Abstract
Chemical imaging probes enable the visualization of dynamic biology; however, engineering high sensitivity in live cells remains challenging. Here we present Sensight, a quantitative multivariate framework that integrates key photophysical and physicochemical descriptors to predict and optimize probe performance. Using a structurally diverse library, we identify five dominant parameters, define their optimal ranges, and unify them into a radar map representation with strong predictive power and intuitive visualization. This framework extends the structure–activity relationship analysis into imaging sensitivity, capturing complex structure–function relationships that shape probe behavior in live cells. Guided by Sensight, we design G3, a superoxide probe with exceptional sensitivity for detecting early oxidative events. The framework’s generality is further demonstrated across distinct systems, including tetrazine–bicyclononyne bioorthogonal chemistry and formaldehyde sensing. Together, these findings establish Sensight as a predictive and generalizable strategy for high-performance probe design, with broad implications for sensing, imaging, and even theranostics.
Similar content being viewed by others
Introduction
Small-molecule tools such as fluorescent probes and bioorthogonal reagents have revolutionized our ability to interrogate biological systems with chemical precision. These tools are indispensable for visualizing biomolecules1,2,3,4,5, tracking dynamic cellular processes6,7,8, and controlling molecular events in situ9,10. Among them, activity-based sensing and bioorthogonal chemistry stand out by leveraging selective chemical reactivity within live cells to generate detectable signals or trigger functional responses11,12,13,14. This reactivity-driven approach underpins key advances in fluorescence imaging, prodrug activation, and biomolecular labeling15,16,17,18, making sensitivity and reaction efficiency critical design priorities.
Achieving high sensitivity is particularly crucial for detecting trace biomolecules or detecting early-stage cellular events. Traditional probe development has focused on maximizing signal gain by developing activatable probes utilizing clever photophysical mechanisms19,20,21, such as Förster resonance energy transfer (FRET)22, photoinduced electron transfer (PET)23, twisted intramolecular charge transfer (TICT)24. While these strategies have yielded powerful activatable probes25,26,27, sensitivity in live cells depends on more than fluorogenic responses. Structural factors such as cell permeability, excitation compatibility, and detection efficiency are equally critical. These properties are interdependent, making rational design inherently complex. Despite the growing demand for high-performance probes, no general framework exists to quantitatively integrate these variables into a predictive, design-oriented model.
To address this gap, we introduce Sensight, a sensitivity engineering strategy integrating multivariate methods for guiding high-performance imaging toolkits. Using a structurally diverse, in-house probe library, we systematically analyzed fifteen descriptors for their contribution to live-cell imaging sensitivity and identified five dominant ones that collectively shape probe performance in cells, including fluorogenic contrast, lipophilicity, polar surface area, excitation mismatch, and quantum efficiency. These parameters were unified into a radar map visualization that enables intuitive assessment and prospective design. The model, termed Sensight, was experimentally validated through single-variable-controlled synthesis and imaging studies. It successfully guided the development of G3, a superoxide-responsive probe with desirable sensitivity for detecting early oxidative bursts in live cells. To demonstrate generality, we further applied Sensight to mechanistically distinct systems, including tetrazine-bicyclononyne (BCN) bioorthogonal conjugation and formaldehyde imaging. In each case, model predictions aligned with experimental performance. Together, Sensight provides a generalizable framework for sensitivity-tuned molecular design. It extends the logic of structure–activity relationships to probe engineering and offers a powerful conceptual and practical toolkit for advancing molecular imaging and chemical biology.
Results
Identifying key descriptors governing imaging sensitivity
To establish a quantitative framework for the rational design of high-sensitivity fluorescent imaging probes, we first sought to define which structural and photophysical parameters most critically influence probe performance in live cells. This required a library of probes with: (1) identical target specificity to eliminate confounding biological factors; (2) consistent sensing chemistry and a single well-defined product to ensure response accuracy; and (3) measurable fluorescence enhancement upon activation to enable quantitative evaluation. 1,2,4,5-Tetrazine (Tz)-based activity-based probes were identified as an ideal system to meet these criteria. These probes react selectively with superoxide to form stable oxadiazoles with robust fluorogenic signals28. Leveraging this reactivity, we designed and synthesized a library of Tz-conjugated probes (A1–J3) covering diverse structural variations (Fig. 1a, Supplementary Table 1).
a The structures of the Tz probes. b Probe sensitivity to superoxide in solution-based (ΔFs) and live-cell imaging (ΔFc) experiments. ΔFs was measured as the fluorescence intensity increase fold of the probe (5 μM) after full conversion to oxadiazole, while ΔFc was the cellular probe fluorescence intensity ratio between cell groups with or without oxidative stress challenge. Probes with relatively high cellular imaging sensitivity, defined as ΔFc ≥ 1.4, were highlighted in red. c Linear regression between normalized (Norm.) ΔFs and normalized ΔFc. d The entropy and the PCC of each descriptor with the sum values annotated. Source data for c, d are provided as a Source Data file.
In solution, all probes underwent quantitative conversion to oxadiazoles upon reaction with superoxide (Supplementary Figs. 1–22), and their fluorogenic fold-change values (ΔFs) were determined (Fig. 1b, Supplementary Figs. 23–44). These ranged from modest to extraordinary enhancements, consistent with tetrazine’s known quenching efficiency29,30,31.
To evaluate their imaging sensitivity in live cells, we employed a H₂O₂-induced superoxide burst model in HepG2 cells32. We first systematically optimized the H2O2 stimulation parameters using probe F-Tz4, a benchmarked Tz-based superoxide probe with well-characterized cellular response behavior28. Titration of H₂O₂ concentration (0–4 mM) and incubation time (0–4 h) revealed that 2 mM for 2 h produced a maximal and reproducible superoxide burst without compromising cell viability (Supplementary Figs. 45, 46). This optimized condition was applied to all subsequent imaging experiments. To quantify live-cell imaging sensitivity, we defined ΔFc as the fold-increase in cellular fluorescence between H2O2-stimulated and unstimulated (Control) conditions (Fig. 1b, Supplementary Figs. 47–56). For each probe, mean fluorescence intensity (MFI) was quantified across three biologically independent replicates. For each replicate, three frames were randomly selected from a single focal plane (non-Z-stack). Fluorescence values were extracted directly from cell-containing regions, and ΔFc was calculated as MFIH2O2/MFIControl. Although all probes showed turn-on behavior, ΔFc values were consistently lower than their ΔFs counterparts, highlighting the limited superoxide availability in cells and underscoring the challenge of achieving high sensitivity in physiological contexts.
