Abstract
CRISPR-Cas12a, an RNA-based DNA targeting system, is widely used for genome editing and biomarker detection. To mitigate the off-target DNA cleavage of Cas12a, we previously developed a Francisella novicida Cas12a variant (FnoCas12aKD2P) by introducing double proline substitutions (K969P/D970P) in a conserved arginine-rich helix called the bridge helix (BH). In this work, we use a combinatorial approach to understand the molecular mechanisms of BH-mediated activation of Cas12a for DNA cleavage. We report five structures of FnoCas12aKD2P that are at different states of conformational activation. Comparison of the variant and wild-type (FnoCas12aWT) structures, along with activity assays and computational simulations, establishes the loop-to-helical transition and bending of the BH as an allosteric trigger for RNA-DNA hybrid propagation. These changes track with the previously reported coupled remodeling of BH and helix 1 of RuvC motif-II as well as the REC lobe movements needed to accommodate the growing hybrid. The transition of the BH is essential for the loop-to-helical transition of the “lid”, which in turn opens the RuvC active site pocket for DNA entry and cleavage. Pairwise 3D structural comparison of the BH and RuvC of Cas12 and Cas9 families provides insight into the diversity of BH’s structural organization in these mechanistically similar enzymes.
Similar content being viewed by others
Introduction
CRISPR-Cas systems are adaptive immune systems of bacteria and archaea that protect them from invading genomes1,2. While there are different types of CRISPR-Cas systems based on the type of nucleic acid target and protein composition, the fundamental principle is RNA-guided DNA/RNA targeting followed by cleavage of the invader genome3. Of the different types, Class 2 systems composed of single multi-domain proteins are widely used for gene editing, including gene therapy4,5. Cas9 (signature protein of the type II system) and Cas12a (signature protein of the type V-A system) are Class 2 proteins widely used for genome applications6,7. These proteins target DNA by recognizing a 20-nucleotide-long complementary region between the crRNA-guide and the target DNA. Both Cas9 and Cas12a use protospacer adjacent motif [PAM; a short DNA motif (2-8 nucleotides) flanking the 20-nucleotide region] present in the target to distinguish self versus non-self DNA3,8,9.
Cas9 and Cas12a have similar protein organization features with two lobes, a recognition (REC) lobe and a nuclease (NUC) lobe to perform the functions of RNA-DNA recognition and DNA cleavage, respectively10. The two lobes are bridged by a conserved arginine/lysine-rich helix, called the bridge helix (BH), and previous studies have shown the importance of BH in RNA-DNA binding and cleavage11,12,13,14,15,16,17,18. Each lobe has multiple domains, and in general, both Cas9 and Cas12a undergo conformational changes to accommodate the RNA-DNA hybrid in a groove between the two lobes, even though the exact conformational changes are different3. Step-by-step R-loop propagation enables both Cas9 and Cas12a to have conformational checkpoints along the conformational cascade towards achieving the pre-catalytic stage such that formation of the catalytically competent state is tightly related to a full RNA-DNA base pairing in the R-loop (17-nucleotides in Cas9 and 16-nucleotides in Cas12a)19,20,21.
Previous structural12,20,21,22,23,24,25,26, computational27,28, and Förster resonance energy transfer (FRET)16,17,20,29,30 studies covering different orthologs have provided in-depth information on the conformational activation process of Cas12a. The apo-form of Cas12a is elongated25 and it transforms into a compact and closed crab-claw shaped structure after binding to crRNA (binary form)23,26,31. After binding to the target DNA (ternary form)12,20,21,23,24,25, the closed structure opens up to accommodate the growing hybrid in between the two lobes16,30. Upon full R-loop formation, there is opening up of the RuvC catalytic pocket through the conversion of a loop form of the “lid” covering the active site to a helix form, which enables passage of single-stranded DNA (ssDNA) into the RuvC active site pocket for cleavage20,21,22,27.
Off-target DNA cleavage is a problem in gene editing and different protein engineering strategies are being used to create high-fidelity Cas proteins for genome applications. Previous studies from our lab have shown that the conserved BH of Cas9 and Cas12a can be used for rationale protein engineering to remove off-target DNA cleavage13,14,32,33. Specifically, we targeted a region of the BH that undergoes a loop-to-helical transition in response to RNA-DNA binding [based on comparing different structures of Cas9 (PDBs: 4OO811, 5F9R34, 4ZT035, 4CMP6) and Cas12a (PDBs: 6GTC20, 6GTG20, 5NFV23, 5MGA22, 5B4312, 5ID626, 6I1L24, 6I1K24 and 5XUS36)] and introduced prolines to disrupt the smooth helical transition. The resulting variants showed higher fidelity in DNA cleavage13,14,32,33.
In this work, we report five cryo-electron microscopy (cryo-EM) structures of a Francisella novicida Cas12a (FnoCas12a) variant with two proline substitutions in the BH (K969P/D970P; referred to as FnoCas12aKD2P). The structures unravel a mechanism by which the BH of Cas12a plays an integral part in the smooth transition of the different intermediate states to reach the final active conformation. Specifically, the loop-to-helical transition and the bending of BH enables the propagation of the RNA-DNA hybrid, which allosterically coordinates several downstream processes required to attain a cleavage-competent state, including the opening of the RuvC-lid. Biochemical and computational analyses show that BH’s helical transition is critical for the rest of the conformational changes, with a positively correlated movement of BH–RuvC-helix 1 (H1) movements with that of RuvC-lid, crRNA, and DNA. Interestingly, structural comparisons of the BH of Cas12 and Cas9 proteins show distinctions in the structural organization of BH and the RuvC-H1 implicating potential differences in the activation of these diverse, yet mechanistically similar, protein families. Since BH is conserved in several types of Cas proteins, as well as in other RNA-binding proteins such as RNA polymerases, our work provides molecular insights into the fidelity of DNA/RNA catalyzing enzymes.
Results
FnoCas12aKD2P samples different conformational states toward the DNA cleavage competent state
To assess how the conserved BH of Cas12a and its helical transitions promote conformational changes needed for DNA cleavage, we determined cryo-EM structures of FnoCas12aKD2P. We reconstituted FnoCas12aKD2P with its crRNA and a 24-nucleotide-long double-stranded DNA (dsDNA) substrate where the target strand (TS) will base pair (bp) with the guide region of the crRNA and the non-target strand (NTS) will exist as a ssDNA in the R-loop. The DNA possesses 4-nucleotides of PAM and 20-nucleotides of protospacer, that is numbered from 0 to 19 in the PDB files. The ternary complex was assembled in the absence of a divalent metal to prevent DNA cleavage and purified over size exclusion chromatography (SEC). The peak corresponding to the ternary complex, peak 2 (Supplementary Fig. 1), was used to prepare cryo-grids. We consistently got better distribution of the particles when we used graphene oxide-coated grids (Supplementary Method 1, Supplementary Fig. 2)37,38. A 3D map with ~2.96 Å resolution was created from ~400 K particles and the map possessed the typical crab-claw shape similar to the wild-type (FnoCas12aWT, PDB: 6GTG20). Further 3D classification generated six subclasses with an almost equal distribution of particles (14% to 18%, Supplementary Fig. 3), five of which ranged in resolution from 3.2 Å to 4 Å and were used for model building and analysis (Supplementary Fig. 4, Supplementary Table 1, and Supplementary Method 2). These five structures have differences in domain placements and represent conformational transitions needed to activate Cas12a for DNA cleavage (Fig. 1).
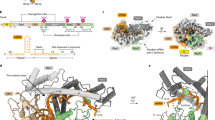
a Schematic representation of the domain architecture of FnoCas12a. The domain color scheme is maintained consistently throughout the manuscript: REC lobe [REC1 (light green), REC2 (dodger blue)] and NUC lobe [WED (tan), PI (PAM-interacting, salmon), RuvC (medium purple), BH (bridge helix, pink), RuvC-H1 (RuvC-helix 1, purple), and Nuc (gray)]. Nucleic acids are shown as: crRNA (orange), target DNA [non-target strand (NTS, deep sky blue), target strand (TS, blue), and protospacer adjacent motif (PAM, black)]. b–f Cryo-EM maps of FnoCas12aKD2P states (b), S1, (c), S2, (d), S3, (e), S4a, and (f), S4b each shown in two orientations. Major conformational difference is in the relative positioning of REC1 and REC2 domains across the different states that enables the opening and closing of the crab-claw structure and the accommodation of the RNA-DNA hybrid in between the two lobes. g–k Corresponding model built to fit the electron density map for states (g), S1, (h), S2, (i), S3, (j), S4a, and (k), S4b. A proposed order for the different states of FnoCas12aKD2P with respect to binary and pre-catalytic states of FnoCas12aWT is shown in Supplementary Fig. 10. Between S1 and S4b of FnoCas12aKD2P, the complex transitions through different amounts of opening of the crab-claw structure, as indicated by the distance between the REC2 and Nuc domains in solid lines (from 12 Å in S1, to 46 Å in S4a and 31 Å in S4b; distance shown is between C-\(\alpha\) atoms of D468 of REC2 domain and V1102 of Nuc domain). Additionally, the separation between the REC1 and REC2 domains also varies in the different states as shown in the dashed lines (from 39 Å in S1, to 77 Å in S4a and 71 Å in S4b; distance shown is between C-\(\alpha\) atoms K85 of REC1 domain and C473 of REC2 domain). See also Supplementary Tables 3 and 4. Figure was made using ChimeraX58.
The maps and models of FnoCas12aKD2P ternary complex from this study were compared with those of FnoCas12aWT from previous studies (Supplementary Table 2), including one that used single-molecule FRET (smFRET) analysis to identify the different conformational states (intermediate states I1-I5) towards attaining the pre-catalytic conformation20. Additionally, we compared FnoCas12aKD2P structures with Acidaminococcus sp. (As) Cas12a structure-series that captured different conformations at different lengths of the R-loop formation21. We named the different states of the FnoCas12aKD2P complex as state 1 (S1) to state 4b (S4b) (Fig. 1b–f). These structures provide mechanistic information about the role of BH in orchestrating the conformational cascade to activate Cas12a for DNA cleavage, as detailed below, including the opening of the lid covering the RuvC active site through direct interactions with the BH residues.
Comparison of the map features of all five states revealed that the major difference between them is in the movements of the contiguous REC1 and REC2 domains (Supplementary Movie 1) and visibility of REC2, which were previously shown to be important to open the closed binary complex of FnoCas12aWT to accommodate the RNA-DNA hybrid in between20,23,26. Notably, in state FnoCas12aKD2P-S1, REC2 lies in close contact with the Nuc domain, giving it a closed conformation (Fig. 1g and Supplementary Table 3), and the PAM-interacting (PI) domain remains open compared to the other states due to some disordered residues. Additionally, FnoCas12aKD2P-S1 does not show a continuous stretch of density for DNA, but there is residual density for positions 4 to 6 of the TS DNA (Supplementary Fig. 5a, b). An analysis of the seed region of the crRNA shows that there is preordering of this region similar to the binary complex (PDB: 5NG623, Supplementary Fig. 5c, d), indicating that the BH substitutions do not impact binary complex formation. These factors suggest that the FnoCas12aKD2P-S1 state represents initiation of the ternary complex formation, where the complex is in very early stages of PAM recognition.
