Abstract
The persistence of CAR T cells and antigen escape remain major barriers to durable therapeutic success in hematologic malignancies. Our study integrates AI-guided design with targeted protein degradation to overcome these challenges. Utilizing an in-silico library of CAR constructs followed by an in vitro screening, we developed a predictive model, CARMSeD, which forecasts constructs prone to self-activation and dysfunction. Optimized bispecific CD20/CD19 CAR T cells demonstrate superior persistence and anti-tumor efficacy. To further improve durability, the platform incorporates a PROTAC-based module that selectively degrades AKT3, promoting FOXO4-driven mitochondrial fitness, central memory differentiation, and reduced mTOR signaling. We extended this strategy to develop a trispecific CAR T platform co-expressing a secretable CD3/CD22 bispecific engager, achieving potent tumor eradication even in CD19/CD20-negative malignancies demonstrates efficacy across patient-derived leukemia samples and solid tumor models. Together, our study introduces a next-generation AI-guided CAR T strategy that integrates structure-based optimization and intracellular modulation to improve persistence, broaden antigen coverage, and ensure durable therapeutic efficacy.
Similar content being viewed by others

Introduction
CD19 CAR T cell therapy has revolutionized the treatment of otherwise incurable B cell malignancies; however, its long-term efficacy is frequently compromised by antigen escape, wherein malignant cells downregulate or lose CD19 expression to evade immune recognition1,2,3. This challenge is particularly pronounced in heterogeneous tumors containing antigen-low or antigen-negative subclones, often leading to relapse2,4. Our in-silico analyses, supported by multiple clinical datasets, consistently show that a significant proportion of patients relapse following CAR T cell therapy due to modulation or complete loss of CD19 expression. To overcome this limitation, there has been increasing interest in dual and triple antigen targeting CAR T cell strategies that simultaneously recognize CD19, CD20, and CD225,6,7,8,9. These multi-antigen approaches aim to mitigate immune escape and offer broader tumor coverage. For instance, bispecific CAR T cells targeting CD19 and CD20 have shown enhanced anti-tumor efficacy and reduced relapse in both preclinical models and clinical trials10,11. In a phase 1 study (NCT04007029), naïve/memory-derived CD19/CD20 CAR T cells achieved a 90% overall response rate and 70% complete remission in patients with relapsed/refractory non-Hodgkin lymphoma, with low toxicity and durable outcomes12. Similarly, trispecific CAR T cells targeting CD19, CD20, and CD22 have demonstrated the potential to overcome antigenic heterogeneity and improve long-term disease control9,13.
However, while these strategies address antigen escape, they introduce a new challenge of insufficient CAR T cell persistence, particularly in complex multi-antigen designs. Although expanding antigen specificity reduces the risk of tumor escape, it often compromises T cell fitness and increases susceptibility to functional exhaustion14,15. This trade-off ultimately limits the sustained efficacy of treatment. As such, next-generation CAR T engineering must develop integrated solutions that tackle both antigen escape and limited persistence in tandem CAR designs, which are comparatively better when translating into the clinical settings due to ease of manufacturing as compared to other approaches including co-transduction methods and bicistronic CAR designs15,16. Thus, in addition to immune evasion, limited CAR T cell persistence itself remains a critical barrier to durable clinical outcomes. Although initial tumor clearance is often achieved, CAR T cells frequently succumb to functional exhaustion, marked by impaired metabolic activity, reduced cytokine production, and poor memory T cell formation17,18,19,20. These factors are strongly associated with CD19-positive relapse over the years21,22,23. To combat this, co-stimulatory domains such as 4-1BB, ICOS and cytokines such as IL-12, IL-15 and IL-18 have been incorporated into CAR constructs to enhance T cell activation, longevity, and memory programming24,25,26,27,28. Third-generation CARs, which integrate multiple co-stimulatory domains, have also demonstrated superior persistence and anti-tumor function compared to earlier CAR designs29,30 Moreover, metabolic reprogramming strategies, particularly those enhancing mitochondrial oxidative phosphorylation (OXPHOS), have been shown to promote the formation of T stem cell memory (Tscm), supporting long-term persistence and durable responses31,32.
A promising approach to boosting CAR T cell durability involves modulation of the AKT signaling pathway. AKT is a central regulator of cell metabolism, differentiation, and survival, and plays a critical role in T cell memory development33,34,35. Sustained AKT activation in CAR T cells, particularly in exhausted states, impairs memory differentiation and favors terminal effector phenotypes. AKT and its downstream mTOR signaling regulate autophagy, mitophagy, and metabolic balance36. Inhibition of AKT or mTOR enhances memory-associated gene expression and supports metabolic fitness. Recent studies have shown that AKT suppresses FOXO1, a transcription factor essential for memory T cell development, by restricting its nuclear localization37. Inhibition of AKT restores FOXO1 activity, promoting T stem cell and central memory cell (Tscm/Tcm) differentiation, increased survival and enhanced anti-tumor function35,38. While the roles of AKT1 and AKT2 have been explored, the specific contribution of AKT3 in T cell memory remains uncharacterized. Similarly, among the four FOXO family members (FOXO1-4), the functions of FOXO1–3 in memory T cell biology are better understood, while FOXO4 remains largely unknown in this context.
To address these gaps and improve CAR T cell survival, we develop a PROTAC-based strategy that selectively degrades AKT3. Transcriptomic profiling reveals that AKT3 is upregulated during CAR T exhaustion, and its targeted degradation promotes Tcm formation and enhances metabolic fitness in multi-antigen CAR T cells. This effect occurs through the upregulation of FOXO4, which functions as a central regulator of memory programming and CAR T persistence. Together, this AKT3-FOXO4 axis establishes a rational framework for engineering long-lasting, metabolically resilient CAR T cells with the potential to overcome antigen escape and functional exhaustion across diverse malignancies.
Results
Design and evaluation of monospecific, bispecific, and trispecific CAR molecules
In an in silico analysis of 4129 patients who received CAR T cell therapy, either monospecific or bispecific, we observed a high relapse rate, consistent with findings from previous meta-analyses (Fig. 1a, Supplementary Data 1 and Supplementary Fig. 1b)1,2,3. Among patients who relapsed (55.7%) after monospecific CAR T cell therapy, a substantial proportion (42.1%) exhibited CD19-negative tumor recurrence, highlighting antigen escape as a dominant mechanism of resistance in this subgroup (Fig. 1a). Although prior studies have explored multi-antigen targeting CAR designs to mitigate antigen escape5,6,7,8,9, these approaches still exhibited a high incidence of CD19 antigen loss (31.52%) and were associated with reduced CAR T cell durability (Supplementary Fig. 1b). Importantly, the structural features of these multi-antigen CARs, and their potential contribution to activation induced cell death (AICD), have not been systematically investigated at the molecular level.
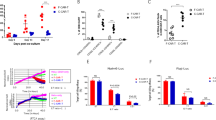
a Histogram showing in-silico analysis of CAR T cell-treated patients (n = 4219) revealed a high relapse rate, with 42.11% (n = 216 of n = 513 overall relapse patients) experiencing CD19-negative recurrence after monospecific CAR Therapy (n = 2916). b Schematic overview of the CAR design strategy showing mono, bi, and trispecific constructs targeting CD19, CD20, and CD22. c Experimental workflow illustrating CAR screening: 1452 CARs were transduced into primary T cells and analyzed for signal-1 (activation), signal-2 (exhaustion), and signal-3 (cell death). Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. d Categorization of CARs into low (L), medium (M), and high (H) levels based on fluorescence intensity cutoffs determined by CD19 CARs as reference. e bar graph showing the distribution of 1452 screened CARs across L-, M-, and H-CARMSeD categories using the CARMSeD scoring system. f AI model development pipeline for CAR dysfunction risk prediction, based on 1,452 CAR constructs with an 80:20 split for training and testing. g–j Performance metrics of AI model predicting CARMSeD scores using 1452 CAR constructs. g ML learning curve of model accuracy over 50 epochs, achieving a training accuracy of 0.98 and validation accuracy of 0.95. h Scatter plot comparing measured versus predicted CARMSeD scores for training (R2 = 0.87) and validation (R2 = 0.83) sets. i Predicted versus measured CARMSeD scores on the validation set, categorized into low (blue), medium (orange), and high (green) CARMSeD. j Box plots show the median (center line), the 25th–75th percentiles (box), and whiskers extending to the minimum and maximum non-outlier values; individual points denote outliers. Numbers above each box indicate sequence counts. k Molecular dynamics simulation of CAR constructs with varying linker lengths, assessing scFv-scFv interaction. Structural conformations at 0 ns, 50 ns and 200 ns for different CAR scFv arrangements highlighting CDR regions (surface transparency 30%), Root Mean Square Deviation (RMSD) plots over 200 ns for both constructs, respectively, indicating structural stability and conformational changes. l Bar graph showing in vitro receptor binding affinity validation for top humanized scFvs of CD19, CD20, and CD22 CARs (n = 6 biologically independent samples). Data represent mean ± SEM. **p < 0.01; ****p < 0.001; ns: not significant. A non-parametric t-test was used for statistical analysis between groups. Source data are provided as a Source Data file.
To overcome structural constraints, we engineered CARs in monospecific, bispecific, and trispecific formats targeting CD19, CD20, and CD22 (Fig. 1b). A comprehensive CAR library comprising 10,824 distinct constructs was generated by systematically varying G4S linker lengths (3x; 4x; 5x), transmembrane domains, and co-stimulatory (co-stim) domains (Supplementary Data 2). To achieve structural diversity, the CAR library incorporated both single and dual domain co-stim domains, combining CD28, 4-1BB, ICOS, and OX40 in multiple configurations with distinct transmembrane domains (CD8/28/ICOS) and a constant CD8 hinge region. This design strategy maximized molecular diversity and functional representation, enabling a comprehensive evaluation of CAR performance across structural variants. From this library, a representative subset of 1452 CARs was selected for in vitro screening (Supplementary Data 3). Selection criteria included coverage of all antigen formats (mono, bi, and trispecific), balanced representation of different co-stim domain combinations, and computational prioritization based on CAR-Mediated Self-Destruction (CARMSeD) scores to ensure that both high and low ranking constructs were evaluated. These CARs were introduced into T cells derived from three healthy donors and assessed for activation (signal-1), exhaustion (signal-2), and cell death (signal-3) using fluorescence-based assays with corresponding markers (Fig. 1c).
Signal intensities were categorized as low (L), medium (M), or high (H) (Fig. 1d), with threshold values established from reference data generated using monospecific CD19 CAR stimulation (Supplementary Fig. 2). Based on a defined scoring system, CARs were grouped according to their CARMSeD classification. Of the 1452 CARs screened, 465 were classified as L-CARMSeD, 359 as M-CARMSeD, and 628 as H-CARMSeD (Fig. 1e), indicating that lower CARMSeD scores correspond to more favorable CAR designs.
Next, an AI model was developed using data from these experiments (Fig. 1f). The model was trained on 1161 CARs and validated on the remaining dataset, achieving a prediction accuracy of 95% (Fig. 1h-j). Using this model, we screened the entire CAR library and identified 2749 candidates with low predicted CARMSeD scores. To experimentally validate the predictions, we tested 20 CARs (top 10 and bottom 10 ranked by CARMSeD) in vitro for anti-tumor activity and CAR T cell durability (as indicated by CAR T cell count over time). The results showed a strong correlation between predicted CARMSeD scores and experimental outcomes (Supplementary Data 4). Building on these findings, we tested these designs in an in vitro tumor rechallenge (TR) model to further assess efficacy32. Among the tested designs, bispecific CD20/19 and CD22/19 CARs incorporating ICOS transmembrane domains and as primary co-stim domain, and 4-1BB as a secondary co-stim domain consistently demonstrated superior activity. The top constructs also shared a domain orientation of VH-VL (linked with 3×G4S linkers) and single-chain fragment variables (scFvs) separated by 4×G4S linkers, which was therefore prioritized.
To establish the broader applicability of this design, we extended the evaluation of ICOS + 4-1BB co-stim domains to CARs targeting additional antigens such as BCMA, Claudin18.2, and EGFR. Across these targets, ICOS + 4-1BB consistently outperformed single co-stim (CD28 or 4-1BB alone), confirming its generalizable advantage in both anti-tumor activity and CAR T cell durability (Supplementary Fig. 3 and Supplementary Data 4).
Since the scFvs were murine-derived, immunogenicity concerns (HAMA responses) were addressed by AI-guided humanization of scFvs, followed by computational evaluation of receptor interactions (Supplementary Fig. 4, 5; Supplementary Data 5, 6). We further applied an AI pipeline to assess intra-CAR interactions, enabling selection of bispecific and trispecific designs with minimal self-interference (Fig. 1k, Supplementary Fig. 6, Movies 1-4). Subsequent in vitro testing of the humanized (Hu) CARs confirmed that the combination of HuCD20 with murine (Mu) CD19 scFv achieved the strongest anti-tumor activity and higher binding affinity (Fig. 1l and Supplementary Data 7). Biolayer interferometry (BLI) confirmed that the AI generated HuCD20 scFv exhibited stronger binding affinity than its murine counterpart (Supplementary Fig. 7). Ultimately, two bispecific CARs and one trispecific CAR (incorporating HuCD20 and HuCD22) were selected for further validation
Preclinical validation of bispecific CAR T cells targeting CD20/19 to overcome antigen escape
To evaluate the anti-tumor efficacy and in vitro durability of the selected CAR T cell constructs, we first established a K562 cell line model, stably expressing CD19, CD20, and/or CD22 receptors with defined phenotypes: CD19⁻/CD20⁺/CD22⁺, CD19⁻/CD20⁻/CD22⁺, and CD19⁻/CD20⁻/CD22⁻ (Fig. 2a and Supplementary Fig. 8). Co-culture assays with these engineered targets revealed potent and specific cytotoxic responses across all antigen settings (Fig. 2b-f). Notably, each CAR design showed strong activity against cells expressing single or multiple antigens, outperforming the second-generation monospecific CD19 (m19) CAR containing a 4-1BB co-stimulatory domain, used as the control (Fig. 2b, -f).
a Schematic illustration of the K562 cell line model expressing individual or triple combinations of CD19 (purple), CD20 (red), and CD22 (yellow) antigens. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. b Bar chart depicting the percentage expression of each antigen in K562 cell lines, both individually and in combination. c–f Line graph of cytotoxicity assays showing antigen-specific killing of K562 target cells. All tested constructs surpassed the performance of second-generation monospecific CD19 (m19) CAR T cells (n = 3 biologically independent samples). g Heatmap showing comparison of proliferation rates for bispecific; b20/19 or b22/19, and trispecific; t20/19/22 CAR T cells, represented as fold expansion up to Day 17 with respect to the baseline at the time of cell seeding. h Schematic of the Raji WT cell line platform expressing CD19 (purple), CD20 (red), and CD22 (yellow) antigens, edited using CRISPR-Cas9 to generate knockout variants. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. i, j Line graph of cytotoxicity assays demonstrating the superior efficacy of b20/19 CAR T cells in eliminating antigen-negative Raji variants, compared to m19 CARs (n = 5 biologically independent samples). k Schematic representation of the tumor rechallenge (TR) model using the Raji WT cell line (RajiWT). Gray circles represent initial engraftment and monitoring phases, pink circle shows the first incubation with RajiWT, while purple circles indicate the timing of the RajiCD19−/− rechallenge. l Heatmap representation of TR model showing IFN-γ secretion (pg/mL), percentage of tumor lysis (1:10; T: E), and the number of CAR T cells detected on days 7, 9, 11, 15, and 17 post-rechallenge (n = 5 biologically independent samples). Data represent mean ± SEM. Source data are provided as a Source Data file.
