Abstract
Cell division is a central process in all living organisms and requires the coordinated action of many proteins and regulatory elements. In most bacteria, the division and cell wall (dcw) gene cluster is regulated by the first gene of the dcw operon, mraZ, a highly conserved DNA-binding transcriptional regulator. Here we report the structural basis of MraZ transcriptional regulation by the resolution of three different cryo-EM structures of MraZ in complex with the upstream promoter region of the dcw cluster from Mycoplasma genitalium at 3.36, 3.57 and 3.87 Å resolution. The structures reveal the specific interactions between MraZ DNA-binding motif and nucleobases of the binding boxes, which induces distortion in the MraZ octamer to enable the interaction with the four repetitive binding boxes of the promoter DNA. The “cradle-like” DNA-binding motif of MraZ exposes three highly conserved basic residues, Lys13, Arg15 and Arg86, which are essential for binding to the consensus sequence of its cognate promoter. Ultimately, the mechanism behind MraZ’s DNA binding and regulation of the dcw operon could be translated to other species, working as a general mechanism for the regulation of dcw gene cluster in bacteria.
Similar content being viewed by others
Introduction
The study of bacterial proteins involved in essential cellular processes provides valuable insights into the molecular mechanisms that govern life. Among the simplest self-replicating organisms, Mycoplasma genitalium possesses a highly reduced genome, consisting only of 580 kb, thus, making it an ideal model for investigating fundamental genetic and biochemical functions1,2,3.
Cell division is a fundamental process in all living organisms, requiring the coordinated action of multiple proteins and regulatory elements4,5,6. In most bacteria, the genes essential for cytokinesis and peptidoglycan biosynthesis are located within the division and cell wall (dcw) gene cluster, a highly conserved operon across various bacterial species7,8,9.
However, mycoplasmas, which lack a cell wall due to extensive genome reduction, typically retain only four genes from the dcw cluster: mraZ, mraW, ftsA, and ftsZ10,11,12,13. mraZ, the first gene in the operon, encodes a transcriptional regulator that controls the expression of genes within the dcw operon9,14,15,16. mraW encodes an RNA methyltransferase that modifies 16S ribosomal RNA17,18,19. The last two genes, ftsZ and ftsA, have been more extensively characterized. ftsZ encodes a tubulin-like protein that assembles into the Z-ring, a crucial structure for bacterial cytokinesis4,20,21. Meanwhile, ftsA functions as a membrane anchor for the FtsZ protein22,23,24,25,26. While ftsZ is essential for division in E. coli27 and B. subtilis28, it is not required for in vitro growth in M. genitalium, making its conservation in these organisms intriguing12,13. Although the absence of cell wall synthesis enzymes in mycoplasmas aligns with their wall-less nature, the conservation of mraZ and mraW suggests that these genes may have crucial roles beyond cell wall biogenesis.
Several recent studies have shown that MraZ functions as a transcriptional regulator by binding to conserved sequence motifs upstream of the dcw promoter9,14,15,29. Across phylogenetically diverse microorganisms, a well-conserved sequence is present, usually consisting of three GTG(G/T) boxes separated by six nucleotides. Interestingly, in M. genitalium, a fourth box is observed, with adenine being the predominant nucleotide in the three bases closest to each box. Thus, the binding sequence for M. genitalium MraZ can be defined as: AAAGTGT-N3-AAAGTGT-N3-AAAGTGT-N3-AAAGTGG. In this context, MraZ has traditionally been described as a repressor of the dcw cluster9,15,16,29. Nonetheless, in M. gallisepticum, it has been reported to function as an activator upon overexpression14. Research on the Neisseriaceae family suggests that the deletion of mraZ, in conjunction with other factors, may have contributed to the development of alternative growth strategies30. Moreover, outside cell division, MraZ also plays a role in bacterial virulence regulation in Staphylococcus aureus31,32, highlighting MraZ’s diverse and complex regulatory functions beyond its traditional association with the regulation of the dcw operon.
Structural studies of MraZ from M. pneumoniae provide insights into its three-dimensional organization33. M. pneumoniae’s MraZ forms an octameric, ring-shaped structure with positively charged outer surfaces, characteristic of DNA-binding proteins. The presence of a conserved DXXXR motif, resembling DNA-binding domains found in transcriptional regulators such as AbrB from B. subtilis and MazE from E. coli (SpoVT-AbrB domain), further supports this function33,34,35,36,37. Interestingly, structural analyses in E. coli suggest that MraZ may assemble into a dodecameric structure34. Despite these structural insights, the mechanisms by which MraZ interacts with the DNA promoter region to regulate transcription remain elusive.
Here, we present two cryoEM structures of four MraZ subunits bound to its cognate 4-box promoter region from M. genitalium at 3.36 Å and 3.87 Å resolution, and one structure of the octameric MraZ ring bound to 1-box at 3.57 Å resolution. Interestingly, MraZ relies on β-strands rather than the more common α-helical motif to interact with and recognize its cognate DNA sequence, resembling the cooperativity and DNA binding mechanism of Arc, CopG and MetJ repressors38,39,40,41,42. Our results shed light on the molecular mechanism behind the MraZ binding to the conserved 4-box promoter region, as well as the role of MraZ oligomerization in DNA binding and thus, in transcription regulation.
Results
MraZ binds to a highly conserved sequence upstream of its promoter
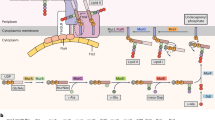
Comparative analysis of the dcw cluster across diverse bacterial species revealed a high degree of conservation in gene organization and regulatory elements. Most microorganisms maintain a well-preserved operon structure, with Mycoplasma exhibiting a more compact arrangement, reflecting its reduced genome (Fig. 1a). Consistent with previous studies9,12,14,15,29,30,43, the sequence alignment of the upstream region of the M. genitalium mraZ gene confirms the presence of highly conserved DNA repeated boxes, which have been associated with regulatory functions9,12,14,15,29,30,43 (Fig. 1b). Whereas in most bacteria this upstream region typically contains three conserved repeated boxes, some Mycoplasma species seem to possess a distinct configuration with the presence of four boxes. Additionally, we also determined the transcription start site (TSS) for the dcw cluster operon, which is located inside the mraZ box 1 promoter (Fig. 1c and Supplementary Fig. 1).
a The four conserved genes in M. genitalium (mraZ, mraW, ftsZ and ftsA*) are highlighted with different colors. The third gene of M. genitalium operon, ftsA is marked with an asterisk as its gene product is a putative ftsA homolog. b Sequence alignment of the upstream region of the dcw cluster comprising the repeated binding boxes of MraZ. In bold, the GTG(G/T) boxes, with six spacer nucleotides. At the bottom, a sequence pattern was generated with Weblogo from the sequence alignment (Crooks et al.69). c Promoter structure in the genome region upstream to cell cell division operon (dcw) of M. genitalium. The transcription start site (TSS, depicted with a black arrow) is located inside the first mraZ box (in brown) and corresponds to the G nucleotide at position 266629 (in green). The − 10 box consensus Pribnow box is highlighted in green. The mraZ gene is highlighted in yellow, and the first encoded amino acids are in red.
