Abstract
The global rise in steatotic liver disease poses a significant public health challenge. While non-contrast computed tomography scans hold promise for opportunistic detection of steatotic liver disease, their potential for staging and risk assessment remains underexplored. Here we present a multimodal AI model trained on a large dataset, comprising of (n=968) histopathologically and (n=1103) radiologically confirmed cases, validated against both histology (n=660) and MRI-PDFF (n=375) gold standards, demonstrating high accuracy in detecting mild to severe steatosis (AUC: 0.904–0.929) and clinically significant fibrosis (AUC: 0.824–0.888). Furthermore, integrating the model into the standard clinical pathway improves primary risk screening in a retrospective patient cohort (n=1192), identifying 36% more patients at risk of fibrosis progression. Using Cox proportional hazard model, we observe that the intermediate-high risk patients identified by the optimized clinical pathway exhibits a significantly higher incidence of cirrhosis (hazard ratio: 5.54: 2.69–11.42), showcasing the model’s potential for early detection and management of steatotic liver disease.
Similar content being viewed by others
Introduction
Steatotic liver disease (SLD) has become one of the most prevalent chronic liver diseases (CLDs)1, affecting an estimated 30.1% of the global population2. Without effective interventions, this figure is projected to increase dramatically, reaching 55.7% by 20403. SLD encompasses a broad clinical spectrum, ranging from isolated steatosis to steatohepatitis, which carry an elevated risk of progressing to advanced liver disease, including cirrhosis and hepatocellular carcinoma2,4. Moreover, SLD is recognized as an independent risk factor for cardiovascular disease2,5,6. Recent advances have led to FDA approval of medications specifically targeting SLD, with several other promising therapies currently under development7,8,9. These therapeutic options underscore the critical need for early detection, as timely intervention can significantly alter the pathophysiological trajectory of the disease and reduce the burden of liver10 and cardiovascular complications11. Moreover, scalable screening modalities can enhance the awareness of SLD in population to allow early intervention and better management4,6.
Despite the increasing prevalence of SLD, current diagnostic and screening tools have significant limitations. Liver biopsy, although the gold standard for assessing hepatic steatosis, is invasive and impractical to detect and monitor large populations12,13. B-mode ultrasound is highly operator dependent and lacks the sensitivity to reliably detect early steatosis. Serum biomarkers, such as the widely used fatty liver index, also face challenges with time demands and suboptimal accuracy2,5. Advanced imaging techniques, such as MRI-proton density fat fraction (PDFF), although being highly sensitive, are often inaccessible due to its high costs, limited availability, and time-consuming nature of each test. Vibration-controlled transient elastography (VCTE) offers a non-invasive alternative but is not widely available, especially in resource-limited settings5.
Computed tomography (CT) has been extensively utilized and investigated for the diagnosis of hepatic steatosis and fibrosis. Conventional non-contrast computed tomography (NCCT) has demonstrated a well-established correlation with hepatic fat quantification, validated across multiple studies14,15. Notably, NCCT exhibits high reliability in detecting moderate-to-severe hepatic steatosis16 and are widely acquired for patients with various health problems. If NCCT scans could be adapted to accurately assess SLD and fibrosis risk, they would provide significant added value beyond their original diagnostic intent, enabling large-scale opportunistic liver disease screening in patients undergoing NCCTs. In contrast, postcontrast CT or contrast enhanced (CECT) poses significant challenges for accurate fat quantification17,18, primarily due to variability in imaging timing relative to contrast administration. Recent advancements in photon-counting CT technology, however, show promise in addressing these limitations through improved spectral resolution18. Despite these challenges, CECT remains widely employed in the evaluation of hepatic fibrosis. Several CT-based biomarkers have emerged as reliable indicators of advanced fibrosis and cirrhosis. These include hepatic texture analysis19, liver surface nodularity (LSN)20, and volumetric parameters such as liver-segmental volume ratios (LSVR) and splenic volume (SV) measurements21,22,23.
Despite encouraging advancements, significant challenges persist in achieving scalable and robust deep learning (DL) systems for simultaneous steatosis and fibrosis grading. First, scalability remains a critical limitation. While DL has been integrated into diagnostic workflows, most studies rely on conventional biomarkers—such as hepatic attenuation for steatosis quantification24,25 or combined volumetric parameters (e.g., liver-segment volume ratio [LSVR], splenic volume [SV], and attenuation deviations) for fibrosis detection21. However, limited research has focused on developing end-to-end frameworks capable of directly mapping CT images to dual diagnostic outcomes (steatosis and fibrosis) without the conventional biomarkers. For instance, Choi et al.26 proposed a 3D convolutional neural network trained on a large-scale dataset but primarily addressed general liver disease etiologies, omitting in detecting SLD patients and SLD with risk of fibrosis progression. Additionally, as highlighted in a recent review27, the scarcity of large-scale datasets with biopsy-confirmed diagnoses—essential for histological validation—severely constrains model generalizability and clinical translation. Second, robustness requires further refinement. Existing studies predominantly focus on isolated data modalities: either imaging biomarkers or serum/proteomic markers28,29,30. Yet, few investigate multimodal integration, despite evidence that combining imaging with clinical data enhances diagnostic accuracy. Composite indices like the FibroScan-AST (FAST) score and magnetic resonance elastography–fibrosis biomarker (MEFIB) index31,32 exemplify the synergistic potential of multimodal approaches. Similarly, integrating CT-derived imaging biomarkers with clinical features, such as serum data, could improve SLD detection and fibrosis progression prediction.
In this study, we sought to develop and validate an end-to-end multi-modal deep model, named Multi-modal AI for Opportunistic hepatic Steatosis Screening (MAOSS), that enables flexible integration of 3D NCCT images, conventional imaging biomarkers, and serum biomarkers (Fig. 1a) for predicting and grading hepatic steatosis and fibrosis. MAOSS was trained on a large-scale, heterogeneous dataset comprising pathologically-confirmed SLD and radiology-reported SLD cases, and validated across multi-centers and real-world datasets. Additionally, we aimed to conduct an AI-in-the-loop reader study to assess the impact of this model on radiologists’ diagnostic accuracy.
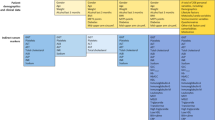
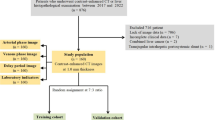
a MAOSS was developed using diverse data modalities, including CT scans, demographic information, physical exams, and serum data. We learn from CLD patients not only with histologically proven steatosis scores but also from a much larger and more accessible general population with radiologically reported steatosis. b Development and evaluation datasets. We trained the MAOSS on single-center data and evaluated its performance on multi-center data including histology or MRI-PDFF proved validation (SHCMU, NDTH and FAHZU) and real-world scenarios (UNIFESP and SIPD). c We integrate MAOSS into the standard of care workflow to stratify patients with steatotic liver disease at risk of progression to more serious liver conditions, such as steatohepatitis and advanced liver fibrosis. Created in BioRender. Li, c. (2025) https://BioRender.com/dtsbzp5.
Finally, by aligning the steatosis and fibrosis staging capability with clinical care pathways, our model not only classifies SLD patients correctly but also stratifies their risk for progression of serious CLDs, such as steatohepatitis, and advanced liver fibrosis.
Results
MAOSS Leverages the power of multimodal data to differentiate hepatic steatosis, using a NCCT volume with cropped liver and numerical data (allowing missing features) as inputs. It outputs a score, named the MAOSS score, to characterize the steatosis grade. We proposed a learning framework (Fig. 8), consisting of an ordinal regression heads designed to learn steatosis and fibrosis grading from liver biopsy-derived histology (gold standards), and a curated distillation head that distills knowledge from the steatosis grades recorded on radiological reports (non-gold standards). Radiology reports indicate four steatosis grades (none, mild, moderate, and severe) based on attenuation values12. Together, these components cover a wide spectrum of study population including those with other CLDs, making MAOSS a generalizable tool for opportunistic screening of SLD patients in a variety of real-world scenarios.
We created three separate models: (i) Unimodal-clinical: the unimodal-clinical model used only scalar-valued clinical variables from demographics, serum laboratory tests, physical testing, and CT biomarkers (supplementary Section 3 for detailed neumerical features) and project each variable into a feature embedding with a linear layer. Then, the feature embeddings are processed with a transformer backbone. Missing features were imputed using the multivariate imputation by chained equations method with predictive mean matching33. (ii) Unimodal-image (image-only model): An image encoder (Fig. 9a) was designed for extracting texture and image features from 3D liver NCCT. The extracted image patch tokens were processed with a transformer backbone to predict steatosis grades. (iii) MAOSS (multimodal model): This framework (supplementary Section 3) used both the imaging and numerical features as inputs and processed them with a transformer structure with interleaved multi-head self-attention blocks and our proposed Missing-Aware Modality Alignment (MAMA) module (Fig. 9b). The MAMA module projects multimodal features into a joint space and predicts the steatosis and fibrosis grade. Further details of model design can be found in the Methods section.
In our experiments, patients from SHCMU (n = 968) with gold standards and patients from SHCMU (n = 1 103) with non-gold standards were included as the development dataset. We conducted a five-fold cross-validation on the subset of gold standards with 80% data for training and 20% data for validation in each fold. MAOSS’s performance for steatosis grading was evaluated on an independent gold-standard internal test set of patients (n = 191) and a MRI-PDFF test set of patients (n = 375) from SHCMU, and an external gold-standard test set of patients (n = 347) from NDTH and FAHZU combined (Fig. 1b), and a multicenter CECT dataset which contains paired precontrast phase i.e. NCCT and portal venous phase scans of patients (n = 122). Both the internal and external datasets were multimodal and contained both imaging and numerical data with histology-derived gold standards for patient’s steatosis grades. To further test MAOSS’s generalizability for an opportunistic screening, we evaluated MAOSS on a public dataset of patients (n = 226) from the UNIFESP and a large-scale real-world dataset of patients (n = 18, 504) collected from different clinical settings in SIPD. The real-world datasets represented the general population and contained single-modality image data only. Histology-derived steatosis grades were not available for this cohort (Table S1). A total of 11 radiologists, including junior, mid-level, and senior professionals, were involved in assessing whether MAOSS could match or enhance their performance in differential diagnosis of hepatic steatosis. The reader study was conducted on the internal gold-standard test set (n = 191). Finally, we integrated MAOSS into the standard clinical pathway ((Fig. 1c)) for evaluating its capability for risk stratification and management of patients with SLD.
MAOSS diagnostic performance for evaluating hepatic steatosis and fibrosis in the internal and external tests
The MAOSS AUCs for evaluating liver steatosis S0 vs ≥S1, ≤S1 vs ≥S2 and ≤S2 vs S3 were 0.917 (95% CI: 0.866-0.958), 0.922 (95% CI: 0.879-0.958), and 0.923 (95% CI: 0.871-0.969) in the internal test, and 0.904 (95% CI: 0.866-0.933), 0.934 (95% CI: 0.912-0.954), and 0.963 (95% CI: 0.943-0.979) in the external test. For evaluating liver fibrosis, AUCs for F0-1 vs ≥F2, ≤F2 vs ≥F3 and ≤F3 vs F3 were 0.888 (95% CI: 0.841-0.931), 0.912 (95% CI: 0.868-0.953), and 0.941 (95% CI: 0.906-0.969) in the internal test, and 0.824 (95% CI: 0.777-0.866), 0.905 (95% CI: 0.850-0.954), and 0.930 (95% CI: 0.855-0.990) in the external test. Corresponding sensitivity, specificity, accuracy, positive and negative predictive values were shown in supplementary Tables S4 and S16. The metrics for 90% sensitivity and specificity were summarized in supplementary Tables S5–S7 and S17–S18. Of note, for diagnosing hepatic steatosis (S0 vs ≥S1), MAOSS demonstrated high specificity for steatosis detection, ranging from 90.8% to 100.0%, with sensitivity between 74.7% and 79.0% across internal and external validation cohorts. In contrast, fibrosis diagnosis ("F0-1 vs ≥F2”) proved more challenging, with specificity ranging from 82.7% to 92.9% and sensitivity between 69.3% and 72.7% for detecting clinically significant fibrosis, at the optimal cutoff.