To further delineate the functional effect of antigen targeting format on CAR T cell expansion, we assessed the proliferative potential of bispecific (b20/19) and trispecific (t20/19/22) CAR T cells. The trispecific construct exhibited reduced proliferation in comparison to the b20/19 configuration, aligning with computational predictions of increased structural rigidity and higher CARMSeD scores (Fig. 2g). Similarly, the b22/19 construct displayed a comparable proliferation rate to b20/19. Given its superior performance profile, we prioritized the b20/19 CAR T cells for downstream in vitro and in vivo studies.
Next, to study these CARs in a clinically relevant model cell line, we generated CD19−/−, CD20−/− and CD19/20−/− knockout variants of the Raji lymphoma cell line using CRISPR-Cas9 genome editing (Fig. 2h)39. Cytotoxicity assays demonstrated that b20/19 CAR T cells effectively eliminated both CD19−/− and CD20−/− Raji cells, whereas m19 CAR T cells were ineffective against antigen-negative targets (Fig. 2i, j). In an in vitro TR model involving rechallenge with CD19-/- Raji cells that retained CD20 expression, b20/19 CAR T cells exhibited durable anti-tumor responses, sustained interferon-gamma (IFN-γ) production, and prolonged cell survival, as shown by bioluminescence imaging and functional assays (Fig. 2k, l).
Following in vitro validation, we performed an in vivo assessment using a xenograft mouse model rechallenged with CD19−/− tumor cells (Fig. 3a). Compared to the m19-treated group, b20/19 CAR T cells achieved superior tumor clearance, shown by a sharp reduction in tumor bioluminescence signal and substantial tumor regression (Fig. 3b, c). However, tumor recurrence was observed over time. Longitudinal assessment of peripheral blood samples from treated mice (n = 5 per group) indicated enhanced survival of CAR T cells in the b20/19 group (Fig. 3d), which was associated with sustained survival benefits (Fig. 3e). Phenotypic characterization of residual tumor cells revealed a marked reduction in both CD19+ and CD20+ Raji cells in the b20/19 CAR T cell treated group. In contrast, mice treated with m19 cells exhibited a progressive accumulation of CD20+ tumor cells, indicating selective-antigen escape. Notably, the b20/19 group showed only a modest increase in CD20+ cells by Day 28, suggesting effective control of antigen escape mechanisms but lower survival of these cells (Fig. 3f). To further validate these findings, CD19 and CD20 expression profiles from untreated Raji control mice were evaluated, showing stable expression of both markers in the absence of CAR T treatment (Supplementary Fig. 9a). Furthermore, analysis of CD19⁻ or CD19low subsets within CD20+ Raji cells across m19 and b20/19 CAR T groups confirmed selective pressure on CD19 while maintaining CD20 expression, consistent with antigen escape (Supplementary Fig. 9b). Functional analysis of human CD8+ CAR T cells isolated from b20/19 treated mice revealed preserved effector function, as indicated by sustained Granzyme B and IFN-γ secretion upon co-culture with CD19⁻/⁻ target cells (Fig. 3g, h). When compared to mice that received mCD19 cells, b20/19 CAR T cells exhibited reduced exhaustion during early treatment phases (Fig. 3i, j).
a Schematic timeline of in vivo lymphoma model for evaluation of monospecific and bispecific CAR T cells. Mice were xenografted with RajiWT cells (expressing CD19, CD20, and CD22) (day 0), followed by administration of m19 or b20/19 CAR T cells on day 5 and subsequent RajiCD19−/− TR on day 12, 19 and 26. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. b Bioluminescent imaging and (c) stacked area plot showing tumor burden quantification show effective tumor control by b20/19 CAR T cells versus m19 CARs (n = 5). d CAR T cell survival over time (n = 5 mice). e Kaplan-Meier survival curves showing survival outcomes over 70 days (n = 5 mice). f Analysis of residual tumor CD19 or CD20 tumor cells over time (n = 5 mice). g, h Bar plot showing Granzyme B and IFN-γ secretion from human CD8+ CAR T cells isolated b20/19 post-treatment to confirm functional cytotoxicity of b20/19 against CD19⁻ targets (n = 5). The CAR T cells isolated from mice that received conventional monospecific (m)CD19 CAR T cells served as the control for comparison. i, j TR induced upregulation of exhaustion markers PD-1 and LAG-3 (n = 5 mice). k Immunophenotyping of CAR T cells post-TR shows loss of central memory (Tcm) populations and increased PD-1 expression, consistent with functional exhaustion and limited engraftment (n = 5 mice). Data represents mean ± SEM. **p < 0.01; ***p < 0.005; ****p < 0.001. A non-parametric t-test was used for statistical analysis between groups, and for (k), a Two-way ANOVA followed by post-hoc testing was applied. Source data are provided as a Source Data file.
Immunophenotypic profiling was performed to assess CAR T cell populations with differential PD-1 expression, representing low versus high exhaustion states. Within these subsets, both Tcm and effector memory (Tem) phenotypes were evaluated. The PD-1high population broadly corresponded to exhausted CAR T cells, marked by a reduced Tcm subset following TR in the b20/19 group (Fig. 3k). This progressive shift toward exhaustion may contribute to a downward trend in CAR T cell survival and a corresponding reduction in overall survival of the mice over time (Fig. 3d, e).
AKT3 is a key regulator of CAR T cell metabolism, memory differentiation, and durability
Limited durability remains a critical barrier to the long-term success of current CAR T cell therapies. Clinical data analyzed shown in Fig. 1A indicate that CAR T cells fail to persist in 40–50% of patients, contributing to disease relapse and reduced therapeutic efficacy1,2,3. To address this limitation, we isolated CAR T cells from control mice (CON; single tumor challenge) and from the TR group that had received three sequential tumor injections, followed by b20/19 CAR T cell administration. Cells from each group (n = 10) were pooled, and transcriptomic profiling via RNA sequencing was performed to identify differentially regulated genes and pathways (Fig. 4a). The analysis revealed notable dysregulation of genes associated with oxidative phosphorylation, mitochondrial metabolism, FOXO signaling and autophagy/mitophagy and genes implicated in T cell exhaustion (Fig. 4b). Importantly, AKT3 was among the most upregulated genes (Supplementary Fig. 10a), which was further confirmed at the protein level by flow cytometry in both b20/19 and m19 CAR T cells following TR (Fig. 4c, d and Supplementary Fig. 10b).
a Schematic timeline of in vivo lymphoma model for isolation of CAR T cells for RNA seq analysis. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. b Bar graph of RNA seq showing pathway analysis of dysregulated genes associated with OXPHOS, mitochondrial metabolism, FOXO signaling, and autophagy in TR group CAR T cells. c Representative flow cytometry histogram showing the fluorescence intensity of FITC-labeled AKT3 (FITC-AKT3) in T cells from different groups. d Bar graph showing the mean fluorescence intensity (MFI) of AKT3 in T cells from the CON and TR groups (n = 6 mice). e Illustration of the plasmids showing pLenti-AKT3 lentiviral vector used for co-transduction of CAR T cells with CAR vectors. f Schematic representation of the 17-day in vitro experimental timeline for evaluating T cell responses in TR model. T cells were isolated, activated, and transduced on Day 0 and further expanded until Day 7, followed by tumor challenge with RajiWT cells on Day 7. g Line graph comparing the survival of CAR T cells transduced with b20/19 CAR and either a control vector (b20/19pVEC, blue circles) or an AKT3-expressing vector (b20/19pAKT3, orange squares) over 15 days (n = 5 biologically independent samples). h Histogram analysis and pie chart showing distribution of Tem and Tcm with PD-1low or PD-1high within these subsets (n = 5 biologically independent samples) at day 17. i Line graph showing Extracellular Acidification Rate (ECAR, mpH/min) over 100 min day 17. j Bar graph of basal ECAR (mpH/min, normalized) between b20/19pVEC (blue) and b20/19pAKT3 (orange) groups (n = 6 biologically independent samples). k, l Similarly line graph showing Oxygen Consumption Rate (OCR, pMoles/min) and corresponding bar graph (n = 6 biologically independent samples). m Schematic of the lentiviral vector delivering shRNA against AKT3 (shAKT3). The lower panel depicts the lentiviral particle transduced target CAR T cells. n Line graph comparing the survival of CAR T cells over 15 days for three groups. Statistical analysis was done for day 15 data points (n = 5 biologically independent samples). o Bar graph and pie charts showing the percentage of various subsets of CD8+ CAR T cells across three groups (n = 5 biologically independent samples). p, q Bar graph of ECAR, and OCR across three groups (n = 8). Data represent mean ± SEM. *p < 0.05; **p < 0.01; ***p < 0.005; ****p < 0.001. A nonparametric t-test was used for statistical analysis between groups. For g, h, and o, a Two-way ANOVA followed by post-hoc testing was applied. Source data are provided as a Source Data file.
To explore the functional impact of AKT3 overexpression, we engineered CAR T cells co-transduced with a lentiviral vector encoding AKT3 (Fig. 4e and Supplementary Fig. 10c, d) and assessed their performance in the in vitro TR model (Fig. 4f). These AKT3 overexpressing CAR T cells exhibited reduced survival over time (Fig. 4g), which was accompanied by immunophenotypic changes characterized by a decline in Tcm and Tem subsets within PD-1low populations and an expansion of PD-1high exhausted-like subsets (Fig. 4h). Given the known role of AKT3 in cellular metabolism, we examined metabolic activity using ECAR and OCR assays. AKT3 overexpression led to a substantial decrease in OXPHOS and a compensatory increase in glycolytic activity (Fig. 4i–l). Consistently, AKT3 overexpression in unstimulated CAR T cells also impaired proliferation and reduced both Tcm and Tem formation, without affecting apoptosis (Supplementary Fig. 10e-j).
To further investigate the causal role of AKT3, we employed shRNA-mediated knockdown of AKT3 in CAR T cells (Fig. 4m and Supplementary Fig. 10d). In contrast to the overexpression model, AKT3 knockdown markedly improved long-term CAR T cell durability and increased the proportion of Tcm subsets within the PD-1low population (Fig. 4n, o). Metabolic profiling of these cells showed restored OXPHOS activity and decrease in glycolysis, indicating a metabolic reprogramming favorable for memory cell formation (Fig. 4p, q). In unstimulated CAR T cells, AKT3 knockdown also enhanced proliferation and promoted both Tcm and Tem differentiation, without affecting apoptosis (Supplementary Fig. 10e-j). The protective effect of AKT3 knockdown was reversed by simultaneous overexpression of AKT3 in the same cells (Fig. 4n-q). Together, these findings establish AKT3 as a key regulator of CAR T cell metabolic state, memory differentiation, and durability. Thus, targeting AKT3 may offer a promising strategy to enhance the durability and efficacy of CAR T cell therapies.