In FnoCas12aKD2P-S2, the ternary complex transitions toward R-loop formation. The protein undergoes a closed-to-open transition, where the REC1 and REC2 domains begin to move away from and upward relative to the NUC lobe to accommodate the R-loop (Fig. 1h, Supplementary Movie 1, Supplementary Tables 3 and 4). During the transition of S1 to S2, the REC1 domain undergoes ~14° rotation with respect to a horizontal axis passing through the PI and Nuc domains (referred to as NUC-lobe axis hereafter, Supplementary Figs. 6 and 7a) and a translation of ~14 Å (distance between PI and REC1 domains). In contrast, the REC2 undergoes only a minor rotation of ~1° away from the NUC-lobe axis and 1 Å away from the Nuc domain, compared to that in S1 (Supplementary Fig. 7a, Supplementary Tables 5a, b). We observe 8-nucleotides of base pairing between crRNA and TS DNA in FnoCas12aKD2P-S2 (Supplementary Fig. 8a, b and Supplementary Table 6). However, due to the disordered PI domain, neither the density for the PAM sequence nor the rest of the NTS is visible. FnoCas12aKD2P-S2 resembles the FnoCas12aWT-I2 structure (PDB: 6GTD20; RMSD 3.6 Å, Supplementary Table 2).
In FnoCas12aKD2P-S3, the movements of the REC1 and REC2 domains increase the accessibility for the RNA-DNA hybrid (Supplementary Fig. 7b). The movement of the REC1 domain in FnoCas12aKD2P-S3 is similar to that observed in FnoCas12aWT-I2 and FnoCas12aWT-I3 states20 (Supplementary Table 5a). We observe a complete absence of the density for the REC2 domain in FnoCas12aKD2P-S3, and hence this domain was omitted from the model (Fig. 1d, i). While the REC2 domain was flexible in the FnoCas12aWT structures, none of them were captured with a complete absence of density for this domain. Similar states with a dislodged REC2 without density were captured for AsCas12a with 8- (PDB: 8SFI21) and 10- (PDB: 8SFJ21) bp of RNA–TS DNA hybrid during R-loop propagation, pointing to large movements needed for REC2 to accommodate the growing R-loop. In FnoCas12aKD2P-S3, 8-nucleotides, out of the total 15 nucleotides that are visible for the TS, are base-paired with the crRNA-guide region (3-nucleotides at the end of R-loop shows distorted pairing), and 3-nucleotides out of 7-nucleotides visible for the NTS exist as a single strand (Supplementary Fig. 8c and Supplementary Table 6). It is interesting to note that the PI domain is not ordered in FnoCas12aKD2P-S2 with 8-nucleotides of base pairing, but it is in FnoCas12aKD2P-S3 with similar base pairing and a visible NTS. This may correlate with the target DNA search mechanism of Cas12a, where establishing a critical amount of base pairing between the TS and crRNA-guide and passage of NTS into the hybrid channel may trigger PI ordering as a cue to further proceed with R-loop formation rather than dissociation and search for another potential DNA target39. The lysine helix-loop (LKL, residue 662-679) that was shown to initiate PAM unzipping in FnoCas12aWT (PDB:5MGA22) is observed in FnoCas12aKD2P-S3 with the lysine(s) interacting with the PAM nucleotides (Supplementary Fig. 9). However, these interactions are not observed in FnoCas12aKD2P-S1 and FnoCas12aKD2P-S2 due to the disordered PI domain. The FnoCas12aKD2P-S3 appears to be an intermediate state between FnoCas12aWT-I2 (3.6 Å RMSD between S3 and PDB: 6GTD20) and FnoCas12aWT-I3 (2.5 Å RMSD between S3 and PDB: 6GTE20) (Supplementary Table 2).
The remaining two states of the FnoCas12aKD2P structures are very close in conformations without significant differences to place them in order towards reaching the active state. These states closely resemble the intermediate state FnoCas12aWT-I320 (S4a: 5.2 Å RMSD and S4b: 4.3 Å RMSD, Supplementary Table 2 and Supplementary Fig. 10) and the two intermediate states of AsCas12a with 15- and 16-bp of R-loop formation (PDB: 8SFL and 8SFN21, respectively, Supplementary Table 2). In FnoCas12aKD2P-S4a, we see less well-defined density for REC2 (similar to the 15-bp complex of AsCas12a), with a density pattern showing stretched features consistent with the high flexibility of this region. With the current positioning of REC2 in FnoCas12aKD2P-S4a, we observe a rotation of ~17° upwards of the NUC-lobe axis and a translation of ~14 Å away from the Nuc domain, compared to that in FnoCas12aKD2P-S2 (Supplementary Fig. 7c and Supplementary Table 5b), with the distance between REC1 and REC2 increasing to ~77 Å (Fig. 1j and Supplementary Table 4). FnoCas12aKD2P-S4a has the most visible length of RNA-DNA hybrid among all five states, with 11 bps of hybrid between crRNA and TS DNA (4-nucleotides at the end of R-loop show distorted pairing) and 4-nucleotides of the single-stranded NTS (Supplementary Fig. 8d and Supplementary Table 6). FnoCas12aKD2P-S4b has the highest resolution (3.21 Å) among all five states and has clear density for the REC2 domain (Fig. 1f, k). The REC2 domain rotates downwards to the central cavity compared to its position in S4a (Supplementary Tables 3 and 5b, Fig. 1k and Supplementary Fig. 7d, e). In S4b, we see 9-bp of crRNA: TS DNA hybrid (5-nucleotides at the end of R-loop distorted) and 7-nucleotides of NTS DNA (Supplementary Fig. 8e and Supplementary Table 6).
Overall, comparing all five states of FnoCas12aKD2P, we observe that the maximum movement of the REC1 domain is the 14 Å translation between PI and REC1 domains between states S1 and S2, with a stable placement of REC1 and Nuc domains across all the states (Supplementary Table 5a, Supplementary Figs. 7 and 10). Comparatively, REC2 moves to a maximum of 34 Å in S4a relative to S1 and then moves closer (by 15 Å) to the NUC-lobe axis in S4b (Supplementary Table 5b, Fig. 1g–k distance indicated by solid line and Supplementary Fig. 7f). These movements increase the distance between the REC1 and REC2 domains ( ~ 35 Å between S1 and S4 states, Supplementary Table 4) as the system advances towards the pre-catalytic state (Fig. 1g–k distance indicated by dashed line, Supplementary Figs. 7 and 10). As the REC lobe opens, the R-loop grows in-between the two lobes.
Sampling of the pre-catalytic state is reduced in FnoCas12aKD2P
We compared the conformational states of FnoCas12aKD2P with those of FnoCas12aWT from different studies (Supplementary Table 2). FnoCas12aKD2P-S1 appears to be a conformation between the FnoCas12aWT-binary23 and the FnoCas12aWT-I1 (PDB: 6GTC20), which is the initial stage of ternary complex formation based on smFRET studies (Supplementary Fig. 10). FnoCas12aKD2P-S2 is similar to FnoCas12aWT-I2 due to the similar positions of REC1 and REC2 domains (Supplementary Table 4). There is a ~ 7° rotation of REC2 towards the NUC-lobe axis in FnoCas12aWT-I4 compared to that in FnoCas12aWT-I320 (Supplementary Table 5b). The additional states of FnoCas12aKD2P-S3 and FnoCas12aKD2P-S4a likely represent intermediate states between FnoCas12aWT-I2 and FnoCas12aWT-I3 that were captured during this rotation of REC2, supported by the weak density for REC2 in these states. While there is ordering of the REC2 density in state FnoCas12aKD2P-S4b, only 9-nucleotides of base pairing with the crRNA guide are visible, compared to 11-nucleotides of base pairing in FnoCas12aKD2P-S4a (Supplementary Fig. 8d, e and Supplementary Table 6). Overall, S4a and S4b of FnoCas12aKD2P appear to be very transient states towards attaining a pre-catalytic state (FnoCas12aWT-I4)20 (Supplementary Fig. 10). While we captured all the different states with an equal probability (Supplementary Fig. 3f), we did not observe a pre-catalytic state where the RNA-DNA hybrid is ordered, along with more visibility for NTS closer to the RuvC active site pocket, and also with the critical loop-to-helical transitioning of the lid region of the RuvC domain (RuvC-lid) to open the catalytic pocket. An impaired BH due to the double proline substitutions appears to reduce the efficiency of conformational transitions to reach the catalytically competent state, which also enabled capturing of sub-intermediate states that were to the best of our knowledge not previously observed. This observation supports the lower rate constant that was observed for DNA cleavage by FnoCas12aKD2P compared to that by FnoCas12aWT (3-fold reduction in on-target DNA cleavage)14.
Impaired BH restricts conformational checkpoints during R-loop formation
Two regions present in the REC1 domain, REC-linker (324-331) and finger (301-309), were shown to sense the RNA-DNA hybrid formation through PAM-proximal and PAM-distal interactions, respectively20. The movement of the REC-linker across the states in FnoCas12aKD2P remains similar to that in FnoCas12aWT (Supplementary Fig. 11). In FnoCas12aKD2P-S1, the REC-linker is positioned away from the crRNA, similar to the R-loop initialization complex of FnoCas12aWT (PDB: 6GTC20) and subsequently shifts closer to the crRNA in states S2 to S4b of FnoCas12aKD2P (G5 to C7, PAM-proximal region, similar to that in FnoCas12aWT-I4, PDB: 6GTG20), with a concomitant R-loop propagation in the later states (Supplementary Fig. 11 and Supplementary Table 7). The finger region moves closer to the PAM-distal region of the RNA-DNA hybrid during the transition of FnoCas12aKD2P from states S1 to S4b. Despite this movement, docking of the finger region onto the crRNA at nucleotide positions 15–17, which was identified as a delayed checkpoint towards the catalytic activation of Cas12a20,22, is not observed in the structural transitions that were captured for FnoCas12aKD2P. Overall, while the interaction of the REC-linker at the PAM-proximal side is intact in FnoCas12aKD2P, the interaction of the finger region at the PAM-distal side is impaired.
