Abstract
β-Strand motifs are essential recognition modules in protein-protein interactions (PPIs), which govern cellular signaling networks and regulate molecular pathway dynamics. Herein we present an unexpected discovery of a previously uncharacterized β-strand insertion mechanism termed as cross-β-strand linking, wherein β-strands within the β-sheet-rich aggregates form inter-β-sheet connections through insertion into adjacent β-sheets. These cross-β-strand linkers comprise <15% of the total β-strands in the amyloidogenic aggregates, but they can mediate a significant proportion of intermolecular interactions, operating as dynamic molecular adapters that regulate the inter-β-sheet packing geometry. Crucially, these linkers exist as conformational ensembles of heterogeneous substates, bestowing remarkable structural diversity to the aggregates. Through promiscuous engagement with multiple conformational substates, cross-β-strand linkers enable the aggregates to balance order and disorder. In this work, we provide a perspective on how low-abundance structural elements can orchestrate complex molecular architectures in assembly systems.
Similar content being viewed by others
Introduction
PPIs are foundational to cellular communications and to orchestrate molecular pathways through precise recognition events1,2,3. These interactions are also critical targets for the therapeutic interventions of diseases4,5,6. While the classical paradigm of interprotein recognition emphasizes shape and chemical complementarity at the interfaces, i.e., the solvent-excluded regions that contribute to binding specificity and affinity7,8. However, our current understanding of PPI mechanisms remains far from complete. Over 95% of human PPIs have not been structurally characterized so far9. Moreover, native proteins usually exist as conformational ensembles comprising multiple interchangeable states, ranging from predominant low-energy conformations to sparsely populated high-energy ones10,11,12. The inherent conformational diversity of protein complexes and their complicated dynamics make the structural characterization of PPIs more challenging13,14,15,16,17. While conventional techniques like cryogenic electron microscopy (cryo-EM), X-ray crystallography, and nuclear magnetic resonance (NMR) spectroscopy can offer us atomic-resolution structural insights, they rely on ensemble-averaged reconstructions and primarily capture the most thermodynamically stable populations10,18,19,20. Obviously, this ensemble-averaged attribute in these techniques fundamentally obscures the characterization of transient and/or low-abundance conformational substates, which are critical to PPI regulation10,17,21.
Short peptide motifs frequently act as recognition elements in PPI, functioning through burial in binding partner cavities, positioning within pockets/grooves, or β-sheet integration at protein interfaces6,22,23. Among PPIs involving peptide recognition elements, β-strand addition emerges as an important mechanism governing interprotein recognition23,24,25. This process involves the donation of ligand β-strand to receptors through three canonical classes (Fig. 1a): (1) β-sheet augmentation via edge-to-edge interactions between a ligand β-strand and a partner β-sheet; (2) β-strand complementation, where a ligand β-strand is embedded into its complementary receptor; and (3) β-strand zippering through antiparallel alignment of complementary flexible regions in both interactors23,26,27,28,29,30. Herein, we describe a previously unrecognized β-strand insertion mechanism: β-strands within the β-sheet-rich aggregates bridge adjacent β-planar sheets by their insertion into neighboring β-sheets (Fig. 1b). We therefore term it a cross-β-strand linking mechanism.
a Three canonical pathways of the donation of β-strands to receptors: β-sheet augmentation, β-strand complementation, and β-strand zippering. b Schematic illustration of the cross-β-strand linking. c–f Cross-β-strand linking observed in the gal-3 225–250 aggregate via the STM imaging. Four different STM images of cross-β-strand linkers are provided. White boxes highlight the structure of the cross-β-strand linker. Scale bars: 3 nm. STM experiments were independently repeated using different samples and tips. At least three high-resolution STM images (60 × 60 nm) were acquired for each sample.
In this work, a cross-β-strand linking mechanism is identified in amyloid aggregation. Amyloid aggregation plays a crucial role in many protein misfolding-related diseases, such as Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis31,32,33. Amyloid fibrils are mainly composed of β-sheet structures stabilized by extensive intermolecular hydrogen bonding and exhibit remarkable structural stability34. A typical feature of amyloid aggregates is conformational heterogeneity, namely, identical peptide sequences can adopt distinct conformational substates coexisting within a heterogeneous conformational ensemble. Such conformational heterogeneity is closely linked to their pathological functions17,21. However, due to the inherent heterogeneity of amyloid structures, conventional structural techniques (e.g., X-ray crystallography, NMR, and cryo-EM) remain limited in resolving coexisting conformations. To directly visualize the local structural features within amyloid fibrils, we employed scanning tunneling microscopy (STM) under ambient conditions. This technique provides exceptional resolution down to the single-molecule level, offering a complementary perspective on the heterogeneous conformational ensemble of amyloid fibrils. STM relies on electron tunneling and requires a conductive, flat substrate; therefore, highy-oriented pyrolytic graphite (HOPG) was used as the supporting substrate. Unlike more reactive STM substrates such as Cu(001) or Cu(110)—which may form covalent bonds with adsorbates—HOPG is chemically inert to interact with adsorbates35. Although STM imaging was performed under nominally dry conditions, a thin residual water film on the HOPG surface helped maintain local hydration, thereby preventing dehydration-induced denaturation36. We use STM-based imaging to detect non-periodic structural signals with submolecular precision and reveal the structural basis of cross-β-strand linking. Despite constituting a minor population (<15%) of the total β-strands in aggregates, these cross-β-strand linkers exhibit conservation across diverse amyloid peptides, promoting the structural order of amyloid fibril architecture through inter-sheet cross-linking.
