Abstract
Spatiotemporal coordination of cellular and molecular events is crucial for cell fate commitment during mouse gastrulation. However, the high-precision mechanisms governing the timing and spatial dynamics remain poorly understood. Here, we present a time-series single-cell multi-omic dataset of the gastrulating mouse embryos and construct a hierarchical gene regulatory landscape. Integrating this with real three-dimensional transcriptomic coordinate, we created ST-MAGIC and ST-MAGIC (+) atlas, dissecting the spatiotemporal logics of regulatory networks and signaling responsiveness underpinning the lineage commitment at gastrulation. Specifically, we delineated the multi-omic basis for left-right symmetry breaking events in the gastrula and also revealed the spatiotemporal molecular relay for axial mesendoderm lineage, where early and intermediate transcription factors first open the chromatin regions and setup the responsiveness to signaling, followed by terminal factors to consolidate the transcriptomic architecture. In summary, our study presents a spatiotemporal regulatory logic framework of mouse gastrulation for advancing our understanding of mammalian embryogenesis.
Similar content being viewed by others
Introduction
Gastrulation is a pivotal phase in embryonic development, at which the multipotent epiblast is allocated to diverse tissue lineages of the three primary germ layers: ectoderm, mesoderm, and endoderm1,2. This stage is also crucial for the spatial organization of the embryo in anterior-posterior, dorsal-ventral and left-right axes1. Gastrulation, therefore, lays the blueprint of embryogenesis, making it a focal point for understanding the molecular and cellular mechanisms that govern embryonic patterning.
The dynamic nature of gastrulation involves rapid changes in cell composition, differentiation and proliferation, all of which are intricately regulated by a combination of epigenetic modifications, transcriptional networks, and intercellular communication. Recent advancements using limited cell numbers or single-cell omics significantly enhance our understanding of these dynamic processes during gastrulation. These technologies have enabled high-resolution analysis of gene expression patterns and cellular heterogeneity within developing embryos, identify that embryogenesis during gastrulation is orchestrated by precise schedules of gene expression within individual cells, regulated by complex intrinsic gene regulatory networks (GRNs) and influenced by intercellular signaling pathways3,4,5,6. These findings highlight the importance of both temporal and spatial regulation in cell fate and tissue patterning.
Despite these advances, a comprehensive characterization of the GRNs that govern the rapid cell fate commitment and spatial patterning during gastrulation remains elusive. Traditional single modality of single-cell omics often disrupts the spatial and temporal continuity of embryonic development, limiting our ability to fully understand the molecular mechanisms underlying embryo patterning. Spatiotemporal coordination of GRNs provides the developmental principles of lineage allocation and tissue patterning in the germ layers of the gastrulating embryo.
One illustrative example is the formation of the axial mesendoderm lineage in mice, a distinct structure emerging from the dorsal organizer during the late-streak stage. The anterior mesendoderm and primitive node cells form the prospective notochordal structure, with the primitive node acting as a reservoir for continuous production of caudal notochord cells7,8,9. The hereby forming notochord structure serves as an organizing center for proper dorsal-ventral and left-right body axes10,11,12. Disruptions in the notochord structure often result in severe embryonic defects, emphasizing its critical role in body axis formation13,14,15,16,17. However, the sequential cellular events and molecular GRNs with spatial-temporal information underlying axial mesendoderm formation are not well characterized.
To address these challenges, we generated a high-resolution, spatiotemporally resolved single-cell multi-omic reference for the mouse gastrulating embryo. Our dataset includes transcriptomics and chromatin accessibility profiles, collected at 6 h intervals across multiple stages of gastrulation. Using the developed Bi-Orientation Cis-Regulatory Elements Predictor (BioCRE) algorithm, we linked genes with their potential regulatory elements, establishing a comprehensive GRNs atlas underlying the rapid cell lineage commitment process. With reference to the three-dimensional spatial transcriptome coordinates, the dataset is rendered into the SpatioTemporal-Multi-omic Atlas of Gastrulating In-silico Cells (ST-MAGIC). By integrating transcription factor (TF)-target gene-target chromatin region cascades as well as signaling effector chromatin immunoprecipitation sequencing (ChIP-seq) data, we developed the ST-MAGIC (+) platform to explore the spatiotemporal dynamics of TF networks and determine intrinsic responsiveness to crucial developmental signals.
The learning gleaned from the ST-MAGIC and ST-MAGIC (+) atlas can inform the spatiotemporal coordination of GRNs for governing lineage allocation and tissue patterning in the germ layers of the gastrulating embryo. Here, from the ST-MAGIC and ST-MAGIC (+) atlas, we have determined the multi-omic basis for left-right symmetry breaking events in the gastrula, identified and experimentally validated the complex GRNs hierarchy involving TFs (EOMES, FOXA2, NOTO/POU6F1) and signaling pathways (such as NODAL and WNT signaling) that regulate mouse axial mesendoderm development, accompanied by a spatial developmental route from the anterior primitive streak to the distal region of the embryo.
Results
Single-cell multi-omics capture the GRNs underlying germ layer formation
To spatiotemporally dissect the intrinsic GRNs at single-cell resolution during mouse gastrulation, we have undertaken: (1) profiling the transcriptome and chromatin accessibility of individual cells of mouse gastrula at early-streak stage (E6.5) to the late-streak stage (E7.5) at 6 h (0.25 embryonic day) intervals; (2) constructing gene-peak linkages, that connect transcriptomic and epigenomic modalities; (3) integrating the inferred gene-peak linkages with the Geo-seq-based spatiotemporal information to reconstruct a real time-series three-dimensional spatiotemporal multiome atlas; and (4) analyzing the hierarchical logics of GRNs in parallel with signaling responsiveness to gain insights into the mechanistic attributes of the gene regulatory networks underpinning lineage development and embryonic patterning at gastrulation (Fig. 1a).
a Schematic overview of the workflow from data generation, peak-to-gene linkage identification to construction of spatiotemporal multi-omic atlas spanning from E6.5 to E7.5 of mouse embryo development, followed by the determination of lineage-specific GRNs and experimental validation. Mouse icon was created in BioRender, Tan, F. (2025) https://BioRender.com/1atcqml. b UMAP projection of cells based on multiome-derived transcriptomic (left) and epigenomic (right) data, colored by cell type annotations. The annotations of cell types were inferred from the published dataset3,6. c Directed acyclic graph showing inferred relationships between cell types defined by transcriptomic data across early mouse embryogenesis. The processed scRNA-seq dataset covering the developmental stages from E3.5 to E6.25 was publicly available from the TOME portal (http://tome.gs.washington.edu/). Each row corresponds to one of the annotated cell types, columns to developmental stages. All edges with weights above 0.2 were shown in grayscale. Of note, parietal endoderm cells were not actively retained during the isolation of embryos in this study and were excluded from analyses in this study. d The schematic diagram showing the capture of gene-peak linkages using BioCRE, in which harnesses a bi-orientation regression model leveraging multi-omics data to decode the linkage between genes and chromatin peaks. e The Jensen-Shannon Similarity (JSS) score evaluates the performance of BioCRE, Signac, and ArchR algorithms in calculating the consistency between cell type transcriptomic features and chromatin accessibility. Wilcox tests were used to calculate the significances. n = 44,644, 43,096, and 52,936 cell type–specific gene regulatory linkages identified by BioCRE, Signac, and ArchR, respectively. Box plots display the median (center line), the first and third quartiles (box limits), and whiskers indicating the minimum and maximum values within 1.5 times the interquartile range. f BioCRE captures a group of distal peaks with the highest consistency with linked gene expression. Distal peaks were annotated as peaks located outside TSS ± 2 kb, proximal peaks were annotated as peaks located within TSS ± 2 kb. Wilcox tests were used to calculate the significances. n = 3537 genes examined for proximal and distal regulatory associations. Box plots display the median (center line), the first and third quartiles (box limits), and whiskers indicating the minimum and maximum values within 1.5 times the interquartile range. g Heatmap showing gene expression (left) and chromatin accessibility (right) for selected cell type-specific genes (top 10 genes) and their linked peaks, with each row representing a pair of one gene and the linked CRE. Each gene can be linked to multiple CREs. Hence, genes can be represented by multiple rows in the corresponding heatmap. h Genome browser snapshot displaying the normalized ATAC-seq signal (top), the distribution of annotated peaks (middle), the linkage with Mesp1 gene for a given cell type (bottom). The expression of Mesp1 across 31 cell types was also shown at the right. Three genomic peaks showing significant linkages with Mesp1 were highlighted in red, two of the peaks (d1, d3) consist the known EME enhancer reported previously20, and the d2 peak was a newly identified Mesp1-linked element and named as Neo_ME element. Sig. represents the significance of the linkage correlation between gene and chromatin peak calculated by BioCRE, the same for the following coverage plot results. For detailed significance calculation, please see the Method section.
In total, we have acquired bi-modal omics data from 35,449 single cells across five embryonic stages (12 E6.5 embryos, 10 E6.75 embryos, 6 E7.0 embryos, 6 E7.25 embryos, and 5 E7.5 embryos) with high cell coverage and high data quality after stringent quality control measurements (Supplementary Fig. 1a–e). Uniform manifold approximation and projection (UMAP) was performed for both transcriptomic and chromatin accessibility (Fig. 1b and Supplementary Fig. 1f, g). To annotate the cell types, iterative transcriptomic clustering for cells from each stage was conducted, and specific gene expression pattern for each sub-cluster was refined. Cells with similar gene expression profiles were then grouped and annotated by inferring known gene expression signatures cross-referenced with the published mouse embryo transcriptome atlases3,6,18. The 31 annotated cell types exhibit consistent composition and high transcriptomic correlation with the published single-cell transcriptome gastrulation atlas3 and delineate the diversifying cell type composition during gastrulation with some cell types selectively enriched for earlier (e.g., anterior primitive streak) or later stages (e.g., cardiac mesoderm, etc.) (Supplementary Fig. 1h, 2a–d). Notably, in contrast to one homogeneous notochord cell cluster recognized in the previous report3,6, here we identified four previously unappreciated distinct subgroups within the notochord lineage, which was named as anterior mesendoderm precursors and node precursors at the E7.25 stage, and anterior mesendoderm and node at the E7.5 stage, respectively (Fig. 1b).
The annotated cell identities were then transferred to cells embedded in the snATAC-seq UMAP based on in silico cell-to-cell matches bridged by the common cellular barcodes incorporated during the preparation of the multi-omic sequencing library (Fig. 1b). Generally, the cells collated in this dataset cover the transitions from early pluripotent epiblast cells at E6.5 to fate-committed neuroectoderm, definitive endoderm and various diversified mesodermal subtypes at the E7.5 stage. We also applied the Trajectories Of Mammalian Embryogenesis (TOME) algorithm18 to systematically analyze the relationship among cell types from two adjacent stages and finally generated a clear relationship graph for all the cell types across gastrulation (Fig. 1c). The inferred lineage trajectories are broadly consistent with previous understandings of mouse early embryogenesis3,18,19, providing a developmental phylogeny to pinpoint the molecular dynamics and hierarchy of transcriptomic and epigenetic regulation during gastrulation.
To map the GRNs governing lineage development during gastrulation, we first determined cell type-specific gene expression, chromatin accessibility and motif enrichment. Intriguingly, despite distinct cell types exhibit distinctive transcriptomic features and epigenomic differences, the motif enrichment remained intermingled for cells within the same pedigree (Supplementary Fig. 3a–c). This observation may indicate that the formation of various cell types and the transcriptomic signatures may be regulated by a common set of TFs, but with frequent turnover of epigenomic landmarks.
Next, to capture the relationship between gene transcriptomic status and peak chromatin accessibility, we developed an algorithm, BioCRE, to capture the linkage between expressed genes and candidate regulatory elements, thereby gaining insight into the relationship between gene expression and chromatin peak accessibility. Distinct from existing tools, such as Signac and ArchR, BioCRE harnesses a bi-orientation regression model leveraging multi-omics data at the chromosome level to identify potential CREs (Fig. 1d). Jensen-Shannon Similarity (JSS) score (see “Methods”), which is calculated based on Jason-Shannon Divergence index, has been developed to measure the consistency between cell type transcriptomic features and linked chromatin accessibility diversities. As shown, the JSS score was higher for BioCRE than for Signac and ArchR (Fig. 1e). Moreover, we found that BioCRE-specific gene-peak linkages are enriched in regulating embryo development-related genes (Supplementary Fig. 4a–f), and show higher cell type-specificity as indicated by Silhouette score (Supplementary Fig. 4g, h). In-depth exploration of performance stability against variable sample sizes demonstrated that BioCRE is more robust with differential sampling sizes than Signac and ArchR, as marked by shorter run time and higher consistency of linkage composition (Supplementary Fig. 4i, j). Cross-validation of predicted gene-peak linkages using publicly available gold standard promoter capture Hi-C dataset further demonstrated the performance superiority of BioCRE over Signac and ArchR (Supplementary Fig. 5a–j). This result indicates that BioCRE is more effective in predicting cell type-specific regulatory elements that modulate target gene expression.
