Abstract
Wild Arachis species exhibit abundant genetic diversity for peanut improvement. However, the evolutionary history of their genomes is unclear. Here, through comparative oligopainting, we establish a one-to-one correspondence between microscopically observed chromosomes and sequenced pseudomolecules of Arachis duranensis, A. ipaensis, and cultivated peanut, and determine the 10 homoeologous groups (Hgs) of the A, B, F, K, and H genomes. Analysis of the telomere-to-telomere (T2T) genome assembly of A. hoehnei reveals that its genome is a diverging form of the A genome and is designated as genome A’. In addition, the unique small chromosome A08 is originated from two inversions and a substantial contraction of A'08. We propose a genome evolution model of the Arachis genus, where A’ bridges the A and B genomes. We further report an artificial hexaploid peanut derived from a hybrid of cultivated peanut and A. hoehnei, and identify differentially expressed genes against web blotch in A. hoehnei.
Similar content being viewed by others
Introduction
Peanut (Arachis hypogaea L., AABB, 2n = 4x = 40), an important food and oil crop, is an allotetraploid and self-pollinating crop with a narrow genetic background. Arachis species, some of which have excellent resistance to pests, diseases, and drought, represent an invaluable gene pool for peanut breeding1. Several genes conferring resistance to nematodes, rust and leafspot have been introgressed from wild species into cultivated peanut2,3,4.
The Arachis genus diverged from other legumes approximately 50 million years ago (Mya)5. The genus originated in South America and is widely distributed from the eastern foothills of the Andes in Bolivia and northern Argentina to the Atlantic coast in Brazil and from the southern limit of the Amazon region to the northern coast of the Rio de la Plata in Uruguay6,7. Arachis comprises 83 species8, which have been classified into nine sections based on their morphology, cross-compatibility, hybrid viability, geographic distribution, and cytogenetic characteristics, namely, Trierectoides, Erectoides, Procumbentes, Rhizomatosae, Heteranthae, Caulorrhizae, Extranervosae, Triseminatae, and Arachis9,10,11. Section Arachis includes cultivated peanut, the tetraploid wild species A. monticola, and 29 diploid wild species. Currently, the genomes of Arachis are preliminarily divided into the A, B, D, F, K, and G genomes12,13,14,15,16,17. However, clear evidence supporting the origin and genome evolution of these species is still lacking.
Genome sequencing facilitates an understanding of the origin and genome evolution of Arachis species. The A genome of A. duranensis and the B genome of A. ipaensis were considered the donors of cultivated peanut, which diverged approximately 2–2.60 Mya18. Species with the A genome harbor a smaller size chromosome A08, also called “small chromosome” or “A chromosome”19. Comparative genomics analyses validated the genomic origin of cultivated peanut, where high collinearity of between wild species and cultivated peanut exists in both A and B genomes. The small chromosome A08 in A-genome species was probably derived from translocations and inversions between chromosomes 7 and 8 of its ancient ancestor, where the repeat-enriched segment of chromosome 8 was translocated to chromosome 7, while the gene-enriched segment of chromosome 7 was translocated to chromosome 820. To date, genome assemblies of Arachis species have been reached to telomere-to-telomere (T2T) level21,22,23,24. Recent pangenome analyses of eight Arachis species and cultivars have revealed frequent structural variations (SVs) during peanut evolution, and 1335 SVs likely associated with domestication have been identified in the A and B genomes25.
Existing studies mainly focused on the A and B genomes of the Arachis species. Limited information is available for D, F, G, H, and K genomes, e.g. genome identity and evolution, homoeologous chromosome pairs or groups. Comparative oligopainting using chromosome-specific single-copy oligonucleotide libraries (CSCOLs) and tandem repeat oligonucleotides provides a fast and efficient tool for determining homoeologous chromosome pairs and large segmental structural chromosome variations (SCVs) during the genome evolution of different species in large samples26,27. In addition, the rapid development of T2T genome assembly techniques facilitates the accurate identification of whole-genome variations28.
In this work, comparative oligopainting not only reveals the correspondence of all the pseudomolecules of A. duranensis to its actual chromosomes but also determines 10 homoeologous groups (Hgs) and SCVs in the A, B, F, K, and H genomes. By combining this information with the T2T genome assembly of A. hoehnei (Zw61), we identify its genome identity, propose the origin of the small chromosome, and elucidate the genome evolution of Arachis species. Further, we report an artificial hexaploid peanut line derived from a hybrid of cultivated peanut and A. hoehnei, and identify differentially expressed genes against web blotch in A. hoehnei.
Results
Development of a consensus karyotype of A. duranensis
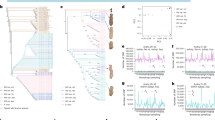
Based on the reference genome sequence of A. duranensis, 10 CSCOLs named L1A-1 to L10A-1 were designed for one or two terminal regions of chromosomes A01 – A10. For each library, 26,365–27,392 unique oligonucleotides (42–48 nt in length) were designed from segments of 4–9 Mb (Fig. 1a and Supplementary Table 1). The oligonucleotides of each library were then mapped in silico onto the corresponding pseudomolecules of A. duranensis, and all sequences were consistent with the target regions (Supplementary Fig. 1).
a Target regions of the 10 CSCOLs on pseudomolecules in A. duranensis. b Sequential FISH karyotype of A. duranensis identified using 10 CSCOLs (first column), SSON multiplex #1 (second column), and 45S (red) and 5S rDNA (green) probes (third column). In SSON, red represents signals from the oligonucleotides DP-1, DP-3, DP-4, and DP-6, whereas green represents signals from the oligonucleotides DP-2, DP-5, and DP-7. c Karyotypes of A. duranensis. d Idiogram of A. duranensis. Source data are provided as a Source Data file.
L1A-1, L2A-1, L3A-1, L8A-1 and L9A-1 were labeled with digoxigenin-11-dUTP and detected using anti-digoxigenin-rhodamine, whereas L4A-1, L5A-1, L6A-1, L7A-1 and L10A-1 were labeled with biotin-16-dUTP and detected using a fluorescein-labeled antibiotin antibody. The CSCOLs were divided into five pairs, each containing one digoxigenin-11-UTP-labeled library and one biotin-16-dUTP-labeled library. We used these probe libraries to paint the chromosomes of A. duranensis (PI 219823). As expected, all 10 CSCOLs produced clear signals in one or two terminal regions of a single pair of chromosomes corresponding to A01 – A10 (Supplementary Fig. 2).
Sequential FISH was conducted using single-strand oligonucleotide (SSON) multiplex #1 (DP-1, DP-2, DP-3, DP-4, DP-5, DP-6, and DP-7) and 45S and 5S rDNA probes following CSCOL painting to determine the correspondence between the pseudomolecules and actual chromosomes of A. duranensis (Fig. 1b, Supplementary Table 2). A karyotype integrating signals from CSCOLs, SSONs and rDNA probes revealed their correspondence to chromosomes and pseudomolecules. The 10 chromosomes were clearly differentiated based on their unique signal patterns and were designated Adu01 through Adu10 (Figs. 1c, d). Analysis of the arm ratios of the actual chromosomes revealed that the six pseudomolecules started from the ends of the short arms of chromosomes Adu02, Adu04, Adu06, Adu07, Adu08 and Adu09, whereas the other four pseudomolecules were from the ends of the long arms of Adu01, Adu03, Adu05 and Adu10.
Each chromosome exhibited a unique signal pattern. Adu01 presented strong green signals for SSON in both the centromeric and short arm terminal regions. Adu02 exhibited very strong green SSON signals in the centromeric and pericentric regions. Adu03 presented only a very weak green telomere signal on its short arm. Adu04 showed strong green SSON signals in the centromeric and pericentric regions. Adu05 exhibited red and green signals in the centromeric and pericentric regions, as well as green signals from the telomeric repeats of DP-5 (a SSON probe) centrally along the long arm. Adu06 presented very strong red and green SSON signals in the centromeric and pericentric regions, along with 5S rDNA sites in the short-arm pericentric region. Adu07 is the only submetacentric (sm) chromosome in A. duranensis and is characterized by very strong SSON signals and red 45S rDNA signals in the centromeric and pericentric regions. Adu08 is the smallest chromosome in A. duranensis, with strong red and green SSON signals in the centromeric and pericentric regions. Adu09 has a very weak green SSON signal in the centromeric region. Adu10 showed weak green SSON signals in the centromeric region and strong 45S rDNA signals in the long-arm pericentric regions (Figs. 1c, d).
Determination of homoeologous chromosome pairs in Arachis species
The method described above was used to analyze 16 other Arachis species/accessions, including three A. hypogaea cultivars, A. monticola, and 12 wild diploid species/accessions, where A. duranensis (PI 219823) was the control (Table 1, Supplementary Figs. 3 to 18). CSCOL painting produced clear signals in all species, allowing consensus karyotypes and ideograms to be established. This not only enabled identification of 10 homoeologous groups (Hgs) named Hg1–10 for Arachis species but also facilitated detection of SCVs that occurred during evolution of Arachis (Figs. 2 and 3a, Supplementary Table 3).
Signals from the CSCOLs, SSON, and 45S and 5S rDNA probes are as described for Fig. 1.
a Idiogram showing homoeologous chromosome groups in Arachis species based on signals from the SSON (green and orange correspond to signals generated by FAM- and TAMRA-labeled SSON probes, respectively; yellow indicates signals generated by both probes simultaneously), 45S (blue) and 5S rDNA (purple) probes. sm and st, submetacentric and subtelocentric chromosomes, respectively. b SCVs observed in chromosomes 7/8 across the A, B, F, K, A’ and H genomes. c SCVs identified in chromosomes 2, 5, 6, 9, and 10 within genome H. “*” indicates heterogeneous sites; sm, submetacentric; st, subtelocentric; bar, 2 μm. Source data are provided as a Source Data file.
Each of the 10 CSCOLs produced clear signals on one pair of chromosomes in all 17 species/accessions, indicating that these unique single-copy sequence blocks are conserved. However, only three CSCOLs (L1A-1, L3A-1, and L4A-1) produced signals at similar positions on the same chromosome pairs in all species. The other seven CSCOLs either hybridized to different positions on the same chromosome pair or to different chromosomes entirely in the B subgenomes of A. hypogaea and A. monticola, as well as in A. ipaensis, A. magna, A. trinitensis, A. hoehnei, A. batizocoi, A. pusilla, and A. dardani. These findings indicate that large segmental SCVs occurred during Arachis evolution. A total of six SCVs were identified in these species, including four types of translocations, one inversion, and one more complex variation (translocation plus inversion) (Fig. 3b, c; Supplementary Fig. 19; Supplementary Table 3).