Results
STM imaging provides direct visualization of β-strand insertion in amyloid β-sheet aggregates
We initiated our investigation with the aggregates formed by the 225–250 segment of galectin-3 (gal-3 225–250), which participates in the PPIs between gal-3 molecules and their phase separation37,38. This peptide undergoes spontaneous self-assembly into amyloid-like fibrils in aqueous solution, as confirmed by transmission electron microscopy (TEM) imaging (Supplementary Fig. 1a). Fourier transform infrared (FTIR) spectroscopy confirms the conformational transition of the fibrils, as evidenced by the increased β-sheet (1632 cm−1) bands relative to its monomeric state (Supplementary Fig. 1b). Time-lapse thioflavin-T (ThT) fluorescence assays show a sigmoidal increase in fluorescence intensity, characteristic of amyloid fibril formation kinetics (Supplementary Fig. 1c). The structure-informed machine learning analysis using ZipperDB [https://zipperdb.mbi.ucla.edu/] predicts39 a high aggregation propensity for gal-3 225–250 with Rosetta energy below −23 kcal/mol, which is well aligned with experimental observations (Supplementary Fig. 1d). These multiscale validation approaches collectively establish gal-3 225–250 as a reliable model system for studying β-sheet-driven self-assembly processes.
To investigate the intermolecular interfaces of β-sheet aggregates, we performed STM imaging of the gal-3 225–250 aggregates after equilibrium incubation (36 h). Our group recently developed an STM-based single-molecule imaging platform, which bypasses the limitation of ensemble averaging and enables direct characterization of conformational ensembles17,21,40,41. The samples were deposited onto freshly cleaved HOPG substrates, a standard choice for ambient STM studies due to its atomically flat surface and chemical inertness42. The STM tips with an atomically sharp radius were homemade from platinum-iridium alloys via the combined processes of mechanical and electric pulsing. Distinct molecular conformations on the HOPG surface result in the change of the local density of electron states, which consequently modulate tunneling current magnitudes. By scanning the tip across the surface, we obtained real-space topographical maps reflecting the molecular structures, dispensing with structural reconstructions.
The single-molecule imaging of gal-3 225–250 aggregates reveals signature β-sheet features (Supplementary Fig. 1e and Supplementary Table 1). The molecules appear as periodic bright linear patterns forming lamellar structures, with protrusions in the local density of electron states corresponding to peptide bonds and aromatic residues21,42,43. The intermolecular spacing between adjacent molecules (4.2 ± 0.1 Å) matches the characteristic β-sheet interstrand distance42,44 (Supplementary Fig. 1f). Individual β-strands within the β-sheet assemblies exhibit an orientation perpendicular to the direction of sheet extension. We also attempted STM imaging of the sample before aggregation (0 h) (Supplementary Fig. 2). However, the result shows that they lack sufficient intermolecular and substrate interactions to form stabilized, ordered structures under ambient conditions, because the unassembled molecules on the HOPG surface undergo rapid thermal diffusion, which precludes the acquisition of single-molecule images45.
More detailed analysis of gal-3 225–250 aggregates via STM imaging uncovers an unprecedented structural feature: stochastic cross-β-strand linkers acting as molecular mortise-and-tenon joints between two β-sheets (Fig. 1c–f). These exceptional β-strand linkers protrude from one β-sheet domain, traverse through the inter-sheet gap, and integrate seamlessly into an adjacent β-sheet domain without any structural discontinuity. Two features are observed for the β-strand linkers. First, the occurrence frequency is low: there are only ~5 instances observed per 15 × 15 nm area containing ~100 molecules. The second one is the conformational plasticity. Individual cross-β-strand linkers exhibit a pronounced length variability (3.0–7.3 nm), indicative of their dynamic conformational flexibility. Our findings represent direct visualization of stochastic β-strand-mediated inter-sheet connections in amyloid aggregates, challenging the conventional models of β-sheet aggregation that typically emphasize the highly ordered arrangement of β-strands46,47,48.
Cross-β-strand linkers form interstrand connections with distinct conformational substates in β-sheet aggregates
Single-molecule imaging allows us to decode the structural organization of cross-β-strand linkers with a nanoscale resolution. In the gal-3 225–250 aggregates, β-strands exhibit length variations from 1.6 to 7.3 nm (average: 4.2 ± 1.2 nm), while cross-β-strand linker lengths span from 3.0 to 7.3 nm. Given the periodicity of 3.25 Å between consecutive residues in the β-strands, we established a quantitative relationship between strand length and amino acid sequence: N = L/3.25 Å, where N is the number of residues and L is β-strand length49. Detailed discussion is provided in Supplementary Note 1.1. This relationship allows us to distinguish the distinct conformational substates based on their residue counts (Supplementary Fig. 3a).
