Abstract
Type 2 diabetes (T2D) exhibits clinical heterogeneity, yet most existing classification models are derived from European populations and face challenges in clinical application. Here, we evaluate the generalizability of a tree-like graph structure from Scottish data to 32,501 newly diagnosed T2D patients from a multi-center Chinese cohort comprising over 8.6 million individuals. We observe similar distribution between the Scottish and Chinese individuals in heart and kidney outcomes, but diabetic retinopathy varies across ancestries even within similar phenotypes. To capture T2D Chinese-specific heterogeneity, we apply a variational autoencoder (VAE) framework to identify key clinical features and construct a tree structure using the Discriminative Dimensionality Reduction Tree (DDRTree) algorithm. This Chinese tree model is validated in two independent external cohorts and revealed longitudinal phenotypic shifts trending toward higher-risk branches. Our findings emphasize the need for population-specific classification frameworks to advance precision diabetology through individualized risk prediction and specialized treatment guidelines.
Similar content being viewed by others
Introduction
Type 2 diabetes (T2D) is a highly heterogeneous disease1. The current classification of diabetes proposed by the World Health Organization, based on hyperglycemia, pancreatic autoimmune antibodies and limited clinical manifestations, roughly classifies diabetes into main types as type 1 diabetes, T2D, et cetera2. For T2D, such classification does not capture the differences observed in clinical manifestations, clinical course and the impact of ethnicity3,4.
Many efforts have been made to re-classify T2D5. Ahlqvist et al. and several studies classify T2D into subgroups using a k-means clustering method with clinically accessible variables at diagnosis and demonstrate that these subgroups are associated with varied risks of complications and responses to antidiabetic regimens6,7. However, the studies replicating such a clustering method in different populations often yielded different subgroups, suggesting that racial disparities should be considered when re-classifying T2D8,9. Also, most of these studies have adopted insulin or C-peptide at diagnosis as clustering variables, which has limited the clinical use of such models as these variables are not always available in primary care settings and often need standardization of their test methods.
More importantly, forcing discrete subtypes may overlook the continuous nature of disease heterogeneity, limiting predictive utility, especially for drug response10. To address this, a recent study has demonstrated that the heterogeneity in T2D can be represented by a tree-like graph structure by using reversed graph-embedded dimensionality reduction of nine simple clinical variables—glycated hemoglobin (HbA1c), body mass index (BMI), total cholesterol (TC), high-density lipoprotein cholesterol (HDL-C), alanine aminotransferase (ALT), creatinine (Cr), systolic blood pressure (SBP), triglycerides, and diastolic blood pressure (DBP)—with position on the tree predicting the long-term probability of incident diabetic complications11. This “Scottish Tree” model enables each individual with T2D to be projected onto a low-dimensional tree-shaped manifold, in which their position correlates with long-term risk of multiple diabetes-related complications including microvascular and macrovascular diseases. A follow-on study mapped deeply phenotyped patients with T2D from the German Diabetes Study (GDS) and the LUdwigshafen RIsk and Cardiovascular Health (LURIC) cohort onto the Scottish tree12. This study demonstrates the pathophysiological foundations of the model, such as insulin resistance and β-cell function that vary across the tree, and offers more granular outcomes, including liver fat and heart failure (HF)12. Compared with rigid subtypes, the tree structure offers a continuous, individualized approach that may better capture disease biology and therapeutic needs. This flexible framework has also been applied to type 1 diabetes and prediabetes, where it enables the delineation of heterogeneous disease trajectories and informs the design of early interventions tailored to specific risk patterns13,14.
However, the selection of features used to construct the tree in these studies was based solely on nine clinically observable variables related to T2D presentation, and it remains unclear whether these features are the most informative or generalizable for capturing disease heterogeneity, particularly in non-European populations. Notably, Asians tend to develop T2D at a lower BMI, often driven by reduced β-cell function rather than prominent insulin resistance, and show a higher propensity for visceral adiposity and ectopic fat accumulation despite lower overall adiposity levels15. To overcome these limitations, we introduce the Variational Autoencoder (VAE), a deep generative model that learns probabilistic latent representations from high-dimensional data by encoding and reconstructing the input through a non-linear manifold structure, which can also facilitate dimensionality reduction and feature extraction16. VAE employs a learned encoder as a feature extractor to generate a latent representation that captures intrinsic data characteristics17. While the learned features are not specifically optimized for downstream predictive modeling tasks, they can still reveal meaningful patterns and help identify clinically relevant variables, particularly in complex datasets. Given the observed disparities in T2D subgroups across different ethnicities, incorporating such deep learning approaches may uncover population-specific features and phenotypic patterns. This highlights the need to investigate ancestry-specific characteristics to gain deeper insights into the underlying drivers of T2D heterogeneity.
In this work, we perform a two-step study in a nationwide electronic health record (EHR)-based cohort of Chinese patients with approximately 613,594 people with T2D. First, we map the Chinese T2D population to the published Scottish tree-like graph to compare the gradient of the included variables and distribution diabetic complications risks. Then we construct a Chinese population-specific tree-like graph: using the VAE framework to select variables, map and estimate the risks and patterns of complications, and validate these findings in independent external Chinese cohorts.
Results
Baseline characteristics of the cohort
We evaluated the disease phenotypes and progression patterns of T2D among Chinese individuals, using the Chinese Renal Disease Data System (CRDS), a comprehensive multicenter retrospective database18 as our primary data source to construct our study population (Fig. 1). It collected inpatient and outpatient data from the electronic medical record systems of 24 medical centers across China from January 1, 2000, to May 26, 2024. The data included all the patients whose data is accessible from the participating centers’ medical record systems, not limited to those who had renal diseases. From the CRDS cohort of 8,632,000 patients, we identified 32,501 individuals who were diagnosed with T2D (ICD-10 codes E11) within 12 months of the index day (Supplementary Fig. 1). To assess T2D heterogeneity, we first mapped individuals onto the previously established Scottish diabetes tree based on nine clinical variables (Fig. 1a), and then constructed a Chinese-specific tree using VAE and machine learning-based feature selection (Fig. 1b), with internal and external validation confirming its robustness (Fig. 1c). Among these 32,501 T2D patients, 9816 had records for one-year follow-up, and 1821 had five-year follow-up data. We observed a worsening trend in lipid-related phenotypes, which including HDL-C, triglycerides, and TC, as well as kidney function markers, such as estimated glomerular filtration rate (eGFR) and Cr, with increasing follow-up duration (Table 1).
a We first extracted a dataset of 32,501 newly diagnosed individuals withT2D from the CRDS, a large-scale electronic health record database comprising over 8.6 million individuals with detailed demographic, laboratory, diagnostic, and surgery data. To evaluate disease progression and heterogeneity, we initially performed model adaptation by mapping each individual to a previously defined “Scottish tree” of diabetes subgroups based on 9 clinical variables: HbA1c, BMI, total cholesterol (TC), HDL-C, alanine aminotransferase (ALT), creatinine (Cr), systolic blood pressure (SBP), triglycerides, and diastolic blood pressure (DBP). b To identify the most informative clinical features for defining T2D subtypes in the Chinese population, we applied a Variational Autoencoder (VAE) model feature selection framework. The process resulted in 10 key variables: HDL-C, triglycerides, SBP, ALT, HbA1c, LDL-C, Cr, heart rate, DBP, and BMI. These original 10 variables were used for constructing a Chinese-specific diabetes tree. c The Chinese tree was developed using 80% of the CRDS T2D cohort as a training set and validated internally in the remaining 20%. External validation was performed using two independent cohorts: the Joint Asia Diabetes Evaluation (JADE) registry and the DR cohort. Consistency of phenotypic and complication profiles across cohorts confirmed the robustness of the Chinese tree. An interactive web-based tool was also developed for clinical application and research usev (https://wenglab-t2d-phenotype.shinyapps.io/wenglab-t2d-phenotype/). (Created in https://BioRender.com).