Among the SCVs, four translocations — between Hpu06 and Hpu10, between Hda05 and Hda09, between Hda02 and Hda10, and between Hda06 and Hda10—occurred in H-genome species (Fig. 3c, Supplementary Fig. 19). The inversion and complex variation involved an inversion of chromosome 8 of A. hoehnei relative to the “small chromosome” A08 and translocations between chromosomes 7 and 8 in the A. hoehnei genome relative to the B, F, K, and H genomes. Based on the variations in chromosomes 7 and 8, the 13 wild diploid species were divided into three groups: 1) Group I - pretranslocation types: it includes all species with the B, F, K, and H genomes, do not exhibit chromosomal translocations involving chromosomes 7/8. 2) Group II - posttranslocation but preinversion type for chromosome 8: includes only A. hoehnei. This species contains a subtelocentric chromosome 8 and a metacentric chromosome 7 formed by translocation between ancestral chromosomes 7 and 8. 3) Group III - postinversion type for chromosome 8: includes all A-genome species. These species are characterized by the small chromosome A08, which is inverted relative to chromosome 8 of A. hoehnei (Fig. 3b).
Additionally, L1A-1 and L3A-1 have extra signals in the pericentric regions of different chromosomes in three cultivars (Tifrunner TIF, Baisha1016 BS and Silihong SLH) and six wild species (A. monticola PI 263393, A. duranensis PI 468202, A. villosa PI 298636, A. herzogii PI 476008, A. simpsonii Grif 14534 and A. diogoi PI 276235), indicating that some unique oligonucleotide sequences in A. duranensis were amplified in other genomes (Supplementary Figs. 3–11).
Cytological characterization of SCVs caused by tandem repeats
Sequential FISH using seven tandem repeat-derived SSON probes was conducted after CSCOL painting in 17 species/accessions to further clarify SCVs during evolution (Figs. 2 and 3a). A consensus karyotype integrating both the SSON and CSCOL signals was established. The karyotypes of the Ahy and Amo subgenomes of cultivated peanut and A. monticola were highly similar to that of the Adu genome of A. duranensis (PI 219823 and PI 468202). The karyotypes of the Bhy and Bmo subgenomes of cultivated peanut and A. monticola were nearly identical to those of the Bip genome of A. ipaensis (PI 468322) and Bma genome of A. magna (PI 468337), except that chromosome Bma08 lacked 45S rDNA sites. In contrast, the karyotypes of the Av, Ahe, Asi, Adi, Kba, Ftr, Hda, and Hpu genomes, as well as the A. hoehnei genome, differed significantly from those of the A and B subgenomes of cultivated peanut and A. monticola (Figs. 2 and 3a).
The seven tandem repeat-derived SSON probes produced unique signals at different positions on the chromosomes: DP-1 signals were consistent with those of the 5S rDNA probe; probes DP-2, DP-3, DP-4, DP-6, and DP-7 produced signals in the centromeric and pericentromeric regions; and DP-5 produced signals in telomeric regions29. A comparison of the karyotypes revealed that chromosomes in the 10 Hgs exhibited variable copy numbers and presence/absence variations (CNVs and PAVs) associated with these seven tandem repeats. Heterogeneous sites of 45S rDNA and SSONs were frequently observed on chromosomes Hg1, Hg8, and Hg10 of Av; Hg6 and Hg10 of Ahe; Hg2, Hg4, Hg6, and Hg7 of Asi; and Hg2, Hg4, Hg5, and Hg8 of Adi. In particular, interstitial telomere repeats (ITRs) occurring at various positions, including telomeric, centromeric and interstitial chromosomal regions, were detected in almost all species (Figs. 2 and 3a).
The signal pattern and intensity variation in the 10 Hgs were as follows:
Hg1 — chromosomes in this group displayed green ITR signals on the short arm in seven genomes (Ahy, Amo, Adu, Av, Ahe, Asi, and A. hoehnei genomes). 45S rDNA sites were detected only in the Av genome, whereas 5S rDNA sites were exclusive to the Kba genome. Green SSON signals were observed in the centromeric and pericentromeric regions of seven genomes (Ahy, Amo, Adu, Av, Ahe, Kba and Hpu), with SSON PAV signals detected in the Av genome. Yellow SSON signals were detected in eight genomes (Asi, Adi, Bhy, Bmo, Bip, Bma, Ftr and A. hoehnei genomes), with no signal in the Hda genome.
Hg2 — this group contained 45S rDNA sites in seven genomes (Av, Ahe, Asi, Adi, Kba, Hda and A. hoehnei genomes) and 5S rDNA sites in the Kba genome. Both PAV and CNV (signal intensity variations) were observed, with PAVs occurring in the Asi and Adi genomes, and CNVs detected in the Av genome. Yellow SSON signals were observed in centromeric regions of Hg2 in all B and K genomes; red SSON signals were detected in the A. hoehnei genome; and green SSON signals were detected in the Asi and Adu genomes; and no signal was detected in the other genomes. Additionally, this chromosome in the Av genome had telomeric signals. SSON PAVs were observed in the Asi genome.
Hg3 — centromeric regions of this group showed strong red SSON signals in all B, K and F genomes; weak red signals in the Ahy, Amo and Adu genomes; green SSON signals in the Av, Ahe, Adi and Hpu genomes; and yellow SSON signals in the Kba genome. CNVs were detected in the Adi genome, but no centromeric signals were detected in the Asi and Hda genomes. Additionally, strong green ITR signals were also present on the short arms of Hg3 in the Kba and Hpu genomes.
Hg4 — green centromeric SSON signals were detected in all A genomes; yellow SSON signals were detected in all B genomes; red signals were detected in the Ftr genome; and no centromeric signals were detected in the Kba genome. 45S rDNA sites were present only in the Hpu genome. In the Hda genome, Hg4 chromosomes exhibited green ITR and SSON signals in the centromeric and pericentric regions.
Hg5 — no signal was observed on this chromosome group in the B, F, and H genomes. The Kba genome showed strong green centromeric SSON signals, whereas all the A genomes and A. hoehnei displayed yellow centromeric SSON signals. Five genomes (Ahy, Amo, Adu, Av and A. hoehnei genomes) had green ITR signals in the middle of the long arm.
Hg6 — 5S rDNA sites were present in all genomes except Hpu. The Bhy, Bmo and Bip genomes also contained 45S rDNA sites. Centromeric SSON signals were green in the A. hoehnei and Hpu genomes but were yellow in all other genomes.
Hg7 — chromosomes in this group were submetacentric (sm) in all A genomes but metacentric (m) in other genomes. 45S rDNA sites were detected in all genomes except H and A. hoehnei. Centromeric signal patterns included strong red SSON signals in A. duranensis PI 468202, B and F genomes; yellow SSON signals in K and A. hoehnei; strong green signals in the A genome of A. duranensis PI 219823 and A. simpsonii; and weak red or green signals in the other genomes.
Hg8 — the unique “small chromosome” A08 of all A-genome species displayed yellow centromeric SSON signals. In the A. hoehnei genome, Hg8 chromosomes were subtelocentric (st) with strong yellow centromeric SSON signals. Red centromeric SSON signals were predominant in all B genomes. 45S rDNA sites were present in the Bhy, Bmo, Bip and Hpu genomes, and the Hpu genome additionally contained 5S rDNA sites.
Hg9 — extensive green ITR signals were observed in the centromeric regions of these chromosomes in the B and F genomes. The short arms of Hg9 chromosomes in the Hpu genome also had green ITR signals, but no centromeric signal was detected in any A genome except Ahe.
Hg10 — 45S rDNA signals were detected in all the A genomes except Av. The A. hoehnei genome contained both 45S rDNA and 5S rDNA sites, whereas no rDNA signal was detected in the B, F, K and H genomes. Centromeric signals were green in the A and K genomes, yellow in the B and A. hoehnei genomes, and absent in the H genome. Large telomeric signals were detected at chromosome ends in the Av and Asi genomes.
Telomere-to-telomere genome assembly of A. hoehnei
Cytological analysis revealed that four types of large segmental SCVs (translocations, inversions, CNVs and PAVs) occurred during evolution. Most of these variations are distributed in tandem repeat-enriched regions, including telomeres, centromeres and nucleolar organizer regions (NORs). Based on the evolution of the small chromosome, the A. hoehnei genome seems to be an evolutionary bridge between the B and A genomes. The genomes of A. duranensis and A. ipaensis were clearly designated and well assembled20, but the genome of A. hoehnei remained unknown (Fig. 4a). Therefore, a T2T genome assembly of A. hoehnei and comparative genomic analysis should not only validate these SCVs but also provide a more precise landscape of the origin of the “small chromosome” and genome evolution of peanut.
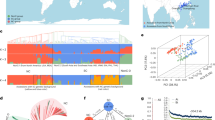
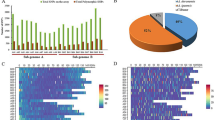
a A. hoehnei plants. b Genome information: chromosome information (a), TE density (b), SSR density (c), gene density (d), and GC content (e). Each term in panels b, c, d, and e represents a scale of 100 kb, and lines within circles illustrate chromosomal collinearity. c Hi-C interaction heatmap indicating that the contigs of A. hoehnei were assembled into 10 chromosomes. d Evolutionary tree showing the relationships among 12 species with differentiation time. The top of the tree is absolute age (million years ago, Mya) and geological time. e Evolutionary tree of the chloroplast genomes of 23 species; outgroups included A. thaliana, C. arietinum, M. truncatula, and G. max.
Genome sequencing resulted in ~67× coverage from raw ultralong Oxford Nanopore Technologies (ONT) data, ~80× coverage from PacBio High Fidelity (HiFi) data, ~49× coverage from Illumina reads, and ~96× coverage from high-throughput chromosome conformation capture (Hi-C) datasets for assembling the A. hoehnei genome (Supplementary Tables 4–6). Based on k-mer analysis of Illumina short reads, the genome size of A. hoehnei was estimated to be ~1.49 Gb, with a heterozygosity rate of ~0.37% (Supplementary Fig. 20a). Hybrid assembly of high-quality HiFi reads and ultralong ONT reads using Hifiasm (v0.19.9) and Verkko (v2.2.1) software to assemble the genome generated 149 contigs with an N50 size of 96.40 Mb. Contaminating sequences were identified by BLAST analysis against bacterial nt databases and organelle sequences and then removed, leaving 37 contigs with an N50 size of 96.39 Mb. Hi-C data were used to map the chromosomes, resulting in a draft A. hoehnei assembly that included 10 pseudomolecules with 20 gaps. These gaps were subsequently filled with HiFi data using QuarTeT software. The ONT data were further assembled using Flye software, and the remaining gaps were filled using TGS-GapCloser and Minimax2 software. A complete T2T assembly of all 10 chromosomes of the A. hoehnei genome was obtained, comprising a total size of 1609.1 Mb (Table 2 and Supplementary Table 7). The basic structural features (gene density, repeat sequence density, and GC density) of the A. hoehnei T2T genome are shown in Fig. 4b.