We next examined whether a probe’s fluorogenic response in solution (ΔFs) directly correlates with its imaging sensitivity in live cells (ΔFc). A linear regression analysis of ΔFc versus ΔFs revealed little correlation (R² = 0.01, Pearson correlation coefficient (PCC) = 0.12) (Fig. 1c), supporting our central hypothesis that additional structural features beyond fluorogenic contrast contribute critically to in-cell performance. To identify these contributing factors, we compiled a broader set of 15 physicochemical and photophysical descriptors encompassing parameters related to probe solubility, membrane permeability, imaging instrument compatibility, and optical signal quality (Supplementary Tables 2, 3). Each descriptor was normalized to a 0–1 scale and plotted against the normalized ΔFc. Two complementary metrics were then calculated for each descriptor: entropy, to capture distributional uncertainty and potential nonlinear effects; and PCC, to assess linear relationship with ΔFc (Fig. 1d, Supplementary Fig. 57, Table 4). To rank the descriptors, we summed their normalized entropy and PCC values, providing a transparent and reproducible way to highlight parameters that contribute both linearly and nonlinearly to imaging sensitivity. This analysis highlighted five descriptors with the strongest associations to imaging sensitivity, including topological polar surface area (tPSA), the calculated octanol–water partition coefficient (ClogP), the difference between the probe’s maximum excitation wavelength and the fixed excitation wavelength of the imaging instrument (Δλex), ΔFs, and the Stokes shift. Notably, although tPSA and cLogP are generally considered opposing descriptors, both showed a weak negative correlation with imaging sensitivity (ΔFc) in our dataset (Supplementary Fig. 57). This trend likely reflects the fact that their influences on membrane permeability are not strictly linear. These observations further support the use of combined PCC and entropy-based criteria in descriptor selection, which together capture both linear and non-linear contributions to probe performance.
The biological rationale behind the five descriptors is compelling. tPSA and ClogP govern passive membrane permeability33,34,35. Δλₑₓ reflects excitation compatibility with common microscopy setups, while Stokes shift affects detection clarity. Notably, even probes with high ΔFs may fail to perform if they cannot efficiently enter cells or be properly excited under imaging conditions. This multifactorial insight emphasizes that fluorescence contrast alone is insufficient as a predictive measure of probe performance. The significant and previously underappreciated roles of tPSA, ClogP, and Δλₑₓ strongly support the need for an integrated, quantitative model that accounts for the interplay of these diverse factors. These findings set the stage for constructing a predictive framework to guide the rational engineering of next-generation imaging probes.
Constructing Sensight for predicting imaging sensitivity
To determine how each descriptor contributes to probe performance, we first classified the probe library into two categories based on their ΔFc: high sensitivity (ΔFc ≥ 1.4) and low sensitivity (ΔFc < 1.4). Recognizing that extreme values of lipophilicity (ClogP) and polarity (tPSA) impair passive membrane diffusion, we empirically selected the median values within the high-sensitivity group as optimal ClogP (~2.0) and tPSA (~70 Å2) (Supplementary Fig. 58), providing a balanced approximation of probe performance. For ΔFs and Stokes shift, the optimal values were taken as the maximal values observed in the dataset, reflecting the fact that stronger fluorogenicity and larger spectral separation universally enhance imaging sensitivity. For excitation mismatch (Δλₑₓ), because minimal mismatch favors efficient excitation, we incorporated this descriptor through min–max normalization. To enable direct comparison across descriptors of different scales and units, we standardized each parameter using min-max normalization or proximity-to-optimum scoring (Supplementary Table 5). This standardization laid the groundwork for a multivariate regression model capable of quantitatively predicting probe sensitivity in live cells.
We first explored logistic regression to differentiate high- and low-performing probes but found that its binary nature lacked granularity. Instead, we used multivariate linear regression, a strategy commonly employed in medical cohort studies, to derive correlation coefficients reflecting the relative contribution of each descriptor to imaging sensitivity (ΔFc). Using SPSS, we performed regression analysis with the five descriptors as independent variables and ΔFc as the dependent outcome (Fig. 2a). The results revealed that tPSA (r = 0.699), Δλₑₓ (r = 0.570), ΔFs (r = 0.277), and ClogP (r = 0.197) were the most influential contributors to live-cell probe performance, while Stokes shift (r = 0.096) was marginal (Fig. 2b). These rankings reinforced the importance of integrating physiochemical properties and excitation compatibility, parameters that are often overlooked in probe design.
a Flowchart of the model development. b The weights of the descriptors, which are absolute values of the correlation coefficients. c Radar map of probes E1, D3, F2 and G2. d Scatter plot of ΔFc versus the radar map area of each probe. ΔFc was normalized to the maximum value among the probes. Source data for c and d are provided as a Source Data file.
To convert these multivariate results into an easy-to-understand visualization, we first re-scaled each descriptor value by its regression-derived weight to reflect its relative importance (Supplementary Table 6). This was carried out by multiplying the standardized value by the descriptor’s corresponding weight. The resulting adjusted descriptor values were then used to plot a five-dimensional radar map (workflow in Supplementary Fig. 59), enabling intuitive visualization of each probe’s predicted imaging performance (Fig. 2c, Supplementary Fig. 60, Table 7). The area enclosed by the radar map served as a quantitative summary metric. Remarkably, we observed a strong exponential correlation (R2 = 0.97) between the radar map area and actual live-cell imaging sensitivity (ΔFc) across the probe library (Fig. 2d). This sharply contrasts with the negligible correlation when ΔFc was plotted against ΔFs alone (Fig. 1c), confirming the effectiveness of a multivariate approach over traditional single-parameter optimization. Overall, these findings establish a robust and interpretable model, Sensight, that not only captures the multifactorial nature of imaging probe performance but also offers a quantitative and visual tool for guiding probe development.
Validating Sensight through single-variable experiments
While Sensight quantitatively explains the imaging sensitivity of the initial probe library, we sought further validation to confirm its reliability. To this end, we adopted a controlled single-variable approach, where new probes were synthesized to selectively test the four most influential descriptors identified, tPSA, Δλₑₓ, ΔFs, and ClogP (Fig. 3a).
Controlled single-variable assessment of the top three influential descriptors, tPSA, Δλex, and ΔFs. Structures of the three pairs of probes are shown in (a). Their key descriptor values are shown in b–d, with key values highlighted in blue to facilitate comparison. Representative confocal imaging results are shown in e–g, and the quantitative data are shown in (h–j). For confocal imaging, HepG2 cells were treated with 2 mM H2O2 for 2 h to induce oxidative stress, stained with the indicated probe (5 μM, 30 min), and then imaged. F2 and W1 channel: λex = 488 nm, λem = 489–600 nm; B1, D3, E1 and W2 channel: λex = 405 nm, λem = 406–550 nm. Scale bars, 20 μm. Data were normalized to their respective control group and presented as mean ± s.d. (n = 9). P values were analyzed by a two-tailed unpaired t-test. Source data for h–j are provided as a Source Data file.
We first evaluated tPSA, which exhibited the strongest correlation with probe performance. Although probe F2 displayed a robust fluorogenic response in vitro (ΔFs = 28.6), it showed low imaging sensitivity in cells (ΔFc = 1.14). Sensight attributed this discrepancy to its suboptimal tPSA value (Fig. 3b). To test this hypothesis, we designed and synthesized probe W1, which maintained a similar Δλₑₓ to F2 but featured a markedly improved tPSA. Both probes showed quantitative conversion to oxadiazoles upon reaction with superoxide, and F2 even had a slightly higher ΔFs (Supplementary Figs. 61–63, Table 8). Nevertheless, W1 achieved significantly greater imaging sensitivity (ΔFc = 1.74) (Fig. 3e, h), substantiating tPSA as a critical determinant for probe permeability and in-cell performance.
