Abstract
O-Linked N-acetylglucosamine (O-GlcNAc) is a nucleocytoplasmic post-translational modification that is tightly regulated by O-GlcNAc transferase (OGT) and O-GlcNAcase (OGA). Dysregulation of O-GlcNAc in human disease has motivated efforts to therapeutically modulate O-GlcNAc. Drug repurposing efforts can accelerate these campaigns and unveil how clinically relevant compounds and pathways intersect with O-GlcNAc. Here we report the results of three parallel drug repurposing screens against the O-GlcNAc cycling enzymes in cells and in vitro that reveal kinase inhibitors GSK690693 and Y-33075 act as splicing modulators that disrupt O-GlcNAc homeostasis and simultaneously downregulate OGT and OGA. These effects are independent of their respective annotated targets, AKT and ROCK, and are distinct from OGT and OGA inhibitors and similar kinase inhibitors. Evaluation of a panel of splicing modulators revealed three additional potent compounds (OTS964, indisulam, GNF2133) that similarly downregulate OGT and OGA with distinct splicing profiles. These findings reveal previously unobserved splicing modulator chemotypes and approaches to disrupt O-GlcNAc homeostasis.
Similar content being viewed by others
Introduction
O-Linked N-acetylglucosamine (O-GlcNAc) is an essential post-translational modification installed on serine or threonine residues across thousands of nucleocytoplasmic proteins (Fig. 1a)1. O-GlcNAc has functional roles in numerous cellular processes, such as the cell cycle, nutrient sensing, and transcription2. Owing to the central role of O-GlcNAc in cell biology, dysregulation of O-GlcNAc has been associated with many diseases including neurodegeneration, X-linked intellectual disability (XLID), cancer, diabetes, and immune dysfunction3. Thus, modulation of O-GlcNAc as a strategy to mitigate these diseases has been explored in preclinical and clinical efforts3,4,5,6,7,8.
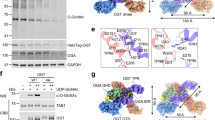
a Schematic of O-GlcNAc regulation by enzymes OGT and OGA with commonly used OGT inhibitor OSMI-4a/b and OGA inhibitor TMG. b Canonical splicing pathways of OGT and OGA transcripts. High O-GlcNAc favors nonproductive OGT transcripts and the productive OGA transcript. Low O-GlcNAc favors the productive OGT transcript and the nonproductive OGA transcript. c Western blot of O-GlcNAc, OGT, and OGA levels in HCT116 cells after indicated treatment for 24 h (n = 3). d Schematic of GFP-β-OGT reporter for OGT splicing. e Schematic of GFP-β-OGT reporter screen. Lower O-GlcNAc correlates with increased GFP signal, while elevated O-GlcNAc correlates with decreased GFP signal. f Sample images of GFP-β-OGT reporter cells following indicated treatment for 24 h. Scale bar 100 μm. g Dose-response change in mean nuclei GFP per well with indicated compound for 24 h (n = 5 biological replicates). Data are represented as mean ± s.d. h Screening results with the Drug Repurposing Library in GFP-β-OGT reporter cells. i Average Δ nuclei mean GFP versus average viability in GFP-β-OGT reporter across n = 2 replicates. Viability determined by nuclei counts. j Schematic of filter for autofluorescent compounds in wild-type HCT116 cells. k Average Δ nuclei mean GFP in wild-type HCT116 cells versus average Δ nuclei mean GFP in GFP-β-OGT reporter cells from n = 3 replicates to filter autofluorescent compounds. l Activity of selective and nonselective ROCK inhibitors in GFP-β-OGT reporter. Source data are provided as a Source Data file.
The O-GlcNAc modification is enzymatically installed by O-GlcNAc transferase (OGT) and removed by O-GlcNAcase (OGA) (Fig. 1a). The cell maintains O-GlcNAc homeostasis in part through regulation of OGT and OGA via a conserved detained intron (DI) feedback mechanism (Fig. 1b). Intron detention is a form of alternative splicing in which immature transcripts containing DIs are sequestered in the nucleus9. This provides a pool of transcripts that can respond rapidly to changes in the cellular environment by channeling OGT and OGA transcripts towards productive or nonproductive pathways, such as nuclear detention or nonsense-mediated decay (NMD), resulting in differential expression of the enzymes to maintain O-GlcNAc homeostasis (Fig. 1b)10,11. For OGT, splicing to join exons 4 and 5 yields the productive transcript, whereas alternative splicing to include a poison cassette exon (PE) introduces a premature termination codon (PTC) into the transcript, leading to NMD12. For OGA, splicing to join exons 10, 11, and 12 yields the productive transcript, whereas splicing to directly join exons 10 and 12 incorporates a PTC into the transcript leading to NMD (Fig. 1b). Through this conserved DI feedback mechanism, the cell sensitively and rapidly responds to changes in O-GlcNAc by tuning OGT and OGA expression and maintains O-GlcNAc homeostasis10,11.
Chemical inhibitors for OGT or OGA, like OSMI-413 and thiamet-G (TMG)14, respectively, have provided important research tools to modulate O-GlcNAc and study the DI feedback mechanism (Fig. 1a). The DI feedback mechanism can dampen the overall cellular response to these inhibitors10,11. Nonetheless, OGA inhibitors have progressed to clinical evaluation for treatment of neurodegenerative diseases4,5 including Alzheimer’s disease6 and progressive supranuclear palsy (PSP)7,8, and preclinical efforts in Parkinson’s disease and amyotrophic lateral sclerosis (ALS)3,15,16. While the exact cellular factors that are involved in the DI feedback mechanism are uncharacterized, identification of additional pharmacological modulators of O-GlcNAc that act by alternate mechanisms will provide valuable tools to study O-GlcNAc and open additional strategies to therapeutically modulate O-GlcNAc.
Here, we report the results of three comprehensive high-throughput screening campaigns for O-GlcNAc modulators utilizing the Broad Institute’s Drug Repurposing Library, a curated collection of 5183 chemical probes, preclinical compounds, and approved drugs17. Drug repurposing efforts can uncover biological relationships between clinically relevant compounds and pathways and O-GlcNAc. We screened this library against a cellular reporter for the OGT DI feedback mechanism and in OGT and OGA activity assays in vitro, leading to the discovery of compounds that inhibit OGA and those that surprisingly downregulate OGT and OGA. Examination of these compounds and additional splicing modulators reveals a previously unreported mechanism to disrupt O-GlcNAc homeostasis through alternative splicing of OGT and OGA transcripts.
Results
Cellular reporter screen for O-GlcNAc modulators
We first evaluated whether clinically relevant compounds affect O-GlcNAc homeostasis by high-throughput screening of the Broad Institute’s Drug Repurposing Library against a GFP-β-OGT cellular minigene reporter derived from the OGT DI region in HCT116 cells18,19. HCT116 cells are sensitive to the O-GlcNAc DI feedback mechanism, as inhibitors of OGT or OGA upregulate or downregulate OGT, respectively (Fig. 1c). The GFP-β-OGT reporter consists of a GFP fusion to OGT exons 4 and 5 with the intervening intronic region flanked by efficiently spliced β-globin sequences to support expression and mimic the OGT DI natural context (Fig. 1d). By co-opting the endogenous OGT transcript elements that are O-GlcNAc-responsive, this reporter links GFP expression to OGT DI splicing such that diminished O-GlcNAc induces splicing to afford the mature transcript, while elevated O-GlcNAc promotes nuclear detention or NMD of the reporter transcript (Fig. 1e). Therefore, the GFP-β-OGT reporter cell line produces GFP signal as a proxy for OGT DI splicing and thereby changes in O-GlcNAc.
To evaluate the Drug Repurposing Library compounds with the GFP-β-OGT reporter, we utilized high-content imaging to measure nuclei count and GFP levels (Fig. 1f). We performed these cellular experiments and later validation experiments at a 24 h timepoint to allow for maximal compound effect. Suitable assay conditions that report on the range of GFP signal were developed using 10 μM OSMI-4b or TMG treatment (Fig. 1f–g). Notably, the assay had a substantially greater dynamic range in the gain-of-signal direction with OSMI-4b than in the loss-of-signal direction with TMG. Changing the media glucose concentration altered basal GFP signal but did not affect the magnitude of GFP signal change upon inhibitor treatment (Supplementary Fig. 1a–b). The assay showed similar results with various nuclear stains, including Hoechst 33342 and SYTO Deep Red (Supplementary Fig. 1c–d). Under these screening conditions, 10 μM OSMI-4b produced an excellent median Z’ of 0.66 while 10 μM TMG produced a lower median Z’ of 0.28 across 34 assay plates (Fig. 1h, and Supplementary Data 1)20. Although the median Z’ for the TMG direction is comparatively low due to limited effect size associated with relatively high basal GFP levels, we proceeded as the assay was nonetheless generally reproducible across independent replicates (Fig. 1h, Supplementary Data 1). Screening of the Drug Repurposing Library was performed in duplicate at a concentration of 5–10 μM, revealing numerous compounds that reproducibly affect GFP signal from the GFP-β-OGT reporter (Fig. 1h, and Supplementary Data 1).
We filtered compounds for orthogonal validation by first examining cell viability and magnitude of GFP signal change. Cell viability was estimated using nuclear counts to identify potential artefactual results arising from cytotoxicity (Fig. 1i)21,22. Focusing on compounds that modulate OGT DI splicing without showing cytotoxicity-related nonspecific assay interference, we set cutoffs of (1) average viability of at least 25% across the two replicates and (2) either a 5σ increase or 3σ decrease in GFP signal, leading to cherrypick of 263 compounds (5.1% of library, Supplementary Data 1). These cherrypicked compounds were counter screened in wild-type HCT116 cells to remove autofluorescent compounds that interfere with the assay readout (Fig. 1j)21,23,24. As expected, OSMI-4b and TMG are not autofluorescent and significantly altered GFP signal in the GFP-β-OGT reporter (Fig. 1k). Most cherrypicked compounds were retained, with 226 of 263 compounds showing minimal autofluorescence (Fig. 1k, and Supplementary Data 1).