A comparison of the RNA-DNA hybrid in different states of FnoCas12aWT and FnoCas12aKD2P shows that the BH is essential for RNA-DNA hybrid propagation and proper base pairing. An analysis of the distance evolution between the BH and the RNA-DNA hybrid is shown in Supplementary Table 8. K969 is at a distance of 34 Å from U4 of the crRNA in the FnoCas12aWT-binary structure (PDB: 5NG6)23, and the corresponding distance is 34 Å between P969 and A4 in FnoCas12aKD2P-S1, showing similar positioning of the crRNA in these two states (Supplementary Table 8). In the pre-catalytic state of FnoCas12aWT-I4, the distance between the RNA-DNA hybrid gets closer to K969 ( ~ 6 Å with U11) when the R-loop has a full 20-nucleotides of RNA-DNA base pairing20 (Fig. 2a, b, and Supplementary Table 8). In FnoCas12aKD2P-S4a and FnoCas12aKD2P-S4b, the RNA-DNA hybrid remains farther (20 Å and 27 Å, respectively) from the BH due to the inability of the BH to undergo the loop-to-helical transition, which prevents its bending towards the hybrid (Fig. 2a, c, Supplementary Fig. 12 and Supplementary Table 8).
a Close-up view of the BH (bridge helix, pink) and RuvC-H1 (RuvC-helix 1, purple) regions and crRNA (orange)-TS DNA (blue) hybrid in FnoCas12aKD2P (state S4b; darker shades) overlaid with the FnoCas12aWT (state I4, PDB: 6GTG20; lighter shades). For FnoCas12WT, nucleotides U6 to C19 of crRNA and dC-5 to dG-19 of the TS DNA are shown; for FnoCas12aKD2P, U6 to U11 nucleotides of crRNA and dC-5 to dT-15 of TS DNA are shown, since the rest are disordered in the structure. Note that the protospacer numbering is from 0 to 19 to cover the 20 nucleotides. In FnoCas12aWT, the BH undergoes a loop-to-helix transition and bends ~28° to latch onto the RNA-DNA hybrid. This transition is absent in FnoCas12aKD2P. Interactions of RNA-DNA hybrid with (b), FnoCas12aWT, (c), and FnoCas12aKD2P at ~60° rotation about the x-axis compared to that in (a). In FnoCas12aWT, the RNA-DNA hybrid maintains a helical structure and lies closer to the BH [K969 at 6 Å to U11 of crRNA and 16 Å to dA(−11) of TS-DNA]. In FnoCas12aKD2P, the RNA-DNA hybrid is distorted and positioned farther ( ~ 20 Å) from P969 of the BH. A twist in the RNA-DNA backbone path at position 11 arises from interactions with the BH segment undergoing the loop-to-helical transition in FnoCas12aWT, but this is absent in FnoCas12aKD2P (See Supplementary Fig. 12 and Supplementary Tables 8 and 9 for details on elongation and bending of the BH towards the R-loop and melting of RuvC-H1 in other states of FnoCas12aKD2P and FnoCas12aWT). Also see Supplementary Fig. 21 and Supplementary Tables 12, 13 and 14 for detailed dynamic cross-correlation network analyses of the BH–RuvC-H1 region and RNA–DNA complex obtained from MD simulations of FnoCas12aKD2P and FnoCas12aWT ternary complexes. Figure was made using ChimeraX58.
In FnoCas12aKD2P states, RNA-TS DNA base pairing pattern is impacted. In FnoCas12aKD2P-S2, a region of the PI domain (S695-I714) is disordered, and there is no clear density for the PAM sequence of the DNA compared to clear visibility of 4-nucleotides of PAM and 9-nucleotides of RNA-DNA hybrid in FnoCas12aWT-I120. In FnoCas12aKD2P-S2 and FnoCas12aKD2P-S3, after dG(−7), the R-loop exhibits unstable base pairing. However, for FnoCas12aKD2P-S4a and FnoCas12aKD2P-S4b, base pairing is stable until position dA(−11) and dA(−10) of TS DNA, respectively, with the corresponding complementary position of the crRNA-guide, and the remaining visible nucleotides (4 in FnoCas12aKD2P-S4a and 3 in FnoCas12aKD2P-S4b) being in a single stranded condition (Supplementary Fig. 8). We observe stacking and distortion of the single-stranded form of the crRNA and the TS DNA in FnoCas12aKD2P structures, beyond the base paired region. This indicates that the BH conformational change is essential for progressive R-loop formation, stable base pairing of the RNA-DNA hybrid, and to enable helix formation of the RNA-DNA hybrid.
Cooperativity in the conformational transitions of BH and RuvC-helix 1 is critical for Cas12a’s activity
It was previously reported that BH and RuvC-helix 1 (RuvC-H1) of RuvC motif-II undergo coordinated conformational transitions while transforming from the binary to the ternary complex16,17,21. A comparison of the 8-bp R-loop state (FnoCas12aWT-I3)20 and the 20-bp R-loop state (FnoCas12aWT-I4)20 of the ternary complex structures shows that there is melting of RuvC-H1 upon partial R-loop formation in state I3, followed by elongation of the BH by 3 turns in state I420. In addition, the BH-end (D970) bends by 28° and moves ~20 Å towards the RNA-DNA hybrid after full R-loop formation compared to the position in the partial R-loop structures (Fig. 2a–c, Supplementary Fig. 12, and Supplementary Tables 8 and 9). Comparing the structure of Cas12a’s BH in the different states of FnoCas12aKD2P in our study with that of wild-type, we find that, while the RuvC-H1 melting is unaffected, the loop-to-helical transition of the BH is impaired due to the proline substitutions. The impairments in the elongation and bending of the BH keep it positioned at a distance of ~20 Å away from the RNA-DNA hybrid in both S4a and S4b of FnoCas12aKD2P (compared to ~17 Å in FnoCas12aWT-I1 and ~6 Å in FnoCas12aWT-I420, Supplementary Table 8).
Another important conformational change towards activation of Cas12a is the insertion of the conserved W971 (residue immediately following the proline substitutions of FnoCas12aKD2P at 969 and 970) into the REC2 hydrophobic pocket (composed of amino acids Y579, K527, K524, R583 of FnoCas12aWT 12,16,22) to enable REC2 docking after hybrid formation. In FnoCas12aKD2P structures, we see insertion of W971 into the REC2 hydrophobic pocket in S1 and S4b (S2 and S4a do not show density in this area to determine sidechain position). However, the interacting distance of W971 with Y579 of the REC2 domain, which is at ~4 Å in FnoCas12aWT-I4, remains farther in S1 and S4b of FnoCas12aKD2P (6 Å and 10 Å, respectively) (Supplementary Fig. 13). It can be envisioned that W971 will be sturdier in a completely elongated BH than when present in an impaired BH. These observations indicate that impairing the loop-to-helical transitioning of BH potentially prevents sampling of the activated state in our cryo-EM classes due to the absence of an elongated and bent BH to latch on to the RNA-DNA hybrid and due to the potential hindrance in coordinating the distinct REC2 movements across the different states through locking of W971 with the REC2 domain.
BH is an allosteric regulator that connects RNA-DNA hybrid formation to the opening of the lid of the RuvC active site
Previous studies have shown that a region called the RuvC-lid (residues 1009 to 1021 in FnoCas12a) occludes the active site pocket of the RuvC during the earlier states of the conformational cycle by existing as a loop. To attain the pre-catalytic state, the RuvC-lid undergoes a loop-to-helical transition, exposing the RuvC active site for DNA access20,21,24. A comparison of the different states of FnoCas12aKD2P with that of FnoCas12aWT provides interesting insights into the cooperative action of the BH and the RuvC-lid to open the RuvC active site for DNA access.
Compared to the conformation in the binary FnoCas12aWT structure23, region comprising E961-D970 of the BH undergoes loop-to-helical transition in response to full R-loop formation in the ternary complex20,23. Similar to AsCas12a, in FnoCas12aWT, the residue E1020 in the RuvC-lid forms an electrostatic interaction with K981 in RuvC-H1, stabilizing the loop form of the RuvC-lid and occluding the RuvC active site in partial R-loop states (until ~15-bp of R-loop formation as observed in AsCas12a)20,21,27. This interaction is disrupted in the pre-catalytic state in FnoCas12aWT-I4 with the concomitant formation of additional interactions between BH and RuvC-lid (R964/R968-E1020) and RuvC-H1 and RuvC-lid (K981-K1021), thus opening the RuvC active site to ssDNA20,21,31 (Fig. 3a–d, Supplementary Figs. 14, 15 and Supplementary Table 10). To understand the importance of these interactions in Cas12a’s activation, we generated alanine substituents of R964, E1020, K1021, and E1020A/K1021A in both FnoCas12aWT and FnoCas12aKD2P backgrounds. R968 was not substituted due to its documented role in binary complex assembly and loss of activity upon alanine substitution7. Recombinantly purified proteins (Supplementary Fig. 16) were used to perform plasmid cleavage assays with different substrates: matched DNA (MCH, complete complementarity with the guide region of the crRNA) and with two different DNA substrates with mismatches (MM2: mismatch at position 2 proximal to the PAM and MM13, mismatch at position 13 from the PAM) (Fig. 3e, f, Supplementary Fig. 17).
a Highlighted ribbon representation of the BH (bridge helix, pink), RuvC-H1 (RuvC-helix 1, purple), and RuvC-lid (medium purple) regions within the overall structure of state S4b of FnoCas12aKD2P shown as a surface depiction. The nucleic acids were omitted for clarity. b Close-up view of the BH, RuvC-H1 and RuvC-lid regions of FnoCas12aKD2P (state S4b; darker shades) overlaid with the corresponding regions from FnoCas12aWT (PDB: 6GTG20; lighter shades). In FnoCas12aWT, both BH and the RuvC-lid have undergone helical transitions, but this is not seen in FnoCas12aKD2P. c, d Close-up view of panel b rotated 120° about the z-axis relative to (b). c In FnoCas12aKD2P S4b, K981 of the RuvC-H1 interacts with E1020 of the RuvC-lid that is in the loop form. d In FnoCas12aWT pre-catalytic state (I420), both BH and RuvC-lid are in helical form. Several distinct contacts are observed in this state compared to that in S4b: E961 of the BH interacts with K1021 of the RuvC-lid, R964 of the BH with E1020 of the RuvC-lid, and R968 of the BH with E1020 of the RuvC-lid. K981 of the RuvC-H1 forms a distinct contact with N1022 of the RuvC-lid and loses its interaction with E1020 of the RuvC-lid in S4b state of FnoCas12aKD2P. Distances for these interactions are shown in the figure (see Supplementary Figs. 14 and 15 for additional interactions between BH, RuvC-lid and RNA/DNA). e, f Graphs representing linearization of DNA substrates over time for the FnoCas12a protein variants: e matched (MCH) and (f), mismatched (MM13, mismatch at position 13). (Also see Supplementary Table 11 and Supplementary Figs. 17 and 18 for additional information on total cleavage including nicked and linear species, and cleavage efficiency of the variants with MM2 substrate having a mismatch at position 2). The different time points are shown in minutes: 1 (light blue), 2.5 (gray), 5 (pink), and 15 (light green). Each column represents the average of three independent replications; black diamond markers indicate values for individual replications; error bars denote S.E.M. Variants: Wild-type (WT), K969P-D970P (KD2P), E1020A (EA), K1021A (KA), R964A (RA), E1020A-K1021A (EA/KA). Figures (a–d), and graphs (e, f), were made using ChimeraX58 and Microsoft Excel respectively. Raw data and calculations used for generating the graphs are provided as a Source Data file.
All FnoCas12aWT variants (E1020A, K1021A, R964A, E1020-K1021A) retain high linearization activity ( ≥90% at 15 min) that is comparable to wild-type for MCH substrate. In the FnoCas12aKD2P background, E1020A and K1021A variants show similar linearization ( ~ 85% at 15 min) to FnoCas12aKD2P–MCH, which has a slower rate compared to FnoCas12aWT–MCH substrate. FnoCas12aKD2P-E1020A/K1021A shows reduced linearization (47% at 15 min), with a 2-fold and 1.8-fold decrease compared to FnoCas12aWT–MCH and FnoCas12aKD2P–MCH, respectively (Fig. 3e, Supplementary Fig. 17a and Supplementary Table 11a). FnoCas12aKD2P-R964A displays a pronounced loss of linearization (12% at 15 min), representing an 8-fold decrease relative to FnoCas12aWT–MCH and a 7-fold decrease relative to FnoCas12aKD2P–MCH. Total cleavage is also low for FnoCas12aKD2P-E1020A/K1021A (68% at 15 min) and FnoCas12aKD2P-R964A (38% at 15 min), with the latter accumulating 26% nicked intermediate (Supplementary Fig. 18a and Supplementary Table 11a). These results indicate that in the absence of a BH loop-to-helical transition, R964A and E1020A/K1021A substitutions in FnoCas12aKD2P variants reduce the efficiency of DNA cleavage, with a more profound effect in R964A. Additionally, it also shows that in FnoCas12aWT, which possesses an intact transitioning of the BH, disrupting the BH–RuvC-lid interactions do not critically impact on-target DNA cleavage.
