Abstract
Stem cells continually self-renew and differentiate to sustain tissue homeostasis, yet the role of post-transcriptional mechanisms in guiding these processes remains incompletely understood. Here, we demonstrate that the regulation of 3’UTR length via alternative mRNA polyadenylation (APA) is essential for stem cell function across diverse tissues. Modulating the APA regulator Nudt21 reveals that stem cell self-renewal and differentiation depend on distinct dosage thresholds and thus can be uncoupled. Specifically, moderate Nudt21 suppression elicits a maturation arrest of stem cells due to 3’UTR-shortening of differentiation-associated mRNAs that escape miRNA regulation and perturb ceRNA networks. By contrast, complete Nudt21 suppression additionally shortens the 3’UTRs of mRNAs encoding essential multiprotein complexes, including the nuclear pore, leading to complex destabilization, proteotoxic stress, DNA damage, and cell cycle arrest. Critically, deletion of the alternative 3’UTRs of individual nucleoporins recapitulates defects observed with Nudt21 loss. We further demonstrate that the co-translational assembly of dozens of protein complexes is impaired in Nudt21-deficient cells, providing a mechanistic framework for compromised complex integrity. Collectively, our results show that APA plays distinct, dose-dependent roles in stem cell homeostasis by fine-tuning the expression of differentiation-associated genes and coordinating the biogenesis of multiprotein complexes essential for cell cycle progression.
Similar content being viewed by others
Introduction
The maintenance of regenerative tissues typically relies on dedicated stem cells that continually self-renew and differentiate to replenish specialized cells lost during homeostasis or injury1. While the self-renewal and differentiation of tissue stem cells are known to be regulated by transcriptional and epigenetic mechanisms, the role of post-transcriptional mechanisms in stem cell regulation remains largely unexplored. Notably, over 50% of mammalian genes contain more than one polyadenylation signal (PAS), allowing mammals to greatly diversify the production of mRNA isoforms via APA2. APA has emerged as a crucial post-transcriptional layer of gene regulation that largely acts in parallel with conventional transcriptional mechanisms of gene expression. Changes in APA typically cause a lengthening or shortening of 3’UTRs, which facilitates the respective inclusion or exclusion of sequences targeted by miRNAs and RNA-binding proteins. This, in turn, can alter the stability and/or translation efficiency of affected transcripts3,4,5,6,7,8,9. More recently, APA has also been implicated in the subcellular localization of certain proteins, particularly surface molecules, as well as in the assembly of alternative cancer-associated protein complexes7,10,11. In addition to generating mRNA isoforms that differ exclusively in their 3’UTR (3’UTR APA), APA within introns (intronic APA) can give rise to mRNA isoforms that differ in their coding sequence due to the selection of alternative terminal exons12. Thus, 3’UTR APA and intronic APA have the potential to profoundly impact the function of proteins.
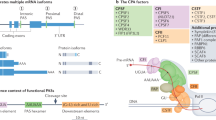
The cleavage and polyadenylation (CPA) machinery, which regulates the 3’ end processing of mRNAs, is composed of multiple subcomplexes with dedicated functions, including CPSF, CstF, CFIm, and CFIIm13,14,15. Of these subcomplexes, only CFIm (cleavage factor I) is dispensable for general mRNA 3’ end processing, but essential for directing 3’UTR APA16,17. Indeed, as many as 1,000 transcripts undergo 3’UTR changes upon siRNA-mediated knockdown of the CFIm components Nudt21 (also known as Cpsf5) or Cpsf618,19, which results in the destabilization of the entire CFIm complex19,20,21. In the absence of either Nudt21 or Cpsf6, the proximal PAS is favored to generate mRNA isoforms with short 3’UTRs, consistent with the CFIm complex functioning as an activator of distal PAS usage22. Accordingly, distal PAS sites are enriched for the Nudt21 binding motif UGUA compared to proximal PAS sites23. However, only a fraction of mRNAs undergoing 3’UTR shortening in the absence of Nudt21 produce increased protein levels, pointing to more complex roles of APA in gene regulation5,24. Together, these data indicate that the 3’ end processing of all mRNAs is controlled by the CPSF/CstF/CFIIm subcomplexes of the CPA machinery, while the APA of mRNAs with multiple PAS sites is additionally controlled by the CFIm complex.
Suppression of CFIm components, particularly Nudt21, has previously been reported to promote6 or suppress25,26 tumorigenesis in diverse cancer contexts. Moreover, the reduction of Nudt21 levels has been suggested to compromise preimplantation development27, certain neurological functions20,28,29 and pulmonary fibrosis30,31. Our lab previously showed that suppression of Nudt21 enhances the reprogramming of somatic cells to induced pluripotent stem cells (iPSCs) by upregulating critical chromatin factors that facilitate cellular plasticity18. Moreover, Nudt21 has been suggested to guide the activation of quiescent hematopoietic stem cells (HSCs) by controlling the switch between two different glutaminase 3’UTR isoforms32, and it reportedly modulates the expression of the B cell surface antigen CD19 in acute lymphoblastic leukemia33. While these studies implicate APA in the modulation of specific genes in particular experimental or disease contexts, its broader physiological roles and associated mechanisms remain largely unexplored.
Here, we investigate the functional significance of APA through conditional deletion of the key CFIm component Nudt21 in mice. Our findings reveal a previously unrecognized, essential role for APA in maintaining actively regenerating tissues, including various epithelia and the hematopoietic system, by ensuring the persistence of proliferative stem and progenitor cell populations. Through mechanistic dissection of Nudt21-deficient stem cells, we uncover a dose-dependent function of APA that (1) ensures lineage-specific gene expression important for differentiation, and (2) coordinates the stability of multiprotein complexes crucial for cell division in the context of stem cell self-renewal and progenitor cell proliferation. We highlight nucleoporins as central Nudt21 targets, whose APA safeguards the stability and function of the nuclear pore complex. Together, our results reveal that APA is indispensable for organismal survival by maintaining tissue homeostasis and suggest an APA-dependent mechanism by which essential multiprotein complexes are coordinately regulated in dividing cells to safeguard stem cell self-renewal and progenitor cell proliferation.
Results
CFIm is enriched in stem and progenitor cells of highly regenerative tissues
To determine whether Nudt21, a key component of the CFIm complex, is dynamically regulated during tissue homeostasis, we performed quantitative immunofluorescence (IF) analysis across three highly regenerative tissues containing defined stem, progenitor, and differentiated cell populations. We first examined the esophagus, which is a simple stratified squamous epithelium comprised of basal stem cells, which continuously self-renew and differentiate into suprabasal and superficial cells. While Nudt21 was detectable across the entire epithelium, we observed a gradient of Nudt21 levels that was highest, albeit variable, in Keratin (Krt) 14+ basal stem cells, lower in Krt14- suprabasal cells and lowest in the superficial layer of the epithelium containing the most differentiated cells (Fig. 1a). We similarly detected differential enrichment of Nudt21 levels in the intestinal epithelium, which is organized into glandular units containing highly proliferative stem and progenitor cells within the crypt compartment, as well as differentiated progeny within the villus compartment. Mirroring our observations in the esophagus, Nudt21 levels were highest in Lgr5+ intestinal stem cells and lowest in mature villus cells, with Lgr5- crypt-based progenitors exhibiting intermediate levels of Nudt21 (Fig. 1b). To explore whether the differential expression of Nudt21 within regenerative epithelia extends to a non-epithelial tissue with high turnover, we measured Nudt21 levels in the hematopoietic system. Indeed, Nudt21 was most abundant in HSCs and multipotent progenitors (MPPs) but less abundant in more committed myeloid progenitors (MyPs) and lineage-positive (Lin+) cells (Fig. 1c). We confirmed the differential expression of the CFIm components Nudt21, Cpsf6, and Cpsf7 across stem/progenitor and mature cell populations using published single-cell RNA-sequencing data for the intestine34 and esophagus35 (Supplementary Fig. 1a-d), as well as our own unpublished data of hematopoietic stem and progenitor cells (HSPCs) (Supplementary Fig. 1e and 1f). NUDT21 mRNA levels were also higher in human HSPCs compared to mature blood cell types using the bloodspot database36 (Supplementary Fig. 2a). Together, these observations indicate that CFIm expression is dynamically regulated in stem and progenitor cells across diverse tissues and species.
a-c: Nudt21 immunofluorescence quantification of various mouse tissues. Single dots represent individual nuclei, solid lines represent the median. a Esophagus. Krt14 marks basal stem cells (n = 3 biol. replicates). b Intestine. Lgr5-GFP marks crypt stem cells (n = 3 biol. replicates). c FACS-purified hematopoietic cells (n = 3 biol. replicates). d Design of Nudt21 floxed (fl) allele (top). Nudt21fl/fl were crossed to R26-CreER or K5-CreER strains to generate ubiquitous or basal cell-specific Nudt21 KO mice (bottom). Nudt21fl/fl lacking the respective CreER allele were used as controls. e Validation of Nudt21 KO by western blot analysis. ETOH (control) or 4OHT (KO) was applied for 3 days (representative of three independent experiments). f Survival curve for conditional Nudt21 KO mice (3-6 months old) after 5 consecutive daily tamoxifen injections. Control: n = 8, Nudt21fl/fl; R26-CreER: n = 5, Nudt21fl/fl; K5-CreER: n = 4. g H&E staining of esophagus from control and Nudt21fl/fl; K5-CreER (KO) mice, analyzed 10 days after receiving 5 consecutive daily tamoxifen injections (top). Dashed line: basal membrane. BrdU IHC (bottom), arrowheads: BrdU-positive nuclei. IHC for Nudt21 (inset). h Quantification of (g) (n = 3 biol. replicates). i H&E of intestinal epithelium from control and Nudt21fl/fl; R26-CreER (KO) mice, analyzed 6 days after receiving 5 consecutive daily tamoxifen injections (top). BrdU IHC (bottom). Nudt21 IHC (inset). j Quantification of (I) (n = 3 biol. replicates). k H&E of bone marrow from control and Nudt21fl/fl; R26-CreER (KO) mice, analyzed 7 days after receiving 5 consecutive daily tamoxifen injections. Arrowheads: hypocellularity. l Design of bone marrow chimeras. m Complete blood count (CBC) of peripheral blood from mice in (l), 14 days after receiving 5 consecutive daily tamoxifen injections (n = 4 biol. replicates). Statistical analysis: Box plots show median (center), 25th–75th percentiles (box), and whiskers to 1.5×IQR; outliers plotted individually. For all experiments, adult mice (3–6 months old) were used. Statistical analyses: Error bars represent the standard deviation of the mean for biological replicates. Statistical significance was determined using a two-tailed unpaired Student’s t-test (a-c, h,j and m), Log-rank (Mantel-Cox) test (f).
CFIm is required for the maintenance of diverse regenerative tissues
To probe the functional role of the CFIm complex in vivo, we generated Nudt21 knockout (Nudt21 KO) murine embryonic stem cells (ESCs) in which the RNA-binding domain essential for Nudt21 function has been flanked by loxP (fl) sites. These ESCs were then used to derive conditional Nudt21 (Nudt21fl/fl) mice (Fig. 1d). We crossed Nudt21fl/fl mice to Rosa26 (R26)-CreER mice, allowing us to delete Nudt21 ubiquitously following administration of tamoxifen. We tested the deletion of the targeted allele by exposing tail keratinocytes explanted from these animals to the tamoxifen analog 4-Hydroxytamoxifen (4OHT), which resulted in a loss of Nudt21 protein, as expected (Fig. 1e). In addition, we crossed Nudt21fl/fl mice to Keratin 5 (Krt5) CreER mice (Fig. 1d) to delete Nudt21 specifically in the basal stem cells of squamous epithelia including the esophagus where we observed elevated Nudt21 levels (Fig. 1a). Strikingly, the ubiquitous, acute loss of Nudt21 with R26-CreER led to the rapid morbidity of animals within 5-7 days. Mice with acute deletion of Nudt21 exclusively in Krt5+ squamous epithelia survived slightly longer and succumbed within 7-14 days (Fig. 1f). Together, these findings point to an essential role of the CFIm complex in the survival of adult mice.
To characterize possible phenotypes underlying the morbidity of Nudt21-deficient mice, we performed histology and marker analyses. Overall, ubiquitous Nudt21 deletion did not result in obvious defects in static tissues such as the kidney, heart and skeletal muscle (Supplementary Fig. 2b). However, when we examined animals with squamous tissue-specific Nudt21 loss, we observed a significant reduction in basal cell density in the esophagus (Fig. 1g top and 1h left), tongue and oral epithelium (Supplementary Fig. 2c), as well as morphologically abnormal cells within the suprabasal layers of these epithelia, suggesting a defect in basal stem cell self-renewal and differentiation. Consistent with this observation, we detected fewer Bromodeoxyuridine-positive (BrdU+) proliferative cells in the basal layer of the esophagus in these animals (Fig. 1g bottom and 1h right). Examination of actively regenerating tissues in ubiquitous Nudt21 KO mice further revealed a defect in the intestinal epithelium over time. Specifically, we observed a reduction in villus length and a decrease in BrdU+ cells in the crypt compartment, leading to the disintegration of the epithelium within 8 days (Figs. 1i and 1j, and Supplementary Fig. 2d). These results suggest a requirement for CFIm activity to sustain stem and progenitor cell populations across diverse epithelial tissues crucial for organismal survival.
We also noted that tamoxifen-induced ubiquitous Nudt21 KO mice exhibited reduced bone marrow cellularity, suggesting a role for CFIm in the hematopoietic system (Fig. 1k). To assess whether CFIm is required for hematopoiesis, we transplanted the bone marrow of untreated Nudt21fl/fl; R26-CreER (experimental) and Nudt21fl/fl (control) mice into irradiated recipient mice and then treated these with tamoxifen (Fig. 1l). While control mice had normal numbers of blood cells two weeks after tamoxifen treatment, most experimental mice became morbid and developed a severe pancytopenia as reflected by the decline in erythrocytes, platelets, and leukocytes in the peripheral blood (Fig. 1m). To confirm the cell autonomy of this phenotype, we generated mixed bone marrow chimeras containing either Nudt21fl/fl; R26-CreER or Nudt21fl/fl bone marrow in combination with wild-type bone marrow. Nudt21 deletion in these chimeras led to a selective loss of Nudt21-deficient HSCs, MPPs and mature blood cells without disrupting wild-type hematopoietic cells in the same animals, confirming an essential and cell-intrinsic requirement for Nudt21 in hematopoiesis (Supplementary Fig. 2e-2h). Collectively, our results establish that CFIm and thus APA are cell-autonomously required for the survival of adult animals by controlling the maintenance of stem and progenitor cell populations in various actively regenerating tissues.
