Abstract
Polycomb repressive complex 2 (PRC2) mediates developmental gene repression as two classes of holocomplexes, PRC2.1 and PRC2.2. EPOP is an accessory subunit specific to PRC2.1, which also contains PCL proteins. Unlike other accessory subunits that collectively facilitate PRC2 targeting, EPOP was implicated in an enigmatic inhibitory role, together with its interactor Elongin BC. We report an unusual molecular mechanism whereby EPOP regulates PRC2.1 by directly modulating its oligomerization state. EPOP disrupts the PRC2.1 dimer and weakens its chromatin association, likely by disabling the avidity effect conferred by the dimeric complex. Congruently, an EPOP mutant specifically defective in PRC2 binding enhances genome-wide enrichments of MTF2 in mouse epiblast-like cells. Elongin BC is largely dispensable for the EPOP-mediated inhibition of PRC2.1. EPOP defines a distinct subclass of PRC2.1, which may uniquely maintain an epigenetic program by preventing the over-repression of key gene regulators along the continuum of early differentiation.
Similar content being viewed by others
Introduction
Polycomb repressive complex 2 (PRC2) provides a crucial epigenetic mechanism for regulating embryonic development1. A major cellular function of PRC2 in development is to catalyze the methylation of histone H3 lysine 27 (H3K27), promote the formation of repressed chromatin domains marked by trimethylated H3K27 (H3K27me3), and thereby impose epigenetic thresholds for the activation of lineage-specific genes2. The gene repression activity of PRC2 is primarily mediated by at least two classes of functional PRC2 assemblies, PRC2.1 and PRC2.2, which share common core subunits and utilize distinct accessory subunits3. The core subunits include the catalytic subunits and paralogs EZH1 and EZH2, EED, SUZ12, and the paralogs RBBP4 and RBBP7. The accessory subunits unique to PRC2.1 are EPOP, the homologs PALI1 and PALI2, and three mammalian homologs of Drosophila PCL protein—PHF1, MTF2, and PHF19, whereas those specific to PRC2.2 are AEBP2 and JARID24. Genomic binding sites of PRC2.1 and PRC2.2 largely overlap in mouse embryonic stem cells (mESCs), suggestive of a functional redundancy5,6; on the other hand, PRC2.1 and PRC2.2 can differentially impact gene expression during mESC differentiation: for example, whereas PRC2.1 preferentially maintains gene repression, PRC2.2 mainly mediates the de novo repression of active genes7. In addition, components of PRC2.1 and PRC2.2 in mESCs display inverse chromatin binding patterns during cell cycle progression: EPOP enrichment is enhanced during the G1 phase over the S and G2/M phases, which contrasts with the case for JARID28. Intriguingly, PRC2.1 but not PRC2.2 reduces sweet responsiveness by changing gene expression in sensory neurons in Drosophila9.
The intricate cellular regulation of PRC2 function is primarily achieved at the level of enzymatic activity and chromatin targeting. A prominent example of the former is the allosteric stimulation of PRC2 catalysis by H3K27me3, which facilitates the spreading of the H3K27me3 repressive histone mark on chromatin through a positive feedback mechanism10,11. In addition, CK2-mediated SUZ12 phosphorylation stabilizes SAM binding at the enzyme active site, promoting the ability of PRC2 in H3K27 methylation and maintenance of the differentiated cell identity12. PRC2 is also directly inhibited by an oncohistone H3 and EZHIP, which contain an H3K27M or H3K27M-like protein sequence known to impede normal histone substrate binding and possibly H3K27me3 spreading13,14,15,16,17,18,19,20. The locus-specific PRC2 targeting and H3K27me3 deposition in mESCs are regulated by the accessory subunits of PRC25,6. Particularly, AEBP2 and JARID2 bind nucleosomes with monoubiquitinated histone H2A lysine 119 (H2AK119ub1)21,22,23, mediating PRC1-dependent chromatin recruitment of PRC2.224,25. In comparison, PHF1, MTF2, and PHF19 link PRC2.1 to chromatin differently, via their functional domains recognizing unmethylated CpG island (CGI) DNAs26,27 or trimethylated histone H3 lysine 36 (H3K36me3) active histone marks28,29,30,31. Importantly, MTF2 and PHF19 can stabilize the intrinsic dimer of the PRC2 core complex, and the dimeric structure of the MTF2 or PHF19-containing PRC2.1 holocomplex enhances CGI binding, likely due to an avidity effect, which refers to the combined affinities of multivalent binding32. Reminiscently, DNA binding by sequence-specific transcription factors can also be profoundly influenced by protein dimerization during active transcription33.
EPOP was previously known as the PRC2 binding protein esPRC2p4834, E130012A19Rik35, or C17orf9636,37. In addition to PRC2, EPOP was shown to interact with the Elongin BC heterodimer and recruit Elongin BC to PRC2 targets38,39,40. This coincided with the reduced chromatin recruitment of PRC2 and maintenance of the low expression of some gene loci in mESCs38,39,40. Conversely, EPOP enhanced histone methylation by PRC2 in an in vitro assay34. In addition, EPOP appears to have PRC2-independent functions in epigenetic and transcriptional regulation at active gene loci38,39. Like MTF2 and JARID2, EPOP is among the most abundant accessory subunits of PRC2 in mESCs34,41, suggesting a dominant role of EPOP in gene regulation by PRC2. EPOP is exceedingly downregulated relative to other PRC2 subunits during neuronal differentiation35,41, and it becomes highly expressed in the adult brain35,38,42 (proteinatlas.org). Although important functional and structural insights on MTF2 or JARID2-containing PRC2 have started to emerge over the years23,32,43, how EPOP impacts PRC2 function remains largely unclear. Here, based on a combination of biochemical, structural, and genomics data, we show that EPOP directly restricts PRC2.1 targeting to chromatin by disrupting the dimeric architecture of the enzyme complex, shaping a unique transcriptional program in mouse epiblast-like cells (EpiLCs), which mimic pregastrulating epiblasts and represent a primed state in stem cell differentiation44. Our results suggest EPOP may provide an efficient mechanism for regulating an epigenetic complex with a diverse array of components, preventing key developmental regulators from being over-repressed during stem cell specification.
Results
EPOP in mESCs forms a major PRC2.1 complex displaying a distinct chromatographic behavior
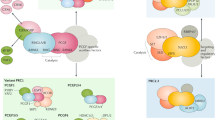
During mESC differentiation, developmental genes are collectively regulated by distinct molecular assemblies of PRC2 (Fig. 1a). To assess the relative abundance of different forms of PRC2.1 complexes, we prepared anti-SUZ12, anti-MTF2, and anti-EPOP antibody affinity beads, which efficiently captured their respective targets in nuclear extracts of E14 mESCs grown in 2i media (Supplementary Fig. 1). Co-immunoprecipitation (co-IP) results showed that substantial fractions of MTF2 and EPOP associate with SUZ12, while a notable portion of MTF2 complexes are also bound by EPOP (Fig. 1b and Supplementary Fig. 2).
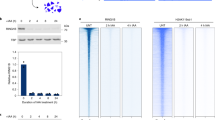
a Components of the PRC2 core complex (PRC2core) and the PRC2.1 and PRC2.2 holocomplexes. b Co-IP assays with mESC nuclear extracts and anti-SUZ12, anti-EPOP, or anti-MTF2 beads. IP, immunoprecipitated; FT, flow through. 20% of input were loaded for the anti-SUZ12 and anti-EPOP blots, and 40% for the anti-MTF2 blot. Representative of two independent replicates. c SEC elution profiles of the nuclear extracts prepared from E14 WT or EPOP KO mESCs grown in 2i media. On the left, representative PRC2 core subunit (SUZ12), PRC2.1 accessory subunits (MTF2), and PRC2.2 accessory subunit (JARID2) were detected by Western blot. On the right, Western blot bands were quantified and shown as the SEC elution profile. Representative of two independent replicates.
We knocked out endogenous EPOP in E14 mESCs, which did not alter the expression of other PRC2 components (Supplementary Fig. 3). The stoichiometry of major accessory subunits relative to SUZ12 in immunoprecipitated PRC2 was not affected in the absence of EPOP (Supplementary Fig. 4). Nuclear extracts of WT and EPOP-knockout (KO) mESCs were subjected to size-exclusion chromatography (SEC) on a Superose 6 column. MTF2 from WT mESCs was eluted as a broad peak, likely corresponding to a mixture of the monomeric and dimeric forms of MTF2-containing PRC2.1 (Fig. 1c and Supplementary Fig. 5)32. This peak partially shifted towards earlier fractions in the absence of EPOP, suggesting accumulation of the dimeric form of PRC2.1 (Fig. 1c and Supplementary Fig. 5). The core subunits EZH2 and SUZ12 displayed a similar shift of the elution peak (Fig. 1c and Supplementary Fig. 5). In stark contrast, the PRC2.2-specific accessory subunits AEBP2 and JARID2 remained unaffected by EPOP KO (Fig. 1c and Supplementary Fig. 5), in line with the mutual exclusivity of PRC2.1 and PRC2.2. Together, EPOP appeared to modulate the molecular architecture of MTF2-containing PRC2.1 in mESCs.