Mapping the Chinese individuals onto the Scottish tree model
The 32,501 individuals were mapped onto the Scottish tree using nine baseline clinical characteristics (HbA1c, BMI, TC, HDL-C, ALT, Cr, SBP, triglycerides, and DBP) described by Nair et al. 11. We assessed the distribution patterns of phenotypes across the tree by regressing each phenotype against the two tree dimensions: triglycerides had the greatest influence on tree coordinates, while HbA1c had the weakest (Supplementary Fig. 2). We allocated the 32,501 individuals to the Scottish tree using a two-step mapping function: first, predictive models estimated the X and Y coordinates for each Chinese individual; second, a distance-based algorithm was applied to calculate the Euclidean distance between each Chinese individual and all nodes in the two-dimensional latent space of the Scottish reference tree. As expected, since the mapping was based on the Scottish reference, the resulting phenotype distribution among Chinese individuals mirrored that of the original Scottish tree (Fig. 2a). However, BMI differences caused substantially fewer Chinese individuals to map to the right side of the tree. Other phenotypes were overlaid: Lower C-peptide levels were in the left-upper part (Fig. 2b), eGFR levels were higher in the upper-left corner (Fig. 2c), and liver indices (Fig. 2d, e) showed similar patterns to the GDS mapping12.
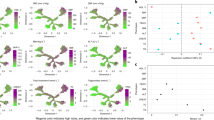
a Mapping nine phenotypic variables adjusted for age and sex onto the Scottish tree structure (n = 32,501). The gray color represents the original tree derived from the Scottish cohort (reference), while magenta indicates higher values and green indicates lower values of the phenotypes from the CRDS dataset. b Fasting C-peptide and serum C-peptide levels used to evaluate insulin secretion function (fasting C-peptide, n = 2510; serum C-peptide, n = 3285). c eGFR at baseline (n = 32,501). d AST/ALT ratio marker for steatotic liver disease (n = 32,501). e FIB-4 marker for liver fibrosis (n = 32,501).
We then examined how baseline phenotypic variation translated into the risk of 13 diabetes-related outcomes, and the definitions of these clinical outcomes are provided in Supplementary Table 1. These included cardiovascular events such as ischemic stroke, hemorrhagic stroke, myocardial infarction (MI), and HF, including both HF with preserved ejection fraction (HFpEF) and reduced ejection fraction (HFrEF). Renal outcomes included chronic kidney disease (CKD) and end-stage renal disease (ESRD). Liver-related outcomes comprised cirrhosis and metabolic dysfunction-associated steatotic liver disease (MASLD). In addition, we assessed the risk of diabetic retinopathy (DR) and diabetic peripheral vascular disease (DPVD). For each outcome, we applied Fine–Gray competing risk models, using death as a competing event. Ten-year event probabilities were estimated for each individual based on their coordinates in the two-dimensional tree space. These predicted risks were then visualized across the Chinese diabetes tree to explore spatial variation in disease progression (Supplementary Fig. 3). The probability of cardiovascular outcomes, kidney outcomes and DPVD resembles those in the Scottish11 and the GDS populations12 (Fig. 3a, b, f–h, Supplementary Fig. 3). Notably, DR risk in the Chinese population appeared to be significantly different, with higher risk in the upper-left driven by elevated HDL-C, fibrosis-4 (FIB-4) index, and Aspartate Aminotransferase to Alanine Aminotransferase Ratios (AST/ALT ratios), contrasting with the Scottish cohort’s upper-right concentration (Fig. 3c). For liver diseases, cirrhosis concentrated on the left (Fig. 3d), whereas MASLD risk aligned with cardiovascular outcomes concentrated on the right part of the tree (Fig. 3e). Additionally, we provide estimates for two types of strokes: Stroke probabilities were highest in the upper-right branch, associated with elevated baseline blood pressure and hyperglycemia, hemorrhagic and ischemic stroke showed similar patterns (Fig. 3g). Interestingly, individuals in lower branches with higher BMI had an increased risk of HFrEF, while HFpEF resembled the distribution pattern of stroke (Fig. 3h). To assess the distribution of event probabilities, we report the sub-hazard ratios (sHRs) from the competing risk models constructed with linear DDRTree dimensions (Supplementary Fig. 3).
a–h Predicted 10-year probability of complications from T2D (n = 32,501): a chronic kidney disease (CKD, 1222 events), b end-stage renal disease (ESRD, 216 events), c diabetic retinopathy (DR, 1856 events), d cirrhosis (911 events), e metabolic dysfunction-associated steatotic liver disease (MASLD, 8240 events), f myocardial infarction (MI, 213 events), g stroke (2187 events), including hemorrhagic stroke (139 events) and ischemic stroke (1984 events), and h heart failure (HF, 1333 events), further classified into HF with reduced ejection fraction (HFrEF, 97 events) and HF with preserved ejection fraction (HFpEF, 344 events) (left ventricular ejection fraction data available: n = 4902).
Overall, while many features and diabetes outcomes overlap between Chinese and Scottish populations, differences in baseline characteristics, such as BMI, led to mismatched tree distributions. T2D-related complications, particularly DR, exhibited varying ten-year incident probabilities. These findings suggest that a tree tailored to Chinese clinical features would enhance diabetes progression prediction in this population.
Development and performance of the Chinese tree model
We developed a feature selection framework integrating VAE and machine learning to identify the most relevant features of T2D, followed by the construction of the Chinese tree using the DDRTree algorithm. The VAE model was trained using data from 45,589 individuals with type 2 diabetes (T2D) and 11,340 healthy individuals (without diabetes). Based on 50 clinical variables (listed in Supplementary Table 2), dimensionality reduction was performed using the VAE framework. To identify the optimal hyperparameters that best captured the underlying structure of the data while maintaining generalizability to unseen individuals, we evaluated both the reconstruction accuracy of the input data and the model’s stability across multiple refitting procedures. The median reconstruction accuracy was 0.8439 on average, and the final models demonstrated high stability when retrained five times, with an average change in cosine similarity within the latent space of 0.015. These results suggest that the VAE models reliably reconstructed the input features with minimal variability across training iterations (Supplementary Fig. 4). To evaluate how well the model captured the structure of the clinical data, we examined the weights connecting the input features to the encoder of the VAE. The analysis revealed that many of the top 15 most influential features were related to cardiovascular, hepatic, renal, and metabolic functions (Fig. 4a). Similar findings were obtained through an adaptation of Shapley Additive exPlanations (SHAP) analysis, which assessed the contribution of each continuous feature to individual positioning within the latent space. Based on the integrated results of the VAE and SHAP analyses, we identified the overlap 10 most important features: HDL-C, triglycerides, SBP, ALT, HbA1c, LDL-C, Cr, heart rate, DBP, and BMI (Fig. 4a).
a Feature importance was estimated from the absolute weights connecting each input variable to the hidden layer of the VAE encoder. SHAP-based perturbation analysis assessed the influence of each input feature on the latent representation by measuring the change in encoded outputs when setting each feature to missing across all individuals. Ten features were consistently identified by both methods (highlighted in red), indicating robust importance across complementary evaluation strategies. b Linear regression coefficients with 95% confidence intervals (CI), estimating associations between tree dimensions and ten clinical phenotypes (n = 26,000). c Chinese Tree visualization from the CRDS validation set (n = 26,000), colored by phenotype values (red = high, blue = low). d Fasting and serum C-peptide levels for insulin secretion assessment (fasting, n = 2012; serum, n = 2633). e, f Distribution of eGFR, AST/ALT ratio, and FIB-4 score (n = 26,000).
To validate the biological and predictive relevance of the ten features jointly selected by VAE-based weighting and SHAP value analysis, we applied multiple machine learning classifiers to evaluate their performance in discriminating T2D status. We first trained several models using all input features and identified the Gradient Boosting Classifier (GBC) as the best-performing model (Supplementary Fig. 5). To evaluate the discriminative power of the VAE-selected features, we compared model performance using all available variables versus the top 5 to 10 ranked variables. The Gradient Boosting model achieved an area-under-curve (AUC) of 0.926 using all features, which remained comparable (AUC = 0.916) when restricted to the 10 variables, with similarly high Precision-Recall AUC(PR-AUC) and F1 scores (Supplementary Fig. 5). These results suggest that the ten selected features are sufficient to capture the key characteristics of T2D in our cohort. Among the tree’s underlying variables, triglycerides had the greatest influence on the tree coordinates, followed by DBP, ALT, HDL-C, LDL-C, Cr, heart rate, BMI, and SBP, HbA1c showed the smallest effect size (Fig. 4b). Then we divided the cohort of 32,501 individuals into an 80% training set (26,000 individuals) for constructing a “Chinese Tree” based on the top ten selected variables and a 20% validation set (6501 individuals). In the Chinese Tree, individuals located in the upper-left corner exhibited high HDL-C and LDL-C levels, low BMI, and ALT levels. Those in the lower-left corner were characterized by higher obesity, with elevated SBP and DBP, while individuals in the lower-right corner showed higher triglycerides and Cr levels (Fig. 4c). For C-peptide levels (Fig. 4d), the lowest values were observed in the top right of the tree, whereas kidney function marker eGFR (Fig. 4e), the AST/ALT ratio (Fig. 4f), and liver fibrosis assessed by the FIB-4 index (Fig. 4g) showed the highest values in the same region.