AKT3-PROTAC bispecific CAR T cells demonstrated higher memory CAR T cells
AKT3 inhibition has previously been shown to be associated with the induction of a memory phenotype in T cells. Recent studies also highlight its role in enhancing effector function and promoting durable memory formation in CAR T cells35,38. However, existing strategies rely heavily on ex vivo application of molecular inhibitors during CAR T cell culture. To circumvent the need for such external supplementation, we developed a novel design to achieve intracellular AKT3 inhibition using a PROTAC-based approach. We first generated a series of AKT3-targeting peptides using RF diffusion modeling and validated them in silico (Fig. 5a and Supplementary Fig. 11). The top five candidates were selected for in vitro screening. These peptides were fused to a cell-penetrating peptide (CPP) and tested in HEK293T cells expressing a GFP-AKT3-mCherry reporter (Supplementary Fig. 12a). Fluorescence analysis revealed dose-dependent AKT3 degradation across all peptides, with peptide 2 (P2) showing the strongest effect, confirmed by immunoblotting at various concentrations (Supplementary Fig. 12b, c). For functional validation in CAR T cells, b20/19 CAR T co-transduced with GFP were incubated with various concentrations of P2, which resulted in a consistent, dose-dependent reduction of the endogenous AKT3 levels (Fig. 5b, c). A non-targeting peptide (NTP) control had no detectable effect (Fig. 5c-e). Subsequently, we engineered b20/19 CAR constructs incorporating the AKT3-PROTAC (AKT3PROTAC) domain after the CAR transgene and evaluated their functional activity on CD19-/- tumor cells upon TR (Fig. 5f). Fluorescence analysis confirmed endogenous AKT3 degradation specifically in PROTAC-modified CAR T cells, while NTP-treated cells retained AKT3 expression (Fig. 5g). These findings were validated by immunoblot and immunofluorescence, confirming endogenous AKT3 proteolysis (Fig. 5h-k).
a Table listing the sequences of peptides designed using RFdiffusion for PROTAC application, targeting AKT3 (AKT3_P1 to AKT3_P5, AKT3_PC) and a non-targeting peptide (NTP), with AKT3_P2 highlighted as the selected peptide. b Schematic illustration of mCherry-AKT3 CAR T cells induced with PROTAC. The PROTAC consists of the AKT3-targeting peptide (AKT3-P) fused to a cell-penetrating peptide (CPP) and via a linker connected to an E3 ligase ligand. c Line graph showing the dose-dependent effect of PROTAC peptide P2 (blue circles) compared to a non-targeting peptide (NTP, orange circles) on the ratio of endogenous AKT3 to GFP fluorescence in CAR T cells (n = 4 biologically independent samples). The AKT3 was detected using primary anti-AKT3 antibody (raised in rabbit) followed by detection with secondary anti-rabbit antibody (Alexafluor 546; red). d Representative fluorescence microscopy images of CAR T cells treated with NTP or P2, showing GFP (green), anti-AKT3 (Alexafluor 546; red), and merged channels with regions of interest (ROIs). ROIs highlight reduced AKT3 signal in P2-treated cells compared to NTP-treated cells. e Violin plot comparing the integrated density of AKT3 in P2 and NTP-treated CAR T cells (n = 15 images). f Schematic of the b20/19 CAR construct, including a signal peptide (SP), furin cleavage (FC), and AKT3PROTAC (P2A linker), designed to target AKT3 for degradation in CAR T cells. Timeline of the 15-day experiment for evaluating b20/19 CAR T cell responses. g Jittered dot plot showing the relative fluorescence units (RFU) of AKT3 in b20/19 CAR T cells treated with NTP or AKT3PROTAC (n = 25 data points from three independent experiments). h Western blot analysis of b20/19 CAR T cells treated with NTPPROTAC or AKT3PROTAC, probing for endogenous AKT3 using anti-AKT3, and GAPDH as a loading control. i Bar graph showing the densitometry analysis of the blots (n = 6 biologically independent samples). j Representative fluorescence microscopy images of b20/19 CAR T cells treated with non-targeting peptide (NTPPROTAC) or AKT3PROTAC, showing GFP (green), anti-AKT3 (Alexafluor 546; red), and merged channels. k Violin plot comparing the integrated density of AKT3 in NTPPROTAC and AKT3PROTAC treated b20/19 CAR T cells (n = 10 images). l Survival curve showing percentage of RajiWT cell survival after co-culture with CAR T cells at different target-to-effector (T:E) ratios (n = 4 biologically independent samples). m Percentage of patient-derived cancer cell survival after 24-h co-culture with CAR T cells at a target-to-effector (T:E) ratio of 1:2.5. b20/19-AKT3PROTAC exhibits higher cytotoxicity in patient derived cells (n = 5 biologically independent samples). n In vitro tumor rechallenge model showing survival of b20/19-AKT3PROTAC CAR T cells over time (n = 5 biologically independent samples). o Flow cytometry histogram showing various T cell subsets (n = 5). p, q Dot plot showing showing ECAR and OCR analysis (n = 12 data points from three independent experiments). Data represent mean ± SEM. **p < 0.01; ***p < 0.005; ****p < 0.001. A nonparametric t-test was used for statistical analysis between groups. For o, Two-way ANOVA followed by post-hoc testing was applied. Scale bar; d: 200 μm; j: 100 μm. Source data are provided as a Source Data file.
Anti-tumor assays demonstrated that b20/19-AKT3PROTAC cells exerted strong cytolytic activity against multiple CD19+ lymphoma and leukemia cell lines (Supplementary Fig. 13a, b). Notably, even at reduced doses, these cells maintained anti-tumor efficacy comparable to both mCD19 and b20/19 CARs (Fig. 5l) and effectively targeted CD19⁻/⁻ Raji cells (Supplementary Fig. 13c). Potent anti-tumor responses were also observed in primary patient-derived leukemia samples (Fig. 5m). Consistent with these findings, in vitro TR assay demonstrated markedly improved survival of b20/19-AKT3PROTAC cells (Fig. 5n). Similar enhancements in antitumor activity and persistence were observed in conventional CAR T cells expressing AKT3PROTAC (Supplementary Fig. 14). Immunophenotypic analysis revealed higher frequencies of Tcm and Tem subsets accompanied by reduced exhaustion marker expression (Fig. 5o). Further, metabolic profiling corroborated these findings, showing OXPHOS and diminished reliance on glycolysis (Fig. 5p, q). Collectively, these results establish a robust and targeted strategy to degrade AKT3 in CAR T cells through PROTACS, thereby enhancing memory phenotype, persistence, and anti-tumor efficacy, which was validated across both cell line and patient derived tumor models.
AKT3PROTAC alleviates CAR T dysfunction by increasing FOXO4 expression and inhibiting mTOR activity
To elucidate the molecular basis by which AKT3 modulation enhances memory T cell formation, we investigated potential downstream targets of AKT3 (Fig. 6a). CAR T cells undergoing TR were harvested for mRNA extraction, and the expression of proteins associated with AKT3 was analyzed at the transcript level. A pronounced upregulation of FOXO4 mRNA was observed in b20/19 AKT3PROTAC cells following TR (Fig. 6b). This transcriptional increase was further confirmed at the total protein level (Fig. 6c, d). To determine whether FOXO4 mediates the observed memory phenotype, we performed shRNA-mediated knockdown of FOXO4 (Supplementary Fig. 15), which resulted in an increase in PD-1high T memory subsets and a concomitant decrease in PD-1low memory cells (Fig. 6e). Functionally, this shift was associated with reduced CAR T cell durability (Fig. 6f).
a Pathway analysis of proteins involved in AKT3 interaction, modifications or regulation of its expression with emphasis on FOXO4. b Relative mRNA expression levels (normalized to beta actin; ACTB) of key genes show upregulation of FOXO4 mRNA in b20/19-AKT3PROTAC CAR T (n = 6 biologically independent samples). c Flow cytometry histograms of total FOXO4 and phosphorylated FOXO4 (p-FOXO4) in CAR T cells after TR with RajiCD19−/− cells. d Histogram analysis of the flow cytometry plots (n = 10 biologically independent samples). e Bar graph shows the percentage of CD8+ CAR T cells expressing different phenotypes. Pie charts illustrate the proportional distribution of these subsets across conditions (n = 5 biologically independent samples). f Survival of CAR T cells over 15 days under various conditions (n = 4 biologically independent samples). g Violin plots showing the percentage of mTOR activity (% mTOR activity) in various conditions, with shRNA based FOXO4 knockdown elevated mTOR activity (n = 6 biologically independent samples). h Bar plots show the percentage of MFI of autophagy from autophagic flux assay (n = 8 data points from three independent experiments). i Dot plot showing ECAR in NTPPROTAC+Scram, NTPPROTAC+shFOXO4, AKT3PROTAC+Scram, and AKT3PROTAC+shFOXO4 conditions, with FOXO4 knockdown increasing shift from OXPHOS to glycolysis (n = 12 data points from three independent experiments). j Similarly, OCR with FOXO4 knockdown decreases mitochondrial respiration. Individual data points are shown for each condition (n = 12 data points from three independent experiments). k Box-and-whisker plot showing percentage of expression of CD19 (yellow), CD20 (blue), and CD22 (purple) across 129 ALL patient samples, with varying expression levels for each marker (n = 63 patient samples). l Bar graph showing the number of patient samples categorized as Negative/Dim, Moderate, or Bright for CD19, CD20, and CD22 expression. m Schematic illustration of K562WT and CD20 expressing K562 stable cells transduced with different MOIs to obtain three populations: CD20L (low), CD20M (medium), and CD20H (high), which were further FACS sorted. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. n Violin plots showing the percentage of CD20 expression (% CD20 expression) in the sorted CD20-expressing K562 cell populations, confirming distinct expression levels (n = 10 data flow cytometry points from three independent experiments). o Representative super-resolution microscopy images of differential CD20 surface expression in K562 cells. Images show CD20 (red) in K562-CD20L (low), K562-CD20M (medium), and K562-CD20H (high) cell. p–r Survival curves of K562 cells expressing varying CD20 expression levels under CAR T cell treatments. The line graph shows the percentage of CD20+ cell survival when treated with Rituximab-based monospecific CAR (Rtx-m20, dark green), in-house humanized anti-CD20 CAR (AB21-m20, green) (n = 4 biologically independent samples). s Survival of CAR T cells with varying CD20-targeting CAR constructs over 15 days (n = 5). Data represents mean ± SEM. ****p < 0.001. A nonparametric t-test was used for statistical analysis between groups. For e, f and s, a Two-way ANOVA followed by post-hoc testing was applied. Scale bar: 5 μm. Source data are provided as a Source Data file.
Given the role of FOXO4 in autophagy via suppression of mTOR signaling, we next assessed mTOR activity. FOXO4 knockdown led to increased mTOR activation and impairment of autophagy (Fig. 6g, h). Metabolic profiling further revealed a shift from OXPHOS to glycolysis in FOXO4-deficient cells (Fig. 6i, j). These findings suggest that AKT3 tightly regulates FOXO4, possibly through phosphorylation and transcriptional repression, although further mechanistic studies are warranted. Overall, these results indicate that AKT3 proteolysis via PROTACs enhances CAR T cell function and long-term survival by promoting FOXO4-mediated autophagy and metabolic fitness.
Unlike CD19, which is robustly and uniformly expressed, CD20 displays heterogeneous and often dim surface expression. Analysis of clinical samples obtained from over 129 patients across multiple hospitals confirmed that CD20 expression is variable and frequently low (Fig. 6k, l and Supplementary Data 8). To model this heterogeneity, we engineered K562 cells expressing low, medium, and high levels of CD20 (Fig. 6m) and validated single-cell expression by flow cytometry (Fig. 6n). Immunofluorescence and super-resolution microscopy further confirmed differences in CD20 surface density (Fig. 6o and Supplementary Fig. 16). We next evaluated the anti-tumor activity of b20/19 AKT3PROTAC CAR T cells against these engineered targets. Compared to standard CD20-specific CAR constructs used in clinical trials, our optimized HuCD20 scFv combined with AKT3PROTAC demonstrated robust cytotoxicity even against cells with very low CD20 expression (Fig. 6p–r). Moreover, b20/19-AKT3PROTAC cells exhibited superior durability when challenged with CD20low targets (Fig. 6s), suggesting their potential for sustained and effective tumor control even in conditions of antigen heterogeneity.
Enhanced in vivo durability of AKT3-targeted PROTAC CAR T cells
To determine the in vivo efficacy of b20/19-AKT3PROTAC CAR T cells, we developed a xenograft mouse model (Fig. 7a). Mice were monitored longitudinally for tumor clearance and survival outcomes. Notably, mice treated with b20/19-AKT3PROTAC CAR T cells showed improved tumor clearance and enhanced survival compared to controls (Fig. 7b-d). In contrast, the group treated with CAR T cells incorporating shRNA-mediated FOXO4 knockdown did not show comparable anti-tumor activity, emphasizing the critical role of FOXO4 in mediating therapeutic benefit. Additionally, longitudinal tracking revealed durable CAR T cell survival in the b20/19-AKT3PROTAC group, with detectable CAR T cells up to 84 days post-infusion, which was the latest timepoint assessed (Fig. 7e, f). Correspondingly, no residual Raji cells were detected in these mice by day 56 (Fig. 7g), confirming complete tumor eradication. Further phenotypic analysis of CAR T cells isolated 28 days post-infusion showed an increased proportion of memory T cell subsets and reduced expression of exhaustion markers (Fig. 7h). Metabolic profiling revealed a predominant reliance on OXPHOS over glycolysis, supporting a metabolically favorable memory phenotype (Fig. 7i, j).
a Schematic timeline of the experiment showing RajiWT cell injection, CAR T cell administration, and Raji CD19−/− TR. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. b In vivo bioluminescence imaging of mice treated with NTPPROTAC+Scram, NTPPROTAC+shFOXO4, AKT3PROTAC+Scram, and AKT3PROTAC+shFOXO4, showing tumor burden (red indicates high tumor load, blue indicates low) over 84 days, with “X” marking deceased mice. c Tumor radiance over 84 days, demonstrating reduced tumor burden in the AKT3PROTAC+scram group. d Kaplan-Meier survival curves, with the AKT3PROTAC+scram group exhibiting the highest survival rate. e Line graph shows the percentage of CAR T cells detected in blood (% CAR T cells) over time. f Bar graph displays the percentage of CAR T cells in blood at day 84, with the AKT3PROTAC+scram group showing detectable levels (~3%), while other groups were not analyzable due to the absence of surviving mice, indicated as not determined (ND) (n = 5 mice). g Tumor burden assessment till day 56. The line graph shows the number of Raji cells over time. The AKT3PROTAC+scram group exhibits no detectable Raji cell burden by day 56, while other groups with some surviving mice show some detectable cells (n = 5 mice). h Bar graph shows the percentage of CD8 CAR T cells (% CD8 T cells) expressing different phenotypes on day 28 post-infusion with corresponding pie charts illustrating the proportional distribution (n = 5 mice). i, j Dot plot showing ECAR and OCR under various conditions on day 28 (n = 12 data points). Data represent mean ± SEM. ***p < 0.005; ****p < 0.001. A non-parametric t-test was used for statistical analysis between groups. The comparison was made between NTPPROTAC+Scram with NTPPROTAC+shFOXO4 and AKT3PROTAC+Scram with AKT3PROTAC+shFOXO4. For h, a Two-way ANOVA followed by post-hoc testing was applied. Source data are provided as a Source Data Source Data file.
To assess the robustness of this approach, we reduced the administered CAR T cell dose from 5 × 106 to 1 × 106 cells. Notably, similar tumor clearance and survival kinetics were observed even at this reduced dose (Supplementary Fig. 17), emphasizing on the potency and durability of AKT3-targeted PROTAC CAR T cells.