We employed electrophoretic mobility shift assays (EMSAs) with different DNA probes to assess the binding of MraZ to its cognate region in M. genitalium (Fig. 2a). The results revealed that purified MraZ binds specifically to the promoter region, as shown by a diminished signal corresponding to unbound DNA (Fig. 2b). To further investigate the contribution of specific binding boxes to the MraZ interaction, we generated mutations within each box sequence and quantified MraZ binding. Interestingly, we observed a stepwise decrease in MraZ binding affinity upon sequentially mutating the binding boxes, underscoring the cooperative behavior and the importance of the four elements for a robust MraZ-DNA interaction (Fig. 2b).
a Cartoon representation of the dcw promoter region showing the binding repeats in yellow and the scheme of the different the promoter variants used for EMSA and repression assays; mutations in the binding boxes are shown in gray and marked with a cross. b, d EMSA assays of increasing concentrations of MraZ with oligonucleotides bearing the different configurations of the binding boxes. Protein concentration is comprised between 0 and 400 nM. dsDNA is at 50 nM and labeled with 5’FAM. Data values represent the mean, n = 3 technical replicates. Data values representing the mean ± SEM are shown in the source data. c Repression assays containing GFP-based transcriptional reporter under the control of WT or mutant promoter binding boxes with overexpression of WT. Data values represent the mean ± SEM, n = 3 technical replicates. Significance was measured by a two-tailed unpaired t test relative to the positive GFP expression control. All data were analyzed with a 95% confidence interval. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. Exact P-values from left to right: 0.0001, < 0.0001, < 0.0024, 0.0003, < 0.0001 and 0.0001.
The repression capability of MraZ was also assessed by a reporter assay in E. coli with the GFP gene under the control of the wild-type or binding boxes deletion mutants of the MraZ cognate promoter of M. genitatium (Fig. 2c). Similar to the EMSA binding analysis, the reporter assay also highlights the importance of the four boxes in the repression, revealing a stepwise increase of the GFP signal upon mutating the binding boxes (Fig. 2c). Moreover, the spacing between boxes also appeared relevant in both the reporter assay and the binding properties of MraZ to its cognate 4-box promoter sequence. Deletion of the two middle boxes resulted in a binding affinity similar to the 2-box contiguous DNA in both EMSA and reporter assays. Moreover, when a longer spacer is introduced between the 4-boxes, MraZ is still able to bind the DNA efficiently at high concentrations (Fig. 2d). These results indicate that MraZ directly engages cooperatively a conserved regulatory 4-box sequence upstream of its own promoter, and that protein-protein interactions contribute to an optimal binding of MraZ to the target DNA.
Oligomeric structural assemblies of MraZ
MraZ is conserved across many bacterial species, with M. genitalium MraZ sharing 28% sequence identity with Escherichia coli, 36% with Bacillus subtilis and 82% with Mycoplasma pneumoniae, among others (Supplementary Fig. 2a). Despite these variations, the structural fold and key structural motifs seems to be preserved in the MraZ family, allowing the formation of a “cradle-like” antiparallel β-barrel domain flanked with some external α-helices (Fig. 3 and Supplementary Fig. 2b)33,34,44,45. Interestingly, structural comparison revealed that the MraZ fold is related to the family of the SpoVT-AbrB transcription factors (SpoVT-AbrB domain), but in this instance, the domain is formed by a homodimer assembly, in contrast to MraZ, which is formed by a unique protein sequence35,36,37,44,45,46. Importantly, the SpoVT/AbrB family members contain a DXXXR motif, highly conserved in MraZ family members, shown to be responsible for the DNA binding35,36,47,48,49,50 (Supplementary Fig. 3).
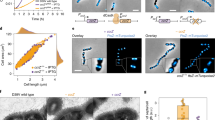
a Two orthogonal cartoon representations of the octamer assembly of MraZ. Each subunit is shown in a different color, and the octamer diameter and thickness are displayed. (below) Close up view of the interdigitated subunits interface formed by helical structures. The C-terminus and helices involved in the interface are labeled. b Two orthogonal cartoon representations of the nonameric assembly of MraZ. Each subunit is shown in a different color, and the nonameric diameter and thickness are displayed. c (left) TEM electron microscopy image of MraZ. 2D reconstruction of a cryoEM micrography of MraZ displaying 3 different Mraz oligomeric assemblies. d Close up view of two orthogonal cartoon representations of the octamer interface of MraZ. Side chains of residues involved in the interface are labeled and shown in stick representation. Secondary structure elements of the MraZ Interface are labeled.
The two crystal structures presented here reveal the existence of different oligomeric states of MraZ in M. genitalium, forming quaternary assemblies as an octamer or as a nonameric structures (Fig. 3and Table 1). The octamer and the nonameric assemblies have an outer diameter of ~ 110 Å and ~ 120 Å, respectively, both with a thickness of ~ 30 Å (Fig. 3a,b). This diverse oligomerization assemblies in MraZ has also been observed in M. pneumoniae and in E. coli, the latter proposed to be composed by 12 subunits33,34. Also, cryo-electron microscopy 2D class averaging identifies octa-, nona-, and decameric assemblies, confirming the structural oligomeric plasticity of the M. genitalium MraZ (Fig. 3c). Among all quaternary assemblies, the octamer appears to be the most prevalent (50%), consistent with MraZ from M. pneumoniae33.
The MraZ “cradle-like” SpoVT-AbrB domain is flanked by four α-helices that are mostly responsible for the protein-protein interactions in the oligomeric assembly. Essentially, the C-terminal α5 and α6 helices of each subunit are interdigitated with the α2-loop-α3 helices of the next two subunits, forming an extended interface involving three different MraZ subunits (Fig. 3a). Most interface contacts are provided by residues emerging from the helices, but also from the N-terminus and two β-hairpin loops of the “cradle-like” domain, namely β3-β4 and β7-β8 (Fig. 3d). Interface contacts include hydrophobic residues, such as Met1, Leu3, Phe36, Phe58, Leu68, Phe73, Phe109, Val133 and Met137, as well as the formation of polar and electrostatic contacts, such as Glu49, Thr61, Gln62, Asp64, Arg66, Arg70, Gln108, Glu113 and Arg136. The global surface extension between subunits in the octamer interface is approximately 1568.9 A2.
The MraZ oligomeric interface is basically identical in the crystal structures of the octamer and nonameric assemblies, and as we will describe in the next section, it is mostly conserved upon DNA binding.
MraZ structure in complex with its cognate DNA promoter
MraZ binding to DNA has been studied previously9,12,14,15,29,30,43, but a structural framework for this interaction remained elusive. Here, we report three cryoEM structures of MraZ in complex with dsDNA segments containing the four binding boxes of the promoter of M. genitatium at 3.36, 3.57 and 3.87 Å resolution. Remarkably, the determination of the cryoEM structure was facilitated by the formation of supra-helix fiber-like assemblies formed by an MraZ oligomer bound to DNA segments containing the cognate 4-box dsDNA sequence (Fig. 4a, b and Supplementary Figs 4–6). In one of the fiber assemblies, at 3.36 Å resolution, the 43 bp dsDNA 4-box segment forms concatemers, in contrast to the second fiber assembly at 3.87 Å resolution, in which the 55 bp dsDNA 4-box segment do not form concatemers and dsDNA breaks are clearly distinct. Importantly, the supra-helical assemblies are not observed either by perturbing the cognate DNA binding boxes or by mutating the MraZ residues responsible for DNA binding or for the MraZ oligomer formation (Supplementary Fig. 7), indicating the specific interaction between the MraZ with the four boxes of the cognate promoter of M. genitatium. Also, an additional cryoEM structure at 3.57 Å resolution was obtained between the MraZ octamer ring bound to 1-box of the cognate 4-box dsDNA sequence, probably indicating an initial step in the MraZ binding mechanism to the promoter (see later).
