Abstract
Brain age prediction has been widely utilized to assess functional connectivity (FC) development, but conventional global brain age indices are limited in capturing spatial heterogeneity across the cortex. This study introduces a regional brain development index to characterize fine-grained FC maturation across cortical regions. We examined its spatial variability and stratified individuals into subtypes with distinct region-wise FC developmental patterns. Using data from the Philadelphia Neurodevelopmental Cohort (ages 8-23 years), we identified three distinct subtypes and found that individuals with advanced FC developmental pattern aligning with the sensorimotor-association axis exhibited superior cognitive performance. Robustness was confirmed through replication in the Human Connectome Project Development cohort. Further analyses revealed associations between FC development and gene expression linked to neural differentiation, synaptogenesis, and myelination. These findings suggest that spatial heterogeneity in FC development reflects cortical microstructure and hierarchical organization, underscoring its critical role in neurocognitive maturation during youth.
Similar content being viewed by others
Introduction
The development of functional networks is a dynamic and complex process closely linked to ongoing maturation of the brain’s functional organization, which underpins the progressive enhancement of cognitive, emotional, and social capacities throughout childhood and adolescence1,2. Research has established the critical role of cortical hierarchy in brain function, and accumulating evidence has demonstrated that the cortical development follows spatial and temporal patterns aligned with this hierarchy from perspectives of macroscale neurobiological properties1,3 and underlying microscale mechanisms in developmental plasticity, such as cortical excitation, inhibition, and glial cell types1,4,5. Recent studies further show that the spatial-temporal patterns of functional network development in youth systematically align with cortical hierarchy organization, particularly along the sensorimotor-association (S-A) axis, with developmental increases in functional connectivity (FC) in sensorimotor cortices and decreases in association cortices6. However, how these FC development patterns vary across individuals remains unclear. Neurocognition undergoes a prolonged development trajectory parallel to the maturation of the brain’s functional organization, with individual differences in youth linked to important outcomes such as academic performance and quality of life2, while deficits in neurocognition are associated with vulnerability to mental illness. Therefore, advancing our understanding of FC development at the individual level may provide valuable insights into typical development processes and deviations from these typical trajectories, with implications for cognitive and psychological functioning as well as mental illness.
Brain age has increasingly been utilized to evaluate brain status and track brain changes at the individual level, leveraging neuroimaging data across various modalities7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22. The difference between brain age and chronological age, known as the brain age gap (BAG), provides a quantitative and interpretable metric for characterizing brain changes associated with development, aging, and brain disorders. BAG measures derived from functional connectome have revealed that individuals with accelerated brain development exhibit enhanced FC in specific functional brain networks, including the default mode network (DMN), during childhood and adolescence12. Existing studies have established a linkage between brain age derived from structural imaging data and cognition, demonstrating that individuals with advanced brain development exhibit superior cognitive processing speed compared to those with delayed development8. While studies have identified associations between FC and neurocognition in youth using predictive modeling23,24,25,26, it remains unclear whether and how brain age estimates derived from the functional connectome are associated with neurocognition in youth.
Most existing studies estimate brain age from a global perspective by mapping whole-brain, high-dimensional FC profiles to a single unitary index, which inherently overlooks the spatial variability of FC changes across brain regions. To address this limitation, several studies have proposed fine-grained modeling at the regional level10,13,15,17,18,20,27. Distinct and heritable patterns of brain aging, derived from structural imaging features, have been linked to brain disorders, such as schizophrenia, multiple sclerosis, and dementia10. Additionally, heterogenous aging patterns across brain regions in schizophrenia have been revealed using multimodal neuroimaging data20. Advanced brain aging in specific brain regions has also been shown to mediate cognitive outcomes in cerebral small vessel disease, highlighting the potential to capture heterogeneous brain changes beyond a unitary measure of global brain age15. Given that variations in FC profiles, functional topography, and structural-functional coupling are associated with brain development and neurocognition2,6,9,11,12,23,24,26,27,28,29, exploring regional patterns of brain development measures derived from FC and their associations with neurocognition could provide valuable insights into the development of brain functional organization and neurocognitive processes.
In this study, we utilized a large sample (total n = 1149) of youths aged 5–23 years from the Philadelphia Neurodevelopmental Cohort (PNC)30 and the Human Connectome Project Development cohort (HCP-D)31 to investigate region-wise FC development using functional MR imaging (fMRI) data. For each cortical region, we developed a region-specific age prediction model based on its regional FC profile. Each model provides an individualized brain age measure, and the difference between this measure and chronological age, referred to as regional brain development (RBD) index, reflects region-wise FC development. We hypothesized that the RBD index would exhibit spatial variability and inter-subject heterogeneity, linked to developmental differences in neurocognition. To explore the spatial variability of regional FC development and its associations with neurocognition, we stratified individuals into subtypes characterized by distinct FC development patterns using a cluster analysis based on individual subjects’ whole-brain RBD indices. Subsequently, we investigated differences in cognitive measures across these subtypes. Our analysis revealed that individuals with advanced FC development parallel to the hierarchical cortical organization defined by the S-A axis exhibited superior cognitive profiles. To investigate the biological mechanism underlying the spatial variability of FC development, we examined the relationship between regional FC development patterns and gene expression. We predicted that regional FC development patterns aligning with the cortical hierarchy organization would show enrichment in the expression of genes associated with neuron development.
Results
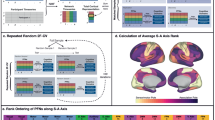
We used fMRI data from the PNC (n = 693; ages 8–23) to examine the spatial variability and inter-individual heterogeneity in regional FC development. The RBD index, defined as brain age minus chronological age and corrected for bias and covariates, was used as a surrogate measure to assess FC development across cortical regions. Specifically, for each cortical region, a dedicated brain age prediction model was built on its FC profile, which was defined as its FC measures to all the other cortical regions, with Pearson’s correlation coefficient serving as the FC measure. Five-fold cross-validation was implemented to predict brain age across all individuals and cortical regions. An overview of the analysis process is illustrated in Fig. 1. Using the RBD index, we began by examining spatial variability in FC development and its inter-individual heterogeneity. Next, we applied clustering to identify covarying patterns of FC development across cortical regions, stratifying individuals into subtypes characterized by distinct regional FC development profiles. We then compared the spatial patterns across subtypes and assessed their alignment with established principles of cortical hierarchy in brain development. To further characterize these subtypes, we analyzed their neurocognitive profiles using cognitive measures and meta-analytic functional decoding. We subsequently explored the biological underpinnings of these FC development patterns by examining their associations with gene expression. Finally, we evaluated the generalizability and robustness of the identified subtypes of FC development using data from an independent cohort.