Similar to FnoCas12aWT–MCH condition, all FnoCas12aWT variants (E1020A, K1021A, R964A, E1020-K1021A) maintain efficient linearization of MM2 substrate (82-90% at 15 min). FnoCas12aKD2P-E1020A and FnoCas12aKD2P-K1021A show moderately reduced linearization (62-70% at 15 min) relative to FnoCas12aWT–MM2 but are similar to FnoCas12aKD2P–MM2 (65% at 15 min). FnoCas12aKD2P-E1020A/K1021A has a reduced linearization (40% at 15 min) and an accumulation of nicked intermediate (38%) compared to FnoCas12aKD2P–MM2 (23% nicking, 65% linearization at 15 min) (Supplementary Figs. 17b, 18b and Supplementary Table 11b). FnoCas12aKD2P-R964A–MM2 exhibits severely impaired linearization (4%), with 22-fold and 16-fold reductions relative to FnoCas12aWT–MM2 and FnoCas12aKD2P–MM2, respectively, with total cleavage at 45% (41% nicking at 15 min) (Supplementary Fig. 18a, b and Supplementary Table 11b). These data suggest that combining FnoCas12aKD2P with R964A or E1020A/K1021A enhances sensitivity to PAM-proximal mismatches.
With an MM13 mismatch, FnoCas12aWT variants E1020A, K1021A, and E1020A/K1021A retain linearization comparable to FnoCas12aWT–MM13 (84-90% at 15 min), but FnoCas12aWT-R964A shows a sharp reduction (12%, 7.5-fold lower than FnoCas12aWT–MM13) with 78% nicked intermediate accumulation, yielding a total cleavage similar to FnoCas12aWT–MM13 ( ~ 90%). FnoCas12aKD2P-E1020A and FnoCas12aKD2P-K1021A, similar to FnoCas12aKD2P, exhibit poor linearization of MM13 (0–5% at 15 min). This represents 29-fold and 43-fold reduction, respectively, for these variants, compared to their efficiency to linearize MCH, and 21-fold and 35-fold reduction, respectively, in their efficiency to linearize MM2 (Fig. 3 f, Supplementary Fig. 17c and Supplementary Table 11c). Similar to FnoCas12aKD2P–MM13, these variants accumulated high nicked intermediate ( ~ 85%), with a total cleavage of 88–90% at 15 min. Notably, FnoCas12aKD2P-R964A and FnoCas12aKD2P-E1020A/K1021A are defective in both strand cleavages, with 11% and 40% nicked intermediates, and linearization limited to 1% and 0%, respectively, at 15 min (Supplementary Fig. 18a and Supplementary Table 11c). These data indicate that R964A severely impairs second-strand cleavage in the FnoCas12aWT background for mid-R-loop mismatches. However, in the FnoCas12aKD2P background, R964A and E1020A/K1021A substitutions impair cleavage of both DNA strands in all substrates that were tested, including on-target and off-target substrates.
Our findings show that contacts between the BH and RuvC-lid (R964-E1020, E961-K1021) are only secondary to the BH’s helical transition for orchestrating the conformational transitions required for double-strand cleavage and mismatch sensitivity. The wild-type protein with an intact conformational change in the BH can linearize (i.e., both NTS and TS cleavages) despite disrupting the BH–RuvC-lid interactions, with the only significant reduction being in the second strand cleavage for MM13 substrate (Supplementary Figs. 17, 18a, and Supplementary Table 11). On the contrary, when BH’s sturdiness is restricted, as in the FnoCas12aKD2P variant, the enzyme becomes highly dependent on these interactions to provide RuvC active site access to ssDNA. In the FnoCas12aKD2P background, while R964A and E1020A/K1021A substitutions impair both strand cleavages, single substitutions (E1020A or K1021A) impede second-strand cleavage (Supplementary Fig. 18a and Supplementary Table 11). These results demonstrate that the BH’s loop-to-helical transition is an allosteric switch coupling R-loop propagation to catalysis and point to the importance of the helicity of the BH in assisting the passage of TS DNA into the RuvC active site pocket. BH, RuvC-H1, and RuvC-lid are potential engineering sites to reduce mismatch tolerance in Cas12a.
Impaired loop-to-helical transition of the BH creates conformational states that are distinct from those of the wild-type protein
Using Molecular Dynamics (MD) simulations and principal component analysis (PCA), we explored the similarities and differences of the conformations of states S2 and S4b of FnoCas12aKD2P with state FnoCas12aWT-I420 (Supplementary Method 3 and Supplementary Fig. 19). The resulting 2D projection of first and second principal components (PC1 and PC2, respectively, Supplementary Movies 2 and 3) reveal distinct clustering for each state, indicating well-defined conformational ensembles (Fig. 4). Furthermore, along PC1 (79.2%), a progression is observed from S2, through S4b, to finally attain the I4 conformation, delineating a pathway consistent with the expected structural transitions toward the pre-catalytic state of Cas12a (Fig. 4a). Also, these MD simulations support the notion that FnoCas12aKD2P-S2 and FnoCas12aWT-I220 states are closely related, while FnoCas12aKD2P-S4b represents a distinct intermediate conformation between FnoCas12aWT-I2 and FnoCas12aWT-I4 (Supplementary Fig. 20). An analysis of the structural differences of the region comprising the BH, RuvC-H1, and RuvC-lid shows the main differences in conformation contributing to PC1, representing 79.2% of the total variance (Fig. 4b and Supplementary Movie 2). In FnoCas12aKD2P-S2, BH is shorter, RuvC-H1 is longer, and RuvC-lid is in the loop form. In FnoCas12aWT-I420, all features of the active state are present: elongated and bent BH, melted RuvC-H1, and a helical RuvC-lid. FnoCas12aKD2P-S4b shows features that are intermediate to FnoCas12aKD2P-S2 and FnoCas12aWT-I420 states. In FnoCas12aKD2P-S4b, BH is longer than that in FnoCas12aKD2P-S2, RuvC-H1 has started to melt, while the RuvC-lid is still in the loop form. The PC2 captures complementary motions of the same regions but accounts for only 16.6% of the total variance (Supplementary Movie 3). These results show that coordinated and complete transitions of the BH and RuvC-H1 are a prerequisite for RuvC-lid’s helical transition. The complete analysis, including state FnoCas12aWT-I220 that lack sidechains in the experimental structure, is available in the SI (Supplementary Method 3 and Supplementary Figs. 19, 20).
a Principal component analysis (PCA) of systems FnoCas12aKD2P states S2 (purple) and S4b (green), and FnoCas12aWT state I4 (orange). b Close-up view of structural representation of the PC1 of Cas12a complex highlighting the motions of BH (bridge helix, pink), RuvC-H1 (RuvC-helix 1, purple), and RuvC-lid (medium purple) between the different states. c–e Correlation networks of internal dynamics derived from 1 μs of MD simulations for FnoCas12aKD2P-S2 in (c), FnoCas12aKD2P-S4b in (d), and FnoCas12aWT-I4 (PDB: 6GTG) in (e). Circular plots represent correlations between four key regions: RuvC-H1 (purple), BH (pink), crRNA (orange), and DNA (blue). Correlation values range from −1 (teal) to +1 (red), with 0 (white) representing no correlation. The color intensity reflects the strength of the correlation, where saturated red or teal corresponds to the most positive or negative correlation, respectively. Visual cutoff was applied for values above −0.4 and below 0.4 to enhance clarity. Selected Cas12a residues in the BH (E961, R964, and R968), RuvC-H1 (K981), and RuvC-lid (E1020, K1021, and N1022) are labeled in the correlation network. As the states progress from S2 to S4b to I4, there is an increase in the positively correlated motion between the BH, RuvC-lid, RuvC-H1, crRNA, and DNA. Due to the absence of sidechains on the experimental structure of FnoCas12aWT-I2 (PDB: 6GTD), the cross-correlation network analysis was not performed for this state (please see Supplementary Method 3 for details).
Loop-to-helical transition of the BH enables a coordinated communication between different regions of Cas12a and nucleic acids
Our structures reveal differences in the interaction of Cas12a with crRNA and DNA at different states (Fig. 4c–e and Supplementary Fig. 21). To delineate potential allosteric communication between BH and nucleic acid, we performed correlation analyses of residues E961, R964, R968, K981, E1020, K1021, and N1022 of the BH, RuvC-H1, and RuvC-lid, as well as nucleotides in crRNA and DNA (Fig. 4c–e). The complete correlation analysis is available in Supplementary Fig. 21 and Supplementary Tables 12–20. Results reveal that BH–RuvC-H1 and RuvC-lid conformations exhibit strongly correlated motions with those of the crRNA and DNA, and that the motions are different between the different states. FnoCas12aKD2P-S2 has fewer correlated motions (i.e., components move in the same direction) between the BH–RuvC-H1 and RuvC-lid, while there are several anti-correlated motions (i.e., components move in the opposite direction) between BH–RuvC-H1 and the RNA and DNA (Fig. 4c and Supplementary Tables 12, 15, and 18). In the intermediate state FnoCas12aKD2P-S4b, however, this correlation pattern shifts; there is an increase in the correlated motions between BH–RuvC-H1 and RuvC-lid and a few additional correlated motions with the crRNA, while there still is a substantial number of anti-correlated motions with crRNA and DNA (Fig. 4d and Supplementary Tables 13, 16, and 19). In FnoCas12aWT-I420, we see only correlated motions between BH–RuvC-H1, RuvC-lid, crRNA, and DNA (Fig. 4e and Supplementary Tables 14, 17, and 20). Interestingly, some of the correlated motions with crRNA and DNA include nucleotides near the middle of the R-loop ( ~ 8) providing mechanistic clues on why mismatches between the crRNA and DNA in the R-loop away from the PAM are particularly sensitive in FnoCas12a. This reorganization of the correlation network in different states of the protein points to the pivotal role that the BH–RuvC-H1 transitions play in organizing the cleavage-competent form of Cas12a.
Helix 1 from the RuvC motif-II of the Cas12 protein family is adopted for coordinated conformational changes with the BH
The coordinated conformational change of BH with the first helix of the RuvC motif-II (RuvC-H1) in Cas12a’s activity was previously proposed17, and its structural mechanisms are revealed in FnoCas12aKD2P structures. To further understand the importance of the coordination of BH and RuvC-H1 and its conservation in other Cas family proteins, an HMM (Hidden Markov Model) profile of these two helices was used to search the HMMER40 database. The result showed hits only from the Cas12a subtype, indicating that the sequence conservation around this region is limited to only Cas12a orthologs (Supplementary Fig. 22).