Multiple methods were used to evaluate the integrity and continuity of the genome. Hi-C interaction heatmaps validated the chromosome order and orientation, revealing strong interactions near the diagonal of each chromosome block, indicating the high-quality assembly of all 10 chromosomes (Fig. 4c). Illumina short reads, HiFi reads, and ultralong ONT reads were mapped to the T2T genome with mapping rates of 99.52%, 99.98%, and 97.23%, respectively. The quality of the T2T genome assembly further assessed using the BUSCO (Benchmarking Universal Single-Copy Orthologs) tool detected 99.13% of the core conserved genes, including 1502 single-copy genes and 98 duplicated genes (Supplementary Fig. 20b). The integrity of long terminal repeat (LTR) sequences was evaluated by calculating the LTR Assembly Index (LAI), which was 27.13, indicating a high-quality genome. A genome consensus quality value (QV) of 49.10 was obtained using Illumina data and the k-mer clustering method in Merqury, confirming that the assembly was highly complete. According to RepeatMasker software, 83.63% (1345.62 Mb) of the A. hoehnei genome consists of repetitive sequences, with transposable elements (TE) accounting for 71.98% (Supplementary Table 8) and tandem repeats accounting for 11.65% (Supplementary Table 9). LTR retrotransposons were the most abundant, comprising 65.02% of the genome, among which Gypsy (45.16%) and Copia (2.81%) were the most prevalent types (Supplementary Table 8).
Telomeres, centromeres, and NORs are key structural components of chromosomes. Using TIDK and FindTelomeres software, we identified 20 telomeric repeat segments on the 10 chromosomes. The telomeres ranged in length from 1004 to 11,265 bp, with an average length of 5721 bp (Supplementary Fig. 20c, Supplementary Table 10). The shortest telomere is located on the short arm of the subtelocentric (st) chromosome 8, and the longest is on chromosome 10. Additionally, the ITR signals from the telomeric probe DP-5 (TTTAGGG)8 were strong in the middle of the short arm of chromosome 1, medium in the middle of the long arm of chromosome 5, and weak near the centromere of chromosome 6. The copy numbers of DP-5 on these chromosomes were 17,546, 5624, and 1262, respectively, indicating that the signal strength increased with increasing copy number (Supplementary Fig. 21a). A BLAST search of the 500-bp subtelomeric sequences near the telomeres of the 10 chromosomes of A. hoehnei against the ITR sequences on chromosomes 1, 5, and 6 showed that these ITRs originated primarily from the telomeres at the starting position of chromosome 3, accounting for 75.22% (45,969 copies), 69.26% (14,919 copies) and 74.75% (3449 copies) of all ITR copies in chromosomes 1, 5 and 6, respectively, followed by the telomeres at the starting position of chromosome 6, accounting for 24.75%, 30.42% and 25.25% of all ITR copies in chromosomes 1, 5 and 6, respectively (Supplementary Table 11).
Centromere positions were predicted using Centromics, quarTeT, and CentIER software. The lengths of the 10 centromeres ranged from ~1.50 Mb to ~11.60 Mb, with an average length of 5.30 Mb. Chromosome 9 has the largest centromere, and chromosome 5 has the smallest, indicating substantial variation in centromere length across the A. hoehnei genome (Supplementary Fig. 21b). In terms of arm ratio, all chromosomes are metacentric (m) except 8, which is subtelocentric (st) (Supplementary Table 12), consistent with FISH karyotyping results. Further analysis of the centromeric sequences revealed a large number of short dispersed repetitive sequences (SINEs) in the centromeric region of chromosome 5, whereas the centromeric regions of the other chromosomes were rich in unknown long terminal repeat sequences (LTRs). Chromosomes 1 and 6 contain no coding genes in the centromeric regions. Among the chromosomes, chromosome 2 has the most coding genes (22) in the centromeric region. The other chromosomes have fewer coding genes in their centromeric regions (17 on chromosome 10, 9 on chromosome 9, 5 on chromosomes 8 and 5, 4 on chromosomes 4, 3 on chromosome 7, and 1 on chromosome 3) (Supplementary Table 12).
We sequenced the transcriptomes of roots, stems, leaves, flowers, and fruits; annotated the protein-coding genes of the A. hoehnei genome; used three prediction methods (homologous protein prediction, de novo prediction, and transcriptome prediction); and ultimately identified 46,625 genes (Supplementary Tables 13 and 14). The average gene length was 3510.12 bp, the average coding sequence (CDS) size was 1059.09 bp, and the average number of exons per gene was 5.06, with an average exon length of 1288.01 bp (Supplementary Table 15 and Supplementary Fig. 22). We also identified 10411 rRNA genes, 909 tRNA genes, 49 miRNAs, 50 snRNAs, and 79 snoRNAs across the genome. The 5S rRNA was transcribed from 5S rDNA, whereas the 18S, 5.8S, and 25S rRNAs were generated through splicing of the single 45S transcript encoded by the 45S rDNA. There were high-frequency 5S rDNA copies on chromosomes 2 (1581 copies, near 46 Mb), 6 (512 copies, near 39 Mb), and 10 (1,504 copies, near 79 Mb; and 3824 copies, near 136 Mb) concentrated near the centromeres and on long arms. In parallel, 5S rDNA FISH signals were strong on chromosome 10, weaker on chromosome 2, and weakest on chromosome 6. 45S rDNA was located mainly on chromosome 2 (566 copies, near 71 Mb; and 31 copies, near 96 Mb), chromosome 10 (390 copies, near 88 Mb), and chromosome 7 (12 copies) (Supplementary Table 16). FISH signals for 45S rDNA were strong on chromosomes 2 and 10, which are high-copy-number locations, whereas no signals were detected for the 31 copies on chromosome 2 and 12 copies on chromosome 7 (Supplementary Fig. 21a). These results show that the signal strength increased with copy number. In silico mapping using the seven tandem repeat-derived SSONs and 45S and 5S rDNA sequences further validated the high quality of the T2T genome assembly, where signals and even signal intensity in target regions corresponded with those in the actual chromosomes and T2T pseudomolecules (Supplementary Fig. 21a). However, some signals in actual chromosomes failed to correspond to their sequences in the pseudomolecules of A. duranensis, A. ipaensis, and cultivar Tifrunner (Supplementary Fig. 23), where five sites in A. duranensis and A. ipaensis, respectively, and two sites in Tifrunner showed clear signals that did not correspond to the probe sequences mapped in the pseudomolecules, thus indicating poor genome assembly of those regions.
Comparative genomic analysis
Eight Arachis species have either chromosome-level or T2T genome assemblies, including A. ipaensis, A. duranensis, A. cardensii, A. stenosperma, A. hypogaea (cultivars Tifrunner, YZ9102 and Shitouqi STQ), and A. monticola. These species, together with Arabidopsis thaliana, Cicer arietinum, Medicag truncatula and Glycine max, were used for analysis of divergence time of A. hoehnei. A phylogenetic analysis based on 390 single-copy genes using A. thaliana, C. arietinum, M. truncatula, and G. max as outgroup species indicates that Arachis species diverged from other leguminous species approximately 64.34 (50.01 – 89.47) Mya. The divergence between the B genome and the clade containing the A and A. hoehnei genomes occurred approximately 2.29 (2.23 – 2.35) Mya, and the A and A. hoehnei genomes diverged approximately 1.78 (1.654 – 1.90) Mya (Fig. 4d). These findings indicate that the A. hoehnei genome is more closely related to the A genome than to the B genome.
Chloroplast assembly and phylogenetic analysis were conducted to validate these results. Using the above outgroup species, phylogenetic analysis showed that Arachis species clustered into a single branch, which diverged into three lineages. Among the 19 species/accessions, two H-genome species formed lineage III, and the remaining 17 species were divided into two major lineages. Lineage II includes species with the Ftr, Bip, Bma, and Kba genomes, with the Ftr genome located at the base branch of the phylogenetic tree, indicating that it originated earlier than the Kba, Bip, and Bma genomes. Lineage I was further divided into two clades: cultivated peanut and two accessions of A. duranensis (PI 468202 and PI 219823) grouped as Clade A, and species with the Av, Ahe, Asi, and Adi genomes and A. hoehnei grouped as Clade B. This suggested that the A. hoehnei genome is closely related to the A genome (Fig. 4e), consistent with results described above.
Using the T2T genome of A. hoehnei as a reference, the whole-genome SNPs of 15 wild diploid species were identified and used for phylogenetic analysis. The species again clustered into three lineages consistent with the chloroplast results, whereby the A. hoehnei genome was closer to the A genome but diverged earlier (Supplementary Fig. 24). Therefore, its genome was designated A’.
Gene family clustering analysis of 12 species revealed 4,398 shared gene families; 367 gene families (1019 genes) were specific to A. hoehnei, a number far greater than that for specific gene families in the A (A. duranensis, 147 families, 354 genes) and B (A. ipaensis, 228 families, 538 genes) genomes (Fig. 5a). These specific genes were distributed across all 10 chromosomes of A. hoehnei, with chromosome 5 having the most, followed by chromosomes 3 and 10. The results of copy number analysis for all gene families across the 12 species are shown in Fig. 5b. Compared with cultivated peanut, A. duranensis and A. ipaensis, A. hoehnei has more multicopy genes, suggesting it underwent more gene duplication events during evolution. In addition, 1697 gene families were expanded, and 948 gene families were contracted (Supplementary Table 17). Sixty-five genes were positively selected during evolution of A. hoehnei. Functional enrichment analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG) revealed that specific genes of A. hoehnei were enriched in pathways such as ether lipid metabolism, glycerophospholipid metabolism, base excision repair, ubiquinone and other terpenoid-quinone biosynthesis, aminoacyl-tRNA biosynthesis, purine metabolism and endocytosis (Supplementary Fig. 25, Supplementary Table 17). Some of the expanded genes and positively selected genes were significantly enriched in pathways such as nucleotide excision repair, DNA replication, mismatch repair, and homologous recombination (Supplementary Figs. 26 and 27). The expanded genes involved in these pathways in A. hoehnei indicate that this species evolved an enhanced DNA repair system to maintain its genome stability.
a Petal map of gene family clusters; the central circle represents common gene families, and the outer petals denote specific gene families. b Distribution of copy numbers for all gene families across the twelve species. c Genome collinearity of A. hoehnei with A. duranensis and A. ipaensis, YZ9102 A and YZ9102 B subgenomes, respectively. d Structural variation (SV) model between chromosomes 7 and 8 across YZ9102 A subgenome, A. hoehnei and YZ9102 B subgenome. e Numbers of different types of TEs in the YZ9102 A subgenome, YZ9102 B subgenome and A. hoehnei. f Compared with those in the YZ9102 A and B subgenomes, SVs involving TE insertions accounted for the total SVs in A. hoehnei. g Oligo GISH using the A genome-specific probe set multiplex #A (A-4 and A-7, green) and the B genome-specific probe set multiplex #B (B-9 and B-10, red) derived from the TE sequences on metaphase chromosomes of A. hoehnei and YZ9102, respectively. All oligo GISH experiments were independently repeated three times with similar results. Bar, 10 μm.
