Abstract
Aeromonads are an ecologically versatile group of bacteria that cause infections in aquatic animals and are recognised as emerging human pathogens. Despite this, our understanding of Aeromonas diversity, especially the relationship between clinical and environmental strains, remains limited. Here, we present a genomic analysis of the Aeromonas genus, comprising 1853 genomes, and a detailed comparison of clinical and environmental strains from South Asia, including 996 newly sequenced genomes from Bangladesh and India. Phylogenetic analyses revealed that Aeromonas is a highly diverse genus, with no distinct clade separating clinical and environmental isolates. We identified 28 Aeromonas species and 905 novel sequence types, comprising 72.5% of the genomes. Notably, we show a high incidence of antimicrobial resistance (AMR) genes across all isolates, including against front and last-line antibiotics. Finally, we highlight frequent misidentification of Aeromonas as Vibrio cholerae, which is relevant to cholera-endemic regions where both genera co-exist and are associated with diarrhoeal disease. Our study underscores Aeromonas as an important environmental AMR reservoir and emerging multi-species pathogen capable of spilling over into human populations.
Similar content being viewed by others
Introduction
Aeromonads (within the Aeromonadaceae family) are Gram-negative, facultatively anaerobic bacteria with oxidase and catalase activity. Aeromonas spp. are abundant in aquatic habitats, with over 30 species identified1. At least 19 of these species are linked to human infections and are capable of causing significant diarrhoeal outbreaks. Aeromonas infections are common in children under 5 years old, especially in low- and middle-income countries (LMICs), where there is inadequate access to water and sanitation, resulting in high environmental exposure. The most common symptoms of Aeromonas infection include diarrhoea (which can be mistaken for cholera2,3), localised soft tissue infection, and bacteraemia4,5,6,7,8. Bacteraemia is most common in patients who have underlying health conditions, such as hepatobiliary disease, cancer, or diabetes9. There appear to be geographic differences in prevalence, with <1% of cases of moderate to severe diarrhoea (MSD), attributable to Aeromonas in four African sites and Kolkata, India, whereas Aeromonas was implicated in >22% of MSD cases in children from Karachi, Pakistan, and Mirzapur in Bangladesh10. Aeromonas spp. comprise important pathogens of fish and aquatic animals, causing diseases such as haemorrhagic septicaemia, furunculosis, and motile Aeromonas septicaemia (MAS) in species like salmonids, catfish, and tilapia, often leading to high mortality and significant economic consequences for aquaculture11,12,13,14,15,16,17. Additionally, Aeromonas spp. have also been isolated from terrestrial animals and are recognised as zoonotic, with human infections commonly occurring through contact with infected animals or contaminated water, especially via wounds18,19,20.
Pathogenicity in Aeromonads is complex; they possess a variety of virulence factors that contribute to biofilm formation, cell adherence, invasion, and cytotoxicity. These include polar and lateral flagella21,22, adhesins23, lipopolysaccharides, iron-binding systems24,25, and numerous extracellular toxins and enzymes26 secreted by various systems such as type 2 and type 3 secretion systems27,28. Additionally, quorum-sensing systems play a crucial role in colonisation and disease development29,30,31.
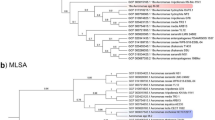
Notwithstanding the above, Aeromonas spp. are frequently misidentified as Vibrio cholerae in clinical and environmental samples. This is explained by sharing many biochemical and microbiological properties with other genera such as Aerobacter, Pseudomonas, Escherichia, Proteus, and Vibrio3,32. Both Vibrio and Aeromonas genera are oxidase-positive, able to thrive in aquatic and marine systems and exhibit similar colony morphologies on Thiosulphate Citrate Bile Salt Sucrose (TCBS) medium routinely used to select for V. cholerae33. These misidentifications pose serious epidemiological challenges, especially in regions where V. cholerae is endemic. Misidentification can not only lead to misreporting of cholera cases, but also a failure to implement necessary public health control measures. Hence, it is important to consider Aeromonas spp. in cholera surveillance and control efforts in endemic regions.
Accurate identification of Aeromonas spp. remains challenging due to overlapping phenotypic traits and the limited specificity of conventional biochemical tests and selective media, which often produce inconclusive results34,35,36. Molecular methods such as 16S rRNA sequencing are helpful but complicated by intragenomic variability, reducing species-level accuracy37,38. Emerging technologies like MALDI-TOF MS offer rapid and more reliable genus- and species-level identification37,39,40, while whole-genome sequencing provides definitive species-level resolution and accurate subtyping. Currently, a tiered diagnostic approach combining initial MALDI-TOF screening with phenotypic assays and, when necessary, molecular confirmation at reference laboratories is recommended to ensure accurate detection and effective epidemiological surveillance of Aeromonas spp.40. It is likely that, as for many other pathogens, genomic approaches will replace these techniques where accessible and in settings with well-developed supply chains.
Aeromonads are known to exhibit high levels of antimicrobial resistance (AMR), especially against β-lactam antibiotics. This resistance is primarily attributed to chromosomally encoded inducible β-lactamases in species such as A. hydrophila, A. sobria, and A. salmonicida, conferring resistance to antibiotics including ampicillin, bacitracin, cefoxitin, and piperacillin/tazobactam41,42,43,44,45. Recent genome data from 447 Aeromonas isolates from Pakistan revealed that human-associated Aeromonas in both clinical cases and age-matched controls were highly diverse and displayed a high level of AMR, particularly extended-spectrum beta-lactamases (ESBLs)46. Moreover, Aeromonas serve as natural reservoirs for mobile colistin resistance genes (mcr)47,48, raising concerns about their role in spreading resistance to last-resort antibiotics like colistin across bacterial populations in aquatic and clinical environments. Looking into determinants that may explain the difference in clinical presentations, this study looked across an array of known or putative virulence factors, of which only two, maf2 and lafT, were weakly associated with the MSD cases. However, by focusing on isolates from humans, this study was unable to establish whether human pathogenic strains were a specialised subset of those found in the environment or if Aeromonas spp. infections are truly opportunistic and linked to environmental exposure46.
Given the ubiquitous nature of Aeromonads in the environment, their potentially emerging role in MSD in South Asia, and the fact that they are often cultured from samples taken from patients with suspected cholera, we created a baseline population genomic snapshot of Aeromonas species from clinical and environmental samples. We sequenced two collections of Aeromonas isolates from V. cholerae endemic regions of Bangladesh (2004–2016), and Northern India (2020–2023). These isolates were obtained from environmental sources—including drinking water, ponds, rivers, lakes and drains, as well as from human faeces, together comprising 996 novel Aeromonas genomes. We compared their genetic diversity, predicted virulence gene profiles, and AMR profiles, and contextualised this with genomes from a global Aeromonas population dataset.
Results
Species diversity and distribution of Aeromonas spp. genomes
To examine the genomic diversity of Aeromonas strains across clinical and environmental settings in South Asia, we included the largest and only available clinical genome datasets from Pakistan and environmental datasets from India and Bangladesh (this study). To place our study into a global context, we also incorporated genomes representing broad geographic and species diversity within the Aeromonas genus. In total, we assembled 1853 Aeromonas genomes, including 996 newly sequenced genomes, of which 198 (10.68%) were mistakenly collected as Vibrio cholerae−133 from suspected cholera cases/households contacts in the ChoBI7 programme in Dhaka city49 and 65 from an environmental surveillance study for V. cholerae conducted in Bangladesh. Further, we generated 798 Aeromonas genomes from isolates taken from 274 water samples collected in Northern India. We contextualised our dataset using public archives, including 441 genomes of clinical origin from children with moderate to severe diarrhoea and corresponding healthy controls, taken in Pakistan46, as well as 416 globally distributed50 Aeromonas genomes. The latter were derived from a wide range of sample types, including human stool, urine, wounds, as well as aquatic environments (e.g., rivers and wastewater), and animals (e.g., fish, duck, dog). The largest proportion of publicly available genomes originated from Denmark (n = 100), followed by the United States (n = 71) and China (n = 62) (see Supplementary Table 1 for detailed information).