We defined the branches of the Chinese diabetes tree based on the spatial coordinates within the tree. The distribution of MASLD, CKD, and ESRD were comparable, with elevated probabilities concentrated in branch 5 (Fig. 5a, b, e). When looking at the complications in the Chinese tree, the highest risk of DR was observed in branch 1, primarily driven by elevated HDL-C levels and increased AST/ALT ratio (Fig. 5c). Cirrhosis cases were more concentrated in branches 3 and 4, accompanied by lower baseline C-peptide and higher HDL-C (Fig. 5d). The probability of cardiovascular outcomes, including MI and HF, was elevated in branches 4 and 5, especially among individuals with obesity, high blood pressure, and reduced eGFR (Fig. 5f, h). Notably, individuals in branches 4 to 6, characterized by the highest BMI, exhibited an increased risk of HFrEF, mirroring the distribution of MI, while the pattern of HFpEF resembled stroke, with elevated SBP and DBP predominantly in branches 1, 5, and 6 (Fig. 5h). Stroke incidence was higher in branches 1, 5, and 6, which featured the highest baseline SBP, DBP and hyperglycemia; both hemorrhagic and ischemic stroke displayed similar patterns (Fig. 5g). Individuals in branches 1, 5, and 6 also exhibited a higher risk of DPVD, paralleling the stroke distribution (Supplementary Fig. 6). The sHRs estimated from competing risk models constructed using linear DDRTree dimensions further quantified these associations, as detailed in Supplementary Fig. 6.
a–h Predicted 10-year probability of T2D complications (n = 26,000): a chronic kidney disease (CKD, 1005 events), b end-stage renal disease (ESRD, 180 events), c diabetic retinopathy (DR, 1503 events), d cirrhosis (740 events), e metabolic dysfunction-associated steatotic liver disease (MASLD, 6593 events), f myocardial infarction (MI, 171 events), g stroke (1766 events), including hemorrhagic stroke (118 events) and ischemic stroke (1608 events), and h heart failure (HF, 1063 events), further classified into HF with reduced ejection fraction (HFrEF, 79 events) and HF with preserved ejection fraction (HFpEF, 276 events) (left ventricular ejection fraction data available: n = 3897 events).
Validation of the Chinese tree
To test the model’s validity, we developed a mapping function to project the 20% validation set, and two external cohorts, separately, onto this Chinese tree using age at diagnosis, sex, and the ten clinical features selected in model development (Supplementary Fig. 7). The results from the 20% validation set, as the internal validation showed consistency with the training set in both phenotype and complication risk estimation, as well as the regression test (Supplementary Fig. 8).
The JADE cohort was extracted from the Joint Asia Diabetes Evaluation (JADE)19,20 program, which originated in Hong Kong SAR, China, where the healthcare system is substantially different from that of the CRDS centers. The JADE external validation cohort included 2328 Chinese individuals diagnosed with T2D selected within the same registering period as the CRDS cohort (Supplementary Table 3). As external validation, we applied the tree models in the JADE cohort, and also had findings consistent with the CRDS cohort (Supplementary Fig. 9). Overall, the performance of the Chinese Tree in the JADE cohort corroborated that in the CRDS: We compared the distribution patterns of key clinical outcomes between the CRDS and JADE cohorts. Similar distribution patterns were observed for HF and MI across both cohorts, whereas the patterns of stroke and ischemic stroke differed. HFpEF showed a distribution pattern similar to stroke in the CRDS cohort. In contrast, within the JADE cohort from the Hong Kong healthcare system, HF and stroke shared similar distribution patterns, while MI and ischemic stroke clustered similarly. For liver cirrhosis, patients in both the CRDS and JADE datasets were concentrated in branches 1 and 3. ESRD and CKD were concentrated in branches 5 and 6 in both cohorts. These differences likely reflect variations in patient characteristics and healthcare delivery systems, underscoring the robustness yet adaptability of our model across diverse populations (Fig. 6a).
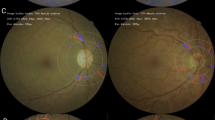
a Visualization of outcomes associated with T2D in the Chinese Tree based on the JADE validation dataset. Each point represents a hazard ratio (HR), and horizontal error bars indicate the corresponding 95% confidence intervals (CI). Predicted 10-year probabilities and HRs with 95% CI of myocardial infarction (MI, n = 28), heart failure (HF, n = 32), stroke (n = 50), cirrhosis (n = 4), chronic kidney disease (CKD, n = 250), and end-stage renal disease (ESRD, n = 38) following T2D diagnosis are shown for the JADE validation dataset. b Predicted probability and hazard ratios (HRs) with 95% CI of DR (87 events) in the DR cohort (n = 1006) on the Chinese Tree. Each point represents a hazard ratio (HR), and error bars denote the 95% confidence intervals. c Sensitivity analysis excluding patients with hepatitis at baseline in the CRDS cohort, FIB-4 for liver fibrosis and AST/ALT ratio marker for steatotic liver disease (n = 24,186). d, e Predicted probability of MASLD (6097 events) and cirrhosis (220 events) at 10 years from the diagnosis of T2D excluding patients with hepatitis (n = 24,186). f Sankey diagram showing the 5-year trajectory of changes classified into seven groups based on the Chinese tree coordinates. Seven groups were manually defined based on approximate regions in the tree coordinates to facilitate visualization of potential transitions.
In addition, we included another independent cohort of 1006 Chinese patients with T2D, identified from EHR in three medical centers with available DR data (DR cohorts), The baseline characteristics of this cohort are shown in Supplementary Table 4. We tested the DR risk predicting performance of the Chinese tree using the DR cohort and found largely consistent outcomes with that of the CRDS cohort (Supplementary Fig. 10). Validation using the DR cohort confirmed this distinct pattern of DR risk in the Chinese population with T2D (Fig. 6b). All this finding indicates that the Chinese tree model demonstrates robustness performance across diverse healthcare systems and medical centers.
Sensitivity analysis of liver-related risks
Furthermore, the high prevalence of MASLD and cirrhosis in our cohort may partly explain the clustering of certain complications such as DR and CVD in liver-related phenotypic branches. As shown in Supplementary Table 5, liver disease commonly co-occurs with a wide range of diabetes complications, including stroke, HF, and DR. These population-specific comorbidity patterns highlight the importance of accounting for liver disease burden when interpreting phenotypic structures in Chinese individuals with T2D. One major cause of liver fibrosis in China is viral hepatitis. To rule out the impact of viral hepatitis-related cirrhosis, we performed a sensitivity analysis in the CRDS cohort, excluding 1814 patients (6.97%) with viral hepatitis at baseline. After the exclusion, no distinct clustering of FIB-4 was observed, the AST/ALT ratio remained primarily concentrated in the first branch as well (Fig. 6c). But concentration of cirrhosis and MASLD stayed in their positions as in the main analysis (Fig. 6d, e). After excluding populations with alcoholic cirrhosis and acute pancreatitis, the distribution of T2D patients with high-risk cirrhosis remained in the third and fourth branches, while patients with high-risk MASLD located in the fifth and sixth branches, reinforcing the association of MASLD with T2D phenotyping in the Chinese tree (Supplementary Figs. 1112).
Migration of individuals within the tree model
Studies suggested that phenotypes of T2D are subject to change due to metabolic control and pharmacotherapy1,3. Given the previously reported 5-year clinical characteristics of T2D patients in our cohort, we further conducted trajectory analyses to evaluate how individual phenotypes evolved over time. We analyzed the follow-up data of the CRDS cohort to investigate such phenotype change in T2D (Supplementary Fig. 13). Five years after diabetes diagnosis, 32.83% of Chinese participants (598 out of 1821, drawn from a total cohort of 32,501 individuals) shifted their position across the Scottish tree, with movements primarily toward the corners—particularly the bottom right—indicating a higher likelihood of developing obesity, liver damage, and heart disease. In the Chinese tree, 6.45% (95 out of 1472, from 26,000 Chinese individuals) showed a shift (distance > 1) from their baseline position. Individuals with cardiovascular and renal disease exhibited the least phenotypic change. After a 5-year follow-up, approximately 3% of individuals with T2D moved out from the central part of the tree in both the Scottish and Chinese trees, suggesting increased disturbances in metabolic indices, poorer cardiac and renal function (Supplementary Fig. 14). We further explored the movement of patients distributed in the middle branches to other groups and found that, regardless of whether in the Scottish tree or the Chinese tree, the number of patients with poorer cardiac and renal function had increased (Fig. 6f, Supplementary Fig. 14). Moreover, over half of the patients in the middle branches remained stable in that region. We examined the changes in metabolic indicators among the different patient groups and found that variations in phenotypes can drive the movement of patients between groups (Supplementary Fig. 14). Individuals transitioning into higher-risk phenotypic branches, especially into branch 6, exhibited a markedly increased burden of diabetes-related complications, underscoring the clinical relevance of the phenotypic branches in disease progression and personalized management (Supplementary Fig. 15).