The T2T genomes of A. hoehnei and cultivar YZ9102 were used as references for comparison with other genomes to elucidate syntenic blocks and structural and sequence variations in the eight Arachis species. Based on the percentage of syntenic block length relative to genome size of each species, the average percentages of syntenic blocks across the three genomes representing these eight species were ordered as follows: B–B (84.01%) > A–A (67.11%) > A’–A (23.56%) > A–B (18.08%) > A’–B (17.45%). This order was consistent with previous phylogenetic analysis results and divergence times of the species (Fig. 4d, Supplementary Fig. 28, and Supplementary Data 1). However, frequent chromosomal rearrangements, including translocations, inversions, duplications, insertions and deletions ( > 50 bp), and sequence variations (SNPs and indels) were observed. Compared with the three reference T2T genomes (A’, YZ9102 A and YZ9102 B), the number of SVs ( > 50 bp) among the different genomes was ordered as A’–A (47,591) > A–B (30,955) > A’–B (28,717) > B–B (23,979) > A–A (22,836) (Supplementary Fig. 29, Supplementary Data 1). The numbers of SNPs and indels ( < 50 bp) were ordered as A’–A (4,317,609) > A–A (3682,424) > A’–B (2,122,313) > A–B (1,994,344) > B–B (1,069,269) (Supplementary Fig. 29, Supplementary Data 1). Compared with the A genomes, the A’ genome had inversions exceeding 5 Mb on chromosomes 1, 3, 5, 8, 9 and 10, whereas compared with all B genomes, the A’ genome had inversions exceeding 5 Mb on chromosomes 1, 5, 6, 7 and 9. Large segmental translocations or complex variations were also observed in chromosomes 7 and 8 (Supplementary Fig. 28). These results indicate that A. hoehnei is quite distinct from species containing both the A and B genomes.
The origin of the small chromosome
Chromosome A08 is the smallest and most obvious chromosome in the A genome15. The length of chromosome A08 in A. duranensis, A. cardensii, A. stenosperma, Tifrunner A, YZ9102 A, STQ A and A. monticola A ranged from 47.73 Mb to 60.52 Mb, with an average of 53.51 Mb. The length of chromosome B08 in Tifrunner B, YZ9102 B, STQ B, A. ipaensis, and A. monticola B ranged from 129.60 Mb to 137.84 Mb, with an average of 135.36 Mb. The length of chromosome A'08 was 115.65 Mb. The average number of genes on chromosome A08 was 3,073, that on chromosome B08 was 3580, and that on chromosome A'08 was 4,131 (Supplementary Table 18). Both cytological and comparative genomic analyses showed that chromosome A08 originated from three major rearrangements. First, segmental translocations between chromosomes A'07 and A'08 in A. hoehnei relative to the B genomes (Fig. 5c); second, two inversions in chromosome A08 relative to chromosome A'08; and third, one substantial genome contraction of chromosome A08 compared with chromosome A'08 (Fig. 5d, Supplementary Fig. 30, Supplementary Table 19).
Collinearity analysis was performed using the T2T genome sequences of YZ9102 and A. hoehnei to precisely determine the rearrangements and relative breakpoints. First, compared with the B genome of YZ9102, there were six major rearrangements in chromosomes 7 and 8 of A. hoehnei. These rearrangements include a segment from 28.50–113.00 Mb of chromosome B08 translocated to two regions (32.00–55.00 Mb and 91.00–101.00 Mb) in chromosome A'07 (Translocations 1 and 2 in Fig. 5d); a segment from the 0.03–1.41 Mb region of chromosome B08 translocated and inverted to the 101.05–102.49 Mb region in chromosome A'07 (Translocation–inversion 2 in Fig. 5d); a segment from the 107.69 – 122.44 Mb region of chromosome B07 translocated to the 64.48 – 80.96 Mb region in chromosome A'08 (Translocation 3 in Fig. 5d); a segment of 123.96–140.30 Mb of chromosome B07 translocated and inverted to the 85.99–103.62 Mb region of chromosome A'08 (Translocation–inversion 1 in Fig. 5d); and an inversion involving the 29.19 – 102.40 Mb region of chromosome B07 and the 55.85–90.76 Mb region of chromosome A'07 (Inversion 3 in Fig. 5d). Second, compared with chromosome A'07, no large SVs were observed on chromosome A07, indicating a strong syntenic relationship between the two chromosomes and an evolutionary connection between the A’ and A genomes. However, compared with chromosome A'08, two large segmental inversions were present in chromosome A08. Inversion 1 was located in the 2.59 – 62.24 Mb region of chromosome A'08, whereas Inversion 2 was identified in the 64.50 – 80.96 Mb region. The breakpoints of Inversion 1 were at 229,789 bp and 4,934,848 bp on chromosome A08, and the breakpoints of Inversion 2 were at 5,708,618 bp and 20,016,695 bp. No breakpoint was located within a gene, but they were all in intergenic regions and involved LTR and TIR sequences. The difference in genome size between chromosomes A'08 (115.65 Mb) and A08 (60.52 Mb) was 55.13 Mb. In Inversion 1, both gene and TE contraction were observed. Specifically, the length of Inversion 1 on chromosome A'08 was 59.65 Mb, but on chromosome A08, it was reduced to only 4.70 Mb - a substantial reduction of approximately 54.95 Mb comprising the loss of approximately 46.35 Mb of TEs along with 495 genes, which have homologous genes in chromosomes B07 or B08 (Fig. 5d, Supplementary Fig. 30, Supplementary Table 19). KEGG enrichment analysis indicates that genes missing from chromosome A08 relative to chromosome A'08 were involved in various pathways, such as homologous recombination, carbon metabolism, nucleotide excision repair, mismatch repair, carbon fixation by the Calvin cycle, and DNA replication (Supplementary Fig. 31). Thus, the chromosomal deletions associated with Inversion 1 are a primary factor contributing to the observed disparity in chromosome lengths between chromosomes A'08 and A08. The 495 genes lost in Inversion 1 of chromosome A08 remained in chromosomes B07 and B08, suggesting that chromosome A'08 originated from translocations between chromosomes 7 and 8, followed by inversion and deletion events that led to the formation of A08.
Overall, the B, A’ and A genomes have undergone significant chromosomal rearrangement events, including large fragment translocations, inversions, and deletions. Chromosomal fragments enriched with repetitive sequences were transferred to chromosomes A'07 and A07, while gene-enriched fragments were transferred to chromosomes A'08 and A08, ultimately leading to the formation of the current small chromosome A08.
TE variations during genome evolution of Arachis species
Both genome survey and assembly indicate that the A’ genome is the largest among the available genomes of Arachis species that mainly contain the A and B genomes (Supplementary Table 20). Analyses of intact TEs in the T2T genomes of YZ9102 (AABB) and A. hoehnei revealed that the A’ genome had the most intact TEs (38,749), followed by the B (28,444) and A (20,689) genomes (Fig. 5e). Compared with YZ9102 A and B genomes, SVs involving intact TE insertions accounted for 28.85% and 24% of the total SVs, respectively (Fig. 5f). Intact TE insertions in genes or neighboring regions can cause differential gene expression. Among the three genomes, the A’ genome had the most intact TE insertions in genes (476) or neighboring regions, followed by the YZ9102 B (199) and YZ9102 A (106) subgenomes. These genes were enriched in similar GO terms (Supplementary Fig. 32). The genome size of YZ9102 A is 1.15 Gb, with 839 Mb of TE sequences, accounting for 72.90% of the genome. In contrast, the A’ genome contained 1159 Mb of total TE sequences, accounting for 71.98% of its genome (Supplementary Table 21). The TE length of the A’ genome was 320 Mb larger than that of YZ9102 A, accounting for most of the difference in size between the two genomes. Although the total TE length of the A’ genome was similar to that of YZ9102 B, the numbers of each intact TE type differed; the A’ genome contained more LTRs, whereas the YZ9102 B genome had more TIRs (Fig. 5e).
SSON FISH was performed on the chromosomes of Tifrunner and A. hoehnei using four oligonucleotide probes derived from the LTR/Gypsy sequences of the A and B genomes to validate the TE differentiation among the A, B and A’ genomes. Oligo A-4 and Oligo A-7 specifically painted A-genome chromosomes but not B-genome chromosomes, whereas Oligo A-9 and Oligo A-10 painted B-genome chromosomes but not A-genome chromosomes. For the A’ chromosomes, Oligo A-4 and Oligo A-7 produced signals, whereas Oligo A-9 and Oligo A-10 produced almost no signal (Fig. 5g). The distribution frequency of the four probes in the genome of A. hoehnei was similar to that in the A subgenome of YZ9102, further confirming that its A’ genome is closely related to the A genome (Supplementary Table 22 and Fig. 5g).
Interspecies hybridization and oligo GISH confirm A’-A genome proximity
The crossability and pollen fertility of hybrids reveal genome affinity of distant species. The pollen viability of a hybrid (named W1612) between cultivar Yuhua 15 and A. hoehnei was 10.76%, and no pods were produced. Chromosomes of the hybrid were doubled by colchicine treatment, resulting in a synthetic hexaploid, designated Am1612. Analysis of simple sequence repeat (SSR) markers C41 and F115, confirmed by multiplex#1 FISH, revealed that Am1612 had 60 chromosomes, comprising chromosomes from the Ahy and Bhy genomes of Yuhua 15 and the A’ genome of A. hoehnei (Supplementary Figs. 33 and 34). Am1612 had significantly increased pollen viability (66.77%) (Fig. 6a), but considerably lower levels observed in cultivated peanut. The height of the main stem, number of branches, leaf length and width on the main stem, petiole length, number of pods per plant, and seed length in Am1612 were significantly lower than for Yuhua 15 (Fig. 6b, Supplementary Table 23). The low pollen fertility and seed setting suggested chromosomal instability among the A’, Ahy and Bhy genomes. Oligo GISH analysis with green probes specific to A-genome repetitive sequences and red probes specific to B-genome repetitive sequences showed that 40 chromosomes of Am1612 displayed green signals, confirming the genomic composition of Am1612 and further suggested that the A’ genome shares a closer genetic relationship with the A subgenome of cultivated peanut (Fig. 6c).
a Pollen viability of the interspecies triploid hybrid W1612 and hexaploid Am1612. Bars, 200 μm. b Leaves, pods and seeds from Am1612 and its parents. Bars, 2 cm. c GISH using multiplex #A (green) and multiplex #B (red) in Yuhua 15 and Am1612. All oligo GISH experiments were independently repeated three times with similar results. Bar, 10 μm.