a left CryoEM image of the helical “fiber-like” formed by MraZ in complex with a 43 bp dsDNA containing four binding boxes. Middle two opposite orientations of the initial cryoEM 3D volume reconstruction of the fiber-like structure forming DNA concatemers. Right local resolution of the final reconstruction of the complex between four binding DNA boxes with four MraZ subunits (unsharpened map). Scale bar indicated resolution range. b left cryoEM image of the helical “fiber-like” formed by MraZ in complex with a 55 bp dsDNA containing four binding boxes. Middle two opposite orientations of the initial cryoEM 3D volume reconstruction of the fiber-like structure that do not form DNA concatemers. Right local resolution of the final reconstruction of the complex between four binding DNA boxes with four MraZ subunits (unsharpened map). Scale bar indicated resolution range. c Final model fitted in the electron microscopy densities of the complex between DNA-binding boxes with four MraZ subunits (unsharpened map). d Three views of the MraZ electrostatic surface representation in complex with the DNA segment, shown in a cartoon representation. e Structural comparison between the crystal structure of the octamer (brown) and the cryoEM structure of the complex with DNA (different color for each subunit chain) after superposition of one subunit. The DNA segment is shown in a cartoon representation. Distance and angle variation between Ser83 from the DNA segment, shown in a cartoon representation two adjacent DNA binding motifs from two subunits are displayed.
Binding to DNA is conducted by the “cradle-like” structural motif of each MraZ subunit, which fits within the major groove of each box of the promoter sequence (Figs. 4, 5). An electrostatic surface representation depicts the positive region from each MraZ DNA binding motif inserted within the major groove of each binding box of the promoter region (Fig. 4d). Besides, the cryoEM complex structure maintains the interdigitated oligomeric helical interface between the four MraZ subunits observed in the unbound MraZ octamer and nonameric rings.
a Cartoon representation of the complex between four binding DNA boxes with four MraZ subunits. DNA boxes and subunits are labeled and shown in different colors. b Surface representation four DNA binding boxes in complex with four MraZ subunits. c Schematic representation of the interactions between side chains and nucleobases of the four binding boxes. Interactions are shown by short lines. d Close up view of the contacts between Lys13, Arg15 and Arg86 with the consensus sequence of the binding box in the DNA major groove. Contact residues with nucleobases and distances are labeled and shown in stick representation. Sharpened volumes are displayed for the contact region.e Opposite view of (d). f Agarose gels of EMSA experiments of MraZ wild type and K13A, R15A and R86A with fluorescent dsDNA containing four boxes. g Repression assays containing GFP-based transcriptional reporter under the control of MraZ promoter binding boxes with overexpression of WT or DNA-binding mutant MraZ protein. Data values represent the mean ± SEM, n = 3 technical replicates. Significance was measured by a two-tailed unpaired t test relative to the positive GFP expression control. All data were analyzed with a 95% confidence interval. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. Exact P values from left to right: 0.0001 and 0.0026. h Bar plot of the RT-qPCR experiment showing the GFP mRNA levels of a GFP-based transcriptional reporter under the control of the MraZ promoter with overexpression of WT or DNA-binding mutant. Data values represent the mean ± SEM, n = 3 technical replicates. Significance was measured by one-way ANOVA with Tukey post-hoc test. Exact P-values from left to right: 0.0004 and 0.1535.
However, structural superposition of the MraZ octamer with the DNA-bound tetramer reveals that the curved octamer or nonameric ring can’t accommodate binding to the four promoter DNA boxes. While maintaining most of the interface contacts, the MraZ DNA-bound tetramer reassembles a distorted open ring octamer. This transformation requires rigid body subunit rearrangements to fit the “cradle-like” binding motif into the four boxes of the cognate promoter. Readjustments of the interdigitated helices result in shifts and angle differences between subunits of up to 8 Å and 22°, respectively (Fig. 4e). Specifically, the α2–loop–α3 helices and the C-terminal α6 helix undergo notable rearrangements. For instance, residues like Phe52 or Phe58 in the DNA-bound MraZ establish contacts in the opposite orientation compared to their positions in the unbound octamer (Supplementary Fig. 8).
Opening the MraZ octamer ring is thus essential in this mechanism, as it provides the proper geometry for the four exposed “cradle-like” binding motifs to engage the cognate promoter’s four binding boxes.
DNA binding by the MraZ “cradle-like” structural motif
The total buried area between the MraZ oligomer and the 4-box dsDNA, calculated by the PISA-PDBe server, is around 3960 Å2,51 whereas the total buried area between the four MraZ subunits is around 7600 Å2. In the cryoEM structure at 3.36 Å resolution, only three conserved DNA binding boxes can be correctly traced in the final cryoEM volume after particle averaging, the fourth DNA box was not well defined, probably due to low local resolution or to volume averaging of the concatemeric DNA boxes (Fig. 5a, b). However, in the structure at 3.87 Å resolution, all 4-boxes are perfectly observed in the volume maps, confirming the engagement of 4 MraZ subunits with the four boxes of the promoter of M. genitatium. Both cryoEM structures can be perfectly superimposed.
The “cradle-like” DNA binding motif in MraZ is conserved in the family of bacterial transcription regulators SpoVT/AbrB (Supplementary Fig. 3), and it is formed by a short antiparallel β2-β6 sheet flanked by two similar loops at opposite sites, resembling the shape of a cradle rocking the DNA (Fig. 5a, b). In our cryoEM complex, each “cradle-like” DNA-binding motif binds a DNA major groove that contains a MraZ-consensus signature TGT/G (Fig. 5c). Three basic side chains emerging from the “cradle-like” DNA-binding motif of each subunit, namely Lys13, Arg15 and Arg86, are engaged with the nitrogen bases of the conserved TGT/G sequence and provide specific contacts with the cognate promoter region (Fig. 5d, e). The structural conformation of this basic DNA binding motif is also maintained by the conserved Asp11 and Asp82 that are engaged in electrostatic contacts with Arg15 and Arg86, thus contributing to keeping a correct side chain conformer orientation to bind the nitrogen bases of the TGT/G consensus sequence.
A detailed analysis of the MraZ subunit B in complex with the DNA binding Box-2 depicts the specific contacts engaged by the three basic residues (Fig. 5d, e and Supplementary Fig. 9). The guanidinium groups of Arg86 and Arg15 forms H-bond interactions with the nucleobases of the pairs T26 and A26’ (3.0 and 3.9 Å, respectively), and G25 and C25’ (3.4 and 4.1 Å, respectively) (Fig. 5d, e). Also, the ε-amino group of Lys13 is at H-bond distance of the nucleobase pair T24 and A24’ (3.9 and 3.6 Å, respectively). The other binding boxes display some variations, such as Box1, in which Lys13 is also capable of interacting with the previous conserved nucleobase pair, G33 (Fig. 5c).
To better understand the specific DNA-protein interactions, we generated single-point mutants of Lys13, Arg15, and Arg86 to alanine and then analyzed their ability to bind DNA. Electrophoretic Mobility Shift Assay (EMSA) with a double-stranded DNA probe containing the four binding boxes revealed that, unlike the wild-type protein, all single-point mutants of these three basic residues failed to bind DNA (Fig. 5f). Furthermore, we observed that a double arginine mutant (Arg15 and Arg86) failed to repress GFP expression due to its inability to bind to the four-box promoter region (Fig. 5g). These results were corroborated by qPCR analysis of the GFP transcript levels, which strongly decay in the presence of wild type MraZ and recover in the presence of the MraZ double arginine mutant (Fig. 5h). These results confirm the crucial role of each individual basic residue, namely Lys13, Arg15, and Arg86, in the specific DNA binding to the promoter region of the dcw cluster in M. genitalium.
