Abstract
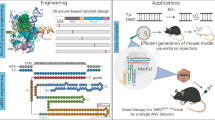
Eukaryotic Fanzor proteins are compact, programmable RNA-guided nucleases with substantial potential for genome editing, although their efficiency in mammalian cells remains suboptimal. Here, we present a combinatorial engineering strategy to optimize a representative Fanzor system, MmeFz2–ωRNA. AlphaFold3-powered rational redesign produced a minimized ωRNA scaffold that is 30% smaller while maintaining up to 82.2% efficiency. Synergistic structure-guided and AI-augmented protein engineering generated two variants, enMmeFz2 and evoMmeFz2, which exhibited an average ~32-fold increase in activity across 38 genomic loci. Moreover, fusion of the non-specific DNA-binding domain HMG-D further enhanced editing performance (enMmeFz2-HMG-D and evoMmeFz2-HMG-D). Notably, evoMmeFz2-HMG-D demonstrated robust in vivo genome editing activity, enabling dystrophin restoration in humanized male Duchenne muscular dystrophy mouse models via single adeno-associated virus (AAV) delivery. This study establishes Fanzor2 as a gene editing platform for genome engineering and therapeutic applications, and underscores the power of AI-guided engineering to accelerate genome editor development while reducing experimental burden.
Similar content being viewed by others
Introduction
The advent of programmable genome editing technologies, particularly the clustered regularly interspaced short palindromic repeats and CRISPR-associated proteins (CRISPR-Cas) system, has revolutionized modern biotechnology and medicine. CRISPR effectors, such as Cas9 and Cas12 nucleases1,2,3, enable precise DNA manipulation across species, making them promising tools for biological research, gene therapy, and agricultural breeding4,5,6,7. Nevertheless, the large sizes of widely used Cas nucleases (Cas9 and Cas12a), typically exceeding 1000 amino acids (aa), pose a significant challenge for efficient delivery, particularly in adeno-associated virus (AAV)-mediated in vivo gene therapy applications8.
Recently, compact CRISPR nucleases and ancestral proteins, derived from prokaryotic organisms, have been discovered and characterized, including miniature Cas12 effectors (Cas12f, Cas12j, and Cas12n, ranging from 400 to 800 aa)9,10,11,12,13,14,15,16,17,18, as well as their ancestral proteins TnpB and IscB (~400 aa)19,20,21,22,23,24,25,26,27,28,29,30. Furthermore, Fanzor (Fz), eukaryotic ωRNA-guided endonucleases evolutionarily conserved across diverse eukaryotes, including fungi, algae, protostomes, metazoans, amorphea, and certain large dsDNA viruses, represent a unique family of RNA-programmable genome-editing enzymes with evolutionary distinctions from prokaryotic systems31,32,33. Notably, phylogenetic and structural studies have identified TnpB, a newly characterized prokaryotic obligate mobile element-guided activity (OMEGA) protein, as the evolutionary precursor to both eukaryotic Fz proteins and prokaryotic CRISPR-Cas12 nucleases. Fanzor proteins are classified into two major groups: Fz1 and Fz231,32,33. While Fz1 proteins range from 600 to 900 amino acids in length, Fz2 proteins are more compact (~480 aa) and share greater structural similarity with TnpB. Fz2’s compact architecture makes it a promising candidate for viral-delivered therapeutic genome editing. However, native Fz2 systems exhibit critically low activity in mammalian cells (<1% editing efficiency), likely due to suboptimal ωRNA scaffold and protein-DNA/RNA interactions. These limitations highlight the need for systematic engineering to optimize Fanzor2 as a practical genome-editing tool.
Traditional approaches to optimizing RNA-guided nucleases involve iterative cycles of directed evolution and structure-guided mutation design, which have successfully enhanced Cas12a, Cas12f, IscB, and TnpB systems. However, these methods are limited by labor-intensive experiments, suboptimal multiparameter optimization, and premature convergence to local fitness maxima. Emerging computational strategies in biology now offer transformative solutions to these challenges. Protein language models (PLMs), trained on evolutionary sequence data, facilitate in silico prediction of functional mutations with minimal experimental validation34. Meanwhile, AlphaFold3 expands structural modeling beyond protein folding to predict ternary complexes of proteins, guide RNA, and target DNA35. While previous studies have applied these tools individually to engineer RNA scaffolds or protein components, no approach has integrated RNA structural optimization with PLM-driven protein engineering to co-optimize an RNA-guided nuclease system. Here, we bridge this gap by developing a combinatorial framework that synergizes AlphaFold3-based ωRNA redesign with structure- and PLM-guided protein engineering (AlphaPLM), bypassing the trial-and-error bottlenecks of conventional methods.
In this study, we engineered MmeFz2, a 479-aa Fanzor2 homolog, into a highly efficient genome editor by synergistically optimizing its ωRNA scaffold and protein sequence. Using AlphaFold3, we first identified structural defects in the wild-type ωRNA (WT-ωRNA) and rationally redesigned a truncated variant with 30% reduced length and significantly improved editing activity. Concurrently, structure-guided mutagenesis of AlphaFold3-informed protein-ωRNA-DNA interfaces was performed, and the validation results were incorporated into a PLM-guided iterative evolution pipeline using EVOLVEpro to predict new functional mutations in silico. Two evolved MmeFz2 variants, enMmeFz2 derived from structure-informed design and evoMmeFz2 from PLM-guided evolution, showed a convergent improvement in editing efficiency. Further fusion of an ssDNA-binding domain (HMG-D) to evoMmeFz2 enhanced editing efficiency, surpassing the performance of engineered TnpB systems. Leveraging its compact size, we packaged the optimized evoMmeFz2 system into a single AAV vector and demonstrated robust dystrophin restoration in a humanized Duchenne muscular dystrophy (DMD) mouse model, achieving therapeutic-level in vivo editing. Our work establishes Fanzor2 as a programmable genome-editing tool and presents a combinatorial engineering strategy for RNA-guided nucleases, underscoring the transformative potential of integrating artificial intelligence with structural biology to advance precision medicine.