Generally, BioCRE results show that one gene is linked to a median of 5 CREs with a median gene-to-peak distance of 127,175 bp (Supplementary Figs. 4b, 6a, b). Moreover, we observed that a considerable number of distal chromatin peaks (TSS ± 2-500 kb) exhibit higher JSS score (distal high group) than the regulated gene promoters, highlighting the distal regulatory regions may play prominent roles in regulating gene expression (Fig. 1f and Supplementary Fig. 6c). Co-variation of cell type-specific DEGs and BioCRE-linked peaks’ chromatin accessibility faithfully distinguished the identified cell types in both transcriptomic and epigenomic modalities (Fig. 1g and Supplementary Data 1). For example, ten distal peaks were linked to the expression of nascent mesoderm marker, Mesp1 (Fig. 1g). Amongst the distal linked peaks, d1 and d3 peaks are related to the previously known EME enhancer for Mesp120, while the potential regulatory logics for the remaining eight distal peaks were newly identified. To validate the regulatory force for the newly-identified Mesp1 linked distal peaks, we analyzed the chromatin accessibility attribute for one of the newly-identified distal peaks (named Neo_ME) and the EME element (Fig. 1h). Detailed exploration revealed that, apart from the co-accessible patterns in nascent mesoderm, lateral mesoderm, paraxial mesoderm 2, mesoderm progenitors and ExE mesoderm cells, the Neo-ME element was accessible in the caudal epiblast, caudal mesoderm cells as well as axial mesendoderm related cells (node and anterior mesendoderm). In contrast, the EME element was more accessible in paraxial mesoderm 1, cardiac mesoderm, and mesenchyme cells (Fig. 1h). Thus, while both Neo_ME and EME elements regulate Mesp1 expression, there may exhibit distinct usage preferences of regulatory elements among distinct cell types.
To further explore the role of the newly-identified element, we performed enhancer reporter assays, demonstrating that the specific distribution of the Neo-ME element was consistent with Mesp1 expression (Supplementary Fig. 6d). In addition, we established ESC cell lines with specific deletion of the Neo-ME element and EME element, respectively (Supplementary Fig. 6e). As expected, genetic removal of these two elements led to specific downregulation of Mesp1 expression during embryoid bodies (EBs) differentiation (Supplementary Fig. 6e). Thus, the newly-identified Neo_ME element is likely a critical regulatory element responsible for Mesp1 expression.
Together, through single-cell co-profiling of gene expression and chromatin accessibility in the mouse gastrula, combined with the gene-peak linkage capturing strategy (BioCRE), we established a comprehensive multi-omic atlas which captures the cell type-specific GRNs encompassing gene expression, chromatin accessibility of regulatory elements, as well as gene-peak linkages underlying mouse gastrulation.
Spatiotemporal multi-omic landscape of the in-silico gastrulating cells
Recent technological advances have enabled the measurement of gene expression across tissue sections using various spatial transcriptomic strategies21,22,23,24,25. However, most studies covered only the transcriptomic data module and provided limited real 3D spatial context. A comprehensive spatiotemporal multi-omic map that contains genuine spatiotemporal coordinates for embryo tissues remains elusive.
To construct a spatiotemporal multi-omic map that incorporates the spatial coordinates and time stamps of the cells of the gastrulating mouse embryo, we leveraged the published spatiotemporal registered transcriptome dataset of mouse embryonic tissues during gastrulation6,19 and the spatial transcriptome reconstruction algorithm-Tangram26. We refined a pipeline (see Methods) to reconstruct and visualize cell type distribution, single-cell based spatial transcriptome, and spatiotemporally resolved chromatin accessibility information from embryonic cells at the registered spatiotemporal resolution (Fig. 2a, Supplementary Fig. 7a and Supplementary Data 2) in a ST-MAGIC atlas (SpatioTemporal reconstructed Multi-omic Atlas of the Gastrulating In-silico Cells). Beyond spatial distribution, ST-MAGIC can be customized to investigate the spatial distribution of specific cell types, gene expression, and chromatin accessible domains (Fig. 2a). It is noteworthy that different from the previous gene activity score based chromatin accessibility spatial mapping5, the integrative usage of common cell barcodes present in both the transcriptome and chromatin accessibility modules would significantly enhance the spatial accuracy of the ST-MAGIC.
a Schematic of the ST-MAGIC, for spatiotemporal projection and reconstruction of multi-omic atlas at the stage-matched spatial coordinates defined by transcriptome6,19. In the reconstructed map (corn-plots), registered positions are labeled in the epiblast/ectoderm domain (A, anterior; L, left lateral; R, right lateral; L1/R1, left/right anterior lateral, L2/R2, left/right posterior lateral), the prospective primitive streak domain (P), the prospective mesoderm domain (M, mesoderm; MA, anterior mesoderm; MP, posterior mesoderm), and the prospective endoderm domain (E, endoderm, EA, anterior endoderm; EP, posterior endoderm) were labeled at the top of each corn-plot. Numbers representing the positions along with the proximal-distal axis in a descending series were labeled on the left side of each corn-plot. Percent represents the ratio of searched cell type, the ratio of expressed genes or the ratio of opened peaks in cells mapped to the indicated spot. The relative abundances of enriched cell type, gene expression level, or chromatin accessibility level were shown on the right side of each corn-plot. Rel. enrichment is short for relative enrichment, Chrom. Acc. is short for chromatin accessibility level. To visualize the multi-omic information, spots in the corn-plot for cell type and gene expression reconstruction were outlined as round circles, while spots in the corn-plot for chromatin accessibility reconstruction were outlined as peak shapes. For sourcing information from the so-constructed ST-MAGIC atlas, we enables data query with a specific cell-type, gene/peak input or start from specific spatiotemporal points. b The corn-plot displaying the spatial distribution of selected mesodermal cell types (mesenchyme, cardiac mesoderm, paraxial mesoderm 1, and paraxial mesoderm 2) as indicated by the ST-MAGIC atlas at the E7.5 mouse embryo. c (i) The spatial distribution of specific mesodermal cell types validated by RNAscope analysis of marker genes expression. Mesen: mesenchyme; Parax M 1: paraxial mesoderm 1; Parax M 2: paraxial mesoderm 2; Cardiac M: cardiac mesoderm. (ii) Schematic summarizing the distribution of mesoderm subtypes. n = 3 replicates for RNAscope staining. d–f ST-MAGIC identifying spatial-specific usage of regulatory elements for widely expressed genes. d Representative genome browser snapshot showing the chromatin accessibility around the Otx2 locus for E6.5 cells. The expression of Otx2 was also shown on the right. e RNAscope validating the localization of Otx2 transcripts in E6.5 embryos. n = 3 replicates for Otx2 staining. f ST-MAGIC visualization of the EPI-specific, VE-specific peak and universal peak linked to Otx2 expression. VE is short for visceral endoderm, Epi is short for epiblast. g, h Region-specific exploration of the ST-MAGIC atlas of E7.5 mouse embryo, identifying the multi-omic basis of left-right symmetry breaking. To capture differences between left and right mesoderm tissues, the left-right resolved Geo-seq transcriptomic coordinate reference was used6. Precise digital spatial sampling of the right (g) and left (h) proximal mesoderm (RPM and LPM) captures the lateral region cell type composition in the E7.5 mouse embryo. i, j Heatmaps showing the chromatin accessibility of the top 1000 specific accessible peaks derived from the RPM across the major cell types enriched in the RPM region (i). The chromatin accessibility profiles of the same peakset from the cell type counterparts of the counter-lateral (LPM here) were also shown. Gene ontology analyses of RPM accessible peaks-linked genes were also performed (j). Meso Prog is short for mesoderm progenitors; Cardiac M is short for cardiac mesoderm; Parax M 1 is short for paraxial mesoderm 1; Mesen is short for mesenchyme. k, l Heatmaps showing the chromatin accessibility of the top 1000 specific accessible peaks derived from the LPM across the major cell types enriched in the LPM region (k). The chromatin accessibility profiles of the same peakset from the cell type counterparts of the counter-lateral (RPM here) were also shown. Gene ontology analyses of LPM accessible peaks-linked genes were shown (l). Nasc M is short for nascent mesoderm; Lateral M is short for lateral mesoderm; Parax M 1 is short for paraxial mesoderm 1; Parax M 2 is short for paraxial mesoderm 2.
To assess the efficacy of ST-MAGIC, we found that cells were mapped evenly across spatial spots in the reconstructed atlas (Supplementary Fig. 7b), and the global gene distributions across five stages in ST-MAGIC were well correlated with Geo-seq results (Supplementary Fig. 7c, d), JSS scoring also revealed a consistent gene expression architecture between ST-MAGIC and the Geo-seq dataset (Supplementary Fig. 7e). Taking the expression of Sox2 as an example, the expression pattern of Sox2, gleaned from the ST-MAGIC, closely matched the Geo-seq resource and also the experimental results (Supplementary Fig. 7f–h). Histone modification H3K27ac has been usually used as an active histone marker frequently deposited at accessible chromatin regions27. Following this, we selected chromatin regions with region-specific H3K27ac distribution, as revealed in our previous study4, and checked their spatial distributions of chromatin accessibility from the ST-MAGIC. For example, the chromatin region (chr10: 63347793-63348989), which locates upstream of the Sirt1 gene and is specially marked by H3K27ac in the anterior epiblast (A) region (Supplementary Fig. 7i), also exhibits anterior epiblast-specific accessible pattern as revealed by the ST-MAGIC (Supplementary Fig. 7i).
We further analyzed the spatial distributions of the embryonic cell types (Supplementary Fig. 8a). Mostly, cell types were assigned to their expected embryonic spatial positions (Supplementary Fig. 8a–c). For example, ectoderm precursors were mapped to the anterior region of the epiblast, while primitive streak cells were specially mapped to the expected posterior region (Supplementary Fig. 8a, c). Interestingly, we observed a clear regionalization of the mesoderm subtypes, particularly in the E7.5 embryos (Fig. 2b and Supplementary Fig. 8d–f). Specifically, the cardiac mesoderm and mesenchyme cells were mostly located in the proximal region, while the paraxial mesoderm 1 cells and the paraxial mesoderm 2 cells were situated in the medial-distal region (Fig. 2b). The spatial distributions for these mesoderm subtypes were validated by determining the related signatural gene expression in vivo and cross-validated in SEU-3D atlas28 (Fig. 2c and Supplementary Fig. 8d–f).
Apart from revealing the formation of cell type-specific spatial territories, ST-MAGIC also unveils the spatial-specific usage of distal linked peaks for gene regulation. For example, Otx2, which is broadly expressed in the epiblast (EPI) and visceral endoderm (VE)29 (Fig. 2d, e), is linked to three spatial types of chromatin peaks: EPI-specific, VE-specific, and universal peaks (Fig. 2f and Supplementary Fig. 8g). To specify, ST-MAGIC revealed that EPI-specific peaks were predominantly located in the epiblast region, while the VE-specific peaks were located in the visceral endoderm region (Fig. 2f and Supplementary Fig. 8g).
ST-MAGIC allows the exploration of specific cell type compositions, gene expression pattern, and chromatin accessibility at any spatial or temporal coordinate across mouse gastrulation (Fig. 2a). Previously, we reported the symmetry breaking event for the left-right body axis first emerges at the late gastrulation stage, manifesting as differential BMP signaling activity and target gene expression in the contralateral proximal mesoderm6. To trace the multi-omic basis for the initiation of left-right asymmetry, we extracted the molecular information from the proximal lateral region of the mesoderm layer using ST-MAGIC (Fig. 2g–l). Consistently, we found that genes related to left-right symmetry breaking also showed laterally biased expression (Supplementary Fig. 9a). Exploration of cell type composition revealed that the mesoderm progenitors, cardiac mesoderm, paraxial mesoderm 1 and mesenchyme cells were over-represented in the right proximal mesoderm region (RPM), while nascent mesoderm, lateral mesoderm, paraxial mesoderm 1 and paraxial mesoderm 2 cells were enriched in the left proximal mesoderm region (LPM) (Fig. 2g, h). Profiling of the top 1000 specific accessible peaks in both RPM and LPM regions revealed that asymmetric levels of chromatin accessibility was discernable between left and right cell type counterparts (Fig. 2i, k, Supplementary Fig. 9b and Supplementary Data 3). To identify the potential biological functions of these asymmetric peaks, we performed gene ontology analyses for these peaks linked genes. Interestingly, BMP signaling pathway-related genes were associated with RPM enriched peaks, while genes related to somitogenesis and heart development were regulated by peaks with higher accessibility in the LPM (Fig. 2j, l). We also traced the emergence of these asymmetric peaks during gastrulation (Supplementary Fig. 9c–n). Intriguingly, both left and right lateral asymmetric peaks became accessible from E6.75 onward (Supplementary Fig. 9c, l), coinciding with the emergence of mesoderm subtypes from the primitive streak cells. For example, two specific peaks, linked with Lefty2 expression, became accessible at the E6.75 nascent mesoderm cells when Lefty2 expression begins, and showed LPM higher distribution at the E7.5 stage when Lefty2 expression is higher on the left side (Supplementary Fig. 9d–g). One of the two peaks is related to a known ASE element30, while the other is a newly-identified Lefty2 regulatory element (Neo_LRE) (Supplementary Fig. 9e, f). Genetic deletion of Neo_LRE in mouse embryonic stem cells showed that Lefty2 expression was severely affected in the knockout (KO) cells, but no severe morphological changes compared to wild-type (WT) controls during gastruloid differentiation in vitro (Supplementary Fig. 9h–k).
Thus, the ST-MAGIC resource established by spatiotemporal reconstruction of the multi-omic atlas to temporal-matched spatial coordinates facilitates in-depth investigation of multi-dimensional molecular architectures of cell types in defined domains of the gastrulating embryo.