While Lys13, Arg15, and Arg86 make specific contacts with the DNA, other non-specific interactions between each MraZ subunit and the double-stranded DNA phosphate groups also participate in DNA binding. These include polar contacts and hydrogen bonds formed by side chain residues like Thr9, Asn14, Ser17, and Ser83, as well as main chain atoms such as the carbonyl oxygens and amino groups of Asn7, Leu18, Ala20, and Leu88 (Fig. 6). Interestingly, the only other nucleobase contact, besides those made by the three key basic residues, is between Ser17 and T28’ (at a distance of 3.8 Å).
a, b Two opposite views of side chain and main chain contacts between MraZ chain B with phosphate groups and nucleobases of the DNA binding Box2 sequence. Contact residues are labeled and shown in stick representation. Contacts (< 4.5 A distance) are shown by a discontinuous red line.
In summary, MraZ’s specificity for its promoter primarily stems from contacts made by Lys13, Arg15, and Arg86 with each of the four conserved GTG(T/G) boxes on the DNA. However, several other non-specific contacts with the DNA phosphate backbone also contribute significantly to the overall binding affinity.
MraZ octamer bound to one box of the cognate promoter
As mentioned before, an additional cryoEM structure at 3.57 Å resolution was obtained between the MraZ octamer ring bound to 1-box of the 4-box dsDNA sequence, probably depicting an initial step in the MraZ binding mechanism to the cognate 4-box promoter sequence (Fig. 7a, b). Interestingly, this cryoEM structure of the MraZ octamer bound to 1-box dsDNA maintains similar interface contacts with the nucleobases and phosphate groups compared to the other two cryoEM structures bound to the 4-box DNA segment. The essential three basic side chains from the “cradle-like” DNA binding motif, namely Lys13, Arg15 and Arg86, are already engaged with the nitrogen bases of the conserved GTG(T/G) sequence to provide specific contacts (Fig. 3c).
a two opposite orientations of the local resolution of the final volumes of the complex between one DNA binding box with the MraZ octamer ring (unsharpened map). Scale bar indicated resolution range. b Final model fitted in the electron microscopy density of the complex between one DNA binding box with the MraZ octamer ring (unsharpened map). c Two opposite close-up views of the contacts between Lys13, Arg15 and Arg86 with the consensus sequence of the binding box in the DNA major groove. Contact residues with nucleobases and distances are labeled and shown in stick representation.
The MraZ octamer ring in this cryoEM-DNA structure perfectly overlaps with the free MraZ octamer, and most interface contacts between subunits are conserved. We believe that this cryoEM complex of the octamer bound to 1-box DNA might represent the initial step in the specific binding of MraZ to the cognate 4-box promoter. Next, conformational changes must occur in the octamer interface to open the MraZ ring and allow binding of the “cradle-like” DNA binding motif of three adjacent MraZ subunits to the specific promoter boxes. Our model considers that the MraZ oligomer might increase the local concentration of “cradle-like” DNA binding motifs in a right orientation and distance to bind all four DNA promoter boxes by opening the ring.
MraZ oligomerization influences the DNA binding affinity
We next questioned to what extent the MraZ oligomerization affects the DNA-binding affinity, thus, we generated a monomeric MraZ variant by perturbing the extended oligomerization interface. To validate that the oligomerization interface was fully perturbed, we mutated Phe36 to arginine, Phe52 to lysine and removed the C-terminal α6 helix (Supplementary Fig. 10). The mutations were designed based on PDB-PISA, FoldX and Aggrescan3D analysis51,52,53. The C-terminal truncated MraZ double mutant, named ΔCTDMraZ-F36R/F52R, elutes as a monomer in a size-exclusion chromatography and does not show the typical ring-shaped oligomer assembly observed by negative staining in TEM analysis (Supplementary Fig. 10).
Interestingly, the ΔCTDMraZ-F36R/F52R mutant is also capable to bind DNA specifically but with a lower affinity. EMSA experiments were conducted with the ΔCTDMraZ-F36R/F52R mutant and various dsDNA probes containing the different boxes of the promoter region of M. genitalium dcw cluster. The ΔCTDMraZ-F36R/F52R mutant binds the 4-box dsDNA probe with a Kd of 94.4 nM, whereas the wild type MraZ binds the 4-box dsDNA probe with a Kd of 43.3 nM (Fig. 8a, b). Furthermore, the ΔCTDMraZ-F36R/F52R mutant shows a lower binding capacity towards all checked dsDNA probes in comparison to the MraZ wild type octamer form, while maintaining a similar repression capacity in our reporter assay and qPCR analysis in E. coli (Supplementary Fig. 11). Moreover, in contrast to the wild type MraZ, the ΔCTDMraZ-F36R/F52R mutant is not able to bind efficiently the DNA at high concentrations when a longer spacer is located between the 4-boxes (Fig. 8c), probably indicating a role of oligomerization to facilitate DNA binding.
a, b EMSA assays of increasing MraZ and MraZΔCTDF36RF52K concentrations with oligonucleotides bearing the wild-type and mutant binding boxes. Protein concentration is comprised between 0 and 400 nM. dsDNA is at 50 nM and labeled with 5’FAM. Data values represent the mean, n = 3 technical replicates. Data values representing the mean ± SEM are shown in the source data. c Similar EMSA assay as (a) but using a long-spacer DNA probe in the middle of the 4-box promoter.
While our findings underscore that MraZ’s oligomerization enhances DNA binding, it is not strictly essential for this process, as it is mediated by a distinct interface. This observation is consistent with what probably occurs in the SpoVT/AbrB family of bacterial transcription regulators (Supplementary Fig. 3), in which the MraZ helical interface, responsible for oligomerization, is absent, and only relies on the DNA-binding motif of the “cradle-like” SpoVT/AbrB domain.
Discussion
MraZ is a conserved bacterial transcriptional regulator found across diverse species, typically located at the beginning of the dcw cluster. It has been implicated in autoregulation and broader control of cell division genes, through binding to conserved repeated sequences upstream of its own promoter. In this study, we focus on MraZ from M. genitalium, a model for minimal bacteria, which features a highly reduced genome and a particularly compact dcw cluster. While previous studies have suggested MraZ’s involvement in DNA binding and oligomerization, the lack of a high-resolution structure in complex with DNA has limited our mechanistic understanding of its function. Here, we combine structural, biochemical, and functional approaches to elucidate how M. genitalium MraZ recognizes its target DNA and whether oligomerization can influence DNA-binding affinity.
In this work, we present the experimentally determined structure of bacterial MraZ in complex with its target DNA. Using cryo-electron microscopy, we resolved the MraZ-DNA complex at near-atomic resolution, revealing the precise interface through which MraZ recognizes and engages the conserved promoter region of the dcw gene cluster upstream of its own gene. This represents a major advance, as previous insights into MraZ-DNA interactions have relied solely on biochemical studies and computational modeling14,15,33,49.