A The regional brain development (RBD) index was derived from functional connectivity (FC). Region-specific brain age prediction models were built upon regional FC profiles comprising FC measures of each region with all other cortical regions, to assess region-wise FC development. These prediction models were trained using a five-fold cross-validation approach to estimate brain age for each individual and cortical region. Spatial variability in prediction performance and inter-individual heterogeneity in the RBD index were then analyzed to gain insights into region-wise FC development. B Spatially covarying patterns of the RBD index were identified through clustering, which stratified individuals into subtypes with distinct FC development profiles. These subtypes were further characterized by examining their associations with cortical hierarchy, neurocognition measures, and gene expression.
Spatial variation and inter-individual heterogeneity in functional connectivity development
To characterize the spatial variation of FC development across cortical regions, we analyzed differences in their capacity for age prediction. Two metrics, accuracy (r) and mean absolute error (MAE), were used to quantify each region’s predictive capacity, with accuracy (r) computed as the Pearson’s correlation coefficient between the predicted age (prior to bias correction) and chronological age. Cortical regions exhibited varying capacity for age prediction (Fig. 2A), with certain functional networks showing higher predictive capacity, as indicated by higher r and lower MAE values. These networks included the somatomotor network, ventral attention network (VAN), and frontoparietal network (Fig. 2B). However, heterogeneity was evident within individual functional networks: for example, the medial prefrontal aspect of the DMN displayed higher predictive capacity compared with lateral elements of the DMN.
A Regional capacity for brain age prediction, quantified by accuracy (r) and mean absolute error (MAE), reflects spatially varying FC development. B Distribution of region-wise age prediction capacity across brain functional networks based on the Yeo 7-network atlas72. The center line of the box plot indicates the median value across cortical regions within a functional network. The box edges represent the interquartile range (IQR, 25th–75th percentiles), whiskers denote the data range within 1.5 \(\times\)IQR, and the “o” markers denote outliers that are more than 1.5 \(\times\)IQR away from the top or bottom of the box. The number of cortical regions in frontoparietal, default mode, dorsal attention, limbic, ventral attention, somatomotor, and visual networks are 52, 91, 46, 26, 47, 77, and 61, respectively. C Maps of RBD indices are shown for six randomly selected individuals (IDs 1-6) at three developmental time points: ages 9, 13, and 17 (two individuals per time point), illustrating brain regions with advanced (positive, red) or delayed (negative, blue) FC development. Source data are provided as a Source Data file.
Building on the observed variation in FC development across cortical regions, we next investigated inter-individual differences in the pace of regional FC development. To assess this, we quantified the brain development for each cortical region and individual using the RBD index, defined as brain age minus chronological age and corrected for bias and covariates32, with positive/negative values indicating advanced/delayed development. Individuals of the same chronological age (9, 13, or 17 years) showed variability in region-wise brain development (Fig. 2C), highlighting inter-individual variability in the pace of regional FC development. Additionally, distinct spatial covariation patterns in RBD were evident at the individual level. For instance, the 13-year-old individual (Fig. 2C, bottom-middle) exhibited consistently advanced development in frontoparietal and DMN regions alongside delayed development in sensorimotor regions, whereas the 9-year-old individual (Fig. 2C, top left) showed a largely opposite pattern.
Subtypes of functional connectivity development and their links to neurocognition
To investigate the observed inter-individual heterogeneity and potential spatial covariation in regional FC development, we identified covarying patterns of FC development and stratified individuals into subgroups based on their whole-brain RBD index maps. Using the K-means clustering algorithm, individuals were grouped into three subtypes, with each subtype characterized by a distinct FC development pattern (P1, P2, and P3) represented by the centroids of their RBD index maps (Fig. 3A). Specifically, P1 showed a global trend of delayed FC development across cortical regions; P2 exhibited advanced FC development localized to the association cortex, prominently within DMN regions; P3 displayed widespread advanced FC development, particularly pronounced in sensorimotor regions.
A Centroid RBD index maps of the three subgroups identified in the PNC cohort. B–D Alignment of the development patterns with the sensorimotor-association (S-A) axis rank across cortical regions, with each dot representing one cortical region. E Region-wise comparison of RBD indices between P2 and P3, with statistical significance determined using the Wilcoxon rank sum test (p < 0.05, one-sided, FDR-corrected). F Statistical comparisons of cognition scores across three subgroups, highlighting differences in cognitive performance. The center line of the box plot indicates the median value across individuals within a subgroup. The box edges represent the interquartile range (IQR, 25th–75th percentiles), whiskers denote the data range within 1.5 \(\times\)IQR, and the “o” markers denote outliers that are more than 1.5 \(\times\)IQR away from the top or bottom of the box. Asterisks indicate statistically significant differences (p < 0.05, Wilcoxon rank sum test, two-sided). The number of individuals in subgroups P1, P2, and P3 are 304, 141, and 248, respectively. G Significant correlation between the development pattern of P2 and the principal functional gradient (p < 0.0001, Spin-test). H Significant negative correlation between the development pattern of P2 and the cortical myelin content (p < 0.0001, Spin-test). I Meta-analytic functional decoding result of the development pattern of P2 based on the NeuroQuery, where larger font size indicates stronger correlation with specific functional term. Source data are provided as a Source Data file.
To further characterize the spatial variation of these distinctive development patterns, we examined their alignment with the S-A axis, which represents a unified cortical organization across diverse neurobiological properties and has been demonstrated to be significantly associated with FC development6. While the development pattern of P1 did not significantly align with the S-A axis (Fig. 3B, Spin-test, r = 0.186, p = 0.188), those of P2 and P3 showed significant correlations, with P2 showing a strong positive correlation (Fig. 3C, Spin-test, r = 0.580, p < 0.0001) and P3 showing a strong negative correlation (Fig. 3D, Spin-test, r = −0.511, p = 0.0001). Notably, the cortical regions with predominantly advanced development in the P2 were located at the high-ranking association end of the S-A axis, while those in P3 were located at the low-ranking sensorimotor end. Given the overall advanced FC development of P2 and P3, we then examined their differences in FC development across regions. Specifically, P2 demonstrated significantly more advanced FC development in DMN regions compared with P3, whereas P3 demonstrated significantly greater advanced development in sensorimotor regions relative to P2 (Fig. 3E).