ST-MAGIC (+) infers the spatiotemporal turn-over of enhancer regulons
The hierarchical activation of gene regulatory networks by developmental signals and TFs is crucial for mammalian development31. To explore the logics of TF GRNs underpinning gastrulation, we applied the SCENIC + 32, a method for the inference of enhancer-driven TF GRNs, to systematically profile the TF distribution, TF target gene expression as well as TF target peak chromatin accessibility across the annotated cell types. Major regulators for germ layer development were recovered (Fig. 3a and Supplementary Figs. 10, 11). Notably, while the distribution of TF expression showed high cell type-specificity, the presence of TF targets, especially for the TF target peaks, often exhibited shared pattern among closely related neighbors in the same pedigree (Figs. 1c, 3a). For example, in the blood cell lineage, which encompasses hematoendothelial progenitors, blood progenitors 1 and blood progenitors 2, a clear TF expression hierarchy from Etv2, Gata2 to Tal1 was observed. However, the chromatin accessibility for these TF targets remains indistinguishable. This phenomenon supports that Etv2 could function as a priming factor responsible for enhancer opening prior to the hematoendothelial fate commitment (Fig. 3a)33. The sharing of accessible TF target peaks (such as Noto TF) was also detected in cells of axial mesendoderm lineage and definitive endoderm lineage (Fig. 3a), both of which were derived from the anterior primitive streak (Fig. 1c).
a Heatmap showing the transcription factor (TF) expression (left), the AUC score of TF-target geneset (middle), and the AUC score of TF-target peakset (right) of the eRegulon on a color scale. Cell types were ordered on the basis of their developmental relationship. Representative TF eRegulons were labeled next to the heatmap. g is short for TF-target genes, p is short for TF-target peaks. b Scheme of the workflow to generate the expanded version of ST-MAGIC. c The visualization of T expression and the downstream regulatory network in E7.5 embryo using ST-MAGIC and ST-MAGIC (+). The distribution of T transcripts was validated by RNAscope (right panel). n = 3 replicates for T staining. d Representative ST-MAGIC snapshot of T targets in E7.5 embryo. Tbx6 and its genomic neighboring peak bound by T were specifically selected. T ChIP-seq data was drawn from the published dataset65. e ST-MAGIC (+) visualization of NOTO eRegulon revealing the spatiotemporal asynchrony among Noto expression, NOTO target gene expression and NOTO target peak chromatin accessibility. f Genome browser snapshot displaying the dynamics of chromatin accessibility of NOTO target peak (blue square marked region) around Noto locus with corresponding gene expression level (right side). g Enrichment of β-Catenin (top) and Smad2/3 (bottom) at NOTO target chromatin peaks. ChIP-seq information was drawn from a dataset of βCatenin ChIP-seq in EpiLCs39 and Smad2/3 ChIP-seq from EBs42.
To explore the spatiotemporal distribution of these eRegulon imputed by SCENIC + , we developed a pipeline called ST-MAGIC (+), which projects the pre-defined TF target gene sets or target peak sets to the ST-MAGIC atlas through Area Under Curve (AUC) scoring (Fig. 3b). Theoretically, the so-constructed ST-MAGIC (+) atlas should be able to visualize the spatiotemporal turnover of TF GRNs during mouse gastrulation. To validate the fidelity of ST-MAGIC (+) profiling, we first checked the spatial distribution of region-specific H3K27ac modified peaks inferred from the epigenetic landscape of the mouse gastrula4. As shown, region-specific peak sets were accurately mapped to their sampling origins (Supplementary Fig. 12a). Next, we examined the spatial distribution of GRNs for the major regulators identified in Fig. 3a. Taken the transcription factor, T, as an example, Geo-seq and ST-MAGIC revealed that T expression is localized to the primitive streak and anterior mesendoderm in E7.5 embryo (Fig. 3c), and T’s target genes showed a similar distribution pattern (Fig. 3c). However, for the target peaks of T, we observed a broader spatial distribution pattern than the TF expression, manifesting as high level of chromatin accessibility in both the primitive streak and neighboring mesodermal regions (Fig. 3c). This observation was verified through determining the direct T binding peak around the gene locus of Tbx6, which is a well-known T target gene, using ST-MAGIC (Fig. 3d). Moreover, considering that T continues to express in the primitive streak throughout gastrulation and that the mesoderm cells are the immediate progeny of the primitive streak cells34, the broad accessible pattern of T binding chromatin regions suggests that T may act as a priming factor with the ability to open a broad spectrum of chromatin regions and instruct the subsequent fate commitment of mesoderm cells.
Finally, we dissected the dynamics of shared accessible patterns among trajectory neighborhoods for the axial mesendoderm lineage. As resolved by ST-MAGIC (+), we found that the target genes of NOTO exhibit a matched spatiotemporal distribution with the TF expression, but the chromatin accessibility of NOTO target peaks was widespread in the endoderm layer of the E7.5 embryo, where Noto is not expressed (Fig. 3e and Supplementary Fig. 12b, c). Further examination revealed that the chromatin accessibility of NOTO target peaks was elevated at early gastrulation stage, especially in the E7.0 anterior primitive streak region, well before Noto expression emerged (Fig. 3e; Supplementary Fig. 12b). Detailed exploration of the direct NOTO target chromatin region around the Noto locus supported the spatiotemporal kinetics of NOTO eRegulon by ST-MAGIC (+) (Fig. 3f). Global examination of the spatiotemporal concordance of TFs and their targets by Pearson correlation and SSIM index highlighted that the pervasive spatiotemporal asynchrony between TFs and their targets (Supplementary Fig. 12d, e).
These results indicate that crucial upstream players would be involved in setting up the chromatin level of TF GRN, such as NOTO, during axial mesendoderm lineage development. It has been reported that the modulations of WNT and NODAL signaling are involved in the patterning of the anterior primitive streak35. We found that both WNT and NODAL signaling are highly enriched at these chromatin regions (Fig. 3g), suggesting crucial inductive signals may participate in establishing the pre-accessible chromatin pattern for NOTO targets.
ST-MAGIC (+) reflects the spatiotemporal responsiveness to developmental signaling
Inductive developmental signals such as WNT and NODAL signaling act as morphogen cues with localized production but imparting distant function, playing crucial roles in elaborating the proper embryo arrangement and related cell lineages. Multi-layered dimensions, including the morphogen dosage (concentration and duration), the completeness of the signaling cascade components, and the intrinsic competence of the receiving cells, collectively influence the interpretation of signals. Currently, a systematic evaluation of the intrinsic chromatin competence for signaling function remains unexplored36. To characterize the spatial context-dependent mechanisms for interpretating developmental signals, we applied ST-MAGIC (+) to investigate the hierarchy of signal and effector distribution, associated chromatin responsiveness and gene-peak linkage inferred transcription output along the lineage trajectory (Fig. 4a). The spatial distributions of key signal components were characterized by determining the expression pattern of related genes, the chromatin responsiveness to signals was delineated by profiling the spatial accessibility of direct signal effector targets, and the spatial transcription output was measured by checking the expression pattern of signal effector chromatin binding regions linked genes (Fig. 4a).
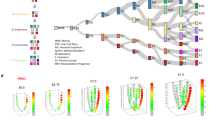
a Schematic overview of the morphogen gradient and the interpretation of signal transduction cascade in embryo patterning, and the in silico visualization using ST-MAGIC (+). b ST-MAGIC visualization of the distribution of WNT signaling ligand-Wnt3 (left) and the antagonist-Dkk1 (right) during mouse gastrulation. c The 2380 EpiLC-specific βCatenin binding peaks and the 801 linked genes were mapped to E6.5 and E7.5 embryo to show the spatiotemporal dynamics of chromatin responsiveness (left) and transcriptional output of WNT signaling (right). d ST-MAGIC visualization of the NODAL signaling ligand-Nodal and the antagonist-Cer1 during mouse gastrulation. e ST-MAGIC (+) visualization of embryoid body (EB)-specific Smad2/3 binding peaks (3635 peaks, left) and the linked genes (320 genes, right) reflecting the spatiotemporal transition of intrinsic chromatin responsiveness to NODAL signaling. f, g The spatiotemporal expression of Nodal (f) and the transition of intrinsic chromatin responsiveness to NODAL signaling (g). h The transition of intrinsic chromatin responsiveness to signaling is mostly related to E6.75_PS region (enriched with primitive streak cells) and E7.0_APS_M region (enriched with anterior primitive streak and nascent mesoderm). Left: the specific spatial locations of E6.75_PS and E7.0_APS_M region; middle: the cell type composition at indicated regions; top right: venn diagram showing the direct comparison of region-specific WNT signaling responsive peaks between E6.75_PS and E7.0_APS_M; bottom right: venn diagram showing the direct comparison of region-specific NODAL signaling responsive peaks between E6.75_PS and E7.0_APS_M. i. j. ST-MAGIC (+) visualization (i) and motif enrichment (j) of the 92 E7.0 APS_M specific WNT signaling responsive peaks during the late stage (E7.0, E7.25, E7.5) of mouse gastrulation. k The model describing the spatial evolution of WNT signaling responsiveness at late gastrulation. l, m ST-MAGIC (+) visualization (l) and motif enrichment (m) of the 291 E7.0 APS_M specific NODAL signaling responsive peaks during the late stage (E7.0, E7.25, E7.5) of mouse gastrulation. n The model describing the spatial evolution of NODAL signaling responsiveness during the late stage of mouse gastrulation.
We first analyzed the global pattern and the cellular responsiveness of WNT signaling across gastrulation (Supplementary Fig. 13a). Consistent with known biology37, the Wnt signal ligand encoded by Wnt3 was predominantly expressed in the primitive streak and adjacent mesoderm tissue, while the Wnt signal antagonist, Dkk1, was specifically expressed in the anterior visceral endoderm region (Fig. 4b and Supplementary Fig. 13b–e). However, the WNT signaling receptor, Lrp6, and the effector, Ctnnb1, (also known as β-Catenin) were widely distributed within the gastrula (Supplementary Fig. 13f).
To systematically characterize the embryo chromatin responsiveness to WNT signaling, we sourced two published ChIP-seq datasets for WNT signaling effector-β-Catenin from the in vitro ESCs38 and EpiLCs39. Through differential binding peaks analyses, the chromatin regions were classified into three groups of common, ESC-specific, and EpiLC-specific peaks (Supplementary Fig. 13g, h). Spatiotemporal reconstruction of these peaks though ST-MAGIC (+) revealed that ESC-specific peaks were accessible in the anterior epiblast, where cells remain in a pluripotent state40 (Supplementary Fig. 13i). Meanwhile, for the peaks with EpiLC-specific β-Catenin binding, ST-MAGIC (+) reported that these peaks were accessible in the embryonic region, largely overlapped with Wnt3 distribution (Fig. 4b, c). In-depth analyses of the ST-MAGIC (+) results for EpiLC-specific peaks across five stages of the gastrula showed that the PS region and neighboring posterior mesoderm region remained consistently accessible across the gastrulation (Fig. 4b, c and Supplementary Fig. 13j, k). Together, these results underscore that in response to WNT signaling input, cells first turnover the Wnt-responsive chromatin landscape during the transition from pluripotent state to lineage primed progenitors, and then modulate the chromatin accessibility to accommodate the specification of posterior mesoderm subtypes during gastrulation (Supplementary Fig. 13l).
We next analyzed the intrinsic cellular responsiveness to NODAL signaling (Supplementary Fig. 14a), a crucial signal for embryo patterning and germ layer formation41. Examination of the components of NODAL signaling cascade in the ST-MAGIC atlas revealed that the ligand-Nodal was initially expressed in the posterior epiblast at early gastrulation stage but abruptly shifted to the distal tip region at the E7.5 stage (Fig. 4d). In the meantime, the NODAL signaling antagonist-Cer1 was constantly expressed in the anterior visceral endoderm region (Fig. 4d and Supplementary Fig. 14b–f). However, the NODAL signaling receptors and the effectors were widely expressed in the gastrula (Supplementary Fig. 14g). The changes of expression domain of the NODAL ligand-antagonist pair may underpin the shift of the NODAL morphogen gradient during mouse gastrulation.
To chart the chromatin responsiveness along with the NODAL pattern shift, we incorporated the published ChIP-seq datasets for NODAL signaling effectors SMAD2 and SMAD342. We found that the chromatin binding ability of the Smad2/3 complex was largely de novo generated in EB cells, an in vitro counterpart of mesoderm cells (Supplementary Fig. 14h, i). GREAT analyses further indicated that these peaks are involved in the cell fate specification, anterior/posterior formation and primitive streak formation (Supplementary Fig. 14i). Then, we used ST-MAGIC (+) to profile the spatial distribution of these peaks during gastrulation. Interestingly, we found that the global chromatin accessibility of these EB-specific Smad2/3 binding peaks first appeared in both the endoderm region and PS regions (Fig. 4e). Subsequently, the accessibility of these peaks in the PS region gradually shifted and formed a distal-to-proximal accessibility gradient in the E7.5 gastrula (Fig. 4e and Supplementary Fig. 14j). Notably, the chromatin responsiveness to Nodal signaling started relocating at an earlier stage between E6.75 and E7.0, ahead of the re-alignment of the Nodal gradient in the germ layers occurring between E7.25 and E7.5 stage (Fig. 4f, g). These results highlight that the spatial turnover of NODAL signaling chromatin responsiveness occurs earlier than the repositioning of Nodal activity.
To determine the intrinsic features for the relocation of signaling chromatin responsiveness, we extracted the specific subset of signaling responsive chromatin peaks which show high accessibility at the E6.75 primitive streak region (E6.75_PS) or at the E7.0 anterior primitive streak and adjacent mesoderm region (E7.0_APS_M) (Fig. 4h). Generally, the signaling responsive peaks for NODAL and WNT signaling were largely independent between E6.75_PS region and E7.0_APS_M region (Fig. 4h and Supplementary Data 4, 5). As shown, the E7.0_APS_M specific WNT signaling responsive peaks were gradually extended to the whole posterior region (Fig. 4i), and enriched with typical WNT signaling co-effector LEF1 and TCF7L1 motifs (Fig. 4j). The posterior enriched WNT signaling responsiveness (Fig. 4k) support the notion that WNT signaling is instrumental for regulating posterior embryo development36. Interestingly, for NODAL signaling, profiling of the E7.0_APS_M specific signal responsive peaks exhibited a gradual shift pattern of these peaks towards the distal region of the E7.5 embryo, where Noto and Nodal-expressing cells reside (Figs. 3e, 4d, l). Functional characterization of E7.0_APS_M NODAL signaling responsive peaks showed enrichment in chordate embryonic development and embryo pattern specification process, and the knockout of related genes can frequently lead to abnormal germ layer morphology, abnormal rostral-caudal axis patterning and absent floor plate (Supplementary Fig. 14k). Motif enrichment analyses of these peaks revealed significant enrichment of major TF regulators especially for axial mesendoderm development such as Zfp281, Foxa2, and Foxj1 (Fig. 4m). These results indicate that the spatial re-distribution of embryonic responsiveness to Nodal signaling may play specific roles in the forthcoming development of axial mesendoderm lineage (Fig. 4n).