CFIm maintains stem and progenitor cells by enabling cell cycle progression
Considering the profound effect Nudt21 loss has on stem and progenitor cells within actively regenerating tissues, we hypothesized that CFIm may be required for cell cycle progression. To this end, we established basal stem cell cultures from the esophageal epithelium of Nudt21fl/fl; R26-CreER mice and measured their distribution in the cell cycle phases, G1, S, and G2/M over time in the presence and absence of 4OHT (Figs. 2a, 2b). Compared to control cells, esophageal stem cells treated with 4OHT exhibited a progressive loss of S phase cells and a concomitant increase in G1 phase and G2/M phase cells (Figs. 2b-2d). By day 6 of 4OHT treatment, the majority of stem cells ceased to synthesize DNA and arrested in either G1 or G2/M. Acute Nudt21 deletion did not lead to increased apoptosis within the examined time window (6 days), although cells began to die shortly thereafter, indicating that cell death is a consequence rather than a driver of cellular defects incurred by the loss of CFIm function (Fig. 2e).
a Scheme outlining the explantation and culture of Nudt21fl/fl; R26-CreER esophageal basal stem cells in self-renewal conditions. Cells were cultured in either ETOH (Control) or 4OHT (Nudt21 KO). b Representative FACS-based cell cycle analysis of esophageal basal stem cells cultured in self-renewal conditions outlined in (a). Cells were pulsed with EdU for 1 hour to label cells in S-phase. c Quantification of representative data shown in (b) using time-course analysis (n = 3 biol. replicates). d Quantification of cell proliferation of samples shown in (c) (n = 3 biol. replicates). e FACS-based quantification of viable cells using Annexin V and propidium iodide stains (n = 3 biol. replicates). f Scheme to produce satellite cell-specific conditional Nudt21 KO mice. Nudt21fl/fl; Pax7-CreER(gaka) (experimental) and Nudt21fl/fl (control) mice received 5 consecutive daily tamoxifen injections before being placed on tamoxifen chow and analyzed 30-40 days after the start of tamoxifen treatment. Satellite cells were subsequently isolated by FACS using indicated surface markers and used for further analysis (h, i and m-o). g Tibia mass-to-tibia length ratio of mice outlined in (f). Nudt21fl/fl: n = 4, Nudt21fl/fl; Pax7-CreER(gaka): n = 3. h Quantification of satellite cells isolated from indicated mice (n = 3 each). i PCR-based assay to confirm Nudt21 excision. j Scheme of muscle injury experiment. Mice treated with tamoxifen as outlined in (f) were subjected to cardiotoxin-induced injury of the tibialis anterior muscles. Regeneration efficiency was assessed 2 weeks post injury. k Evaluation of muscle regeneration at injury site via H&E staining or dystrophin immunofluorescence. Regenerated fibers contain centrally located nuclei (white arrowhead). l Quantification of regenerated muscle fibers with central nuclei from dystrophin immunofluorescent images shown in (k) (n = 3 biol. replicates). m Scheme of experiment to explant and expand satellite cells using satellite expansion media. n Microscopy images of explanted satellite cells. o Quantification of cell numbers of explanted satellite cells shown in (n) (n = 3 biol. replicates). Statistical analysis: Error bars represent the standard deviation of the mean for biological replicates. Statistical significance was determined using a two-tailed unpaired Student’s t-test (d,e,g,h,l, and o).
To examine whether Nudt21 loss disrupts cell cycle progression in other actively regenerating tissues, we explanted intestinal, hematopoietic, and skin cells from Nudt21fl/fl; R26-CreER mice in conditions that support the propagation of resident stem and progenitor cells (Supplementary Fig. 3a-3i). Paralleling our observations in esophageal cultures, intestinal, hematopoietic, and skin keratinocyte cultures exhibited a pronounced reduction of cells in S phase with an accumulation of cells at the G1 and/or G2/M phases of the cell cycle within 6 days of 4OHT treatment. These findings suggest that a CFIm-dependent cell cycle arrest of self-renewing stem cells and proliferative progenitors underlies the observed regenerative defects in animals.
CFIm is dispensable in quiescent stem cells unless they enter the cell cycle
As CFIm loss-of-function in stem cells of actively regenerating tissues elicits a profound self-renewal defect, we next asked whether CFIm is required for the maintenance of stem cells that have transiently exited the cell cycle. Skeletal muscle contains a rare population of stem cells termed satellite cells, which are quiescent in homeostasis but activated upon injury or explantation ex vivo. Once activated, satellite cells enter the cell cycle to proliferate and regenerate the injured muscle37,38,39. Analysis of published single-cell RNA-sequencing data40 revealed that Nudt21 is highly expressed in quiescent and activated satellite cells but downregulated in mature muscle cells (Supplementary Fig. 4a and 4b), consistent with our observations in epithelia and the bone marrow. To study CFIm function in quiescent satellite cells, we crossed Nudt21fl/fl mice to Pax7-CreER(gaka) mice41, which enable satellite cell-specific induction of Cre activity while maintaining endogenous Pax7 expression (Fig. 2f). We observed normal numbers of satellite cells and muscle mass in Nudt21fl/fl; Pax7-CreER mice at 30-40 days after tamoxifen treatment despite efficient Nudt21 deletion (Figs. 2g-2i), and these animals showed no overt muscle pathologies during this experimental time frame. These data suggest that CFIm function is dispensable for satellite cell maintenance in uninjured muscle. By contrast, we detected a severely impaired regenerative response upon cardiotoxin-mediated muscle injury of tamoxifen-treated Pax7-CreER (gaka); Nudt21fl/fl mice (Figs. 2j-2l), with reduced numbers of newly regenerated (centrally nucleated) myofibers and increased scar tissue deposition compared to controls. Consistent with a proliferation-dependent stem cell defect, satellite cells explanted from Nudt21fl/fl; Pax7-CreER mice failed to expand in culture, in contrast to satellite cells explanted from control animals (Figs. 2m-2o). Collectively, our results suggest that CFIm and hence APA are dispensable in quiescent muscle stem cells, but essential in activated muscle stem cells forced to proliferate in response to muscle damage in vivo or external stimuli ex vivo.
Modulation of CFIm levels uncouples self-renewal from differentiation
Previous studies by our lab and others have demonstrated that transient siRNA-mediated suppression of CFIm enhances the reprogramming of fibroblasts into iPSCs, and promotes tumor cell growth6,18,42. However, our findings here indicate that complete loss of CFIm function compromises stem cell self-renewal and drives subsequent failures of tissue regeneration, which are phenotypes that were not observed with siRNA mediated silencing. These differences imply that varying levels of CFIm, as observed in Nudt21 knockdown versus knockout approaches, may have distinct effects on cellular functions. To test this hypothesis, we assessed the cellular and molecular consequences of reducing Nudt21 levels to different extents. To this end, we introduced a doxycycline (dox)-inducible hemagglutinin-tagged Nudt21 (HA-Nudt21) lentiviral transgene into Nudt21fl/fl; R26-CreER esophageal stem cells, allowing us to titrate Nudt21 levels in a Nudt21-deficient background (Fig. 3a). We then determined the effects of different Nudt21 levels on the self-renewal and differentiation potentials of esophageal stem cells. Reassuringly, dox titration in transduced esophageal cultures led to dose-dependent expression of Nudt21 protein (Fig. 3b bottom). Moreover, the expression levels of Cpsf6, another integral subunit of the CFIm complex, scaled with the expression levels of Nudt21 in our titratable system, consistent with Nudt21 being an essential and limiting component of the CFIm complex19,20,21. Leveraging this system, we determined that a reduction of Nudt21 levels down to ~25% of levels in control cells using 50 ng/ml dox is sufficient to support stem cell self-renewal and cell cycle progression (Fig. 3b top) for at least 8 passages. We refer to this condition as Nudt21 hypomorph. However, when Nudt21 levels dropped below ~25%, stem cells could not be maintained due to an irreversible cell cycle arrest.
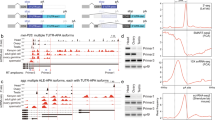
a Generation of Nudt21 conditional KO esophageal basal stem cells with dox-inducible HA-Nudt21 transgene. b Top: FACS-based EdU incorporation assay. Bottom: Western blot analysis for CFIm complex members Nudt21, Cpsf6, Cpsf7, and transgenic HA-Nudt21 (HA-tag), (n = 4 biol. replicates). c Immunofluorescence (top) and proteomic (bottom) analyses for differentiation markers (Krt13, Ppl), stem cell markers (Trp63, Sox2) and CFIm complex members (Nudt21, Cpsf6 and Cpsf7), in cultures exposed to calcium for 6 days (n = 3 biol. replicates). d, e Changes to APA in Nudt21 KO and Hypo cells relative to control, as measured by PAS-seq analysis. Plots display the log2-fold ratios of proximal PAS (“P”) to distal PAS (“D”) usage for each comparison. APA events determined by a significant false discovery rate (FDR) < 0.05 and changes in distal PAS usage > 14.5%. Cells were harvested for analysis after 4 days of ETOH, 4OHT or 4OHT+Dox treatment. f Euler plot showing common and distinct changes to PAS usage. Commonly shortened 3’UTRs (944 mRNAs) were used for subsequent analysis (g-j). g mRNAs undergoing 3’UTR shortening and containing putative miRNA target sites. Only binding sites for expressed miRNAs were considered. h Model: 3’UTR shortening eliminates miRNA binding sites, leading to increased protein levels and a stem cell differentiation block. i Relative mRNA and protein expression levels of Nudt21 targets that follow the model in (h), with the number of cognate miRNAs shown on the right. Analysis was performed after 4 days of ETOH (Control) or 4OHT (KO) or 4OHT+Dox (Hypo) treatment. Cut-offs: log2FC ≥ 1.3, and FDR ≤ 0.05. j PAS-seq gene-tracks for Jmjd1c and Sec24a, including examples of miRNA binding sites. k Effect of Jmjd1c and Sec24a suppression on differentiation. l Quantification of Trp63 signal from images generated in (k) (n = 3 biol. replicates). Statistical analysis: Box plots show median (center), 25th–75th percentiles (box), and whiskers to 1.5×IQR; outliers plotted individually; point inside boxplot signifies geometric mean. Error bars represent standard deviation of the mean for biological replicates. Statistical significance was determined using a non-parametric unpaired, two-tailed, Wilcoxon signed-rank test (l).
To assess the impact of different Nudt21 levels on differentiation, we treated esophageal stem cell cultures with calcium, which triggers a rapid cell cycle exit and subsequent epithelial differentiation43 (Fig. 3c, Supplementary Fig. 4c and d). Specifically, we compared Nudt21 control cultures, Nudt21 KO cultures and Nudt21 hypomorph cultures. We determined that the calcium-induced cell cycle arrest occurs prior to the Nudt21 KO-induced cell cycle arrest, allowing us to assess whether Nudt21 suppression affects differentiation independently from the self-renewal defect noted above. Expression of the differentiation marker Keratin 13 (Krt13) was upregulated in control cells exposed to calcium, as expected. By contrast, Nudt21 hypomorph and KO cells failed to upregulate Krt13, suggestive of a differentiation block (Fig. 3c top and Supplementary Fig. 4d). Consistent with this observation, expression of the basal stem cell marker Trp63 was downregulated in control cells but aberrantly maintained in Nudt21 hypomorph and KO cells. The behavior of these two markers extended to other proteins, as differentiation-associated proteins were less effectively induced, while stem cell-associated proteins were retained in Nudt21 hypomorph and KO cells compared to control (Fig. 3c bottom). Critically, conditional KO mice acutely depleted for Nudt21 showed a similar reduction in Krt13 expression in primary esophagus tissue, demonstrating that our observations in cultured stem cells extend to an in vivo setting (Supplementary Fig. 4e). Together, these titration experiments indicate that Nudt21/CFIm act in a dose-dependent manner, thus uncoupling the differentiation defect from the self-renewal defect in Nudt21-depleted epithelial stem cells.
CFIm regulates polyA site usage in a dose-dependent manner
To understand molecularly how different Nudt21 levels exert distinct phenotypes in stem cells, we performed polyA site-sequencing44 (PAS-Seq) using Nudt21 control, KO and hypomorph esophageal stem cell cultures. Of all detected mRNAs (n = 4,398), 52% or 2,272 mRNAs changed in 3’UTR length (i.e., shortening or lengthening) upon acute deletion of Nudt21 (Fig. 3d). Most of these changes involved a shortening of 3’UTRs from distal to proximal PAS sites (n = 2,173 mRNAs), whereas only a minority of changes involved a lengthening of 3’UTRs from proximal to distal PAS sites (n = 99 mRNAs), consistent with previous observations using Nudt21 knockdown strategies18,45. However, the number of mRNAs undergoing APA in Nudt21 KO cells was two- to three-fold higher than in previous knockdown studies6,18, supporting the dose-dependent effects of Nudt21 depletion on APA patterns. The extent of APA changes in our hypomorph cells was significantly smaller compared to KO cells and resembled the scope of APA changes previously seen in Nudt21 knockdown experiments, with 955 mRNAs undergoing 3’UTR shortening and 41 mRNAs undergoing 3’UTR lengthening (Fig. 3e). Of note, most transcripts undergoing 3’UTR shortening in hypomorph cells (n = 944 genes or 99%) overlapped with transcripts undergoing 3’UTR shortening in Nudt21 KO cells. By contrast, there was minimal overlap between transcripts undergoing 3’UTR lengthening in Nudt21 KO and hypomorph cells (n = 27 genes or 24%), suggesting shortened mRNAs are common, direct targets while lengthened mRNAs are indirect targets of Nudt21 (Fig. 3f). Together, these observations reinforce our conclusion that Nudt21/CFIm acts as a dose-dependent regulator of APA and provide an experimental basis for defining relevant mediators of self-renewal and differentiation.
APA drives differentiation by modulating miRNA access to 3’UTRs
To narrow down CFIm targets underlying the differentiation block, we determined common APA changes between Nudt21 KO and hypomorph cells, considering that both genotypes lead to a marked differentiation block (Fig. 3c). We focused on the 944 transcripts commonly shortened in Nudt21 KO and hypomorph cells (Fig. 3f). Changes in APA usage have been associated with the elimination of miRNA target sites in the 3’UTRs of associated genes, which can lead to increased transcript and protein levels3,4,6,42,46 and consequently altered differentiation18,47,48,49 (Fig. 3h). We thus compared miRNAs expressed in basal stem cells (Supplementary Fig. 4f) with the 944 transcripts undergoing 3’UTR shortening in both Nudt21 KO and hypomorph cells to define potential matches. Strikingly, as many as 75% of shortened 3’UTRs but only ~30% of unchanged 3’UTRs contained binding sites for expressed miRNAs (Fig. 3g), suggesting that the majority of Nudt21 targets undergoing shortening in both Nudt21 KO and hypomorph cells are likely regulated by miRNAs.
To pinpoint relevant candidates among the 944 commonly shortened transcripts that follow this mode of gene regulation, we integrated our RNA-Sequencing, miRNA-Sequencing and proteomics data from Nudt21 KO and hypomorph cells, arriving at 24 high-confidence targets with increased RNA and protein levels (Fig. 3i). For example, we detected binding sites for 11 expressed miRNAs within the 3’UTR of the histone demethylase Jmjd1c previously validated as a relevant Nudt21/APA target during iPSC reprogramming18. Similarly, we detected binding sites for 28 expressed miRNAs within the 3’UTR of the COPII vesicle component Sec24a. Nudt21 KO and hypomorph cells exhibited a shift from distal to proximal PAS usage in the 3’UTRs of Jmjd1c and Sec24a, which is expected to exclude binding sites for cognate miRNAs (Fig. 3j)46,47.
To assess whether the aberrant upregulation of Jmjd1c or Sec24a contributes to the differentiation block of Nudt21-depleted cells, we suppressed each gene using siRNAs in Nudt21 hypomorph esophageal stem cells undergoing differentiation. Remarkably, suppression of either Jmjd1c or Sec24a expression ameliorated the differentiation block, as determined by upregulation of the differentiation marker Krt13, and downregulation of the basal cell marker Trp63 (Figs. 3k and 3l). Of note, the long 3’UTR isoforms of Jmjd1c and Sec24a were detectable throughout differentiation in control cells, implying that miRNAs are continuously required to suppress these targets (Supplementary Fig. 4g). To determine whether specific miRNAs targeting Jmjd1c and Sec24a 3’UTRs underlie the differentiation defect of Nudt21 hypomorph cells, we silenced miR-449a-5p, miR-30d-5p, and miR-34a-5p using anti-miRs in wild-type esophageal cells. The silencing of miR-34a, which targets Jmjd1c, delayed differentiation akin to Nudt21 hypomorphs, as measured by elevated Trp63 expression (Supplementary Fig. 4h). Notably, some anti-miRs decreased p63 levels, pointing to complex regulatory interactions between mRNAs and their cognate miRNAs. Collectively, these findings illustrate that the differentiation block in Nudt21-deficient cells is driven, at least in part, by the dysregulated post-transcriptional control of key miRNA targets including Jmjd1c and Sec24a.