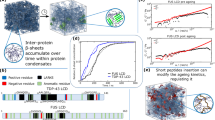
As shown in Fig. 2a, this analysis reveals a highly heterogeneous conformational ensemble in the β-sheet aggregates: 18 coexisting conformational substates (labeled I–XVIII) were identified, corresponding to 5–22 residues. The most abundant substate (IX) contains 13 residues and accounts for 14.1% of total β-strands (Fig. 2b). Among these 18 substates, 14 substates (V–XVIII) participate in the formation of cross-β-strand linkers (Fig. 2b). Importantly, the cross-β-strand linker represents a minor structural element within individual conformational substates (Supplementary Table 2).
Representative STM images (a) and structural models for each conformational substate of gal-3 225–250 (b). Scale bar: 2 nm. Gray: conformations with no cross-β-strand linker. Blue and red indicate the proportion without and with the cross-β-strand linker, respectively. c The inter-conformational interactions involving the cross-β-strand linker (red) within the gal-3 225–250 aggregates relative to the total interactions (overall). d Representative STM images of inter-conformational interactions. White boxes highlight the cross-β-strand linker-mediated interpeptide interactions. Scale bars: 3 nm. STM experiments were independently repeated using different samples and tips. At least three high-resolution STM images (60 × 60 nm) were acquired for each sample. Source data are provided as a Source data file.
In a pleated β-sheet, adjacent β-strands are interconnected laterally through a hydrogen bond network that stabilizes the structure47,49,50. The conformational diversity of β-strands enhances the heterogeneity in the interstrand recognition. Within the gal-3 225–250 aggregates, we identified 139 distinct inter-β-strand interaction types among the 18 conformational substates, with 62 interactions (accounting for ~45% of total interaction types with a population only around ~14%) involving the cross-β-strand linkages (Fig. 2c, d and Supplementary Fig. 3b). The disproportionate contributions of the cross-β-strand linkers to the interactions indicate that they are critical mediators of interstrand interactions. Cross-β-strand linkers exhibit no preferential selection bias toward specific conformational substates (Supplementary Fig. 4), as all conformational substates in the aggregates participate in the interactions with cross-β-strand linkers.
Cross-β-strand linkers are prevalent across diverse β-sheet aggregates
With the cross-β-strand linkers observed, one question naturally arises: Is it unique to gal-3 225–250 aggregates or prevalent in the β-sheet-rich aggregates? To answer this question, we employed STM to characterize a series of aggregation structures formed by β-sheet-containing peptides, including the 206–231 segment of galectin-3 (gal-3 206–231), the 129–154 segment of galectin-3 (gal-3 129–154), amyloid-β 42 (Aβ42), amyloid-β 42 with an N-terminal extension of six residues (−5Aβ42), the 8–37 segment of human islet amyloid polypeptide (hIAPP 8–37), hIAPP 8–37 with a replacement of Ser-to-Gly at site 20 (hIAPP S20G 8–37), and the 341–357 segment of the transactive response DNA binding protein 43 (TDP-43 341–357). Under the same incubation conditions used for STM imaging, the results of TEM, ThT fluorescence, and FTIR analyses coherently confirm that these aggregates are enriched in β-sheets and exhibit characteristic amyloid-like properties51, while AFM imaging further confirms their amyloid-like morphology on the HOPG surfaces (Supplementary Fig. 1a and Supplementary Figs. 5–9). Comparative FTIR spectra of the different peptide aggregates are shown in Supplementary Tables 3 and 4 and a detailed discussion is provided in Supplementary Note 1.2. The STM results confirm that all these peptide aggregates exhibit hallmarks of β-sheet assembly characterized by periodic electron-dense streaks arranged in a lamellar pattern with an orientation perpendicular to the direction of sheet extension (Fig. 3a–h and Supplementary Fig. 10, Supplementary Table 1).
Cross-β-strand linking observed in the aggregates of gal-3 206–231 (a), Aβ42 (b), hIAPP 8–37 (c), KRT-R (d), and hIAPP S20G 8–37 (e). Blue links denote the inter-conformational interactions that do not involve the cross-β-strand linker, whereas red links highlight those mediated by the cross-β-strand linker. The proportion of linker-mediated inter-conformational interactions relative to total interactions within peptide aggregates is quantified at the top of the STM images. White boxes highlight the structure of the cross-β-strand linker. Scale bars: 3 nm. The linker-negative aggregates of -5Aβ42 (f), TDP-43 341–357 (g), and gal-3 129–154 (h). STM experiments were independently repeated using different samples and tips. At least three high-resolution STM images (60 × 60 nm) were acquired for each sample. Source data are provided as a Source data file.
Besides, the occurrence of cross-β-strand linkers exhibits significant molecular specificity. Linker-positive aggregates (Fig. 3a–e) include gal-3 206–231 (12.6% of total β-strand population), KRT-R (4.2%), Aβ42 (4.1%), hIAPP 8–37 (3.0%), and hIAPP S20G 8–37 (2.1%), while linker-negative aggregates (Fig. 3f–h) comprise −5Aβ42, TDP-43 341–357, and gal-3 129–154. We also scanned STM images of Aβ42 aggregates at 80 μM and consistently observed cross-β-strand linkers (Supplementary Fig. 11). Further analysis also confirms the two features shared by the cross-β-strand linkers across different molecular systems. First, the cross-β-strand linkers in all the systems account for a low population but participate in the interaction types in a disproportionate manner, which is similar to the case of the gal-3 225–250 aggregates above (Supplementary Fig. 12, and Supplementary Table 5). Second, these linkers demonstrate exceptional conformational plasticity, with quantitative analysis revealing distinct conformational substates: 20 substates for gal-3 206–231, 14 for Aβ42, 19 for hIAPP 8–37, 19 for hIAPP 8–37, 13 for KRT-R, and 19 for hIAPP S20G 8–37 (Supplementary Figs. 13–17). Collectively, these findings reveal the wide distribution of cross-β-strand linkers in diverse β-sheet assemblies, implying their structural and functional significance across protein aggregation systems. We note that STM imaging can only detect peptide segments within approximately 1 nm of the HOPG substrate, owing to the exponential decay of electron tunneling probability with distance, while regions beyond this range remain invisible. Consequently, STM primarily resolves molecular features in close proximity to the surface. Importantly, STM does not rely on three-dimensional reconstruction but instead directly visualizes the distribution of electronic states on the two-dimensional surface. Detailed discussion is provided in Supplementary Note 1.3.
