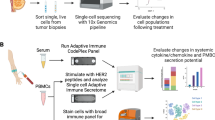
Abstract
Self-amplifying mRNA (saRNA) vectors hold promise for the sustained expression of mRNA vaccines in vivo. However, their inherently high immunogenicity and low-fidelity replication—stemming from the RNA viral genome’s replication mechanisms—limit their efficacy as replacements or adjuncts to protein therapies. Here we report an engineered viral protein genome-linked (VPg) saRNA vector derived from a Norovirus replicon, designed for rapid loading of therapeutic protein mRNAs in vitro. The engineered VPg saRNA is adapted for a range of therapeutic scenarios, including treatment of tumor-associated cachexia under conditions of translational restriction in cap-dependent metabolism, precise encoding of oncolytic mRNAs in vivo to achieve complex functionality, and therapy for graft-versus-host disease in highly auto-immune environments. VPg saRNA addresses key limitations of linear mRNA and conventional saRNA therapies, broadening the potential applications of mRNA-based treatments.
Similar content being viewed by others
Introduction
The demonstrated efficacy of mRNA vaccines against rapidly mutating epidemic RNA virus infections1,2 has established cell-free in vitro transcribed (IVT) mRNA technology as a cornerstone in the development of nucleic acid-based therapeutics3. This technology has expanded into applications for the treatment, replacement, or supplementation of protein therapies4. Notably, mRNA avoids nuclear entry, thereby mitigating the risk of altering the host cell’s genetic material5. However, exogenous mRNA is vulnerable to recognition by cytoplasmic nucleic acid receptors, which can trigger the host’s antiviral response, leading to mRNA degradation and significantly diminishing protein expression6. This vulnerability poses minimal challenges for mRNA vaccines, as even small quantities can elicit robust immune responses, generating sufficient antigen levels7. In contrast, preclinical research focused on encoding active enzymes, cytokines, and therapeutic proteins often necessitates up to a 1000-fold increase in active protein levels to achieve therapeutic thresholds8. This demand has spurred the development of various strategies to enhance protein expression from equivalent mRNA doses, including self-amplifying mRNA (saRNA)9, circular mRNA (circRNA)10, and trans-amplifying mRNA11 constructs.
SaRNA is an engineered mRNA structure that incorporates single-stranded RNA (ssRNA) viral replicon elements, enabling it to replicate itself and achieve long-term, efficient expression of the encoded open reading frame (ORF) sequence. The most extensively studied and utilized saRNA derives from the genomic RNA (gRNA) replicon elements of the alpha-virus (αV) family, including the Venezuelan equine encephalitis virus (VEEV), Sindbis virus (SINV), and Semliki Forest virus (SFV)12. In αV saRNA, the replicase expressed in the nonstructural ORF1 region facilitates self-cloning, while the structural gene region in its subgenomic (sgRNA) is replaced by a gene of interest (GOI), driven by the subgenomic promoter located upstream. This replication and translation mechanism significantly reduces the dosage required for saRNA vaccines compared to conventional linear mRNA vaccines, which require 30–100 μg per dose, saRNA only requires 10 ng–5 μg13. The performance of the gRNA replicon is critical for the saRNA architecture, serving as the foundation for designing effective protein-encoding vectors.
Despite their potential, αV self-amplifying mRNA (αV saRNA) vectors face major engineering challenges in the development of mRNA-encoded therapeutics. First, the lengthy sequence of αV saRNA replicon elements—approximately 10 kb—complicates their separation and purification, affects the performance of lipid delivery carriers, and raises concerns regarding structural stability7. Second, the double-stranded RNA (dsRNA) replicative form (RF) generated during αV saRNA replication can activate cytoplasmic nucleic acid receptors PKR, leading to inhibition of host cell translation by phosphorylated eIF2α14,15, while inducing a severe type I interferon response16,17 and triggering programmed apoptosis through antiviral response mechanisms18,19. This strong immune activation poses risks in treating certain self-limiting diseases. Third, the replication of positive-strand viral gRNA relies on RNA-dependent RNA polymerase (RdRp), which typically lacks a proofreading mechanism20 and operates with low fidelity21. This leads to the incorporation of incorrect bases, potentially resulting in reduced long-term titers of the encoded protein and inducing cytotoxicity. These limitations hinder the research, development, and application of αV saRNA in nucleic acid-based therapeutics.
In this study, we demonstrate that human norovirus gRNA (huNoV gRNA) replicates in human cells with relatively high fidelity. This fidelity is attributed to the interplay between the viral RdRp and the virion gene-binding protein (VPg), which regulates the incorporation of bases during replication. Building on our understanding of the replication mechanism of huNoV-gRNA, we engineer VPg saRNA to achieve high-fidelity replication and low immunogenicity in vivo, while retaining the characteristics of long-term and cap-independent expression. This approach provides a versatile framework with broad potential for the development of mRNA-encoded protein therapeutics.
Results
Characterization of functional elements in the huNoV replicon
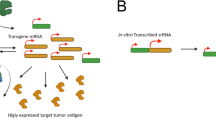
Human norovirus (huNoV) is a small positive-sense RNA virus of the Caliciviridae family, widely recognized as a leading cause of acute gastroenteritis. To date, huNoV remains challenging to culture in mammalian cells22. We constructed huNoV gRNA saRNA using in vitro recombination and IVT technologies (Fig. 1A) to observe its replication in human cell lines and determine whether its replicon could serve as an engineered element for encoding exogenous protein genes in cells. As shown in Fig. 1B, the replicase RdRp and helicase 2C-L from the huNoV genome were expressed in transfected cells. Additionally, huNoV genome (ORF1, VP1, and VP2) mRNAs could be amplified from cells (Fig. 1C). We also detected substantial amounts of dsRNA RF generated by huNoV gRNA saRNA amplification (Fig. 1D). These observations indicate that huNoV genome replication occurs in human cells using IVT technology and lipid nanoparticles (LNPs) delivery systems.
A Schematic illustration of obtaining huNoV gRNA from fecal samples of infected patients and generating T7 IVT-derived huNoV gRNA saRNA containing replicon ORF1 and subgenomic VP1/VP2. The 5′ m7G cap (m7GpppNm) drives the initial translation of genomic saRNA encoding VPg, RdRp and 2C-L. LNPs were used to deliver huNoV gRNA saRNA into Huh-7 cells. B Representative images from three independent experiments showing intracellular RdRp (active form, precursor and full-length ORF1) and helicase 2C-L expression 72 h after transfection, detected by fully automated Western blotting. C qPCR quantification of ORF1, VP1 and VP2 mRNA normalized to GAPDH. D Representative dot blot images from three experiments detecting dsRNA in supernatants of cell lysates, with Ponceau S as control. E Representative automated Western blots from three independent experiments showing p22 and p48 expression in 293T cells transfected with WT, Δp48 or Δp22 huNoV gRNA saRNA-LNPs for 72 h. F qPCR time-course analysis of VP1 mRNA expression following transfection of 293T cells with WT or Δp22 saRNA-LNPs. G Fluorescence microplate-based quantification of VP1 protein levels over time in WT- or Δp22-transfected 293T cells. H Representative confocal images from three independent experiments visualizing ER-associated vesicle structures formed by p48 and p22 in 293T cells transfected with WT, Δp48 or Δp22 saRNA-LNPs, stained with anti-ERp72. I, J Representative automated Western blots from three independent experiments detecting Tlr7 and Tlr3 cleavage in cytoplasmic and lysosomal fractions from 293T cells transfected with WT, Δp48 or Δp22 saRNA-LNPs; LAMP1 marks lysosomes. K Representative automated Western blot showing VPg expression 72 h after transfection. L, M Capillary electrophoresis measurements from three independent experiments quantifying VPg-tagged progeny chains, with recombinant VPg (1 μg) as control. All cell-based experiments were performed independently three times. Data are presented as mean ± s.d. Statistical significance was assessed using two-tailed Welch’s t-test (C, n = 6), two-way ANOVA followed by Sidak’s post-hoc multiple-comparisons test (F, n = 6; G, n = 6), and one-way ANOVA followed by Tukey’s HSD post-hoc test (L, n = 6; M, n = 6). Exact p-values are indicated within the figure. Figure 1A was created in BioRender. Dwad, D. (2025) https://BioRender.com/dfnzple. Source data are provided as a Source Data file.
In host cells, huNoV gRNA saRNA also expressed the N-terminal protein (p48) and 3A-like protein (p22) (Fig. 1E). Deletion of p22 (Δp22) severely impaired the replication of huNoV gRNA saRNA and its ability to drive ORF2 protein expression in host cells (Fig. 1F, G). Confocal microscopy showed that the p22 element of huNoV gRNA saRNA could form vesicle-like structures in the host cell endoplasmic reticulum, recruiting viral genome-linked protein VPg (Fig. 1H), consistent with membrane-bound vesicles formed by positive-strand RNA viruses that facilitate RNA synthesis and protect nascent RNA from nucleases23,24. Endolysosomal TLR3 and TLR7 respond to dsRNA and ssRNA, respectively25,26. The presence of p22 prevented activation of TLR7 and TLR3 in host cell endolysosomes (Fig. 1I, J), although it did not affect TLR9 activation (activated by unmethylated CpG DNA) (Supplementary Fig. 1A). These findings suggest that p22-associated vesicles help protecting nascent ssRNA from recognition and degradation.
The VPg is a 133-amino-acid peptide covalently attached to the 5′ end of huNoV genome27. VPg plays a role in the initiation and elongation phases of picornavirus replication and is proposed to function as a cap analog in the genomes of Caliciviridae and Picornaviridae, distinct from the 5′m7G methylated cap of linear mRNAs28. VPg was also expressed in huNoV gRNA saRNA-transfected host cells (Fig. 1K). However, the precise contribution of huNoV VPg to gRNA replication remains unclear22. Using a biotin-labeled probe to capture the “daughter” replicative chains of huNoV gRNA saRNA (Supplementary Fig. 1B) and analyzing by capillary electrophoresis, we observed that approximately 20% of the negative-sense progeny strands carried VPg at the 5′ end, while 100% of the positive-sense progeny strands were VPg-linked (Fig. 1L, M). The cap-independent translation mechanism of huNoV gRNA saRNA may mitigate limitations traditional linear m7G-capped mRNAs in translation-suppressive disease settings.
We employed huNoV gRNA saRNA to load GOI for replication and expression in eukaryotic cells. In the huNoV gRNA saRNA, we replaced VP1/VP2 with four distinct GOI ORFs of varying lengths: ghrelin (84 nt), Egfp (717 nt), proc1 (1452 nt), and GSDMDENG (1535 nt), each intended for therapeutic application in specific disease contexts in further studies. Four separate IVT plasmids were generated using a two-step Golden Gate reaction, each encoding a distinct huNoV (GOI)-saRNA construct. These plasmids were then transcribed into linear IVT mRNA using T7 polymerase. Each of the four huNoV (GOI)-saRNA constructs was individually encapsulated into LNPs (Supplementary Fig. 1C), yielding uniform and well-defined particle size and morphology across all constructs (Supplementary Fig. 1D). Upon transfecting huNoV (GOI)-saRNA-LNPs into 293T cells, VPg-saRNA concomitantly expressed VPg, RdRp, and 2C-L with endonuclease/helicase activities (Supplementary Fig. 1E), followed by detectable expression of the four respective GOI proteins (Supplementary Fig. 1F).
In brief, we established a huNoV replicon-based self-amplifying RNA system based on the huNoV replicon to drive therapeutic protein expression in eukaryotic cells.
Self-amplifying huNoV saRNA shows relatively high fidelity
The precision of saRNA in encoding is crucial for the execution of protein function, minimizing cytotoxicity and cellular defensive immune responses. Next, we examined the replication fidelity of huNoV gRNA saRNA in vitro. Studies have shown that RdRp acts as the primary executor of RNA viral genome replication29,30. In Coxsackievirus B3 (Picornaviridae), VPg has been observed to bind to the base of the thumb subdomain of RdRp; however, whether huNoV VPg affects RdRp function to participate in or regulate the RNA replication process remains unknown.
HuNoV RdRp comprises seven conserved motifs that form a palm-shaped RNA synthesis center (Fig. 2A). Molecular docking predictions indicated that VPg residues Lys40 and Gly99 form hydrogen bonds with Lys371 and Asp376 in Motif D at the NTP entry port (Fig. 2B). Biolayer interferometry (BLI) experiments revealed that the binding affinity of recombinant VPg to RdRp lacking Motif D was reduced (Supplementary Fig. 2A). Local rearrangements within Motif D are critical for fidelity control in small RNA viruses31. Using the T32/P2 construct32, we evaluated the effect of VPg on the base incorporation error rate of RdRp in vitro. An artificial RNA template-primer complex (T30/P2) was used to observe the extension of the RNA template catalyzed by RdRp. Samples were collected at different time points (time-course) to monitor the conversion rate from 9-mer to 42-mer, which reflects the G:U mismatch incorporation rate (Rmis), in order to analyze the error-incorporation capacity of different RdRp constructs (Fig. 2C).
A, B Predicted interactions between huNoV VPg and RdRp analyzed by molecular docking, highlighting the involvement of the RdRp palm domain in RNA synthesis. C RNA synthesis fidelity assessed using the T32/P2 construct by comparing correctly incorporated 9-mer (P9) with misincorporated 42-mer (P42mis), and calculating mismatch scores. D Recombinant WT VPg or VPg mutants were combined with RdRp in the T32/P2 system, and mismatch scores were determined by capillary electrophoresis. E Time-dependent mismatch fractions (Ratemis) at each UTP concentration were fitted from capillary electrophoresis data, and Rmis values were subsequently fitted to the Michaelis–Menten equation to obtain KMapp and kmis. F Schematic of the T7 endonuclease I (T7EI) digestion assay used to identify mutations in specific mRNA sequences in vitro. G Huh-7 cells transfected with huNoV gRNA saRNA were serially passaged five times; high-fidelity PCR products were digested with T7EI and analyzed by capillary electrophoresis. H Huh-7 cells transfected with LNP-encapsulated VEEV, VEEV–SINV, or SFV EGFP saRNA-LNPs were passaged five times. High-fidelity RT-PCR products amplified with EGFP primers were digested with T7EI, and the digested DNA fragments were identified and quantified by capillary electrophoresis. VPg EGFP saRNA-LNPs served as high-fidelity controls. Representative electropherograms and digestion patterns from three independent experiments are shown. All cell-based experiments were performed independently three times. All data are presented as mean ± standard deviation (s.d.). Statistical significance was assessed using two-way ANOVA followed by Sidak’s post-hoc multiple-comparisons test (D, n = 3), and one-way ANOVA followed by Tukey’s HSD post-hoc test (G, n = 5; H, n = 5). Exact p-values are indicated within the figure. Figure 2A, B were created in BioRender. Dwad, D. (2025) https://BioRender.com/f0yr3ku. Source data are provided as a Source Data file.
Our findings showed that huNoV RdRp generated up to 54.5% G:U mismatch events within 3 h, while the inclusion of recombinant VPg reduced the mismatch rate to 16.9% (Fig. 2D). Based on the molecular docking predictions results, we engineered VPg K40N and VPg G99V mutant huNoV-gRNAs, as shown in Fig. 2D, disruption of the binding interface with Motif D in VPg largely attenuated the reduction of the mismatch rate of RdRp.