Results
Structure-guide optimization of ωRNA using AlphaFold3
Engineering gRNAs and ωRNAs has been demonstrated to effectively enhance genome-editing efficiency across diverse CRISPR-Cas and OMEGA systems14,18,22,26,27,28,29,30,36,37. Nevertheless, most compact RNA-guided systems employ gRNAs and ωRNAs that typically exceed 100 nucleotides (nt) in length and adopt intricate tertiary structures when complexed with their cognate RNA-guided nucleases. Current engineering approaches generally rely on high-resolution structural insights into native ternary complexes—comprising the RNA-guided nuclease, its associated RNA (gRNA or ωRNA), and the target DNA substrate—to inform rational design strategies. The advent of AlphaFold3 bridges critical gaps in structural biology by enabling accurate prediction not only of protein architectures but also of multi-component biomolecular interactions, including protein–nucleic acid interfaces. Leveraging this breakthrough, we used AlphaFold3 to predict the ternary complex structure of the MmeFz2–ωRNA system (Supplementary Fig. 1), providing structural insights into its assembly and interaction dynamics. As shown in Fig. 1a, the MmeFz2 ωRNA scaffold adopts a secondary structure consisting of two distinct stem–loop elements (S1 and S2) and a pseudoknot (PK) motif. The S1 and PK moieties likely engage in primary interactions with MmeFz2 (Supplementary Fig. 1a), whereas the S2 region forms an elongated stem–loop whose distal end exhibits minimal interaction with MmeFz2 and displays notable structural irregularities, such as imperfect base pairing and a polyuridine tract. Therefore, building on established principles from previous gRNA engineering efforts, we implemented two rational modification strategies across the ωRNA scaffold: MS1, structural stabilization by replacing non-canonical base pairs (G–U and A–U) with canonical G–C pairs in stem–loop elements; and MS2, transcriptional enhancement through targeted substitution of uridine residues in polyuridine tracts (notably U36–U40 and U49–U52), where uridines were replaced with non-uridine nucleotides37,38 (Fig. 1a).
a Predicted secondary structure of MmeFz2 ωRNA bound to B2M target dsDNA, with S1, S2, and PK regions highlighted for optimization. NTS, non-target strand; TS, target strand. b Experimental workflow for assessing MmeFz2 genome-editing activity at endogenous loci. c ωRNA optimization by replacing A-U or G-U base pairs in regions S1, S2, and PK. Data represent mean ± s.e.m. of three independent biological replicates. d Increased MmeFz2-mediated genome-editing efficiency resulting from uridine substitutions within the internal uridine-rich region (U49–U52) of S2. The high-efficiency ωRNA variants (UA49CG, UG50CG, UA51GC, and UA52GC) were selected for further optimization. Data represent mean ± s.e.m. of three independent biological replicates. e Combined effects of the top four modifications at positions U49–U52 in S2 on genome-editing efficiency. Data represent mean ± s.e.m. of three independent biological replicates. f Increased MmeFz2-mediated genome-editing efficiency resulting from uridine substitutions within the internal uridine-rich region (U36–U40) of S2. The high-efficiency ωRNA variants (UA36GC, UA37CG, UA37AU, UG38GC, UG38AU, UA39CG, UA39AU, and UA40CG) were selected for further optimization. g Combined effects of the top eight modifications at positions U36-U40 in S2 on genome-editing efficiency. Data represent mean ± s.e.m. of three independent biological replicates. h Synergistic modulation of genome-editing efficiency by combining the ten modifications described in MS1 and MS2. Data represent mean ± s.e.m. of three independent biological replicates. Fold-change represents the ratio of ωRNA variant editing efficiency to WT-ωRNA. The top 30% of mCherry-positive cells were FACS sorted to assess MmeFz2-ωRNA editing efficiency. Source data are provided as a Source Data file.
We first implemented MS1 and evaluated genome-editing efficiency at the endogenous B2M locus—previously identified as the most efficient target—using targeted-amplicon sequencing in HEK293T cells (Fig. 1a, b). Analysis revealed that the GU33GC variant (V1.1) exhibited a 3.8-fold increase in indel efficiency compared to the WT-ωRNA (Fig. 1c). Next, to systematically assess the impact of MS2 modifications, we introduced nucleotide substitutions across the polyuridine tracts. Substitutions converting U49–U52 nucleotides into G–C or C–G base pairs consistently enhanced indel efficiency by 2- to 3-fold (Fig. 1d). Two combinatorial mutants exhibited synergistic effects: UA49CG + UG50CG (V2.1) and UG50CG + UA51GC (V2.2) achieved more than a 4.5-fold increase in indel efficiency compared to the WT-ωRNA (Fig. 1e). In parallel, systematic substitution within the U36–U40 region revealed that more than half of the single-nucleotide variants conferred over a 3-fold improvement in indel efficiency (Fig. 1f). Remarkably, all substitutions at the UG38 position exhibited exceptional performance, resulting in an over 6-fold enhancement in indel efficiency (Fig. 1f). Building on these findings, we designed combinatorial variants by integrating the top-performing substitutions. Seven of these variants (V2.3–V2.9) achieved more than a 9.5-fold improvement in indel efficiency relative to the WT-ωRNA (Fig. 1g, Supplementary Fig. 2). Subsequently, systematic combinatorial integration of all enhancing modifications identified the ωRNA-V3 variant, which incorporates five substitutions (UG50CG, UA51GC, UA36GC, UA37AU, and UG38AU) and exhibits a maximal 16.7-fold enhancement in indel activity compared to the WT-ωRNA (Fig. 1h).
Structural analysis revealed minimal interactions between the distal end of the S2 region and MmeFz2 (Supplementary Fig. 1a), suggesting that truncation of this distal stem region might preserve functionality while potentially enhancing the cellular stability and expression of ωRNA. Building on this structural insight, we conducted a systematic truncation analysis of the distal stem-loop of S2, starting from ωRNA-V3. Initial screening using 33 small tiled deletions (2–3 bp) across the S2 region revealed that most modifications retained comparable or only slightly reduced editing efficiency relative to ωRNA-V3 (Fig. 2a, b). Guided by these findings, we constructed progressively extended truncations (1–19 bp) by sequentially removing nucleotides from the distal end of S2. Remarkably, three truncations (14–16 bp) not only preserved but also enhanced indel efficiency compared to ωRNA-V3 at the DYRK1A-guide1 locus (Fig. 2c, d). Subsequent evaluation across eight endogenous loci revealed that the 15-bp truncation variant (designated as en-ωRNA) outperformed both the other truncation variants (14/16 bp) and ωRNA-V3 (Fig. 2e, Supplementary Fig. 3a). qPCR analysis revealed that en-ωRNA transcripts accumulated to higher levels than both WT-ωRNA and ωRNA-V3 (Supplementary Fig. 3b), which may partially account for the observed enhancement in genome-editing efficiency. Structural remodeling of the optimized en-ωRNA revealed a more compact ternary complex with MmeFz2 and target DNA compared to the original configuration (Fig. 2f). Taken together, by integrating structural insights from AlphaFold3 predictions with rational ωRNA engineering, we achieved synergistic modifications that led to nearly a 20-fold enhancement in indel efficiency across multiple genomic loci, while simultaneously reducing the ωRNA scaffold length by 30%.
a Schematic of 2- and 3-bp truncations in the ωRNA S2 region. b Genome-editing efficiency of MmeFz2-ωRNA with 2- or 3-bp truncated ωRNA variants at the DYRK1A-guide1 locus in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. c Editing efficiency of ωRNA variants with 1-19 bp S2 truncations at the DYRK1A-guide1 locus in HEK293T cells. The three high-efficiency truncated variants (Del-14bp, Del-15bp, and Del-16bp) selected for further validation are indicated by red triangles. Data represent mean ± s.e.m. of three independent biological replicates. d Schematic diagram of 14-16 bp truncations within the S2 region. e Comparison of average genome-editing efficiency of the three high-efficiency truncated variants across eight endogenous loci in HEK293T cells. Each data point represents the average genome-editing efficiency at each target locus. f Schematic of the AlphaFold3-predicted ternary complex of MmeFz2, en-ωRNA, and DYRK1A-guide1 target dsDNA. Fold-change represents the ratio of ωRNA variant editing efficiency to ωRNA-V3. The top 30% of mCherry-positive cells were FACS sorted to assess MmeFz2-ωRNA editing efficiency. Source data are provided as a Source Data file.