Then, we investigated Δλₑₓ, the second-highest ranked descriptor. We compared probe D3 with its analog W2, designed to closely match D3 in both ΔFs and tPSA while significantly reducing Δλₑₓ (Fig. 3c, Supplementary Figs. 64–66, Table 9). As predicted, W2 exhibited higher live-cell imaging sensitivity than D3 (Fig. 3f, i), confirming that excitation compatibility with the imaging system is a major factor influencing probe efficacy.
Next, we assessed the contribution of ΔFs by comparing probe E1 with its isomeric counterpart B1, which shared similar tPSA and Δλₑₓ but exhibited a higher fluorogenic turn-on response (Fig. 3d, Supplementary Figs. 67–69, Table 10). As expected, B1 delivered stronger imaging sensitivity (Fig. 3g, j), validating ΔFs as a relevant driver of performance.
Lastly, the effect of ClogP was evaluated by synthesizing probe W3 and comparing its imaging performance with C2. W3 demonstrated comparable tPSA (108.08 Ų), ΔFs (77.4), and Δλex (66 nm) to that of probe C2 (tPSA 108.08 Ų, ΔFs 78.3, and Δλex 55 nm), but much improved ClogP (2.12 for W3 and 0.72 for C2) (Supplementary Figs. 70–72, Table 11). Under the established imaging conditions, W3 was more sensitive (ΔFc = 1.39 for W3 and 1.10 for C2), highlighting the significant impact of ClogP on probe imaging sensitivity.
Together, these experiments provide orthogonal validation of Sensight’s core principles. By isolating individual variables in structurally related probes, we demonstrated that improvements in tPSA, Δλₑₓ, ΔFs, and ClogP lead to predictable gains in imaging sensitivity. These results not only reinforce the predictive power of Sensight but also underscore its value as a rational design tool for next-generation activity-based probes.
Validating Sensight through de novo probe design
Having established the predictive power of Sensight, we next explored its utility in de novo probe design. We aimed to develop a superoxide-responsive probe with enhanced imaging sensitivity by prioritizing structural features that maximize performance.
Thirteen probe candidates were designed in silico by conjugating 1,2,4,5-tetrazine to various fluorophores via different linkers. For each candidate, key parameters, including tPSA, Δλₑₓ, ClogP, and Stokes shift, were calculated and standardized to their optimal values (Supplementary Tables 12–15). As ΔFs could not be predicted a priori, it was held constant across all designs. We then generated radar plots for each candidate and ranked their predicted imaging sensitivity by radar area (Fig. 4a, Supplementary Table 15).
a In silico-designed probes, which were ranked by their predicted radar area. b The live cell imaging results of K2 and G3, and c the quantified data. HepG2 cells were pretreated with 2 mM H2O2 for 2 h, stained with the probe (5 μM, 30 min), and then imaged. d The key descriptor values of K2 and G3 with radar area values highlighted in blue, and e their radar maps. f Fluorescence images of A431 cells first stimulated with various concentrations of EGF, stained with G3 or DHE (each 5 μM, 30 min), and then imaged. g Fluorescence images of HepG2 cells pretreated with paraquat (2 h), stained with G3 or DHE (each 5 μM, 30 min), and then imaged. h, i The quantitative intensity data for (f). j, k The quantitative intensity data for (g). G3 channel: λex = 488 nm, λem = 489–600 nm, DHE channel: λex = 561 nm, λem = 562–650 nm, K2 channel: λex = 710 nm, λem = 712–834 nm. Scale bars, 20 μm. The quantified data were normalized to their respective control group and presented as mean ± s.d. (n = 9). P values were analyzed by a two-tailed unpaired t-test (for c) or a two-way ANOVA test (for h–k) (ns, no significance). Source data for c, e, h–k are provided as a Source Data file.
To experimentally validate the predictive capability of Sensight, we selected and synthesized six representative probes (Tz-1, Tz-2, Tz-3, Tz-8 (G3), Tz-9, and Tz-13 (K2)). The selection was guided by two considerations. First, their predicted imaging sensitivity values spanned the entire distribution of the virtual design library, ensuring evaluation across low-, medium-, and high-performance regions. Second, they could be readily synthesized. All compounds were confirmed to undergo quantitative conversion to the corresponding oxadiazoles upon reaction with superoxide (Supplementary Figs. 73–88; Supplementary Tables 16–21). In HepG2 cells under H₂O₂-induced superoxide burst, G3 displayed the highest imaging sensitivity with a ΔFc of 2.67, approximately twice that of K2 (ΔFc = 1.24) (Fig. 4b, c). The experimentally observed trend in sensitivity (ΔFc: G3 (2.67) > Tz-3 (1.56) > Tz-9 (1.45) > Tz-1 (1.25) > K2 (1.24)) closely matched the ranking predicted by Sensight, thereby supporting the robustness of the model. Although the experimentally measured ΔFc of Tz-2 (1.20) deviates slightly from the predicted rank order, this difference is expected given the very small separation among the predicted radar map areas of Tz-1 (0.4971), Tz-2 (0.4927), and K2 (0.4857) (Supplementary Table 15). Such minor rank inversions among closely clustered probes fall within the experimental variability of live-cell imaging and do not affect the overall predictive trend of Sensight.
Notably, G3 displayed a near-optimal combination of key descriptors, including tPSA, Δλₑₓ, ΔFs, and ClogP, and showed the largest radar area among all Tz-based probes in our dataset (Fig. 4d, e), consistent with its superior live-cell performance. To evaluate G3’s biological utility, we benchmarked it against dihydroethidium (DHE), a widely used commercial probe for superoxide imaging. While both G3 and DHE responded to superoxide, DHE exhibited broad reactivity to multiple reactive oxygen species (ROS), compromising specificity (Supplementary Figs. 89–91). In contrast, G3 demonstrated higher selectivity and greater storage stability (Supplementary Fig. 92), offering advantages in both precision and practical use.
We then tested G3 in two biological models that involve subtle superoxide bursts. The first is EGF-induced redox signaling. The binding of epidermal growth factor (EGF) to its receptor (EGFR) transiently activates NADPH oxidases (NOXs), generating a localized superoxide burst36. Using A431 cells, which overexpress EGFR37, we observed EGF-dose-dependent fluorescence increases with both G3 and DHE (Supplementary Fig. 93). However, only G3 detected superoxide at lower EGF concentrations (Fig. 4f, h, i), highlighting its superior sensitivity. The second is early-stage oxidative stress from paraquat exposure. Paraquat triggers mitochondrial superoxide production, particularly during early exposure38,39. In HepG2 cells treated with increasing doses of paraquat, G3 detected significant fluorescence even at low concentrations where DHE failed (Fig. 4g, j, k). Complementary JC-1 staining confirmed reduced mitochondrial membrane potential, even at these early stages (Supplementary Fig. 94)40, validating G3’s ability to detect early oxidative damage.
We also benchmarked G3 against DHE and MitoSOX to evaluate its capability to dynamically track H₂O₂-induced superoxide bursts. Cells were first loaded with each probe and then challenged with 1 mM H₂O₂, and the resulting fluorescence changes were recorded in real time using lattice light-sheet microscopy (Supplementary Fig. 95). DHE exhibited minimal fluorescence increase, reflecting its low sensitivity under these conditions. MitoSOX, a mitochondria-targeted analog of DHE, rapidly detected the superoxide burst, consistent with mitochondria being the primary site of superoxide generation in this model, highlighting the importance of organelle targeting for enhancing probe sensitivity. Remarkably, G3, despite lacking mitochondrial targeting, was able to capture the superoxide burst with a ~1.5-fold fluorescence increase within 30 min, demonstrating sufficient sensitivity to monitor cytosolic superoxide dynamics.