The mismatch rate constant (Kmis) and Michaelis constant (KMapp) were determined using supersaturated UTP substrates (Supplementary Fig. 2B, C). The Kmis value for RdRp alone was approximately 3.3 times greater than that of RdRp/VPg (1.806 h−1 vs. 0.555 h−1), while KMapp values remained consistent (451.9 μM vs. 463.3 μM) (Fig. 2E). These data suggest that VPg regulates RdRp fidelity by modulating the catalytic rate, rather than by altering the affinity for the NTP substrate, consistent with its action on Motif D at the active site. PCR products were digested with T7 endonuclease I (T7EI) enzyme to quantify mutations in the daughter chains (Fig. 2F). Briefly, huNoV gRNA saRNA served as a template for in vitro self-replication using recombinant RdRp/2C-L and a His-VPg-pU primer. The resulting mRNA was reverse transcribed and amplified by RT-PCR with VP1/VP2 primers. Replication errors led to mismatched base pairs and heteroduplex formation between wild-type and mutant cDNA strands, which were recognized and cleaved by T7EI. The extent of digestion indirectly reflected the mutation frequency in the amplified products. The replication products of huNoV gRNA saRNA with VPg K40N and VPg G99V mutations exhibited a increased in mutation frequency in cells (Fig. 2G). Compared to egfp sequences loaded by three types of Alphavirus-based saRNA, progeny strands from the huNoV egfp-saRNA platform show virtually no detectable mutation sites (Fig. 2H).
In brief, VPg-dependent initiation represents a high-fidelity replication mechanism, and there exists potential to develop high-fidelity replication saRNA vectors.
Engineering realization of low immunogenicity of huNoV gRNA saRNA
The p22 element ensures that nascent single-stranded huNoV mRNA is not recognized by antiviral responses. However, this does not prevent the activation of double-stranded DNA cytoplasmic receptors by dsRNA RF. Therefore, we next focused on eliminating the dsRNA immunogenicity of huNoV gRNA saRNA.
The replication of the positive strand of huNoV is not sequence-dependent33, but requires guanylylated VPg (VPg-pG) as a peptide primer for initiation34. In contrast, the negative strand requires either de novo initiation or uridylylated VPg (VPg-pU) for replication33. By engineering a system to generate solely VPg-pU primers within the cell, we aim to reduce the VPg-pG-dependent positive strand to inhibit the formation of paired dsRNA intermediates, thereby reducing immunogenicity. To accomplish this, we first examined the mechanism of VPg nucleosidyl acylation. Recombinant huNoV RdRp35 was employed to synthesize VPg-pU products in vitro (Supplementary Fig. 3A, B). We then constructed an in vitro replication system comprising huNoV gRNA mRNA template, VPg-pU primer, recombinant RdRp, and helicase 2C-L (Supplementary Fig. 3C). The replication system containing only the VPg-pU primer effectively inhibited the generation of VPg-independent de novo 5′ gRNA (−) and markedly reduced the synthesis of 5′ VPg gRNA (+) (Supplementary Fig. 3D, E).
To explore methods for spontaneously generating single VPg-pU in cells, we assessed the selectivity of RdRp for VPg nucleotide acylation using mixed NTPs. The results revealed that the covalent attachment of VPg to NTPs did not preferentially favor UTP (accounting for only 10.04%) but rather predominantly produced VPg-pA and VPg-pC (Supplementary Fig. 3F). RdRp consists of a 19.2 kDa NT segment (Pro) and a 57.5 kDa CT segment (pol) connected by a flexible linker36 (Supplementary Fig. 3G). Deletion of the CT region abolished VPg nucleotidylation, while deletion of the NT segment compromised selectivity, resulting in equal ratios of all four nucleosidyl VPg products (Supplementary Fig. 3H). This confirms that RdRp (pro) directly participates in the differential nucleotidylation of huNoV VPg.
Molecular docking predictions indicated that the NT of RdRp contains an NTP binding pocket formed by two large β-folds, displaying distinct preferred binding free energies for different NTPs, consistent with our experimental findings regarding RdRp (pro) NT’s nucleotidylation selectivity (Fig. 3A). Within this pocket, the residues Thr84, Lys146, and Tyr151 of RdRp (pro) tightly interact with the phosphate group of NTP (Fig. 3B). Deleting any of these residues abolished the nucleotidylation function of RdRp (pro) NT (Fig. 3C). Additionally, residues Ala1, Pro2, and Val82 are crucial for base recognition (Fig. 3B). Mutations of Ala1 and Pro2 to polar amino acids Gly (c.3031C > G) and Gln (c.3034C > A), respectively, reduced affinity for A, C, and G, while Val82 was substituted with polar Gly (c.3274T > G) to enhance affinity for polar purine groups. The triple mutant RdRp (pro) nearly completely inhibited the generation of VPg-pA, -pG, and -pC, while increasing VPg-pU production to 93.7 ± 1.79% (Fig. 3D).
A, B Predicted binding free energy (BE) and interaction patterns between huNoV RdRp(pro) NT and four NTPs by Autodock molecular docking (100 predictions each). C VPg nucleosidation using huNoV RdRp-WT or mutants (ΔT84, ΔK146, and ΔY151), with the ratios of nucleosidylated VPg species quantified by high-resolution HPLC. D VPg nucleosidation using RdRp-WT or additional mutants (V82G, A1G/P2Q, A1G/P2Q/V82G), with HPLC-derived peak-area ratios displayed. E, F Representative dot blotting and automated Western blotting results from three independent experiments detecting dsRNA and VP1/VP2 in Huh-7 cells transfected with huNoV gRNA saRNA-LNPs carrying the c.3031C > G, c.3034C > A, or c.3274T > G mutations; Ponceau S and GAPDH served as loading controls. G, H qPCR and ELISA analyses comparing ISG induction (ifit1, Mx2, infa1, and infb1) and cytokine release (IFN-α, IFN-β, and CXCL10) between WT and triM huNoV gRNA saRNA. I Schematic of the huNoV egfp triM-saRNA design: antisense initiation/termination sequences flanking the GOI antisense strand, an upstream replication termination signal (RTS), and 5′ cap1 ensuring ORF1 translation. The production of VPg-pU, triple-mutant RdRp, and 2C-L enables negative-strand synthesis and RTS-mediated release of VPg-linked GOI positive strands for translation. J Representative dot blot images from three independent experiments detecting dsRNA in lysate supernatants of 293T cells transfected with huNoV EGFP triM-saRNA-LNPs or VEEV EGFP-saRNA-LNPs. K, L qPCR and fluorescence microplate quantification of EGFP mRNA and EGFP protein expression over time following transfection of 293T cells with saRNA-LNPs. M Representative automated Western blot images from three experiments showing PKR dimerization, phosphorylation of PKR (Thr446/451) and eIF2α (Ser51), and induction of Rig-I and MDA5 after transfection with saRNA-LNPs. N, O qPCR and ELISA comparing ISG expression and cytokine release between huNoV egfp triM-saRNA and VEEV egfp-saRNA. All cell-based experiments were performed independently three times. All data are expressed as mean ± s.d. P values were calculated using one-way ANOVA followed by Tukey’s HSD post-hoc test (C, n = 5; D, n = 5), two-way ANOVA followed by Sidak’s post-hoc multiple-comparisons test (L, n = 5; K, n = 5), and unpaired two-tailed Student’s t test (G, n = 6; H, n = 6; N, n = 6; O, n = 6). Exact p-values are indicated within the figure. Source data are provided as a Source Data file.
Next, we generated a HuNoV gRNA saRNA triple mutant carrying the c.3031C > G, c.3034C > A, and c.3274T > G substitutions, which we designated as HuNoV gRNA tirM-saRNA (Supplementary Fig. 3I). Transfecting the triple mutant huNoV gRNA tirM-saRNA into Huh-7 cells eliminated to the limit of detection positive-sense 5′ VPg gRNA (+) and 5′ sgRNA (+) (Supplementary Fig. 3J). The triple mutant RdRp reduced the abundance of dsRNA RFs in the cell lysate supernatant, as well as ISG expression of in cells, and the release of IFN-related cytokines, thereby enhancing the translation of VP1/VP2 proteins (Fig. 3E–H).
An antisense sequence of huNoV saRNA based on a triple mutant RdRp (designated huNoV GOI triM-saRNA) was constructed. This structural rearrangement strategy compensates for the loss of the positive-sense progeny strand by generating 5′ VPg GOI mRNA progeny coding strands within the cell (Fig. 3I). To assess the immunogenicity of huNoV triM-saRNA, traditional VEEV-based saRNA was used as a control. At equivalent doses, the amount of paired dsRNA RF from the daughter mRNA of huNoV triM-saRNA was negligible (Fig. 3J). Although huNoV triM-saRNA did not amplify exponentially within cells, it evaded host immune responses, yielding 2.68 times more protein than VEEV saRNA during the same phase (Fig. 3K, L). The low levels of dsRNA produced by huNoV triM-saRNA avoided the activation of the RNA sensor PKR and reduced eIF2α phosphorylation, thereby prevents translation repression. Furthermore, huNoV triM-saRNA exhibited weaker activation of RIG-I and MDA5 (Fig. 3M), as well as lower ISGs expression and IFN-related cytokines release, compared to VEEV egfp saRNA (Fig. 3N, O).
In brief, these RdRp (pro) mutation strategies facilitate the production of supersaturated VPg-pU in cells, reducing dsRNA-induced immunogenicity by simultaneously suppressing positive-sense chain production, without affecting protein coding capacity.
Optimizing 3′UTR to enhance translation efficiency
We employed HuNoV GOI triM-saRNA as a template to assess the integrity of VPg-mediated saRNA replication progeny in vitro. As shown in Fig. 4A, in vitro replication of VPg saRNA produced multiple incompletely amplified VPg ssRNAs, especially during reactions involving longer mRNA templates. These truncated 5′ Vpg-GOI mRNAs contain potential start codons in the +1 or +2 reading frames, which may lead to the translation of truncated frameshift peptides. The potential impact on biosafety and expression efficiency warrants further investigation.
A In vitro amplification reactions contained 80 nM His-tagged VPg-pU, 4 μg recombinant huNoV RdRp, 2 μg huNoV 2C-L, and 400 μM NTPs, together with four different huNoV GOI triM-saRNA templates (1 μg each). VPg-tagged mRNA products were enriched via his-tag purification, and representative capillary electrophoresis traces from three independent experiments show the integrity of 5′ VPg-linked progeny strands. B RNAFold-predicted secondary structures of the huNoV 3′UTR and ghrelin mRNA, with the VPg-dependent replication start site indicated (red arrow). BLI analysis evaluated binding affinities of synthesized huNoV 3′UTR and ghrelin mRNA to recombinant VPg (Kd from global fit). The red dashed box highlights truncated VPg-ssRNA species detected by capillary electrophoresis; representative results from three independent experiments are shown. C Schematic model for truncated ssRNA generation during VPg-dependent replication initiation. VPg-pU binds the huNoV 3′UTR to initiate negative-strand synthesis, but AXXC-containing hairpins can nonspecifically recruit VPg-pU, generating short VPg-ssRNA products. Replacing the huNoV 3′UTR with a 42-nt apt-VPg 3′UTR of higher VPg affinity reduces such nonspecific amplification. Alphafold3 predicts the 3D architecture of the 42-nt apt-VPg 3′UTR in complex with VPg. D Representative capillary electrophoresis results from three independent experiments showing VPg-mRNA amplification abundance and strand integrity in 293T cells 24 h after transfection with four huNoV GOI triM-saRNA-LNPs containing the 42-nt apt-VPg 3′UTR; unmodified triM-saRNAs served as controls. E Representative flow-cytometry analyses from three independent experiments showing expression of Ghrelin, EGFP, GSDMDNT, and proc1 in cells transfected with huNoV GOI triM-saRNA-LNPs harboring the 42-nt apt-VPg 3′UTR, compared with unmodified constructs. All cell-based experiments were performed independently three times. Exact p-values are indicated within the figure. Figure 4C was created in BioRender. Dwad, D. (2025) https://BioRender.com/f0yr3ku. Source data are provided as a Source Data file.
The HuNoV 3′UTR typically functions as a recruitment sequence for VPg-pU primers, driving the replication of the negative-sense strand. However, the synthetic ghrelin(-) mRNA (Kd = 4.6 × 102 nM) forms a secondary structure with slightly weaker affinity for VPg compared to the HuNoV 3′UTR (Kd = 2.1 × 102 nM) (Fig. 4B). We analyzed the RNA secondary structures of sequences near the break points of the progeny VPg ssRNAs. As depicted in Fig. 4B and Supplementary Fig. 4A–C, the AXXU sequence in interior-loops, stem-loops, or bulge-loops appears to serve as a nonspecific recruitment site for VPg-pU primers to initiate replication. This suggests that VPg recruitment is structure-dependent rather than sequence-dependent. The presence of AXXC-containing neck-loops may result in nonspecific de novo replication events. As shown in Supplementary Fig. 4D, removal of the AC site (Δ33A and Δ30U) or disruption of the lock structure (T29A) significantly reduced the synthetic ghrelin mRNA’s affinity for recombinant VPg. Conversely, mutating site 41 from C to A, thereby generating two consecutive AXXU sequences, increased ghrelin mRNA affinity for VPg.
Based on these insights, we designed and screened modified huNoV 3′UTRs with randomly inserted AXXU sequences as VPg-affinity RNA aptamers (apt-VPg) to inhibit the generation of truncated VPg ssRNAs (Supplementary Fig. 4E). A 42-nt high-affinity apt-VPg proved effective in eliminating truncated VPg ghrelin ssRNA production (Supplementary Fig. 4F), even in the context of long-chain template replication (Supplementary Fig. 4G). The apt-VPg-modified huNoV 3′UTR competitively binds to the VPg-pU peptide, thereby suppressing nonspecific short ssRNA amplification (Fig. 4C). Transfecting cells with huNoV saRNA containing the 42-nt apt-VPg-modified 3′UTR reduced the production of nontheoretical length progeny 5′ VPg-GOI mRNA (Fig. 4D). Notably, the inhibition of truncated 5′ VPg-GOI mRNA production was associated with increased GOI protein expression by minimizing waste in mRNA templates and worker proteins (Fig. 4E). In conclusion, optimizing the VPg saRNA 3′UTR reduces nonspecific short-chain amplification and enhances protein production. We simplified the name of the triM-saRNA construct containing the 42-nt apt-VPg-modified 3′UTR to VPg GOI saRNA for the purpose of establishing an in vivo dosing regimen. The four types of GOI saRNA exhibited minimal activation of lysosomal and cytosolic RNA sensors, indicating reduced innate immune stimulation compared to the original huNoV gRNA saRNA sequence (Supplementary Fig. 4H, I). This suggests that the VPg-GOI saRNA self-replicating vector is less prone to innate immune activation within cells.
The studies outlined highlight three key features of VPg-saRNA: cap-independent translation, high-fidelity replication, and low immunogenicity, which collectively support the design of therapeutic applications of huNoV-saRNA. Leveraging these attributes, we designed therapeutic scenarios to illustrate the advantages of huNoV-saRNA.