We next analyzed these 226 compounds by annotated mechanisms. Compounds annotated as ROCK inhibitors showed dichotomous effects, with selective ROCK inhibitors of varying scaffolds such as Y-33075 (also known as Y-39983), hydroxyfasudil, and thiazovivin reducing GFP signal while nonselective ROCK inhibitors showed no trend in GFP signal change (Fig. 1l, Supplementary Data 1). Other classes of compounds such as bromodomain inhibitors and HDAC inhibitors also emerged from this analysis, which may be due to the potential for nonspecific perturbation of the epigenetic machinery upstream of GFP biosynthesis23. Indeed, these compound families have been previously annotated to significantly perturb OGT and OGA and the proteome more broadly25. Inhibitors of transcriptional kinases such as CDK9, which have previously been found to affect OGT splicing26, showed minimal activity (as in LY2857785) in this assay or were unavailable for reordering (as in AT-7519; Supplementary Data 1). We thus focused on ROCK inhibitors for further investigation due to the suppression of productive OGT splicing in the GFP-β-OGT reporter cell line.
In vitro screening for OGT and OGA modulators
In parallel, we investigated whether members of the Drug Repurposing Library act directly on OGT or OGA using in vitro enzyme activity assays (Fig. 2a)27,28. We measured OGT activity using the UDP-Glo assay with hOGT, which contains 4.5 of 13.5 tetratricopeptide repeats (TPRs) and the catalytic domain of OGT27, and a CKIIα peptide substrate (Fig. 2b)29. Compounds were screened at 12.5–25 μM with 10 μM OSMI-4a as a positive control, which gave an excellent median Z’ of 0.70 (Fig. 2c; and Supplementary Data 2)20. Fifty-five inhibitors and 66 activators passed a 3σ or 2σ threshold, respectively, of which 45 inhibitory and 61 activatory primary hits were available for counter screening (2.0% of library). However, substitution of UDP-GlcNAc with UDP revealed that these compounds affect luminescence signal independently of OGT activity (Fig. 2d–e; and Supplementary Fig. 2a; Supplementary Data 2). These data imply that the Drug Repurposing Library compounds that regulate OGT DI splicing in cellular assays do not act directly on OGT.
a Flowchart of in vitro screening results. b Schematic of UDP-Glo assay design. c Screening results from the Drug Repurposing Library using the UDP-Glo assay. d Schematic of UDP-Glo counter screen design. e Counter screening results with cherrypicked OGT hit compounds across n = 2 replicates. f Schematic of 4-MU-GlcNAc assay design. g Screening results from the Drug Repurposing Library using the 4-MU-GlcNAc assay. h Schematic of 4-MU counter screen design. i Counter screening results with cherrypicked OGA hit compounds across n = 2 replicates. j Western blot analysis of OGA hit compounds’ effects on O-GlcNAc, OGT, and OGA levels in HEK293T cells, with indicated compound at 25 μM for 24 h (n = 3). Compounds that elevate O-GlcNAc highlighted in blue. Data are represented as mean ± s.d. One-way ANOVA with Dunnett’s multiple comparisons test with p-values shown. All Western blot data are representative of at least three independent biological replicates. Source data are provided as a Source Data file.
To screen for OGA modulators in vitro, we utilized the 4-MU-GlcNAc fluorogenic assay28 with recombinant human OGA (Fig. 2f). Compounds were screened at 12.5–25 μM with 10 μM TMG as a positive control, giving an excellent median Z’ of 0.89 (Fig. 2g; and Supplementary Data 3). TMG is also a member of the Drug Repurposing Library and was identified as the most potent inhibitor among the collection (Supplementary Data 3). We cherrypicked 54 inhibitory and 356 activatory primary hits from the 4-MU-GlcNAc activity assay that passed a 3σ threshold, of which 46 inhibitory and 338 activatory primary hits were available for counter screening (7.4% of library). Counter screening was performed by substitution of 4-MU-GlcNAc with 4-MU to remove compounds that affected fluorescence signal independently of OGA activity (Fig. 2h; and Supplementary Fig. 2b). Multiple compounds passed this counter screen, indicating that their effects are OGA-dependent (Fig. 2i, and Supplementary Data 3). Interestingly, there was little overlap between these hits and the GFP-β-OGT reporter screen.
The top 24 hits, comprised of 3 activatory hits and 21 inhibitory hits, were further validated by dose-response in the 4-MU assay, the 4-MU-GlcNAc activity assay, and differential scanning fluorimetry (DSF) as an orthogonal measure of OGA binding (Supplementary Fig. 3a–d). Ten compounds showed strong dose-dependent thermal stabilization of OGA and perturbation to OGA activity in vitro and were commercially available for cellular evaluation (Supplementary Fig. 3e). Two of these compounds were cytotoxic at 25 μM. Examination of the remaining eight compounds revealed that AKT inhibitor GSK69069330 and dopamine receptor agonist piribedil31 elevated cellular O-GlcNAc levels (Fig. 2j). Interestingly, while TMG treatment downregulates OGT and upregulates OGA, GSK690693 downregulates both OGT and OGA (Fig. 2j).
In differential scanning fluorimetry (DSF) experiments, piribedil and GSK690693 show dose-dependent thermal stabilization of OGA (2.16 ± 0.18 and 1.58 ± 0.05 °C at 50 µM, respectively) within 1.8–2.5-fold of TMG (3.91 ± 0.11 °C at 50 µM), indicating that these compounds are direct OGA ligands (Supplementary Fig. 3c–d). Piribedil is used in the treatment of Parkinson’s disease31 and we now show that piribedil additionally weakly inhibits OGA in vitro and elevates O-GlcNAc in cells (Supplementary Fig. 4a). However, piribedil bears structural resemblance to more potent OGA inhibitors and PET tracers, and was therefore not investigated further (Supplementary Fig. 4b)6,32,33,34,35,36. By contrast, to our knowledge, no scaffolds resembling GSK690693 have previously been reported to inhibit OGA and the cellular effects on OGT and OGA levels are distinct from the OGA inhibitor TMG. Thus, our in vitro screening efforts against OGA identified piribedil and GSK690693 as OGA inhibitors that elevate O-GlcNAc levels in cells, and we selected GSK690693 for further mechanistic investigation.
GSK690693 and Y-33075 elevate O-GlcNAc and downregulate OGT and OGA
Having identified AKT inhibitor GSK690693 from our OGA screen and ROCK inhibitors from the GFP-β-OGT reporter screen, we focused on these two families of kinase inhibitors for deeper mechanistic evaluation. We first queried the Drug Repurposing Hub for other library members with scaffolds similar to GSK690693 to identify structure–activity relationships17. Three other aminofurazan-azabenzimidazole kinase inhibitors (SB-747651A, SB-772077B, GSK269962) were identified and showed no activity against in the OGA in vitro screen (Fig. 3a, Supplementary Data 3). SB-747651A37,38 is an AKT inhibitor like GSK69069317,39 but shows no OGA inhibition in vitro up to 100 μM (Fig. 3b). Interestingly, GSK690693 but not SB-747651A elevates O-GlcNAc levels while downregulating OGT and OGA levels, suggesting that the O-GlcNAc activity of GSK690693 is independent of AKT inhibition (Fig. 3c). GSK690693 requires elevated concentrations and extended timepoints to affect OGT and OGA, in contrast to its reported activity against AKT30 (Fig. 3d–e, Supplementary Fig. 5a–b). Collectively, these experiments suggest that the O-GlcNAc-perturbing activity of GSK690693 is AKT-independent.
a Structures of GSK690693 and SB-747651A. Aminofurazan-azabenzimidazole core scaffold highlighted in light green. b Dose-response of indicated compound in 4-MU-GlcNAc activity assay (n = 3). c Western blot of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with indicated compound for 24 h (n = 3). d Quantification of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with indicated concentrations of GSK690693 for 24 h (n = 3). e Quantification of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with 50 μM GSK690693 over indicated timepoints (n = 3). f Structures of evaluated ROCK inhibitors. g Western blot of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with indicated compound for 24 h (n = 3). h Quantification of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with indicated concentrations of Y-33075 for 24 h (n = 3). i Quantification of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with 50 μM Y-33075 over indicated timepoints (n = 3). j Western blot analysis of O-GlcNAc, OGT, and OGA levels after treatment of HCT116 cells with indicated compounds for 24 h (n = 3). k Summary schematic of data in (j). Data are represented as mean ± s.d. One-way ANOVA with Dunnett’s multiple comparisons test (c, g) or Šidák’s multiple comparisons test (j) with p-values shown. All data are representative of at least three independent biological replicates. Data normalized to DMSO (b, c, d, g, h, j) or 0 h (e, i). Source data are provided as a Source Data file.
We additionally investigated ROCK inhibitors as a compound class that emerged from the GFP-β-OGT reporter screen (Fig. 1l, Supplementary Data 1). Six ROCK inhibitors of varying scaffolds were repurchased and evaluated for effects on O-GlcNAc, OGT, and OGA in wild-type HCT116 cells (Fig. 3f). Y-33075 slightly elevated O-GlcNAc levels on specific bands while decreasing both OGT and OGA levels, reminiscent of the effects seen with GSK690693 (Fig. 3g). Other ROCK inhibitors, including a closely matched structural analog Y-2763240,41,42, had no measurable effect on O-GlcNAc, OGT, or OGA levels. These results are in line with previous proteomics datasets finding that Y-33075 downregulates OGT but ripasudil does not25. Additionally, while previous reports show Y-33075 inhibits ROCK in vivo within hours41, Y-33075 requires elevated concentrations and extended timepoints to affect OGT and OGA (Fig. 3h, i, and Supplementary Fig. 5c–d). Notably, Y-33075 does not directly inhibit OGT or OGA (Supplementary Data 2; Supplementary Data 3). Collectively, these data indicate that Y-33075 indirectly affects O-GlcNAc and its cycling enzymes in a ROCK-independent manner.
