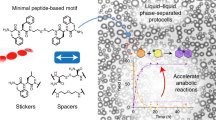
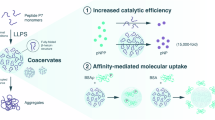
Abstract
The mechanisms governing pH regulation in membrane-less organelles (MLOs) are essentially different from those relying on membrane proteins, yet it remains poorly understood due to the difficulty in directly controlling the conditions across the MLOs interface. Here, we develop a coacervate-based, in vitro model to investigate how liquid-liquid phase separation (LLPS) could contribute to pH regulation in MLOs. We construct peptide-based coacervate droplets using microfluidics and find that charged polymers within the coacervates help create uneven H+/OH- distributions, resulting in pH-regionalized microenvironments similar to the nucleolus. More interestingly, such a pH difference weakens or even disappears following the destruction of LLPS, a phenomenon observed in both nucleolus and coacervate droplets. Based on these findings, we demonstrate the ability to finely tune the local pH of the coacervate droplets over a wide range by incorporating enzymes, which can drive and control cascade reactions, and perform basic molecular biology operations such as polymerase chain reaction (PCR) and in vitro transcription and translation reaction (IVTT). This study highlights the role of LLPS in pH modulation within MLOs and provides insights into the potential of coacervates as protocells for broader applications.
Similar content being viewed by others
Introduction
Cellular functions rely on numerous enzyme-catalyzed reactions that occur spatially and temporally correctly under physiological conditions1,2,3. The maintenance of enzyme activity is essential for metabolism, energy production, and the synthesis of various substances1,3. A key factor that affects enzyme activity is pH, as each enzyme functions optimally within a specific pH range. Deviations from this range can impair the structure and function of enzymes, which can even lead to diseases or disorders. It has been well established that various membrane exchangers and transporters play significant roles in regulating intracellular pH4. In contrast, membrane-less organelles (MLOs), which host diverse biochemical reactions and play essential roles in cellular regulation, lack such membrane-based control mechanisms. Recent studies have revealed that MLOs can exhibit local pH variations, which may influence the kinetics and selectivity of enzymatic reactions5,6. However, given the complexity of living cells, it is challenging to directly investigate the mechanisms of pH regulation in vivo by controlling conditions across the MLOs interface7.
In this study, we found that a natural pH difference exists between the nucleolus and nucleoplasm in several tested cell lines, which can be modulated by drug treatments to alter their structure and properties (Fig. 1a). Since coacervates are liquid-liquid phase separation (LLPS) systems driven by noncovalent interactions, and they share similar properties to MLOs such as molecular organization and compartmentalization7,8,9,10,11,12,13, we then present peptide-based coacervate droplets as protocells to mimic the pH difference found in the nucleus. We use microfluidics to prepare these coacervate droplets to ensure their uniformity in size, thus allowing for the generation of a similar pH difference across the LLPS interface under precisely controlled conditions14,15,16. Through molecular dynamics simulations and quantitative experiments, we conclude that the dense phase of coacervates, rich in charged polymers, tends to attract a higher concentration of H+/OH- ions than the dilute phase. This results in an uneven apparent pH distribution between the interior and exterior of the coacervates. More importantly, such a pH difference weakens or even disappears following the destruction of LLPS, a phenomenon observed in both nucleolus and coacervate droplets. Based on these findings, we demonstrate that the pH-regionalized environment of the coacervate droplets can be customized by selectively accommodating enzymatic reactions (as shown in Fig. 1b), and such feature can be further employed to control cascade reactions and perform polymerase chain reaction (PCR) and in vitro transcription and translation reaction (IVTT), thereby recapitulating various cellular activities (as shown in Fig. 1c)17. Our findings suggest that coacervate-based protocells offer a valuable platform to explore how physicochemical compartmentalization influences biochemical reactions.
a A schematic illustrating the pH difference between the nucleolus and nucleoplasm in a living cell, highlighting changes in the nuclear pH microenvironment induced by drug treatments. b A schematic illustrating that, similar to MLOs, coacervate-based protocells are capable of establishing a pH difference. c The pH-regionalized protocells can be harnessed to finely regulate the efficiency of subsequent biochemical enzymatic reactions.
