Abstract
Maternal mitochondrial inheritance is secured by mechanisms that exclude paternal mitochondrial DNA (mtDNA). While, epigenetic modifications are vital for spermatogenesis and embryo development, their roles in the paternal mitochondrial elimination (PME) remain poorly understood. Here, we identify ALKB-1, a DNA/RNA demethylase, as a pivotal factor for efficient PME in Caenorhabditis elegans (C. elegans), acting through ALKB-1-dependent modulation of tRNA m1A methylation. Mechanistically, ALKB-1 inactivation leads to m1A hypermethylation of tRNA, which subsequently disrupts protein translation, impairs mitochondrial proteostasis, and increases ROS levels. This cascade activates the oxidative stress response factor SKN-1/Nrf2 and initiates the mitochondrial unfolded protein response (UPRmt) through ATFS-1, causing accumulation of mitochondria and mtDNA in sperm, which ultimately impedes efficient paternal mitochondrial removal and negatively impacts male fertility and embryonic development. Our findings describe a mechanism whereby ALKB-1-mediated tRNA m1A epitranscriptomic modifications are necessary for maintaining mitochondrial quality control, thereby influencing PME efficiency, underscoring the importance of this epitranscriptomic stress checkpoint in upholding proper mitochondrial inheritance during reproduction.
Similar content being viewed by others
Introduction
The coordination of nuclear and mitochondrial genome expression is crucial for maintaining mitochondrial function and homeostasis, despite their unique inheritance patterns1,2. Maternal inheritance of the mitochondrial genome (mtDNA) is predominant in eukaryotes, except for certain fungi and mussels. Paternal mitochondria are selectively eliminated during spermatogenesis and early embryogenesis, a key step for maintaining maternal mtDNA3,4,5. Accumulation of mutations and heteroplasmy in mtDNA is linked to a spectrum of mitochondrial pathologies, including diabetes and Parkinson’s disease6. Maternal transmission of mtDNA hinders the spread of harmful mutations to subsequent generations6,7. Multiple mechanisms involving mitochondrial endonuclease G (EndoG) in Caenorhabditis elegans (C. elegans) and Drosophila8,9, the mitochondrial transcription factor A (TFAM) in human sperm3, and autophagy10,11,12,13,14,15,16,17, mitophagy-lysosome18,19,20, and ubiquitin-proteasome pathways21,22,23, have been implicated in the elimination of paternal mitochondria.
Epigenetic modifications, including DNA or RNA methylation, are crucial for regulating spermatogenesis and embryo development24,25,26. Changes in RNA methylation are linked to sperm motility27, while alterations in DNA methylation patterns are associated with increased sperm mtDNA copy number and abnormalities28,29. Sperm cells are characterized by high levels of DNA methylation, which are essential for their development and function30. Nevertheless, the role of methylation-related epigenetic modifications in paternal mitochondrial elimination (PME) and the relevant mechanisms involved are not well understood.
In this study, we investigated the role of ALKBH1, or ALKB-1, in C. elegans PME. ALKBH1 is a dioxygenase with broad demethylation activity that depends on Fe (II) and α-ketoglutarate, and affects both DNA and RNA. Its importance in gene regulation, translation, and mitochondrial function is well-documented31,32,33,34. Studies have shown that Alkbh1 null mice exhibit high embryonic lethality, sexual dimorphism, infertility, and developmental issues in the eyes, skeleton, and genitalia31,35,36. Additionally, ALKBH1 has been shown to modulate gene expression during spermatogenesis and early embryonic stages by changing methylation patterns, underscoring its fundamental biological role in both pre- and post-fertilization processes31,37.
Our study identified ALKB-1 is essential for efficient PME in C. elegans, through its tRNA N1-methyladenosine (m1A) demethylase activity, separate from the DNA N6-methyldeoxyadenosine (6 mA) demethylase activity reported previously38. Disrupting ALKB-1 function perturbed mitochondrial protein homeostasis, which, in turn, activated transcription factors SKN-1/Nrf2 and the mitochondrial unfolded protein response (UPRmt). Loss of ALKB-1 promotes “clonal expansion” of paternal mitochondria and mtDNA, consequently delaying the PME process and negatively impacting sperm fertility and embryo viability. Our study characterizes the role of epigenetic modification in PME and offers insights into the mechanisms of maternal mitochondrial inheritance. These findings advance our understanding of mitochondrial diseases and may inform future therapeutic strategies.
Results
The demethylase ALKB-1 modulates paternal mitochondrial elimination dynamics
To monitor PME in embryos, we employed the mptDf1 mtDNA allele, with a 179 bp deletion, which is smaller than the 3053 bp deletion present in the uaDf5 mtDNA allele used in our previous study (Supplementary Fig. 1a)9,21. We used a nested PCR assay to track the presence of sperm-derived mtDNA during embryogenesis (Supplementary Fig. 1b–d). Crossing mptDf1/+ heteroplasmic males with N2 hermaphrodites revealed that mptDf1 mtDNA is present in early-stage embryos but disappeared before the 64-cell stage (Supplementary Fig. 1e), consistent with previous reports of early paternal mtDNA elimination21,39.
To investigate the function of methylation in the PME process, we conducted an RNA interference (RNAi) screen against potential homologs of the ALKB demethylase family in the C. elegans genome, including B0564.2, F09F7.7, alkb-1, alkb-7, and alkb-8 (Supplementary Fig. 1f). Notably, only alkb-1 knockdown resulted in delayed elimination of sperm-derived mptDf1 mtDNA, which was detectable up to the 1.5-fold stage of embryonic development (Fig. 1a, b, Supplementary Fig. 1f). Subsequent microscopic analyses showed a substantial increase in the number of MitoTracker Red (MTR)-labeled paternal mitochondria in the 32- and 64-cell stage embryos, after alkb-1 RNAi, compared to control RNAi (Fig. 1c, d, Supplementary Fig. 1g, h). These results collectively suggested that alkb-1 knockdown significantly delayed the PME process in cross-fertilized embryos. High mtDNA heteroplasmy has been shown to promote mtDNA replication40, potentially impacting its clearance. To determine whether ALKB-1’s contribution to PME delay involves elevated heteroplasmy, we quantified mptDf1 mtDNA levels in mptDf1/+; him-5 (e1467) males treated with control or alkb-1 RNAi. No significant difference was detected (Supplementary Fig. 1i, j), indicating that ALKB-1’s effect on PME occurs independently of changes in heteroplasmy.
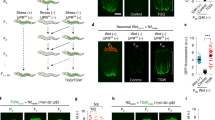
a mptDf1 mtDNA-based PCR assays to monitor persistent mptDf1 paternal mtDNA in several stage cross-fertilized embryos from mating of mptDf1/+; him-5 (e1467) males with wild-type N2 hermaphrodites under empty vector or alkb-1 RNAi conditions. M, DNA marker. b Validation of RNAi-mediated knockdown by qPCR (n = 9 biologically independent samples). EV, empty vector. c Representative differential interference contrast (DIC) and MTR-stained paternal mitochondrial images of the 32- and 64-cell stage cross-fertilized embryos from mating of MTR-stained N2 males with hermaphrodites under empty vector (EV) or alkb-1 RNAi conditions, with scale bars of 10 μm. d Quantification of the number of MTR-labeled mitochondrial clusters in the 32- and 64-cell stage cross-fertilized embryos from mating of MTR-stained N2 males with hermaphrodites under EV or alkb-1 RNAi conditions. n = 11 (EV, 32- and 64-cell), n = 15 (alkb-1 RNAi, 32- and 64-cell). e Protein sequence alignment between C. elegans ALKB-1 and vertebrate ALKBH1. Yellow circle, the conserved ion-coordinating residues. f Schematic diagram showing the mutation site of alkb-1 D247A enzyme mutant. g mptDf1 mtDNA-based PCR assays to monitor persistent mptDf1 paternal mtDNA in several stage cross-fertilized embryos from the indicated crosses. M, DNA marker. h, j Representative DIC and MTR-stained paternal mitochondrial images of the 32- and 64-cell stage embryos from cross-fertilization of the indicated strain males (stained by MTR) with hermaphrodites, with scale bars of 10 μm. i, k Quantification of the number of MTR-labeled mitochondrial clusters in the 32- and 64-cell stage embryos from cross-fertilization of the indicated strain males (stained by MTR) with hermaphrodites in (h) and (j), respectively. In (i), n = 14 (N2, 32-cell), n = 12 (N2, 64-cell), n = 15 (alkb-1D247A, 32- and 64-cell), n = 14 (alkb-1D247A; tbb-2p::ALKB-1, 32-cell), n = 11 (alkb-1D247A; tbb-2p::ALKB-1, 64-cell). In (k), n = 14 (N2, 32-cell), n = 12 (N2, 64-cell), n = 20 (alkb-1D247A, 32-cell), n = 18 (alkb-1D247A, 64-cell), n = 15 (alkb-1D247A males × N2 hermaphrodites, 32-cell), n = 14 (alkb-1D247A males × N2 hermaphrodites, 64-cell), n = 14 (N2 males × alkb-1D247A hermaphrodites, 32- and 64-cell). In (d, i, k), embryos were derived from 3 to 5 independent biological crosses per condition. The experiment was performed twice with similar results. Data were presented as mean ± SD. p-values were determined by unpaired two-tailed t-test (b, d), and Ordinary one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (i, k). Source data are provided as a Source data file.
ALKBH1/ALKB-1 has been reported to localize to the mitochondria, where it is involved in mitochondrial protein synthesis, respiration, and the maintenance of mutant mtDNA in mammals and C. elegans38,41. The probability of ALKB-1 mitochondrial localization by MitoProt is 0.4168 (Supplementary Fig. 2a). Western blot analyses of tbb-2p::ALKB-1::3×FLAG transgenic strains confirmed the presence of ALKB-1 in mitochondria, while deletion of mitochondrial targeting peptide sequence (MTS, Δ2-35) abolished this localization (Supplementary Fig. 2b). Several attempts to directly visualize ALKB-1 in C. elegans have been unsuccessful, likely due to relatively low protein level. To overcome this limitation, we utilized ectopic expression in HeLa cells and observed that ALKB-1 is localized to both the nucleus and mitochondria (Supplementary Fig. 2c, e), with MTS deletion similarly disrupting mitochondrial targeting (Supplementary Fig. 2d, f). Thus, our analyses, implementing both western blotting and direct visualization techniques, demonstrated that ALKB-1 exhibited localization in the nucleus, cytosol, and mitochondria. This finding deviates from a previous report that suggested ALKB-1 was solely localized to mitochondria38. We speculated that this discrepancy might be attributed to the use of different promoters to drive transgene expression in our study, compared to the earlier report38. Nevertheless, our results were in agreement with another study that described similar localization patterns41. Cumulatively, these findings provided compelling evidence for the localization of ALKB-1 in the mitochondria, although not exclusively. The localization in the nucleus, cytosol, and mitochondria indicates a more complex role for ALKB-1 in cellular processes than previously recognized.
Given that ALKBH1 is a known 6mA demethylase dependent on the ferrous iron-coordinating residue D23333,42,43, and sequence alignment revealed its conserved counterpart, D247, in ALKB-1 (Fig. 1e, Supplementary Fig. 3a), we purified both full-length ALKB-1 protein and a catalytically inactive variant (D247A), and tested for demethylase activity. In vitro demethylation assays showed that recombinant wild-type ALKB-1 proteins could decrease 6mA levels in double-stranded oligonucleotides, while the mutant could not (Supplementary Fig. 3b, c), indicating that C. elegans ALKB-1 possesses 6mA demethylase activity in vitro, similar to ALKBH1, and that D247 is the key catalytic residue.