Cap-independent translation VPg saRNA circumvents metabolic translational repression
The VPg can serve as a substitute for the cap structure, facilitating the efficient assembly of the 43S pre-initiation complex on its mRNA. This mechanism is therefore termed cap-independent translation37. HuNoV VPg hijacks host cell machinery for translation through eIF4G, distinguishing it from the linear mRNA Cap1-m7G-dependent initiation translation mode that relies on eIF4E (Supplementary Fig. 5A). eIF4E is regulated by mTORC1 signaling, which releases it from the phosphorylated substrate release of the competitive inhibitor 4E-BP1, thereby mediating Cap1-m7G-dependent translation38. In mTOR-deficient fibroblasts (MEF mTOR−/−), the phosphorylation levels of 4E-BP1 and eIF4E were suppressed (Supplementary Fig. 5B). This suppression led to a reduced in the translation of m7G-cap1-driven linear mRNA in MEF mTOR−/− cells, while VPg saRNA effectively bypassed the translation block in these cells (Supplementary Fig. 5C).
The suppression of mRNA cap-dependent translation due to mTOR inactivation is reminiscent of cancer-associated cachexia syndrome (CACs), an energy-wasting syndrome characterized by protein synthesis repression resulting from mTORC1 impairment39. We observed mTORC1 inactivation and dephosphorylation of 4E-BP1/eIF4E in several postmortem muscle fibers from cancer patients who succumbed to wasting, while the phosphorylation level of eIF4G remained relatively unaffected (Fig. 5A). Inactivation of mTORC1/4E-BP1/eIF4E signaling leads to disrupted skeletal muscle protein fiber and fat synthesis (Fig. 5B), resulting in significant weight loss among CAC patients, which severely impacts the quality of life of approximately 50–80% of cancer patients40. Ghrelin41 and selective androgens42 are considered to counteract the anabolic deficiencies associated with CACs. VPg saRNA is expected to overcome the limitations of these short-chain recombinant peptides, which include short half-life, inconvenient administration, and high cost.
A Immunohistochemical staining of skeletal muscle tissues from 3 healthy, 4 LUAD, 4 PAAD and 3 COAD patients showing phosphorylation of 4E-BP1 (Thr37/46), mTOR (Ser2448), eIF4G (Ser1108) and eIF4E (Ser209). Representative 4× and 20× images from each patient group are shown, with integrated optical density (IOD) quantified across individuals. B Diagram illustrating inhibition of protein synthesis via mTORC1/4E-BP1/eIF4E inactivation, which suppresses m⁷G cap-dependent translation, while persistent eIF4G phosphorylation in CAC muscle allows eIF4G-dependent VPg-GOI mRNA translation. C After 5% body-weight loss, mice received 1 μg/kg intraperitoneal VPg Ghrelin saRNA-LNP injections every 4 weeks; controls received linear Cap1-m7G ghrelin mRNA. Endpoints were >20% weight loss or 48 weeks after treatment initiation. D Representative immunohistochemical staining of phosphorylated 4E-BP1, mTOR, eIF4G and eIF4E in KPC-CAC mice at 5%, 10%, and 20% weight loss. E Plasma ghrelin levels measured by ELISA in KPC-CAC mice treated with VPg Ghrelin saRNA or linear Cap1-m7G ghrelin mRNA. F, G Representative H&E staining of subcutaneous fat and skeletal muscle morphology in monthly VPg Ghrelin saRNA-treated mice versus m⁷G ghrelin mRNA controls. H, I Food intake (g/day) and body-weight changes (%) recorded weekly in VPg Ghrelin saRNA-treated KPC-CAC mice. J Survival curves of KPC-CAC mice receiving monthly VPg Ghrelin saRNA injections. All animal data represent measurements from individual animals. All data are expressed as mean ± standard deviation (s.d.). Statistical significance was assessed using Welch’s ANOVA followed by Games–Howell post-hoc tests (A, n = 18 healthy, 23 PAAD, 23 LUAD, 18 COAD), two-way ANOVA followed by Sidak post-hoc test (E, n = 6), two-way ANOVA followed by Games–Howell post-hoc tests (I, J Cap1-m7G ghrelin mRNA: n = 12 (days 0–85), 9 (days 112–140), 6 (day 160), 3 (days 196–267); VPg ghrelin saRNA: n = 21 (days 0–84), 17 (days 112–140), 13 (day 196), 9 (days 224–267); untreated KPC: n = 8 (days 0–84), 4 (day 112), 1 (day 140). Animals euthanized at humane endpoints were excluded from subsequent analyses), and Log-rank (Mantel–Cox) test (J, Cap1-m7G ghrelin mRNA: n = 12; VPg ghrelin saRNA: n = 20; untreated KPC: n = 8). Exact p-values are indicated within the figure. Figure 5B, C were created in BioRender. Dwad, D. (2025) https://BioRender.com/x77tb3p; and created in BioRender. Dwad, D. (2025) https://BioRender.com/dfnzple; Source data are provided as a Source Data file.
Approximately 80% of pancreatic cancer patients develop CACs40. We employed KPC (Pdx1-cre/LSL-KrasG12D/P53R172H) mice43, a transgenic model that spontaneously develops pancreatic ductal adenocarcinoma (PDAC), to evaluate VPg Ghrelin saRNA. Mouse ghrelin is a 28-amino acid polypeptide (GSSFLSPEHQKAQQRKESKKPPAKLQPR) with 92.9% homology to human ghrelin44. KPC mice begin to exhibit progressive weight loss between 15 and 17 weeks of age, and by over 25 weeks, all surviving mice experience nearly 20% body weight loss (Supplementary Fig. 5D). We administered intraperitoneal injections of VPg Ghrelin saRNA-LNP to KPC mice that had lost more than 5% of their body weight, followed by tissue dissection at the terminal point or upon humane euthanasia (weight loss exceeding 20%) (Fig. 5C). The phosphorylation level of 4E-BP1/eIF4E was inhibited in the muscle tissue of KPC mice after more than 5% body weight loss, while phosphorylated eIF4G only began to decrease when body weight loss exceeded 20% (Fig. 5D).
Due to the cascade inhibition of eIF4E, the serum levels of ghrelin driven by linear cap1-m7G mRNA in KPC mice were lower than those driven by VPg Ghrelin saRNA-LNP (Fig. 5E). Additionally, VPg Ghrelin saRNA did not induce weight loss or organ damage in BALB/c background mice (Supplementary Fig. 5E, F). Repeated administration of VPg Ghrelin saRNA does not affect plasma levels of Ghrelin or antiviral IFN-α in mice (Supplementary Fig. 5G, H). KPC mice progressively exhibited severe loss of subcutaneous white adipose tissue (WAT), muscle fiber atrophy (Fig. 5F, G), feeding disorders, and weight loss (Fig. 5H, I). Monthly intraperitoneal injections of VPg Ghrelin saRNA-LNP alleviated the CAC symptoms in KPC mice (Fig. 5F–H) and improved their metabolic impairments, including blocked liver glycogen synthesis, fatty acid metabolism in WAT, and disrupted myofibril protein synthesis (Supplementary Fig. 5I, J). Moreover, ribosomes, endoplasmic reticulum, and mitochondria in the skeletal muscle of mice treated with VPg Ghrelin saRNA-LNP were found to be active (Supplementary Fig. 5K–N). Notably, while VPg Ghrelin saRNA improved the quality of life and metabolic status of the mice, it also extended the survival rate of KPC mice to a certain extent (median 149 days vs. 214 days) (Fig. 5J).
In brief, the cap-independent and long-lasting expression of VPg saRNA encoding ghrelin can enhance the quality of life and moderately prolong survival in cachectic KPC mice.
High-fidelity VPg saRNA ensures the precise realization of complex protein functions
In previous studies, we constructed an oncolytic mRNA vaccine called GSDMDENG, featuring a precise mRNA structure with long hGSDMD c.825T > A; c.884A > G mutation sequences that encode mitochondrial cardiolipin inner membrane-targeting toxicity. This oncolytic mRNA vaccine can induce mitochondrial autophagy and present tumor antigens in KrasG12D mutant tumor cells45 (Supplementary Fig. 6A). GSDMDENG consists of three key structural elements: a 614 nt HRV2 IRES, a 1455 nt GSDMD ORF, and an 84 nt F1LCT, each serving distinct functions in tumor conditional expression, mitochondrial inner membrane pore toxicity, and mitochondrial signal peptide activity (Fig. 6A). This complex and delicate functional structure necessitates high fidelity in saRNA vector performance.
A Diagram illustrating loading of GSDMDENG into saRNA oncolytic vectors in AsPC1 pancreatic cancer cells to evaluate tumor-cell killing by a low-fidelity replicon. B AsPC1 cells transfected with saRNA were serially passaged for 15 rounds. High-fidelity RT-PCR products were digested with T7EI, and representative capillary electrophoresis results from three independent experiments are shown. C Flow cytometry quantification of GSDMDNT protein after transfection of 10 μg of saRNA-LNPs into 107 AsPC1 or hTERT-hPEN cells for 48 h. D Representative automated Western blot images from three independent experiments showing mitochondrial active GSDMD expression following transfection of saRNA-LNPs into AsPC1 or hTERT-hPEN cells. E, F mtROS leakage and mtDNA release measured by qPCR in AsPC1 and hTERT-hPEN cells transfected with saRNA-LNPs. G Humanized-mouse xenografts bearing AsPC-1 tumors were injected peritumorally with 1 μg/kg saRNA-LNPs, and tumor weights were measured at day 42. H Flow cytometry analysis of co-stimulatory molecule expression in axillary lymph-node APCs from AsPC1-mtOVA-mCherry tumor-bearing mice treated with VPg GSDMDENG saRNA-LNPs. I KPC mice received 1 μg/kg intraperitoneal VPg GSDMDENG saRNA-LNPs, and tumor development and replicon activity were monitored over 24 weeks. J Representative immunohistochemistry images showing VPg, GSDMDNT, and RdRp co-localization with mutant Kras protein in KPC pancreas. K ELISA quantification of IFN-γ, IFN-α, IFN-β and Granzyme B in draining lymph nodes from vaccinated, tumor-bearing mice. L Flow cytometry analysis of proliferating Ki67+ CD4+ and CD8+ T cells, TRP1+ dendritic cells and IFN-γ+ CD8+ T cells in DLNs from vaccinated mice. M IFN-γ+ CD8+ T cells sorted from VPg GSDMDENG saRNA-vaccinated KPC mice showed cytotoxicity against Kras-mutant cancer cells. N Survival curves of KPC mice following VPg GSDMDENG saRNA-LNP vaccination for 24 weeks. All cell-based experiments were performed independently three times, and all animal data represent measurements from individual animals. All data are expressed as mean ± s.d. P values were calculated using two-tailed Mann–Whitney U (Wilcoxon rank-sum) test (B, n = 8; G, n = 8), unpaired two-tailed Welch’s t-test (C, n = 6; E, n = 6; F, n = 6; H, n = 5; K, n = 6; L, n = 6; M, n = 6), and Log-rank Mantel–Cox test (I, Mock LNP: n = 5; VEEV GSDMDENG saRNA: n = 15; VPg GSDMDENG saRNA: n = 24). Exact p-values are indicated within the figure. Figure 6I was created in BioRender. Dwad, D. (2025) https://BioRender.com/x77tb3p. Source data are provided as a Source Data file.
After transfecting AsPC1 cells with VEEV saRNA and VPg saRNA loaded with GSDMDENG, we performed 15 subcultures. T7EI digestion revealed multiple mutations in the daughter mRNA (Fig. 6B). Sanger sequencing46 of the reverse-transcribed mRNA identified a total of 22 random mutation sites in the daughter mRNA, with 7 located in HRV2 IRES, 12 in the ORF region, and 2 in the F1LCT region (Supplementary Fig. 6B). This is because AsPC1 cancer cells are killed by GSDMDNT, while sequence mutations that impair GSDMDENG function tend to be retained. We constructed various mutants based on these random mutations to evaluate their effects on linear Cap1-m7G GSDMDENG mRNA function. Three categories of mutations diminished the functionality of linear GSDMDENG mRNA to varying degrees, including specific expression and driving intensity in tumor cells (Supplementary Fig. 6C), GSDMDNT mitochondrial import (Supplementary Fig. 6D), and cancer cell toxicity (Supplementary Fig. 6E). Since mutation points in the daughter mRNA cannot be exponentially amplified, we found no mutation sites in the high-fidelity VPg GSDMDENG saRNA-transfected cells (Fig. 6B). Therefore, high-fidelity VPg saRNA vectors are essential for realizing the functional potential of long-chain mRNAs with complex roles.
Next, we tested the ablation effect of GSDMDENG integrated into the VPg saRNA vector on tumor tissue (Fig. 6A), using VEEV GSDMDENG saRNA as a control. VPg GSDMDENG saRNA, containing the non-mutated adenocarcinoma-targeting HRV2 IRES sequence, effectively drove GSDMD expression specifically in AsPC-1 cells (Fig. 6C). The F1LCT signal peptide facilitated its transport to mitochondria, activating GSDMDNT with pore-forming activity (Fig. 6D). In contrast, VEEV saRNA loaded with GSDMDENG was less stable and showed reduced conditional selectivity for cancer cells in this experiment. (Fig. 6C, D). Exogenously active GSDMDNT induced mitochondrial damage, leading to mtROS leakage and mtDNA release (Fig. 6E, F). In a mouse model bearing AsPCs-1 with a humanized immune system, peritumoral injection of 1 μg/kg VPg GSDMDENG saRNA increased tumor tissue ablation compared to the same dose of VEEV GSDMDENG saRNA (Fig. 6G). VPg GSDMDENG saRNA prompted the uptake and maturation of the mitochondrial antigen mtOVA by antigen-presenting cells (APCs) (Fig. 6H), resulting in sustained proliferation of OT-I and OT-II cells and the production of IFN-γ in AsPCs-mtOVA tumor-bearing mice (Supplementary Fig. 6F–H). The AsPC-1 model, linked to mitochondrial autophagy and presentation, indicated that VPg GSDMDENG saRNA triggered PINK1- and MHCI-mediated mitophagy and antigen presentation, whereas VEEV GSDMDENG saRNA did not induce mitophagy or antigen presentation (Supplementary Fig. 6I).
KPC mice were utilized to observe specific cytotoxic T responses elicited by mitophagy. This model, characterized by constitutive KrasG12D expression and p53 deletion, spontaneously develops PDAC in adulthood. We intraperitoneally injected VPg GSDMDENG saRNA into newborn mice as a prophylactic measure (Fig. 6I). One week later, VPg saRNA successfully drove GSDMDNT expression in KrasG12D-positive pancreatic cancer cells (Fig. 6J), correlating with the abnormal activation of Raf-MEK-MAPK signaling induced by KrasG12D47, which activated EIF4G2/PTBP1 to mediate HRV2 IRES-driven mRNA expression45. Four weeks later, mice treated with VPg GSDMDENG saRNA exhibited IFN responses, activation of antigen-presenting dendritic cells (DCs), and sustained cytotoxic T cell immunity (Fig. 6K, L). Cytotoxic T lymphocytes sorted from the spleen demonstrated a robust in vitro killing effect on KrasG12D AsPC-1 cells but were unresponsive to other Kras mutant cell lines (Fig. 6M). Ultimately, a single intraperitoneal injection of VPg GSDMDENG saRNA increased the 24-week survival rate of KPC mice from 20 to 62.5% (Fig. 6N). These findings indicate that high-fidelity VPg GSDMDENG saRNA supports sustained specific cytotoxic T cell responses, resulting in lasting tumor protection.