We next analyzed the cognitive performance of individuals across subtypes in three cognitive domains: executive function, social cognition, and memory, to investigate their behavioral implications. Individuals in P2 demonstrated significantly higher cognitive performance than those in P1 and P3 (Fig. 3F), with respect to executive function (Wilcoxon rank sum test, z = 3.73 and p = 0.0002 for P2 vs. P1, z = 2.59 and p = 0.010 for P2 vs. P3), social cognition (Wilcoxon rank sum test, z = 3.80 and p = 0.0001 for P2 vs. P1, z = 2.36 and p = 0.019 for P2 vs. P3), and memory (Wilcoxon rank sum test, z = 2.17 and p = 0.030 for P2 vs. P1, z = 2.24 and p = 0.025 for P2 vs. P3). Interestingly, no significant differences were observed between P1 and P3, despite globally advanced FC development of individuals in P3 (Fig. S1B). There were no significant differences across subgroups regarding chronological age and sex. These findings highlight the critical role of spatial heterogeneity in FC development during youth, particularly within regions that support neurocognitive functions.
To better understand the relationship between distinctive FC development patterns and properties of the brain connectome, we also analyzed their alignment with the principal functional gradient and cortical myelin content. The P2 showed a strong positive correlation with the principal functional gradient (Fig. 3G, Spin-test, r = 0.604, p < 0.0001), consistent with prior findings that hierarchical functional organization supports neurocognition development. Moreover, the P2 exhibited a significantly negative correlation with cortical myelin content (Fig. 3H, Spin-test, r = −0.512, p < 0.0001), suggesting a relationship with the ongoing process of myelination during brain maturation. On the other hand, the P3 exhibited a reversed alignment pattern, showing a negative correlation with the principal functional gradient and a positive correlation with the cortical myelin content (Fig. S2). These findings underscore the critical interplay between functional organization and structural maturation of brain connectome in shaping neurocognitive outcomes.
In addition, we utilized a meta-analytic functional decoding technique33 based on the NeuroQuery database34 to interpret the functional significance of the identified FC development patterns. The decoding results also revealed significant associations between the P2 and neurocognition-related terms, such as cognitive, decision, and context (Fig. 3I).
We next conducted a series of sensitivity analyses to investigate the robustness of the identified FC development patterns. The subtypes of FC development remained robust in terms of spatial patterns, cognitive profiles, and individual-level subtype assignments, when controlling for additional covariates including socioeconomic status (SES) and parental education (Fig. S5), when using Gaussian Mixture Models (GMM) based clustering for the subtyping (Fig. S6), when using the Schaefer-300 atlas and Schaefer-200 atlas for cortical parcellation (Fig. S7), and when using the Fisher’s r-to-z transformed FC measures to derive the RBD index (Fig. S8).
Associations between regional FC development and gene expression
To investigate the biological mechanism underlying the regional FC development, we analyzed the relationship between the FC development patterns and gene expression data from the AHBA35. We first examined the alignment of FC development patterns with the first principal component (PC1) of gene expression36. The P2 exhibited significant negative correlation (Fig. 4A, Spin-test, r = −0.440, p = 0.0017), while no significant correlations were observed for P1 and P3 (Fig. S3). We then used spatial correlation analysis to examine the association between the P2 pattern and the expression of 4112 genes in the cortex. This analysis identified 735 positively correlated genes and 783 negatively correlated genes (q < 0.05, FDR corrected, Spin-test). Among them, the neuronal differentiation 6 (NEUROD6) gene was the most positively correlated, while the Mitochondrial Ribosomal Protein L48 (MRPL48) gene was the most negatively correlated (Fig. 4B). Using Specific Expression Analysis (SEA)37, we found that the set of positively correlated genes was enriched for neurodevelopmental genes with cortical expression spanning stages from early childhood to young adulthood (Fig. 4C), suggesting that these genes may play a role in supporting the regional FC development in youth.
A Alignment of P2 with gene expression PC1 (p = 0.0017, Spin-test). B The top 100 genes with the highest absolute correlations with P2 are displayed; warm and cool colors indicate positive and negative correlations, respectively, and font size is proportional to the magnitude of the absolute correlation. C Neurodevelopmental gene enrichment analysis revealed enrichment for the highly correlated neurodevelopmental genes expressed in the cortex during developmental stages ranging from early childhood to young adulthood. The y-axis represents the statistical significance of the enrichment (FDR-corrected q-value). D Gene Ontology (GO) enrichment analysis for cellular components identified significant enrichment in terms such as “membrane”, “neuron projection”, and “synapse”. Each dot represents an enriched GO term, with size proportional to the number of associated genes and transparency reflecting the significance of enrichment (FDR-corrected q-value, y-axis). E Adult cell-type-specific enrichment analysis demonstrated that the correlated genes were enriched for markers of various cortical cell types in adults, including astrocytes, oligodendrocyte precursor cells (OPCs), and multiple inhibitory and excitatory cortical neurons. Source data are provided as a Source Data file.
We further performed Gene Ontology (GO) enrichment analysis on the set of correlated genes using the online tool of GOrilla38, which revealed that the significantly correlated genes were primarily enriched in GO terms associated with “membrane”, “neuron projection”, and “synapse” (q < 0.05, FDR corrected, Fig. 4D). Given that regional differences in cortical gene expression may be influenced by differences in cellular composition of the cortex, we conducted a cell-type-specific enrichment analysis39 to examine the enrichment of correlated genes in adult cortical cell types. This analysis revealed that the correlated genes were enriched across multiple adult cell types, including astrocytes, oligodendrocyte precursor cells, inhibitory and excitatory neurons (Fig. 4E).
The FC development patterns are generalizable to the HCP-Development cohort
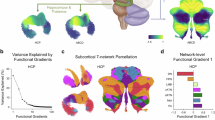
To assess the robustness of our findings, we investigated regional FC development using data from the HCP-D cohort (n = 456; ages 5–22) and evaluated whether the identified FC development patterns could stratify the HCP-D individuals into distinct subgroups. Five-fold cross-validation was utilized to predict brain age for all cortical regions and individuals in the HCP-D cohort, as in the previous analyses of the PNC cohort. The spatial variation of FC development across cortical regions exhibited a pattern consistent with that observed in the PNC cohort (Spin-test, r = 0.467, p < 0.0001), with regions showing higher predictive accuracy primarily located in the somatomotor network, VAN, and the anterior part of the DMN (Fig. 5A, B). Next, HCP-D individuals were stratified into three subgroups based on the FC development patterns identified in the PNC cohort. Specifically, each individual’s RBD index map was compared with the PNC cohort’s centroids of RBD index maps (P1, P2, and P3), and the individual was assigned to the closest matching subgroup (Fig. 5C). We then analyzed cognitive performance of individuals across subgroups using a battery of cognitive assessments relevant to executive function, memory, and general cognition. Overall, individuals in subgroup P2 demonstrated the highest cognitive performance, particularly on the Flanker Test (Wilcoxon rank sum test, z = 2.90, p = 0.004) and the List Sort Task (Wilcoxon rank sum test, z = 2.15, p = 0.032) compared to subgroup P1 (Fig. 5D). No significant differences were observed between subgroups P1 and P3 with respect to these two assessments. Individuals in subgroup P2 also demonstrated the highest cognitive performance in terms of other cognitive domains, including fluid cognition, crystallized cognition, and total cognition (Fig. S9). These findings demonstrate that the three FC development patterns identified in the PNC cohort are robust and generalizable to the HCP-D cohort. Moreover, the results reinforce that the spatial heterogeneity in FC development across individuals is linked to neurocognitive performance. Replication analysis applying the same clustering procedure within the HCP-D cohort revealed consistent FC development patterns and subtyping results (Fig. S10), further supporting the reproducibility of the findings.