To further explore whether this strategy could be extended to solid tumors, we engineered PROTAC CAR T cells targeting Claudin18.2 (CLDN; for gastric cancer) and EGFR (for non-small cell lung carcinoma; NSCLC) (Fig. 8a). Compared to conventional CAR designs, PROTAC-engineered CAR T cells demonstrated a substantial improvement in antitumor activity against gastric and NSCLC tumor models, respectively (Fig. 8b, c). Notably, higher CAR T cell survival was again observed in the PROTAC groups, corroborated by an increased frequency of memory T cell subsets and reduced exhaustion status (Fig. 8d, e). In vivo xenograft models further validated the superior efficacy of PROTAC CAR T cells (Supplementary Fig. 18). Collectively, these results demonstrate the broad utility of AKT3PROTAC engineering to improve CAR T cell durability and function across both hematologic and solid tumors.
a Schematic of CLDN scFv and EGFR scFv CAR constructs with ICOS, 4-1BB, CD3ζ, and AKT3PROTAC domains. b, c Survival curves showing cytotoxicity of CLDN and EGFR CAR T cells with NTPPROTAC or AKT3PROTAC against AGS (K) and A549 (L) cells at varying T:E ratios (n = 5 biologically independent samples). d Line graph showing CAR T cell proliferation over 15 days, showing enhanced expansion with AKT3PROTAC. e Bar graph and pie charts analysis of EGFR-CAR and CLDN-CAR T cells with NTPPROTAC or AKT3PROTAC, showing the percentage of various T cell subsets (n = 5 biologically independent samples). Data represent mean ± SEM. ****p < 0.001. A two-way ANOVA followed by post-hoc testing was applied. Source data are provided as a Source Data file.
Trispecific CAR exhibits safety and sustained cellular durability
To further increase the versatility of the antibodies with three antigens, we developed a strategy combining b20/19-AKT3PROTAC with CD22 secretory BITE based on the CD3 and CD22 nanobody (nb) (Fig. 9a). The humanized nanobodies were designed using the computational approach and the CDR grafting method as described in the “Methods”. Initially to demonstrate the expression of nbCD3/22, we transduced the LVV encoding the b20/19-AKT3PROTAC plasmid encoding nbCD3/22 (hereafter b20/19-AKT3PROTAC+nbCD3/22) in T cells and evaluated the expression on day 7 after transduction (Supplementary Fig. 19a). A dose-dependent increase in the detection of nbCD3 and nbCD22 was observed in the culture supernatant, which was detected using the fluorescent ELISA assay for nbCD3/22 (Supplementary Fig. 19 b, c). To assess the relationship between BITE expression and CAR expression, we inhibited BITE secretion using Brefeldin A and performed intracellular flow cytometry analysis. CD22-PE and CD3-FITC antibodies were used to detect intracellular nbCD22 and nbCD3, respectively. Results demonstrated that nanobody expression positively correlated with surface expression of CD19 and CD20 CARs on T cells (Fig. 9b), indicating coordinated expression and secretion of the engineered constructs.
a Schematic of the strategy for trispecific CAR T cells, integrating b20/19-AKT3PROTAC with a secretory BiTE module consisting of nanobodies targeting CD3 and CD22 (nbCD3/22). b Correlation of expression of nbCD3, nbCD22, CD19 CAR, and CD20 CAR at various MOIs. The cells were treated with Brefeldin, and data were obtained using intracellular flow cytometry (n = 7 data points from three independent experiments). c Experimental setup for T cell activation, using Jurkat-GFP cells and Dynabeads (db) coated with CD3 to assess secreted nbCD3/22 functionality via flow cytometry. d Dose-dependent T cell activation (CD69 expression) in response to culture supernatants (used at various ratios with culture media) with nbCD3/22, using db coated with CD3 for validation (n = 6 data points from three independent experiments). e Line graph of HEK293T synNotch reporter assay showing dose-dependent inhibition of CD22-CAR signaling by nbCD22 in CAR T cell supernatants, confirming BiTE functionality under two condition 1 and condition 2. f Experimental timelines for in vitro T cell engineering, transduction, and co-culture with Raji cells (WT or knockout for CD19, CD20, or CD22). Anti-tumor assays were performed on days 9, 11, and 13. g, h Functional assay of CAR T cells against Raji cells (WT or knockout for CD19, CD20, or CD22) demonstrates that b20/19AKT3PROTAC CAR T cells co-expressing nbCD3/22 exhibit stronger antitumor activity compared to b20/19-AKT3PROTAC or mCD19 CAR T cells at Day 7 and Day 14. Data represent mean ± SEM. ****p < 0.001; ns: not significant. A nonparametric t-test was used for statistical analysis between groups. Source data are provided as a Source Data file.
To determine whether the secretory nbCD3 and nbCD22 are functional, we made a T cell activation model to study nbCD3 (Fig. 9c) and HEK293T reporter system based on synNotch system developed previously40 (Supplementary Fig. 20a). We then tested the supernatant obtained from CAR T cells expressing the b20/19-AKT3PROTAC/+nbCD3/22. A dose-dependent increase in CAR T cell activation was observed with nbCD3, which was also confirmed by Dynabeads (db) for CD3 (Fig. 9d, Supplementary Fig. 20b). Similarly, the reporter system showed a dose-dependent suppression of the CD22 CAR mediated signaling in HEK293T cells, which was neutralized by the supernatant containing nbCD22 (Fig. 9e). Functional testing against Raji cells (WT or knockout for CD19, CD20, or CD22) demonstrated that b20/19-AKT3PROTAC CAR T cells co-expressing nbCD3/22 exhibited better anti-tumor activity compared to counterparts lacking nbCD3/22 or mCD19 CAR T cells (Fig. 9f-h). These findings confirm the functional efficiency of the b20/19-AKT3PROTAC+nbCD3/22 CAR T design.
Next, we assessed in vivo efficacy of the nbCD3/CD22 BITE-secreting CAR T cells using the TR model with Raji or NALM6 cells engineered to lack CD19 and CD20 but retain CD22 expression (Fig. 10a). Mice treated with b20/19-AKT3PROTAC+nbCD3/22 CAR T cells exhibited robust and sustained anti-tumor responses over time (Fig. 9b, c). This was accompanied by enhanced CAR T cell survival (Fig. 9d, e). Importantly, the nbCD3/CD22 BITE was detectable in circulation alongside with the CAR T cells (Fig. 9f). Following TR, both BiTE and CAR T cell exhibited an initial rise, followed by a parallel decline upon tumor regression, suggesting a synchronized regulation of effector function and CAR T cell expansion and survival (Fig. 9b-e). These molecular effects translated into a clear survival advantage in mice, which was further supported by an increased frequency of memory CAR T cell subsets exhibiting reduced exhaustion marker expression (Fig. 9g, h). Importantly, toxicity assessment of the trispecific CAR T cells revealed no notable adverse effects, and cellular engraftment remained intact (Supplementary Figs. 21, 22, and 23).
a Experimental timeline for in vivo study in RajiWT or NALM6WT model followed by CAR T cell administration and TR with double knockout RajiCD19/CD20−/− or double knockout NALM6CD19/CD20−/− cells. Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6. b Bioluminescence imaging of Raji and NALM6 tumor-bearing mice treated with b20/19-AKT3PROTAC or b20/19-AKT3PROTAC+nbCD3/22 CAR T cells, monitored from Day 7 to Day 84. c Line graph of quantified tumor radiance over time, showing sustained tumor control in Raji and NALM6 models with b20/19-AKT3PROTAC+nbCD3/22. d Line graph of percentage of CAR T cells in the blood of Raji and NALM6 tumor-bearing mice treated with b20/19-AKT3PROTAC or b20/19-AKT3PROTAC+nbCD3/22, measured over 56 days. e Bar graph of CAR T cell populations in blood at Day 56. f Levels of nbCD3/22 (pg/mL) in the blood of Raji and NALM6 tumor-bearing mice treated with b20/19-AKT3PROTAC+nbCD3/22, measured over 56 days, showing sustained secretion. g Kaplan-Meier survival curves demonstrating improved survival with nbCD3/22-modified CAR T cells. h Bar graph and pie charts compare b20/19-AKT3PROTAC and b20/19-AKT3PROTAC+nbCD3/22, showing various memory T cell subsets over time (n = 5 mice) in all conditions. Data represent mean ± SEM. ****p < 0.001. A two-way ANOVA followed by post-hoc testing was applied. Source data are provided as a Source Data file.
Discussion
A major challenge in achieving durable responses with CAR T cell therapy is the restricted survival and long-term engraftment of the cells, especially in multi-antigen formats such as bispecific and trispecific CARs. While considerable efforts have focused on optimizing co-stim domains, the structural behavior of these complex designs remains insufficiently characterized. Elements such as multiple antigen-binding domains like scFvs, glycine-serine linkers, and modular co-stim domains can significantly influence CAR stability, surface expression, and function, ultimately impacting therapeutic efficacy41. To address these factors, we systematically screened a comprehensive library of mono, bi, and trispecific CAR constructs, varying antigen specificities, co-stim domains (CD28, 4-1BB, ICOS), and linker lengths. This large-scale screening was enabled through our AI-guided CARMSeD pipeline, which predicts functional vulnerabilities such as self-activation potential, exhaustion signatures, and CAR T cell death. Unlike previous AI models, which focus on single readouts such as activation42,43 or individual domain properties like co-stim44, our platform integrates multiple structural parameters and functional outputs to provide a holistic prediction of CAR behavior and CAR T cell function. Additionally, we developed a computational model to estimate scFv-scFv interaction propensities, addressing an important, previously overlooked mechanism underlying ligand-independent signaling in multiple CAR designs. Based on this extensive in silico screening, we identified CAR designs with reduced self-activation signaling and improved functional properties. Among top candidates, a bispecific CD20/19 CAR consistently showed high expression, low CARMSeD scores, and strong in vitro performance across proliferation, cytokine secretion, and cytotoxicity assays. These findings emphasize the importance of rational structural design in improving CAR T cell efficacy and survival, consistent with previous studies linking CAR architecture to therapeutic outcomes45,46.
To further improve the durability of these multi-antigen CAR T cells, we focused on the pathways that are integral to T cell memory. Based on the RNA-Seq analysis, AKT3 showed differential expression during CAR T cell exhaustion. As previous studies have shown AKT signaling is critical for the T cell memory formation, T cell exhaustion, and metabolic fitness, we explored the specific role of AKT3 further33,34,35. To achieve targeted modulation, we employed a PROTAC-based strategy for the selective degradation of AKT3. PROTACs represent a novel class of molecules that induce the targeted degradation of specific proteins via the ubiquitin-proteasome pathway47,48. Unlike traditional inhibitors that merely block protein function, PROTACs hijack E3 ligases to ubiquitinate the target protein, ensuring complete and irreversible removal47,49. In our study, PROTAC-mediated degradation of AKT3 led to a marked enhancement of CAR T cell survival, metabolic reprogramming, and resistance to exhaustion. Mechanistically, AKT3 degradation promoted a shift in T cell metabolism from glycolysis to OXPHOS, a hallmark of long-lived Tcm and Tscm subsets50,51,52. These subsets play a critical role in maintaining durable anti-tumor control and effective recall responses upon antigen re-exposure. Consistently, AKT3-targeted CAR T cells exhibited a substantial enrichment of Tcm and Tem populations, which closely correlated with sustained cytotoxic activity and prolonged survival in murine models. These memory-enriched CAR T cells demonstrated enhanced metabolic fitness, which translated into substantially improved tumor control across multiple B cell leukemia and lymphoma models. Importantly, this enhanced engraftment, and functionality aligns with emerging evidence that PROTACs, particularly bioPROTACs, can be harnessed to precisely degrade intracellular signaling components such as ZAP70, enabling fine-tuned modulation of CAR signaling53. While AKT3 has previously been targeted in T cells using small molecule inhibitors53, such approaches are often limited by transiently blocking kinase activity, off-target activity, systemic toxicity, and lack of temporal control. In contrast, our PROTAC-based system offers a programmable and modular strategy that couples CAR expression with AKT3 degradation within a single molecular construct, achieving cell-intrinsic specificity while minimizing systemic exposure. This integrated design, besides enhancing therapeutic efficacy, increases safety by preventing sustained or excessive signaling typically observed with small-molecule inhibitors. Thus, PROTAC-engineered CAR T cells provide a novel and clinically translatable platform to overcome exhaustion and enhance persistence.
The most prominent findings in our study were the upregulation of FOXO4 following AKT3 degradation. FOXO4 is a transcription factor involved in autophagy, metabolic homeostasis, and longevity, and plays a key role in downregulating mTOR signaling54,55. Given our previous findings that mTOR suppression is critical for CAR T cell memory formation, we were prompted to investigate the functional contribution of FOXO4 in this context more closely32. Unlike its family members FOXO1 and FOXO3, which are well-established regulators of T cell memory formation37,56,57, FOXO4 has not been extensively studied in the context of T cells. However, our findings reveal that FOXO4 plays a pivotal role in maintaining CAR T cell fitness. We observed that AKT3 suppresses FOXO4 activity through phosphorylation. Targeted degradation of AKT3 led to increased FOXO4 expression, which was associated with enhanced memory T cell formation, improved mitochondrial function, and reduced exhaustion. Importantly, the knockdown of FOXO4 abolished the functional benefits conferred by AKT3 degradation, confirming FOXO4 as a central mediator in this regulatory axis.
We further translated this strategy into a trispecific CAR T platform targeting CD19, CD20, and CD22. This platform included a secretory nanobody-based on CD3/CD22 T cell engager to address dual antigen escape, an emerging mechanism of relapse in B cell malignancies, besides CD19-negative relapse58. When combined with AKT3-targeted PROTAC approach, these trispecific CAR T cells exhibited sustained expansion, superior tumor control, and long-term survival even in antigen-heterogeneous settings. This triple-modality design proved effective in overcoming both structural and functional limitations inherent to complex CAR formats.
Importantly, the benefits of the AKT3-FOXO4 axis were not limited to hematological cancers. We further explored its potential in solid tumors, where CAR T cell therapy has historically been less effective due to hostile microenvironments and poor survival59,60. CAR T cells targeting Claudin18.2 (for gastric cancer) and EGFR (for NSCLC) were modified with AKT3PROTAC treatment. These cells showed improved trafficking, reduced exhaustion markers, and superior tumor regression in xenograft models, suggesting that this approach has broad applicability across cancer types.
