Abstract
DNA damage compromises not only genome stability but also chromatin integrity, which plays a central role in controlling cell identity. Chromatin repair entails the deposition of new histones at sites of DNA damage. How this is coordinated with parental histone dynamics for the maintenance of chromatin states is unknown. To bridge this knowledge gap, here we devise a novel proteomic strategy to characterize dynamic changes in the chromatin landscape during the repair of UV-induced DNA lesions in human cells, in a quantitative, unbiased and time-resolved manner. Thus, we identify the histone chaperones DNAJC9 and MCM2 as central players in chromatin repair. DNAJC9 provides new H3-H4 histones to CAF-1 and HIRA chaperones for deposition into chromatin and stimulates old H3-H4 histone recovery. MCM2 cooperates with DNAJC9 to coordinate old and new histone dynamics during UV damage repair. Together, our proteomic dataset provides a molecular framework for further dissecting epigenome maintenance mechanisms.
Similar content being viewed by others
Introduction
Chromatin is made of DNA wrapped around histone proteins in the cell nucleus and provides epigenetic information that governs cell identity by modulating gene expression1. This pivotal information encompasses multiple layers of regulation, including DNA and histone modifications2,3 as well as histone variants4, which confer specific structural and functional features to the so-called epigenome. Chromatin is in a dynamic state owing to histone deposition and eviction by histone chaperones5, which can be assisted by chromatin remodelers6 in promoting histone dynamics. Histone post-translational modifications further contribute to chromatin plasticity via dedicated factors that deposit, erase, or read these marks2,7.
Epigenome stability is critical for the maintenance of cell function and homeostasis, as underlined by epigenome alterations in disease8,9. Nevertheless, maintaining epigenome stability is a challenge as all DNA metabolic activities entail some level of chromatin reorganization10,11. This challenge is particularly prominent during DNA damage repair because DNA damage can arise in an unscheduled manner at any time, anywhere in the genome12, which poses a major and unpredictable threat to both genome and epigenome stability. As such, chromatin plasticity is an integral part of the DNA damage response, DNA and chromatin repair being tightly coordinated13.
Histone dynamics during DNA repair have been extensively characterized in response to UV damage in mammalian cells and yet our knowledge of these fundamental mechanisms is still incomplete. Chromatin is transiently disrupted at UV damage sites with the eviction of H1 and H2A.Z histones14,15 while H3-H4 histones re-distribute away from damaged sites without dissociating from chromatin16. This local reorganization of chromatin is thought to increase chromatin accessibility, thus facilitating the recruitment of DNA repair factors. DNA repair is accompanied by a restoration of chromatin through the recovery of parental H3-H4 histones16 and the deposition of newly synthesized histones, as shown for H2A, H2A.X, H3.1, and H3.3 histone variants, all deposited de novo by specific histone chaperone complexes that act sequentially during the repair process. Deposition of new H3.3 histones by the histone chaperone complex HIRA occurs at early stages of UV damage repair, coupled to UV damage detection17, while the de novo deposition of H3.1 and H2A variants by the CAF-1 and FACT chaperone complexes, respectively, takes place during late repair steps, coupled to repair synthesis15,18,19. Repaired chromatin is thus a mixture of original and new information, brought about by parental histones that characterize the pre-damage chromatin, and by newly synthesized histones, respectively. Parental and new histone dynamics must be tightly coordinated to ensure the restoration of chromatin states but how this is achieved is a major unresolved issue.
Throughout the repair process, histone chaperones play a central role in controlling histone dynamics. These factors were identified through candidate approaches based on their cognate histone variants, which were shown to accumulate at sites of UV damage repair. We have now reached the limit of those approaches, and yet several important gaps remain in our knowledge of chromatin dynamics during DNA repair. In particular, (i) it is unknown which factors provide new histones to the chaperones identified so far for de novo deposition at repair sites, (ii) the chaperones regulating parental histone redistribution and recovery are uncharacterized, and therefore (iii) how parental and new histone dynamics are coordinated is still an open question. To fill these gaps, we have decided to adopt an unbiased strategy in order to identify new players in chromatin repair in an agnostic manner. We have devised a novel method for the analysis of proteins enriched at sites of UV damage repair in human cells with increased sensitivity and temporal resolution throughout the repair process. This method is based on the isolation of proteins on nascent DNA at repair sites (IPOND-R), combined with their characterization by mass spectrometry (MS)-based proteomics. With this method, we have identified histone chaperones that coordinate new H3-H4 deposition and parental H3-H4 recovery during chromatin repair.
Results
The IPOND-R method captures proteins at sites of DNA damage repair
To profile new chromatin interactors during the repair of DNA damage in human cells, we devised a novel approach based on the established IPOND methodology (Isolation of Proteins On Nascent DNA)20,21, allowing us to specifically identify proteins associated with DNA repair patches in an unbiased, quantitative and time-resolved manner. We called this method IPOND-R for Isolation of Proteins On Nascent DNA at Repair sites (Fig. 1a). In order to capture newly-synthesized DNA associated with DNA repair and not with DNA replication, IPOND-R includes a short-term treatment of cells with replication inhibitors, Hydroxyurea (HU) and Cytosine Arabinoside (Ara-C), for 2 h prior to DNA damage induction by global UVC irradiation. Cells are exposed to 10 J/m2 UVC, which generates UV lesions every 6 kb on average22 and broadly distributed through the genome. Cells are then pulsed with Ethynyl-deoxyUridine (EdU) to label repair patches, crosslinked at different time points after UV irradiation and EdU is biotinylated through a Click-it reaction. After sonication of chromatin, proteins associated with biotinylated nascent DNA are pulled down with streptavidin-coated beads for subsequent proteomic analyses (Fig. 1a). Since there are no major variations in UV damage repair during the cell cycle, this method can be run on asynchronous cell populations.
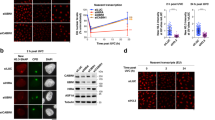
a Scheme of the IPOND-R method. b Inhibition of replication-coupled DNA synthesis measured by EdU incorporation (pulse of 10 min) in U2OS cells treated for 2 h with the replication inhibitor cocktail (Replic. inhib.). c Effect of cell pre-treatment with the replication inhibitor cocktail (Replic. inhib.) on repair-coupled DNA synthesis measured by EdU incorporation (pulse of 1 h) at sites of local UV irradiation (marked by XPA immunostaining) in U2OS cells. S phase cells present in the non-treated population were identified by pan-nuclear EdU staining and excluded from the analysis. d Western blot analysis of proteins captured by IPOND-R 1 h post UV irradiation in U2OS cells. -UV and -Biotin are used as negative controls. Experiment repeated twice with similar results. Dot plots: mean ± SD from n cells scored in 3 independent experiments (the mean of each experiment is shown as a gray triangle). Statistical significance is calculated with a two-sided Student’s t test with Welch’s correction. Scale bars, 10 μm.
We verified that short term cell treatment with the replication inhibitor cocktail was sufficient to erase replicative DNA synthesis (Fig. 1b) while still allowing DNA repair synthesis at sites of UV damage, although with a decrease in efficacy (Fig. 1c). In line with this, we could detect a transient inhibition of gap filling at UV damage sites upon replication inhibition as shown by increased RPA levels 1 h after local UV irradiation returning to normal 2 h post UV (Supplementary Fig. 1a). Cells treated with replication inhibitors also showed signs of replicative stress as illustrated by a transient increase in γH2A.X and phospho-Chk1 signals and an altered cell cycle distribution with an accumulation of cells in G1 24 h post UV (Supplementary Fig. 1b–d). Nevertheless, replication inhibition did not impair the recruitment of the UV damage repair factor XPB (Xeroderma Pigmentosum, complementation group B) to damage sites (Supplementary Fig. 1e), and the kinetics of UV lesion removal (Supplementary Fig. 1f, g). Furthermore, we verified that cell treatment with replication inhibitors allowed efficient and timely recruitment of the histone chaperones CAF-1 and HIRA and subsequent deposition of newly synthesized H3.1 and H3.3 histone variants at UV damage sites (Supplementary Fig. 1h–k).
For EdU biotinylation, we used an enhanced version of biotin-azide conjugate called biotin picolyl azide, which improved in vivo biotinylation efficiency by 4.5-fold as verified by imaging with fluorescently-labeled streptavidin (Supplementary Fig. 2a). Note that DNA breakage arises during the biotinylation reaction due to copper-catalyzed hydrolysis of the DNA21 but as this occurs after cell fixation, it does not introduce any significant bias in the analysis. UV irradiation led to a detectable increase in biotinylated EdU (Supplementary Fig. 2b), thus supporting the sensitivity of the proposed methodology to monitor repair synthesis. Importantly, even though the UV damage repair machinery can excise EdU23, the UV-associated EdU signal was stable for up to 24 h post UVC irradiation in our experimental setting (Supplementary Fig. 2b), arguing against the death of UV-damaged, EdU-labeled cells between 1 h and 24 h post irradiation and allowing us to perform long-term analyses. Damage induction is quite homogeneous in a cell population (Supplementary Fig. 2c) so this does not introduce any bias in subsequent analyses. In addition, we verified that a 1 h EdU pulse immediately after UV irradiation allowed the capture of repair events in heterochromatin regions, known to repair UV lesions more slowly than euchromatin24,25. For this, we employed a validated murine cell line model, NIH/3T3 cells stably expressing the DNA damage sensor DNA damage binding protein 2 (DDB2)14, for studying UV damage repair in DAPI-dense pericentric heterochromatin domains. We observed that the relative levels of EdU in heterochromatin vs. euchromatin were comparable when we labeled early repair events (EdU pulse immediately after UV) or later repair events (EdU pulse 3 h after UV) (Supplementary Fig. 2d), indicating that the IPOND-R analysis is not biased towards repair in euchromatin regions.