Together, these findings demonstrate that G3 is a Sensight-guided, rationally designed probe with unmatched sensitivity and specificity for superoxide detection in live cells. These results affirm the predictive strength and practical utility of Sensight, and establish its potential as a general strategy for engineering high-performance fluorescent probes. In its current form, Sensight deliberately centers on global physicochemical and photophysical descriptors, such as tPSA, ClogP, ΔFs, Δλex, and Stokes shift, that broadly govern intracellular accessibility, excitation efficiency, and fluorogenic turn-on. This choice reflects a rational balance between mechanistic relevance and model generalizability. However, subcellular localization can also modulate observed imaging performance by shaping local analyte availability and modulating microenvironment-dependent fluorescence behaviors. Such localization effects introduce an additional, highly context-dependent layer of complexity that extends beyond the current descriptor space. Future iterations of Sensight could incorporate localization-relevant parameters as more systematic datasets become available.
Sensight generality in the tetrazine-BCN bioorthogonal system
Sensight not only highlighted ΔFs as a central determinant of probe performance but also revealed other underappreciated structural parameters, particularly tPSA and Δλₑₓ, as critical contributors to live-cell imaging sensitivity. Recognizing that both activity-based sensing and bioorthogonal chemistry rely on specific chemical transformations within biological systems to produce a signal or effect, we hypothesized that Sensight might have broader applicability beyond redox-based probes.
To test this, we applied Sensight to the well-established Tz-BCN bioorthogonal system (Fig. 5a). Tz-BCN cycloadditions enable catalyst-free, rapid, and selective labeling in live cells and have seen wide adoption in chemical biology, from biomolecular tracking to prodrug activation41,42,43,44,45. Before proceeding, we verified that BCN treatment alone did not induce oxidative stress in cells, ensuring that the fluorescence responses reflected bioorthogonal conjugation rather than redox activity (Supplementary Fig. 96).
a Tz-BCN reaction scheme. b–d The key descriptor values of indicated probes, with key values highlighted in blue. e Representative confocal images of HepG2 cells after cellular Tz-BCN bioorthogonal reaction. Cells were pre-loaded with various concentrations of BCN, treated with the indicated probe (5 μM, 30 min), and then imaged. B1, D3, E1 and W2 channel: λex = 405 nm, λem = 406–550 nm; F2 and W1 channel: λex = 488 nm, λem = 489–600 nm. Scale bars, 20 μm. f–h The quantitative intensity data for (e). The data were normalized to the respective control group (BCN 0 μM) and presented as mean ± s.d. (n = 9). P values were analyzed by a two-way ANOVA test, and the exact P values for *** were below 0.0001; ns represents no significance. Source data for f–h are provided as a Source Data file.
To assess whether Sensight could predict fluorogenic performance in this context, we first selected two Tz probes, B1 and E1 (Fig. 3a), which have similar tPSA and Δλₑₓ, but differ markedly in their fluorogenic signal change (ΔFs) upon BCN ligation (Fig. 5b; Supplementary Figs. 97–100, Tables 22, 23). Radar map analysis predicted B1 to exhibit superior imaging sensitivity (Supplementary Fig. 101). Live-cell labeling experiments confirmed this. Upon BCN treatment, B1 produced significantly higher ΔFc values than E1, consistent with Sensight predictions (Fig. 5e, f, Supplementary Fig. 102). This comparison reinforces ΔFs as a primary determinant of probe performance in the bioorthogonal context. We next compared probes F2 and W1 (Fig. 3a), which exhibit similar Δλₑₓ and ΔFs values but differ in tPSA, with W1 matching more closely to the predicted optimal range (Fig. 5c, Supplementary Figs. 103–108, Tables 24, 25). Consistent with W1’s larger radar map area, BCN-triggered fluorescence activation in live cells was significantly higher for W1 than for F2 (Fig. 5e, g), supporting tPSA as an independently predictive variable. To further isolate the contribution of Δλₑₓ, we evaluated probes D3 and W2 (Fig. 3a), which differ primarily in Δλₑₓ while maintaining similar values for the other parameters. As predicted, W2 with a smaller Δλₑₓ, displayed improved imaging sensitivity relative to D3 (Fig. 5d, e, h; Supplementary Figs. 109–114, Tables 26, 27), validating Δλₑₓ as a robust predictor across probe scaffolds. We then assessed the influence of lipophilicity and found that ClogP also exerted a strong effect on imaging sensitivity. Probe W3, which possesses a more favorable ClogP, outperformed C2 under identical experimental conditions (Supplementary Figs. 115–120, Tables 28, 29), further demonstrating the generality of Sensight’s predictive framework. Finally, to probe the upper performance limit predicted by Sensight, we tested G3, the highest-ranked structure from the radar map analysis. G3 yielded strong intracellular fluorescence activation even at 25 μM BCN, a level at which the control probe B3 required >100 μM BCN to elicit a similar response (Supplementary Figs. 121–126, Tables 30, 31). Taken together, these results extend the predictive validity of Sensight to the Tz-BCN bioorthogonal platform, confirming its broad utility for rational probe design beyond redox chemistry. The framework could be applied to optimize other reaction-based or triggered probe systems, accelerating the development of next-generation molecular tools for live-cell imaging.
Extending Sensight generality to formaldehyde imaging probes
Building on the successful validation in the Tz–BCN system, we next applied Sensight to a mechanistically distinct platform, formaldehyde (FA) imaging probes. FA plays a critical role in one-carbon metabolism, and its dysregulation has been implicated in diverse pathologies, including neurodegeneration and cancer46,47,48. While numerous FA-responsive probes have been developed49,50,51,52,53, systematic evaluation and predictive modeling of their imaging sensitivity remain unexplored. This case study thus offers a dual opportunity to test the generality of our framework beyond tetrazine chemistry and to provide improved tools for elucidating FA’s biological functions.