Cross-β-strand linkers in amyloidal aggregates promote the structural order
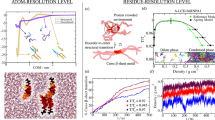
The mortise-and-tenon-like cross-β-strand linkers may function as hierarchical stabilizers that modulate the assembly order of β-sheet aggregates. To quantitatively characterize the structural order of the aggregate morphologies, we conducted the short-time Fourier transform (STFT) analysis of the STM images. The original STM images were processed through binary segmentation to generate 500 × 500-pixel square regions, which were subsequently analyzed via the STFT with a 10-pixel sampling interval (Fig. 4a, Supplementary Figs. 18 and 19). The periodic stripe patterns of β-sheet aggregates manifest as distinct frequency-domain features in the power spectral density (PSD) image obtained via the STFT: (1) the orientation angle (θ) reflecting the directionality of β-sheet alignment, and (2) the spatial frequency (q) quantifying the reciprocal of periodicity (i.e., width/spacing) of β-sheets. For example, the STFT analysis of the red- and green-boxed regions in the STM image revealed distinct local frequency-domain characteristics, with corresponding (q, θ) values of (0.02 nm−1, 45°) for the red-boxed region and (0.01 nm−1, 63.4°) for the green-boxed region (Fig. 4a). Through systematic spatial windowing, color-coded parameter maps were generated to visualize the morphometric value distributions of θ and q across discrete window positions (Fig. 4b, c).
a The upper panel shows one typical binarized STM image for STFT analysis. Scale bar: 5 nm. In the STFT analysis, the window is placed at strictly defined positions, represented by a grid of red dots. Two arbitrary window positions are marked with red and green squares. The lower panel displays the PSD calculated in Cartesian coordinates for the images within the red and green windows. The black dots represent spectral features, where the radial distance (q) indicates the spatial frequency of the β-sheet arrangement, while the angular distance (θ) corresponds to the β-sheet orientation. b Heatmaps of the radial distance q and c angular distance θ of β-sheet arrangement in gal-3 225–250. d Distribution plots of normalized θ (θi−θmin) and e along with the distribution of q values obtained from the windows in the STM images. f Comparison of q values and normalized θ values obtained from STFT analysis of STM images for different samples, where the x-axis represents the CV of normalized θ values, the y-axis displays the CV of q values, the size of the circles represents the proportion of cross-β-strand linker in the aggregates, and different colors represent different samples, Gray dashed circle indicates absence of cross-β-strand linker in the sample. Source data are provided as a Source data file.
Quantitative assessment of morphometric parameters indicates the spatial heterogeneity in aggregate structural ordering. Gal-3 225–250 β-sheets exhibit a multimodal θ distribution with two main peaks at 54° and 107°, yielding a coefficient of variation (CV) of 0.54 (Fig. 4d). Meanwhile, the q value displays a distribution ranging from 0.02 nm−1 to 0.28 nm−1, peaking at 0.12 nm−1 with a CV of 0.29 (Fig. 4e). Correlation analysis between cross-β-strand linker frequency and morphometric parameters shows a clear dependence on peptide type. Circular visualization employs diameter encoding for cross-β-strand proportion and chromatic coding for sample discrimination, with gray dashed circle denoting linker-negative aggregates (Fig. 4f). Notably, the frequency of cross-β-strand linkers exhibits a significant negative correlation with the CV of q (Pearson’s correlation coefficient = −0.7, P = 0.03), suggesting that these linkers promote ordered packing in β-sheet periodic assemblies (Supplementary Fig. 20). Conversely, no statistically significant association can be drawn between linker density and the CV of θ.
Discussion
Our study identifies a cross-β-strand linking mechanism that establishes structural connectivity between adjacent β-sheets through the mortise-and-tenon-like interlocking. Unlike the three previously reported interaction modes, this mechanism generates direct inter-sheet connections through the insertion of β-strands between neighboring β-sheets. Notably, this mechanism has been observed only in β-sheet-rich amyloid aggregates, and whether it could occur in non-amyloid protein assemblies remains to be further explored. These cross-β-strand linkers, constituting less than 15% of total β-strands, function as critical determinants of amyloid fibril architecture. Despite their low abundance (2.1–14.2%), cross-β-strand linkers account for a disproportionately high proportion (17.3–62.0%) of the intermolecular interactions in β-sheet assemblies, acting as molecular adapters that modulate inter-β-sheet packing.