As a proof-of-principle, we performed IPOND-R in human U2OS cells and analyzed by western-blot proteins associated with the repair patch 1 h post UV irradiation (Fig. 1d). Core histones, the damage-induced histone modification γH2A.X, PCNA and the histone chaperone CAF-1 were pulled down only in the presence of biotin and enriched in the capture samples from UV-treated cells compared to non-irradiated cells, demonstrating that IPOND-R efficiently and specifically captures chromatin repair patches. Altogether, these results indicate that the IPOND-R method is suited for studying chromatin dynamics specifically at sites of DNA damage repair, in different chromatin contexts and throughout the repair process, from early to late repair steps.
Identification of chromatin regulators enriched at sites of UV damage repair
In order to identify new players in chromatin repair, we combined IPOND-R with mass spectrometry-based quantitative proteomics. We detected 255 proteins that were significantly enriched at repair patches 1 h post UV irradiation in U2OS cells with a +/-UV fold change ≥ 2 and a limma p-value ≤ 0.01 (Fig. 2a, Supplementary Data 1). Among these proteins, we identified known UV damage repair factors (XPC, DDB1, CUL4A, PCNA) as expected, supporting the validity of the IPOND-R method in capturing UV damage repair patches. Much fewer proteins, 12, were retrieved as enriched at repair sites 24 h post UV irradiation and those were mostly core chromatin components (histones and high-mobility group proteins) (Fig. 2b, Supplementary Data 1). Comparing the 1 h and 24 h time points allows to eliminate factors constitutively bound to chromatin in order to focus on those that transiently bind to chromatin during repair (247 1 h-specific hits) (Fig. 2c, Supplementary Data 1). Gene ontology analysis of the 1 h-specific hits revealed an enrichment of biological processes associated with DNA and RNA metabolic activities, the nuclear periphery (nuclear pores, lamins), the immune system (isotype switching and somatic hypermutation), and chromatin organization (Fig. 2d, Supplementary Fig. 3a). We identified 194 proteins that were not found in previous proteomic studies post UV (Supplementary Fig. 3b, Supplementary Data 2)26,27. There was only a modest overlap with the dataset obtained by Stefos et al.26 in human skin fibroblasts 4 h post UV (41 proteins in common), which can be explained by the different cell types (U2OS osteosarcoma cells vs. 1BR3 skin fibroblasts) and different time points post UV (1 h vs. 4 h) as early repair responses cannot be captured 4 h post irradiation. We also compared the 1 h-specific hits identified by IPOND-R to the proteins identified on nascent replicated strands in undamaged conditions by IPOND in different human cell lines28,29. This analysis revealed several shared replication-associated factors including RFC1-5, PCNA, MCM2-7 but also highlighted important differences between the proteomes at replication forks and at sites of UV damage repair with replication-associated factors that were not retrieved at sites of UV damage repair, such as Cdc45, Claspin, MCM10, TONSL (Supplementary Fig. 3c, Supplementary Data 2). We confirmed by western blot that the replication-associated protein Cdc45 was not enriched in the IPOND-R eluate of the +UV condition (Supplementary Fig. 3d). To further study the chromatin regulators identified by IPOND-R, we crossed our dataset with the EpiFactors database (Fig. 2e). Histone modification activities (writers, readers, erasers) and histone chaperones were the most represented, followed by chromatin remodelers and DNA modifiers (Fig. 2e, f). We validated the presence of the chromatin remodeler catalytic subunit SMARCA5 and of the histone modifying enzyme HDAC1 in the +UV capture by IPOND-R coupled to western blot detection (Fig. 2g).
a, b Volcano plots showing mass spectrometry-based quantitative proteomic results of IPOND-R experiments performed at 1 h (a) and 24 h (b) post global UVC in U2OS cells (n= number of independent experiments). Proteins significantly enriched in irradiated compared to non-irradiated samples (+/-UV fold-change (FC) ≥ 2 and limma p-value ≤ 0.01) are shown in green, non-significant proteins are shown in black, and proteins with a negative fold-change (non-specific) are shown in gray. Nucleotide excision repair proteins reaching significance at 1 h are highlighted. c Venn diagram showing enriched proteins at 1 h and 24 h post UV damage with a +/-UV fold-change (FC) ≥ 2 and limma p-value ≤ 0.01. d Gene ontology analysis of the 1 h-specific hits detected by IPOND-R. Biological process terms with a log2 enrichment ≥ 4 and −log10(False Discovery Rate) ≤ 1.3 are shown and grouped based on their association with DNA replication, DNA repair, nuclear periphery, RNA metabolism, cell division, chromatin, immune system, or other (BER: base excision repair, DSB: double-strand break, NER: nucleotide excision repair, SSB: single-strand break). The −log10-False Discovery Rate is represented by the color, and the log2-fold enrichment of the biological processes by the circle size. e Bar plot showing the enrichment of EpiFactor classes in the 1h-specific IPOND-R dataset colored by categories: chromatin remodelers in purple, chromatin modifiers in orange, histone chaperones in red (TF, transcription factors). f Bar plot showing the EpiFactors in the 1 h-specific IPOND-R dataset ranked by +/-UV fold-change (FC) and colored by categories: chromatin remodelers in purple, chromatin modifiers in orange, histone chaperones in red. SMARCA5 and HDAC1, known EpiFactors recruited to UV-damaged chromatin, are highlighted and validated by Western Blot in (g). g Western blot validation of the EpiFactors HDAC1 and SMARCA5 captured by IPOND-R 1 h post UV irradiation in U2OS cells. −UV is used as negative control. Experiment repeated twice with similar results. Statistical testing was conducted with limma (a–c).
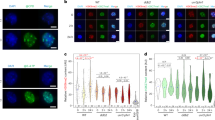
Among histone chaperones, we retrieved most of those known to be involved in chromatin repair following UV damage (Fig. 3a), with the three subunits of the CAF-1 complex (CHAF1A, CHAF1B, RBBP4), which deposits new H3.118, both subunits of the FACT complex (SUPT16H, SSRP1), which promotes new H2A.X deposition and ANP32E, which promotes H2A.Z eviction in UV-damaged chromatin15. In addition, we also detected additional histone chaperones with yet unknown functions in chromatin repair (Fig. 3a), namely Minichromosome Maintenance Complex Component 2 (MCM2) and DnaJ Heat Shock Protein Family Member C9 (DNAJC9). MCM2 was shown to chaperone H3-H4 histones at replication forks30,31 and the heat-shock protein DNAJC932 was recently characterized as an evolutionarily conserved H3-H4 histone chaperone33,34,35. Notably, our optimized IPOND-R methodology permits the unprecedented detection of DNAJC9 (+/-UV fold change = 2.36; limma p-value = 1.53E−3 at 1 h post UV) and MCM2 (+/-UV fold change = 3.07; limma p-value = 6.16E−4 at 1 h post UV) at sites of UV damage repair while these chaperones were not identified in previous proteomic analyses post UV26,27. All the histone chaperones that we identified on UV damaged chromatin were captured at 1 h and not 24 h post UV (Fig. 3a, b), highlighting their dynamic recruitment to chromatin during the repair process. We validated these findings by western blot on the capture material in U2OS cells (Fig. 3c) and in non-cancerous RPE-1 cells (Fig. 3d), which confirmed the transient recruitment of DNAJC9 to repair sites. Western blot analysis of the capture material also enabled the detection of the histone chaperone HIRA (Fig. 3c), which was not identified by mass spectrometry-based proteomics. We further validated DNAJC9 and MCM2 recruitment to sites of UV damage repair through an orthogonal approach that does not rely on using replication inhibitors. For this, we imaged these histone chaperones by immunofluorescence in cells exposed to local UVC irradiation. First, we assessed DNAJC9 enrichment at sites of UVC laser-induced damage in U2OS cells expressing a GFP-tagged form of the UV damage sensor protein DDB2. DNAJC9 showed a distinct and significant enrichment of about 2-fold at UV damage sites (Fig. 3e). We confirmed this result by irradiating cells with a UVC lamp through micropore filters and observed a similar albeit more modest enrichment of DNAJC9 and MCM2 at UV damage sites (1.2-fold, Fig. 3f, g, Supplementary Fig. 4a). The recruitment of DNAJC9 and MCM2 to UV damage sites was observed outside S phase (Supplementary Fig. 4b), pointing to a replication-independent function of these histone chaperones at UV damage sites. Collectively, these findings highlight the potential of IPOND-R in identifying chromatin-associated factors at sites of UV damage repair, including the histone chaperones DNAJC9 and MCM2.
a, b Volcano plots showing mass spectrometry-based quantitative proteomic results of IPOND-R experiments performed at 1 h (a) and 24 h (b) post global UVC in U2OS cells as in Fig. 2. Known histone chaperones reaching significance at 1 h are highlighted. c Western blot detection of some histone chaperones (DNAJC9, MCM2, HIRA and the CAF-1 subunit p60) captured by IPOND-R 1 h post UVC irradiation in U2OS cells. −UV is used as negative control. d Western blot showing the dynamic recruitment of DNAJC9 and CAF-1 p60 to sites of UV damage repair in RPE-1 cells by IPOND-R at 1 h and 24 h post UV damage. −UV is used as negative control. Experiments repeated twice with similar results (c, d). e Immunodetection of DNAJC9 1 h post UVC laser micro-irradiation in U2OS cells stably expressing GFP-DDB2 (UV damage sensor). The enrichment of DNAJC9 at UV damage sites relative to the nucleus is shown on the graph. f Immunodetection of DNAJC9 and MCM2 1 h post local UVC irradiation through micropore filters in U2OS cells. Representative images are shown for DNAJC9. The enrichment of these chaperones at UV damage sites (marked by GFP-DDB2 or CPD) relative to the nucleus is shown on the graphs. Dot plots: mean ± SD from n cells scored in 1–4 independent experiments (the mean of each experiment is shown as a gray triangle). Statistical significance is calculated with a one sample t test compared to a theoretical mean of 1. Scale bar, 10 μm.