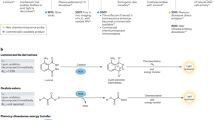
We focused on FA probes leveraging the aza-Cope rearrangement (Fig. 6a), a widely used strategy that enables selective FA sensing via reaction with primary amines49,50,51,52,53. We built a structurally and spectrally diverse probe library, deliberately covering several literature probes for benchmarking purposes (Fig. 6b). For each probe, we computed tPSA and Clog P values. Stokes shift, and Δλₑₓ were determined by the excitation instrument settings and the fluorophore’s spectral properties, both of which are well-defined for standard imaging platforms. While ΔFs remained unpredictable precisely, literature-reported turn-on efficiencies allow reasonable estimation within an appropriate range and were temporarily held constant herein (Supplementary Tables 32–35). Based on these values, radar plots were constructed to generate predicted sensitivity ranking (Fig. 6c), enabling pre-synthetic assessment of probe imaging sensitivity.
a Sensing mechanism of formaldehyde probes (FPs). b Probe structures in this study. c Predicted imaging sensitivity of the probes, which was obtained as the normalized predicted radar map area of each probe by the model. d Validation of LPS-induced inflammation in BV2 cells by quantifying IL-1β secretion. Data shown were mean ± s.d. (n = 3). e Fluorescence images of BV2 cells first stimulated with LPS (24 h) and then stained with FP9 (5 μM, 1 h). Quantification data were shown in g as mean ± s.d. (n = 9). f The measured probe imaging sensitivity, which was the normalized data of ΔFc obtained under the LPS 4 ng/mL condition. h Correlation between predicted ΔFc and measured ΔFc values of the probes, with the bar chart showing the mean difference with 95% confidence intervals. FP1, FP2 and FP5 channel: λex = 405 nm, λem = 406–504 nm, FP3 channel: λex = 405 nm, λem = 450–504 nm, FP4 channel: λex = 405 nm, λem = 450–520 nm, FP6, FP7 and FP8 channel: λex = 488 nm, λem = 489–550 nm, FP9 channel: λex = 561 nm, λem = 565–700 nm. Scale bar, 20 μm. P values were analyzed by a two-way ANOVA test; ns represents no significance. Source data for c, d, f–h are provided as a Source Data file.
To validate Sensight’s predictions, we synthesized the probes, confirmed their full conversion to the products under excess FA, and quantified their fluorogenic responses (Supplementary Figs. 127–135, Tables 36–44). For live-cell imaging, we employed an LPS-induced inflammation model in BV2 microglia, where endogenous FA is upregulated as part of the inflammatory cascade54,55. At 2 ng/mL LPS, which sufficed to trigger IL-1β secretion (Fig. 6d), only FP9 showed a detectable FA response (Fig. 6e, g). At 4 ng/mL, all probes responded (Supplementary Figs. 136–140), and the observed sensitivity ranking (FP9 > FP5 > FP4 > FP8 > FP3 > FP7 > FP6 > FP1 > FP2) closely matched Sensight’s predictions (Fig. 6f). Strikingly, a strong linear correlation (R² = 0.93) was observed between predicted and experimentally measured sensitivities (Fig. 6h), demonstrating Sensight’s quantitative accuracy and generalizability across probe chemistries. Notably, our top-performing probe, FP9, also outperformed FP4 in previous independent studies50, further reinforcing the robustness and translational relevance of our design framework.
Together, these results establish that Sensight, initially developed for superoxide-sensing tetrazine probes, can accurately guide probe optimization in chemically distinct systems. This finding underscores the broad applicability of Sensight for improving imaging sensitivity across mechanistically diverse platforms, offering a powerful approach for small-molecule probe design.
Discussion
Fluorescence imaging with small-molecule probes has become a cornerstone technology for tracking biomolecular dynamics in live systems, enabling high-resolution visualization of key analytes, such as proteins, metabolites, ions, reactive oxygen species, etc. This capability is critical for advancing early diagnostics and therapeutics by uncovering molecular events that signal disease onset. Yet, achieving sufficient imaging sensitivity, particularly for low-abundance or transient species, remains a major challenge, owing to the complex interplay of structural, physicochemical, and photophysical properties that govern probe performance in cells.
To address this unmet need, we introduce Sensight, a quantitative multivariate model that systematically links five key descriptors (tPSA, ΔFs, Δλex, ClogP, Stokes shift) to imaging sensitivity in live cells. Notably, although the initial descriptor–performance relationships were derived from a finite set of probes, the chemical space they occupy spans a broad and representative range of values, including tPSA (52.68–141.26), ClogP (0.35–7.32), ΔFs (1.7–276), Δλₑₓ (5–85), and Stokes shift (22–125). This wide coverage captures diverse photophysical and physicochemical properties relevant to intracellular imaging. Importantly, in single-variable validation experiments and de novo probe design cases, the experimentally measured sensitivities for probes newly synthesized align with Sensight predictions, supporting the model’s reliability and highlighting it as a practical and predictive framework for guiding probe optimization beyond empirical trial-and-error.
Sensight’s utility successfully guided the de novo design of G3, an ultra-sensitive superoxide probe capable of detecting early oxidative bursts in live cells. G3 enabled real-time visualization of superoxide dynamics under both physiological (EGF stimulation) and pathological (paraquat stress) conditions, highlighting the power of sensitivity-focused design for capturing fleeting biochemical signals that are otherwise undetectable. Beyond activity-based sensing of superoxide, Sensight’s generalizability was confirmed in two conceptually distinct systems, the Tz-BCN bioorthogonal reaction platform and a series of formaldehyde-selective probes. In both contexts, predicted sensitivity rankings closely matched experimental outcomes, including a strong linear correlation (R² = 0.93) in the formaldehyde imaging system. These results demonstrate that Sensight can extend beyond a single chemical transformation to inform broader small-molecule probe design across diverse reactivity platforms. Moreover, by using ΔFs as a mechanism-agnostic descriptor that captures the net fluorescence gain rather than any specific modulation mechanism, Sensight is inherently extensible and can, in principle, incorporate probes based on diverse activation chemistries in future studies.
This study quantitatively dissects how multiple physicochemical and photophysical features collectively determine probe sensitivity in live-cell imaging, enabling a shift from empirical tuning to data-driven design. A key limitation is that ΔFs cannot be computed a priori, as it reflects mechanism-specific excited-state processes and microenvironment-dependent quenching pathways. Thus, prospective design is chiefly supported by the four fully predictable descriptors, tPSA, ClogP, Δλex, and Stokes shift, which govern membrane permeability, excitation efficiency, and spectral separation. In practice, Sensight integrates these predictable features with empirically determined ΔFs to provide a pragmatic and mechanistically agnostic framework for rational probe optimization. While the present model was developed from a targeted compound set and a defined descriptor pool, its strong predictive power lies as the foundation for future refinement. As larger and more diverse probe libraries become available, this framework could be extended to incorporate additional descriptors and leverage machine-learning approaches to refine predictive power, opening avenues for real-time molecular imaging, early disease detection, and the broader application of chemical tools in complex biological systems.
Methods
General methods
All chemicals were from commercial supplies and used without further purification except otherwise indicated. 1H NMR spectra were measured on a Bruker 500 Fourier transform spectrometer (500 MHz). 13C NMR spectra were obtained on a Bruker 500 Fourier transform spectrometer (126 MHz) or Asend EVO 600M (151 MHz). Chemical shifts (δ) for 1H and 13C NMR spectra were given in ppm. The residual solvent signals were used as references for 1H and 13C NMR spectra and the chemical shifts were converted to the TMS scale (CDCl3, 7.26 ppm for 1H NMR and 77.16 ppm for 13C NMR; CD3OD, 3.31 ppm for 1H NMR and 49.00 ppm for 13C NMR; (CD3)2SO, 2.50 ppm for 1H NMR and 39.52 ppm for 13C NMR). All chemical shifts were reported in parts per million (ppm) and coupling constants (J) in Hz. The following abbreviations were used to explain the multiplicities: d = doublet, t = triplet, m = multiple, dd = doublet of doublets. High-resolution mass spectra (HRMS) for new compounds were recorded on a Waters TOFMS GCT Premier using ESI ionization. Liquid chromatography equipped with a low-resolution mass detector (LC–MS) was conducted on an Agilent Infinity Lab LC/MSD. High-performance liquid chromatography (HPLC) was measured on an Agilent Technologies 1260 Infinity system. Fluorescence spectra were measured on an Agilent Cary Eclipse Fluorescence Spectrophotometer. Absorption spectra were collected using a Hitachi U-3010 spectrophotometer. The confocal cell imaging experiments were carried out on a Zeiss LSM900 live cell fluorescence microscope, except that K2 was recorded on a Leica STELLARIS 5 confocal microscope. Dynamic imaging was performed with a Zeiss lattice-lightsheet 7 microscope.