Structure-guided engineering of MmeFz2 using AlphaFold3 or EVOLVEpro
Leveraging AlphaFold3-predicted structural models of the MmeFz2–ωRNA–DNA ternary complex, we propose that MmeFz2 consists of four distinct domains: a RuvC nuclease domain (residues 1–61, 265–429, and 457–478) containing an embedded zinc-finger motif (ZF, 429–457); a recognition domain (REC, 72–177); and a wedge domain (WED, 61–72 and 177–265) (Supplementary Fig. 1). We rationally designed 141 single-point variants at the protein–nucleic acid interface to either strengthen interactions between MmeFz2 and nucleic acids or promote conformational flexibility (Fig. 3b). Initial screening at the DYRK1A-guide1 locus identified 15 single-point mutants (C69K, C69R, Q158K, Q158R, S185N, E305N, E309Q, E309R, Y316R, E326N, E326Q, E326K, L356Q, S377N, S377Q) that exhibited >1.2-fold improvement in indel efficiency compared to WT-MmeFz2 (Fig. 3b). Validation across nine genomic loci revealed that C69K and C69R outperformed both WT-MmeFz2 and the other single-point variants (Fig. 3c, Supplementary Fig. 4a). Next, we combined C69K or C69R with secondary mutations (S185N, E305N, E309Q, E309R, Y316R, E326N, E326Q, E326K, L356Q, S377N, S377Q) to assess their synergistic effects. Notably, C69K exhibited superior cooperative potential than C69R in double-mutant combinations (Fig. 3d). Third-round engineering produced the triple mutant C69K + E305N + E326Q (designated en-Pro), which achieved a 2.1-fold increase in indel efficiency compared to WT-MmeFz2 (Fig. 3e).
a Schematic of the evolutionary engineering strategy for the MmeFz2 protein using AlphaFold3 or EVOLVEpro. b Comparison of genome-editing efficiencies mediated by MmeFz2 variants at the DYRK1A-guide1 locus in HEK293T cells. 141 mutations in MmeFz2 residues that may enhance the interaction between MmeFz2-ωRNA and target DNA (predicted by AlphaFold3). c Comparison of average genome-editing efficiencies of the 15 high-efficiency mutants across nine endogenous loci in HEK293T cells. d Comparison of genome-editing efficiencies mediated by combinations of C69K or C69R with the 11 selected mutants at the DYRK1A-guide1 locus in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. e Comparison of genome-editing efficiencies mediated by combinations of C69K and the seven selected mutants at the DYRK1A-guide1 locus in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. f Engineering of MmeFz2 through three rounds of EVOLVEpro. g Comparison of average genome-editing efficiencies of the ten high-efficiency mutants across nine endogenous loci in HEK293T cells. Five mutations (E178S, E178H, E305S, E305D, and E418R) were selected for further validation. h Comparison of genome-editing efficiencies mediated by different combinations of the five mutations at the DYRK1A-guide1 locus in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. i Combination of en-ωRNA with two engineered protein variants (en-Pro and evo-Pro) further increases genome-editing efficiency at the DYRK1A-guide1 locus in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. j Structural basis of the activity-enhancing mutations C69K, E326Q, and E178H. Fold-change represents the ratio of protein variant editing efficiency to WT-MmeFz2. The top 30% of mCherry-positive cells were FACS sorted to assess MmeFz2-ωRNA editing efficiency. For c and g each dot indicates the mean editing efficiency from three independent biological replicates at each endogenous locus. Source data are provided as a Source Data file.
To overcome the limitations of rational design, we implemented the active learning framework EVOLVEpro using indel efficiency data from the initial set of 141 rationally designed single-point mutants (Fig. 3a, b). Three iterative rounds of prediction and experimental validation identified ten variants (E178G, Y316T, E326A, E178H, E178S, E178Q, E178N, E305S, E305D, E418R) that exhibited ≥1.1-fold enhancement in indel efficiency at the DYRK1A-guide1 locus (Fig. 3f). Multi-locus validation demonstrated progressive efficiency gains across iterative evolution rounds, with top performers E178H, E178S, E305D, E305S, and E418R showing consistent improvements (Fig. 3g, Supplementary Fig. 4b). Combinatorial design of these mutations produced the triple mutant E178H + E305S + E418R (designated evo-Pro), which achieved a 2.0-fold increase in indel efficiency compared to WT-MmeFz2 (Fig. 3h).
Integration of the optimized en-ωRNA with engineered en-Pro and evo-Pro resulted in 76.0-fold and 66.1-fold improvements, respectively, over the original MmeFz2–ωRNA system (Fig. 3i). Structural predictions by AlphaFold3 suggest that the beneficial mutations from both approaches likely stabilize critical interactions between MmeFz2 and ωRNA (Fig. 3j). Specifically, C69K introduces new interactions between the WED domain and the sugar-phosphate backbone of the PK region in ωRNA (Fig. 3j). E326Q and E178H are each predicted to reinforce the interaction between the catalytic RuvC domain and the sugar-phosphate backbone of the spacer region (Fig. 3j). Collectively, these results show that AlphaFold3-guided rational design, combined with EVOLVEpro-driven protein evolution via few-shot active learning, synergistically enhances the genome-editing efficiency of the MmeFz2–ωRNA system. Moreover, they demonstrate that the PLM-based EVOLVEpro platform achieves competitive protein-engineering performance comparable to AlphaFold3-based approaches.
Engineered MmeFz2–ωRNA system enables efficient genome editing in human cells
Previous studies have demonstrated that fusing non-sequence-specific DNA-binding domains (ssDBDs) or exonuclease modules to RNA-guided nucleases can enhance genome-editing efficiency26,28,29,39,40,41,42,43. To further enhance the genome-editing activity of the MmeFz2–ωRNA system, we systematically evaluated five ssDBDs (HMG-D, HMGN1, HMGB1, H1G, and Sso7d) and three exonucleases (TREX1, TREX2, and T5 exonuclease) fused to the N- or C-termini of evoMmeFz2. Interestingly, fusion of ssDBDs to evoMmeFz2 overall enhanced genome-editing efficiency, whereas all exonuclease fusions impaired its activity (Fig. 4a, b). Among all tested ssDBDs, the 112-aa HMG-D domain from the high-mobility group family of chromosomal proteins in Drosophila melanogaster outperformed the others (Fig. 4a, b). The C-terminal fusion evoMmeFz2-HMG-D demonstrated maximal indel efficiency of 81.5% across five endogenous loci, providing an average 1.2-fold improvement over evoMmeFz2 (Fig. 4a, b).