Cross-β-strand linkers exist as ensembles of heterogeneously conformational substates (9–25 residues long). This conformational plasticity may enable aggregates to balance order and disorder, facilitating the dynamic regulation of assembly pathways while maintaining ordered morphologies. Through diverse and dynamic interactions with multiple conformational substates, cross-β-strand linkers may serve as molecular connectors that integrate multiple assembly intermediates into functional fibrils. Quantitative analysis reveals that cross-β-strand linkers play a pivotal role in reducing structural entropy during β-sheet assembly. The β-strand donation mechanism observed by STM occurs specifically in pathological protein or peptide aggregates, rather than in natively folded proteins. There is good reason to assume that these aggregates naturally exist in vivo and drive pathological processes (Supplementary Note 1.4). This structural plasticity may represent biologically relevant states. Our results demonstrate that aggregates with a higher content of cross-β-strand linkers can better resist thermal stress and chemical denaturants (Supplementary Figs. 21–23), thereby enabling amyloid aggregates to maintain their stability and aggregation propensity under physiological or pathological conditions (Supplementary Note 1.5). These findings make the classical hierarchical assembly model more complete by demonstrating that rare inter-sheet connectors are also essential architectural regulators in assembly. Future studies are directed to elucidate the mechanistic basis of these interactions and their biological implications for protein misfolding diseases.
Methods
Reconstitution of peptide assemblies in solution
The peptides were obtained from Anhui Guoping Pharmaceutical Co., Ltd, after synthesis and purification, with a confirmed purity of over 98% through high-performance liquid chromatography and mass spectrometry analysis. All lyophilized peptide powders were initially dissolved in 1,1,1,3,3,3-hexafluoro-2-propanol (HFIP, Innochem), a fluorinated solvent known for efficient disruption of secondary structures. After complete evaporation of HFIP, gal-3 129–154, gal-3 206–231, and gal-3 225–250 were re-dissolved in 30 mM Tris buffer (pH 7.2) containing 2% (v/v) DMSO at a final concentration of 400 μM. The solutions were incubated at 37 °C with shaking at 210 rpm for 36 h. Aβ42, hIAPP 8–37, hIAPP S20G 8–37, −5Aβ42, and TDP-43341-357 were dissolved in Milli-Q water containing 2% (v/v) DMSO to a final concentration of 40 μM. These samples were incubated under the same conditions (37 °C, 210 rpm) for 72 h. KRT-R was dissolved in Milli-Q water containing 2% (v/v) DMSO to a final concentration of 1 mM and incubated at 37 °C with shaking at 210 rpm for 48 h. hIAPP and hIAPP S20p were re-dissolved in 100 μM TEA (pH6.0) containing 2% (v/v) DMSO to a final concentration of 500 μM, and incubated at 37 °C with shaking at 210 rpm for 72 h17,21,41,52.
Time-lapse ThT fluorescence assays for monitoring peptide aggregation
To monitor the aggregation kinetics of peptides, a ThT fluorescence assay was performed. ThT stock solution was formulated in deionized water and passed through a 0.22 μm membrane filter to eliminate particulate matter. The final reaction mixtures (200 μL total volume) contained 50 μM ThT and one of the following peptides: 400 μM gal-3 129–154, 400 μM gal-3 206–231, 400 μM gal-3 225–250, 40 μM Aβ42, 40 μM hIAPP 8–37, 1 mM KRT-R, 40 μM hIAPP S20G 8–37, 40 μM -Aβ42, or 40 μM TDP-43341–357.
The gal-3 129–154, gal-3 206–231, and gal-3 225–250 peptides were dissolved in 30 mM Tris buffer (pH 7.2), while the remaining peptides were dissolved in Milli-Q water. Each mixture was transferred to black, clear-bottom 96-well plates and sealed with optically clear adhesive film to prevent evaporation during incubation. Fluorescence readings were acquired using a Synergy H1 microplate reader with an excitation wavelength of 450 nm and an emission wavelength of 485 nm. The plates were incubated at 37 °C with moderate shaking, and fluorescence intensity was measured hourly for 48 h. Control samples lacking peptide were prepared under identical conditions to account for background ThT fluorescence.
Peptide concentration determination
The concentration of the peptide in a 6 M guanidinium chloride solution was quantified through UV-Vis spectroscopic analysis (PerkinElmer, USA). The peptide concentration was calculated using the Beer-Lambert equation53:
where A represents the UV absorbance at 280 nm, ε is the molar extinction coefficient of the peptide, L is the optical path length of the cuvette, and c is the peptide concentration.
TEM measurements
For TEM analysis, 10 μL of the assembled peptide solution was dropped onto a 200-mesh formvar carbon-coated copper grid and incubated at room temperature for 2 min. After removing the excess solution, the sample was air-dried completely at room temperature, followed by negative staining with 1% phosphotungstic acid for 30 s. The excess stain was removed, and the grid was thoroughly dried again at room temperature. TEM imaging was then performed using a Hitachi HT7700 transmission electron microscope (Hitachi, Tokyo, Japan).