Probe synthesis and structure characterization
Tz-based superoxide probes and FP-series formaldehyde probes were synthesized according to procedures outlined in Supplementary Figs. 141–160. Detailed procedures and structure characterization data were described in the supplementary information.
HPLC analysis procedures
To record the reactivity of A1–K2 towards \({{{{\rm{O}}}}}_{2}^{-}\) or BCN, each compound stock solution (5 mM, in DMSO) was diluted in MeCN to make a 50 μM solution. Then KO2 (0–8 eq) or BCN (20 eq) was added. The mixture was then allowed to react either with \({{{{\rm{O}}}}}_{2}^{-}\) for 30 min or with BCN for 2 h at ambient temperature, followed by HPLC analysis. The peak area of the remaining residue and product was used to calculate the conversion rate and yield. The detailed liquid chromatography methods were reported in the supplementary information. The maximum peak area of the remaining residue and the product were normalized and mapped, respectively. The reactivity of FP1–FP9 with formaldehyde (FA) was determined following the procedures described above, except that 20 eq FA was added and incubated at ambient temperature for 2 h. Then, the solution pH was adjusted to 9. After 30 min, the mixture was analyzed by HPLC. Liquid chromatography methods are described in the analytical methods section in the supplementary information.
Optical response analysis
To measure the absorption spectra, the stock solution of A1–K2 was diluted with MeCN containing 1% 18-crown-6. \({{{{\rm{O}}}}}_{2}^{-}\) was added to this solution. After 30 min of incubation, the mixture was diluted with PBS buffer (10 mM, pH = 7.4, F2–K2) or MeCN (A1–E3) and then measured. The final concentration of the probe was 5 μM. The same tube of solution was used to record the fluorescence spectra. The mixture was then measured for fluorescence. To determine the quantum yields, quinine sulfate (Фstandard = 0.577 in 0.1 M H2SO4), fluorescein (Фstandard = 0.95 in 0.1 M NaOH), and SiR 650 (Фstandard = 0.31 in PBS (pH 7.4)) were used as standards according to a published method. For the probe, fluorescein, quinine sulfate, and SiR 650, the absorbance spectra were measured within an absorbance range of 0.01–0.05. The quantum yield was calculated according to Eq. (1):
where Φ is the quantum yield, ΣF is the integrated fluorescence intensity, Abs is the absorbance at the excitation wavelength, and n represents the refractive index of the solvent.
To test the selectivity of the probes, G3 and DHE were diluted with PBS buffer (10 mM, pH 7.4) to make a solution of 5 μM. Aliquots of this solution were then treated with various analytes whose stock solutions were prepared according to the methods described in the supplementary information. The volume change (ca. 1‰) caused by the addition of analytes could be negligible. After incubating at room temperature for 30 min, the fluorescence spectrum was collected. All the fluorescence and absorption spectra data were processed via the software Origin 2022.
Cell culture
The cell lines utilized in this study, including A431, BV2, HeLa, and HepG2, were acquired from the Stem Cell Bank of the Chinese Academy of Sciences. All cultures were maintained at 37 °C in a humidified atmosphere containing 5% CO₂. Cells were grown in high-glucose Dulbecco’s modified Eagle medium (DMEM; Gibco), which was supplemented with 10% (v/v) fetal bovine serum (FBS; PAN) and a 1% antibiotic–antimycotic solution (MeilunBio). To ensure cells remained in an exponential growth phase, cells were passaged when they reached 80–90% confluence.
Fluorescence confocal imaging
All cells were seeded in 15 mm glass-bottomed dishes and cultured under standard conditions for 24 h prior to experimentation. Unless otherwise specified, the following general protocol was applied: following the indicated treatments or incubation periods, cells were washed three times with PBS, then incubated with the respective probe at a final concentration of 5 µM in serum-free medium for 30 min at 37 °C in the dark. Cells were washed with PBS three times before imaging.
For optimizing the oxidative stress condition experiment, HepG2 cells were treated with serum-free medium containing varying concentrations of H₂O₂ (Sinopharm) for specified durations before staining with the F-Tz4 probe. For evaluating the Tz series probes, a standardized oxidative stress condition (2 mM H₂O₂, 2 h) was applied, and cells were fixed before imaging.
For the selectivity experiment, HepG2 cells were pre-incubated with Tiron or TEMPO (Aladdin; 300 µM each) in serum-free DMEM for 1 h, followed by co-treatment with the same scavenger and H₂O₂ (2 mM) for an additional 2 h before staining with probe G3 or DHE (MedChemExpress).
For the experiment on EGF-induced superoxide, A431 cells were co-incubated with serum-free medium containing different concentrations of EGF (MedChemExpress; 0, 8, 16, 50, 100, or 200 nM) together with G3 or DHE for 30 min and imaged.
For the experiment on paraquat-induced superoxide, HepG2 cells were exposed to paraquat (Aladdin; 0, 0.5, or 1 mM) in serum-free medium for 2 h before staining with probe G3, DHE, or JC-1 (MedChemExpress).
For a live-cell kinetic imaging experiment, HeLa cells were first loaded with the probe (G3, DHE, or MitoSOX (MedChemExpress)) for 30 min and washed. Immediately after the addition of H₂O₂ (1 mM), time-lapse imaging was initiated and continued for 30 min.
For bioorthogonal experiment, HepG2 cells were pretreated with BCN (MedChemExpress; 0, 25, 50, 100, or 200 µM) in serum-free medium for 1 h before B1, B3, C2, D3, E1, G3, W1, W2 or W3 staining.
For the endogenous formaldehyde induction experiment, BV2 cells were stimulated with LPS (MedChemExpress; 0, 1, 2, or 4 ng/mL) in serum-free medium for 24 h before FP1-FP9 staining.
Fluorescence images were primarily acquired using a Zeiss LSM900 confocal microscope equipped with ZEISS ZEN 3.6 (blue edition) software, with specific images from K2 (channel: λex = 710 nm, λem = 712–834 nm). The imaging parameters were set as follows: A1–B3, D2–E3, Tz–1–Tz–3 and W2 channel: λex = 405 nm, λem = 406–550 nm, C2, C3, F2-H3, W1 and W3 channel: λex = 488 nm, λem = 489–600 nm, I2–J3, DHE and MitoSOX channel: λex = 561 nm, λem = 562–650 nm. JC-1 channel: λex = 488 nm, λem = 489–560 nm and λex = 561 nm, λem = 562–700 nm. FP1, FP2 and FP5 channel: λex = 405 nm, λem = 406–504 nm, FP3 channel: λex = 405 nm, λem = 450–504 nm, FP4 channel: λex = 405 nm, λem = 450–520 nm, FP6, FP7 and FP8 channel: λex = 488 nm, λem = 489–550 nm, FP9 channel: λex = 561 nm, λem = 565–700 nm.
