, 21985-023) and denatured at 95 °C for 10 min. Samples were resolved by SDS–PAGE on 4–12% Bis-Tris gels and transferred to PVDF membranes. Membranes were blocked in 5% non-fat dry milk in PBST for 1 h at room temperature and incubated with primary antibodies in 5% milk overnight at 4 °C. After three washes in PBST, membranes were incubated with HRP-conjugated secondary antibodies, and protein bands were visualized by chemiluminescent detection reagent (Cytiva, RPN2235) on X-ray film (GE Healthcare). Band intensities were quantified using ImageJ. Primary antibodies used: anti-HA (Roche, 12CA5), anti-FMRpolyG (Sigma-Aldrich, MABN2280, 8FM-2F7), anti-GAPDH (ThermoFisher, MA1-16757), anti-ACTIN (ThermoFisher, MA5-11869), anti-PKC gamma (ThermoFisher, 13-3800).
Study population of FMR1 premutation carriers and WGS
In this study, participant recruitment and sample acquisition were coordinated through the National Fragile X Center at Emory University as described previously21. Once a participant was screened for eligibility and provided consent, a blood or saliva sample was collected, and a general medical history questionnaire was completed. For FXTAS, 159 participants who were carriers of the FMR1 premutation were recruited21. Specifically, 94 males and females with onset of FXTAS motor symptoms before age 65 are enrolled as cases, while 56 males without FXTAS symptoms by age 68 are enrolled as controls21. All samples were mapped to the human genome using PEMapper, and the variants were called using PECaller75. To correct for confounding due to ancestry or batch effects, principal component analysis was performed on a set of 567,720 independent SNPs using PLINK 1.9. The top 2 PCs were retained for covariate adjustment in subsequent studies.
TWAS analysis
We used TIGAR-V276 to train prediction models of gene expression using reference transcriptomic data and genetic data from various tissues collected by the Genotype-Tissue Expression (GTEx) project V8. Expression prediction models were derived from the brain cortex (N = 184), cerebellum (N = 189), and cerebellar hemisphere tissues (N = 158) within GTEx. For each tissue, we performed pre-processing of the tissue expression and genetic data from GTeX using standard QC pipelines. Specifically, TIGAR-V2 considered only genes with gene expression of transcripts per million (TPM) > 0.1 in ≥10 samples and variants with minor allele frequencies >0.01, missing rates <20%, and Hardy-Weinberg equilibrium p > 10−5 for fitting gene expression prediction models. TIGAR-V2 adjusted gene expression for age, body mass index, top five genotype PCs, and top probabilistic estimation of expression residual factors for each gene. TIGAR-V2 then fit prediction models on the adjusted gene expression data treating cis-SNPs (within ±1 Mb region of transcription start sites of the target gene) as predictors. Assuming an additive genetic model for the expression quantitative trait (\({E}_{g}\)) of target gene g, the prediction model can be represented as follows.
where \({G}_{{ref}}\) represents the vector of cis-SNP genotypes for gene g in the GTeX data with corresponding effect sizes w. Within the model, TIGAR-V2 estimates w for each gene using a popular nonparametric Bayesian Dirichlet process regression (DPR) procedure. Second, using estimates of cis-eQTL effect sizes \(\hat{w}\) from each tissue, we applied TIGAR-V2 to impute the genetically regulated gene expression (GReX) of gene g in our FXTAS sample as \(\widehat{{GReX}}={G}_{{test}}\hat{w}\), where Gtest represents the corresponding cis-SNP genotypes for gene g in the FXTAS dataset). Finally, we tested the association between \(\widehat{{GReX}}\) and FXTAS case/control status using a logistic regression model that further adjusted for the covariates of FMR1 repeat length (both linear and quadratic terms) and the top 2 PCs of ancestry.