Differentially expressed genes of A. hoehnei against web blotch
An evaluation of disease resistance revealed that A. hoehnei exhibited high resistance to web blotch (Supplementary Fig. 35a). Transcriptomic analyses identified differentially expressed genes (DEGs) across all periods post inoculation, comprising 233 upregulated DEGs (upregulated DEGs) and 146 downregulated DEGs (downregulated DEGs) (Supplementary Fig. 35b). KEGG enrichment analysis of the upregulated DEGs revealed pathways related to flavonoid biosynthesis, tropane, piperidine and pyridine alkaloid biosynthesis, plant-pathogen interactions, biosynthesis of secondary, and isoflavonoid biosynthesis. In contrast, the KEGG enrichment results for downregulated DEGs revealed pathways related to other types of O-glycan biosynthesis, plant hormone signal transduction, metabolic pathways, and starch and sucrose metabolism (Supplementary Fig. 35c and d). Among all the identified DEGs, 26 up- and 20 downregulated genes were implicated in SVs when compared to those from the A and B genomes. Among them, three genes related to plant hormone signal transduction, plant–pathogen interactions and MAPK signaling pathways (Supplementary Table 24) likely contribute to disease resistance mechanisms previously described in cultivated peanut30. Additionally, a pathogenesis-related protein (PR10) gene, Ast01G035260.1 (Supplementary Table 24), might play an important role in web blotch resistance, as reported in other crops31,32. In the A’ genome, this gene contained a 399 bp insertion compared with its homologs in the A and B genomes of YZ9102 (Supplementary Fig. 35e and f).
Discussion
Previously, only chromosome banding33, rDNA FISH34, and bacterial artificial chromosome (BAC) FISH35 were employed for peanut chromosome identification. However, the distribution of repetitive sequences on chromosomes varies significantly between species and even among accessions within a single species. Thus, no corresponding standard karyotype has been established to distinguish homoeologous chromosome groups among different Arachis species. This deficiency has hindered integration of chromosomal information across studies and the application of chromosome engineering for the development of alien translocations in peanut as in wheat36.
Single-copy sequences are highly syntenic across chromosomes in closely related crop species s37. The development of CSCOLs has resulted in significantly advanced karyotyping capability and accuracy. In this study, 10 CSCOLs were developed from unique single-copy sequences of A. duranensis and used to establish a standard consensus karyotype of Arachis species. This paper reports the connection of all pseudomolecules and actual chromosomes of A. duranensis, and the Hgs of the A, B, F, K and H genomes were determined by sequential FISH using tandem repeat SSONs and 45S and 5S rDNA as probes. Based on the consensus karyotypes, we clarified the chromosomal characteristics of 10 Hgs in 17 species, which not only confirmed the collinear relationships between homoeologous chromosome pairs of the Arachis genus but also enabled identification of SCVs during Arachis genome evolution. Several assembly gaps were also identified in the tandem repeat-enriched regions of the reference genomes of A. duranensis, A. ipaensis, and Tifrunner. These findings demonstrate the potential application of these consensus karyotypes and cytological probes in both peanut chromosome engineering and genome evolution studies.
Although numerous genomes of cultivated and wild peanut, including A. duranensis20, A. ipaensis20, Tifrunner21, Shitouqi22, Fuhuasheng18, A. cardenasii4, and A. monticola24, have been sequenced, few T2T genome assemblies have been reported. Here, we report a gap-free T2T genome assembly of A. hoehnei, a species bridging the B and A genomes. Unlike other available genomes, probes for 45S rDNA, 5S rDNA, ITRs, and repetitive sequences in the centromeric regions of A. hoehnei produced expected signals corresponding to their positions and copy numbers in the T2T A’ genome, further validating its assembly quality. Based on this T2T genome, the telomere lengths of A. hoehnei ranged from 1.0 to 11.3 kb, which are shorter than those of maize (16–48 kb)38 and rice (5.1–10.8 kb)39,40. Additionally, unlike maize and rice, we observed ITRs in chromosomes 3 A’ and 6 A’ of A. hoehnei, indicating telomere breakage and refusion might have occurred in this species.
The current A. hoehnei accession Zw61 described in this study was previously introduced and used as an E-genome accession A. stenophylla PI 46817829,41,42,43. Our results showed that this accession differs from other species carrying the A, B, F, H and K genomes, but it is more closely related to A-genome species. During the peer review of this manuscript, one reviewer noted that its leaf morphology differs substantially from that of A. stenophylla44. Our distant hybridization results also support that it is not an E-genome accession, because cultivated peanuts rarely hybridize with E-genome species14. To determine its identity, phylogenetic analysis with other accessions of known identity representing different Arachis species is a powerful approach, based on the available molecular markers and genome sequences. First, we extracted the internal transcribed spacer (ITS) sequence of Zw61 and performed clustering analysis with sequences from 102 species45. Second, we identified the target SNPs of Zw61 using 28,586 probe sequences from the 47 K Axiom_Arachis v02 array and conducted clustering analysis with 588 accessions representing different species8. Our results indicate that Zw61 formed a monophyletic cluster with A. hoehnei and was located at the basal position within the A-genome clade (Supplementary Fig. 36 and Supplementary Data 2). Combined with previous cytological data41, we deduced that Zw61 was an accession of A. hoehnei8 rather than A. stenophylla PI 468178. As suggested by one reviewer, it may represent a diverging form of the A genome, and has thus been designated as genome A’, indicating that it is closely related to but not identical to the canonical genome A. Our study also clarifies its evolutionary relationship and position relative to those of other Arachis genomes, which represents an ancient form of the A genome with four unique variations. First, it currently has the largest genome among wild diploid Arachis species. Second, chromosome A'08 is 55.13 Mb larger than the typical A08 (YZ9102) and contains 1039 more genes than A08,. Third, A'08 contains translocations of chromosomes 7/8 but lacks the inversions present in A08. Finally, A. hoehnei has 1,019 specific genes and 47,591 SVs ( > 50 bp) and diverged from A-genome species approximately 1.78 (1.654 – 1.90) Mya.
Specific genes in A. hoehnei were enriched in ether lipid metabolism, glycerophospholipid metabolism, base excision repair, ubiquinone and other terpenoid-quinone biosynthesis, aminoacyl-tRNA biosynthesis, purine metabolism and endocytosis, among other processes. Ubiquinone and other terpenoid-quinone biosynthesis and purine metabolism are well-known antioxidants in plants, with established roles in stress resistance and adaptation40. These genes possibly contribute to the resistance of A. hoehnei to diseases and pests, including late leaf spot, peanut rust, corn earworm, leafhoppers and nematodes46. Our results revealed that A. hoehnei was highly resistant to web blotch. Transcriptome analysis identified 46 candidate genes involving SVs associated with web blotch response, as previously reported30. Additionally, compared with those in the A and B genomes, a large number of genomic variations, including SVs, SNPs, and InDels, were identified. A partially fertile synthetic hexaploid peanut accession Am1612, involving cultivated peanut and A. hoehnei, provides a germplasm for resistance gene mining and transfer beneficial genes to cultivated peanut.
Our results revealed that SVs, including large segmental SCVs and TEs, played important roles in the genome evolution of Arachis species. SV is a key driver of genome evolution and species diversification in both animals and plants28,47,48,49,50. Critical SVs directly promote species divergence into different ecological types49. The Consensus karyotyping analysis in this study revealed numerous SCVs, including translocations and inversions, in chromosomes Hg2, Hg5, Hg6, Hg7, Hg8, Hg9, and Hg10 across Arachis species. Comparative genomics revealed extensive SVs in the A’ genome relative to those in the A and B genomes. Diverse and prominent large segmental SCVs in centromeric repetitive sequences and nucleolus organizer regions (NORs), including ITRs and heterogeneous SSON and rDNA sites, led to distinct karyotypes among the 17 species/accessions. ITRs were previously attributed to telomere–telomere fusions of ancestral chromosomes, microsatellite expansion, and repair of double-strand breaks in germ lines51,52,53. Heterogeneous sites on homologous chromosomes are likely caused by mechanisms driving repetitive sequence evolution, such as unequal chromosomal crossing over54. Among these SVs, rearrangements occurring on chromosomes 7 and 8 represent the most significant variations and presumably had the greatest effects on genome evolution that led to the formation of three genome groups. Group I (pretranslocation type) includes all species with the B, F, K, and H genomes with no translocation involving chromosomes 7 and 8; Group II (posttranslocation but preinversion type for chromosome 8) includes A. hoehnei; and Group III (postinversion type for chromosome 8) includes all species carrying the A genome. The primary factor contributing to the SV from chromosome A'08 to chromosome A08 was attributed to a large-segment deletion in the Inversion 1 region, resulting in loss of 495 genes. These genes that were lost are enriched in pathways such as homologous recombination, nucleotide excision repair, mismatch repair, and DNA replication. The loss of these genes may have restricted recombination of A genome chromosomes with homologs in the A genome and other genomes55. This genetic recombination may have facilitated hybridization between A-genome species and geographically adjacent B-genome species, thereby leading to the formation of allotetraploid peanut species characterized by stable chromosomal inheritance. Moreover, some SVs, such as inversions, translocations, insertions and deletions, have also altered the positions and functions of some genes, potentially further driving evolution of the Arachis genus. These results highlight the important role of SVs in the genomic divergence of Arachis species.
TEs are known to contribute to genome differentiation56, SVs and regulatory functions57. Our study showed that wild peanut species contain a large proportion of repetitive sequences. The genome assembly revealed that most of the repetitive sequences in the A, B and A’ genomes were dominated by TEs. Compared with A genomes, TE amplification was the primary driver of expansion of the A’ genome and contributed to the altered sequence composition and structure of the A, B, and A’ genomes. Additionally, many SVs in the A’ genome resulted from TE insertions and might have affected gene expression, altered gene regulation, or generated novel genes58,59.
Inter-species hybridization is a well-documented factor involved in speciation and plant genome evolution. Our study showed almost no heterogeneous chromosomes in the B, F, K, A’, and H genomes. In contrast, four A-genome species (A. simpsonii, A. herzogii, A. villosa, and A. diogoi) displayed heterogeneity in multiple chromosome pairs and higher overall heterozygosity. Cultivated peanut and most wild species are generally considered self-pollinated, with intraspecific outcrossing rates reported at 2% or lower in most studies10,60,61,62. Four additional plants (A. duranensis PI 468202 and three plants of A. diogoi) were randomly selected and analyzed by sequential FISH to further validate the presence of heterogeneous chromosomes (Supplementary Figs. 37–42). No obvious heterogeneous chromosome was detected in A. duranensis, but heterogeneous chromosomes were present in all three plants of A. diogoi (Supplementary Fig. 43). Resequencing of A. duranensis PI 468202 and A. diogoi revealed that the genomic heterozygosity was 0.53% in A. duranensis (PI 468202) and 1.68% in A. diogoi (Supplementary Table 25). These results suggest that some A-genome species are predominantly self-pollinating, whereas others, such as A. diogoi, are known to have higher rates of outcrossing63. The A genome of A. kuhlmannii shows high genetic variability, possibly because of cross-pollination64. Cross-pollination and gene flow may complicate the definitions of genetic relationships among certain Arachis species65. We also observed significant differences in karyotype among A-genome species, suggesting that outcrossing was an important factor in the evolution of A genomes.