To ensure consistency in background signal, identical microscope parameters were applied across all conditions within a single experiment. Each experiment was performed in three independent biological replicates, with three frames captured from a randomly selected single focal plane (without Z-stack) per replicate. Image analysis was conducted using the ImageJ software. Fluorescence intensity was quantified by measuring the mean gray value, without background subtraction or application of a Gaussian blur filter. The final dataset represented the pooled measurements from all parallel samples across the three independent experiments.
Analysis of IL-1β secretion via ELISA
Following the indicated treatments, cell culture supernatants were harvested and clarified by centrifugation at 21,000×g for 15 min at 4 °C. The clarified supernatant was carefully transferred to a new microcentrifuge tube. Total protein concentration in the supernatant was determined using a Bicinchoninic acid (BCA) protein quantification assay.
IL-1β levels were quantified using a commercial enzyme-linked immunosorbent assay (ELISA) kit according to the manufacturer’s protocol (ABclona, RK00006). Briefly, samples and serially diluted standards were loaded into the antibody-precoated wells. Subsequently, biotin-conjugated detection antibody and horseradish peroxidase (HRP)-labeled streptavidin were added sequentially, with thorough washing steps between each incubation. The colorimetric reaction was initiated by adding the 3,3′,5,5′-tetramethylbenzidine (TMB) substrate solution. The optical density (OD) of each well was immediately measured at a wavelength of 450 nm using a SpectraMax M5 microplate reader. Raw OD values from sample and standard wells were corrected by subtracting the average OD of the blank (assay diluent only) wells. A standard curve was generated by plotting the corrected OD values of the standards against their known concentrations. The concentration of IL-1β in each sample was then interpolated from this standard curve.
Descriptors extraction and analysis for model development
ClogP and tPSA were obtained using ChemDraw 23.1.1, MR, fchar, nS, and log S were acquired from the ADMETlab 3.0 (https://admetlab3.scbdd.com/). The extraction of the remaining descriptors was performed in a Jupyter Notebook environment within the Anaconda (25.5.1) platform, utilizing the Python (3.13.5) programming language and the RDKit (2025.03.6) open-source cheminformatics toolkit. Normalized, standardized, and adjusted descriptors were obtained by Origin 2022. Standardized descriptors were obtained as described in Supplementary Table 5. Then, standardized descriptors were used multiple linear regression process in IBM SPSS Statistics 27.0 software for model construction. The non-standardized coefficient is used as the correlation coefficient of each descriptor.
Statistics and reproducibility
All the statistical analysis was performed with Graphpad Prism 8.0 software. A two-tailed unpaired Student’s t-test, or two-way ANOVA test, was used for data statistics. All the cell imaging experiments were performed with three biologically independent replicates.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. All other datasets are included in the Supplementary Information and are available from the corresponding authors upon request. Source data are provided with this paper.
References
Xue, L., Karpenko, I. A., Hiblot, J. & Johnsson, K. Imaging and manipulating proteins in live cells through covalent labeling. Nat. Chem. Biol. 11, 917–923 (2015).
Braselmann, E., Rathbun, C., Richards, E. M. & Palmer, A. E. Illuminating RNA biology: tools for imaging RNA in live Mammalian cells. Cell Chem. Biol. 27, 891–903 (2020).
Paige, J. S., Nguyen-Duc, T., Song, W. & Jaffrey, S. R. Fluorescence imaging of cellular metabolites with RNA. Science 335, 1194–1194 (2012).
Ohata, J., Bruemmer, K. J. & Chang, C. J. Activity-based sensing methods for monitoring the reactive carbon species carbon monoxide and formaldehyde in living systems. Acc. Chem. Res. 52, 2841–2848 (2019).
Grover, K., Koblova, A., Pezacki, A. T., Chang, C. J. & New, E. J. Small-molecule fluorescent probes for binding- and activity-based sensing of redox-active biological metals. Chem. Rev. 124, 5846–5929 (2024).
Huppertz, M.-C. et al. Recording physiological history of cells with chemical labeling. Science 383, 890–897 (2024).
Abdelfattah, A. S. et al. Bright and photostable chemigenetic indicators for extended in vivo voltage imaging. Science 365, 699–704 (2019).
Zhu, W. et al. Chemigenetic indicators based on synthetic chelators and green fluorescent protein. Nat. Chem. Biol. 19, 38–44 (2023).
Wang, X. et al. Machine-learning-assisted universal protein activation in living mice. Cell 188, 3696–3714.e24 (2025).
Schnell, C. Time-resolved protein activation. Nat. Methods 16, 579–579 (2019).
Bruemmer, K. J., Crossley, S. W. M. & Chang, C. J. Activity-based sensing: a synthetic methods approach for selective molecular imaging and beyond. Angew. Chem. Int. Ed. Engl. 59, 13734–13762 (2020).
Chan, J., Dodani, S. C. & Chang, C. J. Reaction-based small-molecule fluorescent probes for chemoselective bioimaging. Nat. Chem. 4, 973–984 (2012).
Prescher, J. A. & Bertozzi, C. R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Scinto, S. L. et al. Bioorthogonal chemistry. Nat. Rev. Methods Prim. 1, 30 (2021).
Hartung, K. M. & Sletten, E. M. Bioorthogonal chemistry: bridging chemistry, biology, and medicine. Chem. 9, 2095–2109 (2023).
Ueno, T. & Nagano, T. Fluorescent probes for sensing and imaging. Nat. Methods 8, 642–645 (2011).
Chang, C. J. Introduction: fluorescent probes in biology. Chem. Rev. 124, 11639–11640 (2024).
Lemieux, G. A., de Graffenried, C. L. & Bertozzi, C. R. A fluorogenic dye activated by the Staudinger ligation. J. Am. Chem. Soc. 125, 4708–4709 (2003).
Devaraj, N. K., Hilderbrand, S., Upadhyay, R., Mazitschek, R. & Weissleder, R. Bioorthogonal turn-on probes for imaging small molecules inside living cells. Angew. Chem. Int. Ed. 49, 2869–2872 (2010).
Wang, L. et al. A general strategy to develop cell permeable and fluorogenic probes for multicolour nanoscopy. Nat. Chem. 12, 165–172 (2020).
Wu, L. et al. Förster resonance energy transfer (FRET)-based small-molecule sensors and imaging agents. Chem. Soc. Rev. 49, 5110–5139 (2020).
Niu, H. et al. Photoinduced electron transfer (PeT) based fluorescent probes for cellular imaging and disease therapy. Chem. Soc. Rev. 52, 2322–2357 (2023).
Wang, C. et al. Twisted intramolecular charge transfer (TICT) and twists beyond TICT: from mechanisms to rational designs of bright and sensitive fluorophores. Chem. Soc. Rev. 50, 12656–12678 (2021).
Yuan, L., Lin, W., Zheng, K. & Zhu, S. FRET-based small-molecule fluorescent probes: rational design and bioimaging applications. Acc. Chem. Res. 46, 1462–1473 (2013).
Sun, W., Li, M., Fan, J. & Peng, X. Activity-based sensing and theranostic probes based on photoinduced electron transfer. Acc. Chem. Res. 52, 2818–2831 (2019).