We next analyzed whether there is structural conservation of these two helices in Cas12 subtypes. We used BH and RuvC domain (motifs I, II, and III) of FnoCas12a (PDB: 5NFV23) as the query in DALI (Distance Matrix Alignment) server41. The structural analysis identified similar structural folds in other Cas12 subtypes, resolvase, and TnpB. This is consistent with previous studies that indicated TnpB as the ancestor for Cas12 proteins42 with conserved structural similarities43. Since Cas9 also has a BH and a RuvC domain (composed of motifs I, II, and III), we included selected Cas9 subtypes along with the results from the DALI output and performed an “all-against-all structure comparison” using the DALI server. A dendrogram based on this analysis groups all the proteins into five clusters, with the Cas9 subtypes occupying a distinct cluster and the Cas12 subtypes grouping into different sub-clusters (Fig. 5a). These observations show that there are some inherent structural differences between Cas12a and Cas9, which may translate to differences in the conformational activation processes. Cas12 structural analysis shows that the relative orientation of BH and RuvC-H1 is similar and that RuvC-H1 overlays very well in all the Cas12 subtypes. Comparing the BH between the Cas12 subtypes, the length and orientation of BH are similar in Cas12e44, Cas12f45, Cas12j46,47, Cas12k48, Cas12g49, Cas12c150, Cas12i151, and Cas12i252 (Fig. 5b and Supplementary Fig. 23). Variations are observed in the BH of Cas12j46,47, Cas12l53, Cas12f 45, and Cas12i151 since their BHs possess an insertion domain or have folded double helices compared to only a loop in FnoCas12a (Fig. 5 and Supplementary Fig. 23).
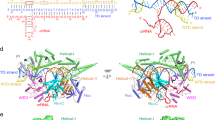
a A DALI41 dendrogram representing the hierarchical clustering of structures of BH (bridge helix) and RuvC domain ( ~ 200 amino acids covering motifs I, II, and III) of Cas12 and Cas9 subtypes, and ancestor proteins. The dendrogram shows five distinct clusters of proteins based on the structural similarities of BH and RuvC domain. PDB IDs with the corresponding proteins are indicated at the tip of each respective leaf. The numerical value above each branch indicates the branch length and longer branches show lower degree of structural similarity. b Structural superposition of FnoCas12a [BH in pink, RuvC-H1 (RuvC-helix 1) in purple, and the remaining part of the RuvC domain in medium purple] with few representative proteins belonging to a cluster (see Supplementary Fig. 23a for related figures). The Z-score, RMSD, and number of equivalent residues aligned (Lalin) based on the preliminary DALI search are shown against each superposition for Cas12 subtypes. Since Cas9 subtypes were manually added, rather than being identified through a DALI search, these parameters are not available for them. Each structure aligned with FnoCas12a is shown in one color, without domain demarcation, and the color matches the highlighting of the associated PDB ID. For Cluster V, BH and the helix from RuvC motif-II of Cas9 that aligns with RuvC-H1 of FnoCas12a are shown for selected structures (see Supplementary Figs. 23 and 24 for the remaining analysis of Cas9’s BH and RuvC domain conformational changes). The PDB IDs and the related structures used in this analysis are listed in Supplementary Table 23. Sequences covering the BH and RuvC domain that were used for generating the dendrogram and the structural comparisons are provided in the Source Data file. The dendrogram was generated using an online tool iTOL73 and structure alignments were made using ChimeraX58.
Structural overlays of the BH and RuvC domain (motifs I, II, and III) of Cas9 (subtypes II-A, II-B, and II-C) with those of FnoCas12a show that the BH and a helix of the RuvC motif-II of Cas9 have a similar structural arrangement to that in Cas12a. While helix in the RuvC motif-II of Cas9 and Cas12a RuvC-H1 shows a better structural overlay, the BH of Cas9 is prominently longer and does not align with that of FnoCas12a (Fig. 5b, Supplementary Figs. 23, 24). Another significant difference between Cas9 and Cas12 is the insertion of the REC lobe between BH and RuvC motif-II in Cas9, compared to their direct connection in Cas12 families (Supplementary Fig. 24). This long separation can potentially disrupt coordinated conformational changes between BH and RuvC-H1 that are present in Cas12s. Within the Cas9, there is a loop-to-helical transition of the BH in response to RNA binding that has been demonstrated to impact DNA cleavage efficiency and DNA mismatch sensitivity6,11,13,32,33,35. These observations show that while the fundamental mechanism of loop-to-helical transition of the BH is conserved in Cas12a and Cas9, the mechanisms for coordinated conformational changes may differ between them. While the RuvC-H1 of RuvC motif-II is co-opted by Cas12 families to undergo coupled transformations with the BH to orchestrate conformational changes needed for assembling the pre-catalytic state, such coupling is absent in Cas9 (Supplementary Fig. 24c). Thus, nature seems to have optimized Cas12a and Cas9 molecular mechanisms differently to attain the same goal of RNA-directed, sequence-specific, DNA cleavage.
Discussion
Previous work from our lab has reported that proline substitutions (K969P/D970P, FnoCas12aKD2P) in a region of the BH that undergoes loop-to-helical transition in Cas12a’s ternary complex impart a higher DNA mismatch sensitivity14. Additionally, FnoCas12aKD2P has a lower efficiency to cleave matched (on-target) plasmid DNA compared to that of FnoCas12aWT14. To get molecular insights into BH-mediated DNA cleavage, we compared the previously reported FnoCas12aWT structures and those from our present work on FnoCas12aKD2P that have captured several intermediate states through which the FnoCas12a-crRNA-DNA complex proceeds towards attaining the pre-catalytic state. The additional results from FnoCas12aKD2P show that an intact BH that can undergo a stable loop-to-helical transition is important for R-loop propagation and in attaining the pre-catalytic stage. It also points to a slowing down of the conformational cascade, with more intermediate states that are captured between FnoCas12aWT-I2 and FnoCas12aWT-I320. We see one state with a complete absence of the REC2 domain, which was also observed in AsCas12a in earlier stages of R-loop formation and was shown to be important in connecting the active state conformation to full R-loop formation21. In addition, the elongation of the BH due to the formation of new helical turns and its bending towards the RNA-DNA hybrid appears to be critical in providing an anchorage for R-loop and to complete the RNA-DNA hybrid formation. FnoCas12aWT-I2 and FnoCas12aWT-I320 have BH in a loop state prior to its the helical conversion. In these structures, BH is at a distance of 17 Å from the RNA-DNA hybrid with a total of 8-nucleotides of base pairing between the crRNA-guide and TS DNA. In comparison, in FnoCas12aWT-I4, we see a fully formed and bent BH that is closer to the RNA-DNA hybrid (6 Å). In FnoCas12aKD2P, which is not capable of undergoing the loop-to-helical transition, we only see up to a maximum of 11-nucleotides of base pairing between the crRNA-guide and TS DNA, with a distance of ~20 Å from the BH. Interestingly, we also see stacking and distortion of bases towards the end of the R-loop, implicating that the RNA and DNA strands are not aligned well with an impaired BH, hinting at the essentiality of the stability provided by the BH in these intermediary stages to drive R-loop propagation. The slowing down of the structural transitions is supported by the lower reaction kinetics of FnoCas12aKD2P when compared to FnoCas12aWT14.
Opening of the lid of the RuvC active site pocket enables passage of DNA into the active site. In all the structures of FnoCas12aKD2P ternary complex, we observed that the RuvC-lid is in a loop form occluding the active site. We propose this to be an allosteric control to prevent non-specific DNA cleavage. The BH-allostery can potentially time the RuvC-lid opening only when full complementarity is present in the RNA-DNA hybrid. In another role, the loop-to-helical transition of RuvC-lid was shown to play an important role in bringing the TS into the active site pocket for DNA cleavage21,27. The AsCas12a structures probing the TS entry mechanism (PDB: 8SFP, 8SFQ and 8SFR21) also support these since the BH is extended towards the RNA-DNA hybrid, establishing interactions with the RuvC-lid helix, which leads to the exposure of the active site pocket in these structures. While we do not have structural features to support the TS entry mechanism due to the presence of partial DNA strands, BH’s helical transition appears to be critical in the passage and cleavage of the TS. This is supported by our previous work and biochemical assays reported in the present work. We observed a 6-fold reduction in TS compared to only a 3-fold reduction in NTS cleavage by FnoCas12aKD2P compared to that by FnoCas12aWT14. Additionally, FnoCas12aKD2P discriminates the RNA-DNA mismatches by preventing TS cleavage since it was able to nick substrates with mismatches but not linearize them14. The efficiency to cleave mismatches is lower for all 20-nucleotides along the RNA-DNA hybrid, with mismatches at positions 12–17 of the target DNA producing higher accumulation of the nicked intermediate (61-77%) compared to that in FnoCas12aWT (3–19%)14. In this study, we observe that variants with disrupted BH–RuvC-lid interactions (R964A, E1020A/K1021A) in the FnoCas12aKD2P background exhibit pronounced reductions in cleavage of both TS and NTS, while the significant effect observed in FnoCas12aWT is the reduction in linearization and accumulation of nicked intermediate for MM13 substrate. Together, these results indicate that the BH’s loop-to-helical transition is the main driving force in R-loop propagation and the opening of the RuvC-lid in response to the total length of the RNA-DNA hybrid. Further studies are needed to find the mismatch-sensing mechanism of FnoCas12aKD2P, especially to understand why positions 12-18 of the RNA-DNA paired region is more sensitive compared to the rest of the R-loop positions.
In our previous work, we have also demonstrated that FnoCas12aKD2P has a lower trans-cleavage efficiency (0% substrate left for FnoCas12aWT at 5 min, compared to 40% remaining in FnoCas12aKD2P after two hours)14. Unlike FnoCas12aWT, which rapidly degraded substrates, FnoCas12aKD2P accumulated linear intermediates and failed to sustain a robust progressive trans-cleavage14. Additionally, FnoCas12aKD2P was unable to perform trans-cleavage when activated by a dsDNA14. The trans-cleavage occurs through keeping the RuvC domain “open” for non-specific ssDNA to access the active site24,54. These observations highlight that impairment of the BH’s helical transition likely blocks the prolonged activated form of the RuvC pocket, preventing trans-cleavage in FnoCas12aKD2P.
Previous MD simulation studies have provided complementary insights into cryo-EM and crystallographic structures of Cas12a by capturing conformational states, such as early R-loop dynamics55, DNA-dependent exposure of the RuvC catalytic core21, and α-helical lid motions that guide DNA movements27. We used MD simulations in our study to provide a mechanistic model that explains our structural and biochemical results. Correlation analyses reveal a reorganization of the allosteric coupling between BH, RuvC-H1, RuvC-lid, and nucleic acids in the different conformational states. There is a change from antagonistic BH–crRNA–DNA motions in FnoCas12aKD2P-S2, to reduced antagonism in FnoCas12aKD2P-S4b, and ultimately to strongly coordinated correlations of BH with crRNA, DNA, RuvC-H1, and RuvC-lid in FnoCas12aWT-I4. Notably, this coordinated coupling is absent in the impaired BH variant, underscoring the requirement of BH’s loop-to-helical transition for establishing the allosteric network that licenses catalytic activation.
Cas12a goes from a closed to an open to slightly closed conformation during the different steps following DNA binding and R-loop propagation (Fig. 6). Our results and previous literature on FnoCas12aWT show that the unwinding of the RuvC-H1 is a prerequisite to the formation of additional turns in the BH. We can envision that the helix melting of RuvC-H1 and helix formation of BH is a method to control the degree to which the crab-claw-shaped protein will open the REC2 and Nuc domains to control DNA’s access into the middle of the bilobed structure. The BH and RuvC-H1 appear to be the two ends of the anchor needed for coordinating the conformational transitions by changing the number of turns in both helices (Supplementary Movie 2). The proposed mechanism is supported by previous work that showed complete absence of Cas12a’s DNA cleavage when RuvC-H1 is deleted, and partial activity when BH is deleted17. Comparison of the BH conformational changes with AsCas12a21 and LbCas12a25,26,31 also shows similar features. While the BH is elongated in AsCas12a even with only 5-nucleotides of base pairing of the crRNA guide and the TS DNA, the bending of the BH to reach the RNA-DNA hybrid only happens after 15-nucleotides of base pairing (Supplementary Fig. 25), supporting our proposal that BH’s conformational change is a trigger to support full R-loop formation, which then translates to opening of the RuvC-lid21. Unwinding of RuvC-H1 is noticed only in the later stages of R-loop formation in AsCas12a and LbCas12a, slightly different from RuvC-H1 melting before BH’s transition in FnoCas12a (Supplementary Fig. 25). Nevertheless, the coordinated structural transitions of BH and RuvC-H1 seem to be a common mechanism for Cas12a’s DNA cleavage in the orthologs. Our computational simulations offer further support for the allosteric mechanism capturing conformational changes of BH–RuvC-H1 and RuvC-lid at different states of Cas12a activation and showing positive cooperativity between these protein regions and RNA/DNA only when the BH is intact.