Genetic screen using Drosophila model
As previously described, the transgenic fly line expressing r(CGG)90 was established20. The GMR-GAL4 and UAS-TRiP lines were obtained from Bloomington Stock Center. All Drosophila lines were maintained, and crosses were performed in the standard medium at 25 °C. To determine the genetic interaction between CGG repeats and candidate genes, eye phenotypes of RNAi lines crossed with the CGG repeat stable line were compared with the eye phenotype of the CGG repeat stable line. The screen was performed by scoring eye phenotype in progeny with n > 10, which was visualized using light microscopy and confirmed with scanning electron microscopy (SEM).
Generation of Drosophila lines overexpressing human PRKCG
Full-length human PRKCG cDNA was obtained from a tagged ORF clone (OriGene Technologies, PRKCG Human Tagged ORF Clone, RG208502). The PRKCG open reading frame subcloned into the pUAST Drosophila expression vector77(Addgene) using standard restriction enzyme–based cloning. Both the PRKCG insert and the pUAST backbone were digested with Not1 and Kpn1 (New England Biolabs, R3189S and R3142S) at 37 °C for 30 min and ligated using T4 DNA ligase (New England Biolabs, M0202) at 16 °C overnight, followed by heat inactivation at 65 °C for 10 min. The resulting construct (pUAST-hPRKCG) was verified by Sanger sequencing. For transgenesis, the validated plasmid was injected into embryos for germline transformation using P-element–mediated insertion. Embryo injections and founder selection were performed by BestGene Inc. (Chino Hills, CA; https://www.thebestgene.com). Transgenic UAS-hPRKCG lines were established and maintained using standard Drosophila genetic crosses. Overexpression of human PRKCG in the FXTAS fly model was achieved by crossing UAS-hPRKCG flies with the appropriate GAL4 driver lines as indicated in the figure legends.
Cell lines and culture
Murine Neuro2A (N2A) cells (ATCC, CCL-131), derived from a male Mus musculus, were cultured in Eagle’s Minimum Essential Medium (EMEM) (Corning; 10-009-CV) with 2.5% fetal bovine serum (FBS). Cells were maintained at 37 °C with 5% CO₂ and passaged using standard trypsin–EDTA dissociation.
Skin biopsy samples were obtained from individual diagnosed with FXTAS (two female and one male) and age- and sex-matched health controls. CGG repeat sizes were verified by short tandem repeat (STR) based genotyping performed at Emory Integrated Genomics Core. All studies were conducted under an approved IRB protocol, and informed consent was obtained from all participants. These fibroblast lines are not commercially available. Fibroblasts from skin biopsies were reprogrammed into iPSCs using a CytoTune-iPS 2.0 Sendai reprogramming kit (ThermoScientific, A16517) according to the manufacturer’s instructions to generate integration-free iPSCs. After transduction, cells were maintained in fibroblast medium [DMEM supplemented with 10% FBS and 2 mM L-glutamine (Thermo Fisher, 25030081). Transduced fibroblasts were transferred onto mouse embryonic fibroblast (MEF) dishes in fibroblast medium. Emerging iPSC colonies were monitored and manually picked for expansion. Established iPSCs were transferred to Matrigel-coated six-well plates, maintained in mTeSR1 medium (STEMCELL Technologies, 85850), and passaged with ReLeSR (STEMCELL Technologies, 05873). NPCs were differentiated from hiPSC as described in refs. 78,79 with some modifications. Colonies of hiPSC colonies were detached using collagenase IV at 1 mg mL⁻¹ (ThermoFisher, 17104019) for 1 h and transferred to low-adhesion dishes in EB medium consisting of DMEM/F12 (Corning, 10-092-CV), 20% KnockOut Serum Replacement (ThermoFisher, 10828028), 2 µM dorsomorphin (Tocris, 3093), and 2 µM A-83-01 (Tocris, 2939). Embryoid bodies (EBs) were cultured for 7 days with daily half-medium changes. EBs were plated onto growth factor–reduced Matrigel-coated dishes (Corning, 354230) and cultured for 7 days in DMEM/F12 supplemented with N2 (ThermoFisher, 17502048) and laminin (µg/mL; Sigma, L2020) to induce neural rosette formation. Neural rosettes were manually isolated on day 14 and expanded for 7 days in NPC medium composed of a 1:1 mixture of DMEM/F12 and Neurobasal medium (ThermoFisher, 21103049) supplemented with N2, B27 minus retinoic acid (ThermoFisher, 12587010), bFGF (20 ng/mL; PeproTech, 100-18B), human LIF (10 ng/mL; PeproTech, 300-05), CHIR99021 (3 µM; Tocris, 4423), and A-83-01 (2 µM; STEMCELL Technologies, 72024). Y-27632 (10 µM; Tocris, 1254) was included only during the first 24 h after replating. The resulting neural progenitor cells were collected and used for downstream assays.