Hanaoka, K. et al. A general fluorescence off/on strategy for fluorogenic probes: Steric repulsion-induced twisted intramolecular charge transfer (sr-TICT). Sci. Adv. 10, https://doi.org/10.1126/sciadv.adi8847 (2024).
Jiang, X. et al. 1,2,4,5-Tetrazine-tethered probes for fluorogenically imaging superoxide in live cells with ultrahigh specificity. Nat. Commun. 14, 1401 (2023).
Chi, W. et al. A unified fluorescence quenching mechanism of tetrazine-based fluorogenic dyes: energy transfer to a dark state. Mater. Chem. Front. 5, 7012–7021 (2021).
Shen, T., Zhang, W., Yadav, P., Sun, X. W. & Liu, X. The fluorescence quenching mechanism of tetrazine-functionalized fluorogenic labels with integrated π-conjugations: internal conversion to a dark state. Mater. Chem. Front. 7, 1082–1092 (2023).
Shen, T., Li, X. & Liu, X. Photoinduced electron transfer endows fluorogenicity in tetrazine-based near-infrared labels. Mater. Chem. Front. 8, 2135–2141 (2024).
Velayutham, M., Hemann, C. & Zweier, J. L. Removal of H2O2 and generation of superoxide radical: role of cytochrome c and NADH. Free Radic. Biol. Med. 51, 160–170 (2011).
Agarwal, P., Huckle, J., Newman, J. & Reid, D. L. Trends in small molecule drug properties: a developability molecule assessment perspective. Drug Discov. Today 27, 103366 (2022).
Hitchcock, S. A. Structural modifications that alter the P-glycoprotein efflux properties of compounds. J. Med. Chem. 55, 4877–4895 (2012).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Weng, M.-S., Chang, J.-H., Hung, W.-Y., Yang, Y.-C. & Chien, M.-H. The interplay of reactive oxygen species and the epidermal growth factor receptor in tumor progression and drug resistance. J. Exp. Clin. Cancer Res. 37, https://doi.org/10.1186/s13046-018-0728-0 (2018).
Ullrich, A. et al. Human epidermal growth factor receptor cDNA sequence and aberrant expression of the amplified gene in A431 epidermoid carcinoma cells. Nature 309, 418–425 (1984).
Zheng, F. et al. Redox toxicology of environmental chemicals causing oxidative stress. Redox Biol. 34, 101475 (2020).
Bus, J. S. & Gibson, J. E. Paraquat: model for oxidant-initiated toxicity. Environ. Health Perspect. 55, 37–46 (1984).
Perelman, A. et al. JC-1: alternative excitation wavelengths facilitate mitochondrial membrane potential cytometry. Cell Death Dis. 3, e430 (2012).
Oliveira, B. L., Guo, Z. & Bernardes, G. J. L. Inverse electron demand Diels-Alder reactions in chemical biology. Chem. Soc. Rev. 46, 4895–4950 (2017).
Thalhammer, F., Wallfahrer, U. & Sauer, J. Reaktivität einfacher offenkettiger und cyclischer Dienophile bei Diels–Alder-reaktionen mit inversem elektronenbedarf. Tetrahedron Lett. 31, 6851–6854 (1990).
Chen, W., Wang, D., Dai, C., Hamelberg, D. & Wang, B. Clicking 1, 2, 4, 5-tetrazine and cyclooctynes with tunable reaction rates. Chem. Commun. 48, 1736–1738 (2012).
Lang, K. et al. Genetic encoding of bicyclononynes and trans-cyclooctenes for site-specific protein labeling in vitro and in live mammalian cells via rapid fluorogenic Diels–Alder reactions. J. Am. Chem. Soc. 134, 10317–10320 (2012).
Kumar, G. S., Racioppi, S., Zurek, E. & Lin, Q. Superfast tetrazole–BCN cycloaddition reaction for bioorthogonal protein labeling on live cells. J. Am. Chem. Soc. 144, 57–62 (2021).
Burgos-Barragan, G. et al. Mammals divert endogenous genotoxic formaldehyde into one-carbon metabolism. Nature 548, 549–554 (2017).
Pham, V. N. et al. Formaldehyde regulates S-adenosylmethionine biosynthesis and one-carbon metabolism. Science 382, https://doi.org/10.1126/science.abp9201 (2023).
Hopkinson, R. J. & Schofield, C. J. Deciphering functions of intracellular formaldehyde: linking cancer and aldehyde metabolism. Biochemistry 57, 904–906 (2018).
Brewer, T. F. & Chang, C. J. An Aza-Cope reactivity-based fluorescent probe for imaging formaldehyde in living cells. J. Am. Chem. Soc. 137, 10886–10889 (2015).
Bruemmer, K. J. et al. Development of a general Aza-Cope reaction trigger applied to fluorescence imaging of formaldehyde in living cells. J. Am. Chem. Soc. 139, 5338–5350 (2017).
Roth, A., Li, H., Anorma, C. & Chan, J. A reaction-based fluorescent probe for imaging of formaldehyde in living cells. J. Am. Chem. Soc. 137, 10890–10893 (2015).
Bruemmer, K. J., Brewer, T. F. & Chang, C. J. Fluorescent probes for imaging formaldehyde in biological systems. Curr. Opin. Chem. Biol. 39, 17–23 (2017).
Brewer, T. F., Burgos-Barragan, G., Wit, N., Patel, K. J. & Chang, C. J. A 2-aza-Cope reactivity-based platform for ratiometric fluorescence imaging of formaldehyde in living cells. Chem. Sci. 8, 4073–4081 (2017).
Pan, Y. et al. Activity-based trapping for multiplex imaging illuminates the hidden role of endogenous formaldehyde in proinflammatory signaling. Cell Biomater. https://doi.org/10.1016/j.celbio.2025.100045 (2025).
Orecchioni, M., Ghosheh, Y., Pramod, A. B. & Ley, K. Macrophage polarization: different gene signatures in M1(LPS+) vs. classically and M2(LPS-) vs. alternatively activated macrophages. Front. Immunol. 10, 1084 (2019).
Acknowledgements
X. Li was supported by the National Program for Support of Top-notch Young Professionals (grant 2021). This study was supported by grants from the National Natural Science Foundation of China (22377106, 82471713), the Fundamental Research Funds for the Central Universities (226-2025-00106), and the National Key Research and Development Program of China (grant numbers 2022YFC2704600, 2022YFC2704601). We thank Ms. S.S. Liu from the Core Facilities, Zhejiang University School of Medicine, for technical support during imaging experiments.
Author information
Authors and Affiliations
Contributions
X. Li designed the project. L.Q. and X. Liu advised the project. C.W., X.J., and S.F. performed organic synthesis and characterization. C.W. performed photophysical measurements and analyzed the data. C.W. and T.Y. conducted confocal cell imaging experiments and analyzed the data. J.Y. validated the imaging results. T.S. advised probe design. X. Li and C.W. drafted the manuscript. All authors have reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wen, C., Jiang, Y., Shen, T. et al. Sensight enables quantitative multivariate engineering of high-performance chemical imaging tools. Nat Commun 17, 2061 (2026). https://doi.org/10.1038/s41467-026-68663-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68663-2