EPOP structurally destabilizes the PRC2.1 dimer containing MTF2 or PHF19
The role of EPOP in PRC2 remains vaguely defined, partly due to the complete absence of structural information. EPOP has been refractory to structural studies for years, given the proline-rich protein sequence and overall lack of protein structure (Supplementary Fig. 6)45. We captured the functional C-terminal domain of EPOP [EPOP(C)] bound to a PRC2.1 subcomplex in a 2.7 Å crystal structure in the form of a heterotetrameric complex, which contains EPOP(C), a modified N-terminal region of SUZ12 [SUZ12(N)], RBBP4, and the reverse chromodomain of PHF19 [PHF19(RC)]32,38 (Fig. 2a and Supplementary Table 1). Importantly, the anomalous signal from the selenomethionine derivative of the L325M mutant EPOP was essential for validating the protein sequence assignment of the structural model (Fig. 2b). The structured portion of EPOP displays sequence homology to the relatively understudied EPOP paralog SKIDA1 and, to a lesser extent, Drosophila Corto, which was also shown to bridge Elongin BC to Polycomb-group (PcG) proteins38,40,46,47 (Fig. 2c). Overall, the EPOP binding site on SUZ12 and RBBP4 is largely shared by other accessory subunits, suggesting competition in assembling distinct holocomplexes (Fig. 2d, e, also see below). In particular, we previously showed that the PRC2 core complex (PRC2core) forms a weak intrinsic dimer via domain swapping and that the dimerization interface is critically stabilized by MTF2 or PHF19 to promote chromatin targeting of PRC2.1, likely via an avidity effect32. However, this dimer stabilization mechanism is disabled in the presence of EPOP (Fig. 2d, e, also see below). Key structural findings are analyzed in detail below.
a Domain structures of the proteins captured in the crystal structure. Domain names are summarized below. SUZ12(N), N-terminal domain of SUZ12, contains ZnB, zinc finger-binding helix; WDB1, WD40-binding domain 1; C2, C2 domain; Zn, zinc finger domain; WDB2, WD40-binding domain 2. RBBP4 contains NT, N-terminal domain; WD40, WD40 domain. PHF19(RC), reversed chromodomain of PHF19. EPOP(C), C-terminal domain of EPOP. Subdomains or secondary structures of the PHF19(RC) and EPOP(C) are shown in brackets. (DS), dimer stabilization helix; (SC), short connecting helix; (C2B), C2-binding domain; (CT), C-terminal tail; (SH), short helix; (L1), loop 1; (βH), β hairpin; (L2), loop 2. The domain structures are color-coded based on proteins, except the DS helix of PHF19(RC), which is disordered in the structure and colored in gray. b 2FoFc electron density map of the EPOP fragment, contoured at 1.0σ, is shown in gray. The anomalous signal of the L325M mutant contoured at 10.0σ is shown in gold. c Sequence alignment of human EPOP, human SKDA1, and Drosophila Corto. Residues deleted in the EPOPD5 mutant are indicated by a dotted box. EPOP residues interacting with SUZ12 are indicated by blue discs, and proline residues contributing to the shape complementarity by blue squares. The secondary structure of the EPOP(C) is shown above the sequence alignment. d Cartoon representation of the overall structure. The SUZ12(N)–RBBP4–PHF19(RC)–EPOP(C) heterotetrameric complex adopts a dimeric structural architecture in the crystal lattice. Protein domains are labeled and color-coded. The two protomers are distinguished by the prime sign. e Overall structure in a different view with the rotation matrix relative to (d) indicated. f Close-up view of the EPOP binding interface. SUZ12 and RBBP4 domains are highlighted as transparent surfaces. Interacting residues are shown as sticks. Hydrogen bonds are indicated by black dotted lines. All proline residues from the EPOP fragment are also shown as sticks. g Structural alignment to PRC2.2. The current structure is aligned to a PRC2.2 subcomplex containing AEBP2 and JARID2 fragments (PDBs 5WAI and 6WKR). SUZ12 and RBBP4 are displayed as surfaces. Protein domains are labeled and color-coded. AEBP2 domains: C2B, C2-binding domain; CC, central connecting helix. JARID2 domain: TR, transrepression domain. Only the EPOP(C) domain from the current structure is shown for clarity. h Structural alignment to PRC2.1. The current structure is aligned to a PRC2.1 subcomplex containing the PHF19(RC) domain (PDB 6NQ3). SUZ12 and RBBP4 are displayed as surfaces. Protein domains are labeled and color-coded. Only one protomer of the dimer and the EPOP(C) domain from the current structure are shown for clarity. i Schematic model of the dimer disruption mechanism. The domain-swapped dimer is shown. The transient PRC2core dimer is locked by PHF19 in PRC2.1PHF19. EPOP unlocks the PRC2.1PHF19 dimer by partially displacing PHF19. The schematic is color-coded based on the crystal structures, with the gray disc representing PRC2 subunits and domains not involved in EPOP-mediated regulation.
The resolved EPOP(C) fragment consists of four structural elements, including a short helix (SH), loop 1 (L1), a β hairpin (βH), and loop 2 (L2) (Fig. 2c, f). EPOP(C)SH and especially residues L315 and F319 are docked on the concave surface jointly formed by the zinc finger-binding helix and zinc finger domain of SUZ12 [SUZ12(ZnB) and SUZ12(Zn)]43 (Fig. 2c, f). Proline-rich EPOP(C)L1 extends across the SUZ12(ZnB) helix, with residue C321 engaging in hydrogen bonding interactions with residues Q95 and R98 of SUZ12 (Fig. 2c, f). EPOP(C)βH sits on the two orthogonally stacked helices, the SUZ12(ZnB) helix and the N-terminal helix of RBBP4 (RBBP4(NT)), and is secured by hydrophobic and charge interactions (Fig. 2c, f). EPOP(C)L2 reaches back to the SUZ12(ZnB) helix, making additional contacts (Fig. 2c, f). Notably, several proline residues of EPOP, including P320, P322, and P323 from L1, and P335 from βH, jointly contribute to the critical shape complementarity of the binding interface (Fig. 2c, f).
Structural alignments with the existing structures of PRC2.1 and PRC2.2 provided additional insights. One concerns the assembly of PRC2.2. EPOP imposes steric clashes with both JARID2 and AEBP2 on the surface of the PRC2 core complex (Fig. 2g). Specifically, EPOP(C)SH and EPOP(C)L2 overlap with the transrepression domain of JARID2 [JARID2(TR)], whereas EPOP(C)L1 and EPOP(C)βH overlap with the central connecting helix of AEBP2 [AEBP2(CC)]32,43 (Fig. 2g). However, EPOP does not appear to influence the functional balance between PRC2.1 and PRC2.2 in cells, at least in mESCs, as PRC2.2 abundance did not increase upon EPOP KO (Supplementary Fig. 4). In comparison, MTF2 or PHF19 only competes with AEBP2 but not JARID2 for PRC2core binding32; correspondingly, loss of AEBP2 induces the formation of a hybrid PRC2 holocomplex containing both MTF2 and JARID2 in mESCs3.
We formerly showed that subdomains of the PHF19(RC) domain function as “molecular glue” to lock together the two protomers of the PRC2core dimer at the domain-swapped dimer interface: whereas the C2-binding (C2B) subdomain of the PHF19(RC) domain is associated with the swapped C2 domain of SUZ12 (SUZ12(C2)) from one protomer, the dimer stabilization (DS) helix from the same PHF19(RC) molecule binds the protein body of the other protomer (Fig. 2h, i)32. MTF2 likely uses the same molecular mechanism to stabilize the PRC2core dimer based on the sequence homology32. Remarkably, although EPOP can coexist with PHF19 in PRC2.1, the EPOP(C) fragment is incompatible with the DS helix of the PHF19(RC) domain, which becomes disordered in the current structure (Fig. 2h, i). The dimeric structural architecture of PRC2.1 is thought to facilitate chromatin targeting via an avidity effect32. In this regard, EPOP is projected to impair PRC2.1PHF19 or PRC2.1MTF2 chromatin targeting by displacing the DS helix, unlocking the stabilized dimer, and thereby promoting the structural transition from the dimeric state into the monomeric state (Fig. 2i).
The PRC2.1 dimer is progressively disrupted by different regions of EPOP
Inspired by the biochemical and structural observations above, we first sought to test whether EPOP can directly disrupt the PRC2.1MTF2 dimer in solution. We performed structure-based alanine-scanning mutagenesis, which identified EPOP residues 317–321 as collectively critical for PRC2 binding (Supplementary Fig. 7). We used WT and mutant EPOP with these five amino acids deleted (EPOPWT and EPOPD5) to test dimer disruption. All components of PRC2.1MTF2—EZH2, EED, SUZ12, RBBP4, and MTF2—were transiently overexpressed in HEK293T cells with or without EPOP. FLAG- and Myc-tagged EZH2 proteins were co-expressed in this context such that anti-Myc signals captured by anti-FLAG co-IP indicated the extent of PRC2.1MTF2 dimer formation (Fig. 3a). The dimeric PRC2.1MTF2 complex containing both FLAG-EZH2 and Myc-EZH2 was readily detected; in comparison, the presence of EPOPWT but not EPOPD5 resulted in greatly reduced dimer formation (Fig. 3b and Supplementary Fig. 8), which was at least in part caused by the loss of dimer stabilization by MTF2 as predicted by the structural analysis.
a Schematic of the co-IP-based dimer disruption assay. FLAG-EZH2 and Myc-EZH2 were co-expressed together with other indicated PRC2 subunits. b Dimer disruption co-IP assay. Subunits of PRC2.1MTF2 were transiently expressed with or without EPOP in HEK293T cells. Representative of two independent replicates. c Mass photometry measurement. Full-length EPOPWT or EPOPD5 was added to the purified PRC2.1MTF2. The mass of major species in solution is indicated. d SEC elution profiles. Full-length EPOP, EPOP(C), and Elongin BC were added to the purified PRC2MTF2 as indicated. Protein standards are shown above the elution profiles.
Data from a reconstituted system with purified components agreed with the co-IP dimerization assay and provided additional insights. Mass photometry measurement showed that PRC2.1MTF2 existed as an equilibrium of the monomeric and dimeric complexes and that the latter was lost upon the addition of full-length EPOPWT (Fig. 3c). In contrast, EPOPD5 did not change the equilibrium (Fig. 3c). We next used an orthogonal approach to assess the effect of EPOP on PRC2.1MTF2 dimerization. PRC2.1MTF2 was eluted as a dimer on a Superose 6 SEC column (Fig. 3d). The EPOP(C) fragment captured in the crystal structure only incompletely shifted the elution peak of the dimeric complex to a lower molecular weight (Fig. 3d), corresponding to a partial dimer disruption. Full-length EPOPWT promoted dimer disruption to a much greater extent (Fig. 3d), indicating EPOP regions beyond the C-terminal fragment also play a role in the dimer-to-monomer transition of PRC2.1MTF2. No additional change was noticed when both EPOP and Elongin BC were supplemented into the system (Fig. 3d), suggesting Elongin BC, known to bind the N-terminus of EPOP, is not involved in the dimer disruption.