MAOSS scores were discriminative between the steatosis grades in both internal and external tests, as shown in supplementary Fig. S7. MAOSS was also validated against MRI-PDFF (supplementary Tables S4 and S7) and demonstrated a good generalization with AUC of 0.929 (95% CI: 0.900-0.955), 0.930 (95% CI: 0.892-0.965), and 0.987 (95% CI: 0.975-0.995) for evaluating hepatic steatosis S0 vs ≥S1, ≤S1 vs ≥S2, and ≤S2 vs S3, respectively. We also trained MAOSS on a subset of the development set comprising patients with CECT scans, specifically selecting the portal venous phase for training and testing, and compared it with the NCCT-trained MAOSS. Table 1 summarizes the results, demonstrating that MAOSS reliably performs on both NCCT and CECT. However, NCCT shows superior performance for assessing moderate and above steatosis (AUC: 0.916 (95% CI: 0.850–0.970) vs. 0.831 (95% CI: 0.755–0.902), p < 0.001) and advanced fibrosis (AUC: 0.856 (95% CI: 0.776–0.927) vs. 0.835 (95% CI: 0.754–0.903), p < 0.01).
Comparison of diagnostic performance between MAOSS, unimodal-clinical, unimodal-image models, and transient elastography
Performance of MAOSS, and the three compared models, i.e., unimodal-clinical, unimodal-image, and transient elastography-controlled attenuation parameter (TE-CAP) for evaluating hepatic steatosis S0 vs ≥S1, ≤S1 vs ≥S2, and ≤S2 vs S3 was shown in Fig. 2. For diagnosing moderate to severe hepatic steatosis ≤S1 vs ≥S2, MAOSS’s AUC is also superior to the three compared models in both the internal and external testing (0.922 (95% CI: 0.879-0.958) vs. 0.886 (95% CI: 0.833-0.932), 0.913 (95% CI: 0.867-0.949), and 0.841 (95% CI: 0.777-0.894), p = 0.007, 0.237, and 0.025, respectively, for internal testing; and 0.934 (95% CI: 0.912-0.954) vs. 0.886 (95% CI: 0.856-0.916), 0.916 (95% CI: 0.889-0.941), and 0.867 (95% CI: 0.834-0.901), p < 0.001, =0.001, and < 0.001, respectively, for external testing). For diagnosing severe hepatic steatosis ≤S2 vs S3, MAOSS AUC was not significantly higher than those of the three compared models (p =0.13, 0.203, and 0.136, respectively) in the internal testing but significantly higher than those of the three compared models (p < 0.001, 0.019 and < 0.001, respectively) in the external testing (Supplementary Table S30). For diagnosing advanced fibrosis to cirrhosis (≤F2 vs ≥F3), MAOSS demonstrated superiority over single-modality models and transient elastography–liver stiffness measurement (TE-LSM), as shown in Fig. 3, in both internal and external testing. In internal validation, MAOSS achieved an AUC of 0.912 (95% CI: 0.868–0.953), outperforming single-modality models (0.844 (95% CI: 0.788–0.895), 0.759 (95% CI: 0.686–0.825), and TE-LSM 0.749 (95% CI: 0.680–0.818))(p <0.001, p <0.001, and p <0.001, respectively). In external validation, MAOSS yielded an AUC of 0.905 (95% CI: 0.850–0.954), surpassing single-modality models (0.858 (95% CI: 0.798–0.907), 0.763 (95% CI: 0.705–0.817), and TE-LSM 0.738 (95% CI: 0.687–0.787)) (p <0.001, p =0.012, and p <0.001, respectively), as detailed in Supplementary Table S31.
Receiver operating characteristic (ROC) analysis of compared models for classification of dichotomized histologically determined hepatic steatosis grades: a on internal test dataset and c. on external test dataset; Individual classification metrics (BACC, F1, Sensitivity, Specificity) were calculated based on 1000 bootstrap replications of the test datasets and reported with mean and 95% CI for compared models: b on internal test dataset and d on external test dataset. BACC: balanced accuracy, F1: F1 score, CI: confidence interval, TE-CAP: transient elastography-controlled attenuation parameter. For more details, please check supplementary Tables S4--S10. Source data are provided as a Source Data file.
Receiver operating characteristic (ROC) analysis of compared models for classification of dichotomized histologically determined hepatic fibrosis grades: a on internal test dataset and c. on external test dataset; Individual classification metrics (BACC, F1, Sensitivity, Specificity) were calculated based on 1000 bootstrap replications of the test datasets and reported with mean and 95% CI for compared models: b on internal test dataset and d on external test dataset. BACC: balanced accuracy, F1: F1 score, CI: confidence interval, TE-LSM: transient elastography-liver stiffness measurement. For more details, please check supplementary Tables S16–S21. Source data are provided as a Source Data file.
Ablation study
We investigate the effectiveness of the main components on the validation set by excluding one of them from the full setting of MAOSS. Table S2 (in supplementary) shows that boosted distillation significantly improves the model performance in each group and overall it brings about 2% and 1.4% increments in mean-BACC and mean-AUC. Random mask of image tokens only gives a moderate increase in mean-AUC but the increase is barely observed in mean-ACC. By removing numerical features, the single modal setting learned with images achieves an mean-BACC and mean-AUC of 85.3% and 89.2% which shows about 0.8% and 1% drop in mean-BACC and mean-AUC respectively, compared with multi-modal learning. Lastly, we found that texture encoding is especially important for improving the detection in the early stage of steatosis i.e. mild-moderate. With texture encoding, it increases the BACC by approx. 3% and 1.8% in identifying mild and moderate steatosis, respectively. To interpret the learned models, we utilize t-SNE34. Figure S4 (in supplementary) display the full setting of MAOSS, we observe that the learned ordinal regression and curated distillation tokens converge towards different vectors, indicating they have learned distinct distribution of the data. This aligns with our expectations since they were trained with the GS and NGS datasets, respectively. By averaging the token embeddings i.e., \(\frac{1}{2}{{\rm{x}}}_{ord}+\frac{1}{2}{{\rm{x}}}_{dist}\), we note the distribution of the joint representation with well-defined and -separated clusters with clearer decision boundaries compared to either of the tokens standing alone.
Comparision with other state of the arts
We compare with three groups of methods: 1) imputing-based methods that are trained with biometrics data in a single modal setting where two classic methods are compared: Multiple Imputation by Chained Equation (MICE)33 and Imputation by K-nearest neighbor35. 2) We compare state-of-the-art image-based methods trained in a single modality setting, including the classic CNN model ResNet36 and the transformer-based model DeiT37. Additionally, we evaluate the texture encoding method, DeepTen38. 3) multi-modal learning methods with missing modalities that are trained on image and biometrics together. MultiPrompt39 is a strong baseline to investigate missing-aware prompt tuning on pre-trained models and we compare it with the input prompt setting in our work. In supplementary Table S3, we observe that multimodal-learning methods overall outperform the single modal learning with imputaion by a large margin, which is as expected since rich information in the image representation is leveraged by joint learning with clinical features. Our proposed method surpasses the strong multimodal baseline presented by ref. 39 on both internal and external tests, demonstrating its superior capability in managing missing modalities and leveraging the relationship between image and clinical features. Notably, due to the unavailability of clinical features in the MRI-PDFF dataset, both our method and the baseline achieve similar AUCs across each group. This underscores the critical importance of clinical features in enhancing model performance. Without these features, both our approach and the baseline are reduced to relying solely on image data, thereby limiting their performance.
Model interpretation
To interpret the learning outcomes of MAOSS, we employ an attribution technique called Integrated Gradients (IG)40. This machine learning-based method is designed to visualize the influence of different input components on the model’s output using heatmaps, where larger values indicate a higher degree of influence. By attributing the output MAOSS score to the input 3D-liver image, IG allows us to visualize the specific alterations caused by steatosis in the liver at the voxel level. Additionally, it demonstrates that our MAOSS model has effectively learned the features associated with steatosis. Given that CT attenuation is a critical feature associated with steatotic changes, we examined the distribution of liver attenuation and integrated gradients. Figure 4a present the density plots for liver attenuation (Hounsfield units) and IG values across different steatosis grades (S0-S3), demonstrating a clear stratification among these grades. Notably, the IG values exhibit distinctive distributions across different steatosis grades, with severe steatosis (S3) showing a distinct peak and narrower spread compared to other grades. This indicates that the IG method effectively captures variations in voxel intensities associated with different degrees of hepatic steatosis. The scatter plot illustrates the joint distribution of liver attenuation and IG, which demonstrates an exponential correlation of IG values and liver attenuation values. Figure 4b provides an intuitive spatial mapping of IG on 3D liver images for representative cases across various steatosis grades, validated against corresponding MRI-PDFF measurements. As depicted, with increasing fat infiltration, the IG exhibit increasingly heterogeneous patterns, with elevated values localized in regions of pronounced fat accumulation. The spatial distribution of fat accumulation closely aligns with the MRI-PDFF reference, not only in uniformly distributed steatosis cases (Fig. 4b) but also in non-uniform steatosis cases (Fig. 4c). This observation suggests that MAOSS is capable not only of distinguishing between steatosis grades but also of effectively identifying regions of interest where steatotic alterations occur within the liver.
a Density plots show histologically-derived testset (internal and external) distribution of mean liver attenuation (in Hounsfield units), mean liver integrated gradients (IG), liver attenuation against integrated gradients for whole liver, showing MAOSS learned an exponential association between IG and liver attenuation with distinctive joint distribution. b The 3D saliency mapping of IGs overlaid on the cropped liver region in NCCTs from different viewpoints demonstrates that IGs spatial distribution in correlation with steatotic alternation on MRI-PDFF. Patient A: 66 year-old male with overall liver MR-PDFF 2% without steatosis. Patient B: 57 year-old female with MR-PDFF 14% (mild-moderate steatosis). Patient C: 53 year-old female with MR-PDFF 37% (severe steatosis). c The saliency map of MAOSS matches findings of steatotic (red arrows) and non-steatotic areas (white arrows) on MRI-PDFF maps in the non-uniformly distributed steatosis cases. None (S0), mild (S1), moderate (S2), and severe (S3) steatosis.