FTIR analysis of peptide samples
FTIR spectra were acquired using a PerkinElmer Spectrum One FTIR Spectrometer (Waltham, Massachusetts, USA). 40 μL aliquot of each peptide solution was deposited onto a CaF₂ window and allowed to dry completely before measurement. Spectra were recorded three times over the range of 400–4500 cm−1 at a resolution of 2 cm−1. All data were processed using PeakFit software (Version 4.12, San Jose, CA, USA). Each spectrum was baseline-corrected, smoothed using a Gaussian instrumental response function, and subjected to second-order derivative calculation, followed by peak fitting on the derivative spectra. Figures of the processed spectra were generated using Prism software (Version 8.01, San Diego, CA, USA).
STM-based single-molecule imaging
For single-molecule imaging experiments using STM, 10 μL of peptide solution was applied to the surface of freshly cleaved HOPG and incubated at 37 °C for 20 min. Following incubation, the excess peptide solution was removed from the substrate in an ambient environment. Single-molecule imaging was performed using a Nanoscope IIIa SPM system (Bruker, USA) with STM tips made from Ir/Pt wires (20/80). To ensure reproducibility, each imaging experiment was repeated with different STM tips and samples. To analyze the β-strands in STM images, we used NanoScope Analysis software (version 1.9, Bruker Corporation, USA).
Interpeptide interaction analysis
The measurement of peptide strand lengths in STM images was performed using Gwyddion software (v2.31, Czech Metrology Institute, Czech Republic). Conformational substates were categorized by dividing β-strands into 3.25 Å stepwise length bins. Adjacent conformational substates were then identified from STM images. The distribution of conformational substates was quantified by counting discrete instances of each category.
Thermal stability assay
The thermal stability of 500 μM hIAPP and hIAPP S20p aggregates was assessed using the integrated UNcle stability platform (Unchained Labs, Norton, MA). Static light scattering was monitored at an excitation wavelength of 266 nm as the temperature increased from 25 to 95 °C at a rate of 0.25 °C per minute.
Chemical denaturation assay
The 500 μM hIAPP and hIAPP S20p aggregates were analyzed using a Zetasizer Nano ZS system (Malvern, UK). Corresponding volume of 6 M guanidine hydrochloride was sequentially added to the aggregates and thoroughly mixed to obtain the concentration gradient of guanidine hydrochloride. The samples were then irradiated with a 4 mW, 633 nm laser, and the scattering intensity was measured at a 90° angle. A digital signal processor was used as a photodetector to record the scattering intensity of the samples.
2D STFT analysis
The STFT was employed to analyze periodic patterns in STM images. Raw STM images were first binarized through threshold segmentation to isolate regions of interest for subsequent analysis. A square window of size N × N (500 × 500) pixels was translated across the preprocessed image with a step size of 10 pixels to ensure full spatial coverage. For each localized subregion, the 2D discrete Fourier transform was computed as54,55:
where \(f(x,y)\) represents the binarized image, and the square window function \(w(x-{x}_{w},y-{y}_{w})\) is defined as54,55:
The PSD was calculated as54,55:
To resolve orientation-specific periodic features, the PSD was converted from Cartesian coordinates \(\left({q}_{x},{q}_{y}\right)\) to polar coordinates \(p(q,\theta )\) via54,55:
where \(q=\scriptstyle\sqrt{{q}_{x}^{2}+{q}_{y}^{2}}\) represents spatial frequency (inverse of periodicity) and \(\theta=\arctan ({q}_{y}/{q}_{x})\) denotes orientation angle. The normalized joint probability density function was derived as54,55:
Marginal distributions \(p(q)\) and \(p(\theta )\), quantifying the dominant spatial frequencies and preferential orientations of β-sheet aggregates, were obtained by integrating over the complementary variable54,55
The spatial frequency (q) and azimuthal angle (θ) distributions were visualized through heatmaps to statistically highlight dominant periodic features and their orientation preferences.
AFM imaging
Incubated peptide solutions (10 μL) were deposited on the freshly cleaved HOPG substrates, followed by air-drying, ddH₂O rinsing, and then air-drying again. Aggregate morphology was examined using a Dimension FastScan AFM (Bruker, Billerica, MA) equipped with ScanAsyst Air probes (Bruker, Camarillo, CA; resonance frequency 70 kHz, spring constant 0.4 N/m). All images were acquired at a resolution of 512 × 512 pixels and analyzed using NanoScope Analysis software.
Visualization
Scientific visualization workflows were implemented in Python, employing Pandas and NumPy frameworks for preprocessing pipelines that enforced data integrity through rigorous cleansing and format standardization prior to visualization. Visual representations were produced using Matplotlib’s computational graphic tools.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper.
Code availability
Custom codes used in this study are available via Zenodo [https://doi.org/10.5281/zenodo.17765773]56.
References
Greenblatt, J. F., Alberts, B. M. & Krogan, N. J. Discovery and significance of protein-protein interactions in health and disease. Cell 187, 6501–6517 (2024).
Gao, Z. et al. Hierarchical graph learning for protein–protein interaction. Nat. Commun. 14, 1093–1104 (2023).
Malinverni, D. & Babu, M. M. Data-driven design of orthogonal protein-protein interactions. Sci. Signal. 16, eabm4484 (2023).
Lu, H. et al. Recent advances in the development of protein–protein interactions modulators: mechanisms and clinical trials. Signal Transduct. Target. Ther. 5, 213 (2020).