We used GTDB-Tk (Genome Taxonomy Database) to perform species assignment, identifying 28 Aeromonas spp. across the 1853 genomes, of which eight species dominated. A. caviae (n = 608) was the most numerous in our collection, followed by A. veronii (n = 566), A. dhakensis (n = 186), A. salmonicida (n = 176), A. hydrophila (n = 124), A. enteropelogenes (n = 105), A. jandaei (n = 25), A. sanarellii (n = 16), with the remaining 47 Aeromonas genomes representing 20 other Aeromonas species (Fig. 1; summarised in Supplementary Fig. 1a). To understand how our species predictions mapped to established species we used Average Nucleotide Identity (ANI) to evaluate their genetic relatedness. The intra-species ANI values ranged from 94.06% to 100%, falling slightly below the established definition for a single species (>95% ANI), in some instances. Conversely, inter-species ANI values ranged from 79.66% to 95.51% with higher-than-expected ANI and lower levels of expected divergence between A. bestiarum and A. piscicola (95.14–95.51%) (Supplementary Fig. 2).
A maximum likelihood phylogenetic tree based on 1969 core genes for Aeromonas spp. genomes. Refer to the key for the interpretation of the colour strips alongside. The scale indicates an evolutionary distance of 0.1 nucleotide substitutions per site. BAPS refers to Bayesian Analysis of Population Structure, used here to delineate genetic clusters.
Looking at the geographic distribution of Aeromonas species across both clinical and environmental samples from India, Pakistan and Bangladesh, where we had the densest sampling (1438 Aeromonas genomes; Supplementary Fig. 1b), Aeromonas spp. found across all three South Asian countries included A. caviae, A. dhakensis, A. enteropelogenes, A. jandaei, A. veronii and A. sanarellii, with A. sanarellii and A. jandaei being rare in all three countries. Whilst overall there were similarities in the relative abundance of the different species, despite differences in sampling depth and strategy (see methods), there were also clear differences. In particular, A. caviae predominated in Bangladesh and Pakistan (31.81%, 63/198 and 64.39%, 284/441; respectively), whereas A. veronii dominated the samples collected in India (48.31%, 386/799 of isolates). Other species, such as A. schubertii, were exclusively seen in Bangladesh (n = 1), while A. media (n = 4) and A. rivipollensis (n = 5) were exclusively seen in India (Supplementary Fig. 1b). What is more, 129 environmental samples from Northern India included between 1–5 different Aeromonas species (Supplementary Fig. 3a). For countries where we had less dense sampling, A. hydrophila was the most common in the United States (42.25%, 30/71), whereas A. salmonicida was the dominant species in the samples from Denmark (99%, 99/100) and China (96.77%, 60/62), (Supplementary Table 1).
In India, all samples were collected as part of a cholera surveillance study. Samples were taken from known cholera hotspots and at times of the year when cholera cases were more common. All water samples were cultured on V. cholerae selective media (TCBS; see methods). Using MALDI-TOF, it was possible to look at the Aeromonas spp. positivity rate, as well as for the presence of a range of other enteric pathogens (Supplementary Fig. 4). Out of 408 collected water samples from Northern India, 274 samples (67.15%) tested positive for Aeromonas spp., 145 (35.53%) for V. cholerae, 112 (27.45%) for E. coli, 198 (48.52%) for Klebsiella spp. and 35 (8.57%) for non-cholera Vibrio spp. (8.57%, 35/408; including species V. fluvialis, V. navarrensis, V. cidicii, V. injensis, and V. metschnikovii).
Of the samples positive for Aeromonas spp., V. cholerae was also isolated from 91 samples (33.57%, 91/274). These co-occurrences were primarily found in river samples (75/91), followed by ponds (13/91) and canals (4/91), across different regions of Northern India. A total of seven Aeromonas species: A. veronii, A. caviae, A. hydrophila, A. dhakensis, A. jandaei, A. enteropelogenes, and A. sanarellii–were identified as cohabiting with V. cholerae. However, there was no clear association between the co-occurrence of V. cholerae with any one specific Aeromonas species; these data largely mirrored the relative isolation rates for Aeromonas species from Northern India (Supplementary Fig. 3b; Supplementary Fig. 3c).
Genomic taxonomy of Aeromonas spp
To infer the phylogeny of the genus, we standardised the annotation of our underlying dataset using PROKKA and then defined the Aeromonas spp. pangenome using Panaroo. This comprised 47,542 coding sequences (CDS), of which 1969 were defined as core (found in more than 99% of genomes) and 118 as soft core (found in 95–99% of genomes), with 45,455 accessory genes (found in 0–95% of genomes). We constructed a maximum likelihood phylogenetic tree from a core gene alignment (Fig. 1; see methods). We used Bayesian Analysis of Population Structure (BAPS, implemented in fastBAPS) to delineate genomic clusters and correlated these with ANI values and the species assignments from GTDB-Tk. From BAPS, whilst most taxa fell on deeply branching nodes in the Aeromonas spp. tree with high concordance with the ANI-defined species blocks (Fig. 1; Supplementary Fig. 2), A. veronii fell across two BAPS clusters (15 and 16). Conversely, BAPS clusters 1, 2, 3, and 7 each span multiple species of Aeromonas (Supplementary Table 2). These inconsistencies may reflect differences between the way the BAPS model clusters genomes, which, unlike ANI, is not distance-based.
Next, to determine how much of the diversity within known taxonomic groups had been previously observed, we constructed in silico MLST51 profiles. Of the 1853 MLST profiles generated, only 495 (26.71%) genomes were assigned to 224 known MLST allelic profiles, with 1343 genomes representing 905 novel sequence types (STs). After assignment by pubmlst.org (https://pubmlst.org/organisms/aeromonas-spp), 1838 of our Aeromonas spp. genomes were assigned to 1129 STs (15 genomes remained unassigned; see Supplementary Data 1), now accessible through the Aeromonas MLST database52. Of note, these novel STs define discrete branches within all species, while 836 STs represented singletons, with clusters of genomes belonging to the same ST being rare. The exceptions to this were ST-2 (n = 103) and the newly defined ST-1799 (n = 60) for A. salmonicida, as well as ST-2398 (n = 20) and ST-251 (n = 19), representing A. caviae and A. hydrophila genomes, respectively.
Virulence gene distribution in clinically and environmentally derived Aeromonas spp
Several virulence genes have been previously described amongst Aeromonas species, including the haemolytic toxins: aerolysin-related cytotoxic enterotoxin (act)41,53,54, heat-labile cytotonic enterotoxin (alt)53,54,55, heat-stable cytotonic toxins (ast)53,55, hemolysin (hlyA)56, cytotoxin aerolysin (aerA)53,55,56 as well as polar flagellum (fla)55,57,58, lateral flagella (laf)59,60, the zinc-and-iron-dependent metallo-endopeptidase elastase (ela)57, the lipase (lip)41,55 and the type III secretion system (TTSS) encoded by ascF-G57. To determine whether any of the known virulence genes are linked to the particular Aeromonas species, we used in silico PCR to understand the distribution of these genes across 1853 Aeromonas species genomes.
We analysed patterns of virulence gene distribution using principal component analysis (PCA) and hierarchical clustering (Fig. 2a; Fig. 2b). The PCA revealed that certain species, such as A. hydrophila, A. dhakensis, A. salmonicida, and A. veronii, formed relatively well-defined clusters, suggesting distinct virulence profiles (Fig. 1; Fig. 2a). Some overlap was observed between the phylogenetically related A. hydrophila and A. dhakensis, as well as between the phylogenetically distant A. hydrophila and A. salmonicida. Genes such as aerA, fla, laf, and lip were found across nearly all species. For instance, aerA was present in over 60% of A. caviae, A. dhakensis, and A. veronii, and in over 95% of A. hydrophila and A. salmonicida genomes. Similarly, fla was widely distributed, occurring in 70–90% of genomes across most species. Consistent with previous studies61,62, hlyA and ast were more species-specific, with hlyA almost exclusively found in A. dhakensis (99.5%) and A. hydrophila (100%), and ast present in 95.2% of A. hydrophila genomes but nearly absent elsewhere. A. caviae (n = 608) and A. salmonicida (n = 176) carried the highest proportions of key virulence genes; all A. caviae genomes encoded alt and lip, with nearly all (99.7%) possessing ela, while A. salmonicida uniformly carried aerA, alt, ela, fla, lip, and act. Genes ela and lip were also prevalent in A. dhakensis (100%), A. hydrophila (100%) and A. sanarellii (93.8%), while A. enteropelogenes and A. jandaei lacked several virulence genes (Fig. 1; summarised in Fig. 3a).
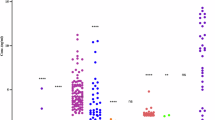
a Principal component analysis (PCA) of virulence genes across different Aeromonas spp. (see key for coloured dots representing species). b Hierarchical clustering of virulence genes across Aeromonas spp., with colours representing the percentage presence of genes (see key for scale).