Results
Identification of the pH difference between the nucleolus and the nucleoplasm
To elucidate whether there is a pH difference across the MLOs interface in living cells, we chose the nucleolus as an example because it is the largest membrane-less condensate and is easily observable18. We assess the pH of nucleoli in different cells using SNARF-4F, a cell-permeable indicator that shows notable pH-dependent dual-emissions at 580 and 640 nm. By measuring the fluorescence intensity ratio at these wavelengths, we can determine the subcellular pH effectively. The spectral shifts of SNARF-4F based on pH changes are described by Eq. (1)19, where Kₐ denotes the acid dissociation constant of the probe, R represents the fluorescence intensity ratio at two wavelengths λ1 (580 nm) and λ2 (640 nm) (Em580/Em640), and RA and RB represent the limiting values of R corresponding to the fully protonated and fully deprotonated states, respectively.
Using Eq. (1), we generate the standard curve for SNARF-4F (Supplementary Fig. 1) after adjusting intracellular pH with calibration buffers and obtain the functional expression given by Eq. (2) (detailed discussion can be found in the Supplementary Note).
With this, we study the pH levels in the nucleolus and nucleoplasm under various physiological conditions, as shown in Fig. 2a. Based on the results, we find that the nucleolus is slightly more acidic than the surrounding nucleoplasm in various cell types, including 3T3, OC1, HELA, and AML12 Fig. 2b–f and 2k–n20.
a A schematic illustrating the relationship between nucleolar pH, POL1 (RNA polymerase I) activity, DNA replication, and the phase separation status of the nucleolus. b Representative confocal fluorescence images of different cells after treatment with either DMSO or CX-5461, incubated with SNARF-4F. c-j The 580/640 nm emission signal profile for the nucleolus and nucleoplasm in 3T3, OC1, HELA, and AML12 cells treated with (c–f) DMSO or (g–j) CX-5461. The white lines in (b) denote the region where the data points are taken. The light blue area in c-j highlights the location of the nucleolus. Violin plots showing nucleolar pH quantification in (k) 3T3, (l) OC1, (m) HELA, and (n) AML12 cells treated with DMSO or CX-5461. Dots represent the average pH values of both the nucleolus and the nucleoplasm. n = 30 independent biological replicates. Data are presented as mean ± standard deviation (SD). Statistical significance is determined using a two-sided Student’s t-test. o Confocal fluorescence images of HELA cells stained with SNARF-4F after treatment with different concentrations of Act D. p Profile of the 580/640 nm emission signal for the nucleus in HELA cells treated with different concentrations of Act D. The white lines in (o) denote the region where the data points are taken.
We next examine whether the pH difference between the nucleolus and nucleoplasm is linked to the nucleolar functions and the LLPS status. To this end, we used CX-5461, a well-known inhibitor of RNA polymerase I, to suppress rRNA synthesis in the nucleolus. Previous studies have also shown that CX-5461 can disrupt the native organization of the nucleolar structure20,21. We find that inhibiting rRNA synthesis with CX-5461 deacidifies both the nucleolus and nucleoplasm, indicating a close relationship between nucleolar transcriptional activity and its pH (Fig. 2a, 2b, and 2g–2n). Additionally, the pH difference between the nucleolus and nucleoplasm is smaller in CX-5461-treated cells than in untreated cells regardless of the cell type (Supplementary Fig. 2). Moreover, we treated HeLa cells with different concentrations of actinomycin D (Act D), an inhibitor of RNA and protein synthesis, and found that low concentrations of Act D can also induce nucleolar deacidification (Fig. 2o and 2p). Furthermore, after nucleolar disassembly in HeLa cells treated with high concentrations of Act D, no detectable pH difference was observed (Fig. 2o and 2p). Taken together, these observations clarify that a pH difference exists across the nucleolus interface and suggest that this difference is closely linked to the LLPS status and biological function of the nucleolus. These experiments imply that the transcriptional activity and structural integrity of the nucleolus might both contribute to maintaining this pH difference. However, as transcriptional inhibition and structural disruption may take place simultaneously under drug treatment, the available data are not sufficient to distinguish which of these processes predominantly drives the observed pH changes.