Cell viability assay
Predesigned shRNAs targeting the coding sequence of mouse Prkcg (Sigma; TRC Clone IDs TRCN0000022684, TRCN0000022687, TRCN0000022688) in the pLKO.1 lentiviral vector were obtained. N2a cells were plated in EMEM (Corning, 10-009-CV) containing 2.5% FBS. Twenty-four hours after plating, cells were transfected with the 5′UTR CGG 99× FMR1-EGFP plasmid, which expresses 99 CGG repeats within the human FMR1 5′UTR fused in-frame with EGFP, or with the empty vector control (pcDNA3-EGFP), using GenJet (SignaGen SL100488; 300 ng per well of a 96-well plate). After additional 24 h, cells were transfected with either the PRKCG expression vector or a mixture of the three shRNAs (300 ng per well). Four hours after transfection, the media was replaced with new EMEM. Ninety-six hours after transfection, cell viability was measured using Cell Titer Blue (Promega, 8081) according to the manufacturer’s instructions. The fluorescence of resorufin converted from resazurin by live cells was recorded with FLUOstar Omega microplate reader (BMG LABTECH) at fluorescence (560 Excitation/590 Emission). The average reading value of triplicate empty wells and blank wells with culture media without cells were subtracted from the reading value. The fluorescence values of treated groups were further normalized with empty control vector or control shRNA value.
Quantitative real-time RT-PCR
Total RNAs from mouse tissues or cells were isolated by TRIzol™ Reagent (Invitrogen) according to the manufacturer’s instructions. cDNAs were synthesized from 1 μg of total RNA using the SuperScript@ III First-Strand Synthesis System. All primers and TaqMan Assays used for RT-PCR are listed in Supplementary Data 7. Primer sequences for quantifying transgene expression in the conditional mouse models were obtained from Sellier et al.7 and synthesized by IDT. PRKCG expression in mouse cortex and NPCs were quantified using TaqMan Assays. To measure expression of the transgene, qPCR was performed using PowerUp™ SYBR™ Green Master Mix (Applied Biosystems, A25742) on a QuantStudio 3 instrument (Applied Biosystems) with the following cycling conditions: 94 °C for 15 min followed by 50 cycles of 94 °C for 15 s, 58 °C for 20 s, and 72 °C for 20 s. PRKCG expression was quantified using TaqMan™ Universal PCR Master Mix (Applied Biosystems, 4364340) with standard parameters: 95 °C for 10 min C followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. The mRNA levels of Rplpo or Gapdh were used as an internal control. Relative expression was calculated using the ΔΔCT method with GraphPad Prism 9 software.
Statistics and reproducibility
Statistical analysis was performed using GraphPad Prism 9.0 software. Sample sizes were selected based on prior experience with similar mouse, cellular, and genetic studies and are reported in the figure legends and Methods. To assess for differences between two groups, unpaired Welch’s t-test or two-way ANOVA test followed by Šídák’s multiple correction was performed otherwise stated. Correlations between logFCs of DEGs in bulk sequencing (cortex) and logFC of snRNA-seq data within cell types were assessed using two-sided Spearman’s rank correlation tests. All data in bar charts were presented as mean ± SD unless otherwise stated. No data were excluded from the analyses. Within each genotype, sex- and age- matched mice were assigned to experimental groups randomly. Unless otherwise specified, all experiments were performed with at least three independent biological replicates, and all results were successfully reproduced across independent replicates. Histological analyses and behavioral assessments were performed blinded to treatment groups.
