Abstract
This work presents a general strategy for engineering cell spheroids with capillary-like structures using intercellular self-assembly of peptide nanofibers. These nanofibrous materials induce mechanical changes in the extracellular matrix (ECM), activate mechanotransduction pathways, and enhance cellular morphogenesis, resulting in dynamic 3D spheroids with improved cell-cell and cell-matrix interactions. By promoting the formation of capillary-like structures within tumor spheroids, we develop models that closely mimic human tissue physiology. Our results demonstrate that tumor spheroids with capillary-like structures display gene expression profiles that closely match those of patient-derived tumors, underscoring their relevance for cancer research. Furthermore, these spheroids, including those derived from an islet cell line, exhibit significantly increased functionality, such as enhanced insulin secretion in response to glucose stimulation, highlighting their potential for diabetes research and regenerative medicine applications. This work advances our understanding of tissue engineering and provides a robust platform for studying complex cellular interactions and therapeutic responses. By highlighting the critical role of capillary-like structure formation in engineered tissues, our findings pave the way for innovative strategies to address significant challenges in drug delivery and cancer therapy, ultimately enhancing patient care and treatment outcomes.
Similar content being viewed by others
Introduction
The extracellular matrix (ECM) is a complex, three-dimensional (3D) network that plays a fundamental role in tissue remodeling and cellular activities1,2,3,4. It provides essential structural support, facilitates cell signaling, and regulates mechanical properties crucial for modulating cell behavior. By interacting with cell surface receptors, the ECM influences cell processes such as growth, differentiation, and communication5,6,7. Recent studies underscore the ECM’s significance in controlling tissue mechanics, where its composition and organization are vital for mediating cellular responses to mechanical forces8,9. Cells utilize integrins and other membrane proteins to adhere to the ECM, enabling them to sense and respond to its mechanical properties. This mechanotransduction process involves conformational changes in mechanosensitive proteins, which activate downstream signaling pathways affecting cellular behavior10,11,12,13,14. Additionally, the ECM’s role in supporting angiogenesis—the formation of new blood vessels—is critical for maintaining tissue integrity and facilitating nutrient transport. Angiogenesis ensures adequate oxygen and nutrients for tissue growth and function15.
Despite the importance of the ECM in these processes, the development of effective biomaterials that can replicate its dynamic properties remains a significant challenge in tissue engineering16,17,18,19,20,21,22. Traditional natural scaffolds, such as those derived from decellularized tissues and purification (i.e. silk, chitosan, collagen, gelatin, proteoglycan, etc.), offer some advantages but often come with limitations, including high production costs, variability in composition, and a lack of spatially controlled biochemical cues23. Furthermore, while synthetic biomaterials provide tunable properties24, they frequently fail to mimic the complex interactions and mechanical behaviors characteristic of natural ECM4,25. The construction of artificial scaffolds that can support angiogenesis has primarily focused on engineered 3D printing techniques, designed to create artificial channels and shapes. However, these approaches often fall short of replicating the spontaneous and multifaceted nature of angiogenesis observed in vivo26,27. The engineered vascular structures typically lack the functional diversity and adaptability necessary for realistic tissue integration. Moreover, the mechanical properties of the ECM profoundly influence tissue morphology and functional behavior. Alterations in ECM mechanics are frequently associated with pathological processes in various diseases25,28, highlighting the need for materials that can dynamically modulate mechanical properties to reflect physiological conditions. The challenge of in situ construction of matrix materials that can reshape ECM mechanics remains a significant barrier to advancing tissue engineering8.
This study describes the intercellular formation of peptide nanofibrous materials, which assemble rapidly in the spaces between cells through enzymatic self-assembly. The in situ self-assembly of these nanofibers alters mechanical cues within the ECM, activating signaling pathways that remodel cytoskeletal actin and facilitate YAP nuclear translocation (Fig. 1). This process induces cell morphogenesis and enables the formation of dynamic 3D cellular spheroids. Notably, co-culturing these spheroids with endothelial cells promotes angiogenesis, resulting in capillary-like structures that enhance substance transport and cellular communication. This work demonstrates that the strategy of in situ formed nanofibrous materials effectively mimics ECM architecture while integrating mechanical cues to promote cellular morphogenesis and functional capillary-like structure formation. This innovative approach holds significant promise for advancing tissue engineering and organoid development, offering a paradigm for creating functional, vascularized tissues.
Intercellular peptides (CSP) self-assemble into a nanofibrous network surrounding the cells, modulating mechanical cues in the ECM. This mechanical signaling, characterized by YAP activation and actin remodeling, drives cellular morphogenesis and the formation of 3D spheroids. Subsequent co-culture with endothelial cells and peptides (CSP-A) leads to the development of spheroids with capillary-like structures, enhancing substance exchange and signal communication. Created in BioRender. Li, Y. (2025) https://BioRender.com/sqlourl.
Results
In situ self-assembled peptide nanofibers driving the cells into spheroids
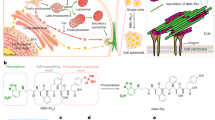
As shown in Fig. 2a, the peptide CSP features a backbone of “Tryptophan-phenylalanine-Tyrosine (W-F-Y), a well-studied composition for self-assembly29,30. The modification at N-terminal by Pyrene increases the self-assembly ability of the whole system by providing strong aromatic-aromatic interactions31, while phosphorylated tyrosine provides enzymatic responsiveness that mimics the processes of biocatalytic ECM formation. Additionally, lysine at the C-terminus promotes cell adhesion through electrostatic interactions32. The peptide solution was clear after 24 h, and cryo-EM imaging revealed a few of self-assembled nanofibers with a diameter of 6.3 nm (Fig. 2b). Treatment with alkaline phosphatase (ALP) for 2 h initiated the formation of a nanofiber network (Supplementary Fig. 1a), demonstrating the peptides’ responsiveness to enzymatic cues. Extending the incubation to 24 h significantly increased fiber density, suggesting that enzymatic hydrolysis enhances the assembly process. Circular dichroism (CD) spectra confirmed that these nanofibers exhibit a β-sheet secondary structure (Supplementary Fig. 1b), indicating a stable and organized structure critical for cellular interactions. The same time points as cryo-EM were chosen for HPLC and MS analysis (Supplementary Fig. 1c, d), the peaks of the hydrolysis products gradually increased.
a The molecular structure of the peptide CSP. b Cryo-EM images of nanostructures formed by CSP (50 μM) with or without the treatment of ALP (1 U/mL). c The illustration and microscopic images of U87MG cell spheroids formation induced by peptide CSP (50 μM) at 24 and 96 h. Created in BioRender. Li, Y. (2025) https://BioRender.com/dmp2u6m. d The projected area of U87MG cell spheroids induced by peptide CSP at different concentrations (n = 10). Data are presented as plots: center lines = mean, error bar = SD, and individual data points are shown as circles. e Live/dead fluorescence staining images of cell spheroids induced by CSP for 96 h. f SEM images of U87MG cell spheroid, with filopodia marked in yellow, peptide nanofibers in cyan, and microvilli in purple, respectively. g Bio-EM images of U87MG cells treated with CSP for 6 and 24 h (The cell membrane is depicted by blue lines), respectively. Source data are provided as a Source Data file.