MAOSS performance for real-world opportunistic screening
Figure 5 demonstrates the performance of MAOSS in opportunistic screening on a large-scale real-world NCCT dataset. We collected real-world data from a public source (UNIFESP) as well as in a hospital with inpatient, outpatient, emergency, and physical examination cohort. Overall (Fig. 5a), the physical examination scenario exhibited the highest volume of total scans (n = 6 051), followed by emergency (n = 4 546), outpatient (n = 4 231), and inpatient (n = 3 450). The public dataset UNIFESP has the fewest number of scans (n = 299). The distribution of chest and abdominal scans also varied significantly across the considered clinical scenarios. For instance, nearly 98.9% of the scans (n = 5 985) in the physical examination scenario were chest scans; similarly, chest scans were dominant in the other scenarios. The results (Fig. 5b, c) illustrate the concordance between MAOSS and radiological reports for non-hepatic steatosis (Non-HS) and hepatic steatosis (HS) classifications. We found a high concordance rate between the HS cases reported by MAOSS and radiological reports across different scenarios. The concordance rate i.e. recall (HS) in physical examination, inpatient, outpatient, emergency, and UNIFESP was 96.6% (821 out of 850 scans), 93.5% (272 out of 291 scans), 92.8% (450 out of 485 scans), 92.3% (632 out of 685 scans), and 92.1% (186 out of 202 scans), respectively. For non-HS identification, there was also an almost perfect agreement between MAOSS and radiological reports: the concordance rate i.e. precision (Non-HS) in physical examination, inpatient, outpatient, emergency, and UNIFESP was 99.4% (4573 out of 4602 scans), 99.3% (2544 out of 2563 scans), 98.9% (3274 out of 3309 scans), 98.3% (2985 out of 3038 scans), and 82.4% (75 out of 91 scans), respectively. Cohen’s kappa reflected the overall inter-reader agreement between MAOSS and radiological reports, varying across different scenarios ranging from 0.383 (inpatient) to 0.704 (UNIFESP). Most disagreements occurred in cases where radiologists reported non-HS scans, but MAOSS identified mild steatosis in some of them. Radiologists appeared to be less sensitive to mild steatosis (see next session) and might have misreported these cases, which could account for the observed discrepancies. We bootstrapped test samples in each scenario 1000 times and computed the MAOSS AUC. Figure 5d showed that the distribution of AUCs was notably high across all scenarios, indicating excellent discriminatory power. Particularly, the physical examination scenario showed the highest AUC, suggesting the superior performance of MAOSS screening in the general population. When stratified by scan type (Fig. 5e, f), chest CT scans yielded higher AUCs or Cohen’s kappa compared to abdominal scans in most scenarios. Overall, MAOSS demonstrated reliability for examining NCCTs across multiple real-world clinical scenarios, with moderate to substantial inter-reader agreement with radiologists’ reports. This affirms its potential utility in augmenting radiological assessments in clinical practice.
a distribution of NCCT scans (chest/abdomen) collected from different real-world scenarios and prevalence of hepatic steatosis (HS). b Contingency tables for comparing the MAOSS diagnosis with the radiologist’s diagnosis. c Recall and precision of MAOSS. d Overall area under ROC curve (AUC) of MAOSS under different real-world scenarios using radiology report as reference. e AUC and f Cohen’s kappa, measuring the inter-reader agreement between MAOSS and radiology report, are stratified by the scan type. AUCs and Cohen’s kappas were calculated based on 1000 bootstrap replications of the test datasets and reported with median, IQR: 25th-75th percentile and whisker bounds. Source data are provided as a Source Data file.
MAOSS-augmented reader study for improving radiologists’ assessment of hepatic steatosis
We invited radiologists (n = 11) at different levels of the profession to differentiate the steatosis grades on NCCTs with and without the assistance of MAOSS. In the multi-reader multi-case study (Fig. 6a), radiologists with the assistance of MAOSS had significantly higher AUC than radiologists’ standalone diagnoses in two conditions: S0 vs ≥S1 (0.798 (95% CI: 0.741–0.855] vs 0.709 [0.70–0.749]; p < 0.001), and ≤S1 vs ≥S2 (0.851 [95% CI: 0.806–0.895] vs 0.821 [0.774–0.870]; p = . 043). Similar results were observed across all radiologist’s subgroups (Figs. S10–S12). MAOSS assistance was not significant for improving the radiologist’s diagnosis (p = 0.494) of severe steatosis ≤S2 vs S3, as severe steatosis can be easily diagnosed on NCCT.
a Primary comparison of the area under ROC curves (AUCs) in the multi-reader multi-case study: Delong tests were performed and a two-sided p value less than 0.05 indicates the significant difference between the compared groups. b MAOSS ROC curves (mean and 95%CI band) and individual reader operation point when staging HS at different stages. c Individual reader balanced accuracy with or without MAOSS assistance. d Balanced accuracy of grouped readers, both with and without MAOSS assistance, was reported using the mean and standard deviation (SD) of individual’s balanced accuray across all readers (Average, n = 11), mid-seniors (n = 6), and juniors (n = 5). Source data are provided as a Source Data file.
Figure 6b shows the MAOSS ROC curves with 95% CI and individual radiologist performance for grading steatosis with and without the assistance of MAOSS. We found that without the aid of MAOSS, radiologists generally had very high specificity of diagnosis regardless of their levels of the profession. Still, their sensitivity was insufficient for differentiating steatosis at different stages. With the assistance of MAOSS, radiologists’ sensitivity was generally improved by a large margin while maintaining a high specificity. Overall, mid-senior groups perform generally better than the junior groups with the aid of MAOSS for diagnosis. Figure 6c shows the balanced accuracy for the individual radiologists. An improvement in balanced accuracy with the aid of MAOSS was observed for each of the 11 radiologists, ranging from 1.8% to 16.5%, 1.2% to 18.1%, and 1.9% to 14.9% for differentiating S0 vs ≥S1, ≤S1 vs ≥S2 and ≤S2 vs S3, respectively. Figure 6d grouped the balanced accuracy by levels of radiologist’s profession. Generally, the trend of balanced accuracy for grouped radiologists was similar to the results observed for individual performance.
Primary risk stratification of patient identified with hepatic steatosis
To evaluate the potential of MAOSS to transform the current standard of care of steatosis patients, we embed MAOSS into the American Association for the Study of Liver Diseases (AASLD) clinical practice guidance41 for screening and referring patients established with steatosis who may be at risk of progression to more serious CLDs. We also compare it with the standard clinical flow that uses transient elastography. Of note, although the emphasis of risk stratification in the primary care setting is to exclude advanced fibrosis i.e., ≥F3 with a high negative predictive value, it is also of great significance for identifying patients with at-risk steatohepatitis i.e., NAFLD activity score (NAS) ≥4 and clinically significant liver fibrosis ≥F2, who may benefit from a therapeutic intervention29,41.
We established a large cohort of 1192 patients (see method section) in which liver biopsy confirmed 145 patients with NAS≥4 and fibrosis ≥F2, i.e., at risk of steatohepatitis progression. The rest 1,047 patients are at low risk. We compared the AASLD standard practice, referred to as the AASLD pathway, with our proposed MAOSS pathway (Fig. 7a and supplementary Fig. S13) for evaluating patients with hepatic steatosis who are at risk of steatohepatitis progression. The proportion of intermediate-high risk populations identified by AASLD pathway and MAOSS pathway (Fig. 7b) was 6.9% (82/1192) and 21.9% (261/1192), respectively. Figure 7c demonstrates that MAOSS pathway (MAOSS + FIB-4 + FAST) exhibited a sensitivity of 52.4% (95% CI: 44.0–60.8%), outperformed the 16.6% (95% CI: 10.6–22.8%) sensitivity observed for AASLD pathway (CAP+FIB-4+LSM) by a large margin (p < 0.001). The corresponding negative predictive value for the MAOSS pathway was 92.6% (95% CI: 90.9–94.3%) compared to 89.1% (95% CI: 87.4–90.9%) for AASLD pathway (p = 0.003). We also found that the MAOSS pathway improved the odds (supplementary Table S35) for identifying patients at risk of steatohepatitis (odds ratio 5.12 (95% CI: 3.51-7.49); p < 0.001) compared to the AASLD pathway (odds ratio: 3.38 (95% CI: 1.93-5.76); p < 0.001). These results verified that the MAOSS pathway was superior to the AASLD pathway for identifying steatosis patients who are at risk of steatohepatitis progression.
a Current AASLD clinical workflow41 for screening steatotic liver disease patients who are at risk of progression (AASLD pathway), with MAOSS embedded (MAOSS pathway). b SLD patients stratified into low, intermediate and high risk with AASLD/MAOSS pathway. c Comparison AASLD and MAOSS pathway for identifying SLD patients at risk of fibrosis progression. Competing risks analysis shows the cumulative incidence of a subgroup of patients (n = 236) who developed cirrhosis at the end (n = 39). Kaplan Meier plots. d Show stratification of patients according to the AASLD pathway. e show stratification of patients according to the MAOSS pathway. Shaded areas represent 95% CIs for the cumulative incidence curves. Gray’s test was used to assess differences in rates of cirrhosis incidence between groups. Two-sided p value less than 0.05 was considered statistically significant. Source data are provided as a Source Data file. Created in BioRender. Li, c. (2025) https://BioRender.com/ohxalky.
Identifying patients at-risk of advanced liver fibrosis
A total of 122 patients, accounting for about 10% (122/1192) of the whole screened population, were confirmed with liver biopsy who developed advanced liver fibrosis ≥F3. Again, we applied the AASLD and MAOSS pathway for a risk stratification to detect patients at risk of advanced fibrosis. The results were summarized in supplementary Table S37. We found the MAOSS pathway again was significantly superior to the AASLD pathway for identifying patient at risk of advanced fibrosis. The sensitivity of the MAOSS pathway was 59.1% (95% CI: 50.7–67.4%), significantly higher than that of the AASLD pathway’s 23% (95% CI: 15.6–30.5%) (p < 0.001). MAOSS pathway also demonstrated a superior (p = 0.002) capability for excluding advanced fibrosis patients with a higher NPV of 94.6% (95% CI: 93.2–95.9%), compared with that of the AASLD pathway’s 91.5% (95% CI: 89.9–93.0%).
Association between different risk stratification pathways and incidence of hepatic cirrhosis
We followed up a subgroup of patients (n = 236) from the risk stratification cohort with liver fibrosis stages F0-F3 at baseline. Among these, 39 (16.5%) progressed to cirrhosis during follow-up. The progression rates to cirrhosis were 9.6% (8/83) in F0, 10.2% (9/88) in F1, 30.8% (12/39) in F2, and 38.5% (10/26) in F3 cohorts, during a median follow-up of 2.3, 2.2, 1.7, and 2.1 years, respectively (supplementary Figs. S14 and S16). Cause-specific proportional subhazards analysis, based on the Fine and Gray method42, was used to assess the association between the different risk stratification pathways (AASLD and MAOSS pathways) and the time to developing cirrhosis, accounting for competing events. We conducted an ablation study on both pathways to investigate the single and combined predictors for identifying at-risk patients. The median duration of follow-up was 2.3 years (IQR, 1.7–3.3 years) in the low-risk group and 1.6 months (IQR, 0.2–2.8 months) in the intermediate-high-risk group when stratified using the MAOSS pathway. In these groups, 24 out of 203 patients in the low-risk group and 15 out of 33 patients in the intermediate-high-risk group progressed to cirrhosis, corresponding to cirrhosis rates of 11.8% and 45.5%, respectively. Figure 7d and e depicted the cumulative incidence of hepatic cirrhosis across risk groups, showing a significantly longer time to cirrhosis in the predicted low-risk group compared to the intermediate-high risk group (AASLD pathway, p = 0.026; MAOSS pathway, p < 0.001). In univariable Cox regression analysis, intermediate-high risk identified by the MAOSS pathway was associated with a higher incidence of cirrhosis (hazard ratio, 5.54; 95% CI: 2.69-11.42; p < 0.001). Multivariable Cox regression analysis also revealed that intermediate-high risk identified by the MAOSS pathway independently predicted subsequent cirrhosis in baseline patients (hazard ratio, 5.56; 95% CI: 2.41-12.81; p < 0.001) (Supplementary Tables S38 and S39).