At this juncture, we compared GSK690693 and Y-33075 for their effects on OGT, OGA, and O-GlcNAc levels in several additional studies. Simultaneous downregulation of OGT and OGA upon GSK690693 or Y-33075 treatment were also observed in HEK293T cells, demonstrating that these effects are generalizable beyond HCT116 cells (Supplementary Fig. 6a). As elevation of O-GlcNAc may result from upregulation of UDP-HexNAc, we quantified UDP-HexNAc from HCT116 cells treated with GSK690693 or Y-33075, or their respective controls SB-747651A and Y-27632. GSK690693 did not significantly alter UDP-HexNAc levels whereas Y-33075 treatment produced an 80% increase (Supplementary Fig. 6b), which may contribute to O-GlcNAc changes seen with Y-33075 (Fig. 3g). By contrast, GSK690693 likely elevates O-GlcNAc by directly inhibiting OGA and altering relative OGT and OGA levels. Interestingly, the downregulation of OGT and OGA due to GSK690693 and Y-33075 remains in the presence of OSMI-4b and TMG, indicating that the downregulation circumvents the endogenous DI feedback mechanism triggered by OGT or OGA inhibitors (Fig. 3j, k)11. Additionally, co-treatment with GSK690693 or Y-33075 with TMG potentiates upregulation of global O-GlcNAc, possibly due to suppression of TMG-induced OGA upregulation (Fig. 3j, k). In summary, GSK690693 and Y-33075, but not their structurally matched negative controls SB-747651A and Y-27632, downregulate OGT and OGA with concomitant O-GlcNAc elevation.
GSK690693 and Y-33075 regulate OGT and OGA transcripts
The downregulation of OGT and OGA induced by GSK690693 and Y-33075 could occur at the protein level or at the transcript level. To examine the protein-level angle, we evaluated OGT and OGA protein half-life by pulse-chase experiments with ribosome inhibitor cycloheximide (CHX). OGA is moderately destabilized by GSK690693 or Y-33075, while OGT stability is unaffected (Supplementary Fig. 7a). OGA destabilization by GSK690693, but not Y-33075, was partially rescued by neddylation inhibitor MLN4924 (Supplementary Fig. 7b). Unfortunately, parallel examination of the effects on the proteasome with MG132 or on E1 ubiquitin-activating enzymes with TAK-243 were not possible due to confounding cytotoxicity in the presence of GSK690693 or Y-3307543,44,45. These data indicate that there is some post-translational contribution to GSK690693- and Y-33075-induced downregulation of OGA but not on OGT. By contrast, analysis of effects from GSK690693 and Y-33075 on OGT and OGA transcripts by RT-qPCR showed substantial dose-dependent downregulation (Fig. 4a). Maximal downregulation (10–50-fold) of these transcripts was achieved at inhibitor concentrations that are similar to those required for efficacy at the protein level (Figs. 3d, h). We therefore further evaluated transcript-level regulation of OGT and OGA by GSK690693 and Y-33075.
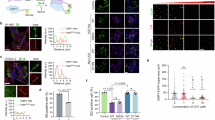
a RT-qPCR analysis of transcript abundance in HCT116 cells with indicated treatments in dose-response for 24 h (n = 3). b–d RT-qPCR analysis of transcript abundance for OGT (b), OGA (c), or with matched negative controls (d) in HCT116 cells treated as indicated for 24 h (n = 3). e Time course RT-qPCR analysis of transcript abundance in HCT116 cells treated with indicated compound (n = 3). f Schematic of ΔOGT-ISS cell line generation by CRISPR engineering. g Western blot of O-GlcNAc, OGT, and OGA levels in wild-type HCT116 cells after indicated treatments for 24 h (n = 3). h Western blot of O-GlcNAc, OGT, and OGA levels in ΔOGT-ISS HCT116 cells after indicated treatments for 24 h (n = 3). i RT-qPCR analysis of productive OGT transcript abundance in ΔOGT-ISS HCT116 cells after indicated treatments for 24 h (n = 3). y-Axes plotted on log scale (a–e, i) and linear scale (g, h). Data are represented as mean ± s.d. One-way ANOVA with Dunnett’s multiple comparisons test (b–d, g–i) with p-values for RT-qPCR experiments computed based on ACTB-normalized Cq values shown. All data are representative of at least three independent biological replicates. Data normalized to DMSO (a–d, g–i) or 0 h (e). Source data are provided as a Source Data file.
Deeper investigation of the effects of GSK690693 and Y-33075 on other OGT and OGA transcript isoforms revealed distinct effects in comparison to each other and OGT and OGA inhibitors. For OGT, OSMI-4b treatment downregulates DI1, DI2, and NMD transcripts while upregulating the productive transcript, whereas TMG acts on these transcripts in the opposite direction, in agreement with prior analyses (Fig. 4b)11. By contrast, GSK690693 and Y-33075 broadly downregulate DI1, DI2, and productive OGT transcripts. The NMD transcript was downregulated by GSK690693 but upregulated by Y-33075. These data demonstrate that GSK690693 and Y-33075 have distinct transcript-level effects on OGT in comparison to known OGT and OGA inhibitors.
Similarly, these compounds elicited distinct effects on OGA transcripts. For OGA, OSMI-4b treatment upregulates the NMD transcript and downregulates DI1/2, DI1, and productive transcripts, whereas TMG downregulates the NMD transcript and induces accumulation of the DI1 and productive OGA transcripts (Fig. 4c), in line with previous studies11. By contrast, GSK690693 and Y-33075 strongly downregulate NMD and productive transcripts, with more muted effects on the DI1/2 and DI1 species. These data also show that Y-33075 upregulates the OGT NMD transcript but downregulates the OGA NMD transcript, indicating that Y-33075 is not a general activator or inhibitor of NMD. Notably, GSK690693 and Y-33075 downregulate the productive OGA transcript in a manner similarly as OSMI-4b, despite having an opposite effect on cellular O-GlcNAc levels.
Further mechanistic evaluation with GSK690693 and Y-33075 was performed against the NMD and productive transcripts for OGT and OGA. Structurally matched negative control compounds SB-747651A and Y-27632 minimally perturb the NMD and productive transcript isoforms of OGT and OGA (Fig. 4d), in agreement with protein-level measurements (Figs. 3c, g). GSK690693 and Y-33075 affect OGT and OGA transcripts following siRNA knockdown of their respective primary targets AKT1/2 (AKT3 is not expressed in HCT116 cells) or ROCK1/2, further indicating that effects on OGT and OGA transcripts are independent of AKT and ROCK inhibition (Supplementary Fig. 8). Effects on OGT and OGA transcripts may be mediated through additional kinases inhibited by GSK690693 and Y-33075 but not SB-747651A and Y-27632. Indeed, GSK690693 and Y-33075 lower phosphorylation of multiple serine-arginine-rich (SR) splicing factors, whereas SB-747651A and Y-27632 show no effect (Supplementary Fig. 9). These data indicate that GSK690693 and Y-33075 possess splicing modulator activity against OGT and OGA transcripts that is independent of their primary annotated kinase targets.
We next investigated dependence of compound activity on the DI feedback mechanism. GSK690693 and Y-33075 achieve maximal effect on the NMD and productive transcripts after 3 h, in contrast to the DI feedback mechanism that is active within 2 h (Fig. 4e)11. The DI feedback mechanism is triggered for OGT by a cis-acting intronic splicing silencer (ISS) region between exons 4 and 5 that when removed in HCT116 cells (ΔOGT-ISS) lead to constitutive generation of the productive OGT transcript even in elevated O-GlcNAc conditions (Fig. 4f)10. As expected, OGT levels in ΔOGT-ISS cells are insensitive to OSMI-4b or TMG treatment relative to wild-type cells, yet OGT levels in ΔOGT-ISS cells remain sensitive to GSK690693 and Y-33075 (Fig. 4g, h), indicating that these compounds downregulate the productive OGT transcript in a distinct manner from the DI feedback mechanism activated by OGT and OGA inhibitors. Indeed, RT-qPCR experiments demonstrate that OSMI-4b and TMG do not affect productive OGT transcript levels in ΔOGT-ISS cells, but GSK690693 and Y-33075 downregulate the productive OGT transcript to a similar extent as in wild-type cells (Figs. 4b, i). Thus, GSK690693 and Y-33075 suppress the productive OGT transcript via a mechanism orthogonal to the previously reported cis-regulatory ISS, and this activity is independent of effects on the OGT NMD transcript, which does not exist in ΔOGT-ISS cells.
Y-33075 modulates splice site selection within OGT and OGA
Upon inspection of the DNA melt curves following RT-qPCR, we noticed that Y-33075 treatment induces a second peak in the melt curves for the RT-qPCR reactions for the productive OGT transcript and the OGA DI1 transcript (Fig. 5a). This activity is dose- and time-dependent (Supplementary Fig. 10a–b) and does not occur with Y-27632 treatment (Supplementary Fig. 10c). A minor shoulder in some RT-qPCR reactions targeting the productive OGA transcript may also be connected to the observations below but was not evaluated further (Fig. 5a, and Supplementary Fig. 10a–c).
a DNA melt curve analysis following RT-qPCR reactions using displayed primers from HCT116 cells treated as indicated for 24 h. One representative curve shown from three independent biological replicates. b– d Gel electrophoresis of RT-qPCR reactions using displayed primers from HCT116 cells treated as indicated for 24 h (n = 1). Secondary products generated by Y-33075 indicated with red asterisks. c– e Schematic of secondary products identified in (b) and (d). f Canonical (top) and alternative 5’ splicing (bottom) pathways for OGA. g Predicted splice acceptor and donor sites in OGA between exons 11 and 12. Borders of exon 11, DI2, exon 12, and sub-DI2 indicated. Sub-DI2 alternative 5’ splice site indicated with pink arrow. Figure generated using SpliceAI-Visual (https://mobidetails.chu-montpellier.fr/)47. h RT-qPCR analysis of transcript abundance in HCT116 cells treated as indicated for 24 h (n = 3). Data are represented as mean ± s.d. Two-tailed paired samples t-test with p-values computed based on ACTB-normalized Cq values shown. Data normalized to DMSO. i Gel electrophoresis of RT-qPCR reactions from HCT116 cells treated as indicated for 24 h (n = 1). Expected sub-DI2-containing products denoted by pink asterisks. ΔCq (vs. ACTB) values normalized to DMSO. For sequencing results, see Supplementary Fig. 11–14. Source data are provided as a Source Data file.