To investigate the physiological impact of the peptide nanofibers, we treated U87MG cells (a human glioblastoma cell expressing ecto-phosphatase) with CSP for 24 h, leading to spheroid formation (Fig. 2c). Over 96 h, these spheroids increased in size, as shown in a video of the spheroid formation process (Supplementary Fig. 2, Supplementary Movie 1). Concentration-dependent experiments revealed that 50 μM CSP yielded the largest spheroids, with volume positively correlated to peptide concentration and incubation time within the 10–50 μM range (Fig. 2d, Supplementary Fig. 3), suggesting the importance of optimizing peptide concentration to induce cell spheroid formation. The decrease in spheroid size at higher CSP concentrations (>50 µM) and the failure to form spheroids at 200 µM are governed by the balance between the generation rate of the hydrolyzed product and cell aggregation dynamics. While the enzymatic hydrolysis rate per se may remain constant (as enzyme concentration is fixed at the same density as the cells), increasing the CSP concentration elevates the amount of hydrolyzed product due to greater substrate availability. The hydrolyzed product forms at a moderate rate at 50 µM CSP, allowing cells to aggregate into larger spheroids before becoming entrapped in the assembled matrix. At 100 µM CSP, the higher precursor concentration increases the cumulative hydrolyzed product over time, leading to earlier and more extensive matrix formation. This restricts cell-cell contact at a smaller spheroid size. At 200 µM CSP, the rapid accumulation of hydrolyzed product overwhelms the system, entrapping individual cells before they can establish stable contacts. This prevents spheroid formation entirely, as the matrix solidifies around dispersed cells. Therefore, the total hydrolyzed product generated determines whether cells form large spheroids, small spheroids, or remain isolated. The threshold concentrations reflect a kinetic competition between cell aggregation and matrix entrapment. Cellular viability assays indicated that peptide-induced spheroids maintained similar proliferation levels to monolayer U87MG cells. Live-dead staining showed strong green fluorescence in the spheroids, indicating good cell viability (Fig. 2e).
Congo Red staining visualized the β-sheet nanofibers33, showing strong red fluorescence around the cell spheroids but minimal fluorescence in monolayer cells (Supplementary Fig. 4), confirming that the peptide nanofibers are preferentially formed locally around the cell spheroids, enhancing their structural integrity and functional capabilities. Scanning electron microscopy (SEM) and transmission electron microscopy (TEM) provided further insights into the morphology of in situ formed peptide assemblies induced spheroids. SEM images (Fig. 2f) displayed extensive cell aggregation with huge clusters of nanofibers distributed between the cells. These nanofibers adhered to the surface of the cell membranes, promoting cell-cell interactions. Filopodia formed by the cells extended towards the nanofibers, integrating into the fiber network. In contrast, the control group showed only independent cells with an elliptical appearance and microvillus structures on their membranes, lacking any associated nanofibers. Bio-EM (Fig. 2g) images revealed that after 6 h, the nanofibers aggregate around the cells, which distribute along cell membranes to help cells form cell spheroids. Prolonging incubation to 24 h resulted in tighter cell connections and dense fibrous networks, resembling the nanofibers formed by enzymatic assembly of CSP in vitro. This dynamic correlation suggests that the nanofibers not only support structural assembly but also could facilitate the mechanical and signaling interactions necessary for the cell spheroid development.
Beyond U87MG cells, we successfully induced spheroid formation in other cell lines (SK-OV-3, HepG2, Saos-2) that naturally expressed ALP34,35,36 using the peptide, demonstrating the broad applicability of CSP (Supplementary Figs. 5, 6). Notably, CSP successfully induced spheroid formation in MDA-MB-231 cells (Supplementary Fig. 7), which showed 2D structure in low-adhesion plates and low expression levels of ALP. This finding further highlights the unique capability of our enzyme-responsive strategy to promote 3D spheroid formation even in challenging cell models. However, CSP alone could not induce HUVEC which didn’t express ALP to form spheroids (Supplementary Fig. 8). To investigate the influence of molecular structure on cell spheroid formation, we synthesized similar peptide variants (Supplementary Fig. 9). The peptide lacking the pyrene group (CSP1) resulted in single adherent cells, with no cell spheroid formation, highlighting the critical role of aromatic-aromatic interactions in self-assembly for inducing cell spheroid formation. The variant without the phosphatase-responsive motif (CSP2) yielded smaller aggregates with reduced growth activity, suggesting that enzymatic responsiveness is vital for effective spheroid formation. This process reflects the natural mechanisms of ECM remodeling and cellular aggregation, highlighting that enzymatic peptide self-assembly is crucial for the dynamic assembly of ECM components that support cell spheroid formation. Such parallels emphasize the potential of our peptide design to replicate physiological processes in tissue engineering. Peptides containing glycine (CSP3) and glutamic acid (CSP4) showed significantly diminished spheroid-inducing ability, indicating that charge and structural integrity are crucial for optimal cellular aggregation.
Cytoskeletal remodeling drives YAP translocation and cell spheroids formation induced by in situ formed peptide nanofibers
The ECM is composed of dense fibrillar protein complexes that provide structural support and facilitate cell communication37. To visualize the distribution of peptides at the cellular level, we used nitrobenzoxadiazoles (NBD) to label the peptide at the N-terminal, resulting in NBD-CSP. An optimized ratio of CSP:NBD-CSP of 9:1 was used in our experiment to visualize the distribution of peptides without affecting the assembly of CSPs. Immunofluorescence imaging (Fig. 3a, Supplementary Fig. 10) revealed strong green fluorescence from self-assembled peptide nanofibers co-localized with red fluorescence from ECM proteins such as collagen III and fibronectin1,38. To determine co-localization, we employed Pearson’s correlation coefficient, which quantifies the degree of overlap between the two fluorescent signals. A high coefficient value would indicate a strong correlation between the peptide nanofibers and ECM proteins, confirming their spatial proximity. The observed overlap suggests that peptide nanofibers interact closely with ECM components, facilitating enzyme-instructed self-assembly at the cell surface. This interaction is crucial for establishing a supportive microenvironment that enhances cellular functions, promoting not only structural integrity but also dynamic signaling pathways essential for cell behavior and morphology.
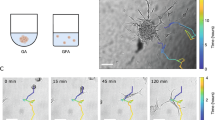
a Immunofluorescence staining of U87MG cells treated with CSP/NBD-CSP. The cell spheroid and 2D cells stained with antibodies of ECM protein Collagen III and Fibronectin. b, c The measurement of mechanical force of U87MG cells with or without the treatment of CSP (The blue area reflects cell-to-cantilever adhesion and the red reflects cell-to-plate adhesion.) (n = 5). Data are presented as mean value ± SD, and individual data points are shown as circles. Two-tailed unpaired Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). d Schematic illustration of the effects of inhibitors (Latrunculin B, CK666) on cellular actin remodeling and spheroid formation. e Optical microscopic images and projected areas of U87MG cells treated with CSP (50 μM) in the presence of Latrunculin B and CK666 (n = 10). Data are presented as plots: center lines = mean, error bar = SD, and individual data points are shown as circles. f The immunofluorescence staining of U87MG cells, and (g) statistical localization (n = 100) of YAP protein in U87MG cells treated with the peptides CSP compared to a control group. Data are presented as mean value ± SD. Two-tailed unpaired Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). h Mechanistic scheme of cell spheroid formation with CSP treatment. Created in BioRender. Li, Y. (2025) https://BioRender.com/utjk4z9. i, j The transcriptomics analysis of genes related to the mechanoregulation and morphogenesis in the spheroids and control cells (n = 3) Data are presented as mean value ± SD, and individual data points are shown as circles. Two-tailed unpaired Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). Source data are provided as a Source Data file.