Arachis species include nine sections and 15 genomes, but the differences in the genomic relationship among these genomes remain unclear. Based on the “small chromosome” and rDNA FISH karyotypes, the A genome was further subdivided into different groups15, and non-A genomes were classified into B, F, K, and other genomes16. Our karyotyping data support these classifications. From the perspectives of SCVs on chromosomes 7 and 8, as well as the evolution of the chloroplast genome, the lineages of the B, F, K, and H genomes diverged prior to the differentiation of the A’ genome, with the A genomes emerging last. Among the B, F, K, and H genomes, the H genome diverged earliest. H genome species cannot hybridize with other wild species of section Arachis14, reflecting its more distant relationship. Following divergence of the H genome, the F, K, and B genomes diverged. Phylogenetic analysis based on ITS intron sequences and microsatellite markers confirmed close evolutionary relationships among the B, F, and K genomes56,64.
In the Section Arachis species included in this study, 5S rDNA sites were distributed in Hg6, whereas 45S rDNA sites were in Hg7. Fewer loci detected in Hg1, Hg2, Hg8, and Hg10 may have been inserted or translocated as a result of later genomic SVs. Therefore, we hypothesize that early wild Arachis species had 5S rDNA and 45S rDNA loci only on chromosomes in Hg6 and Hg7. Combining the findings of this study with those of previous research16, only the Ftr genome conforms to these ancestral characteristics. Samoluk et al11. reported that the evolutionary dynamics of repetitive sequences were less dynamic in F-genome species A. trinitensis than in other species of Section Arachis, further supporting the early divergence of the F genome.
A. duranensis, A. cardenasii, and A. villosa were earlier considered candidate donors of the A genome, whereas A. trinitensis, A. batizocoi, and A. ipaensis were proposed as potential B-genome donors66,67,68. More recently, interspecies hybridization, cytology, and genome sequencing suggested that A. ipaensis and A. duranensis are the most likely donor parents of cultivated peanut18,21,35,69. In this study, the consensus karyotypes of A. ipaensis and A. duranensis were most similar to those of the Bhy and Ahy subgenomes. Moreover, both genomic sequencing and resequencing-based phylogenetic analyses supported A. duranensis and A. ipaensis as the donor parents of cultivated peanut. In addition, the chloroplast phylogenetic tree showed that BS and SLH were located on different branches from TIF, A. duranensis accessions (PI 219823 and PI 468202) and A. monticola, suggesting that A. duranensis was the recipient parent and that cultivated peanuts could have multiple origins35,69.
Based on the current findings, we propose a genome evolution model for Arachis species (Fig. 7). The ancestral A’ + F + H genome first diverged into the A’ + F and H genomes, after which the A’ + F genome split into the A’ and F genomes. The F genome further diverged into the B and K genomes, whereas the A’ genome diverged into the A genome. Finally, the Adu, Av, Ahe, Asi, Adi, Bip, Bma, Ftr, Kba, A’, Hpu, and Hda genomes emerged from the A, B, K, F, A’, and H genomes. Multiple hybridization events between A. duranensis genotypes with different chloroplast origins and A. ipaensis and chromosome doubling gave rise to domesticated peanut.
A, A’, B, K, F, and H represent the genomes of Arachis species. Adi, Asi, Av, Adu, Ahe, A’, Bma, Bip, Kba, Ftr, Hpu, Hda, and AhyAhyBhyBhy denote the genomes of A. diogoi, A. simpsonii, A. villosa, A. duranensis, A. herzogii, A. hoehnei, A. magna, A. ipaensis, A. batizocoi, A. trinitensis, A. pusilla, A. dardani, and A. hypogaea, respectively. A’+F+H refers to the ancestral genome of the A’, F, and H genomes, while A’+F refers to the ancestral genome of the A’ and F genomes.
The two H genomes are the smallest, followed by the A genome; the A’ genome is the largest, followed by the B and K genomes (Supplementary Table 25). Older plant lineages generally tend to have smaller genomes, as plant genomes tend to evolve in a “fatter” direction70. SCVs, repetitive sequence variations, and gene family expansion and contraction also contribute to differences in genome size. Due to the current limitations in genome assembly for wild peanut species, we cannot fully explain genome size variations among some wild species or precisely determine the placement of all species at each differentiation node. Moreover, the sequence of chromosome A'08 from the centromere to the end of the short arm shows almost no collinearity with any other chromosome; it contains scattered small patches that are collinear with regions flanking the centromere of chromosome B07. This observation raises questions regarding whether the centromere of chromosome A'08 was derived from other genomes or if other variations occurred. A future super-pangenome analysis of Arachis species11,67,68. It will be necessary to address these issues.
Methods
Plant materials
This study utilized 18 Arachis accessions, including 4 tetraploid peanut cultivars, 1 tetraploid wild species (A. monticola) and 13 diploid Arachis accessions (Table 1), which were all provided by the Henan Academy of Agricultural Sciences.
Interspecies hybridization and chromosome doubling
Interspecies hybridization of Yuhua 15 and A. hoehnei was performed in a Yuanyang experimental field. F1 (W1612) pods and embryos were cultured on 1/2 MS medium supplemented with 3% sucrose. The seedlings underwent chromosome doubling (MS + 0.1 mg L–1 NAA + 0.4 mg L–1 6-BA + 5% sucrose+0.05% colchicine for 12 days) and recovery culture (MS + 0.8 mg L−1 NAA + 3% sucrose), yielding synthetic polyploid Am1612. The pollen viabilities of W1612 and Am1612 were observed.
Bioinformatic pipeline for CSCOL design and labeling
Ten CSCOLs were developed from 12 target regions at the distal end of each of chromosomes Adu01 to Adu10. The methods followed Han et al71. and Du et al29. and were based on the reference genome of A. duranensis20 in PeanutBase (https://peanutbase.org/). Briefly, after excluding repetitive sequences with RepeatMasker, the sequences in the target region were divided into SSONs of 42–48 nt. Each single-copy oligonucleotide was aligned to the reference sequence of A. duranensis using the BLAST tool72 and screened for homologs with >75% similarity. Single-copy SSONs with dTm >10 °C (dTm = Tm – hairpin Tm) were retained. Ultimately, approximately 27,000 single-copy oligonucleotides in each chromosome were selected to construct 10 libraries (Supplementary Table 1). The libraries were then synthesized by MYcroarray (Ann Arbor, MI, USA), amplified, and labeled with biotin-16-dUTP or digoxigenin-11-dUTP73.
Sequential FISH and karyotyping analysis
Chromosomes were prepared according to the protocol reported by Du et al29. Briefly, healthy lateral root tips were excised and treated with 2 mM 8-hydroxyquinoline at 25 °C for 3.5 h. The root tips were then fixed in a 3:1 (v/v) solution of absolute ethanol: glacial acetic acid for 1 h. For chromosome slide preparation, the meristematic tissue of the root tips was placed onto slides, squashed in 45% glacial acetic acid, and then frozen at -80 °C for 12 h.
Clones of wheat 5S and 45S rDNA were labeled with biotin-16-dUTP or digoxigenin-11-dUTP (Roche) by nick translation and detected with a fluorescein-conjugated antibiotin antibody or anti-digoxigenin-rhodamine antibody (Roche). Multiplex #1, which included TAMRA (6-carboxytetramethylrhodamine)-modified oligonucleotides DP-1, DP-3, DP-4, and DP-6 and the FAM (6-carboxyfluorescein)-modified oligonucleotides DP-2, DP-5 and DP-7, was obtained from Du et al29. Multiplex #A and multiplex #B were obtained from Du et al74. All oligonucleotides were synthesized and modified by General Biosystems Company (Chuzhou, Anhui).
FISH was performed on chromosome spreads using CSCOLs, followed by the removal of chromosome signals. Next, GISH was performed using multiplex #A and multiplex #B as probes to identify A and B subgenome chromosomes, respectively74. Subsequent FISH procedures using repetitive multiplex #1 (SSONs) and rDNA probes were then performed on the same slides to develop the karyotypes. A fluorescence microscope (Leica DM6000, Germany) was used to detect signals. Images were optimized for contrast and brightness using Adobe Photoshop.
Investigation, statistics and reproducibility
Five mature plants of Am1612 and its cultivated parent were selected for investigation and analysis of eight phenotypic traits, including height of the main stem, length of the first primary branch, number of branches, number of pods per plant, length and width of the top 3rd and 4th leaves on the main stem, petiole length, and seed length (10 seeds). Statistical analyses of the eight trait values were performed using Excel. Prior to difference analysis, a one-tailed F test (95% confidence interval) was conducted on each data set to verify the homogeneity of variance. Based on the results of the F test, a one-tailed t test (95% confidence interval) was applied to determine differences between Am1612 and its cultivated parent.
For GISH and FISH analyses, three independent biological replicates were established. In each replicate, 3–5 cells were observed to verify the consistency of the signals. For karyotyping analysis, 3–5 cells of each accession were observed and analyzed. Chromosome size was measured in at least three metaphase cells of each species using Image-Pro Plus 6.0. Excel was used to calculate three key karyotypic parameters: arm ratio (long arm length/short arm length), individual chromosome length, and karyotype length. Arm ratio was used to classify the chromosomes of each species as metacentric (m, 1.00–1.70), submetacentric (sm, 1.71–3.00) or subtelocentric (st, 3.01–7.00).
For RNA-seq analyses of infected plants, third leaves of A. hoehnei at the 6–8-leaf stage were sampled at 0 (mock-inoculated control), 1, 3, 5, 7, and 14 days after inoculation with the causative pathogen Phoma arachidicola (strain YY187)30. Three biological replicates were analyzed at each sampling time point. Transcriptome sequencing and subsequent bioinformatics analyses were performed at BGI (Beijing Genomics Institute). RT‒qPCR (with the same RNA) was performed using a PrimeScript™ II Kit (TaKaRa, Dalian) and PowerUp SYBR Master Mix (Applied Biosystems, CA, USA), with ADH3 used as the reference gene (2−ΔΔCt method; 3 biological replicates/4 technical replicates).
Analysis of molecular markers
Codominant SSR markers C41 (F: CACGAGAAGCAGAGGAAGAAGA; R: GCAGCAACAACTCAGGGAAGT) and F115 (F: GAGGGATTCGCTCCACAG; R: CACTCCAATAATTCAAACA) were used to confirm the interspecies hybrid plants. The amplified products were separated by capillary electrophoresis on an Applied Biosystems 3500/3500 XL system. The data were analyzed, and polymorphic markers were screened using Genographer software (https://sourceforge.net/projects/genographer/).