Low immunogenic VPg saRNA protects against autoimmune organ injury
The long-term expression, low dosage, and low immunogenicity of VPg saRNA are may be advantageous for treating chronic autoimmune-related diseases. Activated protein C (APC, also known as Proc1) is a plasma serine protease, as depicted in Fig. 7A. It cleaves protease-activated receptor 1 at the classical Arg41 or nonclassical Arg46 sites48. The latter site is located on the surface of allogeneic T cells49. Its activation promotes follicular TFH cell differentiation, germinal center (GC) reactions, and GC B cell maturation, leading to the deposition of nonspecific immunoglobulin (Ig)G in the lungs, liver, and colon, ultimately causing chronic fibrosis. Pulmonary fibrosis is the most serious complication of cGVHD, and utilizing Proc1 to protect alveolar cells from autologous IgG attacks represents a potential strategy for mitigating cGVHD50.
A Diagram depicting how VPg Proc1 saRNA activates PAR1 to modulate allogeneic T-cell responses and protect alveolar epithelial cells from autologous IgG-mediated injury, thereby alleviating chronic GVHD. B Mouse tracheal epithelial cells (TEC; 107 cells) were transfected with 10 μg saRNA- or mRNA-LNP, and Proc1 protein expression was monitored over 28 days to estimate APC protein half-life. C, D Kaplan–Meier survival analysis and clinical scoring (weight loss, activity, posture, fur, and skin integrity) in aGVHD mice monitored for 14 weeks following BMT. E Schematic of chronic GVHD lung-injury model: B10.BR recipients received 0.2 μg/kg VPg Proc1 saRNA by intratracheal inhalation, followed by 12.5 Gy conditioning and transplantation with B6 bone marrow (4 × 106) and T cells (1 × 106). Lung fibrosis was assessed over 8 weeks. F Representative 3D immunofluorescence images from three independent experiments showing VPg and PROC1 protein in lung airways on day 3 post-BMT, confirming VPg Proc1 saRNA-LNP replication in vivo. G, H Flow cytometry analysis of GC B cells, TFH cells and TFR cells in lung-draining lymph nodes at day 7 post-BMT; representative plots from independent biological replicates are shown. I Autoantibodies in bronchoalveolar lavage fluid (anti-dsDNA IgG/IgM/IgA/IgE, anti-SSB, anti-SSA IgG/IgA) measured by ELISA at day 14 post-BMT. J Representative H&E, Masson and immunohistochemistry images showing lung tissue injury, fibrosis and immunoglobulin deposition at day 56 post-BMT, demonstrating the therapeutic efficacy of VPg Proc1 saRNA-LNP. All cell-based experiments were performed independently three times, and all animal data represent measurements from individual animals; representative immunofluorescence. All data are presented as mean ± standard deviation (s.d.). AUC comparison of aGVHD clinical scores among four treatment groups was performed using one-way ANOVA followed by Tukey’s multiple-comparisons test (D, n = 15). One-way ANOVA with Tukey’s post hoc test was used for flow cytometry and ELISA analyses (G, n = 7; H, n = 7; I, n = 7). Proc1 protein half-life was analyzed using the Welch t-test (B, n = 6). Survival analysis used the Gehan–Breslow–Wilcoxon test (C, n = 15). Exact p-values are indicated within the figure. Figure 7A, E were created in BioRender. Dwad, D. (2025) https://BioRender.com/xwlgkhh. Source data are provided as a Source Data file.
We used Proc1 mRNA loaded with Alphavirus saRNA as a control to evaluate the therapeutic effect of VPg saRNA encoding Proc1 on cGVHD. Both VPg Proc1 saRNA and VEEV Proc1 saRNA effectively drive PROC1 expression in mouse airway epithelial cells (Supplementary Fig. 7A) and enable its secretion into the cell culture supernatant for detection (Supplementary Fig. 7B, C). Compared to an equivalent amount of Cap1-m7G linear mRNA transfected, the half-life of VEEV Proc1 saRNA and VPg Proc1 saRNA increased by 3–5 times, respectively (Fig. 7B). VPg Proc1 saRNA exhibited lower immunogenicity in vivo. Intravenous injection of VEEV Proc1 saRNA induced systemic innate immune responses in C57BL/6 (B6) background mice, including activation of PKR, eIF2α, MDA5, RIG-I, and expression of ISG in the lungs, liver, and intestines (Supplementary Fig. 7D, E). In contrast, VPg Proc1 saRNA did not detectably activate dsRNA receptors or induce an IFN-I response. In an acute GVHD model using MHC-incompatible allo-HSCT mice (C57BL/6 (B6) donor mice, B6D2F1 recipient mice) (Supplementary Fig. 7F), pre-injection of VPg Proc1 saRNA improved survival rates and reduced aGvHD scores, while VEEV Proc1 saRNA worsened mortality and aGvHD scores (Fig. 7C, D).
To assess the protective effects of VPg Proc1 saRNA on the organs of chronic GVHD mice, we used an MHC-matched allo-HSCT mouse model (C57BL/6 (B6) as donor mice and B10.BR as recipient mice). A dose of 0.2 μg/kg VPg Proc1 saRNA was administered via inhalation three days prior to observation of protection against chronic pulmonary fibrosis induced by cGVHD (Fig. 7E). VPg Proc1 saRNA was able to drive the replication and expression of 5′VPg mRNA in the airway cells (VPg+ cells), with secreted Proc1 (Proc1+ cells) distributed throughout the airway bronchioles and terminal alveoli (Fig. 7F). Immunological examinations showed that VPg Proc1 saRNA reduced the proportion of allogeneic GC B cells and TfHelp in the mouse lung lymph nodes while increasing the proportion of Tfreg cells (Fig. 7G, H). Additionally, it reduced the abundance of nonspecific IgG antibodies against various self-antigens in the alveolar lavage fluid (Fig. 7I). In functional tests, VPg Proc1 saRNA decreased lung resistance and improved lung compliance in mice (Supplementary Fig. 7G). Pathological assessments revealed that VPg Proc1 saRNA reduced lung damage, collagen deposition, and accumulation of IgG immunoglobulins in the lungs (Fig. 7J).
In conclusion, VPg saRNA functions as a low-immunogenicity carrier suitable for mRNA therapy targeting active autoimmune diseases.
Discussion
Due to the absence of RdRp proofreading enzymes—except for the unique case of coronaviruses, which possess a proofreading exonuclease—RNA virus replication is susceptible to base mismatches, resulting in mutations in the viral genome approximately once per replication cycle51. αV is a high-mutation virus that is prone to genetic recombination events, including deletions and/or duplications52,53. Identifying additional saRNA replicon vectors may help address the inefficient expression of linear mRNA and the low fidelity and high immunogenicity observed with αV saRNA. This consideration is relevant to the long-term safety of mRNA-based therapeutics. In this study, we found that approximately one-fifth of the negative-sense progeny chains produced by huNoV gRNA replication are guided by a polypeptide primer known as the virion gene-binding protein VPg. This VPg-dependent replication mode exhibits relatively high fidelity, with the VPg K40 and G99 amino acid residues modulating the RdRp motif D catalytic rate to influence correct base incorporation. Consequently, huNoV operates through two modes: a high-fidelity replication mechanism reliant on VPg and a low-fidelity replication process that initiates independently of VPg. This dual strategy may support stable transmission of viral information while maintaining a reservoir of genetic diversity.
The huNoV gRNA replication process generates dsRNA pairing intermediates; however, VPg serves as a cap structure substitute, interacting with the host cell’s eIF4G27, thereby facilitating cap-independent translation28. We opted to introduce a replication termination signal at the 3′ end of the template chain, along with an antisense start codon element and GOI sequence at the 5′ end, which enables the production of 5′VPg mRNA via direct replication of saRNA. Previous studies indicate that the synthesis of the antisense chain during the “pioneer round” of huNoV can occur through two mechanisms: de novo initiation and VPg-pU-dependent initiation, although the precise mechanisms underlying RdRp selection during replication remain unclear44. Our findings revealed that the presence of a single VPg-pU in the system substantially reduced the de novo synthesis and replication mechanism of RdRp. To facilitate the production of high-fidelity encoding 5′VPg mRNA in cells, we engineered a triple-mutant RdRp enzyme capable of synthesizing uridylylated VPg through molecular docking predictions and point mutation experiments. Elevated levels of VPg-pU directly inhibited the generation of low-fidelity negative-sense chains in cells. Additionally, the absence of specific VPg-pG markedly reduced the production of VPg-pG-dependent progeny positive chains. While this limitation hinders exponential replication of huNoV gRNA, its low immunogenicity mitigates the reduction in coding efficiency caused by host cell immune responses. Given the critical roles of VPg in three aspects: (i) uridylated VPg serving as a peptide primer for extending the template chain; (ii) binding to motif D of RdRp to regulate fidelity; and (iii) acting as a cap structure alternative to initiate translation, we have designated this huNoV replicon-based vector as VPg GOI saRNA. This vector effectively mediates saRNA replication and the expression of its genetic information in cells.
Single-stranded viral RNA (ssRNA) can still activate receptors, such as cytoplasmic and endolysosomal TLR9/7, triggering IFN-I responses that lead to translational shutdown and systemic inflammation54. COVID-19 mRNA vaccines utilize N1-methylpseudouridine (N1mΨ) chemical modification to mitigate IFN-I responses and enhance efficacy55. This chemical modification technology has also been applied to improve the expression efficiency of αV saRNA, using fully modified nucleotides such as 5-hydroxymethylcytidine (hm5C), 5-methylcytidine (m5C), or 5-methyluridine (m5U)56. While these modifications may be gradually lost during replication, they can still enhance the translation efficiency and immune stability of αV saRNA. For VPg saRNA, immune stability can be achieved without requiring additional chemical modifications beyond the incorporation of N1mΨ during parent chain IVT. This is because the daughter 5′-VPg mRNA generated from the replication of the parent chain template is encapsulated in vesicle-like structures associated with the replicon23. These vesicles associate with the endoplasmic reticulum for direct protein encoding and evade recognition by TLR7/9 in the endolysosome. The formation of this vesicle relies on the p22 protein expressed in huNoV ORF1, which may contribute to the low immunogenicity of VPg GOI saRNA.
In subsequent treatment scenarios across various mouse disease models, VPg GOI saRNA showed practical advantages, such as low-dose administration, long-term self-replication, and flexible administration routes. A single monthly intraperitoneal injection of 1 μg/kg VPg Ghrelin saRNA significantly improved the quality of life in mice with tumor-associated cachexia and indirectly prolonged survival in cancer-bearing mice. The translation-driven capability of VPg saRNA, independent of host cell eIF4E, is particularly important. It broadens treatment scenarios where traditional Cap1-m7G mRNA may struggle, such as encoding therapeutic antibodies in severe infectious diseases (e.g., FMDV Lpro, Rhinovirus 2Apro, and SARS-CoV-2 Mpro enzymes, all of which disrupt the host eIF4E-eIF3 interaction to limit cap-dependent translation)26,57,58 and in mTOR dysregulation-related energy wasting diseases. In the context of autoimmune diseases, a single pulmonary inhalation of 0.2 μg/kg VPg Proc1 saRNA prior to bone marrow transplantation alleviated chronic pulmonary fibrosis caused by aGvHD. The low immunogenicity of the VPg saRNA vector was particularly valuable in this setting, as it offset the IFN-I response that was associated with the αV saRNA control and thereby maintained the therapeutic effect of Proc1. The high fidelity of VPg saRNA vectors is critical for the treatment of complex oncolytic mRNA, as random mutations during replication can lead to a partial loss of oncolytic function.
The structural characteristics of VPg saRNA offer practical advantages. The huNoV gRNA is only 3.9 kb long, achieving the core functions of self-replication and driving GOI expression without imposing strain on existing purification methods and delivery vector technologies. VPg-dependent translation does not generate redundant protein fragments, nor does it itself connect to protein products. The infectivity and host toxicity of huNoV primarily stem from structural capsid proteins VP1 and VP2, as well as the virulence factor VF159. The replacement of VP1/VP2-VF1 suggests a favorable biosafety profile for VPg saRNA. No studies have indicated that huNoV nonstructural replicase-related proteins are toxic to human cells, and our research did not observe organ toxicity from VPg saRNA.
In summary, the investigation of the huNoV replication mechanism is essential for developing new saRNA structures. Observing the functional mechanisms of VPg saRNA vectors in host cells further clarifies the infection cycle and immune escape mechanisms of huNoV. As illustrated in the schematic diagram (Fig. 8), the use of huNoV gRNA enables the development of an of saRNA vector characterized by safety, reliability, high expression efficiency, low immunogenicity, low dosing requirements, plug-and-play functionality, and broad applicability, suitable for therapeutic mRNA-encoded protein drug development.
Figure 8 was created in BioRender. Dwad, D. (2025) https://BioRender.com/xwlgkhh.
Methods
Cell culture
AsPC-1 human pancreatic adenocarcinoma cells (ATCC CRL-1682), hTERT-immortalized human pancreatic ductal epithelial cells (hTRET-hPNE; ATCC CRL-4023), HEK293T cells (ATCC CRL-3216), and HuH-7 human hepatocellular carcinoma cells (Procell CL-0120) were obtained from the indicated repositories and authenticated by the suppliers. AsPC-1, hTRET-hPNE, and HuH-7 cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; high glucose) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin (P/S). HEK293T cells were cultured under the same conditions unless otherwise specified. All cells were incubated at 37 °C in a humidified atmosphere containing 5% CO₂ and were routinely tested to confirm the absence of mycoplasma contamination.
Acquisition of norovirus genomic cDNA
Fecal samples from patients suspected of norovirus infection were tested using multiplex real-time quantitative PCR (qRT-PCR) to confirm positivity. Subsequently, the genogroup and genotype of the norovirus were determined by Sanger sequencing. A BLAST search against the NCBI nucleotide database confirmed that the obtained strain was Hu/GII.P4 New Orleans 2009_GII.4 Sydney 2012/NSW789Z/2016/AU (99.71% match, GenBank: KY905331.1). Additionally, blood samples were collected from the patients, and total RNA was extracted for reverse transcription to synthesize first-strand cDNA. The norovirus gRNA region, encompassing the 5′ UTR, ORF1, VP1, VP2, and 3′ UTR, was amplified using the following primers: forward primer: 5′-GACTCAGGCGCCctagaaagcacttggcctctaa-3′ (the underlined sequence indicates the KasI restriction site); reverse primer: 5′-CAGTCACCTGCAGGtaaagaaaaagaaagataatcaattttgtcttttca-3′ (the underlined sequence indicates the SbfI restriction site). The amplified product was digested with KasI and SbfI and ligated into a similarly digested pUC19 cloning vector (New England Biolabs) using T4 DNA ligase. The resulting construct, designated pUC19-hNoV-gRNA, was further verified by Sanger sequencing to confirm the correct insertion of the target DNA fragment.