Beyond AKT3, the PROTAC platform offers flexibility to target other regulators of CAR T cell exhaustion and immune evasion. For example, PROTACs directed against PD-1, TIGIT, or LAG-3 may enable real-time modulation of inhibitory checkpoint receptors, thereby enhancing T cell activity without the need for systemic checkpoint blockade61,62. Moreover, transient PROTAC exposure can fine-tune CAR T cell phenotypes without permanently altering the genome, offering a safer alternative to CRISPR or other gene-editing tools. Our findings position PROTAC technology as a versatile and powerful tool for next-generation CAR T cell engineering. When integrated with rational CAR design, co-stimulatory optimization, and antigen multiplexing, PROTAC-mediated protein degradation and recently developed synthetic gene switches hold the potential to surmount long-standing limitations of CAR T therapy63. These include interconnected molecular events such as antigen escape, exhaustion, poor metabolic fitness, and short-lived memory responses32,64.
In conclusion, our study introduces a multifaceted strategy combining high-throughput CAR screening, AI-assisted design, and PROTAC-based intracellular signaling modulation to generate functionally superior CAR T cells. This integrative approach represents a critical step forward in addressing the twin challenges of antigen escape and CAR T cell exhaustion and has the potential to extend CAR T efficacy across both hematologic and solid malignancies.
Methods
Cell line details
The cell lines K562, HEK293T, Raji, NALM6, Jurkat-GFP, AGS, A549 and Daudi were purchased from the American Type Culture Collection (ATCC) and cultured following ATCC-recommended protocols. HEK293T, AGS cells were maintained in DMEM (Gibco, #11965092) with 10% FBS (Gibco, #16140-071) at 37 °C, 5% CO₂, and 95% humidity, with subculturing at 70–80% confluency using trypsin (Gibco, #25200056). Raji, K562, Jurkat, NALM6, and Daudi cells were cultured in RPMI-1640 medium (Gibco, #11875093) with 10% FBS (Gibco, #16140-071) under similar conditions, with Raji and Daudi cells sub-cultured at 2–3 × 106 cells/mL and Jurkat, K562 and NALM6 at 1–2 × 106 cells/mL. Similarly, A549 cells were cultured under same conditions in F12/K media (Gibco, #21127022) with 10% FBS (Gibco, #16140-071).
Human samples
Peripheral blood mononuclear cells (PBMCs) were obtained from healthy adult volunteers (20 males and 23 females; aged 20–55 years) at Apollo Indraprastha Hospital, New Delhi. Sample collection and processing were conducted in accordance with the Declaration of Helsinki and approved by the Institutional Review Board of the CSIR-Institute of Genomics & Integrative Biology (Ethical Clearance Ref No: CSIR-IGIB/IHEC/2022-23/01). Written informed consent was obtained from all participants prior to sample collection. Additionally, PBMCs were collected from five patients, following written informed consent (as described in our previous article32 and Supplementary Note 1) and adherence to the hospital’s ethical and regulatory standards managed through Apollo Hospitals Educational and Research Foundation Institutional Ethics Committee (IRB-Ethics Waiver-IRB Application No. IAH-BMR-064/07-23). The blood samples utilized were surplus specimens that would otherwise have been discarded after routine marker analysis. These procedures adhered strictly to the hospital’s ethical and regulatory standards and were overseen by Dr. Gaurav Kharya, Senior Consultant in Pediatric Hematology, Oncology, and Immunology at Indraprastha Apollo Hospital.
Integrated meta-analysis of CAR T therapy resistance mechanisms
A meta-analysis was conducted to assess relapse patterns and structural vulnerabilities in CAR T cell therapy, specifically targeting CD19 and CD20 antigens in mono and bispecific constructs. Existing clinical data from 4129 patients treated with CAR T cells were compiled from peer-reviewed publications indexed on PubMed, registered clinical trial databases (including studies from the USA, China, and Europe), and preprint servers such as bioRxiv (Supplementary Data 1). Patient-level and cohort-level information were extracted to identify relapse rates, with a specific focus on CD19-negative tumor recurrences. Studies involving both monospecific and multi-antigen (bi and trispecific) CAR T therapies were included. The data compiled was analyzed to evaluate the frequency of antigen escape and CAR T cell engraftment. For automated retrieval, Python packages Biopython (Entrez module) and BeautifulSoup were used to query and parse PubMed records, while the requests library was applied to access clinical trial databases. Preclinical studies, incomplete reports, and duplicate datasets were excluded. All included records were manually curated and harmonized across studies to enable pooled estimation of treated, relapsed, and resistant cases stratified by mono- versus bispecific CAR T constructs.
Profiling of CD19, CD20, and CD22 receptor expression in ALL patients
Flow cytometry data from ALL patients were collected from Sir Ganga Ram Hospital (Delhi), Medanta Hospital (Gurugram), and Shalby Sanar International Hospital (Gurugram), India, following ethical approval and informed consent from all patients. Samples from over 129 ALL patients were included to assess the surface expression of CD19, CD20 and CD22 antigens (Supplementary Data 8). The data were analyzed and categorized based on intensity and distribution into four groups: positive (strong, uniform expression), dim (low-intensity expression), negative (no detectable expression), and heterogeneous (variable expression within the cell population).
Animal experiments
NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) (Stock No # 005557) and NOD-Prkdcem26Cd52Il2rgem26Cd22/NjuCrl (NCG) (Strain Code: 572) both male and female mice aged 6-8 weeks, were purchased from The Jackson Laboratory and Charles River Laboratories, respectively and housed in the Laboratory. These animals were kept under a 12 h light/dark cycle at 22 °C with free access to food and water. All animal experiments were conducted following approval from the respective Ethics Committee for Animal Experiments. All animal experiments were approved by the Institutional Animal Ethics Committee; CSIR-Institute of Chemical Biology, Kolkata, West Bengal, India; Regional Centre for Biotechnology, Faridabad, Haryana, India; and Adgyl Lifesciences, Hyderabad, India.
All transgenic animals were housed in a specific pathogen-free (SPF) barrier facility equipped with individually ventilated cages (IVCs), high-efficiency particulate air (HEPA) filtration, and restricted access to maintain pathogen exclusion. Cage changing and handling were performed in Class II biosafety laminar flow workstations to minimize contamination risks. Experimental and control animals were co-housed whenever possible and maintained under identical husbandry, enrichment, and environmental conditions to minimize cage or microbiome-related variability. Mice were euthanized using CO2 inhalation followed by cervical dislocation, in accordance with CPCSEA and institutional ethical guidelines.
Measurement of tumor burden
Tumor size was not assessed through in vivo BLI and corroborated by post-mortem tumor cell analysis, which together served as the primary endpoints for monitoring disease progression. Animals were monitored daily for clinical signs, including body weight loss, reduced mobility, and any indication of distress. Early endpoint criteria included a rapid increase in bioluminescent signal suggestive of uncontrolled tumor progression, marked clinical deterioration, or the appearance of pain or distress as defined by CPCSEA and institutional guidelines. Animals meeting these criteria were humanely euthanized using CO2 inhalation followed by cervical dislocation, in accordance with approved ethical protocols.
Reporter cell lines and peptide screening
Reporter systems were established as follows: A GFP-AKT3-mCherry reporter construct was generated in HEK293T cells via transduction with a lentiviral vector encoding mCherry-AKT3. The mCherry-AKT3 fusion plasmid was assembled using synthetic gene fragments (Genscript) and cloned using Gibson Assembly. The lentiviral vector used for transduction was prepared as previously described32. Stable GFP-AKT3-mCherry-expressing cell lines were established through single-cell sorting using FACS (BD Aria). Peptides targeting AKT3 were tested in these cells. Transfections were performed in 12-well plates at a density of 2 × 105 cells per well using 1 µg of plasmid DNA and Lipofectamine 3000 (Thermo Fisher Scientific, #L3000008). Cells were treated with PROTAC peptides (0.1–10 µM) for 24 hours, followed by assessment of AKT3 degradation using confocal microscopy (Zeiss LSM 880) and western blotting with anti-AKT3 (Cell Signaling Technology, #14982) and anti-GAPDH (Sigma-Aldrich, G8795) antibodies.
A SynNotch reporter system was developed using SynNotch plasmids obtained from Addgene, as described previously40. Synthetic gene fragments encoding the CD22 receptor were synthesized and cloned into the pLenti vector and transduced into HEK293T-GFP cells to create sender cells. A stable clone (CD22R-HEK293T-GFP) was selected by FACS (BD Aria). A separate synthetic fragment encoding CD22-scFv was cloned into the SynNotch vector (Plasmid #79125), and the resulting lentiviral construct was used to transduce HEK293T cells. Additionally, a vector encoding pHR_Gal4UAS_tBFP_PGK_mCherry (Plasmid #79130), enabling inducible BFP expression and constitutive mCherry expression, was used to generate receiver cells. Stable receiver HEK293T cells co-expressing both plasmids were isolated by FACS (BD Aria).
For the co-culture assay, sender and receiver cells were plated at a 1:1 ratio and treated with b20/19-AKT3PROTAC+nbCD3/22 supernatants (dilutions ranging from 1:2 to 1:10) for 48 h. BFP expression was assessed via confocal microscopy to confirm nbCD22-mediated inhibition of CD22-CAR signaling.
A Jurkat-GFP reporter assay was developed to assess nbCD3 functionality. Jurkat-GFP cells (1 × 105 per well) were plated in 96-well plates and incubated with the same supernatants (1:2 to 1:10 dilutions) for 24 h. Dynabeads CD3/CD28 (Thermo Fisher Scientific, #11132D) served as positive controls. Cells were stained for the T cell activation marker CD69 using an APC-conjugated anti-CD69 antibody (BD Biosciences, #555751). Flow cytometry was used to measure CD69 expression, confirming T cell activation.
Further, to model CD20 heterogeneity, K562 cells were transduced with lentiviral vectors encoding CD20 at varying expression levels (low, medium, high), sorted by FACS (BD Aria) based on CD20 expression (anti-CD20-PE, BD Biosciences, #556633). The cells were analyzed by super-resolution microscopy. Imaging was performed using a Zeiss Elyra PS.1 super-resolution microscope equipped with a 63× oil immersion objective. Structured Illumination Microscopy (SIM) was conducted using a 561 nm laser line. Image stacks were acquired and processed using the Zeiss Zen software, and final image analysis, including intensity profiling and comparative visualization of CD20 expression levels (low, medium, and high), was performed using FIJI (ImageJ). Similarly, normal fluorescent imaging was done using EVOS cell imaging with 40× objective (Thermo Fischer Scientific)
CAR library construction and in vitro screening
A library of 10,824 chimeric antigen receptor (CAR) constructs was systematically generated, encompassing mono, bi, and trispecific formats targeting CD19, CD20, and CD22 antigens. Constructs were designed with variability in glycine-serine (G4S) linker lengths (3, 4, or 5 G4S repeats connecting scFvs and VH/VL domains), a constant CD8 hinge domain, a transmembrane domain (CD8 or CD28 or predominantly ICOS-derived), co-stimulatory domains (CD28, 4-1BB, OX40, or ICOS) and a CD3ζ signaling domain for T cell activation. From this library, 1452 constructs were selected, and synthetic fragments or gblocks were ordered. These were then cloned into our in-house optimized lentiviral backbone plasmid using Gibson and Infusion assembly methods and subsequently used for in-vitro functional screening based on diversity in structural configurations.
Primary human T cells were isolated from PBMCs of three healthy donors using magnetic bead separation with CD3/CD28 Dynabeads (Thermo Fisher Scientific, #11132D). These T cells were then activated using anti-CD3/CD28 stimulation (T cell TransAct, Miltenyi Biotec) and cultured in TexMACS T cell culture medium supplemented with 100 IU/mL of human recombinant IL-7 and 200 IU/ml of human IL-15 (Miltenyi Biotec). 0.2 × 106 millions of these T cells were further transduced (for each CAR) with lentiviral concentrate at an MOI of 5 followed by a half media change the following day.
Functional screening evaluated three parameters: T cell activation (signal-1) via CD69 expression, exhaustion (signal-2) via PD-1 expression, and apoptosis (signal-3) via 7-AAD staining, all measured by flow cytometry acquiring 20,000 cells for each individual CARs using anti-CD69-FITC (BD Biosciences, # 555748), anti-PD-1-PE (BD Biosciences, #560795), and 7-AAD apoptosis detection kit (Thermo Fisher Scientific, # V35123. Importantly, these assays were conducted without co-culture with tumor cells to confirm that observed changes in activation, exhaustion, or death were antigen-independent, and instead reflected intrinsic effects such as CAR structure, aggregation, or CAR-CAR self-interactions. Signal intensities were categorized into low (L), medium (M), and high (H) levels based on cutoffs established using a second-generation monospecific CD19 CAR as a reference with CD19 ECD-coated wells of a 96-well plate for the activation of the CAR T cells. CD19 ECD was used at various concentrations (0–40 μg/mL) to measure the three signals to generate the reference signals: Signal-1 (CD69; activation; base-line value: <1), Signal-2 (PD-1; exhaustion; <0.25), and Signal-3 (7-AAD; cell death; <0.25). A scoring system was applied to classify CARs into low, medium, and high CARMSeD categories, reflecting their propensity for self-activation and dysfunction.
AI-guided screening with CARMSeD model
To facilitate downstream classification, signal values were discretized into categorical bins: Signal-1 was divided into low, medium, and high using thresholds at 1.0; Signal-2 and Signal-3 were binned using thresholds at 0.5. Each CAR sequence was then assigned to one of three functional synergy categories, L-CARMSeD, M-CARMSeD, or H-CARMSeD based on a rule-based mapping of all valid signal triplets. Specifically, 27 possible combinations of the discretized signals (e.g., low-low-high) were heuristically grouped into these three classes to reflect combinatorial synergy and downstream activation potency, informed by empirical distribution of in vitro measurements. The grouping was based on a consensus rule wherein triplets with higher aggregate signal strength and presence of multiple high-signal axes (e.g., H-H-M or H-M-H) were mapped to the H-CARMSeD class, while weaker combinations (e.g., L-L-L, L-M-L) were assigned to L-CARMSeD. This rule-based mapping was finalized in consultation with experimental collaborators based on activation potential observed in vitro. CAR sequences were tokenized using a fixed amino acid vocabulary of 20 standard residues, with unknown or non-canonical residues assigned to a special out-of-vocabulary token. All sequences were truncated or zero-padded to a uniform length of 1024 residues. The model, termed CARMSeD, was implemented in TensorFlow and structured as a multi-task deep neural network. The architecture began with a trainable embedding layer (dimensionality 128), followed by two one-dimensional convolutional layers, each with 256 filters and a kernel size of 5, using ReLU activations. A max-pooling operation was followed by a global max pooling layer to condense spatial representations. The resulting latent feature map was branched into two task-specific heads: a classification head with SoftMax activation to predict the CARMSeD class, and a regression head with linear activation to predict the continuous values of Signal-1, Signal-2, and Signal-3.