The interactions between the self-assembled peptide nanofibers and the ECM likely play a critical role in modulating the mechanical environment of the cells. To assess the mechanical properties of the cells, we employed atomic force microscopy (AFM) (Fig. 3b, c). The basal adhesion of cells in the spheroid group was 72.5 nN, significantly lower than the control group (343.4 nN). This reduction indicates that the fibrous network formed by the peptide assembly substantially alters the mechanical microenvironment of the cells.
Cytoskeletal protein remodeling is crucial for filopodia formation39, so we explored the role of cytoskeletal protein remodeling in spheroids formation by incubating cells with the peptide and the small molecule inhibitors Latrunculin B (which inhibits actin polymerization40) and CK666 (which stabilizes the ARP2/3 complex in an inactive state and blocks actin nucleation and branching41). Results indicate that cells treated with the inhibitors formed significantly smaller aggregates compared to the peptide-only group (Fig. 3d, e, Supplementary Fig. 11). This suggests that the introduction of Latrunculin B and CK666 markedly impaired the peptide-induced spheroid-forming activity. Our viability assays (Alamar Blue) and 2D culture retention data collectively demonstrate that CSP and cytoskeletal inhibitors (Latrunculin B/CK666) impair 3D spheroid formation primarily through adhesion/migration disruption rather than proliferation effects. The maintained cell viability coupled with significant 2D cell accumulation (Supplementary Fig. 12) suggests these treatments interfere with cell-cell contact initiation—a critical first step in spheroid assembly. This aligns with known roles of actin dynamics (Latrunculin B) and Arp2/3 (CK666) in cellular adhesion. The diminished size of the cell spheroids underscores the critical role of cytoskeletal protein remodeling in spheroid formation. Specifically, through actin-mediated mechanosensing, cells can respond to the mechanical alterations induced by peptide nanofibers by forming structures such as protrusions and filopodia that facilitate cell-cell interactions and enhance aggregate stability. This highlights how the cytoskeleton is essential for translating mechanical signals from the nanofiber network into morphological changes necessary for spheroid development.
YAP proteins are key mediators of cellular responses to mechanical cues, translocating to the nucleus to activate downstream signaling pathways13,42. In our study, 76.5% of YAP proteins in cell spheroids were localized in the nucleus (Fig. 3f, g, Supplementary Fig. 13), compared to only 26.0% in the control group, indicating that self-assembled peptides significantly enhanced YAP nuclear translocation, an almost fourfold increase. CSP treatment triggered YAP nuclear localization, consistent with its mechanosensitive role in spheroid growth and survival. While low-adhesion controls were technically unfeasible (spheroid loss during staining), CSP-induced YAP activation suggests matrix stiffness or ligand cues override adhesion-dependent mechanosignaling. These findings underscore YAP’s role in 3D spheroid maintenance. We investigated the expression levels of the relevant proteins in the upstream signaling pathway of YAP - the Hippo pathway (Supplementary Fig. 14). After CSP treatment, cell spheroids were formed, the expression level of LATS1 slightly decreased, the expression levels of YAP and MOB1 remained almost unchanged, but the phosphorylation levels decreased. Among them, MOB1 acts as the molecular switch of the Hippo pathway. It transmits the upstream signals (MST1/2) to LATS1/2 through a phosphorylation-dependent mechanism, ultimately regulating the activity of YAP/TAZ. Then, YAP detaches from the cytoplasmic retention protein due to dephosphorylation and relocates to the nucleus, where it binds to the transcription factor TEAD and activates downstream target genes. We also applied Verteporfin (VP), an inhibitor of YAP protein43, finding that 0.2 μM VP substantially reduced cell spheroid formation, while 1 μM VP nearly completely inhibited spheroid formation (Supplementary Fig. 15). Statistical analysis revealed a negative correlation between spheroid area and VP concentration. These findings suggest that peptide CSP modulates the mechanical microenvironment of cells and activates the YAP pathway.
These results together indicate that the peptide CSP undergoes enzyme-instructed self-assembly to form a nanofiber network on the cell membrane, significantly influencing the mechanical properties of the ECM. This alteration induces mechanotransduction, promotes cytoskeletal remodeling, and facilitates the nuclear translocation of YAP proteins, leading to the activation of downstream signaling pathways that drive spheroid morphogenesis (Fig. 3h). Additionally, gene transcriptomics analysis (Fig. 3i, j) of cell spheroids revealed expression of genes associated with mechanoregulation and morphogenesis, confirming the proposed induction mechanism. These findings underscore the potential of peptide nanofibers in tissue engineering, where they can modulate cellular behavior through mechanosensing pathways.
Peptide nanofibers induce capillary-like structure formation in spheroids
Vascularization is essential for the functional performance of tissues44,45,46. To mimic in vivo angiogenesis, we designed the self-assembled peptide sequence CSP-A (Fig. 4a), incorporating the cNGR fragment known to target angiogenesis47,48. In the co-culture system of cell spheroids induced by the original peptide CSP and endothelial cells49,50, the self-assembled peptide CSP-A was treated for inducing capillary-like structure formation of cell spheroids. We co-cultured CSP-formed spheroids with HUVECs in three matrices (CSP-A, Matrigel, or agarose) to assess vascularization potential (Supplementary Figs. 16, 17). While control groups (spheroids alone or with HUVECs) showed no immature vascular networks formation, CD31 immunofluorescence revealed endothelial localization around spheroids in HUVEC-containing groups. Notably, CSP-A supported progressive immature capillary-like structure formation around spheroids, with CD31 tubular structures indicating functional capillary-like structure formation. In Matrigel, we observed initial tubular networks between spheroids that later matured into perfusable immature capillary-like structures. By contrast, agarose cultures exhibited passive spheroid-HUVEC aggregation due to low adhesion, with minimal immature vascular network morphogenesis. These results demonstrate that matrix properties critically regulate vascular patterning, with CSP-A uniquely enabling organized peri-spheroidal capillary-like structure formation. As a comparison, we also used Matrigel as a substrate and the Agarose method to culture U87MG cells and HUVECs to form 3D cell spheroids with endothelial tubes. The results (Supplementary Fig. 18, Supplementary Table 1) revealed the presence of outgrowth structures around the spheroids. Similar structures were also observed in the Matrigel group, which provided a substrate environment for cell growth, adhesion, and differentiation under 3D conditions, but these structures were absent in both the control group and the Agarose group. Immunofluorescence (IF) confirmed a high expression of CD31, a hallmark of angiogenesis51, indicating the formation of immature capillary-like structures induced by CSP-A (Fig. 4b). Compared with angiogenic spheroids cultured with Matrigel, the immature capillary-like structures were also distributed inside the spheroids when cultured with CSP/CSP-A, likely due to the combined effect of CSP-A on U87MG cells and HUVECs (Supplementary Fig. 19, Supplementary Table 1). Microstructural studies (Fig. 4c, d) showed that endothelial cells surrounding the spheroids developed sprouting and elongated structures, consistent with the newly formed immature capillary-like structure observed by IF. Magnified images of these immature capillary-like structures revealed dense nanofibers adhering to the surfaces of elongated cells, resembling the self-assembled peptide nanofibers seen in the spheroids. This suggests that CSP-A co-assembles with the original CSP to form nanofibers featuring the angiogenic motif cNGR, enhancing interactions between U87MG spheroids and endothelial cells, ultimately facilitating endothelial tube formation and immature capillary-like structure formation. We subsequently used FITC-Dextran to simulate nutrient transport within the spheroids52,53. The results showed that fluorescent signals were primarily localized at the edges of cell spheroid without immature capillary-like structure (Fig. 4e, f, Supplementary Fig. 20), indicating impaired transport in the inner regions. Notably, in spheroids containing immature capillary-like structures, there was deeper penetration of the fluorescently labeled dextran, with strong signals observed in the interior. We cultured CSP-treated tumor spheroids with CSP-A (without HUVECs) and observed their growth over time (Supplementary Fig. 21). The spheroids gradually increased in size, showing no significant differences between experimental groups. Furthermore, neither structural loosening of the spheroids nor significant penetration of FITC-dextran into the spheroid cores were observed (Supplementary Fig. 22). These results suggest that nutrient transport is significantly enhanced in spheroid tissues with capillary-like structures, highlighting the importance of capillary-like structures in facilitating nutrient distribution within cellular constructs. As the culturing time increases to 7 d, the spheroids with capillary-like structures do gradually grow larger, and multiple cell spheroids can merge to form a larger structure (Supplementary Fig. 23).