The histone chaperone DNAJC9 is not involved in DNA repair following UV damage
To study the function of DNAJC9 and MCM2 in the DNA damage response, we first focused on DNAJC9 as it was the least characterized histone chaperone. We employed siRNA-mediated depletion because DNAJC9 knock-out was reported to be lethal in human cells33,34, and we reached an effective depletion of DNAJC9 of over 50% in U2OS cells (Supplementary Fig. 5a). Since DNAJC9 was recruited early during the DNA damage repair process, we addressed whether this histone chaperone may participate, directly or indirectly, in the repair of DNA damage. However, we did not observe any significant effect of DNAJC9 depletion on the recruitment of the UV-damage repair factor XPB, involved in the unwinding of the DNA double-helix prior to damage excision (Supplementary Fig. 5b), nor on DNA repair synthesis, monitored by EdU incorporation at UV damage sites (Supplementary Fig. 5c). Furthermore, the kinetics of UV photoproduct removal (CPD and 6,4-PP) were also unchanged upon DNAJC9 knock-down while depletion of the repair factor XPG prevented UV lesion removal (Supplementary Fig. 5d). In line with this, DNAJC9 depletion did not impair the survival of cells to UV irradiation (Supplementary Fig. 5e). Knocking down DNAJC9 also did not impair transcription recovery at sites of UV damage as monitored by ethynyl-uridine incorporation into nascent transcripts, whereas depleting the transcription-coupled repair factor CSB (Cockayne Syndrome group B) inhibited transcription recovery, as expected (Supplementary Fig. 5f). Taken together, these results rule out a function for DNAJC9 in the global genome and transcription-coupled repair of DNA damage and prompted us to explore a direct role of this chaperone in histone dynamics at sites of UV damage repair.
DNAJC9 promotes the deposition of newly synthesized H3 variants at sites of UV damage repair
Given the H3-H4 histone binding activity of DNAJC9, we investigated whether this histone chaperone may impact the deposition of newly synthesized H3 histone variants at UV damage sites. We employed the SNAP-tag technology to track new H3.1 and H3.3 histones in U2OS cells stably expressing SNAP-tagged H3 variants36 and monitored their accumulation at sites of local UVC irradiation upon depletion of DNAJC9 (Fig. 4a). Given that H3.1 is also de novo deposited at replication foci37, the incorporation of new H3.1 histones was analyzed outside S-phase to focus on repair-coupled deposition. We can easily discriminate between cells in and outside S phase because the nuclear levels of new H3.1 are significantly higher in S phase (Supplementary Fig. 6a). We observed a marked and reproducible decrease of new H3.1 and H3.3 levels at sites of UV damage upon DNAJC9 depletion (Fig. 4b, c), which we confirmed with a second siRNA targeting DNAJC9 (Supplementary Fig. 6b, c). Exogenous expression of siRNA-resistant DNAJC9 rescued new H3.3 deposition at UV damage sites in DNAJC9-knocked down cells (Supplementary Fig. 6d) which was not observed upon expression of a histone binding mutant form of DNAJC934 although both wild type and mutant DNAJC9 were similarly recruited to damage sites (Fig. 4d). These experiments establish that DNAJC9, through its histone binding domain, promotes the deposition of newly synthesized H3 variants at UV damage sites. We recapitulated these results in non-cancerous RPE-1 cells stably expressing SNAP-tagged H3 variants (Supplementary Fig. 6e, f). Contrary to what observed with H3 histone variants, DNAJC9 depletion did not affect the de novo deposition of the H2A.X variant at UV damage sites (Supplementary Fig. 6g), which is fully consistent with the H3-H4 specificity of this histone chaperone33,34. Importantly, DNAJC9 depletion did not affect the total protein levels of the histone chaperones CAF-1 and HIRA (Supplementary Fig. 6b, c), known to promote the de novo deposition of H3.1 and H3.3 in UV-damaged chromatin (Fig. 4e), and also did not impair the recruitment of these chaperones to UV damage sites (Fig. 4f, g). From these results, we conclude that DNAJC9 promotes new H3 variant deposition at sites of UV damage repair without affecting their cognate histone chaperones CAF-1 and HIRA.
a Protocol for new H3-SNAP histone detection at UV damage sites. b, c, h, i Detection and quantification of newly synthesized H3.1-SNAP (b, h) and H3.3-SNAP (c, i) histones at sites of UVC irradiation (marked by CPD) in U2OS cells stably expressing SNAP-tagged H3 variants and treated with the indicated siRNAs (siLuc, control). The incorporation of new H3.1-SNAP histones at UV sites is quantified in cells outside S-phase, S-phase cells showing a typical focal pattern of new H3.1 deposition throughout the nucleus. The percentages of signal decrease observed in siDNAJC9-treated cells are indicated. d Detection and quantification of newly synthesized H3.3-SNAP histones at sites of UVC irradiation (marked by CPD) in U2OS cells stably expressing SNAP-tagged H3.3 variants and HA-DNAJC9 wild type (WT) or histone binding mutant (4A) and treated with the indicated siRNAs (siLuc, control). New H3.3 deposition at UV sites in cells that do not express HA-DNAJC9 (pcDNA) is also included on the graph as a reference (data from Supplementary Fig. 6d). Nuclear levels of H3.3-SNAP and HA-DNAJC9 and the recruitment of HA-DNAJC9 to UV damage sites are measured in both DNAJC9 wild type and mutant cell lines (right graphs). Nuclear levels of H3.3-SNAP and HA-DNAJC9 are measured in the absence of DNAJC9 knockdown. e Histone chaperones (white) promoting the deposition of newly synthesized H3 variants (red) in UV-damaged chromatin (yellow star, UV lesion). f, g Immunodetection of CAF-1 (f) and HIRA (g) chaperones at sites of UVC irradiation (marked by XPA and XPB, respectively) in U2OS cells treated with the indicated siRNAs (siLuc, control). CAF-1 signal at damage sites is quantified in non-S phase cells. Dot plots: mean ± SD from n cells scored in 1–5 independent experiments (the mean of each experiment is shown as a gray triangle). Statistical significance is calculated with a two-sided Student’s t test with Welch’s correction (b–d, f, g), one sample t test compared to a theoretical mean of 1 (d), or one-way ANOVA with multiple comparisons (d, h, i). Scale bars, 10 μm.
In addition to a role at damage sites, we suspected that DNAJC9 may also promote new histone deposition in undamaged chromatin, during replication in particular, because this histone chaperone sustains cell cycle progression into S phase (Supplementary Fig. 6h), which is consistent with its reported function in cell proliferation33. Similar to what was observed at UV damage sites, DNAJC9 depletion impaired the incorporation of newly synthesized H3.1 histones at replication foci in S phase and also the replication-independent deposition of H3.3 in undamaged conditions (Supplementary Fig. 6a, i). These results highlight a functional role of DNAJC9 in chromatin assembly in vivo via the deposition of new H3.1 and H3.3 histones, both in basal conditions and in response to DNA damage.
We next sought to characterize whether DNAJC9 may function in the same histone deposition pathways as the CAF-1 and HIRA chaperones at repair sites. To do so, we compared single and double knock-downs of DNAJC9 and CAF-1 or HIRA (Supplementary Fig. 6j–m) for their impact on new H3 histone deposition at UV damage sites (Fig. 4h, i). Single depletions of DNAJC9 and CAF-1 similarly impaired the incorporation of new H3.1 histones at damage sites (Fig. 4h) and single depletions of DNAJC9 and HIRA had comparable effects on new H3.3 deposition at UV sites (Fig. 4i). The impact on new H3 variant deposition was partial in each condition, and we did not observe any additive effect upon double depletion of DNAJC9 and the other chaperones (Fig. 4h, i). These results establish that DNAJC9 functions in the same histone deposition pathways as CAF-1 and HIRA to promote the incorporation of new H3.1 and H3.3 histones at sites of UV damage repair. In contrast to CAF-1 and HIRA, DNAJC9 is loosely bound to UV-damaged chromatin, as it is mostly solubilized by detergents (Supplementary Fig. 6n). Together, these findings suggest an upstream function of DNAJC9 in the supply of newly synthesized histones to CAF-1 and HIRA chaperones.