Dimer disruption by EPOP weakens the chromatin binding of PRC2.1
How exactly the dimeric PRC2.1MTF2 complex binds chromatin is unclear. The extended homologous domain of MTF2 [MTF2(EH)] in PRC2.1MTF2 can mediate direct binding to the linker DNA in pairs in the context of the dimeric PRC2.1MTF2 holocomplex26,32. A biotinylated 100 bp CGI DNA from the mouse Lhx6 gene (CGILhx6) was designed to mimic the linker DNA at PRC2 binding sites and served as the bait in a pull-down assay with WT and EPOP KO mESC nuclear extracts32. Bound PRC2.1 was released by restriction enzyme digestion. Noticeably more MTF2-containing PRC2.1 were captured when EPOP was absent (Fig. 4a and Supplementary Fig. 9), indicating EPOP restricts linker DNA binding by PRC2.1MTF2.
a Nuclear extract pull-down assay. Biotinylated CGILhx6 DNA was used as the bait. The bound PRC2.1 was released by enzyme cleavage of a restriction site inserted between biotin and the CGILhx6 sequence. Representative of two independent replicates. b DNA binding EMSA. The serial dilution of PRC2.1MTF2 and PRC2.1MTF2–EPOP is labeled. The dotted red line in the lower panel indicates the position of the DNA– PRC2.1MTF2 supercomplex to compare with that of the DNA–PRC2.1MTF2–EPOP supercomplex. Representative of three independent replicates. c Apparent Kd for DNA binding by the holocomplexes. Graphs display mean ± SEM and are based on three independent replicates. d Nucleosome binding EMSA. The same as (b) except that a mononucleosome was used as the probe. Representative of three independent replicates. e Apparent Kd for nucleosome binding by the holocomplexes. Graphs display mean ± SEM and are based on three independent replicates.
To examine DNA binding quantitatively, we performed an electrophoretic mobility shift assay (EMSA) with purified PRC2.1MTF2 and PRC2.1MTF2–EPOP (Supplementary Fig. 10). A fluorescently labeled CGILhx6 DNA at 0.2 nM was used as the probe, and nonspecific competitor yeast tRNA at 8 µM was added to suppress nonspecific binding. PRC2.1MTF2 displayed an apparent Kd for DNA binding of 3.4 nM, and the apparent Kd was reduced by 2 to 3 folds to 9.2 nM for PRC2.1MTF2-EPOP (Fig. 4b, c, and Supplementary Fig. 11). Importantly, in line with the distinct oligomerization states of the enzyme complexes, DNA-bound PRC2.1MTF2 and PRC2.1MTF2-EPOP migrated differently on the gel, with the latter behaving as a smaller supercomplex (Fig. 4b and Supplementary Fig. 11).
Nucleosome binding was assessed by an EMSA using a ‘601’ mononucleosome with a 30 bp DNA extension on both ends. Shifted bands were clearly visible with 0.5 nM of the nucleosome probe. The measured apparent Kd values for PRC2.1MTF2 and PRC2.1MTF2–EPOP were 122.9 nM and 295.4 nM, respectively (Fig. 4d, e, and Supplementary Fig. 12), indicating nucleosome binding by PRC2.1MTF2 is weakened by the presence of EPOP. The eight-member PRC2.1MTF2–EPOP–Elongin BC complex displayed a binding affinity comparable to that of PRC2.1MTF2–EPOP (Supplementary Figs. 10 and 13), suggesting Elongin BC is unlikely involved in the EPOP-mediated inhibition of chromatin binding by PRC2.1MTF2. In addition, like the case for the CGILhx6 probe, PRC2.1MTF2 ran as a considerably larger supercomplex when bound to the nucleosome compared to PRC2.1MTF2–EPOP (Fig. 4d and Supplementary Fig. 12), highlighting that EPOP restricts the nucleosome binding of PRC2.1MTF2, likely by disrupting its dimeric structure. Consistently, compared to EPOPWT, the PRC2-binding-defective mutant EPOPD5 did not alter nucleosome binding by PRC2.1MTF2 in a biotinylated mononucleosome pull-down assay (Supplementary Fig. 14). Together, these results prompted us to investigate whether PRC2.1MTF2 and PRC2.1MTF2–EPOP display distinct chromatin binding in cells.
EPOP confers unique genome-wide profiles of PRC2.1 in EpiLCs
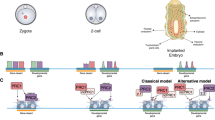
The enrichment of SUZ12, JARID2, and H3K27me3 on chromatin was previously shown to be increased in the presence of EPOP KO or knockdown (KD) in mESCs grown in serum-containing media38,39,40. It remains unclear whether EPOP exerted its influence directly or indirectly. The structure of the EPOP–PRC2 complex provided a guide for a specific disruption of the complex without disturbing the PRC2-independent function of EPOP. We established isogenic mESC lines by re-expressing comparable levels of exogenous WT and mutant EPOP proteins, EPOPWT and EPOPD5, in the EPOP KO mESCs in 2i media (Fig. 5a). As expected, the re-expressed EPOPD5 mutant displayed a severe defect in binding to endogenous PRC2 (Fig. 5b and Supplementary Fig. 15). To understand the direct role of EPOP in regulating PRC2.1 function during early development, the EPOPWT and EPOPD5 mESCs were differentiated into EpiLCs, which mimics the cell fate transition of naïve ESCs to pregastrulating epiblasts (Fig. 5c)44. EpiLCs also represent a transient time window for induction of the primordial germ cell-like cells (PGCLCs) in vitro44. Unlike in further differentiated states, EPOP is still highly expressed in EpiLCs, which makes these cells a suitable model system for studying the direct PRC2-dependent role of EPOP in gene regulation.
a Re-expression of WT and mutant EPOP in mESCs. Representative of two biological replicates. b Co-IP assay of re-expressed EPOP. The re-expressed 3 × FLAG-EPOP-HA protein was used as the bait, and the bound endogenous SUZ12 was detected. Representative of two replicates. c Schematic of the EpiLC differentiation. Some parts of the figure were created in BioRender. Liu, X. (2026) (https://biorender.com/gvraoqv). d Heatmaps of ChIP-seq replicates. Using the FDR < 0.05 threshold, the gain-of-signal MTF2 (purple) and H3K27me3 (green) peaks from individual replicates (WT-1, WT-2, D5-1, and D5-2) are shown in the heatmaps. The centers of the consensus binding sites are aligned. The number of the consensus binding sites is labeled. e Metaplots of ChIP-seq replicates. The mean ChIP-seq signal of the differential MTF2 (upper panel) and H3K27me3 (lower panel) peaks shown in (d) is plotted. The individual replicates are color-coded. f Genome browser tracks of selected gene loci. Tracks were generated by SparK (https://github.com/harbourlab/SparK). Genomic coordinates are provided. The difference between the replicates, calculated as the standard deviation, is indicated as the shaded areas surrounding the tracks. g Scatter plots of MTF2 and H3K27me3. MTF2 (left panel) and H3K27me3 (right panel) reads were normalized by the total reads. The mean read concentration corresponding to log2 (normalized ChIP-seq reads with input reads subtracted) was calculated by DiffBind.
EPOPWT and EPOPD5 mESCs cultured in 2i media were stimulated for 48 h to generate EpiLCs according to the established protocol44. Downregulation of the naïve pluripotency genes, such as Prdm14, Zfp42, and Esrrb, and upregulation of EpiLC marker genes, such as Fgf5, Dnmt3b, and Wnt3, were observed as expected (Supplementary Fig. 16 and Supplementary Data 1). No global effect on H3K27me3 or PRC2 subunit binding on chromatin was observed between the EPOPWT and EPOPD5 EpiLCs (Supplementary Fig. 17). To study the specific impact of EPOP on PRC2.1, we performed chromatin immunoprecipitation followed by sequencing (ChIP-seq) to map the genome-wide distribution of MTF2 and H3K27me3. MTF2 is a dedicated accessory subunit of PRC2.1, and H3K27me3 directly correlates with the PRC2.1 function in gene repression. The consistency of ChIP-seq replicates was confirmed by the correlation plot (Supplementary Fig. 18). Differential binding analysis of ChIP-seq peaks indicated that 2317 out of 3575 consensus binding sites showed statistically different MTF2 signals at FDR < 0.05 and that MTF2 was more enriched in the EPOPD5 EpiLCs than the EPOPWT EpiLCs at all the 2317 sites (Fig. 5d–f and Supplementary Data 2), which is in line with the general inhibitory role of EPOP in the chromatin binding of PRC2.1MTF2 as indicated by the in vitro data above.
Similarly, H3K27me3 gained occupancy at 5400 out of 20742 consensus regions in the EPOPD5 EpiLCs compared to the EPOPWT EpiLCs (Fig. 5d–f and Supplementary Data 2), indicating a negative impact of EPOP in H3K27me3 deposition. In contrast, 594 regions lost H3K27me3 occupancy for unknown reasons (Supplementary Data 2). The gain of the MTF2 and H3K27me3 signals on the chromatin of the EPOPD5 EpiLCs was further assessed by the read count scatter plots (Fig. 5g). In addition, ChIP coupled with quantitative PCR (ChIP-qPCR) was used to confirm the gain of MTF2 and H3K27me3 enrichment in two independent pairs of the EPOPWT and EPOPD5 clones (Supplementary Fig. 19). Interestingly, genome-wide JARID2 ChIP-seq signals were elevated in EPOPD5 relative to EPOPWT EpiLCs (Supplementary Figs. 19 and 20), a phenomenon also observed in mESCs lacking EPOP40. This increase may mirror the seemingly interdependent chromatin binding of MTF2 and JARID2 previously seen in mESCs27, possibly contributing to the higher H3K27me3 signal as well. Co-IP analyses indicated that the hybrid PRC2 complex containing both MTF2 and JARID2 was not promoted by EPOPD5 compared to EPOPWT, despite the structural competition between EPOP and JARID2 for PRC2 association (Supplementary Fig. 21). This contrasts with the loss of AEBP2, which appeared to result in hybrid complex formation3.