a Schematic representation of spheroids with capillary-like structures formed by co-culturing U87MG spheroids with HUVECs treated with peptides CSP-A. b Immunofluorescence imaging showing capillary-like structure formation (CD31 expression) in the spheroids. c, d Scanning electron microscopy (SEM) images illustrating the structure of spheroids with capillary-like structures, with U87MG spheroids marked in yellow and HUVECs in red. e, f) Schematic and confocal images depicting the distribution of FITC-Dextran (70 kDa, 10 μM in PBS for 15 min) in spheroids and spheroids with capillary-like structures. Created in BioRender. Li, Y. (2025) https://BioRender.com/1yi5qot.
Transcriptomic analysis of tumor spheroids with capillary-like structures reveals enhanced similarity to patient-derived GBM
To assess the similarity of tumor spheroids with capillary-like structures to clinical tumor tissue, we analyzed the transcriptomics of these cells. Gene differential analysis (Supplementary Figs. 24, 25) revealed that compared with cells and spheroids, the spheroid +HUVEC group was located closer to the patient-GBM, suggesting that the transcriptional characteristics of the two are more similar, indicating that the spheroid model with capillary-like structures better simulates the clinical GBM samples at the transcriptional level. Gene Set Enrichment Analysis (GSEA) (Fig. 5a) indicated that spheroids with capillary-like structures shared a gene signature with patient GBM, particularly in pathways related to ECM protein interactions.
a Gene Set Enrichment Analysis (GSEA) comparing spheroids with capillary-like structures to patient-GBM using GO and KEGG methods. Two-tailed unpaired Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). b Gene expression levels of angiogenesis-related genes in HUVEC and spheroids with capillary-like structures (n = 3). Data are presented as mean value ± SD, and individual data points are shown as circles. c Gene expression levels of GBM oncogenes in mono U87MG cells, spheroids with capillary-like structures, and clinical GBM (n = 3). Data are presented as mean value ± SD, and individual data points are shown as circles. Source data are provided as a Source Data file.
Specifically, angiogenic gene hallmarks in the spheroids with capillary-like structures showed significantly higher expression levels comparable to those in HUVECs (Fig. 5b), which proved that the presence of immature capillary-like structures enhanced spheroid growth and proliferation. In terms of GBM-associated oncogenic pathways (Fig. 5c), such as EGFR and JAK-STAT54, spheroids with capillary-like structures closely mimicked patient GBM gene expression, surpassing that of 2D tumor cells. Our transcriptomic analyses of spheroids with capillary-like structures (U87MG+HUVECs) and patient GBM samples (GSE119834) reveal two key findings: First, HUVEC-specific endothelial tube formation signatures in mixed cultures are distinguishable from tumor cell profiles, confirming capillary-like structure contributions. Second, conserved oncogenic pathways in patient GBMs are recapitulated in pure U87MG cells and spheroids with capillary-like structures, supporting their clinical relevance. While bulk sequencing cannot fully resolve cell-type-specific signals, our controlled comparisons-using HUVEC-only groups and tumor-only controls-provide critical context for interpreting mixed-population data. These results highlight the importance of microenvironmental crosstalk in GBM biology. These findings confirm that peptide nanofibers effectively modulate the ECM environment and induce capillary-like structure formation, resulting in tumor spheroids with immature capillary-like structures that more accurately represent clinical tumor tissues compared to conventional 2D cell models.
Enhancing tumorigenicity and functionality in engineered spheroids through capillary-like structure formation and intercellular self-assembly
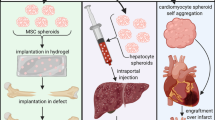
Angiogenesis plays a crucial role in tumor development and progression, as it enables the formation of new blood vessels to supply nutrients and oxygen to growing tumors. In tumor models, the presence of vascularization is essential for accurately mimicking the physiological conditions found in human cancers. Angiogenesis not only supports tumor growth but also facilitates the dissemination of cancer cells, contributing to metastasis. Therefore, establishing a tumor model with robust angiogenic features is vital for studying tumor biology, assessing therapeutic responses, and developing effective treatments. By incorporating angiogenesis into our tumor models, we can better understand the complex interactions between tumor cells and the vascular environment, ultimately leading to more effective cancer therapies. To investigate the tumorigenicity of peptide-induced U87MG cell spheroids and immature capillary-like structures containing spheroids, we established a subcutaneous tumor model in immunodeficient mice (Fig. 6a). Throughout the growth period, there were no significant changes in body weight among the four groups (Supplementary Fig. 26a). Tumor weights were similar in the single-cell and spheroid groups, while the matrigel group showed higher weights. Notably, the spheroids with capillary-like structures exhibited the highest tumor weights, indicating greater tumorigenicity (Supplementary Fig. 26b).
a Schematic representation of the U87MG tumor model in mice, illustrating the grafting of cells, cells in Matrigel, spheroids, and spheroids with capillary-like structures, with subsequent evaluation of tumor growth and capillary-like structure formation. b Tumor volume growth curves for the four experimental groups (n = 5). Data are presented as mean value ± SD. Two-way ANOVA (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). c Immunofluorescent images showing capillary-like structure formation (CD31) in tumors collected on days 16 and 33. d Quantitative assessment of capillary-like structure formation (CD31) in tumors from days 16 and 33 (n = 10). Data are presented as mean value ± SD, and individual data points are shown as circles. Two-way ANOVA (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). e Schematic illustrating MIN6 cell spheroids and the capillary-like structure formation activity induced by self-assembled peptides, highlighting cellular signaling communication. f Differential interference contrast (DIC) images of MIN6 monocellular cultures, pseudoislets (spheroids) induced by incubation with peptides (CSP/CSP-RGD = 10:1, 50 μM) for 24 h and angio-pseudoislets formed by MIN6 spheroids and HUVECs induced by peptides (CSP-A) for 24 h. g Schematic and evaluation of glucose-stimulated insulin secretion in MIN6 cells (n = 4). Data are presented as mean value ± SD, and individual data points are shown as circles. Two-way ANOVA (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001). Source data are provided as a Source Data file.