DNAJC9 promotes the recovery of parental H3 histones at sites of UV damage repair
We next investigated whether, besides its activity in promoting new histone deposition at repair sites, DNAJC9 may also control the recovery of old H3-H4 histones that redistribute away from damaged chromatin regions16. The only factor known so far to control parental histone dynamics at UV sites is the UV damage sensor protein DDB2, whose binding to damaged chromatin and subsequent release promote the redistribution and recovery of parental histones, respectively16. To test DNAJC9 contribution to these dynamics, we used the SNAP-tag technology to mark old H3.3 histones in U2OS cells stably expressing H3.3-SNAP, and quantified their levels at UV damage sites compared to the entire nucleus in cells fixed at different time points post irradiation as a proxy for measuring old histone redistribution and recovery (Fig. 5a). The depletion of old H3.3 histones at UV damage sites early after irradiation (10 min) and their subsequent enrichment at late time points (8 h) reflect parental histone redistribution and recovery, respectively (Fig. 5b, siLuc condition). The ratio of old histones at UV damage sites relative to the nucleus 8 h post UV in siLuc cells sets the level of recovery of old histones in control condition. Comparatively, although not completely abrogated, this recovery was impaired upon siRNA-mediated knock-down of the late repair factor XPG, as expected, because UV damage repair is compromised by reduced XPG levels (Fig. 5b, siXPG condition). Importantly, DNAJC9-knocked down cells showed a similar phenotype with an impaired recovery of old histones (Fig. 5b). This phenotype cannot be attributed to defective UV damage repair because these cells were fully proficient for the repair of UV lesions (Supplementary Fig. 5). These results thus demonstrate that DNAJC9 promotes parental H3.3 histone recovery at sites of UV damage repair. To test whether the defective recovery of parental histones may result from impaired DDB2 release from chromatin, we examined DDB2 dynamic binding to UV-damaged chromatin in the absence of DNAJC9 in U2OS cells stably expressing GFP-tagged DDB2. While XPG depletion prevented DDB2 release by inhibiting DNA repair progression, as expected, the depletion of DNAJC9 had no effect on DDB2 release from chromatin (Fig. 5c). Together, these findings demonstrate that the histone chaperone DNAJC9 promotes parental H3 histone recovery at repair sites independently of the UV damage sensor DDB2. Given that DNAJC9 promotes both new histone deposition and old histone recycling at UV-damaged sites, we reasoned that DNAJC9 may control the overall histone density at repair sites. We used the SNAP-tag technology to mark all H3.3-SNAP histones 8 h post irradiation (Fig. 5d) and observed a modest but significant decrease in the total levels of H3.3-SNAP histones at UV sites when DNAJC9 was depleted (Fig. 5e, f). Therefore, DNAJC9 maintains histone density at sites of UV damage repair through new histone deposition and old histone recycling.
a Protocol for old H3.3-SNAP histone detection at UV damage sites. b Detection and quantification of old H3.3-SNAP histone enrichment at sites of UVC irradiation (marked by CPD) relative to the nucleus in U2OS cells stably expressing SNAP-tagged H3.3 and treated with the indicated siRNAs (siLuc, negative control; siXPG, positive control inhibiting old histone recovery). c GFP-DDB2 levels at UVC damage sites (marked by CPD) measured in U2OS GFP-DDB2 cells treated with the indicated siRNAs (siLuc, negative control; siXPG, positive control inhibiting DDB2 release). d Protocol for total H3.3-SNAP histone detection at UV damage sites. e, f Detection and quantification of total H3.3-SNAP histones at sites of UVC irradiation (marked by CPD) in U2OS cells stably expressing SNAP-tagged H3.3 and treated with siDNAJC9#1 (f) and siDNAJC9#3 (g) (siLuc, negative control). Dot plots: mean ± SD from n cells scored in 3 independent experiments. Statistical significance is calculated by one-way ANOVA with multiple comparisons (b, c), a two-sided Student’s t test with Welch’s correction (e, f). Scale bars, 10 μm.
DNAJC9 cooperates with MCM2 to coordinate old and new histone dynamics at sites of UV damage repair
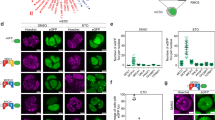
We next investigated how DNAJC9 may promote old histone recovery at repair sites. We envisioned that DNAJC9 could cooperate with the MCM2 histone chaperone in this process because MCM2 is engaged in histone-dependent interactions with DNAJC934, and promotes the recycling of parental H3-H4 histones at replication forks30. We first tested by immunofluorescence in cells exposed to local UVC irradiation whether MCM2 was still enriched on damaged chromatin at the time of old histone recovery. MCM2 enrichment at UV damage sites was clearly detectable 8 h post irradiation, even more pronounced than at 1 h post irradiation (Fig. 6a, siLuc conditions), and we verified that this was not due to bleed-through from the CPD channel (Supplementary Fig. 7a). MCM2 nuclear levels and recruitment to sites of UV damage repair were unaffected by DNAJC9 knock-down and MCM2 depletion did not impair DNAJC9 nuclear levels and enrichment at repair sites (Supplementary Fig. 7b, Fig. 6a, b), arguing that both histone chaperones are independently recruited to UV damage sites. We next tested if MCM2 could function together with DNAJC9 in repair-coupled histone dynamics. Similar to DNAJC9 depletion, MCM2 depletion by siRNA impaired old H3.3 recovery at sites of UV damage repair (Fig. 6c) and the dual knock-down of both chaperones did not show any additive effect (Fig. 6d and Supplementary Fig. 7c). Thus, MCM2 cooperates with DNAJC9 to promote old histone recovery at repair sites. We obtained comparable results when examining new H3.3 deposition at earlier times following UV irradiation, arguing that MCM2 also cooperates with DNAJC9 to stimulate new histone deposition at sites of UV damage repair (Fig. 6e and Supplementary Fig. 7d). This is consistent with the reported interaction of MCM2 with both new and old H3-H4 histones30. Similar to what was observed in cycling cells, DNAJC9 and MCM2 knock down in cells arrested in G1 by palbociclib treatment impaired new H3.3 deposition at sites of UV damage repair (Supplementary Fig. 7e), arguing that the function of these chaperones in new histone deposition coupled to repair does not require passage through S phase. We validated the contribution of MCM2 to new histone deposition at repair sites in functional rescue experiments, showing that exogenous expression of siRNA-resistant MCM2 restored at least partially new H3.3 deposition at UV damage sites in MCM2 knocked down cells (Supplementary Fig. 7f). This is an important finding because it demonstrates the role of MCM2 in new H3.3 deposition at sites of UV damage repair. We also ruled out the hypothesis that MCM2 and DNAJC9 contribution to old H3.3 recovery may be an indirect consequence of these chaperones stimulating new H3.3 deposition. Indeed, depletion of the H3.3 chaperone HIRA, known to deposit new H3.3 histones at sites of UV damage repair (Fig. 4i)17 did not impair old H3.3 recovery (Supplementary Fig. 7g). From these results, we conclude that the DNAJC9 and MCM2 histone chaperones play a central function in chromatin repair by coordinating old and new histone dynamics.
a Immunodetection of MCM2 8 h post local UVC irradiation in U2OS cells (UV damage sites are marked by CPD). The graph shows MCM2 enrichment at UV damage sites relative to the nucleus in U2OS cells treated with the indicated siRNAs (siLuc, control). The first condition (siLuc, 1 h post UVC) is also presented in Fig. 3g. b Immunodetection of DNAJC9 1 h post local UVC irradiation in U2OS cells treated with the indicated siRNAs (siLuc, control). The enrichment of DNAJC9 at UV damage sites (marked by CPD) relative to the nucleus is shown on the graph. c, d Detection and quantification of old H3.3-SNAP histone enrichment at sites of UVC irradiation (marked by CPD) relative to the nucleus in U2OS cells stably expressing SNAP-tagged H3.3 and treated with the indicated siRNAs (siLuc, negative control) at 10 minutes and 8 h post-irradiation (c) or at 8 h only (d). e Detection and quantification of newly synthesized H3.3-SNAP histones at sites of UVC irradiation (marked by CPD) in U2OS cells stably expressing SNAP-tagged H3.3 and treated with the indicated siRNAs (siLuc, control). f Working model. The histone chaperones DNAJC9 and MCM2 are recruited to sites of UV damage repair and jointly coordinate old and new histone H3-H4 dynamics during chromatin repair. Dot plots: mean ± SD from n cells scored in 3 independent experiments. Statistical significance is calculated by one-way ANOVA with multiple comparisons (a, c–e) or with a two-sided Student’s t test with Welch’s correction (b). Scale bars, 10 μm.
Discussion
Preserving chromatin integrity together with genome stability in response to genotoxic stress is key for safeguarding cell identity and protecting cells from pathological development. Yet, our understanding of chromatin repair mechanisms is still very incomplete. Here, we bridge this knowledge gap and identify new players in these mechanisms through the proteomic profiling of repair patches following UV irradiation. Our methodology could be further applied to explore the proteome at other DNA lesions that require DNA synthesis for their repair.
While similar methods for labeling sites of DNA damage repair in non-replicating cells have been employed in recent works26,38, our proteomic approach presents several notable improvements compared to previous proteomic studies in UV-damaged cells26,27: (i) the present approach is focused on repair sites rather than a global analysis of the chromatin-bound proteome post damage infliction, (ii) our method is time-resolved with access to the whole repair process, from early to late repair steps, allowing us to discriminate proteins that dynamically bind to chromatin in response to damage from constitutively bound proteins, and (iii) the increased sensitivity of our methodology allows us to detect low abundant or weakly bound proteins that were not identified in previous datasets.
Among them, we focused our attention on the recently characterized histone chaperone DNAJC933,34. Thus, we uncover a function for DNAJC9 in chromatin assembly in vivo, which was previously characterized only in vitro34. Our data support an upstream function of DNAJC9 in the histone supply chain, DNAJC9 providing new H3-H4 histones to CAF-1 and HIRA chaperones for deposition into chromatin. Whether DNAJC9 directly provides histones to CAF-1 and HIRA is still unclear, as the latter chaperones were not systematically identified as DNAJC9 interactors in basal conditions34,39,40. By ensuring H3-H4 histone supply, DNAJC9 prevents mislocalization of the centromeric H3 variant CENP-A to chromosome arms41. It is thus tempting to speculate that DNAJC9 may similarly prevent ectopic CENP-A deposition at sites of UV damage repair.
In addition to the reported function of DNAJC9 in controlling histone protein folding in the cytoplasm33,34, our work highlights a nuclear function of DNAJC9 in coordinating old and new H3-H4 histone dynamics during the repair process. Indeed, we show that DNAJC9 cooperates with MCM2 to stimulate both new histone deposition and old histone recovery during repair, which constitutes the first proof of a functional cooperation between these two chaperones. Our discoveries not only reveal a function of these histone chaperones in chromatin repair but also instruct us on their activity and central position in the histone supply chain, at the crossroads of new histone deposition and old histone recycling (Fig. 6f). The mechanism whereby DNAJC9 and MCM2 stimulate parental histone recovery during UV damage repair warrants further investigation.