Ethics statement
The protocols and consent forms of this study were approved by the Institutional Review Board at Emory University, and written informed consent was obtained from all subjects (IRB00074941). The animal study was approved by the Emory University Institutional Animal Care and Use Committee (IACUC) for all mouse procedures under the protocol PROTO201700104.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq, snRNA-seq, and TRAP-seq data generated in this study have been deposited in the Gene Expression Omnibus (GEO) database under accession code GSE286188. The processed data are available at Supplementary Data 1–3. The scRNA-Seq reference datasets for adult mouse brain were obtained under accession code GSE124952 and GSE116470. The raw WGS data for FMR1 premutation carriers are protected and are not available due to data privacy. The processed WGS summary statistics used in this study are available under restricted access because they contain patient-level genetic information that cannot be publicly released due to privacy and consent requirements. Access can be obtained by submitting a request to the corresponding author (peng.jin@emory.edu) and completing a data use agreement. Source data are provided with this paper.
Code availability
Code used for the snRNA-seq analysis in this study is available on GitHub at: https://github.com/marvinquiet/analysis_code_for_FXTAS. Analyses for bulk RNA-seq and TRAP-seq were performed using publicly available software packages and established pipelines, including TopHat and FeatureCounts (https://github.com/johanzi/RNA-seq_pipeline), and DESeq2 (https://cran.r-project.org/web/packages/mpmi/index.html). Details of the analysis workflows, parameters, and functions used are provided in the corresponding sections of the Methods. The TWAS analysis code includes components that interact with patient-level genetic data and, therefore, cannot be publicly released due to privacy and consent restrictions. Access to the TWAS analysis scripts can be obtained by contacting the corresponding author (peng.jin@emory.edu) and completing a data use agreement.
References
Hagerman, R. J. & Hagerman, P. Fragile X-associated tremor/ataxia syndrome-features, mechanisms and management. Nat. Rev. Neurol. 12, 403–412 (2016).
Rodriguez-Revenga, L. et al. Penetrance of FMR1 premutation associated pathologies in fragile X syndrome families. Eur. J. Hum. Genet. 17, 1359–1362 (2009).
Jin, P. et al. Pur α binds to rCGG repeats and modulates repeat-mediated neurodegeneration in a Drosophila model of fragile X tremor/ataxia syndrome. Neuon 55, 556–564 (2007).
Li, Y. & Jin, P. RNA-mediated neurodegeneration in fragile X-associated tremor/ataxia syndrome. Brain Res. 1462, 112–117 (2012).
Todd, P. K. et al. CGG repeat-associated translation mediates neurodegeneration in fragile X tremor ataxia syndrome. Neuon 78, 440–455 (2013).
Oh, S. Y. et al. RAN translation at CGG repeats induces ubiquitin proteasome system impairment in models of fragile X-associated tremor ataxia syndrome. Hum. Mol. Genet. 24, 4317–4326 (2015).
Sellier, C. et al. Translation of expanded CGG repeats into FMRpolyG is pathogenic and may contribute to fragile X tremor ataxia syndrome. Neuon 93, 331–347 (2017).
Tassone, F., Iwahashi, C. & Hagerman, P. J. FMR1 RNA within the intranuclear inclusions of fragile X-associated tremor/ataxia syndrome (FXTAS). RNA Biol. 1, 103–105 (2004).
Pretto, D. I. et al. Reduced excitatory amino acid transporter 1 and metabotropic glutamate receptor 5 expression in the cerebellum of fragile X mental retardation gene 1 premutation carriers with fragile X-associated tremor/ataxia syndrome. Neurobiol. Aging 35, 1189–1197 (2014).
