Abstract
Cancer cell fate has been widely ascribed to mutational changes within protein-coding genes associated with tumor suppressors and oncogenes. In contrast, the mechanisms through which the biophysical properties of membrane lipids influence cancer cell survival, dedifferentiation and metastasis have received little scrutiny. Here, we report that cancer cells endowed with high metastatic ability and cancer stem cell-like traits employ ether lipids to maintain low membrane tension and high membrane fluidity. Using genetic approaches and lipid reconstitution assays, we show that these ether lipid-regulated biophysical properties permit non-clathrin-mediated iron endocytosis via CD44, resulting in significant increases in intracellular redox-active iron and enhanced ferroptosis susceptibility. Using a combination of in vitro three-dimensional microvascular network systems and in vivo animal models, we show that loss of ether lipids from plasma membranes also strongly attenuates extravasation, metastatic burden and cancer stemness. These findings illuminate a mechanism whereby ether lipids in carcinoma cells serve as key regulators of malignant progression while conferring a unique vulnerability that can be exploited for therapeutic intervention.
Similar content being viewed by others
Introduction
Cancer cells have the capacity to undergo dynamic changes in identity, structure, and function, making them remarkably versatile and adaptable. Alterations in the lipid composition of cell membranes contribute to the phenotypic plasticity of cells. The distinctive physicochemical properties and subcellular localization of various lipids within cell membranes influence a range of biological processes, including cellular trafficking, signaling and metabolism1. Despite growing knowledge of lipid biology, an understanding of how specific lipid subtypes impact cancer cell fate remains limited.
Emerging studies demonstrate that therapy-resistant mesenchymal-like carcinoma cells exhibit an elevated vulnerability to ferroptosis2,3,4, an iron-dependent form of cell death characterized by an unrestricted accumulation of oxidized membrane phospholipids5,6,7,8. In previous work we showed that the natural product salinomycin can selectively eliminate otherwise therapy-resistant, mesenchymal-enriched cancer stem cells (CSC) by targeting lysosomal iron to promote an iron-dependent cell death4,9,10. In this context, we found that such CSC-enriched cells exhibit a high intracellular iron load compared to their non-CSC-like counterparts, rendering them especially vulnerable to elimination by induced ferroptosis4.
Ferroptosis can also be instigated by pharmacologic inhibition of ferroptosis suppressors, such as glutathione peroxidase 4 (GPX4)5,11, ferroptosis-suppressor protein 1 (FSP1, previously known as AIFM2)12,13, as well as through downregulation of reduced glutathione (GSH)14,15. Activated CD8+ T cells may also induce ferroptosis in cancer cells16,17. Beyond cancer, ferroptosis has been implicated in the pathogenesis of several neurodegenerative diseases and acute injury of the kidney, liver and heart18,19,20,21.
In previous work, we undertook an unbiased, genome-wide CRISPR/Cas9 screen with the goal of identifying genes that govern ferroptosis susceptibility in high-grade human serous ovarian cancer cells22. This screen revealed a role for ether lipid-synthesizing enzymes, such as alkylglycerone phosphate synthase (AGPS), in modulating ferroptosis susceptibility. The ether phospholipids generated by these enzymes represent a unique subclass of glycerophospholipids characterized by an ether-linked hydrocarbon group formed at the sn-1 position of the glycerol backbone23. This phospholipid subtype constitutes ~20% of the total phospholipid pool in many types of mammalian cells.
The significance of ether lipid species in human health is underscored by the severe inherited peroxisomal disorders caused by their deficiency23. This often manifests as profound developmental abnormalities, such as neurological defects, vision and hearing loss, and reduced lifespan. In the context of cancer, elevated ether lipid levels have been correlated with increased metastatic potential of carcinoma cells24,25,26,27. Despite these pathological associations, the mechanism(s) by which ether lipids affect cancer progression remain(s) elusive. Furthermore, the mechanism(s) by which loss of ether lipids results in decreased ferroptosis susceptibility required further investigation.
Our previous work, along with that of others, ascribed a role to polyunsaturated ether phospholipids as chemical substrates prone to the iron-mediated oxidation that triggers ferroptotic cell death22,28. Here, we demonstrate that ether lipids also play an unrelated biophysical role, facilitating iron endocytosis in carcinoma cells. This represents a mechanism of intracellular signaling by which ether lipids contribute to intracellular levels of a critical metal catalyst. In addition, our findings highlight the functional importance of this poorly studied lipid subtype in enabling a variety of malignancy-associated cell phenotypes, including metastasis and tumor-initiating abilities. Taken together, these results establish a role for ether lipids as critical effectors of cancer cell fate.
Results
Ether lipids play a key role in maintaining a ferroptosis-susceptible cell state
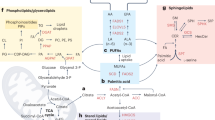
To investigate the mechanism(s) by which ether lipid deficiency reduces ferroptosis susceptibility, we employed CRISPR/Cas9 to knockout (KO) the AGPS gene in ferroptosis-sensitive TGF-β-treated PyMT-1099 murine breast carcinoma cancer cells29 (Supplementary Fig. 1a). The AGPS gene encodes a rate-limiting enzyme critical for ether lipid biosynthesis23 (Fig. 1a). Consistent with our prior studies22, loss of ether lipids via AGPS KO significantly decreased the susceptibility of these cancer cells to ferroptosis induced by treatment with the GPX4 inhibitors RSL3 or ML210 (Fig. 1b, Supplementary Fig. 1b).
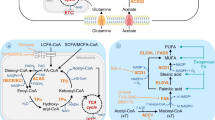
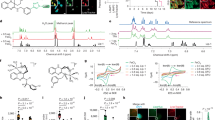
a Schematic of peroxisomal-ether lipid biosynthetic pathway. b Cell viability following treatment with the GPX4 inhibitor RSL3 for 72 h. PyMT-1099 WT, AGPS KO, or AGPS addback cells were pretreated with TGF-β (2 ng/mL) for 10 days prior to assay. Data shown as mean of n = 3 technical replicates and is representative of three independent biological replicates. c Bar graph showing percent of total lipids constituted by ether lipids following AGPS KO in untreated wildtype (WT) or TGF-β-treated (2 ng/mL;10 d) PyMT-1099 cells. d Pie chart showing the relative proportion of ether lipids with various total numbers of double bonds. e Amount in pmol of oxidized phosphatidylethanolamine (Oxi. PE) ether and ester phospholipids in PyMT-1099 TGF-β cells treated with ML210 or vehicle control for 24 h. Five biological replicates per condition. 100,000 cells were used for lipid extraction in each condition. Box plots: interquartile range, center lines = medians and whiskers = the minimum and maximum values. Statistical significance was calculated by unpaired, two-tailed t-test; ns, not significant. f Volcano plot showing the log2 fold change in the relative abundance of various lipid species upon knockout of AGPS in PyMT-1099 TGF-β-treated cells. Blue indicates non-ether linked polyunsaturated phospholipids (PUFA-PLs), red indicates all ether lipids identified in lipidomic analysis and black denotes all other lipids identified. g Volcano plot showing the log2 fold change in the relative abundance of various lipid species upon knockout of AGPS in pB3 cells. Blue indicates non-ether linked polyunsaturated phospholipids (PUFA-PLs), red indicates all ether lipids identified in lipidomic analysis and black denotes all other lipids identified. h Bar graph showing the percent of total lipids constituted by ether lipids in pB3 WT, pB3 AGPS KO and pB3 AGPS addback cells. i Bar graphs showing the effects of ether lipids on the relative abundance of selected polyunsaturated diacyl phospholipids in pB3 cells. All lipidomic data represent n = 3 technical replicates shown as the mean +/− SEM. Unless otherwise stated statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test; ns, not significant. For panel b: PyMT-1099 WT and AGPS KO cells were transduced with the respective vector control plasmids. For panels h-i: pB3 WT and AGPS KO cells were transduced with the respective vector control plasmids.
Lipidomic analysis confirmed that AGPS knockout significantly reduces total ether lipid abundance, while only marginally affecting most other membrane-associated lipid species (Fig. 1c, Supplementary Fig. 2a). More than half of the identified ether lipids contained polyunsaturated fatty acyl groups, which are highly prone to hydrogen abstraction by reactive free radicals (Fig. 1d). Based on this observation, we speculated that loss of ether lipids may attenuate ferroptosis susceptibility by depleting the pool of available ether lipid substrates for lipid peroxidation. Thus, we performed oxidized lipidomic analysis on two ferroptosis-sensitive breast cancer cell lines treated with either ML210 or RSL3. These experiments confirmed that ether lipids are oxidized following ferroptosis induction (Fig. 1e, Supplementary Fig. 1c).
In order to test whether ether lipid deficiency leads to a global reduction of cellular pools of oxidation prone lipid species, we also investigated whether the relative abundance of pro-ferroptosis non-ether-linked polyunsaturated phospholipids was impacted by loss of ether lipids. Our analyses revealed that while total levels of phospholipid unsaturation are only marginally changed, ether lipid deficiency increased the relative abundance of several polyunsaturated diacyl phosphoethanolamine (PE) and phosphatidylcholine (PC)-derived lipids with putative pro-ferroptosis function30,31 (Fig. 1f, Supplementary Fig. 2b-d). To ensure that these findings were not an idiosyncrasy of our TGF-β-treated PyMT-1099 AGPS KO cells, we confirmed this observation in PyMT-MMTV-derived pB3 murine AGPS KO breast carcinoma cells32 (Fig. 1g, h, Supplementary Figs. 1d, 3a–f). Importantly, re-expression of AGPS (addback) restored the relative levels of these non-ether-linked polyunsaturated diacyl phospholipids to levels comparable to pB3 wildtype (WT) cells (Fig. 1i). These findings dispel the notion that ether lipid deficiency attenuates ferroptosis susceptibility simply by decreasing the global level of polyunsaturated phospholipids, further underscoring the importance of polyunsaturated ether phospholipids in maintaining a ferroptosis susceptible cell state.
Ether lipids regulate levels of cellular redox-active iron in cancer cells
The above observations, together with our oxidized ether phospholipidomic analysis, supported the notion that ether lipids could modulate ferroptosis susceptibility—at least in part by serving as substrates for lipid peroxidation. However, these observations failed to address the possibility that alterations in ether lipid composition could also affect intracellular levels of redox-active iron, the central mediator of the lipid peroxidation that drives ferroptosis. Therefore, we used two orthogonal analyses to assess intracellular iron levels. Since the endolysosomal compartment is a key reservoir of reactive iron within cells7,33,34,35, we used a lysosomal iron (II)-specific fluorescent probe, HRhoNox-M36, to gauge levels of lysosomal redox active iron. In addition, we used inductively coupled plasma mass spectrometry (ICP-MS) to quantify total intracellular iron levels7,35,37,38,39.