Generation of huNoV gRNA mutations and deletions
Site-directed mutagenesis or gene deletion was performed on the pUC19-hNoV gRNA plasmid template using the Q5 site-directed mutagenesis kit (New England Biolabs). The following primers were designed to construct various mutants: RdRp Mutants, RdRp ΔT84-his forward: 5′-actactcatcaaaaggtccac-3′, reverse: 5′-taaccacggtgccctc-3′; RdRp ΔK146-his: forward: 5′-aaggggtaatgactatgtg-3′, reverse: 5′-tgtagatgtaggggc-3′; RdRp ΔY151-his: forward: 5′-tgtggtcattggagtc-3′, reverse: 5′-agtcattaccccttttgtag-3′; RdRp A1Q/P2Q-his: forward: 5′-catcatcatcatgcaacagcaagcatttg-3′, reverse: 5′-atgatgatgatgcatgaattc-3′; RdRp V82G-his: forward: 5′-gagggcaccgatattacactactcatcaaaag-3′, reverse: 5′-aggtgcgccttcttc-3′; HuNoV gRNA Deletions, HuNoV gRNA Δp22: forward: 5′-ggcaagaaggggaag-3′, reverse: 5′-ctgcagttcaaattcatc-3′; HuNoV gRNA Δp48: forward: 5′-gctgcaagacctgag-3′, reverse: 5′-gtaaagaaaaagaaagataatcaattttgtc-3′; VPg Mutants, VPg K40N (K915N in full-length sequence): forward: 5′-aaggaatggcaactactctatagaag-3′, reverse: 5′-aaggaatggcaactactctatagaag-3′; VPg G99V (G974V in full-length sequence): forward: 5′-tttggtcacagtgtctgaaattaggaaaag-3′, reverse: 5′-ccgagagagactctttc-3′; All constructs were verified by Sanger sequencing to confirm the desired mutations or deletions.
In vitro transcription (IVT)
The PUC19-hNoV gRNA plasmid and its mutant derivatives were digested with KasI/SbfI, followed by PCR amplification of the gRNA region using: forward primer: 5′-GACTCAGAATTCctagaaagcacttggcctctaa-3′ (EcoRI site underlined, start codon in lowercase), reverse primer: 5′-GTCACTGCGGCCGCtcaGTGGTGGTGGTGGTGGTGCTCtaaagaaaaagaaagataatcaattttgtcttttca-3′ (NotI site underlined, stop codon in lowercase, His-tag in bold). The amplified product was digested with EcoRI/NotI and ligated into the similarly digested pIVTRup-T7 expression vector (Biovector) using T4 DNA ligase, generating the construct pIVTRup-his-HuNoV-gRNA. The plasmid pIVTRup-his-HuNoV gRNA was linearized by PCR using primers containing the T7 promoter and terminator sequences: forward primer (T7 promoter): 5′-taatacgactcactataggggaagcacacggcc-3′, reverse primer (terminator): 5′-ctagcataaccccttggggcctctaaacgggtcttgaggggttttttaaagaaaaagaaagataatcaattttgtctttt-3′. After 30 PCR cycles, the linearized DNA was column-purified and used as a template for IVT with T7 RNA polymerase (NEB). Each 20 µL IVT reaction contained 1 µg DNA template and was incubated at 37 °C for 2 h with shaking at 1000 x g. 4 mM hm5CTP (5-hydroxymethylcytidine triphosphate) was used to replace CTP to incorporate methylation modifications into IVT saRNA. Post-IVT, residual DNA was degraded using 2 µL DNase I (37 °C, 20 min, 1000 × g). The unmodified linear mRNA was purified using the MEGAclear transcription clean-up kit (Ambion). Capillary electrophoresis was used to assess the integrity of IVT products, as shown in Supplementary Information Table 1 and Supplementary Fig. 8.
For the three linear mRNAs, a co-transcriptional capping strategy was employed using the HiScribe™ T7 ARCA mRNA Kit (NEB #E2065). After purification, the transcripts were converted from Cap 0 to Cap 1 using the mRNA Cap 2′-O-Methyltransferase (NEB #M0366), following the manufacturer’s protocol. In addition, 4 mM N1Ψ (N1-Pseudouridine) substituted for UTP to enhance mRNA stability. For all self-amplifying RNAs (saRNAs), an enzymatic capping approach was adopted. The Cap 1 structure was introduced using the Vaccinia capping system (NEB #M2080) in combination with the mRNA Cap 2′-O-Methyltransferase (NEB #M0366), according to the technical guidelines provided by the manufacturer. In addition, 4 mM hm5CTP (5-hydroxymethylcytidine triphosphate) was used to replace CTP to incorporate methylation modifications into IVT saRNA. The final processed saRNA (HuNoV gRNA saRNA) and mRNA (Cap1-m7G mRNA) were repurified and quantified.
LNP encapsulation and characterization
Self-assembly of saRNA-LNPs: a lipid mixture containing ionizable lipid, DSPC, cholesterol, and PEG-lipid (50:10:38.5:1.5 molar ratio) was combined with HuNoV gRNA saRNA in citrate buffer (1:2 ratio) using a microfluidic mixing system. The mixture was filtered (0.22 µm) and stored at 4 °C. Particle size, concentration, PDI, and zeta potential were measured using NanoSight NS300 (Malvern Instruments, UK). Encapsulation efficiency (EE%) was determined via ultrafiltration centrifugation. TEM imaging (JEM-1230, JEOL, JPN) was performed after staining with 2% uranyl acetate. RNA quantification was conducted using a GeneQuant pro RNA/DNA analyzer (Cytiva). All HuNoV gRNA saRNA doses refer to the mass of the GOI. For example, 1 µg HuNoV gRNA saRNA corresponds to 7.53 × 1011 copies of VP1/VP2 mRNA. Details of all saRNA variants, including sequence modifications and physical characteristics, are summarized in Supplementary Information Table 1.
Detection of progeny 5′ VPg mRNA
To determine the proportion of progeny strands carrying 5′ VPg in the HuNoV gRNA saRNA replicons, 1 μg of huNoV gRNA saRNA-LNP was transfected into 107 Huh-7 cells. After 72 h, cells were harvested by scraping into 10 mL of ice-cold phosphate-buffered saline (PBS) and pelleted by centrifugation at 1500 × g for 5 min at 4 °C. The cell pellet was resuspended in 5 mL of cytoplasmic lysis buffer containing: 25 mM HEPES (pH 7.9), 5 mM KCl, 0.5 mM MgCl2, 0.5% (v/v) NP-40, protease inhibitors (Halt Protease and Phosphatase Inhibitor Cocktail, Thermo Fisher Scientific), RNase inhibitor (RiboLock RNase Inhibitor, Thermo Fisher Scientific). After 5 min of incubation on ice, cell lysates were clarified by centrifugation at 14,000 × g for 10 min at 4 °C in a microcentrifuge. For RNA capture, 100 nM of biotinylated VP1 sense or antisense primer was added to the lysate and hybridized using a thermal cycler with the following program: 85 °C for 15 min, 9–65 °C at 2 °C/s, 65–25 °C at 0.1 °C/s. Subsequently, 500 μL of pre-equilibrated Pierce™ Streptavidin Magnetic Beads (Thermo Fisher Scientific) were added and incubated at room temperature (RT) for 30 min with gentle mixing. The beads were then washed three times with cytoplasmic lysis buffer and resuspended in 300 μL of resuspension buffer from the Thermo Scientific GeneJet RNA Cleanup and Concentration Kit for RNA purification according to the manufacturer’s protocol.
Progeny mRNA analysis was performed by capillary gel electrophoresis as described by Denise A. Warzak60. Briefly: mRNA samples were diluted to 100 ng/μL in nuclease-free water. 1 μL of intercalating dye was added per 10 mL of RNA separation gel to prepare the gel/dye mixture. Appropriate markers and 2 μL of sample (or Agilent RNA Ladder, 200–6000 nt) were loaded into an Eppendorf 96-well Twin.Tec PCR plate. Samples were analyzed using an Agilent 2100 Bioanalyzer system with a 12-capillary short array.
Predicted structure of VPg/RdRp and autodocking analysis
The sequences of huNoV RdRp and huNoV VPg were retrieved from the GenBank database (GenBank: KY905331.1). Structural predictions for huNoV RdRp and VPg were generated using AlphaFold2 (https://colab.research.google.com/drive/1uAJDgAu-ejsG70h6cG-glm8YYU3-GHGt#scrollTo=kOblAo-xetgx). The resulting PDB files were visualized using PyMOL software. To assess the interaction between VPg and RdRp, the PDB files were submitted to the HDOCK online server (http://hdock.phys.hust.edu.cn/data) for docking analysis. The 376–384 region, corresponding to the A motif D, was designated as the RdRp binding site for VPg. Docking predictions were ranked by docking scores, calculated using the knowledge-based ITScorePP or ITScorePR scoring functions. Based on docking results, 10 VPg mutants were selected for further experimental analysis. Mutations at the binding sites K40 and G99 significantly affected the interaction with motif D in RdRp.
T32/P2 construct and in vitro misbase incorporation assay
The T32 template strand was chemically synthesized and purified using 10% polyacrylamide gel electrophoresis, followed by electroelution with an Elu-Trap device (GE Healthcare). After annealing at 95 °C for 3 min, the template was stored in RNA storage buffer containing 50 mM NaCl, 5 mM Tris (pH 7.5), and 5 mM MgCl₂. The T32/P2 construct was formed by annealing the T32 strand with UU dinucleotide primers at a molar ratio of 1:1.25, incubated at 45 °C for 3 min, then cooled to 4 °C for storage.
In vitro RNA replication reactions were performed in 20 µl reactions at 30 °C for 3 h, containing 4 µM T32/P2 construct, 50 µM synthetic VPg, 1 µM purified RdRp, 300 µM ATP, 300 µM UTP, 20 mM NaCl, 50 mM Tris (pH 7.0), 5 mM MgCl₂, and 2.5 mM DTT. Following the reaction, RNA was analyzed by high-resolution capillary gel electrophoresis, and the misincorporation fraction (rmis) of mismatch products P9 and P42 was calculated. The rmis values were fitted to a Michaelis–Menten model: Rmis = kmis × [S]/(KMapp + [S]), where kmis is the rate constant, and KMapp is the Michaelis constant for UTP.
Biolayer interferometry (BLI) experiments
BLI experiments were conducted using an Octet 8-channel system (Sartorius) in HBS buffer supplemented with 0.05% (w/v) Tween-20 as the blocking buffer. VPg (30 nM) was immobilized onto Octet protein A biosensors. Subsequently, the biosensors were immersed in wells containing purified RdRp proteins, lacking specific motifs, at concentrations of 320, 160, 80, 40, 20, and 10 nM for 200 s. The biosensors were then transferred to wells containing HBS buffer supplemented with 0.05% (w/v) Tween-20 for 300 s to measure dissociation. The data were reference-subtracted and curve-fitted using the “association-dissociation” model in GraphPad Prism.
Determination of progeny mRNA mutation frequency by T7 endonuclease I cleavage assay
Huh-7 human hepatoma cells were maintained in DMEM supplemented with 10% FBS and 1% penicillin/streptomycin at 37 °C under 5% CO2. For transfection, 1 × 10⁷ cells were seeded in 6-well plates and transfected at 70–80% confluency with 1 μg of HuNoV gRNA saRNA-LNP and the corresponding VPg mutant strains. Untreated cells and empty LNP-transfected controls were included. Cells were harvested 72 h post-transfection for RNA extraction. Total RNA was isolated using TRIzol Reagent (Invitrogen). Briefly, cells were lysed in TRIzol, mixed with chloroform, and centrifuged at 12,000 × g for 15 min at 4 °C. The aqueous phase was precipitated with isopropanol, washed with 75% ethanol, and dissolved in RNase-free water. RNA concentration and purity (A260/A280 ≥ 1.8) were verified by NanoDrop™ spectrophotometry. Reverse transcription was performed with 1 μg RNA using a PrimeScript™ RT Reagent Kit (Takara) with oligo(dT)18 and random hexamer primers in a 20 μL reaction (25 °C for 5 min, 42 °C for 60 min, 70 °C for 5 min). VP1 and VP2 genes were amplified using gene-specific primers (VP1-F: 5′-GGGACCTGGGTGGCGACTA-3′, VP1-R: 5′-CAGCCACACCTTCGGCATA-3′; VP2-F: 5′-TGGGTGCTGGAGTTGATGG-3′, VP2-R: 5′-CCTCGGTGGCAATCTCTTC-3′). GAPDH (F: 5′-GAAGGTGAAGGTCGGAGTC-3′, R: 5′-GAAGATGGTGATGGGATTTC-3′) served as the endogenous control. PCR reactions (25 μL) contained 2 μL cDNA, 10 μM primers, and 2× Taq PCR Master Mix (Thermo Fisher), with cycling conditions: 95 °C for 5 min; 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 45 s; final extension at 72 °C for 5 min. The reverse transcription product was subjected to denaturation and hybridization in a PCR thermocycler (95 °C for 5 min, followed by temperature reduction from 95 to 85 °C at 2 °C/s, then from 85 to 25 °C at 0.1 °C/s) to ensure generation of mutant chimeras. Subsequently, 10 U of T7 Endonuclease I (Beyotime) was added to the hybridization product and incubated at 37 °C for 30 min for enzymatic digestion. Finally, the digestion results were analyzed by capillary gel electrophoresis (Agilent 2100 Bioanalyzer system), and the number of resulting cleavage fragments was quantified.
RdRp and VPg recombinant proteins
To obtain recombinant RdRp and its mutant variants for in vitro experiments, the PUC19-hNoV gRNA plasmid (digested with KasI/SbfI) and corresponding RdRp mutant constructs were used as templates for PCR amplification. The RdRp region (nt 3025-5097) was amplified using the following primers: forward primer: 5′-GACTCAGAATTCatgGCCCCGCCAAGCATTTGGT-3′ (EcoRI site underlined, start codon in lowercase), reverse primer: 5′-GTCACTGCGGCCGCtcaGTGGTGGTGGTGGTGGTGCTCTCACTCGACGCCATCTTCATTTCA-3′ (NotI site underlined, stop codon in lowercase, His-tag in bold). The PCR product was digested with EcoRI/NotI and ligated into the similarly digested pET-28b(+) expression vector (Novagen) using T4 DNA ligase, generating the construct pET-his-RdRp. For VPg protein expression, the VPg region (nt 2626–3024) was amplified from KasI/SbfI-digested PUC19-hNoV gRNA using: forward primer: 5′-CAGTCAGGTACCatgGGCAAGAAGGGGAAGAACA-3′ (Acc65I site underlined, start codon in lowercase), reverse primer: 5′-CAGTCACTCGAGtcaAAAACTGAGTTTCACATT-3′ (PaeR7I site underlined, stop codon in lowercase). The amplified product was digested with Acc65I/PaeR7I and cloned into the pE-SUMO vector (LifeSensors) via T4 DNA ligase, yielding pE-His-SUMO-VPg.