MCM2 is enriched in the IPOND-R dataset together with the other MCM subunits (MCM3 to 7), showing that the whole MCM hexamer is present at sites of UV damage repair, like at replication forks. However, the observed recruitment of MCM2 to sites of UV damage repair does not require the passage of replication forks because it is detected outside of S phase, highlighting a function of MCM2 that is disconnected from DNA replication. In line with this, we do not retrieve other components of the replicative helicase complex at repair sites like Cdc45 and GINS subunits, which indicates that MCM2 likely operates independently of the replicative helicase complex at repair sites. Notably, an abundant pool of MCM proteins is not engaged with active replisomes in human cells42 and those could fulfill replication-independent roles.
Several pieces of evidence also suggest that DNAJC9 may coordinate new and parental histone dynamics in contexts other than DNA repair. Indeed, DNAJC9 is found enriched on nascent replicated chromatin28 and we show that DNAJC9 stimulates new H3.1 deposition at undamaged replication forks. A potential role of DNAJC9 in parental histone recycling at replication forks, possibly in coordination with MCM230, deserves to be investigated.
How MCM2 and DNAJC9 are recruited to sites of UV damage repair is not yet elucidated. It is tempting to speculate that their recruitment may be mediated by direct or indirect interactions with UV damage repair factors or with other histone chaperones enriched at repair sites. In this respect, MCM2 is involved in histone-dependent interactions with the FACT histone chaperone39 that is recruited to UV sites coupled to repair synthesis15. An additional candidate that could promote DNAJC9 recruitment to UV-damaged chromatin is RAD23A, which was identified in the DNAJC9 interactome in human cells33 and binds the early UV damage repair factor XPC43,44. UV-damage responsive post-translational modifications of these histone chaperones may also regulate their function in chromatin repair27. Our proteomic approach does not distinguish between repair in euchromatin and heterochromatin. However, we envision that histone chaperones may play distinct roles in different chromatin compartments. The use of murine cellular models suited for the analysis of heterochromatin repair of UV lesions14 will be instrumental to address this issue.
We do not rule out the possibility that DNAJC9 and MCM2 may respond to other types of DNA lesions than UV photoproducts, and thus play a more general role in maintaining epigenome stability following genotoxic stress. Interestingly, a fungal ortholog of DNAJC9, Dnaj4, which also exhibits H3-H4 histone chaperone activity, was shown to safeguard genome integrity at least in part through the control of gene expression in response to DNA damage45. Together with our findings, this hints towards an evolutionarily conserved function of DNAJC9 in genome and epigenome maintenance. DNAJC9 is a dual heat shock molecular and histone chaperone that recruits heat shock proteins of the HSP70 family via its J domain to promote histone protein folding33,34. Whether DNAJC9 functions in chromatin repair depend on binding to HSP70 family proteins remains to be investigated, which could be tested using separation of function mutants in the J domain and the histone binding domain of DNAJC934. Notably, HSPA4/HSP74 is significantly enriched at sites of UV damage repair in the 1 h IPOND-R dataset and thus represents a potential candidate that could cooperate with DNAJC9 in controlling histone dynamics during repair.
While in this study we focused on the histone chaperones DNAJC9 and MCM2, our proteomic analysis uncovered a number of additional interesting candidates as potential new players in chromatin dynamics during DNA repair. Among them, several histone modifiers and readers highlight a fine-tuned regulation of histone modifications at repair sites, which deserves to be investigated in future studies. The histone modifiers and readers identified in our dataset and by others26,27 will provide clues to the underlying mechanisms. Along the same lines, the DNA modifiers enriched at sites of UV damage repair will help characterize the machinery for repair-coupled DNA methylation maintenance, thus broadening our understanding of epigenome maintenance mechanisms in response to DNA damage. Intriguingly, several nuclear pore components and lamins were also enriched at sites of UV damage repair, as seen in a previous study26. There is growing evidence for a role of the nuclear periphery in the repair of DNA double-strand breaks, with DSBs relocalizing to the nuclear periphery for repair in yeast and flies, while in human cells, the nuclear envelope forms intra-nuclear tubules that capture DSBs for repair, and nuclear pores and lamina-associated chromatin domains regulate DSB repair pathway choice46,47,48,49,50. However, the connection of UV damage repair with the nuclear periphery is still elusive and will be interesting to explore to further dig into how nuclear architecture may govern genome maintenance and reciprocally. Overall, our proteomic dataset provides a molecular foundation for further exploration of the fundamental mechanisms underpinning chromatin plasticity and integrity following genotoxic stress.
Methods
Cell culture
U2OS (ATCC HTB-96, human osteosarcoma, female), RPE-1 hTERT (ATCC CRL-4000, human retinal pigment epithelial cell, female), and NIH/3T3 cells (ATCC CRL-1658, mouse embryonic fibroblast, male) were grown at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen) supplemented with 10% fetal bovine serum (EUROBIO) and antibiotics (100 U/ml penicillin and 100 µg/ml streptomycin, Invitrogen) and the appropriate selection antibiotics for stable cell lines (Euromedex, Table 1). For seeding NIH/3T3 cells on coverslips, coverslips were first coated with 20 µg/ml Collagen Type I (MERCK Millipore) and 2 µg/ml fibronectin (Sigma-Aldrich) to increase cell adhesion.
In order to inhibit DNA replication, we use 10 mM Hydroxyurea (HU, Sigma-Aldrich) and 1 mM cytosine arabinoside (Ara-C, Sigma-Aldrich) in fresh growth medium for 2 h prior to DNA damage induction. To prevent S-phase entry, we use the cdk4/6 inhibitor palbociclib (Selleck Chemicals) at 150 nM final for 48 h. Palbociclib is applied to RPE-1 cells only because U2OS cells are resistant due to high cdk4 expression.
All cell lines generated in this study were authenticated by verifying the expression of the transgenes by immunofluorescence and western blot. They are available for reuse and can be provided upon request.
siRNA and plasmid transfections
siRNA purchased from Eurofins MWG Operon, Sigma Aldrich and Dharmacon (Table 2) were transfected into cells using Lipofectamine RNAiMAX (Invitrogen) following the manufacturer’s instructions. The final concentration of siRNA in the culture medium was 50 nM. Cells were harvested 48-72 h after transfection.
Cells were transfected with plasmid DNA (Table 3) using Lipofectamine 2000 (Invitrogen) or by nucleofection with the Neon™ Transfection System (Thermo Fisher Scientific) according to the manufacturer’s instructions. For stable cell line establishment (Table 1), plasmid DNA was transfected into cells at 1 µg/ml final, 48 h before antibiotic selection of clones. For rescue experiments, cells were co-transfected with plasmid DNA (3 µg/ml final) and siRNA (50 nM final), using the NeonTM Transfection System (Thermo Scientific) according to the manufacturer’s instructions and harvested 72 h later.
All plasmids generated in this study are available for reuse and can be provided upon request.
UVC irradiation
Cells grown on glass coverslips (12 mm diameter, thickness No. 1.5, Thorlabs) were irradiated with UVC (254 nm) using a low-pressure mercury lamp. Conditions were set using a VLX-3W dosimeter (Vilber Lourmat). For global UVC irradiation, cells in Phosphate Buffer Saline (PBS) were exposed to UVC doses ranging from 4 to 12 J/m2 for survival assays and to 10 J/m2 in other experiments. For local UVC irradiation51,52, cells were covered with a polycarbonate filter (5 μm pore size, Millipore) and irradiated with 150, 300 or 500 J/m2 UVC. Irradiated cells were allowed to recover in culture medium for the indicated times before fixation.
For UVC laser micro-irradiation53, cells were grown on quartz coverslips (25 mm diameter, thickness No. 1, SPI supplies) and nuclei were stained by adding Hoechst 33258 (10 μg/mL final, Sigma-Aldrich) to the culture medium 30 min before UVC irradiation. Quartz coverslips were transferred to a Chamlide magnetic chamber (Gataca-systems) on a custom stage insert (Live Cell Instrument) and cells were irradiated for 100 ms using a 2 mW pulsed diode-pumped solid-state laser emitting at 266 nm (RappOptoElectronics, Hamburg GmbH) directly connected to a Zeiss LSM900 confocal microscope adapted for UVC transmission with all-quartz optics. The laser was attenuated using a 0.1% neutral density filter and focused through a 40x/0.6 Ultrafluar glycerol objective with quartz lenses. The laser is controlled by a UGA Firefly module with SysCon2 software (RappOpto). The laser impact has an average size of 1 μm in diameter and damages around 1-2% of the total nuclear volume. The corresponding UVC dose, not directly measurable but estimated by comparing the intensity of the CPD damage generated by the laser and the UVC lamp, is 800 J/m2.
IPOND-R (Isolation of Proteins On Nascent DNA at Repair sites)
EdU labeling of repair sites
Cells are seeded in 15-cm dishes to harvest at least 70 million cells per condition on the day of the experiment. In order to label only repair patches, replication is inhibited with 10 mM HU and 1 mM Ara-C (Sigma-Aldrich) in fresh growth medium for 2 h at 37 °C prior to global UVC irradiation (10 J/m2) in PBS supplemented with 10 mM HU and 1 mM Ara-C to maintain the inhibition of replication. A non-irradiated sample is used as negative control. After irradiation, cells are incubated in fresh medium containing the replication inhibitor cocktail and 10 µM Ethynyl-deoxyUridine (EdU, Euromedex) for 1 h at 37 °C. At this step, a chase can be performed in fresh medium supplemented with 10 µM Thymidine (Sigma-Aldrich).