APA rewiring impacts global ceRNA networks
In addition to mediating individual mRNA-miRNA interactions, 3’UTRs have previously been implicated in the co-regulation of mRNAs with shared miRNA binding sites. As these co-regulated mRNAs typically compete for the same miRNAs, they are often referred to as competing endogenous RNAs (ceRNAs)50. To test whether widespread 3’UTR shortening via Nudt21 modulation influences ceRNA networks, we grouped mRNAs based on common miRNA binding sites using wild-type esophageal stem cells. RNA pairs were defined as ceRNAs if they were co-regulated across biological replicates (>0.6 Pearson’s correlation coefficient). In wild-type cells, 41.9% of mRNA pairs behaved as ceRNAs based on these criteria (Supplementary Fig. 5a). However, in Nudt21 KO cells, only 27.7% of mRNA pairs were classified as ceRNAs (Supplementary Fig. 5b), while in Nudt21 hypomorph cells, 36.3% of mRNA pairs qualified as ceRNAs (Supplementary Fig. 5c). Inspection of mRNAs co-regulated in this fashion revealed the aforementioned Sec24a transcript as a bona fide ceRNA. Sec24 shows strong co-regulation with dozens of other mRNAs targeted by the same miRNAs including the chromatin regulators Dnmt3a and Baz2b (Supplementary Fig. 5d). However, this co-regulation is severely disrupted in Nudt21 KO and hypomorph cells (Supplementary Fig. 5e and f). Thus, loss of the long 3’UTR isoform of Sec24a appears to disrupt a strong ceRNA network by altering the corresponding miRNA target landscape. More generally, these observations reinforce our conclusion that APA modulates miRNA-dependent gene expression and associated ceRNA networks critical for differentiation42.
APA sustains stem cell self-renewal by maintaining essential multiprotein complexes
To define APA targets underlying the self-renewal defect of Nudt21-deficient cells, we performed multiplexed mass spectrometry-based proteomics51 of control and Nudt21 KO cells at different time points after 4OHT treatment (0, 2, 4, 6 days), as well as Nudt21 hypomorph cells at day 6 (Hd6). As endogenous Nudt21 protein levels were fully depleted only by day 4 of 4OHT treatment (Fig. 4a), we determined differentially expressed proteins (DEPs) that were commonly deregulated on days 4 and 6 (n = 1,677) in Nudt21 KO cells (Fig. 4b). Moreover, since Nudt21 hypomorph cultures self-renew normally, we focused only on those 353 candidates that changed protein levels and 3’UTR length in Nudt21 KO cells but not in hypomorph cells. Given Nudt21’s crucial role in self-renewal, we selected essential genes from the remaining 353 candidates based on the publicly available ‘essentialome’ dataset DepMap52, leading to 73 essential candidates for final analysis (Fig. 4b). Of note, the majority of these candidate proteins (55 out of 73 proteins (75%) at day 4, and 58 out of 73 proteins (80%) at day 6) were downregulated in the Nudt21 KO cells (Fig. 4c), suggesting a strong disruption of essential cellular proteins (Fig. 4b). Indeed, when we examined the expression levels of all essential genes detected in our proteomics dataset (n = 4,386) regardless of whether they are Nudt21 targets, we observed a progressive downregulation of these genes in Nudt21 KO cells starting on day 4, but stable expression in Nudt21 hypomorph cells (Supplementary Fig. 6a). These results imply that Nudt21 deficiency impacts diverse essential cellular processes beyond direct Nudt21 targets. Conversely, non-essential genes (n = 21,768) exhibited a subtle yet significant upregulation in Nudt21 KO and hypomorph cells (Supplementary Fig. 6b), consistent with an escape from miRNA-mediated suppression (see Fig. 3). In line with this interpretation, non-essential genes were more likely to be targeted by miRNAs than essential genes (Supplementary Fig. 6c). Consistent with their essentiality, the 73 candidate genes have more PAS sites than expected by chance, and this enrichment is conserved between mouse and human, pointing to common regulatory roles of associated 3′UTRs (Supplementary Fig. 6d and e). Together, these data suggest that APA plays distinct and opposing roles in the regulation of proteins driving self-renewal (downregulated with Nudt21 loss) and differentiation (upregulated with Nudt21 loss via miRNAs).
a Western blot analysis using a Nudt21-specific antibody. Asterisk marks unspecific band persisting in Nudt21 KO cells (representative of three independent experiments). b Strategy to narrow down potential CFIm targets driving the self-renewal defect of Nudt21 KO cells, by comparing differentially polyadenylated mRNAs (PAS-seq) and differentially expressed proteins (DEPs) (quantitative multiplexed mass spectrometry-based proteomics) between Nudt21 KO and Nudt21 hypomorph cells, and by considering only essential genes for further analysis. Boxplots show relative protein levels over time. c Pie charts illustrating the fractions of up-regulated and down-regulated proteins in esophageal basal stem cells on day 4 and day 6 of Nudt21 deletion when applying the filters shown in (b). d Heatmap displaying relative mRNA and protein levels for the 73 candidates identified in (b) using esophageal basal stem cells on day 0 (control) and day 4 (experimental) after Nudt21 deletion, alongside Nudt21 hypomorph cells (Hypo). Note that the translation efficiency (TE) of the 73 candidate genes cannot explain the observed changes in protein levels. To the right of the heatmap, gene ontology terms are indicated for enriched groups of proteins. e Experimental outline to determine how long after 4OHT-induced Nudt21 deletion, expression of an exogenous, dox-inducible HA-Nudt21 transgene can rescue the self-renewal defect of esophageal stem cells. f Effect of exogenous HA-Nudt21 expression on the continual self-renewal of Nudt21-deficient basal stem cells, as determined by measuring the fraction of cells in S phase (n = 4 biol. replicates). g Global enrichment analysis for protein complexes detected in esophageal stem cells (Nudt21 KO vs control, day 4) based on the EBI Complex Portal106 (mouse paralogs of human proteins were searched). Statistical analysis: Box plots show median (center), 25th–75th percentiles (box), and whiskers to 1.5×IQR; outliers plotted individually. Error bars represent standard deviation of the mean for biological replicates. Statistical significance was determined using a two-tailed unpaired Student’s t-test (f).
To gain insights into the regulation of the 73 candidates associated with the self-renewal defect of Nudt21 KO stem cells, we compared the dynamics of their RNA and protein levels using our RNA-Sequencing and proteomics data, respectively (Fig. 4d). We focused on day 4 after Nudt21 deletion since cells have not yet undergone cell cycle arrest (Fig. 2b), yet endogenous Nudt21 levels are fully depleted (Fig. 4a) and the global 3’UTR landscape has been rewired (Fig. 3d and e). Further emphasizing the significance of day 4 as a critical time point, we found that a dox-inducible Nudt21 transgene could rescue the cell cycle arrest of conditional Nudt21 KO cells when dox was added on day 1, 2, or 3, but not when added on day 4 after Nudt21 deletion (Fig. 4e and f). This result indicates that day 4 Nudt21 KO cells have embarked on an irreversible path towards cell cycle arrest, which manifests at day 6. Importantly, protein levels for the 73 essential candidate genes decreased at day 4 even though associated RNA levels remained unchanged in Nudt21 KO cells (Fig. 4d). Hypomorph cells largely retained RNA and protein expression of these candidates, as expected. We further showed that protein downregulation in Nudt21 KO cells was not due to an impaired translational efficiency of associated RNAs using polysome sequencing (Fig. 4d). These results indicate that acute disruption of the CFIm complex elicits downregulation of candidate proteins through 3’UTR shortening of corresponding mRNAs, but without altering mRNA levels or translational efficiency.
Gene Ontology (GO) analysis of the 73 deregulated proteins revealed several protein subunits of the nuclear pore (e.g., Nup160, Seh1l, Nup107), the mitochondrial ribosome (e.g., Mrpl33, Mrpl 35, Mrpl 37), the nucleolus (e.g., Ddx54, Dhx33, Nifk) and the spliceosome (e.g., Alyref, Srsf1, Hnrnpm) among the downregulated proteins (Fig. 4d right). These results suggest that multiprotein complexes are uniquely sensitive to destabilization upon complete loss of Nudt21 function. By contrast, GO analysis of upregulated proteins revealed pathways associated with “Regulation of protein-containing complex assembly”, “Autophagy” and “Protein folding”, consistent with defects in protein homeostasis (Supplementary Fig. 7a). Indeed, chaperones involved in “protein folding” were strongly upregulated at the protein level in Nudt21 KO cells, without major changes in the 3’UTR lengths of associated mRNAs, consistent with the activation of a cellular stress response caused by imbalanced protein levels, rather than APA (Supplementary Fig. 7b-d).
As the integrated stress response (ISR) pathway is typically activated in contexts of proteotoxic stress, we treated Nudt21 KO cells with the ISR inhibitor ISRIB53,54,55. This led to a subtle yet significant increase in proliferation, suggesting that proteotoxic stress indeed contributes to the observed cell cycle defects of Nudt21 depleted cells (Supplementary Fig. 7e).
To confirm and expand this analysis with an orthogonal approach, we queried our unfiltered DEPs using the EBI Complex Portal database56, which encompasses all annotated protein complexes, and then calculated enrichment scores for each complex in Nudt21 KO cells compared to wild-type controls (Fig. 4g). Strikingly, proteins downregulated in Nudt21 KO cells were assigned to 21 different protein complexes, whereas upregulated proteins were assigned to only 6 different protein complexes, reinforcing the notion that CFIm disruption leads to the preferential downregulation of multiprotein complexes. Top-scoring downregulated complexes included again the 39S mitochondrial large ribosomal subunit, the 40S cytosolic small ribosomal subunit and the nuclear pore, as well as several histone acetyltransferase complexes (TFTC, NSL, NuA4) and the telomerase holoenzyme. Of these, the nuclear pore stood out in that it contained the most subunits (40%) with APA changes compared to the other two top-scoring ribosomal complexes (9% each) in Nudt21 KO cells. Moreover, the integrity of the nuclear pore is critical for the assembly of ribosomes and could thus function upstream of the other top-scoring complexes57. The observed disruption of entire protein complexes despite the 3’UTR shortening of only some of its subunits in Nudt21 KO cells is consistent with cellular protein quality mechanisms, whereby the elimination of individual subunits elicits the proteasome-dependent degradation of remaining, unpaired orphan subunits58. Together, these observations indicate that essential APA targets such as nucleoporins are destabilized specifically in Nudt21 KO cells while they remain intact in Nudt21 hypomorph cells, providing a plausible mechanism for the observed self-renewal defect of Nudt21 KO stem cells.
CFIm stabilizes nuclear pores to facilitate mRNA export and genomic stability
Focusing on the nuclear pore complex, we observed that in total, 14 nucleoporins exhibited 3’UTR shortening with Nudt21 loss, which accounts for almost half of all nucleoporins (n = 30) (Figs. 5a and 5b). A disproportionate number of these 14 nucleoporins belong to the so-called outer ring structure (e.g., Ahctf1, Nup160, Seh1l, Nup107, and Ndc1) required for nuclear pore stability and mRNA export59. Associated proteins were downregulated upon Nudt21 loss, suggesting a major nuclear pore defect (Fig. 5c). To determine whether nuclear pores are dysfunctional in Nudt21-depleted cells, we stained Nudt21 KO, hypomorph and control esophageal stem cell cultures with the nuclear pore antibody mAb414. Indeed, Nudt21 KO cells showed a severe diminution of nuclear pore signal (Figs. 5d and 5e), while hypomorph cells retained control-like signal (Supplementary Fig. 8a and 8b). Supporting a functional defect in nuclear pores, we also detected a marked decrease in cytoplasmic mRNA signal in Nudt21 KO cells using RNA fluorescence in situ hybridization (FISH) analysis, suggesting disrupted APA leads to a pronounced nucleocytoplasmic mRNA export defect (Figs. 5f and 5g). To rule out the possibility that Nudt21 directly interacts with and stabilizes nuclear pores, we performed co-IP/MS for HA-Nudt21 in self-renewing Nudt21 hypomorph cells (Supplementary Fig. 8c). Although we captured the entire CFIm complex including Nudt21, Cpsf6, and Cpsf7, as expected, we detected no nucleoporins in association with Nudt21, suggesting that the RNA-binding function of Nudt21 rather than physical interactions with nucleoporins underlie the defect in nuclear pore stability in Nudt21 KO cells.
a Nucleoporins undergoing APA upon Nudt21 KO. b Example of PAS-seq tracks. c Western blot for nucleoporins upon Nudt21 KO. Asterisk: unspecific band (representative of three independent experiments). d Immunofluorescence with mAb414 (nuclear pores). e Quantification of nuclear signal in (d) (n = 3 biol. replicates). f Poly(A) mRNA FISH. g Quantification of poly(A) foci in (f) (n = 3 biol. replicates). h Setup of pooled nucleoporin-focused 3’UTR CRISPR screen. Dual gRNAs (dgRNAs) flank regions undergoing APA upon Nudt21 KO. Basal stem cells were derived from a dox-inducible Cas9 mouse (iCas9) and cultured without dox (Cas9-off) or with dox (Cas9-on) for three passages. i Quantification of dgRNAs enrichment. j Gene tracks of PAS-seq peaks (top) at the Nup160 locus in control cells, Nudt21 KO and hypomorph cells, revealing 3 different isoforms (short, medium, long). Dashed line highlights region targeted by dual dgRNAs. k Excision efficiency of region highlighted in (j) after culturing basal cells for 4 days in the presence or absence of dox (representative of three independent experiments). l Effect of Nup160 3’UTR deletion on proliferation (n = 3 biol. replicates). m Immunofluorescence staining with mAb414 antibody. n Quantification of nuclear pore signal in (m) (n = 3 biol. replicates). o Poly(A) mRNA FISH. p Quantification of mRNAs detected by poly(A) mRNA FISH shown in (o) (n = 3 biol. replicates). q Immunofluorescence of basal cells undergoing normal cytokinesis (control) or cytokinesis with chromatin bridges (dgNup160L) (representative of three independent experiments). r Effect of various treatments on chromatin bridge formation, with at least 100 cells counted per condition (n = 3 biol. replicates). Statistical analysis: Every dot represents one nucleus. The line represents the mean of biological replicates. Error bars represent the standard deviation of the mean for biological replicates. Statistical significance was determined using a two-tailed unpaired Student’s t-test (e,g,l,n,p, and r).