The x-axis represents the most abundant Aeromonas species, with less abundant species grouped as Aeromonas spp., and the y-axis denotes virulence genes in (a) and antimicrobial classes corresponding to AMR genes in (b). Each dot is coloured based on the gene abundance per species (see scale), and the size of the dots corresponds to the number of genomes harbouring the gene.
The Type III secretion system genes (ascF-G) were found more often in A. salmonicida (80%, 141/176), A. veronii (51%, 288/566), A. dhakensis (42%, 78/186), and A. hydrophila (17%, 21/124). Gene hlyA was present in all isolates of A. salmonicida, A. dhakensis, and A. hydrophila, but absent in A. jandaei, A. enteropelogenes, A. sanarellii, A. caviae, and A. veronii, representing a potential molecular marker for differentiating between species (Fig. 2b). Similarly, the enterotoxin-encoding ast gene was found only in A. hydrophila (95%, 118/124) and at low levels in A. dhakensis (6%, 11/186), highlighting its potential for species-specific identification. We noted, a high prevalence of the combination of the cytotoxic enterotoxin (act) and cytotonic heat-labile enterotoxin (alt) in A. salmonicida (alt = 100%, 176/176; act = 99%, 175/176), A. hydrophila (alt = 100%, 124/124; act = 72%, 90/124), and A. dhakensis (alt = 94%, 176/186; act = 29%, 55/186). We also observed variation in the distribution of laf, suggesting that this gene had been gained or lost multiple times across the phylogeny (Fig. 1); for example, it was present in A. salmonicida ST-2 but absent from ST-1799.
Lastly, we investigated whether virulence gene presence varied within species depending on sample origin. For this, we focused on South Asia (Bangladesh, India and Pakistan), since here we had 1438 Aeromonas spp. genomes confirmed to originate from clinical or environmental samples. As shown in Supplementary Fig. 5a and 5b, no obvious association was observed between species, sample origin, and virulence gene presence. This was also true when looking at the phylogeny of A. caviae taken from South Asia (n = 577), where we had almost equal representation from clinical (n = 286; of which 153 were from individuals with moderate to severe diarrhoea, while 133 were asymptomatic) and environmental (n = 291) Aeromonas genomes (Supplementary Fig. 6). Of the 11 BAPS clusters, 10 contained both clinical and environmental Aeromonas genomes; the only exception was cluster-4.10, which included just five isolates in total, all from environmental sources. Sublineages containing a mixture of both clinical and environmental isolates were also identified in the other abundant Aeromonas species (A. veronii, A. dhakensis, A. enteropelogenes, and A. hydrophila; Supplementary Table 3).
In silico prediction of AMR
To understand if there were differences in AMR profiles across or within species, we inferred the genotypic AMR profiles for all the genomes included in this study. Overall, we found 162 discrete AMR genes, conferring resistance to 16 different antimicrobial classes, including β-lactams, sulphonamides, tetracycline, and aminoglycosides (Fig. 3b, summarised in Supplementary Table 4). Of the 162 AMR genes identified, those conferring resistance to β-lactam antibiotics were the most common, representing 40.74% (66/162) of the genes. Within this group, Class D oxacillinases were the most common, present in 94.5% of genomes, with genes such as blaOXA-427 (44.6%, 826/1853) and blaOXA-12 (32.7%, 605/1853) being particularly abundant. This was followed by Class C cephalosporinases, which were frequently found in 71.3% of genomes, including blaMOX-6 (18.9%, 350/1853) and blaFOX-2 (9.6%, 177/1853). Class B metallo-β-lactamases were detected in 55.4% of genomes, dominated by blaCEPH-A3 (22.1%, 410/1853) and blacphA4 (9.6%, 177/1853). In contrast, Class A β-lactamases were less common, identified in 5.07% of genomes, with genes such as blaKPC-1 (0.8%, 15/1853) and blaRSA-1 (1.1%, 20/1853) among the more notable types (Supplementary Table 5). Of note, mobile colistin resistance genes (mcr) were detected in several species. These included A. jandaei (100%, 25/25), all of which carried mcr-7.1; A. piscicola (100%, 1/1) harbouring mcr-3 and mcr-3.8; A. salmonicida (59.1%, 104/176) carrying both mcr-3 and mcr-3.12; A. media (28.6%, 2/7) with mcr-3 and mcr-3.6; A. sobria (16.7%, 1/6) harbouring mcr-3 and mcr-3.12; A. veronii (1.94%, 11/566) with mcr-3, mcr-3.12, mcr-3.3, mcr-3.6, and mcr-3.8; A. caviae (1.32%, 8/608) carrying mcr-3, mcr-3.12, mcr-3.3, and mcr-3.6; and A. hydrophila (0.81%, 1/124), which harboured mcr-5 (Supplementary Table 4; see Supplementary Data 1). Only three genomes in our collection lacked any known AMR gene.
Looking more closely at the subset of 1438 South Asian genomes, we found that genotypic AMR patterns varied significantly across India, Bangladesh, and Pakistan (Fig. 4a). Aeromonas isolates from Pakistan exhibited the highest prevalence of resistance to β-lactams (cephamycin: 65.8%, 290/441), sulphonamides (24.3%, 107/441), diaminopyrimidines (22.4%, 99/441), aminoglycosides (22.2%, 98/441), tetracyclines (18.4%, 81/441), phenicols (16.3%, 72/441), and fluoroquinolones (12.9%, 57/441). Indian isolates showed comparatively lower prevalence of resistance gene presence amongst the three South Asian countries, with the highest resistance observed for carbapenems (69.2%, 553/799); macrolides (7.8%, 62/799); rifamycins (3.3%, 26/799); and oxazolidinones, pleuromutilins, and streptogramins (4.4%, 35/799). Aeromonas from Bangladesh had the lowest prevalence of resistance, with no genes conferring resistance to lincosamides, oxazolidinones, pleuromutilins, streptogramins, and nucleosides. Notably, all genomes across the three countries were predicted to carry cephalosporin resistance genes. Among these, 4.5% of the isolates (65/1438; 14 clinical and 51 environmental) carried genes encoding third-generation extended-spectrum β-lactamases (ESBLs), such as blaCTX-M, blaGES, blaPER, blaTEM, and blaVEB.
The x-axis represents the country of origin in (a) and the sample type (clinical vs. environmental) in (b), while the y-axis denotes antimicrobial classes corresponding to resistance genes. The heatmap is colour-coded to indicate the percentage presence of AMR genes per country and sample type, with numeric indicators above each tile denoting the number of genomes carrying the gene within that category. c Principal component analysis (PCA) illustrates the clustering of genes in clinical and environmental isolates (see key for coloured dots). d Absolute difference in gene presence between clinical and environmental isolates, with yellow-green indicating minimal or no difference and blue-violet representing genes with significant variation in presence across sample types.
To further refine our understanding beyond drug class-level resistance, we assessed the genetic differences in AMR gene profile between clinical and environmental isolates among the 1438 genomes using PCA and NMDS. This revealed considerable overlap between both groups within each species (Fig. 4c and Supplementary Fig. 7a). Notably, species like A. caviae and A. veronii formed distinct clusters, likely due to differences in AMR gene diversity (Supplementary Fig. 7b and Supplementary Table 6). Environmental isolates generally exhibited a greater diversity in AMR genes carriage than the clinical isolates across Aeromonas species (Supplementary Table 7), except in A. caviae, where clinical isolates contained more unique determinants (18 vs. 7) but also had largest overlap (48 shared genes; Supplementary Fig. 8). Notably, A. veronii and A. dhakensis environmental isolates carried extensive AMR repertoires (45 and 22 genes, respectively) with several shared with the clinical strains, suggesting potential environmental reservoirs (Supplementary Table 7). Shared genes across species commonly included tet(A), tet(E), blaOXA and blaMOX variants, sul1/sul2, and qnr variants, indicating frequent exchange between strains occurring in the aquatic and clinical settings, while certain ESBLs and carbapenemases (e.g., blaCTX-M-15, blaVIM-6) remained confined to the clinical isolates included in this study. Furthermore, certain AMR genes exhibited notable differences in prevalence between the two groups (Fig. 4d); clinical isolates had a higher proportion of blaOXA-427 (64.7%, 288/445), blaMOX-6 (35.7%, 159/445), and sul1 (17.3%, 77/445), compared to environmentally derived isolate genomes. In contrast, environmental isolates showed higher proportions of blaOXA-12 (43.9%, 435/990), blaOXA-724 (18.3%, 181/990), blaCEPHA3 (28.7%, 284/990) and blacphA4 (12.7%, 126/990) (Fig. 4d). Of note, one Indian environmentally derived A. sanarellii isolate was predicted to be multidrug resistant by possessing genes to seven drug classes including: aminoglycoside, β-lactam, diaminopyrimidine, macrolide, phenicol, rifamycin, and sulphonamide (see Supplementary Data 1).