Molecular hierarchy underlying axial mesendoderm lineage development
The axial mesendoderm has been reported to be the direct developmental antecedent for the midline notochord cells, which instructs the following somitogenesis and neural patterning7. To better understand axial mesendoderm lineage development, it is essential to document the spatiotemporal context of the stepwise appearance of molecular features and cellular states during the lineage formation process, with accurate spatiotemporal information.
Here, based on the inferred lineage trajectory for the gastrula (Figs. 1c, 5a) and the ST-MAGIC reconstructed cell distribution for the mouse gastrula (Supplementary Fig. 8a), we found that cells residing in the E7.5 distal tip, where the prospective notochord cells first emerge43, are composed of the node cells and anterior mesendoderm cells (Fig. 5a and Supplementary Fig. 15a). Expression of known notochord markers, Shh, Noto, and Foxj1, revealed that the presence of two distinct cell subtypes (Shh+Noto-Foxj1- and Shh+Noto+Foxj1+) at the ventral distal surface for the E7.5 embryo, and the two distinct cell subtypes can persist through organogenesis (Fig. 5b and Supplementary Fig. 15b–d).
a ST-MAGIC revealing the spatiotemporal lineage trajectory of axial mesendoderm (top) from the anterior primitive streak cells of E7.0 embryo to the derivatives – the node and anterior mesendoderm of E7.5 embryo. Bottom: ST-MAGIC results showing the changes in the location of axial mesendoderm lineage-related cells during gastrulation. PS is short for primitive streak cells, APS is short for anterior primitive streak cells. b ST-MAGIC showing the expression of known markers for node and anterior mesendoderm cells (Shh, Noto, Foxj1), and validated by whole-mount RNAscope. AME is short for anterior mesendoderm. c Violin plot showing the AUC scoring for E7.0_APS_M NODAL responsive peaks (top) and WNT signaling responsive peaks (bottom) accessibility during axial mesendoderm development. Node P is short for node precursors, AME P is short for anterior mesendoderm precursors. d UMAP of mouse axial mesendoderm during gastrulation with scVelo-projected velocities, shown as streamlines. Top: UMAP representation with RNA velocity stream plots overlaid for axial mesendoderm lineage; bottom: UMAP representation with inferred trajectories colored by pseudo-time ordering. e Heatmap showing the direct comparison of differentially expressed genes for the two axial mesendoderm subtypes (node and anterior mesendoderm). f, g Line chart describing the smoothed trends of gene expression in pseudotime (f). The trend for each gene was shown for each trajectory leading to the terminal populations of node (Dynlrb2 and Rsph9) or anterior mesendoderm (Col2a1). The gene expression pattern was validated by RNAscope (g). 3 replicates for each probe with similar results. h–k Heatmaps displaying the smoothened gene expression trends (left) and chromatin accessibility levels of linked peaks (right) in node (h, i) and anterior mesendoderm development (j, k). Cells ordered by the pseudo-time were marked (top of each heatmap). Three major gene groups were identified, with the G3 group sub-dividing into two sub-groups. l SCENIC + analyses revealing the transcription factor hierarchy during axial mesendoderm development. The numbers for selected TF-target genes and TF-target peaks were labeled, where g was short for TF-target genes, p was short for TF-target peaks. m Line plot describing the TF expression, TF-target gene expression, as well as TF-target peak accessibility of Eomes (top panel) and Foxa2 (bottom panel) during axial mesendoderm lineage. The spatiotemporal expression dynamics for both TF was validated at the right. n = 3 RNAscope replicates for each probe were performed with similar results. n Line plot describing the TF expression, TF-target gene expression, as well as TF-target peak accessibility for Foxj1, Noto, Pou6f1, and Sox9 during axial mesendoderm development.
The inferred lineage trajectory indicates that the node and the anterior mesendoderm cells are spatially derived from the E7.0 APS cells (Fig. 5a). Concurrently, chromatin responsiveness to NODAL and WNT signaling gradually increased in the E7.0 anterior primitive streak cells and is maintained at high accessibility levels till the E7.5 stage (Fig. 5c). To chart the molecular dynamics along the developmental trajectory, we applied CellRank44 to assign the cell fate probabilities for all the related cells at single-cell resolution. RNA velocity and pseudotime ordering revealed that the cells at an early stage (E6.5 primitive streak, E6.75 anterior primitive streak) are unspecified for cell fate directions, while cells at more advanced stages show directed flow towards the E7.5 node and E7.5 anterior mesendoderm (Fig. 5d). In-depth analyses revealed that the two cell subtypes showed distinct transcriptomic and epigenomic features (Supplementary Fig. 15e). To specify, the node cells express high levels of cilia and dynein-related genes, such as Foxj1 and Dnah11, while the anterior mesendoderm cells express high level of collagen-related genes, such as Col2a1 and Srd5a2 (Fig. 5e and Supplementary Fig. 15f–i). Molecularly, smoothed trends of gene expression along the pseudotime order showed that cilium related genes, such as Dynlrb2 and Rsph9, were gradually upregulated down the road to node, while the collagen gene Col2a1 was expressed till the terminal stage of the anterior mesendoderm (Fig. 5f, g).
Pseudotime-based tracking of gene expression cascades and associated chromatin peaks revealed three successive stages (G1, G2, G3) enriched in the early, intermediate and terminal populations, respectively (Fig. 5h–k). Intriguingly, for both lineages, a subset of the terminal stage expressed genes (G3a) was regulated by chromatin peaks which get accessible at an earlier stage (Fig. 5h–k). For example, in node cells, Foxj1 expression was evidently upregulated at the E7.25 stage, but the chromatin accessibility for the linked distal peak-Foxj1-DRE was accessible in the E7.0 anterior primitive streak cells (Fig. 5i). Similarly, for anterior mesendoderm cells, genes like Sox9 showed the same pattern (Fig. 5k). The presence of these pre-accessible chromatin peaks points to early epigenomic priming for the development of axial mesendoderm lineage.
To systematically determine the molecular hierarchy of the enhancer-driven TF GRNs for the axial mesendoderm lineage, we used SCENIC+ to infer the potential candidate TFs and the dynamics of downstream target genes and linked chromatin peaks (Fig. 5l). A clear hierarchy of TF usage turnover was captured. To specify, pluripotency-related TFs, such as Nanog, were enriched at the E6.5 stage; early lineage factors, such as Eomes, were enriched at the E6.75 stage; subsequently, for the intermediate stage factors, such as Mixl1, Foxa2 and Lhx1, were enriched in the E7.0 anterior primitive streak cells, E7.25 node precursors, and E7.25 anterior mesendoderm precursors, respectively; and terminal stage factors, such as Noto and Sox9, were abundant in E7.5 node and anterior mesendoderm cells, respectively (Fig. 5l). Detailed analyses of these eRegulons revealed that the early and intermediate TFs first upregulate TFs gene expression, followed by increasing the target gene expression as well as target peak chromatin accessibility (Fig. 5m and Supplementary Fig. 16a). In contrast, for the terminal stage enriched TFs, the chromatin of TF target peaks becomes accessible at an earlier stage, followed by TF expression and TF target gene expression (Fig. 5n). This temporal turnover of TF GRNs may reflect a molecular relay underlying the sequential cascade of cell fate commitment during axial mesendoderm development. Notably, among multiple known TF GRNs, we identified a novel TF, POU6F1, which exhibits highly specific expression of both the TF itself (adjusted P-value, 8.4e-29) and its target genes (adjusted P-value, 1.3e-55) in the node (Fig. 5l and Supplementary Fig. 16b). However, the chromatin regions targeted by POU6F1 get accessible prior to the expression of the TF and its target genes (Fig. 5n).
Distinct roles of stage-related TFs in regulating gene expression and setting up chromatin accessibility
To determine the relevance of TF hierarchy and signal responsiveness during axial mesendoderm lineage development, we first checked the distribution of the TF GRNs for early stage TF (EOMES), intermediate stage TFs (MIXL1, FOXA2, LHX1), and terminal stage TFs (NOTO and POU6F1) during gastrulation using ST-MAGIC (+). Consistent with the SCENIC+ results (Fig. 5m, n), we found that for the early and intermediate stage TFs, expression of these TFs commences prior to or concurrently with the target gene expression and target peak accessibility (Supplementary Fig. 16c–f). In contrast, for the terminal stage TFs-NOTO and POU6F1, we observed that their target chromatin peaks first become accessible at the expected APS region and endoderm region at the E7.0 stage, when TFs and their target genes only show minimal expression (Figs. 4e, 6a, b and Supplementary Fig. 17a–g). With the progressing of embryo development, once Noto and Pou6f1 reached the expression summit at the E7.5 node region, the target genes show abundant enrichment at the same region (Figs. 4e, 6a, b and Supplementary Fig. 17a–g). These results suggest that the early and intermediate stage TFs may function as priming factors by opening a broad spectrum of chromatin regions, thereby setting up a permissive chromatin environment for the following signaling and the terminal stage TFs-NOTO and POU6F1. The expression of TFs-NOTO and POU6F1 at the terminal stage can then interact with the pre-accessible chromatin regions, and enhance the expression of target genes to establish the requisite transcriptomic state for further lineage differentiation.
a, b ST-MAGIC visualizing the distribution of Pou6f1 eRegulon during mouse gastrulation (a). Top: ST-MAGIC of Pou6f1 gene expression; middle: ST-MAGIC (+) of POU6F1 target geneset expression; bottom: ST-MAGIC (+) of POU6F1 target peak chromatin accessibility. RNAscope experiment was performed to validate the expression of Noto and Pou6f1 in the gastrula (b). n = 3 RNAscope replicates for each probe were performed with similar results. c Heatmaps showing the enrichment of Smad2/3, EOMES, and FOXA2 binding at the prospective target peak regions of NOTO (left) and POU6F1 (right). Information was sourced from a dataset of Smad2/3 ChIP-seq42, EOMES ChIP-seq65, ATAC-seq for both WT EB cells and Eomes-KO EB cells66, FOXA2 ChIP-seq67, ATAC-seq for both WT mesendoderm (MEN) cells and Foxa2-KO MEN cells67. Eo-KO is short for Eomes knockout cells, F2-KO is short for Foxa2 knockout cells. Statistical significances were shown in Supplementary Fig. 18d. d The workflow to determine the biological functions of Noto and Pou6f1. Mouse icon was created in BioRender, Tan, F. (2025) https://BioRender.com/1atcqml. e UMAP plotting of the acquired single cell multi-omic data for the wildtype (WT), Pou6f1 KO and Noto KO E7.5 embryos. Top: The 10xscMultiome-snRNA-seq data, bottom: 10xscMultiome-snATAC-seq data. f Bar plot showing the relative abundance of cell types in the mutants. g Violin plot showing the AUC score for the down-regulated node-related genes in WT, Noto KO and Pou6f1 KO embryos. h RNAscope analyses validating the expression change of ciliogenesis-related gene-Foxj1 in Noto KO and Pou6f1 KO embryos. i, j The cilia in E7.5 mouse embryos of different genotypes (i). Scanning electron micrographs of the node (white arrowhead) in the embryos (left panels) and the cilia (white arrowhead) in the node (right panels) (j). Violin plot showing the measured length of the cilia in WT, Noto KO and Pou6f1 KO embryos. *** significant difference in cilia length across between genotypes by Welch’s t test tests. Box plots display the median (center line), the first and third quartiles (box limits), and whiskers indicating the minimum and maximum values within 1.5 times the interquartile range. k Heatmaps profiling the chromatin accessibility around the prospective NOTO target peakset (left) and POU6F1 target peakset (right).
To demonstrate this molecular framework, we integrated datasets of both EOMES ChIP-seq (from embryoid bodies in vitro) and FOXA2 ChIP-seq (from mesendoderm cells differentiated in vitro) and the chromatin accessibility data of Eomes-KO and Foxa2-KO cells. By profiling the enrichment around the pre-opening Foxj1-DRE element (Fig. 5i), we found that the chromatin accessibility for Foxj1-DRE element was markedly downregulated in Eomes-KO and Foxa2-KO cells (Supplementary Fig. 18a), which strongly suggests the roles of EOMES and FOXA2 in opening Foxj1-DRE chromatin region. We also systematically analyzed the enrichment of EOMES and FOXA2 around the E7.0_APS_M signaling responsive peaks, the NOTO target peaks, and also the POU6F1 target peaks, we found that both EOMES and FOXA2 were enriched around these pre-defined genomic regions (Fig. 6c and Supplementary Fig. 18b, c). Moreover, the knockout of either Eomes or Foxa2 led to the reduction of chromatin accessibility around these loci (Fig. 6c and Supplementary Fig. 18b–d). Therefore, EOMES and FOXA2 may play roles in pre-opening signaling responsive elements and NOTO and POU6F1 target chromatin regions.