Cell fixation and permeabilization
Cells are fixed with 1% Formaldehyde (Sigma-Aldrich) for 15 minutes on a rocking platform. The fixation is stopped by adding 1.33 M Glycine (Sigma-Aldrich) for 5 minutes on the rocking platform. Cells are washed twice in cold PBS, scrapped in 1% BSA solution in PBS and pelleted for 5 min at 300 g at 4 °C. Cell pellets are permeabilized with 1% Triton X-100 (1 mL/ 10 million cells, Euromedex) for 30 min at room temperature and washed twice with cold 1% BSA solution.
EdU biotinylation by Click-it chemistry
Cell pellets are resuspended in a Click-it reaction cocktail containing 2 mM CuSO4 (Sigma-Aldrich), 1 mM Tris((1-hydroxy-propyl-1H-1,2,3-triazol-4-yl)methyl)amine (THPTA, Euromedex), 10 mM Sodium Ascorbate (Sigma-Aldrich), 5 mM Amino-guanidine hydrochloride (Sigma-Aldrich), and 10 μM Biotin Picolyl azide or Biotin azide (Sigma-Aldrich). Cell suspensions are left on a rotating wheel at room temperature for 1 h. Cells are then washed twice in cold 1% BSA solution containing 0.5X EDTA-free Protease inhibitor cocktail (PIC, Roche), and once in cold PBS. A negative control without biotin reagent was performed to check for the specificity of the Streptavidin pull-down.
Cell lysis and sonication
Cell pellets are resuspended in lysis buffer (200 µl per 15 million cells) containing 1% SDS, 50 mM Tris pH 7.5, and 1X PIC. The cell suspensions are transferred to 1.5 ml sonication tubes (Diagenode) and sonication is performed using a Bioruptor Pico (Diagenode) with 5-10 cycles of 30 sec ON and 30 sec OFF. The sonicated samples are transferred to new Eppendorf tubes and centrifuged 10 min at 16,000 g at 4 °C. The supernatants are transferred to Low Protein Binding tubes (Thermo Fisher Scientific).
Analysis of DNA shearing
A 20 μl aliquot of each cell lysate is incubated with 200 mM NaCl and 200 μg/ml RNAse A (Millipore) overnight at 65 °C. After addition of 100 μg/ml Proteinase K (Sigma-Aldrich), samples are incubated for a further 2 h at 45 °C prior to DNA purification using the PCR purification kit from Macherey-Naegel following the manufacturer’s instructions. DNA samples are analyzed with a TapeStation (Agilent) to check that the average fragment size is about 500 bp.
Streptavidin capture of biotin-labeled nascent DNA and associated proteins
The remaining solution is resuspended in an equal volume of cold PBS supplemented with 1 X PIC to dilute the SDS present in the sonication buffer. At this stage, input samples are collected, diluted in Laemmli buffer (50 mM Tris-HCl pH 6.8, 1.6% Sodium Dodecyl Sulfate (SDS), 8% glycerol, 4% β-mercaptoethanol, 0.0025% bromophenol blue) and decrosslinked by heating for 5 min at 95 °C.
For streptavidin capture, Dynabeads MyOne Streptavidin-C1 beads (Invitrogen) are washed in lysis buffer and in PBS supplemented with 1X PIC and then are added to cell lysates (1 µl beads per 1 million cells) and incubated overnight at 4 °C on a rotating wheel. After capture, beads are washed once in lysis buffer (1 ml wash solution per 10 million cells), once in 1 M NaCl supplemented with 1X PIC, followed by 3 additional washes in lysis buffer.
For western blot analysis, the beads are resuspended in Laemmli buffer and decrosslinking is achieved by heating for 5 min at 95 °C.
For mass spectrometry analysis, beads are transferred to new Low Protein binding tubes and washed in 50 mM Tris-HCl pH 8 before resuspension in Laemmli buffer and decrosslinking by heating for 5 min at 95 °C. Ten percent of the protein solution is kept for a Silver staining analysis.
Mass spectrometry-based proteomic analyses
Proteins from three and four IPOND-R biological replicates, purified 1 h and 24 h after UV irradiation, respectively (-UV, negative control) were solubilized in Laemmli buffer (50 mM Tris-HCl pH 6.8, 1.6% Sodium Dodecyl Sulfate (SDS), 8% glycerol, 4% β-mercaptoethanol, 0.0025% bromophenol blue) and separated by SDS-PAGE (Mini Protean TGX, 4-20%, Biorad), before staining with Coomassie blue R-250 (Bio Basic Canada Inc.). The band corresponding to streptavidin was discarded and the remaining proteins were digested in-gel using modified trypsin (Promega, sequencing grade) as previously described54. The resulting peptides were analyzed by online nanoliquid chromatography coupled to MS/MS (Ultimate 3000 RSLCnano and Q-Exactive HF, Thermo Fisher Scientific, for the 1 h dataset, and Ultimate 3000 RSLCnano and Orbitrap Exploris 480, Thermo Fisher Scientific, for the 24 h dataset) using a 120-min gradient. For this purpose, the peptides were sampled on a precolumn (300 μm × 5 mm PepMap C18, Thermo Scientific) and separated in a 75 μm × 250 mm C18 column (Reprosil-Pur 120 C18-AQ, 1.9 μm, Dr. Maisch, for the 1 h dataset, and Aurora Generation 2, 1.6 µm, IonOpticks, for the 24 h dataset). The MS and MS/MS data were acquired using Xcalibur (Thermo Fisher Scientific).
Peptides and proteins were identified by Mascot (version 2.8.0, Matrix Science) through concomitant searches against the Uniprot database (20230719 download, 207’981 sequences) and a homemade database containing the sequences of classical contaminant proteins found in proteomic analyses (human keratins, trypsin…, 250 sequences). Trypsin/P was chosen as the enzyme and two missed cleavages were allowed. Precursor and fragment mass error tolerances were set at respectively at 10 and 20 ppm. Peptide modifications allowed during the search were: Carbamidomethyl (C, fixed), Acetyl (Protein N-term, variable) and Oxidation (M, variable). The Proline software55 version 2.2.0 was used for the compilation, grouping, and filtering of the results (conservation of rank 1 peptides, peptide length ≥ 6 amino acids, false discovery rate of peptide-spectrum-match identifications <1%56, and a minimum of one specific peptide per identified protein group). MS data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository57 with the dataset identifier PXD055129. Proline was then used to perform an MS1 label-free quantification of the identified protein groups based on razor and specific peptides.
Statistical analysis was performed using the ProStaR software58 based on the quantitative data obtained with the different biological replicates analyzed for each condition. Proteins identified in the contaminant database, proteins identified by MS/MS in less than two replicates of one condition, and proteins quantified in less than all replicates of one condition were discarded. After log2 transformation, abundance values were normalized using the variance stabilizing normalization (vsn) method applied condition-wise, before missing value imputation (SLSA algorithm for partially observed values in the condition and DetQuantile algorithm for totally absent values in the condition). Statistical testing was conducted with limma, whereby differentially expressed proteins were selected using a log2 (Fold Change) cut-off of 1 and a p-value cut-off of 0.01, allowing to reach a false discovery rate inferior to 3% according to the Benjamini-Hochberg estimator. Proteins found differentially abundant but identified by MS/MS in less than two replicates, and detected in less than all replicates, in the condition in which they were found to be more abundant were invalidated (p-value = 1).
Data visualization and gene ontology analysis
Biological process gene ontology terms were extracted from the PANTHER database (v.18 2023-08-01 release) and analyzed through the PANTHER Overrepresentation Test (released 2023-07-05). Significance was calculated using a Fisher test with a false-discovery rate correction. The list of EpiFactors was extracted from https://epifactors.autosome.ru/public_data version 2.059,60 curated based on most recent literature. The proteins detected by IPOND-R coupled to mass spectrometry were visualized using Volcano plots generated with R studio 2021.09.0 using ggplot2 version 3.3.5, ggrepel version 0.9.1, and dplyr version 1.1.2. Venn diagrams were generated with eulerr version 6.1. Statistical analyses and all other graphs were generated using GraphPad Prism version 8.4.3, and test details and p-values are referred to in the figure legends.
EdU-labeling of replicating cells and repair sites
To visualize replication foci, 10 µM Ethynyl-deoxyUridine (EdU) was incorporated into cells on glass coverslips during 15 min at 37 °C and revealed using the Click-It EdU Alexa Fluor 488 or 594 Imaging kit (Invitrogen) according to the manufacturer’s instructions. To localize the sites of UV damage repair, cells were incubated with 10 µM EdU for 1 h after local UVC irradiation and EdU was revealed using the Click-It EdU Alexa Fluor 488, 594 or 647 Imaging kit (Invitrogen).
EU-labeling of nascent transcripts at UV damage sites
Cells were exposed to local UVC irradiation (300 J/m2) 48 h post siRNA transfection and allowed to recover for 2 h (to monitor transcription inhibition at UV sites) or 24 h (to monitor transcription recovery). For nascent transcript labeling, cells were incubated in DMEM supplemented with 0.5 mM ethynyl-uridine (EU) for 45 min at 37 °C, and rinsed in PBS before fixation in 2% paraformaldehyde (Electron Microscopy Sciences). EU incorporation was revealed with Click-iT RNA Imaging kit (Thermo-Fisher Scientific) using AlexaFluor 594 dye according to the manufacturer’s instructions. Cells were post-fixed in 2% paraformaldehyde before CPD immunostaining with AlexaFluor 488 dye as described in the immunofluorescence section below. Coverslips were mounted in Vectashield medium with DAPI (Vector laboratories). The mean EU fluorescence intensity was measured at UV damage sites (delineated based on CPD staining) relative to the entire nucleus (delineated based on DAPI staining).