Cas12a undergoes largescale conformational changes that makes the bilobed protein to take a closed conformation after binding to crRNA, followed by an open conformation to enable R-loop growth, and finally attaining a slightly-less open state after full R-loop formation to reach the pre-catalytic state. Our work illuminates the molecular mechanisms by which BH (bridge helix, pink) orchestrates these conformational changes. a In the binary complex and during the initial phase of DNA binding, part of the BH is shorter with a disordered loop. b After DNA binding progresses ∼9 bp of RNA-DNA hybrid formation, (Supplementary Fig. 25), the long RuvC-H1 (RuvC-helix 1, purple) of RuvC motif-II unwinds a few turns and the BH remains shorter with the loop region. c When the R-loop attains ∼16 bp of hybridization, the BH elongates by gaining a few helical turns with its concomitant bending towards the crRNA-DNA hybrid. These changes of the BH are correlated to the loop-to-helical transitioning of the RuvC-lid (medium purple) through direct interactions between amino acids in the region of the BH that has undergone the helical transition, enabling opening of the RuvC active site pocket. RuvC-H1, especially K981, acts as a pivot by changing the residue that it interacts within the RuvC-lid before and after the helical transition of the RuvC-lid. In FnoCas12aKD2P, where BH transition is hindered, the protein attains the open conformation (second state in the top panels) and the interactions in the pre-catalytic state are absent, with the RuvC-lid exiting in a loop form. The R-loop is only partially formed and distorted in FnoCas12aKD2P, which demonstrates the allosteric cooperation of BH and RuvC-H1 to open the RuvC active site pocket based on the number of base pairs in the RNA-DNA hybrid. In this figure, interactions for panels a and b are for FnoCas12aKD2P (with K969P and D970P substitutions), while that in (c) is for FnoCas12aWT20 (with K969 and D970). Figure was prepared using Microsoft PowerPoint.
Our structural comparison of the BH and RuvC-H1 of the diverse Cas12 subtypes also hints towards the conservation of BH and RuvC-H1, even in Cas12 subtypes where there is a domain insertion in between these helices (e.g., Cas12j46,47). Interestingly, Cas9’s BH is longer with more conserved Arg residues and does not show a coordinated helical transition with a RuvC helix similar to that in Cas12a. While the BH’s helical transition and its allosteric effect of DNA cleavage seem to be conserved mechanisms in Cas9 and Cas12a, the structural features contributing to this allostery seem to be evolutionarily divergent. Further studies targeting these helices of Cas9 and Cas12a will help elucidate the differences and similarities of the BH-mediated conformational activation and DNA mismatch-sensing mechanisms of these two proteins that evolved differently to perform the same RNA-guided DNA cleavage activity.
In conclusion, cryo-EM studies of our variant, FnoCas12aKD2P, unravel five intermediary steps that broaden our understanding of the critical role played by BH towards attaining the pre-catalytic state. The structures show that impairment of BH through proline substitutions arrests the conformational cascade, illuminating the allosteric role that BH exerts on the RuvC catalytic pocket by keeping the length of the RNA-DNA hybrid as a checkpoint. The structures and the simulation results also show direct evidence on the cooperativity of BH and RuvC-H1 in enabling R-loop propagation and opening of the RuvC-lid. These findings, together with the earlier14,16,18 and current biochemical evidence and structural studies of AsCas12a21 and LbCas12a31, support a model in which BH dynamics couple R-loop propagation to catalytic activation, thereby enforcing mismatch discrimination. The importance of BH in Cas12a is accentuated by the presence of several naturally existing anti-CRISPR proteins that block several aspects of BH, including conserved Arg(s) and loop-to-helical transitioning of the BH31. The BH being a conserved single helical domain in Cas9 and Cas12 families, as well as in other RNA-binding proteins such as the RNA polymerase, our study reveals the molecular mechanisms by which BH’s conformational changes regulate communication in multi-domain proteins to assist enzyme function.
Methods
Plasmid preparation and mutagenesis
The Francisella novicida (Fno) cas12a wild-type and K969P-D970P variant (cas12aKD2P) genes that were cloned in a pET28a-based vector with an N-terminal Maltose Binding Protein (MBP) tag were used as backbones for generating all the protein variants9,14. Variants were constructed by site-directed mutagenesis using the primers listed in Supplementary Table 21. Resultant PCR products were transformed into Escherichia coli DH5\(\alpha\) cells, and selected clones for each variant were sequence confirmed by nanopore sequencing (whole plasmid sequence for all the clones are available in Source Data). One sequence-confirmed clone for each variant was subsequently transformed into E. coli Rosetta (DE3) cells for protein overexpression.
Protein expression and purification
The plasmids containing the Fno cas12a gene with the different substitutions were transformed into E. coli Rosetta (DE3) cells and purified using established procedures9,14,56. Briefly, Rosetta cells with the expression plasmid were grown in 2x Yeast Extract Tryptone (2xYT) medium supplemented with 50 μg mL−1 Kanamycin and 34 μg mL−1 Chloramphenicol. The cultures were induced with 0.2 mM IPTG at an optical density (OD) 600 of 0.6–0.9, followed by an overnight incubation at 18 °C. Cells were harvested by centrifugation (5000 × g for 20 min at 4 °C) and resuspended in buffer containing 20 mM HEPES pH 7.5, 1 M NaCl, and 20 mM Imidazole. For cryo-EM studies, FnoCas12aKD2P was purified using a two-step method comprising Ni2+-NTA (nitrilotriacetic acid) and cation exchange chromatography. Cell lysis was achieved by sonication, and insoluble material was removed by centrifugation (30,000\(\times g\) for 45 min at 4 °C). The clarified supernatant was applied to a 5 mL Ni-NTA Superflow cartridge (Qiagen) equilibrated with wash buffer. After washing the resin with 20 column volumes of wash buffer, the bound protein was eluted from the Ni-NTA column using a continuous imidazole gradient, increasing to 100% buffer B (1X composition: 20 mM HEPES pH 7.5, 1 M NaCl, and 500 mM imidazole) over 15 column volumes. Eluted fractions were pooled and mixed with TEV protease for MBP-tag cleavage while dialyzing against cation exchange buffer A (1X composition: 20 mM HEPES pH7.5, 150 mM KCl, 2 mM EDTA, and 1 mM DTT), overnight with slow stirring at 4 °C. Dialyzed protein was clarified by centrifugation and applied to a HiTrap SP HP cation-exchange column equilibrated in cation exchange buffer A. After washing with 10 column volumes of cation exchange buffer A, protein was eluted using a 0–100% cation exchange buffer B (1X composition: 20 mM HEPES pH7.5, 1 M KCl, 2 mM EDTA, and 1 mM DTT) gradient over 20 column volumes, with FnoCas12a eluting at ~350 mM KCl. Fractions were assessed by 7.5% SDS-PAGE to confirm TEV cleavage (expected mass ~152 kDa). The purified protein was concentrated using the Thermo Scientific PierceTM protein concentrator PES, 30 kDa MWCO (2–6 mL) and the Cytiva Vivaspin 500 \({{\rm{\mu }}}\)L, 10 kDa MWCO centrifugal protein concentrators and stored on ice until complex preparation for cryo-EM. For variants used in biochemical assays, fractions eluted from cation exchange chromatography were pooled and further purified using size-exclusion chromatography (HiPrep 16/60 Sephacryl S-300 HR column; GE Healthcare) in 20 mM HEPES pH7.5, 150 mM KCl, 2 mM EDTA and 1 mM TCEP. Peak fractions were concentrated using Amicon® Ultra-centrifugal filters (30 kDa MWCO, 15 mL), followed by final concentration to ~13 mg/mL with Cytiva Vivaspin filters (10 kDa MWCO, 500 µL). Proteins were aliquoted, flash-frozen in liquid nitrogen, and stored at −80 °C. Aliquots were thawed on ice immediately prior to biochemical assays.
RNA and DNA preparation (for oligo sequences see Supplementary Table 22)
RNA was prepared by in vitro transcription and purified using urea polyacrylamide gel electrophoresis14,32,56. The template and T7 promoter DNA oligos (Supplementary Table 22) were ordered from Integrated DNA Technologies (IDT) and annealed in annealing buffer (10 mM HEPES, pH 8, 50 mM KCl) in a 1:1.5 molar ratio (2 \({{\rm{\mu }}}\)M: 3 \({{\rm{\mu }}}\)M) of template: T7 promoter strands. Approximately 400 ng of annealed template was used for in vitro transcription using 40 μg T7 RNA polymerase and NTPs (GTP: 9 mM, ATP: 8 mM, CTP: 8 mM, and UTP: 8 mM) in transcription buffer (1X composition: 40 mM Tris-HCl pH 8, 1 mM spermidine, 50 μg bovine serum albumin, 20 mM MgCl2, and 5 mM DTT)56. The transcription reaction was incubated at 37 °C for 4 h and then treated with 0.01 mg/mL DNase I to remove the DNA template. The crRNA product was precipitated through ethanol precipitation and further purified using a 12% polyacrylamide-8M urea gel. The purified crRNA was suspended in annealing buffer and heated to 95 °C for 2 min, then slowly cooled to room temperature for correct folding and quantification. The folded crRNA was stored at 4 °C until use, up to a maximum of one week.
For cryo-EM, a 24-mer target strand (TS) and non-target strand (NTS) (Supplementary Fig. 8a) were ordered as ssDNA oligos from Sigma-Aldrich. The TS and NTS were annealed by mixing them in a 1:1 ratio (45 \({{\rm{\mu }}}\)M each) in annealing buffer, followed by heating at 95 °C for 5 min and allowing it to slowly cool to room temperature.
Cryo-EM grid preparation and data collection
Freshly purified 10 μM FnoCas12aKD2P protein and 12 μM crRNA were incubated at 37 °C for 30 min to form the ribonucleoprotein (RNP) complex in the complexing buffer (20 mM HEPES, pH 7.5, 150 mM KCl, 1% glycerol, 1 mM TCEP, 2 mM EDTA). For formation of the ternary complex, 15 μM DNA duplex was added to the assembled RNP and incubated at 37 °C for another 30 min. The resulting ternary complex was loaded onto a Superdex 200 increase 10/300 GL column (GE Healthcare), which was pre-equilibrated with 1X complexing buffer at 4 °C, to separate complexes from excess nucleic acid. Peak fractions were analyzed on a 6% native polyacrylamide gel (40% (w/v) 29:1 acrylamide: bis-acrylamide) (Supplementary Fig. 1), and concentration was determined using absorbance at 280 nm as measured on a Nanodrop 8000 Spectrophotometer (Thermo Scientific), considering 1 Abs = 1 mg/mL. Peak fraction ( ~ 0.147 mg/mL, peak 2) containing the band for the ternary complex was vitrified on the same day of purification using a Leica EM GP2 plunger at 4 °C and 98% humidity. A 4 μL of sample was applied to graphene oxide (Sigma-Aldrich, 2 mg/mL)- coated Quantifoil R1.2/1.3 300 mesh Cu grid, which was glow discharged using PELCO easiGlowTM at 15 mA for 30 s (Preparation of graphene oxide grids is mentioned in Supplementary Method 1). After the sample application, the grid was immediately blotted for 2 s, followed by plunge freezing.