Unexpectedly, loss of ether lipids (via AGPS KO) reduced intracellular iron levels in all murine cancer cell lines tested, whereas AGPS addback restored intracellular iron to levels comparable to those seen in parental ferroptosis-sensitive cancer cells (Fig. 2a–d, Supplementary Fig. 4a). These findings provided the first indication that changes in ether phospholipids directly affect the levels of intracellular iron. Indeed, there was no direct precedent for the ability of a membrane-associated phospholipid to serve as a key regulator of the levels of an intracellular metal ion.
a Relative lysosomal iron levels (see methods) of PyMT-1099 WT or AGPS KO cells (+/−) pretreatment with TGF-β (n = 5 independent experiments). Fold change calculated relative to the average of untreated PyMT-1099 wild-type (WT) cells. Statistical significance was calculated by one-way ANOVA with Holm-Šídák multiple comparisons test. b Relative lysosomal iron levels of WT, AGPS KO, and AGPS addback pB3 cells (n = 5 independent experiments). Fold change calculated relative to the average of pB3 WT cells. Statistical significance was calculated by one-way ANOVA with Holm-Šídák multiple comparisons test. c Inductively coupled plasma-mass spectrometry (ICP-MS) of cellular iron in PyMT-1099 WT or AGPS KO cells (+/−) pretreatment with TGF-β (n = 6 independent experiments). Statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test. d ICP-MS of cellular iron in WT, AGPS KO, and AGPS addback pB3 cells (n = 4 independent experiments). Statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test. e Relative lysosomal iron levels in OVCAR8 NT sg, FAR1 KO or AGPS KO cells pretreated with ferric ammonium citrate (FAC, 50 μg/mL) (n = 4 independent experiments). Fold change is calculated relative to the average of NT sg cells. Statistical significance was calculated by one-way ANOVA with Holm-Šídák multiple comparisons test. f ICP-MS of cellular iron in OVCAR8 NT sg, FAR1 KO or AGPS KO cells pretreated with FAC (50 μg/mL) (n = 5 independent experiments). Statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test. g Cell viability of OVCAR8 NT sg, FAR1 KO or AGPS KO cells (+/−) FAC pretreatment followed by ML210 treatment for 72 h. Liproxstatin-1 (0.2 μM) was added at the time of ML210. Data shown as mean of n = 3 technical replicates and is representative of three independent biological replicates. h Cell viability in response to ML210 treatment of PyMT-1099 WT or AGPS KO (+/−) pretreatment with TGF-β (10 days) followed by FAC treatment (100 µg/mL) for an additional 24 h. Cells were then treated with ML210 (+/−) liproxstatin-1 (0.2 µM) and cell viability was assessed after 72 h. Data shown as mean of n = 3 technical replicates and is representative of three independent biological replicates. i ICP-MS of cellular iron from primary tumors derived from WT, AGPS KO, and AGPS addback pB3 cells. Mean +/− SEM from 3 independent tumor samples per condition. Group differences were tested by permutation ANOVA followed by one tailed Welch’s t-tests with Holm correction for multiple comparisons. For panels b, d, i: pB3 WT and AGPS KO cells were transduced with the respective vector control plasmids. NT nontargeting, ns not significant.
Supporting these findings, we found that the depletion of ether lipids, achieved via knockout of the fatty acid reductase 1 (FAR1) enzyme23 (Supplementary Fig. 4b), also resulted in a significant decrease in intracellular iron levels, in this case in the OVCAR8 human high-grade serous ovarian cancer cell line, which exhibits mesenchymal characteristics40 (Fig. 2e, f). Furthermore, we noted that treatment of AGPS KO cells with ferric ammonium citrate (FAC)41,42, which provides an exogenous source of ferric ions, re-sensitized cultured AGPS KO mesenchymal breast and ovarian carcinoma cells to ferroptosis induction (that could be rescued by the ferroptosis inhibitor liproxstatin-1), doing so even in the absence of elevated ether phospholipids (Fig. 2g, h).
We further extended this analysis by studying the behavior of mammary carcinoma tumors in vivo. Consistent with our in vitro data, ICP-MS indicated that total iron levels are reduced in breast tumors derived from implanted pB3 AGPS KO cells relative to those arising from either pB3 WT or pB3 AGPS addback cells (Fig. 2i). These observations reinforced the notion that ether lipids are critical regulators of intracellular iron levels, a rate-limiting component of ferroptosis7.
Ether lipids facilitate CD44-mediated iron endocytosis
We proceeded to investigate the mechanism(s) by which membrane-associated ether lipids could regulate intracellular iron levels. This led us to examine the behavior of two proteins that are known to act as major mediators of cellular iron import—transferrin receptor 1 (TfR1)43 and CD4435—and whether their functioning was altered in response to loss of ether phospholipids. While CD44 is best known as a cell-surface cancer stem-cell marker44,45, recent research has revealed its critical role in mediating endocytosis of iron-bound hyaluronates in CSC-enriched cancer cells and in activated immune cells35,39. To monitor these two alternative iron import mechanisms, we performed endocytosis kinetics experiments using fluorescently labeled transferrin as a proxy for TfR1 internalization and fluorescently labeled hyaluronic acid (HA)—whose main plasma membrane receptor is CD44—as a marker for CD44 internalization46,47.
We observed that the rate of TfR1 endocytosis was marginally affected by ether lipid deficiency in pB3 cells (Fig. 3a, Supplementary Fig. 5a). In stark contrast, CD44-mediated endocytosis was significantly impaired in these ether lipid-deficient cancer cells (Fig. 3b, Supplementary Fig. 5b). Conversely, CD44-dependent iron import in AGPS KO cells could be restored to WT levels upon AGPS addback (Fig. 3b, Supplementary Fig. 5b). Reduction in the rate of internalization of CD44 but not TfR1 was also observed in ether lipid-deficient PyMT-1099 TGF-β-treated cells (Fig. 3c, d, Supplementary Fig. 5c, d). Therefore, ether lipids play a critical role in modulating intracellular iron levels by regulating endocytosis of CD44 but not TfR1. These findings were consistent with our previous observations that TfR1 and CD44 localize to distinct endocytic vesicles in CSC-enriched cancer cells35,48, making plausible that their internalization was governed by distinct endocytic mechanisms.
a Endocytic transport of fluorescently labeled transferrin as assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT, AGPS KO, and AGPS addback cells. b Endocytic transport of fluorescently labeled hyaluronate probe as assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT, AGPS KO, and AGPS addback cells. c Endocytic transport of fluorescently labeled transferrin as assessed by quantitative colocalization with an early endosomal marker (EEA1) in PyMT-1099 WT or AGPS KO cells +/− pretreatment with TGF-β. d Endocytic transport of fluorescently labeled hyaluronate probe as assessed by quantitative colocalization with an early endosomal marker (EEA1) in PyMT-1099 WT or AGPS KO cells +/− pretreatment with TGF-β. e Inductively coupled plasma-mass spectrometry (ICP-MS) of cellular iron following treatment with either hyaluronic acid or hyaluronidase in PyMT-1099 WT or CD44 KO cells +/− pretreatment with TGF-β (n = 4 independent experiments). Box plots: interquartile range, center lines = medians and whiskers = the minimum and maximum values. f Endocytic transport of dextran as assessed by quantitative colocalization with the early endosomal marker EEA1 in pB3 WT, AGPS KO, and AGPS addback cells. g Endocytic transport of dextran as assessed by quantitative colocalization with the early endosomal marker EEA1 in PyMT-1099 WT or AGPS KO cells +/− pretreatment with TGF-β. h Endocytosis of EGFR as assessed by quantitative colocalization of internalized fluorescently labeled EGF with an early endosomal marker (EEA1) in PyMT-1099 WT or AGPS KO cells +/− pretreatment with TGF-β. Cells were treated with 2 ng/mL Alexa 555-conjugated EGF. i Endocytic transport of hyaluronate probe assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT or AGPS KO cells +/− pretreatment (16-18 hr) with polyunsaturated fatty acid (PUFA) BSA conjugate C(22:6). j Endocytic transport of hyaluronate probe assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT or AGPS KO +/− pretreatment (16-18 hr) with liposomes composed of the following: PE (18:0_20:4), PE (18:1p_20:4), and PC (18:1p_20:4). All data shown as mean +/− SEM and statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test; ns, not significant. For all endocytosis assays n = 10 fields were examined for each timepoint, and data are representative of two independent experiments with similar results. In some cases, error bars are smaller than the symbol size and not visible. Panels a, b, f: pB3 WT and AGPS KO cells were transduced with the respective vector control plasmids.
To further support the role of CD44 in promoting iron uptake—acting via endocytosis of HA—we demonstrate that knocking out the gene encoding CD44 or, alternatively, treating cancer cells with the HA-degrading enzyme, hyaluronidase, led to a reduction in intracellular iron levels (Fig. 3e). Conversely, supplementing WT cells with HA significantly increased intracellular iron levels (Fig. 3e). Similar observations were seen in human OVCAR8 cells (Supplementary Fig. 7f). Taken together, these observations further supported the central role of CD44 in mediating iron uptake in these cancer cells35.
We then studied whether the defect in CD44 endocytosis observed in ether lipid-deficient cells was limited specifically to CD44 or other cell-surface proteins internalized by a similar mechanism. CD44 is known to undergo a type of clathrin- and dynamin-independent form of endocytosis48,49,50. To test whether loss of ether phospholipids had a wider effect on the clathrin- and dynamin-independent mode of endocytosis, we examined the rate of uptake of dextran (70 kDa), a branched polysaccharide known to undergo endocytosis by a clathrin-independent mechanism49 (pinocytosis). Similar to CD44, we observed that loss of AGPS also induced a significant reduction in the rate of dextran endocytosis; this behavior could be reversed by restoration of ether phospholipid levels achieved by AGPS addback (Fig. 3f, g, Supplementary Fig. 5e, f).
The internalization mechanism of CD44 and dextran differs from that of many plasma-membrane proteins, including TfR1 and EGFR, which undergo clathrin-dependent endocytosis, in which small invaginations of clathrin-coated pits undergo scission facilitated by the GTPase dynamin49,51. As predicted by the distinct actions of these two mechanisms, we observed that loss of ether lipids had a negligible effect on the rate of EGFR endocytosis (Fig. 3h, Supplementary Figs. 6a, 7a, b). Taken together, these observations indicate that internalization of extracellular and cell-surface molecules is mediated by distinct mechanisms which differ in their dependence on plasma membrane ether phospholipids.
To further characterize the clathrin-independent mechanism used by CD44 and dextran, we next pursued both genetic and pharmacologic approaches that target against either the CLIC/GEEC pathway or pinocytosis. We found that inhibition of the CLIC/GEEC pathway via siRNA-mediated knockdown of PICK152 or treatment with 7-keto-cholesterol53,54 significantly reduced CD44 internalization, which in turn led to decreased total cellular and lysosomal iron (Supplementary Fig. 8a–c, e, g, h). We also show that blocking pinocytosis via knockdown of CtBP155,56,57 or treatment with Cytochalasin-D58 is capable of impairing CD44 internalization (albeit, to a lesser extent than CLIC/GEEC inhibition), resulting in lower but non-significant reductions in total cellular and lysosomal iron (Supplementary Fig. 8a–c, d, f, h). These findings suggest that CD44 internalization depends more on the CLIC/GEEC pathway than on pinocytosis for iron delivery into cells.
Given that polyunsaturated phospholipids have been shown to promote endocytosis59, we investigated whether treatment with exogenous polyunsaturated fatty acids (PUFAs) could compensate for loss of ether lipids in regulating CD44 endocytosis. We observed that supplementation with exogenous PUFAs could not fully rescue impaired CD44 endocytosis induced by loss of ether lipids (Fig. 3i, Supplementary Fig. 6b). Furthermore, while liposomes containing ester-linked PUFA-PE phospholipids could only partially rescue CD44 endocytosis, ether-linked PUFA-PE or -PC liposomes achieved an even greater degree of rescue, with ether-linked PUFA-PE liposomes fully restoring CD44 endocytosis to levels exceeding those of WT cells (Fig. 3j, Supplementary Fig. 6c). These findings demonstrate that both the presence of the ether linkage in phospholipids as well as the type of headgroup present within phospholipids play important roles in supporting efficient CD44-mediated endocytosis.
Ether lipid deficiency impairs membrane biophysical properties
The above observations did not provide mechanistic insights into how changes in the composition of membrane ether lipids could exert an effect on CD44 internalization. As observed by others, non-clathrin-mediated endocytosis, which is employed by CD44, is particularly sensitive to changes in the physicochemical properties of the lipid bilayer forming plasma membranes50,60,61,62,63,64,65. Such changes can influence membrane tension and fluidity and the stability and formation of lipid rafts, all of which impact the assembly and dynamics of clathrin-independent, cell-surface endocytic structures. Hence, we hypothesized that ether lipids alter the biophysical properties of the plasma membrane to facilitate elevated iron endocytosis via CD44. To substantiate these hypotheses, we quantified the abundance of ether lipids in the plasma membrane. Lipidomics profiling of pB3-derived giant plasma membrane vesicles (GPMV) showed that ether lipids make up ~23% of plasma membrane lipids and are depleted to ~5% upon AGPS KO (Supplementary Fig. 9a-f).
Alterations in membrane tension have long been demonstrated to affect endocytosis66,67,68,69,70,71. This prompted us to assess the effects of ether lipid deficiency on plasma membrane tension. Membrane tension measures the forces exerted on a defined cross-section of the plasma membrane, which is influenced by both the in-plane tension of the lipid bilayer and the attachment of the plasma membrane to the underlying cell cortex72,73.