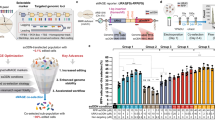
a Genome-editing efficiencies of evoMmeFz2 with N- or C-terminal ssDBD and exonuclease fusions at five loci in HEK293T cells. b Comparison of average genome-editing efficiencies of 16 evoMmeFz2 variants across five endogenous loci in HEK293T cells. c Comparison of genome-editing efficiencies induced by WT-MmeFz2, enMmeFz2, enMmeFz2-HMG-D, evoMmeFz2, and evoMmeFz2-HMG-D at 38 endogenous loci in HEK293T cells. Data represent mean ± s.e.m. of three independent biological replicates. d The summary dot plot compares the activities of these five variants in HEK293T cells. P values were calculated using an unpaired two-tailed Student’s t test, with adjusted P (Padj) values of 7 × 10−8, 1 × 10−9, 7 × 10−9, and 9 × 10−10, respectively. For b and d each dot represents the mean editing efficiency of three independent biological replicates per endogenous locus. All mCherry-positive cells were FACS sorted to assess the MmeFz2-ωRNA editing efficiency. Source data are provided as a Source Data file.
Next, we comprehensively evaluated genome-editing efficiencies of engineered MmeFz2–ωRNA systems across a panel of 38 endogenous loci spanning nine genes. All engineered MmeFz2–ωRNA systems exhibited significantly enhanced genome-editing efficiencies compared to the wild-type system (Fig. 4c, d). Specifically, enMmeFz2-HMG-D, evoMmeFz2-HMG-D, enMmeFz2, and evoMmeFz2 exhibited 40.2-, 37.1-, 30.1-, and 32.3-fold improvements, respectively, over WT-MmeFz2 (Fig. 4c, d). Notably, the HMG-D–containing variants (enMmeFz2-HMG-D and evoMmeFz2-HMG-D) showed slightly higher average indel efficiencies than the non-fusion enzymes across all tested loci (Fig. 4d). To benchmark the performance of engineered MmeFz2–ωRNA systems against other established compact editors, we compared these MmeFz2 variants with two previously characterized IS200/IS605 transposon-encoded TnpB nucleases, IsTfu1 and IsDge10. While all MmeFz2 variants recognize a 5’-TAG target adjacent motif (TAM), IsTfu1 and IsDge10 recognize distinct TAM sequences, 5’-TGAT and 5’-TTAT, respectively. For direct comparison, we selected endogenous loci with overlapping TAM sequences that are compatible with all systems (Supplementary Fig. 5). All engineered MmeFz2 variants exhibited significantly higher genome-editing efficiency than IsTfu1 TnpB and comparable efficiency to IsDge10 TnpB (Supplementary Fig. 6a, b). Collectively, these findings establish the engineered MmeFz2–ωRNA systems as potent mammalian genome-editing tools, offering broad target compatibility and markedly enhanced efficiency compared to both WT-MmeFz2 and established TnpB systems.
Evaluation of genome-editing specificity of the engineered MmeFz2–ωRNA system
To comprehensively evaluate the specificity profiles of evoMmeFz2 and its HMG-D fusion variant in human cells, we first identified potential ωRNA-dependent off-target sites at six endogenous loci—including KRAS, CXCR4, DYRK1A, and B2M—using the Cas-OFFinder algorithm44. Targeted-amplicon sequencing analysis revealed that the on-target editing efficiency of evoMmeFz2-HMG-D was higher than that of evoMmeFz2 across almost all six sites (Fig. 5). The engineered variants evoMmeFz2 and evoMmeFz2-HMG-D exhibited minimal but detectable off-target editing, with evoMmeFz2 showing 6.15% and 7.15% at DYRK1A-guide1 OT4 and B2M-guide2 OT7, respectively, and evoMmeFz2-HMG-D showing comparable levels of 5.81% and 6.04% at the same sites (Fig. 5). These findings revealed that the engineered MmeFz2 variants maintain reasonable genome-editing specificity in human cells, and HMG-D fusion does not exacerbate non-specific editing.
On- and off-target analyses of evoMmeFz2, evoMmeFz2-HMG-D, and WT-MmeFz2 were performed at six genomic loci (KRAS-guide1, CXCR4-guide2, DYRK1A-guide1, B2M-guide1, B2M-guide2, and B2M-guide5), with off-target sites containing 2 to 5 mismatches identified by Cas-OFFinder. Values are expressed as the mean of three independent biological replicates. Source data are provided as a Source Data file.
Engineered MmeFz2 variants restore dystrophin expression in a humanized Duchenne mouse model
DMD is a fatal muscular disease caused by dystrophin deficiency, affecting 1 in 3500–5000 newborn males, resulting from various pathogenic mutations in the human X chromosome-linked DMD gene45. In Duchenne muscular dystrophy, pathogenic variants of the DMD gene are enriched within exons 45–55, a region encoding the dystrophin rod domain46. Loss of one or more exons caused by DMD gene mutations frequently disrupts translational continuity, resulting in a shortened, nonfunctional dystrophin protein and progressive muscle pathology. Exon skipping can restore the open reading frame (ORF) by introducing small insertions or deletions through a single cut at a splice acceptor site (SAS) or splice donor site (SDS), leading to the removal of the targeted exon47. The engineered MmeFz2–ωRNA system can be packaged into a single rAAV vector owing to its compact size, making it a promising genome editor for in vivo correction of DMD.
Previously, we generated and validated a genetically humanized male DMD mouse model carrying a human-specific exon deletion mutation by knocking in the human exon 50 sequence to replace mouse exons 50 and 51 (designated as DMDΔmE5051, KIhE50/Y)48 (Fig. 6a). To evaluate the in vivo activity of evoMmeFz2 and evoMmeFz2-HMG-D, we first screened for efficient target sites in HEK293T cells and found that only the DMD-guide3 site lies within the SDS region of exon 50 (Fig. 4c). Next, we performed intramuscular (IM) injections of AAV9 particles carrying the expression elements for engineered MmeFz2-ωRNA systems into the tibialis anterior (TA) of 3-week-old male DMDΔmE5051, KIhE50/Y mice (Fig. 6a). Three weeks post-injection, we collected TA muscle samples for subsequent analysis (Fig. 6a). The DMD ORF can be restored by disrupting the SDS of exon 50, enabling the splicing of exon 49 to 52 or exon 50 to 52 in cases of exon 50 skipping or reframing (Fig. 6b). PCR-based detection across the transcript confirmed successful splicing alteration to skip human DMD exon 50 following evoMmeFz2 or evoMmeFz2-HMG-D-mediated indel formation, as verified by gel electrophoresis (Fig. 6c). Genomic editing analysis revealed average indel rates of approximately 0.22% for evoMmeFz2 and 0.57% for evoMmeFz2-HMG-D (Fig. 6d). Among these editing events, the productive 3n + 1 indels in exon 50 occurred at frequencies of 0.08% for evoMmeFz2 and 0.24% for evoMmeFz2-HMG-D, resulting in therapeutic-level dystrophin expression (Fig. 6d). RT-PCR analysis of mRNA extracted from whole muscle revealed that, in the evoMmeFz2-HMG-D group, the out-of-frame efficiency was 9.38 ± 0.82%, the in-frame efficiency was higher at 7.73 ± 0.46%, and the skipping efficiency was also elevated at 9.62 ± 0.43%, compared to the evoMmeFz2 group (Fig. 6e). Western blotting and immunostaining results further confirmed that dystrophin protein expression was efficiently rescued by evoMmeFz2 or evoMmeFz2-HMG-D (Fig. 6f, h; Supplementary Fig. 7). Additionally, evoMmeFz2-HMG-D treatment restored a higher number of dystrophin-positive muscle fibers and greater protein expression compared to the evoMmeFz2 group (Fig. 6g, i). Overall, our results demonstrate that the engineered MmeFz2-ωRNA system is not only effective in mammalian cells but also capable of efficiently restoring the causative protein expression via single-AAV delivery in vivo.