Discussion
We introduce an end-to-end multimodal deep learning model named Multi-modal AI for Opportunistic Hepatic Steatosis Screening (MAOSS). This model seamlessly integrates data from 3D NCCTs, conventional imaging biomarkers, and serum biomarkers to detect and grade the severity of hepatic steatosis. Recognizing the potential unavailability of serum biomarkers for some patients, MAOSS is designed to accept flexible input, accommodating missing features. Traditionally, diagnosing steatosis and fibrosis with NCCTs has depended on conventional biomarkers, which often lack accuracy. Given the limited availability of the gold standard (liver biopsy), there have been few attempts to build deep learning models, and the impact of multimodal representation on diagnostic accuracy remains under-explored. MAOSS leverages data from patients with CLDs, where gold standards are available, and enhances model performance with diagnostic information from broader and more accessible radiological reports. We extensively validated MAOSS on both histologically and MRI-PDFF-confirmed steatotic liver disease patients, achieving high performance with an AUC of 0.904-0.929 for detecting mild to severe steatosis—significantly surpassing standalone radiologist performance (AUC 0.709) and diagnosis with MAOSS assistance (AUC 0.798). Beyond its performance, MAOSS provides explainability through Integrated Gradients, aligning steatotic alterations with MRI-PDFF mappings. Furthermore, by integrating MAOSS with standard clinical workflow, we conducted a simulated primary risk screening on a large retrospective CLD cohort. Compared to AASLD risk stratification pathway, our optimized approach identified significantly more SLD patients at risk of steatohepatitis and advanced fibrosis while maintaining a high negative predictive value for excluding low-risk patients.
MAOSS demonstrates high performance in large-scale, gold-standard-validated datasets for differentiating steatosis, outperforming conventional CT-based markers by a significant margin (as detailed in supplementary Table S28) and potentially matching the accuracy of advanced proteomics approaches. Previous studies have demonstrated the potential of NCCT scans being used for opportunistic screening of hepatic steatosis in large population24. However, one limitation of these NCCT studies is that they were not thoroughly validated against gold standards, and their accuracy remains uncertain. Until a recent radiomics study, LARI43, developed logistic regression models that investigated conventional CT biomarkers and shape- and texture-based radiomics features. These models were validated against both proton magnetic resonance spectroscopy (1HMRS, n = 350) and liver biopsy gold standards (n = 159). The study demonstrated strong discriminative performance for detecting hepatic steatosis (S0 vs ≥S1), with AUCs ranging from 0.82 to 0.91 when validated against 1HMRS and from 0.71 to 0.76 when validated against liver biopsy. Another deep learning study on ultrasound images44 validated against with a small gold standard dataset (n = 199) and reported AUCs of 0.85, 0.73 and 0.67 in differentiating steatosis for S0 vs ≥S1, ≤S1 vs ≥S2 and ≤S2 vs S3. In addition to imaging, non-imaging data also shows great promise for identifying steatosis. A study published in ref. 30 investigated proteomic markers for detecting steatosis and demonstrated excellent discriminative ability for differentiating between S0 and ≥S1, with an AUC of 0.91 based on a small dataset referenced against histology gold standards. Despite the potential of proteomics for diagnosing steatosis, it is not an ideal tool for large-scale screening due to limited scalability and high costs. Other studies45 have attempted to combine imaging data with more easily acquired serum markers, showing very promising results, especially in improving the detection of mild steatosis. These findings motivated us to develop and validate an AI tool, MAOSS, which achieves high precision for screening steatosis in large populations using easily accessible data modalities, specifically NCCT scans and serum tests. We have extensively validated MAOSS using both liver biopsy and non-invasive MRI-PDFF gold-standard datasets. The results demonstrated high accuracy in detecting steatosis for S0 vs ≥S1, achieving AUCs ranging from 0.90 to 0.92 in internal and external liver biopsy validations, and an AUC of 0.93 in MRI-PDFF validation. This performance is on par with proteomics AUC 0.9130 and outperforms the state-of-the-art radiomics model LARI43- AUC 0.76, and notably MAOSS leverages a much larger scale of gold standards validation. Additionally, we compared MAOSS with conventional CT-based markers used in clinical practice, assessing steatosis through liver attenuation, liver-to-spleen ratio, and liver-to-spleen difference, as well as fibrosis-related markers like liver segmental volume ratio (LSVR), liver surface nodularity (LSN), and splenic volume (SV). We reported a logistic regression model trained on the conventional markers collectively (supplementary Table S28), achieving AUCs ranging 0.75-0.78 for differentiating S0 vs ≥S1 when validated against liver biopsy. This performance is comparable to the result reported by LARI43- AUC 0.71 and significantly higher than the AUCs of individual CT markers validated separately, which ranged from 0.55 to 0.67. Specifically, the results (in supplementary Table S26) show that MAOSS has significantly higher balanced accuracy (internal: 87.5% [95% CI: 84.3–90.8%], external: 88.0% [95% CI: 84.0–91.0%]) compared to these conventional CT-based markers in both internal (range from 59.1% to 61.9%) and external validation (range from 70.3% to 74.6%). This advantage extends to fibrosis assessment, as indicated in Supplementary Table S29, where MAOSS consistently achieves significantly higher AUCs compared to multivariable logistic regression models21 using conventional markers, regardless of fibrosis stage. Furthermore, incorporating LSN, LSVR, and SV substantially enhances MAOSS’s performance in identifying cirrhosis, as demonstrated in Supplementary Table S32. To evaluate the robustness of MAOSS various clinical and CT settings, we evaluated internally and externally the influence of the patient and CT characteristics on the performance of MAOSS for liver steatosis and fibrosis staging (Supplementary Tables S11–15 and S22–25) and also the real-world scenarios for liver steatosis staging (Supplementary Table S33). MAOSS demonstrated stable and generalizable performance, with no clinically relevant differences in model performance across key clinical and CT characteristics. MAOSS showed slightly better discrimination of liver fibrosis/cirrhosis in abdomen scans compared to chest scans. This may be attributed to the fact that chest scans often include incomplete segmentation of the liver and spleen, leading to inaccurate estimates of volumetric markers such as the LSVR and SV. Furthermore, thin slices (3 mm or less) were found to perform better than thick slices (5 mm) for detecting early-stage liver fibrosis, a finding consistent with those reported in a recent study46. This could be due to the higher spatial resolution provided by thin slices, which enhances the detection of subtle structural changes in liver tissue architecture.
Our analysis reveals that radiologists’ diagnoses using conventional CT biomarkers lack sensitivity in detecting steatosis, whereas MAOSS demonstrates the potential to significantly improve diagnostic performance. Our findings indicate that when radiologists diagnose steatosis independently, they demonstrate a high specificity for diagnosing steatosis (≥S1) cases, with an average specificity of 99.5% ± 1.4%. However, the sensitivities for detecting steatosis (≥S1) remain low, with juniors achieving 41.0% ± 7.8% and mid-seniors 43.5% ± 6.2% (Supplementary Fig. S20). Despite the low sensitivities, these results still outperform diagnoses made using single CT-based biomarkers alone. Specifically (supplementary Table S26), liver attenuation achieved a sensitivity of 24.1% (95% CI: 17.8–30.6%) at 100% specificity, while the liver-spleen ratio had a sensitivity of 18.1% (95% CI: 12.7–23.9%) at the same specificity. The liver-spleen difference demonstrated significantly higher sensitivity at 80.6% (95% CI: 74.9–86.1%), but its specificity was markedly lower at 74.6% (95% CI: 53.6–93.3%). Radiologists, when collectively assessing these markers, achieve high specificity, but the sensitivity remains unsatisfactory. This implies a limitation in the current standard practice for diagnosing steatosis from CT scans, where conventional CT biomarkers are insensitive in differentiating between S0 and ≥S1. Park et al.47 proposed a 4-point visual grading criterion for scoring the steatosis grade on NCCTs and conducted a reader study with 6 radiologists involved and with the criterion, average AUC for differentiating the moderate to severe steatosis i.e., ≤S1 vs ≥S2 was ≥0.90 and the specificity for ruling out S0-S1 steatosis was high ranging 99.4%- 99.8% similar to our finding. However, this work did not investigate the more challenging differentiating mild-severe steatosis i.e., S0 vs ≥S1. We demonstrate for differentiating S0 vs ≥S1, MAOSS not only achieved a high AUC of 0.917 but also helps radiologists improve the AUC significantly, overall from 0.709 to 0.798 for a multi-reader multi-case study in all 11 radiologists with different levels of experience. Moreover, although MAOSS was trained using liver biopsy-derived steatosis grades, which may be subject to sampling error, we further validated the model on a large dataset (n = 375) with MRI-PDFF as the reference standard. In this context, MAOSS achieved a comparable AUC of 0.929 for differentiating S0 vs ≥S1, indicating its effectiveness and suggesting that it is not biased towards biopsy-derived gold standards.
The efficacy of MAOSS is further demonstrated through population screening using NCCT data collected from diverse clinical settings. Our findings underscore MAOSS’s potential as a highly accurate and scalable tool for the opportunistic screening of hepatic steatosis in large populations utilizing NCCT scans. Among the identified steatosis patients, we compared the performance of MAOSS with the Controlled Attenuation Parameter (CAP) obtained via transient elastography to further identify SLD patients at risk of progressing to more severe conditions such as steatohepatitis and advanced hepatic fibrosis. We proposed a clinical pathway, termed the MAOSS pathway, which integrates the MAOSS score with hepatic fibrosis-related biomarkers, specifically FIB-4 and FAST. This pathway was compared to the current standard of practice, referred to as the AASLD pathway, for primary risk screening. To validate the effectiveness of both the MAOSS and AASLD risk stratification pathways, we assembled a large cohort of CLD patients (n = 1192) with histologically proven stages of hepatic fibrosis and inflammatory activity. Of these, 236 patients with fibrosis stages F0–F3 were followed longitudinally to assess the development of cirrhosis. Our results indicate that the MAOSS pathway correctly referred a significantly higher number of patients at risk of steatohepatitis and advanced fibrosis compared to the AASLD pathway, while maintaining high specificity and a negative predictive value (NPV) for ruling out low-risk SLD patients. Notably, longitudinal analysis using Cox regression demonstrated that the MAOSS pathway has a generally high hazard ratio for predicting patients at intermediate risk of cirrhosis. To our knowledge, no previous studies have addressed longitudinal risk stratification for SLD patients at risk of progressing to cirrhosis, highlighting the clinical significance of our findings. Maria et al.48 investigated the identification of advanced fibrosis patients using FIB-4 and the Enhanced Liver Fibrosis (ELF) test. Their study demonstrated that the sequential application of FIB-4 followed by ELF identified a substantial proportion (sensitivity of 37%) of participants with advanced hepatic fibrosis requiring specialist care, while achieving an impressive NPV of 92%. Although this approach has been extensively studied across various populations (including the general population, individuals at risk for alcoholic liver disease, and those at risk for non-alcoholic fatty liver disease), its reference standard is problematic. The use of transient elastography (TE) with a Liver Stiffness Measurement (LSM) ≥8 kPa as the reference standard may not serve as a gold standard, thereby questioning its true performance in large-scale screening. Devon et al.29 explored the identification of steatohepatitis patients using traditional machine learning models combined with serum data, validated against histological gold standards. For identifying patients with NAS≥4 and fibrosis stage F2 or higher, their model achieved specificity around 86–87% and sensitivity ranging from 25% to 56% at a 90% specificity threshold. These metrics are comparable to the MAOSS pathway (specificity of 82.3% and sensitivity of 52.4%). However, the NPV exhibited large variation, ranging from 33% to 83%, and the accuracy was generally lower than that of the MAOSS pathway (78.7% compared to 55–75%). Chen et al.49 developed a deep learning model, FIB-NET, based on B-mode ultrasound images, which was combined with the serum biomarker FIB-4 and LSM in a sequential manner for screening patients with advanced hepatic fibrosis. Utilizing a two-step process—FIB-4 followed by the FIB-NET score—the model achieved 77% accuracy, 68% sensitivity, and 91% NPV in a large cohort of 1165 patients. A three-step process (FIB-4, FIB-NET, LSM) yielded 85% accuracy, 54% sensitivity, and 87% NPV in a cohort of 447 patients. These results are comparable to our MAOSS pathway, which demonstrated sensitivity and NPV of 59.1% and 94.6%, respectively, in a large cohort of 1192 patients.