EPOP restricts PRC2.1 independently of Elongin BC
EPOP was shown to bridge the Elongin BC heterodimer to PRC2.139,40. Elongin BC likely lost the association with PRC2.1 in the EPOPD5 EpiLCs. It remained ambiguous if Elongin BC contributed to the observed PRC2.1 inhibition by EPOP. To answer this question, Elongin B was stably knocked down in the EPOPWT and EPOPD5 EpiLCs (Supplementary Fig. 22), and the genome-wide enrichment of MTF2 was compared. In the presence of the Elongin B KD, MTF2 was differentially bound to chromatin at 3259 out of 4157 consensus binding sites, and it became more enriched at 3251—almost all the differential binding sites—in the EPOPD5 EpiLCs compared to the EPOPWT EpiLCs (Fig. 6a, b). Notably, these sites overlapped with 85.8% of the 2813 similarly upregulated MTF2 sites in the case of the control KD (Fig. 6a–c), indicating Elongin BC is largely dispensable for the EPOP-mediated PRC2.1 inhibition. Unexpectedly, the Elongin B KD hampered MTF2 binding at a fraction of gene loci in the EPOPWT EpiLCs, which suggests Elongin BC may promote PRC2.1 targeting in EpiLCs (Supplementary Fig. 23). This phenomenon was largely lacking in the EPOPD5 EpiLCs (Supplementary Fig. 23), highlighting the mediating role of EPOP in the PRC2.1–Elongin BC crosstalk. It is unclear how Elongin BC may facilitate the chromatin recruitment of PRC2.1MTF2–EPOP in EpiLCs; one intriguing possibility is that Elongin BC may stabilize the monomeric PRC2.1MTF2–EPOP holocomplex and thereby protect it from proteasomal degradation, as shown for some other binding partners of Elongin BC48,49.
a Heatmaps of ChIP-seq replicates. The differential MTF2 peaks between the EPOPWT and EPOPD5 EpiLCs were grouped into three categories based on the response to shRNA KD, using the FDR < 0.05 threshold. Differential peaks shared by the control and Elongin B KD are labeled as n1, differential peaks unique to the control KD are labeled as n2, and differential peaks unique to the Elongin B KD are labeled as n3. b Genome browser tracks of selected gene loci. Tracks were generated by SparK. Two gene loci associated with the shared differential MTF2 peaks between the control KD and the Elongin B KD are shown. c Venn diagram of the differential MTF2 peaks in the three categories.
EPOP prevents over-repression of key developmental genes in EpiLCs
RNA-seq results revealed that the expression of a set of genes was markedly changed in the EPOPD5 EpiLCs (Fig. 7a and Supplementary Data 1). To directly correlate the change in gene expression with PRC2.1 targeting regulated by EPOP, the MTF2 enrichment around the transcription start site (TSS) of differentially expressed genes was compared between the EPOPWT and EPOPD5 EpiLCs. 130 transcriptionally downregulated genes in the EPOPD5 EpiLCs were found to be associated with increased MTF2 signals around their TSSs, which were also accompanied by apparently higher H3K27me3 enrichments (Fig. 7b, Supplementary Fig. 24, and Supplementary Data 3). The gene ontology (GO) analysis of these PRC2.1-repressed, EPOP-maintained genes using the DAVID (Database for Annotation, Visualization, and Integrated Discovery) server indicated that sequence-specific DNA binding proteins were highly overrepresented, on the background of all the MTF2-associated genes observed (Fig. 7c)50. These DNA-binding proteins include Rel homology transcription factors, basic leucine zipper transcription factors, winged helix/forkhead transcription factors, and homeodomain transcription factors, which are associated with biological processes like regulation of transcription, stem cell maintenance, and organ development (Supplementary Fig. 25).
a Volcano plot of differential gene expression. RNA-seq results of the EPOPD5 EpiLCs in triplicates were compared to those of the EPOPWT EpiLCs. The number of upregulated and downregulated genes is indicated. Data passing the FDR < 0.05, FC > 1.5, and average TPM of WT or mutant > 0.5 thresholds were analyzed. b Correlation of RNA-seq and ChIP-seq. One ChIP-seq replicate is shown here, and the other replicate is shown in the supplemental materials. The differential gene expression was aligned with the differential MTF2 enrichment around the transcription start site (TSS). The corresponding H3K27me3 signals are also displayed. Compared to the WT counterpart, 130 genes in the EPOPD5 EpiLCs were downregulated and associated with enhanced MTF2 signals around the TSS. The other 318 downregulated genes were not associated with EPOP-regulated MTF2 targeting. 187 upregulated genes are shown as well. c Gene ontology analysis. The PRC2.1-repressed, EPOP-maintained genes were subjected to gene ontology analysis on the DAVID server. The top 5 overrepresented terms in molecular function are shown. The p-value is one-tail Fisher Exact probability value used for gene-enrichment analysis by the DAVID server. d Schematic model of developmental gene repression by PRC2. On the left, the transient intrinsic dimer of the PRC2 core complex is illustrated. In the middle, distinct oligomerization states of various PRC2.1 and PRC2.2 holocomplexes are highlighted. MTF2 mediates direct chromatin binding, and it also stabilizes the intrinsic dimer, promoting chromatin targeting of the dimeric PRC2.1, likely via an avidity effect. EPOP disrupts the dimeric architecture of PRC2.1, containing MTF2, restricts PRC2.1 targeting, and thereby maintains the limited expression of PRC2.1-repressed developmental regulators. On the right, the PRC2.1-dependent role of EPOP in early development is illustrated. Black solid curve: during the ESC differentiation, a set of key gene regulators is repressed by PRC2.1, with limited expression being maintained by the EPOP-mediated inhibition of PRC2.1 targeting, which is followed by upregulation of the same set of gene regulators, leading to cell fate 1, e.g., PGCLCs. Gray dotted curve: the absence of EPOP results in the over-repression of these gene regulators by PRC2.1, which may change stem cell differentiation trajectories and result in an alternative cell fate 2. Some parts of the figure were created in BioRender. Liu, X. (2026) (https://biorender.com/l3ji249).
Although targeted for repression during the EpiLC differentiation, some of the PRC2.1-repressed, EPOP-maintained genes may still play a crucial role in the next step of cell fate specification. For example, in agreement with the prediction, five of these genes, Esrrb, Gjb3, Plagl1, Prdm1, and Prdm14, which are repressed during the ESC-to-EpiLC transition, were found among the top 100 upregulated genes during the EpiLC-to-PGCLC transition, whereas none in the downregulated gene list (Supplementary Table 4)44. Therefore, along the continuum of the ESC-to-EpiLC-to-PGCLC differentiation, EPOP appears to act as a “brake” on the repressive system to prevent abrupt over-repression of some key developmental regulators by directly disrupting the dimeric form of PRC2.1 and thereby restricting its chromatin targeting (Fig. 7d)44. This mechanism may help establish a critical time window for efficient induction of PGC fate (Fig. 7d)44.
Discussion
PRC2.1 and PRC2.2 are the two distinct classes of the PRC2 holocomplexes. Emerging evidence indicates that PRC2.1 and PRC2.2 are involved in non-overlapping processes in cell development, which is largely dictated by their respective accessory subunits. Although these accessory subunits are believed to collectively contribute to the chromatin recruitment of PRC2 and H3K27me3 deposition5,6, EPOP, an accessory subunit of PRC2.1, is an exception. EPOP was formerly noted to somehow limit PRC2 binding to chromatin39,40, and the present work provides a molecular mechanism for this puzzling observation. Our previous work showed that PRC2.1 containing only MTF2 or PHF19 forms a stabilized domain-swapped dimer, which facilitates chromatin targeting of the holocomplex likely via an avidity effect32. Here, we find EPOP can directly disrupt the dimeric form of PRC2.1 and thereby constrains its chromatin binding both in vitro and in vivo. Both transcription factors and epigenetic complexes carry out their functions by interacting with chromatin; whereas the role of transcription factor dimerization in transcription has been well recognized33, the present study exemplifies how the oligomerization state of a central epigenetic complex can be regulated to impact biological output during development (Fig. 7d).
The crystal structure of an EPOP-bound PRC2.1 subcomplex lays an important foundation for understanding the mechanistic model of the dimer disruption. Anomalous signals are essential for removing some ambiguities in the structural model of the C-terminal proline-rich region of EPOP. This EPOP region disables the mechanism of the MTF2 or PHF19-mediated dimer stabilization of PRC2.1 by displacing the dimer stabilization helix of MTF2 or PHF19 from the core complex, which allows other EPOP regions to complete the dimer disruption process. Despite the partial structural displacement of MTF2 and PHF19 by EPOP, they can still be stably and stoichiometrically associated with the PRC2 core complex. How the largely unstructured EPOP region beyond the PRC2-interacting C-terminal region contributes to the dimer disruption is unknown. In contrast, PHF1 lacks the structural element responsible for dimer stabilization32, and it remains to be determined if PHF1-containing PRC2.1 is similarly regulated by EPOP in cells.
In the reconstituted system, the direct consequence of PRC2.1MTF2 dimer disruption by EPOP is the substantially weakened binding of linker DNA and nucleosome in vitro, which is also largely recapitulated by ChIP-seq in vivo in EpiLCs. The structure-based EPOP mutant defective in PRC2 binding brings invaluable specificity to the in vivo study, particularly considering EPOP may also function in a non-PRC2 context39. The dimeric PRC2.1MTF2 and monomeric PRC2.1MTF2–EPOP represent two major pools of the PRC2.1 holocomplex in mESCs. In the EpiLCs, the loss of the EPOP–PRC2.1 interaction converts PRC2.1MTF2–EPOP into PRC2.1MTF2, resulting in the genome-wide increase of the MTF2 enrichment, likely due to the enhanced chromatin binding of the dimeric complex as compared to the monomeric complex. Accordingly, the genomics data may now be understood mechanistically on the molecular level. It is possible that some of the observed differential MTF2 binding peaks arise from indirect effects, such as those occurring during cell manipulation or ESC-to-EpiLC differentiation. Approaches with temporal resolution, like time-course induction of EPOP, may provide further insight into this regulation. In addition, while EPOP appears to have a genome-wide inhibitory effect on PRC2.1 targeting, whether it exerts locus-specific effects under certain circumstances remains to be investigated.