7232 micrographs were collected at the Stanford-SLAC Cryo-EM Center (S2C2), an NIH National Cryo-EM facility on TEMGamma, a Titan Krios G3i (Thermo Fisher Scientific) cryogenic electron microscope operating at 300 keV and equipped with an X-FEG electron source, fringe-free optics, a BioQuantum (Gatan) energy filter and a K3 (Gatan) direct electron detector. Movies were recorded using a 100 μm objective aperture, an energy filter slit of 20 eV, and an electron dose of 1.25 e−/Å2 per frame at a magnification of 105,000\(\times\) with a corresponding pixel size of 0.86 Å for a total exposure of 50 e−/Å2 (40 frames over 2.13 s, 0.053 s per frame) (Supplementary Table 1).
Single particle cryo-EM data processing and model building
The dataset was processed using the standard workflow on cryoSPARCv4.4.157. After patch motion correction and contrast transfer function (CTF) estimation, 5538 micrographs were selected for further processing. Particle picking, conducted with blob picker, identified 6,576,476 particles across all selected micrographs. Following particle inspection, 5,142,814 particles were extracted with a box size of 280 pixels (details in Supplementary Method 2).
We began 2D classification with 400 classes (details in Supplementary Method 2). After three rounds of iterative 2D classification, each with ~200 classes, we selected 399,609 particles for ab-initio reconstruction and homogeneous refinement. To assess conformational heterogeneity, 3D variability analysis was performed with a filter resolution of 4.5 Å. The variability observed in this step guided a subsequent 3D classification, resulting in six different classes (Supplementary Fig. 4, details in Supplementary Method 2), of which five, with up to 4 Å resolution, were used for structural analysis.
An initial model was built using PDB: 6GTG20 for each class. The models were first aligned with their respective maps on ChimeraX58, followed by manually rebuilding in Coot59 and refinement using Phenix.real_space_refine60,61. The REC1 and REC2 domains were individually isolated and fitted manually into maps, followed by rigid body fitting and real space refinement. The models were iteratively improved based on the validation metrics provided by MolProbity62 and Phenix refinement. The model building procedure for the FnoCas12aKD2P-S4a map, which exhibited stretched density for the REC2 domain and a different placement of REC2 compared to the corresponding position of this domain in PDB: 6GTE20, is described in section Supplementary Method 2.
In vitro DNA cleavage assays
Plasmid cleavage reactions (10 μL) were carried out in 1X plasmid reaction buffer (20 mM HEPES, pH 7.5, 150 mM KCl, 2 mM TCEP, 10% glycerol, 5 mM MgCl₂). FnoCas12a and annealed crRNA were mixed to final concentrations of 25 nM and 50 nM, respectively, and pre-incubated at 37 °C for 10 min to form the ribonucleoprotein (RNP) complex. Reactions were initiated by the addition of 100 ng (∼5 nM) substrate plasmid and incubation at 37 °C for 1–15 min, as indicated. Reactions were quenched by adding 10 μL of a 2X stop dye (1X composition: 50 mM EDTA, 1% SDS, 10% glycerol, 0.0125% orange G). Products were resolved on 1.5% agarose gels, post-stained with ethidium bromide (0.25 μg mL−1) and imaged using a Bio-Rad ChemiDoc system (Supplementary Fig. 17). Band intensities corresponding to nicked (\({{{\rm{I}}}}_{{{\rm{N}}}}\)), linear (\({{{\rm{I}}}}_{{{\rm{L}}}}\)), and supercoiled (\({{{\rm{I}}}}_{{{\rm{SC}}}}\)) species were quantified in ImageJ software63. The no-enzyme lane (consisting of DNA, buffer, and metal) of the same gel was used as a control, designated as ‘C’. Nicked, linear, and total cleavage percentages for each protein variant were calculated as:
The fraction of total cleavage was defined as:
Standard deviation (SD) and standard error of the mean (SEM) were calculated as described previously13.
Where \(R\) is the value form each replication, \({R}_{{AV}}\) is average of data values of all of the replications, and \({{n}}\) is the number replications; \({{n}}=3\).
Where \({{n}}\) is the number replications; \({{n}}=3\).
Three independent replications using proteins from two independent purifications were used in this work. Raw data (intensities of different DNA bands) and calculations are provided in the Source Data.
Computational modeling
We used structures of FnoCas12aWT states I2 (PDB: 6GTC20) and I4 (PDB: 6GTG20) and FnoCas12aKD2P states S2 and S4b for computational studies. The missing protein segments were added using Modeller10.664, protonation states were assigned by PROPKA65,66, and systems were solvated, neutralized, and prepared, maintaining consistency with experimental conditions. Each system underwent 1 μs NVT simulation. The resulting trajectories, totaling 4 μs after including all the systems, were concatenated and analyzed using the Principal Component Analysis (PCA). For the cross-correlation analysis, the first 100 ns of the initial NVT production trajectory was excluded and only the last 900 ns of each trajectory were used. Both PCA and cross-correlation analyses were performed using CPPTRAJ67. Simulations were performed using the AMBER2068 employing AMBER/ff14SB, OL15, ff99bsc0 + χOL3, and TIP3P parameters for protein, DNA, RNA, and water, respectively69,70,71. Protonation states were assigned using PROPKA, and systems were solvated in TIP3P water using a nearly cubic simulation box with at least 12 Å of solvent beyond the protein/crRNA/DNA complex in all directions, neutralized with Na+ counterions, and prepared without divalent metals to match experimental conditions. Each system was heated to 300 K and equilibrated through sequential NVT and NPT stages (1 ns NVT heating followed by NPT equilibration at 1 bar, continued until the system density reached ~1.01 g/cm3) before initiating the 1 μs NVT production run. Additional computational details are provided in Supplementary Method 3.
Bioinformatics
HMM (Hidden Markov Model) profile for the BH–RuvC-H1 region of FnoCas12a (residues 953-1000; Uniprot ID A0Q7Q2) was built using HMMER40 based on the sequence alignment using Clustal Omega72 for 313 Cas12a annotated sequences retrieved from NCBI (National Centre for Bioinformatics Information). The profile was used to search for similar sequences using hmmsearch (a tool in HMMER) with a default e-value of 0.01. Structural similarity search for the BH–RuvC domain (motifs I, II, and III; PDB: 5NFV23) was performed using DALI server41. The top hits belong to Cas12a orthologs with Z-scores ranging from 24.9 to 13.9, while hits belonging to other Cas12 subtypes, bacterial and viral Resolvases, Recombinase, and TnpB have Z-scores ranging from 13.5 to 8.0 indicating there is probable structural homology40. To assess the structural similarities of Cas9, the BH and RuvC domain of eight Cas9 structures belonging to subtypes II-A, II-B, and II-C were added to the Cas12 structures that were identified by DALI search. Structures that belong to similar bacterial species were removed to avoid redundancy in the structure comparison. This was followed by an “all-against-all structure comparison” for the selected Cas12 and Cas9 structures covering the BH and RuvC domain (sequences covering this region are available in Source Data). The output from DALI was visualized as a dendrogram in a rectangular mode (unrooted) using an online tool iTOL73 (Interactive Tree of Life). The PDB codes and the corresponding structures used for structural comparison are indicated in Fig. 5, Supplementary Fig. 23, and Supplementary Table 23.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the manuscript, Supplementary Information, and public data repositories as mentioned below. The atomic coordinates and cryo-EM density maps are available in the Protein Data Bank and Electron Microscopy Data Bank under accession codes 9MKT/ EMD-48337 for FnoCas12aKD2P-state 1, 9MKU/ EMD-48338 for FnoCas12aKD2P-state 2, 9MKV/ EMD-48339 for FnoCas12aKD2P-state 3), 9MKW/ EMD-48340 for FnoCas12aKD2P-state 4a, and 9MKX/ EMD-48341 for FnoCas12aKD2P-state 4b. The molecular dynamics data generated in this study have been deposited in the GitHub database74. DNA sequencing files for the engineered Cas12a variants, the full raw dataset used to generate Fig. 3e, f, Supplementary Fig. 18a, b, Supplementary Table 11, and the sequences used to generate Fig. 5 and Supplementary Fig. 23 are provided in the Source Data file. Uncropped gel images for all the replications of the DNA cleavage assay are included in Supplementary Information (Appendix, pages 96 to 104). Source data are provided with this paper.
References
Mojica, F. J., Diez-Villasenor, C., Garcia-Martinez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Ganguly, C., Rostami, S., Long, K., Aribam, S. D. & Rajan, R. Unity among the diverse RNA-guided CRISPR-Cas interference mechanisms. J. Biol. Chem. 300, 107295 (2024).
Stella, S., Alcon, P. & Montoya, G. Class 2 CRISPR-Cas RNA-guided endonucleases: Swiss Army knives of genome editing. Nat. Struct. Mol. Biol. 24, 882–892 (2017).
Adashi, E. Y., Gruppuso, P. A. & Cohen, I. G. CRISPR Therapy of Sickle Cell Disease: The Dawning of the Gene Editing Era. Am. J. Med 137, 390–392 (2024).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Murugan, K., Babu, K., Sundaresan, R., Rajan, R. & Sashital, D. G. The Revolution Continues: Newly Discovered Systems Expand the CRISPR-Cas Toolkit. Mol. Cell 68, 15–25 (2017).
Sundaresan, R., Parameshwaran, H. P., Yogesha, S. D., Keilbarth, M. W. & Rajan, R. RNA-Independent DNA Cleavage Activities of Cas9 and Cas12a. Cell Rep. 21, 3728–3739 (2017).
Makarova, K. S. et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol 18, 67–83 (2020).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Yamano, T. et al. Crystal Structure of Cpf1 in Complex with Guide RNA and Target DNA. Cell 165, 949–962 (2016).
Babu, K. et al. Bridge Helix of Cas9 Modulates Target DNA Cleavage and Mismatch Tolerance. Biochemistry 58, 1905–1917 (2019).
Parameshwaran, H. P. et al. The bridge helix of Cas12a imparts selectivity in cis-DNA cleavage and regulates trans-DNA cleavage. FEBS Lett. 595, 892–912 (2021).
Bratovic, M. et al. Bridge helix arginines play a critical role in Cas9 sensitivity to mismatches. Nat. Chem. Biol. 16, 587–595 (2020).
Worle, E., Jakob, L., Schmidbauer, A., Zinner, G. & Grohmann, D. Decoupling the bridge helix of Cas12a results in a reduced trimming activity, increased mismatch sensitivity and impaired conformational transitions. Nucleic Acids Res. 49, 5278–5293 (2021).
Worle, E., Newman, A., D’Silva, J., Burgio, G. & Grohmann, D. Allosteric activation of CRISPR-Cas12a requires the concerted movement of the bridge helix and helix 1 of the RuvC II domain. Nucleic Acids Res. 50, 10153–10168 (2022).