The cryo-EM structure reveals that MraZ still binds DNA as an oligomer, preserving the characteristic C-terminal docking interactions between subunits but no longer maintains a closed ring-shaped conformation (Fig. 4). Within this complex, DNA is clamped along the positively charged outer surface of the oligomer, with direct contacts mainly mediated by the unique residues conserved in the SpoVT/AbrB domain (Supplementary Fig. 2). Arg15, Arg86 and Lys13 play a central role in establishing sequence-specific interactions, corroborated by mutagenesis, EMSA, reporter assay and qPCR data, where substitution of either residue abolished DNA binding, transcriptional repression and GFP-reporter expression (Fig. 5). These findings not only validate previous predictions9,14,15,33,34 but now anchor them in a defined structural context, confirming the central role of the “cradle-like” DNA-binding motif in the transcriptional regulation by MraZ.
Functionally, MraZ binds with high specificity to a cluster of repeated DNA boxes upstream of its promoter region. Our experiments demonstrated that mutations in these boxes reduce binding affinity, indicating that each repeat contributes to a stable complex formation (Fig. 2). Interestingly, even when the spacing between boxes was altered or when two central boxes were deleted, MraZ maintained a certain ability to bind DNA at high concentration, suggesting structural flexibility in recognizing diverse DNA configurations.
Our structural findings are further substantiated by the analysis of MraZ oligomerization. Consistent with prior studies33,34, we found that MraZ assembles into various oligomeric forms, with octamers being predominant in the absence of DNA (Fig. 3c). To dissect the functional relevance of oligomerization for DNA binding and transcriptional regulation, we engineered a non-oligomeric MraZ variant by disrupting key interfaces, specifically by mutating Phe36, Phe52 and deleting the last 14 residues. While this mutant retained the capacity to bind DNA, its affinity was markedly reduced, and binding to multiple DNA constructs was significantly weakened in vitro (Fig. 8and Supplementary Fig. 11). Notably, although oligomerization enhances DNA affinity, it is not strictly essential for DNA recognition, as the MraZ mutant still binds to the 4-box DNA probe in EMSAs and still represses GFP RNA transcription in qPCR analysis. This aligns with the SpoVT/AbrB domain family of transcriptional regulators, which are capable to bind DNA binding boxes in the absence of known oligomerization interfaces in this domain36,47,48,49.
These findings suggest that MraZ recognizes DNA primarily through its β-barrel architecture, with oligomerization modulating affinity and possibly cooperativity. Interestingly, structural superposition of the DNA-bound and unbound states reveals that the curvature of the closed-ring octamer is incompatible with the linear geometry of the DNA-bound complex, implying significant conformational rearrangements required to accommodate DNA binding. This observation raises the possibility that, in the bacterial cellular context, MraZ exists in a dynamic equilibrium between a closed-ring oligomeric state and a DNA-bound cooperative assembly (Fig. 9a). Our cryoEM structure of the MraZ octamer bound to 1-box of the promoter might be the first event of the binding mechanism, which strictly relies on the affinity of the MraZ “cradle-like” DNA binding motif for each specific promoter box. Next, the disruption and remodeling of the pre-formed ring is necessary to interact with the remaining promoter boxes. Our model suggests that oligomerization contributes to increase the local concentration of four MraZ DNA binding motifs in a right orientation to bind the four boxes of the promoter.
a Scheme of the oligomer transition between the unbound octamer to the DNA-bound MraZ structures through the open ring mechanism described in the text. PDB codes are shown below each structure. b Cartoon representation of two orthogonal views of Alphafold-2 models of MraZ from M. gallisepticum, E. coli, B. subtilis and N. gonorrhoeae. Conserved arginine residues of the “cradle-like” DNA binding motif are depicted in stick representation.
This dynamic assembly mechanism may represent a broader regulatory strategy among bacterial transcription regulators, at least for MraZ homologs, in which the structural architecture is conserved (Fig. 9b), whereby oligomerization is used not as a strict prerequisite for DNA binding, but as a means of fine-tuning affinity and specificity in response to DNA availability or cellular signals. Future studies examining MraZ dynamics in vivo and identifying cofactors or environmental signals that influence its oligomerization state will be crucial to fully understanding its role in bacterial gene regulation.
Methods
Plasmids, cloning and point mutation
mraZ gene was amplified and cloned into a pET28a vector. Point mutants were performed by site directed mutagenesis54. The primers used for these experiments are all listed in the Supplementary Table 1. Final constructs were verified by DNA sequencing. E. coli XL1Blue strain was used for cloning purposes.
Protein expression and purification
MraZ expression constructs were expressed in E. coli BL21 cells O/N at 20 °C after induction with 0.5 mM IPTG. Harvested cells were resuspended in Lysis Buffer (350 mM NaCl, 20 mM Tris-HCl, pH 8.0, 10 mM imidazole, 20% sucrose, 1 mM β-mercaptoethanol, 10 U/mL DNase and 0.1% IGEPAL CA-630) and disrupted by sonication. Nickel affinity chromatography (GE-Healthcare) was used to purify MraZ via the N-terminal His-tag from the pET28 vector. Proteins were further separated by gel filtration chromatography (Superdex 200 26/60; Cytiva) with buffer 250 mM NaCl, 20 mM Tris 8.0, 1 mM BME. Finally, protein was concentrated using an Amicon (Millipore) until the desired concentration was reached.
Electron microscopy (Negative staining)
Protein or DNA-protein sample were applied to a glow-discharged 300 mesh carbon-coated copper grids (Electron Microscopy Science) for 1 min. Then, excess of liquid was blotted with a Whatman filter paper, and the sample was negatively stained with 5 µL of 1 % uranyl acetate (Polysciences Inc.) for 1 min and blotted again. Grids were finally dried at room temperature. Micrographs were collected in a TEM Jeol 1400 operating at 80 kV and equipped with a Gatan Orius 8 9 SC200 CCD camera or a Hitachi H-7000 operating at 75 kV and equipped with a CCD GATAN ES500W Erlangshen camera.
Crystallization and data collection
Sample protein was concentrated to 9.5 mg/mL for crystallization in a buffer composed of 250 mM NaCl, 20 mM Tris 8.0, 1 mM β-mercaptoethanol. Crystallization was performed at 18 °C by the sitting drop vapor diffusion method by mixing protein with an equal volume of a screen condition solution containing 2 M Ammonium sulfate, 100 mM Sodium cacodylate, pH 6.5 and 200 mM Sodium chloride for the octamer, or 12% w/v Polyethylene glycol 6000 and 2 M Sodium chloride for the nonamer. After 1 week, the crystals were harvested and soaked for 10 s in 15% ethylene glycol before flash freezing in liquid nitrogen.
Diffraction data were collected to 1.96 Å at the ALBA synchrotron beamline BL13-XALOC (Barcelona, Spain) for the octameric assembly55 (Table 1) and collected to 3.85 Å at the ID30B beamline at ESRF synchrotron (Grenoble, France) for the nonameric form56. Data was processed, scaled, reduced, and further analyzed with the automatic autoPROC software package57, including XDS, Pointless, Aimless, CCP4 and Staraniso. Crystallographic details are summarized in Table 1.
Structure determination and refinement (X-ray crystallography)
The structure of MraZ was solved by molecular replacement using MraZ from M. pneumoniae (1N0E) or the AlphaFold-2 model as search models with Phaser. Following rounds of model building and refinement were carried out with Coot and Phenix58,59 (Table 1). The crystal structures of MraZ have been deposited in the Protein Data Bank with the PDB codes 9QLG and 9QLR for the octamer and nonameric respectively.