To quantify membrane tension directly, we generated a membrane tether using an optically trapped bead and measured the pulling force (\(f\)) and the tube radius (R) to calculate membrane tension (σ) of living cells59,74 (Fig. 4a, b). We found that depletion of ether phospholipids led to a significant increase in membrane tension in pB3 AGPS KO cells relative to the corresponding pB3 WT cells (Fig. 4c). This shift was largely attenuated upon restoration of AGPS expression in pB3 AGPS KO cells or upon exposure of cultured cells to liposomes composed of ether phospholipids (Fig. 4c). Consistent with earlier measurements, treatment of pB3 AGPS KO cells with ether PUFA-PE liposomes increased the rate of CD44 endocytosis to levels comparable to those of pB3 WT cells. In contrast, treatment of pB3 AGPS KO cells with liposomes derived from monounsaturated phosphatidylethanolamine (MUFA-PE) ether lipids only partially rescued CD44 endocytosis (Fig. 4d, Supplementary Fig. 6d). No changes were observed in the rate of clathrin-dependent TfR1 endocytosis under these conditions (Fig. 4e, Supplementary Fig. 6e) and loss of CD44 showed modest effects on membrane tension (Supplementary Fig. 7d, e). Taken together, these results provide evidence that ether lipids facilitate CD44-mediated iron endocytosis in cancer cells, in part by decreasing membrane tension.
a Schematic of membrane tether pulling assay and fluorescence image showing a tether pulled from the plasma membrane of a pB3 cell using an optically trapped 4 µm anti-Digoxigenin coated polystyrene bead. b Graph showing tether radius (R) and tether force measurements (\(f\)) in pB3 WT, AGPS KO, and AGPS addback cells. All data shown as mean +/− SD, data is representative of three independent experiments which yielded similar results. c Membrane tension measurements in pB3 WT, AGPS KO pretreated with the indicated ether phospholipid liposomes, and AGPS addback cells. All data shown as mean +/− SEM of WT: n = 14, AGPS KO: n = 16, AGPS addback: n = 11, AGPS KO + PE (18:1p_18:1): n = 13, AGPS KO + PE (18:1p:20:4): n = 12 independent measurements. d Endocytic transport of fluorescently labeled hyaluronate probe as assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT or AGPS KO cells pretreated with the indicated ether phospholipid liposomes. All data shown as mean +/− SEM. e Endocytic transport of fluorescently labeled transferrin as assessed by quantitative colocalization with an early endosomal marker (EEA1) in pB3 WT or AGPS KO cells pretreated with 20 µM of the indicated ether phospholipid liposomes. All data shown as mean +/− SEM. f Generalized polarization (GP) values of C-laurdan-labeled plasma membranes from pB3 WT (n = 13 technical replicates), AGPS KO (n = 7 technical replicates), and AGPS addback cells (n = 7 technical replicates). Data is shown as mean GP + / − SD. Experiment was repeated three times, yielding similar results. g GP values of C-laurdan-labeled intracellular membranes from pB3 WT (n = 21), AGPS KO (n = 19) and AGPS addback cells (n = 29 technical replicates) Data is shown as mean GP + / − SD. Experiment was repeated three times, yielding similar results. h GP values of C-laurdan-labeled intracellular membranes from PyMT-1099 WT or AGPS KO cells treated with or without 2 ng/mL TGF-β for 10 days. Data shown as mean GP + / − SD of n = 38 (WT), n = 19 (WT + TGF-β), n = 26 (AGPS KO), n = 19 (AGPS KO + TGF-β) technical replicates. Experiment was repeated three times, yielding similar results. i Representative giant plasma membrane vesicle (GPMV) phase separation curves from AGPS KO versus WT pB3 cells. Curves were generated by counting ≥ 20 vesicles/temperature/condition at > 4 temperatures. The data was fit to a sigmoidal curve to determine the Tmisc. Data shown as the mean +/− SD of 3 independent experiments with the average fit shown. Inset shows mean Tmisc +/− SD upon loss of AGPS in pB3 cells. Statistical significance was calculated using unpaired, two-tailed t-test. Unless otherwise noted statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test; ns, not significant. Examined n = 10 fields for all endocytosis-related experiments and two independent replicates were performed with similar results. For panels b, c, f, g: pB3 WT and AGPS KO cells were transduced with the respective vector control plasmids.
Membrane lipid packing can also impact endocytosis75,76,77. This is related to the fluidity or viscosity of the lipid bilayer, with higher lipid packing correlating with higher viscosity. This affects the ease with which proteins and lipids undergo lateral diffusion and conformational changes within a lipid bilayer, thereby affecting endocytosis-related signaling59. This prompted us to investigate the contribution of ether lipids to membrane lipid packing. We used C-laurdan, a lipid-based, polarity-sensitive dye, which yields a spectral emission shift dependent on the degree of lipid packing78. These measurements are used to calculate a unitless index, termed generalized polarization (GP), where a higher GP indicates increased lipid packing78. Our measurements using C-laurdan indicated that a reduction in ether lipid levels resulted in a measurable and significant increase in membrane packing (Fig. 4f–h, Supplementary Fig. 7c) which, like increases in membrane tension, negatively affects membrane deformability75.
The association of CD44 with lipid rafts, which are dynamic plasma membrane nanodomains, is known to be critical for CD44-mediated HA endocytosis79. Previous work has shown that PUFA-containing lipids can stabilize lipid rafts by increasing the packing contrast between ordered and disordered phases80,81,82,83. Thus, we hypothesized that lipid raft stability might be reduced in cells lacking AGPS and therefore PUFA-containing ether lipids which would result in impairment of CD44 endocytosis84. Accordingly, we measured the miscibility transition temperature (Tmisc) which is hypothesized to be related to the stability membrane domains of isolated cellular plasma membranes. We observed a decrease in Tmisc upon loss of AGPS in pB3 cells, which indicate a decrease in lipid raft stability (Fig. 4i). This finding supports a role for ether lipids in maintaining the plasma membrane organization through lipid raft microdomains, revealing yet another biophysical property of lipid bilayers that can influence CD44 endocytosis.
It is noteworthy that clathrin-independent endocytosis exhibits a greater dependency on the membrane biophysical properties assessed above50,60,61,62,63,64,65. This may explain why loss of ether lipids can exert a significant effect on the rate of clathrin-independent, CD44-mediated iron endocytosis but negligible effects on the clathrin-dependent TfR1 endocytosis. Furthermore, these findings revealed a mechanism by which membrane-associated ether lipids could govern a major mechanism of iron internalization, which may impact the vulnerability of cancer cells to ferroptosis inducers.
Loss of ether lipids decreases metastasis and cancer cell stemness
Prior studies have demonstrated that reduced membrane tension and elevated intracellular iron can promote cancer metastasis85,86,87,88,89. These findings prompted us to investigate whether changes in the ether lipid composition of cancer cells impact key steps of the multi-step invasion-metastasis cascade, notably extravasation efficiency, post-extravasation proliferation90 and importantly stemness as manifested by tumor-initiating ability.
We measured extravasation efficiency by employing an in vitro three-dimensional microvascular network system composed of human umbilical vein endothelial cells (HUVECs) and normal human lung fibroblasts. This system has been shown to accurately model some of the complex biological processes associated with cancer cell extravasation91,92,93,94,95. Using this defined experimental system, we found that loss of ether lipids significantly decreased extravasation efficiency (Fig. 5a–c). Furthermore, we observed a strong reduction in overall metastatic burden following intracardiac injection in syngeneic hosts of the pB3 AGPS KO cells relative to corresponding WT cells (Fig. 5d–f). We note that ether lipid deficiency had a modest effect on primary tumor growth kinetics (Fig. 5g–i). A decrease in metastatic burden was also observed upon knockout of AGPS or FAR1 in OVCAR8 cells, and upon loss of CD44 in pB3 cells (Supplementary Fig. 10a–c).
a Representative confocal images of extravasated tdTomato-labeled pB3 WT and AGPS KO cells from an in vitro microvascular network (see methods), over 24 h. Data representative of two independent replicates (scale bar = 100 µm). b Quantification of extravasated pB3 WT and AGPS KO cells from microvascular network. Data represent the mean percentage of extravasated cells per device +/− SEM (n = 5 technical replicates). Significance was calculated using unpaired, two-tailed t-test. Data representative of two independent experiments. c Quantification of extravasated tdTomato-labeled PyMT-1099 cell line derivatives from microvascular network. Data represent the mean percentage of extravasated cells per device +/− SEM (n = 6 technical replicates). Significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test; ns, not significant. Data representative of two independent experiments. d Representative in vivo imaging system (IVIS) images of overall metastatic burden in C57BL/6 female mice following intracardiac injection of pB3 WT (n = 5) and AGPS KO (n = 5) cells. e Quantification of metastatic burden in C57BL/6 female mice following intracardiac injection of pB3 WT and AGPS KO cells (n = 5). Graph shows the mean +/− SEM and statistical significance was calculated using two-tailed Mann-Whitney test. f Representative images of H&E-stained sections of harvested kidneys from C57BL/6 female mice following intracardiac injection of pB3 WT or pB3 AGPS KO cells (scale bar = 1,000 µm). g Gross images of primary tumors derived from pB3 WT control cells and pB3 AGPS KO cells. h Tumor growth kinetics of primary tumors derived from pB3 WT control cells and pB3 AGPS KO cells (n = 5 mice per group). Data shown as mean +/− SEM. i Bar graph showing the average weight of primary tumors derived from pB3 WT and AGPS KO cells (n = 5 mice). Graph shows the mean +/− SEM and statistical significance was calculated using two-tailed Mann-Whitney test; ns, not significant. j Estimated number of cancer stem cells (CSCs) per 10,000 cells as calculated by extreme limiting dilution analysis (ELDA) software. Tumor-initiating capacity was assessed following implantation of pB3 WT or pB3 AGPS KO cells into the mammary fat pad of C57BL/6 mice. P values, two-sided χ2 pairwise test. k Table showing the number of mice with palpable primary tumors at 121 d post orthotopic implantation of PyMT-1099 WT or AGPS KO cells +/− pretreatment with TGF-β into female NSG mice. l Quantification of lung metastases for aforementioned experiment. Data represents mean number of lung metastases +/− SEM (n = 8 for WT and AGPS KO + TGF-β, n = 7 for WT + TGF-β). Statistical significance was calculated by one-way ANOVA with Tukey’s multiple comparisons test; ns, not significant. m Representative images of H&E-stained lungs harvested from female NSG mice from aforementioned experiment. Arrows indicate metastases. Scale bar = 2000 µm.
Given that high CD44 expression and elevated intracellular iron levels are positively correlated with cancer cell stemness35,44,96, we investigated whether ether lipid deficiency could modulate CSC properties, by employing mammosphere formation assays97 and experimental limiting dilution tumor-implantation studies. These experiments indicated that loss of ether lipids in pB3 cells decreases cancer cell stemness (Fig. 5j, Supplementary Fig. 10d, e), which, as we have found in other investigations, serve as a reliable marker of metastasis-initiating capacity98.
Additionally, we show that treatment of human MCF7 breast adenocarcinoma cells with oncostatin M (OSM) increases the sub-population of CD44hi/CD24lo cells (Supplementary Fig. 11a, b), resulting in elevated PUFA-containing ether phospholipids and increased intracellular iron (Supplementary Fig. 11c–g). This is consistent with our TGFβ-treated PyMT-1099 model (Supplementary Fig. 12a–c).
Moreover, we found that loss of ether lipids significantly attenuates the tumor-initiating potential and metastatic capacity of PyMT-1099 AGPS KO TGF-β-treated cells following orthotopic implantation into the mammary stromal fat pad compared to PyMT-1099 WT TGF-β-treated cells (Fig. 5k–m). Collectively, our findings indicate that ether lipids play critical roles in promoting cancer cell stemness and resulting post-extravasation colonization.