Probe mapping
The sequences of the multiplex #1 (SSON) rDNA probes exhibiting >80% homology were identified in the reference genomes of A. duranensis, A. ipaensis, A. hoehnei, and Tifrunner by BLAST searches (http://mcgb.uestc.edu.cn/b2dsc). Genome positions with more than 50 matches of DP-1, DP-2, DP-3, DP-4, DP-5, DP-6, and DP-7 or more than 1 match of 5S rDNA or 45S rDNA per Mb were mapped.
Genome sequencing, assembly and assessment of A. hoehnei
The A. hoehnei genome was sequenced using the PacBio HiFi, ONT ultralong, Hi-C, and Illumina NovaSeq 6000 systems at BGI. The ONT ultralong and HiFi reads were combined and assembled using Hifiasm (v 0.19.9)75 and Verkko (v 2.2.1)76, with the optimal assembly selected. Contaminants (mitochondrial, chloroplast, and bacterial sequences) were removed by BLAST alignment. The contigs within each chromosome group were ordered and oriented, and the interactions between contigs were transformed into a specified binary file using 3D-DNA77 and Juicer78. Juicebox78 was used to visualize and guide the manual sequencing and orientation of the contigs. First, HiFi sequencing data and quarTeT79 software were used to preliminarily fill the gap region. Second, ONT ultralong sequencing data were assembled using Flye, and the gap region was filled with TGS-GapCloser80 and Minimap2. Finally, NextPolish281 was used for error correction to obtain the final genome assembly.
BUSCO11 was used to evaluate completeness and integrity of the assembled A. hoehnei genomes and the uniformity of the sequencing coverage. Sequencing data were mapped onto the genome using BWA and Minimap2. Consistency quality values (QVs) of the assembled genomes were assessed with Merqury82. LAIs of the assembled genomes were determined using repetitive sequences83. Potential telomere repeat units in the genome were identified using TIDK (https://github.com/tolkit/telomeric-identifier). Potential centromere sequences and their positions were identified using third-generation reads and Centromics (https://github.com/JanaSperschneider/FindTelomeres).
Genome annotation and comparative genomic analysis
De novo prediction of the repetitive sequence database of A. hoehnei was performed using RepeatModeler284, and the database was then used to predict TEs in the genome with RepeatMasker85. Tandem repeats were identified mainly using Microsatellite86 and Tandem Repeat Finder87, with all predicted repetitive sequences subsequently integrated.
For gene coding predictions, the root, stem, leaf, flower, fruit needle, fruit and other tissues of the plant were sampled for transcriptomic sequencing, with three biological replicates included for each sampling group. Three prediction strategies were applied: de novo prediction using Augustus88 and SNAP and homologous species-based prediction using GeMoMa. These predictions were integrated using EVM, modified with PASA, and annotated against databases such as NR, eggNOG, GO, KEGG, TrEMBL, KOG, SWISS-PROT, and Pfam.
Circlize software was used to generate circle plots. Minimap2 was used to align the A. hoehnei genome with previously published genomes, and SyRI identified syntenic regions, structural rearrangements, and sequence variations. Centromere regions were analyzed using StainedGlass. Orthoender v2.4 and the PANTHER V15 database enabled gene family identification and annotation. GO and KEGG enrichment analyses were performed for species-specific gene families, and gene copy numbers were analyzed within each family across species.
MAFFT was used to align each individual single-copy gene sequence, IQ-TREE’s ModelFinder was used to select optimal models, and maximum likelihood (ML) was used to construct an evolutionary tree. The divergence time was calculated using MCMCTREE with fossil calibration times retrieved from the TimeTree website (http://www.timetree.org/). CAFE (v4.2.1) software was used to analyse gene family contraction and expansion. MCScanX was used to obtain genes in collinear blocks, and JCVI was used to draw images of collinearity.
EDTA software was used to identify, annotate, and count complete transposons from the A and B subgenomes of the YZ9102 reference genome and the complete genome of A. hoehnei. Python was employed to statistically analyze genes adjacent to transposons and transposon-associated SVs, followed by GO enrichment analysis of these genes. A resequencing-based phylogenetic tree was constructed using A. hoehnei as the reference genome: BWA and GATK were used to align the resequencing data and generate a SNP-containing VCF file, which was converted to PHYLIP format prior to tree construction with IQ-TREE.
A maximum likelihood (ML) phylogenetic tree was constructed based on the aligned ITS sequences45 using RAxML version 8. BLASTN was employed to align 28,586 probe sequences8 against the Zw61 genome to generate Fastaseq sequences of Zw61. Another phylogenetic tree was constructed using the neighbor-joining (NJ) method, based on the aligned SNP Fastaseq sequences of Zw61 and 588 Arachis species/accessions.
Chloroplast genome assembly and evolution analysis
Total genomic DNA from the 17 species/accessions was extracted using a Plant Genomic DNA Kit (Tiangen Biotech, Beijing) and sequenced on the Illumina HiSeq Xten platform (20× depth). Chloroplast genomes were de novo assembled using the GetOrganelle toolkit89. For Arachis phylogenetic reconstruction and maternal progenitor identification, complete plastomes of the 17 species and 4 outgroups (A. thaliana, C. arietinum, M. truncatula, and G. max) were aligned with MAFFT90, and ML phylogenetic trees were constructed using FigTree91 with the 21 sequences. Jellyfish software was used to analyze the K-mer frequency distribution for heterozygosity, estimate genome size and calculate SNP proportions from Arachis genome sequencing data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data of genome sequencing of Arachis hoehnei (Zw61), genome resequencing data and transcriptome sequencing data were deposited in the National Center for Biotechnology Information under accession PRJNA605106. The telomere-to-telomere (T2T) genome assembly, gene annotations, and chloroplast genomes of A. hoehnei (Zw61) were available at Zenodo [https://doi.org/10.5281/zenodo.18161068]. Additionally, data on chromosomes, agronomic traits data, and sequences of 10 chromosome-specific single copy oligonucleotide libraries (CSCOLs) can be accessed at Zenodo [https://doi.org/10.5281/zenodo.18168257]. Source data are provided with this paper.
References
Stalker, H. T. Utilizing wild species for peanut improvement. Crop Sci. 57, 1102 (2017).
Prine, G. M., French, E. C., Blount, A. R., Williams, M. J. & Quesenberry, K. H. Registration of Arblick and Ecoturf rhizoma peanut germplasms for ornamental or forage use. J. Plant Regist. 4, 145–148 (2010).
Moss, J. P. et al. Registration of ICGV-SM 86715 peanut germplasm. Crop Sci. 38, 572 (1998).
Bertioli, D. J. et al. Legacy genetics of Arachis cardenasii in the peanut crop shows the profound benefits of international seed exchange. Proc. Natl. Acad. Sci. USA. 118, e2104899118 (2021).
Garg, V. et al. Chromosome-length genome assemblies of six legume species provide insights into genome organization, evolution, and agronomic traits for crop improvement. J. Adv. Res. 42, 315–329 (2022).
Valls, J. F. & Simpson, C. E. New species of Arachis (leguminosae) from Brazil, Paraguay and Bolivia. Bonplandia 23, 35–63 (2005).
Simpson, C. E., Krapovickas, A. & Valls, J. History of Arachis including evidence of A. hypogaea L. progenitors. Peanut Sci. 28, 78–80 (2001).
Leal Bertioli, S. C. et al. Relationships of the wild peanut species, section Arachis: A resource for botanical classification, crop improvement, and germplasm management. Am. J. Bot. 111, e16357 (2024).
Santana, S. H. & Valls, J. F. Arachis veigae (Fabaceae), the most dispersed wild species of the genus, and yet taxonomically overlooked. Bonplandia 24, 139–150 (2015).
Krapovickas, A. & Gregory, W. C. Taxonomia Del Genero “Arachis (Leguminosae)”. Bonplandia 1–186 (1994).
Samoluk, S. S. et al. Comparative repeatome analysis reveals new evidence on genome evolution in wild diploid Arachis (Fabaceae) species. Planta 256, 50 (2022).
Koppolu, R., Upadhyaya, H. D., Dwivedi, S. L., Hoisington, D. A. & Varshney, R. K. Genetic relationships among seven sections of genus Arachis studied by using SSR markers. BMC Plant Biol. 10, 15 (2010).
Friend, S. A., Quandt, D., Tallury, S. P., Stalker, H. T. & Hilu, K. W. Species, genomes, and section relationships in the genus Arachis (Fabaceae): a molecular phylogeny. Plant Syst. Evol. 290, 185–199 (2010).
Gregory, M. P. & Gregory, W. C. Exotic germ plasm of Arachis L. interspecific hybrids. J. Hered. 70, 185–193 (1979).
Robledo, G., Lavia, G. I. & Seijo, G. Species relations among wild Arachis species with the A genome as revealed by FISH mapping of rDNA loci and heterochromatin detection. Theor. Appl. Genet. 118, 1295–1307 (2009).
Robledo, G. & Seijo, G. Species relationships among the wild B genome of Arachis species (section Arachis) based on FISH mapping of rDNA loci and heterochromatin detection: a new proposal for genome arrangement. Theor. Appl. Genet. 121, 1033–1046 (2010).
Silvestri, M. C., Ortiz, A. M. & Lavia, G. I. rDNA loci and heterochromatin positions support a distinct genome type for ‘x= 9 species’ of section Arachis (Arachis, Leguminosae). Plant Syst. Evol. 301, 555–562 (2015).
Chen, X. et al. Sequencing of cultivated peanut, Arachis hypogaea, yields insights into genome evolution and oil improvement. Mol. Plant. 12, 920–934 (2019).
Husted, L. Cytological studies on the peanut, Arachis. Cytologia. (Tokyo) 5, 109–117 (1933).
Bertioli, D. J. et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat. Genet. 48, 438 (2016).
Bertioli, D. J. et al. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat. Genet. 51, 877–884 (2019).
Zhuang, W. et al. The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication. Nat. Genet. 51, 865–876 (2019).
Wang, X. et al. A telomere-to-telomere genome assembly of the cultivated peanut. Mol. Plant. 18, 5–8 (2025).
Xue, H. et al. A near complete genome of Arachis monticola, an allotetraploid wild peanut. Plant Biotechnol. J. 22, 2110–2112 (2024).
Zhao, K. et al. Pangenome analysis reveals structural variation associated with seed size and weight traits in peanut. Nat. Genet. 57, 1250–1261 (2025).
Liu, X. et al. Dual-color oligo-FISH can reveal chromosomal variations and evolution in Oryza species. Plant. J. 101, 112–121 (2020).
Braz, G. T. et al. Comparative oligo-FISH mapping: an efficient and powerful methodology to reveal karyotypic and chromosomal evolution. Genetics 208, 513–523 (2018).