A Regional prediction accuracy for brain age, measured by correlation coefficient (r). B Spatial correspondence between prediction accuracy in the PNC and HCP-D cohorts, revealing consistent pattens of regional FC development (p < 0.0001, Spin-test). C Assignment of HCP-D individuals to the FC development subtypes defined in the PNC cohort. For each HCP-D individual, the distances between their RBD index map and the centroid RBD index maps of the PNC-defined subtypes were computed, and the individual was assigned to the subtype with the smallest distance. For example, individuals i and i + 1 were assigned to subtypes P2 and P1, respectively. D Statistical comparisons of cognition scores across the three subtypes in HCP-D, highlighting subtype-related differences in cognitive profiles. The center line of the box plot indicates the median value across individuals within a subgroup. The box edges represent the interquartile range (IQR, 25th–75th percentiles), whiskers denote the data range within 1.5 \(\times\)IQR, and the “o” markers denote outliers that are more than 1.5 \(\times\)IQR away from the top or bottom of the box. Asterisks indicate statistically significant differences (p < 0.05, Wilcoxon rank sum test, two-sided). The number of individuals in subgroups P1, P2, and P3 are 207, 84, and 164, respectively. CardSort, Dimensional Change Card Sort Task; Flanker, Flanker Test; ListSort, List Sort Working Memory Task; Reading, Oral Reading Test; Pattern, Pattern Comparison Processing Speed Task; PicSeq, Picture Sequence Memory Task; PicVocab, Picture Vocabulary Test. Source data are provided as a Source Data file.
Discussion
In this study, we investigated the spatial heterogeneity of brain FC development at the individual level by leveraging machine learning-based brain age prediction and fMRI data of large cohorts of youths. Using a region-specific brain age prediction approach, we estimated an RBD index for each brain region and individual, uncovering significant variability in FC development across cortical regions, which reflects the spatial heterogeneity of brain FC development at the individual level. By clustering these individualized RBD index maps, we identified three distinct development patterns and stratified individuals into corresponding subgroups. Notably, individuals with a FC development pattern positively correlated with the hierarchical cortical organization exhibited superior cognitive performance, a relationship that could not be captured using global BAG measures. These findings underscore the utility of RBD indices in providing fine-grained insights into the complex relationships between brain development and behavior phenotypes, offering a more nuanced understanding than global measures can achieve.
Previous research has investigated brain development in children and adolescents using imaging-based brain age estimation. Multimodal brain development indices have revealed that elevated brain age is associated with significantly lower gray matter volume, higher white matter volume in the prefrontal cortex, and enhanced FC in networks such as the DMN12. Additionally, brain age estimates have been linked to various cognitive functions8 and psychiatric disorders, including schizophrenia12, autism14, and attention-deficit/hyperactivity disorder16. However, most of these studies focus on brain development from a global perspective by estimating BAG, a single univariate index, for each individual. This approach overlooks the spatial heterogeneity of brain development, which reflects the possibility that brain regions develop at varying rates within the same individual. While cluster-wise brain development indices derived from structural imaging data are promising to capture brain changes that are not detectable by BAG17, the region-wise heterogeneity of FC development within individuals remains largely unexplored. This study systematically characterizes the individualized RBD index of FC in children and adolescents, providing new insights into the spatial heterogeneity of FC development. Our findings reveal that this heterogeneity reflects differences in developmental pacing across brain regions and is significantly linked to individual differences in cognitive performance.
Normative modeling40,41 is increasingly used neuroimaging studies to derive deviation scores from normative trajectories for the investigation of inter-individual heterogeneity. However, it is typically based on element-wise regression, which poses challenges for modeling FC due to its high dimensionality. Prior studies applying normative modeling to FC have either used summary measures such as FC strength42 or surrogate scores derived from dimension reduction techniques43, which may not fully capture the richness of FC profiles. Although multivariate normative modeling methods exist, their application to FC data remains relatively limited. In contrast, the predictive modeling of brain age provides an interpretable, composite score that reflects the development status of each cortical region based on its full FC profiles, without relying on FC strength or dimension reduction. Moreover, the calibration of predicted brain age against chronological age is an analogy to normative modeling, enabling the estimation of individual deviations in brain development.
We observed spatial heterogeneity in FC development at both the population and individual levels. At the population level, different brain regions exhibited varying capacities for age prediction, with regions within the somatomotor network and VAN showing higher prediction accuracy. At the individual level, brain regions displayed heterogenous advanced or delayed brain development across cortical regions, independent of an individual’s chronological age. The RBD indices facilitated a fine-grained, individualized characterization of the FC development, enabling the stratification of individuals into subgroups based on distinct FC development patterns. While it might seem intuitive that individuals with advanced FC development would outperform those with overall delayed development in cognitive tasks, our findings highlight the critical role of specific brain regions in this relationship: individuals with advanced development in the DMN and VAN regions demonstrated superior cognitive performance, whereas those with advanced development in the somatomotor regions did not, despite showing greater overall advancement in FC development. These results underscore the importance of accounting for spatial heterogeneity in brain age-related studies, providing compelling evidence that region-specific patterns of development significantly influence cognitive outcomes.
The subtypes of FC development identified by the global BAG exhibited distinct cognitive profiles compared to those derived based on the RBD index maps. Notably, its subgroup with the best cognitive profile corresponded to the subgroup in-between the delayed and advanced developmental groups (Fig. S1A). Similarly, a recent study found that the global BAG had limited utility in capturing fluid cognition in older individuals21. Our findings further reveal that the distributions of global BAG measures were largely overlapping across subgroups identified based on the RBD indices (Fig. S1B), indicating that the RBD indices provide additional insights into brain development beyond what is captured by global brain development index alone. These results highlight the potential of incorporating the spatial heterogeneity in brain age to better capture the associations between brain age and neurocognition.