Discussion
Our findings underscore the importance of integrating membrane biophysical properties into existing genetic and biochemical frameworks for understanding cancer cell fate. While phospholipids have long been implicated in cellular transformation, the focus thus far has primarily been to study the actions of specific lipids (e.g. inositol phospholipids) and their signaling derivatives99. Our study reveals a distinct and underappreciated mechanism by which lipids influence malignancy. Specifically, we identify ether lipids as key modulators of plasma membrane biophysical properties that regulate iron uptake and neoplasia-related phenotypes such as metastasis and cancer cell stemness/tumor-initiating potential. Importantly, this biophysical configuration creates a unique vulnerability of cancer cells to ferroptosis and suggests that targeting lipid metabolism and iron homeostasis could be exploited to suppress subpopulations of highly metastatic and drug-tolerant carcinoma cells7,100.
Ether phospholipids have been widely characterized as participants in ferroptosis through their role as substrates prone to iron-catalyzed oxidation22. However, our findings indicate an entirely distinct biochemical mechanism is operative here, whereby ether lipids directly modulate the levels of intracellular iron, a rate-limiting parameter governing ferroptosis susceptibility4,35,86. By emphasizing the role of membrane biophysical properties in mediating iron uptake, we depart from the conventional portrayal of phospholipids simply as substrates of oxidation. This shift in perspective has the potential to open new avenues for research, as it challenges researchers to explore the biophysical aspects of membranes as a new dimension in the regulation of this cell death program.
Although most lipid classes remained largely unchanged upon AGPS KO, it is plausible that subtle changes in membrane lipid composition beyond ether lipid levels may also be responsible for the phenotypes described here. Moreover, in addition to perturbed membrane receptor dynamics promoted by changes in membrane tension, the observed decrease in metastatic capacity of AGPS KO cells may also result from changes in oncogenic signaling lipids24, mechanisms that are not addressed here.
Alterations in intracellular iron level can impact gene expression via various mechanisms, including modulation of chromatin-modifying enzyme activity35,101,102. For example, an increase of intracellular iron levels has been shown to promote the activity of iron-dependent histone demethylases35,101,102, impacting gene expression profiles underlying cell plasticity35 and immune cell activation39. Our finding that ether lipid deficiency reduces intracellular iron levels explains, at least in part, how loss of ether lipids may affect cancer-associated transcriptional programs, acting at the epigenetic level and enabling a variety of malignancy-associated cell phenotypes including metastasis and cancer stemness. Such mechanisms may act in concert with processes independent from iron, which are also regulated by ether lipids to affect cancer malignancy traits.
In the longer term, the implications of our findings may reach well beyond cancer pathogenesis. Thus, we suggest that the interplay between membrane biophysics and iron biology could represent a fundamental determinant of cell fate, influencing diverse processes such as differentiation, immune activation, wound healing, and embryonic development.
Methods
Cell lines
The pB2 and pB3 MMTV-PyMT-derived murine breast cancer cell lines were a kind gifts from the laboratory of Harold L. Moses32. 687 g cells (also called EpCAMLoSnail-YFPHi) were originally established from tumors that developed in the MMTV-PyMT-Snail-IRES-YFP reporter mouse model, previously developed by the Weinberg lab103. pB3 and 687 g cell lines were cultured in 1:1 DMEM/F12 medium containing 5% adult bovine serum with 1% penicillin-streptomycin and 1% non-essential amino acids32. The PyMT-1099 murine breast cancer cell line was a kind gift from the laboratory of Gerhard Christofori and cultured in DMEM supplemented with 10% fetal bovine serum, 1% penicillin-streptomycin and 1% glutamine29. For select experiments, these cells were treated with 2 ng/mL of TGF-β for 10 days prior to performing subsequent analyses. OVCAR8 cells were obtained from the laboratory of Joan Brugge and cultured in 1:1 MCDB 105 medium/Medium 199 Earle’s Eagles medium supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin. MCF7 (ATCC, HTB-22) cells were cultured in Dulbecco’s Modified Eagle Medium GlutaMAX (DMEM, ThermoFisher Scientific, 61965059) supplemented with 10% Fetal Bovine Serum and Penicillin-Streptomycin mixture (BioWhittaker/Lonza, DE17-602E) and treated with oncostatin M (OSM, R&D systems, 295-OM-050, 100 ng/mL, 72 h). All cells were cultured in a humidified incubator at 37 °C with 5% CO2. All cells were negative for mycoplasma. Human cell line authentication (CLA) analysis of OVCAR8 cells were performed by the Duke University DNA Analysis facility. Established murine lines have not been STR profiled.
Animal studies
All animal experiments were performed in the animal facility at the Koch Institute for Integrative Cancer Research at MIT. Mice we housed under a 12-h light/12-h dark cycle, temperatures in the range 20–22 °C and humidity between 30–70%. Animal experiments performed in this study received approval by the MIT Institutional Animal Care and Use Committee. Tumor burden, where measurable, was not allowed to exceed 3 cm. Animals showing discomfort or distress were humanely euthanized following the MIT Institutional Animal Care and Use Committee guidelines. For primary tumor growth studies: 1 million cells were resuspended in 20% Matrigel/PBS and injected into the mammary fat pad of 6-8 week old female mice. C57BL/6 mice (Jackson Laboratories; strain name: C57BL/6 J; stock number: 000664) were used for in vivo experiments with pB3 cells. NSG mice (Jackson Laboratories; strain name: NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ; stock number: 005557) were used for in vivo experiments with PyMT-1099 cells. These cells were pretreated with TGF-β (2 ng/mL) for 10 days prior to injection. Tumor size was measured once a week using a vernier caliper and tumor volume was calculated using the formula: Tumor volume = (length x width2)/2, where length represents the largest tumor diameter, and width represents the perpendicular tumor diameter. For limiting dilution tumor-initiating assays, pB3 WT or pB3 AGPS KO cells were resuspended in 20% Matrigel/PBS and injected into the mammary fat pad of 6-8 weeks old female C57BL/6 mice at the following dilutions: 100,000, 10,000, 1000, 100 cells. Animals were assessed for palpable tumors after 39 days post injection. The estimated number of CSCs was calculated using the extreme limiting dilution analysis (ELDA) software104. For experimental metastasis involving pB3 cell lines, 0.2 million GFP-luciferized cells were resuspended in 200 µl of PBS and injected into the left ventricle of 6-8 weeks old female C57BL/6 mice. Metastatic burden was measured after 10 d post-injection via bioluminescence in live animals using the spectrum in vivo imaging system (IVIS)105. Images were analyzed using Living Image software (PerkinElmer, version 4.8.2). For OVCAR8 cells, 1.5 million cells were resuspended in PBS and implanted into 6-8 weeks old female athymic nude mice (Jackson Laboratories; strain name: NU/J; stock number: 002019) via intraperitoneal injections. Metastatic burden was assessed after 6 weeks using a fluorescence dissecting microscope.
Generation of gene-edited cell lines using CRISPR/Cas9
Except for OVCAR8 cells, all AGPS KO single-cell clones were generated via transient transfection with mouse AGPS CRISPR/Cas9 KO Plasmids (Catalog no. sc-432759, Santa Cruz) according to the manufacturer’s instructions. GFP-positive cells were sorted via fluorescence-activated cell sorting (FACS) into 96-well plates with one cell per well and single-cell clones were subsequently expanded. AGPS KO single-cell clones were assessed for loss of AGPS expression via western blot analysis. pB3 and PyMT-1099 AGPS addback cells were generated by transducing an AGPS KO single-cell clone with pLV[Exp]-Puro-EF1A-mAgps lentiviral vector (VectorBuilder). pB3 and PyMT-1099 AGPS KO cells expressing pLV[Exp]-Puro-EF1A-Stuffer_300bp (VectorBuilder) were established as controls for AGPS addback experiments and noted in the figure legends where used. Lentivirus was produced by transfecting HEK293T cells with viral envelope (VSVG, Addgene) and packaging plasmids (psPAX2, Addgene). Viral supernatant was collected after 48 h and filtered through a 0.45 µm filter. Stably transduced cells were selected with 2 µg/mL puromycin. OVCAR8 FAR1 KO and AGPS KO single cell clones as well as nontargeting control cells were established as previously described22. pB3 CD44 KO cells (bulk) were generated using human CD44 CRISPR/Cas9 KO Plasmids (Catalog no. sc-419558, Santa Cruz) according to the manufacturer’s instructions. After 48 h post-transfection, cells were sorted by flow cytometry for GFP positive cells, expanded in culture, and re-sorted twice for CD44 negative cells using Alexa Fluor® 647 anti-mouse/human CD44 Antibody. Cells were maintained as bulk CD44 KO cells. A list of all plasmids used to generate gene edited cell lines can be found in Supplementary Table 1.
Lipidomics analysis
Sample preparation (whole cell preparations)
Briefly, 1 × 106 cells were plated in individual wells of a 6 well dish. No more than 16 hours later, cells were washed with DPBS (Dulbecco’s PBS without Mg and Ca), trypsinized, and washed twice again with before resuspending in 300 μL of DPBS and storing at −80 °C until further processing.
Sample preparation (GPMV preparations)
Giant plasma membrane vesicles (GPMVs) were isolated from pB3 WT, AGPS KO, and AGPS addback cells as previously described with minor modifications106. Briefly, three 10-cm dishes of approximately 70% confluency prepared for each cell line were washed once with phosphate-buffered saline (PBS) and twice with GPMV buffer (10 mM HEPES, 150 mM NaCl, 2 mm CaCl2 – pH 7.4). Fresh N-ethylmaleimide (Millipore Cat. No. E3876-5G) was added to GPMV buffer to yield a final concentration of 5 mM, then 4 mL of this buffer was added to each culture dish followed by incubation for 3 hours at 37 °C. Supernatants containing GPMVs were then collected and pooled for each condition and passed through 5 µm centrifugal filters (Millipore Sigma Cat. No. UFC30SV00) by centrifuging at 500 x g to remove any whole cells. The pooled filtrates were subsequently centrifuged at 20,000 × g for 2 hours at 4 °C and resuspended in 300 µL of DPBS. The presence of vesicles was confirmed by light microscopy before storing at −80 °C until further processing. Three independent biological replicates were collected and used for lipidomic analysis.
Lipid extraction for mass spectrometry lipidomics
Mass spectrometry-based lipid analysis was performed by Lipotype GmbH (Dresden, Germany) as described107. Lipids from whole cells or isolated GPMVs were extracted using a chloroform/methanol procedure108. Samples were spiked with internal lipid standard mixture containing: cardiolipin 14:0/14:0/14:0/14:0 (CL), ceramide 18:1;2/17:0 (Cer), diacylglycerol 17:0/17:0 (DAG), hexosylceramide 18:1;2/12:0 (HexCer), lyso-phosphatidate 17:0 (LPA), lyso-phosphatidylcholine 12:0 (LPC), lyso-phosphatidylethanolamine 17:1 (LPE), lyso-phosphatidylglycerol 17:1 (LPG), lyso-phosphatidylinositol 17:1 (LPI), lyso-phosphatidylserine 17:1 (LPS), phosphatidate 17:0/17:0 (PA), phosphatidylcholine 17:0/17:0 (PC), phosphatidylethanolamine 17:0/17:0 (PE), phosphatidylglycerol 17:0/17:0 (PG), phosphatidylinositol 16:0/16:0 (PI), phosphatidylserine 17:0/17:0 (PS), cholesterol ester 20:0 (CE), sphingomyelin 18:1;2/12:0;0 (SM), triacylglycerol 17:0/17:0/17:0 (TAG). After extraction, the organic phase was transferred to an infusion plate and dried in a speed vacuum concentrator. The dry extract was re-suspended in 7.5 mM ammonium formate in chloroform/methanol/propanol (1:2:4, V:V:V). All liquid handling steps were performed using Hamilton Robotics STARlet robotic platform with the Anti Droplet Control feature for organic solvents pipetting.