The combination of the ChIP-seq and RNA-seq data reveals a list of PRC2.1-repressed, EPOP-maintained genes in EpiLCs, which become over-repressed when EPOP no longer binds PRC2.1. Sequence-specific transcription factors are overrepresented on the background of all the detected MTF2-associated genes, indicating the EPOP-mediated inhibition of PRC2.1 maintains the partial expression of a group of key gene regulators. These genes start to be repressed by PRC2.1 during the ESC-to-EpiLC differentiation, but may remain needed for downstream developmental processes. Indeed, some of these genes are known to be activated during the EpiLC-to-PGCLC cell fate transition44. In this regard, we speculate that EPOP may impact early development by tempering the sharp change in the gene expression profile (Fig. 7d). Apparently, compared to the coordinated transcriptional regulation of multiple subunits of PRC2.1, the single EPOP protein provides an economical mechanism to modulate the PRC2.1 activity in cells in a specific and efficient way. Although we are focused on EpiLCs here, it is interesting to posit that EPOP may similarly regulate gene expression in the nervous system, where this protein is also found to be expressed.
Elongin BC is an interactor of EPOP39,40, and the role of Elongin BC in gene regulation by PRC2 remains vaguely defined. Our data suggest that Elongin BC is not involved in the PRC2.1 dimer disruption by EPOP and is largely dispensable for the EPOP-mediated inhibition of PRC2.1 targeting both in vitro and in EpiLCs. Interestingly, Elongin BC appears to promote MTF2 binding to chromatin at a set of gene loci in an EPOP-dependent manner. Although experimental evidence is lacking, one possibility is that Elongin BC may help stabilize the monomeric PRC2.1MTF2–EPOP in cells, which may become less stable than the dimeric PRC2.1MTF2 due to dimer disruption. Similarly, Elongin BC was previously shown to stabilize the SOCS1 suppressor of cytokine signaling protein and VHL tumor suppressor protein by forming a protein complex and protecting the respective protein from proteasomal degradation48,49.
Methods
Antibodies
Antibodies used in this study are listed below: SUZ12 antibody (CST, Cat. # 3737S), MTF2 antibody (Proteintech, Cat. # 162081AP), EPOP antibody (Active Motif, Cat. # 61753), JARID2 antibody (CST, Cat. # 13594S), AEBP2 antibody (CST, Cat. # 14129S), Elongin B antibody (Abcam, Cat. # ab168836), EZH2 antibody (CST, Cat. # 5246S), FLAG antibody (Sigma, Cat. # F1804), Myc antibody (CST, Cat. # 2276S), H3 antibody (CST, Cat. # 9715S), H3K27me3 antibody (CST, Cat. # 9733S), β-Tubulin antibody (CST, Cat. # 2128S).
mESC culture
The parental E14 mESCs (ES-E14TG2a) were obtained from ATCC (Cat. # CRL-1821). Cells were cultured on 0.2% gelatin-coated plates in 2i/LIF medium (1:1 mix of DMEM/F-12 and Neurobasal medium supplemented with 1 × Pen/Strep, 0.05% BSA, 1 mM GlutaMAX, 100 μM β-mercaptoethanol, 0.5 × N2 supplement, 1 × B27 supplement, 1 μM MEK inhibitor PD0325901, 3 μM GSK3 inhibitor CHIR99021, and home-made LIF). Medium was changed each day, and cells were passaged every 2 or 3 days. Cells were tested routinely for mycoplasma (Bulldog Bio Inc.) e-Myco PLUS Mycoplasma PCR Detection Kit (Bulldog Bio Inc).
EpiLC differentiation
EpiLC induction was performed according to a previous publication44. Briefly, 1.5 × 106 mESCs were plated on a 10 cm dish coated with human fibronectin (16.7 μg/ml) in N2B27 medium (1:1 mix of DMEM/F-12 and Neurobasal medium supplemented with 1 × Pen/Strep, 2 mM GlutaMAX, 50 μM β-mercaptoethanol, 0.5 × N2 supplement, 0.5 × B27 supplement, 20 ng/ml activin A, 12 ng/ml bFGF, and 1% KSR). The medium was replaced every day until 48 h before cell collection. The differentiation was confirmed using RT-qPCR for pluripotency and EpiLC-specific markers.
Establishment of EPOP KO mESC lines
EPOP KO mESC lines were generated by transfecting E14 mESCs with the pSpCas9 (BB)−2A-GFP (PX458) vector (Addgene) containing the gRNA targeting EPOP Exon 1 (CGAGCAGGGAGACCCCCGCG)5. Electroporation was performed using the Lonza transfection system. GFP-positive cells were sorted on day 3 and plated on 96-well plates after dilution of the sorted cells for single-cell clone selection. Western blot and Sanger sequencing were used to confirm the EPOP KO.
Re-expression of EPOPWT and EPOPD5
WT and D5 mutant EPOP proteins were re-expressed in EPOP KO mESCs using the pCDH-EF1a-3 × FLAG-EPOP-HA construct with a blasticidin selection marker. The Kozak sequence was inserted to improve the expression. HEK293T cells (ATCC Cat. # CRL-3216) were transfected with the EPOPWT or EPOPD5 lentiviral constructs, together with psPAX2 and Vsvg plasmids using Xtreme Gene 9 DNA transfection reagent (Roche). The culture medium containing the lentivirus was collected and concentrated using a Lenti-XTM concentrator (Takara) overnight. Concentrated lentivirus was added to EPOP KO mESCs in the presence of polybrene (8 μg/ml) overnight. Forty-eight hours post-transfection, cells were selected in the presence of 10 μg/ml blasticidin for 7 days. Single-cell clones were generated in 96-well plates. Cells were collected for Western blot analysis to pick clones expressing comparable EPOPWT and EPOPD5.
shRNA KD
The shRNA targeting the Elongin B gene TCEB2 (GATGTGATGAAGCCACAGGAT) was purchased from Sigma. For TCEB2 KD, the lentivirus was produced as described above. EPOPWT and EPOPD5 mESCs were transfected with the concentrated lentivirus in the presence of polybrene (8 μg/ml). 48 h post-transfection, puromycin (1 μg/ml) was added to the medium for selection. After 7 days of selection, cells were collected for Western blot analysis.
ChIP-qPCR and ChIP-Seq
Cells were trypsinized with TrypLE™ Express Enzyme and washed once with PBS before crosslinking with 1% formaldehyde for 10 min. Formaldehyde was quenched with 0.125 M Glycine for 5 min before two PBS washes. To check the global levels of H3K27me3, MTF2, and JARID2 on chromatin by Western blot, chromatin-bound proteins were extracted as previously described and then reverse-crosslinked for Western blot51. For ChIP, the crosslinked cells were lysed in Farnham lysis buffer (5 mM PIPES pH 8.0, 85 mM KCl, 0.5% NP40, 1 mM DTT, and 1 × protease inhibitor cocktail) to collect nuclei. Nuclei were resuspended with the lysis buffer (50 mM Tris-HCl, pH 7.9, 10 mM EDTA, 1% SDS, 1 mM DTT, and 1 × protease inhibitor cocktail) and sonicated (Diagenode Bioruptor) using 30 s ON/30 s OFF for 4 cycles. Insoluble chromatin was pelleted by centrifugation at 15,000 × g at 4 °C for 10 min. The supernatant was harvested to determine the chromatin concentration and sonication efficiency. The sheared chromatin was diluted 10-fold with ChIP dilution buffer (20 mM Tris-HCl, pH 7.9, 2 mM EDTA, 150 mM NaCl, 0.5% Triton X-100, 1 mM DTT, and 1 × protease inhibitor cocktail) for ChIP. 50 μg chromatin was used for MTF2 ChIP and 20 μg for H3K27me3 ChIP. Chromatin was incubated with 5 μg antibodies while rotating at 4 °C overnight. The next day, the antibody-chromatin complexes were incubated for 2 h at 4 °C with Protein A/G Dynabeads. After incubation, the beads were washed with 1 mL of each of the following wash buffers in this order: low salt buffer (20 mM Tris-HCl, pH 8.0, 2 mM EDTA, 1% Triton X-100, 0.1% SDS, and 150 mM NaCl), high salt buffer (20 mM Tris-HCl, pH 8.0, 2 mM EDTA, 1% Triton X-100, 0.1% SDS, and 500 mM NaCl), LiCl buffer (10 mM Tris-HCl, pH 8.0, 1 mM EDTA, 1% NP40, 1% sodium deoxycholate, and 250 mM LiCl), and TE buffer (20 mM Tris-HCl, pH8.0, and 1 mM EDTA). Samples were eluted with the elution buffer (1% SDS and 100 mM NaHCO3) followed by reverse crosslinking overnight. RNase treatment and Proteinase K digestion were performed before phenol: chloroform: isoamyl alcohol (25:24:1) and chloroform extraction.
For deep sequencing, 8 ng of eluted DNA was used to build the library with the NEBNext Ultra II kit (NEB) according to the manufacturer’s protocol, except that Ampure XP beads were used for the size selection. Libraries were sequenced on Illumina NextSeq 2000 (UT Southwestern) or Novaseq X plus (Novogene) using the paired-end strategy. For quantification by qPCR, the LightCycler 480 Instrument II system and SYBRTM Green PCR master mix were used. 3 μl of diluted ChIP DNA was used for each PCR reaction. Statistical analysis was performed using Microsoft Excel and GraphPad Prism. Primer sequences are provided below:
ESRRB_TSS_F; TTCTCCTCCAACTGGGAATG
ESRRB_TSS_R; CTAGGTCCCTGCCACTTCAG
HOXD13_F; GTGGAACAGCCAGGTGTACT
HOXD13_R; TAACCACTCCCAAATAGGGGC
OLIG2_F; TCTGTTGCAAGAAGGGGAGC
OLIG2_R; AGGTTCTGGAACGAACGCAT
Negative_F; AGAGAAGTGCTGTGCAGACG
Negative_R; CATGCACAGTTCAGCAGCTC
RNA-Seq
Total RNA was extracted using TRIzolTM reagent (Invitrogen) according to the manufacturer’s instructions. The concentration and purity of RNA were assessed using a NanoDrop device (Life Technologies). RNA integrity was checked on a TapeStation system (Agilent Technologies). The RNA-seq library was prepared using NEBNext Ultra™ II RNA Library Prep Kit for Illumina (NEB), and deep sequencing was conducted using the PE150 strategy on the NovaSeq 6000 platform.