The RBD pattern of the subgroup with superior cognitive performance aligns significantly with the brain’s hierarchical organization. Advanced development predominantly occurs in regions with high hierarchy ranks, such as DMN and VAN regions, and delayed development occurs in regions with low hierarchy ranks, such as sensorimotor regions. This finding is consistent with previous research showing that the development of hierarchical functional system is supportive to neurocognitive development1,2,44,45. It also corroborates findings that individual FC patterns in the DMN and VAN are linked to cognitive functions in youth23,24,25. Furthermore, the RBD pattern shows a significant correlation with cortical myelination, suggesting that changes in cortical microstructure may influence FC development by enhancing neural signal transmission and regulating plasticity46. These associations converge to support the idea that advanced FC development parallel to the hierarchical axis of cortical organization underpins better cognition in youth.
We further investigated the biological basis of the advanced FC development pattern through gene expression data analysis, identifying a set of co-located genes, including NEUROD6 and THRB, known to play critical roles in brain development and neurocognition. These genes regulate and influence key processes such as neural cell migration and differentiation, synaptogenesis, and myelination47,48,49,50,51. Additionally, these co-located genes were enriched for those expressed in cortical regions during mid-to-late childhood and adolescence, consistent with known developmental trajectories37. GO enrichment analysis revealed that these genes were associated with multiple cellular components, including membrane, neuron projection, and synapse. Cell-type-specific enrichment analysis further demonstrated correlations between the advanced FC development pattern and genes related to astrocytes as well as multiple excitatory and inhibitory neuron types. These findings suggest that the co-located genes play essential roles in supporting the inter-neuron connectivity and brain circuit development52. It is speculated that the identified FC development pattern reflects underlying changes in gene transcription and its influence on brain cytoarchitecture.
We also observed that the spatial heterogeneity of FC development was both consistent and replicable across different datasets. Predictive analysis on the HCP-D cohort demonstrated that the region-wise predictive capacity closely aligned with that observed in the PNC cohort. Additionally, the FC development patterns identified in the PNC cohort were generalizable to the HCP-D cohort. Stratification results from the HCP-D cohort demonstrated that individuals in the subgroup P2 exhibited superior cognitive performance. Notably, this was observed despite differences in the cognition domains assessed in the PNC and HCP-D cohorts, indicating that the identified FC development patterns were robust and reliably captured underlying brain development processes.
Although the RBD index has demonstrated potential for capturing fine-grained and reproducible patterns of brain development, several limitations should be acknowledged. First, though brain age has been validated to be sensitive to clinical outcomes and other biomarkers53,54,55, concerns have been raised regarding the inflated prediction accuracy due to statistical bias-correction and its interpretation56. While brain age prediction accuracy is not the focus of this study, alternative measures merit further investigation. Second, the use of a linear modeling approach for region-wise brain age prediction may constrain its ability to characterize non-linear developmental effects in certain brain regions. Developing a principled framework to select the optimal predictive model (linear or non-linear) for each region could better characterize the complex developmental changes in FC. Third, the associations between FC development patterns and gene expression data are correlational and rely on the AHBA, which represents transcriptomic data from adult brains. Replication studies using transcriptome data from youth populations are needed to confirm and extend these findings. Moreover, future analysis using causal inference approaches may help elucidate mechanistic links between FC development, molecular factor, and cognition. Fourth, while sex was treated as a covariate and controlled in this study, investigating sex differences in the spatial heterogeneity of brain FC development offers a compelling avenue for future research. Fifth, the associations between FC development and cognition were investigated using a relatively narrow cognition battery. Future studies incorporating a broader spectrum of behavioral and psychological measures are needed to comprehensively capture the multidimensional relevance of FC development to youth behavior and cognition. It would also be beneficial to conduct further validation using external datasets with harmonized behavioral and cognitive measures. In addition, the current analysis was based on cross-sectional data to characterize the development differences across individuals. Future research incorporating longitudinal data is needed to investigate intra-individual changes in FC and cognition throughout development. Lastly, while this study focuses on FC development in youth, a promising direction for future research involves integrating multi-modal data, including brain imaging, behavioral measures, and other relevant measures, to investigate the heterogeneity of brain development from a multi-dimensional perspective. Extending the imaging analysis to include subcortical regions is also important, as it would enable investigation of the subcortical-cortical FC development and its relationship to neurocognition. Such approaches could offer a more comprehensive understanding of the complex mechanisms underlying brain development and maturation and provide valuable insights into the convergence and divergence of developmental heterogeneity revealed across different data modalities.
In summary, we investigated the spatially fine-grained development of FC in youth using the RBD index, derived by region-specific brain age predictive modeling. Our findings reveal replicable spatial heterogeneity in FC development at the individual level and establish links between the spatial patterns of region-wise brain development and neurocognition. Additionally, these results suggest region-wise brain development patterns align with the brain’s hierarchical organization and are potentially associated with gene expression. These findings underscore the critical role of spatial heterogeneity in FC development for supporting neurocognition development in youth.
Methods
Participants and data processing
Data were drawn from two datasets: the PNC (n = 693) and the Human Connectome Project Development (HCP-D; n = 456).
The PNC30 is a community sample of children and adolescents from the greater Philadelphia area, collected for studying brain development, including 1601 participants with neuroimaging data. For this study, demographic, cognitive assessment, and neuroimaging data from 693 participants (301 males and 392 females) aged 8–23 years were included after applying the inclusion criteria as previously described11. Informed consent was obtained from all participants or their parents/guardians, and minors provided assent. All study procedures were approved by the institutional review boards of both the University of Pennsylvania and the Children’s Hospital of Philadelphia. One resting-state and two task-based (n-back and emotion identification) fMRI scans for each participant were used in this study. The neuroimaging preprocessing followed previously described procedures and methods11. Cognitive assessment was conducted using the Penn computerized neurocognitive battery (Penn CNB), and three factor-analysis based composite scores57, including executive and complex cognition, social cognition, and episodic memory, were included for analysis in this study.
The HCP-D31 includes 652 participants recruited across four sites in the U.S. to study healthy brain development in children and adolescents. A sample of 456 participants (214 males and 242 females) aged 5–22 years with complete demographic, cognitive assessment, and neuroimaging data were included in this study after applying the inclusion criteria as previously described6. All study procedures were approved by a central Institutional Review Board at Washington University in St. Louis. Resting-state and task-based (carit, emotion, and guessing tasks) fMRI scans were used for all participants. The neuroimaging acquisition and preprocessing procedures have been previously described6. Participants completed a battery of cognitive assessments, including segments of the NIH toolbox and others31. Cognitive scores from the Dimensional Change Card Sort Task, the Flanker Test, the List Sort Working Memory Task, the Oral Reading Test, the Pattern Comparison Processing Speed Task, the Picture Sequence Memory Task, and the Picture Vocabulary Test were included in this study. Composite scores, including fluid cognition, crystalized cognition, and total cognition from the HCP-D data release31 were also included. Demographic information and variables of interest for both datasets may be found in Tables 1 and 2.