Tumor volumes were also greater in the Matrigel group compared to the single-cell and spheroid groups (Fig. 6b). However, tumor growth in these three groups plateaued within 18 days post-inoculation, likely due to hypoxic conditions limiting nutrient and oxygen delivery. In contrast, the spheroids with capillary-like structures consistently formed larger tumors (Supplementary Fig. 27). The matrigel group produced smaller tumors, while enhancing cell-matrix interactions, but it lacked endothelial cell angiogenesis55. We evaluated capillary-like structure formation (CD31 and α-SMA) at 16- and 33-day post-inoculation (Fig. 6c, Supplementary Fig. 28). After 16 days, the single-cell, spheroid, and matrigel groups showed low capillary-like structure formation, consistent with the growth plateau. The spheroids with capillary-like structures demonstrated the highest level of endothelial tube formation, suggesting that immature capillary-like structures within the spheroids facilitate essential transport for tumor growth, including nutrient and oxygen delivery, as well as metabolic waste removal.
To assess whether the ability to promote capillary-like structure formation by intercellular self-assembly of the CSP-A/CSP peptides is also applicable in organoid-like spheroid formation of functional cells other than tumors, we co-cultured MIN6 cells, a pancreatic islet insulin-secreting β-cell line with HUVECs and our peptide nanofibers. The pancreatic islets are highly vascularized in vivo since they have to produce insulin in a rapid and dynamic fashion in response to external stimuli56. Since the level of vascularization is highly important to the regulation of insulin secretion of the pancreatic islet β-cells, the ability to produce insulin in response to glucose stimulation would be directly reflective of the induction of immature capillary-like structures by the CSP-A/CSP peptide nanofibers. As a result, considering the nature of MIN6 cells, peptide CSP-RGD was initially introduced to enhance cellular adhesion. Successful formation of MIN6 cell spheroids was observed without compromising cellular function (Fig. 6e, f, Supplementary Fig. 29). The MIN6 cell spheroids were then treated with peptide CSP-A and co-cultured with the HUVECs for potential establishment of immature capillary-like structures as already detailed previously. Subsequent functional analysis showed that the CSP-A/CSP-RGD MIN6/HUVEC spheroids exhibited the highest level of glucose-stimulated insulin secretion (Fig. 6g), with significantly higher stimulation index than the no spheroid single-cell MIN6 group and the non-vascularized MIN6 spheroid group.
These results demonstrate that (1) the CSP-based peptide self-assembly induction ability in cell spheroid formation could be used in multiple cell types, both cancerous cells as well as functional cell types. (2) The facilitated capillary-like structure formation by the CSP-based peptide nanofibers not only enhances cell-cell interactions and cell proliferation, but also significantly improves the functional performance of the organoid-like spheroids. Our work underscores the potential of intercellular self-assembly techniques to create complex tissue architectures that can be tailored for specific therapeutic outcomes.
Discussion
This work introduces a unique strategy for engineering cell spheroids with capillary-like structures through in situ self-assembly of peptide nanofibers. By facilitating capillary-like structure formation within tumors and pancreatic islet spheroids, we have developed models that closely mimic the physiological characteristics of human tissues, significantly enhancing their utility in biomedical research. A critical aspect of our approach is the role of in situ self-assembly in inducing mechanotransduction. The formation of capillary-like structures not only improves nutrient and oxygen delivery but also enhances intercellular signaling through mechanosensitive pathways. This dual action is vital for maintaining cellular viability and function in three-dimensional environments. The significant increase in insulin secretion observed in pancreatic islet spheroids with capillary-like structures illustrates the practical implications of our strategy for regenerative medicine, particularly in developing advanced therapies for diabetes. Moreover, the gene expression profiles of tumor spheroids with capillary-like structures showed similarities to those of patient-derived tumors, underscoring the relevance of our engineered models for cancer research. This alignment enables more effective evaluations of therapeutic strategies and provides deeper insights into tumor biology. By bridging the gap between in vitro models and clinical realities, our work offers a powerful platform for investigating the complexities of tumor microenvironments and their responses to treatment.
However, it is important to acknowledge the limitations of the current model. While we observe the formation of CD31-positive, capillary-like structures and enhanced molecular transport, these in vitro networks may not fully recapitulate the complexity, stability, or perfusive functionality of true in vivo vasculature. In this project, our structures are better described as pro-angiogenic endothelial networks or capillary-like assemblies that represent an important step toward vascularization. Future work will focus on incorporating additional cell types, including immune, stromal, and mural cells, to achieve long-term stability, perfusion, and hierarchical organization of these networks for a more accurate mimicry of native vasculature.
Despite these limitations, the ability to leverage intercellular self-assembly for capillary-like structure formation and mechanotransduction represents a significant advancement in tissue engineering. This approach not only enhances the structural and functional capabilities of engineered tissues but also opens an avenue for exploring intricate tissue interactions and cellular dynamics. The mechanotransduction induced by immature capillary-like structure formation is particularly promising, as it could lead to improved drug delivery systems and more effective cancer therapies. In summary, this research establishes a framework for engineering vascularized spheroids that emphasizes the importance of in situ self-assembly and mechanotransduction. By focusing on the interplay between vascularization and cellular function, this work paves the way for innovative strategies to address some of the most pressing challenges in tissue regeneration and cancer treatment. The insights gained from this work have the potential to significantly impact the fields of tissue engineering and regenerative medicine, ultimately improving therapeutic outcomes and patient care.
Methods
Materials sources
All reagents were of analytical grade and used as received without further purification. Fluorenylmethyloxycarbonyl (Fmoc-) protected amino acids, as well as 2-(1H-benzotriazole-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate (HBTU) and O-(benzotriazol-1-yl)-N,N,N’,N’-tetramethyluronium hexafluorophosphate (HATU), were obtained from GL Biochem Ltd. (China). 1-Pyrenebutyric acid and 4-chloro-7-nitrobenzofurazan (NBD-Cl) were sourced from Sigma-Aldrich. Piperidine and trifluoroacetic acid (TFA), along with N,N-diisopropylethylamine (DIPEA), were supplied by Sinopharm Chemical Reagent Co., Ltd. (Shanghai). Triisopropylsilane (TIPS), N,N-dimethylformamide (DMF), and dichloromethane (DCM) were purchased from J&K Scientific.
Peptide synthesis and characterization
The peptides used in this study were synthesized via solid-phase peptide synthesis (CSP as an example, Supplementary Fig. 30). 2-Chlorotrityl chloride resin was selected. The resin was fully swollen in DCM for 15 min in a peptide synthesis tube. The first amino acid (2 eq) was weighed and dissolved in DCM. After adding DIPEA (6 eq), it was transferred to the above-mentioned peptide synthesis tube containing the resin and reacted for 1 h. The reaction solution was then removed. A mixed solution (DCM: MeOH: DIPEA = 17: 2: 1, v/v/v) was prepared and added to the synthesis tube to block the unreacted resin. The resin was washed with DCM and DMF successively. A DMF solution containing piperidine (20%) was added to the above synthesis tube and reacted for 30 min to completely remove the protecting group (Fmoc). Then the resin was washed with DCM and DMF. The second amino acid (3 eq) and HBTU (3 eq) were weighed and dissolved in DMF. After adding DIPEA (3 eq), the solution was transferred to the synthesis tube and reacted for 2 h. The resin was washed six times with DCM and DMF. After a 30 min deprotection reaction, the resin was washed with DCM and DMF. Then the next amino acid was reacted in the same way as the second one. Repeat this process until the last amino acid was reached. Cleavage was performed using a reagent mixture of 2.5% H₂O, 2.5% triisopropylsilane, and 95.0% trifluoroacetic acid for 30 min.