Electrophoresis of the crude RT-qPCR reactions and sequencing revealed the secondary OGT transcript product as the OGT NMD transcript (Fig. 5b, c, and Supplementary Fig. 11–12)11,46. Supporting this identification, the analogous DNA melt curves from ΔOGT-ISS cells that lack this region treated with Y-33075 do not show this second peak (Supplementary Fig. 10d). This observation agrees with previous results showing Y-33075-induced accumulation of the OGT NMD transcript (Fig. 4b– d, e). This secondary product is observable to a lesser extent with TMG treatment, which also upregulates the OGT NMD transcript and downregulates the productive OGT transcript (Figs. 4b, and 5a, b). Redesign of primers targeting the exon 4–exon 5 junction in the productive OGT transcript with higher fidelity shows a 1.7-fold greater reduction in productive OGT transcript levels induced by Y-33075 than previously measured (Supplementary Fig. 10e,f). Analogously, we isolated and sequenced the secondary product in the RT-qPCR reaction targeting the OGA transcript containing only DI1 (Fig. 5d,e, Supplementary Fig. 13–14). This product retains a 227 bp region of DI2, which we term sub-DI2, corresponding to an alternative 5’ splicing event within DI2 (Fig. 5f). This alternative splice site has been computationally predicted but not previously detected experimentally (Fig. 5g)47,48. The sub-DI2-containing OGA transcripts are low abundance endogenously and contain DI1, implying nuclear detention of these transcripts (Fig. 5h,i). In summary, these data demonstrate that Y-33075 affects splice site selection, promoting the inclusion of the OGT poison exon and inducing an alternative 5’ splice site in OGA.
GSK690693 inhibits intron processing and Y-33075 modulates exon inclusion
The effects of GSK690693 and Y-33075 on the O-GlcNAc cycling enzymes at the protein and transcript levels were compared to previously characterized splicing modulators. Namely, we evaluated kinase activators and inhibitors that affect splicing factor phosphorylation (RECTAS, SRPIN340, TG003), SMN2 RNA–U1 snRNP molecular glue branaplam, RBM39 molecular glue degrader indisulam, PRMT5 inhibitor GSK3326595, CDK11 inhibitor OTS964, SF3B1 degrader PROTAC-O4I2, and DYRK inhibitors EHT1610 and GNF213349,50. Indisulam, OTS964, and GNF2133 downregulate both OGT and OGA at the protein level (Fig. 6a). Interestingly, at the transcript level, indisulam and GNF2133 upregulate the OGT NMD transcript similarly as Y-33075, while OTS964 shows downregulation across the nonproductive and productive transcript isoforms of OGT and OGA similarly as GSK690693 (Fig. 6b). We evaluated if knockdown of the mechanistic target of the most potent compound OTS964, CDK11, could recapitulate the effects of the molecule (Fig. 6c). CDK11 knockdown resulted in a comparatively minor effect on the OGT NMD transcript but did not downregulate the other OGT and OGA transcripts profiled, indicating that the effects of OTS964 on OGT and OGA are not mediated through inhibition of CDK11 activity alone (Fig. 6d).
a Western blot of O-GlcNAc, OGT, and OGA levels in HCT116 cells after indicated treatments for 24 h (n = 3). b RT-qPCR analysis of transcript abundance in HCT116 cells with indicated treatments for 24 h (n = 3). c Western blot of CDK11 levels in HCT116 cells (n = 2). d RT-qPCR analysis of HCT116 cells with or without CDK11 knockdown (n = 2). Data are represented as mean ± s.d. HCT116 cells were transfected with siRNA for 48 h. One-way ANOVA with Dunnett’s multiple comparisons test with p-values shown (a, b). p-values computed based on ACTB-normalized Cq values for RT-qPCR experiments (b). All Western blot data are representative of at least two independent biological replicates. Data normalized to DMSO (a, b) or average of siCTRL #1 and siCTRL #2 (c, d). Source data are provided as a Source Data file.
To holistically profile the effects of GSK690693 and Y-33075 on the transcriptome, we performed RNA-seq on HCT116 cells treated with 50 μM GSK690693, SB-747651A, Y-33075, or Y-27632 for 24 h. In differential expression analysis, GSK690693 and Y-33075 result in significantly greater perturbations to the transcriptome than SB-747651A and Y-27632 (Supplementary Fig. 15, Supplementary Data 4). In agreement with previous RT-qPCR data, RNA-seq data showed downregulation of the productive OGT and OGA transcripts, elevated inclusion of the OGT poison exon with Y-33075, elevated intron retention within OGA with GSK690693, and minimal changes with SB-747651A and Y-27632 (Figs. 4b–d, 7a, and Supplementary Fig. 16). Additionally, RNA-seq reveals that Y-33075 promotes both the previously characterized sub-DI2 (Fig. 5d−i), and an additional isoform arising from a mutually exclusive exon splicing pattern incorporating a 59 bp segment of OGA DI2 between exons 10 and 12 (Fig. 7a, and Supplementary Fig. 17), corresponding to the aforementioned computationally predicted cryptic splice sites (Fig. 5g). This transcript is low abundance in DMSO control (Fig. 7a) and contains a PTC that would subject the transcript to NMD (Supplementary Fig. 17). GSK690693 and Y-33075 do not grossly affect the expression or splicing of housekeeping gene transcripts such as ACTB and GAPDH (Supplementary Fig. 18). Thus, RNA-seq data recapitulated our previous observations by RT-qPCR for effects on OGT and OGA transcripts and improved our understanding of the Y-33075-induced cryptic splice sites within OGA.
a Sashimi plots of OGT and OGA from HCT116 cells treated with 50 μM of indicated compound for 24 h (n = 3). b Significant events identified from rMATs alternative splicing analysis of HCT116 cells with indicated treatments for 24 h (n = 3). RI, retained intron; MXE, mutually exclusive exon; A3SS/A5SS, alternative 3’/5’ splice site; SE, skipped exon. For tabulation of all events, see Supplementary Data 5. c Sashimi plots of GNL3. HCT116 cells treated with DMSO or 50 μM GSK690693 for 24 h (n = 3). d MAJIQ analysis of retained introns induced by 50 μM GSK690693 treatment in HCT116 cells for 24 h (n = 3). e Volcano plot of PEs affected by 50 μM Y-33075 treatment in HCT116 cells for 24 h (n = 3). Select genes containing PEs labeled. f Sashimi plots of DDX17 and SMNDC1 from HCT116 cells treated with DMSO or 50 μM Y-33075 for 24 h (n = 3). g Endpoint RT-PCR analysis of OGT, DDX17, SMNDC1, ACTB transcripts in HCT116 cells treated with indicated concentrations of Y-33075 for 24 h (n = 3). h Schematic summary of effects of GSK690693, Y-33075, and piribedil on OGT and OGA at the transcript and protein level. All data are representative of at least three independent biological replicates. Source data are provided as a Source Data file.
We further analyzed global changes to alternative splicing induced by these compounds by rMATS analysis (Fig. 7b, and Supplementary Data 5). Strikingly, GSK690693 and Y-33075 result in 5211 and 4530 significant events, respectively, whereas negative controls SB-747651A and Y-27632 result in only 48 and 11 significant events, respectively. A large portion of alternative splicing events induced by GSK690693 are retained introns (GSK690693, 34%; Y-33075, 7%; Fig. 7b, c, and Supplementary Fig. 19, Supplementary Data 5). Using MAJIQ analysis to include unannotated de novo retained intron events, GSK690693 induces 1597 retained introns of which 1063 (67%) are de novo (Fig. 7d, and Supplementary Data 6). Thus, GSK690693 is likely a general splicing inhibitor rather than a modulator of specific intron retention processes, resembling the CDK11 inhibitor OTS964 that also induces widespread intron retention and disrupts O-GlcNAc homeostasis (Fig. 6a, b)51, although the activity profile of GSK690693 does not completely overlap with OTS964 (Supplementary Fig. 20).
In contrast to GSK690693, Y-33075 affects skipping or inclusion of specific cassette exons (GSK690693, 54%; Y-33075, 82%; Fig. 7b). Analysis of the splice sites flanking exons skipped or included upon Y-33075 treatment did not reveal any notable differences versus genome-wide splice site consensus sequences, suggesting Y-33075 does not directly modulate interactions between RNA and the spliceosome (Supplementary Fig. 21)52. Notably, many of the Y-33075-induced exons are PEs53, including that of OGT which is among the most strongly induced (Fig. 7e). Y-33075 additionally induces PEs within splicing regulators such as the splicing factors SRSF3 and HNRNPDL (Fig. 7e, and Supplementary Data 7). Remarkably, Y-33075 promotes inclusion of 30 of the 31 PEs that are significantly affected (Fig. 7e). NMD inhibition is known to promote PE inclusion as these transcripts are no longer degraded; however, for many of the genes in which Y-33075 induces PE inclusion, as in the case of OGT or DDX17, the total transcript level decreases, in line with previous data that Y-33075 is not a general NMD activator or inhibitor (Fig. 7a, f, and Supplementary Data 4,7)54,55.