A web-based tool for visualizing individual risk
We summarized the patterns of characteristics and complications distribution for Chinese individuals with T2D in the Chinese tree model. The Chinese “tree” identified six branches: Branch 1 linked to DR and peripheral vascular disease with elevated HDL-C and LDL-C; Branch 2 largely complication-free with higher HDL-C; Branch 3 associated with high HDL-C, Cr, and liver cirrhosis risk; Branch 4 marked by higher BMI, heart rate, triglycerides, and Cr accompanied with elevated HF and MI risks; Branches 5 and 6 featured elevated blood pressure, HbA1c, LDL-C, and risks of kidney disease, HF, and stroke. As a potential tool for assisting clinicians and their patients in visualizing individual patient profiles at the diagnosis of T2D and understanding associated risks for disease progression and complications, we have summarized our findings in the graph. We developed a computational tool to systematically visualize the associations between clinical phenotypes and complications for individuals with T2D using the Chinese tree model (https://wenglab-t2d-phenotype.shinyapps.io/wenglab-t2d-phenotype/). With this tool, each newly diagnosed T2D Chinese individuals can be mapped to the Chinese tree and their ten-year risks of diabetes-related complications will be estimated based on their ten baseline characteristics needed for the model.
Discussion
We demonstrated heterogeneity in baseline characteristics and T2D-related complication risk distribution between the Chinese and white European populations by analyzing data from a nationwide cohort of 32,501 newly diagnosed Chinese patients with T2D using a Scottish tree model. We then constructed a unique Chinese tree model based on ten diabetes-related variables derived from a deep learning method from this nationwide cohort and validated it in external cohorts of Chinese individuals with T2D.
Our findings suggest that the tree structure and distribution possess heterogeneity among different populations, but consistency within populations of the same ancestry. This has been underpinned by previous studies trying to subtype T2D, which also revealed population-specific patterns and unique association with diabetes-related complications8,9. Ethnic-specific genetic backgrounds may be attributed to the heterogeneity, such as the differences in DR risk patterns between the populations observed in this study. A previous gene network study revealed pathways underlying proliferative DR in African Americans, which was not observed in Europeans21, supporting this possibility. In addition to genetic factors, variations in healthcare systems and routine clinical practices may also contribute to the observed disparities. Unlike many European countries, where structured DR screening programs are well established, routine clinical care in China may not consistently detect mild or early-stage DR. As a result, the prevalence and pattern of DR subtypes captured in our real-world EHR data may underrepresent milder cases, thereby influencing risk estimates and their distribution across the subtype branches.
Interestingly, in our study, higher HDL-C levels were associated with an elevated risk of DR, which appears to contradict the traditional view of HDL-C being protective due to its anti-inflammatory and antioxidant properties. However, the association between HDL-C levels and DR remains inconclusive across studies. A cross-sectional study from the NO BLIND cohort reported that higher HDL-C levels were independently associated with an increased risk of DR in patients with T2D22. A large population-based study using National Health and Nutrition Examination Survey (NHANES) data found an inverted U-shaped association between HDL-C levels and DR risk among adults with diabetes, with increased risk at both low and high HDL-C levels23. These findings suggest that under certain metabolic or inflammatory contexts, high HDL-C may reflect dysfunctional HDL-C particles that have lost their vasoprotective function24. Furthermore, the strong link we observed between liver-related phenotypes and DR risk may also provide indirect mechanistic clues. Earlier research demonstrated that liver dysfunction is known to alter HDL-C metabolism and composition, this may lead to accumulation of pro-inflammatory HDL-C subtype, which may contribute to microvascular damage25,26. Future studies are warranted to explore the role of HDL particle functionality and liver-derived metabolic factors in the development of DR.
More insight has been provided into the varied drivers of T2D-related complications in this study. The tree structures in both Scottish and Chinese populations revealed consistent phenotypic clusters associated with elevated risks of cardiovascular and renal diseases, such as high blood pressure, HbA1c, obesity, and lower eGFR levels. Differences between HFpEF and HFrEF were evident, consistent with findings from the German cohort12. Moreover, we provided estimates of probabilities of stroke in both the Scottish and the Chinese tree models. Stroke remains one of the primary causes of mortality and long-term disability27. We found that both ischemic and hemorrhagic stroke exhibited similar distribution patterns, primarily concentrated in regions with elevated BMI, HDL-C, and LDL-C levels, corroborating with the findings from the Global Burden of Disease Study 201927. Notably, in the Chinese populations, a different pattern for MASLD and cirrhosis were evident in both the Scottish and Chinese trees, suggesting potential metabolic differences. Compared with the European population, Asian people tend to develop MASLD at a lower BMI, which may be driven by different genetic backgrounds28,29. While live is one of the central organs of insulin resistance and dyslipidemia30, our findings highlight the potential role of liver-related mechanisms in population-specific T2D heterogeneity31. Furthermore, the substantial burden of liver disease in China may partly explain the strong influence of liver-related phenotypes on certain diabetes complications such as DR and cardiovascular disease. Liver disease, which including MASLD, viral hepatitis, and cirrhosis, affect an estimated 300 million people in China, representing a major public health challenge32. To better contextualize this, we quantified the prevalence of MASLD and cirrhosis across diabetes complication subgroups within our cohort, demonstrating frequent co-occurrence with cardiovascular, renal, and microvascular complications. This co-morbidity likely reflects shared metabolic pathways and highlights the complex interplay between liver health and T2D heterogeneity in this population. Importantly, emerging data from an international individual participant data meta-analysis demonstrated that T2D was associated with a markedly increased risk of liver decompensation and hepatocellular carcinoma (HCC) among individuals with biopsy-proven or elastography-defined non-alcoholic fatty liver disease (NAFLD)33. These findings strongly support a mechanistic link between T2D and progressive liver dysfunction, emphasizing the need to integrate liver health into models of diabetes heterogeneity and risk stratification.
Approximately one-third of patients shifted branches over the five-year follow-up, often moving toward higher-risk branches, which underscores the importance of continuous monitoring and timely adjustments to stratified treatment to address evolving risk factors and improve patient outcomes. Interestingly, when comparing tree position changes over five years, Chinese participants showed greater positional shifts on the Scottish tree than on the Chinese tree, despite a common assumption that East Asian populations are more metabolically vulnerable. This may reflect the mismatch between population-specific structures: the Scottish tree, derived from the Caucasian population, may exaggerate transitions when applied to external data. Additionally, underrepresentation of certain phenotypes, such as those with very high BMI in the Chinese baseline, may have led to initial central clustering, making later movements appear more substantial. Besides, within the Chinese cohorts involved in this study, we consistently found that hyperglycemia at onset, as reflected by HbA1c, ranked relatively low in the variable listed to characterizing T2D phenotype. This pattern was also observed in the Scottish study11, and may reflect the constrained variability in HbA1c at the time of diagnosis, where most individuals meet similar diagnostic thresholds. This may also suggest a differentiated role and trajectory of glucose in the development of T2D across populations. Current therapeutic strategies for T2D predominantly prioritize glycemic control, often at the expense of understating the progression of metabolic dysfunction-related complications34. Effective management of T2D requires an integrated approach that emphasizes comprehensive care to safeguard target organs susceptible to damage from metabolic disorders. Collectively, considering the heterogeneity in different populations, it is worth developing population-specific models and further studies into the drivers of such heterogeneity is warranted.