For the cyclic peptide CSP-A (Supplementary Fig. 31), the thiol groups of the two cysteines were reacted through the air oxidation method to form a disulfide bond. The linear crude peptide synthesized by solid-phase peptide synthesis (50 mg) was dissolved in MeCN/H2O (40:60, v/v), and the pH of the reaction solution was adjusted to 8.5-9.0 using ammonia. After stirring for 48 h, the solvent was removed by vacuum rotary evaporation, and the peptide product was purified using HPLC, yielding approximately 70%.
The crude peptides were then purified through high-performance liquid chromatography (HPLC) and lyophilized into powder form. Each peptide was analyzed using LC-MS (Agilent 1260 Infinity) (Supplementary Figs. 32–39) with a C18 column, employing a flow rate of 1 mL/min and a mobile phase composed of acetonitrile (0.5‰ TFA) and deionized water (0.5‰ TFA). The molecular structures of the peptides were confirmed by nuclear magnetic resonance (NMR) spectroscopy (1H NMR, Bruker 500 MHz AVANCE NEO), utilizing d6-DMSO as the solvent for analysis (Fig. S40-47).
Liquid chromatography-mass spectrometry (LC-MS)
Synthesized peptides were dissolved in methanol at a concentration of 0.2 mg/mL. The purity and molecular weight of the samples were analyzed using LC-MS (Agilent 1260 Infinity) equipped with a C18 column. The mobile phase consisted of acetonitrile (0.5‰ TFA) and deionized water (0.5‰ TFA).
Circular dichroism (CD)
A 100 μL aliquot of the assemblies was added to a quartz cell with a 0.1 cm path length, and the CD signal was recorded from 180 nm to 280 nm at a scan speed of 100 nm/min using circular dichroism spectroscopy (Applied Photophysics Ltd, UK).
Transmission Electron Microscopy (TEM)
A 5 μL solution of peptides was placed on 200 mesh carbon-coated copper grids. After a 1-min incubation, excess solution was removed with filter paper. A 5 μL solution of 2% (w/v) uranyl acetate was added for staining for 1 min. Samples were imaged using a Talos L120C transmission electron microscope (Thermo Fisher), and diameters were analyzed using ImageJ.
Cryo-electron microscopy (Cryo-EM)
Ten microliters of assemblies were placed on a grid, and excess samples were removed with filter paper. The grid was then plunged into precooled liquid ethane. Observations were conducted using a 200 kV Cryo-EM (Glacios, US).
Bio-EM
Bio-EM was utilized to investigate the microstructure of U87MG cells and spheroids. U87MG cells were treated with peptides CSP at a concentration of 50 µM for varying durations, with blank medium as a control. Cells and spheroids were collected using a scraper, centrifuged, and washed three times with cell living buffer. They were then incubated overnight at 4 °C in a mixture of 2% PFA and 2.5% glutaraldehyde. Subsequent steps involved washing with PB buffer (0.1 M) three times, fixation with 1% osmium acid for 1 h at 4 °C, and further washing with PB buffer and H₂O. Cells were stained with 1% uranyl acetate for 1 h, dehydrated through a series of ethanol concentrations, and prepared for embedding in “812” resin. The embedded samples were sliced and stained before observation with a 120 kV TEM.
Cell culture
U87MG, Saos-2, SK-OV-3, HepG2, and HUVEC cells were obtained from ATCC, while MIN6 cells were sourced from AddexBio Technologies. U87MG cells were cultured in DMEM supplemented with 10% heat-inactivated fetal bovine serum (FBS). Saos-2 cells were maintained in McCoy’s 5A medium with 15% heat-inactivated FBS. SK-OV-3 cells were also cultured in McCoy’s 5 A medium, but with 10% heat-inactivated FBS. HepG2 cells were grown in minimum essential medium supplemented with 10% heat-inactivated FBS. HUVEC cells were cultured in Vascular Cell Basal Medium supplemented with the Endothelial Cell Growth Kit-BBE. MIN6 cells were maintained in DMEM supplemented with 10% heat-inactivated FBS and 55 μM β-mercaptoethanol. All cell lines were supplemented with 100 U/mL penicillin and 100 µg/mL streptomycin and were cultured in a humidified atmosphere of 5% CO₂ at 37 °C.
Cell viability assay
To assess cell viability following peptide treatment, 10 μL of Alamar Blue reagent (Invitrogen) was added to each well at specified time points. After 4 h of incubation at 37 °C with 5% CO₂, the culture medium was collected, and fluorescence was measured with an excitation wavelength of 560 nm and emission at 590 nm.
Cell spheroid culture
Cells were seeded in a 96-well culture plate at a density of 2 × 105 cells/mL for 24 h to allow attachment. Peptide-containing fresh culture medium was prepared. Following incubation with the culture medium for varying durations, cells gradually formed spheroids. Additionally, cells diluted directly into peptide-containing medium without prior attachment were also assessed for spheroid formation. The projected area of the spheroids was measured using ImageJ.
Spheroid with capillary-like structure Construction
In our experiment, the tumor spheroids were mechanically isolated without enzymatic treatment. All the CSP-treated cells which contain spheroids and single cells in the wells are collected. The harvested cells are mixed with HUVEC cells (2 × 105 cells/mL) and seeded into a 96-well culture plate for 24 h to facilitate attachment. CSP-A (50 µM) is added 24 h after seeding into a new plate. For Matrigel method, 100 μL Matrigel is added to a 96-well culture plate. Then, U87MG cells (2 × 105 cells/mL) or CSP-treated spheroids and HUVECs (2 × 105 cells/mL) are mixed and seeded into a 96-well culture plate. The cells grow on the top of the Matrigel. After culturing for 96 h, the projected area of the spheroids is measured using Image J. For Agarose method, U87MG cells (2 × 105 cells/mL) or CSP-treated spheroids and HUVECs (2 × 105 cells/mL) are mixed and seeded into a 1% agarose pre-treated 96-well culture plate. After culturing for 96 h, the projected area of the spheroids is measured using Image J.
Immunofluorescent imaging
After incubating U87MG cells for 72 h with peptide-containing medium (50 μM) at 37 °C in a humidified atmosphere of 5% CO₂, cells were washed three times with PBS buffer. They were then fixed with 4% formaldehyde for 15 min at 37 °C and permeabilized with 0.1% Triton X-100 in PBS for 10 min, followed by PBS washing. Cells were blocked with 3% bovine serum albumin (BSA) in PBS for 1 h at room temperature and stained with primary antibodies diluted in 1% BSA in PBS overnight at 4 °C. After incubating with Alexa Fluor-conjugated secondary antibody for 1 h at room temperature, nuclei were stained with 1 µg/mL Hoechst 33342 in PBS for 15 min. After washing with PBS, confocal laser scanning microscopy (CLSM) was used for detection.
AFM measurement
U87MG cells were treated for 12 h in 6-well culture plates before assessing their mechanical properties using atomic force microscopy (AFM). The adhesion force was measured using micropipette cantilevers, which were not coated with any ligand or PLL, with an 8 μm diameter aperture and a nominal spring constant of 2N/m. The force exerted on the cantilever, proportional to its deflection, was recorded by the photodetector. Ten cells per condition were analyzed.
GSIS test of MIN6 cells
MIN6 cells were plated on 6 cm plates for GSIS analysis. After peptide treatment for 2 days, cells were washed twice with KRBH buffer. They were then incubated with 1 mL of KRBH buffer containing 2.8 mM glucose for 1 h at 37 °C. Following this, the buffer was replaced with either 1 mL of KRBH with 2.8 mM glucose or 1 mL of KRB with 16.8 mM glucose for 1 h. The supernatant was collected, centrifuged, and 1 mL of the upper supernatant was prepared for insulin measurement using a mouse insulin enzyme-linked immunosorbent assay (Mouse Insulin ELISA, ALPCO, NH, USA) according to the manufacturer’s instructions. The stimulation index (SI) was calculated as the insulin level under high-glucose conditions divided by that under low-glucose conditions.