To validate the RNA-seq results, we performed endpoint RT-PCR for some of the most strongly included PEs: OGT, DDX17, and SMNDC1 (Fig. 7g). DDX17 is an RNA helicase56, and SMNDC1 is a splicing factor57. These proteins and other family members are implicated in splicing regulation during biological processes such as differentiation56 and spliceosome formation57, as well as diseases such as cancer58 and spinal muscular atrophy (SMA)49,50. In agreement with the RNA-seq analysis, Y-33075 promotes PE inclusion in each of these transcripts at 25–50 μM (Figs. 7f, g). By contrast, indisulam and GNF2133 promote PE inclusion in OGT but not DDX17 and SMNDC1 (Supplementary Fig. 22). Thus, Y-33075 is a mechanistically distinct splicing modulator that specifically remodels the exome and opens opportunities for small molecule control over transcript-level regulation of biologically and therapeutically relevant targets.
Discussion
We report the results of three parallel high-throughput screening campaigns using the Broad Institute Drug Repurposing Library aimed at modulating O-GlcNAc (Fig. 7h). From an engineered minigene reporter cell line screen, we found that ROCK inhibitor Y-33075 is a modulator of OGT DI splicing. Complementary in vitro enzymatic assay screens revealed that dopamine receptor agonist piribedil and AKT inhibitor GSK690693 are OGA inhibitors, the latter of which represents a previously unknown OGA ligand scaffold. Further cellular evaluation revealed that GSK690693 and Y-33075 disrupt a homeostatic DI feedback mechanism that regulates OGT and OGA protein levels, resulting in the downregulation of both OGT and OGA while elevating global O-GlcNAc. These effects are driven by alternative splicing events that are independent of AKT and ROCK, as shown through structurally matched negative controls SB-747651A and Y-27632 and siRNA knockdown of AKT and ROCK. Transcriptome-wide profiling and alternative splicing analyses demonstrated that GSK690693 leads to widespread intron retention while Y-33075 modulates inclusion of specific cassette exons. We additionally find that Y-33075 promotes the usage of cryptic splice sites within OGA leading to previously uncharacterized transcript isoforms, motivating future studies of whether these splice sites have a regulatory role that can be triggered by endogenous stimuli. Thus, GSK690693 and Y-33075 add two chemotypes to the growing repertoire of small molecule splicing modulators. Further investigation of other small molecule splicing modulators revealed a set of compounds—OTS964, indisulam, and GNF2133—that affect OGT and/or OGA transcripts with mid-nanomolar to low-micromolar potency, demonstrating that O-GlcNAc homeostasis is sensitive to diverse splicing perturbations. Collectively, these investigations add to previous literature connecting O-GlcNAc to RNA splicing10,11,18,19,26 and kinase inhibitors to transcriptional modulation49,50, and survey small molecules that can disrupt the conserved alternative splicing mechanism in OGT and OGA transcripts that governs O-GlcNAc homeostasis.
The splicing modulators profiled here are unlikely to be selective probes for O-GlcNAc biology prior to mechanistically untangling their molecular targets. While this is likely to be challenging for GSK690693 and Y-33075, as these compounds display weak splicing modulatory activity that could arise from polypharmacology, such efforts may be facilitated by focusing on and comparing profiles to the more potent splicing modulators identified. For example, OTS964 inhibits intron removal in a manner similar to GSK690693 and also shows similar effects on OGT and OGA transcripts, while indisulam and GNF2133 promote PE inclusion in OGT similar to Y-33075, indicating that there may be shared mechanistic targets. However, the activity profiles of OTS964, indisulam, and GNF2133 do not completely overlap with those of GSK690693 and Y-33075, which may be of interest to the development of splicing modulators more broadly. For example, Y-33075, but not indisulam and GNF2133, promotes PE inclusion in SMNDC1 and DDX17, which have been implicated in splicing regulation and disease. Examination of these more potent splicing modulators together with structure–activity relationship studies on GSK690693 and Y-33075 are important future directions to fully illuminate the mechanisms of these splicing modulators. Our characterization of SB-747651A and Y-27632 as matched negative controls provides a starting point for these campaigns.
As mechanistic investigation into splicing modulators of OGT and OGA progresses, disrupting the cellular DI feedback mechanism may become a strategy to increase the therapeutic efficacy of targeting O-GlcNAc. Preclinical and clinical studies of OGA inhibitors to treat neurodegenerative diseases have noted that a high degree (> 80%) of OGA target occupancy is requisite for significant O-GlcNAc elevation6,33,59, which may be a consequence of OGA upregulation in response to OGA inhibition via the DI feedback mechanism. Thus, antagonizing the cellular DI feedback response could be synergistic with increasing O-GlcNAc with OGA inhibitors. Alternative splicing of OGT has been associated with maintaining cell viability in colorectal cancer10, which could be another angle to explore therapeutic applications of targeting O-GlcNAc homeostasis. More extensive characterization and optimization of small molecule splicing modulators to target O-GlcNAc homeostasis will enable future studies in relevant models to test these hypotheses15. In conclusion, our discoveries here point to the potential in targeting OGT and OGA splicing and will spur additional investigations into the interplay between splicing and O-GlcNAc homeostasis more broadly.
Methods
Protein purification
OGT
Plasmid was transformed into BL21 (DE3) E. coli cells. Overnight starter cultures were used to inoculate cultures in fresh LB media supplemented with 50 μg/mL kanamycin sulfate at a 1:50 ratio then grown at 37 °C to an OD600 of 1.1. Cells were then grown at 16 °C for 30 min and induced with 0.2 mM IPTG for overnight protein expression. Cultures were pelleted and resuspended in lysis/wash buffer (20 mM Tris pH=7.4, 150 mM NaCl, 40 mM imidazole) supplemented with 100 μg/mL chicken egg white lysozyme, DNAse I, and 1X cOmplete™ EDTA-free Protease Inhibitor Cocktail then incubated at 4 °C for 1 hour with rotation. Lysates were sonicated on ice (10 seconds on, 5 seconds off, 7.5 min total, 25% amplitude) then centrifuged at 15000 rcf for 10 min at 4 °C. Clarified were filtered through 0.45 μm filters then purified by fast protein liquid chromatography (FPLC). Specifically, nickel affinity chromatography was performed with HiTrap Chelating HP using a gradient of 20 mM Tris pH=7.4, 150 mM NaCl with imidazole concentrations ranging from 40 mM to 500 mM and size exclusion chromatography with HiLoad® 16/600 Superdex® 200 pg in 20 mM Tris pH=7.4, 150 mM NaCl. 1 mM tris(3-hydroxypropyl)phosphine (THPP) was added to the eluted protein. Protein was concentrated using Vivaspin 20 MWCO 50 000 then aliquoted, flash frozen, and stored at -80 °C until use. Protein aliquots were thawed on ice and centrifuged at 21,330 rcf for 10 min at 4 °C prior to use.
Plasmid for His-tagged hOGT4.5 construct was a gift from Suzanne Walker.
OGA
Purification of His-tagged full-length OGA was performed as described for OGT with minor modifications. Specifically, the cultures were cooled to 16 °C at an OD600 of 0.8 instead of 1.1, and lysates were sonicated for 10 min total at 30% amplitude. Protein aliquots were thawed on ice and centrifuged at 21,330 rcf for 10 min at 4 °C prior to use.
Plasmid for His-tagged full-length OGA was a gift from David Vocadlo.
In vitro activity assay for OGT
Reactions were carried out using 750 nM His-hOGT, 100 μM Ultra Pure UDP-GlcNAc, and 50 μM CKIIα peptide KKKYPGGSTPVSSANMM in 20 mM Tris (pH=7.4), 150 mM NaCl, 1 mM THPP, 12.5 mM MgCl2, and 0.24 mg/mL bovine serum albumin (BSA). 10 μL of 1.5 μM His-hOGT was plated into white 384-well plates using a Multidrop Combi plate dispenser, 100 nL of compound stock solution was added with a CyBio Well vario, then 10 μL of 200 μM Ultra Pure UDP-GlcNAc and 100 μM CKIIα peptide was added using a Multidrop Combi plate dispenser. Reactions were incubated at room temperature for 1 h in the dark.
UDP-Glo enzyme was diluted 75X with enzyme dilution buffer to prepare UDP-Glo working solution. UDP-Glo working solution was diluted 100X with nucleotide detection reagent, and this mixture was in turn diluted 3X with PBS to generate UDP detection reagent. 10 μL UDP detection reagent was added to each well using a Multidrop Combi plate dispenser. After 1 hour, luminescence measurements were taken on an EnVision 2105 plate reader. Luminescence measurements were corrected for background and cross-talk by subtracting the average signal from blank wells on each plate. Negative control experiments were performed with UDP in place of UDP-GlcNAc. See also Supplementary Table 1 for screening information.
In vitro activity assay for OGA
Reactions were carried out using 100 nM His-OGA and 500 μM 4-MU-GlcNAc in 20 mM Tris (pH=7.4), 150 mM NaCl, 1 mM THPP, and 0.1 mg/mL BSA. 10 μL of 200 nM His-OGA was plated into black 384-well plates using a Multidrop Combi plate dispenser, 100 nL of compound stock solution was added with a CyBio Well vario, then 10 μL of 1 μM 4-MU-GlcNAc was added using a Multidrop Combi plate dispenser. Reactions were incubated at room temperature for 1 h in the dark, then quenched with 20 μL 3 M glycine/NaOH buffer (pH=10.3) added using a Multidrop Combi plate dispenser. Fluorescence measurements were taken on a SpectraMax M5 plate reader or SpectraMax iD5 plate reader using a 360 nm excitation wavelength and 460 nm emission wavelength. Negative control experiments were performed with 4-MU in place of 4-MU-GlcNAc. See also Supplementary Table 1 for screening information.