Compared with traditional clustering methods used for subtyping T2D, such as k-means clustering35, the DDRTree algorithm provides an alternative framework that captures both the complexity and continuity of disease progression, organizing individuals along trajectories rather than assigning them into discrete clusters, and thereby preserving transitional states between subtypes36. This structure allows for visualizing the metabolic evolution of diabetes, aiding in personalized risk estimation at the individual level. Furthermore, DDRTree produces a low-dimensional and biologically plausible manifold that supports trajectory-informed interpretation of phenotypic branches. This continuous model structure may better capture gradual transitions between phenotypes and reflect the spectrum of T2D heterogeneity. A major strength of our study lies in the data-driven selection of clinically meaningful variables using a VAE combined with machine learning framework. Previous tree-based or clustering studies, such as the seminal work by Nair et al., and subsequent studies using the LURIC cohort—typically selected 9 variables without clearly justifying their selection rationale11,12. For instance, China Cardiometabolic Disease and Cancer Cohort (4C) study adopted a fixed set of 12 variables for prediabetes classification, but did not explain the biological or statistical basis for this choice13. In contrast, we implemented a VAE-informed dimensionality reduction framework to capture latent features that represent the intrinsic structure of high-dimensional clinical data. We then used a gradient-boosting classifier using the 10 VAE-selected features to verify that this subset retained predictive performance comparable to the full feature set. This approach enhances the biological relevance and statistical robustness of the selected features by accounting for nonlinear interactions and multicollinearity between variables. Importantly, this data-driven approach reduces selection bias associated with manually choosing clinically relevant variables.
The strength of this study is that we utilized a nationwide EHR database with follow-up and comprehensive, structuralized data. This large sample size enabled us to construct a robust two-dimensional tree structure using the DDRTree algorithm, capturing the complexity and continuity of T2D progression. Importantly, we enhanced the DDRTree model by selecting phenotype-defining features through a deep learning method, which improved the biological relevance while reducing feature-selection bias. The continuous manifold generated by DDRTree offers an interpretable framework for identifying transitional states and visualizing disease dynamics. This is particularly valuable for informing risk stratification and personalized care, especially in high-risk subpopulations predisposed to specific complications such as liver, renal, and heart diseases. For these individuals, DDRTree-guided prevention strategies may help inform early, targeted interventions. Our findings also underscore that tree structures derived from one population may not be directly transferable to others. The distinct subtype architecture and complication risk patterns observed in the Chinese population highlight the importance of developing population-specific models. Here, the web-based tool we developed with the Chinese tree model comes into use; it facilitates clinical application of Chinese diabetes subtyping and complication risk estimation, which may, in the future, realize individualized treatment.
Several limitations should be acknowledged. First, the use of real-world EHR data introduces potential selection bias and may limit data completeness or accuracy for certain variables. For example, due to limitations in the EHR structure and data availability, we were unable to stratify DR by severity in this analysis. Second, the classification imbalance in training data which is an inherent issue when working with observational cohorts, may have influenced the performance of our unsupervised model. Third, ten-year complication risks in our study were estimated through probability-based modeling. The declining number of individuals with extended follow-up may have slightly reduced the statistical power for accurate estimation of individual-level complication risks. Fourth, although we performed external validation using the JADE cohort from a distinct healthcare system, our findings should be further validated in prospective cohort studies to confirm their generalizability and clinical relevance. Finally, genetic data were not available in this study. Given the known contribution of genetic factors to T2D heterogeneity, this limited our ability to explore genotype-phenotype associations. Future studies integrating genomic data will be essential to further elucidate the biological underpinnings of the identified subtypes.
In conclusion, this study demonstrates the utility of phenotype-based tree models in stratifying T2D patients and predicting the risk of complications. We highlighted the heterogeneity of diabetes in the Chinese population compared to the Caucasian population, highlighting that T2D subtyping is not a one-size-fits-all process and reinforcing the need for ethnic-specific tools. This approach enhances the understanding of their unique phenotype and has the potential to contribute to the precision management of T2D in China.
Methods
Ethics statement and study population
This study was approved by the Medical Ethics Committee of Nanfang Hospital, Southern Medical University (approval number: NFEC-2019-213). The requirement for individual informed consent was waived by the Medical Ethics Committee of Nanfang Hospital, Southern Medical University, in compliance with the Measures for the Ethical Review of Biomedical Research Involving Humans (National Health Commission of China), which permits informed-consent waivers for minimal-risk retrospective studies using anonymized data, as in our study. For DR cohort and the healthy individuals, ethical approval was granted by the First Affiliated Hospital of USTC (2021-KY-034). For the JADE cohort, ethical approval was granted by The Chinese University of Hong Kong (CUHK) Clinical Research Ethics Committee (CREC 2007.339) and the respective local institutional ethics boards involved in the implementation and evaluation of the JADE Program.
We used the Chinese Renal Disease System (CRDS) as our main data source. The CRDS18 is a comprehensive multicenter database. It collected inpatient and outpatient data from the EHR systems of 24 medical centers across China from January 1, 2000, to May 26, 2024. The system collected anonymized health-related data from all the patients, not limited to those with renal diseases, who visited the participating hospital during this period, such as outpatient and hospitalization notes, diagnosis, surgical procedures, laboratory results, medications, and in-hospital mortality. The system now possesses a database encompassing 8,632,000 patients, with follow-up data available for 2,550,000 individuals by the time the current analysis was performed.
For this study, we included individuals aged 18 years or older from the CRDS who had T2D (ICD-10 code E11) with a disease duration of no more than 12 months at baseline. Baseline was defined as the first recorded diagnosis of T2D in the database, and we confirmed the disease duration at baseline by reviewing the clinical notes. We excluded individuals (a) diagnosed with type 1 diabetes or other types of diabetes (ICD-10 codes E10 or E13); (b) under 18 years; (c) pregnant at baseline; and (d) with major medical conditions, including malignancy, organ transplantation, or ESRD, as these conditions may significantly impact metabolic status and clinical profiles, thereby impact the clustering of T2D-specific phenotypes (Supplemental Table 1).
We construct two external validation cohorts with the same inclusion and exclusion criteria as the CRDS cohort. For the first cohort, we used data of eligible participants from the JADE program between 2001 and 2019, to validate the models in population treated under a different healthcare system (JADE cohort). The JADE Program19,20, developed from the Hong Kong Diabetes Register, JADE retrieves data on medications, laboratory tests, hospitalizations, and outpatient visits, by linking participants’ unique identity numbers with the territory-wide Hospital Authority electronic medical record system.
As the incident time of DR was unavailable from the JADE cohort, another cohort (DR cohort) was constructed to validate the findings in the DR distribution pattern in the Chinese population. The data sources were EHRs between 2018 and 2023 from three medical centres: the First Affiliated Hospital of Bengbu Medical University, the Third Affiliated Hospital of Sun Yat-sen University, and Nanjing Drum Tower Hospital. We collected data needed to validate the Scottish and the Chinese models.
Diabetes-related outcomes
MI: Myocardial infarction was defined as acute or chronic myocardial injury due to ischemia, identified using ICD-10 codes I21 (acute MI), I22 (subsequent MI), and I25.2 (old MI)37.
Stroke: Stroke was defined as the first appearance of ICD-10 codes I60–I64, including ischemic stroke (I63), intracerebral hemorrhage (I61), subarachnoid hemorrhage (I60), other and unspecified nontraumatic intracranial hemorrhage (I62), and unspecified stroke (I64)37. Ischemic Stroke: Defined by ICD-10 code I63. Hemorrhagic Stroke: Defined by ICD-10 codes I60–I62.
HF: Hospitalization due to HF was defined by the first inpatient diagnosis coded as I50. HFpEF was defined as heart failure with a left ventricular ejection fraction (LVEF) ≥ 50% documented by echocardiography37. HFrEF was defined as heart failure with a left ventricular ejection fraction (LVEF) < 40%.
CKD: Chronic kidney disease was identified through electronic health records by detecting an eGFR of ≤60 ml/min/1.73 m², confirmed by at least two separate measurements taken 3 months apart38. The eGFR was determined using the CKD-EPI equation based on serum creatinine levels39.
ESRD: ESRD was defined as a requirement for dialysis longer than 3 months (either hemodialysis or peritoneal dialysis) or preemptive renal transplantation.
DR: Diabetic retinopathy was identified via ICD-10 codes E11.371, and E14.371. Due to limitations in EHR coding, grading granularity was not available. These codes primarily reflect moderate to severe forms of DR with clinical significance, including moderate non-proliferative diabetic retinopathy (NPDR) with diabetic macular edema (DME), severe non-proliferative diabetic retinopathy, and proliferative diabetic retinopathy (PDR)40.
Liver Cirrhosis: Liver cirrhosis was diagnosed according to the ICD-10 codes K74.1 to K74.6 and K70.2 to K70.3.