To delineate the precise role of ALKB-1 in PME, we employed CRISPR/Cas9 to generate a series of alkb-1 mutant C. elegans strains, including an alkb-1 knockout strain (alkb-1 (ajz209)/+), an MTS deletion strain (alkb-1 (ajz102)/+), and a demethylase inactive mutant strain, with the D247A substitution (alkb-1 (ajz103)) (Fig. 1f, Supplementary Fig. 3d). Utilizing an mptDf1 mtDNA-based PCR assay, we observed that strains ajz209/+ and ajz103, but not ajz102/+, showed a significant delay in PME (Fig. 1g, Supplementary Fig. 3e). Microscopic analyses further confirmed these findings (Fig. 1h, i, Supplementary Fig. 3f). As alkb-1-/- is not viable, we focused on the ajz103 (alkb-1D247A mutant) to elucidate the regulatory mechanisms of ALKB-1 demethylase activity in PME. Re-expression of either wild-type ALKB-1 or its MTS-deleted variant rescued the PME defect in alkb-1D247A mutant, whereas the catalytically inactive D247A variant did not (Supplementary Fig. 4a, b). These findings collectively underscore the pivotal role of ALKB-1’s demethylase activity, independent of its mitochondrial localization, in preventing transmission of paternal mitochondria to offspring.
To further explore the requirement for ALKB-1 in PME across different lineages, we conducted microscopic assays. Paternal alkb-1 mutations resulted in a significant increase in paternal mitochondria in 64-cell stage embryos compared with wild-type controls. Embryos from mothers carrying alkb-1 mutations showed a minor delay in PME, with increased MTR-labeled mitochondrial clusters, visible at the 32-cell stage only (Fig. 1j, k). To evaluate the relative contribution of ALKB-1 in the sperm, we conducted rescue experiments in alkb-1D247A mutants. Sperm-specific expression of ALKB-1 significantly alleviated PME defects, whereas germline-specific expression had a minor and statistically insignificant effect (Supplementary Fig. 4c, d). These findings suggest that ALKB-1 catalytic activity contributes to PME in both lineages, with a more pronounced impact in the paternal lineage, particularly in sperm. This observation prompted our hypothesis that ALKB-1 deficiency might interfere with the typical decrease in sperm mitochondria and mtDNA content during spermatogenesis. Indeed, there was a substantial increase in the number of mitochondria and mtDNA in the sperm of alkb-1D247A mutants, compared to wild-type controls (Supplementary Fig. 4e, f). Overall, our results demonstrate the critical role of ALKB-1 demethylase activity in the PME process, during spermatogenesis and early embryogenesis.
ALKB-1 regulates mitochondrial proteostasis through tRNA m1A modification
In humans, ALKBH1 is known to catalyze DNA 6 mA demethylation43, which we confirmed through in vitro biochemical assays. However, Dot blot and LC-MS analyses showed no significant changes in 6 mA levels in genomic DNA (gDNA) or mtDNA from the alkb-1D247A mutants (Supplementary Fig. 5a–c), contradicting a recent report that identified ALKB-1 as a mtDNA 6 mA demethylase38. This discrepancy could be attributed to the sensitivity limitations of our detection methods, which may potentially be insufficient to detect changes in 6 mA levels in the alkb-1D247A mutants. To further ascertain the role of 6 mA in PME, we overexpressed the methyltransferase DAMT-1, which is known to increase 6 mA levels in both gDNA and mtDNA38. Surprisingly, DAMT-1 overexpression did not affect the PME process (Supplementary Fig. 5d). Collectively, these data indicate that the ALKB-1-mediated delay in PME is not related to its DNA 6 mA demethylase activity.
Considering the role of mammalian ALKB family members in RNA demethylation44, we measured N6-methyladenosine (m6A) mRNA levels in the alkb-1D247A mutant and found no significant differences compared to the wild-type (Supplementary Fig. 5e, f). In light of a recent study demonstrating the demethylation of m1A in tRNA by ALKBH1 in mammals33, we investigated the potential role of ALKB-1 in tRNA m1A demethylation. LC-MS analyses of tRNA from the alkb-1D247A mutant showed significant elevation of m1A modification (Fig. 2a, b). These findings were further corroborated by dot blot assays, using two distinct m1A-specific antibodies from distinct companies, yielding consistent results (Fig. 2c, d, Supplementary Fig. 5g, h). Additionally, in vitro demethylation assays demonstrated that recombinant wild-type ALKB-1 protein significantly reduced m¹A modification in stem-loop tRNA probes, whereas the D247A variant showed no such activity (Supplementary Fig. 5i). Taken together, these findings demonstrated that ALKB-1 demethylates tRNA m1A in C. elegans.
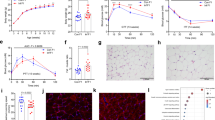
a Workflow for isolating and extracting tRNA from N2 and alkb-1D247A worms for tRNA modification analyses using LC-MS. b Quantification of the tRNA m1A modifications in N2 and alkb-1D247A worms using LC-MS (n = 2). Dot blot (c) and quantification (d) of tRNA m1A levels from N2 and alkb-1D247A worms (n = 3). tRNA loading was detected using methylene blue (Meth-Blue), and m1A levels were quantified using an m1A antibody (D345-3, MBJ Life Science). e Heatmap of cluster analysis for differentially modified m1A tRNAs in N2 vs alkb-1D247A worms from tRNA m1A seq analysis. |ΔScore | > 0.05. GSEA plots showing the Oxidative phosphorylation (OXPHOS) (f) and TCA cycle (g) pathways from RNC-seq data. h Principal component analysis (PCA) of proteins from N2 and alkb-1D247A worms. i Heatmap of cluster analysis for differentially expressed proteins in N2 vs alkb-1D247A worms from proteomics analysis. j Volcano plot of differentially expressed proteins in alkb-1D247A worms compared with N2 worms, including 121 upregulated and 47 downregulated proteins. MRPL-13 (deep blue circle) and 15 other proteins (deep red circle) were identified as significantly dysregulated proteins with mitochondrial localization. In (f, g), differential expression analysis was performed using the DESeq2 Wald test (two-sided), p-values were adjusted for multiple testing using the Benjamini–Hochberg false discovery rate (FDR) method. In (j), Fisher’s exact test was used to analyze the significance of functional enrichment of differentially expressed proteins. Functional terms with Fold enrichment >1.3 and p-values < 0.05 were considered as significant. FDR was adjusted to <1%. Data were represented as mean ± SD. p-values were determined by an unpaired two-tailed t-test (b, d). Source data are provided as a Source data file.
To further elucidate the role of ALKB-1 in tRNA m1A modification, we performed tRNA m1A sequencing. These analyses revealed significantly elevated m¹A levels in 11 tRNA isodecoders and marked reductions in 4 others. Further analyses of the upregulated isodecoders revealed 9 cytosolic and 2 mitochondrial tRNAs (Fig. 2e), with 8 of the 9 cytosolic species being tRNA-Gly (TCC/GCC) isoacceptors (Supplementary Fig. 5j, k), indicating ALKB-1 preferentially demethylates m¹A in tRNA-Gly. Collectively, these findings indicated that ALKB-1 demethylates m¹A on both cytosolic and mitochondrial tRNAs, with a primary preference for cytosolic substrates. Concurrently, tRNA sequencing detected no significant change in the abundance of any identified tRNAs (Supplementary Fig. 5l), a finding validated by biotin-labeled Northern blot analyses of six selected tRNAs (Gly-TCC, iMet-CAT, Asp-GTC, Gln-TGG, Lys-CTT, and Ser-AGA) (Supplementary Fig. 5m), which revealed comparable levels between wild-type and alkb-1D247A worms.
The m1A modification in tRNA is crucial for tRNA stability, translation, and protein synthesis45. We conducted ribosome nascent-chain complex sequencing (RNC-seq) and proteomics analysis, revealing a substantial impact of ALKB-1 on translation. RNC-seq analysis revealed a significant translation efficiency (TE) alteration in alkb-1D247A worms compared to the wild-type N2, with 471 translated transcripts upregulated and 140 transcripts downregulated (Supplementary Fig. 5n). These findings indicate that hypermethylation of tRNA at position m1A might contribute to TE alterations. Gene Ontology (GO) analysis showed that the upregulated genes are primarily involved in cGMP biosynthesis, metabolic processes, and the regulation of cellular and biological processes (Supplementary Fig. 5o). Gene Set Enrichment Analysis (GSEA) revealed an enrichment of gene sets associated with oxidative phosphorylation (OXPHOS) (NES = −2.209, p-value = 0.008) and citrate cycle (TCA cycle) (NES = −1.925, p-value = 0.008), in wild-type animals (Fig. 2f, g). Proteomics analysis identified 121 upregulated and 47 downregulated proteins in alkb-1D247A worms, versus wild-type N2 worms (Fig. 2h, i). Importantly, MRPL-13, a component of the mitochondrial ribosome, was significantly less abundant in alkb-1D247A worms (Fig. 2j), hinting at a possible impairment of mitochondrial translation. Consistent with this, RNA-seq and qPCR validation showed elevated mRNA levels of most mtDNA-encoded genes (Supplementary Fig. 6a, b), consistent with a compensatory response to translational inhibition, as previously reported46,47,48. Subcellular localization analyses, using the WoLF PSORT software, identified 15 nuclear-encoded proteins that localize in mitochondria and were significantly upregulated (Fig. 2j), indicating enhanced mitochondrial protein import. This upregulation may be associated with disrupted proteostasis and potential abnormal protein aggregation. To investigate this, we employed a well-established NS-mtGFP reporter system in C. elegans by generating a transgenic strain with the construct myo-3p::SDHB-1 MTS::gfp (S65C) (no stop codon)::unc-54 3’UTR AAAAn (Supplementary Fig. 6c), a method validated in yeast and mammalian models49,50,51. Imaging analyses revealed punctate mitochondrial aggregates in alkb-1D247A body wall muscle, contrasting with the uniform GFP distribution in controls (Supplementary Fig. 6d), providing direct evidence of aberrant protein aggregation. Collectively, these findings indicated that ALKB-1-mediated m¹A modification of tRNA influences both cytoplasmic and mitochondrial translation. ALKB-1 inhibition enhances cytoplasmic translation efficiency while simultaneously impairing mitochondrial translation, thereby disrupting the balance of protein homeostasis between the cytoplasm and mitochondria.
ALKB-1 impairment delays PME through the induction of UPRmt
Imbalances in mitochondrial protein homeostasis commonly trigger UPRmt 52. To explore the effects of ALKB-1 on UPRmt, we employed the hsp-6p::gfp strain reporter in C. elegans and found that alkb-1D247A mutants exhibited robust UPRmt induction in both males (Fig. 3a, b) and hermaphrodites (Supplementary Fig. 7a, b). This effect was recapitulated by alkb-1 RNAi (Supplementary Fig. 7c, d). Silencing atfs-1, a central UPRmt regulator, effectively abolished the UPRmt triggered by alkb-1 loss, with no effect observed in wild-type backgrounds (Fig. 3a, b, Supplementary Fig. 7a–d). Thus, we deduced that UPRmt activation contributed to the delayed PME phenotype observed in alkb-1D247A mutants. Consistent with this hypothesis, deletion of atfs-1 rescued the PME defect in alkb-1D247A embryos (Fig. 3c, d), while atfs-1 mutants displayed normal PME (Supplementary Fig. 7e, f).