While the MAOSS demonstrates substantial promise for the opportunistic screening and risk stratification of steatotic liver disease and its progression, several limitations must be acknowledged. First, the high prevalence of hepatic steatosis in the general population poses challenges for real-world evaluation. Specifically, it is impractical to retrieve and extensively evaluate all individuals identified by MAOSS who may be missed by human experts, potentially leading to underestimation of the model’s false-negative rate. Second, although the MAOSS pathway exhibits high specificity and negative predictive value, the sensitivity for risk stratification requires further enhancement. Improved sensitivity is crucial to ensure that a greater proportion of at-risk patients are accurately identified and can receive timely interventions to prevent progression to more severe liver conditions. Lastly, the longitudinal component of our study was limited by a relatively small cohort size. A larger longitudinal cohort with more uncensored cases would provide greater statistical power, enabling more definitive conclusions regarding the predictive capabilities of the MAOSS pathway. Future studies incorporating broader and more diverse longitudinal data are necessary to strengthen the evidence for MAOSS’s effectiveness in predicting disease progression.
Methods
Ethics approval
The retrospective collection of de-identified patient imaging and clinical data was obtained from internal and external study institutions, in accordance with ethical guidelines. Each participating institution secured ethics approval from their respective Institutional Review Board (IRB) and obtained a waiver for informed consent. The institutions involved included: the Shengjing Hospital of China Medical University (SHCMU) IRB, the Nanjing Drum Tower Hospital (NDTH) IRB, the First Affiliated Hospital of Zhejiang University (FAHZU) IRB, and the Shanghai Institution of Pancreatic Diseases (SIPD) IRB.
Study population
This multi-center retrospective study involved six patient cohorts (Table S1): an internal development cohort(n = 2071), on which MAOSS model was built; an internal test cohort (n = 191), on which the model performance and reader study for hepatic steatosis grading were assessed; a MRI-PDFF test cohort (n = 375) in which patients have paired NCCTs and MRIs; an external multi-center test cohort (n = 347), on which the generalization across multiple centers was assessed; a contrast-enahnced CT (CECT) test cohort (n = 122) in which patients have both non-contrast and contrast enhanced CTs; a real-world test cohort (n = 18,504), on which the generalization to large-scale real-world data for an opportunistic screening was assessed; and a risk-stratification cohort (n = 1192), on which patient’s with hepatic steatosis at risk of progression to more serious CLDs, such as steatohepatitis and advanced liver fibrosis, were assessed.
Internal development cohort
The internal development cohort was collected from SHCMU patients recruited between 2008 March and 2021 June, consisted of a gold-standard (GS) subset of 968 patients who have histology-derived steatosis severity (S0: 472, S1: 242, S2: 165, and S3: 89), fibrosis severity (F0-1: 546, F2: 130, F3: 47, and F4: 245), and a non-gold standard (NGS) subset of 1103 patients (none: 438, mild: 307, moderate: 112, and severe: 246) with radiologists’ reported steatosis severity.
Internal and external test cohort
The internal test cohort was collected from SHCMU and is composed of two groups. (a) The first group includes 191 patients recruited between July 2021 and April 2024, with histology-derived steatosis severity in 191 patients (S0: 20, S1: 82, S2: 57, and S3: 32), fibrosis severity (F0-1: 99, F2: 20, F3: 16, and F4: 56). (b) The second group includes 375 patients recruited between June 2022 and October 2024, with MRI-PDFF-derived steatosis severity in 375 patients (S0: 75, S1: 214, S2: 55, and S3: 31). The external test cohort was collected from NDTH and FAHZU, which involved 347 patients who had histology-derived steatosis severity (S0: 17, S1: 153, S2: 77, S3: 100), fibrosis severity (F0-1: 214, F2: 45, F3: 52, and F4: 35).
Contrast-enhanced CT (CECT) test cohort
The CECT test cohort was collected from multicenters including SHCMU, NDTH and FAHZU, which involved 122 patients recruited from May 2024 onwards, who had contrast enhanced CT scans and histology-derived steatosis severity (S0: 8, S1: 85, S2: 21, S3: 8), fibrosis severity (F0-1: 60, F2: 24, F3: 24, F4: 14). We selected non-contrast phase and portal venous phase scans for measurements.
Real-world cohort
The real-world cohort was a imaging-based dataset that combined a public dataset UNIFESP and data collected from SIPD. UNIFESP included 226 patients (none: 79, mild-severe steatosis: 147) who have undergone Chest or Abdomen CT scans. Data collected from SIPD involved patients from various scenarios, including physical examination 6051 patients (None: 5201, mild-severe steatosis: 850), inpatient dept. 3450 patients (none: 3159, mild-severe steatosis: 291), outpatient dept. 4231 patients (none: 3746, mild-severe steatosis: 485), Emergency dept. 4546 patients (None: 3861, mild-severe steatosis: 685) who have undergone CT scans.
Risk stratification cohort
The risk stratification cohort was a subset of SHCMU patients (n = 1192) recruited between 2008 March and 2021 June, who had complete set of liver function tests and VCTE, specifically for assessing if MAOSS can be embedded in the standard of care practice for identifying SLD patients who at risk of progression to more serious CLDs. We managed retrieved (n = 236) patients with their longitudinal record of disease progression and their clinical end that (n=39) diagnosed cirrhosis (supplementary Figs. S14–16 and Table S1).
Reference standard
Gold Standards (GS): liver biopsies were interpreted using NAS score50, including steatosis grade, ballooning grade, lobular inflammation grade, and fibrosis stage. Biopsies were performed via a transcutaneous approach with US guidance using 16- or 18-gauge core needles51. We extracted the grades from pathology reports, in which steatosis grade (S) assessed the percentage of hepatocytes containing large or medium-sized lipid droplets according to the following ordinal scale: none (< 5%, grade S0), mild (5–33%, grade S1), moderate (34–66%, grade S2), and severe (> 66%, grade S3). The MRI-PDFF derived steatosis scale was graded as none (< 5%, grade S0), mild (5%–15%, grade S1), moderate (15–25%, grade S2), and severe (≥25%, grade S3)12. The steatosis scores were dichotomized as: S0 vs ≥S1; ≤S1 vs ≥S2; and ≤S2 vs S344. Fibrosis stage is F0 (none fibrosis), F1 (perisinusoidal or periportal fibrosis), F2 (perisinusoidal and periportal fibrosis), F3 (bridging fibrosis), F4 (cirrhosis). And the fibrosis scores were dichotomized as: F0-1 vs ≥F2; ≤F2 vs ≥F3; and ≤F3 vs F4. Non-gold standards (NGS): for real-world NCCTs’ assessment, hepatic steatosis was diagnosed with the AASLD practice guideline52. Specifically, the following standards were considered: (1) the attenuation of the liver is at least 10 Hounsfield units (HU) lower than that of spleen i.e. liver-spleen HU < -10; (2)the liver/spleen HU ratio is ≤ 0.8; (3) the attenuation of the liver is < 40 HU. Specifically for SIPD data, each scan was examined by a two-round review process with a junior radiologist giving the initial assessment and a senior radiologist making the final report.
Inclusion and exclusion criterion
The inclusion criteria for the development and validation were patients suspected or confirmed with SLD who had undergone CTs and must have NCCT series and had undergone liver biopsy or MRI-PDFF within one month of each other, for the general population who has no history of CLD, must have at least one NCCT series and available radiological reports. The following patients were excluded: (a) those under 18 years of age, (b) those NCCTs missing or NCCT with quality issues e.g. artifacts (c) those with a history of liver transplant or treatment for liver cancer, and (d) those with focal liver lesions.
MAOSS model
Figure 8 illustrates the complete diagram of our proposed framework. The input data, i.e., liver CT image, biomarkers, and clinical factors, are first tokenized and linearly projected to the embedding space, with an image encoder (Fig. 9a) and biometric encoder respectively. The resulting tokens are concatenated and preattached with a learnable class token and a distillation token. All tokens are then fed into a ViLT53 like encoder consisting of multi-head self-attention (MSA) and MLP layers. We design a missing-aware modality alignment (MAMA) module (Fig. 9b) that is adaptable and pluggable and can be inserted anywhere inside a pre-trained multi-modal encoder.
a MAOSS takes as inputs the imaging and numerical features (allowing for missing features) and outputs the MAOSS score indicating the severity of hepatic steatosis. We propose a learning strategy that formulates the gold-standards learning (from histology results) as an ordinal regression problem and non-gold standards learning (from radiology report) with a curated distillation process. b A transformer consisting of Missing-aware Multimodal Alignment (MAMA) Adaptor and Multi-head Self-Attention (MSA) blocks, served as the scaffold for the model. Each feature was processed into a fixed-length vector using a modality-specific encoder and fed into the transformer as input. Three linear layers \({F}_{or{d}_{st}}\), \({F}_{or{d}_{fib}}\) and Fdist were used to process the data with gold standard ordinal regression and curate distillation accordingly, based on the annotation type either histology report (gold standards of steatosis and fibrosis grades) or radiological report (non-gold standards of steatosis grades). The final prediction was the average of the outputs from the \({F}_{or{d}_{st}}\) and Fdist heads for steatosis grading (MAOSS score) and \({F}_{or{d}_{fib}}\) for fibrosis grading (FIBRO score) at inference time. Supplementary material.
a Image Encoder Design detail. b Missing-Aware Modality Alignment (MAMA) Adaptor.