Immunofluorescence
Cells grown on coverslips were either fixed directly with 2% paraformaldehyde (Electron Microscopy Sciences) for 10 min and permeabilised for 5 min with 0.5% Triton X-100 in PBS or cells were pre-extracted before fixation with 0.5% Triton X-100 in CSK buffer (Cytoskeletal buffer: 10 mM PIPES pH 7.0, 100 mM NaCl, 300 mM sucrose, 3 mM MgCl2) for 5 min at room temperature to remove soluble proteins. For the detection of UVC photoproducts, the DNA was denatured with 2 N HCl for 10 min at 37 °C (6,4-PP detection) or with 0.5 M NaOH for 5 min at room temperature (CPD detection). Since this denaturation quenches GFP fluorescence, when CPD detection was combined with the visualization of GFP-DDB2, immunofluorescence was performed with GFP immunodetection using a rat anti-GFP antibody (Supplementary Table 1) combined with CPD immunodetection. Samples were blocked for 10 min in 5% BSA (Bovine Serum Albumin, Sigma-Aldrich) in PBT (PBS 0.1% Tween-20), followed by 45 min incubation with primary antibodies and 30 min incubation with secondary antibodies coupled to AlexaFluor dyes (Supplementary Tables 1 and 2) diluted in blocking buffer. Coverslips were mounted in Vectashield medium with DAPI (Vector laboratories).
SNAP-tag labeling of histones
For specific labeling of newly synthesized histones36,61, cells were grown on glass coverslips and pre-existing SNAP-tagged histones were first quenched by incubating cells with 10 μM of the non-fluorescent substrate SNAP-cell Block (New England Biolabs) for 30 min followed by a 30 min-wash in fresh medium and a 2 h (U2OS) or 4 h (RPE-1) chase. The new SNAP-tagged histones synthesized during the chase were fluorescently labeled with 2 μM of the red-fluorescent reagent SNAP-cell TMR star (New England Biolabs) during a 15 min-pulse step followed by 30 min wash in fresh medium. Cells were subsequently permeabilised with Triton X-100, fixed and processed for immunostaining. Cells were irradiated with a UVC lamp before the pulse step.
For specific labeling old histones, cells were grown on glass coverslips and pre-existing SNAP-tagged histones were fluorescently labeled with 2 μM of the red-fluorescent reagent SNAP-cell TMR star (New England Biolabs) during a 30 min-pulse step followed by 30 min wash in fresh medium. Cells were irradiated with a UVC lamp 48 h after the pulse step. Cells were subsequently permeabilised with Triton X-100, fixed and processed for immunostaining.
For labeling all histones, cells were grown on glass coverslips and SNAP-tagged histones were fluorescently labeled with 2 μM of the red-fluorescent reagent SNAP-cell TMR star (New England Biolabs) during a 30 min-pulse step. Cells were subsequently permeabilised with Triton X-100, fixed and processed for immunostaining. Cells were irradiated with a UVC lamp before the pulse step.
Image acquisition and analysis
Fluorescence imaging was performed with a Leica DMI6000 epifluorescence microscope using a Plan-Apochromat 40x/1.3 oil objective. Images were captured using a CCD camera (Photometrics) and Metamorph software (version 7.10.2.240). Images were assembled with Adobe Photoshop. Live cell imaging coupled to UVC laser micro-irradiation was performed using a 40x/0.6 Ultrafluar Glycerol objective on a Zeiss LSM900 confocal microscope. Images were captured using Zen blue software version 3.5, and analyzed with ImageJ v.2.0.0-rc-69/1.52.p (U. S. National Institutes of Health, Bethesda, Maryland, USA, http://imagej.nih.gov/ij/). The subtract background function was applied to all images prior to quantification, and images were smoothed using the median filter function. Nuclei and heterochromatin domains were segmented based on DAPI or Hoechst staining, and UVC-damaged regions based on GFP-DDB2 fluorescence or immunostaining for UV damage or repair factors using the default threshold function with manual adjustment. Mean intensities were measured in each region of interest using the measure and set measurement functions. When calculating the ratios of histone signals at UV sites relative to the nucleus, a correction factor of 1.19 was applied to nuclear histone signals to take into account lower histone content in nucleoli as measured in independent experiments with histone and nucleolin co-staining.
Cell extracts and Western blot
Total extracts were obtained by scraping cells on plates or resuspending cell pellets in Laemmli buffer (50 mM Tris-HCl pH 6.8, 1.6% Sodium Dodecyl Sulfate (SDS), 8% glycerol, 4% β-mercaptoethanol, 0.0025% bromophenol blue) followed by 5 min denaturation at 95 °C.
For western blot analysis, extracts were run on 4%–20% Mini-PROTEAN TGX gels (Bio-Rad) in running buffer (200 mM glycine, 25 mM Tris, 0.1% SDS). Proteins were transferred onto nitrocellulose membranes (0.2 μm, Amersham Protran) for 30 min at 15 V with a Trans-Blot SD semidry transfer cell (Bio-Rad) in transfer buffer (50 mM Tris, 40 mM glycine, 0.04% SDS, 15% ethanol). Total proteins were revealed by Reversible Stain (Pierce). Proteins of interest were probed using the appropriate primary and Horse Radish Peroxidase (HRP)-conjugated secondary antibodies (Supplementary Tables 1 and 2), detected using SuperSignal West Pico or Femto chemiluminescence substrates (Pierce) on hyperfilms MP (Amersham) or with an Odyssey Fc-imager (LI-COR Biosciences). When fluorescence detection was used instead of chemi-luminescence, secondary antibodies were conjugated to IRDye 680RD or 800CW (Supplementary Table 2), membranes were scanned with the Odyssey Fc-imager (LI-COR Biosciences) and analyzed with Image Studio Lite software (version 5.2.5) using total protein stain for normalization.
Flow cytometry
For cell cycle analysis, cells were fixed in ice-cold 70% ethanol before DNA staining with 50 ug/ml propidium iodide (Sigma-Aldrich) in PBS containing 0.05% Tween and 0.5 mg/ml RNase A (USB/Affymetrix). DNA content was analyzed by flow cytometry using a BD FACScalibur flow cytometer equipped with CellQuest Pro version 6.0 (BD Biosciences) and FlowJo software version 9.7.5 (TreeStar).
Cell survival assays
Cells were replated 48 h after siRNA transfection and exposed to global UVC irradiation (4, 8, and 12 J/m2) the following day. Colonies were stained 12 days later with 0.5% crystal violet/20% ethanol and counted. Results were normalized to plating efficiencies.
Statistical analyses
Percentages of positively stained cells were obtained by scoring at least 150 cells in each experiment. Statistical tests were performed using GraphPad Prism (versions 8 to 10). We run the statistical analyses on the means of the biological replicates in conditions of global UV irradiation, which leads to homogeneous damage induction in the cell population, while analyses are performed on all data points and not on the means of the biological replicates in conditions of local UV irradiation because in this case each nucleus receives a different UV dose, which depends on the pore distribution in the micropore filters used for local UVC irradiation, so each nucleus should be considered independently. P-values for mean comparisons between two groups were calculated with a Student’s t test with Welch’s correction when necessary. Multiple comparisons were performed by one-way or two-way ANOVA. Comparisons of clonogenic survival were based on non-linear regression with a polynomial quadratic model. ns: non-significant, *: p < 0.05, **: p < 0.01, ***: p < 0.001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated during this study are included in this article and its supplementary information files. Mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD055129. Source data have been deposited in the Figshare Repository (https://doi.org/10.6084/m9.figshare.30665291). Source data are provided with this paper.
References
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Millán-Zambrano, G., Burton, A., Bannister, A. J. & Schneider, R. Histone post-translational modifications — cause and consequence of genome function. Nat. Rev. Genet. 23, 563–580 (2022).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Martire, S. & Banaszynski, L. A. The roles of histone variants in fine-tuning chromatin organization and function. Nat. Rev. Mol. Cell Biol. 21, 522–541 (2020).
Hammond, C. M., Strømme, C. B., Huang, H., Patel, D. J. & Groth, A. Histone chaperone networks shaping chromatin function. Nat. Rev. Mol. Cell Biol. 18, 141–158 (2017).
Clapier, C. R., Iwasa, J., Cairns, B. R. & Peterson, C. L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 18, 407–422 (2017).
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Flavahan, W. A., Gaskell, E. & Bernstein, B. E. Epigenetic plasticity and the hallmarks of cancer. Science 357 (2017).
Brookes, E. & Shi, Y. Diverse epigenetic mechanisms of human disease. Annu. Rev. Genet. 48, 1–32 (2014).
Ferrand, J., Plessier, A. & Polo, S. E. Control of the chromatin response to DNA damage: histone proteins pull the strings. Semin. Cell Dev. Biol. 113, 75–87 (2021).
Stewart-Morgan, K. R., Petryk, N. & Groth, A. Chromatin replication and epigenetic cell memory. Nat. Cell Biol. 22, 361–371 (2020).
Hoeijmakers, J. H. J. DNA damage, aging, and cancer. N. Engl. J. Med. 361, 1475–1485 (2009).
Dabin, J., Mori, M. & Polo, S. E. The DNA damage response in the chromatin context: a coordinated process. Curr. Opin. Cell Biol. 82, 102176 (2023).
Fortuny, A. et al. Imaging the response to DNA damage in heterochromatin domains reveals core principles of heterochromatin maintenance. Nat. Commun. 12, 2428 (2021).
Piquet, S. et al. The histone chaperone FACT coordinates H2A.X-dependent signaling and repair of DNA damage. Mol. Cell 72, 888–901.e7 (2018).
Adam, S. et al. Real-time tracking of parental histones reveals their contribution to chromatin integrity following DNA damage. Mol. Cell 64, 65–78 (2016).
Adam, S., Polo, S. E. & Almouzni, G. Transcription recovery after DNA damage requires chromatin priming by the H3.3 histone chaperone HIRA. Cell 155, 963 (2013).