Transcriptomics study
To evaluate gene expression changes in U87MG cells, spheroids, and spheroids with capillary-like structures compared to clinical patient GBM samples, RNA-seq analysis was performed. The reported gene expression set of patients GBM samples (GSE119834) was collected. U87MG cells are seeded in a 96-well culture plate at an initial density of 2 × 105 cells/mL for 24 h to allow attachment. Then, peptide CSP is given to form spheroids. Harvest the spheroids in one well, mix them with HUVECs at an initial density of 2 × 105 cells/mL. Seed them into a new 96-well culture plate. After 24 h, add CSP-A to induce the formation of spheroids with capillary-like structures. All of the spheroids in one well are harvested to do this experiment. Replicate this experiment three times. Total RNA was isolated from all samples using TRIzol RNA Isolation reagent (Invitrogen) following the manufacturer’s protocol. RNA quality was assessed using the TapeStation RNA Assay (Agilent Technologies). Libraries were constructed with the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina, amplifying 25 to 75 ng of RNA using 14 cycles of PCR. Final library quality was evaluated by TapeStation DNA HS Assay (Agilent Technologies). Equimolar pooling of libraries was performed based on Qubit values and loaded in triplicates onto an Illumina NextSeq 500 platform for gene expression analysis. Expression levels were calculated using RSEM and processed with the Bioconductor package DESeq2 (version 1.24.0 in R). Data normalization was performed using Trimmed Mean of M values (TMM), and differentially expressed genes were identified with a false discovery rate (FDR)-adjusted P-value of <0.05. Principal component analysis (PCA) and gene set enrichment analysis (GSEA) were conducted with KEGG and GO methods.
Transport experiment in spheroids
Cells, spheroids, and spheroids with capillary-like structures were prepared with peptides and blank controls. Cells were washed with living buffer solution and treated with FITC-dextran (70 kDa, 10 µM in PBS) for 15 min, followed by three washes with PBS. Fluorescence of FITC in different layers of spheroids was examined using CLSM.
In vivo experiment
Female NSG mice (6–8 weeks) were obtained from Westlake University Animal Center and maintained under sterile conditions. Animal experiments were approved by the Animal Care and Use Committee of Westlake University (License number: AP#22-050-WHM-2). Mice were inoculated subcutaneously with U87MG cells (2 × 106/100 μL in PBS or Matrigel), spheroids, and spheroids with capillary-like structures. Tumor volume and body weight were measured every three days, with tumor volume calculated as width² × length/2. Mice were sacrificed 16 days post-inoculation, and tumors were collected for immunofluorescent imaging of vascularization markers (CD31 and α-SMA). All tumor-bearing mice were sacrificed at the end of the experiment, and tumor capillary-like structure formation was evaluated through immunofluorescence. Images (n = 10) were taken at 40× magnification and analyzed using ImageJ.
Statistical analysis
All experimental data are expressed as mean ± standard deviation (SD). Statistical comparisons between two groups were performed using a two-tailed unpaired Student’s t test. Statistical comparisons between more than two groups were performed using One-way ANOVA. Two-way ANOVA was used for comparisons among groups influenced by two factors, with statistical significance set at P < 0.05. Significance levels were indicated as *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Ethical statement
All mice studies were approved by the Institutional Animal Care and Use Committee of Westlake University (AP# 22-050-WHM-2). Female NSG mice (6-8 weeks) were obtained from Westlake University Animal Center. All male mice were fed with non-fluorescent chow. Housing conditions: 12 h dark/12 h light cycle, ambient temperature: 20–25 °C, Humidity: 45–55%. The maximal tumor burden is defined as a mass reaching 10% of body weight or a maximum diameter of 2 cm.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data that support the findings are available within the main text and the supplementary materials. Source data are provided with this paper. The RNA-seq data generated in this study have been deposited in the NCBI database under accession number PRJNA1367769. Source data are provided with this paper.
References
Mouw, J. K., Ou, G. & Weaver, V. M. Extracellular matrix assembly: a multiscale deconstruction. Nat. Rev. Mol. Cell Biol. 15, 771–785 (2014).
Theocharis, A. D., Skandalis, S. S., Gialeli, C. & Karamanos, N. K. Extracellular matrix structure. Adv. Drug Deliv. Rev. 97, 4–27 (2016).
Bonnans, C., Chou, J. & Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 15, 786–801 (2014).
Rosales, A. M. & Anseth, K. S. The design of reversible hydrogels to capture extracellular matrix dynamics. Nat. Rev. Mater. 1, 1–15 (2016).
Kyriakopoulou, K., Piperigkou, Z., Tzaferi, K. & Karamanos, N. K. Trends in extracellular matrix biology. Mol. Biol. Rep. 50, 853–863 (2023).
Hynes, R. O. Stretching the boundaries of extracellular matrix research. Nat. Rev. Mol. Cell Biol. 15, 761–763 (2014).
Geiger, B., Bershadsky, A., Pankov, R. & Yamada, K. M. Transmembrane crosstalk between the extracellular matrix and the cytoskeleton. Nat. Rev. Mol. Cell Biol. 2, 793–805 (2001).
Chaudhuri, O., Cooper-White, J., Janmey, P. A., Mooney, D. J. & Shenoy, V. B. Effects of extracellular matrix viscoelasticity on cellular behaviour. Nature 584, 535–546 (2020).
Kechagia, J. Z., Ivaska, J. & Roca-Cusachs, P. Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 20, 457–473 (2019).
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Stowers, R. S. et al. Matrix stiffness induces a tumorigenic phenotype in mammary epithelium through changes in chromatin accessibility. Nat. Biomed. Eng. 3, 1009–1019 (2019).
Stupack, D. G. & Cheresh, D. A. ECM remodeling regulates angiogenesis: endothelial integrins look for new ligands. Sci. STKE 2002, pe7–pe7 (2002).
Wade, R. J. & Burdick, J. A. Engineering ECM signals into biomaterials. Mater. Today 15, 454–459 (2012).
Xu, Y. et al. Convergent synthesis of diversified reversible network leads to liquid metal-containing conductive hydrogel adhesives. Nat. Commun. 12, 2407 (2021).
Zhu, M. et al. In vivo engineered extracellular matrix scaffolds with instructive niches for oriented tissue regeneration. Nat. Commun. 10, 4620 (2019).
Ai, S. et al. A SupraGel for efficient production of cell spheroids. Sci. China Mater. 65, 1655–1661 (2022).
Zhang, Y. et al. Mesenchymal stem cell spheroids induced by supramolecular nanofibers for diabetic wound healing. Adv. Funct. Mater. 34, 2314607 (2024).
Wang, H. et al. An in situ dynamic continuum of supramolecular phosphoglycopeptides enables formation of 3D cell spheroids. Angew. Chem., Int Ed. 56, 16297–16301 (2017).
Guo, J. et al. Cell spheroid creation by transcytotic intercellular gelation. Nat. Nanotechnol. 18, 1094–1104 (2023).
Hinderer, S., Layland, S. L. & Schenke-Layland, K. ECM and ECM-like materials—biomaterials for applications in regenerative medicine and cancer therapy. Adv. Drug Deliv. Rev. 97, 260–269 (2016).