For both samples, preprocessed fMRI time series were parcellated into 400 cortical regions using the Schaefer-400 atlas58, and time series from resting-state and task fMRI scans were concatenated to compute FC measures for the analysis.
Brain development index
For region-specific brain age prediction, a dedicated linear ridge regression model to predict chronological age was built for each cortical region on its FC profile that was defined as its FC values to all the other cortical regions, with each value measured as a Pearson’s correlation coefficient between the two regions. Five-fold cross-validation was utilized to obtain predicted brain age for all individuals and cortical regions.
For global brain age prediction, the whole-brain functional connectome (upper triangle entries of the FC matrix, containing FC values between all pairs of cortical regions) was used as features, and a linear ridge regression model was trained for the prediction of chronological age.
Both the region-specific and global prediction models were built and evaluated with the same five-fold cross-validation, with identical training and evaluation data splits. For each prediction model, the regularization parameter in ridge regression was optimized in the range [\({2}^{-10}\), \({2}^{-9}\), …, \({2}^{4}\), \({2}^{5}\)] using nested cross-validation within the training sample. The prediction models were implemented using the scikit-learn library (v1.4.1) in Python (v3.9.18).
Given the predicted brain age for each cortical region, the difference between this measure and chronological age was referred as the RBD index, reflecting region-specific FC development. To alleviate the known systematic bias in the brain age prediction due to the “regression to the mean” effect, the RBD index was adjusted by a linear bias correction procedure32. In addition, covariates including sex, head motion, and site information (for the HCP-D sample) were regressed out from the RBD index for all subsequent analysis. The same procedure was applied to obtain global BAG measures based on the predicted global brain age.
Subtyping FC development
Given the RBD index maps for all participants of the PNC cohort, the K-means clustering algorithm was utilized to identify subtypes of FC development and stratify the participants into different subgroups. Specifically, the clustering was performed with whole-brain RBD indices as features and cosine distance as the distance metric, as implemented in MATLAB (R2021a).
The number of clusters was set to 3, determined based on both reproducibility and within-cluster point-to-centroid distances of the clustering results. Specifically, all participants were randomly split into two halves, and clustering was performed on each half separately with the number of clusters \(k\) set from 2 to 7. This procedure was repeated for 100 times. The clustering results for each half was denoted as \({C}_{k,1}\) and \({C}_{k,2}\). For each \(k\), the clustering centroids of the first half were then applied to samples in the second half to obtain the clustering results \({C}_{k,1\to 2}\), and vice versa to obtain \({C}_{k,2\to 1}\). The average of \({RandIndex}({C}_{k,1},{C}_{k,2\to 1})\) and \({RandIndex}({C}_{k,2},{C}_{k,1\to 2})\) was adopted as the reproducibility measure for clustering results with \(k\) clusters, where \({RandIndex}\)(\(C\),\({C}^{{\prime} }\)) refers to the Rand index for two sets of clustering results \(C\) and \({C}^{{\prime} }\). Additionally, to evaluate the data-fitting performance of the clustering results, the distance of each data point to its corresponding cluster centroid was calculated and averaged across all data points. The optimal number of clusters was determined by reaching a balance between reproducibility and data-fitting performance, where the two measures interacted as shown in Fig. S4A.
In addition to reproducibility based and data-fitting based measures (Fig. S4A), we further justified the choice of number of clusters using the NbClust package59 and according to the data distribution characteristics. Particularly, NbClust uses a wide variety of indices to determine the optimal number of clusters using Hierarchical clustering or K-means methods. The most frequently selected cluster numbers across clustering methods were 2 and 3 (Fig. S4B), suggesting them as optimal choices for our dataset. To further verify this finding, we examined the distribution of individual RBD patterns in a 2D embedding space using t-SNE projection (Fig. S4C). It was observed that clusters P1 and P3 in the 3-cluster solution aligned closely with clusters P1 and P2 in the 2-cluster solution, while cluster P2 in the 3-cluster solution captured fine-grained inter-individual variability between P1 and P3. Based on all these quantitative and qualitative assessments, the number of clusters was set to 3 in our analysis.
The HCP-D individuals were stratified into distinctive subgroups based on the FC development patterns identified in the PNC cohort for generalization analyses. Specifically, each individual’s RBD index map was compared to the PNC cohort’s centroid maps, and the individual was assigned to the subgroup with the closest match. For instance, individuals i and i + 1 were assigned to subgroups P2 and P1, respectively, as their RBD index maps showed the highest similarity to the corresponding centroids (Fig. 5C). To further assess the robustness of the identified FC development patterns, the clustering procedure was also replicated within the HCP-D cohort.
Meta-analytic functional decoding of FC development pattern
The meta-analytic decoding tool33 was used to link the FC development pattern to functional/behavioral profiles. Specifically, the functional terms and term-based meta-analytic maps from the NeuroQuery database34 was utilized with the correlation decoder33 for the functional decoding. The functional decoding results were illustrated by a word cloud, with larger font size indicating higher functional relevance.
Gene enrichment analysis
Gene expression data was obtained from the Allen Human Brain Atlas (AHBA)35 and processed with the abagen toolbox60 and the Schaefer-400 atlas in the MNI space to obtain regional gene expression. Since not all genes in the AHBA dataset are assumed to be expressed in the brain, only brain-expressed genes were retained for analysis, following an established procedure61. Additionally, the analysis focused on the left hemisphere with 200 brain regions to minimize sample variability across brain regions, as the gene expression data were sampled from both hemispheres of only two of the six donors62. A total of 4112 genes were included in our analysis.
The spatial alignment of regional FC development patterns and the expression patterns of each gene was quantified using Peason’s correlation coefficient, and its significance was tested using the spin test with 1,0000 permutations63,64. The set of significantly correlated genes was included in the subsequent enrichment analysis. A SEA tool37 was first used to examine the enrichment for cortical-expressed genes at different developmental stages through childhood to young adulthood. A rank-based GO enrichment analysis using Gorilla38 was then conducted to test the enrichment for the ontology of cellular component. As the regional differences in gene expression may reflect underlying differences in cellular composition across cortical regions, a median rank-based approach61 was adopted to assess the enrichment for the adult cell types39.