If nucleoporins are relevant targets of the CFIm complex, we would expect their depletion to phenocopy aspects of Nudt21 loss. We therefore suppressed the nucleoporins Nup107, Nup160 and Seh1l using siRNAs in esophageal stem cells and measured effects on cell cycle progression (Supplementary Fig. 8d-8f). Indeed, suppression of Nup160 and to a lesser extent that of Nup107 and Seh1l led to an S phase arrest and a concomitant loss of nuclear pore staining in esophageal stem cells akin to Nudt21 KO cells. Of note, transient, near-complete suppression of Nudt21 with potent siRNAs also impaired Nup160 expression, but not Nup107 and Seh1l expression, suggesting that Nup160 stability is directly controlled by the CFIm complex. Moreover, Nudt21 deletion as well as nuclear pore depletion by siNup107 or siNup160 displayed an increased frequency of chromatin bridges, consistent with chromosomal instability, a known downstream consequence of nuclear envelope defects60,61 (Supplementary Fig. 8g and h), thus further strengthening the connection between APA and the nuclear pore. Together, these results uncover a previously unknown functional link between APA and the nuclear pore complex by placing specific nucleoporins downstream of CFIm.
Nucleoporin 3’UTRs are essential for nuclear pore stability and function
To assess whether the APA change of any particular nucleoporin(s) in Nudt21 KO cells drives the self-renewal arrest, we conducted a 3’UTR-focused CRISPR/Cas9 screen. Briefly, we lentivirally introduced a pool of 32 dual guide RNAs (dgRNA) targeting the distal (i.e., alternative) 3’UTRs of the 14 candidate nucleoporins undergoing APA in Nudt21 KO cells (see Fig. 5a) into wild-type esophageal stem cells containing a dox-inducible Cas9 transgene. Transduced cultures were propagated for 3 passages in the absence or presence of dox before quantifying dgRNA abundance by dgRNA sequencing (Fig. 5h). Depletion of specific dgRNAs indicates that associated 3’UTR elements are required for the self-renewal of esophageal stem cells akin to the Nudt21 KO phenotype. Interestingly, dgRNAs targeting the distal 3’UTRs of the outer ring nucleoporins Nup160 and Nup98/96 showed the strongest depletion, indicating these non-coding elements are required for continual self-renewal (Fig. 5i).
Considering that dgRNAs targeting Nup160 scored the highest in our self-renewal assay, we focused on this particular protein for the remainder of the study. Closer examination of our PAS-Seq data revealed that the Nup160 gene produces 3 different APA isoforms (Fig. 5j), which we designated Nup160 short (Nup160S), medium (Nup160M) and long (Nup160L). In asynchronously dividing esophageal stem cells, Nup160M was the dominant isoform while Nup160S and Nup160L were minor isoforms. To determine whether the low expression levels of Nup160L are due to cell cycle dependent effects, we purified esophageal basal cells at the G1, S and G2/M phases of the cell cycle and performed Nup160 isoform-specific qPCR analysis. Indeed, Nup160L levels transiently increased during the S and G2/M phases (Supplementary Fig. 9a and 9b), pointing to a cell cycle-dependent regulation of APA62. Of relevance, the dynamic expression of the Nup160L isoform coincides with the upregulation of nucleoporins and the assembly of nuclear pores during the late S and G2/M phases of the cell cycle (Supplementary Fig. 9c-9f), suggesting a possible regulatory role of Nup160L in nuclear pore homeostasis.
Nudt21 deletion led to a slight increase in Nup160S and a decrease in Nup160M expression but a complete loss of Nup160L expression, which was the isoform that scored in our pooled CRISPR/Cas9 screen (Fig. 5j). Accordingly, deletion of the distal-most 3’UTR exclusively associated with Nup160L with distinct dgRNA pairs led to a pronounced defect in S phase progression, nuclear pore signal and mRNA export (Figs. 5j-5p). By contrast, dgRNAs targeting the centrally located 3’UTR associated with Nup160M yielded cells with only a mild defect in cell cycle progression and nuclear pore signal intensity, suggesting this region is largely dispensable for cell proliferation (Supplementary Fig. 9g-9k). Similar to Nudt21 KO cells, deletion of the 3’UTR associated with the Nup160L isoform also resulted in the downregulation of the nuclear pore outer ring proteins Nup160, Nup107, and Seh1L, consistent with a destabilization of the entire outer ring structure (Supplementary Fig. 9l). We confirmed the crucial role of the Nup160L isoform in maintaining cell cycle progression and nuclear pore integrity using isoform-specific siRNAs (Supplementary Fig. 9m-9r). Interestingly, depletion of the Nup160L isoform via either CRISPR or siRNA approaches was also sufficient to elicit chromatin bridges typically observed in Nudt21 KO cells (Figs. 5q and 5r). However, deletion of the Nup160M 3’UTR did not cause chromatin bridges, underscoring the specific role of the long isoform in maintaining the self-renewal of esophageal stem cells by preserving nuclear and genomic integrity61,10. Notably, loss of Nudt21 in quiescent satellite cells did not diminish nuclear pore signal intensity, suggesting that dividing cells are uniquely sensitive to nuclear pore loss upon Nudt21 KO (Supplementary Fig. 9s and 9t). Together, these results establish that the outer ring nucleoporin Nup160 is a direct and crucial target of the CFIm complex responsible for maintaining nuclear pore integrity and consequently genomic integrity and stem/progenitor cell maintenance via regulation of its 3’UTR.
Finally, we tested whether the re-expression of specific Nup160 mRNA isoforms is sufficient to rescue the cell cycle defect of Nudt21 KO cells. Briefly, we introduced in vitro transcribed (IVT) RNAs encoding Nup160S, Nup160M and Nup160L into wild-type and Nudt21 KO cells. None of these IVT isoforms was able to rescue the cell cycle phenotype, and exogenous Nup160L RNA even exacerbated the Nudt21 KO-dependent cell cycle defect. This finding implies that Nup160 isoform levels need to be tightly regulated in cells, with aberrant up- or downregulation disrupting Nup160 function in dividing cells (Supplementary Fig. 10a and 10b). We tested the role of these 3’UTR isoforms in protein localization by leveraging chimeric fusion transcripts between Nup160 or eGFP coding regions and the three Nup160 3’UTR variants (Supplementary Fig. 10c). Notably, eGFP-Nup160L mRNAs transfected into esophageal stem cells yielded lower eGFP signal compared to eGFP-Nup160S or eGFP-Nup160M mRNAs 24 hours post transfection, pointing to a possible role of these RNA isoforms in the translational control of Nup160 (Supplementary Fig. 10d-g). In agreement with this model, we discovered that a biotinylated Nup160L mRNA probe associates with ribosomal proteins more efficiently than biotinylated Nup160M and Nup160S mRNA probes (Supplementary Fig. 11a-c). Of note, transfection of cells with the eGFP-Nup160L mRNA isoform but not the eGFP-Nup160M or eGFP-Nup160S mRNA isoforms triggered the formation of ectopic peri-nuclear foci 2-3 hours after transfection, which is reminiscent of condensates previously associated with nuclear pore homeostasis63,64 and warrants further investigation (Supplementary Fig. 11d and 10e).
APA impacts co-translational protein complex assembly
The above-mentioned observations raised the possibility that global 3′UTR shortening might affect the assembly of protein complexes in Nudt21 KO cells. Large protein complexes tend to assemble via co-translational mechanisms whereby a nascent polypeptide encoding for one subunit interacts with another nascent polypeptide (co-co-translational assembly) or with a fully folded polypeptide (co-post-translational assembly) encoding for a different subunit65,66.To determine whether disruption of APA patterns alters the co-co-translational mode of protein complex assembly, we performed Disome Selective Profiling (DiSP)65,66 of ribosomes using control and Nudt21 KO cells. Of 4,986 mRNAs actively translated in control cells, 143 mRNAs (3%) met the criteria for co-co-translational assembly, which is in line with previous observations using transformed cell lines65 (Fig. 6a). Strikingly, nearly 44% (n = 63) of these interactions were reduced in Nudt21 KO cells and about half of the associated mRNAs were Nudt21 targets, suggesting APA plays a crucial role in their co-translational regulation (Fig. 6b and c). Supporting the aforementioned role of Nudt21 in cell cycle progression, we identified pivotal regulators of mitosis among the most significantly altered mRNAs (Fig. 6d), including the Nudt21 targets Cenpf and Smc5 involved in chromosome segregation (Fig. 6e). We note that Nup160 itself did not score in our DiSP assay, which is likely due to an alternative co-translational or post-translational mode of assembly67,68. Collectively, these results suggest that Nudt21-dependent APA dynamics coordinate mitotic progression and protein complex homeostasis, processes that are crucial for maintaining the proliferative state and regenerative potential of several adult tissues.
a Rank-ordered disome-selective profiling (DiSP) co-co-assembly scores in control cells. 143/4,843 translated mRNAs meet the criterion for co-co-translational assembly. b Scatterplot of co-co-assembly score changes (Nudt21 KO vs control). The orange quadrant indicates co-co-translational assemblies impacted by Nudt21 KO. c Dot plot showing all factors from the orange quadrant in (b), indicating whether they are Nudt21 targets based on PAS-seq analysis. d Gene Ontology (GO) enrichment of the 63 disrupted mRNAs in (b), highlighting cell cycle relevant terms. e Heatmap of co-co-assembly scores for representative nuclear envelope-associated factors and their functions. Nudt21 targets are shown in bold font. The data represent two independent experiments.
Discussion
Here, we demonstrate that APA is essential for the survival of adult mice by maintaining the homeostasis of diverse regenerative tissues undergoing continuous turnover or injury-induced cell replacement. This finding was unexpected considering previous reports, which concluded that suppression of the CFIm complex using siRNAs promotes cell growth and tumorigenesis6,26,42,69,70,71,72. Our data help to reconcile these seemingly disparate conclusions by showing that APA exerts dose-dependent effects on stem cell homeostasis, affecting either self-renewal or differentiation, or both, depending on the extent to which CFIm/Nudt21 levels are modulated.
Specifically, our data indicate that hypomorph levels of Nudt21 (~25% of wilde-type levels) are compatible with stem cell self-renewal and lead to the 3’UTR shortening of ~1,000 transcripts (Fig. 7a). These transcripts encode for genes with roles in chromatin regulation and developmental processes and undergo aberrant upregulation at the protein level due to loss of miRNA target sequences, which then elicits a differentiation arrest in both Nudt21 hypomorph and KO stem cells (Fig. 7b). This mechanism is consistent with, and significantly extends, previous Nudt21 siRNA-based in vitro studies by our lab and others, highlighting the important role for APA in guiding cell fate transitions across diverse cellular contexts18. Our results also support the notion that APA modulates larger ceRNA networks42,73,74,75 to further solidify stem cell and differentiation-associated regulatory programs.
a Nudt21/CFIm acts in a dose-dependent manner to modulate APA in stem cells. High Nudt21/CFIm levels (wild-type) promote the formation of long 3’UTRs, while intermediate levels (Nudt21 hypomorph) result in the shortening of approximately 1,000 mRNAs. Complete loss of CFIm/Nudt21 leads to more extensive shortening, impacting around 2,000 3’UTRs in total. b Nudt21 hypomorphic and KO stem cells display 3’UTR shortening of mRNAs associated with development and chromatin regulation, resulting in their escape from miRNA-mediated repression and a subsequent arrest in differentiation. c Nudt21 KO stem cells, but not Nudt21 hypomorph stem cells, exhibit additional 3’UTR shortening of mRNAs encoding for subunits of essential multiprotein complexes, leading to arrested self-renewal.
By contrast, severe reduction or complete loss of Nudt21 levels (<25% of WT levels) is incompatible with the continuous self-renewal of stem cells and leads to more accentuated APA changes of transcripts than in Nudt21 hypomorphs. Moreover, complete Nudt21 loss elicits 3’UTR shortening of an additional ~1,000 transcripts enriched for subunits of multiprotein complexes with essential roles in cell proliferation, genome integrity and cell cycle progression, processes that are key for continuous stem cell self-renewal and progenitor cell proliferation (Fig. 7c). In contrast to miRNA-dependent targets undergoing APA, which are typically upregulated with Nudt21 deletion, these essential APA targets are less enriched for miRNA binding sites and tend to be downregulated at the protein level when their 3’UTRs are shortened. Consequently, affected protein complexes disassemble, resulting in DNA damage and cell cycle arrest. These observations cannot be explained by the classical model(s) of APA-dependent gene regulation and thus point to an expanded role for APA in the coordinated regulation of multiprotein complexes essential for cell cycle progression.
The most striking example following the latter mechanism is the nuclear pore, which, among all deregulated multiprotein complexes, contains the largest number of subunits undergoing APA in Nudt21 KO cells and plays an essential role in proliferative cells by coordinating nucleo-cytoplasmic mRNA transport, chromatin organization and genome integrity76,77,78,79,80,81. Indeed, deletion of the Nup160 mRNA isoform with the longest 3’UTR, which represents only a minor isoform in asynchronously dividing basal cells, disrupts stem cell self-renewal and nucleocytoplasmic mRNA transport, and leads to DNA damage. Of relevance, nucleoporins and Nudt21 independently scored in previous loss-of-function screens for modulators of DNA damage response, lending further support to the molecular and functional link between CFIm and the nuclear pore complex identified in this study82,83. To our knowledge, these results represent the first demonstration that the stability of multiprotein complexes such as the nuclear pore are regulated downstream of the CFIm complex via APA modulation of individual subunits. This finding not only contributes to our fundamental understanding of how the nuclear pore complex is regulated but should also inform the study of physiological and pathological contexts with altered nuclear pore function such as aging76,78,84.
Our results significantly build on prior studies in transformed cell lines showing that APA of individual mRNAs increase protein synthesis due to the elimination of miRNA binding sites18 or endow cells with new protein functions by facilitating the formation of additional protein complexes10. It has remained unclear from this prior work whether APA also controls protein complex stability in a physiologically relevant context in addition to directing the formation of alternative complexes in cancer cell lines. Based on our results, we propose that CFIm plays a central role in coordinating the integrity of diverse multiprotein complexes essential for cell division including but not limited to the nuclear pore, spliceosome, and the nucleolus. This model explains why stem cells have elevated Nudt21 levels compared to differentiated progeny, with Nudt21 enhancing the stability of essential protein complexes critical for self-renewal, while simultaneously regulating differentiation-related programs via the classical miRNA-dependent pathway.
Beyond tissue homeostasis, our study has implications for understanding the role of APA in cancer. Our finding that intermediate Nudt21 levels inhibit stem/progenitor cell differentiation while permitting proliferation may help to explain why certain cancers with poor prognosis and increased metastatic potential have reduced CFIm levels42,72,85,86,87,88,89. CFIm downregulation in such cancer contexts may suppress lineage-specific programs, akin to our observations in basal stem cells, and lock cancer cells into a less differentiated state required for aberrant proliferation and metastasis. Similar to our results in normal epithelial cells, we predict that malignant cells with CFIm downregulation are hypersensitive to genetic or pharmacological inhibition of CFIm, which should lead to the disintegration of essential multiprotein complexes and cell cycle arrest. It should thus be valuable to develop small molecule or peptide inhibitors of the CFIm complex to test this concept in the future.
Finally, a key mechanistic question raised by our model is how APA controls the stability of protein complexes in dividing cells. One possibility is that the 3’UTRs of specific subunits facilitate the co-translational assembly of entire protein complexes. In support of this hypothesis, many multiprotein complexes including the nuclear pore are regulated by co-translational assembly mechanisms65,90,91,92, and we detected enriched binding of ribosomal proteins to the long Nup160 isoform. Indeed, Nudt21 loss reduced nearly half of all co-co-translational assembly events and impacted key factors associated with cell division. As the DiSP assay we employed here cannot capture co-post-translational assembly events including nascent nuclear pores65,67,68, it will be important to assess whether Nudt21 depletion affects not only co-co-translational but also co-post-translational assembly events. Similarly, it should be informative to determine whether global APA rewiring via Nudt21 loss also impacts the biogenesis of proteins with intrinsically disordered domains recently linked to 3’UTR-dependent assembly mechanisms65,67,68,90,91,92. In summary, our study reveals a crucial and previously unrecognized role of APA in maintaining stem/progenitor cell and tissue homeostasis and highlights the indispensable function of CFIm in coordinating the integrity of essential protein complexes as exemplified by the nuclear pore.