a Schematics of in vivo intramuscular (IM) injection of single AAV9-evoMmeFz2 or evoMmeFz2-HMG-D construct into the tibialis anterior (TA) muscle of the right leg of 3-week-old male DMDΔmE5051, KIhE50/Y mice. b Schematic representation of the exon-skipping strategy used by evoMmeFz2 or evoMmeFz2-HMG-D to restore the correct open reading frame (ORF) of the DMD transcript. c Gel electrophoresis was performed to analyze RT-PCR products from the muscle tissue of DMDΔmE5051, KIhE50/Y mice, and the experiment was repeated three times with similar results. Genomic (d) and RNA (e) indel editing events were analyzed by targeted-amplicon sequencing 3 weeks after intramuscular injection. f Immunofluorescence staining for DMD showed the restoration of dystrophin expression 3 weeks after TA injection of evoMmeFz2 or evoMmeFz2-HMG-D. The staining of dystrophin and spectrin proteins is depicted in green and purple, respectively. Scale bar: 100 μm. g Quantification of Dys+ fibers in cross sections of TA muscles. h Western blotting analysis was performed to assess dystrophin and vinculin expression in TA muscles 3 weeks after injection with AAV9-evoMmeFz2, AAV9-evoMmeFz2-HMG-D, or saline. Vinculin protein levels were used as an internal loading control. i The percentage of recovered dystrophin was quantified by grayscale intensity analysis. Data represent mean ± s.e.m. of three independent biological replicates. The p-value was determined using an unpaired two-tailed Student’s t test. Each dot represents an individual mouse for (d, e, g, and i). Source data are provided as a Source Data file.
Discussion
Fanzor proteins (Fanzor1 and Fanzor2) and prokaryotic CRISPR-Cas12 systems share evolutionary origins from distinct OMEGA-TnpB systems, despite having undergone independent evolutionary trajectories24,31,32,33,49,50. Structural analyses indicate that Fanzor2 retains greater homology to ancestral TnpB proteins than Fanzor1 (refs. 31,32,33). This enhanced structural conservation, coupled with its compact size (479 aa), makes Fanzor2 orthologs attractive candidates for single-AAV in vivo delivery in therapeutic applications31,32,33. However, the translational potential of most hypercompact Fanzor2 orthologs remains constrained by markedly low genome-editing efficiency in mammalian systems31.
Our study addresses this critical limitation through systematic engineering of both the ωRNA and MmeFz2 protein components. Building on established evidence that guide RNA architecture profoundly influences genome-editing efficiency across CRISPR and OMEGA systems14,22,27,28,29,37,51,52, we identified structural deficiencies in the WT-ωRNA, including internal poly-uridinylate tracts and destabilizing G–U wobble base pairs, based on the predicted ternary complex structure generated by AlphaFold3. Through base-pair substitutions and iterative truncations, we developed en-ωRNA, which exhibited nearly a 20-fold increase in genome-editing efficiency compared to the WT-ωRNA in human cells, along with a 30% reduction in scaffold size (Figs. 1h, 2d). Removal of the polyuridine tract in ωRNA-V3 and subsequent truncation of the stem-loop in en-ωRNA were associated with increased transcript abundance, as confirmed by qPCR (Supplementary Fig. 3b), indicating that these combined structural and transcriptional modifications contribute, at least in part, to the enhanced genome-editing activity.
Protein engineering is essential to biology and medicine, but current methods—such as directed evolution—are labor-intensive, prone to local optima, and poorly suited for complex objectives. Structure-guided rational design using AlphaFold and active machine learning based on protein language models (PLMs) have demonstrated the ability to efficiently enhance RNA-guided nuclease function with minimal experimental validation, thereby overcoming key limitations of traditional directed evolution. Together, the two approaches synergistically achieved approximately a 2.0-fold improvement in MmeFz2 activity over the wild type (Fig. 3e, h), with PLM-driven optimization showing particular promise for researchers without extensive structural biology expertise. As our study evaluated only a limited subset of predicted variants—potentially omitting more potent candidates—further exploration of a broader variant pool may yield deeper insights into the enzyme’s full functional potential. Comparative analysis with related systems (Cas12f and TnpB) suggests that ωRNA architecture represents the primary bottleneck limiting the efficiency of Fanzor nucleases13,22,36,37. By contrast, engineering the MmeFz2 protein had a comparatively modest effect, potentially due to the greater structural complexity of ωRNA in Fanzor systems compared to the sgRNA or crRNA used in Cas9 and Cas12a systems. Moreover, fusion of the HMG-D domain to the C terminus of evoMmeFz2 enhanced editing efficiency while maintaining specificity, underscoring its strong potential for practical applications. The mechanism by which the ssDBDs (HMG-D) enhance MmeFz2 activity remains to be elucidated, despite their known chromatin-binding function in native contexts.
Conventional CRISPR systems such as Cas9 and Cas12a, along with their derivative editors, exceed the AAV cargo capacity (>4.7 kb), whereas the compact MmeFz2–ωRNA system (1.4 kb coding sequence) enables single-AAV in vivo delivery. In this study, we explored targeting the SDS site adjacent to the end of exon 50 for exon skipping in the DMD gene using evoMmeFz2 or evoMmeFz2-HMG-D as a potential treatment for Duchenne muscular dystrophy. The results demonstrate that the engineered MmeFz2–ωRNA system can efficiently restore dystrophin expression in a humanized Duchenne mouse model via single-AAV delivery in vivo. In addition, their eukaryotic origin may also help reduce immunogenicity in humans. The broad distribution of Fanzor nucleases implies that numerous RNA-guided systems may still be undiscovered in eukaryotes, representing a valuable resource for future biotechnological applications. However, the smaller REC domain in MmeFz2, which can stabilize only a 12–15 bp heteroduplex compared with the ~20 bp stabilized by Cas12a, is likely a major contributor to its potential off-target effects. Therefore, strategies such as extending the REC domain via grafting from structurally similar nucleases or developing high-fidelity variants represent promising approaches to reduce off-target activity in therapeutic applications. Collectively, this study establishes a paradigm for integrating computational biology tools—such as AlphaFold3 and PLMs (AlphaPLM)—with experimental validation to optimize compact RNA-guided nuclease systems. While current efforts have focused on a limited subset of variants, expanding this approach could unlock more substantial functional enhancements. The demonstrated success in optimizing both the RNA and nuclease components underscores the versatility of this framework for advancing next-generation genome editors toward clinical translation.