Representative bright-field and GFP fluorescence images (a) and quantification (b) of the hsp-6p::gfp reporter in wild-type and alkb-1D247A males treated with control or atfs-1 RNAi (n = 30 per group), with scale bars of 100 μm. The experiment was performed three times with similar results. c, e Representative DIC and MTR-stained paternal mitochondrial images of the 32- and 64-cell stage cross-fertilized embryos from the indicated crosses with scale bars of 10 μm. d, f Quantification of the number of MTR-labeled mitochondrial clusters in (c, e), respectively. In (d), n = 19 (N2, 32-cell), n = 15 (N2, 64-cell), n = 23 (alkb-1D247A, 32-cell), n = 19 (alkb-1D247A, 64-cell), n = 22 (alkb-1D247A;atfs-1, 32-cell), n = 18 (alkb-1D247A;atfs-1, 64-cell), n = 22 (alkb-1D247A;atfs-1 males × alkb-1D247A hermaphrodites, 32-cell), n = 14 (alkb-1D247A;atfs-1 males × alkb-1D247A hermaphrodites, 64-cell), n = 21 (alkb-1D247A males × alkb-1D247A;atfs-1 hermaphrodites, 32-cell), n = 17 (alkb-1D247A males × alkb-1D247A;atfs-1 hermaphrodites, 64-cell). In (f), n = 16 (N2, 32-cell), n = 15 (N2, 64-cell), n = 15 (alkb-1D247A, 32- and 64-cell), n = 14 (alkb-1D247A;dve-1, 32- and 64-cell), n = 14 (alkb-1D247A;lin-65, 32-cell), n = 13 (alkb-1D247A;lin-65, 64-cell), n = 13 (alkb-1D247A;ubl-5, 32- and 64-cell). g Schematic diagram illustrating the relationship among LONP-1, mitochondrial ATFS-1 accumulation, and mtDNA replication. h Western blot analyses showing the accumulation of mitochondrial ATFS-1 in atfs-1p::atfs-1::TY1::egfp::3×flag worms treated with control, alkb-1, cco-1, and lonp-1 RNAi. cco-1 and lonp-1 RNAi served as the positive control, and N2 worms were used as the negative control. n = 3 biological repeats. i Quantification of the levels of total mtDNA in both N2 and alkb-1D247A, including males and early embryos (eggs) (n = 9 biologically independent samples). j Quantification of the levels of total mtDNA in males of both N2 and alkb-1D247A treated with control or atfs-1 RNAi (n = 9 biologically independent samples). In (d, f), embryos were derived from 3 to 5 independent biological crosses per condition. The experiment was performed twice with similar results. Data were represented as mean ± SD. p-values were determined by Ordinary one-way ANOVA with Tukey’s multiple comparisons test (b, d, f, j), and unpaired two-tailed t-test (i). Source data are provided as a Source data file.
In C. elegans, UPRmt activation typically involves the cooperative function of the transcription factor ATFS-1, together with DVE-1, UBL-5, and LIN-65, to facilitate retrograde signaling from the mitochondria to the nucleus, during mitochondrial stress53. Consistent with this role of ATFS-1, loss of dve-1, ubl-5, or lin-65 restored normal PME in alkb-1D247A embryos (Fig. 3e, f), and mutants of these factors displayed PME comparable to wild-type worms (Supplementary Fig. 7e, f). Repression of dve-1, ubl-5, or lin-65 by RNAi also counteracted UPRmt activation, induced by alkb-1 lesion (Supplementary Fig. 7g, h). Additionally, alkb-1D247A mutants showed increased protein levels and nuclear translocation of DVE-1::GFP and UBL-5::GFP (Supplementary Fig. 7i–l). Collectively, these results demonstrate that ALKB-1 regulates the PME processes through its impact on UPRmt activation and retrograde signaling to the nucleus.
ATFS-1 possesses both mitochondrial and nuclear localization signals. Nuclear ATFS-1 activates UPRmt, while mitochondrial ATFS-1 promotes mtDNA replication, to facilitate mtDNA “clonal expansion” in dysfunctional mitochondria40,45. The protein level of ATFS-1 is primarily regulated by the ATP-dependent protease LONP-154. Upon mitochondrial damage, ATFS-1 accumulates in the mitochondria and enhances mtDNA replication by facilitating the binding of POLG to mtDNA (Fig. 3g)45. We observed that the knockdown of alkb-1 significantly increased both the mitochondrially localized form (the lower band produced after MTS cleavage54) and the unprocessed form of ATFS-1 (Fig. 3h). Remarkably, the alkb-1D247A mutation resulted in a significant elevation of mtDNA copy number in males, early embryos, and sperm (Fig. 3i, Supplementary Fig. 4f), which was dependent on ATFS-1, as evidenced by the mitigation of this effect upon atfs-1 RNAi (Fig. 3j). Additionally, absence of paternal ATFS-1 was sufficient to rescue the PME defect caused by ALKB-1 inactivation, while maternal ATFS-1 had no effect (Fig. 3c, d), highlighting the key role of ATFS-1 in pre-fertilization mtDNA amplification and PME suppression, in alkb-1D247A mutants. Taken together, these findings indicate that the delay in PME, resulting from ALKB-1 inactivation, depends on UPRmt induction and ATFS-1-mediated mtDNA replication.
ALKB-1 impairment delays PME, dependent on transcription factor SKN-1
Results from translatomics and proteomics, in conjunction with UPRmt activation upon mitochondrial proteostasis imbalance, indicated that ALKB-1 inactivation led to mitochondrial dysfunction. It is well-established that compromised mitochondrial function can increase reactive oxygen species (ROS) production in cells55. GO analysis of RNC-seq data indicated that alkb-1 inactivation affected ETC function and the oxidative stress response (Supplementary Fig. 8a), hinting at a potential link between ALKB-1-mediated PME and increased ROS production. Utilizing the ROS sensor H2DCF-DA, we observed a significant increase in ROS levels in both alkb-1D247A mutant hermaphrodites and males, compared to wild-type controls (Fig. 4a). Additionally, mitochondrial ROS (mtROS) levels were higher in the alkb-1D247A mutant, as indicated by mtSOX Red staining (Supplementary Fig. 8b, c). Treatment with the antioxidants N-Acetyl-L-cysteine (NAC) or Pyrroloquinoline quinone (PQQ) effectively reversed the PME delay (Fig. 4b, c) and reduced mtROS levels in the alkb-1D247A mutant (Supplementary Fig. 8b, c). Notably, induction of oxidative stress by paraquat treatment delayed the PME process (Supplementary Fig. 8d, e). Collectively, these findings suggest an association between the ALKB-1-mediated PME process and ROS production.
a Quantification of relative intracellular ROS fluorescence intensity in N2 and alkb-1D247A hermaphrodites (Top) and males (Bottom). N2 hermaphrodites treated with 5 mM NAC and 1 mM paraquat (Para) served as negative and positive controls (n = 30 per group), respectively. The experiment was performed three times with similar results. b, d Representative DIC and MTR-stained images of paternal mitochondrial in the 32- and 64-cell stage embryos from cross-fertilization of the indicated strain males (stained by MTR) with hermaphrodites under the indicated treatment, with scale bars of 10 μm. c, e Quantification of the number of MTR-labeled mitochondrial clusters in (b, d) (n = 12 per group). Representative images (f, h) and corresponding quantification (g, i) of ges-1p::mitoGFP fluorescence were shown for hermaphrodites (f, g) or males (h, i) under the indicated condition (n = 30 per group), with scale bars of 50 μm. In (b, d), embryos were derived from 3 to 5 independent biological crosses per condition. The experiment was performed twice with similar results. Data were represented as mean ± SD. p-values were determined by unpaired two-tailed t-test (a-Bottom), and Ordinary one-way ANOVA with Tukey’s multiple comparisons test (a-Top), (c, e, g, i). Source data are provided as a Source Data file.
Elevated ROS levels typically activate the oxidative stress response transcription factor SKN-1/Nrf2, which subsequently upregulates genes involved in antioxidant and phase II detoxification pathways56. A significant upregulation of the SKN-1 target gene gst-4 was observed in the alkb-1D247A mutant, indicating that impairment of ALKB-1 leads to SKN-1 activation (Supplementary Fig. 8f, g). Microscopic assays confirmed that skn-1 knockdown in alkb-1D247A embryos abrogated the PME delay (Fig. 4d, e). Notably, unlike atfs-1, skn-1 knockdown in either the paternal or maternal lineage only moderately alleviated the PME defect induced by ALKB-1 inactivation (Fig. 4d, e), suggesting that both paternal and maternal SKN-1 play roles in the PME delay following ALKB-1 inactivation. These findings support a model in which SKN-1 offers broad antioxidant and mitochondrial buffering capabilities, contributing to PME regulation in both maternal and paternal lineages, whereas ATFS-1, although induced in both gametes, likely acts as a mitochondria-centric, sperm-biased surveillance mechanism, predominantly regulating PME in the paternal lineage.
SKN-1 also facilitates mitochondrial biogenesis through interaction with outer mitochondrial membrane proteins, a process that may require coordination with ATFS-157. We investigated whether mitochondrial biogenesis mediated by SKN-1 contributes to the delay in PME, induced by ALKB-1 inactivation. By using the ges-1p::mitoGFP reporter strain, we detected a significant increase in mitochondrial content, in alkb-1D247A mutants of both sexes compared to wild-type controls, with a corresponding increase observed in male sperm by MTR staining (Supplementary Fig. 4e). This phenotype was reversed by silencing skn-1 or atfs-1 (Figs. 4f–i, 5f). Together, these findings suggest that ALKB-1 inactivation initiates a mitochondrial retrograde signaling pathway that regulates mitochondrial biogenesis, pre- and post-fertilization, mainly through the coordination of the transcription factors SKN-1 and ATFS-1.
a Representative DIC and MTR-stained images of paternal mitochondrial in the 32- and 64-cell stage embryos from cross-fertilization of the indicated strain males (stained by MTR) with hermaphrodites under the indicated treatment, with scale bars of 10 μm. b Quantification of the number of MTR-labeled mitochondrial clusters in (a). n = 12 (N2, 32- and 64-cell), n = 21 (alkb-1D247A, 32- and 64-cell), n = 22 (alkb-1D247A; skn-1 RNAi, 32-cell), n = 18 (alkb-1D247A; skn-1 RNAi, 64-cell), n = 12 (alkb-1D247A; atfs-1(et17); skn-1 RNAi, 32- and 64-cell). c Representative images (Left) and quantification (Right) of the hsp-6p::gfp reporter in wild-type, alkb-1D247A, and alkb-1D247A worms treated with 5 mM NAC or 1 mM PQQ (n = 50 per group), with scale bars of 100 μm. d Representative images (Left) and quantification (Right) of the hsp-6p::gfp reporter in wild-type and alkb-1D247A worms exposed to control RNAi, atfs-1 RNAi, skn-1 RNAi, or atfs-1 and skn-1 double RNAi (n = 30 per group), with scale bars of 100 μm. The experiment was performed three times with similar results. e Representative images (Top) and quantification (Bottom) of ges-1p::mitoGFP fluorescence in wild-type, alkb-1D247A, and alkb-1D247A treated with 5 mM NAC or 1 mM PQQ (n = 60 per group), with scale bars of 50 μm. f Representative images (Left) and quantification (Right) of the ges-1p::mitoGFP fluorescence in wild-type worms treated with control RNAi (n = 50), or alkb-1D247A worms treated with control RNAi (n = 56), atfs-1 RNAi (n = 59), skn-1 RNAi (n = 60), or atfs-1 and skn-1 double RNAi (n = 60), with scale bars of 50 μm. The experiment was performed three times with similar results. Quantification of ges-1p::mitoGFP fluorescence in the skn-1 RNAi group is also presented in Fig. 4i. g Quantification of total mtDNA levels in wild-type and alkb-1D247A males treated with control RNAi, atfs-1 RNAi, skn-1 RNAi, or a combination of atfs-1 and skn-1 double RNAi (n = 9 biologically independent samples). Quantification of total mtDNA levels in the atfs-1 RNAi group is also presented in Fig. 3j. In (b), embryos were derived from 3 to 5 independent biological crosses per condition. The experiment was performed twice with similar results. Data were represented as mean ± SD. p-values were determined by Ordinary one-way ANOVA with Tukey’s multiple comparisons test (b–g). Source data are provided as a Source data file.