Methods
Inducible HA-Nudt21 lentiviral constructs
To construct an inducible HA-Nudt21 lentiviral vector, we synthesized an N-terminally tagged HA-Nudt21 gBlock (IDT) and cloned it into a BamHI and EcoRI-linearized pCW57 (#78933) vector using Gibson Assembly (NEB). This method allowed for the insertion of HA-Nudt21 downstream of the TRE promoter, resulting in the deletion of Turbo RFP. Lentiviral particles were produced as previously described93.
Mice
The following mice strains were purchased from The Jackson Laboratory (jax.org): K5CreER (Jax 029155), Lgr5-EGFP-IRES-CreER (Jax 008875), Rosa26-CreER (Jax 008463), KH2-iCas9(Jax 029415) Nudt21fl/fl (this study, see below). Conditional Nudt21 KO mice were generated by crossing Nudt21fl/fl with one of the following Cre allele bearing mice: Rosa26CreER (ubiquitous KO), K5CreER (squameous epithelia KO) and Pax7-CreER (satellite cell KO). Control mice were either Nudt21 wt/fl and the corresponding CreER or Nudt21fl/fl without any CreER. For experiments, we administered tamoxifen into adult mice between 3 and 6 months of age. Both male and female mice (approximately equal proportions) were used in all experiments. No sex-specific differences were observed or analyzed, and no sex-related effects were expected for the biological processes under investigation. Mice were housed in ventilated cages on a standard 12 h:12 h light cycle. All procedures involving mice adhered to the guidelines of the approved Massachusetts General Hospital Institutional Animal Care and Use Committee (IACUC) protocol no. 2006N000104.
To generate bone marrow chimeric mice through bone marrow transplantation, 6-8 week-old B6.SJL (CD45.1) recipient mice were irradiated with a total dose of 1,200 rad (600 rad twice, with a 3-hour interval). Bone marrow was harvested from uninduced donor mice (Nudt21fl/fl; Rosa26CreER (KO) or Nudt21fl/fl (control)). Each recipient received 2,000,000 bone marrow cells from the donor mouse, injected retro-orbitally in 100 μl of FBS-free PBS. The mice were then housed for 10 weeks to ensure complete engraftment before tamoxifen injection). For competitive bone marrow transplantation, 6-8 week-old B6.SJL (CD45.1) recipient mice were irradiated with a total dose of 1,200 rad (600 rad twice, with a 3-hour interval). Bone marrow was harvested from uninduced donor mice (Nudt21fl/fl; Rosa26CreER or Nudt21 fl/wt; Rosa26CreER, or Nudt21fl/fl) and a competitor mouse (B6.SJL (CD45.1)). Each recipient received 1,000,000 bone marrow cells from the donor mouse and 1,000,000 cells from the competitor mouse, or the specified number of HSCs from the donor mouse and 500,000 bone marrow cells from the competitor mouse. The cells were injected retro-orbitally in 100 μl of PBS. The mice were then housed for 8 weeks to ensure complete engraftment. Control mice were either Nudt21 wt/fl with the corresponding CreER or Nudt21fl/fl without any CreER. Both KO and control groups were treated with tamoxifen in the same manner.
ES cell culture and Nudt21 gene targeting
Mouse embryonic stem cells (Nudt21fl/fl, HEPD0770_7_B01 from IMPC) were grown on irradiated mouse embryonic fibroblasts in a growth medium consisting of KO-DMEM (Life Technologies) with 15% fetal bovine serum (FBS) (Hyclone), 1X L-glutamine (Life Technologies), 100 U/mL Penicillin, 100 μg/mL Streptomycin (Life Technologies), 1X MEM Non-Essential Amino Acids Solution (Life Technologies), and 50 μM β-mercaptoethanol (Life Technologies). For selection casette removal, cells were grown to 80% confluence, then trypsinized and collected. The cells were washed once in PBS and resuspended in 500 μL of PBS. Subsequently, 40 μg of pCAGGS-flpE-puro were added, and the mixture was transferred to a 4 mm electroporation cuvette (Biorad). Electroporation was performed using a Gene Pulser Xcell system (Biorad) in time constant mode with the parameters set to voltage=800 V and time constant=0.2 ms. The cells were then plated at clonal density onto DR4 irradiated mouse embryonic fibroblasts (GlobalStem) in a 6 cm dish and cultured in ES cell medium for 24 hours. Afterward, the medium was replaced with growth medium containing 200 μg/mL hygromycin (Gibco) to select for targeted clones. Individual colonies were picked, expanded, and tested via PCR to confirm proper integration.
Blastocyst Injection and generation of conditional Nudt21 KO mice
To create transgenic mice, Nudt21fl/fl ES cells were injected into E3.5 blastocysts as described in previous studies52,53. The resulting high-grade chimeras from Nudt21fl/fl ES cells were bred with B6/C57 wildtype mice, and the offspring were genotyped to confirm germline transmission. These mice were then crossed with mice carrying Rosa26CreER or K5-CreER on a B6/C57 background. Mice of both genders, aged 8-16 weeks, were used in the study and induced, unless otherwise specified, with 5 daily doses of 20 mg/ml tamoxifen in corn oil. The mice in this study were housed and bred in Specific Pathogen Free (SPF) rooms at the AAALAC-accredited Center for Comparative Medicine vivarium at Massachusetts General Hospital. They were kept in ventilated cages with a standard 12:12 light cycle. All procedures involving mice followed the guidelines of the approved Massachusetts General Hospital Institutional Animal Care and Use Committee (IACUC) protocol no. 2006N000104.
Tamoxifen treatment
For in vivo tamoxifen treatment, both KO and control mouse were treated with tamoxifen. Tamoxifen (20 mg/ml, T5648-1G, Sigma-Aldrich) was prepared in corn oil (Sigma-Aldrich), and each mouse received a daily intraperitoneal injection of 100 µl for five consecutive days. For the satellite cell experiments, the mice received the same five doses of tamoxifen and were subsequently maintained on a diet containing 500 mg/kg tamoxifen (Envigo) for 30-40 days. For in vitro tamoxifen treatment, cells were derived from conditional Nudt21 KO or control mice and cultured under specific conditions (detailed below). The media contained either ethanol (ETOH) as a control or (Z)−4-Hydroxytamoxifen (4OHT, Sigma-Aldrich, H7904-5MG) to induce Nudt21 KO. The 4OHT stock solution was prepared by dissolving 4OHT powder in ETOH to a concentration of 5 mg/ml (13 mM). To test for Cre toxicity94, new 4OHT batches were titrated on Nudt21 wt/wt; Rosa26CreER esophageal basal cells, skin keratinocytes, intestinal organoids, and hematopoietic progenitors. The effect of 4OHT on the cell cycle was assessed by EdU staining and FACS. The maximum tolerable concentration of 4OHT without impacting the cell cycle was approximately 0.1-0.05 µM for all cell types. This concentration was used throughout all in vitro experiments, with 4OHT added at each media change. The 4OHT stock solution remained stable for up to 3 months.
Histology
Mice or tissues were fixed in Bouin’s solution (Sigma-Aldrich) or 10% formalin solution (Sigma-Aldrich), respectively. The fixed tissues were then prepared for IF, immunohistochemistry and H&E staining.
Immunofluorescence (tissues)
Immunohistochemistry staining was performed by iHisto Inc. (ihisto.io) Samples were processed, embedded in paraffin, and sectioned at 4 μm. Paraffin sections were then deparaffinized and hydrated using the following steps: 15 min in xylene twice; 5, 5, and 5 min in 100%, 100%, and 75% ethanol, respectively; and 5 min in PBS at RT repeated three times. Antigen retrieval was achieved by boiling the sections in Citrate-based unmasking buffer for 115 min in pressure cooker and 20 min of cooling at RT. Slides were washed and blocked with bovine serum albumin (BSA) at RT for 20 min. Following blocking, sections were incubated with the primary antibodies diluted in BSA overnight at 4 °C. The following day, slides were washed as above. Secondary antibodies were applied to sections and allowed to incubate for 1 h at RT. Slides were washed as above. Slides were stained with Sudan Black for 15 min to quench background and autofluorescence. Slides were washed under running water for 15 min. Slides were washed three times for 5 min each in TBS, TBS, and TBST, respectively. Slides were counterstained with DAPI and coverslipped using Fluoroshield (SIGMA).
Primary antibodies: goat anti-GFP(abcam, ab6673)1:500, Nudt21 (Santa Cruz, 2203C3) 1:50, rabbit anti-Krt14 1:1000 (biolegend, 905304). Secondary antibodies (all from Thermofisher): donkey anti-goat Alexa flour 488, Donkey anti-mouse AF555, Goat anti, mouse Alexa flour 488, Goat anti-rabbit AF555
Immunohistochemistry
Immunohistochemistry (IHC) staining was performed by iHisto Inc. (ihisto.io) Samples were processed, embedded in paraffin, and sectioned at 4 μm. Paraffin sections were then deparaffinized and hydrated using the following steps: 15 min in xylene twice; 5, 5, and 5 min in 100%, 100%, and 75% ethanol, respectively; and 5 min in PBS at RT repeated three times. Antigen retrieval was achieved by boiling the sections in Citrate-based unmasking buffer for 115 min in pressure cooker and 20 min of cooling at RT. A peroxidase blocking solution was applied to sections and allowed to incubate for 10 min and 2.5% normal horse serum for 30 min. After 30 min, excess serum was removed from the sections and primary antibody diluted in 2.5% normal horse serum was applied. Sections were incubated with diluted primary antibody overnight at 4 °C. Sections were incubated in secondary antibody for one hour at RT. Slides were washed as before. The DAB working solution was applied to sections and the stain was allowed to develop. Slides were counterstained with hematoxylin. Slides were dehydrated in an oven set to 37 °C and coverslipped using a xylene-based mounting medium. Primary Antibodies: mouse anti-Brdu(GB12031), (Santa Cruz, 2203C3) 1:50.
Secondary antibody: HRP Horse Anti-Mouse IgG Polymer Kit (ImmPRESS), Peroxidase (MP-7402, Vector Laboratories).
H&E staining
H&E staining was performed by iHisto Inc. (ihisto.io). Samples were processed, embedded in paraffin, and sectioned at 4 μm. Paraffin sections were then deparaffinized and hydrated using the following steps: 15 min in xylene twice; 5, 5, and 5 min in 100%, 100%, and 75% ethanol, respectively; and 5 min in PBS at RT repeated three times. After deparaffinization, 4‑µm‑sectioned samples were placed on glass slides and stained with hematoxylin for 3 min and Eosin for 5 min. The slides were dehydrated by graded alcohol and coverslipped using a xylene-based mounting medium.
Excision PCR
To quantify knockout efficiency, cells were treated as described in the results section and genomic DNA was isolated using the Monarch Genomic DNA Purification Kit (NEB). PCR amplification was performed with the GoTaq Master Mix (Promega) by combining 5 µM primers with 10–50 ng of genomic DNA. The primers used for the reactions were as follows:
Nudt21_1: 5’-ACCAGTGAGGTGGCTCTAAGG-3’
Nudt21_2: 5’-AGGTCCCTGAAATGCTTAGCC-3’
Nudt21_3: 5’-AGGATCCAAGTTTTCAGATATTCTCCCC-3’
Nup160_long_1: 5’-AAACTTACGGCTCTCCACTACAGCATAAAAGGC-3’
Nup160_long_2: 5’-GTACCTGTATGTATGTGGCGGCGAGAGG-3’
Nup160_med_1: 5’-ACAGAAACCCTGTCTCAAGGC-3’
Nup160_med_2: 5’-GATTGAGGGCCTTGGATCTGCT-3’
The PCR program consisted of an initial denaturation step at 98 °C for 2 min, followed by 35 cycles of denaturation at 98 °C for 15 s, annealing at 58 °C for 20 s, and extension at 72 °C for 5 min. A final extension was carried out at 72 °C for 10 min. The PCR products were then electrophoresed alongside a 1 kb DNA Ladder (NEB) on a 1% agarose gel.
2D epithelial cell culture
long-term expansion of diverse epithelial basal cells was conducted as previously described95. Briefly, adult esophagi or tails were removed from mice, cleaned in ice-cold PBS, and rocked for no more than 10 h at 4 °C in dissociation medium composed of DMEM/F12 (Gibco), 200 μg/ml Pronase (Sigma-Aldrich), 10 μg/ml DNase I (Worthington), 1% Fraction V BSA (Gibco), 1× pen/strep, and 10 μM Y-27632 (Tocris). The epithelium from the esophagi or tail was dissociated from the dermis using angled forceps (FST 00276-13). The epidermis was cut into ~2–5 mm pieces, suspended in cold PBS, pipetted up and down, and passed through a 100 μm mesh placed on a 50 ml conical tube. The epithelium on the mesh was ground with a syringe plunger, and PBS was flushed through the mesh to recover epithelial cells. Dissociated epithelial cells were explanted onto T25 flasks pre-coated with conditioned medium collected from 804G cells. Complete SAGM medium, supplemented with 10 μM Y-27632 (Tocris), 1 μM CHIR99021 (Tocris), 1 μM A83-01, and 1 μM DMH-1, was used for expansion. Cells were kept in the exponential growth phase (maximum 8000 cells/cm²). Media changes were performed every 48 h. Cells were dissociated from the flasks and dishes either by scraping or treatment with TrypLE (Gibco) for 10 min. Epithelial cells were collected in 1% BSA/PBS (Invitrogen) and spun down at 300 × g. Cells were kept in culture for a maximum of 6 passages.
Intestinal organoid culture
Intestinal crypts were derived from Nudt2fl/fl; R26-CreER (mice and cultured as described before96.
Esophageal cell differentiation
Esophageal epithelial cells were isolated as described above. For experiments including the Nudt21 hypomorph, cNudt21 KO cells were transduced with the inducible HA-Nudt21 virus, selected with blasticidn and pre-treated for 3 days with low dose of of DOX(0.2 µg/ml). For the differentiation assay, cNudt21 KO cells were seeded at 8e3/cm2, and cultured for 4 days in self-renewal media containing either ETOH (Control) or 4OHT (KO) or 4OHT + DOX (Hypomoprh) until a confluency of 8e3/cm2 was reached at day 4. The media was changed every two days. Cells were collected at day 4 for the self-renewal timepoint before initiation of differentiation. Differentiation was induced by first washing the cells with PBS and then changing to CnT-PR-D media (Fisher Scientific) containing 1.8 mM CaCl2. Cells were collected for IF, RNA and protein at days 3 and 6.