Methods
Structure prediction by AlphaFold3
The sequences of the wild-type MmeFz2 protein and its cognate/engineered full-length ωRNA containing 20-nt B2M or DYRK1A guides, together with the corresponding 40-bp endogenous B2M or DYRK1A target DNA, were submitted to the AlphaFold3 web server (https://golgi.sandbox.google.com/) to predict the ternary complex structure. The resulted structure was fine-tuned by using COOT. Molecular visualization figures were generated using CueMol (http://www.cuemol.org).
EVOLOVEpro-driven enhancement of enzymatic activity
EVOLVEpro (https://github.com/mat10d/EvolvePro) was utilized for protein engineering involved a few-shot active learning framework integrated with structural insights from AlphaFold3 predictions. Initially, a dataset comprising 141 AlphaFold3-informed single-point mutants located at the interaction interfaces between MmeFz2 and nucleic acids, along with their corresponding activity data in mammalian cells, was collected (Fig. 3b). This dataset served as the initial input for EVOLVEpro’s regression model, which combines protein language model (PLM) embeddings (ESM-2 15B) with a random forest–based top-layer regression to map all MmeFz2 variations to relative activity compared to WT-MmeFz2. Then, the top 20 mutations predicted to exhibit high activity in the first-round analysis were selected for experimental validation of indel efficiency at the DYRK1A-guide1 locus in mammalian cells. The resulting activity data were subsequently incorporated to refine the model’s activity landscape predictions. This process was iteratively repeated for two additional rounds.
Plasmid vector construction
Plasmids were cloned using standard molecular cloning techniques. Human codon-optimized wild-type MmeFz2 and ωRNA scaffold were synthesized by HuaGene Co. Ltd. For MmeFz2-ωRNA plasmid construction, Phanta Max Super-Fidelity DNA Polymerase (Vazyme) was used for PCR, and the Basic Seamless Cloning and Assembly Kit (TransGen) for fragment assembly. Each plasmid includes a CMV enhancer, chicken β-actin promoter, 3×FLAG, SV40 NLS, MmeFz2 protein, nucleoplasmin NLS, bGH poly(A) signal, U6 promoter, and ωRNA in sequence. The ωRNA target oligonucleotides were ordered from Tsingke Biotechnology Co. Ltd, annealed and ligated into BsaI-digested backbone vectors using T4 DNA Ligase (Thermo). The spacer sequences of the ωRNAs used in this study are listed in Supplementary Data 1. The generated MmeFz2 ωRNA and protein mutants with efficient genome-editing activities are shown in Supplementary Data 2 and Data 3.
Cell culture, transfection, and flow cytometry analysis
Human HEK293T cells (CRL-3216) were purchased from the American Type Culture Collection (ATCC) and cultured in Dulbecco’s Modified Eagle Medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (Gibco), 1% non-essential amino acids (Gibco), and 1% Penicillin-Streptomycin-Glutamine (Gibco). All cell types were cultured at 37 °C with 5% CO2 and routinely passaged every 2 days before reaching 80% confluency. For screening experiments involving protein and ωRNA variants at the endogenous locus, 2 × 105 HEK293T cells were seeded in 24-well plates and transfected at approximately 80% confluence with MmeFz2 expression plasmids (1 µg) using 2 µl of polyethylenimine (PEI) at a 1:2 DNA (µg) to PEI (µl) ratio per well. After 18 h of incubation, 1 μg of plasmid was transfected using PEI (Polysciences) according to the manufacturer’s instructions. After 60–72 h, the transfected cells were digested with 0.05% trypsin (Gibco) for fluorescence-activated cell sorting (FACS), and top 30% or all mCherry-positive cells were used for genome extraction (Supplementary Fig. 8).
DNA extraction and indel efficiency analysis
Approximately 10,000 sorted cells were lysed with 20 μl of lysis buffer (10 mM Tris-HCl, pH 8.0; 0.05% SDS; 20 μg/ml proteinase K). The lysate was incubated at 55 °C for 30 min and then at 95 °C for 5 min, after which 1 μl of the cell lysate was used as the PCR amplification template. To evaluate the in vivo gene editing efficiencies of MmeFz2 variants, DNA was extracted from muscle tissues of successfully born MmeFz2-edited mice that had been treated with AAV9-evoMmeFz2-DMD ωRNA, using TIANamp Genomic DNA Kit (TIANGEN). To perform targeted-amplicon sequencing analysis, we amplified genomic regions ranging from 200 to 250 bp using nested PCR with Phanta Max Super-Fidelity DNA Polymerase P505 (Vazyme) and primers containing barcodes. The PCR products were pooled and then purified using a Gel extraction kit (Omega). The amplicon-seq libraries were prepared using the VAHTS Universal DNA Library Prep Kit (Vazyme), followed by purification and sequencing on an Illumina NovaSeq 6000 platform with 150-bp paired-end reads. The sequencing data were initially demultiplexed using Cutadapt (v.2.8) and subsequently processed with CRISPResso2 (v.2.0.20b) to quantify indel efficiency. Refer to Supplementary Data 1 for information on the target site sequence and primers.
Reverse transcription and quantitative real-time PCR (qPCR)
HEK293T cells transfected with plasmids expressing different ωRNA variants were harvested 48 h post-transfection. Total RNA was extracted using RNAiso Plus (Takara) according to the manufacturer’s instructions. RNA concentration and purity were assessed with a NanoDrop 2000 spectrophotometer (Thermo). Reverse transcription was performed using the HiScript IV 1st Strand cDNA Synthesis Kit (+gDNA wiper) (Vazyme) with 500 ng of total RNA per reaction. Quantitative PCR was carried out on a QuantStudio 1 Real-Time PCR System (Thermo) using the SupRealQ Purple Universal SYBR qPCR Master Mix (U+) (Vazyme) in a 20 µL reaction volume. The thermal cycling conditions were as follows: 50 °C for 2 min, 95 °C for 10 min, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. A melt-curve analysis was conducted following amplification to confirm the specificity of each reaction. RNA expression levels were normalized to GAPDH, and relative transcript abundance was calculated using the 2^–ΔΔCt method. Each sample was analyzed with three biological replicates and three technical replicates. Primer sequences are listed in Supplementary Data 5.
Animals
The mice were housed in a controlled barrier facility with a 12-h light/dark cycle, maintained at 18 to 23 °C and 40% to 60% humidity, with food and water available at all times. DMDΔmE5051, KIhE50/Y mice were generated on the C57BL/6J background using the CRISPR/Cas9 system. Given that Duchenne muscular dystrophy (DMD) is the most common sex-linked lethal disease in human male patients, male mice were chosen for this study.