Differential scanning fluorimetry (DSF)
DSF experiments were performed using 1 µM His-OGA in 100 mM imidazole (pH=7.0), 150 mM NaCl, 1 mM THPP and 10X SYPRO Orange in sealed MicroAmp Optical 384-Well Reaction Plates. Experiments were performed on a ThermoFisher QuantStudio 7 with heating from 25.0 °C to 65.0 °C at a rate of 2 °C min-1. Fluorescence was detected using a 470 nm excitation wavelength and 586 nm emission wavelength. ΔTm values calculated based on difference versus DMSO-treated wells. Data was analyzed using Protein Thermal Shift Software 1.4.
Cell culture and Western blotting
HEK293T or HCT116 cells were cultured in DMEM supplemented with 10% heat-inactivated fetal bovine serum (FBS) and 1X penicillin/streptomycin (P/S). Cells were grown in a 37 °C humidified incubator with 5% CO2. 1.5e6 cells were plated in each well of treated 6-well plates (Western blot, RNA experiments) or 6e5 cells were plated in each well of treated 12-well plates (RNA experiments) in DMEM supplemented with 10% FBS and 1X P/S then allowed to grow for 24 hours. Cells were treated with indicated compounds for indicated timepoints then collected by trypsinizing and one PBS wash. Cell pellets for RNA analysis were washed with PBS an additional time. Cell pellet was lysed or frozen at -80 °C. Wild-type HEK293T and HCT116 cell lines sourced from ATCC. GFP-β-OGT HCT116 cells18,19 and ΔOGT-ISS HCT116 cells10 prepared as described.
For Western blot analysis, cell pellets were lysed with 2% sodium dodecyl sulfate (SDS) in PBS supplemented with 50 μM TMG and 1X protease/phosphatase inhibitor cocktail. Protein concentration was determined using bisinchoninic acid (BCA) assay, then lysates were supplemented with 5X Laemmli buffer to a final concentration of 50 mM Tris, 2% (m/v) SDS, 7% (v/v) glycerol, 0.0044% (m/v) bromophenol blue, and 0.14 M β-mercaptoethanol. Lysates were boiled at 95 °C for 5 min then 15 μg was run on 12% Criterion™ TGX™ Precast Midi Protein Gel or 6% –12% tris-glycine gels. Proteins were transferred to nitrocellulose membranes then blocked with either 5% BSA or 5% milk for 1 h at room temperature then incubated with primary antibody overnight at 4 °C. Membranes were washed 3 times with 1X TBST for 5 minutes each, then incubated with secondary antibody for 1 h at room temperature, washed 3 times with 1X TBST for 5 minutes each again, then imaged on Azure 400, 600, or c600.
High-content screening
For screening, 2500 GFP-β-OGT HCT116 cells were plated in 50 μL DMEM supplemented with 10% FBS and 1X P/S in each well of black clear bottom 384-well plates using a Multidrop Combi plate dispenser. For pilot experiments, 10,000 cells were plated in 50 μL DMEM supplemented with 10% FBS and 1X P/S in each well of black clear bottom 96-well plates. Cells were grown in a 37 °C humidified incubator with 5% CO2. For screening, 24 hours later, 100 nL of compound stock solution was added with a CyBio Well vario then plates were returned to the incubator. For pilot experiments, 24 hours later, 50 μL of 2X compound stocks in DMEM supplemented with 10% FBS and 1X P/S were added then plates were returned to the incubator. Following 24 hours, media was aspirated using a BioTek 405 TS microplate washer. Cells were fixed with 50 μL neutral buffered formalin, washed with PBS two times, stained with Hoechst 33342 or SYTO Deep Red, then washed with PBS two times. Plates were imaged on an Operetta CLS system using a 20X air objective. Images were analyzed on Harmony 4.9 software. Hoechst 33342 or SYTO Deep Red staining was used to identify nuclei regions and count total nuclei and average nucleus GFP signal in each well. Cell viability was estimated with nuclei counts per well. Edge wells (columns 1 and 24) were filled with cells and media and excluded from analysis. Top and bottom rows were filled with compounds and were included in analysis. See also Supplementary Table 1 for screening information.
UDP-HexNAc quantification
10e6 HCT116 cells were plated in 10 mL DMEM supplemented with 10% FBS and 1X P/S in 10 cm plates. Cells were grown in a 37 °C humidified incubator with 5% CO2. 24 h later, cells were treated with compound or vehicle. After 24 h, cells were collected by trypsinizing and two PBS washes then flash frozen and stored at -80 °C. Pellets were lysed in 1 mL PBS by sonicating 15% amplitude for 1 min, boiling at 95 °C for 15 min, freezing at -80 °C for 30 min, then boiling at 95 °C for 15 min. 500 μL of each sample was transferred to new 1.5 mL tubes and protein was precipitated in 4:4:1 methanol:water:chloroform mixture. The aqueous supernatants (lysate extracts) were transferred to new 1.5 mL tubes and dried by speedvac then resuspended in 800 μL milliQ water. 125 μL methanol was added to protein pellets, samples were centrifuged 10,000 rcf for 1 min, then supernatant was removed. Protein was resuspended in 2% SDS in PBS and quantified by BCA assay.
Volume of lysate extracts corresponding to 200 μg protein were transferred to new 1.5 mL tubes and dried by speedvac. Samples were resolubilized in 400 µL milliQ water by vortexing for 2 min. The redissolved samples were passed through a 0.2 µm filter prior to UPLC–MS/MS analysis with a Waters Acquity UPLC H-Class System and a Waters Xevo TQ-S mass spectrometer. Chromatography was performed using an Acquity UPLC BEH C18 1.7 µm (2.1 ×50 mm) column with mobile phase A 30 mM ammonium acetate in water and mobile phase B 0.1% formic acid in acetonitrile. The gradient used was as follows: 0-3 min 2-70% B, 3-4 min 70-90% B, 4-4.5 min 90-2% B, 4.5-5.5 min 2% B isocratic. Column temperature was 40 °C, flow rate was 0.3 mL/min, and injection volume was 6 µL. Chromatography conditions adapted from ref.60.
UDP-HexNAc was detected in positive mode using electrospray ionization (MRM parent ion m/z 607.7928, daughter ion m/z 203.9614). Cone voltage was 38 V and collision voltage was 14 V. Standards of UDP-GlcNAc and UDP-GalNAc were used to optimize UPLC-MS/MS conditions and eluted at 0.45 min with this method. Peak integrations were obtained using MassLynx v4.2 software. Response plotted for each sample is the average peak area from two technical duplicate injections.
Reverse transcriptase-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was isolated using a RNeasy Mini Kit according to the manufacturer’s protocol. RT-qPCR was performed using Luna® Universal One-Step RT-qPCR Kit according to the manufacturer’s protocol using 200 ng template RNA. Experiments were performed on a CFX Connect Real-Time PCR Detection System instrument in sealed 96-well PCR plates. Reaction plates were stored at -20 °C until further analysis. Cq values were normalized to actin (ACTB) as an internal control. Primers utilized are indicated in Supplementary Table 2. Primers targeting OGT and OGA were based on previous reports11.
Sequencing
Crude RT-qPCR reaction mixtures were mixed with 6X loading dye then separated on a 2% agarose gel and stained with ethidium bromide. DNA was purified using Monarch® DNA Gel Extraction Kit. Sanger sequencing was performed using Quintara Biosciences services (www.quintarabio.com).
siRNA knockdown
7.5e5 cells were plated in each well of treated 6-well plates in DMEM supplemented with 10% FBS and 1X P/S then allowed to grow for 24 h. 10 µL Lipofectamine™ RNAiMAX Transfection Reagent was incubated with 250 µL Opti-MEM™ for 15 min at room temperature to form liposomes. 10 µL of 20 µM siRNA resuspended in nuclease water was diluted into 250 µL Opti-MEM™ and added to the liposomes then incubated for 15 min at room temperature. 520 µL of the siRNA-containing liposomes was added dropwise to the cells. Cells were grown in a 37 °C humidified incubator with 5% CO2 then treated with indicated compounds for indicated timepoints then collected by trypsinizing and one PBS wash. Cell pellet was lysed or frozen at -80 °C. The list of siRNA products used is provided in Supplementary Table 3.
RNA-seq
Libraries were prepared using a SciClone G3 NGSx workstation (PerkinElmer) using the Kapa mRNA HyperPrep kit (Roche). Polyadenylated mRNAs were captured using oligo-dT-conjugated magnetic beads (Kapa mRNA HyperPrep kit) from 400 ng of total RNA on a Revvity SciClone G3 NGSx automated workstation. Poly-adenylated mRNA samples were immediately fragmented to 300–400 bp using heat and magnesium. First-strand synthesis was completed using random priming followed by second-strand synthesis and A-tailing. A dUTP was incorporated into the second strand to allow strand-specific sequencing of the library. Libraries were enriched and indexed using 10 cycles of amplification (Kapa mRNA HyperPrep kit) with PCR primers, which included dual 8 bp index sequences to allow for multiplexing (IDT for Illumina unique dual 8 bp indexes). Excess PCR reagents were removed through magnetic bead-based cleanup using Kapa Pure magnetic beads on a SciClone G3 NGSx workstation (PerkinElmer). The resulting libraries were assessed using a 4200 TapeStation (Agilent) and quantified by QPCR (Roche). Libraries were pooled and sequenced on one Illumina NovaSeq X Plus 10B lane using paired-end, 150 bp reads.
Paired-end RNAseq reads were trimmed with Trim Galore v 0.6.4 (https://github.com/FelixKrueger/TrimGalore) with a quality value cutoff of 20.