Next, we investigated whether the TFs-NOTO and POU6F1 play roles in establishing the terminal transcriptomic state but not the chromatin setup. Cross-referencing published atlas3 confirmed the enrichment of these two genes in the notochord cells (Supplementary Fig. 19a). We then generated two mouse mutants with the genetic deletions of the Noto or Pou6f1 genes (Fig. 6d and Supplementary Fig. 19b). No visible morphological phenotypes were detected in either mutants at E7.5 stage (Supplementary Fig. 19c). To probe the potential molecular abnormalities, we collected E7.5 WT control, Noto KO, and Pou6f1 KO embryos and performed 10xscMultiome sequencing (snRNA-seq + snATAC-seq). Cells were annotated by computationally mapping their transcriptome onto our E7.5 WT atlas (Fig. 6e). The cell type compositions in the KO embryos remained comparable to WT, except for the axial mesendoderm-related cells, in which Noto and Pou6f1 are expressed and now successfully removed (Fig. 6f and Supplementary Fig. 19d, e). Moreover, in both Noto KO and Pou6f1 KO embryos, the expression of node cell-related genes and ciliogenesis was severely disrupted (Fig. 6g–j, Supplementary Fig. 19f, g and Supplementary Data 6). Phenotypically, by E11.5, directional axis turning, which is related to proper notochord function17, was randomized (Supplementary Fig. 19h, i). Importantly, the chromatin accessibility of the TF target chromatin peaks remained unchanged (Fig. 6k and Supplementary Fig. 19j). These results demonstrate that NOTO and POU6F1 function primarily as the final-step transcriptional regulators in the TF relay during the sequential development of the mouse axial mesendoderm.
Together, through systematic exploration of the ST-MAGIC (+) resource, we unraveled the cellular events and associated highly-organized regulatory cascades of TF and signal GRNs, and demonstrated the differential roles for TFs from different stages in shaping transcriptomic architecture and chromatin accessibility landscape at different milestones of the lineage trajectory (Fig. 7).
Model summarizing the spatiotemporal logics of cell type regionalization, epigenomic programming and transcriptional patterning during mouse gastrulation and the hierarchical relay of gene regulatory networks underlying axial mesendoderm development.
Discussion
Gastrulation is a critical developmental stage responsible for generating the three primary germ layers and related derivatives. This process involves scheduling the spatiotemporal sequence of events that leads to the formation and positioning of tissue and organ progenitors in the body plan45. Understanding the sequential allocation of cell lineages and acquisition of cell fates during gastrulation is challenging due to the intricate spatiotemporal dynamics involved. In this study, by performing time-series single-cell multi-omic profiling and integrating the dataset with true 3D spatial transcriptome reference data, we established the ST-MAGIC and ST-MAGIC (+) resources, and uncovered an unprecedented level of details of the spatiotemporal GRN dynamics that underlie the sequential cell fate commitment process. This comprehensive integration empowers the exploration of intricate interplays among transcriptomic, epigenomic, and signaling in a spatiotemporal context, providing a detailed overview of the molecular attributes during gastrulation.
The refined gene-peak linkage identification method, BioCRE, was developed to address the challenge of linking genes with their regulatory elements. While existing methods can predict regulatory elements, they often lack the gene-peak dual regulation precision required for dynamic processes. BioCRE integrates transcriptomic and chromatin accessibility data to construct robust gene-peak linkages, enabling a more accurate and context-specific understanding of GRNs. Although standard datasets are not yet available to benchmark the efficacy of BioCRE, preliminary results using external datasets indicate its ability to identify key regulatory elements and their interactions at high precision. The incorporation of more external multiome data and matched gold standard dataset will foster a more systematic evaluation of BioCRE. This method offers an advantage in uncovering the molecular mechanisms driving cell fate decisions, which can be further validated experimentally.
Single-cell omic technologies have significantly broadened the molecular understanding of vertebrate embryogenesis5,36,46,47,48,49. Joint analyses of various data modalities, including spatiotemporal dimensions, transcriptome, epigenome, proteome, metabolome, etc, hold promise for achieving a comprehensive understanding of the principles of developmental biology50,51. Cross-modal data integration using a bioinformatic strategy is essential to maximize the value of existing data resources. However, integrating different data modalities remains challenging due to the lack of reliable anchors and toolkits. Here, taking transcriptomic information as an anchor to integrate 10xscMultiome with the stage-matched spatial transcriptome coordinates, we constructed the ST-MAGIC atlas with genuine spatiotemporal information, transcriptome architecture, and epigenomic landscape for diverse cell types in the gastrula. Moreover, by incorporating the eRegulon datasets and published ChIP-seq data of signaling effector, we expanded the ST-MAGIC into ST-MAGIC (+), which enables the delineation of spatiotemporal dynamics of developmental TFs and the characterization of spatiotemporal transition of chromatin responsiveness to developmental signaling during sequential cell fate commitment for a broad spectrum of cell lineages at single-cell resolution in the mouse gastrula.
The ST-MAGIC and ST-MAGIC (+) platforms provide a versatile resource for dissecting the multifaceted mechanisms of embryonic development. Despite the extensive data generated, the scope of this study is limited by the need to focus on specific aspects due to length constraints. In this study, we have unveiled the multi-omic basis for the left-right symmetry breaking event by identifying the potential involvement of distal regulatory elements in regulating the expression of symmetry-breaking genes. We next focused on the axial mesendoderm lineage, particularly the development of the node, to demonstrate the utility of our resource. As known, the axial mesendoderm lineage gives rise to the prechordal plate, anterior head process, and the node-derived notochordal precursors, ultimately forming the notochordal plate on the ventral surface of the mouse embryo7,52,53,54. We identified the spatiotemporal trajectory of the axial mesendoderm lineage, which originates from cells in the anterior primitive streak and gradually features at the distal tip in E7.5 embryos. By tracing the molecular routes for the two cell subtypes (anterior mesendoderm cells and node cells), we found that both are derived from E7.0 anterior primitive streak cells, when and where the intrinsic chromatin setup for developmental signaling starts to relocate. Furthermore, we identified sequential relays of TF GRNs from the early stage, intermediate stage, to the terminal stage for each sub-lineage. Through detailed exploration of TF expression, TF-target gene expression, and TF-target peak chromatin accessibility, we characterized the distinct kinetics for early, intermediate, and terminal stage TFs.
Transcription factors emerging at different stages of lineage development may be responsible for distinct functions. Previous studies have revealed genetic deletion of TFs such as Eomes55 and Foxa256 in mouse embryos leads to the complete absence of node structures. Mutation of Noto in the mouse embryo only shows moderate defects related to ciliogenesis in the node cell, without affecting the formation of the node57. In contrast, the loss of Noto homolog, flh, in zebrafish embryos leads to the absence of notochord-related cells58,59. These results strongly suggest that the mouse NOTO TF is involved in a lower hierarchy during the notochordal development than EOMES, FOXA2, and even its homolog in zebrafish embryos. Here, as revealed in our study, we found that deletion of early stage TF (EOMES) and intermediate stage TF (FOXA2) severely affects the downstream GRNs, whereas ablation of late stage TFs in mouse embryos only affects the expression of target genes but not the pre-established chromatin setup. Phenotypical characterization showed that knockout mice for terminal stage TFs only leads to the down-regulation of ciliogenesis genes and shortened cilium length. These discrepancies suggest that the post-established chromatin landscape for early and intermediate stage TFs and the pre-established chromatin landscape for terminal stage TFs may build up the molecular basis of developmental competency for node cell formation. Thus, we propose that these distinctions may reflect a molecular ‘priming-specification-determination’ cascade that underlies cell fate commitment. Further exploration of GRN hierarchies across various lineages could define fundamental rules as well as the lineage-specific logics that guide lineage development during embryogenesis.
Several limitations of the current ST-MAGIC and ST-MAGIC (+) resource should be acknowledged. First, the current resources are limited to spatiotemporal information, transcriptome, and chromatin accessibility at Geo-seq spatial resolution. Transcriptome coordinate profiling using Stereo-seq24 or MERFISH60, incorporation of additional data types, such as histone modifications, proteomics, and metabolomics, and optimizations of bioinformatic toolkits would further enhance the utility of the combined dataset in molecular understanding of early embryogenesis. Second, most GRNs for TFs and signaling pathways are defined by integrating consensus targets from public datasets or state-matched in vitro counterparts. Comprehensive profiling of TF or signal targets from precisely matched cell types would improve the fidelity of GRN delineation. Third, while the refined BioCRE method provides useful insights into regulatory elements and gene-peak linkages, further experimental validation and systematic benchmarking are needed to verify the predicted regulatory elements and interactions. This includes validating the roles of linked regulatory elements through functional genomics study in embryo models and cell differentiation models.
In summary, this study establishes a comprehensive spatiotemporal multi-omic resource, that recapitulates the sequential cellular events, and dissecting the continuous molecular cascades of TF GRNs and signaling responsiveness during mouse gastrulation. This work opens an avenue toward a systematic learning of the molecular principles that govern early mammalian embryogenesis in a spatiotemporal context.
Methods
Ethics statement
Mouse used in this study were housed and bred in the SPF facilities of Guangzhou National Laboratory. All animal experiments were performed in compliance with the guidelines of the Animal Core Facility.
Mouse embryo collection and multi-omic profiling
For embryo sampling, C57BL/6 J embryos were harvested from pregnant mice at day 6.5, 6.75, 7.0, 7.25, and 7.5 of gestation (day of vaginal plug detection = Day E0.5). Plugged female mice were picked after mating and marked as embryonic day 0.5 (E0.5). Female mice were sacrificed for embryos collection at specific gestational stages. Embryos were isolated from the uterus and carefully transferred into pre-cool PBS in petri dishes, and surrounding decidua and parietal endoderm tissues were removed using needles under an Olympus stereoscope. Sex of the embryos was not considered in this study.
Careful morphological staging of the acquired embryos was performed before single-cell multi-omic data preparation. The developmental time points of embryos were staged by the proximal-distal span of the PS and the anterior-posterior span of the mesoderm layer. To meet the loading requirements for 10x genomics, embryos from the same stage were pooled together and subjected to TrypLE Express enzyme (Gibco, 12604) incubation at 37 °C for 7–10 min. The acquired single-cell suspension was carefully washed and filtered to ensure proper integrity and avoid cell clumping of each single cell. Cell counts were then assessed with a haemocytometer counted under a microscope. Nuclei isolation and multiome library preparation were performed by following the manufacturer’s instructions (https://www.10xgenomics.com/cn/support/single-cell-multiome-atac-plus-gene-expression/documentation).
For the single-cell multi-omic profiling of the Noto KO and Pou6f1 KO embryos, the parents of the heterozygotic mutation were crossed and checked for the vaginal plug. E7.5 embryos were freshly harvested, and a tiny portion of extraembryonic tissues were collected and genotyped. During the genotyping process, the embryos were freshly frozen in liquid nitrogen. Embryos with the same genotypes (WT, Noto KO homozygote, Pou6f1 KO homozygote) were grouped and dissociated into single nuclei. Approximately 15,000 nuclei for each group were collected and loaded for further single-cell multiome library preparation, following the manufacturer’s instructions.
Enhancer activity reporter assay
Generally, the enhancer reporter assay was performed as previously reported4. In brief, DNA fragments for potential regulatory elements were cloned from the C57BL/6 J mouse genome and then ligated into the plasmid construct containing the minimal Hsp68 promoter and LacZ. The acquired purified plasmids were then linearized and used for pronuclear injections of PN4 stage zygotes with a FemtoJet Microinjection System (Eppendorf). The injected embryos were cultured to the 2-cell stage in KSOM medium with amino acids at 37 °C under 5% CO2, and then transferred to the oviduct of pseudo-pregnant ICR females and marked as 0.5 dpc. Embryos were collected at the corresponding stage for LacZ staining. LacZ staining was performed using a commercialized β-gal staining kit (Beyotime, RG0039).
RNAscope staining for whole mount or sections of embryo
RNAscope probes including mm-Lefty2-C2 (436291-C2), mm-Hand1-C1 (429651), mm-Irx5-C2 (513871-C2), mm-Isl1-C3 (451931-C3), mm-Sox2-C1 (401041-C1), mm-Otx2-C3 (444381-C3), mm-T-C3 (423511-C3), mm-Eomes-C2 (429641-C2), mm-Foxa2-C4 (409111-C4), mm-Shh-C2 (314361-C2), mm-Noto-C3 (1253281-C3), mm-Foxj1-C3 (317091-C3), mm-Dynlrb2-C3 (1243011-C3), mm-Col2a1-C4 (407221-C4), mm-Rsph9-C2 (430201-C2), and mm-Pou6f1-C1 (801931-C1) were bought from the Advanced Cell Diagnostics.
For wholemount RNAscope staining, embryos were fixed in 4% PFA overnight. After serial dehydration and rehydration of embryos using gradient methanol-PBS solution, embryos were subject to the RNAscope protocol following the manufacturer’s instruction(https://acdbio.com/sites/default/files/MK%2050016%20Zebrafish_WISH_Tech%20Note_12042017.pdf). Images of stained embryos were acquired using the LiTone XL system (Light Innovation Technology Limited).
For RNAscope staining with embryo sections, dissected embryos were first fixed in 4% PFA, then dehydrated in 20% Sucrose-PBS and 30% Surcrose-PBS solution, respectively. The dehydrated embryos were embedded in OCT matrix and then cryo-sectioned using Leica CM1950. RNAscope workflow was performed by following the manufacturer’s instructions (https://acdbio.com/ebook/introduction/materialsmethod). Images were acquired by using the Carl Zeiss LSM980 system.