Analysis of off-target sites predicted by Cas-OFFinder
To evaluate the specificity of MmeFz2 system, potential off-targets were predicted using CRISPR RGEN Tools (Cas-OFFinder, http://www.rgenome.net/cas-offinder/). We entered the 23-nt sequence of interest, comprising the 20-nt target sequence and the 3-nt TAM (5’-TAG), into the box, as no option was available for the TAM of the MmeFz2 protein. The mismatch was limited to five nucleotides, and the PAM sequence was set to 5’-NNN, consistent with SPRY Cas9. Potential off-target sites with one or more mismatches were selected for primer design using the online Primer-BLAST tool (https://www.ncbi.nlm.nih.gov/tools/primer-blast). The top 10 predicted potential off-target sites were PCR-amplified and sequenced to assess the genome-editing specificity of WT-MmeFz2, evoMmeFz2, and evoMmeFz2-HMG-D. All predicted off-target site sequences and their corresponding primers are provided in Supplementary Data 4.
Intramuscular injection
The adeno-associated virus 9 (AAV9) serotype was used in this study. The evoMmeFz2 and evoMmeFz2-HMG-D plasmids with ωRNA were sequenced before being packaged into the AAV9 vehicle, and the AAV vectors were subsequently packaged by transfecting HEK293T cells with pHelper, pRepCap, and GOI plasmids. AAVs were harvested and purified using iodixanol density gradient centrifugation after a three-day incubation. For intramuscular administration, 3-week-old DMDΔmE5051, KIhE50/Y mice were anesthetized, and the tibialis anterior (TA) muscle was injected with either 50 μl of AAV9 (1 × 1012 vg) preparations or an equivalent volume of saline solution. Mice were anesthetized and euthanized three weeks after injection, and tissues were dissected into distinct segments for targeted assessment. The distal region was analyzed for DNA editing and exon skipping efficiency, the middle for dystrophin expression via immunoblotting, and the proximal for dystrophin levels by immunofluorescence.
Western blot analysis
Muscle samples were incubated in RIPA Lysis and Extraction Buffer for lysis. Lysate supernatants were first quantified with the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific) and then adjusted to a uniform concentration with water. Ten micrograms of total protein per lane were separated using SDS-polyacrylamide gel electrophoresis. Samples were transferred to PVDF membranes for 3.5 h at 350 mA under wet conditions and then blocked with 5% non-fat milk in TBST for 1 h at room temperature. Immunoblots were incubated overnight with primary antibodies against dystrophin (Sigma, D8168) or vinculin (CST, 13901S) in TBST containing 0.05% BSA. Immunoblots were washed three times for 5 min each in TBST on a shaker, then incubated with HRP-conjugated IgG secondary antibodies for 1 h at room temperature. Finally, target proteins were visualized with chemiluminescent substrates (Invitrogen).
Histology and immunofluorescence
For histological analysis, paraffin-embedded tissue samples were first deparaffinized in xylene, then rehydrated through a gradient of ethanol from 100% to 50%. After washing in distilled water, the sections were stained with hematoxylin and eosin (H&E) and 0.1% picrosirius red solution for histological examination.
For Sirius red staining, slides were stained with picrosirius red for 1 h and then washed twice with acidified water. Most of the water was physically removed from the slides by vigorous shaking. After dehydration in three changes of 100% ethanol, the slides were cleared in xylene and mounted in neutral resin.
For immunofluorescence, tissues were embedded in optimal cutting temperature (OCT) compound and snap-frozen in liquid nitrogen. Sequential frozen sections (10 μm thick) were fixed at 37 °C for 2 h, permeabilized with 0.4% Triton-X in PBS for 30 min, and then blocked with 10% goat serum for 1 h at room temperature. Slides were incubated overnight at 4 °C with primary antibodies for dystrophin (Abcam, ab15277) and spectrin (Millipore, MAB1622). After washing with PBS thoroughly, samples were incubated for 3 h at room temperature with compatible secondary antibodies (Alexa Fluor 488 donkey anti-rabbit IgG or Alexa Fluor 647 donkey anti-mouse IgG from Jackson ImmunoResearch) and DAPI. Following a 15-min wash in PBS, the slides were sealed with Fluoromount-G mounting medium. All images were captured using a Nikon C2 microscope. The percentage of dystrophin-positive (Dys+) muscle fibers was calculated relative to the total number of spectrin-positive fibers.
Statistics and reproducibility
Data are presented as mean ± SEM from three independent biological replicates. Statistical analyses were performed using GraphPad Prism 9 (v.9.5.1) with an unpaired two-tailed Student’s t test. A p value < 0.05 was considered statistically significant. Each experiment was independently repeated at least three times with similar results. No statistical method was used to predetermine sample size. No data were excluded from the analyses. Cells were randomly assigned to test or control groups, and DMD mice undergoing gene-editing therapy were randomly allocated to control or AAV9-treated groups.
Ethics statement
Only male mice were used for all AAV injection experiments. All animal experiments were conducted in accordance with relevant ethical regulations and were approved by the Institutional Animal Care and Use Committee (IACUC) of HuidaGene Therapeutics Inc., Shanghai, China.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Next-generation sequencing data are available at the National Center for Biotechnology Information (NCBI) Sequence Read Archive database under the BioProject accession code PRJNA1259048. Source data are provided with this paper.
References
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Fellmann, C., Gowen, B. G., Lin, P. C., Doudna, J. A. & Corn, J. E. Cornerstones of CRISPR-Cas in drug discovery and therapy. Nat. Rev. Drug Discov. 16, 89–100 (2017).
Chen, K., Wang, Y., Zhang, R., Zhang, H. & Gao, C. CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu. Rev. Plant Biol. 70, 667–697 (2019).
Wang, J. Y. & Doudna, J. A. CRISPR technology: a decade of genome editing is only the beginning. Science 379, eadd8643 (2023).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362, 839–842 (2018).
Karvelis, T. et al. PAM recognition by miniature CRISPR-Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res. 48, 5016–5023 (2020).
Takeda, S. N. et al. Structure of the miniature type V-F CRISPR-Cas effector enzyme. Mol. Cell 81, 558–570.e553 (2021).
Wu, Z. et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat. Chem. Biol. 17, 1132–1138 (2021).
Wang, Y. et al. Guide RNA engineering enables efficient CRISPR editing with a miniature Syntrophomonas palmitatica Cas12f1 nuclease. Cell Rep. 40, 111418 (2022).
Kong, X. et al. Engineered CRISPR-OsCas12f1 and RhCas12f1 with robust activities and expanded target range for genome editing. Nat. Commun. 14, 2046 (2023).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Pausch, P. et al. DNA interference states of the hypercompact CRISPR-CasΦ effector. Nat. Struct. Mol. Biol. 28, 652–661 (2021).
Carabias, A. et al. Structure of the mini-RNA-guided endonuclease CRISPR-Cas12j3. Nat. Commun. 12, 4476 (2021).
Chen, W. et al. Cas12n nucleases, early evolutionary intermediates of type V CRISPR, comprise a distinct family of miniature genome editors. Mol. Cell 83, 2768–2780.e2766 (2023).
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692–696 (2021).