Sang, P. et al. Inhibition of β-catenin/B cell lymphoma 9 protein−protein interaction using α-helix–mimicking sulfono-γ-AApeptide inhibitors. Proc. Natl. Acad. Sci. USA 116, 10757–10762 (2019).
Scott, D. E., Bayly, A. R., Abell, C. & Skidmore, J. Small molecules, big targets: drug discovery faces the protein–protein interaction challenge. Nat. Rev. Drug Discov. 15, 533–550 (2016).
Dang, L. T. et al. Receptor subtype discrimination using extensive shape complementary designed interfaces. Nat. Struct. Mol. Biol. 26, 407–414 (2019).
Borgia, A. et al. Extreme disorder in an ultrahigh-affinity protein complex. Nature 555, 61–66 (2018).
Burke, D. F. et al. Towards a structurally resolved human protein interaction network. Nat. Struct. Mol. Biol. 30, 216–225 (2023).
Stiller, J. B. et al. Structure determination of high-energy states in a dynamic protein ensemble. Nature 603, 528–535 (2022).
Gianni, S. et al. Fuzziness and frustration in the energy landscape of protein folding, function, and assembly. Acc. Chem. Res. 54, 1251–1259 (2021).
Adamcik, J. & Mezzenga, R. Amyloid polymorphism in the protein folding and aggregation energy landscape. Angew. Chem. Int. Ed. 57, 8370–8382 (2018).
Wei, G., Xi, W., Nussinov, R. & Ma, B. Protein ensembles: How does nature harness thermodynamic fluctuations for life? The diverse functional roles of conformational ensembles in the cell. Chem. Rev. 116, 6516–6551 (2016).
Berkeley, R. F. et al. Capturing the conformational heterogeneity of HSPB1 chaperone oligomers at atomic resolution. J. Am. Chem. Soc. 147, 4c18668 (2025).
Lama, D. et al. A druggable conformational switch in the c-MYC transactivation domain. Nat. Commun. 15, 1865 (2024).
Jiang, T., Thielges, M. C. & Feng, C. Emerging approaches to investigating functional protein dynamics in modular redox enzymes: nitric oxide synthase as a model system. J. Biol. Chem. 301, 108282 (2025).
Zhang, W. et al. Single-molecule visualization determines conformational substate ensembles in β-sheet–rich peptide fibrils. Sci. Adv. 9, eadg7943 (2023).
Fraser, J. S. et al. Accessing protein conformational ensembles using room-temperature X-ray crystallography. Proc. Natl. Acad. Sci. USA 108, 16247–16252 (2011).
Torchia, D. A. NMR studies of dynamic biomolecular conformational ensembles. Prog. Nucl. Magn. Reson. Spectrosc. 84–85, 14–32 (2015).
Bock, L. V. & Grubmüller, H. Effects of cryo-EM cooling on structural ensembles. Nat. Commun. 13, 1709 (2022).
Yu, L. et al. Experimental insights into conformational ensembles of assembled β-sheet peptides. ACS Cent. Sci. 9, 1480–1487 (2023).
Stanfield, R. L. & Wilson, I. A. Protein-peptide interactions. Curr. Opin. Struct. Biol. 5, 103–113 (1995).
Remaut, H. & Waksman, G. Protein–protein interaction through β-strand addition. Trends Biochem. Sci. 31, 436–444 (2006).
Bhat, A. S., Kinch, L. N. & Grishin, N. V. -Strand-mediated interactions of protein domains. Proteins Struct. Funct. Bioinform. 88, 1513–1527 (2020).
Cheng, P.-N., Pham, J. D. & Nowick, J. S. The supramolecular chemistry of β-sheets. J. Am. Chem. Soc. 135, 5477–5492 (2013).
Gulbis, J. M., Kelman, Z., Hurwitz, J., O’Donnell, M. & Kuriyan, J. Structure of the C-terminal region of p21WAF1/CIP1 complexed with human PCNA. Cell 87, 297–306 (1996).
Deshaies, R. J. SCF and cullin/RING H2-based ubiquitin ligases. Annu. Rev. Cell Dev. Biol. 15, 435–467 (1999).
Yu, X., Ge, P., Jiang, J., Atanasov, I. & Zhou, Z. H. Atomic model of CPV reveals the mechanism used by this single-shelled virus to economically carry out functions conserved in multishelled reoviruses. Structure 19, 652–661 (2011).
Korotkov, K. V., Pardon, E., Steyaert, J. & Hol, W. G. J. Crystal structure of the N-terminal domain of the secretin GspD from ETEC determined with the assistance of a nanobody. Structure 17, 255–265 (2009).
Puorger, C. et al. Infinite kinetic stability against dissociation of supramolecular protein complexes through donor strand complementation. Structure 16, 631–642 (2008).
Willbold, D., Strodel, B., Schröder, G. F., Hoyer, W. & Heise, H. Amyloid-type protein aggregation and prion-like properties of amyloids. Chem. Rev. 121, 8285–8307 (2021).
Teunissen, C. E. et al. Blood-based biomarkers for Alzheimer’s disease: towards clinical implementation. Lancet Neurol 21, 66–77 (2022).
Wilson, D. M. et al. Hallmarks of neurodegenerative diseases. Cell 186, 693–714 (2023).
Lührs, T. et al. 3D structure of Alzheimer’s amyloid-β (1–42) fibrils. Proc. Natl. Acad. Sci. USA 102, 17342–17347 (2005).