CryoEM sample preparation and Data Collection
MraZ protein was incubated with (5′- ATAAAAGTGTTTAAAAGTGTCGCAAA-GTGTGACAAAGTGGAAA-3′ and its complementary strand) at RT for 30 min. Complex was formed at 5 µM with 2:1 dsDNA:Protein ratio. 3 µL of sample were applied onto a holey gold grid (UltrAuFoil R1.2/1.3 300 mesh) rendered hydrophilic by a 120-s treatment in a Fischione 1070 plasma cleaner operating at 34% power with a gas mixture of 80% argon:20% oxygen. Grids were blotted for 3.0 s with a blot force of 3 at 6 °C, 100% humidity, and flash frozen in liquid ethane using a FEI Vitrobot Mark IV. Images of the first dataset were collected on a Titan Krios G4 electron microscope (Thermo Fisher Scientific) operated at an acceleration voltage of 300 kV and equipped with a Selectris X energy filter, using a 10 eV slit width in zero-loss. Images were acquired at a nominal magnification of 165,000×, corresponding to a pixel size of 0.73 Å. Movies were recorded on a detector in counting mode at an electron dose rate of 7.88 e − /pixel/s with a total exposure time of 3 s, for an accumulated electron dose of 44.48 e − /Å2. For the second dataset, the complex was formed at 70 μM in the presence of 0.05% DDM detergent. Images were collected on a Glacios microscope (Thermo Fisher Scientific) operated at an acceleration voltage of 200 kV. Images were acquired at a nominal magnification of 45,000 ×, corresponding to a pixel size of 0.901 Å. Movies were recorded on a K2 Summit (Gatan) detector in counting mode at an electron dose rate of 5.06 e − /pixel/s with a total exposure time of 6 s, for an accumulated electron dose of 37.4 e − /Å2. Acquisitions were performed semi-automatically with Serial EM software60.
Structure determination and refinement (CryoEM)
For the first dataset, patch motion correction of the movies frames was performed in Cryosparc v.4.2.161 and CTF correction was performed with Ctffind462 in RELION 5.063,64 (Supplementary Fig. 4). crYOLO 1.965 was used for picking particles after training a model on a manually selected subset of particles. From 9195 images, a total of 914,851 particles were extracted using a 384px box size, which ensured the inclusion of the complex with at least one dsDNA and were used for 2D classification in Cryosparc. 830,419 particles were selected and 120,000 of these particles were used for Ab initio reconstruction, followed by two rounds of heterogeneous refinement using the entire selected dataset. The class showing the most promising secondary structural features was kept and the corresponding 393,254 particles yielded an electron microscopy map with a global resolution of 3.3 Å after non-uniform refinement in Cryosparc. To further improve the map, we decided to focus on a single dsDNA molecule, corresponding to the 4-box promoter region, binding an MraZ tetramer. To this aim, we generated masks to separate between the DNA ends in the helical complex. Then, the tetramers present in the helicoidal segment in the map were extracted and merged in a single dataset. This was done by particle subtraction in RELION. Briefly, micrographs from Cryosparc were imported into RELION, and 898,615 particles from crYOLO picking were selected by 2D classification. Particle selection was further refined by 3D classification using as a reference the map refined previously in Cryosparc. 646,688 particles were kept for Refine 3D in order to obtain a consensus alignment of the particles on the helix. To identify the boundaries of the tetramers in this helix, an additional round of 3D classification (without alignment) was performed using 20 classes. Classes were regrouped according to the position of the DNA ends in the map, to define masks for isolation of the tetramer by particle subtraction. The particles were finally joined resulting in a subset of 1,937,745, which was subsequently 3D classified and refined. The resulting electron microscopy map, generated from 928,675 particles, yielded a resolution of 3.36 Å. This map showed significantly enhanced structural details of the complex compared to the Cryosparc map of the larger helix, which seemed to have an overestimated resolution, despite a similar global resolution. Finally, postprocessing of the map was performed with EMready 2.166.
For the second dataset, patch motion correction and CTF estimation were performed in Cryosparc v.4.2.161 (Supplementary Fig. 5). Warp67 was used for picking particles using the general model. From 4,002 movies, a total of 701,330 particles were extracted using a 360px box size, further used for 2D classification in Cryosparc. 217,456 particles were selected and 2D classified. Particles corresponding to the octameric structure and particles corresponding to the fiber could be unambiguously identified from the 2D classes and were selected separately. Ab Initio reconstruction was performed for both set of particles. 115,463 particles were selected for the fiber and 82,151 for the octamer and used for hetero refinement. For the DNA-Octamer structure, non-uniform refinement was performed with 49,625 particles, yielding a map with a resolution of 3,57 Å. To process the fiber, 56,929 particles were selected after Ab Initio and hetero refinement and further refined using homogenous refinement. As the fiber appeared flexible, particle subtraction was used to focus the refinement on a segment corresponding to 2 DNA molecules and further refined. As the picking using the Warp general model seemed incomplete on the fibers in the micrographs, templates were generated from the map and used for template picking. 710,303 particles were selected, 2D classified, and 183,021 particles were selected for further Ab-Initio and 3 rounds of hetero refinement. 46,906 particles were refined using non-uniform refinement, yielding a map at 3.82 Å resolution. Particles were re-extracted with a box size of 512px to incorporate a larger portion of the fiber. Those particles yielded an electron microscopy map with a global resolution of 3.87 Å after a last round of non-uniform refinement. Finally, postprocessing of the maps was performed with EMready 2.166.
Model-building of DNA-MraZ atomic in all structures was performed manually based on the cryo-EM density map by using COOT58. The structure obtained by crystallography of the MraZ oligomer was used as a template to generate the structure. The model was then refined against the cryo-EM density maps using phenix.real space_refine59. More details are shown in Supplementary Figs. 4–6 and Table 2. The PDB structures of MraZ have been deposited in the Protein Data Bank with the PDB code 9R4J, 9SXZ and 9SZ7 and the maps have been deposited in EMDB with the codes EMD-53569, 55332 and 55361.
Electrophoretic mobility shift assay
For DNA binding experiments, the buffer contained 50 nM NaCl, 20 mM Tris pH 8, 50% Glycerol and 1 mM EDTA. 50 nM DNA probes labeled with 3’-Fluorescein were incubated with increasing protein concentrations, ranging from 10 to 400 nM, at RT for 30 min and then loaded into a 5% Acrylamide (40% Acrylamide/Bis 37.5:1)-TAE gel. Electrophoresis was performed for 120 min, 80 V at 4 °C. For imaging, ChemiDoc MP (Biorad) was used. Image Lab was used for quantification.
Reporter Assay
GFP gene was cloned under the Lac promoter fused to the wild type or mutant MraZ binding boxes. BL21 cells were co-transformed with the different GFP plasmids and a plasmid expressing MraZ. Cultures were grown O/N at 37 °C 250 rpm. Cells were centrifuged and washed twice with PBS, finally resuspended at the same concentration. Fluorescence emission was measured using a Jasco FP-8200 spectrofluorometer at 395 nm excitation and the measurement range was 460 – 560 nm.
Transcription start site (TSS) determination
TSS were determined using the Cappable-seq method described by Ettwiller et al., 2016. Briefly, 2,95 µg of each one of three RNA samples purified from separate M. genitalium cultures were submitted to the capping with desthiobiotin-GTP. Capped RNA were fragmented and enriched with two rounds of streptavidin AMPure beads. Samples were decapped using RppH before library preparation using the NEBNext Small RNA library. Libraries were sequenced in an Illumina MySeq. Reads were aligned to the M. genitalium genome and TSS were finally determined by using the pipeline in https://github.com/Ettwiller/TSS68.