Sensitivity analysis
To investigate whether our findings were robust and consistent across analytic choices, such as inclusion of additional covariates, clustering algorithms, atlas used for cortical parcellation, and computation of FC measures, a series of sensitivity analyses were conducted. Specifically, FC development subtyping was evaluated by further including SES and parental education as covariates to derive the RBD index. Neighborhood SES measure quantified by geocoded variables as in previous studies65,66 was adopted for the PNC cohort, and SES measure quantified as the natural log of the income-to-needs ratio was adopted for the HCP-D cohort as in ref. 67. Furthermore, subtyping results using GMM based clustering were evaluated, as suggested that K-means and GMM perform well in terms of stability and generalizability by a prior study on MRI based subtyping68. Finally, subtyping analyses were evaluated using additional cortical parcellations, including the Schaefer-200 atlas and Schaefer-300 atlas. The subtyping results using Fisher’s r-to-z-transformed correlation values as FC measures were also evaluated.
Cortical organization maps
The S-A axis1 and the first principal component of gene expression36 used in this study are available at https://github.com/PennLINC/S-A_ArchetypalAxis. The principal functional gradient map69 and the myelin content map70 are available at https://github.com/PennLINC/Brain_Organization.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data used in this study are publicly available. The Philadelphia Neurodevelopmental Cohort is accessible from the Database of Genotypes and Phenotypes (phs000607.v3.p2) at https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000607.v3.p2. Data from the Human Connectome Project-Development are available for download through the NIMH Data Archive (https://nda.nih.gov/). Source data are provided with this paper.
Code availability
Codes used for data analysis71 are available at: https://github.com/MLDataAnalytics/FC-SpatialHeterogeneity.
References
Sydnor, V. J. et al. Neurodevelopment of the association cortices: patterns, mechanisms, and implications for psychopathology. Neuron 109, 2820–2846 (2021).
Keller, A. S. et al. Hierarchical functional system development supports executive function. Trends Cogn. Sci. 27, 160–174 (2023).
Felleman, D. J. & Van Essen, D. C. Distributed hierarchical processing in the primate cerebral cortex. Cereb. Cortex 1, 1–47 (1991).
Toyoizumi, T. et al. A theory of the transition to critical period plasticity: inhibition selectively suppresses spontaneous activity. Neuron 80, 51–63 (2013).
Larsen, B. & Luna, B. Adolescence as a neurobiological critical period for the development of higher-order cognition. Neurosci. Biobehav. Rev. 94, 179–195 (2018).
Luo, A. C. et al. Functional connectivity development along the sensorimotor-association axis enhances the cortical hierarchy. Nat. Commun. 15, 3511 (2024).
Dosenbach, N. U. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Erus, G. et al. Imaging patterns of brain development and their relationship to cognition. Cereb. Cortex 25, 1676–1684 (2015).
Li, H., Satterthwaite, T. D. & Fan, Y. Brain age prediction based on resting-state functional connectivity patterns using convolutional neural networks. Proc. IEEE Int. Symp. Biomed. Imaging 2018, 101–104 (2018).
Kaufmann, T. et al. Common brain disorders are associated with heritable patterns of apparent aging of the brain. Nat. Neurosci. 22, 1617–1623 (2019).
Cui, Z. et al. Individual variation in functional topography of association networks in youth. Neuron 106, 340–353 e8 (2020).
Truelove-Hill, M. et al. A multidimensional neural maturation index reveals reproducible developmental patterns in children and adolescents. J. Neurosci. 40, 1265–1275 (2020).
Popescu, S. G. et al. Local brain-age: a U-net model. Front Aging Neurosci. 13, 761954 (2021).
Wang, Q. et al. Predicting brain age during typical and atypical development based on structural and functional neuroimaging. Hum. Brain Mapp. 42, 5943–5955 (2021).
Lee, P.-L. et al. Regional rather than global brain age mediates cognitive function in cerebral small vessel disease. Brain Commun. 4, fcac233 (2022).
Lund, M. J. et al. Brain age prediction using fMRI network coupling in youths and associations with psychiatric symptoms. Neuroimage Clin. 33, 102921 (2022).
Niu, X. et al. Multidimensional brain-age prediction reveals altered brain developmental trajectory in psychiatric disorders. Cereb. Cortex 32, 5036–5049 (2022).
Gianchandani, N. et al. A voxel-level approach to brain age prediction: a method to assess regional brain aging. J. Mach. Learn. Biome. Imaging 2, 761–795 (2024).
Zhou, Z. et al. Multiscale functional connectivity patterns of the aging brain learned from harmonized rsfMRI data of the multi-cohort iSTAGING study. Neuroimage 269, 119911 (2023).
Zhu, J. D. et al. Investigating brain aging trajectory deviations in different brain regions of individuals with schizophrenia using multimodal magnetic resonance imaging and brain-age prediction: a multicenter study. Transl. Psychiatry 13, 82 (2023).
Tetereva, A. & Pat, N. Brain age has limited utility as a biomarker for capturing fluid cognition in older individuals. Elife 12, RP87297 (2024).
Antoniades, M. et al. Relationship between MRI brain-age heterogeneity, cognition, genetics and Alzheimer’s disease neuropathology. eBioMedicine 109, 105399 (2024).
Sripada, C. et al. Prediction of neurocognition in youth from resting state fMRI. Mol. Psychiatry 25, 3413–3421 (2020).
Chen, J. et al. Shared and unique brain network features predict cognitive, personality, and mental health scores in the ABCD study. Nat. Commun. 13, 2217 (2022).
Lombardo, D. & Kaufmann, T. Different patterns of intrinsic functional connectivity at the default mode and attentional networks predict crystalized and fluid abilities in childhood. Cereb. Cortex Commun. 4, tgad015 (2023).
Lopez, K. C. et al. Development of network topology and functional connectivity of the prefrontal cortex. Cereb. Cortex 30, 2489–2505 (2020).
Shafiei, G. et al. Generalizable links between borderline personality traits and functional connectivity. Biol. Psychiatry 96, 486–494 (2024).
Baum, G. L. et al. Development of structure-function coupling in human brain networks during youth. Proc. Natl. Acad. Sci. USA 117, 771–778 (2020).
Sanders, A. F. P. et al. Age-related differences in resting-state functional connectivity from childhood to adolescence. Cereb. Cortex 33, 6928–6942 (2023).
Satterthwaite, T. D. et al. The Philadelphia Neurodevelopmental Cohort: a publicly available resource for the study of normal and abnormal brain development in youth. Neuroimage 124, 1115–1119 (2016).
Somerville, L. H. et al. The lifespan human connectome project in development: a large-scale study of brain connectivity development in 5-21 year olds. Neuroimage 183, 456–468 (2018).
de Lange, A. G. & Cole, J. H. Commentary: correction procedures in brain-age prediction. Neuroimage Clin. 26, 102229 (2020).
Peraza, J. A. et al. Methods for decoding cortical gradients of functional connectivity. Imaging Neuroscience 2, 1–32 (2024).