Ma, E. et al. Improved genome editing by an engineered CRISPR-Cas12a. Nucleic Acids Res. 50, 12689–12701 (2022).
Dagdas, Y. S., Chen, J. S., Sternberg, S. H., Doudna, J. A. & Yildiz, A. A conformational checkpoint between DNA binding and cleavage by CRISPR-Cas9. Sci. Adv. 3, eaao0027 (2017).
Stella, S. et al. Conformational Activation Promotes CRISPR-Cas12a Catalysis and Resetting of the Endonuclease Activity. Cell 175, 1856–1871.e1821 (2018).
Strohkendl, I. et al. Cas12a domain flexibility guides R-loop formation and forces RuvC resetting. Mol. Cell 84, 2717–2731.e2716 (2024).
Stella, S., Alcon, P. & Montoya, G. Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 546, 559–563 (2017).
Swarts, D. C., van der Oost, J. & Jinek, M. Structural Basis for Guide RNA Processing and Seed-Dependent DNA Targeting by CRISPR-Cas12a. Mol. Cell 66, 221–233.e224 (2017).
Swarts, D. C. & Jinek, M. Mechanistic Insights into the cis- and trans-Acting DNase Activities of Cas12a. Mol. Cell 73, 589–600.e584 (2019).
Jianwei, L. et al. Structures of apo Cas12a and its complex with crRNA and DNA reveal the dynamics of ternary complex formation and target DNA cleavage. PLoS Biol. 21, e3002023 (2023).
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 532, 522–526 (2016).
Saha, A. et al. An alpha-helical lid guides the target DNA toward catalysis in CRISPR-Cas12a. Nat. Commun. 15, 1473 (2024).
Baranova, S. V. et al. Thermodynamic parameters obtained for the formation of the Cas12a-RNA/DNA complex. Biochem Biophys. Res Commun. 743, 151176 (2025).
Jeon, Y. et al. Direct observation of DNA target searching and cleavage by CRISPR-Cas12a. Nat. Commun. 9, 2777 (2018).
Naqvi, M. M., Lee, L., Montaguth, O. E. T., Diffin, F. M. & Szczelkun, M. D. CRISPR-Cas12a-mediated DNA clamping triggers target-strand cleavage. Nat. Chem. Biol. 18, 1014–1022 (2022).
Zhang, H. et al. Structural Basis for the Inhibition of CRISPR-Cas12a by Anti-CRISPR Proteins. Cell Host Microbe 25, 815–826.e814 (2019).
Babu, K. et al. Coordinated Actions of Cas9 HNH and RuvC Nuclease Domains Are Regulated by the Bridge Helix and the Target DNA Sequence. Biochemistry 60, 3783–3800 (2021).
Wu, D. et al. Structural integrity and side-chain interaction at the loop region of the bridge helix modulate Cas9 substrate discrimination. Nucleic Acids Res. 53, https://doi.org/10.1093/nar/gkaf557 (2025).
Jiang, F. et al. Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. STRUCTURAL BIOLOGY. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Yamano, T. et al. Structural Basis for the Canonical and Non-canonical PAM Recognition by CRISPR-Cpf1. Mol. Cell 67, 633–645.e633 (2017).
Patel, A., Toso, D., Litvak, A. & Nogales, E. Efficient graphene oxide coating improves cryo-EM sample preparation and data collection from tilted grids. bioRxiv, 2021.2003.2008.434344 https://doi.org/10.1101/2021.03.08.434344 (2021).
Kumar, A., Sengupta, N. & Dutta, S. Simplified Approach for Preparing Graphene Oxide TEM Grids for Stained and Vitrified Biomolecules. Nanomaterials (Basel) 11, https://doi.org/10.3390/nano11030643 (2021).
Gao, P., Yang, H., Rajashankar, K. R., Huang, Z. & Patel, D. J. Type V CRISPR-Cas Cpf1 endonuclease employs a unique mechanism for crRNA-mediated target DNA recognition. Cell Res. 26, 901–913 (2016).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Holm, L. Using Dali for Protein Structure Comparison. Methods Mol. Biol. 2112, 29–42 (2020).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR-Cas systems. Nat. Rev. Microbiol 15, 169–182 (2017).
Sasnauskas, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023).
Tsuchida, C. A. et al. Chimeric CRISPR-CasX enzymes and guide RNAs for improved genome editing activity. Mol. Cell 82, 1199–1209.e1196 (2022).
Hino, T. et al. An AsCas12f-based compact genome-editing tool derived by deep mutational scanning and structural analysis. Cell 186, 4920–4935.e4923 (2023).
Pausch, P. et al. DNA interference states of the hypercompact CRISPR-CasPhi effector. Nat. Struct. Mol. Biol. 28, 652–661 (2021).
Carabias, A. et al. Structure of the mini-RNA-guided endonuclease CRISPR-Cas12j3. Nat. Commun. 12, 4476 (2021).
Xiao, R. et al. Structural basis of target DNA recognition by CRISPR-Cas12k for RNA-guided DNA transposition. Mol. Cell 81, 4457–4466.e4455 (2021).
Li, Z., Zhang, H., Xiao, R., Han, R. & Chang, L. Cryo-EM structure of the RNA-guided ribonuclease Cas12g. Nat. Chem. Biol. 17, 387–393 (2021).
Zhang, B. et al. Structural insights into target DNA recognition and cleavage by the CRISPR-Cas12c1 system. Nucleic Acids Res. 50, 11820–11833 (2022).
Zhang, B. et al. Mechanistic insights into the R-loop formation and cleavage in CRISPR-Cas12i1. Nat. Commun. 12, 3476 (2021).
Huang, X. et al. Structural basis for two metal-ion catalysis of DNA cleavage by Cas12i2. Nat. Commun. 11, 5241 (2020).
Sun, A. et al. The compact Caspi (Cas12l) ‘bracelet’ provides a unique structural platform for DNA manipulation. Cell Res. 33, 229–244 (2023).
Chen, J. S. et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Cofsky, J. C. et al. CRISPR-Cas12a exploits R-loop asymmetry to form double-strand breaks. Elife 9, https://doi.org/10.7554/eLife.55143 (2020).
Martin, L., Rostami, S. & Rajan, R. Optimized protocols for the characterization of Cas12a activities. Methods Enzymol. 679, 97–129 (2023).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D. Biol. Crystallogr 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr D. Biol. Crystallogr 68, 352–367 (2012).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Webb, B. & Sali, A. Comparative Protein Structure Modeling Using MODELLER. Curr. Protoc. Bioinforma. 54, 5 6 1–5 6 37 (2016).
Olsson, M. H., Sondergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: Consistent Treatment of Internal and Surface Residues in Empirical pKa Predictions. J. Chem. Theory Comput 7, 525–537 (2011).
Sondergaard, C. R., Olsson, M. H., Rostkowski, M. & Jensen, J. H. Improved Treatment of Ligands and Coupling Effects in Empirical Calculation and Rationalization of pKa Values. J. Chem. Theory Comput 7, 2284–2295 (2011).
Roe, D. R. & Cheatham, T. E. 3rd PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data. J. Chem. Theory Comput 9, 3084–3095 (2013).
Case, D. A. et al. AmberTools. J. Chem. Inf. Model 63, 6183–6191 (2023).
Zgarbova, M. et al. Refinement of the Cornell et al. Nucleic Acids Force Field Based on Reference Quantum Chemical Calculations of Glycosidic Torsion Profiles. J. Chem. Theory Comput 7, 2886–2902 (2011).
Zgarbova, M. et al. Refinement of the Sugar-Phosphate Backbone Torsion Beta for AMBER Force Fields Improves the Description of Z- and B-DNA. J. Chem. Theory Comput 11, 5723–5736 (2015).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128 (2007).
Alberto Monteiro dos Santos, Y. S., and Rakhi Rajan Bridge helix of Cas12a is an allosteric regulator of R-loop formation and RuvC activation. https://doi.org/10.5281/zenodo.17834622 (2025).
Acknowledgements
We thank the OU Protein Production and Characterization Core (PPCC) facility for protein purification services and instrument support and the Biomolecular Structure Core (BSC)-Norman for cryo-EM experimental support. The OU PPCC and BSC-Norman are supported by IDeA grants from NIGMS [grant number P20GM103640 and P30GM145423]. The data for FnoCas12a-crRNA-DNA complex was collected at Stanford-SLAC Cryo-EM Center (S2C2), which is supported by the National Institutes of Health Common Fund Transformative High-Resolution Cryo-Electron Microscopy program (grant number U24 GM129541) and National Institute of General Medical Sciences (1R24GM154186). Some of this work was performed at the National Center for CryoEM Access and Training (NCCAT) and the Simons Electron Microscopy Center located at the New York Structural Biology Center, supported by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy program (U24 GM129539 and NIGMS R24 GM154192) and by grants from the Simons Foundation (SF349247) and NY State Assembly. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The authors would also like to thank S2C2 members (Dr. Wah Chiu, Dr. Nathan D. Burrows, and Dr. Xu Yang) and NCCAT members (Dr. Edward Eng, Dr. Christina Zimanyi, and Dr. Aaron Owji) for their invaluable support and guidance for establishing cryo-EM studies in the Rajan laboratory. We also thank Dr. Martin Lawrence and Luke Findlay for valuable advice on cryo-EM troubleshooting. We acknowledge Dr. Fares Najar at the High-Performance Computing Center of the Oklahoma State University for building the HMM profile for the BH and RuvC-H1 region. Work reported here was supported in part by grants from the National Science Foundation [MCB-1716423 and MCB-2424888], grants from the Research Council of the University of Oklahoma Norman Campus to R.R. and through the Dodge Family College of Arts and Sciences (DFCAS) Dissertation Research Fellowship awarded to C.G. and DFCAS Dissertation Completion Fellowship awarded to L.M. The computational work was financed in part by the National Institutes of Health (NIH) grants R35GM153297 and R44GM133270 awarded to Y.S. We also acknowledge the access to the computational resources of the PETE Supercomputer provided by the High-Performance Computing Center of Oklahoma State University (OSU) and the OU Supercomputing Center for Education & Research (OSCER) at the University of Oklahoma. Financial support for publication was partly provided by the University of Oklahoma Libraries’ Open Access Fund.
Author information
Authors and Affiliations
Contributions
R.R. conceived the idea. R.R., C.G., and L.M. designed and performed the cryo-EM experiments. L.T. assisted in cryo-EM experiments and model building. C.G., S.D.A., and R.R. designed and performed the biochemical experiments. S.R.A. performed the bioinformatics and structural comparisons of BH and RuvC-H1. A.M.S. and Y.S. performed computational simulations and data analysis. All authors contributed to data analysis, manuscript writing, and editing.
Corresponding author
Ethics declarations
Competing interests
R.R. is an inventor on US patents related to FnoCas12a bridge helix variants (US11459552B2 and US12163166B2; status = granted) that were filed by the University of Oklahoma. These patents describe how amino acid substitutions in the bridge helix of Cas12a increase DNA cleavage selectivity. This manuscript presents the structure and mechanism of one of the bridge helix variants (FnoCas12a-K969P/D970P, abbreviated as FnoCas12aKD2P) that was described in the patents. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chunyi Hu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ganguly, C., Aribam, S.D., dos Santos, A.M. et al. Bridge helix of Cas12a is an allosteric regulator of R-loop formation and RuvC activation. Nat Commun 17, 2126 (2026). https://doi.org/10.1038/s41467-026-68657-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68657-0