The constructed pET- and pSUMO-based plasmids were transformed into E. coli BL21(DE3) cells (Novagen). Protein expression was induced with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG), and recombinant proteins were successfully expressed in the E. coli lysate. All recombinant proteins were purified using Ni-NTA affinity chromatography. Elution was performed using a buffer containing 300 mM NaCl, 50 mM NaH2PO4, 10% glycerol, and 60 mM imidazole (pH 8.0). The recombinant his-tagged proteins—his-RdRp ΔT84, his-RdRp ΔK146, his-RdRp ΔY151, his-RdRp A1Q/P2Q, and his-RdRp V82I—were dialyzed overnight at 4 °C in a buffer containing 150 mM KCl, 50 mM Tris (pH 8.0), 2 mM DTT, and 20% glycerol. The dialyzed proteins were aliquoted and stored at −80 °C.
For His-SUMO-VPg, the proteins were dialyzed overnight in PBS (pH 7.4) with 10% glycerol and 2 mM DTT at 4 °C. SUMO tags were removed by incubating the purified fusion proteins at 1 mg/ml with His-tagged SUMO protease (1 U/20 μg) (LifeSensors, Inc.) overnight at 4 °C. Following tag removal, the protein mixtures were diluted to 2 ml and incubated with 250 μl of a 50% Ni-NTA resin suspension (Qiagen) for 1 h at 4 °C. After incubation, the supernatant containing untagged VPg K40N or G99V was collected by centrifugation at 10,000 × g for 10 min.
The supernatant was concentrated using a 10 kDa cutoff Centricon column (Millipore) by centrifugation at 3000 × g for 45 min, reducing the final volume to 1 ml. The concentrated proteins were then dialyzed overnight at 4 °C in a buffer containing 150 mM KCl, 50 mM Tris-HCl (pH 8.0), 2 mM DTT, and 20% glycerol. This was followed by a second round of concentration using a 10 kDa cutoff Microcon column (Millipore) by centrifugation at 10,000 × g for 30 min. Protein concentrations were determined using the Bradford assay, and samples were stored at −80 °C.
In vitro generation of nucleosidylated VPg
Purified HuNoV RdRp and VPg peptides (>99% purity, confirmed by Coomassie staining) were used to nucleosidylate VPg in a 20 µl reaction mixture containing 50 mM HEPES buffer (pH 7.0–7.5), 3.5 mM magnesium acetate, 6–8% glycerol, 1 µl NTP mix (10 mM), 50 µM recombinant VPg, and 1 µM recombinant RdRp. The reaction was incubated at 33 °C for 60 min. The formation of VPg-pU was assessed by liquid chromatography–Tandem mass spectrometry (LC–MS/MS)46, and the selectivity of VPg nucleosidylation was evaluated using high-performance liquid chromatography (HPLC).
HPLC separation and LC–MS/MS analysis of nucleosidylated VPg
LC–MS/MS analysis of nucleosidylated VPg was performed on a total of 2 samples, including unmodified VPg (n = 1) and nucleosidylated VPg (n = 1) species with 3 technical replicates. Samples were prepared by diluting purified VPg or in vitro nucleotidylation reaction mixtures in LC–MS-grade water, centrifuging at 12,000 × g to remove particulates, and injecting 1 μl of each preparation onto a CAPCELL CORE AQ S2.7 column (2.1 × 150 mm). HPLC separation was carried out on a Waters Alliance 2795 system equipped with a 2996 PDA detector using 10 mmol/L ammonium phosphate as the mobile phase at a flow rate of 400 μl/min, allowing up to 200 μg of VPg per injection without loss of resolution. Fractions corresponding to individual nucleosidylated VPg species were manually collected and stored at −80 °C for subsequent LC–MS analysis. Mass spectrometry was performed on a Waters Q-TOF Micro instrument equipped with a dual electrospray ionization lockspray source, acquiring data in positive-ion mode under identical conditions for sample eluates and a VPg reference infused at 2 μl/min. Source parameters included a source temperature of 120 °C, desolvation temperature of 300 °C, and cone voltages of 25 V (reference) and 20 V (sample). Raw spectra were processed and deconvoluted using MassLynx 4.1 with the MaxEnt1 Transform algorithm to obtain zero-charge intact molecular masses. Because nucleotidylated VPg yields intact mass shifts rather than proteolytic peptides, peptide-database searches, missed-cleavage criteria, FDR filtering and peptide-scoring parameters were not applicable. Instead, VPg nucleosidylation states were identified by theoretical-to-observed mass matching, with VPg-pU confirmed by an intact mass increase of 303.17 Da corresponding to uridylylation at Tyr30 (Y30). All LC–MS analyses were repeated independently at least 3 times to ensure reproducibility.
In vitro extension reaction system for HuNoV gRNA mRNA
The purity of HuNoV 2C-L helicase and RdRp proteins (>99%, confirmed by Coomassie staining) was assessed prior to their use in in vitro extension reactions. Reactions were carried out in a 25 µl system containing 50 mM HEPES buffer (pH 7.0–7.5), 3.5 mM magnesium acetate, 6–8% glycerol, 50 µM VPg-pU peptide primer or 80 nM pU22 primer, 0.5–16 µM HuNoV gRNA saRNA template, and 1 µM each of purified 2C-L helicase and RdRp. After incubation at 37 °C for 60 min, the reaction mixture was applied to a Ni-NTA column and eluted with 60 mM imidazole buffer. The production of progeny his-5′VPg mRNA was finally assessed by capillary gel electrophoresis (Agilent 2100 Bioanalyzer system).
Structural rearrangement of huNoV gRNA saRNA and insertion of GOI sequences
To construct the huNoV gRNA saRNA backbone, the ORF1 (nt 1–5105) regions was amplified from the PUC19-hNoV gRNA template using BsmBI-restricted primers, followed by Golden Gate assembly into the pRC0569 vector. Concurrently, synthetic DNA fragments containing: Rts+polyT, GFP1(-), Kozak(-), HuNoV 3′ UTR were similarly assembled into pRC0569 via BsmBI-mediated Golden Gate cloning (all sequence details in Supplementary Information Table 2). The reaction mixture (2 µL) was transformed into Turbo Competent Cells (NEB), plated on carbenicillin agar, and nongreen colonies were selected for miniprep and sequencing. For secondary assembly, the above fragments were cloned into the pIVTRup-T7 vector using a BsaI-based Golden Gate reaction, generating pIVTRup-T7 VPg saRNA. Transformed cells (2 µL plasmid product) were plated on kanamycin agar, and nongreen colonies were verified by sequencing. The Ghrelin, proc1, and GSDMDENG sequences were inserted into pIVTRup-T7 VPg saRNA via BamHI/MfeI restriction sites. The plasmid was PCR-amplified using restriction-site-flanked primers, followed by column purification. 1 µg linearized template per 20 µL reaction with T7 RNA polymerase (NEB).
Optimization of the 3′UTR sequence
To enhance the translation efficiency of huNoV gRNA, the 3′UTR was optimized by designing randomized insertions of the AXXU sequence into a VPg-binding RNA aptamer (apt-VPg). Among seven different-length apt-VPg candidates theoretically predicted to exhibit high VPg-binding affinity, in vitro experiments involving 2C-L RdRp and VPg-pU elongation on a ghrelin mRNA template confirmed that the 42-nt apt-VPg (tgtgaaaagacaaaattgattatctattcatttattcattta) demonstrated superior secondary structure stability and higher VPg-binding activity. For the construction of huNoV gRNA saRNA with an optimized 3′UTR sequence, the 42-nt apt-VPg was used to replace the original 3′UTR in the triple-mutant PUC19-hNoV triM-gRNA template plasmid using the Q5 Site-Directed Mutagenesis Kit (New England Biolabs). To generate replace, the following primers were employed: forward primer: 5′-atctattcatttattcatttactagaaagcacacgg-3′, reverse primer: 5′-aatcaattttgtcttttcacattaagcccgtgactc-3′. Subsequently, standard procedures were carried out, including IVT, methylation modification, tailing, capping, and LNP encapsulation.
Fully automated Western blot analysis
Total protein concentration in the cell lysates was determined using a Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. Briefly, a standard curve was generated using bovine serum albumin (BSA) 0–2000 μg/mL, and sample absorbance was measured at 562 nm using a microplate reader (BioTek Synergy H1). For Western blot analysis, 5 μg of total protein per sample was mixed with 1× fluorescent master mix (ProteinSimple) containing a fluorescent molecular weight marker and reducing agent (DTT). The samples were heated at 95 °C for 5 min to ensure complete denaturation before loading. Protein separation and immunodetection were performed using the Jess™ Fully Automated Western blot system (Bio-Techne) with the following parameters: separation module: 12–230 kDa (SM-W008, 25-capillary cartridge), separation time: 30 min (optimized for high-resolution separation). Automated data collection: the Jess system performed capillary-based size separation, immunoprobing, washing, and chemiluminescent detection without manual intervention. Raw data were processed using Compass for Simple Western™ (ProteinSimple, v6.1.0). Peak area quantification was used to determine relative protein expression levels.
Dot blot analysis of viral replication intermediate dsRNA
To detect dsRNA replication intermediates generated during viral infection, a dot blot assay was performed as follows: cells were lysed in TRIzol reagent (Invitrogen) for total RNA extraction, followed by phase separation with chloroform. The aqueous phase containing RNA was mixed with an equal volume of 2 × RNA loading buffer (95% formamide, 0.025% SDS, 0.5 mM EDTA, 0.02% bromophenol blue) and heated at 65 °C for 5 min to denature secondary structures. Ten microliters of the RNA-containing supernatant was spotted directly onto a nitrocellulose membrane (0.45 µm pore size, Millipore) using a Bio-Dot microfiltration apparatus (Bio-Rad). The membrane was air-dried for 15 min to allow RNA binding. UV cross-linking (254 nm, 120 mJ/cm², Stratagene Stratalinker) was performed to immobilize RNA onto the membrane. To confirm equal sample loading, the membrane was stained with Ponceau S (Beyotime) for 5 min, followed by destaining in PBS-T (0.1% Tween-20) until protein spots were clearly visible. The membrane was blocked in 5% nonfat milk in PBS-T for 1 h at RT to prevent nonspecific antibody binding. Anti-dsRNA monoclonal antibody (Scicons, 1:1000 dilution in blocking buffer) was applied overnight at 4 °C with gentle agitation. HRP-conjugated anti-mouse IgG (Cell Signaling, 1:5000 dilution) was applied for 1 h at RT. Enhanced chemiluminescence (ECL, Thermo Fisher) was used for visualization. Images were acquired using a ChemiDoc MP Imaging System (Bio-Rad) with Image Lab software.
Confocal microscopy
293T cells were seeded at low density in 35 mm glass-bottom dishes (MatTek) and cultured overnight at 37 °C in 5% CO2. Non-adherent cells were washed with DMEM (HyClone) supplemented with 10 mM HEPES, 100 µg/ml penicillin, 100 µg/ml streptomycin, and 10% heat-inactivated FBS. Following transfection with huNoV gRNA saRNA-LNPs, cells were fixed with paraformaldehyde. After fixation, primary antibodies (rabbit anti-his-VPg, mouse anti-p22, goat anti-erp72) were applied in the dark, followed by fluorescent secondary antibodies. Imaging was performed using a Leica STELLARIS DIVE confocal system (Wetzlar) according to the excitation wavelength specified by the fluorescent antibody instructions.
qPCR
Gene-specific TaqMan primers were used for qPCR, and reactions were carried out using LightCycler 480 Probe Master Mix and a LightCycler 480 instrument (Roche). Each sample was analyzed in technical replicates for data reliability. Gene expression levels were normalized to GAPDH for accurate comparisons using the comparative Ct method. The required primer sequences are as follows: VP1 primer 5′-atgaagatggcgtcgagtgacg-3′, 5′-ttataccacacgtctacgcc-3′; VP2 primer 5′-atggctggagctttctttgctg-3′, 5′-ttaagcccgtgactcccctc-3′; ORF1 primer 5′-cacctccgagtggaagtggt-3′, 5′-tcgacctcgacagccgagtc-3′; ifit1 primer 5′-cagctgcagaggtgtgaaga-3′, 5′-tggtgcagattccaggtgat-3′; mx2 primer 5′-gctgggacttcttcgacctg-3′, 5′-cagccacattcggttcctta-3′; infa1 primer 5′-atggctgtccttgctcagac-3′, 5′-ggcagacaggaggtccaatc-3′; infb1 primer 5′-aaactcatgagcagtctgca-3′, 5′-aggagatcttcagtttcggagg-3′.
ELISA assay to detect cytokine release
Huh-7 cells were cultured in DMEM supplemented with 10% FBS and 1% penicillin–streptomycin at 37 °C under 5% CO2. Cells were seeded in 6-well plates (1 × 106 cells/well) and treated according to experimental conditions. Supernatants were collected at specified time points, centrifuged (300 × g, 5 min, 4 °C) to remove debris, and stored at −80 °C until analysis. Cytokine levels were quantified using commercial ELISA kits (IFN-α: Invitrogen; IFN-β: R&D Systems; CXCL10: Abcam) following manufacturers’ protocols. Briefly, standards and samples were loaded in duplicate onto pre-coated plates, incubated with detection antibodies, and developed using TMB substrate. Absorbance was measured at 450 nm (reference 570 nm) using a microplate reader (BioTek Synergy H1). Standard curves were generated using a 4-parameter logistic model, and sample concentrations were interpolated from the standard curve. Assay validity was confirmed by R2 > 0.99 for standard curves, intra-assay CV < 10%, and inter-assay CV < 15%. For normalization, total protein concentration was determined by BCA assay when required.
Posthumous muscle fiber analysis from cancer patients with cachexia
The CRISPR/Cas9 system was delivered into target cells using third-generation lentiviral vectors (pLenti-CRISPRv2, Addgene). The lentiviral transfer plasmid was co-transfected with packaging plasmids (psPAX2 and pMD2.G) into HEK293T cells using Lipofectamine 3000 (Thermo Fisher). Viral supernatants were collected at 48 and 72 h post-transfection, concentrated via ultracentrifugation (25,000 × g, 2 h, 4 °C), and titered using a qPCR-based lentivirus titration kit (Takara). Single-guide RNAs (sgRNAs) targeting key GOI were designed using the CRISPR Design Tool (Broad Institute) and CHOPCHOP v3, with selection based on high on-target efficiency and minimal off-target effects. The following sgRNA sequences were cloned into the BsmBI-digested pLenti-CRISPRv2 vector via Golden Gate Assembly (NEB): target gene sgRNA sequence PAM mTOR: attagttcctggccgggttt; eif4e: agaaatctagtagactccta; Eif4g1: cttgccctgttgattactca; pink1:actggcggaagaagcggaga; MHC-I(HLA-A): acggccatcctcggcgtctg. Cloning success was confirmed by Sanger sequencing (Tsingke Biotechnology) using the U6-F primer (5′-gagggcctatttcccatgatt-3′). Target cells were transduced with lentiviral particles at an MOI of 5–10 in the presence of 8 µg/mL polybrene (Sigma). After 48 h, stable integrants were selected using 2 µg/mL puromycin (Thermo Fisher) for 7 days. GFP-positive cells were further enriched via fluorescence-activated cell sorting (FACS, BD FACSAria III) to generate a polyclonal knockout pool.