Model training was conducted for up to 50 epochs using the Adam optimizer with a batch size of 64. An early stopping criterion based on validation classification accuracy was employed to prevent overfitting. The composite loss function combined categorical cross-entropy (for classification) and mean squared error (for regression), with loss weighting of 3:1 to prioritize accurate functional classification. All signal values were normalized to the [0, 1] range using MinMaxScaler prior to training. The dataset was split into training and validation cohorts in an 80:20 ratio using stratification based on the CARMSeD class and discretized Signal-3 values to preserve class balance. Model performance was evaluated using classification accuracy, confusion matrices, mean squared error, and the coefficient of determination (R²) for regression tasks. After training, the CARMSeD model was applied to held-out test datasets to generate predictions for both categorical class labels and signal values. Model outputs were visualized using Python libraries matplotlib and seaborn. Visual inspection included measured vs. predicted scatter plots for each signal axis, residual error distributions, and class-specific confusion matrices to characterize error structure and functional resolution across the prediction space. Finally, the trained model was benchmarked across multiple random initialization seeds to confirm stability and was then integrated into the downstream CAR design workflow to prioritize constructs predicted to exhibit high functional synergy.
Primary cell isolation and transduction
The PBMCs were extracted from whole blood samples by density gradient centrifugation using Ficoll-Paque (Cytiva, #17144002) as per the manufacturer’s guidelines. Subsequently, CD4/CD8 magnetic beads were used to isolate T cells. Similarly, CD4 and CD8 magnetic beads (Miltenyi Biotec, #130-116-480) were used to isolate pure CD4+ and CD8+ T cell populations. These T cells were then activated using anti-CD3/CD28 stimulation (T cell TransAct, Miltenyi Biotec, #130-128-758) and cultured in TexMACS T cell culture medium (Miltenyi Biotec, #130-097-196) supplemented with 100 IU/mL of human recombinant IL-7 and IL-15 (Miltenyi Biotec, #130-095-367 and #130-095-760). Following 24 h of activation, the T cells were transduced with lentiviral concentrate at an MOI of 5 and incubated for an additional 48 hrs before media change. Fresh medium was replenished every 2 days throughout the experiment. The percentage of transduced T cells was determined using the CD19 CAR detection reagent (Miltenyi Biotec, #130-129-550) following the manufacturer’s instructions, and flow cytometry analysis was done after day 5 of transduction, unless otherwise specified.
Patient blood samples were processed to isolate PBMCs using Ficoll density gradient centrifugation. PBMCs were further separated into T cells and B cells. For T cell isolation, standard procedures were followed, while for B cell isolation, magnetic beads conjugated to CD19 antibodies (Miltenyi Biotec, #130-091-151) were utilized. The isolated B cells were confirmed via flow cytometry using CD19-specific antibodies. After isolation, the cells were cultured in RPMI medium (Gibco, #11875093) supplemented with 10% FBS (Gibco, #16140-071) and subsequently co-cultured with CAR T cells, which were generated from the patient-derived T cells. All procedures adhered strictly to ethical guidelines and regulatory standards to ensure patient safety and data integrity.
Humanization and computational validation of scFvs
To evaluate the structural and dynamic properties of bispecific (anti-CD19/CD20) and trispecific (anti-CD19/CD20/CD22) CAR constructs, a computational pipeline was established. Initial murine scFv sequences and structures for anti-CD19 and anti-CD20 were retrieved from the Protein Data Bank (PDB IDs: 7URV and 6VJA, respectively), while the murine anti-CD22 (clone - RFB4) scFv sequence was obtained from the literature65. All possible bispecific and trispecific constructs were generated using AlphaFold2 (DeepMind) for ab initio structure prediction, followed by structural refinement and visualization in UCSF Chimera (v1.16). Molecular dynamics (MD) simulations were performed to investigate intramolecular interactions between scFvs within these constructs using Amber2266 with the AMBER ff14SB force field and TIP3P water model. The simulation protocol consisted of three stages: energy minimization, heating, and production. Systems were minimized over 10,000 steps, with 2000 steps of steepest descent followed by 8000 steps of conjugate gradient. The minimized systems were heated to 300 K over 150 ps under NVT conditions, followed by a 200 ns production run under NPT conditions at 1 bar and 300 K, using a 2 fs timestep. Trajectory analysis was conducted using CPPTRAJ (AmberTools67) to calculate root mean square deviation (RMSD) and intramolecular hydrogen bond interactions, focusing on complementarity-determining regions (CDRs).
Humanization of the murine anti-CD19, and anti-CD20 scFvs was performed using BioPhi68 (v1.0), a web-based tool offering two approaches: CDR grafting and the Sapiens iterative humanization method. Both CDR grafting and Sapiens method were used to generate library of humanized sequences. From those, the one with high humanness in both approaches were selected to iteratively optimize framework regions while preserving CDR integrity, generating humanized scFv variants with high sequence identity to human germline frameworks. The resulting humanized scFvs were modeled using the corresponding murine structures (PDB IDs: 7URV, and 6VJA) as templates in MODELLER (v10.4), producing 10 homology models per scFv, with the best model selected based on the DOPE score, while for humanized anti-CD22 only scFv was modeled using AlphaFold and docked to the previously mentioned C-like domain-2 region of CD2269. These models were docked against their respective antigens (CD19, CD20) using the HADDOCK webserver (v2.4)70, employing default parameters to predict/generate best-fitting complexes. To compare the binding affinities of murine and humanized scFv variants, MD simulations were conducted using the same AMBER ff14SB force field and TIP3P water model. Systems were prepared as described above, but production runs were extended to 300 ns to ensure convergence of binding interactions. Trajectory analysis included RMSD, hydrogen bond analysis, per-residue decomposition, and enthalpy calculations using the MM-GBSA method in AmberTools2367, validating structural stability, interaction profiles, and binding affinity of the humanized variants.
The bispecific and trispecific combinations were then modeled with optimized humanized scFvs via homology modeling on previously described structures, to preserve spatial orientation between scFvs in the bispecific construct. The resulting bispecific structure was subjected to MD simulation following the same protocol as above, with a 200 ns production run. Final trajectory analysis focused on RMSD and intramolecular hydrogen bond interactions within CDRs to confirm structural stability and inter-scFv interactions critical for antigen recognition.
In vitro functional validation
K562 cells were engineered to stably express CD19, CD20, and CD22 individually, as well as in combination, using lentiviral transduction. Synthetic gene fragments encoding CD19 (NM_001385732.1), CD20 (NM_021950.4), and CD22 (NM_001185099.2) were cloned into a previously described lentiviral backbone32. Stable cell lines were established by FACS (BD Aria) sorting using anti-CD19, anti-CD20, and anti-CD22 antibodies, and expression of each antigen was confirmed by flow cytometry. Cytotoxicity assays were conducted by co-culturing CAR T cells (b20/19, b22/19, and t20/19/22) with K562 target cells at effector-to-target (E:T) ratios of 1:1, 2.5:1, 5:1, and 10:1 for 24 h. Tumor cell lysis was quantified using 7-AAD staining (BD Biosciences #559925). T cell proliferation was assessed as previously described32. Raji cell knockout clones were generated using gene-specific gRNAs targeting CD19, CD20, and CD22. A detailed protocol for the generation of these stable Raji knockout cell lines is published in our recent article39.
CAR T cell proliferation, durability, and antitumor activity
CAR T cell proliferation was assessed using the CellTrace CFSE Cell Proliferation Kit (Thermo Fisher Scientific, #C34554). Briefly, CAR T cells were stained with CFSE dye, and proliferation was monitored at various time points in the presence of the tumor cells (Days 7, 9, 11, 13, 15, 17, 19, and 21), as previously described32. Initially, CAR T cells were harvested and washed to ensure viability and purity. CAR T cell survival was measured by counting cell numbers on Day 0 (prior to co-culture with tumor cells) and on Days 7, 13, 21, and 28 following co-culture with tumor cells. Co-cultures were performed on alternate days (Days 6, 8, 10, 12, 14, 16, 18, and 20). The assay methodology followed our previously published protocol32.
The data shown in Cell death analysis was conducted using 7-AAD staining (BD Biosciences #559925), followed by acquisition on a flow cytometer, as previously described32. Data were presented as either representative flow cytometry histograms showing the percentage of mean fluorescence intensity of 7-AAD, or as bar graphs indicating percent cell death.
Transcriptomic and functional analysis of AKT3
CAR T cells were isolated from control and tumor-rechallenged (TR) mice (n = 5 per group) treated with b20/19 CAR T cells. RNA was extracted using QIAGEN RNeasy Mini Kit, following further enrichment and heat fragmentation, RNA was reverse-transcribed into cDNA, employing RNase H-Reverse Transcriptase and primers for strand-specific ligation. and transcriptomic profiling was performed via RNA sequencing on an Illumina NovaSeq6000. Differential gene expressions were analyzed using iGeak_RNA-seq, focusing on pathways such as oxidative phosphorylation, FOXO signaling, and autophagy. The RNA Seq analysis was done as described by us previously32. AKT3 upregulation was confirmed by flow cytometry using anti-AKT1/2/3-FITC (Thermo Fisher, #BS-6951R-FITC). AKT3 overexpression was achieved by co-transducing CAR T cells with a lentiviral vector encoding AKT3 via pLenti-AKT3, while knockdown was performed using shRNA (Supplementary Data 9).
PROTAC design and validation
AKT3-targeting PROTAC peptides were designed using RF diffusion modeling with AlphaFold (https://github.com/RosettaCommons/RFdiffusion), yielding a library of peptides of which top five candidate peptides (AKT3_P1 to AKT3_P5) and and a non-targeting peptide (NTP) as a control were selected based on the sequence diversity, secondary structure and complex structure confidence score. The designed PROTACs were checked for binding to AKT3 and VHL E3 ligase, respectively. In addition to this, the stability of the designed PROTACs was checked in silico. Each peptide was fused to a TAT-derived cell-penetrating peptide (CPP) and a VHL E3 ligase ligand via a flexible linker to facilitate intracellular delivery and ubiquitin-mediated degradation. Degradation efficacy was evaluated in HEK293T cells (ATCC, #CRL-3216) stably expressing a GFP-AKT3-mCherry reporter, seeded at 2 × 10⁵ cells per well in 12-well plates. Cells were treated with peptides at concentrations ranging from 0.1 to 10 µM for 24 h in DMEM (Gibco, #11965092) supplemented with 10% FBS (Gibco, #16140-071). The peptides were synthesized from GenScript at a concentration of 2–4 mg each. AKT3 degradation was quantified by fluorescence microscopy on a Zeiss LSM 880 confocal microscope, measuring the mCherry/GFP fluorescence ratio with ImageJ (v1.53), and by western blotting using antibodies against AKT3 (1:1000, Cell Signaling Technology, #14982), GFP (1:2000, Abcam, #ab290), and GAPDH (1:5000, Sigma-Aldrich, #G8795) as a loading control. The top-performing peptide, P2, was initially tested in GFP-expressing CAR T cells and endogenous AKT3 was detected using rabbit anti-AKT3 primary antibody (Cell Signaling Technology, #14982) followed by staining with Alexafluor 546-conjugated secondary anti-rabbit antibody (ThermoFischer Scientific, # A-11035). Notably, AKT3-PROTAC (without CPP) was integrated into the b20/19 CAR construct downstream of the CAR sequence using a P2A self-cleaving peptide linker, cloned into a pLenti vector. Primary T cells transduced with this construct were co-cultured with CD19-/- Raji cells at an effector-to-target (E:T) ratio of 2.5:1 for 24 h, and AKT3 degradation was confirmed by fluorescence microscopy and western blotting. Cytotoxicity was assessed against wild-type Raji cells and patient-derived leukemia samples (n = 5) at the same E:T ratio for 24 h, with target cell survival quantified by flow cytometry using 7-AAD staining (Thermo Fisher Scientific #A1310). CAR T cell survival was monitored over 15 days in a tumor rechallenge model with CD19−/− Raji cells, while T cell subsets (Tcm, Tem) and exhaustion markers (PD-1) were analyzed by flow cytometry using anti-CD62L-FITC (BD Biosciences # 561917), anti-CD45RO-PE (BD Biosciences, # 555493), and anti-PD-1-APC (BD Biosciences, # 558694). Metabolic profiles, including extracellular acidification rate (ECAR) and oxygen consumption rate (OCR), were measured as described by us previously32.
Functional and biochemical assays
Immunoblotting for AKT3 and GAPDH was performed as previously described32. Similarly, RT-qPCR assays were conducted following our established protocols. Primers used for AKT3 and FOXO4 are listed in Supplementary Data 9. Total RNA was extracted using the RNeasy Mini Kit (Qiagen, #74104) according to the manufacturer’s instructions, and RNA integrity was assessed using a NanoDrop One or Qubit Instrument (Thermo Fisher Scientific). FOXO4 expression was quantified by quantitative real-time PCR (qRT-PCR) using a QuantStudio 3 system (Applied Biosystems), with primers designed via Primer-BLAST (Supplementary Data 9).
FOXO4 and AKT3 knockdown was performed using pLKO.1 lentiviral vector carrying FOXO4-shRNA and AKT3-shRNA, with efficiency confirmed by qRT-PCR and western blotting as described by us previously. The mTOR activity assay was performed using the K-LISA mTOR Activity Kit (Merck Millipore; #CBA104) as per the manufacturer’s instructions and as per our previously described method. Absorbance at 450 nm was measured on a VICTOR Nivo Multimode Reader (PerkinElmer) and reported as percentage mTOR activity32.