SKN-1 acts upstream of ATFS-1 during ALKB-1-mediated PME
To elucidate how ATFS-1 and SKN-1 interact in the PME delay caused by ALKB-1 inactivation, we conducted epistasis analyses. Overexpression of ATFS-1 using the gain-of-function mutant atfs-1 (et17), known for its constitutive UPRmt activation58, partially restored the delayed PME phenotype of the alkb-1D247A; skn-1 RNAi mutant (Fig. 5a, b), suggesting that ATFS-1 functions downstream of SKN-1 to mediate PME delay upon ALKB-1 inactivation. Notably, atfs-1 (et17) harbors a disrupted mitochondrial targeting sequence, resulting in exclusive nuclear localization. Our study unveiled that PME delay, induced by ALKB-1 impairment, depends on the subcellular localization of ATFS-1 in both mitochondria and the nucleus. Due to a lack of mitochondrial localization, the atfs-1 (et17) mutant was incapable of fully restoring the phenotype of alkb-1D247A; skn-1 RNAi, to levels comparable to the alkb-1D247A mutant. Furthermore, treatment with NAC, PQQ, and skn-1 RNAi significantly inhibited the induction of UPRmt and the mitochondrial content increase, induced by the alkb-1D247A mutant (Fig. 5c–f), indicating that ROS modulates the effect of ALKB-1 on the PME process. We also found that skn-1 RNAi, atfs-1 RNAi, or their combination suppressed the increase in mitochondrial biogenesis and mtDNA copy number caused by the alkb-1D247A mutation, with atfs-1 RNAi exerting a stronger inhibitory effect (Fig. 5d, f, g, Supplementary Fig. 4f). This further supports the involvement of both ATFS-1 and SKN-1 in mtDNA amplification and mitochondrial biogenesis, induced by the alkb-1 mutant, with ATFS-1 acting downstream of SKN-1 in this process.
ALKB-1 facilitates PME to maintain sperm fertility and embryo viability
Previous studies have shown that an abnormally high number of sperm mitochondria and PME defects can negatively impact sperm and early embryonic development9,59,60,61. We assessed sperm development using GFP::NKB-2 reporter62 and quantified sperm numbers via DAPI staining. Neither alkb-1 RNAi nor alkb-1D247A mutation affected sperm differentiation at the L4 or adult stages (Supplementary Fig. 9a, b), nor altered sperm counts (Supplementary Fig. 9c, d), indicating that ALKB-1 is dispensable for male sperm development. However, we observed reduced fertility of alkb-1D247A sperm, as evidenced by a significant decrease in the number of progeny from N2 hermaphrodites, mated with alkb-1D247A males (Fig. 6a, b). Additionally, we observed significantly increased embryonic lethality in embryos with delayed PME, caused by the paternal alkb-1 mutation, compared to wild-type controls (Fig. 6c). These findings suggest that ALKB-1 inactivation leads to reduced male fertility and hinders embryo development, possibly due to delayed PME.
The average daily cross progeny production (a) and the total number of cross progeny (b) produced by mating of him-5 (e1467) or alkb-1D247A; him- 5(e1467) males with N2 hermaphrodites. n = 45 (him-5 (e1467) males × N2 hermaphrodites), n = 38 (alkb-1D247A; him- 5(e1467) males × N2 hermaphrodites). The experiment was performed twice with similar results. c Quantification of embryonic lethality in N2 and alkb-1D247A mutants (n = 9). d Schematic model depicting the regulation of PME by the demethylase ALKB-1. Data were represented as mean ± SD. p-values were determined by an unpaired two-tailed t-test (b, c). Source data are provided as a Source data file.
Discussion
Maternal mitochondrial inheritance provides evolutionary benefits by limiting the accumulation of mtDNA mutations in offspring3. Unraveling the regulatory mechanism underlying the rapid elimination of paternal mitochondria could yield important insights for developing innovative strategies to prevent and treat infertility and mitochondrial disorders63. Given the multifaceted roles of methylation patterns in spermatogenesis and embryonic development24,25,26, it is of great interest to study how epigenetic modifications affect the transmission of paternal mitochondria to offspring. Our study reveals that ALKB-1 impairment delays PME in C. elegans, primarily driven by its m¹A tRNA demethylase activity rather than its mitochondrial localization. ALKB-1 inactivation elevates tRNA m¹A levels, enhancing cytoplasmic protein translation, which increases mitochondrial protein influx and results in dysfunction. The resulting rise in ROS, triggered by this disruption, serves as a key driver of retrograde signaling, activating transcription factors SKN-1 and UPRmt. This cascade facilitates mitochondrial biogenesis and mtDNA amplification, delaying paternal mitochondrial clearance during spermatogenesis and early embryogenesis (Fig. 6d).
Our findings indicate that C. elegans ALKB-1 possesses iron-coordinating residues similar to those found in mammalian ALKBH1, and exhibits catalytic activity in demethylating tRNA m1A, corroborating previous insights into ALKBH1 function in mammals33,64. Furthermore, our study highlights a significant correlation between the demethylase activity of ALKB-1 on tRNA m1A and its effect on translation. Inactivation of ALKB-1 disrupts both cytoplasmic and mitochondrial translation, causing mitochondrial protein imbalance and dysfunction. This is in line with previous studies, indicating that ALKBH1 knockout affects mitochondrial translation and oxidative phosphorylation, leading to mitochondrial dysfunction32,34.
Our investigation has shown that ALKB-1 inactivation delays the PME process by increasing ROS generation, aligning with prior studies reporting that high ROS levels impair sperm cells and impede mtDNA removal28,65. Additionally, cells augment mtDNA synthesis in response to mtDNA damage to cope with the increased respiratory demands of ROS detoxification66. In our study, we found that excess ROS activates the oxidative stress response factors SKN-1 and UPRmt, leading to upregulation of antioxidant defense genes and the restoration of mitochondrial function, through mtDNA amplification and mitochondrial biogenesis. This cascade of events, ultimately, impedes the clearance of paternal mitochondria during pre- and post-fertilization stages.
Various endogenous and exogenous stressors can induce UPRmt in C. elegans, including mutant mtDNA accumulation, oxidative phosphorylation dysfunction, increased mtROS, disrupted mitochondrial protein import, and bacterial infections53. However, not all UPRmt inducers resulted in PME delay. For instance, PME was not delayed by various interventions known to strongly activate UPRmt in C. elegans, including pharmacological treatment with doxycycline (Supplementary Fig. 10a, b), the atfs-1 (et17) gain-of-function mutants (Supplementary Fig. 10c, d), and mtDNA deletion mutants mptDf1 or uaDf59(Fig. 1a, b), as demonstrated by our observations. This suggested that the PME defect is specifically linked to ALKB-1 impairment-induced disruption of mitochondrial homeostasis, particularly via m¹A-driven translational imbalance, rather than a general UPRmt induction. Moreover, comparable UPRmt activation was observed in alkb-1ΔMTS/+ and alkb-1D247A worms, without PME delay (Supplementary Fig. 2g–i), supporting the decoupling of UPRmt from PME delay. These findings reinforce that the PME defect induced by ALKB-1 impairment specifically relies on its m¹A-driven translational imbalance, with UPRmt acting as a downstream response rather than a primary driver. Future studies could explore how ALKB-1 inactivation triggers UPRmt and its specific impact on PME, as well as the distinct regulatory roles of UPRmt and ROS in this process.
Autophagy is recognized as the main mechanism for the elimination of paternal mitochondria in C. elegans post-fertilization10,14. Our findings, using the autophagy reporter GFP::LGG-1, indicated that alkb-1 knockdown significantly impaired heat-induced autophagy (Supplementary Fig. 11a–c), suggesting a potential role for autophagy in the PME impairment caused by ALKB-1 inactivation. Additionally, ENDOG/CPS-6, a mitochondrial endonuclease involved in paternal mtDNA degradation post-fertilization, may become inhibited by high levels of ROS9,67. Elevated ROS levels in alkb-1D247A worms raise the possibility that ALKB-1 inactivation might impede PME by affecting CPS-6-mediated mtDNA degradation. Supporting this, the cps-6 mutation or lgg-1 RNAi in alkb-1D247A mutants worsened PME defects to levels comparable to those of single impairments (Supplementary Fig. 12a–d), indicating that ALKB-1 acts upstream of both CPS-6 and autophagy.
In conclusion, our study provides compelling evidence for the important regulatory function of m1A, a conserved epitranscriptomic modification on tRNAs, in the rapid clearance of paternal mitochondria. These findings significantly advance our understanding of maternal mitochondrial inheritance and indicate potential directions for future research.
Methods
C. elegans strains and cultures
The wild-type strain Bristol N2, and the mutant strains CB1467, him-5(e1467)V; SJ4100, zcIs13 [hsp-6p::GFP + lin-15( + )]; SJ4143, zcIs17 [ges-1::GFP (mit)]; SJZ328, foxSi75 [eft-3p::tomm-20::mKate2::HA::tbb-2 3’ UTR (oxti185)I]; CL2166, dvIs19 [(pAF15) gst-4p::GFP::NLS]III; OP675, wgIs675 [atfs-1::TY1::EGFP::3×FLAG + unc-119 ( + )]; TM4919, atfs-1 (tm4919)V; QC117, atfs-1 (et17)V; TM4803, dve-1 (tm4803)X; MT13232, lin-65 (n3441)I; VC2654, ubl-5 (ok3389)I; and DA2123, adIs2122 [lgg-1p::GFP::lgg-1 + rol-6B (su1006)], were obtained from the Caenorhabditis Genetics Center (Minneapolis, MN, USA) and the National BioResource Project (Tokyo, Japan). The strains SJ4197, zcIs39 II [dve-1p::dve-1::GFP] and SJ4151, zcIs19 [ubl-5p::ubl-5::GFP], were the gifts from Dr. Ye Tian of the Genetics and Developmental Biology, Chinese Academy of Sciences. The strain LMW467, ibp96 [gfp::nkb-2] was the gifts from Dr Long Miao of the Key Laboratory of RNA Biology, Chinese Academy of Sciences. The strains JNU102, alkb-1 (ajz102)/+II; JNU103, alkb-1 (ajz103)II; JNU105, ajzIs105 [pie-1p::ALKB-1::3xFlag::tbb-2 3’UTR; unc119 (ed3)]; JNU111, ajzIs111 [trp-3p:: ALKB-1::3xFlag::trp-3 3’UTR; unc-119 (ed3)]; JNU209, alkb-1 (ajz209)/+II; JNU213, ajzIs213 [pMD18-T::damt-1p::DAMT-1::3xHA::3×Flag::tbb-2 3’UTR, myo-2p::mCherry] and JNU214, ajzIs214 [tbb-2p::ALKB-1::3×Flag::tbb-2 3’UTR] were generated by SunyBiotech (Fuzhou, Fujian, China). The double mutants were constructed using standard genetic techniques. All worms were cultured at 20 °C on standard nematode growth medium (NGM) plates seeded with Escherichia coli (E. coli) strain OP50 (unless otherwise stated).
Plasmid construction
The coding sequences of ALKB-1, ALKB-1D247A, ALKB-1(Δ2-35), SDHB-1 MTS, and DAMT-1 was amplified by PCR from wild-type N2 or alkb-1D247A mutant cDNAs. The sequences of tbb-2/damt-1/myo-3 promoter and tbb-2/unc-54 3’UTR were amplified by PCR from N2 genome. The sequence of GFP (S65C) was cloned from plasmid pPD95_75 (Addgene: 1494). The sequence of mCherry was cloned from plasmid Pmyo-2::mCherry::unc-and C. e ALKB-1(Δ2-35)::mCherry fusion proteins in HeLa cells. The C. e ALKB-1::mCherry and C. e ALKB-1(Δ2-35)::mCherry constructs were cloned using the ClonExpress II One Step Cloning Kit (Vazyme, Nanjing, China) into the eukaryotic expression plasmid pcDNA3.1(+). To create the plasmids expressing ALKB-1::3×Flag, ALKB-1D247A::3×Flag, ALKB-1(Δ2-35)::3×Flag, DAMT-1::3xHA::3×Flag, and SDHB-1 MTS::gfp (S65C) (no stop codon) AAAAn (NS-mtGFP) fusion protein in C. elegans, the tbb-2p::ALKB-1::3×Flag::tbb-2 3’UTR, tbb-2p::ALKB-1D247A::3×Flag::tbb-2 3’UTR and tbb-2p::ALKB-1(Δ2-35)::3×Flag::tbb-2 3’UTR cassette were constructed and cloned into the pCFJ356-MCS vector (Addgene: 34871), the damt-1p::DAMT-1::3xHA::3×Flag and myo-3p::SDHB-1 MTS::gfp (S65C) (no stop codon)::unc-54 3’UTR AAAAn cassette was constructed and cloned into the pMD 18-T vector (Takara, 6011), using the ClonExpress II One Step Cloning Kit. All plasmids were verified by Sanger sequencing before use.