Wholemount in situ hybridization
Wholemount in situ hybridization of RNA transcripts was performed by following the published protocol61. Briefly, DNA fragments encoding the probes of Lety2 were firstly PCR amplified by using the oligos in Supplementary Data 7 using a mouse embryo cDNA library. Embryos at relevant stages were collected in DMEM media and then fixed in 4% PFA at 4 °C overnight. Fixed embryos were then washed in PBS solution at room temperature to remove residual PFA, followed by serial dehydration and rehydration in PBS, 25% Methanol/PBS, 50% Methanol/PBS, 75% Methanol/PBS, and 100% Methanol. Afterward, the embryos were treated with 10 μg/mL proteinase K and incubated with DIG-labeled RNA probes at 70 °C overnight. To remove the remaining RNA probes, embryos were washed in TBST buffer with frequent buffer changing for at least 2–4 h. Then the embryos were incubated with anti-DIG-AP antibody at 4 °C overnight. The embryos were subjected to sufficient TBST wash before final staining with NBT and BCIP solution. The final images of stained embryos were collected using an Olympus SZX16 microscope.
The generation of genome edited mouse embryonic stem cell or mouse model
In this study, we have prepared the genome-edited mouse embryonic stem cell lines with Mesp1 Neo_ME element, Mesp1 EME element, and Lefty2 Neo_LRE element knockout, respectively. The CRISPR-Cas9 system was used to generate the genome-edited mouse embryonic stem cell lines. Guided RNAs were designed by using the online tool Chop-chop (http://chopchop.cbu.uib.no/). The synthesized sgRNA DNA fragments were then ligated into the px330 plasmid with a Cas9 protein expression cassette. The acquired purified plasmids were then transfected into the WT cells using Lipofectamine 3000. The genome-edited clones were then selected and genotyped as previously reported62. The potential off-target editing sites were tested based on the website’s indication.
To determine the roles of Pou6f1, we also generated a mouse model with Pou6f1 deletion. The specific sequences for sgRNA pairs were included in Supplementary Data 7. DNA fragments for the sgRNA and T7-Cas9 were transcribed and purified in vitro using MMESSAGE MMACHINE T7 Ultra Kit (Invitrogen, AM1345) and MEGAclear kit (Invitrogen, AM1908). To prepare a sufficient number of fertilized zygotes, C57BL/6 J female mice (4 weeks old) were superovulated and mated with the male C57BL/6 mice. Twenty-four hours later, fertilized embryos were collected from the oviducts. RNA for Cas9 protein (100 ng/µl) and corresponding sgRNA (100 ng/µl) were mixed in HEPES-CZB medium containing 5 μg/ml cytochalasin B (CB) and injected into the cytoplasm of fertilized eggs using a FemtoJet microinjector (Eppendorf) with constant flow settings. The injected embryos were cultured in KSOM with amino acids at 37 °C under 5% CO2 in the air to reach the 2-cell stage after 24 h in vitro. Two-cell embryos were transferred into pseudo-pregnant ICR female mice. The acquired mouse individuals were then subjected to genotyping for successful Pou6f1 deletion. Mice with Pou6f1 deletion were retained and recorded as F0. To prepare the stable Pou6f1 deletion mouse line, F0 mice were crossed, and the acquired F1 mice were checked. Pou6f1 KO heterozygotes F1 mice were maintained, and the population was expanded for the following experiments.
The Noto KO mouse line was purchased from GemPharmatech (Strain ID: T017401).
Scanning electron microscopy
Embryo samples were freshly collected and fixed in 2.5% glutaraldehyde overnight at 4 °C. Subsequently, the samples were thoroughly washed with PBS four times at 10-minute intervals to ensure the complete removal of the fixative. Fixed embryos were treated with osmium tetroxide for 1 h to enhance the contrast and stability of cellular structures under the electron beam. The samples were then washed with ddH2O five times at 10 min intervals to remove any residual osmium tetroxide. Dehydrate the embryos in an ethanol series for 5–10 min each: 50%, 70%, 85%, and three times in 100%. Transferred the embryos in ethanol to baskets for critical point drying (CPD) in a critical point dryer machine (Quorum K850 critical point dryer). After that, the samples were mounted on sample stubs and sputter-coated with gold using a Quorum Q150R S sputter coater. Finally, the samples were imaged using the Zeiss GeminiSEM 300 scanning electron microscope.
The ST-MAGIC atlas
Pre-processing of Geo-seq and 10x scMultiome: The Geo-seq data utilized in this study were obtained from our previously published datasets, specifically including E6.5, E7.0, and E7.5 from GSE120963, and E6.75, E7.25, and E7.5 (left-right resolved) from GSE171588. The five stages of 10x scMultiome data were newly generated in this study. For the Geo-seq data, the quantification of the FASTQ files was performed using RSEM (v 1.3.3), leveraging annotations from the CellRanger GTF file. The normalized TPM values were processed using the ‘NormalizeData’ function in the Seurat (v 4.1.1) package and were then used for further analytical procedures. For the scMultiome data, the normalized snRNA-seq and snATAC-seq data were processed using Seurat and Signac (v 1.6.0) and were subsequently employed for further analytical procedures.
Alignment of 10x scMultiome to Geo-seq: The alignment of snRNA-seq data to Geo-spatial spots was achieved using Tangram (v 1.0.4), with the precise spatial location of each cell determined from the mapping matrix with the highest probability. The gene expression of each reconstructed spot was obtained by averaging the gene expression levels of the cells mapped to the indicated spot, and the gene expression percentage of a spot represents the proportion of cells within the indicated spot that express the gene (gene reads > 0). For snATAC-seq data mapping, the spatial location of each cell from snRNA-seq is shared. The peak accessibility of each reconstructed spot was obtained by averaging the peak accessibility level of the cells mapped to the indicated spot, and the peak accessibility percentage of a spot represents the ratio of cells within the indicated spot with an open peak (peak reads > 0). The cell type distribution was obtained using the tg.project_cell_annotations function within the Tangram toolkit, and the cell type percentage of a spot represents the ratio of cells of that type within the spot.
In the application of Tangram, the training genes are specified as the zipcode genes relevant to each stage, and the mapping strategy is customized for individual cells. For the E6.5 and E6.75 embryos, the top 100 cell-type marker genes identified by the COSG (v 0.9.0) package63 were designated as zipcode genes. For the E7.0, E7.25, and E7.5 embryos, the PC loading genes calculated by the Seurat package (E7.0, top 500 highest and 500 lowest genes of PC1-10; E7.25, top 50 highest and 50 lowest genes of PC1-10; E7.5, top 500 highest and 500 lowest genes of PC1-5) were designated as zipcode genes. The zipcode list has been provided in Supplementary Data 2.
Since Geo-seq primarily captures the proximal to distal embryonic regions of the gastrula, cell types from the scMultiome data that is not present within these zones were excluded from the analysis. Specifically, the following cell types were filtered out: ExE endoderm, ExE ectoderm, Haematoendothelial progenitors, Blood progenitors 1, and Blood progenitors 2.
Smooth of corn plot: The smoothing of mapped features in ST-MAGIC is achieved through Gaussian weighting of neighboring spots, including the mapped feature smooth denoted as F and the percent smooth denoted as P, defined as:
For the selection of neighboring spots, ST-MAGIC provides three types of input: column neighboring spots along the proximal-distal axis by default, row neighboring spots along the anterior-posterior axis, and grid neighboring spots on the corn plot.
Customized query of corn plot: The customized query is designed to recognize user-defined consistent patterns, whether they are genes, peaks, or cell types. The spatial pattern of interest was defined in binary, with the value of the corresponding spot set to 1 and the value of all other spots set to 0. Similar omics data matching this binary pattern were selected using cosine similarity, as implemented in the COSG package. For example, ST-MAGIC detects the specified peaks in customized RPM and LPM regions (distal peaks in the left and right proximal mesoderm regions shown in Fig. 2g, h). This process will facilitate uncovering multi-omic patterns underlying coordinated biological processes or spatially regulated activities.
The ST-MAGIC (+) atlas
Calculation of enrichment score: Building on the single gene or peak analysis capabilities of ST-MAGIC, ST-MAGIC (+) extends these features by enabling the spatial monitoring of gene sets and peak sets from SCEINIC + or public resources through enrichment analysis by the AUCell (v 1.18.1) package. The enrichment of gene sets and peak sets of each spot was obtained by averaging the AUC score of the cells mapped to that spot, and the enrichment percentage of a spot represents the average ratio of expressed genes or accessible peaks within that spot.
Spatial signaling responsiveness pattern: The binding of signaling molecules to multiple genomic regions highlights the complexity and interconnectedness of gene regulatory networks. It highlights how single factors can coordinate with diverse downstream targets to orchestrate cellular responses and developmental programs. ST-MAGIC (+) enables the depiction of the spatial activities of signaling responsiveness, such as WNT and Nodal (Fig. 4), using data from public ChIP-seq datasets. Before calculating the AUC score, the peaks from public datasets must first be aligned to those identified in our own dataset using the GenomicRanges (v 1.48.0) package to ensure a consistent genomic context for comparative analysis.
BioCRE model
The construction of BioCRE: Unlike published tools that detect CREs using only a single peak or a single gene, BioCRE employs a systematic approach to calculate these linkages. BioCRE harnesses a bi-orientation regression model leveraging multi-omics data decoding GRN at the chromosomal level to identify potential CREs.
To formulate the linear optimization procedure using paired snRNA-seq and snATAC-seq data, let \({G}^{\theta }\in {{\mathbb{R}}}^{g\times n}\) denotes snRNA-seq data profiling g genes across n cells on a given chromosome θ, let \({P}^{\theta }\in {{\mathbb{R}}}^{p\times n}\) denotes snATAC-seq data with p peaks for the same set of n cells on the chromosome θ. We assumed the linkage matrix \({L}^{\theta }\in {{\mathbb{R}}}^{g\times p}\) is the cis-regulation between genes and peaks on chromosome θ. Our goal is to learn the linkage matrix L that captures cis-regulation under two conditions: (1) the predicted snRNA-seq matrix \(\hat{{G}^{\theta }}\), generated using the linkage matrix \({L}^{\theta }\) and snATAC-seq matrix \({P}^{\theta }\), should be similar to the real snRNA matrix \({G}^{\theta }\); (2) the predicted snATAC-seq matrix \(\hat{{P}^{\theta }}\), generated using the linkage matrix \({L}^{\theta }\) and snRNA matrix \({G}^{\theta }\), should be similar to the real snATAC-seq matrix \({P}^{\theta }\). We minimize the following cost function as:
The cost function consists of three parts:
-
1)
The expected loss between the real snRNA-seq matrix \({G}^{\theta }\) and the predicted snRNA-seq matrix \(\hat{{G}^{\theta }}\):
$${\mathbb{E}}\left[{({{G}^{\theta }}-\hat{{G}^{\theta }})}^{2}\right]=\mathop{\sum }\limits_{i=1}^{g}\mathop{\sum }\limits_{k=1}^{n}{({G}_{i,k}^{\theta }-\hat{{G}_{i,k}^{\theta }})}^{2}=\mathop{\sum }\limits_{i=1}^{g}\mathop{\sum }\limits_{k=1}^{n}{({G}_{i,k}^{\theta }-\mathop{\sum }\limits_{j=1}^{p}{L}_{i,j}^{\theta }\times {P}_{j,k}^{\theta })}^{2}$$Where \({G}_{i,k}^{\theta }\) is the expression level of gene i in cell k on chromosome θ, \({P}_{j,k}^{\theta }\) is the chromatin accessibility of peak j in cell k on chromosome θ, the \({L}_{i,j}^{\theta }\) is the strength of linkage between gene i and peak j on chromosome θ.
-
2)
The expected loss between the real snATAC-seq matrix \({P}^{\theta }\) and the predicted snATAC-seq matrix \(\hat{{P}^{\theta }}\):
$${\mathbb{E}}\left[{({{P}^{\theta }}-\hat{{P}^{\theta }})}^{2}\right]=\mathop{\sum }\limits_{j=1}^{p}\mathop{\sum }\limits_{k=1}^{n}{({{P}_{j,k}^{\theta }}-\hat{{P}_{j,k}^{\theta }})}^{2}=\mathop{\sum }\limits_{j=1}^{p}\mathop{\sum }\limits_{k=1}^{n}{\left({{P}_{j,k}^{\theta }}-\mathop{\sum }\limits_{i=1}^{g}{L}_{i,j}^{\theta }\times {G}_{i,k}^{\theta }\right)}^{2}$$ -
3)
The L2 regularization loss of the linkage matrix \({L}^{\theta }\) with hyperparameter \({\lambda }^{\theta }\):
Under this model, we systematically considered all combinations of genes and peaks within the same chromosome, effectively simulating the complexity of GRN.
To mitigate the risk of gradient vanishing in a model with a large number of parameters, ‘He’ initialization was employed for the weight matrices:
Where p is the number of peaks. The L-BFGS algorithm is employed to achieve the minimization of the cost function, the training process is configured for 500 epochs with a learning rate of 0.5.
After the optimization of L, the extraction of significant linkages is required. Firstly, we computed bi-orientation p-values, which involved assessing the significance of a given CRE in relation to all CREs linked to the same gene by z-score, followed by evaluating its significance among all CREs associated with the same peak, the p-values were adjusted using the Bonferroni correction method applied separately to each set of comparisons. Then, the two p-values are combined into a single value through Pearson’s method in the ‘SciPy’ package:
Finally, the CREs are constrained to those within a 1 mega base range, and only those with a combined p-value less than 10-7 are reserved.
Benchmark analysis of BioCRE
Performance on gastrula multi-omic dataset: In the absence of a genome-wide gold standard for gene-to-peak linkages in the mouse gastrula, we used the Jensen-Shannon Similarity score, which is calculated based on the Jensen-Shannon Divergence index, to evaluate the concordance between genes and peaks in linkage for BioCRE, compared to linkages established using default parameters in ArchR and Signac. The JSS score of each linkage was calculated using the normalized gene expression \({G}_{i}\) from snRNA data and the peak accessibility \({P}_{j}\) from snATAC data across n cells.