Numerical features used in MAOSS development
We divide the numerical features into three groups: demographics & physical data, laboratory results and CT-based biomarkers. Demographics & physical data used in the study includes: Age, Gender, Body mass index (BMI), Height, Weight. Laboraty results used in the study includes hematology, liver function tests, lipid profile, cardiac markers etc: White blood cells (WBC), Red blood cells (RBC), Hemoglobin (Hgb), Hematocrit (HCT), Platelet (PLT), Total protein (TP), Albumin (ALB), Albumin/Globulin (A/G), Aspartate aminotransferase (AST), Alanine transaminase (ALT), Gamma-glutamyl transferase (GGT), Alkaline phosphatase (ALP), Prealbumin (PALB), Cholinesterase (CHE), Total bilirubin (TBIL), Direct bilirubin (DBIL), Indirect bilirubin (IDBIL), Total bile acid (TBA), Creatine kinase (CK), CKMB (Creatine Kinase-MB), Lactate dehydrogenase (LDH), Hydroxybutyrate dehydrogenase (HBDH), Blood urea nitrogen (BUN), Uric acid (UA), Cystatin C (CysC), Blood cholesterol (CHOL), Triglycerides (TG), High-density lipoprotein (HDL-C), Low-density lipoprotein (LDL-C), Apolipoprotein A-1 (Apoa1), Apolipoprotein B-100 (apoB), Blood glucose (GLUO), Lipase (LiPA), Creatinine (CREA), Prothrombin time (PT), plasma thromboplastin antecedent (PTA), International normalised ratio (INR), Prothrombin time Test and INR (PT/INR), activated partial thromboplastin time (APTT), Fibrinogen (FIB), Thrombin time (TT), D-dimer (DD), Procollagen type III amino terminal peptide (PIIIP), Hyaluronic acid (HA), Laminin (LN), AST to Platelet Ratio Index (APRI), Fibrosis-4 (FIB-4) Index. We conducted a series of conventional CT-based biomarkers over both the liver and spleen. For deriving the liver regions of interest (ROIs), we extended the 2D biometry method54 to 3D for automatically assessing regional attenuation in the liver. Specifically, three spherical liver ROIs were generated using morphological operations. First, the 3D liver mask was eroded to locate the central slice, and then dilation was performed from the center in three directions - laterally, anterior-posterior- to define a central area of liver where major vessel structures are located. Thereafter, three spherical ROIs were placed between the central area and the original liver mask to avoid the hepatic vasculatures or other causes of heterogeneity. We computed the histogram of Hounsfield units (HU) in the three 3D ROIs and derived the liver attenuation by the mean HU across the three ROIs. We derived the liver-spleen ratio (LSR) by dividing the mean HU of the three liver ROIs by the mean HU of the spleen. Similarly, the liver-spleen difference (LSD) is determined by subtracting the mean HU of the spleen from the mean HU of the three liver ROIs. Furthermore, we trained a 3D deep learning model to segment the liver Couinaud segments using both the public dataset55 and our in-house annotated data. This enabled us to extract liver volumetric CT-based markers, such as the liver segmental volume ratio, LSVR (liver segmental volume ratio, LSVR = volumes of Couinaud segments I-III/IV-VIII) and liver segmental proportion21. Figure S5 (in supplementary) shows that the model segmented the liver Couinaud segments in various grades of liver steatosis and fibrosis. Additionally, we quantified liver surface nodularity (LSN) following the approach described in ref. 20. And splenic volumes (SV) were also measured using 3D spleen segmentation masks.
Numerical feature embeddings
We assign dummy inputs (filled with ones) to handle the missing feature. The numerical features are categorized into three groups: (1) Demographic and CT-biomarker features, (2) Serum and laboratory test features, and (3) Liver functional test and transient elastography features. A complete summary of the missingness statistics for these numerical features across different study cohorts is provided in Supplementary Figs. S1–S3. Min-max scaling was first employed to normalize every component of numerical features. A linear projection layer was applied to the features to acquire their embeddings, denoted as \({\{{x}_{n}^{nim}\in {{\mathbb{R}}}^{C}\}}_{n=1:{N}_{nim}}\) (C-embedding dimensionality, Nim - number of non-image features).
Imaging feature embeddings
We built the unimodal-image model with an image encoder and a transformer-based architecture (in Fig. 9a). It took the cropped liver image \({x}^{im}\in {{\mathbb{R}}}^{H\times W\times D}\) as input and fed it into a patch encoder (implemented with a 3D ResNet-34) to get 3D-patch embeddings \({\{{x}_{n}^{pat}\in {{\mathbb{R}}}^{C}\}}_{n=1:{N}_{im}}\) (C-embedding dimensionality, Nim - number of image patches). The patch embeddings were then fed into a texture encoder which resembles radiologists’ long-term attention to observe texture variations over the liver region. Inspired by ref. 38, we designed the texture encoder as:
where (⋅)T was the transpose operator, \({\{{d}_{k}\in {{\mathbb{R}}}^{C}\}}_{k=1:K}\) was a learned texture codebook with K codewords and \({\{{s}_{k}\in {{\mathbb{R}}}^{C}\}}_{k=1:K}\) were learned scaling factors. We formed the final image embeddings as \({x}_{n}^{im}={x}_{n}^{pat}+{x}^{tex}\). Note that we applied random zero masking on the image tokens as a way of data augmentation and simulated partially missing image patches. This was motivated by driving the model’s attention to discriminative regions caused by disease alternations.
Multimodal representation
Last, we construct the multimodal representation by concatenating all acquired embeddings as \({h}^{0}=\left[{x}^{ord};{x}^{im};{x}^{nim};{x}^{dist}\right]\). Of note, except for an ordinal regression token xord, an extra distillation token xdist is introduced as part of the optimization purpose.
Missing-aware multi-modality alignment (MAMA)
The multimodal embedding h0 is fed into an encoder consisting of stacked MAMA adapter blocks and transformer blocks, as depicted in Fig. 8. Due to the heterogeneity between the visual and biometric features, directly applying self-attention and forcing the embeddings to interact with each other could lead to degraded learning results. We therefore design a modality alignment method that employs a learnable modality alignment prompt \({P}_{A}^{l}\) to prepend to hl and partially blinded attention \({M}_{A}^{l}\) that enforces the prompt vectors to only access each other but be blind to the inputs, as illustrated in Fig. 9b. This emphasizes that the prompt vectors should be focusing on the alignment between modalities rather than on specifics of the content of a modality56. Likewise, we address the missing inputs issues via introducing another missing-aware prompt \({P}_{M}^{l}\)39 and masked attention \({M}_{M}^{l}\) that is specifically designed to blind to the missing contents but enables the interaction within the missing-aware prompt vectors.
where Q, K, V are the query, key, and value vectors obtained by linear projections of the prompted inputs \(\left[{P}_{A}^{l-1};{h}^{l-1}\right]\) for modality alignment QA, KA, VA and \(\left[{P}_{M}^{l-1};{h}_{A}^{l-1}\right]\) for missing-aware QM, KM, VM, respectively. M is the masking operator that sets the blinded elements of QKT to − ∞. τ is a learnable scaling factor and l is the layer index.
Gold standard learning & curated distillation
In this step, optimization proceeds via three heads (Fig. 8): steatosis staging, fibrosis staging and distillation. The steatosis and fibrosis staging is trained on XGS, a histology-proven dataset with gold standard labels \({y}_{G{S}_{st}}\) and \({y}_{G{S}_{fib}}\), respectively. Meanwhile, the distillation head is trained on a larger non-biopsy proven dataset, XNGS. To address the issue of inaccurate or noisy labeling of non-gold standard labels yNGS, we propose a curated distillation strategy that only enables the gradients back-propagation when the ensemble of pretrained models outputs yPME agrees with the radiological report yNGS on any patient case. We formulate the steatosis and fibrosis staging problem as an ordinal regression task57. Ordinal regression also encourages the model to learn a soft decision boundary for distinctions between disease stages and avoid overfitting. Overall, the optimization process is formulated as follows:
where \({Z}_{G{S}_{st}}^{i}\), \({Z}_{G{S}_{fib}}^{i}\) and \({Z}_{NGS}^{i}\) are the output logits of the linear projections \({F}_{or{d}_{st}}\), \({F}_{or{d}_{fib}}\) and Fdist. \(\psi \left(\cdot \right)\) is a sigmoid function. LMSE is the mean squared error loss, KL is the Kullback-Leibler divergence, and \({\bf{1}}\left\{\cdot \right\}\) indicates the agreement on staging. During inference, we average the outputs of the heads \({F}_{or{d}_{st}}\) and Fdist to generate the final prediction on liver steatosis, and the head \({F}_{or{d}_{fib}}\) to generate the prediction on liver fibrosis.
Liver-spleen 3D segmentation& liver image preprocessing
To segment the liver and spleen from CTs, we utilized a framework described in ref. 58. Briefly, region of interest of the abdominal region was extracted at first, which is a key step towards fine segmentation of liver and spleen, given the diversity of CT scans exhibiting large variations in the field of view (FoV). This framework enables highly efficient detection and segmentation of the organs of interest. We refined and tailored it using a pre-trained model on FLARE dataset, expanded the training dataset by incorporating a large number of unenhanced CTs, and performed z-direction sub-volume cropping as a data augmentation to simulate the large variations in FoV. As a result, we had a highly efficient and accurate segmentation model that achieves a dice score of 98.0% and 97.2% for liver and spleen segmentation on the FLARE202259 validation. We cropped liver in 3D based on the segmentation and truncated voxels to \(\left[-175,275\right]\) HU and normalized to \(\left[0,1\right]\) and then resampled to the input size (384, 256, 64) for training and inference. We also applied data augmentation during training, by randomly selecting from a pool of operations, including: 1) randomly shifting the volume within 5 voxels in x, y, z direction; 2) randomly rotating across the slice within ± 25° in x, y direction; 3) randomly flipping the slice horizontally in x, y direction; 4) randomly adding gaussian noise to the image.
Training and implementation details
We developed our models using the gold-standard GS dataset and ablation studies were conducted via five-fold cross-validation. We held 20% data for validation and 80% data for training at each fold. For training the MAOSS, we conducted two stages of training. In the first stage, we pretrained the backbone on the modality-complete subset i.e. paired image and numerical data and only the ordinal regression heads (steatosis and fibrosis) were optimised during the first training stage with gold standards. Then, we kept all the multi-head self-attention (MSA) blocks frozen in the second stage, only optimizing the plugged Missing-aware Multi-Modality Alignment (MAMA) blocks by training on both the modality complete subset and the modality-incomplete i.e. with missing numerical features of the GS dataset. At the same time, we also incoprated the non-gold standard (NGS) dataset for learning the curated distllation head. The image feature encoder was implemented with a 3D ResNet-34 architecture. The number of code words K was set to 8 for texture encoding. The length of prompt vectors in MAMA was set to 16. C-the feature dimensionality was set to 512 for both image and texture embedding. All models were trained using Adam with an initial learning rate of 10−4 and batch size of 60, using four NVIDIA Tesla V100 GPUs. For curated distillation training, batch size for the GS and NGS samples denoted as BGS and BNGS were both 30. λ in the loss function was set to 0.5.
Interpretation of MAOSS
We flattened a 3D liver image \({x}^{im}=({x}_{1},\ldots .,{x}_{n})\in {{\mathbb{R}}}^{n}\) (n= number of voxels), score mapping saliency map could be obtained by calculating the gradient of the MAOSS score denoted as S(x) at the input xim: \(g=\frac{\partial S(x)}{\partial x}=({g}_{1},...,{g}_{n})\in {{\mathbb{R}}}^{n}\), where gi represents the contribution of individual voxel xi to the MAOSS score. The back-propagated gradients were potentially sensitive to unimportant features of the models. To deal with these limitations, we employed a feature attribution method named “Integrated Gradients”40 that assigns an importance score ϕi(S(x), x) (similar to voxel-wise gradients) to the ith pixel representing how much the pixel value adds or subtracts from the network output. To compute the importance score, it needs to introduce a baseline input representing “absence” of the feature input, denoted as \({x}^{{\prime} }=({x}_{1}^{{\prime} },...,{x}_{n}^{{\prime} })\in {{\mathbb{R}}}^{n}\), which in our study, was a null image (filled with zeros) with the same shape as input image x integrated gradients can obtain importance scores by accumulating gradients on images interpolated between the baseline value and the current input. This can be efficiently approximated via a summation of the gradients as:
where m is the number of steps in the Riemann approximation of the integral. We compute the approximation in a loop over the set of inputs for n = 1, …, m.