Bioinformatics analysis
For the initial ChIP-seq data without the shRNA KD sequenced at UT Southwestern, the paired-end 40 bp read length FASTQ files were checked for quality using FastQC and FastQ Screen. Reads were aligned to the mm10 reference genome with Bowtie 252, and alignments with mapping quality scores less than 10 were filtered out. Duplicates were removed using SAMtools53. ChIP-seq peaks were called using input ChIP samples as controls54. BAM coverage BigWig files were generated by bamCoverage in deepTools55. The differential binding analysis was performed using DiffBind56, with normalization by total reads and blacklist regions removed. For the shRNA KD ChIP-seq data sequenced by Novogene, the raw sequencing reads were trimmed by Cutadapt and then mapped by Bowtie 2 with the “--very-sensitive” parameter. The aligned reads were filtered by SAMtools using the “-F 1804 -f 2 -q 30” parameter. Duplicates were removed by Picard. ChIP-seq peaks were called by MACS2 using the “-f BAMPE -q 0.05 --broad” parameter. Differential peaks were identified by DiffBind using FDR < 0.05 as the cutoff.
The read densities ± 10 kb surrounding the peak center or TSS were generated and visualized as heatmaps using deepTools. Metagene analyses were performed to show the distribution of average ChIP-seq read densities ± 10 kb surrounding the peak center or TSS using deepTools. Genome browser tracks with the standard deviation of replicates were plotted with Spark (https://github.com/harbourlab/SparK), an NGS visualization tool. The mean read concentration of each condition calculated by DESeq2 was applied to generate scatter plots using the R package ggplot2. The Venn diagram was generated using the R package VennDiagram. For RNA-seq, differentially expressed genes between the EPOPWT and EPOPD5 EpiLCs were associated with differential MTF2 binding around TSSs. The expression fold changes were used to plot the heatmap using the R package ggplot2. The R package ggplot2 was used to generate the volcano plot and the MA plot. The deepTools suite was used to generate the Pearson correlation heatmap. The read coverage statistics for genomic regions for each BAM file are calculated by multiBamSummary, and the correlation matrix using Pearson’s correlation method is plotted with plotCorrelation.
mESC nuclear extract preparation
WT and EPOP KO E14 mESCs were harvested at around 70-80% confluence from 10 cm dishes and washed twice with cold PBS. 5 volumes of hypotonic buffer A (10 mM HEPES, pH 7.9, 10 mM KCl, 1.5 mM MgCl2, 0.5% NP40, 1 mM DTT, 1 mM PMSF, and 1× protease inhibitors) were added to the pellet to lyse the cells. Nuclei were collected by centrifugation at 1000 × g at 4 °C for 10 min. 5 volumes of ice-cold high-salt buffer (20 mM HEPES, pH 7.9, 420 mM KCl, 1.5 mM MgCl2, 0.2 mM EDTA, 25% glycerol, 1 mM PMSF, and 1 × protease inhibitors) were added to lyse the nuclei. The mixture was incubated on a rotator at 4 °C for 1 h, followed by centrifugation at 15,000 × g at 4 °C for 10 min. Nuclear extracts in the supernatant were kept.
IP-MS and iBAQ
For immunoprecipitation, nuclear extracts from EPOP WT and KO mESCs were incubated with anti-SUZ12 antibody or control IgG in three biological replicates. Bound proteins were eluted under acidic conditions, reduced and alkylated, digested with trypsin, and the resulting peptides were analyzed by LC-MS/MS on an Orbitrap Eclipse mass spectrometer coupled to an Ultimate 3000 RSLC-Nano liquid chromatography system. Protein abundances were quantified using the intensity-based absolute quantification (iBAQ) method within MaxQuant 2.0.3.0, as previously described36. iBAQ values from IgG control IPs were subtracted from those of anti-SUZ12 IPs. Stoichiometries of individual PRC2 components were determined by normalizing their iBAQ values to that of SUZ12. Statistical comparisons of PRC2 component enrichment between EPOP WT and KO mESCs were carried out using an unpaired t-test.
Size exclusion chromatography
For the SEC of nuclear extracts, nuclear extracts were dialyzed against the SEC buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, and 2 mM DTT), followed by centrifugation at 15,000 × g at 4 °C for 10 min. The supernatant containing 200 μg nuclear extracts was loaded onto a Superose 6 10/300 GL column (GE Healthcare) in the SEC buffer. Fractions were collected and analyzed with a Western blot. For the SEC of purified PRC2.1 complexes, PRC2.1MTF2 was mixed with EPOP (residues 311–379), full-length EPOP-1-379, or full-length EPOP plus Elongin BC at the 1:5, 1:5, or 1:5:5 molar ratio, respectively. The mixed proteins were dialyzed against the reconstitution buffer (20 mM Tris-HCl, pH 8.0, 300 mM NaCl, 10% glycerol, and 2 mM DTT), at 4 °C overnight. Reconstituted complexes were loaded onto a Superose 6 10/300 GL column in the SEC buffer (20 mM Tris-HCl, pH 8.0, 300 mM NaCl, and 2 mM DTT), and the elution profile was recorded.
Expression and purification of PRC2.1MTF2, PRC2.1MTF2–EPOP, and PRC2.1 MTF2–EPOP–Elongin BC
PRC2.1MTF2 (EZH2–EED–SUZ12–RBBP4–MTF2) was expressed in Sf9 insect cells as described previously32. Cell pellets were harvested, resuspended in the lysis buffer (50 mM Tris-HCl, pH 8.0, 1 M NaCl, 1 mM PMSF, 1 × protease inhibitors, 0.1% NP40, 10% glycerol, and 2 mM DTT), and sonicated. The cell lysate was clarified, and proteins were captured using the IgG affinity resin. The bound proteins were released by the TEV cleavage overnight at 4 °C. The eluted proteins were dialyzed against the ion exchange low salt buffer (50 mM sodium citrate, pH 6.0, 50 mM NaCl, and 2 mM DTT) and further purified by the Mono S 5/50 GL column, with a gradient wash from the ion exchange low buffer to the ion exchange high salt buffer (50 mM sodium citrate, pH 6.0, 1 M NaCl, and 2 mM DTT). PRC2.1MTF2 was finally purified on a Superose 6 10/300 GL column in the SEC buffer (20 mM Tris-HCl, pH 8.0, 300 mM NaCl, and 2 mM DTT). PRC2.1MTF2–EPOP was expressed and purified similarly. PRC2.1MTF2–EPOP was mixed with bacterially expressed Elongin BC heterodimer to reconstitute PRC2.1MTF2–EPOP–Elongin BC.
Mass photometry
170 nM PRC2.1MTF2 was incubated with or without 850 nM full-length EPOPWT or EPOPD5 in the fresh MP buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, and 1 mM DTT) on ice for 60 min. Sample chambers were assembled with a 2 × 4 well gasket on the clean cover slide and were placed on the Two MP mass photometer (Refeyn). 9 μl MP buffer was loaded onto one sample well and focused, followed by adding 9 μl sample and mixing in the well. The AcquireMP software package was used to record contrast dots for 1 min, and the data were analyzed with the DiscoverMP software.
Co-IP of PRC2.1 complexes in nuclear extracts
The antibody-conjugated resin (anti-MTF2, anti-EPOP, or anti-SUZ12) was made using cyanogen bromide-activated-Sepharose 4B (Sigma) following the manufacturer’s instructions. Antigen binding efficiency was checked using the immunodepletion method. Nuclear extracts of WT or EPOP KO E14 mESCs were mixed with the homemade antibody resin and washed with the wash buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 0.1% NP40, 2 mM DTT, and 10% Glycerol). Captured protein complexes were released with 200 mM glycine, pH 2.8, and analyzed using Western blot.
Co-IP dimerization assay
pCS2-Myc-EZH2, pCS2-FLAG-EZH2, pCS2-His6-EED, pCS2-HA-SUZ12, pCS2-HA-RBBP4, pCS2-ProteinA-3C-MTF2, and pCS2-ProteinA-TEV-EPOP plasmids were transiently co-transfected into HEK293T cells to express PRC2.1MTF2 in the absence or presence of EPOP. Cells were harvested after 48 h of transfection. Nuclear extracts were captured by the IgG resin, washed with the wash buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 0.1% NP40, 2 mM DTT, and 10% Glycerol), and released by the TEV cleavage or the TEV plus 3 C combined cleavage overnight at 4 °C. The eluted proteins were bound to the anti-FLAG resin and washed 3 times with the same wash buffer. The bound proteins were released by the wash buffer supplemented with 1 mg/ml FLAG peptide. The dimerization efficiency was assessed based on the anti-FLAG and anti-Myc signals in the Western blot.
EMSA
For the DNALhx6 binding assay, the 100 bp CGILhx6 DNA was amplified from mouse genomic DNA through PCR with fluorescein-labeled primers. CGILhx6 DNA (0.2 nM) was incubated with purified PRC2.1MTF2 or PRC2.1MTF2–EPOP (2-fold serial dilution starting from 1 μM) in a 20 μl reaction system, also containing 20 mM Tris-HCl, pH 7.5, 100 mM KCl, 10% Glycerol, 1 mM DTT, 0.05% NP40, and 0.2 mg/ml yeast tRNA, for 30 min on ice. Samples were next loaded onto 4% native polyacrylamide gel (acrylacrylamide/bis 60:1) and resolved by electrophoresis in 1 × TGE buffer (25 mM Tris, 190 mM Glycine, and 1 mM EDTA) at 100 V for 1 h on ice. The native gel was imaged by a Typhoon scanner. For the nucleosome binding assay, a mononucleosome was assembled with the “601” DNA sequence using the standard salt dialysis method. Nucleosome (0.5 nM) was incubated with PRC2.1MTF2, PRC2.1MTF2–EPOP, or PRC2.1MTF2–EPOP–Elongin BC (2-fold serial dilution starting from 1 μM) in a 20 μl reaction system, also containing 10 mM Tris-HCl, pH 8.0, 100 mM NaCl, and 10% Glycerol, on ice for 30 minutes. The mixture was resolved on a 4% native polyacrylamide gel in 1× TGE buffer at 100 V for 1 h on ice. The native gel was stained with SYBR Gold. The binding assays were repeated 3 times for each complex. Gel band quantification was performed using Image J, and the apparent dissociation constant (Kd) was calculated by fitting binding curves in GraphPad Prism.