Intron-exon read mapping
Genome and annotation for human hg38 were downloaded from Encode (release 40). To account for splice variants and avoid over-counting, bedtools merge was used with the settings -s -c 4,5,6 -o distinct to combine overlapping introns and exons into “intron regions” and “exon regions”. Using aligned BAM files from STAR 2-pass alignment step, the number of reads mapping to intron regions vs. exon regions vs. whole gene regions were calculated using bedtools coverage (v 2.31) with the setting -F 1.0 to only count reads that were completely mapped within a given region. Custom Python scripts were used to aggregate read mapping counts for each region (intron region, exon region, whole gene) per gene (https://github.com/harvardinformatics/wooLab_RNAseq/tree/main). In cases where exon read counts were larger than gene read counts from this analysis, due to merging of overlapping or bookended exons from adjacent genes, exon read counts were set to the gene counts minus the intron read counts in order to appropriately apportion merged counts to the respective source gene. Percents of total gene reads mapped to exon regions and intron regions are plotted in Supplementary Fig. 19 (for genes with at least 1000 reads in each replicate of each treatment condition) and enumerated in Supplementary Data 8.
Differential expression analysis
To quantify transcript expression, Salmon61 (v 1.10.2; https://salmon.readthedocs.io/en/latest/) was used in the mapping-based mode and using the refgenie (http://refgenomes.databio.org/) pre-built “full-sa” index for hg38, which includes full decoy sequences to improve quantification accuracy. Differential expression was calculated using the DESeq262 library in R; differential expression for individual transcripts was calculated and the dataset was truncated to only include MANE Select transcripts (https://ftp.ncbi.nlm.nih.gov/refseq/MANE/MANE_human/current/) to minimize biasing due to minor alternatively spliced transcripts (see Supplementary Fig. 19, Supplementary Data 8, Fig. 7b)63. Each of our treatment conditions (GSK690693, SB-747651A, Y-33075, Y-27632) was compared in a pairwise fashion versus DMSO control to get differential expression. To determine p-values, DESeq2 uses two-tailed Wald test with Benjamini-Hochberg FDR correction.
Alternative splicing analysis
For rMATS64 analysis, trimmed reads were aligned using STAR in ‘two pass basic’ mode using additional rmats recommended parameters ‘--chimSegmentMin 2 --outFilterMismatchNmax 3 --alignEndsType EndToEnd --outSAMstrandField intronMotif --alignIntronMax 299999.’ Differentially spliced events were computed using rmats-turbo (v4.3.0) using the --novelss and --variable-read-length options. Only reads that span the splice junction were included for enumeration of isoform counts. High-confidence significant alternative splicing events were considered to have (1) an absolute inclusion level difference of at least 20% versus DMSO control, (2) a false discovery rate (FDR) less than 0.05, and (3) at least 50 junctional reads averaged across 3 replicates for both alternatively spliced isoforms in either the DMSO control or treatment condition. Visualization of events in sashimi plots as the average of three independent biological replicates was performed using rmats2sashimiplot (v3.0.0; https://github.com/Xinglab/rmats2sashimiplot). rMATS uses a binomial distribution to model the read count associated with each splice isoform given the inclusion level in each individual replicate and a logit-normal distribution to model the variation among replicates within each sample group. p-values were then determined using a two-tailed likelihood-ratio test with Benjamini-Hochberg FDR correction.
For MAJIQ65 analysis, trimmed reads were aligned to the primary assembly of GENCODE human reference genome (GRCh38 v40) using STAR66 2.7.11a in ‘two pass basic’ mode. The ‘build’ step of MAJIQ (using v2.5.1) was performed requiring events to be detectable in at least 2 replicates and supported by at least 50 reads. Local splicing variations from MAJIQ were modulized to obtain binary events. Events with probability_changing of ≥0.9 and at least 20% change in median dpsi were considered significant.
Exon splice site motif analysis
bedtools getfasta was used to extract the 20 bp upstream and downstream of the exon start site and exon end site for each exon in the hg38 annotation of the genome and generated consensus sequences for the splice site regions using WebLogo (https://weblogo.berkeley.edu/logo.cgi)67. The same was done for each of the skipped exons significantly affected by Y-33075 treatment. Duplicate sequences representing the same exon start or end site arising from the same exon being included in multiple transcript isoforms were excluded from the analysis.
Poison exon analysis
Skipped exon events induced by Y-33075 treatment were filtered for at least 50 junctional reads averaged across 3 replicates for both alternatively spliced isoforms in either the DMSO control or treatment condition. Events were annotated as likely representing poison exon events if the start side and end site of the skipped exon event were each within 100 bp of previously annotated poison exons53. Genes were annotated as involved in splicing based on cross-referencing to a curated list of splicing-related proteins68.
For analysis of poison exons, events with overlapping genomic coordinates were further curated to remove duplicates. The event with the greatest alignment to previously annotated genomic coordinates was included in downstream analyses. Alternatively, if the same skipped exon was identified from multiple transcript isoforms, the event with more associated reads, i.e., the major isoform, was included in downstream analyses.
Endpoint reverse transcriptase-polymerase chain reaction (RT-PCR)
RNA was isolated as in RT-qPCR method. RT-PCR was performed on a T100 Thermal Cycler using 200 ng template RNA with the following protocol: 55 °C for 10 min; 95 °C for 1 min; 35 cycles of 95 °C for 30 seconds, 60 °C for 30 seconds, and 68 °C for 45 seconds; 72 °C for 5 min. Reaction plates were stored at -20 °C until further analysis. PCR products were mixed with 6X loading dye then separated on a 2% agarose gel and stained with ethidium bromide. Primers utilized are indicated in Supplementary Table 3.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study have been deposited in the ENA database under accession code PRJEB101908. Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper.
Code availability
Custom code used in this study can be found in GitHub repositories from the respective Methods section. These include previously reported code for trimming RNA-seq reads (https://github.com/FelixKrueger/TrimGalore) and for generating sashimi plots (https://github.com/Xinglab/rmats2sashimiplot), as well as code developed expressly for this study to map RNA-seq reads to exon/intron/gene regions (https://github.com/harvardinformatics/wooLab_RNAseq/tree/main).
References
Wulff-Fuentes, E. et al. The human O-GlcNAcome database and meta-analysis. Sci. Data 8, 25 (2021).
Zachara, N. E. et al. Ch. 19, (Cold Spring Harbor Laboratory Press, 2022).
Cheng, S. S., Mody, A. C. & Woo, C. M. Opportunities for therapeutic modulation of O-GlcNAc. Chem. Rev. 124, 12918–13019 (2024).
15th Conference Clinical Trials Alzheimer’s Disease, November 29- December 2, 2022, San Francisco, CA, USA: Symposia - Oral Communications - Late Breaking Abstracts (Clinical Trial Alzheimer’s Disease). The Journal of Prevention of Alzheimer’s Disease 9, 8–50 (2022).
Kielbasa, W. et al. Brain target occupancy of LY3372689, an inhibitor of the O-GlcNAcase (OGA) enzyme, following administration of single and multiple doses to healthy volunteers. Alzheimer’s. Dement. 17, e057774 (2021).
Kielbasa, W. et al. Discovery and clinical translation of ceperognastat, an O-GlcNAcase (OGA) inhibitor, for the treatment of Alzheimer’s disease. Alzheimers Dement (N. Y.) 10, e70020 (2024).
Ryan, J. M. et al. O1-12-05: phase 1 study in healthy volunteers of the o-glcnacase inhibitor asn120290 as a novel therapy for progressive supranuclear palsy and related tauopathies. Alzheimer’s. Dement. 14, P251–P251 (2018).
Smith, S. M. et al. O2-13-04: Early clinical results and preclinical validation of the O-glcnacase (OGA) inhibitor Mk-8719 as a novel therapeutic for the treatment of tauopathies. Alzheimer’s. Dement. 12, P261–P261 (2016).
Boutz, P. L., Bhutkar, A. & Sharp, P. A. Detained introns are a novel, widespread class of post-transcriptionally spliced introns. Genes Dev. 29, 63–80 (2015).
Park, S. K. et al. A conserved splicing silencer dynamically regulates O-GlcNAc transferase intron retention and O-GlcNAc homeostasis. Cell Rep. 20, 1088–1099 (2017).
Tan, Z. W. et al. O-GlcNAc regulates gene expression by controlling detained intron splicing. Nucleic Acids Res 48, 5656–5669 (2020).
Neil, C. R. et al. Poison exons: tuning RNA splicing for targeted gene regulation. Trends Pharm. Sci. 46, 264–278 (2025).
Martin, S. E. S. et al. Structure-based evolution of low nanomolar o-glcnac transferase inhibitors. J. Am. Chem. Soc. 140, 13542–13545 (2018).
Yuzwa, S. A. et al. A potent mechanism-inspired O-GlcNAcase inhibitor that blocks phosphorylation of tau in vivo. Nat. Chem. Biol. 4, 483–490 (2008).
Pratt, M. R. & Vocadlo, D. J. Understanding and exploiting the roles of O-GlcNAc in neurodegenerative diseases. J. Biol. Chem. 299, 105411 (2023).
Bartolome-Nebreda, J. M., Trabanco, A. A., Velter, A. I. & Buijnsters, P. O-GlcNAcase inhibitors as potential therapeutics for the treatment of Alzheimer’s disease and related tauopathies: analysis of the patent literature. Expert Opin. Ther. Pat. 31, 1117–1154 (2021).
Corsello, S. M. et al. The Drug Repurposing Hub: a next-generation drug library and information resource. Nat. Med 23, 405–408 (2017).
Scarborough, A. M., Govindan, A. & Conrad, N. K. Genome-wide CRISPR screening to identify mammalian factors that regulate intron retention. Methods Mol. Biol. 2537, 263–284 (2022).
Govindan, A. & Conrad, N. K. SFSWAP is a negative regulator of OGT intron detention and global pre-mRNA splicing. eLife 13, RP104439 (2025).
Zhang, J. H., Chung, T. D. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen 4, 67–73 (1999).