Subgroup analysis based on clinical factors
The subgroup analysis was conducted based on demographic, cardiovascular, and chronic liver disease risk factors60. Specifically, for demographic factors, the variables included age, gender, and body mass index (BMI). Age was categorized as either < 50 years or ≥50 years. Gender was classified into male and female. BMI was categorized into two groups: without obesity (BMI < 25.0 kg/m2) and with obesity (BMI ≥25.0 kg/m2). For cardiovascular factors, several metabolic characteristics were considered. Prediabetes/diabetes was identified based on fasting blood glucose levels (≥5.6 mmol/L)61 or a self-reported diagnosis of prediabetes/diabetes. Patients were classified as having dyslipidemia if they demonstrated any of the following lipid abnormalities: total cholesterol (TC) ≥6.20 mmol/L, triglycerides (TG) ≥2.30 mmol/L, high-density lipoprotein cholesterol (HDL-C) < 1.00 mmol/L, or low-density lipoprotein cholesterol (LDL-C) ≥4.20 mmol/L. Additionally, the presence of metabolic syndrome (MetS) was defined as meeting at least two of the following criteria: (1) hypertension; (2) fasting blood glucose ≥5.6 mmol/L; (3) HDL-C < 1.03 mmol/L for men or < 1.29 mmol/L for women; and (4) TG ≥1.7 mmol/L. For chronic liver disease factors, elevated levels of alanine aminotransferase (ALT) and aspartate aminotransferase (AST) were assessed. Elevated ALT was defined as ALT > 40 IU/L, and elevated AST was defined as AST > 40 IU/L.
MAOSS-augmented radiologist assessments
The aim of the first reader study was to assess the readers’ performance in detecting hepatic steatosis and grading on NCCT. The study (supplementary Fig. S17) was conducted in two rounds. The first round compared performance with that of radiologists with varying levels of expertise in radiology. The second round investigated whether MAOSS would be capable of assisting radiologists. There was a washout period of 2 weeks between the two rounds for each reader. A total of 11 readers were recruited in this study, consisting of 5 junior radiologists (experience < 2 yrs), 3 mid-level radiologists (experience > 5 yrs), and 3 senior radiologists (experience > 10yrs) (supplementary Table S34). In the first round each reader was trained to use the in-house annotation-viewing software (supplementary Fig. S18) for the visualization of the CT images. Basic functions of this software include but are not limited to HU (Hounsfield unit) windowing, zooming in and out, and axial, sagittal and coronal view simultaneous display. In interpreting the 191 randomly ordered cases from the internal test cohort, NCCT images and information on age and sex, clinical features (if available) were provided. The readers were informed that the study dataset was enriched with more positive patients than the standard prevalence of HS in daily practice. However, they were not informed about the proportions of different grades of HS. Each reader interpreted the image without time constraints and classified each case as 0 None, 1 mild HS, 2 moderate HS, 3 severe HS. In the second round the same group of readers interpreted the 191 cases again using the same software but augmented with MAOSS measurements (supplementary Fig. S19).
Primary risk stratification for patients identified with hepatic steatosis
According to the AASLD practice guidance, patients suspected to have SLD on the basis of metabolic risk factors or incidentally identified as having hepatic steatosis by imaging should undergo further evaluation for risk progression of steatohepatitis or advanced liver fibrosis, those identified at risk will be a referral for further risk assessment with more sophisticated examination, such as magnetic resonance elastography or corrected T1 at gastroenterology/hepatology clinics41. The AASLD pathway for evaluation of steatosis patients at risk involves transient-elastography and laboratory-based biomarkers, specifically fibrosis-4 index (FIB-4)62, which consists of 3 steps (Supplementary Fig. S13a): first, TE-CAP ≥275 dB/m was used to identify the patients with presence of steatosis52, who were further classified with FIB-4. Those with a FIB-4 of 2.67 higher, 1.3–2.67 or 1.3 lower were divided into high-, intermediate- or low-risk groups. The intermediate risk group was further divided into low-risk ( < 8 kPa) or intermediate-high risk (≥8 kPa) groups based on the TE-LSM values. We embedded MAOSS into the primary risk stratification setting and proposed a new screening pathway, referred to as the MAOSS pathway (Supplementary Fig. S13b) that is also a three-step sequential algorithm that involves MAOSS score, FIB-4 and FibroScan-AST score (FAST)63. First, we applied a MAOSS score ≥1.6 and FIBRO score ≥ 1.7 to identify the patients with hepatic steatosis, who were then divided into the three subgroups i.e. low-, intermediate-, high-risk, based on the FIB-4, same as the AASLD pathway. Finally, the intermediate risk group was further divided into low-risk (<0.35) or intermediate-high risk (≥0.35) groups based on the FAST values. These algorithms were evaluated using patients with liver biopsy
Performance metrics
Histologic scores were dichotomized as: S0 vs S1 or higher; S0 or S1 vs S2 or S3; and S2 or lower vs S3. We evaluate model’s overall performance with the area under the receiver operating characteristic curve (AUC), balanced accuracy (BACC) and F1-score. Additionally, we computed sensitivity, specificity, accuracy, negative predictive value (NPV) and positive predictive value (PPV). The F1-score considers both precision and recall of a test whereas the BACC considers sensitivity and specificity as a holistic measurement of classification performance. We also calculated inter-annotator agreement using Cohen’s kappa (κ), as the ratio of the number of times MAOSS and radiologists agreed on a diagnosis.
Statistical analysis
Classification performance was evaluated by calculating the AUC, accuracy, sensitivity, specificity, positive predictive value, and negative predictive value with 95% confidence intervals (CIs). The 95% CIs were calculated based on 1000 bootstrap replications of the test data. The Kruskal-Wallis test was used to compare the MAOSS scores between the different histology stages of hepatic steatosis. The Dorfman-Berbaum-Metz method64 (OR-DBM MRMC version 2.52) with the empirical AUC estimation was used to analyze multi-reader diagnostic performance with AI assistance and without AI assistance. The significance of the difference between the AUCs of models was assessed using the Delong test. Permutation tests65 were performed, and a two-sided p-value was calculated using 10,000 permutations to compare the subgroups’ AUCs and determine whether the difference between them was statistically significant. Odds ratio use Pearson’s chi-squared test. Competing risks analysis was used to assess the association of different screening pathway with development of hepatic decompensation. Cause-specific proportional subhazards analysis was used to assess the association between screened high risk cohort and time to development of hepatic cirrhosis after competing events were taken into account42. Statistical analysis was conducted in R version 4.3.1 and python 3.7.12. Comparisons with a two-sided p value less than 0.05 were considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets analyzed in this study are not publicly available due to restrictions imposed by the respective IRBs. Researchers may request access to the anonymized data and supporting clinical documentation by contacting corresponding author Y.S., J.L. and J.C. Access will be granted subject to IRB approval, a signed data-use agreement and will be for noncommercial academic purposes only. Requests will be processed within 6 weeks. Source data are provided with this paper.
Code availability
The code used for the implementation of MAOSS has dependencies on internal tooling and infrastructure and is under patent protection (application number CN 202310814888 and CN 202411205413), and thus the full custom codes is not able to be publicly released. The codes used for inference and result evaluation is available at: https://github.com/YGOX/MAOSS. All experiments and implementation details are described in sufficient detail in Methods and Supplementary information (‘Training and implementation details’) to support replication with nonproprietary libraries. Several principal components of MAOSS are available in open-source repositories: ViT-Pytorch (https://github.com/lucidrains/vit-pytorch), ViLT (https://github.com/dandelin/ViLT), missing aware prompts (https://github.com/YiLunLee/missing_aware_prompts).
References
Rinella, M. E. et al. A multisociety delphi consensus statement on new fatty liver disease nomenclature. Hepatology 78, 1966–1986 (2023).
Younossi, Z. M. et al. The global epidemiology of nonalcoholic fatty liver disease (nafld) and nonalcoholic steatohepatitis (nash): a systematic review. Hepatology 77, 1335–1347 (2023).
Le, M. H. et al. Forecasted 2040 global prevalence of nonalcoholic fatty liver disease using hierarchical bayesian approach. Clin. Mol. Hepatol. 28, 841 (2022).
Rinella, M. E. Nonalcoholic fatty liver disease: a systematic review. Jama 313, 2263–2273 (2015).
Panel, C. P. G. et al. Easl clinical practice guidelines on non-invasive tests for evaluation of liver disease severity and prognosis–2021 update. J. Hepatol. 75, 659–689 (2021).
Le, M. H. et al. 2019 global nafld prevalence: a systematic review and meta-analysis. Clin. Gastroenterol. Hepatol. 20, 2809–2817 (2022).
Noureddin, M. et al. Expert panel recommendations: practical clinical applications for initiating and monitoring resmetirom in patients with mash/nash and moderate to noncirrhotic advanced fibrosis. Clin. Gastroenterol. Hepatology. 22, 2367–2377 (2024).
Harrison, S. A. et al. A phase 3, randomized, controlled trial of resmetirom in nash with liver fibrosis. N. Engl. J. Med. 390, 497–509 (2024).
Loomba, R. et al. Tirzepatide for metabolic dysfunction–associated steatohepatitis with liver fibrosis. N. Engl. J. Med. 391, 299–210 (2024).
Chalasani, N. et al. The diagnosis and management of nonalcoholic fatty liver disease: practice guidance from the american association for the study of liver diseases. Hepatology 67, 328–357 (2018).
Duell, P. B. et al. Nonalcoholic fatty liver disease and cardiovascular risk: A scientific statement from the american heart association. Arteriosclerosis, Thrombosis, Vasc. Biol. 42, e168–e185 (2022).
Starekova, J., Hernando, D., Pickhardt, P. J. & Reeder, S. B. Quantification of liver fat content with ct and mri: state of the art. Radiology 301, 250–262 (2021).
Pouwels, S. et al. Non-alcoholic fatty liver disease (nafld): a review of pathophysiology, clinical management and effects of weight loss. BMC Endocr. Disord. 22, 63 (2022).
Guo, Z. et al. Liver fat content measurement with quantitative ct validated against mri proton density fat fraction: a prospective study of 400 healthy volunteers. Radiology 294, 89–97 (2020).
Pickhardt, P. J., Graffy, P. M., Reeder, S. B., Hernando, D. & Li, K. Quantification of liver fat content with unenhanced mdct: Phantom and clinical correlation with mri proton density fat fraction. Am. J. Roentgenol. 211, W151–W157 (2018).
Haghshomar, M. et al. Diagnostic accuracy of ct for the detection of hepatic steatosis: A systematic review and meta-analysis. Radiology 313, e241171 (2024).
Kim, D. Y. et al. Contrast-enhanced computed tomography for the diagnosis of fatty liver: prospective study with same-day biopsy used as the reference standard. Eur. Radiol. 20, 359–366 (2010).
Dell, T. et al. Photon-counting ct–derived quantification of hepatic fat fraction: a clinical validation study. Radiology 314, e241677 (2025).