The JSS score was also used to evaluate the robustness of the ST-MAGIC atlas, which maps snRNA-seq data to Geo-seq data. Specifically, it assessed the concordance between the reconstructed gene expression in spot from the snRNA-seq and the raw gene expression in spot from the Geo-seq data.
In addition, we computed the silhouette score to evaluate the effectiveness of cell type-specific gene-linked peaks in clustering cells in UMAP visualizations. Using pre-annotated cell types as clustering labels, the silhouette score was calculated based on the 2nd to 10th latent semantic indexing components via the ‘cluster’ package. A higher silhouette score indicates that cells of the same type are more closely grouped.
To systematically evaluate the stability and robustness of BioCRE in predicting CREs across varying sequencing depths, comprehensive down-sampling experiments were conducted. Initially, paired single-cell transcriptomic (snRNA-seq) and single-cell chromatin accessibility (snATAC-seq) datasets were jointly down-sampled to 20, 40, 60, 80, and 100% of their original sequencing depth. This allowed for an assessment of prediction consistency across methods under increasingly sparse data conditions. Subsequently, two complementary experimental paradigms were implemented: first, snATAC-seq data were maintained at full depth while the sequencing depth of snRNA-seq was progressively reduced; second, snRNA-seq data were preserved in their entirety while the depth of snATAC-seq data was incrementally decreased. In all experimental conditions, the stability of predicted regulatory linkages was quantitatively evaluated based on the proportion of shared linkages identified across different sequencing depths.
Performance on public multi-omic datasets with corresponding promoter capture Hi-C data: To systematically compare the predictive performance of BioCRE, we conducted additional evaluations using publicly available data. We analyzed published multi-omics datasets (PBMC: https://www.10xgenomics.com/cn/datasets/pbmc-from-a-healthy-donor-granulocytes-removed-through-cell-sorting-10-k-1-standard-1-0-0 and GM12878: GSE166797), with corresponding promoter capture Hi-C data (PBMC: EGAS00001001911 and GM12878: https://zenodo.org/records/3255048) as reference datasets for validating predicted linkages.
Standardized genomic scope of predicted linkages across all three tools were restricted to ±500 kb upstream and downstream of the gene body. Notably, although the total numbers of predicted linkages are theoretically consistent across methods, minor discrepancies may arise due to each tool’s internal preprocessing steps—such as gene and peak filtering—which may exclude certain elements prior to linkage inference. The filtered geneset and peakset shared by all three tools were used for further linkage prediction.
For comparison between the public multi-omic datasets and the reference dataset, any predicted linkages overlapping the reference set was considered as positives, while non-overlapping linkages were treated as negatives. Due to the distinct features for different data modules between the predicted linkages and the reference dataset (PCHiC), the overlap is inherently limited, leading to a sparse set of positive linkages. Here, to ensure balanced and robust evaluation, we randomly sampled negative linkages ten times at each positive-to-negative ratio. Specifically, for the PBMC dataset, we evaluated the AUPRC through ‘sklearn.metrics’ package under varying positive-to-negative ratios: 1:1, 1:2, 1:3, 1:4, and 1:5. For GM12878, we used ratios of 1:1 and 1:2, as the larger number of positive linkages—relative to PBMC—limits the feasible extent of negative down-sampling. Besides, tool-specific metrics to calculate AUPRC were applied: the combined p-value for BioCRE, the p-value for Signac, and the FDR for ArchR.
Multi-omics data pre-processing
Raw FASTQ files of snRNA-seq and snATAC-seq were processed by Cell Ranger ARC (v2.0.1) with default mapping parameters. Reads were mapped to the mouse reference genome (mm10) and quantified with gene annotation (GRCm38.98). The quality control of snRNA-seq and snATAC-seq data was performed separately, retaining only high-quality cells that passed both criteria for subsequent analyses. For snRNA-seq data, the following quality control criteria were applied: (1) cells that had fewer than 1000 expressed genes, or over 10% of UMIs derived from mitochondrial genome were removed; (2) cells identified as doublet (DoubletFinder, v2.0.3) were removed; (3) clusters generated by Seurat using default parameters were removed if AY036118 and Gm42418 genes were highly expressed. For snATAC-seq data, cells were required to have a minimum of 3000 fragments, a minimum fraction of fragments in peaks of 0.15, a minimum of TSS enrichment of 9, a maximum of blacklist ratio of 0.05 and a maximum of nucleosomal signal strength of 4.
snRNA-seq processing
After converting the filtered snRNA-seq count matrix to a Seurat object, the standard procedures were performed using the Seurat package. For each stage (E6.5, E6.75, E7.0, E7.25, and E7.5), the following analysis steps were applied: (1) the count matrix was log-normalized for each cell with a scaling factor of 10,000; (2) 2000 variable genes were selected by ‘vst’ method; (3) the ‘nFeature_RNA’ was regressed out during the scaling process. For the combined dataset, the ‘stage’, ‘nFeature_RNA’, and ‘percent.mt’ were regressed out; (4) The scaled variable genes were projected into a low-dimensional space using principal component analysis; (5) The shared nearest neighbor(SNN) graph was built based on the distance of PC space (nPCs = 50); (6) The Louvain algorithm was used to identify clusters based on SNN graph (resolution = 1); (7) The UMAP embedding was calculated using ‘uwot’ with the top 10 PCs.
snATAC-seq processing
As the precise boundaries of peaks differed across embryonic stages, a common set of peaks (20 bp < length of peak < 10,000 bp) across all stages was generated using the ‘reduce’ function in GenomicRanges (v1.48.0) package. The fragment file was then remapped to these common peaks to get the chromatin accessibility matrix. Then, the remapped chromatin accessibility matrix and correspondent fragment file were processed using the Signac package. The matrix was normalized by Term Frequency Inverse Document Frequency, and all peaks were used for dimension reduction using Singular Value Decomposition. The UMAP embedding was computed utilizing dimensions from 2 to 10, with min.dist set to 0.2.
Clustering and cell type annotation
The annotation of multi-omics is based on snRNA-seq data, while snATAC-seq annotation is facilitated via the consistent cellular barcodes between the transcriptome and chromatin accessibility data, ensuring harmonized identification across datasets. Given that cell type diversification during gastrulation, the cell types of snRNA-seq data were annotated in a stage-by-stage manner. Initially, the snRNA-seq data is clustered through the standard Seurat pipeline (vars.to.regress = ‘nFeature_RNA’, k.param = 20, dims = 1:10, resolution = 1) with a preliminary annotation to the three primary groups: Embryonic, ExE endoderm, and ExE ectoderm, then cells within the Embryonic group were subjected to an iterative clustering process to achieve more finely detailed cell types. The iterative clustering also employs the Seurat pipeline (vars.to.regress = c(‘nFeature_RNA’, ‘percent.mt’), k.param = 30, dims = 1:30, resolution = 0.9, these parameters remain constant throughout the iterative process) to conduct a clustering process independently on each cluster, with the stipulation that clusters cannot be further subdivided or any cluster containing fewer than 50 cells will not be subjected to additional clustering. Following several iterations, the definitive clusters are established. The integration of cluster-specific genes (defined by the COSG algorithm63) and marker genes from the published gastrulation transcriptome atlas3,6 were mainly used to annotate the clusters. Ambiguous clusters were clarified by correlating them with the closest matching cell type found in existing public datasets (GSE87038). The integration of the public dataset with our snRNA-seq data is facilitated through the use of Canonical Correlation Analysis (CCA) within the Seurat package. The Pearson correlation between the pseudobulk transcriptomic profiles of our iterative clusters and those of cell types in the public dataset is employed to refine annotations.
The similarity between snRNA-seq data and published datasets
To verify the reliability of the annotation, we compared it with the public dataset (GSE87038) at three levels: the proportion of cell types, the correlation of marker genes, and the correlation of the whole transcriptome. First, the shared stages and corresponding cell type data were extracted from the public dataset. Then, the CCA method in Seurat was used to integrate these two datasets. The proportion of each cell type and the expression of marker genes (Sox2, Mesp1, Sox17, and Hnf4a) were statistically compared between the two datasets, along with the pseudo-bulk expression of 2000 high-variance genes across cell types.
Cellular trajectory analysis with TOME
The cellular trajectory spanning from E3.5 to E7.5 was reconstructed based on scRNA/snRNA-seq data using the TOME. The processed dataset covering the developmental stages from E3.5 to E6.25 was made available by the TOME. For the subsequent stages from E6.5 to E7.5, we employed our snRNA-seq data. The identification of the nearest ancestral node for each subsequent stage node is repeated for 500 times based on 50 nearest neighbors. Only those edges with a probability greater than 0.2 will be preserved. Given that the annotated cell types do not align perfectly with those in the pre-stage dataset, we have designated “Embryonic visceral endoderm” as “visceral endoderm,” and “Extraembryonic visceral endoderm” as “ExE endoderm” within our dataset for consistency.
WNN embedding for embryonic cells
The RNA and ATAC modality from embryonic cells, mapping to Geo-seq data, across five development stages, were integrated to construct a weighted nearest neighbor graph by the ‘FindMultiModalNeighbors’ method in the Seurat package. In the snRNA-seq data process, the ‘stage’, ‘nFeature_RNA’, and ‘percent.mt’ were regressed out during normalizing. In snATAC-seq data process, the ‘min.cutoff’ was set to ‘q0’ in finding top features.
Inference of the TF regulatory network with SCENIC +
The pre-processed multi-omic RNA and ATAC matrices, encompassing the full dataset and axial mesendoderm lineages, are utilized as inputs for SCENIC + (v 0.1.dev452), facilitating the construction of the TF regulatory network. SCENIC+ was executed with the majority of its settings at their default values, with adjustments made only to parameters specific to the dataset. For the full dataset, the peak matrix is re-generated utilizing stage-specific cell type pseudobulk data through MACS2 (v 2.2.7.1). During the cisTopic (v 1.0.2) module, an optimized set of 16 topics is selected for further analysis. Peaks within a 2 kb~ 500 kb range upstream or downstream of genes are incorporated into the SCENIC + modeling process. eRegulons featuring positive region-to-gene associations, along with a correlation exceeding 0.15 between the TF cistrome and region-based activity, were retained, resulting in a set of 302 high-quality eRegulons. For axial mesendoderm lineages, stage-specific peaks for each cell type are also generated. In cisTopic, the optimized number of topics is set at 16. The range for peaks is consistent with that of the full datasets. eRegulons are retained based on a correlation threshold of 0.15 between the TF cistrome and region-based activity, yielding a total of 118 high-quality eRegulons.
Differential ChIP-seq peaks of the NODAL and WNT signaling
For each ChIP-seq dataset, raw sequencing data was downloaded from the relative resource (NODAL signaling: GSE70486; WNT signaling: GSE43565, GSE162774) and then mapped to the mouse genome (mm10). Peak calling was performed against the corresponding input sample using MACS with the parameters “--shiftsize = 100 --nomodel --keep-dup = all”. Subsequently, we merged all resulting peaks of samples into a consensus list of genomic regions and counted the reads within those regions using MAnorm2_utils (v 1.0.0) with the parameters “--min-peak-gap = 150 --typical-bin-size = 2000 --shiftsize = 100 --filter = blacklist”. The blacklist regions of mm10 were obtained from Amemiya et al. 64. Then, MAnorm2 (v 1.2.2) were used to perform differential peak analysis between samples with default parameters (refer to https://cran.r-project.org/web/packages/MAnorm2/vignettes/MAnorm2_vignette.html). The differential peaks were identified using the cutoffs adjusted p-value < 0.01 and fold change > 2.
Axial mesendoderm lineage cells
The selection of axial mesendoderm lineage cells was guided by trajectory analysis depicted in Fig. 1c. This lineage originates from the primitive streak cells at E6.5, and subsequently progresses through the Anterior primitive streak at E6.75 and E7.0. It then developed into Node precursors and Anterior mesendoderm precursors at E7.25. The lineage culminates at E7.5 with Node and Anterior mesendoderm. To ensure more reliable lineage-related cells, we retain only those cells from ancestral stages that are found among the nearest neighbors as calculated by TOME, except for cells in the E6.5 Primitive streak.
Driver genes of axial mesendoderm lineage with CellRank
The sorted BAM files derived from CellRanger were processed using Velocyto (v 0.17.17) to compute RNA velocities for individual cells. The mask GTF file was obtained from the UCSC database, and the genome annotation file originates from CellRanger. This procedure enables detailed analysis of transcriptional dynamics at the single-cell level by leveraging the spliced and unspliced transcript information contained within the BAM files. The velocity matrix specific to the axial mesendoderm lineage cells has been extracted from the loom file utilizing the SeuratWrappers (v 0.3.0) R package. Subsequently, the velocity matrix was employed for velocity estimation through the application of the ‘dynamical’ model in scVelo (v 0.2.5). The cell-cell transition matrix was calculated employing the ‘cellrank.tl.transition_matrix’ function in CellRank (v 1.5.1). Terminal states were set as Node and Anterior mesendoderm manually. Following this, the GPCCA estimator was utilized to compute both the fate probabilities and driver genes. The top 1000 smoothed driver genes, which exhibited the highest correlation with pseudotime, were selected as the key drivers. The driver genes identified in the BioCRE linkage analysis of the axial mesendoderm lineage and their linked peaks were selected to plot the heatmap. The pseudotime for the lineage was computed utilizing the ‘scanpy.tl.dpt’ function (scanpy, v 1.9.1), incorporating the eRegulon RNA and ATAC AUC matrices obtained from the SCENIC + .