Peptide coacervate droplets recapitulating the pH difference in MLOs
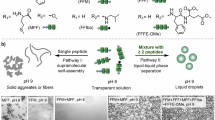
For simplified modeling of MLOs in vitro, we utilize a coacervate composed of decapeptides arginine (R10) and aspartic acid (D10). These amino acids are representative sequences of phase-separating proteins found in cells and can form coacervates through ion-pairing interactions between cationic R10 and anionic D10 (Fig. 3a)22. We generate R10/D10 coacervate microdroplets using the direct mixing method. A state diagram is plotted to indicate the optimal concentration ranges of each component, within which coacervate droplets can be readily formed, albeit with polydispersity (Fig. 3b and Supplementary Fig. 3). Based on the phase diagram, we set 10 mM R10 and 10 mM D10 as the standard experimental conditions (the formed coacervate is denoted as 10 mM R10/D10 coacervate), unless otherwise specified. Additionally, to assess the stability of these coacervates under different pH conditions, we observed them using bright-field microscopy (Supplementary Fig. 4) and measured their turbidity (Fig. 3c) with a UV-Vis spectrophotometer across a pH range of 6.0–9.0. The results suggest that the coacervates remain stable in pH environments close to physiological conditions, making them suitable for modeling the environments of MLOs within cells.
a Schematic illustration showing the opposite effects of coacervate and salt concentrations on the H⁺ or OH⁻ concentration difference between phases. b Phase diagram of the R10/D10 mixing solution as a function of the concentrations of each component. c Turbidity of 10 mM R10/D10 coacervate solution at different pH values. n = 3 independent biological replicates. Data are presented as mean ± SD. d pH changes in the condensate phase and dilute phase upon the addition of different amounts of H+ or OH-. n = 30 independent biological replicates. Data are presented as mean ± SD. Statistical significance is determined using a paired two-sided Student’s t-test. e State diagram showing the effects of salt and coacervate concentrations on the pH difference between phases after H⁺ or OH⁻ addition. f Turbidity of R10/D10 coacervate solutions at different concentrations. n = 3 independent biological replicates. Data are presented as mean ± SD. g pH difference between the condensate phase and the dilute phase of coacervate upon mixing 2 μmol H+ or OH- with 2 mL R10/D10 coacervate solution, at various coacervate concentrations. n = 9 independent biological replicates. Data are presented as mean ± SD. h Turbidity of the R10/D10 coacervate solution at different salt concentrations. n = 3 independent biological replicates. Data are presented as mean ± SD. i pH difference between the condensate phase and the dilute phase of coacervate after mixing 2 μmol H+ or OH- with 2 mL R10/D10 coacervate solution, at different salt concentrations. n = 9 independent biological replicates. Data are presented as mean ± SD. j (Upper) Snapshots of the state of different pairs of R10 and D10 after 5 μs of CG molecular dynamic simulations. (Lower) Snapshots of 300 pairs of R10 and D10 after 5 μs of simulations under different NaCl concentrations. Red color represents R10, while blue color represents D10. Water molecules are not shown. k Simulated RDF of peptide-H+ at varying coacervate concentrations. l Simulated contact number between peptides and ions at various coacervate concentrations. n = 10 independent biological replicates. Data are presented as mean ± SD. Statistical significance is determined using a two-sided Student’s t-test. m Simulated RDF of peptide-H+ at different salt concentrations. n Simulated contact number between peptides and ions at different salt concentrations. n = 10 independent biological replicates. Data are presented as mean ± SD. Statistical significance is determined using a two-sided Student’s t-test.