RNA extraction
Total RNA was extracted from bacterial pellets using Trizol (Invitrogen), according to the manufacturer’s recommendations. Briefly, bacterial pellets are resuspended in TriZOL and homogenized using a 20 G syringe 10 times/sample, followed by an additional 10 times/sample using a 10 G syringe. Incubate 5 mins and add 200 µL of chloroform. Centrifuge for 15 mins at maximum speed and 4 °C and keep the aqueous superior phase. Add approximately the same volume of isopropanol and incubate 10 mins. Centrifuge for 25 mins, maximum speed, 4 °C. Wash pellet twice with cold ethanol 75% and let it dry at RT at least one hour. Reconstitute pellets with 50 μL of RNase-free water. Quantify by Nanodrop N−1000 (Thermo Fisher).
RT-qPCR
500 ng of RNA were used to generate cDNA using the qScript™ cDNA Supermix according to the manufacturer’s protocol instructions (Quanta Bioscience). cDNA was used to analyze mRNA expression levels by RT-qPCR using Fast SYBR Green Master Mix (Applied Biosystems) and specific primers (Supplementary Table 1), following the manufacturers recommendations. CysG was used as a normalization control. Statistical analyses were performed using GraphPad Prism 5 (GraphPad Software Inc.). One-way analysis of variance (ANOVA) with Tukey post hoc test was used for the comparison of multiple groups. Exact p-values are indicated in the figures.
Western blot
SDS/PAGE, proteins were transferred for 30 min onto PVDF membranes (Millipore), which were blocked with 5% (v/v) non-fat milk in TBS-T [TBS containing 0.1% (v/v) Tween20] for 1 h. Membranes were incubated overnight at 4 °C with the primary antibody (see below) in TBS-T containing 5% (v/v) non-fat milk, washed three times with TBS-T, and incubated for a further 1 h in TBS-T containing 5% (v/v) non-fat milk containing the appropriate secondary antibody. After three washes with TBS-T, membranes were processed with Immobilon® Forte Western Blotting HRP Substrate (Merck Millipore, WBLUF0100) in the Li-Cor Odyssey XF imager. Primary antibodies used in Western blotting were anti-His tag antibody (ThermoFisher, MA1-21315, 1:2000), HRP-conjugated Goat Anti-Mouse (H + L) (Biorad, 1706516, 1:5000).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Coordinates from crystal structures reported here have been deposited in the Protein Data Bank under accession codes 9QLG (MraZ octamer) and 9QLR (MraZ nonamer). CryoEM coordinates and maps have been deposited in the Protein Data Bank under accession codes 9R4J and EMD-53569 (MraZ Fiber 1), 9SZ7 and EMD-55361 (MraZ Fiber 2), and 9SX6 and EMD-55332 (MraZ Octamer-DNA). Other Protein Data Bank accession codes used in this study: 1N0E (MraZ M.pneumoniae), 2W1T (SpoVT), 1Z0R (AbrB) and 2FY9] (AbH). All other data supporting the findings of this study are available within the article and its supplementary information files. Source data are provided in this paper.
References
Hutchison, C. A. et al. Global transposon mutagenesis and a minimal mycoplasma genome. Science 286, 2165–2169 (1999).
Glass, J. I. et al. Essential genes of a minimal bacterium. Proc. Natl. Acad. Sci. USA 103, 425–430 (2006).
Lluch-Senar, M. et al. Defining a minimal cell: essentiality of small ORFs and ncRNAs in a genome-reduced bacterium. Mol. Syst. Biol. 11, 780 (2015).
Errington, J. et al. Cytokinesis in bacteria. Microbiol. Mol. Biol. Rev. 67, 52–65 (2003).
Weiss, D. S. Bacterial cell division and the septal ring. Mol. Microbiol. 54, 588–597 (2004).
Goehring, N. W. & Beckwith, J. Diverse paths to midcell: assembly of the bacterial cell division machinery. Curr. Biol. 15, R514–R526 (2005).
Tamames, J. et al. Bringing gene order into bacterial shape. Trends Genet. 17, 124–126 (2001).
Vicente, M. et al. Regulation of transcription of cell division genes in the Escherichia coli dcw cluster. Cell. Mol. Life Sci. 54, 317–324 (1998).
Eraso, J. M. et al. The highly conserved MraZ protein is a transcriptional regulator in Escherichia coli. J. Bacteriol. 196, 2053–2066 (2014).
Alarcón, F. et al. Genes involved in cell division in mycoplasmas. Genet. Mol. Biol. 30, 174–181 (2007).
Benders, G. A. et al. Transcriptional Analysis of the Conserved ftsZ Gene Cluster in Mycoplasma genitalium and Mycoplasma pneumoniae. J. Bacteriol. 187, 4542–4551 (2005).
Martínez-Torró, C. et al. Functional characterization of the cell division gene cluster of the wall-less bacterium mycoplasma genitalium. Front. Microbiol. 12, 695572 (2021).
Lluch-Senar, M. et al. Cell division in a minimal bacterium in the absence of ftsZ. Mol. Microbiol. 78, 278–289 (2010).
Fisunov, G. Y. et al. Binding site of MraZ transcription factor in Mollicutes. Biochimie 125, 59–65 (2016).
White, M. L. et al. MraZ Transcriptionally controls the critical level of FtsL required for focusing Z-rings and kickstarting septation in Bacillus subtilis. J. Bacteriol. 204, e00243-22 (2022).
Maeda, T. et al. RNase III mediated cleavage of the coding region of mraZ mRNA is required for efficient cell division in Corynebacterium glutamicum. Mol. Microbiol. 99, 1149–1166 (2016).
Xu, X. et al. Beyond a ribosomal RNA methyltransferase, the wider role of MraW in DNA methylation, motility and colonization in Escherichia coli O157:H7. Front. Microbiol. 10, 2520 (2019).
Kyuma, T. et al. Ribosomal RNA methyltransferases contribute to Staphylococcus aureus virulence. FEBS J. 282, 2570–2584 (2015).
Kimura, S. & Suzuki, T. Fine-tuning of the ribosomal decoding center by conserved methyl-modifications in the Escherichia coli 16S rRNA. Nucleic Acids Res. 38, 1341–1352 (2010).
Adams, D. W. & Errington, J. Bacterial cell division: assembly, maintenance and disassembly of the Z ring. Nat. Rev. Microbiol. 7, 642–653 (2009).
Busiek, K. K. & Margolin, W. Bacterial actin and tubulin homologs in cell growth and division. Curr. Biol. 25, R243–R254 (2015).
Fujita, J. et al. Crystal structure of FtsA from Staphylococcus aureus. FEBS Lett. 588, 1879–1885 (2014).
Vicente, M. & Rico, A. I. The order of the ring: assembly of Escherichia coli cell division components. Mol. Microbiol. 61, 5–8 (2006).
Dai, K. & Lutkenhaus, J. The proper ratio of FtsZ to FtsA is required for cell division to occur in Escherichia coli. J. Bacteriol. 174, 6145–6151 (1992).
Dewar, S. J. et al. Inhibition of cell division initiation by an imbalance in the ratio of FtsA to FtsZ. J. Bacteriol. 174, 6314–6316 (1992).