Microinjection and transgenic strains
Transgenic C. elegans strains were generated by standard microinjection into the syncytial gonads of young adult hermaphrodites. The DNA injection mixture contained an expression plasmid at 50 ng/μL and a fluorescent co-injection marker at 5 ng/μL. Specifically, we used plasmids to express a catalytically inactive ALKB-1 (tbb-2p::ALKB-1D247A::3×Flag::tbb-2 3’UTR), an ALKB-1 variant lacking its N-terminal mitochondrial targeting sequence (tbb-2p::ALKB-1(Δ2-35)::3×Flag::tbb-2 3’UTR), or an NS-mtGFP reporter (myo-3p::SDHB-1 MTS::gfp (S65C) (no stop codon)::unc-54 3’UTR AAAAn). These were co-injected with either Pmyo-2::GFP::H2B or Pmyo-3::mCherry as selection markers into either wild-type N2 or alkb-1D247A mutant backgrounds. F1 progeny exhibiting marker fluorescence were subsequently selected, which maintained the respective transgenes as heritable, extrachromosomal arrays.
Cell culture and transfection
HeLa cells were maintained in DMEM medium (Gibco, C11995500BT) with 10% fetal bovine serum (Gibco, A5256701) and 1% penicillin and streptomycin (Gibco, SH30030.02). Cells were incubated at 37 °C in 5% CO2. For the expression of C. e ALKB-1::mCherry or C. e ALKB-1(Δ2-35)::mCherry fusion proteins, HeLa cells were transfected with plasmids using Lipofectamine LTX reagent (Invitrogen, 11668-019) according to the manufacturer’s protocol.
Immunofluorescence analysis in HeLa cells
After transfection of 300 ng of ALKB-1-mCherry or ALKB-1(Δ2-35)-mCherry plasmids into HeLa cells cultured on 10 cm dishes coated with poly-lysine, the cells were subjected to two cold PBS washes 48 h post-transfection. Subsequently, the cells were fixed with 4% paraformaldehyde (PFA) for 20 min at room temperature, followed by three additional PBS washes. The cells were then treated with 0.1% Triton X-100 for 10 min. Following three more PBS washes, the cells were blocked with 5% bovine serum albumin (BSA) for 1.5 h at room temperature. Antibodies against TOM20 (1:200, Proteintech, 11802-1-AP) were then added at a 1:400 dilution and allowed to incubate for 4 h at room temperature. Following this, the cells underwent four PBS washes before being incubated with 488-AffiniPure Goat Anti-Rabbit (diluted 1:1000 in 5% BSA) for 1 h. After six additional PBS washes, the slides were mounted with DAPI-containing mounting medium and imaged using a Leica confocal microscope to capture single-slice images. The co-localization of mCherry with GFP or DAPI was quantified using ImageJ software.
RNA interference (RNAi)
All RNAi strains, with the exception of the alkb-1 RNAi strain, were obtained from the Ahringer library. To generate the RNAi strain targeting alkb-1, the alkb-1 cDNA sequence was amplified from C. elegans N2 cDNA and inserted into a Hind III-digested L4440 vector. Subsequently, the L4440-alkb-1 (cDNA) plasmids were transformed into HT115 (DE3) chemically competent cells through a CaCl2 transformation protocol. The bacteria expressing dsRNA were cultured overnight in LB medium supplemented with 100 mg/mL of ampicillin, after which the bacteria were spread onto NGM plates containing 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG). L1 larvae were grown on plates containing gene-specific RNAi bacteria until they reached the L4 or young adult stage for subsequent experiments.
RNAi Screen
To identify methylation-related genes involved in PME, we performed an RNAi screen to pinpoint genes predicted to encode dealkylating enzymes in C. elegans. We used the nested PCR method to detect the sperm-derived mptDf1 mtDNA in the 1.5-fold stage cross-progeny embryos. Briefly, mptDf1/+; him-5 (e1467) and N2 worms were grown on IPTG-added NGM plates containing HT115 (DE3) bacteria harboring specific-gene dsRNA expression constructs for two generations. Afterward, F1 adults mptDf1/+; him-5 (e1467) males and N2 hermaphrodites were transferred to fresh RNAi plates for mating experiments. Following mating, 1.5-fold stage cross-progeny embryos dissected from the gravid hermaphrodites and then subjected to the nested PCR to ascertain the presence of mptDf1 deletion.
Detection of mptDf1 mtDNA
To determine the heteroplasmy level of the mptDf1 mtDNA in males and to detect sperm-derived mptDf1 mtDNA in cross-progeny samples, nested PCR assays were performed. For heteroplasmy analyses in males, approximately 20 L4-stage worms were collected and lysed in 20 μL of lysis buffer at 65 °C for 60 min. For the detection in cross-progeny, total lysates were prepared from different developmental stages of embryos or larvae. Wild-type mtDNA and mptDf1 mtDNA were distinguished using the following primers: mptDf1 Outer F, 5′-GGTATAATTGGGGCCATCCGT-3′ and mptDf1 Outer R, 5′-TCAAAAGTGGAGCGGTGCTAC-3′; mptDf1 Inner F, 5′-GAACTTAACCGGGCGCCATT-3′ and mptDf1 Inner R, 5′-ACGCTACAGCAGCATAAACAA-3′. The heteroplasmy level in males was subsequently quantified by assessing the relative band intensity (grayscale value) of the wild-type and mptDf1-specific PCR products.
Microscopy and quantification of paternal mitochondria
Young adult males were stained on OP50 NGM plates containing 50 μM MitoTracker® Red CMXRos (Invitrogen, M7512) for 12 h and then transferred to a fresh OP50 NGM plates for 30 min to eliminate residual MTR. The MTR-stained males were subsequently mated with unstained young adult hermaphrodites for 8–10 h. Following mating, the hermaphrodites were dissected to release the cross-fertilized embryos onto an agarose pad. A small volume of M9 buffer was added to the agar pad, and a coverslip was placed on top. The embryos were systematically imaged using a Zeiss Axio Imager Z2 with Apotome.2 microscope at 1000× magnification, capturing images from top to bottom in Z-stacks. Deconvolution was performed on the Z-stacks, and the images were projected onto a single plane to provide a comprehensive visualization of all MTR-labeled mitochondria in the embryo. The number of MTR-stained mitochondrial clusters in cross-fertilized embryos was counted using ImageJ software. At least 10 embryos were scored in each group. Statistical significance was determined using the unpaired student’s t-test or one-way ANOVA to calculate the p-value.
RNA extraction and quantitative RT-PCR
Young adult worms were collected using M9 buffer and subjected to multiple washes. The worm samples were then resuspended in AG RNAex PRO reagent (Accurate Biology, Changsha, China) for RNA extraction. Total RNA was isolated using chloroform extraction followed by isopropanol precipitation. Subsequently, 500 ng of RNA was utilized for cDNA synthesis utilizing a high-capacity cDNA transcription kit (RK20400, ABclonal, Wuhan, China). Quantitative RT-PCR was conducted utilizing SYBR Green Select Master Mix (RK21203, ABclonal, Wuhan, China) on a LightCycler480 real-time system (Roche, USA), with each assay being replicated three times. The mRNA expression levels of the genes were evaluated after normalization to the reference gene cdc-42, and statistical significance was assessed using the unpaired student’s t-test. Details of the primer sequences used in this study were provided in Supplementary Table 1.
mRNA isolation
The mRNA was isolated from the total RNA using the Dynabeads mRNA purification kit (Cat No. 61006, Invitrogen) according to the manufacturer’s instructions. In brief, 75 μg of total RNA was heated at 65 °C for 2 min to disrupt the secondary structure. The superparamagnetic DynabeadsTM coupled with oligonucleotide-(dT)25 in the binding buffer, were equilibrated and mixed with the total RNA. The beads were then washed to remove RNA contaminants, and the mRNA was eluted using 5 µL of 10 mM Tris-HCl. The isolated mRNA samples were subsequently utilized for dot blot analyses.
tRNA isolation
Young adult hermaphrodite worms were collected using M9 buffer and underwent several washes. The small RNA fraction (<200 nucleotides, primarily consisting of mature tRNA) was extracted from the worm samples using the mirVanaTM miRNA Isolation Kit (AM1561, Life Technologies) according to the manufacturer’s instructions. The isolated tRNA samples were subsequently used for dot blot analyses.
Protein purification
The fragments sequences of ALKB-1 and ALKB-1D247A were cloned into the pET-30a (+) vector and then introduced into BL21 (DE3) chemically competent cells using the CaCl2 transformation protocols. Subsequently, these proteins with N-terminal His-tag were expressed in the BL21 (DE3) cells following overnight induction with 0.2 mM IPTG. The cells were collected by centrifugation and resuspended in lysis buffer containing 20 mM Tris-HCl (pH 8.0) and 100 mM NaCl. Afterward, the cells were homogenized using an Emulsiflex C3 (Avestin) homogenizer, and the centrifugation-cleared supernatant was loaded onto a HisTrap nickel column (GE Healthcare). The protein bound to the nickel column was eluted using a linear imidazole gradient ranging from 20 to 500 mM. The eluted protein underwent His-tag removal by PreScission protease and was subsequently subjected to ion-exchange chromatography using HiTrap QHP and HiTrap Heparin HP columns (GE Healthcare). Finally, the protein was purified to homogeneity via size exclusion chromatography on a Superdex 200 increase 10/300 column (GE Healthcare) and concentrated to 15 mg/mL in a buffer consisting of 20 mM Tris-HCl (pH 8.0) and 100 mM NaCl for subsequent experiments.
Demethylation assays in vitro
The modified 6 mA oligo used in this study was obtained from The Beijing Genomics Institute BGI (Shenzhen, Guangdong, China). The sequence of the oligo was as follow: 6 mA, 5′-GGGAATTTCCCGGCGATTTGA (N6-Me-dA) TCAAATCGCCGGGAAATTCCC-3′, as previously described68. The stem-loop structured tRNA probe modified with m1A was obtained from Sangon Biotech (Shanghai, China), and the sequence of the probe was as follow: 5′-CACGGUUCG (m1A) UUCAAAG-3′, as previously described33. To assess the demethylation activity of ALKB-1 in vitro, each substrate was annealed under distinct conditions. The 6 mA oligos were denatured at a concentration of 0.1 nmol/μL in a denaturation buffer (10 mM Tris-HCl, 100 mM NaCl, 10 μM EDTA, pH 8.0) by heating at 95 °C for 2 min followed by a gradual cooling to 25 °C over 1 h. For the m1A-tRNA probe, annealing was carried out at 0.1 μM in RNA annealing buffer (100 mM Tris-HCl, 1 M NaCl, 50 mM MgCl₂, 0.1 mM EDTA, pH 7.5) using DEPC-treated water, with a protocol involving heating to 95 °C for 2 min, rapid cooling to 45 °C, a 30 min hold, and gradual cooling to 4 °C at 0.1 °C/s. The demethylation reaction was then conducted in a 20 μL volume containing 0.1 nM of the respective annealed substrate and 1 μg of either ALKB-1 or the mutant ALKB-1D247A protein in reaction buffer (50 μM HEPES pH 7.0, 50 μM KCl, 2 mM MgCl₂, 2 mM ascorbic acid, 1 mM α-ketoglutarate, 200 μM (NH₄)₂Fe(SO₄)₂). Notably, reactions involving the m1A-tRNA substrate were supplemented with 0.2 U/μL RNase Inhibitor. The reactions were then incubated at 25 °C for 6 h and terminated by adding EDTA to a final concentration of 5 mM, followed by incubation at 95 °C for 10 min. The reaction products were subsequently utilized for dot blot analyses.