The condensate and dilute phases of freshly prepared coacervates exhibited comparable pH values of approximately 8.0 (as shown in Fig. 3d, p > 0.05). Then, to investigate whether a pH difference can be established across the coacervate interface in response to environmental variables, we introduce different concentrations of H⁺ or OH⁻ into the R10/D10 coacervate system through the addition of HCl and NaOH. A micro-pH probe is used to separately measure the pH of the condensed and diluted phases obtained after centrifugation (Supplementary Fig. 5). We observe the establishment of a notable pH difference, with the condensate phase responding to pH changes: it becomes more acidic upon H⁺ addition and more alkaline upon OH⁻ addition, compared with the dilute phase (Fig. 3d and Supplementary Fig. 6). In addition, we utilized the SNARF-4F dye to cross-compare the pH change trends of the condensate and dilute phases, further validating the same observations (Supplementary Fig. 7).
We then investigate whether varying the coacervate and salt concentrations affects the pH response behavior of the coacervates. The results indicate that decreasing the coacervate concentration reduces the tolerance of the system to added salts, making the pH difference (defined as the pH of the condensed phase minus that of the dilute phase) detectable within a narrower range of salt concentrations (Fig. 3e). We prepare coacervate solutions with different concentrations and measure their turbidity, as shown in Fig. 3f. The turbidity of the coacervate solution decreases as the concentration of R10 and D10 is reduced. Then, we measure the pH difference between the condensate phase and the dilute phase of coacervate after adding 1 μM HCl or NaOH to the coacervate system. We find that the pH difference between the two phases is larger at higher coacervate concentrations (Fig. 3g, Supplementary Fig. 8a and 8b). Additionally, adding NaCl into the 10 mM R10/D10 coacervate solution with increasing concentrations leads to decreased turbidity, suggesting that high salt concentrations can lead to disassembly of the coacervate due to the screening effect on the peptide charges (Fig. 3h). Accordingly, a smaller pH difference is found between the two phases (Fig. 3i, Supplementary Fig. 8c and 8d). These results suggest that the condensate and dilute phases of the coacervate can respond differently to external acid or base, and this effect becomes more pronounced when the stability of the coacervate is higher. To further validate the above findings, we prepared coacervates in Tris-HCl buffer and added varying concentrations of acid or base. We then measured the pH of both the condensate and dilute phases. The results showed that, compared with coacervates prepared in water at pH 8.0, those prepared in pH 8.0 Tris-HCl buffer exhibited a similar trend in pH difference between the two phases (Supplementary Fig. 9).
We reason that this phenomenon might be attributed to the difference in H+ or OH- distribution in the two phases of the coacervates, as illustrated in Fig. 3a. To support our hypothesis, we conduct molecular dynamics simulations using a coarse-grain (CG) model of the R10/D10 coacervate in 20 × 20 × 20 nm³ simulation boxes containing equal amounts of R10 and D10 molecules23,24. Meanwhile, 100 H₃O⁺ (referred to as “H⁺” for simplicity below) or OH⁻ are introduced into the simulation box, accompanied by Na⁺ or Cl⁻ to maintain electrostatic balance. We first investigate the effect of coacervate concentration on H⁺ or OH⁻ distribution. Starting with a random distribution of the decapeptides, we perform simulations with 100, 200, and 300 pairs of peptides. The energy curve of the system indicates that it has reached a relatively stable state at 5 μs (Supplementary Fig. 10a and Supplementary Fig. 11). Figure 3j shows snapshots of the simulation box at 5 μs, illustrating the status of coacervation of the system at increasing concentrations of R10 and D10. Supplementary Fig. 12 shows the density profiles of the decapeptides, water, H+, and OH-. We find that both H⁺ and OH⁻ are enriched in the condensate phase (Supplementary Fig. 13). The peptide–ion (H⁺ or OH⁻) radial distribution functions (RDFs) at different coacervate concentrations are shown in Fig. 3k and Supplementary Fig. 14. The pronounced first peaks around 0.5 nm indicate direct contact between the peptide and ions. The contact numbers