Discussion
We initiated this study to provide a genomic framework for understanding the relationship between Aeromonas spp. from the natural aquatic environment and those isolated from clinical samples linked to disease. We also wanted to understand the nature and diversity of Aeromonas spp. co-inhabiting the same natural environment with Vibrio cholerae, in endemic settings. We included a range of isolates and genomes generated here, as well as published Aeromonads genomes from moderate to severe diarrhoea cases and matched controls from Pakistan46. We investigated the genetic diversity, prevalence of virulence and antibiotic resistance genes amongst Aeromonads genomes from households and freshwater in multiple countries. In doing so, we have provided the most comprehensive view of this genus thus far.
Among 1853 genomes included in this study, we identified 28 Aeromonas species, including A. veronii, A. caviae, A. dhakensis, A. enteropelogenes, A. salmonicida, and A. hydrophila, several of which are known human or fish pathogens13,16,17,20,34,63,64. A. dhakensis and A. enteropelogenes were prevalent in Bangladesh, comprising approximately 22% and 29% of isolates, respectively, but were less common in the neighbouring countries. Other notable geographic signals include the psychrophilic A. salmonicida, seen largely in cooler countries such as Denmark (56%) and certain regions of China (35%, primarily from salmon facilities), where most commercial fish farming is done65. Here, the majority of the A. salmonicida were collected from infections of trout, Oncorhynchus mykiss, known to be highly susceptible to A. salmonicida infections. Similarly, 20% of A. hydrophila described here originated from diverse sources, including wastewater, fish and humans, and were from the United States of America, where A. hydrophila causes significant economic losses in catfish farming66.
Despite being considered an emerging human and animal pathogen linked to a wide array of diseases, there is limited available genomic data for the Aeromonas genus, particularly from environmental sources. Our core gene phylogeny highlights that this genus is remarkably diverse; this was particularly evident from the high number of novel MLST STs (905), with 1343 (72.47%) genomes assigned to these previously unreported profiles (see Supplementary Data 1). Perhaps unsurprisingly, given the extent of the previously unknown diversity, ANI showed the intra-species ANI values for A. sobria and A. veronii fell below the standard 95% threshold, challenging current boundaries for species delineation and suggesting a need for reclassification within these species.
Looking at the species distribution between human and environmental samples, from previous studies of children with moderate-to-severe diarrhoea in Karachi, Pakistan, A. caviae (64.2%) was seen to be the most numerous, followed by A. veronii (19.2%), A. dhakensis (9.8%), and A. enteropelogenes (4.9%)46. Other studies have also shown that A. hydrophila is also common in causing intestinal and extra-intestinal infections67. From our data, comparing diarrhoea-associated clinical and environmentally derived isolates from South Asia, we showed a similar distribution across both human and environmental samples, with A. caviae, A. veronii, A. dhakensis, and A. enteropelogenes being the most abundant.
To identify signatures of intra- and interspecific ecological separation within and between clinical and environmental isolates, we examined all genomes for differences in virulence potential and AMR profiles, linked to taxonomic affiliation and source of isolation. From the virulence gene profiles, there were no significant genetic differences between clinical and environmental strains. However, these profiles allowed us to observe distinct clusters formed by species such as A. hydrophila, A. veronii, A. dhakensis, and A. salmonicida, implying that virulence gene profiles are strongly associated with species-level taxonomy, and could be used to classify Aeromonas spp. into different groups, serving as a useful marker. For instance, the nearly exclusive presence of hlyA in A. hydrophila and A. dhakensis, as well as the conserved presence of ela and lip in multiple species, could inform the development of molecular assays for rapid species identification. Furthermore, the consistent clustering patterns observed in PCA and hierarchical analyses highlight the potential of virulence gene profiling as a complementary approach to traditional phylogenetic or multi-locus sequence typing methods.
Most Aeromonas genomes, irrespective of source, carried AMR genes, particularly those conferring resistance to β-lactams, aminoglycosides, and sulphonamides, consistent with previous clinical studies46,65. Notably, in our study, we identified a wide range of β-lactamase genes, highlighting the problem of widespread β-lactam resistance, consistent with global trends42,46,68. These included chromosomally encoded class B metallo-β-lactamases (blaCphA), class C cephalosporinases (AmpC β-lactamases: blaFOX, blaMOX), and class D β-lactamases (oxacillinases: blaOXA variants), underscoring the well-documented intrinsic resistance mechanisms of Aeromonas spp.43,69,70. Additionally, a notable proportion of isolates carried acquired ESBLs (n = 52, 2.8%; e.g., blaCTX-M, blaGES, blaPER, and blaVEB) and carbapenemases (n = 16, 0.9%; blaKPC, blaVIM)71,72. This is particularly concerning given the lack of standardised treatment guidelines for Aeromonas infections, where fluoroquinolones, particularly ciprofloxacin and, in some cases, third-generation cephalosporins are commonly used empirically32,34,73,74,75. Our analysis showed considerable overlap in AMR gene profiles between clinical and environmental isolates, with few genes showing distinct patterns. For example, blaOXA-427, a class D carbapenemase previously identified in several Enterobacteriaceae clinical strains76, appeared more frequently in clinical Aeromonas isolates. In contrast, The class D carbapenemases gene blaOXA-12, first identified in A. jandaei77 and most abundant in environmental isolates, was found in all A. jandaei, 565 A. caviae, and two A. hydrophila isolates, as well as in less common species like A. allosaccharophila (n = 5), A. australiensis (n = 1), A. fluvialis (n = 1), and A. sobria (n = 6). The gene blaOXA-12 was widely distributed across several countries and has been previously reported in environmental A. veronii from the United States78 and in strains from humans and animals in India79. The presence of these genes in both clinical and environmental isolates raises concern, as carbapenems are critical last-resort antibiotics80,81.
We also observed the presence of mobile colistin resistance genes in eight different Aeromonas species, spanning diverse geographical regions including Bangladesh, China, Denmark, India, Pakistan, Spain, the United Kingdom, and the United States. Colistin remains a last-resort antibiotic for treating clinical infections caused by multidrug-resistant Gram-negative bacteria82. The presence of mcr genes, many of which are plasmid-encoded, raises significant public health concerns, as they can facilitate horizontal gene transfer to other clinically important pathogens82. The widespread occurrence of mcr in both clinical and environmental Aeromonas83,84,85 isolates further suggests that this genus may act as both a reservoir and conduit for global colistin resistance dissemination.
Notably, the AMR patterns observed in Aeromonas spp. from South Asia highlight significant regional variability, with clinical isolates from Pakistan (collected from individuals with moderate to severe diarrhoea) 46 exhibiting resistance across a broader range of antibiotic classes compared to environmental isolates from India and Bangladesh. Nevertheless, environmental isolates often harbour a greater diversity of unique AMR genes within several species, underscoring their role as reservoirs of antimicrobial resistance genes and that antimicrobial contamination of the environment is likely driving their selection and ultimately evolution. Whilst it is clear that many resistance genes are shared between clinical and environmental isolates, with higher proportions of these genes in clinical isolates, it is important to note that the lack of both environmental and clinical isolates from the same geographic region is a limitation of this study. It is also important to note that we lacked phenotypic resistance data; therefore, our findings are based purely on genotypic inference. Overall, this suggests that while environmental isolates contribute to the overall AMR gene pool, selective pressures in clinical settings likely drive the greater prevalence of resistance genes in clinical isolates.
Taken together, the widespread occurrence of clinically relevant resistance genes in Aeromonas, a genus found in aquatic and wastewater environments, highlights its role as a key environmental sentinel in the One Health context. This aligns with the broader understanding that various environmental sources contaminated with residual antimicrobial substances, such as wastewater, agricultural runoff, and pharmaceutical discharges86,87,88, select for AMR in important bacteria such Aeromonas and members of the Enterobacteriaceae. These contaminated environments not only support the survival of resistant organisms but also promote their genetic exchange and adaptation, potentially accelerating the spread of resistance across both clinical and environmental settings and harbouring genes conferring resistance to first and last-line antimicrobials.