Biotinylated DNA pull-down assay
The biotinylated CGILhx6 DNA was prepared as previously reported with minor modifications32. Briefly, a PCR primer was labeled with biotin and contained an ECoRV cleavage site before the CGILhx6 sequence. 1 μg PCR-amplified biotinylated CGILhx6 was incubated with avidin beads for 2 hours at 4 °C and subjected to 3 × wash with the wash buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 0.1% NP40, 2 mM DTT, and 10% Glycerol) to remove the unbound DNA. Nuclear extracts from WT or EPOP KO E14 mESCs were mixed with the DNA-bound beads and incubated at 4 °C for 4 hours. Next, the beads were washed 3 times with the wash buffer and 2 times with the NEB CutSmart buffer. 1 μl ECoRV was added to the resin and incubated at 16 °C overnight to cleave the DNA and release the bound protein.
Expression and purification of the SUZ12(N)-RBBP4 binary complex
Human SUZ12 (residues 76–545 Δ146–155) and human RBBP4 (residues 1–425) were transiently co-expressed in HEK293T cells with the following plasmids: pHEK293Ultra-kozak-ProteinA-TEV-SUZ12 (residues 76–545 Δ146–155) and pHEK293Ultra-His6-thrombin-RBBP4 (residues 1–425). Transfection was performed when HEK293T cells reached 70% confluence, and transfected cells were harvested after 60 hours. For purification, the cell pellet was resuspended in lysis buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 2 mM DTT, and 5% Glycerol) and sonicated to lyse the cells. The cell lysate was clarified by centrifugation at 24,600 x g for 30 min. An affinity pull-down was performed by incubating the cell lysate with IgG resin on a nutator at 4 °C for 2 h. The IgG resin was washed with the wash buffer 1 (20 mM Tris-HCl, pH 8.0, 500 mM NaCl, 2 mM DTT, 5% Glycerol, and 0.1% NP-40), wash buffer 2 (20 mM Tris-HCl, pH 8.0, 1 M NaCl, 2 mM DTT, 5% Glycerol), and wash buffer 3 (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 2 mM DTT, 5% Glycerol). TEV protease was added for overnight cleavage. The binary complex released from the IgG resin was eluted, concentrated, and further purified by size-exclusion chromatography (SEC) using a Superdex200 10/300 GL column in the SEC buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 2 mM DTT). Protein from peak fractions was collected, concentrated, aliquoted, flash-frozen, and stored in a − 80 °C freezer.
Expression and purification of the EPOP fragment
EPOP was expressed in the Rosetta2 E. coli strain. Plasmid pET28a-His6-SUMO-TEV-EPOP (residues 311-379)-FLAG was transformed into Rosetta2 competent cells, plated on LB agar plates containing kanamycin and chloramphenicol, and incubated at 37 °C overnight. The following day, the colonies on plates were resuspended by 20 ml of LB medium, and the entire 20 ml cell suspension was inoculated into 1 liter of LB medium containing kanamycin and chloramphenicol. The cell culture was cultured at 37 °C in a 250 rpm shaker until OD 600 reached 0.6. Protein expression was induced using 0.5 mM IPTG at 20 °C for 16 h. For purification, the cell pellet was resuspended in the lysis buffer (100 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 20 mM Imidazole, and 5% Glycerol) and sonicated to lyse the cells. The cell lysate was clarified by centrifugation at 24,600 x g for 30 min. The clarified cell lysate was mixed with Ni-NTA resin and incubated on a nutator at 4 °C for 1 h. The Ni-NTA resin was washed with wash buffer 1 (50 mM HEPES, pH 7.4, 1 M NaCl, 2 mM BME, 20 mM Imidazole, 0.1% NP-40, and 5% Glycerol) and wash buffer 2 (50 mM HEPES, pH 7.4, 1 M NaCl, 2 mM BME, 20 mM Imidazole, and 5% Glycerol). The resin was collected in a filter column, and 10 bed volumes of elution buffer (20 mM HEPES, pH 7.4, 150 mM NaCl, 2 mM BME, 250 mM Imidazole, and 10% Glycerol) were applied to the Ni-NTA resin. SUMO protease was added to the eluted protein for overnight digestion. The next day, the digested protein solution was incubated with magnetic anti-FLAG resin on a nutator at 4 °C for 2 h. The anti-FLAG resin was washed using wash buffer 1 (20 mM Tris-HCl, pH 8.0, 500 mM NaCl, 2 mM BME, 5% Glycerol, and 0.1% NP-40) and wash buffer 2 (20 mM Tris, pH 8.0, 1 M NaCl, 2 mM BME, and 5% Glycerol). EPOP was eluted using the elution buffer containing the FLAG peptide (20 mM Tris-HCl, pH 8.0, 1 M NaCl, 2 mM BME, 5% Glycerol, and 0.5 mg/ml FLAG peptide). The protein was concentrated, aliquoted, flash-frozen, and stored at a − 80 °C freezer.
Expression and Purification of the PHF19 fragment
PHF19 (residues 531-580) was expressed in the Rosetta2 strain. Plasmid pGex4T1-GST-TEV-PHF19 (residues 531–580) was transformed into Rosetta2 competent cells, plated on an LB agar plate with ampicillin and chloramphenicol, and incubated at 37 °C overnight. The next morning, the plate was washed with 20 ml of LB medium, and the entire 20 ml cell suspension was inoculated into 6 liters of LB medium with ampicillin and chloramphenicol. The cell culture was grown at 250 rpm in a 37 °C shaker until the O.D.600 reached 0.8. Protein expression was induced using 0.5 mM IPTG at 20 °C for 16 h. For purification, the cell pellet was resuspended in the lysis buffer (20 mM Tris-HCl, pH 8.0, 1 M NaCl, 5 mM DTT, and 5% Glycerol) and sonicated to lyse the cells. The cell lysate was clarified by centrifugation at 24,600 x g for 30 min. The cell lysate was mixed with GST resin and incubated on a nutator at 4 °C for 1 h. The GST resin was washed with wash buffer 1 (20 mM Tris-HCl, pH 8.0, 1 M NaCl, 5 mM DTT, 0.1% NP-40, and 5% Glycerol) and wash buffer 2 (20 mM HEPES, pH 7.4, 1 M NaCl, 5 mM DTT, and 5% Glycerol). The resin was collected in a filter column, and 10 bed volumes of elution buffer (20 mM HEPES, pH 7.4, 1 M NaCl, 5 mM DTT, 25 mM Glutathione, and 5% Glycerol) were applied to the GST resin for elution. TEV protease was added to the protein elution to cleave the GST tag overnight. The resulting protein mixture was mixed with guanidine HCl powder to a final concentration of 7 M and incubated at 4 °C for 1 h. The denatured protein mixture was dialyzed against a urea buffer (20 mM sodium acetate, pH 5.2, 10 mM NaCl, 5 mM BME, and 7 M urea) using a 3.5 kD cutoff dialysis bag. After 4 hours, the dialysis bag was transferred to a new urea buffer, and the dialysis continued overnight. After dialysis, the protein mixture was loaded onto an SP column with low salt buffer (10 mM sodium acetate, pH 5.2, 10 mM NaCl, and 5 mM BME). It was washed with low salt buffer and then directly eluted using high salt buffer (10 mM Sodium acetate, pH 5.2, 1 M NaCl, and 5 mM BME) to obtain the highly concentrated PHF19 fragment. Finally, the PHF19 fragment was dialyzed against water containing 5 mM BME overnight to remove urea. The protein solution was aliquoted, flash-frozen, and kept at − 80 °C.
Reconstitution of the heterotetrameric complex and crystal structure determination
The SUZ12(N)–RBBP4 binary complex was mixed with the PHF19 fragment at a molar ratio of 1:1.2 and incubated on ice for 1 hour. The EPOP fragment was next added such that a molar ratio of SUZ12(N)–RBBP4:PHF19:EPOP became 1:1.2:2.0, and the protein complex was incubated on ice for another 1 hour. The protein complex was purified using a Superdex200 10/300 GL column in the SEC buffer (20 mM Tris-HCl, pH 8.0, 100 mM NaCl, and 2 mM DTT). Peak fractions were collected, concentrated, aliquoted, and flash-frozen at − 80 °C freezer. Crystals of the heterotetrameric complex were obtained using hanging drop vapor diffusion against a reservoir buffer (100 mM sodium citrate, pH 5.5, 200 mM sodium acetate, 9% PEG4000) at 20 °C for 1-2 weeks. The crystals were harvested and frozen in a cryo-protecting buffer (100 mM sodium citrate, pH 5.5, 200 mM sodium acetate, 12% PEG4000, 20% glycerol). Single crystal diffraction data were collected at a synchrotron light source and processed by HKL-200057. The structure was solved by molecular replacement in Phaser using PDB 5WAK as the search model58. The structural model was built in Coot and refined in autoBUSTER59,60. The structure refinement statistics were generated in Phenix61. Structural images were rendered in PyMOL62.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The crystal structure described in this work has been deposited in the Protein Data Bank (PDB) under the accession number 9XZI. Genomics data have been deposited in NCBI’s Gene Expression Omnibus under the accession numbers GSE264672 and GSE264673. LC-MS/MS data files have been deposited to the ProteomeXchange via the MassIVE partner repository with the accession code PXD072233. Source data are provided in this paper.
References
Schuettengruber, B., Bourbon, H. M., Di Croce, L. & Cavalli, G. Genome regulation by polycomb and trithorax: 70 years and counting. Cell 171, 34–57 (2017).
Comet, I., Riising, E. M., Leblanc, B. & Helin, K. Maintaining cell identity: PRC2-mediated regulation of transcription and cancer. Nat. Rev. Cancer 16, 803–810 (2016).
Grijzenhout, A. et al. Functional analysis of AEBP2, a PRC2 Polycomb protein, reveals a Trithorax phenotype in embryonic development and in ESCs. Development 143, 2716–2723 (2016).