between H⁺ and OH⁻ and peptides are also shown in Fig. 3l. It is evident that as the coacervate concentration increases, the contact number between the peptide and H⁺ or OH⁻ also rises. To further analyze the movement of H⁺ or OH⁻ within the coacervate solution, we assess the diffusive properties by calculating the mean square displacement (MSD) of H⁺ and OH⁻ (Supplementary Fig. 15a and 15b). We find that the diffusion coefficients of H⁺ and OH⁻ decrease significantly with increasing coacervate concentration (Supplementary Fig. 15c and 15d). It indicates reduced mobility of ions within the coacervate, likely due to stronger molecular interactions or limited space in the coacervate network. Notably, in the coacervate solution, the diffusion coefficient of H⁺ decreases more significantly than that of OH⁻ when compared to their respective values in water. This finding aligns with the RDF results, which reveal greater recruitment of H⁺ to the peptide condensates and a higher contact number of H⁺ with peptides relative to OH⁻. Together, these results suggest that H⁺ may have an even more preferential partitioning affinity in the R10/D10 coacervate system over OH⁻.
Similarly, to analyze the influence of salt concentration on the pH compartmentalization behavior of the D10/R10 coacervates, we set up identical simulation boxes with 300 pairs of randomly distributed R10 and D10 and perform simulations with salt concentrations ranging from 0 to 1.27 M (Supplementary Fig. 10b). Among the snapshots at 5 μs with varying NaCl concentrations, coacervation behavior occurs only at lower salt concentrations (0 M and 0.26 M) (Fig. 3j and Supplementary Fig. 16). The peptide–ion (H⁺ or OH⁻) RDFs with different salt concentrations are presented in Fig. 3m and Supplementary Fig. 17. We also find that as the salt concentration increases, the contact numbers between the peptides and H⁺ or OH⁻ are correspondingly decreased (Fig. 3n). Moreover, with increasing salt concentration, the diffusion coefficients of H⁺ and OH⁻ are enhanced (Supplementary Fig. 18). As salt concentration increases, the electrostatic interactions between charged peptides are screened, weakening the coacervate formation and reducing pH compartmentalization. This suggests that Na⁺ and Cl⁻ disrupt the attractive forces between oppositely charged peptides, allowing H⁺ and OH⁻ ions to distribute more evenly between the condensate and dilute phases. In summary, coacervates can dynamically respond to external environments and create regionalized pH difference.
Construction of pH-regionalized protocell protocells
The pH regionalization feature of the coacervate droplets makes them promising candidates as protocells by integrating pH-dependent biological reactions. Prior to this, we first examine two key points: whether enzymes can be accommodated within the coacervates and whether their activity can be sustained. To this end, we choose two common enzymes, glucose oxidase (GOx) and urease, for encapsulation in the R10/D10 coacervates. Similar to many biochemical reactions within cells, these two enzymes generate acidic or alkaline byproducts in the presence of their substrates. One of the key features of coacervates, and a significant factor in their potential role as protocells, is their ability to capture a wide range of guest molecules and concentrate them at high levels to facilitate reactions. Because the uniformity of the coacervate droplets might influence the reaction rate, we employ our previously reported microfluidic technology25 to prepare uniform R10/D10 coacervate droplets for subsequent experiments (Fig. 4a). Then, we assess the ability of the R10/D10, along with other types of coacervate droplets, poly (dimethyl diallyl ammonium chloride) (PDDA)/poly (sodium acrylate) (PAA) and DEAE-dextran/dsDNA to concentrate enzymes using fluorescence microscopy. As shown in Fig. 4b and Supplementary Fig. 19, both FITC and RBITC display a uniform distribution within three types of homogeneous coacervate droplets, indicating that both enzymes have been successfully encapsulated within the droplets. Next, we examined the turbidity of solutions after mixing coacervates with varying concentrations of enzymes to assess whether the enzymes affect coacervate stability. As shown in Supplementary Fig. 20, the results indicated that neither GOx nor urease had a significant impact on the stability of the coacervates.