KO-omics data processing
The preprocessing for KO-omics data (WT, NotoKO, Pou6f1 KO embryos at the E7.5 stage) is consistent with those previously mentioned. The annotation method for WT snRNA-seq data remains consistent with the previous approach, whereas for Noto KO and Pou6f1 KO samples, snRNA-seq annotation is performed using the ‘MapQuery’ in Seurat, with annotated WT snRNA-seq data serving as a reference. The annotation for snATAC-seq data was aligned with that of snRNA-seq by leveraging shared cell barcodes.
Webserver implementation
The 10 x scMultiome data and reconstructed Geo-seq data are available through the ST-MAGIC webserver, which was implemented using Shiny apps from R. This web service provides three main functionalities: (1) Transcriptomics in snRNA-seq data, which illustrates gene expression on UMAP and cell type grouping; (2) Epigenomics in snATAC-seq data, which shows peak accessibility, linkage, and motif information; (3) Spatialomics of multiome data, which includes gene, peak, cell type, gene set, and peak set enrichment visualized on a corn plot. ST-MAGIC enables users to explore spatial and temporal patterns in a user-friendly manner. For transcriptomics, profiled genes can be searched for their expression patterns over space and time. For epigenomics, profiled peaks and motifs can be used to investigate spatial and temporal accessibility patterns. In addition, ST-MAGIC provides a selection list of gene sets or peak sets from SCENIC+ to TF enrichment analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data generated in this study have been deposited in the Genome Sequence Archive of China National Center for Bioinformation with the accession number: CRA020056. All other data are available from the corresponding authors upon request. Source data are provided in this paper.
Code availability
Our resources can be explored via an interactive web portal: https://stmagic.miracle.ac.cn:29909/. The code for BioCRE has been deposited in https://github.com/SuoLab-GZLab/BioCRE and also linked to Zenodo (https://doi.org/10.5281/zenodo.17907133).
References
Tam, P. P. & Behringer, R. R. Mouse gastrulation: the formation of a mammalian body plan. Mech. Dev. 68, 3–25 (1997).
Arias, A. M., Dias, A., Wehmeyer, A. E., Arnold, S. J. & Fiuza, U. M. A modular organization of mammalian gastrulation and the Spemann-Mangold organizer. Cells Dev. 184, 204031 (2025).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Yang, X. et al. Distinct enhancer signatures in the mouse gastrula delineate progressive cell fate continuum during embryo development. Cell Res. 29, 911–926 (2019).
Jiang, S. et al. Single-cell chromatin accessibility and transcriptome atlas of mouse embryos. Cell Rep. 42, 112210 (2023).
Wang, R. et al. Time space and single-cell resolved tissue lineage trajectories and laterality of body plan at gastrulation. Nat. Commun. 14, 5675 (2023).
Balmer, S., Nowotschin, S. & Hadjantonakis, A. K. Notochord morphogenesis in mice: Current understanding & open questions. Dev. Dyn. 245, 547–557 (2016).
Yamanaka, Y., Tamplin, O. J., Beckers, A., Gossler, A. & Rossant, J. Live imaging and genetic analysis of mouse notochord formation reveals regional morphogenetic mechanisms. Dev. Cell 13, 884–896 (2007).
Tam, P. P. L. & Masamsetti, P. Functional attributes of the anterior mesendoderm in patterning the anterior neural structures during head formation in the mouse. 184, Cells Dev. 203999 (2025).
Sulik, K. et al. Morphogenesis of the murine node and notochordal plate. Dev. Dyn. 201, 260–278 (1994).
Stemple, D. L. Structure and function of the notochord: an essential organ for chordate development. Development 132, 2503–2512 (2005).
Masamsetti, V. P. et al. Lineage contribution of the mesendoderm progenitors in the gastrulating mouse embryo. Dev. Cell 60, 1991–2006 (2025).
Davidson, B. P., Kinder, S. J., Steiner, K., Schoenwolf, G. C. & Tam, P. P. Impact of node ablation on the morphogenesis of the body axis and the lateral asymmetry of the mouse embryo during early organogenesis. Dev. Biol. 211, 11–26 (1999).
Cleaver, O. & Krieg, P. A. Notochord patterning of the endoderm. Dev. Biol. 234, 1–12 (2001).
Cheng, C. et al. Yap controls notochord formation and neural tube patterning by integrating mechanotransduction with FoxA2 and Shh expression. Sci. Adv. 9, eadf6927 (2023).
Kahane, N. & Kalcheim, C. Neural tube development depends on notochord-derived sonic hedgehog released into the sclerotome. Development 147, dev183996 (2020).
Tsukui, T. et al. Multiple left-right asymmetry defects in Shh(-/-) mutant mice unveil a convergence of the shh and retinoic acid pathways in the control of Lefty-1. Proc. Natl. Acad. Sci. USA 96, 11376–11381 (1999).
Qiu, C. et al. Systematic reconstruction of cellular trajectories across mouse embryogenesis. Nat. Genet. 54, 328–341 (2022).
Peng, G. et al. Molecular architecture of lineage allocation and tissue organization in early mouse embryo. Nature 572, 528–532 (2019).
Haraguchi, S. et al. Transcriptional regulation of Mesp1 and Mesp2 genes: differential usage of enhancers during development. Mech. Dev. 108, 59–69 (2001).
Rao, A., Barkley, D., Franca, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
You, Y. et al. Systematic comparison of sequencing-based spatial transcriptomic methods. Nat. Methods 21, 1743–1754 (2024).
Qu, F. et al. Three-dimensional molecular architecture of mouse organogenesis. Nat. Commun. 14, 4599 (2023).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Jiang, F. et al. Simultaneous profiling of spatial gene expression and chromatin accessibility during mouse brain development. Nat. Methods 20, 1048–1057 (2023).
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Xie, P. et al. Digital reconstruction of full embryos during early mouse organogenesis. Cell 188, 4754–4772 (2025).
Perea-Gomez, A. et al. Otx2 is required for visceral endoderm movement and for the restriction of posterior signals in the epiblast of the mouse embryo. Development 128, 753–765 (2001).
Saijoh, Y. et al. Left–Right Asymmetric Expression of lefty2 and nodal Is Induced by a Signaling Pathway that Includes the Transcription Factor FAST2. Mol. Cell 5, 35–47 (2000).
Levine, M. & Davidson, E. H. Gene regulatory networks for development. PNAS 102, 4936–4942 (2005).
Bravo Gonzalez-Blas, C. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023).
Steimle, J. D. et al. ETV2 primes hematoendothelial gene enhancers prior to hematoendothelial fate commitment. Cell Rep. 42, 112665 (2023).
Sheng, G., Martinez Arias, A. & Sutherland, A. The primitive streak and cellular principles of building an amniote body through gastrulation. Science 374, abg1727 (2021).
Robles-Garcia, M. et al. In vitro modelling of anterior primitive streak patterning with human pluripotent stem cells identifies the path to notochord progenitors. Development 151, dev202983 (2024).
Chen, Y. et al. Gastrula-premarked posterior enhancer primes posterior tissue development through cross-talk with TGF-beta signaling pathway. Adv. Sci. 36, e00895 (2025).
Morgani, S. M. & Hadjantonakis, A. K. Signaling regulation during gastrulation: Insights from mouse embryos and in vitro systems. Curr. Top. Dev. Biol. 137, 391–431 (2020).
Zhang, X., Peterson, K. A., Liu, X. S., McMahon, A. P. & Ohba, S. Gene regulatory networks mediating canonical Wnt signal-directed control of pluripotency and differentiation in embryo stem cells. Stem Cells 31, 2667–2679 (2013).
Blassberg, R. et al. Sox2 levels regulate the chromatin occupancy of WNT mediators in epiblast progenitors responsible for vertebrate body formation. Nat. Cell Biol. 24, 633–644 (2022).
Pera, M. F. & Rossant, J. The exploration of pluripotency space: Charting cell state transitions in peri-implantation development. Cell Stem Cell 28, 1896–1906 (2021).
Shen, M. M. Nodal signaling: developmental roles and regulation. Development 134, 1023–1034 (2007).
Wang, Q. et al. The p53 Family Coordinates Wnt and Nodal Inputs in Mesendodermal Differentiation of Embryonic Stem Cells. Cell Stem Cell 20, 70–86 (2017).
Lee, J. D. & Anderson, K. V. Morphogenesis of the node and notochord: the cellular basis for the establishment and maintenance of left-right asymmetry in the mouse. Dev. Dyn. 237, 3464–3476 (2008).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Bardot, E. S. & Hadjantonakis, A. K. Mouse gastrulation: Coordination of tissue patterning, specification and diversification of cell fate. Mech. Dev. 163, 103617 (2020).
Guo, F. et al. Single-cell multi-omics sequencing of mouse early embryos and embryonic stem cells. Cell Res. 27, 967–988 (2017).
Calderon, D. et al. J. The continuum of Drosophila embryonic development at single-cell resolution. Science 377, eabn5800 (2022).
Shu, M. et al. Single-cell chromatin accessibility identifies enhancer networks driving gene expression during spinal cord development in mouse. Dev. Cell 57, 2761–2775 (2022).
Argelaguet, R. et al. Multi-omics profiling of mouse gastrulation at single-cell resolution. Nature 576, 487–491 (2019).
Kiessling, P. & Kuppe, C. Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases. Genome Med. 16, 14 (2024).
Vandereyken, K., Sifrim, A., Thienpont, B. & Voet, T. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Tam, P. P. & Beddington, R. S. The formation of mesodermal tissues in the mouse embryo during gastrulation and early organogenesis. Development 99, 109–126 (1987).
Kinder, S. J. et al. The organizer of the mouse gastrula is composed of a dynamic population of progenitor cells for the axial mesoderm. Development 128, 3623–3634 (2001).
Camus, A. et al. The morphogenetic role of midline mesendoderm and ectoderm in the development of the forebrain and the midbrain of the mouse embryo. Development 127, 1799–1813 (2000).
Arnold, S. J., Hofmann, U. K., Bikoff, E. K. & Robertson, E. J. Pivotal roles for eomesodermin during axis formation, epithelium-to-mesenchyme transition and endoderm specification in the mouse. Development 135, 501–511 (2008).
Weinstein, D. C. et al. The winged-helix transcription factor HNF-3 beta is required for notochord development in the mouse embryo. Cell 78, 575–588 (1994).
Abdelkhalek, H. B. et al. The mouse homeobox gene Not is required for caudal notochord development and affected by the truncate mutation. Genes Dev. 18, 1725–1736 (2004).
Talbot, W. S. et al. A homeobox gene essential for zebrafish notochord development. Nature 378, 150–157 (1995).
Saunders, L. M. et al. Embryo-scale reverse genetics at single-cell resolution. Nature 623, 782–791 (2023).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Yang, X., Chen, Y., Song, L., Zhang, T. & Jing, N. Wholemount in situ Hybridization for Spatial-temporal Visualization of Gene Expression in Early Post-implantation Mouse Embryos. Bio. Protoc. 11, e4229 (2021).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Dai, M., Pei, X. & Wang, X. J. Accurate and fast cell marker gene identification with COSG. Brief Bioinform. 23, bbab579 (2022).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Tosic, J. et al. Eomes and Brachyury control pluripotency exit and germ-layer segregation by changing the chromatin state. Nat. Cell Biol. 21, 1518–1531 (2019).
Schüle, K. M. et al. Eomes restricts Brachyury functions at the onset of mouse gastrulation. Dev. Cell 58, 1626–1642 (2023).
Cernilogar, F. M. et al. Pre-marked chromatin and transcription factor co-binding shape the pioneering activity of Foxa2. Nucleic Acids Res. 47, 9069–9086 (2019).
Acknowledgements
We are grateful to Prof. Chi-Chung Hui for his constructive suggestions. We thank the imaging and animal care core facilities at Guangzhou National Laboratory. We also would like to thank Jiaheng Chen and Heying Li from the core facility at Guangzhou Institutes of Biomedicine and Health, CAS for the help of scan electron microscopy. This work was supported in part by the National Key Basic Research and Development Program of China (2025YFA1804200 to X.Y., 2025YFE0200600 to X.Y. and N.J., 2018YFA0800100 to X.Y., 2019YFA0801402 to X.Y.), the Major Project of Guangzhou National Laboratory (GZNL2023A02005 to X.Y. and N.J., GZNL2023A02007 to S.S., GZNL2023A03005 to S.S.), the National Natural Science Foundation of China (32130030 to N.J., 32470866 to X.Y., 31900454 to X.Y., 32370972 to S.S.), the Union Project from Guangzhou National Laboratory and State Key Laboratory of Respiratory Disease, Guangzhou Medical University (GZNL2024B01007 to N.J., GZNL2024B01004 to S.S.), the Guangdong Basic and Applied Basic Research Foundation (2024B1515020052 to S.S., 2023A1515011783 to S.S.).
Author information
Authors and Affiliations
Contributions
X.Y., S.S., and N.J. conceived the study. X.Y., N.J. designed the experiments, X.Y., P.S., Y.C., Yun Yang, Z.L., and P.M. collected the sequencing data and performed the experiments, M.W. performed animal husbandry. B.X. and S.S. designed the bioinformatic model and pipeline for data analyses and visualization. B.X., C.L., F.T., Yumeng Yang, and R.S. performed the bioinformatic analyses. X.Y., B.X., P.T., S.S., and N.J. wrote the manuscript with the help of all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yang Liu, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, X., Xie, B., Shen, P. et al. Integrated multi-omic atlas reveals the hierarchy of spatiotemporal regulatory networks of mouse gastrulation. Nat Commun 17, 1572 (2026). https://doi.org/10.1038/s41467-026-68291-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68291-w