MS data acquisition
Samples were analyzed by direct infusion on a QExactive mass spectrometer (Thermo Scientific) equipped with a TriVersa NanoMate ion source (Advion Biosciences). Samples were analyzed in both positive and negative ion modes with a resolution of Rm/z=200 = 280000 for MS and Rm/z=200 = 17500 for MSMS experiments, in a single acquisition. MSMS was triggered by an inclusion list encompassing corresponding MS mass ranges scanned in 1 Da increments109. Both MS and MSMS data were combined to monitor CE, DAG and TAG ions as ammonium adducts; LPC, LPC O-, PC, PC O-, as formate adducts; and CL, LPS, PA, PE, PE O-, PG, PI and PS as deprotonated anions. MS only was used to monitor LPA, LPE, LPE O-, LPG and LPI as deprotonated anions; Cer, HexCer and SM as formate adducts.
Data analysis and post-processing
Data were analyzed with in-house developed lipid identification software based on LipidXplorer110,111. Only lipid identifications with a signal-to-noise ratio >5, and a signal intensity 5-fold higher than in corresponding blank samples were considered for further data analysis. Cholesterol was removed from each data set before analysis. Simple imputation was performed by replacing missing values with 0.2 * the minimum non-zero value for each lipid assayed, a modification from previously described methods112. Relative mol% levels of individual lipid species were determined by dividing each pmol value by the sum pmol of all lipids detected in that sample and multiplying by 100. Total ether lipid (and other lipid classes presented) were calculated by summation of mol% values of each lipid within that class. Lipid fold changes were calculated by dividing the average mol% of each lipid species between groups and applying a Log2 transformation. Corresponding p-values were calculated using the student’s t-test.
Oxidized lipidomics
Sample preparation
100,000 cells per condition were plated in 6-well plates 24 h prior to the experiment. For PyMT-1099, cells were treated with TGF-β (2 ng/mL) for 10 days. pB3 cells were treated with 500 nM RSL3, OVCAR8 cells with 2 µM ML210 and TGF-β-treated PyMT-1099 cells with 10 µM ML210 for 24 h. Cells were subsequently washed with PBS and then with 150 mM ammonium bicarbonate. Cells were then scraped and resuspended in 150 mM ammonium bicarbonate and centrifuged at 300 x g for 5 min. The supernatant was removed, and cells were resuspended in 1 mL of 150 mM ammonium bicarbonate. The solutions were centrifuged at 13,500 x g for 10 min and the supernatant was removed. 200 µL of 150 mM sodium bicarbonate was added to the pellet and samples were flash frozen in liquid nitrogen. Cells were prepared in 5 independent biological replicates and lipidomics analysis was performed on the same day for all the replicates.
Lipid extraction for mass spectrometry lipidomics
For lipidomics analysis, the 200 µL cell lysates were spiked with 1.4 μL of internal standard lipid mixture containing 300 pmol of phosphatidylcholine 17:0-17:0, 50 pmol of phosphatidylethanolamine 17:0-17:0, 30 pmol of phosphatidylinositol 16:0-16:0, 50 pmol of phosphatidylserine 17:0-17:0, 30 pmol of phosphatidylglycerol 17:0-17:0 and 30 pmol of phosphatidic acid 17:0-17:0 and subjected to lipid extraction at 4 °C, as previously described113. The sample was then extracted with 1 mL of chloroform-methanol (10:1) for 2 h. The lower organic phase was collected, and the aqueous phase was re-extracted with 1 mL of chloroform-methanol (2:1) for 1 h. The lower organic phase was collected and evaporated in a SpeedVac vacuum concentrator. Lipid extracts were dissolved in 100 μL of infusion mixture consisting of 7.5 mM ammonium acetate dissolved in propanol:chloroform:methanol [4:1:2 (vol/vol)].
MS data acquisition
Samples were analyzed by direct infusion in a QExactive mass spectrometer (Thermo Fisher Scientific) equipped with a TriVersa NanoMate ion source (Advion Biosciences). 5 µL of sample were infused with gas pressure and voltage set to 1.25 psi and 0.95 kV, respectively. PC, PE, PEO, PCOx and PEOx were detected in the 10:1 extract, by positive ion mode FTMS as protonated adducts by scanning m/z = 580–1000 Da, at Rm/z=200 = 280 000 with lock mass activated at a common background (m/z = 680.48022) for 30 s. Every scan is the average of 2 micro-scans, automatic gain control (AGC) was set to 1E6 and maximum ion injection time (IT) was set to 50 ms. PG and PGOx were detected as deprotonated adducts in the 10:1 extract, by negative ion mode FTMS by scanning m/z = 420–1050 Da, at Rm/z=200 = 280 000 with lock mass activated at a common background (m/z = 529.46262) for 30 s. Every scan is the average of 2 micro-scans. Automatic gain control (AGC) was set to 1E6 and maximum ion injection time (IT) was set to 50 ms. PA, PAOx, PI, PIOx, PS and PSOx were detected in the 2:1 extract, by negative ion mode FTMS as deprotonated ions by scanning m/z = 400–1100 Da, at Rm/z=200 = 280 000 with lock mass activated at a common background (m/z = 529.46262) for 30 s. Every scan is the average of 2 micro-scans, automatic gain control (AGC) was set to 1E6 and maximum ion IT was set to 50 ms. All data was acquired in centroid mode. All lipidomics data were analyzed with the lipid identification software, LipidXplorer111. Tolerance for MS and identification was set to 2 ppm. Data were normalized to internal standards.
Immunoblotting
Cells were washed with ice-cold PBS and lysed in cell lysis buffer (Cell Signaling Technology, Cat. #9803S) containing 1 mM PMSF protease inhibitor (Cell Signaling Technology, Cat. #8553S). Protein samples were prepared with NuPAGE LDS Sample Buffer (Thermo Fischer Scientific, Cat. #NP0007) and NuPage Sample Reducing Agent (Thermo Fischer Scientific, Cat. #NP0004) or 2X Laemmli buffer and heated at 70 °C or 94 °C for 10 minutes. Samples were resolved by SDS-PAGE, transferred to nitrocellulose (Bio-Rad) and blocked in 5% milk/TBST or 5% milk/PBST for 1 h at room temperature. Membranes were incubated overnight with the respective primary antibodies at 4 °C in 5% milk/PBST or in 5% BSA/TBST, washed with TBST or PBST, incubated with HRP-conjugated secondary antibodies and developed using SuperSignalTM West Dura, SuperSignalTM West Pico PLUS, or SuperSignalTM West Femto kits (ThermoFisher Scientific, Cat. 34076, Cat. 34580 and Cat. 34096). A list of all antibodies used in this study can be found in Supplementary Table 2.
Cell viability assay
Cells were seeded in 96-well black clear bottom plates (Corning) at 2000 or 3000 cells (PyMT-1099 + /- TGF-β) and 6000 cells (OVCAR8) per well. Approximately, 12-16 h post-seeding, cells were treated with various drug concentrations using an HP D300e Digital Dispenser unless stated otherwise. Cell viability was assessed at 72 h post-treatment by performing CellTiter-Glo Luminescent Cell Viability Assay (Promega) according to the manufacturer’s instructions. Relative viability was calculated by normalizing to untreated controls unless stated otherwise. Non-linear regression models (four parameter, variable slope) were applied to generate the regression fit curves using GraphPad Prism version 10.4.1. Drug compounds were purchased as indicated: RSL3 (Selleck Chem), ML210 (Sigma Aldrich), and Liproxstatin-1 (Fisher Scientific). For experiments involving ferric ammonium citrate (FAC), FAC (Sigma) was prepared fresh in sterile 1× PBS and manually added directly to cell culture media at the indicated concentrations at the time of seeding into 96-well plates. Unless stated otherwise, cells were pretreated with FAC for 24 h prior to ML210 treatment.
Inductively coupled plasma mass spectrometry (ICP-MS)
For select experiments, cells were treated for 24 h with hyaluronic acid (Carbosynth, FH45321, 600-1000 kDa, 1 mg/mL) or hyaluronidase (HD, Sigma-Aldrich, H3884, 0.1 mg/mL) as indicated. Glass vials equipped with Teflon septa were cleaned with nitric acid 65% (VWR, Suprapur, 1.00441.0250), washed with ultrapure water (Sigma-Aldrich, 1012620500) and dried. Cells were harvested and washed twice with 1× PBS. Cells were then counted using an automated cell counter (Entek) and transferred in 200 µL 1× PBS to the cleaned glass vials. The same volume of PBS was transferred into separate vials for the background subtraction, at least in duplicate per experiment. For tumor samples, small pieces of the tumors were added into pre-weighed cleaned glass vials. Samples were lyophilized using a freeze dryer (CHRIST, 22080). Glass vials with lyophilized tumor samples were weighed to determine the dry weight for normalization. Samples were subsequently mixed with nitric acid 65% and heated at 80 °C overnight. Samples were diluted with ultrapure water to a final concentration of 0.475 N nitric acid and transferred to metal-free centrifuge vials (VWR, 89049-172) for subsequent ICP-MS analyses. Amounts of metals were measured using an Agilent 7900 ICP-QMS in low-resolution mode, taking natural isotope distribution into account. Sample introduction was achieved with a micro-nebulizer (MicroMist, 0.2 mL/min) through a Scott spray chamber. Isotopes were measured using a collision-reaction interface with helium gas (5 mL/min) to remove polyatomic interferences. Scandium and indium internal standards were injected after inline mixing with the samples to control the absence of signal drift and matrix effects. A mix of certified standards was measured at concentrations spanning those of the samples to convert count measurements to concentrations in the solution. Values were normalized against cell number or dry weight.
Iron measurements using HRhoNox-M
The lysosome-specific fluorescent Fe(II) probe HRhoNox-M was synthesized in 3 steps according to a previously published procedure36. 1H NMR (300 MHz, CDCl3) δ 7.93 (1H, d, J = 2.0 Hz), 7.45 (1H, dd, J = 8.5 Hz, 2.0 Hz), 7.40–7.30 (3H, m), 7.05 (1H, d, J = 8.5 Hz), 6.90 (1H, d, J = 7.0 Hz), 6.80 (1H, d, J = 8.0 Hz), 6.50–6.44 (2H, m), 5.28–5.35 (2H, m), 3.62 (6H, m), 2.97 (6H, s). MS (ESI) m/z: calcd. for C24H25N2O3 [M + H]+ 389.19, found: 389.35. Cells were incubated with 1 µM HRhoNox-M for 1 h or lysotracker deep red (Thermo Fisher Scientific L12492) according to the manufacturer’s instructions for 1 h. Cells were then washed twice with ice-cold PBS and suspended in incubation buffer prior to being analyzed by flow cytometry. For each condition, at least 10,000 cells were counted. Data were recorded on a BD Accuri C6 (BD Biosciences) and processed using Cell Quest (BD Biosciences) and FlowJo (FLOWJO, LLC). The signal for HRhoNox-M was normalized against the signal of lysotracker of cells treated in parallel.
Endocytosis experiments
Antibody against EEA1 was purchased from BD Biosciences (Catalog no. 610456). Conjugated transferrin (mouse)-Alexa546, dextran-Alexa555, and EGF-Alexa555 were purchased from Invitrogen. Cy3-conjugated hyaluronate was synthesized in-house. Cy2- or Cy3-conjugated donkey antibodies against mouse IgG were purchased from Jackson ImmunoResearch. For receptor-mediated endocytosis, cells were washed with serum-free medium and then incubated in this medium with Cy3-conjugated hyaluronate (0.1 mg/mL), Alexa 555-conjugated EGF (either 2 ng/mL or 200 ng/mL), or Alexa 546-conjugated transferrin (5 μg/mL) for 1 h at 4 °C. Cells were then washed to clear unbound ligand, and shifted to 37 °C for times indicated in the figures. Cells were stained for EEA1, followed by confocal microscopy to assess the arrival of ligand to the early endosome. To assess fluid-phase uptake, Alexa 555-conjugated dextran (0.2 mg/mL) was added to complete medium and cells were incubated 37 °C for times indicated in the figures. Cells were then stained for EEA1, followed by confocal microscopy to assess the arrival of this probe to the early endosome. In select assays cells were pretreated with 20 μM C(22:6) BSA conjugate for 16-18 h.