Yan, X. Peptide Self-Assembly and Engineering (Wiley-VCH, 2024).
Garcia, R. Interfacial liquid water on graphite, graphene, and 2D materials. ACS Nano 17, 51–69 (2023).
Chiu, Y.-P. et al. Liquid-liquid phase separation and extracellular multivalent interactions in the tale of galectin-3. Nat. Commun. 11, 1229 (2020).
Suthahar, N. et al. Galectin-3 activation and inhibition in heart failure and cardiovascular disease: an update. Theranostics 8, 593–609 (2018).
Goldschmidt, L., Teng, P. K., Riek, R. & Eisenberg, D. Identifying the amylome, proteins capable of forming amyloid-like fibrils. Proc. Natl. Acad. Sci. USA 107, 3487–3492 (2010).
Yu, L. et al. Molecular recognition of human islet amyloid polypeptide assembly by selective oligomerization of thioflavin T. Sci. Adv. 6, eabc1449 (2020).
Wang, R. et al. Submolecular resolution of β-sheet plasticity: decoding mutations and PTMs in protein aggregation disorders. ACS Cent. Sci. 11, 927–937 (2025).
Mao, X. et al. Beta structure motifs of islet amyloid polypeptides identified through surface-mediated assemblies. Proc. Natl. Acad. Sci. USA 108, 19605–19610 (2011).
Ma, X. et al. Amyloid β (1–42) folding multiplicity and single-molecule binding behavior studied with STM. J. Mol. Biol. 388, 894–901 (2009).
Makin, O. S. & Serpell, L. C. Structural characterisation of islet amyloid polypeptide fibrils. J. Mol. Biol. 335, 1279–1288 (2004).
Qiu, X. et al. Alkane-assisted adsorption and assembly of phthalocyanines and porphyrins. J. Am. Chem. Soc. 122, 5550–5556 (2000).
Kollmer, M. et al. Cryo-EM structure and polymorphism of Aβ amyloid fibrils purified from Alzheimer’s brain tissue. Nat. Commun. 10, 4760 (2019).
Riek, R. & Eisenberg, D. S. The activities of amyloids from a structural perspective. Nature 539, 227–235 (2016).
Yang, Y. et al. Cryo-EM structures of amyloid-β 42 filaments from human brain. Science 375, 167–172 (2022).
Garrett, R. H. & Grisham, C. M. Biochemistry 2nd edn (Saunders College Publishing, 1999).
Nelson, R. et al. Structure of the cross-β spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Chiti, F. & Dobson, C. M. Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu. Rev. Biochem. 86, 27–68 (2017).
Jian, Z. et al. Structural evolution of the conformational ensembles in peptide fibrillar aggregates. Mater. Today Phys. 44, 101437 (2024).
Gill, S. C. & Von Hippel, P. H. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182, 319–326 (1989).
Ciężar, K. & Pochylski, M. 2D fourier transform for global analysis and classification of meibomian gland images. Ocul. Surf. 18, 865–870 (2020).
Ciężar, K. & Pochylski, M. 2D short-time fourier transform for local morphological analysis of meibomian gland images. PLoS ONE 17, e0270473 (2022).
BoyceL. BoyceL/STM_PostProcess: First Version. Code related to the paper, available at https://doi.org/10.5281/zenodo.17765773 (2025).
Acknowledgements
The work is supported by the National Natural Science Foundation of China (32471451 Chenxuan Wang, 92353302 Chenxuan Wang, 32201243 L.Y., 32201142 W.Z.), the Beijing Nova Program (20240484521 L.Y.), the CAMS Innovation Fund for Medical Sciences (2025-I2M-KJ-010 Chenxuan Wang, 2025-I2M-XHXX-076 L.Y.), the Fundamental Research Funds for the Central Universities (3332024154 S.M.), State Key Laboratory Special Fund (2060204 Chenxuan Wang). The authors thank the National Center for Nanoscience and Technology, China, for STM facilities. The authors also thank State Key Laboratory of Common Mechanism Research of Major Diseases Platform and the Core Laboratories at the Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences & Peking Union Medical College for consultation and instrument availability that supported this work. M.W. acknowledges the support of the Global Research Assistant Professor Scheme of the CityUHK.
Author information
Authors and Affiliations
Contributions
Chenxuan Wang and L.Y. conceived the project and designed the experiments. R.W., S.M., Z.J., and F.Z. performed the STM tests. S.M., R.W., Z.J., M.L., and Z.D. analyzed the STM data. S.M. and R.W. performed the ThT assay, FTIR, and TEM tests. M.W. and M.L. also contributed to STM and FTIR measurements and analysis. S.M. and L.Y. performed AFM imaging. S.M. collected most of the data in the revision stage. Chen Wang and Y.Y. provided the STM facilities and contributed to data analysis. W.Z. contributed to data analysis. S.M., R.W., M.W., L.Y., and Chenxuan Wang wrote the manuscript with inputs from all the authors. Chenxuan Wang and L.Y. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Antonino Natalello and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mo, S., Wang, R., Jian, Z. et al. Atypical β-strand insertion mediates the noncovalent cross-linking in amyloid aggregates. Nat Commun 17, 1449 (2026). https://doi.org/10.1038/s41467-025-68185-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68185-3