a A schematic illustrating the generation of uniform R10/D10 coacervate microdroplets containing FITC-GOx via microfluidics, where the encapsulation of FITC-GOx is facilitated by interactions between the R10 (positively charged) and FITC-GOx (negatively charged). b Fluorescence microscopy images of FITC-GOx and RBITC-urease co-encapsulated in microfluidic-generated R10/D10 coacervate microdroplets. c MST binding curves of FITC-GOx with R10, DEAE-dextran, and PDDA. n = 3 independent biological replicates. Data are presented as mean ± SD. d MST binding curves of FITC-GOx with D10, dsDNA, and PAA. n = 3 independent biological replicates. Data are presented as mean ± SD. e Snapshots of R10/D10 coacervate and GOx system after 5 μs of CG molecular dynamics simulations. The light color represents the components of the coacervate, while the purple color represents GOx, and water molecules are not shown. f Snapshot of the final configuration of the R10 and GOx solution after 5 μs of CG molecular dynamics simulations. Water molecules are not shown. g R10-GOx RDF and PMF obtained from the simulation. h Snapshots of R10/D10 coacervate and urease system after 5 μs of CG molecular dynamics simulations. The light color represents the components of the coacervate, while the purple color represents urease. Water molecules are not shown. i Snapshot of the final configuration of the R10 and urease solution after 5 μs of CG molecular dynamics simulations. Water molecules are not shown. j R10-urease RDF and PMF obtained from the simulation. k Snapshot of the final configuration of the D10 and GOx system after 5 μs of CG molecular dynamics simulations. Water molecules are omitted for clarity. l D10-GOx RDF obtained from the simulation. m Snapshot of the final configuration of the D10 and urease system after 5 μs of CG molecular dynamics simulations. Water molecules are omitted for clarity. n D10-urease RDF obtained from the simulation.
To determine the partition coefficients of FITC-GOx and RBITC-urease, the R10/D10 coacervates were incubated with each fluorescently labeled enzyme, and the fluorescence intensities in the condensate and dilute phases were subsequently quantified (Supplementary Fig. 21). In addition, with the condensate volume fraction already determined (Supplementary Fig. 5b), these parameters enabled us to estimate the percentage of the total enzyme recruited into the condensate phase, yielding PGOx = 80.45% and Purease = 73.14%. To further elucidate the preferential partitioning of enzymes within the coacervates, we employ microscale thermophoresis (MST) to assess the interactions between GOx and the components of the coacervates, i.e., D10 and R10, respectively. As shown in Fig. 4c and 4d, R10 interacts directly with FITC-GOx at Kd = 737.07 ± 104.96 nM, whereas no detectable binding between D10 and FITC-GOx is observed. We hypothesize that the partitioning affinity of FITC-GOx into the D10/R10 coacervate is driven by the interactions between the positively charged R10 and the negatively charged FITC-GOx. Therefore, we test other coacervate systems, including DEAE-dextran/dsDNA and PDDA/PAA. Based on the results from MST, it is evident that similar to R10, there are interactions between GOx and the positively charged polyelectrolytes DEAE-dextran (Kd = 1377.90 ± 197.96 nM) and PDDA (Kd = 1938.61 ± 255.71 nM). However, no interactions were observed between GOx and the negatively charged dsDNA and PAA. These results suggest that the interaction between the charged polymer and the enzymes plays a crucial role in the affinity partitioning of enzymes in coacervates.