Sofola, O. A. et al. RNA-binding proteins hnRNP A2/B1 and CUGBP1 suppress fragile X CGG premutation repeat-induced neurodegeneration in a Drosophila model of FXTAS. Neuon 55, 565–571 (2007).
Sellier, C. et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J. 29, 1248–1261 (2010).
Sellier, C. et al. Sequestration of DROSHA and DGCR8 by expanded CGG RNA repeats alters microRNA processing in fragile X-associated tremor/ataxia syndrome. Cell. Rep. 3, 869–880 (2013).
He, F., Krans, A., Freibaum, B. D., Taylor, J. P. & Todd, P. K. TDP-43 suppresses CGG repeat-induced neurotoxicity through interactions with HnRNP A2/B1. Hum. Mol. Genet. 23, 5036–5051 (2014).
Kearse, M. G. et al. CGG repeat-associated non-AUG translation utilizes a cap-dependent scanning mechanism of initiation to produce toxic proteins. Mol. Cell. 62, 314–322 (2016).
Krans, A., Skariah, G., Zhang, Y., Bayly, B. & Todd, P. K. Neuropathology of RAN translation proteins in fragile X-associated tremor/ataxia syndrome. Acta Neuropathol. Commun. 7, 1–17 (2019).
Tassone, F. et al. Intranuclear inclusions in neural cells with premutation alleles in fragile X associated tremor/ataxia syndrome. J. Med. Genet. 41, e43-e43 (2004).
Greco, C. et al. Neuropathology of fragile X-associated tremor/ataxia syndrome (FXTAS). Brain 129, 243–255 (2006).
Valor, L. M., Morales, J. C., Hervás-Corpión, I. & Marín, R. Molecular pathogenesis and peripheral monitoring of adult fragile X-associated syndromes. Int. J. Mol. Sci. 22, 8368 (2021).
Wenzel, H. J., Hunsaker, M. R., Greco, C. M., Willemsen, R. & Berman, R. F. Ubiquitin-positive intranuclear inclusions in neuronal and glial cells in a mouse model of the fragile X premutation. Brain Res. 1318, 155–166 (2010).
Jin, P. et al. RNA-mediated neurodegeneration caused by the fragile X premutation rCGG repeats in Drosophila. Neuon 39, 739–747 (2003).
Kong, H. E. et al. Identification of PSMB5 as a genetic modifier of fragile X–associated tremor/ataxia syndrome. Proc. Natl. Acad. Sci. USA 119, e2118124119 (2022).
Sun, Y. et al. A transcriptome-wide association study of Alzheimer’s disease using prediction models of relevant tissues identifies novel candidate susceptibility genes. Genome Med. 13, 1–11 (2021).
Tang, S. et al. Novel Variance-Component TWAS method for studying complex human diseases with applications to Alzheimer’s dementia. PLoS. Genet. 17, e1009482 (2021).
Liu, N. et al. Hippocampal transcriptome-wide association study and neurobiological pathway analysis for Alzheimer’s disease. PLoS. Genet. 17, e1009363 (2021).
Yao, S. et al. A transcriptome-wide association study identifies susceptibility genes for Parkinson’s disease. npj. Parkinsons. Dis. 7, 79 (2021).
Li, Y. I., Wong, G., Humphrey, J. & Raj, T. Prioritizing Parkinson’s disease genes using population-scale transcriptomic data. Nat. Commun. 10, 994 (2019).
Shi, J.-J. et al. Joint analysis of proteome, transcriptome, and multi-trait analysis to identify novel Parkinson’s disease risk genes. Aging 16, 1555 (2024).
Xiao, L., Yuan, Z., Jin, S., Huang, S. & Zeng, P. Multiple-tissue integrative transcriptome-wide association studies discovered new genes associated with amyotrophic lateral sclerosis. Front. Genet. 11, 587243 (2020).
Park, S., Kim, D., Song, J. & Joo, J. W. J. An integrative transcriptome-wide analysis of amyotrophic lateral sclerosis for the identification of potential genetic markers and drug candidates. Int. J. Mol. Sci. 22, 3216 (2021).