Dahlin, J. L. et al. Nuisance compounds in cellular assays. Cell Chem. Biol. 28, 356–370 (2021).
Escher Beate, I., Henneberger, L., König, M., Schlichting, R. & Fischer Fabian, C. Cytotoxicity burst? differentiating specific from nonspecific effects in tox21 in vitro reporter gene assays. Environ. Health Perspect. 128, 077007 (2020).
Johnson, R. L. et al. A quantitative high-throughput screen identifies potential epigenetic modulators of gene expression. Anal. Biochem 375, 237–248 (2008).
Ibanez, G. et al. Evaluation of compound optical interference in high-content screening. SLAS Discov. 23, 321–329 (2018).
Mitchell, D. C. et al. A proteome-wide atlas of drug mechanism of action. Nat. Biotechnol. 41, 845–857 (2023).
Itkonen, H. M. et al. Inhibition of O-GlcNAc Transferase renders prostate cancer cells dependent on CDK9. Mol. Cancer Res 18, 1512–1521 (2020).
Lazarus, M. B., Nam, Y., Jiang, J., Sliz, P. & Walker, S. Structure of human O-GlcNAc transferase and its complex with a peptide substrate. Nature 469, 564–567 (2011).
Dorfmueller, H. C. & van Aalten, D. M. Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds. FEBS Lett. 584, 694–700 (2010).
Jackson, E. G. et al. 4-Deoxy-4-fluoro-GalNAz (4FGalNAz) is a metabolic chemical reporter of O-GlcNAc modifications, highlighting the notable substrate flexibility of O-GlcNAc transferase. ACS Chem. Biol. 17, 159–170 (2022).
Heerding, D. A. et al. Identification of 4-(2-(4-amino-1,2,5-oxadiazol-3-yl)-1-ethyl-7-{[(3S)-3-piperidinylmethyl]oxy}-1H-imidazo[4,5-c]pyridin-4-yl)-2-methyl-3-butyn-2-ol (GSK690693), a novel inhibitor of AKT kinase. J. Med Chem. 51, 5663–5679 (2008).
Jenner, P. Parkinson’s disease: pathological mechanisms and actions of piribedil. J. Neurol. 239, S2–S8 (1992).
Li, X. et al. Structure-based discovery and development of novel O-GlcNAcase inhibitors for the treatment of Alzheimer’s disease. Eur. J. Med Chem. 238, 114444 (2022).
Permanne, B. et al. O-GlcNAcase inhibitor ASN90 is a multimodal drug candidate for tau and α-synuclein proteinopathies. ACS Chem. Neurosci. 13, 1296–1314 (2022).
Nag, S. et al. Development of a Novel [11C]CO-labeled positron emission tomography radioligand [11C]BIO-1819578 for the detection of O-GlcNAcase enzyme activity. ACS Chem. Neurosci. 14, 2560–2568 (2023).
Martinez-Viturro, C. M. et al. Diazaspirononane nonsaccharide inhibitors of O-GlcNAcase (OGA) for the treatment of neurodegenerative disorders. J. Med Chem. 63, 14017–14044 (2020).
Cheng, Z. et al. Discovery of 4-(Arylethynyl)piperidine Derivatives as Potent Nonsaccharide O-GlcNAcase Inhibitors for the Treatment of Alzheimer’s Disease. J. Medicinal Chem. https://doi.org/10.1021/acs.jmedchem.4c01132 (2024).
Bamford, M. J. et al. 1H-imidazo[4,5-c]pyridin-2-yl)-1,2,5-oxadiazol-3-ylamine derivatives: a novel class of potent MSK-1-inhibitors. Bioorg. Med Chem. Lett. 15, 3402–3406 (2005).
Bamford, M. J. et al. 1H-imidazo[4,5-c]pyridin-2-yl)-1,2,5-oxadiazol-3-ylamine derivatives: further optimisation as highly potent and selective MSK-1-inhibitors. Bioorg. Med Chem. Lett. 15, 3407–3411 (2005).
Naqvi, S. et al. Characterization of the cellular action of the MSK inhibitor SB-747651A. Biochem J. 441, 347–357 (2012).
Ishizaki, T. et al. Pharmacological properties of Y-27632, a specific inhibitor of Rho-associated kinases. Mol. Pharmacol. 57, 976 (2000).
Tokushige, H. et al. Effects of topical administration of Y-39983, a selective rho-associated protein kinase inhibitor, on ocular tissues in rabbits and monkeys. Invest Ophthalmol. Vis. Sci. 48, 3216–3222 (2007).
Davies, S. P., Reddy, H., Caivano, M. & Cohen, P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem J. 351, 95–105 (2000).
Li, X. et al. OGT controls mammalian cell viability by regulating the proteasome/mTOR/ mitochondrial axis. Proc. Natl. Acad. Sci. USA 120, e2218332120 (2023).
Butkinaree, C. et al. Characterization of β-N-acetylglucosaminidase cleavage by caspase-3 during apoptosis. J. Biol. Chem. 283, 23557–23566 (2008).
Schwalm, M. P. et al. Functional characterization of pathway inhibitors for the ubiquitin-proteasome system (UPS) as tool compounds for CRBN and VHL-mediated targeted protein degradation. ACS Chem. Biol. https://doi.org/10.1021/acschembio.4c00450 (2025).
Parra, M. et al. An important class of intron retention events in human erythroblasts is regulated by cryptic exons proposed to function as splicing decoys. RNA 24, 1255–1265 (2018).
de Sainte Agathe, J. M. et al. SpliceAI-visual: a free online tool to improve SpliceAI splicing variant interpretation. Hum. Genom. 17, 7 (2023).
Jaganathan, K. et al. Predicting Splicing from Primary Sequence with Deep Learning. Cell 176, 535–548 e524 (2019).
Barraza, S. J., Bhattacharyya, A., Trotta, C. R. & Woll, M. G. Targeting strategies for modulating pre-mRNA splicing with small molecules: Recent advances. Drug Discov. Today 28, 103431 (2023).
Bouton, L., Ecoutin, A., Malard, F. & Campagne, S. Small molecules modulating RNA splicing: a review of targets and future perspectives. RSC Med Chem. 15, 1109–1126 (2024).
Hluchy, M. et al. CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1. Nature 609, 829–834 (2022).
Palacino, J. et al. SMN2 splice modulators enhance U1-pre-mRNA association and rescue SMA mice. Nat. Chem. Biol. 11, 511–517 (2015).
Felker, S. A. et al. Poison exon annotations improve the yield of clinically relevant variants in genomic diagnostic testing. Genet Med 25, 100884 (2023).
Ni, J. Z. et al. Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay. Genes Dev. 21, 708–718 (2007).
Lareau, L. F., Inada, M., Green, R. E., Wengrod, J. C. & Brenner, S. E. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007).
Dardenne, E. et al. RNA helicases DDX5 and DDX17 dynamically orchestrate transcription, miRNA, and splicing programs in cell differentiation. Cell Rep. 7, 1900–1913 (2014).
Enders, L. et al. Pharmacological perturbation of the phase-separating protein SMNDC1. Nat. Commun. 14, 4504 (2023).
Dardenne, E. et al. Splicing switch of an epigenetic regulator by RNA helicases promotes tumor-cell invasiveness. Nat. Struct. Mol. Biol. 19, 1139–1146 (2012).
Wang, X. et al. MK-8719, a novel and selective O-GlcNAcase inhibitor that reduces the formation of pathological Tau and ameliorates neurodegeneration in a mouse model of tauopathy. J. Pharm. Exp. Ther. 374, 252–263 (2020).
Sunden, M. et al. Enzymatic assay for UDP-GlcNAc and its application in the parallel assessment of substrate availability and protein O-GlcNAcylation. Cell Rep. Methods 3, 100518 (2023).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Morales, J. et al. A joint NCBI and EMBL-EBI transcript set for clinical genomics and research. Nature 604, 310–315 (2022).
Wang, Y. et al. rMATS-turbo: an efficient and flexible computational tool for alternative splicing analysis of large-scale RNA-seq data. Nat. Protoc. 19, 1083–1104 (2024).
Vaquero-Garcia, J. et al. RNA splicing analysis using heterogeneous and large RNA-seq datasets. Nat. Commun. 14, 1230 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res 14, 1188–1190 (2004).
Scott, K. A. et al. Covalent targeting of splicing in T cells. Cell Chem. Biol. https://doi.org/10.1016/j.chembiol.2024.10.010 (2024).
Acknowledgements
We thank A.G. Reidenbach and R. Devkota for technical support, G.W.C. Thomas for assistance in sashimi plot generation, the Liau lab for support with RT-qPCR, and the Bauer Core Facility at Harvard University for support with next-generation sequencing. Support from the National Institutes of Health (1RF1AG081475, C.M.W.; U01 CA242115, A.G., N.K.C.; R01 AI153175, A.G., N.K.C.), and the NSF GRFP (S.S.C., A.C.M.) is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
S.S.C. and C.M.W. conceptualized the research. S.S.C., A.C.M., and Y.L. designed and performed experiments and analyzed data, with input from N.K.C., B.K.W., and C.M.W. S.S.C. performed high-throughput screening and in vitro experiments. S.S.C., A.C.M., and Y.L. performed Western blotting and RT-qPCR experiments. A.V. and B.K.W. supported Drug Repurposing Library screening and related experiments. A.G. and N.K.C. supplied engineered cell lines and advised on RNA experiments. A.G., N.K.C., D.E.K., and T.B.S. advised and analyzed RNA-seq data. R.N. performed UDP-HexNAc quantification experiments. S.S.C. and C.M.W. wrote the initial manuscript, which was reviewed and edited by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, S.S., Mody, A.C., Vetere, A. et al. Small molecule splicing modulators that disrupt O-GlcNAc homeostasis. Nat Commun 17, 1548 (2026). https://doi.org/10.1038/s41467-025-68271-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68271-6