Human skeletal muscle tissue collection
Postmortem skeletal muscle tissue samples were retrospectively collected from the biobank of pathology at the Red Cross Body Donation Center, Wannan Medical College (Wuhu, China). The tissues were resected, formalin-fixed, and paraffin-embedded (FFPE). Samples were obtained from 14 body donors aged 35–58 years, of whom 42.8% were female. These donors exhibited significant weight loss and were pathologically diagnosed with cancer between 2015 and 2021, forming an independent cohort that included 4 cases of lung adenocarcinoma (LUAD), 4 cases of pancreatic adenocarcinoma (PAAD), and 3 cases of colon adenocarcinoma, as well as 3 healthy donors who died from accidental trauma.
Posthumous muscle fiber analysis from cancer patients with cachexia
Muscle tissues were fixed in 10% neutral buffered formalin for 24–48 h, followed by standard paraffin embedding. Serial sections (4 μm thickness) were prepared using a rotary microtome (Leica RM2235). Slides were subjected to heat-induced epitope retrieval in citrate buffer (pH 6.0) at 95 °C for 20 min. Primary antibodies (all diluted in PBS with 1% BSA): phosphorylated 4E-BP1 (Thr37/46); phospho-mTOR (Ser2448); phospho-eIF4E (Ser209); phospho-eIF4G (Ser1108). HRP-conjugated secondary antibodies (Dako EnVision+ System), DAB chromogen and Hematoxylin counterstaining. Images of sections were collected using an (Olympus IX73) microscope and quantified using ImageJ.
KPC mouse model for cachexia, weight loss, and organ damage
KPC mice (Trp53LSL-R172H; Kras+/LSL-G12D; Pdx1-Cre) were used as a spontaneous pancreatic cancer model to monitor cachexia. Body weight, food intake, and survival were recorded weekly starting at 8 weeks of age. When KPC mice lost >5% body weight, they were administered VPg Ghrelin saRNA-LNP intraperitoneally. Mice were euthanized with pentobarbital (200 mg/kg, intraperitoneally) once body weight loss reached 20%. Muscle, subcutaneous fat, liver, and blood samples were collected post-mortem.
Blood was analyzed for liver and kidney damage (AST/ALT, creatinine, and urea nitrogen levels) and platelet counts to assess hematopoietic damage. Muscle and subcutaneous white adipose tissue were fixed in formaldehyde for 48 h, embedded in paraffin, and sectioned for H&E staining. Immunohistochemistry was performed to evaluate phosphorylated 4E-BP1Thr37/46, mTORSer2448, eIF4ESer209, and eIF4GSer1108 expression. Images were captured with an Olympus BX51 microscope equipped with a DP72 camera. RNA was extracted from muscle, fat, and liver tissue for qPCR analysis of ribosome, endoplasmic reticulum, and mitochondria-associated gene expression.
Construction of linear mRNA and VEEV saRNA controls
The construction of linear Cap1-m7G GSDMDENG mRNA-LNP was performed as previously described45. Briefly, the coding sequences of hGSDMDFL and hGSDMDNT were amplified from human fibroblasts. The IRES and F1LCT sequences were chemically synthesized by GenScript (New Jersey, USA). Gibson assembly was performed using the NEBuilder HiFi DNA Assembly Kit (NEB) on a linearized pUC19 vector containing the T7 promoter. The ATG start codon and hGSDMD mutations (c.884A > G / c.825T > A) were introduced using the Q5 site-directed mutagenesis kit (NEB). The assembled product was transformed into 5-α competent E. coli (NEB), and single clones were selected after carbenicillin screening and verified by Sanger sequencing. IVT was performed using the HiScribe T7 high-yield RNA synthesis kit (NEB), with 4 mM hm5CTP substituted for CTP to enhance mRNA stability. The IVT product was digested with DNase I to remove the DNA template, and unmodified linear mRNA was purified using the MEGAclear transcription clean-up kit (Ambion). Capping with m7G-cap and poly(A) tailing were conducted at 70 °C (5 min) and 4 °C (3 min), respectively. Further purification by column chromatography yielded high-purity Cap1-m7G-modified GSDMDENG mRNA for subsequent LNP encapsulation.
The VEEV, VEEV-SINV, and SFV backbone saRNA T7-IVT vectors were all provided by VectorBuilder. Using VEEV GSDMDENG saRNA as an example, the full-length GSDMDENG oncolytic sequence was amplified from the linear GSDMDENG mRNA vector using restriction enzyme digestion primers: forward primer 5′-GGACATATGgggtcggcctttgagcggg-3′ (the underlined sequence indicates the NdeI site) and reverse primer 5′-CTCCTCGAGttatcctatcatgtatttgagagtttt-3′ (the underlined sequence indicates the XhoI site). The amplified product was digested with NdeI/XhoI restriction enzymes and ligated into the NdeI/XhoI-digested VEEV-saRNA T7-IVT plasmid (VectorBuilder, Guangzhou, CHN) using T4 DNA ligase. Subsequently, standard procedures were performed, including IVT, methylation modification, tailing, capping, and LNP encapsulation.
Mitochondrial analysis
Mitochondrial and cytosolic fractions were separated using the mitochondria/cytosol isolation kit (Biovision, CA, USA) following the manufacturer’s protocol. Approximately 106 cells were resuspended in 2 ml cytosol extraction buffer and disrupted using a Potter-Elvehjem PTFE grinder (Sigma). The homogenate was centrifuged at 1200 × g for 10 min at 4 °C, and the supernatant was further centrifuged at 12,000 × g for 10 min at 4 °C. The supernatant served as the cytosolic fraction, while the pellet (mitochondrial fraction) was washed and collected. GSDMD expression in both fractions was analyzed by immunoblotting, with TOM20 serving as a mitochondrial marker. Total DNA was extracted from 300 µl of cytosol using the QIAquick Nucleotide Removal Kit (Qiagen) and eluted in 30 µl. Mitochondrial DNA (mtDNA) was quantified using the QX200 Droplet Digital PCR System (Bio-Rad) with ND1 as a marker and β-actin as an internal control.
Animal and animal ethics
All animal procedures were approved by the Animal Ethics Committee of Wenzhou Medical University (approval no. XMSQ2024-0471) and conducted in accordance with institutional and national guidelines. All mice were obtained from Cyagen Biosciences or Jackson lab and housed in a specific pathogen-free facility under controlled temperature (20–24 °C), humidity (40–60%) and a 12-h light/dark cycle, with ad libitum access to sterilized water and a standard rodent chow diet (LabDiet 5001, catalog no. 0007688; protein 23.9%, fat 5.0%, carbohydrate 55.2%, fiber 5.0%; 3.36 kcal/g). Animals were maintained in individually ventilated cages with autoclaved corncob bedding and provided with environmental enrichment. NSG mice (NOD.Cg-Prkdcscid Il2rgtm1Wjl) were used for immunodeficient studies. KPC mice (LSL-KrasG12D; LSL-Trp53R172H; Pdx1-Cre) were used as a spontaneous PDAC model. Wild-type C57BL/6 mice were used as background controls. Age- and sex-matched mice were used for all in vivo experiments, and humane endpoints were implemented according to the approved protocol to minimize animal distress.
Tumor-bearing mouse model with humanized immune system
CD34+ cells were isolated using the direct CD34+ precursor cell isolation kit (Miltenyi Biotec). Newborn NSG mice were sublethally irradiated with 100× cGy and subsequently received CD34+ stem cells injected into the liver. Human CD45+ cells in peripheral blood were assessed by flow cytometry at 8 and 12 weeks post-transplantation to confirm engraftment. Beginning on day 8 post-transplant, mice were administered 30 µg of recombinant human FLT3-L protein intraperitoneally each day. At 13 weeks post-transplantation, 5 × 105 AsPC-1 tumor cells were implanted subcutaneously in humanized mice. VPg GSDMDENG saRNA or VEEV GSDMDENG saRNA was injected peritumoral around the tumor on day 5 post-implantation. Tumor size was measured every 2–4 days with an electronic digital caliper, and tumor volume was calculated as length × width × height (mm3). Mice were euthanized when tumor volume exceeded 1000 mm³.
Construction and application of AsPC1-mtOVA-mCherry cells
To generate AsPC1 cells expressing mitochondrially-targeted ovalbumin (OVA257-264) fused with mCherry, we first cloned the OVA257-264 sequence into the PHR-SIN-mCherry vector to create an N-terminal mCherry fusion construct. Lentiviral particles were then produced using this plasmid and subsequently transduced into AsPC1 human pancreatic cancer cells. For in vivo studies, humanized immune system mice bearing AsPC1-mtOVA-mCherry tumors received weekly VPg GSDMDENG saRNA or VEEV GSDMDENG saRNA peritumoral injections (1 μg/kg). On day 42 post-implantation (with analysis performed at day 14), mice were sacrificed and draining lymph node cells were harvested. These cells were then stained and analyzed for CD80, CD86, and CD40 expression by flow cytometry to assess antigen-presenting cell (APC) (CD45+ CD103+ CD11c+ CD11b−) uptake of the mCherry-tagged antigen.
IFN-γ ELISPOT analysis
Fourteen days after peritumoral administration of either VPg GSDMDENG saRNA or VEEV GSDMDENG saRNA, tumor-bearing mice were euthanized, and spleens were aseptically harvested. Single-cell suspensions were prepared by mechanically dissociating splenic tissues through a 70-µm nylon cell strainer (BD Biosciences). Erythrocytes were subsequently lysed using ACK lysis buffer (BioWhittaker), and the remaining leukocytes were washed and resuspended in complete RPMI-1640 medium supplemented with 10% FBS, 1% penicillin-streptomycin, and 50 µM β-mercaptoethanol. For IFN-γ detection, 96-well ELISPOT plates (pre-coated with PVDF membranes containing anti-IFN-γ capture antibody, Biosciences) were seeded with 2 × 105 splenocytes per well and incubated for 24 h at 37 °C in a 5% CO2 humidified incubator. Following stimulation, cells were removed, and plates were washed extensively before incubation with a biotinylated anti-IFN-γ detection antibody (Biosciences). After additional washing, spots were developed using streptavidin-conjugated fluorescent secondary antibody and visualized under a fluorescence microscope (Olympus IX73). Spot enumeration was performed using an automated ELISPOT reader (AID iSpot) and analyzed with AID ELISPOT software.
In vivo T cell proliferation assay
OT-I cells originated from transgenic mice expressing CD8+ T cell receptors specific for MHC-I-restricted OVA peptide (257–264), while OT-II cells originated from mice with CD4+ T cell receptors specific for MHC-II-restricted OVA peptide (323–339). Two days before VPg GSDMDENG saRNA treatment, 2 × 106 OT-I cells or OT-II cells were purified and transferred intravenously into tumor-bearing mice. Four days later, draining lymph nodes were isolated, and OT-I or OT-II cell proliferation was analyzed by flow cytometry. Cells were stained with anti-CD16/CD32 antibodies to block Fc receptors, followed by labeling with anti-CD4, CD8, CD3, CD19, and MHC-I dextramer H-2 Kb/SINFEKL. After permeabilization, Ki67 expression levels were measured to determine proliferation activity.
In vitro CLTs cell killing assay
Tumor-draining lymph nodes were harvested from KPC mice treated with VPg-GSDMDENG or VEEV-GSDMDENG saRNA-LNP. Antigen-specific CD8+ T cells were isolated via H-2Db KrasG12D tetramer sorting or IFN-γ capture after AsPC-1 lysate stimulation, then co-cultured with IFN-γ-primed KRAS-mutant lines (AsPC-1, MDA-MB-231, HT-29, NCI-H358, SW480) in real-time impedance (xCELLigence), flow cytometric (CFSE/7-AAD), and MHC-I-blockade-controlled cytotoxicity assays at E:T ratios of 20:1-2.5:1.
Establishment of mouse GvHD model
Pan T cells were isolated from mouse spleens by depleting non-T cells using the Mouse Pan T Cell Isolation Kit II (Miltenyi Biotec) according to manufacturer guidelines. The purity of T cells (95–98%) was confirmed by flow cytometry. Bone marrow cells were flushed from the tibiae and femurs, with RBCs lysed using ACK buffer (BioWhittaker). For MHC-incompatible GvHD induction, C57BL/6-derived bone marrow and splenic T cells were transplanted into B6D2F1 recipients. For MHC-matched GvHD induction, C57BL/6-derived bone marrow and T cells were transplanted into B10.BR recipients. Recipient mice (8–10 weeks old) underwent total body irradiation with a 12.5 Gy single dose using a BioBeam 8000 (Gamma Service Medical).
Mice were anesthetized with 100 µl of a solution containing Ketavet (20 mg/ml) and Rompun (1 mg/ml) and immobilized in an irradiation chamber. Four hours post-irradiation, recipients were injected intravenously with 4 × 106 mismatched bone marrow cells and 4 × 106 purified mismatched Pan T cells. Body weight was monitored on day 1 and weekly thereafter. Survival was assessed daily. GvHD clinical scoring followed an established system61, including weight loss, posture, activity, hair texture, and skin integrity, with weekly evaluations and a cumulative score based on the sum of the five parameters.
Flow cytometric analysis of GC B and T follicular cells
Single-cell suspensions from lymphoid tissues were prepared by mechanical dissociation, followed by RBC lysis (ACK buffer) and viability staining (Ghost Violet™ 510). Cells were Fc-blocked (anti-CD16/32) and stained for surface markers: GC B cells (CD19+CD95hiGL7+), TFH cells (CD4+PD-1+CXCR5+FoxP3−), and TFR cells (CD4+PD-1+CXCR5+FoxP3+). Intracellular FoxP3 staining was performed using a fixation/permeabilization kit. Samples were acquired on a BD LSRFortessa™, with doublets excluded (FSC-H vs. FSC-W) and live cells gated (Ghost Violet™ 510).
Detection of autoantibodies in mice
Serum levels of IgM, IgG, IgA, and IgE were quantified using commercial ELISA kits (eBioscience) with absorbance measurements performed on a SpectraMax M5 microplate reader (Molecular Devices). For autoantibody detection, ELISA plates were pre-coated with either 2.5 μg/ml calf thymus double-stranded DNA (Sigma) or recombinant/isolated autoantigens, including La protein, Ro52, and Ro60 (10 μg/ml for recombinant proteins; 5 μg/ml for native Ro60; Arotec Diagnostics). Serially diluted serum samples were incubated with the coated antigens, followed by detection using alkaline phosphatase (AP)-conjugated goat anti-mouse secondary antibodies specific for IgM, IgG, IgA, or IgE (Southern Biotech).
3D immunofluorescence staining of VPg and PROC1 in mouse lung tissue
Mouse lung tissues were fixed in 4% paraformaldehyde at 4 °C for 24 h, followed by three washes in PBS (pH 7.4). Tissues were permeabilized with 0.5% Triton X-100 in PBS for 1 h at RT and blocked with 5% BSA containing 0.1% Triton X-100 for 2 h at RT. For immunostaining, tissues were incubated with primary antibodies against VPg (1:200) and PROC1 (1:200) diluted in blocking buffer at 4 °C for 48 h with gentle agitation. After washing with PBST (PBS + 0.1% Tween-20) three times (1 h each), samples were incubated with fluorophore-conjugated secondary antibodies (Alexa Fluor 594 for VPg, AlexaFluor 488 for PROC1; 1:500 and DAPI for nuclear counterstaining at RT for 4 h. For 3D visualization, tissues were cleared using the CUBIC method: Dehydration in CUBIC-1 solution (25% urea, 25% N,N,N′,N′-tetrakis(2-hydroxypropyl)ethylenediamine) for 3 days at 37 °C. Refractive index matching in CUBIC-2 solution (50% sucrose, 25% urea) for 1 day. Cleared tissues were imaged using a light-sheet fluorescence microscope (e.g., Zeiss Lightsheet Z.1) with a Z-step size of 2 μm. 3D reconstruction and colocalization analysis were performed using Imaris 9.0 (Bitplane).