Western blot assay
Cells grown in culture were lysed in 200–500 μl of RIPA buffer (Thermo Scientific) supplemented with Complete Mini Protease Inhibitors (Roche), PhosSTOP (Roche), and 0.01 U/µl benzonase (Sigma-Aldrich). The lysates were then incubated on ice for 10 min and sonicated 5 times for 30 s with a 30 s pause in between pulses. The lysates were then cleared for 5 min at maximum speed in a bench top centrifuge to remove cell debris. The supernatant was boiled together with Laemmli sample buffer (Biorad) and loaded into 5–20% Mini-PROTEAN TGX Precast Protein Gels (Biorad). Protein was transferred to nitrocellulose membranes (Biorad) and blocked for 1 h in 5% powdered milk in TBST (Santa Cruz). Unless otherwise specified, antibodies were used following the manufacturer’s recommendations. Primary antibodies used were: Nudt21 (Santa Cruz, sc-81109, 1:100), β-Actin (Santa Cruz, sc-47778, 1:1000), Cpsf7 (Santa Cruz, sc-393880), Cpsf6 (Bethyl, 50-156-0173), Krt13 (abcam, ab92551), P63 (abcam, ab53039), Seh1l (abcam, ab218531), Nup107 (Invitrogen, PA5-20562), Nup133 (abcam, ab155990), Nup155 (Proteintech, 66359-1-1 g), Nup96 (Bethyl, A301-784A), Nup35 (abcam, ab169162), Gapdh (Fisher Scientific, NC0330823), Phospho-ATR (Ser428, Cell Signaling, 2853), ATR (C-1, sc-515173, Santa Cruz). The secondary antibodies used were: goat, anti-rabbit-HRP-conjugated (Thermo Fisher Scientific 31460, 1:2000 dilution) or anti-mouse-HRP-conjugated (Thermo Fisher Scientific 31430, 1:2000 dilution). Proteins were detected using Clarity Western ECL Substrate (Biorad). Images were taken using a ChemiDox Imaging System with enhanced chemiluminescence detection.
Cell cycle and viability assay
Cells were cultured as described above, with specific attention to preventing over-confluence. For the incorporation of EdU in proliferating cells, 1 µM EdU was added to the media and incubated for 1 h in the tissue cell incubator. Epithelial cells were harvested using TrypLE (Gibco), and intestinal organoids were harvested using Trypsin (Thermo Fisher Scientific). Both cell types were fixed and stained according to the manufacturer’s recommendations of the Click-iT Edu-488 kit (Thermo Fisher Scientific). To assess the viability of epithelial cells, they were harvested with TrypLE (Gibco), incubated with 1× Annexin V Binding Buffer (Biolegend, 556454), and stained with FITC-conjugated anti-Annexin V (Biolegend, 640905) and propidium iodide (Thermo Fisher Scientific, BMS500PI) following the manufacturer’s instructions. The gating strategy for EdU-based cell-cycle analysis is shown in Supplementary Fig. 12.
siRNA transfection of epithelial cells
Epithelial cells were cultured as described above. For transfection, cells were seeded at 2e3/cm2 and cultured for one day. On the day of transfection medium was exchanged with fresh pre-warmed medium. The transfection mix was prepared by diluting 30 µl siRNA (20 µM) with 1.1 ml Genmute transfection buffer (SignaGen), followed by brief vortexing and addition of 20 µl Genmute transfection reagent (SignaGen). The mixture was incubated at RT for 15 min and then dropwise added to the cells (4 µl/cm2). For example, 228 µl transfection mixture was added to 10 cm dish containing 10 ml media. Cells were harvested for analysis after three days. All siRNAs were reconstituted in 1× siRNA Buffer (Horizon Discovery) at 20 µM.
ON-TARGETplus Mouse Jmjd1c (108829, Horizon Discovery)
ON-TARGETplus Mouse Sec24a (77371, Horizon Discovery)
ON-TARGETplus Mouse Nup160 (59015, Horizon Discovery)
ON-TARGETplus Mouse Nup107 (103468, Horizon Discovery)
ON-TARGETplus Mouse Seh1l (72124, Horizon Discovery)
siGENOME Non-Targeting siRNA (D-001210-01-05, Horizon Discovery)
siNudt21 Nudt21 (EMU094771, Sigma-Aldrich)
The following siRNAs were purchased as Custom siRNA Standard from Horizon Discovery and pooled at 20 µM for each siRNA.
Nup160_distal_pool:
distal _Nup160_1: AAGCATTTCCTGTAAATCCTA
distal _Nup160_2: AAGAACCGGAGTACGAGTCCT
distal_Nup160_3: AAGGTTCTGTTAGGCTAGGAG
Nup160_Short_pool:
med_isof_Nup160_1: AAGACATCCTTTATCTGCAAA
med_isof _Nup160_2: AACTACCGAAGACATCCTTTA
Immunofluorescence assay (tissue culture cells)
Satellite cells were plated without coating on chamber slides (µ-Slide 8 Well, 80806, ibidi). For epithelial cells, a 30-min 8O4G coating was performed prior to plating. Cells were cultured in their specific media and treated as described in the results section. For immunofluorescence, cells were fixed inside the chamber slides with Paraformaldehyde Solution (PFA), 4% in PBS (AAJ19943K2, Fisher Scientific) for 20 min, followed by three washes with PBS and subsequent permeabilization with 0.5% Triton X-100/PBS (X100-500ML, Sigma-Aldrich) for 5 min. After permeabilization, the slides were washed three times with PBS and blocked for 1 h with 1% BSA/PBS. Primary antibodies were diluted in 1% BSA/PBS, added to the slides, and incubated overnight at 4 °C, followed by three washes with PBS. Anti-Nudt21 (sc-81109, Santa Cruz) was diluted 1:100, mAb414 (902901, Biolegend) was diluted 1:1000, Anti-Cytokeratin 13 (ab92551, abcam) was diluted 1:50,000, Trp63/p63 (ab53039, abcam) was diluted 1:1000. Staining with secondary antibodies and DAPI (BD Biosciences, 564907) was performed for 1 h at RT, followed by three washes with PBS. The staining solution contained isoform and species-specific antibodies coupled to Alexa-Fluor-488 (A32766, Thermo Fisher Scientific) and DAPI (both diluted 1:1000 in 1% BSA/PBS). Cells were imaged on a Leica DMi8 Fluorescent microscope (Objective: N PLAN L 40x/0.55, CORR PH2, 11506298).
Nuclear IF signal quantification
Nuclear signal was quantified using Fiji Macro code97. First, a Gaussian filter was applied to the DAPI channel using the function run(“Gaussian Blur…”, “sigma=1”). A mask for nuclei was then created with the following code: setAutoThreshold(“Default dark”); run(“Threshold…”); setThreshold(45, 255); run(“Convert to Mask”). Subsequently, a region of interest (ROI) for each nucleus was generated with the code: run(“Analyze Particles…”, “size=0.1-100 show=Masks display exclude clear include summarize record add in_situ”); run(“Create Selection”). Finally, the desired channel was selected and quantified using roiManager(“multi-measure measure_all”).
RNA FISH assay
Cells were seeded in 8-well chamber slides as outlined in the “Immunofluorescence” section. Treatment conditions are detailed in the “Results” section. For fixation, cells were treated with 4% paraformaldehyde (PFA) for 20 min at RT and subsequently washed three times with phosphate-buffered saline (PBS). For dehydration, cells underwent a graded ethanol (ETOH) series. Cells were incubated in increasing concentrations of ethanol for 1 min each at 30%, 70%, and finally 100%. After the final step, cells were stored in 100% ethanol at 20 °C for up to two weeks to ensure preservation. Rehydration was performed by reversing the ethanol concentrations in the same sequence, with each step lasting 1 min, concluding with a PBS wash. Cells were prepared for FISH using the RNAscope™ HiPlex v2 Assay according to the manufacturer’s protocol. The specific probe used was the RNAscope® Probe - PolyA-C3, targeting the poly(A) mRNA tail. For quantification of the poly(A) signal, images were analyzed using a custom Fiji Macro. Initially, the poly(A) channel was duplicated and a Gaussian filter (sigma = 2) was applied to enhance feature visibility. The image was then processed to create a binary mask: automated thresholding (Percentile dark), hole filling, watershed algorithm application, and particle analysis (size range 80–1800 square pixels) were performed. For individual cell analysis, selections were created for each cell in an image using the “Create Selection” function. The unaltered poly(A) channel of each ROI was duplicated and saved as a separate TIFF file. Each image file was subsequently opened, and a nuclear mask was applied as detailed in the “Nuclear Signal Quantification” section. Foci quantification within the nucleus and the whole cell was accomplished using the “Find Maxima…” function with a prominence setting of 100. The number of foci detected in the cytoplasm was calculated by subtracting the number of nuclear foci from the total cell detected foci. The notably strong signal within the nucleus suggested active transcription sites, while the cytoplasmic signal predominantly consisted of distinct, clearly visible foci. Cells were imaged on a Leica DMi8 Fluorescent microscope (Objective: N PLAN L 40x/0.55, CORR PH2, 11506298).
3’UTR focused CRISPR screening
Guide RNAs were designed to delete the genomic locus within the 3’UTR that undergoes APA, mimicking the 3’UTR shortening observed in Nudt21 KO. These dgRNA libraries were cloned into a lentiviral plasmid by VectorBuilder (https://en.vectorbuilder.com/vector/Lib240216-1778swe.html). The plasmid harbored two guide RNAs, cloned directly after either the hU6 or macU6 promoter plus scaffold. For selection the plasmid contained and hPGK promotor driving mCherry:T2A:Bsd expression. Lentiviral particles were produced as previously described93. Esophageal basal cells were isolated from the iCas9 mouse and transduced with lentiviruses containing dgRNA library at a multiplicity of infection (MOI) of less than 0.1, as assessed by FACS for mCherry. Subsequently, the transduced cells were split into two 15 cm plates: one containing blasticidin (InvivoGen) and dox (treatment), and the other without dox (control). The cells were passaged twice, and genomic DNA was isolated using the Genomic DNA Purification Kit (NEB). The dgRNA sequences were PCR amplified with the primers actatcatatgcttaccgtaacttgaaagtatttcg and agccaaattaattctaattatctctctaacattcttgc, followed by purification using the PCR & DNA Cleanup Kit (NEB). Sequencing was performed by the MGH CCIB DNA Core. Statistical analysis was performed with MAGeCK98.
Satellite cell isolation and in vitro expansion
Satellite cells from control or Nudt21 KO mice treated with tamoxifen (see above), were isolated from mice as previously described37. Satellite cells were seeded on µ-Slide 8 Well, (80806, ibidi) coated with 1 μg/mL collagen type I from rat tail (Sigma) and 10 μg/mL mouse Laminin (Invitrogen) and were cultured for 5 days in F10 (Gibco) supplemented with 20% horse serum (Gibco), 100 U/mL penicillin, 100 μg/mL streptomycin, and GlutaMAX (Thermo), bFGF (5 ng/mL; Sigma) was added daily.
Muscle injury and regeneration
Nudt21fl/fl (control) or Nudt21fl/fl; Pax7-CreER (KO) were treated with tamoxifen as described above. Cardiotoxin (CTX) injury for in vivo regeneration studies was performed by injecting 50 µl of 10 mM CTX (Latoxan) into the tibialis anterior (TA) muscle of animals. Injured TA muscles were collected 14 days post injury and embedded in optimal cutting temperature (OCT) compound (Sakura Tissue-Tek) and frozen in liquid nitrogen-chilled isopentane for 30 s. The frozen muscles were then sectioned using a cryostat set at −20 °C to obtain 10 μm thick mid-belly cross-sections. For dystrophin and Hoechst double staining, the mid-belly sections were first fixed with 1% paraformaldehyde (PFA) in PBS for 10 min at room temperature (RT). Following fixation, the sections were washed with PBS three times. The sections were blocked using a staining buffer composed of 0.5% BSA and Triton-X100 in PBS for 20 min at RT. After blocking, the sections were incubated with primary antibodies in the staining buffer for 1 hour at RT. The primary antibody used was anti-dystrophin (Mandys8, Santa Cruz, sc-58754) at a dilution of 1:100. Following primary antibody incubation, the sections were washed with three times with PBS. Subsequently, the sections were incubated with secondary antibodies in the staining buffer for 1 h at RT. The secondary antibody used was anti-mouse IgG2b Alexa 594 (Thermo Fisher Scientific) at a dilution of 1:250. After secondary antibody incubation, the sections were washed with PBS three times. The sections were then stained with Hoechst dye at a dilution of 1:1000 in PBS. Finally, the sections were washed with PBS, repeated two times, and prepared for imaging.
Complete blood counts with differential
Whole blood was collected via retro-orbital bleeding using heparinized micro-hemocrit capillary tubes (Thermo Fisher Scientific) and transferred into EDTA-coated microvette tubes (Sarstedt). Complete blood counts with differential were then analyzed using an Element HT5 machine (Heska).
Flow cytometry (bone marrow)
Whole blood was processed for flow cytometry by lysing red blood cells in RBC lysis buffer (BioLegend) for 8 min on ice. Cells were then centrifuged at 1200 rpm for 5 min, resuspended in FACS buffer (2% fetal bovine serum (Hyclone) in PBS), and filtered through a 0.2 μm mesh. After another centrifugation at 1200 rpm for 5 min, the cells were stained on ice for 1 h. The following antibodies were used: CD3e (APC; BD Biosciences; clone 145-2C11), CD45.1 (PE; BioLegend; clone A20), CD45.2 (Pacific Blue; BioLegend; clone 104), Mac1 (FITC; eBiosciences; clone M1/70), B220/CD45R (APC eFluor 780; Life Technologies; clone RA3-6B2), Gr1 (PE or PE/Cy5; eBiosciences; clone RB6-8C5), Fc block (BD Biosciences), propidium iodide (BD Biosciences), and DAPI (BD Biosciences). After staining, cells were washed with FACS buffer. Propidium iodide or DAPI was used for viability staining. Bone marrow was harvested from euthanized mice by flushing femurs and tibiae with FACS buffer using a 27½ gauge needle, followed by dissociation with a 22 gauge needle. Cells were centrifuged at 1200 rpm for 5 min, resuspended in 5 ml of RBC lysis buffer for 8 min on ice, and the lysis was quenched with 20 ml of culture media before another centrifugation at 1200 rpm for 5 min. Cells were resuspended in 10 ml of FACS buffer, filtered through a 0.2 μm mesh, and counted. Cells were aliquoted and stained for the detection of HSPCs with the following panel: SLAM panel (3 million cells): Lineage markers [B220/CD45R (PE/Cy5; BioLegend; clone RA3-6B2), Ter119 (PE/Cy5; eBiosciences; clone TER-119), CD3e (PE/Cy5; BioLegend; clone 145-2C11), Mac1 (PE/Cy5; eBiosciences; clone M1/70), Gr1 (PE/Cy5; eBiosciences; clone RB6-8C5)], Sca1 (PE/Cy7; eBiosciences; clone D7), CD150 (PE or BV605; BioLegend; clone TC15-12F12.2), CD48 (APC; BioLegend; clone HM48-1), CD45.1 (PE; BioLegend; clone A20), CD45.2 (Pacific Blue; BioLegend; clone 104), cKit (eFluor 780; eBiosciences; clone 2B8), Fc block (BD Biosciences). DAPI was used as a viability dye.
Hematopoietic progenitor in vitro culture
Ten thousand LSK cells were sorted and cultured in IMDM (Gibco) supplemented with 10% FBS, 1× GlutaMax (Life Technologies), 100 U/mL Penicillin, 100 μg/mL Streptomycin (Life Technologies), and 50 μM β-mercaptoethanol (Life Technologies). 10 ng/ml IL3 (Peprotech), 10 ng/ml IL6 (Peprotech), 10 ng/ml SCF (Peprotech), 5 ng/ml GM-CSF (Peprotech), 50 ng/ml TPO (Peprotech), and 3 U/ml EPO (Bio legend) were added. Where indicated, 4OHT (0.05 µM) or ETOH was added.
PAS-seq analysis
Total RNA from cells was isolated with the RNeasy Kit (QIazol). PAS-seq library preparation and data analyses were performed as previously described99. APA events were considered significant if they met both criteria: a false discovery rate (FDR) < 0.05 and a change in distal PAS usage greater than 14.5%. Statistical significance was assessed using edgeR, and FDR values were calculated using the Benjamini–Hochberg (BH) method to correct for multiple testing.