MASLD: Diagnosis defined as the presence of hepatic steatosis in conjunction with one CMRF and no other discernible cause41 (Supplementary Table 1).
DPVD: Identified by ICD-10 code E11.5, which includes I79.2, indicating peripheral angiopathy due to diabetes42. Detailed definitions of these outcomes of interest are summarized in Supplementary Table 1.
Mapping the Chinese populations to the “Scottish tree”
Following the methods11 described by Nair et al., the individuals from the CRDS cohort were mapped onto the Scottish tree using baseline clinical characteristics of HbA1c, BMI, TC, HDL-C, ALT, Cr, SBP, triglycerides and DBP, ensuring alignment with the distribution of these nine phenotypic variables within the Scottish tree. Individuals mapped to the “tree” were required to have all the aforesaid nine variables available within one year from the baseline. Outliers were excluded based on values beyond 5 standard deviations15. The data were transformed using rank normalization. Then each participant was assigned a position on the Scottish tree, adjusted for age and sex.
Construction of the “Chinese Tree”
To better capture the key features for constructing the phenotyping tree for the Chinese population, we employed a VAE model to learn a compressed latent representation of clinical input features (Supplementary Table 2). After excluding individuals with missing values in key variables required for model training (e.g., HbA1c, BMI, blood pressure, lipid profiles), a total of 45,589 patients with T2D from the CRDS cohort and 11,340 non-diabetic individuals from the Anhui Provincial Health Center were included in the analysis.
Design of VAE
The VAE framework was designed with a flexible architecture comprising fully connected layers in both the encoder and decoder networks17. Each hidden layer is followed by batch normalization and LeakyReLU activation, along with dropout for regularization. We implemented a standard variational autoencoder (VAE) with fully connected encoder and decoder networks. The encoder network \({q}_{\phi }(z|x)\) maps input \(x\) to a latent representation through a series of hidden layers with batch normalization, nonlinear activations, and dropout regularization. The encoder outputs parameters of a diagonal Gaussian posterior distribution over latent variables \(z\in {{\mathbb{R}}}^{L}\):
where \({\mu }_{\phi }(x)\) and \({\sigma }_{\phi }(x)\) denote the mean and standard deviation predicted by the encoder. The decoder network \({p}_{\theta }({x|z})\) reconstructs the input from the latent representation. We employed the reparameterization trick to enable backpropagation through stochastic sampling:
The latent prior was set to a standard Gaussian \(p(z)={{{\mathscr{N}}}}(0,I)\). For downstream analyses, we used the encoder mean \({\mu }_{\phi }(x)\) as the deterministic latent representation.
Loss Function and Optimization
The model was trained by maximizing the evidence lower bound (ELBO), equivalently minimizing the negative ELBO:
The reconstruction loss \({{{{\mathscr{L}}}}}_{{\mbox{recon}}}\) was computed as the Mean Squared Error (MSE) between input and reconstruction:
The Kullback-Leibler divergence \({{{{\mathscr{L}}}}}_{{\mbox{KL}}}\) regularizes the posterior toward the prior:
where \({\mu }_{i}\) and \({\sigma }_{i}^{2}\) are the mean and variance of the \(i\)-th latent dimension. The hyperparameter \(\beta\) controls the trade-off between reconstruction fidelity and latent regularization. We specifically used a β value of 0.001 for the final model. All losses were normalized by batch size and input dimensionality.
Model training
The model was implemented in PyTorch (v1.9.0) and trained on a GPU with CUDA (v11.8). Model optimization was performed using the Adam optimizer with a fixed learning rate of 1 × 10⁻⁴ and a batch size of 1024. Training was conducted for 150 epochs, and a fixed random seed was used to ensure reproducibility. After training, the latent representation of each sample was obtained by extracting the mean vector μ from the encoder output.
Hyperparameter optimization
We initially divided the dataset into a training set (90%) and a test set (10%) to identify the optimal hyperparameter settings to capture the data structure without losing the ability to generalize on the test data. All metrics were computed only on non-missing entries. To identify the optimal combination of hyperparameters that best captured the underlying structure of the data while maintaining good generalization performance, we systematically tested various configurations of the VAE. These included: hidden layer sizes (256, 512, and 1024), number of hidden layers (1 to 3), latent dimension (24, 32, and 48), dropout rate (10%, 20%, and 30%), and KLD weight (1 × 10⁻3, 1 × 10⁻⁴, and 1 × 10⁻5). Model performance was evaluated based on the ELBO loss (reconstruction error plus KL divergence) on the test set and the stability of the latent space. Reconstruction quality was evaluated by calculating the cosine similarity between the input and reconstructed feature vectors for each individual. Latent space stability was further assessed for the top-performing configurations by repeatedly training models with identical hyperparameters and computing the variation in cosine similarity of each individual’s latent vector with all others across runs. A smaller average variation indicates greater stability. The model with the most consistent representations was selected with the average change closest to zero. We observed that models with more complex architectures—such as those with additional hidden layers or a greater number of neurons—tended to perform worse. In particular, models with more than one hidden layer failed to achieve good reconstruction performance without overfitting. The only exception was the dimensionality of the latent space: models with a small latent dimension 16 underperformed, whereas those with dimensions between 32 and 48 showed comparable performance. Among evaluated architectures, simpler models with a single hidden layer performed better in generalization compared to deeper or wider models, which tended to overfit. Specifically, the configuration with one hidden layer of 512 neurons, a latent dimension of 32, dropout rate of 10%, and KLD weight of 0.0001 yielded the best balance between accuracy and stability.
Feature importance analysis
Feature importance was estimated from the absolute weights connecting each input feature to the hidden layer. Specifically, for each input feature, importance is calculated as:
where \({I}_{i}\) represents the importance of the \({I}_{i}\) th input feature and \({w}_{{ij}}\) denotes the weight from the \({I}_{i}\) th input to the \(j\) th hidden neuron. To assess the actual impact on the latent representation an adaptation of the SHAP16 analysis was applied. For the perturbation-based approach, we systematically set each feature to “missing” for all individuals and measure the resulting change in latent representations. Missing values are encoded as mean-substitution (0 after z-score standardization). The combination of these two methods provided a comprehensive view of feature relevance in the clinical data. A set of 10 overlapping features was consistently identified as the most influential: HDL-C, triglycerides, SBP, ALT, HbA1c, LDL-C, creatinine, heart rate, BMI, and DBP.
Comparison for feature selection and model performance
Beyond unsupervised representation learning, we sought to validate the discriminatory utility of the features selected by the VAE using supervised machine learning classifiers. Multiple classifiers were implemented in Python using the scikit-learn package, including Logistic Regression (LR), Random Forest (RF), Support Vector Machine (SVM), Linear SVM (LSVM), Gaussian Naive Bayes (GNB), Decision Tree Classifier (DTC), Gradient Boosting Classifier (GBC), Least Absolute Shrinkage and Selection Operator (LASSO) and Elastic net, which trained and evaluated using the full original clinical variables. Model performance was comprehensively assessed using 10-fold cross-validation and quantified by multiple metrics, including accuracy, area under the ROC AUC, and PR AUC, to ensure robustness across imbalanced classes. All models were trained using default hyperparameters provided by scikit-learn, except for LASSO and Elastic Net, which employed LogisticRegressionCV to automatically tune the regularization strength (λ), which corresponds to the inverse of C. Among all classifiers, GBC achieved the best overall performance (ROC AUC > 0.9), followed closely by RF, and was thus selected as the final classifier. These two classifiers were further applied to evaluate the predictive performance of models reconstructed using the top 5-10 VAE-derived features (Supplementary Fig. 5). The comparable results between reduced and full feature sets confirmed the stability and sufficiency of the VAE-selected variables in capturing the intrinsic heterogeneity of T2D.
Data dimensionality reduction with DDRTree
We utilized data from the CRDS cohort and divided the dataset into an 80% training set and a 20% validation set. Using the top ten features identified, we applied the DDRTree algorithm43 to construct a “Chinese Tree” model as Nair et al did11, to visualize the clustering of individuals with T2D based selected features distribution in a dimensionality reduced graph. Each phenotype was then residualized for age and sex via linear regression. The resulting matrix was input into the DDRTree algorithm, which embedded the high-dimensional phenotype data into a two-dimensional space. Each individual was then mapped onto the Chinese tree, representing the data-driven organization of phenotypic heterogeneity among individuals with T2D. All data management and statistical analyses were conducted using R version 3.5.2. The DDRTree algorithm was implemented using the ‘monocle’ package from Bioconductor.