Pain, O. et al. Harnessing Transcriptomic Signals for Amyotrophic Lateral Sclerosis to Identify Novel Drugs and Enhance Risk Prediction. Heliyon. 10, 15 (2023).
Li, P. et al. Identifying candidate genes associated with sporadic amyotrophic lateral sclerosis via integrative analysis of transcriptome-wide association study and messenger RNA expression profile. Cell. Mol. Neurobiol. 43, 327–338 (2023).
Ross-Inta, C. et al. Evidence of mitochondrial dysfunction in fragile X-associated tremor/ataxia syndrome. Biochem. J. 429, 545–552 (2010).
Park, J. et al. RAS–MAPK–MSK1 pathway modulates ataxin 1 protein levels and toxicity in SCA1. Nature 498, 325–331 (2013).
Dard, L., Bellance, N., Lacombe, D. & Rossignol, R. RAS signalling in energy metabolism and rare human diseases. Biochim. Biophys. Acta Bioenerg. 1859, 845–867 (2018).
Alvarez-Mora, M. et al. Impaired mitochondrial function and dynamics in the pathogenesis of FXTAS. Mol. Neurobiol. 54, 6896–6902 (2017).
Gohel, D., Berguerand, N. C., Tassone, F. & Singh, R. The emerging molecular mechanisms for mitochondrial dysfunctions in FXTAS. Biochim. Biophys. Acta Mol. Basis Dis. 1866, 165918 (2020).
Duffney, L. J. et al. Epigenetics and autism spectrum disorder: a report of an autism case with mutation in H1 linker histone HIST1H1E and literature review. Am. J. Med. Genet. B Neuropsychiatr. Genet. 177, 426–433 (2018).
Fröhlich, A. S. et al. Single-nucleus transcriptomic profiling of human orbitofrontal cortex reveals convergent effects of aging and psychiatric disease. Nat. Neurosci. 27, 2021–2032 (2024).
Bhattacherjee, A. et al. Cell type-specific transcriptional programs in mouse prefrontal cortex during adolescence and addiction. Nat. Commun. 10, 4169 (2019).
Salem, N. A. et al. Cell-type brain-region specific changes in prefrontal cortex of a mouse model of alcohol dependence. Neurobiol. Dis. 190, 106361 (2024).
Kaczmarczyk, L. et al. Tagger—A Swiss army knife for multiomics to dissect cell type–specific mechanisms of gene expression in mice. PLoS. Biol. 17, e3000374 (2019).
Wong, M. M. et al. Neurodegeneration in SCA14 is associated with increased PKCγ kinase activity, mislocalization and aggregation. Acta Neuropathol. Commun. 6, 1–14 (2018).
Riso, V. et al. NGS-based detection of a novel mutation in PRKCG (SCA14) in sporadic adult-onset ataxia plus dystonic tremor. Neurol. Sci. 41, 2989–2991 (2020).
Vlak, M. H., Sinke, R. J., Rabelink, G. M., Kremer, B. P. & van de Warrenburg, B. P. Novel PRKCG/SCA14 mutation in a Dutch spinocerebellar ataxia family: expanding the phenotype. J. Mov. Disord. 21, 1025–1028 (2006).
Dev, K. K., van der Putten, H., Sommer, B. & Rovelli, G. Part I: parkin-associated proteins and Parkinson’s disease. Neuropharmacol 45, 1–13 (2003).
Shimura, H. et al. Familial Parkinson disease gene product, parkin, is a ubiquitin-protein ligase. Nat. Genet. 25, 302–305 (2000).
Cookson, M. R. et al. RING finger 1 mutations in Parkin produce altered localization of the protein. Hum. Mol. Genet. 12, 2957–2965 (2003).
Niaz, S. et al. Mutation analysis of RING1 domain of Parkin in early onset of Parkinson’s disease in Pakistani patients-a pilot study. Punjab Univ. J. Zool. 31, 249–254 (2016).