One of the drivers for this study was that Aeromonas spp. are frequently misidentified as V. cholerae in clinical and environmental samples3,89. Our data shows that 67.15% of the environmental isolates cultured on TCBS in Northern India were Aeromonas spp. and not V. cholerae. Similarly, of the environmental isolates that were originally collected and identified as V. cholerae through culture from the Dhaka household study, ~70% were subsequently reclassified as Aeromonas spp. This contrasts with the prevalence of this genus among the 239 stool samples positive for V. cholerae from the same study, where only two Aeromonas spp. were misidentified as V. cholerae, both of which were cultured from asymptomatic individuals, Dr Munir Alam, personal communication (ref. 49, this study). This suggests that misidentification of Aeromonas for V. cholerae is unlikely to overestimate cholera prevalence in clinical settings (e.g., from human stool). Still, environmental sampling and identification based on microbiological growth and appearance on selective media may overinflate apparent V. cholerae prevalence in the environment, especially during a cholera epidemic, if culture is the only confirmatory method employed.
In conclusion, our data brings together the different ecological views of this bacterium; it shows the environment to be an important reservoir of a highly diverse range of species capable of overspilling into humans and, in some instances, causing disease. It is also clear that they carry an important array of therapeutically relevant AMR genes. Here, by taking a broader approach to understanding questions such as sources and sinks of AMR and considering non-traditional organisms, we show that Aeromonas has seemingly been hidden in plain sight – both as an emerging human pathogen and clearly as an important environmental reservoir of AMR.
Methods
Aeromonas isolates and genome sequences
We used a collection of 996 novel Aeromonas isolates. Of these, 198 had been originally cultured and identified as V. cholerae using conventional microbiological and biochemical techniques90 before they were sequenced and classified genomically as Aeromonas. These included 133 isolates from a study entitled the Cholera-Hospital-Based-Intervention-for-7-Days (ChoBI7) programme in Dhaka city49 and 65 isolates from an environmental surveillance study for V. cholerae conducted in Bangladesh. These 198 genomes were obtained from diverse sources, including drinking water, plankton, zooplankton, sediment, and human stool or rectal swabs, collected between 2004 and 2016. The remaining 798 genomes were obtained from 408 water samples collected for isolation of V. cholerae from ponds, rivers, and canals across six different states—Delhi, Haryana, Himachal Pradesh, Punjab, Uttar Pradesh and Uttarakhand, as well as Chandigarh in Northern India—between 2020 and 2023. The collected water samples were first enriched in alkaline peptone water and Selenite F broth. They were then sub-cultured onto various media. Blood agar and Thiosulphate Citrate Bile Salt Sucrose (TCBS; Difco Laboratories, Detroit, Michigan, USA) agar were used for the isolation of V. cholerae and Aeromonas spp. MacConkey agar (HiMedia Laboratories Private Limited, Mumbai, India) and Xylose Lysine Deoxycholate (XLD; Difco Laboratories, Detroit, Michigan, USA) were used for isolating other Enterobacteriaceae. The presumptively selected colonies from the media listed above were identified by matrix-assisted laser desorption ionisation-time of flight mass spectrometry (MALDI-TOF)91, and isolates that were identified as Aeromonas spp. underwent DNA extraction and whole genome sequencing as described below. Between 1 and 9 colonies were retained from each Aeromonas-positive water sample in India for downstream processing, including DNA extraction and whole genome sequencing.
DNA was extracted from bacterial isolates using the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) and Qiagen DNeasy Blood & Tissue Kits (Qiagen, Hilden, Germany). Illumina sequencing libraries were prepared using 0.5 µg DNA and sheared using an S2 ultrasonicator (Covaris), before adaptor ligation and dual index barcoding (NEBNext UltraII, New England Biolabs)92. Libraries were sequenced on the HiSeq X10 and NovaSeq 6000 platform (Illumina, San Diego, CA, USA) at the Wellcome Sanger Institute (Cambridge, UK), generating 150 bp paired-end reads with a mean depth of coverage of 68× per genome (range: 21–118×, excluding outliers).
A total of 1853 genomes were analysed, including 996 genomes sequenced in this study. These genomes were contextualised using 416 Aeromonas genomes (including one genome from India) downloaded from a library of 661,405 previously assembled bacterial genomes held in the European Nucleotide Archive (ENA)50, along with 441 genomes from a study on children with moderate to severe diarrhoea and corresponding controls in Pakistan46.
Quality control and genome assembly
We performed initial quality control of sequencing reads using FastQC (v0.11.4.)93 and MultiQC (v1.17)94. To further screen for contamination in the raw sequencing reads, we used Kraken2 (v2.2)95 and Bracken (v2.6.2)96 (Standard RefSeq database containing archaea, bacteria, viruses, plasmids and human, downloaded from https://benlangmead.github.io/aws-indexes/k2 on 9th October 2023). No trimming of raw reads was performed prior to de novo assembly using SPAdes (v3.9.0)97 to obtain contiguous scaffolded assemblies. Additional quality control was performed on the de novo assemblies using Quast (v5.0.2)98 and CheckM (v1.1.2)99 to calculate the genome contamination and completeness. Assemblies were filtered to retain high-quality assemblies based on genome length (4.5-5.2 Mb), genome contamination (<5%) and genome completeness (>95%). Overall, 5% (97/1950) of the genomes were excluded from the study based on these criteria, including six genomes from the study in Pakistan.
Species identification and distinction
For species-level classification, we employed GTDB-Tk (v. 2.4.1)100 with the Genome Taxonomy Database (GTDB; R226, released on 16 April 2025), see Supplementary Data 2. GTDB-Tk integrates marker gene analysis, phylogenetic placement, and average nucleotide identity (ANI) to provide robust and consistent taxonomic assignments. This was followed by assessment of genome-relatedness indices, where we used FastANI (v1.3)101 for the pairwise comparison of the Aeromonas genomes. For a subset of genomes (n = 49), digital DNA–DNA hybridisation (dDDH) assays were also performed to further refine species-level resolution (Supplementary Data 3). The FastANI threshold was set at 90% based on pairwise sequence mapping101, and heatmaps based on estimated ANI for all Aeromonas genomes were created using the Complex heatmap package in R (v4.4.3)102.
Prediction of AMR and virulence genes
AMR genes were identified from the assembled contigs using ABRicate (v1.0.1, available at https://github.com/tseemann/abricate) to query the Comprehensive Antimicrobial Resistance Database (CARD, v3.2.4, last updated in August 2022)103, where the CARD identification threshold was set at ≥80% and <80% nucleotide identity, both with ≥80% coverage (Supplementary Data 1). For the identification of the putative virulence genes, we initially used ABRicate with the Virulence Finder Database104, but this did not yield any hits. We subsequently used in_silico_pcr (v1.0.0, available at https://github.com/sanger-pathogens/sh16_scripts/blob/master/legacy/in_silico_pcr.py) with previously described primer sequences (see Supplementary Table 8)105,106 to detect virulence genes. To explore the patterns of AMR and virulence gene distribution in relation to sample origin (clinical vs. environmental) and Aeromonas species, we conducted a Principal Component Analysis (PCA) using the prcomp function from the stats package in R (v4.4.3)102, based on presence/absence matrices. The first two components, PCA1 and PCA2, were used. Furthermore, we applied Non-metric Multidimensional Scaling (NMDS) with the metaMDS function from the vegan package in R (v4.4.3)102, to investigate whether the AMR gene diversity of sample clusters based on the species or the source of isolation (clinical and environmental), using Bray-Curtis dissimilarity to achieve higher resolution. The two-dimensional ordinations from both PCA and NMDS were visualised using the ggplot2 package.
Genome annotation and phylogenetic analysis
Draft Aeromonas genomes were annotated using PROKKA (v1.4.5)107 with default parameters. We used Panaroo (v1.3.0)108 to infer the pangenome of 1853 genomes and used the resulting core gene alignment (comprising 1969 core genes) for phylogenetic analysis. We extracted 1,282,613 variable sites using SNP-sites (v2.5.1)109, and the resulting multiple sequence alignment file was used to infer a phylogeny with IQ-TREE (v2.1)110, using a generalised time reversible substitution model with a FreeRate model of heterogeneity111, incorporating invariant sites using the -fconst parameter, and with 1000 UltraFast Bootstraps112. Phylogenetic trees and associated metadata were visualised in iTOL (v.5.6)113. Similarly, a separate phylogenetic tree was constructed for genomes from India, Bangladesh, and Pakistan (n = 1438), consisting of 2067 core genes and 1,197,118 variable sites.