Kyburz, K. A. & Anseth, K. S. Synthetic mimics of the extracellular matrix: how simple is complex enough? Ann. Biomed. Eng. 43, 489–500 (2015).
Elosegui-Artola, A. The extracellular matrix viscoelasticity as a regulator of cell and tissue dynamics. Curr. Opin. Cell Biol. 72, 10–18 (2021).
Datta, P., Ayan, B. & Ozbolat, I. T. Bioprinting for vascular and vascularized tissue biofabrication. Acta Biomater. 51, 1–20 (2017).
Grebenyuk, S. et al. Large-scale perfused tissues via synthetic 3D soft microfluidics. Nat. Commun. 14, 193 (2023).
Jiang, Y. et al. Targeting extracellular matrix stiffness and mechanotransducers to improve cancer therapy. J. Hematol. Oncol. 15, 34 (2022).
Yang, X., Lu, H., Tao, Y., Zhou, L. & Wang, H. Spatiotemporal control over chemical assembly in living cells by integration of acid-catalyzed hydrolysis and enzymatic reactions. Angew. Chem. Int. Ed. 60, 23797–23804 (2021).
Liu, Z., Guo, J., Qiao, Y. & Xu, B. Enzyme-instructed intracellular peptide assemblies. Acc. Chem. Res. 56, 3076–3088 (2023).
Zhang, Y. et al. Molecular recognition remolds the self-assembly of hydrogelators and increases the elasticity of the hydrogel by 106-fold. J. Am. Chem. Soc. 126, 15028–15029 (2004).
Ciucurel, E. C. & Sefton, M. V. A poloxamine–polylysine acrylate scaffold for modular tissue engineering. J. Biomater. Sci. Polym. Ed. 22, 2515–2528 (2011).
Aksenov, M. Y., Aksenova, M. V., Mactutus, C. F. & Booze, R. M. HIV-1 protein-mediated amyloidogenesis in rat hippocampal cell cultures. Neurosci. Lett. 475, 174–178 (2010).
Millán J. L. Mammalian Alkaline Phosphatases: From Biology to Applications in Medicine and Biotechnology (Wiley, 2006).
Fedde, K. N., Lane, C. C. & Whyte, M. P. Alkaline phosphatase is an ectoenzyme that acts on micromolar concentrations of natural substrates at physiologic pH in human osteosarcoma (SAOS-2) cells. Arch. Biochem. Biophys. 264, 400–409 (1988).
Zhou, J. et al. Enzyme-instructed self-assembly for spatiotemporal profiling of the activities of alkaline phosphatases on live cells. Chem 1, 246–263 (2016).
Nicolas, J. et al. 3D extracellular matrix mimics: fundamental concepts and role of materials chemistry to influence stem cell fate. Biomacromolecules 21, 1968–1994 (2020).
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Craig, S. W. & Chen, H. Lamellipodia protrusion: moving interactions of vinculin and Arp2/3. Curr. Biol. 13, R236–R238 (2003).
Spector, I., Shochet, N. R., Kashman, Y. & Groweiss, A. Latrunculins: novel marine toxins that disrupt microfilament organization in cultured cells. Science 219, 493–495 (1983).
Hetrick, B., Han, M. inS., Helgeson, L. ukeA. & Nolen, B. radJ. Small molecules CK-666 and CK-869 inhibit actin-related protein 2/3 complex by blocking an activating conformational change. Chem. Biol. 20, 701–712 (2013).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Kandasamy, S. et al. The YAP1 signaling inhibitors, verteporfin and CA3, suppress the mesothelioma cancer stem cell phenotype. Mol. Cancer Res. 18, 343–351 (2020).
Auger, F. A., Gibot, L. & Lacroix, D. The pivotal role of vascularization in tissue engineering. Annu. Rev. Biomed. Eng. 15, 177–200 (2013).
Laschke, M. W. et al. Angiogenesis in tissue engineering: breathing life into constructed tissue substitutes. Tissue Eng. 12, 2093–2104 (2006).
Shirure, V. S., Hughes, C. C. W. & George, S. C. Engineering vascularized organoid-on-a-chip models. Annu Rev. Biomed. Eng. 23, 141–167 (2021).
Bieker, R. et al. Infarction of tumor vessels by NGR-peptide–directed targeting of tissue factor: experimental results and first-in-man experience. Blood 113, 5019–5027 (2009).
Wang, X. et al. NGR-modified micelles enhance their interaction with CD13-overexpressing tumor and endothelial cells. J. Controlled Release 139, 56–62 (2009).
Walter-Yohrling, J., Pratt, B. M., Ledbetter, S. & Teicher, B. A. Myofibroblasts enable invasion of endothelial cells into three-dimensional tumor cell clusters: a novel in vitro tumor model. Cancer Chemother. Pharm. 52, 263–269 (2003).
Ehsan, S. M., Welch-Reardon, K. M., Waterman, M. L., Hughes, C. C. W. & George, S. C. A three-dimensional in vitro model of tumor cell intravasation. Integr. Biol. 6, 603–610 (2014).
Batlle, R. et al. Regulation of tumor angiogenesis and mesenchymal–endothelial transition by p38α through TGF-β and JNK signaling. Nat. Commun. 10, 3071 (2019).
Sokolova, V. et al. Transport of ultrasmall gold nanoparticles (2 nm) across the blood–brain barrier in a six-cell brain spheroid model. Sci. Rep. 10, 18033 (2020).
Zheng, Y. et al. In vitro microvessels for the study of angiogenesis and thrombosis. Proc. Natl. Acad. Sci. 109, 9342–9347 (2012).
Henrik Heiland, D. et al. Tumor-associated reactive astrocytes aid the evolution of immunosuppressive environment in glioblastoma. Nat. Commun. 10, 2541 (2019).
Benton, G., Arnaoutova, I., George, J., Kleinman, H. K. & Koblinski, J. Matrigel: from discovery and ECM mimicry to assays and models for cancer research. Adv. Drug Deliv. Rev. 79-80, 3–18 (2014).
Yang, J. et al. Bioengineering and vascularization strategies for islet organoids: advancing toward diabetes therapy. Metabolism 152, 155786 (2024).
Acknowledgements
This project was supported by the National Key Technologies Research and Development Program of China (grant number: 2020YFA0803701 to D.L. Kong), the National Natural Science Foundation of China (82272145 to H.M. Wang) and the Foundation of Westlake University. We thank the Instrumentation and Service Center for Molecular Sciences, Instrumentation and Service Center for Physical Sciences, General Equipment Core Facility, Biomedical Research Core Facilities, and Laboratory Animal Resource Center at Westlake University for the assistance with measurements.
Author information
Authors and Affiliations
Contributions
Z.Q.Q.F. and H.M.W. conceptually designed the strategy for this study, provided intellectual input, supervised the studies, and revised the manuscript. H.L.L. designed the study and perform most of the experiments. Y.T.L. designed the study and perform most of the cell experiments in different models. X.J.Y. performed some in vitro characterization of self-assembly. B.H.W. and H.L.L. did animal experiments. C.L. and D.L.K. provide the model building and guide the model comparison. H.M.W., Z.Q.Q.F., Y.T.L., and H.L.L. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, H., Li, Y., Yang, X. et al. Mechanical forces from intercellular peptide self-assembly drive spheroid formation. Nat Commun 17, 1801 (2026). https://doi.org/10.1038/s41467-026-68513-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68513-1