Statistics and reproducibility
No statistical method was used to predetermine sample size in all the highly controlled in vitro or in vivo experiments, but our sample sizes are similar to those reported in previous publications. For each experiment, we aimed for at least three samples or animals per group to allow basic statistical significance. The exact sample sizes used for each experiment are provided in the corresponding figure legends.
No data were excluded from the analyses.
All experiments were reliably reproduced, and data are presented as mean ± s.d. unless otherwise stated. All experiments were repeated with at least three independent biological donors or performed independently three times. Statistical significance between experimental groups was assessed using two-tailed Student’s t tests or one-way or two-way analysis of variance (ANOVA) followed by the appropriate post hoc test or other specified statistical methods, as indicated in the respective figure legends. A P value of 0.05 or less was considered statistically significant.
For in vitro experiments, cells were randomly allocated into control and experimental groups. For in vivo experiments, age- and sex-matched mice were randomized into control and experimental groups.
For therapeutic animal experiments, investigators responsible for data collection were blinded to treatment allocation. No blinding was used for in vitro experiments. The majority of data collection involved quantifiable endpoints for which blinding would not influence measurement bias.
Ethical statement
All cadaveric specimens were obtained through the Wannan Medical College Red Cross Body Donation Center with full ethical compliance. Each donation was accompanied by properly executed informed consent documents, signed by the donors during their lifetime, explicitly permitting the use of their remains for scientific research. The study protocol was approved by the Medical Ethics Committee of the First Affiliated Hospital of Wannan Medical College (approval number: 2022-LSX-12). In accordance with the body donation agreement, all tissue samples were destroyed after analysis, as they are permitted for research use for only 2 years.
All protocols involving animals in this study were approved by the Institutional Animal Care and Use Committee of Wenzhou Medical University and were carried out in accordance with the approved guidelines (approval no. XMSQ2024-0471).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for Figs. 1–7 and Supplementary Figs. 1–7 are provided as Source Data files. LC–MS/MS raw datasets and Sanger sequencing chromatograms generated in this study have been deposited in Figshare and are available at https://doi.org/10.6084/m9.figshare.3078216546 and https://doi.org/10.6084/m9.figshare.3069415146. Raw fluorescence microscopy, cryo-transmission electron microscopy (cryo-TEM), immunohistochemistry (IHC), and hematoxylin and eosin (H&E) staining images are not publicly available due to large file sizes, instrument-specific formats, and institutional data management restrictions; however, all processed and representative images supporting the findings of this study are included in the paper and its supplementary materials. All other data, including raw imaging data, are available from the corresponding author upon request. Source Data are provided with this paper.
References
Skowronski, D. M. & De Serres, G. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 384, 1576–1577 (2021).
Qiu, X. et al. Development of mRNA vaccines against respiratory syncytial virus (RSV). Cytokine Growth Factor Rev. 68, 37–53 (2022).
Liu, C. et al. mRNA-based cancer therapeutics. Nat. Rev. Cancer 23, 526–543 (2023).
Kim, Y. K. RNA therapy: rich history, various applications and unlimited future prospects. Exp. Mol. Med. 54, 455–465 (2022).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines-a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Anderson, B. R. et al. Nucleoside modifications in RNA limit activation of 2’-5’-oligoadenylate synthetase and increase resistance to cleavage by RNase L. Nucleic Acids Res. 39, 9329–9338 (2011).
Blakney, A. K., Ip, S. & Geall, A. J. An update on self-amplifying mRNA vaccine development. Vaccines 9, 97 (2021).
Rohner, E., Yang, R., Foo, K. S., Goedel, A. & Chien, K. R. Unlocking the promise of mRNA therapeutics. Nat. Biotechnol. 40, 1586–1600 (2022).
Minnaert, A. K. et al. Strategies for controlling the innate immune activity of conventional and self-amplifying mRNA therapeutics: getting the message across. Adv. Drug Deliv. Rev. 176, 113900 (2021).
Chen, R. et al. Engineering circular RNA for enhanced protein production. Nat. Biotechnol. 41, 262–272 (2023).
Perkovic, M. et al. A trans-amplifying RNA simplified to essential elements is highly replicative and robustly immunogenic in mice. Mol. Ther. 32, 257–259 (2024).
Bloom, K., van den Berg, F. & Arbuthnot, P. Self-amplifying RNA vaccines for infectious diseases. Gene Ther. 28, 117–129 (2021).
Schmidt, C. & Schnierle, B. S. Self-amplifying RNA vaccine candidates: alternative platforms for mRNA vaccine development. Pathogens 12, 138 (2023).
Ventoso, I. et al. Translational resistance of late alphavirus mRNA to eIF2alpha phosphorylation: a strategy to overcome the antiviral effect of protein kinase PKR. Genes Dev. 20, 87–100 (2006).
García, M. A. et al. Impact of protein kinase PKR in cell biology: from antiviral to antiproliferative action. Microbiol. Mol. Biol. Rev. 70, 1032–1060 (2006).
Rudd, P. A. et al. Interferon response factors 3 and 7 protect against Chikungunya virus hemorrhagic fever and shock. J. Virol. 86, 9888–9898 (2012).
Wauquier, N. et al. The acute phase of Chikungunya virus infection in humans is associated with strong innate immunity and T CD8 cell activation. J. Infect. Dis. 204, 115–123 (2011).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15, 87–103 (2015).
Pindel, A. & Sadler, A. The role of protein kinase R in the interferon response. J. Interferon Cytokine Res. 31, 59–70 (2011).
Castro, C., Arnold, J. J. & Cameron, C. E. Incorporation fidelity of the viral RNA-dependent RNA polymerase: a kinetic, thermodynamic and structural perspective. Virus Res. 107, 141–149 (2005).
Patterson, E. I. et al. Measuring alphavirus fidelity using non-infectious virus particles. Viruses 12, 546 (2020).
Thorne, L. G. & Goodfellow, I. G. Norovirus gene expression and replication. J. Gen. Virol. 95, 278–291 (2014).
Doerflinger, S. Y. et al. Membrane alterations induced by nonstructural proteins of human norovirus. PLoS Pathog. 13, e1006705 (2017).
Wolff, G., Melia, C. E., Snijder, E. J. & Bárcena, M. Double-membrane vesicles as platforms for viral replication. Trends Microbiol. 28, 1022–1033 (2020).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-kappaB by toll-like receptor 3. Nature 413, 732–738 (2001).
Leen, E. N. et al. A conserved interaction between a C-terminal motif in norovirus VPg and the HEAT-1 domain of eIF4G is essential for translation initiation. PLoS Pathog. 12, e1005379 (2016).
Belliot, G., Sosnovtsev, S. V., Chang, K. O., McPhie, P. & Green, K. Y. Nucleotidylylation of the VPg protein of a human norovirus by its proteinase-polymerase precursor protein. Virology 374, 33–49 (2008).
Goodfellow, I. The genome-linked protein VPg of vertebrate viruses-a multifaceted protein. Curr. Opin. Virol. 1, 355–362 (2011).
Högbom, M., Jäger, K., Robel, I., Unge, T. & Rohayem, J. The active form of the norovirus RNA-dependent RNA polymerase is a homodimer with cooperative activity. J. Gen. Virol. 90, 281–291 (2009).
Yang, X. et al. Motif D of viral RNA-dependent RNA polymerases determines efficiency and fidelity of nucleotide addition. Structure 20, 1519–1527 (2012).
Gong, P. & Peersen, O. B. Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase. Proc. Natl. Acad. Sci. USA 107, 22505–22510 (2010).
Liu, W., Shi, X. & Gong, P. A unique intra-molecular fidelity-modulating mechanism identified in a viral RNA-dependent RNA polymerase. Nucleic Acids Res. 46, 10840–10854 (2018).
Rohayem, J., Robel, I., Jäger, K., Scheffler, U. & Rudolph, W. Protein-primed and de novo initiation of RNA synthesis by norovirus 3Dpol. J. Virol. 80, 7060–7069 (2006).
Subba-Reddy, C. V., Goodfellow, I. & Kao, C. C. VPg-primed RNA synthesis of norovirus RNA-dependent RNA polymerases by using a novel cell-based assay. J. Virol. 85, 13027–13037 (2011).
Paul, A. V. & Wimmer, E. Initiation of protein-primed picornavirus RNA synthesis. Virus Res. 206, 12–26 (2015).
te Velthuis, A. J. Common and unique features of viral RNA-dependent polymerases. Cell. Mol. Life Sci. 71, 4403–4420 (2014).
Chung, L. et al. Norovirus translation requires an interaction between the C terminus of the genome-linked viral protein VPg and eukaryotic translation initiation factor 4G. J. Biol. Chem. 289, 21738–21750 (2014).
Qin, X., Jiang, B. & Zhang, Y. 4E-BP1, a multifactor regulated multifunctional protein. Cell Cycle 15, 781–786 (2016).
Geremia, A. et al. Activation of Akt-mTORC1 signalling reverts cancer-dependent muscle wasting. J. Cachexia Sarcopenia Muscle 13, 648–661 (2022).
Baracos, V. E., Martin, L., Korc, M., Guttridge, D. C. & Fearon, K. C. H. Cancer-associated cachexia. Nat. Rev. Dis. Primers 4, 17105 (2018).
Currow, D. C. & Abernethy, A. P. Anamorelin hydrochloride in the treatment of cancer anorexia-cachexia syndrome. Future Oncol. 10, 789–802 (2014).
Dalton, J. T., Taylor, R. P., Mohler, M. L. & Steiner, M. S. Selective androgen receptor modulators for the prevention and treatment of muscle wasting associated with cancer. Curr. Opin. Support. Palliat. Care 7, 345–351 (2013).
Renz, B. W. et al. β2 adrenergic-neurotrophin feedforward loop promotes pancreatic cancer. Cancer Cell 33, 75–90(2018).
Kojima, M. et al. Ghrelin is a growth-hormone-releasing acylated peptide from stomach. Nature 402, 656–660 (1999).
Feng, Z. et al. An in vitro-transcribed circular RNA targets the mitochondrial inner membrane cardiolipin to ablate EIF4G2+/PTBP1+ pan-adenocarcinoma. Nat. Cancer 5, 30–46 (2024).
Feng, Z. Engineered VPg saRNA achieves cap-independent, low-immunogenic and precise encoding of therapeutic proteins in vivo. figshare https://doi.org/10.6084/m9.figshare.30694151.v1 (2025).
Yan, L. et al. Targeting glucose metabolism sensitizes pancreatic cancer to MEK inhibition. Cancer Res. 81, 4054–4065 (2021).
Griffin, J. H., Zlokovic, B. V. & Mosnier, L. O. Activated protein C: biased for translation. Blood 125, 2898–2907 (2015).
Ranjan, S. et al. Activated protein C protects from GvHD via PAR2/PAR3 signalling in regulatory T-cells. Nat. Commun. 8, 311 (2017).
Sinha, R. K. et al. Activated protein C ameliorates chronic graft-versus-host disease by PAR1-dependent biased cell signaling on T cells. Blood 134, 776–781 (2019).
Sanjuán, R., Nebot, M. R., Chirico, N., Mansky, L. M. & Belshaw, R. Viral mutation rates. J Virol 84, 9733–9748 (2010).
Poirier, E. Z. et al. Low-fidelity polymerases of alphaviruses recombine at higher rates to overproduce defective interfering particles. J. Virol. 90, 2446–2454 (2015).
Langsjoen, R. M., Muruato, A. E., Kunkel, S. R., Jaworski, E. & Routh, A. Differential alphavirus defective RNA diversity between intracellular and extracellular compartments is driven by subgenomic recombination events. mBio 11, e00731–20 (2020).
Anderson, B. R. et al. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 38, 5884–5892 (2010).
Kormann, M. S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 29, 154–157 (2011).
McGee, J. E. et al. Complete substitution with modified nucleotides in self-amplifying RNA suppresses the interferon response and increases potency. Nat. Biotechnol. 43, 720–726 (2025).
Lei, J. & Hilgenfeld, R. RNA-virus proteases counteracting host innate immunity. FEBS Lett. 591, 3190–3210 (2017).
Koudelka, T. et al. N-terminomics for the identification of in vitro substrates and cleavage site specificity of the SARS-CoV-2 main protease. Proteomics 21, e2000246 (2021).
Chhabra, P. et al. Updated classification of norovirus genogroups and genotypes. J. Gen. Virol. 100, 1393–1406 (2019).
Warzak, D. A., Pike, W. A. & Luttgeharm, K. D. Capillary electrophoresis methods for determining the IVT mRNA critical quality attributes of size and purity. SLAS Technol. 28, 369–374 (2023).
Cooke, K. R. et al. An experimental model of idiopathic pneumonia syndrome after bone marrow transplantation: I. The roles of minor H antigens and endotoxin. Blood 88, 3230–3239 (1996).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (32471002, 82401829, 82472617, and 82404553; F.Z.Y.), the National University of Singapore (NUHSRO/2020/133/Startup/08, NUHSRO/2023/008/NUSMed/TCE/LOA, NUHSRO/2021/034/TRP/09/Nanomedicine, NUHSRO/2021/044/Kickstart/09/LOA, 23-0173-A0001; C.X.Y.), the National Medical Research Council (MOH-001388-00, CG21APR1005, MOH-001500-00, MOH-001609-00; C.X.Y.), the Singapore Ministry of Education (MOE-000387-00; C.X.Y.), the National Research Foundation (NRF-000352-00; C.X.Y.), the Leading Innovative and Entrepreneur Team Introduction Program of Zhejiang (2023R01002; Z.G.W.), the Distinguished Young Scientists Fund of Zhejiang (LR25H250001; Z.G.W.), and the National Science and Technology Major Project of China (No. 2025ZD1802201; Z.G.W.).
Author information
Authors and Affiliations
Contributions
Z.Y.F. conceived and designed the overall study, supervised key experiments, and led data analysis and interpretation. Z.Y.F. and L.X.C. performed most molecular and cellular experiments. J.Z. and P.W. assisted with cell culture and animal studies. Q.L., Z.L.X., L.Y., and Y.J.H. contributed to data processing, statistical analysis, and figure preparation. X.B.Z., J.H.Z., and Q.C. provided essential reagents, technical guidance, and methodological support. Y.B.P. performed imaging and histopathological examinations. X.K.L., Z.G.W., and X.Y.C. jointly supervised the project, contributed to conceptual refinement, and provided funding and resources. Z.Y.F. wrote the manuscript with input from all authors. All authors discussed the results, revised the manuscript, and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
Xiaoyuan Chen is a co-founder of and holds shares in Yantai Lannacheng Biotechnology Co., Ltd. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, Z., Chu, L., Li, Q. et al. Engineered VPg saRNA achieves cap-independent, low-immunogenic and precise encoding of therapeutic proteins in vivo. Nat Commun 17, 1666 (2026). https://doi.org/10.1038/s41467-026-68364-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68364-w