Dockes, J. et al. NeuroQuery, comprehensive meta-analysis of human brain mapping. Elife 9, e53385 (2020).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Burt, J. B. et al. Hierarchy of transcriptomic specialization across human cortex captured by structural neuroimaging topography. Nat. Neurosci. 21, 1251–1259 (2018).
Xu, X. et al. Cell type-specific expression analysis to identify putative cellular mechanisms for neurogenetic disorders. J. Neurosci. 34, 1420–1431 (2014).
Eden, E. et al. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinforma. 10, 48 (2009).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Marquand, A. F. et al. Understanding heterogeneity in clinical cohorts using normative models: beyond case-control studies. Biol. Psychiatry 80, 552–561 (2016).
Rutherford, S. et al. The normative modeling framework for computational psychiatry. Nat. Protoc. 17, 1711–1734 (2022).
Sun, X. Y. et al. Mapping neurophysiological subtypes of major depressive disorder using normative models of the functional connectome. Biol. Psychiatry 94, 936–947 (2023).
Kessler, D., Angstadt, M. & Sripada, C. Growth charting of brain connectivity networks and the identification of attention impairment in youth. Jama Psychiatry 73, 481–489 (2016).
Pines, A. R. et al. Dissociable multi-scale patterns of development in personalized brain networks. Nat. Commun. 13, 2647 (2022).
Yang, H. et al. Connectional axis of individual functional variability: patterns, structural correlates, and relevance for development and cognition. Proc. Natl. Acad. Sci. USA 122, e2420228122 (2025).
Baum, G. L. et al. Graded variation in T1w/T2w ratio during adolescence: measurement, caveats, and implications for development of cortical myelin. J. Neurosci. 42, 5681–5694 (2022).
Rogge, B. et al. Changes in brain structure in subjects with resistance to thyroid hormone due to THRB mutations. Thyroid Res. 16, 34 (2023).
Tutukova, S., Tarabykin, V. & Hernandez-Miranda, L. R. The role of neurod genes in brain development, function, and disease. Front. Mol. Neurosci. 14, 662774 (2021).
Thompson, C. C. & Potter, G. B. Thyroid hormone action in neural development. Cereb. Cortex 10, 939–945 (2000).
Bernal, J. Thyroid hormone receptors in brain development and function. Nat. Clin. Pr. Endocrinol. Metab. 3, 249–259 (2007).
Smith, J. W. et al. Thyroid hormones, brain function and cognition: a brief review. Neurosci. Biobehav. Rev. 26, 45–60 (2002).
Tau, G. Z. & Peterson, B. S. Normal development of brain circuits. Neuropsychopharmacology 35, 147–168 (2010).
Wagen, A. Z. et al. Life course, genetic, and neuropathological associations with brain age in the 1946 British Birth Cohort: a population-based study. Lancet Healthy Longev. 3, e607–e616 (2022).
Cole, J. H. et al. Brain age predicts mortality. Mol. Psychiatry 23, 1385–1392 (2018).
Gaser, C., Kalc, P. & Cole, J. H. A perspective on brain-age estimation and its clinical promise. Nat. Comput. Sci. 4, 744–751 (2024).
Butler, E. R. et al. Pitfalls in brain age analyses. Hum. Brain Mapp. 42, 4092–4101 (2021).
Moore, T. M. et al. Psychometric properties of the penn computerized neurocognitive battery. Neuropsychology 29, 235–246 (2015).
Schaefer, A. et al. Local-global parcellation of the human cerebral cortex from intrinsic functional connectivity MRI. Cereb. Cortex 28, 3095–3114 (2018).
Charrad, M. et al. Nbclust: an R package for determining the relevant number of clusters in a data set. J. Stat. Softw. 61, 1–36 (2014).
Markello, R. D. et al. Standardizing workflows in imaging transcriptomics with the abagen toolbox. Elife 10, e7212 (2021).
Seidlitz, J. et al. Transcriptomic and cellular decoding of regional brain vulnerability to neurogenetic disorders. Nat. Commun. 11, 3358 (2020).
Arnatkeviciute, A., Fulcher, B. D. & Fornito, A. A practical guide to linking brain-wide gene expression and neuroimaging data. Neuroimage 189, 353–367 (2019).
Vasa, F. et al. Adolescent tuning of association cortex in human structural brain networks. Cereb. Cortex 28, 281–294 (2018).
Alexander-Bloch, A. F. et al. On testing for spatial correspondence between maps of human brain structure and function. Neuroimage 178, 540–551 (2018).
Murtha, K. et al. Associations between neighborhood socioeconomic status, parental education, and executive system activation in youth. Cereb. Cortex 33, 1058–1073 (2023).
Moore, T. M. et al. Characterizing social environment’s association with neurocognition using census and crime data linked to the Philadelphia Neurodevelopmental Cohort. Psychol. Med. 46, 599–610 (2016).
Weissman, D. G. et al. Family income is not significantly associated with T1w/T2w ratio in the Human Connectome Project in development. Imaging Neurosci. 1, 1–10 (2023).
Zhang, S. R. et al. A comparative analysis of subtyping methodologies on cross-sectional sMRI data. Neurosci. Bull. 41, 1689–1695 (2025).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl. Acad. Sci. USA 113, 12574–12579 (2016).
Glasser, M. F. & Van Essen, D. C. Mapping human cortical areas in vivo based on myelin content as revealed by T1- and T2-weighted MRI. J. Neurosci. 31, 11597–11616 (2011).
Li, H. Spatial heterogeneity and subtypes of functional connectivity development in youth. FC-SpatialHeterogeneity (v0.0.1). Zenodo https://doi.org/10.5281/zenodo.17805407 (2025).
Yeo, B. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Acknowledgements
This study was supported by grants from the National Institute of Health: R01EB022573 (Y.F. and T.D.S.), R01AG066650 (Y.F.), U24NS130411 (C.D. and Y.F.).
Author information
Authors and Affiliations
Contributions
H.L. and Y.F. designed study; H.L. analyzed data and wrote the initial draft of the manuscript; R.E.G., R.C.G., and T.D.S. provided resources and supervised collection of PNC. Z.C., M.C., T.S., T.M.M., T.D.S., and R.T.S. contributed to data pre-processing and analytic tools; R.T.S., D.J.O., C.D,. and T.D.S. contributed to scientific guidance; Y.F. supervised the project; all authors (H.L., Z.C., M.C., T.S., T.M.M., R.E.G., R.C.G., R.T.S., D.J.O., C.D., T.D.S., and Y.F.) reviewed and revised the final draft of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Shaozheng Qin, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, H., Cui, Z., Cieslak, M. et al. Spatial heterogeneity and subtypes of functional connectivity development in youth. Nat Commun 17, 1956 (2026). https://doi.org/10.1038/s41467-026-68707-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68707-7