Confocal microscopy
Colocalization studies were performed with the Zeiss Axio Observer Z1 Inverted Microscope having a Plan-Apochromat 63x objective, the Zeiss LSM 800 with Airyscan confocal package with Zeiss URGB (488- and 561-nm) laser lines, and Zen 2.3 blue edition confocal acquisition software. For quantification of colocalization, ten fields of cells were examined, with each field typically containing about 5 cells. Images were imported into the NIH ImageJ v.1.50e software, and then analyzed through a plugin software (https://imagej.net/Coloc_2). Under the ‘image’ tab, the ‘split channels’ option was selected. Under the ‘plugins’ tab, ‘colocalization analysis’ option was selected, and within this option, the ‘colocalization threshold’ option was selected. Manders Coefficient was used for colocalization analysis. Colocalization values were calculated by the software and expressed as the fraction of protein of interest colocalized with EEA1. Colocalization experiments were performed at least twice, and representative data is shown.
Mammosphere assay
Mammospheres were generated in accordance to a previously described protocol with minor modifications114. Briefly, single cell suspensions containing 1,200 WT or AGPS KO pB3 cells were plated in serum free DMEM/F12 supplemented with 2nM L-glutamine, 20 ng/mL recombinant human epidermal growth factor (EGF; Peprotech Cat. No. AF-100-15), and B27 supplement (Gibco, Cat. No. 17504-044) in individual wells of an ultra-low attachment 6 well plate (Corning Cat. No. 3471). After 4 days, the total number of spheres greater than 40 µm were quantified before dissociation in trypsin at 37 °C. Dissociated cells were then passed through a 40 µm cell strainer to yield a single cell suspension, before replating 1,200 cells. Sphere formation efficiency ((# of mammospheres per well) / (# of cells seeded per well) x 100) was calculated over 3 passages. Data shown is representative of 3 independent biological replicates.
Synthesis of HA-Cy3 probe
Hyaluronic acid (HA, 2 mg, Sigma 75044, Lot #BCBM2884) was dissolved in a 1:1 solution of dimethylsulfoxide (DMSO) and water (0.4 mL) for a stock concentration of 5 mg/mL. The polymer was sonicated under heating to ensure full solubilization. The HA solution was then diluted into HEPES (50 mM final HEPES concentration for a total reaction volume of 2 mL once all components are combined). Sulfo-Cyanine3 amine (2.36 mg, Lumiprobe) was separately dissolved in DMSO (0.236 mL) for a stock concentration of 10 mg/mL. N-(3-Dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride (EDC, 0.253 mg, Sigma) was separately dissolved in 50 mM HEPES (0.051 mL) for a stock concentration of 5 mg/mL. The HA and EDC solutions were then combined under stirring, followed by addition of the dye solution. The reaction was stirred, protected from light, at room temperature for 12 h. Following, unreacted dye was removed via Amicon Ultra-0.5 Centrifugal Filters (Millipore Sigma). Manufacturer guidelines were followed to select purification spin speeds and times: 14000 xg, 15 min per wash step (water) until washes were clear and colorless. The purified HA-Cy3 probe was stored in water at 4 °C until used.
Preparation of liposomes
Ether lipid liposomes were prepared as previously described22. PE (18:1p_20:4) (Catalog no. 852804), PE (18:1p_18:1) (Catalog no. 852758), PC (18:1p_20:4) (Catalog no. 852469) and PE (18:0_20:4) (Catalog no. 850804) were purchased from Avanti Polar Lipids Inc. A Malvern ZS90 Particle Analyzer was used to analyze the diameter and polydispersity index of liposomes used in this study and these are reported in Supplementary Table 3.
Liposome reconstitution assays
Adherent cells were treated with liposomes 16-18 h prior to performing respective membrane tension or endocytosis assays. Lipid liposomes were added directly to the culture medium for a final concentration of 20 µM. Cells were switched from liposome-containing media to extracellular imaging buffer (HEPES buffer with dextrose, NaCl, KCl, MgCl2, CaCl2) during membrane tension experiments.
Preparation of BSA conjugated DHA C(22:6)
In brief, BSA (FA-free; Fisher Scientific BP9704100) was dissolved in ddH2O to give a 1 mM solution. Exactly 25 mg of docosahexaenoic acid (DHA, C22:6; Sigma Aldrich D2534) was added to 38.05 mL of 1 mM BSA solution to yield a 2:1 DHA:BSA ratio in a glass vial. The vial was then purged with nitrogen and stirred at room temperature until dissolved. The conjugate was then filter sterilized and concentration verified via BCA assay. Aliquots were then stored at −80 °C until use.
Membrane tension
Tether pulling experiments were performed on a home-built optical trap, following principles described elsewhere74,115. Briefly, 4 µm anti-Digoxigenin-coated polystyrene beads (Spherotech) were trapped with a 1064 nm, Ytterbium laser (IPG Photonics) focused through a 60 × 1.2 NA objective (Olympus). Forces on the beads were measured by the deflection of backscattered trapping laser light onto a lateral effect position sensor (Thorlabs) and calibrated using the viscous drag method116. To measure tether radii (R), cell lines were transiently transfected with a membrane-targeted fluorescent protein (glycosylphosphatidylinositol-anchored eGFP, Addgene #32601) using a TransIT-X2 transfection kit (Mirus). Tether radius was obtained by comparing tether fluorescence to fluorescence counts from a known area of the parent cell membrane, as described59. Tether force \((f)\) and fluorescence measurements were performed simultaneously. Membrane tension was calculated using the following equation:
Miscibility transition temperatures (Tmisc) measurements
Miscibility transition temperatures (Tmisc) measurements were performed as previously reported106,117. Briefly, cells were washed in PBS, and cell membranes were labeled with 5 µg/mL fluorescent disordered/nonraft phase marker FAST DiO (Thermo Fisher Scientific) for 10 min on ice. Cells were then washed twice in GPMV buffer (10 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH 7.4), and then incubated with GPMV buffer supplemented with 25 mM paraformaldehyde (PFA) and 2 mM dithiothreitol (DTT) for 1 h at 37 °C. Vesicles were imaged at 40× on an inverted epifluorescence microscope (Leica DMi8) under temperature-controlled conditions using a microscope stage equipped with a Peltier element (Warner Instruments). GPMVs were imaged from 4 °C-28 °C, counting phase-separated and uniform vesicles at each temperature. For each temperature, 25-50 vesicles were counted, and the percent of phase-separated vesicles were calculated, plotted versus temperature, and a fitted to a sigmoidal curve to determine the temperature at which 50% of the vesicles were phase-separated (Tmisc).
C-laurdan spectral imaging
C-Laurdan imaging was performed as previously described80,82,106,117,118. Briefly, cells were washed with PBS and stained with 10 µg/mL C-Laurdan for 10 min on ice, then imaged using confocal microscopy on a Leica SP8 with spectral imaging at 60× (water immersion, NA = X) and excitation at 405 nm. The emission was collected as two images: 420–460 nm and 470–510 nm. MATLAB (MathWorks, Natick, MA) was used to calculate the two-dimensional (2D) GP map, where GP for each pixel was calculated as previously described118. Briefly, each image was background subtracted and thresholded to keep only pixels with intensities greater than 3 standard deviations of the background value in both channels. The GP image was calculated for each pixel using Equation 2, where G is the G-factor. The G-factor was determined before each experiment using the protocol in ref. 119. GP maps (pixels represented by GP value rather than intensity) were imported into ImageJ. To calculate the average PM GP, line scans drawn across individual cells. PM GP values were taken as peak GP values from the periphery of the cell, whereas internal membranes were calculated as the average of all values outside the PM peak. The average GP of the internal membranes was calculated by determining the average GP of all pixels in a mask drawn on each cell just inside of the PM.
Extravasation assay
Cells and reagents
Immortalized human umbilical vein endothelial cells (HUVECs) expressing BFP95 were cultured in VascuLife VEGF Endothelial Medium (Lifeline Cell Technology). Normal human lung fibroblasts (FBs) (Lonza, P7) were cultured in FibroLife S2 Fibroblast Medium (Lifeline Cell Technology).
Microfluidic device
3D cell culture chips (AIM Biotech) were used to generate in vitro microvascular networks (MVNs). The AIM chip body was made of cyclic olefin polymer (COP) with a type of gas-permeable plastic serving as the bottom film. AIM Biotech chips contained three parallel channels: a central gel channel flanked by two media channels. Microposts separated fluidic channels and serve to confine the liquid gelling solution in the central channel by surface tension before polymerization. The gel channel was 1.3 mm wide and 0.25 mm tall, the gap between microposts was 0.1 mm, and the width of media channels was 0.5 mm.
Microvascular network formation
To generate perfusable MVNs, HUVECs and FBs were seeded into the microfluidic chip using a two-step method120. Briefly, HUVECs and FBs were concentrated in VascuLife containing thrombin (4 U/mL). For the first step seeding, the outer layer EC solution was made with a final concentration of 10 × 106/mL. After mixed with fibrinogen (3 mg/mL final concentration) at a 1:1 ratio, the outer layer EC solution was pipetted into the gel inlet, immediately followed by aspirating from the gel outlet, leaving only residual solution around the microposts. For the second step, another solution with final concentrations of 5 × 106/mL HUVECs and 1.5 × 106/mL fibroblasts were similarly mixed with fibrinogen and then pipetted into the same chip through the gel outlet. The device was placed upside down to polymerize in a humidified enclosure and allowed to polymerize at 37 °C for 15 min in a 5% CO2 incubator. Next, VascuLife culture medium was added to the media channels and changed daily in the device. After 7 days, MVNs were ready for further experiments.
Tumor cell perfusion in MVNs
PyMT-1099 or pB3 cell line derivatives expressing pCDH-EF1-Luc2-P2A-tdTomato (Plasmid #72486, Addgene) were resuspended at a concentration of 1 × 106/mL in culture medium. To perfuse these tumor cells into in vitro MVNs, the culture medium in one media channel was aspirated, followed by injection of a 20 µL tumor cell suspension in the MVNs and repeated twice. Microfluidic devices were then placed at 37 °C for 15 min in a 5% CO2 incubator. After that, the tumor cell medium was aspirated from the media channels to remove the unattached cells, and Vasculife was replenished. Devices were then placed back to the incubator. 24 h later, devices were fixed, washed, and imaged using an Olympus FLUOVIEW FV1200 confocal laser scanning microscope with a 10× objective and an additional 2× zoom-in function. Z-stack images were acquired with a 5 µm step size. All images shown are collapsed Z-stacks, displayed using range-adjusted Imaris software, unless otherwise specified. Extravasation percentage was calculated by dividing the cell number of extravasated tumor cells with the total number of tumor cells in the same imaging region of interest.
CLIC/GEEC and micropinocytosis inhibition
7-keto-cholesterol was obtained from Cayman Chemical (Cat. 16339). Cytochalasin D was obtained from Sigma Aldrich (C8273). pB3 cells were pre-treated for 30 minutes with either of the inhibitors before assessing endocytic rate.
RNA interference
In brief, cells were seeded in 6-well plates at the density of 2 × 105 cells/well and transfected 24 hr later with the specified siRNA using jetPRIME (Polyplus, 114-15) or PepMute™ (SignaGen, Cat. SL100566) according to the manufacturer’s protocol with 100 nM siRNA. The medium was replaced after 6 h. Analysis was performed 72 h after transfection. Suitable small interfering RNAs were designed by Dharmacon for specific down-regulation of PICK1 (5’-cuuagacuaugacaucgaa-3) and CtBP1 (5’-ugucucaucugcuugacagu-3).
Flow cytometry
MCF7 cells were washed with ice-cold PBS. For antibody staining, cells were incubated with Fc block (Human TruStain FcX, Biolegend, 422302, 1:20) for 15 min, then incubated with the following antibodies: CD24-BV605 (Biolegend, 311124) and CD44-AF647 (Novus Biologicals, NB500-481AF647) in ice cold PBS with 10% FBS and 2 mM EDTA for 20 min at 4 °C and then washed with PBS and resuspended in PBS with 10% FBS and 2 mM EDTA before analysis using a Attune flow cytometer. Data were analyzed with FlowJo software v. 10.10.0.