Lordén, G. & Newton, A. C. Conventional protein kinase C in the brain: repurposing cancer drugs for neurodegenerative treatment?. Neuronal Signal. 5, NS20210036 (2021).
Yao, B. et al. Genome-wide alteration of 5-hydroxymethylcytosine in a mouse model of fragile X-associated tremor/ataxia syndrome. Hum. Mol. Genet. 23, 1095–1107 (2014).
Kong, H. E. et al. Metabolic pathways modulate the neuronal toxicity associated with fragile X-associated tremor/ataxia syndrome. Hum. Mol. Genet. 28, 980–991 (2019).
Holm, K. N. et al. Human cerebral cortex proteome of fragile X-Associated Tremor/Ataxia syndrome. Front. Mol. Biosci. 7, 600840 (2021).
Garcia-Arocena, D. & Hagerman, P. J. Advances in understanding the molecular basis of FXTAS. Hum. Mol. Genet. 19, R83–R89 (2010).
Schwartz, J. L., Jones, K. L. & Yeo, G. W. Repeat RNA expansion disorders of the nervous system: post-transcriptional mechanisms and therapeutic strategies. Crit. Rev. Biochem 56, 31–53 (2021).
Hoem, G., Bowitz Larsen, K., Sjøttem, E. & Johansen, T. The FMRpolyGlycine protein mediates aggregate formation and toxicity independent of the CGG mRNA hairpin in a cellular model for FXTAS. Front. Genet. 10, 449501 (2019).
Gohel, D. et al. FMRpolyG alters mitochondrial transcripts level and respiratory chain complex assembly in Fragile X associated tremor/ataxia syndrome (FXTAS). Biochim. Biophys. Acta Mol. Basis Dis. 1865, 1379–1388 (2019).
Iwahashi, C. et al. Protein composition of the intranuclear inclusions of FXTAS. Brain 129, 256–271 (2006).
Hoem, G. et al. CGG-repeat length threshold for FMR1 RNA pathogenesis in a cellular model for FXTAS. Hum. Mol. Genet. 20, 2161–2170 (2011).
Le Merre, P., Ährlund-Richter, S. & Carlén, M. The mouse prefrontal cortex: Unity in diversity. Neuon 109, 1925–1944 (2021).
Dias, C. M. et al. Glial dysregulation in the human brain in fragile X-associated tremor/ataxia syndrome. Proc. Natl. Acad. Sci. USA 120, e2300052120 (2023).
Rozycka, A. & Liguz-Lecznar, M. The space where aging acts: focus on the GABA ergic synapse. Aging Cell. 16, 634–643 (2017).
D’Hulst, C. et al. Expression of the GABAergic system in animal models for fragile X syndrome and fragile X associated tremor/ataxia syndrome (FXTAS). Brain. Res. 1253, 176–183 (2009).
Conde, V. et al. Abnormal GABA-mediated and cerebellar inhibition in women with the fragile X premutation. J. Neurophysiol. 109, 1315–1322 (2013).
Giulivi, C., Napoli, E., Tassone, F., Halmai, J. & Hagerman, R. Plasma biomarkers for monitoring brain pathophysiology in FMR1 premutation carriers. Front. Mol. Neurosci. 9, 71 (2016).
Giulivi, C., Napoli, E., Tassone, F., Halmai, J. & Hagerman, R. Plasma metabolic profile delineates roles for neurodegeneration, pro-inflammatory damage and mitochondrial dysfunction in the FMR1 premutation. Biochem. J. 473, 3871–3888 (2016).
Craig, N. J. et al. A candidate gene for human neurodegenerative disorders: a rat PKCγ mutation causes a Parkinsonian syndrome. Nat. Neurosci. 4, 1061–1062 (2001).
Chen, Y. et al. Early-onset Parkinson’s disease with atypical molecular imaging abnormalities in a patient carrying the de novo PRKCG mutation. Park. Relat. Disord. 95, 100–102 (2022).