Purification of worm genomic DNA
Young adult worms were harvested using M9 and then lysed in a worm gDNA lysis buffer containing 200 mM NaCl, 100 mM Tris-HCl (pH 8.5), 50 mM EDTA (pH 8.0), 0.5% SDS, and proteinase K (0.1 mg/mL). The lysate was incubated at 65 °C for 1 h with intermittent vortexing. Proteinase K was then inactivated by heating at 95 °C for 20 min. Afterward, RNase A was added and incubated at 37 °C for 1 h. The sample was then mixed with an equal volume of phenol: chloroform: isopentanol (25:24:1) and centrifuged at 13,000 rpm for 5 min at room temperature. The aqueous phase was transferred to a new tube and mixed with sodium acetate and 100% ethanol before being incubated at −20 °C overnight. After centrifugation at 16,000 × g for 10 min at 4 °C, the DNA pellet was washed with 70% EtOH, air-dried, and resuspended in TE buffer. Following this, the purified gDNA underwent treatment with RNase A/T1 (Thermo Fisher Scientific) and RNase H (New England Biolabs) at 37 °C for 1 h. The process concluded with another purification step, starting with the phenol: chloroform: isoamyl alcohol extraction. The isolated gDNA samples were then prepared for dot blot analyses.
Dot blot
Oligos, gDNA, mRNA, and tRNA samples were denatured at 95 °C for 10 min and snap-cooled on ice. A total of 2 μL of each sample, containing Oligos (all reaction products), 300 ng of DNA, 120 ng of mRNA, and 100 ng of tRNA, was spotted onto a membrane (Amersham, Hybond-N + , GE), air-dried, and UV cross-linked for 1.5 h. The membranes were then incubated in a blocking buffer (5% milk, PBS-T) for 2 h at room temperature, followed by an overnight incubation with antibodies against 6 mA (1:1000, Synaptic Systems, 202-003) or m1A (1:1000, MBJ Life Science, D345-3 or 1:1000, abcam, ab208196) at 4 °C. After three subsequent washes, the membrane was incubated with a secondary antibody at room temperature for 1 h, and the signals were visualized utilizing a Bio-Rad ChemiDoc MP imaging system.
LC-MS based tRNA modification Detection
The LC-MS based tRNA modification detection was conducted by Aksomics (Shanghai, China). Total RNA from each sample was assessed using a NanoDrop ND-1000 instrument, and approximately 20 μg of total RNA was utilized for tRNA isolation using the Urea-PAGE method. The purified tRNA was enzymatically digested into single dephosphorylated nucleosides. Subsequently, LC-MS analysis was performed on an Agilent 6460 QQQ mass spectrometer coupled with an Agilent 1260 HPLC system using Multi Reaction Monitoring (MRM) mode for detection. Data acquisition for LC-MS was performed using Agilent Qualitative Analysis software. MRM peaks corresponding to each modified nucleoside were extracted and normalized to the quantity of injected tRNA.
Isolation of mitochondria
To isolate mitochondria from C. elegans, approximately 40,000–60,000 L4 worms were collected using M9 buffer with 0.01% Triton X-100, washed three times, and snap-frozen in liquid nitrogen before storage at −80 °C. The collected worm pellets were suspended in a mitochondria isolation buffer (10 mM Hepes-KOH pH 7.5, 210 mM mannitol, 70 mM sucrose, 1 mM EDTA). The worm cuticle was disrupted by grinding using an acid-washed mortar and pestle, followed by further homogenization with a Dounce homogenizer. The homogenate underwent low-speed centrifugation at 200 × g for 5 min to remove cellular debris. Subsequently, the resulting supernatant was subjected to a higher-speed centrifugation at 12,000 × g for 10 min to isolate the cytosolic fraction in the final supernatant, while the pellet contained the mitochondria. The obtained supernatant and mitochondrial samples were then used for subsequent Western blot analyses.
Western blot analysis
Young adult worms were collected using M9 buffer, washed three times, and then immediately frozen in liquid nitrogen for storage at −80 °C. The worm pellets were lysed using RIPA buffer and homogenized twice using a TissueLyser at 75 Hz for 6 min at 4 °C. This was followed by centrifugation at 10,000 × g at 4 °C to obtain supernatants. Protein concentrations in these supernatants were quantified using a BCA Protein Assay Kit. Samples were boiled at 95 °C for 5 min to denature the proteins prior to electrophoresis on either 6% or 10% SDS-polyacrylamide gels. Following electrophoresis, the separated proteins were transferred onto nitrocellulose membranes. The membranes were then blocked with 5% milk and incubated with primary antibodies against the FLAG (1:3000, Sigma-Aldrich, F1804), HA (1:3,000, Cell Signaling Technology, 3724S), GFP (1:5000, ROCHE, 11814460001), β-actin (1:5,000, Sigma-Aldrich, A1978), or α-tubulin (1:5,000, Cell Signaling Technology, 2125S). The primary antibody was visualized using a horseradish peroxidase-conjugated anti-rabbit or anti-mouse secondary antibody (anti-rabbit, 1:5000, Jackson ImmunoResearch, 111-035-144; anti-mouse, 1:5000, Jackson ImmunoResearch, 115-035-068) and ECL Western Blotting Substrate.
Biotin-labeled Northern Blot analysis
Total RNA samples were analyzed by Northern blot using the Biotin Northern Blot Kit for Small RNA (Beyotime). Briefly, 10 μg of total RNA was denatured, separated on a 15% urea-polyacrylamide gel, and transferred to a nylon membrane. After UV crosslinking, the membrane was hybridized overnight at 42 °C with biotin-labeled probes. Following hybridization, the membrane was incubated with HRP-conjugated streptavidin and detected using ECL chemiluminescence. 5S rRNA was used as a reference control for tRNA abundance, and all probe sequences are provided in Supplementary Table 2.
Isolation and lysis of C. elegans sperm
To isolate the sperm, at least 100 virgin males were transferred into a lysis buffer droplet. Incisions were made at one-third of the body length from the tail, between the midpoint and tail, using syringe needles to release the sperm. The broken worms were then carefully removed from the lysis buffer using needles. The buffer containing the sperm was incubated at 65 °C for 1 h. The lysed samples were then utilized for subsequent qPCR to measure mtDNA levels.
Measurement of ROS levels
Endogenous ROS levels were evaluated by measuring the fluorescence intensity of 2′7′-Dichlorofluorescein diacetate (H2DCF-DA) according to established protocols69. Briefly, young adult worms were exposed to 10 mM H2DCF-DA on OP50 NGM plates for 1 h, then transferred to a fresh OP50 NGM plate for 30 min to eliminate residual H2DCF-DA. Subsequently, the H2DCF-DA labeled animals were mounted on agarose pads and imaged at 20× magnification utilizing a Nikon Ti2-U fluorescence microscope. Each assessment involved a minimum of 30 worms, and each assay was replicated at least twice. The fluorescence intensity in the images was analyzed and quantified using ImageJ software.
Measurement of mtROS
To evaluate mitochondrial ROS levels, MitoSox staining was carried out on worms with minor modifications to an established protocol70. Briefly, young adult worms were collected and subjected to triple washing with M9 buffer to remove bacteria before being stained with 5 mM MitoSox in M9 for 30 min. Afterward, the stained worms underwent additional washes in M9 and were mounted on agarose slides for imaging. The head regions of the worms were imaged using a Nikon Ti2-U fluorescence microscope at 20× magnification, and fluorescence intensity measurements were carried out using ImageJ software.
Fluorescent microscopic imaging
To quantify the expression of hsp-6p::gfp, ges-1p::mitoGFP, gst-4p::gfp, and ubl-5p::ubl-5::gfp, at least 30 animals per group were anesthetized with 10 µM levamisole and placed on 2% agar pads for imaging with a Nikon Ti2-U fluorescence microscope. The fluorescence intensity was measured using ImageJ software. For assessing DVE-1::GFP intestine nuclear location, at least 30 dve-1p::dve-1::gfp animals were anesthetized in M9 buffer containing sodium azide (50 mM), mounted onto 2% agarose pads, and intestine cells with DVE-1::GFP signal were counted directly at 200× magnification using a Zeiss Axio Imager Z2 with Apotome.2. To quantify mitochondria in C. elegans sperm, L4 stage male worms were stained overnight with 10 μM MTR and imaged at 1000× magnification using a Zeiss Axio Imager Z2 with Apotome.2 after crawling on fresh OP50 plates for 1 h; fluorescence intensity was quantified using ImageJ software, including at least 10 worms and 3-5 sperm per worm. For imaging and puncta quantification in the seam cells of strain DA2123, animals were immobilized with 10 µM levamisole, and GFP::LGG-1 puncta were directly counted at 630× magnification using a Zeiss Axio Imager Z2 with Apotome.2. For imaging of GFP::NKB-2, males at both L4 and adult stages were anesthetized in M9 buffer containing sodium azide and mounted on agar pads. The entire male germline was then imaged at 630× magnification using a Leica confocal microscope to capture the fluorescence signal. All experiments were conducted at least two times, and statistical analyses were performed using an unpaired Student’s t-test or one-way ANOVA.
C. elegans mtDNA quantification
Approximately 20 L4 stage worms were collected and suspended in 20 μL of lysis buffer before being incubated at 65 °C for 60 min. The levels of mtDNA were quantified using qPCR-based methods as previously described45. qPCR was conducted using SYBR Green Select Master Mix (RK21203, ABclonal, Wuhan, China) on a LightCycler480 real-time system (Roche, USA), with each assay being replicated three times. The relative levels of mtDNA were determined using primer pairs targeting mtDNA and gDNA, as previously described45. The primer sequences are as follows: wild-type mtDNA F, 5′-GCTTTTTCTTTATATGTTTTGTG-3′; wild-type mtDNA R, 5′-TCACCTTCAGAAAAATCAAATGG-3′; ges-1 (nuclear genome DNA) F, 5′-AGGCTAAGCCGGGGTAAGTT-3′; ges-1 (nuclear genome DNA) R, 5′-GCCAAAAGCTTAAACTGCGG-3′.
Embryonic lethality assays
The experiments were performed at 25 °C following a previously established protocol9. L4 larvae were allowed to mature for 12 h before being transferred to fresh plates for a 6-h egg-laying period. Subsequently, the worms were removed from the plates, and the total number of embryos on the plates was counted. After an additional 24-h incubation period, any embryos that had not hatched were recorded as non-viable. The embryonic lethality rate was calculated by dividing the number of non-viable embryos by the total number of embryos counted.
C. elegans sperm numbers analysis
L4 virgin males were isolated onto fresh NGM plates and incubated at 20 °C for 18–24 h before staining. Subsequently, the worms were washed with PBST and fixed in 40% isopropanol for 3 min. The samples were then incubated in a DAPI staining solution (10 μg/mL in 40% isopropanol) for 5 min in the dark, followed by a 30 min destaining procedure in PBST. The entire spermatid region was systematically imaged using a Zeiss Axio Imager Z2 microscope at 630× magnification, capturing images from top to bottom in Z-stacks. Total sperm counts were quantified from maximum intensity projections using the ImageJ cell counter plugin.
Male fertility assays
To assess male fertility, a single L4 stage wild-type or alkb-1D247A virgin male was paired with one N2 hermaphrodite at the 24 h post-young adult stage on each 35 mm NGM plate. These pairs were subsequently transferred to fresh plates every 24 h until the hermaphrodite stopped producing eggs. Progeny count was then conducted. Hermaphrodites that failed to mate with males, identified by the absence of male offspring, were excluded from the dataset.