Single-cell RNA-seq data analysis
Chromium single cell 3′ reagent v3.0 kit (10x Genomics), followed by sequencing on Illumina NextSeq 2000 instrument, which resulted in approximately ranging from 250 to 400 million reads per sample. Primary processing of scRNA-Seq sequencing Single-cell RNA-seq library construction was performed on the Chromium 10× instrument using data was performed using CellRanger (v6.0.0) (https://support.10xgenomics.com/single-cell-gene-expression/software/overview/welcome) [https://doi.org/10.1038/ncomms14049]. In brief, reads were aligned to mm10 mouse reference genome (mapping rate of ~95%), followed by the generation of read counts per gene in each cell. Further analysis was performed using the Seurat 3.2.3 package (https://satijalab.org/seurat/). We filtered out cells with <200 expressed genes and genes expressed in <3 cells, followed by the exclusion of cells with high content of mitochondrial transcripts (>10% of total reads). Counts across all cells for each sample were normalized using the NormalizeData function in Seurat. Using the FindVariableFeatures function, 2000 features were selected to be used in principal component analysis (PCA). All individual samples were integrated using Seurat canonical correlation analysis (CCA) method. Integration anchors were determined using FindIntegrationAnchors function and then used in IntegrateData function. UMAP plots and cell clusters were generated using RunUMAP and FindClusters functions, followed by manual cell type annotation based on type-specific gene markers.
RNA-sequencing and bioinformatic analysis
mRNA-seq libraries were constructed with the SMARTer v3 kit (Clontech) and sequenced on the Illumina HiSeq2500 as PE50, producing approximately 20–30 million reads per sample. microRNA-seq libraries were constructed with the NEBNext® Small RNA Library Prep Set (NEB) and sequenced on the Illumina HiSeq2500, producing approximately 10 million reads per sample as SR50. Transcriptome mapping was performed using STAR with the UCSC annotation of the mm10 reference genome. To align microRNAs to the reference database miRbase (mature.fa). The mature.fa file was U to T converted via: “sed ‘/^[^>]/s/U/T/g’ mature.fa > mature_conv.fa”. And aligned with bowtie100 using the following attributes “-v 1 -a --best –strata”. Read counts for individual genes were generated using featureCounts from the Subread package101. microRNA counts were generated with samtools idxstas. Differential expression analysis for mRNA and microRNA was carried out with EdgeR after normalizing read counts and including only genes with CPM > 1 in at least one sample. Differentially expressed genes or microRNAs were identified based on a >2-fold change in expression value, unless otherwise specified.
Proteomics
Protein concentration of cell lysates was determined by DC Assay (Bio-Rad). Reduction alkylation, precipitation and TMT labeling and Liquid chromatography coupled to mass spectrometry were performed as previously described51. Differential analysis was performed by the DEP package102.
Translational efficiency
Conditional Ribosome profiling cDNA library preparation for deep sequencing was performed as previously described90. Translation efficiency (TE) was calculated using ribosome protected fragments (RPF) and RNA-seq reads per kilobase per million mapped reads (RPKM).TE = (RPKM[RPF]/RPKM[mRNA]). We obtained transcript, CDS and 3′UTR length for mouse genes from UCSC (GRCm38_UCSC_refGene.gtf).
IVT mRNA synthesis and transfection
The mRNA constructs were generated by VectorBuilder using a standardized workflow. Briefly, Nup160 CDS fused N-terminally to eGFP (or eGFP alone) and carrying either Short, Mid, or Long Nup160 3′UTRs were cloned into pmRVac plasmid backbones under control of a T7 promoter. Following sequence validation, plasmids were linearized and used as templates for in vitro transcription with complete substitution of uridine by N1-methylpseudouridine (m1Ψ), co-transcriptional Cap 1, and poly(A) addition. The resulting mRNAs were purified, quality-controlled, and delivered as RNase-free, ready-to-use solutions.
mRNA transfection was performed with MessengerMAX Transfection Reagent and Opti-MEM Reduced Serum Medium (both purchased from Thermo Fisher Scientific). One day before transfection (day −1), cells were seeded at 3 × 104 cells per well in 24-well plastic or glass-bottom plates. Equimolar concentrations of distinct IVT mRNAs were used for every transfection, with maximum mRNA dose not exceeding 25 ng per 1 × 104 cells. On the day of transfection (day 0), culture medium was exchanged with pre-warmed culture media 30 min prior transfection. The transfection mixture was prepared in two separate tubes. In the first tube, IVT mRNAs were diluted in 50 µl OPTI-MEM and left at room temperature. In the second tube, the transfection reagent was prepared by diluting 1.5 µl MessengerMAX in 50 µl Opti-MEM followed by brief vortexing (2–3 s). Both reactions were incubated separately at room temperature for 10 min before they were combined. Both reactions were incubated for 5 min at room temperature and combined followed by an additional 10 min incubation at room temperature. Before transfection, the brought 900 µl Opti-MEM were added to the mix. The resulting mRNA–lipid complexes were added to cells dropwise to ensure even distribution. For 24-well plates, 150–300 µl of complex were dispensed per well. Transfected cells were maintained under standard culture conditions. On day 1, cells were gently rinsed with PBS to remove phenol red and examined microscopically to assess cell health and transfection performance prior to downstream procedures. eGFP Expression was assessed by microscopy or FACS 24 h post transfection.
RNA-IP/MS
N-Terminally Biotinylated RNAs were purchased from VectorBuilder and all reagents for the RNA-immune precipitation were from Thermo Fisher. For RNA-IP, Biotinylated IVT RNAs were separately resuspended in equimolar concentrations of 1 µM in 500 µl ice-cold binding buffer (20 mM Tris-HCl pH 8.0, 250 mM NaCl, 1 mM EDTA, RNase inhibitor) and denatured at 65 °C for 10 min, followed by cooling on ice. For coupling, 25 µl Dynabeads MyOne Streptavidin C1 (Thermo Fisher) were washed 5× in 1 ml washing buffer (20 mM Tris-HCL pH 8, 250 mM NaCl, 50 mM KCl, 2.5 mM MgCl, RNase inhibitor, protease inhibitor) and incubated for 1–2 h at 4 °C with rotation. After washing, RNA-bead complexes were incubated with cell lysates. Cell lysates were prepared by resuspending 10⁷ cells in 1 ml lysis buffer (20 mM Tris-HCl pH 8.0, 250 mM NaCl, 50 mM KCl, 2.5 mM MgCl2, 0.5% Triton X-100, 0.2% NP-40, RNase inhibitor, protease inhibitor, 0.6% RNase-free DNase), followed by 3x needle passaging and sonication (Bioruptor Pico; 30 s on/off, 5 cycles at 4 °C) and clarification through centrifugation. Cell lysate supernatants were combines with RNA-bead complexes and incubated for 4 h at 4 °C with rotation. Beads were washed in 3× wash buffer and 1× rinse buffer (20 mM Tris-HCL pH 8, 250 mM NaCl, 50 mM KCl, 2.5 mM MgCl2) to remove residual inhibitors. Bound proteins were eluted in 200 µl elution buffer (10 mM Tris-HCl pH 8.0, 15 mM NaCl containing 0.1 mg/mL RNase A) for 10 min at room temperature followed by 20 min at 30 °C. Beads were removed via and eluates were transferred to fresh tubes, further incubated on ice, and stored at –80 °C until submitted for label-free mass spectrometry.
ceRNA analysis
This analysis was carried out as reported previously42. Briefly, the datasets PAS-seq (APA), bulk RNA-seq (three biological replicates per condition: Ctrl, Nudt21 KO, Hypomorph) and small-RNA-seq (miRNA) were used to determine putative ceRNAs. For each miRNA, we quantified all unordered RNA-RNA pairs among its targets. To ensure robustness, only genes with >20 distinct targeting miRNAs were further considered. Pairwise Pearson correlations were computed across the three replicates. Pairs with r > 0.6 were classified as ceRNAs.
Integrative analysis
The list of essential genes was derived by combining two datasets: CRISPR_Inferred_essential (23q4) and CRISPR_common_essentials (22q2), both available from the DepMap portal (https://depmap.org/portal/data_page/). miRNA targets were identified by extracting annotated miRNA-mRNA interactions from TarBase103, specifically using the Mus_musculus_TarBase_v9.tsv file. Only miRNA-mRNA interactions positioned at the 3’UTR with a stringent microT score ≥0.7, as previously recommended104, were considered. We collected protein complex information from the EBI databases to compile a comprehensive dataset of known protein complexes. All processed ‘Omics’ data are available as tables in the Supplementary Data.
Disome selective profiling (DiSP)
Cells were lysed and subjected to RNase1 digestion, ribosome-nascent chain complexes were separated to disome (D) and monosome on sucrose density gradient91,105. Ribosome profiling reads from D and monosome (M) fractions were processed using a custom R pipeline. For each coding sequence (CDS), aligned reads were mapped to the nucleotide grid and log2(D/M) ratios were computed. To quantify positional changes in co-translational assembly (co-co-assembly), each CDS was divided into three windows: early (5–25%), mid (30–70%), and tail (last 30%), with enforced minimum lengths. For each window, reads were summed, and window-level log₂(D/M) were calculated. The co-co assembly score was defined as the maximum of (mid–early) or (tail–early), capturing either mid-body or C-terminal enrichment.
Genes were excluded if they lacked sufficient CDS coverage, had fewer than the required number of codons per window, or failed adaptive read-support thresholds (scaled to library depth). This pipeline was applied to Nudt21 knockout (KO) and control (WT) datasets (two replicates each). Of ~13,800 coding sequences, ~6400–7300 genes per replicate passed filtering (52–56% retention), consistent with lower sequencing depth compared to benchmark HEK293T data. Replicates showed moderate reproducibility (Spearman ρ ≈ 0.43; ~50% overlap among the top 200 genes). Consistent with Bertolini et al.91, genes encoding transmembrane domain (TMD) proteins were excluded, as membrane proteins can yield false-positive DiSP signals due to aberrant sedimentation. For KO vs WT comparison, mean co-co assembly scores were computed per gene, and the difference (Δ = KO − WT) was used as the effect size.
Statistical analysis
Statistical evaluation was performed with GraphPad Prism 5 or R using linear regression and two-tailed tests: t-test, Anova test or Wilcoxon rank test as specified in the Figure legends. The number of independent data points n always refers to biological replicates. Bonferroni correction was applied to the gene expression analysis to correct for multiple comparisons. *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001, n.s., not significant, p > 0.05.
Statistical analyses were conducted using Prism software (GraphPad) and R software. Specific details for each analysis, including the number of replicates, are provided in the Figure legends. Analyses were performed comparing genotypes and treatment conditions (wild-type, Nudt21 knockout, and hypomorph). Where applicable, experimental replicates and time points were included as covariates in linear models. Sex was balanced across groups (50% male, 50% female), but it was not included as a covariate, as no sex-specific effects were anticipated or observed. No additional covariates were tested.
Data availability
The transcriptomics data are available at the Gene Expression Omnibus (GEO) webserver under accession ID GSE280706 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE280706]. All mass spectrometry RAW data can be accessed through the MassIVE data repository (massive.ucsd.edu) under the accession number MSV000096277 [ftp://massive-ftp.ucsd.edu/v11/ MSV000096277 /]. The following publicly available single-cell RNA-seq data sets were analyzed in this study: Muscle (GSE143437): [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE143437]. Esophagus (GSE193376): [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE193376]. Intestine (GSE92332): [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE92332] Source data are provided with this paper.
References
Beumer, J. & Clevers, H. Hallmarks of stemness in mammalian tissues. Cell Stem Cell 31, 7–24 (2024).
Tian, B., Hu, J., Zhang, H. & Lutz, C. S. A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res. 33, 201–212 (2005).
Thivierge, C. et al. Alternative polyadenylation confers Pten mRNAs stability and resistance to microRNAs. Nucleic Acids Res. 46, gky666 (2018).
Boutet, S. C. et al. Alternative polyadenylation mediates microRNA regulation of muscle stem cell function. Cell Stem Cell 10, 327–336 (2012).
Tian, B. & Manley, J. L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 18, 18–30 (2017).
Masamha, C. P. et al. CFIm25 links alternative polyadenylation to glioblastoma tumour suppression. Nature 510, 412–416 (2014).
Horste, E. L. et al. Subcytoplasmic location of translation controls protein output. Mol. Cell 83, 4509–4523.e11 (2023).
Luo, Y. et al. mRNA interactions with disordered regions control protein activity. Preprint at https://doi.org/10.1101/2023.02.18.529068 (2023).
Ma, W., Zhen, G., Xie, W. & Mayr, C. In vivo reconstitution finds multivalent RNA–RNA interactions as drivers of mesh-like condensates. eLife 10, e64252 (2021).
Lee, S.-H. & Mayr, C. Gain of additional BIRC3 protein functions through 3ʹ-UTR-mediated protein complex formation. Mol. Cell 74, 701–712.e9 (2019).
Berkovits, B. D. & Mayr, C. Alternative 3′ UTRs act as scaffolds to regulate membrane protein localization. Nature 522, 363–367 (2015).
Mayr, C. What are 3′ UTRs doing? Cold Spring Harb. Perspect. Biol. 11, a034728 (2019).
Mandel, C. R., Bai, Y. & Tong, L. Protein factors in pre-mRNA 3′-end processing. Cell. Mol. Life Sci. 65, 1099–1122 (2008).
Shi, Y. et al. Molecular architecture of the human pre-mRNA 3′ processing complex. Mol. Cell 33, 365–376 (2009).
Zhang, Y., Sun, Y., Shi, Y., Walz, T. & Tong, L. Structural insights into the human pre-mRNA 3′-end processing machinery. Mol. Cell 77, 800–809.e6 (2020).
Schmidt, M. et al. Reconstitution of 3′ end processing of mammalian pre-mRNA reveals a central role of RBBP6. Genes Dev. 36, 195–209 (2022).
Boreikaite, V., Elliott, T. S., Chin, J. W. & Passmore, L. A. RBBP6 activates the pre-mRNA 3′ end processing machinery in humans. Genes Dev. 36, 210–224 (2022).
Brumbaugh, J. et al. Nudt21 controls cell fate by connecting alternative polyadenylation to chromatin signaling. Cell 172, 106–120.e21 (2018).
Gruber, A. R., Martin, G., Keller, W. & Zavolan, M. Cleavage factor I m is a key regulator of 3′ UTR length. RNA Biol. 9, 1405–1412 (2012).
Alcott, C. E. et al. Partial loss of CFIm25 causes learning deficits and aberrant neuronal alternative polyadenylation. eLife 9, e50895 (2020).
Yang, Q., Gilmartin, G. M. & Doublié, S. Structural basis of UGUA recognition by the Nudix protein CFI(m)25 and implications for a regulatory role in mRNA 3’ processing. Proc. Natl. Acad. Sci. USA 107, 10062–10067 (2010).
Zhu, Y. et al. Molecular mechanisms for CFIm-mediated regulation of mRNA alternative polyadenylation. Mol. Cell 69, 62–74.e4 (2018).
Kim, H. & Lee, Y. Interaction of poly(A) polymerase with the 25-kDa subunit of cleavage factor I. Biochem. Biophys. Res. Commun. 289, 513–518 (2001).
Mitschka, S. & Mayr, C. Context-specific regulation and function of mRNA alternative polyadenylation. Nat. Rev. Mol. Cell Biol. 1–18 https://doi.org/10.1038/s41580-022-00507-5 (2022).