Nakagawa, R. et al. Cryo-EM structure of the transposon-associated TnpB enzyme. Nature 616, 390–397 (2023).
Sasnauskas, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023).
Li, Z. et al. Engineering a transposon-associated TnpB-ωRNA system for efficient gene editing and phenotypic correction of a tyrosinaemia mouse model. Nat. Commun. 15, 831 (2024).
Xiang, G. et al. Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors. Nat. Biotechnol. 42, 745–757 (2024).
Altae-Tran, H. et al. The widespread IS200/IS605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57–65 (2021).
Schuler, G., Hu, C. & Ke, A. Structural basis for RNA-guided DNA cleavage by IscB-ωRNA and mechanistic comparison with Cas9. Science 376, 1476–1481 (2022).
Han, D. et al. Development of miniature base editors using engineered IscB nickase. Nat. Methods 20, 1029–1036 (2023).
Yan, H. et al. Assessing and engineering the IscB-ωRNA system for programmed genome editing. Nat. Chem. Biol. 20, 1617–1628 (2024).
Han, L. et al. Engineering miniature IscB nickase for robust base editing with broad targeting range. Nat. Chem. Biol. 20, 1629–1639 (2024).
Xue, N. et al. Engineering IscB to develop highly efficient miniature editing tools in mammalian cells and embryos. Mol. Cell 84, 3128–3140.e3124 (2024).
Xiao, Q. et al. Engineered IscB-ωRNA system with expanded target range for base editing. Nat. Chem. Biol. 21, 100–108 (2025).
Saito, M. et al. Fanzor is a eukaryotic programmable RNA-guided endonuclease. Nature 620, 660–668 (2023).
Jiang, K. et al. Programmable RNA-guided DNA endonucleases are widespread in eukaryotes and their viruses. Sci. Adv. 9, eadk0171 (2023).
Xu, P. et al. Structural insights into the diversity and DNA cleavage mechanism of Fanzor. Cell 187, 5238–5252.e20 (2024).
Jiang, K. et al. Rapid in silico directed evolution by a protein language model with EVOLVEpro. Science 387, eadr6006 (2025).
Hennig, J. Structural biology of RNA and protein-RNA complexes after alphaFold3. Chembiochem 26, e202401047 (2025).
Su, M. et al. Molecular basis and engineering of miniature Cas12f with C-rich PAM specificity. Nat. Chem. Biol. 20, 180–189 (2024).
Kim, D. Y. et al. Efficient CRISPR editing with a hypercompact Cas12f1 and engineered guide RNAs delivered by adeno-associated virus. Nat. Biotechnol. 40, 94–102 (2022).
Gao, Z., Herrera-Carrillo, E. & Berkhout, B. Delineation of the exact transcription termination signal for type 3 polymerase III. Mol. Ther. Nucleic Acids 10, 36–44 (2018).
Zhang, X. et al. Increasing the efficiency and targeting range of cytidine base editors through fusion of a single-stranded DNA-binding protein domain. Nat. Cell Biol. 22, 740–750 (2020).
Yin, S. et al. Engineering of efficiency-enhanced Cas9 and base editors with improved gene therapy efficacies. Mol. Ther. 31, 744–759 (2023).
Yang, C. et al. HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing. Nat. Commun. 14, 2430 (2023).
Ding, X. et al. Improving CRISPR-Cas9 genome editing efficiency by fusion with chromatin-modulating Peptides. CRISPR J. 2, 51–63 (2019).
Yin, J. et al. Cas9 exo-endonuclease eliminates chromosomal translocations during genome editing. Nat. Commun. 13, 1204 (2022).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Roberts, T. C., Wood, M. J. A. & Davies, K. E. Therapeutic approaches for Duchenne muscular dystrophy. Nat. Rev. Drug Discov. 22, 917–934 (2023).
Flanigan, K. M. et al. Mutational spectrum of DMD mutations in dystrophinopathy patients: application of modern diagnostic techniques to a large cohort. Hum. Mutat. 30, 1657–1666 (2009).
Min, Y. L., Bassel-Duby, R. & Olson, E. N. CRISPR correction of Duchenne muscular dystrophy. Annu. Rev. Med. 70, 239–255 (2019).
Lin, J. et al. Adenine base editing-mediated exon skipping restores dystrophin in humanized Duchenne mouse model. Nat. Commun. 15, 5927 (2024).
Bao, W. & Jurka, J. Homologues of bacterial TnpB_IS605 are widespread in diverse eukaryotic transposable elements. Mob. DNA 4, 12 (2013).
Altae-Tran, H. et al. Diversity, evolution, and classification of the RNA-guided nucleases TnpB and Cas12. Proc. Natl. Acad. Sci. USA 120, e2308224120 (2023).
Dong, C., Gou, Y. & Lian, J. SgRNA engineering for improved genome editing and expanded functional assays. Curr. Opin. Biotechnol. 75, 102697 (2022).
Wu, Z. et al. Structure and engineering of miniature Acidibacillus sulfuroxidans Cas12f1. Nat. Catal. 6, 695–709 (2023).
Acknowledgements
We are grateful for the support from the Gene Editing Scientific Teaching (NWAFU-GEST), High-Performance Computing (HPC), and Life Science Research Core Service platforms (K.R. Huang, X.R. Liu, L. Chen, M. Zhou, and L.Q. Li) at Northwest A&F University (NWAFU). The authors also wish to express their gratitude to the members of HuidaGene Therapeutics Co., Ltd. for their contributions in supplying experimental materials and insightful discussions. This work is supported by the National Natural Science Foundation of China (32441080, 32301251 to Y.W. and 22207074 to Z.W.), the Biological Breeding-Major Projects (2023ZD04074 to K.X., 2023ZD04051 to Y.W., and 2022ZD04014 to X.W.), the National Key Research and Development Program of China (2023YFF1000904 to X.W.), the National Science and Technology Major Project of China (2023ZD0500500 to Z.W.), the China Agricultural Research System (CARS-39-03 to X.W.), and local grants (2024A02004-1-3, 2025NC-YBXM-109, and QCYRCXM-2023-104 to Y.W. and 2023A02011-2 to X.W.).
Author information
Authors and Affiliations
Contributions
Y.W., Z.W.W., K.X., and X.W. conceived the project. Y.W., Z.W.W., K.X., and X.L. designed the experiments. Y.W., S.L., P.G., and G.L. performed data analysis. Z.W.W. conducted the structural prediction analysis. S. L., P.G., Z.M.W., Y.Y., H.J., Y.F.C., Z.L., B.Z., and M.Z. performed cell transfection and FACS. Y.W. and G.L. performed animal experiments. Y.W. and Z.W.W. wrote the manuscripts. Y.W., X.W., Z.W.W., Y.L.C., K.X., and X.L. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chunyi Hu, Jun-Jie (Gogo) Liu and Hidetoshi Sakurai for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, S., Xu, K., Li, G. et al. Engineering the MmeFz2-ωRNA system for efficient genome editing through an integrated computational-experimental framework. Nat Commun 17, 1867 (2026). https://doi.org/10.1038/s41467-026-68644-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68644-5