CAR binding analysis
Binding affinity analysis of CAR scFvs targeting CD19, CD20, and CD22 was performed using a flow cytometry-based method (BD FACSLyric). T cells were transduced with lentiviral vectors encoding the respective CAR scFvs. Following transduction, cells were fixed and stained using anti-CAR antibodies that recognize the extracellular domain (ECD) of the respective receptors: CD19 (Miltenyi Biotec, #130-129-550) and CD22 (Miltenyi Biotec, #130-126-727). The CAR ECDs were biotin-conjugated, and detection was carried out using PE-conjugated anti-biotin REAfinity antibody (clone REA746, Miltenyi Biotec, #130-110-951). For CD20 binding analysis, a plasmid encoding the CD20 extracellular domain (ECD) was generated. A synthetic CD20 ECD fragment was cloned into the vector pD649-HAsp-CD22-Fc (DAPA)-AviTag-6xHis (Addgene Plasmid #156940). This construct was used to purify recombinant CD20 protein conjugated to biotin. The CD20-biotin protein was then used for assessing binding to CD20-targeting CAR scFvs. Protein purification, expression and isolation of recombinant CD20 were performed according to our previously published protocol71. Binding affinity data were analyzed using FlowJo software and presented as percentage binding.
To identify the various CAR components and the BiTE, detection reagents were applied to the respective scFvs: CD19 CAR detection for CD19 scFv, CD20 CAR detection for CD20 scFv, and CD22 CAR detection to identify the BiTE containing CD22 scFv linked to the CD3 nanobody.
Binding kinetics between the CD20 receptor and biotinylated CD20 CAR detection reagent were analyzed using a Biolayer Interferometry platform (Gator Pilot) with streptavidin (SA) biosensors (PN: 160002, Gator R Probes). Biotinylated ligand was immobilized on pre-hydrated SA probes (10 min), followed by a 120 s baseline equilibration. Recombinant murine and HuCD20 antibodies (200 µg/mL stock) served as analytes, prepared as a five-point, two-fold serial dilution (12.5–50 nM) in running buffer [PBS (pH 7.4), 0.1% BSA, 0.02% Tween-20]. Association and dissociation phases were monitored in real time, with buffer-only wells included as 0 nM controls. Bare SA probes, irrelevant biotinylated protein, and buffer-only wells were used as controls for non-specific binding and bulk refractive index shifts. Sensorgrams were double-referenced, and data globally fitted to a 1:1 Langmuir model to calculate Kon, Koff, and KD values.
Flow cytometry analysis
Flow cytometry was performed using BD FACS Aria, Accuri, Lyric systems, and Beckman Coulter CytoFLEX Flow Cytometer and data were analyzed with FlowJo or CytExpert software. The flow cytometry was performed as described by us previously32. Briefly, CD19, CD22 CAR expression was evaluated using CD19 and CD22 CAR detection reagents (Miltenyi Biotec, #130-129-550, #130-126-727) and CD20 CAR expression was evaluated using Protein L-APC (CST #29480) or biotinylated anti CD20 antibody (Acro biosystems) followed by PE-conjugated anti-biotin secondary antibodies (Miltenyi Biotec, #130-110-951). As described previously, T cell immunophenotyping and functional marker analyses (e.g., Granzyme B, PD-1, LAG-3) were performed using fluorophore-conjugated antibodies. Antibodies against CD45RO (BD Biosciences, # 555493), CCR7 (BD Biosciences, #561271), and PD-1 (BD Biosciences, # 558694) were used to identify memory and exhausted T cell subsets. Cell death was determined using 7-AAD staining (BD Biosciences #559925). For FOXO4 and phospho-FOXO4 (Ser193) levels, cells were fixed/permeabilized with the Foxp3/Transcription Factor Staining Buffer Set (eBioscience, #00-5523-00) and stained with anti-FOXO4-APC (Abcam, ab211689) and anti-phospho-FOXO4-PE (Cell Signaling Technology, #9471). The data was analyzed using FlowJo software and presented as mean fluoresce intensity (MFI) unless specified. The list of antibodies with catalog number is given in Supplementary Data 10.
A detailed analysis of the flow cytometry data and the complete sequential gating strategy has been performed in accordance with our previously published study32. As the gating workflow for CAR T cell phenotyping and functional profiling remains consistent with that publication, the comprehensive gating schematic provided in Supplementary Fig. 1 of the earlier study fully represents the gating strategy used here.
Super-resolution microscopy
Super-resolution microscopy was done to visualize low-level CD20 expression in K562 cells. A total of 1.5 × 106 cells were harvested from actively growing cultures and washed twice with 1× phosphate-buffered saline (PBS) (Thermo Fisher Scientific, #10010023). Cells were then incubated with a fluorophore-conjugated CD20 antibody (Sigma-Aldrich, #SAB4700118) prepared at a dilution of 2 µL antibody per 100 µL 1×PBS, following the manufacturer’s instructions. The staining was carried out at 4 °C for 20 min. Following staining, cells were washed twice with 1×PBS and subsequently fixed with 4% paraformaldehyde at room temperature for 15 min. Fixed cells were washed and mounted onto poly-L-lysine-coated glass slides (Sigma-Aldrich, #P0425-72EA) using ProLong Gold Antifade Mountant (Thermo Fisher Scientific, #P36930). The slides were allowed to dry overnight at room temperature in the dark. Imaging was performed using a Zeiss ELYRA PS.1 3D PALM, equipped with a 63× oil immersion objective (Plan-Apochromat 63×/1.40 Oil DIC; Filter: MBS-561 + EF BP 570–650/LP750; 100 mW laser). Structured Illumination Microscopy (SIM) was carried out using a 561 nm laser line. Image stacks were acquired and processed using Zeiss Zen software. Final image analysis, including intensity profiling and comparative visualization was performed using ImageJ.
In vivo studies
For B cell malignancy xenograft tumor models, mice were injected intravenously with 0.5 × 106 Raji or NALM-6 cells expressing luciferase on day 0, followed by 5 × 106 CAR T cells (b20/19, b20/19-AKT3PROTAC, or b20/19-AKT3PROTAC+nbCD3/22) on day 5, unless specified. Tumor rechallenge with 0.5 × 106 CD19−/− or CD19/CD20−/− cells was performed on days 12, 19, and 26 and so on. Tumor burden was monitored by bioluminescence imaging using an IVIS Spectrum, PerkinElmer, after intraperitoneal injection of D-luciferin (Thermo Fisher Scientific, # 88293) as per the protocol previously described32. CAR T cell survival and nbCD3/22 levels in peripheral blood were quantified by flow cytometry and ELISA, respectively. Survival was analyzed using Kaplan-Meier curves with the Mantel-Cox test. The tumor models were repeated twice, independently.
For solid tumor xenograft models, gastric cancer (AGS, Claudin18.2+) and NSCLC (A549, EGFR+) tumors were established by subcutaneous injection of 3 × 105 ASG cells or 5 × 106 A549 cells into NSG mice, and once tumors reached ~100 mm³ (around day 10), mice were randomized and treated intravenously with 1 × 106 CAR T cells targeting Claudin18.2 or 3 × 107 targeting EGFR. Tumor progression was monitored by caliper measurements and expressed as tumor volume (mm3), while body weight and survival kinetics were recorded throughout the study.
Trispecific CAR T cell engineering
Trispecific CAR T cells were engineered by integrating the b20/19-AKT3PROTAC construct with a secretory CD3/CD22 bispecific T cell engager (BiTE, nbCD3/22). The nbCD3 and nbCD22 nanobodies were humanized via CDR grafting using BioPhi68 (v1.0) to minimize immunogenicity while preserving antigen-binding affinity. Humanized nanobody sequences were synthesized (GenScript) and cloned into a pLenti vector downstream of the b20/19-AKT3PROTAC sequence, separated by a P2A self-cleaving peptide to ensure co-expression, using Gibson Assembly (New England Biolabs). Constructs were sequence-verified by Sanger sequencing. Primary human T cells, isolated and activated as described above, were transduced with the lentiviral concentrate at MOI ranging from 2 to 10 and incubated for an additional 48 h before media change. Fresh medium was replenished every 2 days throughout the experiment. Cells were cultured in TexMACS T cell culture medium (Miltenyi Biotec, #130-097-196) supplemented with 100 IU/mL of human recombinant IL-7 and IL-15 (Miltenyi Biotec, #130-095-367 and #130-095-760) and cultured for 7 days.
Expression of nbCD3/22 was quantified on day 7 post-transduction by fluorescent ELISA using a custom anti-CD3/anti-CD22 detection system with a fluorescence VICTOR Nivo Multimode Microplate Reader (Revvity). Intracellular expression was confirmed by flow cytometry using anti-CD3-FITC (1:100, BD Biosciences, #570821) and anti-CD22-PE (1:100, BD Biosciences #562859) antibodies after treating cells with Brefeldin A (5 µg/mL, Sigma-Aldrich, #B6542) for 4 h to block secretion, followed by fixation and permeabilization. Functionality of the secreted nbCD3/22 BiTE was assessed in two systems: (1) a Jurkat-GFP activation model, where Jurkat-GFP cells (1 × 105 cells/well in 96-well plates) were incubated with CAR T cell supernatants (1:2 to 1:10 dilution) for 24 h, and T cell activation was measured by CD69 expression using anti-CD69-APC (1:100, BD Biosciences, #555533) via flow cytometry, with Dynabeads Human T-Activator CD3/CD28 (Thermo Fisher Scientific) as a positive control; and (2) a HEK293T SynNotch reporter assay, where sender cells (expressing CD22R-HEK293T-GFP), and receiver cells (expressing anti-CD22-SynNotch, GAL4-UAS-BFP, and constitutively expresing mCherry) were co-cultured at a 1:1 ratio with CAR T cell supernatants (1:2 to 1:10 dilution) for 48 h, and BFP expression was quantified by confocal microscopy to confirm nbCD22-mediated inhibition of CD22-SynNotch signaling (Leica EM UC7, Leica, Wetzlar, Germany) as described by us previously72.
Image illustration
All illustrated schematic images in Fig. 1C, 2 (A, H), 3A, 4A, 6M, S2A, S17A, and S18 (A, E), were either completely or partially created using BioRender. (Created in BioRender. Chauhan, V. (2025) https://BioRender.com/uj3gas6)
Statistical analysis
Statistical analyses were performed using GraphPad Prism version 10.3 (GraphPad Software). Group differences were evaluated using nonparametric t-tests. Bar charts and associated P values were generated with Prism. For comparisons involving more than two groups, a nonparametric One-way ANOVA or Two-way ANOVA was applied, while survival analyses between two groups were performed using the Mantel-Cox (log-rank) test. Data are presented as the mean ± standard error of the mean (SEM). Statistical significance was indicated as follows: ****P < 0.001, ***P < 0.005, **P < 0.01, and *P < 0.05. Unless mentioned otherwise in figure legends, experiments included a minimum of three biological replicates and were repeated independently at least three times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Individual clinical data cannot be publicly shared to protect patient privacy, but further de-identified participant data are available from the corresponding authors on request. The RNA-seq data have been deposited in the Github: 10.5281/zenodo.17852612. Source data are provided with this paper.
Code availability
All computational scripts and AI-based modeling workflows used in this study are available at the following GitHub repository: https://github.com/NishaChaudhary23/CAR-AI. Additional code supporting the analyses is available from the authors upon reasonable request.
References
Cappell, K. M. & Kochenderfer, J. N. Long-term outcomes following CAR T cell therapy: what we know so far. Nat. Rev. Clin. Oncol. 20, 359–371 (2023).
Mishra, A., Maiti, R., Mohan, P. & Gupta, P. Antigen loss following CAR-T cell therapy: mechanisms, implications, and potential solutions. Eur. J. Haematol. 112, 211–222 (2024).
Maude, S. L. et al. Tisagenlecleucel in children and young adults with B-cell lymphoblastic leukemia. N. Engl. J. Med. 378, 439–448 (2018).
Majzner, R. G. et al. Low CD19 antigen density diminishes efficacy of CD19 CAR T cells and can be overcome by rational redesign of CAR signaling domains. Blood 132, 963–963 (2018).
Wang, L. et al. Bispecific CAR-T cells targeting CD19/20 in patients with relapsed or refractory B cell non-Hodgkin lymphoma: a phase I/II trial. Blood Cancer J. 14, 130 (2024).
Zhang, Y. et al. Long-term activity of tandem CD19/CD20 CAR therapy in refractory/relapsed B-cell lymphoma: a single-arm, phase 1–2 trial. Leukemia 36, 189–196 (2022).
Cordoba, S. et al. CAR T cells with dual targeting of CD19 and CD22 in pediatric and young adult patients with relapsed or refractory B cell acute lymphoblastic leukemia: a phase 1 trial. Nat. Med. 27, 1797–1805 (2021).
Spiegel, J. Y. et al. CAR T cells with dual targeting of CD19 and CD22 in adult patients with recurrent or refractory B cell malignancies: a phase 1 trial. Nat. Med. 27, 1419–1431 (2021).
Schneider, D. et al. Trispecific CD19-CD20-CD22-targeting duoCAR-T cells eliminate antigen-heterogeneous B cell tumors in preclinical models. Sci. Transl. Med. 13. (2021).
Wang, L. et al. Bispecific CAR-T cells targeting CD19/20 in patients with relapsed or refractory B cell non-Hodgkin lymphoma: a phase I/II trial. Blood Cancer J. 14. https://doi.org/10.1038/S41408-024-01105-8 (2024).
Shah, N. N. et al. Bispecific anti-CD20, anti-CD19 CAR T cells for relapsed B cell malignancies: a phase 1 dose escalation and expansion trial. Nat. Med 26, 1569–1575 (2020).
Larson, S. M. et al. CD19/CD20 bispecific chimeric antigen receptor (CAR) in naive/memory T cells for the treatment of relapsed or refractory non-Hodgkin lymphoma. Cancer Discov. 13, 580–597 (2023).
Shah, N. N. et al. Phase 1 Study of trispecific anti-CD20, anti-CD19, anti-CD22 (CAR20.19.22) T cells for relapsed, refractory B-cell malignancies. Blood 144, 3448–3448 (2024).
Jayaraman, J. et al (2020). CAR-T design: elements and their synergistic function. EBioMedicine 58. https://doi.org/10.1016/j.ebiom.2020.102931 (2020).