Zhou, Y. et al. Nudt21-mediated alternative polyadenylation of MZT1 3′UTR contributes to pancreatic cancer progression. iScience 27, 108822 (2024).
Zhu, Y. et al. NUDT21 promotes tumor growth and metastasis through modulating SGPP2 in human gastric cancer. Front. Oncol. 11, 670353 (2021).
Li, N. et al. CFIm-mediated alternative polyadenylation safeguards the development of mammalian pre-implantation embryos. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2022.11.016 (2022).
Gennarino, V. A. et al. NUDT21-spanning CNVs lead to neuropsychiatric disease and altered MeCP2 abundance via alternative polyadenylation. Elife 4, e10782 (2015).
Pelka, G. J., Watson, C. M., Christodoulou, J. & Tam, P. P. L. Distinct expression profiles of Mecp2 transcripts with different lengths of 3′UTR in the brain and visceral organs during mouse development. Genomics 85, 441–452 (2005).
Ko, J. et al. Transforming growth factor β1 alters the 3′-UTR of mRNA to promote lung fibrosis. J. Biol. Chem. 294, 15781–15794 (2019).
Weng, T. et al. Cleavage factor 25 deregulation contributes to pulmonary fibrosis through alternative polyadenylation. J. Clin. Investig. 129, 1984–1999 (2019).
Sommerkamp, P. et al. Differential alternative polyadenylation landscapes mediate hematopoietic stem cell activation and regulate glutamine metabolism. Cell Stem Cell https://doi.org/10.1016/j.stem.2020.03.003 (2020).
Witkowski, M. T. et al. NUDT21 limits CD19 levels through alternative mRNA polyadenylation in B cell acute lymphoblastic leukemia. Nat. Immunol. 23, 1424–1432 (2022).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Kabir, M. F. et al. Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium. Nat. Commun. 13, 2167 (2022).
Bagger, F. O. et al. BloodSpot: a database of gene expression profiles and transcriptional programs for healthy and malignant haematopoiesis. Nucleic Acids Res. 44, D917–D924 (2016).
Maesner, C. C., Almada, A. E. & Wagers, A. J. Established cell surface markers efficiently isolate highly overlapping populations of skeletal muscle satellite cells by fluorescence-activated cell sorting. Skelet. Muscle 6, 35 (2016).
Almada, A. E. et al. FOS licenses early events in stem cell activation driving skeletal muscle regeneration. Cell Rep. 34, 108656 (2021).
Yagi, M. et al. Dissecting dual roles of MyoD during lineage conversion to mature myocytes and myogenic stem cells. Genes Dev. 35, 1209–1228 (2021).
Micheli, A. J. D. et al. Single-cell analysis of the muscle stem cell hierarchy identifies heterotypic communication signals involved in skeletal muscle regeneration. Cell Rep. 30, 3583–3595.e5 (2020).
Murphy, M. M., Lawson, J. A., Mathew, S. J., Hutcheson, D. A. & Kardon, G. Satellite cells, connective tissue fibroblasts and their interactions are crucial for muscle regeneration. Development 138, 3625–3637 (2011).
Park, H. J. et al. 3′ UTR shortening represses tumor-suppressor genes in trans by disrupting ceRNA crosstalk. Nat. Genet. 50, 783–789 (2018).
Hennings, H. et al. Calcium regulation of growth and differentiation of mouse epidermal cells in culture. Cell 19, 245–254 (1980).
Shepard, P. J. et al. Complex and dynamic landscape of RNA polyadenylation revealed by PAS-Seq. RNA 17, 761–772 (2011).
Kowalski, M. H. et al. Multiplexed single-cell characterization of alternative polyadenylation regulators. Cell https://doi.org/10.1016/j.cell.2024.06.005 (2024).
Sun, M. et al. NUDT21 regulates 3′-UTR length and microRNA-mediated gene silencing in hepatocellular carcinoma. Cancer Lett. 410, 158–168 (2017).
Lackford, B. et al. Fip1 regulates mRNA alternative polyadenylation to promote stem cell self-renewal. EMBO J. 33, 878–889 (2014).
de Morree, A. et al. Alternative polyadenylation of Pax3 controls muscle stem cell fate and muscle function. Science 366, 734–738 (2019).
Porpiglia, E. et al. Elevated CD47 is a hallmark of dysfunctional aged muscle stem cells that can be targeted to augment regeneration. Cell Stem Cell 29, 1653–1668.e8 (2022).
Sumazin, P. et al. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 147, 370–381 (2011).
Lapek, J. D. et al. Detection of dysregulated protein-association networks by high-throughput proteomics predicts cancer vulnerabilities. Nat. Biotechnol. 35, 983–989 (2017).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
Xu, S. et al. Chemical inhibition of the integrated stress response impairs the ubiquitin-proteasome system. Commun. Biol. 7, 1282 (2024).
Cervia, L. D. et al. A ubiquitination cascade regulating the integrated stress response and survival in carcinomas. Cancer Discov. 13, 766–795 (2022).
Rabouw, H. H. et al. Small molecule ISRIB suppresses the integrated stress response within a defined window of activation. Proc. Natl. Acad. Sci. USA 116, 2097–2102 (2019).
Tsitsiridis, G. et al. CORUM: the comprehensive resource of mammalian protein complexes–2022. Nucleic Acids Res. 51, D539–D545 (2022).
Delavoie, F., Soldan, V., Rinaldi, D., Dauxois, J.-Y. & Gleizes, P.-E. The path of pre-ribosomes through the nuclear pore complex revealed by electron tomography. Nat. Commun. 10, 497 (2019).
Yagita, Y., Zavodszky, E., Peak-Chew, S.-Y. & Hegde, R. S. Mechanism of orphan subunit recognition during assembly quality control. Cell 186, 3443–3459.e24 (2023).
Walther, T. C. et al. The conserved Nup107-160 complex is critical for nuclear pore complex assembly. Cell 113, 195–206 (2003).
Rodriguez-Muñoz, M., Anglada, T. & Genescà, A. A matter of wrapper: defects in the nuclear envelope of lagging and bridging chromatin threatens genome integrity. Semin. Cell Dev. Biol. 123, 124–130 (2022).
Gunn, A. L., Yashchenko, A. I., Dubrulle, J., Johnson, J. & Hatch, E. M. A high-content screen reveals new regulators of nuclear membrane stability. Sci. Rep. 14, 6013 (2024).
Wang, J. et al. Comprehensive mapping of alternative polyadenylation site usage and its dynamics at single-cell resolution. Proc. Natl. Acad. Sci. USA 119, e2113504119 (2022).
Hampoelz, B. et al. Nuclear pores assemble from nucleoporin condensates during oogenesis. Cell 179, 671–686.e17 (2019).
Shi, K. et al. Nucleoporins shape germ granule architecture and balance small RNA silencing pathways. Nat. Commun. 16, 4295 (2025).
Seidel, M. et al. Co-translational assembly orchestrates competing biogenesis pathways. Nat. Commun. 13, 1224 (2022).
Galmozzi, C. V., Merker, D., Friedrich, U. A., Döring, K. & Kramer, G. Selective ribosome profiling to study interactions of translating ribosomes in yeast. Nat. Protoc. 14, 2279–2317 (2019).
Lautier, O. et al. Co-translational assembly and localized translation of nucleoporins in nuclear pore complex biogenesis. Mol. Cell 81, 2417–2427.e5 (2021).
Kamenova, I. et al. Co-translational assembly of mammalian nuclear multisubunit complexes. Nat. Commun. 10, 1740 (2019).
Ran, Y. et al. CFIm25 regulates human stem cell function independently of its role in mRNA alternative polyadenylation. RNA Biol. 19, 686–702 (2022).
Chang, J.-W., Yeh, H.-S. & Yong, J. Alternative polyadenylation in human diseases. Endocrinol. Metab. 32, 413–421 (2017).
Sandberg, R., Neilson, J. R., Sarma, A., Sharp, P. A. & Burge, C. B. Proliferating cells express mRNAs with shortened 39 untranslated regions and fewer microRNA target sites. Science 320, 1643–1647 (2008).
Huang, J. et al. Suppression of cleavage factor Im 25 promotes the proliferation of lung cancer cells through alternative polyadenylation. Biochem. Biophys. Res. Commun. 503, 856–862 (2018).
Liu, Y. et al. Competitive endogenous RNA is an intrinsic component of EMT regulatory circuits and modulates EMT. Nat. Commun. 10, 1637 (2019).
Bosson, A. D., Zamudio, J. R. & Sharp, P. A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 56, 347–359 (2014).
Li, X., Ding, J., Wang, X., Cheng, Z. & Zhu, Q. NUDT21 regulates circRNA cyclization and ceRNA crosstalk in hepatocellular carcinoma. Oncogene 39, 891–904 (2020).
D’Angelo, M. A., Raices, M., Panowski, S. H. & Hetzer, M. W. Age-dependent deterioration of nuclear pore complexes causes a loss of nuclear integrity in postmitotic cells. Cell 136, 284–295 (2009).
Sakuma, S. et al. Inhibition of nuclear pore complex formation selectively induces cancer cell death. Cancer Discov. 11, 176–193 (2021).
Cho, U. H. & Hetzer, M. W. Nuclear periphery takes center stage: the role of nuclear pore complexes in cell identity and aging. Neuron 106, 899–911 (2020).
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140, 372–383 (2010).
Ibarra, A., Benner, C., Tyagi, S., Cool, J. & Hetzer, M. W. Nucleoporin-mediated regulation of cell identity genes. Gene Dev. 30, 2253–2258 (2016).
Taniguchi, R. et al. Nuclear pores safeguard the integrity of the nuclear envelope. Nat. Cell Biol. 27, 762–775 (2025).
Huang, M. et al. FACS-based genome-wide CRISPR screens define key regulators of DNA damage signaling pathways. Mol. Cell 83, 2810–2828.e6 (2023).
Paulsen, R. D. et al. A genome-wide siRNA screen reveals diverse cellular processes and pathways that mediate genome stability. Mol. Cell 35, 228–239 (2009).
Liu, J. & Hetzer, M. W. Nuclear pore complex maintenance and implications for age-related diseases. Trends Cell Biol. 32, 216–227 (2021).
Hoffman, Y. et al. 3’UTR shortening potentiates microRNA-based repression of pro-differentiation genes in proliferating human cells. PLoS Genet. 12, e1005879 (2016).
Gruber, A. J. & Zavolan, M. Alternative cleavage and polyadenylation in health and disease. Nat. Rev. Genet. 20, 599–614 (2019).
Subramanian, A. et al. Alternative polyadenylation is a determinant of oncogenic Ras function. Sci. Adv. 7, eabh0562 (2021).
Li, L. et al. An atlas of alternative polyadenylation quantitative trait loci contributing to complex trait and disease heritability. Nat. Genet. 1–12 https://doi.org/10.1038/s41588-021-00864-5 (2021).
Pagani, G. & Gandellini, P. Cleavage and polyadenylation machinery as a novel targetable vulnerability for human cancer. Cancer Gene Ther. 1–4 https://doi.org/10.1038/s41417-024-00770-y(2024).
Shiber, A. et al. Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling. Nature 561, 268–272 (2018).
Bertolini, M. et al. Interactions between nascent proteins translated by adjacent ribosomes drive homomer assembly. Science 371, 57–64 (2021).
Bernardini, A. & Tora, L. Co-translational assembly pathways of nuclear multiprotein complexes involved in the regulation of gene transcription. J. Mol. Biol. 436, 168382 (2024).
Tsopoulidis, N. et al. T cell receptor–triggered nuclear actin network formation drives CD4+ T cell effector functions. Sci. Immunol. 4, eaav1987 (2019).
Loonstra, A. et al. Growth inhibition and DNA damage induced by Cre recombinase in mammalian cells. Proc. Natl. Acad. Sci. USA 98, 9209–9214 (2001).
Huebner, A. J. et al. Dissection of gastric homeostasis in vivo facilitates permanent capture of isthmus-like stem cells in vitro. Nat. Cell Biol. 25, 390–403 (2023).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Yoon, Y., Soles, L. V. & Shi, Y. PAS-seq 2: A fast and sensitive method for global profiling of polyadenylated RNAs. Methods Enzym 655, 25–35 (2021).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Zhang, X. et al. Proteome-wide identification of ubiquitin interactions using UbIA-MS. Nat. Protoc. 13, 530–550 (2018).
Skoufos, G. et al. TarBase-v9.0 extends experimentally supported miRNA–gene interactions to cell-types and virally encoded miRNAs. Nucleic Acids Res. 52, D304–D310 (2023).
Tastsoglou, S. et al. DIANA-microT 2023: including predicted targets of virally encoded miRNAs. Nucleic Acids Res. 51, W148–W153 (2023).
Koubek, J. et al. A simple, fast, and cost-efficient protocol for ultra-sensitive ribosome profiling. Nucleic Acids Res. 53, gkaf902 (2025).
Meldal, B. H. M. et al. Complex Portal 2018: extended content and enhanced visualization tools for macromolecular complexes. Nucleic Acids Res. 47, D550–D558 (2019).
Acknowledgements
We thank members of the Hochedlinger laboratory for their suggestions, and members of the MGH CRM/HSCI Flow Core, the Harvard Rodent Histopathology Core, the Taplin Mass Spectrometry Facility, and the MGH Next Generation Sequencing Core for their support. We also thank Panagiotis Kastritis, Ioannis Sanidas for constructive comments and feedback. N.T. was supported by the German Research Foundation (DFG) Research Grant. A.J.W. was supported by funds from NIH (R01 AR077695). H.Ho. is the Brant Carleton Endowed Chair in Acute Myeloid Leukemia Research. K.Ho. was supported by funds from the MGH, the NIH (R01 HD058013, R01 AR077695, and P01 GM099134), and the Milky Way Research Foundation and the Gerald and Darlene Jordan Chair in Regenerative Medicine. Y.Y. and Y.S. were supported by R01AI170840. B.B. acknowledges a research grant of the European Union (ERC - SyG − 101072047 - CoTransComplex). Views and opinions expressed are however, those of the authors only and do not necessarily reflect those of the European Union or the European Research Council. Neither the European Union nor the granting authority can be held responsible for them. M.S.H. was supported by Deutsche Krebshilfe (70115739; Max-Eder Program).
Author information
Authors and Affiliations
Contributions
N.T. performed the majority of experiments and bioinformatic analyses, and wrote the manuscript together with K.Ho. K.Ho. conceived and supervised the study, provided overall guidance, and secured funding. M.Y., K.A.M., K.-H.L., and A.J.W. performed all muscle-related experiments and analyses. J.B. generated the Nudt21 floxed mouse line. R.M., E.F.Z., and W.Ha. conducted the mass spectrometry–based proteomics experiments and assisted with bioinformatic analyses and data interpretation. R.G. performed single-cell RNA-sequencing of murine hematopoietic cells. A.Hu. established intestinal organoid cultures. H.Ho. provided conceptual guidance for the hematopoietic studies and interpretation of the overall phenotype. G.Kr., B.B., M.S.H., S.I., and V.L.S. performed DiSP assays and analyses of co-translational protein complex assembly. Y.Y. and Y.S. performed PAS-seq experiments, analyzed alternative polyadenylation data, and provided expert consultation on APA biology. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
N.T. is a founder and equity holder in Tector Therapeutics LLC. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Bin Tian, Eric Wagner, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tsopoulidis, N., Yagi, M., Brumbaugh, J. et al. Modulation of Nudt21 levels reveals dose-dependent roles of alternative polyadenylation in tissue regeneration. Nat Commun 17, 2005 (2026). https://doi.org/10.1038/s41467-026-68630-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68630-x