Bisson-Filho, A. W. et al. Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division. Science 355, 739–743 (2017).
Dai, K. & Lutkenhaus, J. ftsZ is an essential cell division gene in Escherichia coli. J. Bacteriol. 173, 3500–3506 (1991).
Beall, B. & Lutkenhaus, J. FtsZ in Bacillus subtilis is required for vegetative septation and for asymmetric septation during sporulation. Genes Dev. 5, 447–455 (1991).
Trespidi, G. et al. Molecular characterization of the Burkholderia cenocepacia dcw operon and FtsZ interactors as new targets for novel antimicrobial design. Antibiotics 9, 841 (2020).
Nyongesa, S. et al. Evolution of longitudinal division in multicellular bacteria of the Neisseriaceae family. Nat. Commun. 13, 4853 (2022).
Wang, C. et al. Structural insights into the PrpTA toxin–antitoxin system in Pseudoalteromonas rubra. Front. Microbiol. 13, 1053255 (2022).
Sultan, M. et al. Targeting the G-quadruplex as a novel strategy for developing antibiotics against hypervirulent drug-resistant Staphylococcus aureus. J. Biomed. Sci. 32, 15 (2025).
Chen, S. et al. Crystal structure of a protein associated with cell division from Mycoplasma pneumoniae (GI: 13508053): a novel fold with a conserved sequence motif. Proteins 55, 785–791 (2004).
Adams, M. A. et al. MraZ from Escherichia coli: cloning, purification, crystallization and preliminary X-ray analysis. Acta Crystallogr. F Struct. Biol. Cryst. Commun. 61, 378–380 (2005).
Bobay, B. G. et al. Revised structure of the AbrB N-terminal domain unifies a diverse superfamily of putative DNA-binding proteins. FEBS Lett. 579, 5669–5674 (2005).
Sullivan, D. M. et al. Insights into the nature of DNA binding of AbrB-like tanscription factors. Structure 16, 1702–1713 (2008).
Coles, M. et al. AbrB-like transcription factors assume a swapped hairpin fold that is evolutionarily related to double-Psi β barrels. Structure 13, 919–928 (2005).
Raumann, B. E. et al. DNA recognition by beta-sheets in the Arc repressor-operator crystal structure. Nature 367, 754–757 (1994).
Somers, W. S. & Phillips, S. E. V. Crystal structure of the met represser–operator complex at 2.8 Å resolution reveals DNA recognition by β-strands. Nature 359, 387–393 (1992).
Costa, M. et al. Plasmid transcriptional repressor CopG oligomerises to render helical superstructures unbound and in complexes with oligonucleotides1. J. Mol. Biol. 310, 403–417 (2001).
Gomis-Rüth, F. X. et al. The structure of plasmid-encoded transcriptional repressor CopG unliganded and bound to its operator. EMBO J. 17, 7404–7415 (1998).
Stefanucci, A. et al. A novel β-hairpin peptide derived from the ARC repressor selectively interacts with the major groove of B-DNA. Bioorg. Chem. 112, 104836 (2021).
Weber, L. et al. The conserved Dcw gene cluster of R. sphaeroides is preceded by an uncommonly extended 5’ leader featuring the sRNA UpsM. PLoS ONE 11, e0165694 (2016).
Alva, V. et al. A vocabulary of ancient peptides at the origin of folded proteins. ELife 4, e09410 (2015).
Coles, M. et al. Common evolutionary origin of swapped-hairpin and double-psi beta barrels. Structure 14, 1489–1498 (2006).
Asen, I. et al. Crystal structure of SpoVT, the final modulator of gene expression during spore development in Bacillus subtilis. J. Mol. Biol. 386, 962–975 (2009).
Dong, T. C. et al. DNA-binding studies on the Bacillus subtilis transcriptional regulator and AbrB homologue, SpoVT. FEMS Microbiol. Lett. 233, 247–256 (2004).
Chumsakul, O. et al. Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation. Nucleic Acids Res. 39, 414–428 (2011).
Olson, A. L. et al. Structure and DNA-Binding Traits of the Transition State Regulator AbrB. Structure 22, 1650–1656 (2014).
Vaughn, J. L. et al. The DNA-binding domain in the Bacillus subtilis transition-state regulator AbrB employs significant motion for promiscuous DNA recognition. J. Mol. Biol. 305, 429–439 (2001).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–W388 (2005).
Kuriata, A. et al. Aggrescan3D (A3D) 2.0: prediction and engineering of protein solubility. Nucleic Acids Res. 47, W300–W307 (2019).
Bachman, J. Chapter Ninteen - Site-Directed Mutagenesis. In Methods in Enzymology 529, 241–248, (Academic Press, 2013)
Juanhuix, J. et al. Developments in optics and performance at BL13-XALOC, the macromolecular crystallography beamline at the Alba Synchrotron. J. Synchrotron. Rad. 21, 679–689 (2014).
McCarthy, A. A. et al. ID30B – a versatile beamline for macromolecular crystallography experiments at the ESRF. J. Synchrotron. Rad. 25, 1249–1260 (2018).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Cryst. D 67, 293–302 (2011).
Emsley, P. et al. Features and development of Coot. Acta Cryst. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Cryst. D 66, 213–221 (2010).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A. et al. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kimanius, D. et al. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Kimanius, D. et al. Data-driven regularization lowers the size barrier of cryo-EM structure determination. Nat. Methods 21, 1216–1221 (2024).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 1–13 (2019).
He, J. et al. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Ettwiller, L. et al. A novel enrichment strategy reveals unprecedented number of novel transcription start sites at single base resolution in a model prokaryote and the gut microbiome. BMC Genom. 17, 199 (2016).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2024).
Acknowledgements
This work was supported by the Spanish Ministry of Science and Innovation (MICINN) PID2024-160233OB-I00 to D.R., PID2024-156687OB-I00 to V.A. and PID2024-159663OB-C21 to J.P.; and by ICREA to DR, ICREA-Academia-2022, from Generalitat de Catalunya. L.S.A. acknowledges her FPI fellowship from the Spanish Government (PRE2019-088509). D.R. acknowledges support from the Serra Hunter program from Generalitat de Catalunya. X-ray experiments were performed at BL-13 XALOC beamline at ALBA Synchrotron with the collaboration of ALBA staff. This work benefited from access to IGBMC, an Instruct-ERIC Center. Financial support was provided by Instruct-ERIC internship APPID 3074 and PID 27465. We thank the Microscopy Services (UAB). We thank Nils Marechal (IGBMC-CBI) for his help with cryoEM sample preparation, data collection and insightful discussions. We also thank Pablo Guerra (IBMB-CSIC, JEMCA, ALBA Synchrotron) for his help in obtaining preliminary cryoEM data.
Author information
Authors and Affiliations
Contributions
L.S.A. conducted crystallization experiments. A.D, L.S.A., and J.G.P. performed the cryoEM analysis. J.P. analyzed the promotor sequences. L.S.A. and N.V. conducted all in vitro activity assays. M.C-C. and V.A. conducted qPCR analysis. D.R., N.V., and L.S.A. contributed to the correction and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Naruhiko Adachi, Vadim Govorun and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sánchez-Alba, L., Varejão, N., Durand, A. et al. Structural basis for transcriptional regulation by the cell division regulator MraZ in Mycoplasma genitalium. Nat Commun 17, 2132 (2026). https://doi.org/10.1038/s41467-026-68809-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68809-2