Histology
Harvested tissues were fixed by incubating with 10% neutral-buffered formalin (VWR Scientific) at 4 °C for 16–18 h. Fixed samples were then transferred to 70% ethanol and submitted to Hope Babette Tang Histology Facility at the Koch Institute at MIT for paraffin-embedding and H&E staining. Metastatic burden was quantified using QuPath software121 and Image J122.
Statistical analysis
For statistical analyses, Mann-Whitney U test, two-tailed t-test, and one-way ANOVA with Dunnett, Tukey, or Holm-Šídák multiple test corrections were performed using GraphPad Prism Version 10.4.1. Permutation ANOVA was performed in R (v4.4.2) using the coin package with 10,000 permutations123. Planned contrasts were tested with one-sided Welch’s t-tests (t.test, base R), and exploratory pairwise comparisons were adjusted for multiple testing using Holm’s method (p.adjust).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Processed data file from lipidomic analysis of ether-lipid-deficient cells has been deposited in Zenodo (https://doi.org/10.5281/zenodo.17808937). All other data is provided within source data. Correspondence and requests for materials should be addressed to: Raphaël Rodriguez, PhD (raphael.rodriguez@curie.fr), Robert A. Weinberg, PhD (weinberg@wi.mit.edu) and Whitney S. Henry, PhD (wshenry@mit.edu). Source data are provided with this paper.
References
Levental, I. & Lyman, E. Regulation of membrane protein structure and function by their lipid nano-environment. Nat. Rev. Mol. Cell Biol. 24, 107–122 (2023).
Viswanathan, V. S. et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 547, 453–457 (2017).
Hangauer, M. J. et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 551, 247–250 (2017).
Mai, T. T. et al. Salinomycin kills cancer stem cells by sequestering iron in lysosomes. Nat. Chem. 9, 1025–1033 (2017).
Stockwell, B. R. et al. Ferroptosis: A Regulated Cell Death Nexus Linking Metabolism, Redox Biology, and Disease. Cell 171, 273–285 (2017).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Caneque, T. et al. Activation of lysosomal iron triggers ferroptosis in cancer. Nature 642, 492–500 (2025).
Yang, W. S. & Stockwell, B. R. Ferroptosis: Death by Lipid Peroxidation. Trends Cell Biol. 26, 165–176 (2016).
Gupta, P. B. et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138, 645–659 (2009).
Antoszczak, M. et al. Iron-Sensitive Prodrugs That Trigger Active Ferroptosis in Drug-Tolerant Pancreatic Cancer Cells. J. Am. Chem. Soc. 144, 11536–11545 (2022).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Doll, S. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 575, 688–692 (2019).
Yang, W. S. & Stockwell, B. R. Synthetic lethal screening identifies compounds activating iron-dependent, nonapoptotic cell death in oncogenic-RAS-harboring cancer cells. Chem. Biol. 15, 234–245 (2008).
Dixon, S. J. et al. Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. Elife 3, e02523 (2014).
Wang, W. et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569, 270–274 (2019).
Liang, Y., Zhao, Y., Qi, Z., Li, X. & Zhao, Y. Ferroptosis: CD8(+)T cells’ blade to destroy tumor cells or poison for self-destruction. Cell Death Discov. 11, 128 (2025).
Li, Q. et al. Inhibition of neuronal ferroptosis protects hemorrhagic brain. JCI Insight 2, e90777 (2017).
Fang, X. et al. Ferroptosis as a target for protection against cardiomyopathy. Proc. Natl. Acad. Sci. USA 116, 2672–2680 (2019).
Yu, Y. et al. Hepatic transferrin plays a role in systemic iron homeostasis and liver ferroptosis. Blood 136, 726–739 (2020).
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014).
Zou, Y. et al. Plasticity of ether lipids promotes ferroptosis susceptibility and evasion. Nature (2020).
Dean, J. M. & Lodhi, I. J. Structural and functional roles of ether lipids. Protein Cell 9, 196–206 (2018).
Benjamin, D. I. et al. Ether lipid generating enzyme AGPS alters the balance of structural and signaling lipids to fuel cancer pathogenicity. Proc. Natl. Acad. Sci. USA 110, 14912–14917 (2013).
Snyder, F. & Wood, R. Alkyl and alk-1-enyl ethers of glycerol in lipids from normal and neoplastic human tissues. Cancer Res 29, 251–257 (1969).
Albert, D. H. & Anderson, C. E. Ether-linked glycerolipids in human brain tumors. Lipids 12, 188–192 (1977).
Snyder, F., Blank, M. L. & Morris, H. P. Occurrence and nature of O-alkyl and O-alk-I-enyl moieties of glycerol in lipids of Morris transplanted hepatomas and normal rat liver. Biochim Biophys. Acta 176, 502–510 (1969).
Cui, W., Liu, D., Gu, W. & Chu, B. Peroxisome-driven ether-linked phospholipids biosynthesis is essential for ferroptosis. Cell Death Differ. 28, 2536–2551 (2021).
Saxena, M., Kalathur, R. K. R., Neutzner, M. & Christofori, G. PyMT-1099, a versatile murine cell model for EMT in breast cancer. Sci. Rep. 8, 12123 (2018).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Kagan, V. E. et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat. Chem. Biol. 13, 81–90 (2017).
Ye, X. et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 525, 256–260 (2015).
Pantopoulos, K., Porwal, S. K., Tartakoff, A. & Devireddy, L. Mechanisms of mammalian iron homeostasis. Biochemistry 51, 5705–5724 (2012).
Muckenthaler, M. U., Rivella, S., Hentze, M. W. & Galy, B. A Red Carpet for Iron Metabolism. Cell 168, 344–361 (2017).
Muller, S. et al. CD44 regulates epigenetic plasticity by mediating iron endocytosis. Nat. Chem. 12, 929–938 (2020).
Niwa, M., Hirayama, T., Okuda, K. & Nagasawa, H. A new class of high-contrast Fe(II) selective fluorescent probes based on spirocyclized scaffolds for visualization of intracellular labile iron delivered by transferrin. Org. Biomol. Chem. 12, 6590–6597 (2014).
Houk, R. S. Mass spectrometry of inductively coupled plasmas. Anal. Chem. 58, 97A–105A (1986).
Figueroa, J. A., Stiner, C. A., Radzyukevich, T. L. & Heiny, J. A. Metal ion transport quantified by ICP-MS in intact cells. Sci. Rep. 6, 20551 (2016).
Solier, S. et al. A druggable copper-signalling pathway that drives inflammation. Nature 617, 386–394 (2023).
Haley, J. et al. Functional characterization of a panel of high-grade serous ovarian cancer cell lines as representative experimental models of the disease. Oncotarget 7, 32810–32820 (2016).
Hoepken, H. H., Korten, T., Robinson, S. R. & Dringen, R. Iron accumulation, iron-mediated toxicity and altered levels of ferritin and transferrin receptor in cultured astrocytes during incubation with ferric ammonium citrate. J. Neurochem 88, 1194–1202 (2004).
Bauckman, K. A., Haller, E., Flores, I. & Nanjundan, M. Iron modulates cell survival in a Ras- and MAPK-dependent manner in ovarian cells. Cell Death Dis. 4, e592 (2013).
Gammella, E., Buratti, P., Cairo, G. & Recalcati, S. The transferrin receptor: the cellular iron gate. Metallomics 9, 1367–1375 (2017).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl. Acad. Sci. USA 100, 3983–3988 (2003).
Yan, Y., Zuo, X. & Wei, D. Concise Review: Emerging Role of CD44 in Cancer Stem Cells: A Promising Biomarker and Therapeutic Target. Stem Cells Transl. Med 4, 1033–1043 (2015).
Aruffo, A., Stamenkovic, I., Melnick, M., Underhill, C. B. & Seed, B. CD44 is the principal cell surface receptor for hyaluronate. Cell 61, 1303–1313 (1990).
Hua, Q., Knudson, C. B. & Knudson, W. Internalization of hyaluronan by chondrocytes occurs via receptor-mediated endocytosis. J. Cell Sci. 106, 365–375 (1993).
Bendris, N. et al. SNX9 promotes metastasis by enhancing cancer cell invasion via differential regulation of RhoGTPases. Mol. Biol. Cell 27, 1409–1419 (2016).
Thottacherry, J. J., Sathe, M., Prabhakara, C. & Mayor, S. Spoiled for Choice: Diverse Endocytic Pathways Function at the Cell Surface. Annu Rev. Cell Dev. Biol. 35, 55–84 (2019).
Saric, A. & Freeman, S. A. Endomembrane Tension and Trafficking. Front Cell Dev. Biol. 8, 611326 (2020).
McMahon, H. T. & Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 12, 517–533 (2011).
Sathe, M. et al. Small GTPases and BAR domain proteins regulate branched actin polymerisation for clathrin and dynamin-independent endocytosis. Nat. Commun. 9, 1835 (2018).
Rentero, C. et al. Functional implications of plasma membrane condensation for T cell activation. PLoS One 3, e2262 (2008).
Chaudhary, N. et al. Endocytic crosstalk: cavins, caveolins, and caveolae regulate clathrin-independent endocytosis. PLoS Biol. 12, e1001832 (2014).
Kalin, S. et al. Macropinocytotic uptake and infection of human epithelial cells with species B2 adenovirus type 35. J. Virol. 84, 5336–5350 (2010).
Gilleron, J. et al. Image-based analysis of lipid nanoparticle-mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat. Biotechnol. 31, 638–646 (2013).
Liberali, P. et al. The closure of Pak1-dependent macropinosomes requires the phosphorylation of CtBP1/BARS. EMBO J. 27, 970–981 (2008).
Brenner, S. L. & Korn, E. D. Substoichiometric concentrations of cytochalasin D inhibit actin polymerization. Additional evidence for an F-actin treadmill. J. Biol. Chem. 254, 9982–9985 (1979).
Pinot, M. et al. Lipid cell biology. Polyunsaturated phospholipids facilitate membrane deformation and fission by endocytic proteins. Science 345, 693–697 (2014).
Galbiati, F., Razani, B. & Lisanti, M. P. Emerging themes in lipid rafts and caveolae. Cell 106, 403–411 (2001).
Doherty, G. J. & McMahon, H. T. Mechanisms of endocytosis. Annu Rev. Biochem 78, 857–902 (2009).
Boucrot, E. et al. Endophilin marks and controls a clathrin-independent endocytic pathway. Nature 517, 460–465 (2015).
Holst, M. R. et al. Clathrin-Independent Endocytosis Suppresses Cancer Cell Blebbing and Invasion. Cell Rep. 20, 1893–1905 (2017).
Thottacherry, J. J. et al. Mechanochemical feedback control of dynamin independent endocytosis modulates membrane tension in adherent cells. Nat. Commun. 9, 4217 (2018).
Baba, T. et al. Clathrin-dependent and clathrin-independent endocytosis are differentially sensitive to insertion of poly (ethylene glycol)-derivatized cholesterol in the plasma membrane. Traffic 2, 501–512 (2001).
Wu, X. S. et al. Membrane Tension Inhibits Rapid and Slow Endocytosis in Secretory Cells. Biophys. J. 113, 2406–2414 (2017).
Gottlieb, T. A., Ivanov, I. E., Adesnik, M. & Sabatini, D. D. Actin microfilaments play a critical role in endocytosis at the apical but not the basolateral surface of polarized epithelial cells. J. Cell Biol. 120, 695–710 (1993).
Dai, J. & Sheetz, M. P. Regulation of endocytosis, exocytosis, and shape by membrane tension. Cold Spring Harb. Symp. Quant. Biol. 60, 567–571 (1995).
Hirama, T. et al. Membrane curvature induced by proximity of anionic phospholipids can initiate endocytosis. Nat. Commun. 8, 1393 (2017).
Loh, J. et al. An acute decrease in plasma membrane tension induces macropinocytosis via PLD2 activation. J. Cell Sci. 132, jcs232579 (2019).
Bucher, D. et al. Clathrin-adaptor ratio and membrane tension regulate the flat-to-curved transition of the clathrin coat during endocytosis. Nat. Commun. 9, 1109 (2018).
Kozlov, M. M. & Chernomordik, L. V. Membrane tension and membrane fusion. Curr. Opin. Struct. Biol. 33, 61–67 (2015).