Guedan, S., Calderon, H., Posey, A. D. & Maus, M. V. Engineering and design of chimeric antigen receptors. Mol. Ther. Methods Clin. Dev. 12, 145–156 (2019).
Brillembourg, H. et al. The role of chimeric antigen receptor T cells targeting more than one antigen in the treatment of B-cell malignancies. Br. J. Haematol. 204, 1649–1659 (2024).
Yin, C. et al. CAR T cell exhaustion but not ex vivo cytotoxicity is predictive of patient clinical response: an interim analysis of ACIT001/EXC002, a phase Ib/II trial of decentralized production of CAR T cells for treatment of relapsed/refractory aggressive NHL and ALL. Blood 142, 2092–2092 (2023).
Sheih, A. et al. (2020). Clonal kinetics and single-cell transcriptional profiling of CAR-T cells in patients undergoing CD19 CAR-T immunotherapy. Nat. Commun. 11. https://doi.org/10.1038/S41467-019-13880-1
Stewart, C. M. et al. IL-4 drives exhaustion of CD8 + CART cells. Nat. Commun. 15, 7921 (2024).
Tao, Z. et al. Impact of T cell characteristics on CAR-T cell therapy in hematological malignancies. Blood Cancer J. 14, 213 (2024).
Ghorashian, S. et al. Enhanced CAR T cell expansion and prolonged persistence in pediatric patients with ALL treated with a low-affinity CD19 CAR. Nat. Med. 25, 1408–1414 (2019).
Park, J. H. et al. Long-term follow-up of CD19 CAR therapy in acute lymphoblastic leukemia. N. Engl. J. Med. 378, 449–459 (2018).
Wang, Y. et al. CD19-specific CAR-T cell treatment of 115 children and young adults with acute B lymphoblastic leukemia: long-term follow-up. Cancer Res. Treat. 56, 945–955 (2024).
Guedan, S. et al. ICOS-based chimeric antigen receptors program bipolar TH17/ TH1 cells. Blood 124, 1070–1080 (2014).
Lee, E. H. J. et al. Antigen-dependent IL-12 signaling in CAR T cells promotes regional to systemic disease targeting. Nat. Commun. 14, 4737 (2023).
Svoboda, J. et al. Enhanced CAR T-cell therapy for lymphoma after previous failure. N. Engl. J. Med. 392, 1824–1835 (2025).
Alizadeh, D. et al. IL15 enhances CAR-T cell antitumor activity by reducing mTORC1 activity and preserving their stem cell memory phenotype. Cancer Immunol. Res. 7, 759–772 (2019).
Philipson, B.I. et al. 4-1BB costimulation promotes CAR T cell survival through noncanonical NF-κB signaling. Sci. Signal. 13. https://doi.org/10.1126/SCISIGNAL.AAY8248 (2020).
Derigs, P. et al. Third-generation anti-CD19 CAR T cells for relapsed/refractory chronic lymphocytic leukemia: a phase 1/2 study. Leukemia 38. https://doi.org/10.1038/S41375-024-02392-7 (2024).
Schubert, M.L. et al. Treatment of adult ALL patients with third-generation CD19-directed CAR T cells: results of a pivotal trial. J. Hematol. Oncol. 16. https://doi.org/10.1186/S13045-023-01470-0 (2023).
Lakhani, A. et al. Extracellular domains of CARs reprogramme T cell metabolism without antigen stimulation. Nat. Metab. 6, 1143–1160 (2024).
Akhtar et al. Bioengineering the metabolic network of CAR T cells with GLP-1 and Urolithin A increases persistence and long-term anti-tumor activity. Cell Rep. Med. 6. https://doi.org/10.1016/J.XCRM.2025.102021 (2025).
He, Y. et al. Targeting PI3K/Akt signal transduction for cancer therapy. Signal Transduct. Target. Ther. 6. https://doi.org/10.1038/S41392-021-00828-5 (2021).
Jones, N. et al. Akt and STAT5 mediate naïve human CD4 + T-cell early metabolic response to TCR stimulation. Nat. Commun. 10. https://doi.org/10.1038/S41467-019-10023-4 (2019).
Klebanoff, C.A. et al. Inhibition of AKT signaling uncouples T cell differentiation from expansion for receptor-engineered adoptive immunotherapy. JCI Insight 2. https://doi.org/10.1172/JCI.INSIGHT.95103 (2017).
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat. Rev. Immunol. 12, 325–338 (2012).
Doan, A. E. et al. FOXO1 is a master regulator of memory programming in CAR T cells. Nature 629, 211–218 (2024).
Mehra, V. et al. (2023). AKT inhibition generates potent polyfunctional clinical grade AUTO1 CAR T-cells, enhancing function and survival. J. Immunother. Cancer. 11. https://doi.org/10.1136/JITC-2023-007002
Ramdas Iyer, A. et al. CRISPR-mediated generation of a tumor-associated antigen-deficient Raji platform to investigate antigen loss in CAR-T cell therapy. Front. Genome Ed. 7, 1649993 (2025).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell 164, 780–791 (2016).
Mazinani, M. & Rahbarizadeh, F. CAR-T cell potency: from structural elements to vector backbone components. Biomark. Res. 10, 1–24 (2022).
Qiu, S. et al. CAR-Toner: an AI-driven approach for CAR tonic signaling prediction and optimization. Cell Res. 34, 386–388 (2024).
Barden, M. et al. Integrating binding affinity and tonic signaling enables a rational CAR design for augmented T cell function. J. Immunother. Cancer 12. (2024).
Daniels, K. G. et al. Decoding CAR T cell phenotype using combinatorial signaling motif libraries and machine learning. Science 378, 1194–1200 (2022).
Simon, S. et al. Design of sensitive monospecific and bispecific synthetic chimeric T cell receptors for cancer therapy. Nat. Cancer 6, 647–665 (2025).
Krokhotin, A. et al. Computationally guided design of single-chain variable fragment improves specificity of chimeric antigen receptors. Mol. Ther. Oncol. 15, 30–37 (2019).
Békés, M., Langley, D. R. & Crews, C. M. PROTAC-targeted protein degraders: the past is prologue. Nat. Rev. Drug Discov. 21, 181–200 (2022).
Hsia, O. et al. Targeted protein degradation via intramolecular bivalent glues. Nature 627, 204–211 (2024).
Li, K. & Crews, C. M. PROTACs: past, present and future. Chem. Soc. Rev. 51, 5214–5236 (2022).
Vardhana, S. A. et al. Impaired mitochondrial oxidative phosphorylation limits the self-renewal of T cells exposed to persistent antigen. Nat. Immunol. 21, 1022–1033 (2020).
Chen, C. et al. Mitochondrial metabolic flexibility is critical for CD8 + T cell antitumor immunity. Sci. Adv. 9. https://doi.org/10.1126/SCIADV.ADF9522 (2023).
Wu, H. et al. Mitochondrial dysfunction promotes the transition of precursor to terminally exhausted T cells through HIF-1α-mediated glycolytic reprogramming. Nat. Commun. 14. https://doi.org/10.1038/S41467-023-42634-3 (2023).
Kim, M. S. et al. Degron-based bioPROTACs for controlling signaling in CAR T cells. ACS Synth. Biol. 13, 2313–2327 (2024).
Han, K. et al. Fasting-induced FOXO4 blunts human CD4 + T helper cell responsiveness. Nat. Metab. 3, 318–326 (2021).
Schäffner, I. et al. FoxO function is essential for maintenance of autophagic flux and neuronal morphogenesis in adult neurogenesis. Neuron 99, 1188–1203.e6 (2018).
Sullivan, J.A., Kim, E.H., Plisch, E.H., Peng, S.L., and Suresh, M. FOXO3 regulates CD8 T cell memory by T cell-intrinsic mechanisms. PLoS Pathog. 8. https://doi.org/10.1371/JOURNAL.PPAT.1002533 (2012).
Chan, J. D. et al. FOXO1 enhances CAR T cell stemness, metabolic fitness and efficacy. Nature 629, 201–210 (2024).
Shalabi, H. et al. Sequential loss of tumor surface antigens following chimeric antigen receptor T-cell therapies in diffuse large B-cell lymphoma. Haematologica 103, e215–e218 (2018).
Albelda, S. M. CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn. Nat. Rev. Clin. Oncol. 21, 47–66 (2024).
Hou, A. J., Chen, L. C. & Chen, Y. Y. Navigating CAR-T cells through the solid-tumour microenvironment. Nat. Rev. Drug Discov. 20, 531–550 (2021).
Dai, M.Y. et al. High-potency PD-1/PD-L1 degradation induced by Peptide-PROTAC in human cancer cells. Cell Death Dis. 13. https://doi.org/10.1038/S41419-022-05375-7 (20220.
Cotton, A. D., Nguyen, D. P., Gramespacher, J. A., Seiple, I. B. & Wells, J. A. Development of antibody-based PROTACs for the degradation of the cell-surface immune checkpoint protein PD-L1. J. Am. Chem. Soc. 143, 593–598 (2021).
Lu, L. et al. Enhancing the safety of CAR-T cell therapy: synthetic genetic switch for spatiotemporal control. Sci. Adv. 10. https://doi.org/10.1126/SCIADV.ADJ6251 (2024).
Guerrero, J. A. et al. GLUT1 overexpression in CAR-T cells induces metabolic reprogramming and enhances potency. Nat. Commun. 15, 8658 (2024).
Mansfield, E., Amlot, P., Pastan, I. & FitzGerald, D. J. Recombinant RFB4 immunotoxins exhibit potent cytotoxic activity for CD22-bearing cells and tumors. Blood 90, 2020–2026 (1997).
Case, D. A. et al. The Amber biomolecular simulation programs. J. Comput. Chem. 26, 1668–1688 (2005).
Case, D. A. et al. AmberTools. J. Chem. Inf. Model 63, 6183–6191 (2023).
Prihoda, D. et al. BioPhi: a platform for antibody design, humanization and humanness evaluation based on natural antibody repertoires and deep learning. MAbs https://doi.org/10.1101/2021.08.08.455394 (2021).
Bannister, D. et al. Epitope mapping and key amino acid identification of anti-CD22 immunotoxin CAT-8015 using hybrid β-lactamase display. Protein Eng. Des. Sel. 24, 351–360 (2011).
Honorato, R. V. et al. The HADDOCK2.4 web server for integrative modeling of biomolecular complexes. Nat. Protoc. 19. https://doi.org/10.1038/S41596-024-01011-0 (2024).
Azmi, I. et al. A saliva-based RNA extraction-free workflow integrated with Cas13a for SARS-CoV-2 detection. Front. Cell Infect. Microbiol. 11. https://doi.org/10.3389/FCIMB.2021.632646 (2021).
Faizan, M.I. et al. NSP4 and ORF9b of SARS-CoV-2 induce pro-inflammatory mitochondrial DNA release in inner membrane-derived vesicles. Cells 11. https://doi.org/10.3390/CELLS11192969 (2022).
Acknowledgements
The authors gratefully acknowledge the support of Dr. Satya Prakash Yadav (Medanta Hospital, Gurugram, Haryana, India), Dr. Satyendra Katewa (SHALBY Sanar International Hospital, Gurugram, Haryana, India), and Dr. Manas Kalra (Sir Ganga Ram Hospital, Delhi, India) for their assistance in arranging the flow cytometry data of ALL patients. We also thank Dr. Garima Nirmal for her assistance in obtaining the human samples. We are deeply thankful to Professor Avinash Bajaj and his research team at the Regional Centre for Biotechnology, Faridabad, Haryana, India, for their contributions to the initial animal studies; the staff of CSIR-Indian Institute of Chemical Biology (CSIR-IICB) for their essential support in the pre-clinical studies; and the team at Adgyl Lifesciences, Hyderabad, India, for their help with animal studies. Special thanks are extended to Dr. Anurag Agrawal (Former Director, CSIR-IGIB) for his encouragement and support during the early phases of the project at CSIR-IGIB and to the present Director of CSIR-IGIB, Dr. Souvik Maiti. The authors also appreciate the valuable contributions of Dr. Nedun Murugaiyan, Ankita Rajput, Aswathi C.B., Deeksha Joshi, Jahnvi Hora, Aparna Ramachandran, Syed Ahmad, and Dr. Suresh Madheswaran during the preparation of the manuscript. A patent application related to this work has been filed by Cellogen Therapeutics. We appreciate the support of Optical Microscopy Facility of the Advanced Technology Platform Centre (ATPC), Regional Centre for Biotechnology (RCB) for help with super-resolution imaging. MSA acknowledges ICMR JRF-2020 (31) for the funding support. We also acknowledge the use of ChatGPT 5.1 (OpenAI) for grammar and language refinement during manuscript preparation; all scientific content, analyses, and interpretations were generated and verified by the authors.
Author information
Authors and Affiliations
Contributions
T.A. conceived the idea, T.A., S.R., and G.K. supervised the study. M.S.A., V.C., A.S., A.A., N.C., R.T., D., K.H., S.S., R.A., M.S., M.P., S.A., M.N., P.T., M.I.F., I.A., S.A., A.R.I., P.G., M.N., I.M.U., M.J., and Z.A.B. performed experiments, generated data, and carried out analyses. Computational modeling, AI-guided design, and structural analyses were performed by the integrated computational team, V.C., D., N.C., M.N., and Z.A.B. Animal studies were conducted by collaborating institutional teams under the guidance of senior authors. T.A., S.R., G.K., M.H. and senior contributors interpreted the data and provided strategic direction. U.M. and A.K.S. provided the pre-clinical studies and data analysis. G.K., V.M. and H. K.P. helped with the patient data analysis and clinical sample collection. M.S.A., V.C., and T.A. drafted the manuscript, and all authors reviewed and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
T.A. and Cellogen Therapeutics have filed a patent application related to the CAR T cell engineering strategies described in this study. All other authors declare no competing financial or non-financial interests.
Peer review
Peer review information
Nature Communications thanks Haopeng Wang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ansari, M.S., Chauhan, V., Singh, A. et al. AI-guided CAR designs and targeted pathway modulation to enhance multi-antigen CAR T cell durability and overcome antigen escape. Nat Commun 17, 1549 (2026). https://doi.org/10.1038/s41467-025-68272-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68272-5