Multi-locus sequence typing (MLST) and subtyping of Aeromonas isolates
We used the MLST tool (v2.9, available at https://github.com/tseemann/mlst) to infer MLST against the Aeromonas scheme developed by Martino and colleagues51 and hosted at https://pubmlst.org/organisms/aeromonas-spp114. This MLST scheme uses six housekeeping genes (gyrB, groL, gltA, metG, ppsA, and recA) to subtype Aeromonas below the species level into Sequence Types (STs). Novel alleles and STs were submitted to the Aeromonas MLST database via pubmlst.org for naming.
Given the broad distribution of genomes across diverse MLST types among species, we also employed Bayesian Analysis of Population Structure (BAPS) using fastBAPS (v1.0.6)115 to define higher-level groupings.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Short-read sequence data (150 bp paired-end reads) are available in the EMBL Nucleotide Sequence Database (European Nucleotide Archive, http://www.ebi.ac.uk/ena); accession numbers are listed in Supplementary Data 1. Additional data supporting the findings of this study are available in the Supplementary Information. Additional data supporting the findings of this study, including R scripts and documented command-line workflows for genomic analyses, are available at Figshare (v4, https://doi.org/10.6084/m9.figshare.30833858).
References
Bastos, R. et al. The genus Aeromonas: a general approach. Micro. Pathog. 130, 81–94 (2019).
Klontz, E. H. et al. Clinical and epidemiologic features of diarrheal disease due to Aeromonas hydrophila and Plesiomonas shigelloides infections compared with those due to Vibrio cholerae Non-O1 and Vibrio parahaemolyticus in Bangladesh. ISRN Microbiol 2012, 654819 (2012).
van Zwetselaar, M. et al. Aeromonas caviae mimicking Vibrio cholerae infectious enteropathy in a cholera-endemic region with possible public health consequences: two case reports. J. Med Case Rep. 12, 71 (2018).
Devos, M. et al. Skin and soft-tissue infections associated with Aeromonas species in French Guiana: an 11-year retrospective study. J. Eur. Acad. Dermatol. Venereol. 34, e414–e416 (2020).
Sadeghi, H., Alizadeh, A., Vafaie, M., Maleki, M. R. & Khoei, S. G. An estimation of global Aeromonas infection prevalence in children with diarrhoea: a systematic review and meta-analysis. BMC Pediatr. 23, 254 (2023).
Kitagawa, H. et al. Aeromonas dhakensis is not a rare cause of Aeromonas bacteremia in Hiroshima, Japan. J. Infect. Chemother. 26, 316–320 (2020).
Yuwono, C., Wehrhahn, M. C., Liu, F. & Zhang, L. Enteric Aeromonas infection: a common enteric bacterial infection with a novel infection pattern detected in an Australian population with gastroenteritis. Microbiol. Spectr. 11, e0028623 (2023).
Parker, J. L. & Shaw, J. G. Aeromonas spp. clinical microbiology and disease. J. Infect. 62, 109–118 (2011).
Ugarte-Torres, A., Perry, S., Franko, A. & Church, D. L. Multidrug-resistant Aeromonas hydrophila causing fatal bilateral necrotizing fasciitis in an immunocompromised patient: a case report. J. Med. Case Rep. 12, 326 (2018).
Qamar, F. N. et al. Aeromonas-associated Diarrhea in children under 5 years: the GEMS experience. Am. Soc. Trop. Med. Hyg. 95, 774–780 (2016).
Igbinosa, I. H., Beshiru, A., Odjadjare, E. E., Ateba, C. N. & Igbinosa, E. O. Pathogenic potentials of Aeromonas species isolated from aquaculture and abattoir environments. Micro. Pathog. 107, 185–192 (2017).
Maldonado-Miranda, J. J., Castillo-Pérez, L. J., Ponce-Hernández, A. & Carranza-Álvarez, C. Summary of economic losses due to bacterial pathogens in aquaculture industry. Bacterial Fish Dis. 2022, 399–417 (2022).
Rasmussen-Ivey, C. R. et al. Classification of a hypervirulent aeromonas hydrophila pathotype responsible for epidemic outbreaks in warm-water fishes. Front. Microbiol. 7, 216817 (2016).
Dalsgaard, I. et al. Identification of atypical Aeromonas salmonicida: Inter-laboratory evaluation and harmonization of methods. J. Appl. Microbiol. 84, 999–1006 (1998).
Park, S. Y., Han, J. E., Kwon, H., Park, S. C. & Kim, J. H. Recent Insights into Aeromonas salmonicida and Its bacteriophages in aquaculture: a Comprehensive Review. J. Microbiol. Biotechnol. 30, 1443 (2020).
Ofek, T., Izhaki, I. & Halpern, M. Aeromonas hydrophila infection in tilapia triggers changes in the microbiota composition of fish internal organs. FEMS Microbiol. Ecol. 99, 1–11 (2023).
Briones, V. et al. Haemorrhagic Septicaemia by Aeromonas salmonicida subsp. Salmonicida in a Black-tip Reef Shark (Carcharhinus melanopterus). J. Vet. Med. Ser. B 45, 443–445 (1998).
Borella, L. et al. Motile aeromonads from farmed and wild freshwater fish in northern Italy: An evaluation of antimicrobial activity and multidrug resistance during 2013 and 2016. Acta Vet. Scand. 62, 1–8 (2020).
Bartie, K. L. & Desbois, A. P. Aeromonas dhakensis: a zoonotic bacterium of increasing importance in aquaculture. Pathogens 13, 465 (2024).
Pessoa, R. B. G., de Oliveira, W. F., Correia, M.T.D.S., Fontes, A. & Coelho, L.C.B.B. Aeromonas and human health disorders: clinical approaches. Front. Microbiol. 13, 868890 (2022).
Rabaan, A. A., Gryllos, I., Tomás, J. M. & Shaw, J. G. Motility and the polar flagellum are required for Aeromonas caviae adherence to HEp-2 cells. Infect. Immun. 69, 4257–4267 (2001).
Gavín, R. et al. Lateral flagella are required for increased cell adherence, invasion and biofilm formation by Aeromonas spp. FEMS Microbiol. Lett. 224, 77–83 (2003).
Kirov, S. M., O’donovan, L. A. & Sanderson, K. Functional characterization of type IV pili expressed on diarrhea-associated isolates of Aeromonas species. Infect. Immun. 67, 5447–5454 (1999).
Byers, B. R., Arceneaux, J. E. L., Barghouthi, S., Massad, G. & Zywno, S. Iron acquisition and microbial virulence: potential uptake systems in the Aeromonas species. Iron Biomineral. 409–415. https://doi.org/10.1007/978-1-4615-3810-3_30 (1991).
Massad, G., Arceneaux, J. E. L. & Byers, B. R. Acquisition of iron from host sources by mesophilic Aeromonas species. J. Gen. Microbiol. 137, 237–241 (1991).
Braun, M. et al. Characterization of an ADP-Ribosyltransferase Toxin (AexT) from Aeromonas salmonicida subsp. salmonicida. J. Bacteriol. 184, 1851 (2002).
Sha, J. et al. The type III secretion system and cytotoxic enterotoxin alter the virulence of Aeromonas hydrophila. Infect. Immun. 73, 6446–6457 (2005).
Burr, S. E., Stuber, K., Wahli, T. & Frey, J. Evidence for a type III secretion system in Aeromonas salmonicida subsp. salmonicida. J. Bacteriol. 184, 5966–5970 (2002).
Swift, S. et al. Quorum sensing in Aeromonas hydrophila and Aeromonas salmonicida: identification of the LuxRI homologs AhyRI and AsaRI and their cognate N-acylhomoserine lactone signal molecules. J. Bacteriol. 179, 5271–5281 (1997).
Kozlova, E. V. et al. Mutation in the S-ribosylhomocysteinase (luxS) gene involved in quorum sensing affects biofilm formation and virulence in a clinical isolate of Aeromonas hydrophila. Micro. Pathog. 45, 343–354 (2008).
Khajanchi, B. K., Kozlova, E. V., Sha, J., Popov, V. L. & Chopra, A. K. The two-component QseBC signalling system regulates in vitro and in vivo virulence of Aeromonas hydrophila. Microbiology 158, 259 (2012).
Fernández-Bravo, A. & Figueras, M. J. An Update on the Genus Aeromonas: taxonomy, epidemiology, and pathogenicity. Microorganisms 8, 129 (2020).