RNA-seq
The RNA-seq libraries were prepared and sequenced at Chi-Biotech (Shenzhen, China). Synchronized young adult worms were harvested using cold PBS buffer for total RNA extraction with AG RNAex PRO reagent (Accurate Biology, Changsha, China). Subsequently, RNA integrity was assessed using the RNA Nano 6000 Assay Kit on the Bioanalyzer 2100 system from Agilent Technologies, CA, USA. For library construction and sequencing, 1 μg of RNA per sample was utilized, and sequencing was conducted using the cBot Cluster Generation System with the TruSeq PE Cluster Kit v3-cBot-HS (Illumina) following the manufacturer’s protocols. Differential expression analysis was carried out using the DESeq2 R package (version 1.20.0), with genes showing a fold change greater than twofold and a false discovery rate <0.01 considered as differentially expressed genes (DEGs). The DEGs were functionally annotated using various databases, including Gene Ontology (GO), Kyoto Encyclopedia of Gene and Genomes (KEGG).
RNC-seq
The RNC-seq libraries were prepared and sequenced at Chi-Biotech (Shenzhen, China). Young adult worms were collected using M9 buffer, washed three times, and rapidly froze in liquid nitrogen. The frozen worms were crushed into a power using a pre-chilled mortar and then mixed with lysis buffer (1% Triton X-100, 20 mM HEPES-KOH (pH 7.4), 15 mM MgCl2, 200 mM KCl, 100 μg/mL cycloheximide, and 2 mM dithiothreitol) for extraction. The resulting lysate was subjected to ultracentrifugation in a 30% sucrose solution. The resulting precipitate was collected, re-suspended, and subjected to chloroform extraction and isopropanol precipitation to isolate total RNA. The integrity of the total RNA was evaluated using the Agilent 4200 Bioanalyzer. mRNA was enriched using oligo (dT) magnetic beads to isolate poly (A) tailed mRNA. The isolated mRNA was fragmented using divalent cations, and these fragments were utilized as templates for synthesizing the first cDNA strand, with random oligonucleotides as primers within the 1st Strand Enzyme reverse transcription system. The second cDNA strand was then synthesized using the 2nd Strand Enzyme and dNTPs. The purified double-stranded cDNA underwent end repair, A-tailing, and adapter ligation. Subsequently, cDNA fragments ranging in size from 200 to 300 bp were selected, PCR amplified, and purified to form the final library. The library concentration was determined using Qubit or qPCR, while the library quality was assessed using the Agilent 4200 Bioanalyzer to verify the insert size. The library was sequenced on the Illumina NovaSeq 6000 platform using PE150 sequencing. To assess changes in translation efficiency, the Translation Ratio (TR) for each gene was calculated as the ratio of its translational abundance (measured by TPM from RNC-seq) to its transcriptional abundance (measured by TPM from RNA-seq), using the formula: TR = (TPM in RNC-seq)/(TPM in RNA-seq). This normalization corrects for changes in mRNA levels that are independent of translation, thereby isolating genuine alterations in translational regulation. Genes exhibiting significant changes in TR between conditions were identified as differential translation efficiency genes. The differential translation efficiency analysis was carried out using DESeq2 with a filter threshold of p-value < 0.05 and |log2 FC | > 1. The differential translation transcripts were functionally annotated using Gene Ontology (GO) and Kyoto Encyclopedia of Gene and Genomes (KEGG).
Proteomics analysis
The proteomic experiment was performed by Jingjie PTM BioLab (Hangzhou, China) utilizing 4D label-free proteomics analysis. Young adult worms were collected using M9 buffer, washed, and then snap-frozen in liquid nitrogen before storage at −80 °C. Proteins were extracted using lysis buffer containing 8 M urea and 1% protease inhibitor cocktail, followed by trypsin digestion. The resulting tryptic peptides, dissolved in solvent A (0.1% FA in water), were loaded onto an Evotip according to the manufacturer’s protocol. Peptides were separated using the 60-SPD method on the EvoSep One liquid chromatography system, with the mobile phase composed of solvent A and solvent B (0.1% FA in ACN). A gradient increase in solvent B was performed, starting from 4 to 22% over 70 min, then 22 to 30% over 14 min, followed by a quick increase to 80% over 3 min, and a subsequent 3 min equilibration period between gradient changes, all at a flow rate of 450 nL/min. Subsequently, the peptides underwent capillary source analysis followed by analysis using timsTOF Pro mass spectrometry. The timsTOF Pro was operated in dia-PASEF mode with a full MS scan range of 300–1500 m/z and 20 PASEF scans acquired per cycle. The MS/MS isolation window was set at 7 m/z within the range of 400–850 m/z. The DIA data were analyzed using the DIA-NN search engine (v.1.8), and tandem mass spectra were searched against a combined target and decoy database of Blast_Caenorhabditis_elegans_6239_PR_20230529.fasta (26706 entries). Trypsin/P was designated as the cleavage enzyme, allowing for up to one missed cleavage. N-terminal Met excision and carbamidomethyl modification of Cys were indicated as fixed modifications. The false discovery rate (FDR) was adjusted to be less than 1%. Subcellular localization predictions and protein annotation were conducted using WolF PSORT software.
tRNA seq
tRNA m1A seq was performed on the Illumina NextSeq 500 system by Aksomics (Shanghai, China). Total RNA was extracted from C. elegans samples using a standard RNA extraction kit. RNA quality and concentration were assessed using an Agilent 2100 Bioanalyzer. Total RNA (≥2 μg) was separated on a 7.5% urea-PAGE gel, and tRNA fragments (60–100 nt) were excised and eluted. To analyze m1A modifications, a modified tRNA-seq approach was employed, incorporating a demethylation (Demethylation (−)) control group. Each sample was divided into two groups: Demethylation (+) and Demethylation (−). For the Demethylation (+) group, the RNA Pretreatment kit (Arraystar) was used to remove m1A modifications, facilitating efficient cDNA reverse transcription. In the Demethylation (−) group, the demethylation step was omitted, allowing for the identification of m1A-associated mismatches or truncations during sequencing and analysis.
tRNA fragments were subjected to partial hydrolysis using a Na2CO3/NaHCO3 buffer to generate 5′-monophosphate ends. Dephosphorylation with calf intestinal alkaline phosphatase (CIP) was followed by rephosphorylation using T4 polynucleotide kinase. Purified tRNA fragments were converted into barcoded small RNA libraries using the NEBNext Small RNA Library Prep Set for Illumina (New England Biolabs). Library quality and concentration were assessed using an Agilent 2100 Bioanalyzer. Libraries were pooled in equal molar ratios and sequenced on the Illumina NextSeq 500 system.
For tRNA m¹A sequencing data analysis, raw sequencing reads underwent quality assessment using FastQC (v0.11.7) to evaluate base call accuracy and overall read quality, followed by adapter trimming with Cutadapt (v1.14) to remove 5′ and 3′ adapter sequences and filtering to discard reads shorter than 18 nucleotides or longer than 45 nucleotides, retaining only high-quality trimmed reads for downstream analysis. Trimmed reads were aligned to a reference set of mature cytoplasmic and mitochondrial tRNA sequences obtained from GtRNAdb and mitotRNAdb, respectively, using BWA (v0.7.12-r1039) with a maximum of one mismatch allowed per read to ensure high specificity, and the resulting BAM files were used for quantification of m¹A modification sites. To identify and quantify m¹A modification sites, aligned reads were analyzed at each nucleotide position of the reference tRNA sequences: for each position, the total number of mapped reads (coverage) was recorded separately for untreated and treated samples; the number of reads containing a non-reference base (e.g., A → T, A → G, A → C substitutions) or truncations was counted, as these are indicative of potential m¹A modifications due to reverse transcription misincorporation; the modification ratio at each position was calculated as ratio_untreated = Count_untreated/Coverage_untreated and ratio_treated = Count_treated/Coverage_treated; and a differential score was computed for each site as score = ratio_untreated - ratio_treated, reflecting the relative abundance of m¹A modifications assuming chemical treatment reduces misincorporation at modified sites (e.g., a score of 0.1 corresponds to approximately 10% of molecules being modified). To exclude background noise from low-abundance modifications, a change threshold (|ΔScore | >0.05, equivalent to approximately 5% modification ratio change) was applied to filter tRNA species with significantly differential modification levels. Differential analysis was performed using a t-test, with fold change and P-values used to identify differentially modified m¹A sites.
To profile tRNA expression abundance and identify differentially expressed tRNAs, the purified tRNA underwent a critical pre-treatment step involving m¹A and m³C demethylation to reduce sequencing biases, and was then partially hydrolyzed using the Hydro-tRNA seq protocol. These processed fragments were re-phosphorylated and converted into sequencing libraries with the NEBNext® Multiplex Small RNA Library Prep Set for Illumina®. The resulting libraries underwent stringent size selection (~140–155 bp, corresponding to ~19–35 nt tRNA fragments) and were rigorously qualified and quantified on an Agilent 2100 BioAnalyzer prior to being pooled in equimolar amounts for sequencing on an Illumina platform. For bioinformatic analysis, sequencing quality was assessed with FastQC, adapter sequences were trimmed using cutadapt, and the cleaned reads were aligned to both cytoplasmic (GtRNAdb) and mitochondrial (mitotRNAdb) tRNA reference sequences with BWA software, allowing a maximum of two mismatches. Finally, tRNA expression profiles were generated from mapped reads, and differential expression analysis was conducted based on read counts using the R package edgeR.
Data availability
The data supporting the findings of this study are available from the corresponding authors upon request. The RNA-seq, RNC-seq, tRNA m1A seq, and tRNA seq analysis data generated in this study have been deposited in the Sequence Read Archive (SRA) database under accession code PRJNA1104793, PRJNA1204926, PRJNA1358275 and PRJNA1358279. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository with the dataset identifier PXD070509. All data necessary for confirming the conclusions of the article are present within the article and/or the Supplementary information. Source data for the figures and Supplementary Figs. are provided as a Source Data file. Source data are provided with this paper.
References
Mottis, A., Herzig, S. & Auwerx, J. Mitocellular communication: shaping health and disease. Science 366, 827–832 (2019).
Isaac, R. S., McShane, E. & Churchman, L. S. The multiple levels of mitonuclear coregulation. Annu. Rev. Genet. 52, 511–533 (2018).
Lee, W. et al. Molecular basis for maternal inheritance of human mitochondrial DNA. Nat. Genet. 55, 1632–1639 (2023).
Carelli, V. Keeping in shape the dogma of mitochondrial DNA maternal inheritance. PLoS Genet. 11, e1005179 (2015).
Patel, M. R. Inheritance: male mtDNA just can’t catch a break. Curr. Biol. 27, R264–R266 (2017).
Wei, W. & Chinnery, P. F. Inheritance of mitochondrial DNA in humans: implications for rare and common diseases. J. Intern. Med. 287, 634–644 (2020).
Jiang, M. et al. Increased total mtDNA copy number cures male infertility despite unaltered mtDNA mutation load. Cell Metab. 26, 429–436.e424 (2017).
DeLuca, S. Z. & O’Farrell, P. H. Barriers to male transmission of mitochondrial DNA in sperm development. Dev. cell 22, 660–668 (2012).
Zhou, Q. et al. Mitochondrial endonuclease G mediates breakdown of paternal mitochondria upon fertilization. Science 353, 394–399 (2016).
Al Rawi, S. et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science 334, 1144–1147 (2011).
Al Rawi, S. et al. Allophagy: a macroautophagic process degrading spermatozoid-inherited organelles. Autophagy 8, 421–423 (2012).
Djeddi, A. et al. Sperm-inherited organelle clearance in C. elegans relies on LC3-dependent autophagosome targeting to the pericentrosomal area. Development 142, 1705–1716 (2015).