Evaluating the distribution of phenotypes and outcomes
To assess phenotype distribution across the tree structure, we performed linear regression analyses of each phenotype against the tree dimensions (X and Y), and visualized the regression coefficients alongside their 95% confidence intervals (CIs). To evaluate the risk of diabetes-related complications, we employed a competing risk model (Fine–Gray model) and derived sub-distribution hazard ratios (sHRs), treating death as a competing event. This analysis was conducted in the full CRDS cohort, with separate models developed for each complication, including DR, HF (HFrEF, HFpEF), Stroke (ischemic and hemorrhagic stroke), MI, ESRD, CKD, MASLD, liver cirrhosis, and DPVD. Individuals who had already experienced the outcome of interest at baseline or who lacked follow-up data were excluded from the corresponding models. Those who died during follow-up were treated as having experienced a competing event, whereas individuals who reached the end of follow-up or were lost to follow-up were censored. We constructed Fine–Gray models for each outcome of interest, and developed three model versions: the first using the tree dimensions from DDRTree, the second integrating DDRTree dimensions with spline functions as covariates, and the third incorporating age, sex, ten clinical characteristics, and duration of T2D diagnosis as covariates11. These models enabled us to examine the association between the identified latent subgroups and long-term risk of complications while appropriately accounting for competing mortality. To obtain the individual probability for developing each diabetes outcome (for example, CKD or other outcomes) for each study participant at a 10-year follow-up period, we used the previously constructed competing risk models with dimensions from DDRTree as covariates. For example, to estimate the probability of stroke incidence for a study participant, we used the stroke competing risk model (event of interest: stroke; competing event: death) constructed with DDRTree dimensions as covariates. Using this model and each individual’s position in reduced space, we predicted event probability for that individual. Similarly, we calculated the individual-level probability of other outcomes using corresponding competing risk models. These event probabilities were overlaid on the tree diagram to visualize the heterogeneity in diabetes progression.
Constructing a mapping function for external validation
To map individuals with newly diagnosed T2D to the Chinese tree, we constructed a mapping function using 10 diabetes-related clinical variables: HDL-C, triglycerides, SBP, ALT, HbA1c, LDL-C, Cr, heart rate, BMI, and DBP. The function comprises two components: (1) two generalized additive models (GAMs) with smooth terms fitted using cubic regression splines to predict the DDRTree-derived two-dimensional coordinates (Dimension 1 and Dimension 2) from the 10 phenotypes; and (2) a distance-estimating algorithm that calculates the Euclidean distance between points in the two-dimensional space. Given a new individual with all 10 phenotypes, the trained GAMs first predict the provisional DDRTree coordinates (Dimension 1 and Dimension 2). Next, the distance between this provisional point and all individuals in the reference Chinese tree (n = 26,000) is computed. The final mapping assigns the new individual to the nearest neighbor in the reference tree based on the minimum Euclidean distance. This process allows for identifying the most phenotypically similar reference individual for the newly diagnosed case.
We applied the mapping function to evaluate the alignment and consistency of positioning for the remaining 20% of newly diagnosed T2D patients within the established tree structure, as an internal validation. Secondly, to investigate the performance of the “Chinese tree” model across different healthcare systems, we validated the model in the JADE cohort and the DR cohort. In the JADE cohort, individuals were mapped to the reference tree using age at diagnosis, sex, and the 10 clinical phenotypes. After mapping, we overlaid these phenotypes to assess their distribution across the tree and evaluated the consistency with the reference population. Diabetes progression was assessed by examining the occurrence of seven clinical outcomes during follow-up: MI, stroke, ischemic stroke, HF, cirrhosis, CKD, and ESRD. The definitions of CKD and ESRD are provided in Supplementary Table 3. In the external JADE cohort, we similarly applied Cox proportional hazard model using the DDRTree-derived tree dimensions as covariates to estimate the individual probability of each complication.
In the DR cohort, we specifically validated the model’s performance in predicting DR. DR cases were defined based on clinical diagnosis of moderate NPDR with DME, severe NPDR, or PDR, consistent with the CRDS cohort definition. In DR cohort, the predicted event probabilities were overlaid on the tree to assess the spatial distribution of complications across phenotypic subgroups. For the probability of the incidence of DR, we derived the hazard ratio (HR) and 95% CIs from the Cox proportional hazard model for deriving event probabilities.
Sensitivity analysis
To examine the potential impact of hepatitis virus infection on the associations between diabetes and primary liver outcomes in the Chinese population, we conducted a sensitivity analysis by excluding individuals with a diagnosis of viral hepatitis from the CRDS cohort. Then we excluded the population with alcoholic cirrhosis and acute pancreatitis (Supplementary Table 1). After removing these individuals, we re-assessed the phenotypes and complication outcomes using the Chinese tree framework to ensure the robustness and stability of our findings.
Phenotype changing with time in the Scottish Tree and Chinese Tree models
To assess changes in these variables during the first 5 years after a diabetes diagnosis, we examined the gradient of individual slopes (deltas) across the two three-dimensional. Deltas with a gradient greater than 1.0 were considered indicative of significant positional changes across the tree. We also divided the coordinate regions of the Scottish and Chinese trees into distinct parts to visualize individual changes over the five-year follow-up period.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The individual-level data used in this study are not publicly available due to privacy regulations. Access to the CRDS data subset used in this article via the corresponding author (Prof. Jianping Weng, wengjp@ustc.edu.cn). Applicants must provide a research proposal, evidence of ethical approval, and a data protection plan. Requests will be evaluated within 8 weeks, and approved applicants will receive access through a secure, time-limited environment for a period of 12 months, with possible renewal upon review. Data used to generate the Scottish tree model can be obtained upon request from the Scottish Care Information-Diabetes Collaboration, Tayside & Fife, Scotland unit (https://www.sci-diabetes.scot. nhs.uk/). For access to other datasets, including the JADE and DR cohorts, or to request scripts adapted to those datasets, please apply through the Asia Diabetes Foundation (enquiry@adf.org.hk) for JADE data, or through the corresponding author (wengjp@ustc.edu.cn) for DR data. Inquiries will receive a response within 1 month. These datasets are available under controlled access because they contain individual-level clinical and biomarker information governed by participant consent and institutional privacy regulations. Aggregated statistics and summary-level data that support the main findings of this study are provided in the Source Data file. Source data are provided with this paper.
Code availability
The code for training the VAE, machine learning methods and constructing the Chinese tree is available on GitHub at: https://github.com/Tongyue1999/Chinese-DDrtree. This repository includes annotated scripts for data preprocessing, model training, and evaluation, as well as parameters used in the final model. To facilitate reproducibility, we also provide the original plotting scripts and access to publicly available datasets used in the analysis. The R scripts used for phenotype trajectory mapping and validation are specific to the CRDS database and contain sensitive field-level references; therefore, they are not publicly available. The trajectory analysis was performed using the publicly available DDRTree method implemented in the monocle R package (v0.1.5), which can be accessed via the Bioconductor repository: https://www.bioconductor.org/packages/release/bioc/html/monocle.html.
References
Roden, M. & Shulman, G. I. The integrative biology of type 2 diabetes. Nature 576, 51–60 (2019).
World Health Organization. Classification of diabetes mellitus. (2019).
Pearson, E. R. Type 2 diabetes: a multifaceted disease. Diabetologia 62, 1107–1112 (2019).
Tuomi, T. et al. The many faces of diabetes: a disease with increasing heterogeneity. Lancet 383, 1084–1094 (2014).
Herder, C. & Roden, M. A novel diabetes typology: towards precision diabetology from pathogenesis to treatment. Diabetologia 65, 1770–1781 (2022).
Ahlqvist, E. et al. Novel subgroups of adult-onset diabetes and their association with outcomes: a data-driven cluster analysis of six variables. Lancet Diab. Endocrinol. 6, 361–369 (2018).
Zaharia, O. P. et al. Risk of diabetes-associated diseases in subgroups of patients with recent-onset diabetes: a 5-year follow-up study. Lancet Diab. Endocrinol. 7, 684–694 (2019).
Anjana, R. M. et al. Novel subgroups of type 2 diabetes and their association with microvascular outcomes in an asian indian population: A data-driven cluster analysis: the INSPIRED study. BMJ Open Diab. Res Care 8, e001506 (2020).
Xiong, X. F. et al. Identification of two novel subgroups in patients with diabetes mellitus and their association with clinical outcomes: A two-step cluster analysis. J. Diab. Investig. 12, 1346–1358 (2021).