Mehrabadi, J. F., Morsali, P., Nejad, H. R. & Imani Fooladi, A. A. Detection of toxigenic Vibrio cholerae with new multiplex PCR. J. Infect. Public Health 5, 263–267 (2012).
Janda, J. M. & Abbott, S. L. The genus Aeromonas: taxonomy, pathogenicity, and infection. Clin. Microbiol. Rev. 23, 35–73 (2010).
Abbott, S. L., Seli, L. S., Catino, M., Hartley, M. A. & Janda, J. M. Misidentification of unusual Aeromonas species as members of the genus Vibrio: a continuing problem. J. Clin. Microbiol. 36, 1103–1104 (1998).
Lamy, B. et al. Accuracy of 6 commercial systems for identifying clinical Aeromonas isolates. Diagn. Microbiol. Infect. Dis. 67, 9–14 (2010).
Nalbone, L., Forgia, S., Pirrone, F., Giarratana, F. & Panebianco, A. Use of matrix-assisted and laser desorption/ionization time-of-flight technology in the identification of Aeromonas strains isolated from retail Sushi and Sashimi.Pathogens13, 432 (2024).
Nhung, P. H. et al. Use of the novel phylogenetic marker dnaJ and DNA-DNA hybridization to clarify interrelationships within the genus Aeromonas. Int. J. Syst. Evol. Microbiol. 57, 1232–1237 (2007).
Benagli, C. et al. A Rapid MALDI-TOF MS identification database at genospecies level for clinical and environmental Aeromonas strains. PLoS ONE 7, e48441 (2012).
Vávrová, A., Balážová, T., Sedláček, I., Tvrzová, L. & Šedo, O. Evaluation of the MALDI-TOF MS profiling for identification of newly described Aeromonas spp. Folia Microbiol. 60, 375–383 (2015).
Nagar, V., Shashidhar, R. & Bandekar, J. R. Prevalence, characterization, and antimicrobial resistance of Aeromonas strains from various retail food products in Mumbai, India. J. Food Sci. 76, M486-92 (2011).
Bakken, J. S., Sanders, C. C., Clark, R. B. & Hori, M. Beta-lactam resistance in Aeromonas spp. caused by inducible beta-lactamases active against penicillins, cephalosporins, and carbapenems. Antimicrob. Agents Chemother. 32, 1314 (1988).
Chen, P. L., Ko, W. C. & Wu, C. J. Complexity of β-lactamases among clinical Aeromonas isolates and its clinical implications. J. Microbiol. Immunol. Infect. 45, 398–403 (2012).
Lu, J. et al. Whole-genome sequencing-based species classification, multilocus sequence typing, and antibiotic resistance mechanisms of the clinical Aeromonas complex. Front. Microbiol. 16, 1473150 (2025).
Neil, B., Cheney, G. L., Rosenzweig, J. A., Sha, J. & Chopra, A. K. Antimicrobial resistance in aeromonads and new therapies targeting quorum sensing. Appl. Microbiol. Biotechnol. 108, 205 (2024).
Klemm, E. J. et al. Genomic analysis of clinical Aeromonas isolates reveals genetic diversity but little evidence of genetic determinants for diarrhoeal disease. Micro. Genom. 10, 1211 (2024).
Yu, K. et al. The definition and global epidemiology of nonmobile colistin resistance (NMCR-3) determinants in Aeromonas from 1968 to 2022. Drug Resist. Updates 71, 101006 (2023).
Guo, Y. et al. Characterization of NMCR-3, NMCR-4 and NMCR-5, three novel non-mobile colistin resistance determinants: implications for MCR-3, MCR-7, and MCR-5 progenitors, respectively. Drug Resist. Updates 75, 101088 (2024).
George, C. M. et al. Randomized controlled trial of hospital-based hygiene and water treatment intervention (CHoBI7) to reduce Cholera. Emerg. Infect. Dis. 22, 233–241 (2016).
Blackwell, G. A. et al. Exploring bacterial diversity via a curated and searchable snapshot of archived DNA sequences. PLoS Biol. 19, e3001421 (2021).
Martino, M. E. et al. Determination of microbial diversity of Aeromonas strains on the basis of multilocus sequence typing, phenotype, and presence of putative virulence genes. Appl. Environ. Microbiol. 77, 4986–5000 (2011).
Public Databases for Molecular Typing and Microbial Genome Diversity. Reassigned_ST_Aeromonas. https://pubmlst.org/bigsdb?db=pubmlst_aeromonas_isolates&page=query&genomes=1 (2025).
Chopra, A. K. & Houston, C. W. Enterotoxins in Aeromonas-associated gastroenteritis. Microbes Infect. 1, 1129–1137 (1999).
Sha, J., Kozlova, E. V. & Chopra, A. K. Role of various enterotoxins in Aeromonas hydrophila-induced gastroenteritis: generation of enterotoxin gene-deficient mutants and evaluation of their enterotoxic activity. Infect. Immun. 70, 1924 (2002).
Nawaz, M. et al. Detection and characterization of virulence genes and integrons in Aeromonas veronii isolated from catfish. Food Microbiol. 27, 327–331 (2010).
Heuzenroeder, M. W., Wong, C. Y. F. & Flower, R. L. P. Distribution of two hemolytic toxin genes in clinical and environmental isolates of Aeromonas spp.: correlation with virulence in a suckling mouse model. FEMS Microbiol Lett. 174, 131–136 (1999).
Bhowmick, U. D. & Bhattacharjee, S. Bacteriological, clinical and virulence aspects of aeromonas-associated diseases in humans. Pol. J. Microbiol. 67, 137 (2018).
Umelo, E. & Trust, T. J. Identification and molecular characterization of two tandemly located flagellin genes from Aeromonas salmonicida A449. J. Bacteriol. 179, 5292 (1997).
Senderovich, Y. et al. A molecular study on the prevalence and virulence potential of Aeromonas spp. recovered from patients suffering from diarrhea in Israel. PLoS ONE 7, e30070 (2012).
Gavín, R. et al. Lateral flagella of Aeromonas species are essential for epithelial cell adherence and biofilm formation. Mol. Microbiol 43, 383–397 (2002).
Talagrand-Reboul, E., Colston, S. M., Graf, J., Lamy, B. & Jumas-Bilak, E. Comparative and evolutionary genomics of isolates provide insight into the pathoadaptation of Aeromonas. Genome Biol. Evol. 12, 535–552 (2020).
Lee, H. J. et al. Whole genome sequence analysis of Aeromonas spp. isolated from ready-to-eat seafood: antimicrobial resistance and virulence factors. Front. Microbiol. 14, 1175304 (2023).
Truong, N. H. M. et al. Genomic characterization of Aeromonas spp. isolates from striped catfish with motile Aeromonas septicemia and human bloodstream infections in Vietnam. Microb. Genom. 10, 001248 (2024).
Erickson, V. I. et al. Comparative genomic analysis of Aeromonas dhakensis and Aeromonas hydrophila from diseased striped catfish fingerlings cultured in Vietnam. Front. Microbiol. 14, 1254781 (2023).
Shuang, M. et al. Genetic diversity, antimicrobial resistance, and virulence genes of Aeromonas isolates from clinical patients, tap water systems, and food. Biomed. Environ. Sci.33, 385–395 (2020).
Bebak, J., Wagner, B., Burnes, B. & Hanson, T. Farm size, seining practices, and salt use: risk factors for Aeromonas hydrophila outbreaks in farm-raised catfish, Alabama, USA. Prev. Vet. Med. 118, 161–168 (2015).
Zhou, Y. et al. Taxonomy, virulence genes and antimicrobial resistance of Aeromonas isolated from extra-intestinal and intestinal infections. BMC Infect. Dis. 19, 158 (2019).
Drk, S., Puljko, A., Dželalija, M. & Udiković-Kolić, N. Characterization of third generation cephalosporin- and carbapenem-resistant Aeromonas isolates from municipal and hospital wastewater. Antibiotics 12, 513 (2023).
Sakurai, A. et al. Multiplex PCR assay to identify clinically important Aeromonas species. Microbiol. Spectr. 13, e0333124 (2025).
Sakurai, A. et al. Clinical features, genome epidemiology, and antimicrobial resistance profiles of Aeromonas spp. causing human infections: a multicenter prospective cohort study. Open Forum Infect. Dis. 10, ofad587 (2023).
Wu, C. J. et al. Bacteremia due to extended-spectrum-β-lactamase-producing Aeromonas spp. at a medical center in Southern Taiwan. Antimicrob. Agents Chemother. 55, 5813 (2011).