Polo, S. E., Roche, D. & Almouzni, G. New histone incorporation marks sites of UV repair in human cells. Cell 127, 481–493 (2006).
Dinant, C. et al. Enhanced chromatin dynamics by FACT promotes transcriptional restart after UV-induced DNA damage. Mol. Cell 51, 469–479 (2013).
Sirbu, B. M. et al. Identification of proteins at active, stalled, and collapsed replication forks using isolation of proteins on nascent DNA (iPOND) coupled with mass spectrometry*. J. Biol. Chem. 288, 31458–31467 (2013).
Dungrawala, H. & Cortez, D. Purification of proteins on newly synthesized DNA using iPOND. Methods Mol. Biol. 1228, 123–131 (2014).
Zeeland, A. A. van, Smith, C. A. & Hanawalt, P. C. Sensitive determination of pyrimidine dimers in DNA of UV-irradiated mammalian cells Introduction of T4 endonuclease V into frozen and thawed cells. Mutat. Res.Fundam. Mol. Mech. Mutagen. 82, 173–189 (1981).
Wang, L. et al. Nucleotide excision repair removes thymidine analog 5-ethynyl-2′-deoxyuridine from the mammalian genome. Proc. Natl. Acad. Sci. USA 119, e2210176119 (2022).
Hu, J., Adebali, O., Adar, S. & Sancar, A. Dynamic maps of UV damage formation and repair for the human genome. Proc. Natl. Acad. Sci. 114, 6758–6763 (2017).
Adar, S., Hu, J., Lieb, J. D. & Sancar, A. Genome-wide kinetics of DNA excision repair in relation to chromatin state and mutagenesis. Proc. Natl. Acad. Sci. 113, E2124–E2133 (2016).
Stefos, G. C., Szantai, E., Konstantopoulos, D., Samiotaki, M. & Fousteri, M. aniFOUND: analysing the associated proteome and genomic landscape of the repaired nascent non-replicative chromatin. Nucleic Acids Res. 49, gkab144 (2021).
Boeing, S. et al. Multiomic analysis of the UV-induced DNA damage response. Cell Rep. 15, 1597–1610 (2016).
Wessel, S. R., Mohni, K. N., Luzwick, J. W., Dungrawala, H. & Cortez, D. Functional analysis of the replication fork proteome identifies BET proteins as PCNA regulators. Cell Rep. 28, 3497–3509.e4 (2019).
Alvarez, V. et al. Proteomic profiling reveals distinct phases to the restoration of chromatin following DNA replication. Cell Rep. 42, 111996 (2023).
Huang, H. et al. A unique binding mode enables MCM2 to chaperone histones H3–H4 at replication forks. Nat. Struct. Mol. Biol. 22, 618–626 (2015).
Richet, N. et al. Structural insight into how the human helicase subunit MCM2 may act as a histone chaperone together with ASF1 at the replication fork. Nucleic Acids Res. 43, 1905–1917 (2015).
Han, C. et al. HDJC9, a novel human type C DnaJ/HSP40 member interacts with and cochaperones HSP70 through the J domain. Biochem. Biophys. Res. Commun. 353, 280–285 (2007).
Piette, B. L. et al. Comprehensive interactome profiling of the human Hsp70 network highlights functional differentiation of J domains. Mol. Cell 81, 2549–2565.e8 (2021).
Hammond, C. M. et al. DNAJC9 integrates heat shock molecular chaperones into the histone chaperone network. Mol. Cell 81, 2533–2548.e9 (2021).
Ding, Y. et al. The ortholog of human DNAJC9 promotes histone H3-H4 degradation and is counteracted by Asf1 in fission yeast. Nucleic Acids Res 53, gkaf036 (2025).
Adam, S., Dabin, J., Bai, S.-K. & Polo, S. E. Imaging local deposition of newly synthesized histones in UVC-damaged chromatin. Methods Mol. Biol. 1288, 337–347 (2015).
Ray-Gallet, D. et al. Dynamics of histone H3 deposition in vivo reveal a nucleosome gap-filling mechanism for H3.3 to maintain chromatin integrity. Mol. Cell 44, 928–941 (2011).
Reid, D. A. et al. Incorporation of a nucleoside analog maps genome repair sites in postmitotic human neurons. Science 372, 91–94 (2021).
Carraro, M. et al. DAXX adds a de novo H3.3K9me3 deposition pathway to the histone chaperone network. Mol. Cell 83, 1075–1092.e9 (2023).
Smith, C. L. et al. A separable domain of the p150 subunit of human Chromatin Assembly Factor-1 promotes protein and chromosome associations with nucleoli. Mol. Biol. Cell 25, mbc.E14-05–1029 (2014).
Balachandra, V. et al. DNAJC9 prevents CENP-A mislocalization and chromosomal instability by maintaining the fidelity of histone supply chains. EMBO J 43, 2166–2197 (2024).
Polasek-Sedlackova, H., Miller, T. C. R., Krejci, J., Rask, M.-B. & Lukas, J. Solving the MCM paradox by visualizing the scaffold of CMG helicase at active replisomes. Nat. Commun. 13, 6090 (2022).
Marteijn, J. A., Lans, H., Vermeulen, W. & Hoeijmakers, J. H. J. Understanding nucleotide excision repair and its roles in cancer and ageing. Nat. Rev. Mol. Cell Biol. 15, 465–481 (2014).
Sugasawa, K. et al. Two human homologs of Rad23 are functionally interchangeable in complex formation and stimulation of XPC repair activity. Mol. Cell. Biol. 17, 6924–6931 (1997).
Horianopoulos, L. C. et al. A J domain protein functions as a histone chaperone to maintain genome integrity and the response to DNA damage in a human fungal pathogen. mBio 12, e03273-21 (2021).
Lemaître, C. et al. Nuclear position dictates DNA repair pathway choice. Genes Dev. 28, 2450–2463 (2014).
Lemaître, C. et al. The nucleoporin 153, a novel factor in double-strand break repair and DNA damage response. Oncogene 31, 4803–4809 (2012).
Ryu, T. et al. Heterochromatic breaks move to the nuclear periphery to continue recombinational repair. Nat. Cell Biol. 17, 1401–1411 (2015).
Schep, R. et al. Impact of chromatin context on Cas9-induced DNA double-strand break repair pathway balance. Mol. Cell 81, 2216–2230.e10 (2021).
Shokrollahi, M. et al. DNA double-strand break-capturing nuclear envelope tubules drive DNA repair. Nat. Struct. Mol. Biol. 31, 1319–1330 (2024).
Katsumi, S. et al. In situ visualization of ultraviolet-light-induced DNA damage repair in locally irradiated human fibroblasts. J. Investig. Dermatol. 117, 1156–1161 (2001).
Moné, M. J. et al. Local UV-induced DNA damage in cell nuclei results in local transcription inhibition. EMBO Rep. 2, 1013–1017 (2001).
Dinant, C. et al. Activation of multiple DNA repair pathways by sub-nuclear damage induction methods. J. Cell Sci. 120, 2731–2740 (2007).
Casabona, M. G., Vandenbrouck, Y., Attree, I. & Couté, Y. Proteomic characterization of Pseudomonas aeruginosa PAO1 inner membrane. Proteomics 13, 2419–2423 (2013).
Bouyssié, D. et al. Proline: an efficient and user-friendly software suite for large-scale proteomics. Bioinformatics 36, 3148–3155 (2020).
Couté, Y., Bruley, C. & Burger, T. Beyond target–decoy competition: stable validation of peptide and protein identifications in mass spectrometry-based discovery proteomics. Anal. Chem. 92, 14898–14906 (2020).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47, gky1106 (2018).
Wieczorek, S. et al. DAPAR & ProStaR: software to perform statistical analyses in quantitative discovery proteomics. Bioinformatics 33, 135–136 (2017).
Medvedeva, Y. A. et al. EpiFactors: a comprehensive database of human epigenetic factors and complexes. Database 2015, bav067 (2015).
Marakulina, D. et al. EpiFactors 2022: expansion and enhancement of a curated database of human epigenetic factors and complexes. Nucleic Acids Res. 51, D564–D570 (2022).
Bodor, D. L., Rodríguez, M. G., Moreno, N. & Jansen, L. E. T. Analysis of protein turnover by quantitative SNAP-based pulse-chase imaging. Curr. Protoc. Cell Biol. 55, 8.8.1−8.8.34 (2012).
Dunleavy, E. M., Almouzni, G. & Karpen, G. H. H3.3 is deposited at centromeres in S phase as a placeholder for newly assembled CENP-A in G1 phase. Nucleus 2, 146–157 (2011).
Acknowledgements
The authors thank Colin Hammond and Anja Groth for sharing plasmids, the EPI2 Imaging platform and in particular Sandra Piquet—UMR7216 Epigenetics and Cell Fate Centre for access to instruments and technical advice. This work was supported by the European Research Council (ERC-2018-CoG-818625 “REMIND” to S.E.P.). The proteomic experiments were partially supported by Agence Nationale de la Recherche under projects ProFI (Proteomics French Infrastructure, ANR-10-INBS-08) and GRAL, a program from the Chemistry Biology Health (CBH) Graduate School of University Grenoble Alpes (ANR-17-EURE-0003) to Y.C.
Author information
Authors and Affiliations
Contributions
A.P., A.C., E.P., and S.E.P. designed and performed experiments, analyzed the data, and wrote the manuscript. J.N.D. and Y.C. performed MS-based proteomic analyses. S.E.P. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Plessier, A., Chansard, A., Petit, E. et al. Proteomic profiling of UV damage repair patches uncovers histone chaperones with central functions in chromatin repair. Nat Commun 17, 2127 (2026). https://doi.org/10.1038/s41467-026-68781-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68781-x