Dennis, J. M. Precision medicine in type 2 diabetes: Using individualized prediction models to optimize selection of treatment. Diabetes 69, 2075–2085 (2020).
Nair, A. T. N. et al. Heterogeneity in phenotype, disease progression and drug response in type 2 diabetes. Nat. Med 28, 982–988 (2022).
Schön, M. et al. Analysis of type 2 diabetes heterogeneity with a tree-like representation: insights from the prospective german diabetes study and the LURIC cohort. Lancet Diab. Endocrinol. 12, 119–131 (2024).
Lin, H. et al. Elucidating the heterogeneity of prediabetes through subphenotyping with a two-dimensional tree structure. Cell Rep. Med. 6, 102212 (2025).
Fagherazzi, G. et al. Heterogeneity of glycaemic phenotypes in type 1 diabetes. Diabetologia 67, 1567–1581 (2024).
Coral, D. E. et al. Subclassification of obesity for precision prediction of cardiometabolic diseases. Nat. Med. 31, 534–543 (2025).
Nissen, J. N. et al. Improved metagenome binning and assembly using deep variational autoencoders. Nat. Biotechnol. 39, 555–560 (2021).
Allesøe, R. L. et al. Discovery of drug-omics associations in type 2 diabetes with generative deep-learning models. Nat. Biotechnol. 41, 399–408 (2023).
Wu, C. et al. Predicting in-hospital outcomes of patients with acute kidney injury. Nat. Commun. 14, 3739 (2023).
Chan, J. C. N. et al. From hong kong diabetes register to JADE program to RAMP-DM for data-driven actions. Diab. Care 42, 2022–2031 (2019).
Ko, G. T. et al. From design to implementation-the joint asia diabetes evaluation (JADE) program: a descriptive report of an electronic web-based diabetes management program. BMC Med Inf. Decis. Mak. 10, 26 (2010).
Pollack, S. et al. Multiethnic genome-wide association study of diabetic retinopathy using liability threshold modeling of duration of diabetes and glycemic control. Diabetes 68, 441–456 (2019).
Sasso, F. C. et al. High HDL cholesterol: A risk factor for diabetic retinopathy? Findings from NO BLIND study. Diab. Res Clin. Pr. 150, 236–244 (2019).
Xu, W., Xu, X., Zhang, M. & Sun, C. Association between HDL cholesterol with diabetic retinopathy in diabetic patients: a cross-sectional retrospective study. BMC Endocr. Disord. 24, 65 (2024).
Zhao, L., Ma, J., Wang, S. & Xie, Y. Relationship between β-cell function, metabolic control, and microvascular complications in type 2 diabetes mellitus. Diab. Technol. Ther. 17, 29–34 (2015).
Nicholls, S. J. et al. Consumption of saturated fat impairs the anti-inflammatory properties of high-density lipoproteins and endothelial function. J. Am. Coll. Cardiol. 48, 715–720 (2006).
Patel, S. et al. Acute hypertriglyceridaemia in humans increases the triglyceride content and decreases the anti-inflammatory capacity of high density lipoproteins. Atherosclerosis 204, 424–428 (2009).
Ma, Q. et al. Temporal trend and attributable risk factors of stroke burden in China, 1990-2019: an analysis for the global burden of disease study 2019. Lancet Public Health 6, e897–e906 (2021).
Fracanzani, A. L. et al. Liver and cardiovascular damage in patients with lean nonalcoholic fatty liver disease, and association with visceral obesity. Clin. Gastroenterol. Hepatol. 15, 1604–1611.e1601 (2017).
Wei, J. L. et al. Prevalence and severity of nonalcoholic fatty liver disease in non-obese patients: A population study using proton-magnetic resonance spectroscopy. Am. J. Gastroenterol. 110, 1306–1314 quiz 1315 (2015).
Stefan, N., Lonardo, A. & Targher, G. Role of steatotic liver disease in prediction and prevention of cardiometabolic diseases. Nat. Rev. Gastroenterol. Hepatol. 21, 136–137 (2024).
Mak, L. Y. et al. Liver diseases and hepatocellular carcinoma in the asia-pacific region: burden, trends, challenges and future directions. Nat. Rev. Gastroenterol. Hepatol. 21, 834–851 (2024).
Wang, F. S., Fan, J. G., Zhang, Z., Gao, B. & Wang, H. Y. The global burden of liver disease: the major impact of China. Hepatology 60, 2099–2108 (2014).
Huang, D. Q. et al. Type 2 diabetes, hepatic decompensation, and hepatocellular carcinoma in patients with non-alcoholic fatty liver disease: an individual participant-level data meta-analysis. Lancet Gastroenterol. Hepatol. 8, 829–836 (2023).
Lu, X. et al. Type 2 diabetes mellitus in adults: pathogenesis, prevention and therapy. Signal Transduct. Target Ther. 9, 262 (2024).
Li, X. et al. Trajectories of clinical characteristics, complications and treatment choices in data-driven subgroups of type 2 diabetes. Diabetologia 67, 1343–1355 (2024).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Tsao, C. W. et al. Heart disease and stroke statistics-2022 update: A report from the american heart association. Circulation 145, e153–e639 (2022).
Andrassy, K. M. Comments on ‘KDIGO 2012 clinical practice guideline for the evaluation and management of chronic kidney disease. Kidney Int. 84, 622–623 (2013).
Levey, A. S. et al. A new equation to estimate glomerular filtration rate. Ann. Intern Med. 150, 604–612 (2009).
Hou, X. et al. Prevalence of diabetic retinopathy and vision-threatening diabetic retinopathy in adults with diabetes in China. Nat. Commun. 14, 4296 (2023).
Kanwal, F., Neuschwander-Tetri, B. A., Loomba, R. & Rinella, M. E. Metabolic dysfunction-associated steatotic liver disease: Update and impact of new nomenclature on the american association for the study of liver diseases practice guidance on nonalcoholic fatty liver disease. Hepatology 79, 1212–1219 (2024).
ElSayed, N. A. et al. 12. Retinopathy, neuropathy, and foot care: Standards of care in diabetes-2023. Diab. Care 46, S203–s215 (2023).
Qi, M., Li, W., Tsang, I. W. & Yijun, S. Principal graph and structure learning based on reversed graph embedding. IEEE Trans. Pattern Anal. Mach. Intell. 39, 2227–2241 (2017).
Acknowledgements
This study is funded by Noncommunicable Chronic Diseases-National Science and Technology Major Project (grant No. 2023ZD0509100; awarded to J.W.), Institute of Health and Medicine, Hefei Comprehensive National Science Center 2023 Annual Research Project of the Institute of Pioneer Medicine and Advanced Technologies (grant no. 2023IHM01060; awarded to J.W.), and Anhui Province Clinical Medical Research Translation Special Project (grant no. 202204295107020026; awarded to J.W.). We thank Mr. Fei Wang (IT Technician, Hefei Comprehensive National Science Center) and Ms. Minzhe Zhao (IT Technician, First Affiliated Hospital of USTC) for technical support. The funding sources had no role in study design, data collection, data analysis, data interpretation, or writing of the report. The corresponding authors had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Author information
Authors and Affiliations
Contributions
T.Y., W.Z., and Y.D. contributed equally to the manuscript. T.Y. wrote the first draft of the manuscript and contributed to methodology, data analysis, and data interpretation. W.Z. contributed to methodology, data analysis, data interpretation, and visualization, reviewed and revised the manuscript. Y.D. contributed to the study conceptualization and data interpretation, reviewed, and revised the manuscript. X.Z. contributed to administrative support, data interpretation, and discussion of the manuscript. Y.M. and Z.W. contributed to methodology, data analysis, and discussion of the manuscript. J.C.N.C., E.S.H.L., J.N.M.L., G.J., W.X., Y.B., and S.N. contributed to data collection, data interpretation, and discussion of the manuscript. S.N. also provided administrative support. M.G. contributed to data collection and discussion of the manuscript. E.R.P. contributed to methodology, data analysis, and data interpretation, reviewed and revised the manuscript. SL contributed to study conceptualization, administrative support, data analysis, data interpretation, and reviewed and revised the manuscript. JW contributed to study conceptualization, administrative support, fund acquisition, data interpretation, and reviewed and revised the manuscript. The corresponding authors attest that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Aravind Lathika Rajendrakumar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yue, T., Zhang, W., Ding, Y. et al. Precision phenotyping of type 2 diabetes in chinese populations using a variational autoencoder-informed tree model. Nat Commun 17, 1475 (2026). https://doi.org/10.1038/s41467-025-68211-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68211-4