Liu, X. & Liu, X. PRC2, Chromatin regulation, and human disease: insights from molecular structure and function. Front. Oncol. 12, 894585 (2022).
Hojfeldt, J. W. et al. Non-core subunits of the PRC2 complex are collectively required for its target-site specificity. Mol. Cell 76, 423–436 (2019).
Healy, E. et al. PRC2.1 and PRC2.2 Synergize to coordinate H3K27 trimethylation. Mol. Cell 76, 437–452 (2019).
Petracovici, A. & Bonasio, R. Distinct PRC2 subunits regulate maintenance and establishment of Polycomb repression during differentiation. Mol. Cell 81, 2625–2639 (2021).
Asenjo, H. G. et al. Polycomb regulation is coupled to cell cycle transition in pluripotent stem cells. Sci. Adv. 6, eaay4768 (2020).
Vaziri, A. et al. Persistent epigenetic reprogramming of sweet taste by diet. Sci. Adv. 6, https://doi.org/10.1126/sciadv.abc8492 (2020).
Margueron, R. et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009).
Jiao, L. & Liu, X. Structural basis of histone H3K27 trimethylation by an active polycomb repressive complex 2. Science 350, aac4383 (2015).
Gong, L. et al. CK2-mediated phosphorylation of SUZ12 promotes PRC2 function by stabilizing enzyme active site. Nat. Commun. 13, 6781 (2022).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Justin, N. et al. Structural basis of oncogenic histone H3K27M inhibition of human polycomb repressive complex 2. Nat. Commun. 7, 11316 (2016).
Ragazzini, R. et al. EZHIP constrains polycomb repressive complex 2 activity in germ cells. Nat. Commun. 10, 3858 (2019).
Jain, S. U. et al. PFA ependymoma-associated protein EZHIP inhibits PRC2 activity through a H3 K27M-like mechanism. Nat. Commun. 10, 2146 (2019).
Hubner, J. M. et al. EZHIP/CXorf67 mimics K27M mutated oncohistones and functions as an intrinsic inhibitor of PRC2 function in aggressive posterior fossa ependymoma. Neuro Oncol. 21, 878–889 (2019).
Piunti, A. et al. CATACOMB: An endogenous inducible gene that antagonizes H3K27 methylation activity of Polycomb repressive complex 2 via an H3K27M-like mechanism. Sci. Adv. 5, eaax2887 (2019).
Harutyunyan, A. S. et al. H3K27M induces defective chromatin spread of PRC2-mediated repressive H3K27me2/me3 and is essential for glioma tumorigenesis. Nat. Commun. 10, 1262 (2019).
Jain, S. U. et al. H3 K27M and EZHIP Impede H3K27-Methylation Spreading by Inhibiting Allosterically Stimulated PRC2. Mol. Cell 80, 726–735 (2020).
Kalb, R. et al. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat. Struct. Mol. Biol. 21, 569–571 (2014).
Cooper, S. et al. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nat. Commun. 7, 13661 (2016).
Kasinath, V. et al. JARID2 and AEBP2 regulate PRC2 in the presence of H2AK119ub1 and other histone modifications. Science 371, https://doi.org/10.1126/science.abc3393 (2021).
Blackledge, N. P. et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 157, 1445–1459 (2014).
Cooper, S. et al. Targeting polycomb to pericentric heterochromatin in embryonic stem cells reveals a role for H2AK119u1 in PRC2 recruitment. Cell Rep. 7, 1456–1470 (2014).
Li, H. et al. Polycomb-like proteins link the PRC2 complex to CpG islands. Nature 549, 287–291 (2017).
Perino, M. et al. MTF2 recruits Polycomb Repressive Complex 2 by helical-shape-selective DNA binding. Nat. Genet. 50, 1002–1010 (2018).
Ballare, C. et al. Phf19 links methylated Lys36 of histone H3 to regulation of Polycomb activity. Nat. Struct. Mol. Biol. 19, 1257–1265 (2012).
Musselman, C. A. et al. Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1. Nat. Struct. Mol. Biol. 19, 1266–1272 (2012).
Brien, G. L. et al. Polycomb PHF19 binds H3K36me3 and recruits PRC2 and demethylase NO66 to embryonic stem cell genes during differentiation. Nat. Struct. Mol. Biol. 19, 1273–1281 (2012).
Cai, L. et al. An H3K36 methylation-engaging Tudor motif of polycomb-like proteins mediates PRC2 complex targeting. Mol. Cell 49, 571–582 (2013).
Chen, S., Jiao, L., Liu, X., Yang, X. & Liu, X. A dimeric structural scaffold for PRC2-PCL targeting to CpG Island chromatin. Mol. Cell 77, 1265–1278 (2020).
Amoutzias, G. D., Robertson, D. L., Van de Peer, Y. & Oliver, S. G. Choose your partners: dimerization in eukaryotic transcription factors. Trends Biochem. Sci. 33, 220–229 (2008).
Zhang, Z. et al. PRC2 complexes with JARID2, MTF2, and esPRC2p48 in ES cells to modulate ES cell pluripotency and somatic cell reprogramming. Stem Cells 29, 229–240 (2011).
De Cegli, R. et al. Reverse engineering a mouse embryonic stem cell-specific transcriptional network reveals a new modulator of neuronal differentiation. Nucleic Acids Res. 41, 711–726 (2013).
Smits, A. H., Jansen, P. W., Poser, I., Hyman, A. A. & Vermeulen, M. Stoichiometry of chromatin-associated protein complexes revealed by label-free quantitative mass spectrometry-based proteomics. Nucleic Acids Res. 41, e28 (2013).
Alekseyenko, A. A., Gorchakov, A. A., Kharchenko, P. V. & Kuroda, M. I. Reciprocal interactions of human C10orf12 and C17orf96 with PRC2 revealed by BioTAP-XL cross-linking and affinity purification. Proc. Natl. Acad. Sci. USA 111, 2488–2493 (2014).
Liefke, R. & Shi, Y. The PRC2-associated factor C17orf96 is a novel CpG island regulator in mouse ES cells. Cell Discov. 1, 15008 (2015).
Liefke, R., Karwacki-Neisius, V. & Shi, Y. EPOP Interacts with Elongin BC and USP7 to modulate the chromatin landscape. Mol. Cell 64, 659–672 (2016).
Beringer, M. et al. EPOP Functionally links elongin and polycomb in pluripotent stem cells. Mol. Cell 64, 645–658 (2016).
Kloet, S. L. et al. The dynamic interactome and genomic targets of Polycomb complexes during stem-cell differentiation. Nat. Struct. Mol. Biol. 23, 682–690 (2016).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Chen, S., Jiao, L., Shubbar, M., Yang, X. & Liu, X. Unique structural platforms of Suz12 dictate distinct classes of PRC2 for chromatin binding. Mol. Cell 69, 840–852 e5 (2018).
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011).
Meszaros, B., Erdos, G. & Dosztanyi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 46, W329–W337 (2018).
Hauri, S. et al. A high-density map for navigating the human polycomb complexome. Cell Rep. 17, 583–595 (2016).
Rougeot, J., Renard, M., Randsholt, N. B., Peronnet, F. & Mouchel-Vielh, E. The elongin complex antagonizes the chromatin factor Corto for vein versus intervein cell identity in Drosophila wings. PLoS ONE 8, e77592 (2013).
Kamura, T. et al. The Elongin BC complex interacts with the conserved SOCS-box motif present in members of the SOCS, ras, WD-40 repeat, and ankyrin repeat families. Genes Dev. 12, 3872–3881 (1998).
Schoenfeld, A. R., Davidowitz, E. J. & Burk, R. D. Elongin BC complex prevents degradation of von Hippel-Lindau tumor suppressor gene products. Proc. Natl. Acad. Sci. USA 97, 8507–8512 (2000).
Dennis, G. Jr. et al. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003).
Gillotin, S. Isolation of chromatin-bound proteins from subcellular fractions for biochemical analysis. Bio. Protoc. 8, e3035 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Bricogne, G. et al. BUSTER version 2.10.4. Cambridge, United Kingdom: Global Phasing Ltd. (2017).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Schrodinger, L. The PyMOL Molecular Graphics System, Version 2.5.5 (2010).
Acknowledgements
The cDNA of EPOP was kindly provided by Dr. Robert Liefke and Dr. Yang Shi. The cDNAs of human PRC2 core subunits were kindly provided by Dr. Robert Kingston. This research was supported by NIH grants R35GM136308 and R21NS137323 to X.L. X.L. is a W.W. Caruth, Jr., Scholar in Biomedical Research. L.G. was supported by an American Heart Association postdoctoral fellowship 19POST34450043, the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB1230000), NSFC-Joint Foundation of Yunnan Province (U24A20806), Introducing Talents Start-up Fund of Kunming Institute of Botany, Chinese Academy of Sciences (51E46G0111K1), and Fund of State Key Laboratory of Phytochemistry and Natural Medicines (52E46P5211Z1). S.C. was supported by the National Natural Science Foundation of China grant 32100464. Some data presented in this report were acquired with a mass photometer that was supported by an award S10OD030312-01 from the NIH. Results shown in this report are derived from work performed at Argonne National Laboratory, Structural Biology Center (SBC) at the Advanced Photon Source. SBC-CAT is operated by UChicago Argonne, LLC, for the U.S. Department of Energy, Office of Biological and Environmental Research under contract DE-AC02-06CH11357. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (P30GM133894). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH.
Author information
Authors and Affiliations
Contributions
Xin.L. conceived the study; L.G., Xiuli.L., M.Y.B, X.Y., S.C., and Xin.L. designed and performed the experiments; X.Y. and Xin.L. analyzed the structural data; Xiuli.L., Z.Y., C.X., and Xin.L. analyzed the genomics data; A.L. analyzed the proteomics data; L.G., Xiuli.L., X.Y., Z.Y., C.X., and Xin.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tatiana Kutateladze and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gong, L., Liu, X., Yang, X. et al. EPOP restricts PRC2.1 targeting to chromatin by directly modulating enzyme complex dimerization. Nat Commun 17, 1565 (2026). https://doi.org/10.1038/s41467-025-68280-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68280-5