Lubner, M. G., Malecki, K., Kloke, J., Ganeshan, B. & Pickhardt, P. J. Texture analysis of the liver at mdct for assessing hepatic fibrosis. Abdom. Radiol. 42, 2069–2078 (2017).
Pickhardt, P. J., Malecki, K., Kloke, J. & Lubner, M. G. Accuracy of liver surface nodularity quantification on mdct as a noninvasive biomarker for staging hepatic fibrosis. Am. J. Roentgenol. 207, 1194–1199 (2016).
Lee, S. et al. Fully automated and explainable liver segmental volume ratio and spleen segmentation at ct for diagnosing cirrhosis. Radiology: Artif. Intell. 4, e210268 (2022).
Pickhardt, P. J. et al. Hepatosplenic volumetric assessment at mdct for staging liver fibrosis. Eur. Radiol. 27, 3060–3068 (2017).
Pickhardt, P. J. & Lubner, M. G. Noninvasive quantitative ct for diffuse liver diseases: Steatosis, iron overload, and fibrosis. RadioGraphics 45, e240176 (2024).
Graffy, P. M., Sandfort, V., Summers, R. M. & Pickhardt, P. J. Automated liver fat quantification at nonenhanced abdominal ct for population-based steatosis assessment. Radiology 293, 334–342 (2019).
Modanwal, G. et al. Deep-learning-based hepatic fat assessment (dehft) on non-contrast chest ct and its association with disease severity in covid-19 infections: A multi-site retrospective study. EBioMedicine 85, 104315 (2022).
Choi, K. J. et al. Development and validation of a deep learning system for staging liver fibrosis by using contrast agent–enhanced ct images in the liver. Radiology 289, 688–697 (2018).
Kutaiba, N. et al. Deep learning for computed tomography assessment of hepatic fibrosis and cirrhosis: a systematic review. Mayo Clin. Proc.: Digital Health 1, 574–585 (2023).
Sarvestany, S. et al. Development and validation of an ensemble machine learning framework for detection of all-cause advanced hepatic fibrosis: A retrospective cohort study., 2022, 4, pp. e188-e199. https://doi.org/10.1016/S2589-7500(21) 00270–3.
Chang, D. et al. Machine learning models are superior to noninvasive tests in identifying clinically significant stages of nafld and nafld-related cirrhosis. Hepatology 77, 546–557 (2023).
Niu, L. et al. Noninvasive proteomic biomarkers for alcohol-related liver disease. Nat. Med. 28, 1277–1287 (2022).
Ianiro, G. et al. Correction to lancet gastroenterol hepatol. Gastroenterol. Hepatol. 5, 430–31 (2020).
Jung, J. et al. Mre combined with fib-4 (mefib) index in detection of candidates for pharmacological treatment of nash-related fibrosis. Gut 70, 1946–1953 (2021).
Van Buuren, S. & Groothuis-Oudshoorn, K. mice: Multivariate imputation by chained equations in r. J. Stat. Softw. 45, 1–67 (2011).
Van der Maaten, L. & Hinton, G. Visualizing data using t-sne. J. Machine Learning Res. 9, 2579–2605 (2008).
Troyanskaya, O. et al. Missing value estimation methods for dna microarrays. Bioinformatics 17, 520–525 (2001).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, 770–778 (2016).
Touvron, H. et al. Training data-efficient image transformers & distillation through attention. In International conference on machine learning, 10347–10357 (PMLR, 2021).
Zhang, H., Xue, J. & Dana, K. Deep ten: Texture encoding network. In Proceedings of the IEEE conference on computer vision and pattern recognition, 708–717 (2017).
Lee, Y.-L., Tsai, Y.-H., Chiu, W.-C. & Lee, C.-Y. Multimodal prompting with missing modalities for visual recognition. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 14943–14952 (2023).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. In International conference on machine learning, 3319–3328 (PMLR, 2017).
Rinella, M. E. et al. Aasld practice guidance on the clinical assessment and management of nonalcoholic fatty liver disease. Hepatology 77, 1797–1835 (2023).
Fine, J. P. & Gray, R. J. A proportional hazards model for the subdistribution of a competing risk. J. Am. Stat. Assoc. 94, 496–509 (1999).
Modanwal, G. et al. Opportunistic hepatic steatosis assessment in low-dose coronary artery calcium ct using liver adipose-radiomic index (lari). EBioMedicine 121, 105982 (2025).
Vianna, P. et al. Comparison of radiologists and deep learning for us grading of hepatic steatosis. Radiology 309, e230659 (2023).
Qian, L. -l et al. Serum biomarkers combined with ultrasonography for early diagnosis of non-alcoholic fatty liver disease confirmed by magnetic resonance spectroscopy. Acta Pharmacologica Sin. 41, 554–560 (2020).
Hu, P. et al. Effects of slice thickness on ct radiomics features and models for staging liver fibrosis caused by chronic liver disease. Jpn. J. Radiol. 40, 1061–1068 (2022).
Park, H. J. et al. Ct-based visual grading system for assessment of hepatic steatosis: diagnostic performance and interobserver agreement. Hepatol. Int. 16, 1075–1084 (2022).
Kjaergaard, M. et al. Using the elf test, fib-4 and nafld fibrosis score to screen the population for liver disease. J. Hepatol. 79, 277–286 (2023).
Chen, L.-D. et al. Us-based sequential algorithm integrating an ai model for advanced liver fibrosis screening. Radiology 311, e231461 (2024).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Neuberger, J. et al. Guidelines on the use of liver biopsy in clinical practice from the british society of gastroenterology, the royal college of radiologists and the royal college of pathology. Gut 69, 1382–1403 (2020).
Sterling, R. K. et al. Aasld practice guideline on imaging-based non-invasive liver disease assessments of hepatic fibrosis and steatosis. Hepatology 10–1097 (2024).
Kim, W., Son, B. & Kim, I. Vilt: Vision-and-language transformer without convolution or region supervision. In International Conference on Machine Learning, 5583–5594 (PMLR, 2021).
Huo, Y. et al. Fully automatic liver attenuation estimation combing cnn segmentation and morphological operations. Med. Phys. 46, 3508–3519 (2019).
Tian, J., Liu, L., Shi, Z. & Xu, F. Automatic couinaud segmentation from ct volumes on liver using glc-unet. In International workshop on machine learning in medical imaging, 274–282 (Springer, 2019).
Liang, S., Zhao, M. & Schuetze, H. Modular and parameter-efficient multimodal fusion with prompting. In Muresan, S., Nakov, P. & Villavicencio, A. (eds.) Findings of the Association for Computational Linguistics: ACL 2022, 2976–2985 https://aclanthology.org/2022.findings-acl.234 (Association for Computational Linguistics, Dublin, Ireland, 2022).
Cheng, J., Wang, Z. & Pollastri, G. A neural network approach to ordinal regression. In 2008 IEEE international joint conference on neural networks, 1279–1284 (IEEE, 2008).
Sun, M., Jiang, Y. & Guo, H. Semi-supervised detection, identification and segmentation for abdominal organs. In MICCAI Challenge on Fast and Low-Resource Semi-supervised Abdominal Organ Segmentation, 35–46 (Springer, 2022).
Ma, J. et al. Unleashing the strengths of unlabelled data in deep learning-assisted pan-cancer abdominal organ quantification: the flare22 challenge. Lancet Digital Health 6, e815–e826 (2024).
Man, S. et al. Prevalence of liver steatosis and fibrosis in the general population and various high-risk populations: a nationwide study with 5.7 million adults in china. Gastroenterology 165, 1025–1040 (2023).
Association, A. D. Standards of medical care in diabetes—2022 abridged for primary care providers. Clin. Diab. 40, 10–38 (2022).
Sterling, R. K. et al. Development of a simple noninvasive index to predict significant fibrosis in patients with hiv/hcv coinfection. Hepatology 43, 1317–1325 (2006).
Newsome, P. N. et al. Fibroscan-ast (fast) score for the non-invasive identification of patients with non-alcoholic steatohepatitis with significant activity and fibrosis: a prospective derivation and global validation study. lancet Gastroenterol. Hepatol. 5, 362–373 (2020).
Smith, B., Hillis, S. & Pesce, L. Mrmcaov: multi-reader multi-case analysis of variance. R package version 0.1 16 (2021).
Lange, K. Permutation tests and the bootstrap. In Numerical Analysis for Statisticians, Statistics and Computing (Springer, New York, NY, 2010).
Acknowledgements
We thank all the investigators and participants in this study. Y.S. and Y.H. were supported by Shengjing Hospital of China Medical University, grants from the National Natural Science Foundation of China (No. 82471971), the Liaoning Province Science and Technology Joint Plan (2023JH2/101700127), and the Leading Young Talent Program of Xingliao Yingcai in Liaoning Province (XLYC2203037). J.L. was supported by Nanjing Drum Tower Hospital, Affiliated with Nanjing University, grants from the National Natural Science Foundation of China (No. 82170609), and the Nanjing Drum Tower Hospital Academic Innovation Peak Project (No. 2024-DF-03). X.Y. was supported by The First Affiliated Hospital, Zhejiang University’s Zhejiang Provincial “Spearhead & Pathfinder + X” R&D Breakthrough Program under Grant No. 2024C03043, and Zhejiang Provincial Natural Science Foundation of China under Grant No. 2024-KYI-00I-I05.
Author information
Authors and Affiliations
Contributions
Y.G., C.L., L.Z., and Y.S. conceived and designed the study. Y.G. and Y.H.Y. drafted the original manuscript, while Y.S., L.Z., and K.Y. provided critical advice and refined the manuscript for submission. Y.G., L.Z., and Y.S. contributed to subsequent revisions. W.X.C. created and refined the figures. Y.G. designed the architecture and developed the MAOSS framework. C.L. and W.X.C. contributed to the experimental design, execution, and data analysis. Y.S., Y.H., and C.L. organized and conducted the reader study. Y.S., Y.H., C.L., and B.D. managed the collection, organization, cleaning, and de-identification of internal training and testing data, and provided data for additional revision experiments. X.Y., J.L., and J.C. collected and organized the external validation data, including CT scans, clinical data, and liver biopsy histology. K.C. provided real-world CT data. Y.X., H.G., X.Z., W.L., R.B., B.L., J.Y., and L.L. provided machine learning or clinical expertise and participated in technical discussions. L.Z. and Y.K. co-supervised the technical development of the project, while Y.S., Y.H., J.L., J.C., and Y.H.Y. supervised the clinical aspects.
Corresponding authors
Ethics declarations
Competing interests
Alibaba Group has filed patent applications CN 202310814888 (on behalf of Y.G., K.Y., H.G., L.Z., J.Y., L.L., Y.S., and C.L.) and CN 202411205413 (on behalf of Y.G., C.L., L.Z., K.Y., H.G., W.L., X.Z., L.L., and Y.S.) for work related to the MAOSS methods. Y.G., W.X.C., Y.X., H.G., X.Z., W.L., J.Y., K.Y. and Z.L. are employees of Alibaba Group and own Alibaba stock as part of the standard compensation package. All other authors have no competing interests.
Peer review
Peer review information
Nature Communications thanks Amir Borhani, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gao, Y., Li, C., Chang, W. et al. Multi-modal AI for opportunistic screening, staging and progression risk stratification of steatotic liver disease. Nat Commun 17, 1562 (2026). https://doi.org/10.1038/s41467-026-68414-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68414-3