Zhang, Y. et al. Telomere-to-telomere Citrullus super-pangenome provides direction for watermelon breeding. Nat. Genet. 56, 1750–1761 (2024).
Du, P. et al. High-resolution chromosome painting with repetitive and single-copy oligonucleotides in Arachis species identifies structural rearrangements and genome differentiation. BMC Plant Biol. 18, 240 (2018).
Wu, X. et al. Cytological and transcriptomic analysis to unveil the mechanism of web blotch resistance in Peanut. BMC Plant Biol 23, 518 (2023).
Guo, J. et al. Revealing the contribution of GbPR10. 5D1 to resistance against Verticillium dahliae and its regulation for structural defense and immune signaling. Plant. Genome 15, e20271 (2022).
Park, C. J. et al. Pathogenesis-related protein 10 isolated from hot pepper functions as a ribonuclease in an antiviral pathway. Plant. J. 37, 186–198 (2004).
Cai, Q., Lu, S. & Chinnappa, C. C. Analysis of karyotypes and Giemsa C-banding patterns in eight species of Arachis. Genome 29, 187–194 (1987).
Seijo, J. G. et al. Physical mapping of the 5S and 18S-25S rRNA genes by FISH as evidence that Arachis duranensis and A. ipaensis are the wild diploid progenitors of A. hypogaea (Leguminosae). Am. J. Bot. 91, 1294–1303 (2004).
Zhang, L., Yang, X., Tian, L., Chen, L. & Yu, W. Identification of peanut (Arachis hypogaea) chromosomes using a fluorescence in situ hybridization system reveals multiple hybridization events during tetraploid peanut formation. New Phytol 211, 1424–1439 (2016).
Bhullar, R. et al. Silencing of a metaphase I-specific gene results in a phenotype similar to that of the Pairing homeologous 1 (Ph1) gene mutations. Proc. Natl. Acad. Sci. USA. 111, 14187–14192 (2014).
Lou, Q. et al. Single-copy gene-based chromosome painting in cucumber and its application for chromosome rearrangement analysis in Cucumis. Plant. J. 78, 169–179 (2014).
Chen, J. et al. A complete telomere-to-telomere assembly of the maize genome. Nat. Genet. 55, 1221–1231 (2023).
Mizuno, H. et al. Sequencing and characterization of telomere and subtelomere regions on rice chromosomes 1S, 2S, 2L, 6L, 7S, 7L and 8. S. Plant. J. 46, 206–217 (2006).
Valenzuela, C. E. et al. Salt stress response triggers activation of the jasmonate signaling pathway leading to inhibition of cell elongation in Arabidopsis primary root. J. Exp. Bot. 67, 4209–4220 (2016).
Du, P. et al. Chromosome painting of telomeric repeats reveals new evidence for genome evolution in peanut. J. Integr. Agr. 15, 2488–2496 (2016).
Du, P. et al. Development and characterization of bacterial wilt-resistant synthetic polyploid peanuts. Crop J. 13, 125–134 (2025).
Du, P. et al. Development of an oligonucleotide dye solution facilitates high throughput and cost-efficient chromosome identification in peanut. Plant. Methods 15, 69 (2019).
Krapovickas, A. G. W. C. Taxonomy of the genus Arachis (Leguminosae). Bonplandia 16, 7–205 (2007).
Inglis, P. W. et al. A slow march along the South American Dry Diagonal: a multi-gene molecular phylogeny and biogeographical history of the peanut genus, Arachis L. Ann. Bot. 136, 795–820 (2025).
Stalker, H. T. et al. The value of diploid peanut relatives for breeding and genomics. Peanut Sci. 40, 70–88 (2013).
One Thousand Plant Transcriptomes Initiative. One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574, 679–685 (2019).
Liu, Y. et al. Pan-genome of wild and cultivated soybeans. Cell 182, 162–176 (2020).
Hager, E. R. et al. A chromosomal inversion contributes to divergence in multiple traits between deer mouse ecotypes. Science 377, 399–405 (2022).
Levy, A. A. & Feldman, M. Evolution and origin of bread wheat. Plant. Cell 34, 2549–2567 (2022).
Lin, K. W. & Yan, J. Endings in the middle: current knowledge of interstitial telomeric sequences. Mutat. Res. Rev. Mutat. Res. 658, 95–110 (2008).
Ruiz-Herrera, A. et al. Telomeric repeats far from the ends: mechanisms of origin and role in evolution. Cytogenet. Genome Res. 122, 219–228 (2008).
Uchida, W. et al. Interstitial telomere-like repeats in the Arabidopsis thaliana genome. Genes Genet. Syst. 77, 63–67 (2002).
He, L. et al. Interstitial telomeric repeats are enriched in the centromeres of chromosomes in Solanum species. Chromosome Res. 21, 5–13 (2013).
Chambers, S. R. et al. The mismatch repair system reduces meiotic homeologous recombination and stimulates recombination-dependent chromosome loss. Mol. Cell. Biol. 16, 6110–6120 (1996).
Mehrotra, S. & Goyal, V. Repetitive sequences in plant nuclear DNA: types, distribution, evolution and function. Genom. Proteom. Bioinf. 12, 164–171 (2014).
Garrido-Ramos, M. A. Satellite DNA in plants: more than just rubbish. Cytogenet. Genome Res. 146, 153–170 (2015).
Kidwell, M. G. & Lisch, D. Transposable elements as sources of variation in animals and plants. Proc. Natl. Acad. Sci. USA. 94, 7704–7711 (1997).
Kashkush, K., Feldman, M. & Levy, A. A. Transcriptional activation of retrotransposons alters the expression of adjacent genes in wheat. Nat. Genet. 33, 102–106 (2003).
Hammons, R. O. & Leuck, D. B. Natural cross-pollination of the peanut, Arachis hypogaea L., in the Presence of Bees and Thrips. Agron. J. 58, 396 (1966).
Coffelt, T. A. Natural crossing of peanut in Virginia. Peanut Sci. 16, 46–48 (1989).
Gibbons, R. W. & Tattersfield, J. R. Out-crossing trials with groundnuts (Arachis hypogaea) L. Rhod. J. Agric. Res. 7, 71–85 (1969).
Oliveira, J. C. D. et al. Inferring mating system parameters in forage peanut, Arachis pintoi, for Brazilian Amazon conditions. Acta Amazon 49, 277–282 (2019).
Moretzsohn, M. C. et al. A study of the relationships of cultivated peanut (Arachis hypogaea) and its most closely related wild species using intron sequences and microsatellite markers. Ann. Bot. 111, 113–126 (2013).
Tian, X. et al. Chloroplast phylogenomic analyses reveal a maternal hybridization event leading to the formation of cultivated peanuts. Front. Plant Sci. 12, 804568 (2021).
Raina, S. N. & Mukai, Y. Genomic in situ hybridization in Arachis (Fabaceae) identifies the diploid wild progenitors of cultivated (A. hypogaea) and related wild (A. monticola) peanut species. Plant. Syst. Evol. 214, 251–262 (1999).
Kochert, G. et al. RFLP variability in peanut (Arachis hypogaea L.) cultivars and wild species. Theor. Appl. Genet. 81, 565–570 (1991).
Singh, A. K. & Smartt, J. The genome donors of the groundnut/peanut (Arachis hypogaea L.) revisited. Genet. Resour. Crop Evol. 45, 113–116 (1998).
Zheng, Z. et al. Chloroplast and whole-genome sequencing shed light on the evolutionary history and phenotypic diversification of peanuts. Nat. Genet. 56, 1975–1984 (2024).
Ma, J. & Bennetzen, J. L. Rapid recent growth and divergence of rice nuclear genomes. Proc. Natl. Acad. Sci. USA. 101, 12404–12410 (2004).
Han, Y. et al. Chromosome-specific painting in Cucumis species using bulked oligonucleotides. Genetics 200, 771–779 (2015).
Kent, W. J. BLAT—The BLAST-Like Alignment Tool. Genome Res. 12, 656–664 (2002).
Murgha, Y. E., Rouillard, J. & Gulari, E. Methods for the preparation of large quantities of complex single-stranded oligonucleotide libraries. PLoS One 9, e94752 (2014).
Du, P. et al. Development of Oligo-GISH kits for efficient detection of chromosomal variants in peanut. Crop J. 11, 238–246 (2023).
Cheng, H. et al. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Rautiainen, M. et al. Telomere-to-telomere assembly of diploid chromosomes with Verkko. Nat. Biotechnol. 41, 1474–1482 (2023).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Lin, Y. et al. quarTeT: a telomere-to-telomere toolkit for gap-free genome assembly and centromeric repeat identification. Hortic. Res. 10, uhad127 (2023).
Xu, M. et al. TGS-GapCloser: a fast and accurate gap closer for large genomes with low coverage of error-prone long reads. Gigascience 9, giaa94 (2020).
Hu, J. et al. NextPolish2: a repeat-aware polishing tool for genomes assembled using HiFi long reads. Genom. Proteom. Bioinf. 22, qzad009 (2024).
Rhie, A. et al. Reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 1–27 (2020).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic. Acids. Res. 46, e126 (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Tarailo Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Current Protocols in Bioinformatics 25, 4–10 (2009).
Beier, S. et al. MISA-web: a web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic. Acids. Res. 27, 573–580 (1999).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Jin, J. et al. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 1–31 (2020).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Acknowledgements
We thank Yonghua Han (Jiangsu Normal University, Xuzhou, China) for help in designing and labeling the oligonucleotide libraries. We thank Robert McIntosh, University of Sydney, Australia, for his kind review, suggestions, and language editing. We are also grateful to Dawn M. Schmidt (www.editbydawn.com) for English editing. This project is supported by Henan Province Science and Technology R&D Joint Fund (232301420025 to P.D.), National Natural Science Foundation of China (32272153 to P.D. and 32501918 to L.F.), the Key Research Project of the Shennong Laboratory (SN01-2022-03 to X.Z.), China Agriculture Research System (CARS-13 to X.Z.), Henan Provincial Agriculture Research System (S2012-5 to W.D.), and innovation team of peanut breeding (2024TD01 to X.Z.). The funding agencies played no role in the design of the study and collection, analysis, and interpretation of data or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
X.Z., Z.Q. and P.D. designed and supervised the project. P.D., L.F., G.C., Q.W., C.L., Zhiyuan Z., and L.M. performed the experiments. S.H., X.W., Z.S., Zhiyuan Z., X.X., H.L., L.Q., J.X. and Zhongxin Z. analyzed the data. P.D., G.C. and L.F. wrote the manuscript. X.Z., Z.Q., W.D., and Zheng Z. provided critical guidance for the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks David Bertioli, Chuanzhi Zhao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Du, P., Fu, L., Chen, G. et al. Origin of small chromosome A08 and genome evolution of Arachis species. Nat Commun 17, 2029 (2026). https://doi.org/10.1038/s41467-026-68884-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68884-5