Ben-Hur, S. et al. Egg multivesicular bodies elicit an LC3-associated phagocytosis-like pathway to degrade paternal mitochondria after fertilization. Nat. Commun. 15, 5715 (2024).
Sato, M. & Sato, K. Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 334, 1141–1144 (2011).
Sato, M., Sato, K., Tomura, K., Kosako, H. & Sato, K. The autophagy receptor ALLO-1 and the IKKE-1 kinase control clearance of paternal mitochondria in Caenorhabditis elegans. Nat. cell Biol. 20, 81–91 (2018).
Merlet, J., Rubio-Peña, K., Al Rawi, S. & Galy, V. Autophagosomal sperm organelle clearance and mtDNA inheritance in C. elegans. Adv. Anat., Embryol. Cell Biol. 231, 1–23 (2019).
Sasaki, T. et al. ALLO-1- and IKKE-1-dependent positive feedback mechanism promotes the initiation of paternal mitochondrial autophagy. Nat. Commun. 15, 1460 (2024).
Lim, Y. et al. FNDC-1-mediated mitophagy and ATFS-1 coordinate to protect against hypoxia-reoxygenation. Autophagy 17, 3389–3401 (2021).
Rojansky, R., Cha, M. Y. & Chan, D. C. Elimination of paternal mitochondria in mouse embryos occurs through autophagic degradation dependent on PARKIN and MUL1. eLife 5, e17896 (2016).
Rubio-Peña, K. et al. Mitophagy of polarized sperm-derived mitochondria after fertilization. iScience 24, 102029 (2021).
Zhou, Q., Li, H. & Xue, D. Elimination of paternal mitochondria through the lysosomal degradation pathway in C. elegans. Cell Res. 21, 1662–1669 (2011).
Sutovsky, P. et al. Ubiquitin tag for sperm mitochondria. Nature 402, 371–372 (1999).
Molina, P., Lim, Y. & Boyd, L. Ubiquitination is required for the initial removal of paternal organelles in C. elegans. Dev. Biol. 453, 168–179 (2019).
Dura, M. et al. DNMT3A-dependent DNA methylation is required for spermatogonial stem cells to commit to spermatogenesis. Nat. Genet. 54, 469–480 (2022).
Vallet-Buisan, M., Mecca, R., Jones, C., Coward, K. & Yeste, M. Contribution of semen to early embryo development: fertilization and beyond. Hum. Reprod. update 29, 395–433 (2023).
Xu, R., Li, C., Liu, X. & Gao, S. Insights into epigenetic patterns in mammalian early embryos. Protein cell 12, 7–28 (2021).
Guo, H. et al. Alteration of RNA modification signature in human sperm correlates with sperm motility. Mol. Hum. Reprod. 28, 1360–9947 (2022).
Tian, M. et al. Association of DNA methylation and mitochondrial DNA copy number with human semen quality. Biol. Reprod. 91, 101 (2014).
Houshdaran, S. et al. Widespread epigenetic abnormalities suggest a broad DNA methylation erasure defect in abnormal human sperm. PLoS ONE 2, e1289 (2007).
Zeng, Y. & Chen, T. DNA methylation reprogramming during mammalian Development. Genes 10, 2073–4425 (2019).
Nordstrand, L. M., Furu, K., Paulsen, J., Rognes, T. & Klungland, A. Alkbh1 and Tzfp repress a non-repeat piRNA cluster in pachytene spermatocytes. Nucleic acids Res. 40, 10950–10963 (2012).
Kawarada, L. et al. ALKBH1 is an RNA dioxygenase responsible for cytoplasmic and mitochondrial tRNA modifications. Nucleic acids Res. 45, 7401–7415 (2017).
Liu, F. et al. ALKBH1-mediated tRNA demethylation regulates translation. Cell 167, 1897 (2016).
Müller, T. A., Struble, S. L., Meek, K. & Hausinger, R. P. Characterization of human AlkB homolog 1 produced in mammalian cells and demonstration of mitochondrial dysfunction in ALKBH1-deficient cells. Biochem. Biophys. Res. Commun. 495, 98–103 (2018).
Pan, Z. et al. Impaired placental trophoblast lineage differentiation in Alkbh1(-/-) mice. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 237, 316–327 (2008).
Nordstrand, L. M. et al. Mice lacking Alkbh1 display sex-ratio distortion and unilateral eye defects. PLoS ONE 5, e13827 (2010).
Liu, Y., Yuan, Q. & Xie, L. The AlkB family of Fe (II)/Alpha-ketoglutarate-dependent dioxygenases modulates embryogenesis through epigenetic regulation. Curr. Stem Cell Res. Ther. 13, 136–143 (2018).
Hahn, A. et al. Misregulation of mitochondrial 6mA promotes the propagation of mutant mtDNA and causes aging in C. elegans. Cell Metab. 36, 2528–2541.e2511 (2024).
Wang, Y. et al. Kinetics and specificity of paternal mitochondrial elimination in Caenorhabditis elegans. Nat. Commun. 7, 12569 (2016).
Gitschlag, B. L. et al. Homeostatic responses regulate selfish mitochondrial genome dynamics in C. elegans. Cell Metab. 24, 91–103 (2016).
Wagner, A. et al. Mitochondrial Alkbh1 localizes to mtRNA granules and its knockdown induces the mitochondrial UPR in humans and C. elegans. J. Cell Sci. 132, jcs223891 (2019).
Wu, T. P. et al. DNA methylation on N6-adenine in mammalian embryonic stem cells. Nature 532, 329–333 (2016).
Xiao, C.-L. et al. N6-methyladenine DNA modification in the human genome. Mol. cell 71, 306–318.e307 (2018).
Zhang, L. S. et al. ALKBH7-mediated demethylation regulates mitochondrial polycistronic RNA processing. Nat. cell Biol. 23, 684–691 (2021).
Yang, Q. et al. LONP-1 and ATFS-1 sustain deleterious heteroplasmy by promoting mtDNA replication in dysfunctional mitochondria. Nat. cell Biol. 24, 181–193 (2022).
Gabrielli, A. P. et al. Inhibiting mtDNA transcript translation alters Alzheimer’s disease-associated biology. Alzheimer’s. Dement. 20, 8429–8443 (2024).
Selwood, S. P., McGregor, A., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Inhibition of mitochondrial protein synthesis promotes autonomous regulation of mtDNA expression and generation of a new mitochondrial RNA species. FEBS Lett. 494, 186–191 (2001).
Quirós, P. M. et al. Multi-omics analysis identifies ATF4 as a key regulator of the mitochondrial stress response in mammals. J. Cell Biol. 216, 2027–2045 (2017).
Izawa, T., Park, S. H., Zhao, L., Hartl, F. U. & Neupert, W. Cytosolic protein Vms1 links ribosome quality control to mitochondrial and cellular homeostasis. Cell 171, 890–903.e818 (2017).
Su, T. et al. Structure and function of Vms1 and Arb1 in RQC and mitochondrial proteome homeostasis. Nature 570, 538–542 (2019).
Bertram, N. et al. Delayed protein translocation protects mitochondria against toxic CAT-tailed proteins. Mol. cell S1097-2765, 00815–00819 (2025).
Topf, U., Wrobel, L. & Chacinska, A. Chatty mitochondria: keeping balance in cellular protein homeostasis. Trends Cell Biol. 26, 577–586 (2016).
Anderson, N. S. & Haynes, C. M. Folding the mitochondrial UPR into the integrated stress response. Trends Cell Biol. 30, 428–439 (2020).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Su, L., Zhang, J., Gomez, H., Kellum, J. A. & Peng, Z. Mitochondria ROS and mitophagy in acute kidney injury. Autophagy 19, 401–414 (2023).
Paek, J. et al. Mitochondrial SKN-1/Nrf mediates a conserved starvation response. Cell Metab. 16, 526–537 (2012).
Palikaras, K., Lionaki, E. & Tavernarakis, N. Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521, 525–528 (2015).
Rauthan, M., Ranji, P., Aguilera Pradenas, N., Pitot, C. & Pilon, M. The mitochondrial unfolded protein response activator ATFS-1 protects cells from inhibition of the mevalonate pathway. Proc. Natl. Acad. Sci. USA 110, 5981–5986 (2013).
Liu, P. et al. Mitopherogenesis, a form of mitochondria-specific ectocytosis, regulates sperm mitochondrial quantity and fertility. Nat. cell Biol. 25, 1625–1636 (2023).
Zhang, H., Zhu, Y. & Xue, D. Moderate embryonic delay of paternal mitochondrial elimination impairs mating and cognition and alters behaviors of adult animals. Sci. Adv. 10, eadp8351 (2024).
Wang, S. & Xue, D. Asymmetric partitioning of persistent paternal mitochondria during cell divisions safeguards embryo development and mitochondrial inheritance. Dev. Cell 60, 1730–1750.e1710 (2025).
Wang, P. et al. RNA-binding protein complex AMG-1/SLRP-1 mediates germline development and spermatogenesis by maintaining mitochondrial homeostasis in Caenorhabditis elegans. Sci. Bull. 68, 1399–1412 (2023).
Otten, A. B. & Smeets, H. J. Evolutionary defined role of the mitochondrial DNA in fertility, disease and ageing. Hum. Reprod. update 21, 671–689 (2015).
Zhang, M. et al. Mammalian ALKBH1 serves as an N(6)-mA demethylase of unpairing DNA. Cell Res. 30, 197–210 (2020).
Song, G. J. & Lewis, V. Mitochondrial DNA integrity and copy number in sperm from infertile men. Fertil. Steril. 90, 2238–2244 (2008).
Zhang, G. et al. Associations of ambient air pollutant exposure with seminal plasma MDA, sperm mtDNA copy number, and mtDNA integrity. Environ. Int. 136, 105483 (2020).
Lin, J. L. J. et al. Oxidative stress impairs cell death by repressing the nuclease activity of mitochondrial endonuclease G. Cell Rep. 16, 279–287 (2016).
Greer, E. L. et al. DNA methylation on N6-adenine in C. elegans. Cell 161, 868–878 (2015).
Zheng, J. et al. Lycium barbarum glycopetide prolong lifespan and alleviate Parkinson’s disease in Caenorhabditis elegans. Front. Aging Neurosci. 15, 1156265 (2023).
Aspernig, H. et al. Mitochondrial perturbations couple mTORC2 to autophagy in C. elegans. Cell Rep. 29, 1399–1409.e1395 (2019).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32430049) (Q.Z.), National Key R&D Program of China (2021YFA0804903) (Q.Z.), Programme of Introducing Talents of Discipline to Universities (B14036) (Q.Z.), the Guangdong Basic and Applied Basic Research Foundation (2023B1515120089) (Q.Z.). Qinghua Zhou also gratefully acknowledges the support of the K.C. Wong Education Foundation. We thank the Caenorhabditis Genetic Center (CGC) and the National BioResource Project (NBRP) for strains; Dr Tian Y (the Genetics and Developmental Biology, Chinese Academy of Sciences) and Dr Miao L (the Key Laboratory of RNA Biology, Chinese Academy of Sciences) for providing the worm strains; Dr Yan GR (Biomedicine Research Center, the Third Affiliated Hospital of Guangzhou Medical University) for helping with the demethylation activity analysis of ALKB-1.
Author information
Authors and Affiliations
Contributions
Q.Z. and Q-L.W. conceptualized and conceived the study. Q. Z., Q-L.W. and Z.L. designed the study. Z.L., Y.L., C.H., D.W., W.D., L.H. and J.Y. performed the experiments. Z.L. and Y.L. analyzed the data. Q-L.W. and Z.L. wrote the manuscript. Q.Z., Q-L.W., Z.J., N.T. and H.W. reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, Z., Li, Y., He, C. et al. ALKB-1-dependent tRNA methylation is required for efficient paternal mitochondrial elimination. Nat Commun 17, 2144 (2026). https://doi.org/10.1038/s41467-026-68813-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68813-6