Zhang, Q. et al. Molecular epidemiological characteristics of carbapenem resistant Aeromonas from hospital wastewater. Infect. Drug Resist. 17, 2439 (2024).
Ko, W. C., Wu, H. M., Chang, T. C., Yan, J. J. & Wu, J. J. Inducible β-lactam resistance in Aeromonas hydrophila: therapeutic challenge for antimicrobial therapy. J. Clin. Microbiol. 36, 3188 (1998).
Piotrowska, M. & Popowska, M. Insight into the mobilome of Aeromonas strains. Front. Microbiol. 6, 138626 (2015).
Rhee, J. Y., Jung, D. S. & Peck, K. R. Clinical and therapeutic implications of Aeromonas bacteremia: 14 years nation-wide experiences in Korea. Infect. Chemother. 48, 274 (2016).
Bogaerts, P. et al. OXA-427, a new plasmid-borne carbapenem-hydrolysing class D β-lactamase in Enterobacteriaceae. J. Antimicrob. Chemother. 72, 2469–2477 (2017).
Alksne, L. E. & Rasmussen, B. A. Expression of the AsbA1, OXA-12, and AsbM1 beta-lactamases in Aeromonas jandaei AER 14 is coordinated by a two-component regulon. J. Bacteriol. 179, 2006–2013 (1997).
Estrada, R. & Ruiz, C. First report of a carbapenem-resistant Aeromonas veronii environmental isolate in the United States co-harboring two carbapenemase genes. J. Glob. Antimicrob. Resist. 39, 119–121 (2024).
Dubey, S. et al. Aeromonas species isolated from aquatic organisms, insects, chicken, and humans in India show similar antimicrobial resistance profiles. Front. Microbiol. 13, 1008870 (2022).
Queenan, A. M. & Bush, K. Carbapenemases: the versatile β-Lactamases. Clin. Microbiol. Rev. 20, 440–458 (2007).
McCarley, A., Espejo, M. L., Harmon, D. E. & Ruiz, C. Freshwater and marine environments in California are a reservoir of carbapenem-resistant bacteria. Microorganisms 12, 802 (2024).
Mondal, A. H. et al. A review on colistin resistance: an antibiotic of last resort. Microorganisms 12, 772 (2024).
Shen, Y. et al. Prevalence and genetic analysis of mcr-3-positive Aeromonas species from humans, retail meat, and environmental water samples. Antimicrob. Agents Chemother. 62, e00404–18 (2018).
Ma, S. et al. Mobile colistin resistance gene mcr-5 in porcine Aeromonas hydrophila. J. Antimicrob. Chemother. 73, 1777–1780 (2018).
Gonzalez-Avila, L. U., Loyola-Cruz, M. A., Hernández-Cortez, C., Bello-López, J. M. & Castro-Escarpulli, G. Colistin Resistance in Aeromonas spp. Int. J. Mol. Sci. 22, 5974 (2021).
Manyi-Loh, C., Mamphweli, S., Meyer, E. & Okoh, A. Antibiotic use in agriculture and its consequential resistance in environmental sources: potential public health implications. Molecules 23, 795 (2018).
La Rosa, M. C. et al. The impact of wastewater on antimicrobial resistance: a scoping review of transmission pathways and contributing factors. Antibiotics 14, 131 (2025).
Polianciuc, S. I., Gurzău, A. E., Kiss, B., Ștefan, M. G. & Loghin, F. Antibiotics in the environment: causes and consequences. Med. Pharm. Rep. https://doi.org/10.15386/mpr-1742 (2020).
Saini, A., Kaur, H., Purwar, S., Kholkute, S. D. & Roy, S. Discrepancies in identification of Vibrio cholerae strains as members of Aeromonadaceae and Enterobacteriaceae by automated microbial identification system. Lett. Appl. Microbiol. 55, 22–26 (2012).
Chakraborty, S. et al. Adaptation of a simple dipstick test for detection of Vibrio cholerae O1 and O139 in environmental water. Front. Microbiol. 4, 65995 (2013).
Singhal, N., Kumar, M., Kanaujia, P. K. & Virdi, J. S. MALDI-TOF mass spectrometry: an emerging technology for microbial identification and diagnosis. Front. Microbiol. 6, 791 (2015).
Quail, M. A., Corton, C., Uphill, J., Keane, J. & Gu, Y. Identifying the best PCR enzyme for library amplification in NGS. Micro. Genom. 10, 001228 (2024).
Andrews, S. FASTQC. A quality control tool for high throughput sequence data. https://www.bibsonomy.org/bibtex/f230a919c34360709aa298734d63dca3 (2010).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Wood, D. E. & Salzberg, S. L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, 1–12 (2014).
Lu, J., Breitwieser, F. P., Thielen, P. & Salzberg, S. L. Bracken: estimating species abundance in metagenomics data. PeerJ Comput Sci. 2017, e104 (2017).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. https://doi.org/10.1101/gr.186072.114. (2015).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Jain, C., Rodriguez-R. L.M., Phillippy A. M., Konstantinidis K. T. & Aluru S. High-throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. https://doi.org/10.1101/225342. (2017).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.r-project.org/ (2025).
Alcock, B. P. et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the comprehensive antibiotic resistance database. Nucleic Acids Res. 51, 691 (2023).
Chen, L. VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res. 33, D325–D328 (2004).
Kingombe, C. I. et al. PCR detection, characterization, and distribution of virulence genes in Aeromonas spp. Appl. Environ. Microbiol. 65, 5293–5302 (1999).
Nhinh, D. T. et al. Prevalence, virulence gene distribution and alarming the multidrug resistance of Aeromonas hydrophila associated with disease outbreaks in freshwater aquaculture. Antibiotics 10, 532 (2021).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 1–21 (2020).
Page, A. J. et al. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2, e000056 (2016).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. Modelfinder: fast model selection for accurate phylogenetic estimates. 14, 587–589 (2017).
Hoang, D. T., Chernomor, O., Von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast Bootstrap Approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Jolley, K. A. & Maiden, M. C. J. BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinforma. 11, 1–11 (2010).
Tonkin-Hill, G., Lees, J. A., Bentley, S. D., Frost, S. D. W. & Corander, J. Fast hierarchical Bayesian analysis of population structure. Nucleic Acids Res. 47, 5539–5549 (2019).
Acknowledgements
This research was funded in whole, or in part, by the Wellcome Trust [to NS, ROG, MAB, MJD, AC, NRT: Grant #206194, 220540/Z/20/A; CT, NT, MA, NRT: Grant 215704/Z/19/Z]. This research was also supported by the US National Institutes of Health (Grant 1R01AI3912901) and by USAID (Grant AID-OAA-F-15-00038) [to FTJ, MR, SM, FZ, TP, SIB, MS, MA]. For the purpose of open access, the authors have applied a CC-BY public copyright licence to any author-accepted manuscript version arising from this submission. The authors thank the teams involved in sample collection in Bangladesh and India, as well as the Parasites and Microbes Programme Sequencing and Informatics teams at the Wellcome Sanger Institute. We also thank the National Biodiversity Authority of India. MA acknowledges with gratitude the contribution of collaborating colleagues involved in the grant in Bangladesh and the USA, including the laboratory and field team members of icddr,b. icddr,b acknowledges the donors who provide unrestricted support for its operations and research.
Author information
Authors and Affiliations
Contributions
Conceptualisation: N.S., R.O.G., M.A.B., M.J.D., A.C., A.T.B., N.R.T.; Formal analysis: N.S., R.O.G., M.A.B., N.R.T.; Investigation: N.S., C.T., F.T.J., M.R., S.M., F.Z., T.P., S.I.B., M.S., M.A., N.T., N.R.T.; Resources: M.A., N.T., N.R.T.; Writing and Visualisation: N.S., R.O.G., M.A.B., N.R.T.; Supervision: M.A.B., M.J.D., A.C., A.T.B., S.K., N.T., M.A., N.R.T.; Review and Editing: N.S., R.O.G., C.T., M.A.B., M.J.D., A.C., A.T.B., F.T.J., M.R., S.M., F.Z., T.P., S.I.B., M.S., B.M., D.D., C.M.G., S.K., M.A., N.T., N.R.T. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Izzet Burcin Saticioglu, Changwei Lei, and the other anonymous reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, N., Golicha, R.O., Thakur, C. et al. Aeromonas in South Asia: genomic insights into an environmental pathogen and reservoir of antimicrobial resistance. Nat Commun 17, 2214 (2026). https://doi.org/10.1038/s41467-026-68712-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68712-w