Ghoumari, A. M., Wehrle, R., De Zeeuw, C. I., Sotelo, C. & Dusart, I. Inhibition of protein kinase C prevents Purkinje cell death but does not affect axonal regeneration. J. Neurosci. 22, 3531–3542 (2002).
Schrenk, K., Kapfhammer, J. & Metzger, F. Altered dendritic development of cerebellar Purkinje cells in slice cultures from protein kinase Cγ-deficient mice. Neuroscience 110, 675–689 (2002).
Takahashi, H. et al. Identification and characterization of PKCγ, a kinase associated with SCA14, as an amyloidogenic protein. Hum. Mol. Genet. 24, 525–539 (2015).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuon 71, 995–1013 (2011).
Nguyen, A. Q., Dela Cruz, J. A., Sun, Y., Holmes, T. C. & Xu, X. Genetic cell targeting uncovers specific neuronal types and distinct subregions in the bed nucleus of the stria terminalis. J. Comp. Neurol. 524, 2379–2399 (2016).
Knoedler, J. R. et al. A functional cellular framework for sex and estrous cycle-dependent gene expression and behavior. Cell 185, 654–671.e622 (2022).
Ma, W., Lu, J. & Wu, H. Cellcano: supervised cell type identification for single cell ATAC-seq data. Nat. Commun. 14, 1864 (2023).
Johnston, H. R. et al. PEMapper and PECaller provide a simplified approach to whole-genome sequencing. Proc. Natl. Acad. Sci. USA 114, E1923–E1932 (2017).
Parrish, R. L., Gibson, G. C., Epstein, M. P. & Yang, J. TIGAR-V2: efficient TWAS tool with nonparametric Bayesian eQTL weights of 49 tissue types from GTEx V8. HGG. Adv. 3, 100068 (2022).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Wen, Z. et al. Synaptic dysregulation in a human iPS cell model of mental disorders. Nature 515, 414–418 (2014).
Xu, R. et al. OLIG2 drives abnormal neurodevelopmental phenotypes in human iPSC-based organoid and chimeric mouse models of Down syndrome. Cell Stem Cell 24, 908–926.e908 (2019).
Acknowledgements
We are grateful to Nicolas Charlet-Berguerand from the Innstitut de Génétique et de Biologie Molélaire et Cellulaire (IGBMC) at University of Strasbourg for generously sharing the FMRpolyG conditional lines and W. Jackson from the Department of Biomedical and Clinical Sciences at Linköping University for generously sharing the Tagger mouse line used in this study. We thank S. Sloan, J. Hu, Y. Hou, and B. Yao from the Department of Human Genetics at Emory School of Medicine for their valuable assistance with snRNA-Seq data analysis and immunostaining. We are also thankful to M. Gearing and K. M. Kelly from the Neuropathology Core of the Emory Center for Neurodegenerative Disease for preparing the paraffin-embedded brain sections used in this study. The support and resources provided by the High-Performance Computing facilities at Emory University are gratefully acknowledged. This work was supported by the U.S. National Institutes of Health grants NS111602 and HD104458 to P.J.
Author information
Authors and Affiliations
Contributions
P.J. conceived the study. Y.J. and P.J. wrote the manuscript. Y.J. and Y.C. and performed mouse breeding and most experimental procedures. Y.J. and W.M. performed most of the data analysis. Y.J. performed image analysis for microscope data. Y.J. and R.L. and performed immunostaining staining. Y.L., Y.K., and X.G. performed the eCLIP experiment and data analysis. J.H. and M.P.E. performed TWAS analyses. N.R. and J.L. performed fly screening. Y.K. performed cell viability assay. Y.Z. and Y.K. performed cell culture. E.G.A. and Z.W. provided advice and technical assistance to the study. All the authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jin, Y., Cao, Y., Ma, W. et al. Integrative transcriptome-wide association analyses reveal PRKCG-linked GABAergic dysfunction in Fragile X-associated tremor/ataxia syndrome. Nat Commun 17, 1418 (2026). https://doi.org/10.1038/s41467-025-68163-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68163-9