Sitarska, E. & Diz-Munoz, A. Pay attention to membrane tension: Mechanobiology of the cell surface. Curr. Opin. Cell Biol. 66, 11–18 (2020).
Dai, J. & Sheetz, M. P. Mechanical properties of neuronal growth cone membranes studied by tether formation with laser optical tweezers. Biophys. J. 68, 988–996 (1995).
Levental, K. R. et al. Lipidomic and biophysical homeostasis of mammalian membranes counteracts dietary lipid perturbations to maintain cellular fitness. Nat. Commun. 11, 1339 (2020).
Halbleib, K. et al. Activation of the Unfolded Protein Response by Lipid Bilayer Stress. Mol. Cell 67, 673–684 e678 (2017).
Ernst, R., Ejsing, C. S. & Antonny, B. Homeoviscous Adaptation and the Regulation of Membrane Lipids. J. Mol. Biol. 428, 4776–4791 (2016).
Levitan, I. Evaluating membrane structure by Laurdan imaging: Disruption of lipid packing by oxidized lipids. Curr. Top. Membr. 88, 235–256 (2021).
Thankamony, S. P. & Knudson, W. Acylation of CD44 and its association with lipid rafts are required for receptor and hyaluronan endocytosis. J. Biol. Chem. 281, 34601–34609 (2006).
Levental, K. R. et al. Polyunsaturated lipids regulate membrane domain stability by tuning membrane order. Biophys. J. 110, 1800–1810 (2016).
Shaikh, S. R. et al. Oleic and docosahexaenoic acid differentially phase separate from lipid raft molecules: a comparative NMR, DSC, AFM, and detergent extraction study. Biophys. J. 87, 1752–1766 (2004).
Levental, K. R. et al. omega-3 polyunsaturated fatty acids direct differentiation of the membrane phenotype in mesenchymal stem cells to potentiate osteogenesis. Sci. Adv. 3, eaao1193 (2017).
Lin, X. et al. Domain Stability in Biomimetic Membranes Driven by Lipid Polyunsaturation. J. Phys. Chem. B 120, 11930–11941 (2016).
Veatch, S. L. et al. Critical fluctuations in plasma membrane vesicles. ACS Chem. Biol. 3, 287–293 (2008).
Li, P. et al. The iron chelator Dp44mT suppresses osteosarcoma’s proliferation, invasion and migration: in vitro and in vivo. Am. J. Transl. Res 8, 5370–5385 (2016).
Basuli, D. et al. Iron addiction: a novel therapeutic target in ovarian cancer. Oncogene 36, 4089–4099 (2017).
Guo, W. et al. An important role of the hepcidin-ferroportin signaling in affecting tumor growth and metastasis. Acta Biochim Biophys. Sin. (Shanghai) 47, 703–715 (2015).
Tsujita, K. et al. Homeostatic membrane tension constrains cancer cell dissemination by counteracting BAR protein assembly. Nat. Commun. 12, 5930 (2021).
Chi, Y. et al. Cancer cells deploy lipocalin-2 to collect limiting iron in leptomeningeal metastasis. Science 369, 276–282 (2020).
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging Biological Principles of Metastasis. Cell 168, 670–691 (2017).
Chen, M. B. et al. On-chip human microvasculature assay for visualization and quantification of tumor cell extravasation dynamics. Nat. Protoc. 12, 865–880 (2017).
Chen, M. B., Whisler, J. A., Jeon, J. S. & Kamm, R. D. Mechanisms of tumor cell extravasation in an in vitro microvascular network platform. Integr. Biol. (Camb.) 5, 1262–1271 (2013).
Frose, J. et al. Epithelial-Mesenchymal Transition Induces Podocalyxin to Promote Extravasation via Ezrin Signaling. Cell Rep. 24, 962–972 (2018).
Chen, M. B., Lamar, J. M., Li, R., Hynes, R. O. & Kamm, R. D. Elucidation of the Roles of Tumor Integrin beta1 in the Extravasation Stage of the Metastasis Cascade. Cancer Res 76, 2513–2524 (2016).
Wan, Z. et al. A robust vasculogenic microfluidic model using human immortalized endothelial cells and Thy1 positive fibroblasts. Biomaterials 276, 121032 (2021).
Prince, M. E. et al. Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc. Natl. Acad. Sci. USA 104, 973–978 (2007).
Dontu, G. et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 17, 1253–1270 (2003).
Lambert, A. W. et al. DeltaNp63/p73 drive metastatic colonization by controlling a regenerative epithelial stem cell program in quasi-mesenchymal cancer stem cells. Dev. Cell 57, 2714–2730.e2718 (2022).
Fruman, D. A., Meyers, R. E. & Cantley, L. C. Phosphoinositide kinases. Annu Rev. Biochem 67, 481–507 (1998).
Rodriguez, R., Schreiber, S. L. & Conrad, M. Persister cancer cells: Iron addiction and vulnerability to ferroptosis. Mol Cell (2021).
Suzuki, T. et al. Crucial role of iron in epigenetic rewriting during adipocyte differentiation mediated by JMJD1A and TET2 activity. Nucleic Acids Res (2023).
Herr, C. Q. & Hausinger, R. P. Amazing Diversity in Biochemical Roles of Fe(II)/2-Oxoglutarate Oxygenases. Trends Biochem Sci. 43, 517–532 (2018).
Dongre, A. et al. Epithelial-to-Mesenchymal Transition Contributes to Immunosuppression in Breast Carcinomas. Cancer Res 77, 3982–3989 (2017).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Contag, C. H. & Bachmann, M. H. Advances in in vivo bioluminescence imaging of gene expression. Annu Rev. Biomed. Eng. 4, 235–260 (2002).
Levental, K. R. & Levental, I. Isolation of giant plasma membrane vesicles for evaluation of plasma membrane structure and protein partitioning. Methods Mol. Biol. 1232, 65–77 (2015).
Surma, M. A. et al. Mouse lipidomics reveals inherent flexibility of a mammalian lipidome. Sci. Rep. 11, 19364 (2021).
Ejsing, C. S. et al. Global analysis of the yeast lipidome by quantitative shotgun mass spectrometry. Proc. Natl. Acad. Sci. USA 106, 2136–2141 (2009).
Surma, M. A. et al. An automated shotgun lipidomics platform for high throughput, comprehensive, and quantitative analysis of blood plasma intact lipids. Eur. J. Lipid Sci. Technol. 117, 1540–1549 (2015).
Herzog, R. et al. A novel informatics concept for high-throughput shotgun lipidomics based on the molecular fragmentation query language. Genome Biol. 12, R8 (2011).
Herzog, R. et al. LipidXplorer: a software for consensual cross-platform lipidomics. PLoS One 7, e29851 (2012).
Frolich, N., Klose, C., Widen, E., Ripatti, S. & Gerl, M. J. Imputation of missing values in lipidomic datasets. Proteomics 24, e2300606 (2024).
Sampaio, J. L. et al. Membrane lipidome of an epithelial cell line. Proc. Natl. Acad. Sci. USA 108, 1903–1907 (2011).
Shaw, F. L. et al. A detailed mammosphere assay protocol for the quantification of breast stem cell activity. J. Mammary Gland Biol. Neoplasia 17, 111–117 (2012).
Ashkin, A. Acceleration and Trapping of Particles by Radiation Pressure. Phys. Rev. Lett. 24, 156–159 (1970).
Kuo, S. C. & Sheetz, M. P. Optical tweezers in cell biology. Trends Cell Biol. 2, 116–118 (1992).
Sezgin, E. et al. Elucidating membrane structure and protein behavior using giant plasma membrane vesicles. Nat. Protoc. 7, 1042–1051 (2012).
Sezgin, E., Waithe, D., Bernardino de la Serna, J. & Eggeling, C. Spectral imaging to measure heterogeneity in membrane lipid packing. Chemphyschem: a Eur. J. Chem. Phys. Phys. Chem. 16, 1387–1394 (2015).
Owen, D. M., Rentero, C., Magenau, A., Abu-Siniyeh, A. & Gaus, K. Quantitative imaging of membrane lipid order in cells and organisms. Nat. Protoc. 7, 24–35 (2011).
Wan, Z. et al. A Robust Method for Perfusable Microvascular Network Formation In Vitro. Small Methods 6, e2200143 (2022).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Hothorn, T., Hornik, K., van de Wiel, M. A. V. & Zeileis, A. Implementing a Class of Permutation Tests: The coin Package. J. Stat. Softw. 28, 1–23 (2008).
Acknowledgements
We thank all members of the Weinberg, Farese and Walther labs for insightful discussions and reagents. We are grateful to Brent Stockwell, Laurie Boyer, Edward Lyman, and Fred Heberle for helpful discussions. We acknowledge technical support from Caroline A. Lewis, Zon W. Lai and Marina Plays, the CurieCoreTech Metabolomics and Lipidomics Technology Platform at the Institut Curie, the following facilities at the Whitehead Institute: Metabolite Profiling Core, W.M. Keck Microscopy Core, Flow Cytometry Core, as well as the Koch Institute’s Robert A. Swanson (1969) Biotechnology Center, specifically the MIT Koch Institute Animal Imaging and Pre-Clinical Testing Core Facility and Hope Babette Tang (1983) Histology Core Facility. We also thank Naviya Kapadia for assistance with proofreading and providing helpful comments on the manuscript and figures. This work was funded in part by the National Institutes of Health: Koch Institute Support (core) Grant P30CA014051 from the National Cancer Institute (W.S.H.), National Institute Of General Medical Sciences R37GM058615 (V.W.H.), the National Cancer Institute K99CA255844/R00CA255844 (N.B.), and the National Cancer Institute (R.D.K.). Additional funding support was received from MIT Stem Cell (R.A.W.), Brendan Bradley Gift (R.A.W.), Nile Albright Research Foundation (RAW), Samuel Waxman Cancer Research Foundation (R.A.W.), Virginia and D.K. Ludwig Fund for Cancer Research Center (R.A.W.), Marble Center for Cancer Nanomedicine (P.T.H.), Jane Coffin Childs Memorial Fund (W.S.H.), Ludwig Center at MIT’s Koch Institute for Integrative Cancer Research (W.S.H.), Emerald Foundation, Inc. (W.S.H.), National Institute Of General Medical Sciences 1R21CA300756 (K.R.L.), National Institute Of General Medical Sciences R35GM134949 (I.L.), National Institute Of General Medical Sciences T32GM136540 (R.P.M. & J.S.) and the New Horizon UROP Fund/MIT (V.V.P.). R.R. is supported by Ligue Contre le Cancer (Equipe Labellisée), Fondation Charles Defforey-Institut de France, Fondation Bettencourt Schueller, Institut National du Cancer, Agence Nationale de la Recherche and the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant agreement no. 647973). R.R. also received support from INCa 2022-1-PL BIO-05-ICR-1 and INCa_16712.
Author information
Authors and Affiliations
Contributions
W.S.H. and R.R. conceived the project. W.S.H, R.P.M. and S.M. designed and performed experiments with input from R.A.W. and R.R. S.M. conducted iron measurement studies. S.M. and J.L.S. performed oxidized lipidomics analysis. J.Y. performed endocytosis assays, with critical insight from V.W.H. Membrane tension studies were conducted by S.I., with support from A.E.C. on data analysis. K.R.L. performed membrane tension experiments and together with I.L. provided key insights on interpretation. W.S.H., S.D. and F.R. conducted animal experiments. Z.W. performed extravasation assays with support from R.D.K. A.D. and C.B.C provided significant evaluation and interpretation of lipidomics data. Liposomal nanoparticles and HA-Cy3 probes were synthesized by N.B. and A.V. G.W.B. and P.T. provided support for data analysis. K.J.S., V.V.P., J.S., D.H., F.S., L.S., N.S. and E.N.E. contributed to data acquisition and analysis. P.T.H. provided helpful discussion and scientific input. W.S.H., and R.P.M. wrote the manuscript. W.S.H., R.A.W., and R.R. edited the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mansell, R.P., Müller, S., Yang, JS. et al. Ether lipids influence cancer cell fate by modulating iron uptake. Nat Commun 17, 1835 (2026). https://doi.org/10.1038/s41467-026-68547-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68547-5