We further conduct molecular dynamics simulations of R10/D10 coacervate systems containing GOx or urease. Starting from random placements in the solution, GOx and urease gradually partition into the condensate phase and tend to move to the inside (Figs. 4e, 4h, Supplementary Fig. 22a, and 22c). Additionally, we separately performed molecular dynamics simulations for solutions containing R10 and GOx, as well as R10 and urease. Snapshot of the final configuration for the solution containing R10 and GOx is shown in Fig. 4f and Supplementary Fig. 22b, while that for the solution containing R10 and urease is presented in Fig. 4i and Supplementary Fig. 22d. Over time, R10 molecules interact with GOx or urease molecules, as evidenced by the peptide-protein RDF and the potential of mean force (PMF) obtained from the simulations (Figs. 4g and 4j). In both R10-GOx and R10-urease systems, the peak in the RDF around 0.5 nm suggests the presence of interaction zones between the peptides and the enzyme. The PMF provides insight into the energetic landscape of these interactions, implying that R10 contacts with both GOx and urease. For comparison, the interactions between D10 and the enzymes were also analyzed using molecular dynamics simulations. The results showed that, unlike the R10 peptide, D10 did not exhibit significant interactions with either GOx or urease (Fig. 4k–n). These observations align with the MST measurements.
Next, we use R10/D10 coacervate droplets as protocells accommodating enzyme reactions, as shown in Fig. 5a. Glucose or urea is added to the system, which acts as a substrate for GOx and urease, respectively. Glucose can be oxidized to produce gluconic acid, thus creating a continuous H+-supplying environment. Similarly, urease catalyzes the hydrolysis of urea to produce NH3, a basic substance in the solution, thereby establishing a continuous OH- supply. We measured the reaction rates of GOx and urease in both coacervate and bulk solutions. The reaction rate of GOx as a function of glucose concentration follows typical Michaelis–Menten kinetics in both bulk solution (pH 8.0) and R10/D10 coacervate solutions (Fig. 5b, 5c, Supplementary Figs. 23, and 24). Similarly, the reaction rate of urease as a function of urea concentration also follows typical Michaelis–Menten kinetics in both bulk solution and R10/D10 coacervate solutions (Fig. 5d, 5e, Supplementary Figs. 25, and 26). The Vmax and Km values, representing the maximum reaction rate and substrate binding affinity, respectively, were obtained by fitting the Michaelis–Menten plots (Fig. 5f and Supplementary Fig. 27). In the absence of additional H⁺ or OH⁻ ions, the lower Vmax values observed for both enzymes in coacervates, compared with those in solution, could possibly arise from restricted enzyme mobility within the dense coacervate matrix, which might limit catalytic turnover. The decrease in Km indicates an enhanced apparent affinity for the substrate, which may arise from the microenvironmental effects inside the coacervates. Moreover, as shown in Fig. 5f, upon the addition of extra H⁺ ions, the Vmax values of GOx and urease (with optimal pH values of 5.5 and 7.4, respectively) in the condensate phase became more comparable to those in solution, likely because the preferential partitioning of ions toward the condensate phase resulted in a locally elevated concentration of H⁺26,27. In contrast, as OH⁻ ions were added, the difference in Vmax between the coacervate and solution phases increased accordingly.
a Schematic illustrating the R10/D10 coacervate droplets loaded with GOx or urease as protocells to establish a pH difference. b Michaelis−Menten plots showing GOx activity in bulk solution with different concentrations of H+/OH-. n = 5 independent biological replicates. Data are presented as mean ± SD. c Michaelis−Menten plots showing GOx activity in R10/D10 coacervates with different concentrations of H+/OH-. n = 5 independent biological replicates. Data are presented as mean ± SD. d Michaelis−Menten plots showing urease activity in bulk solution with different concentrations of H+/OH-. n = 5 independent biological replicates. Data are presented as mean ± SD. e Michaelis−Menten plots showing urease activity in R10/D10 coacervates with different concentrations of H+/OH-. n = 5 independent biological replicates. Data are presented as mean ± SD. f Vmax of GOx and urease in solution and within R10/D10 coacervates upon the addition of different concentrations of H+ or OH-. n = 5 independent biological replicates. Data are presented as mean ± SD. pH values of both the condensate phase and the dilute phase of g R10/D10 coacervate, h DEAE-dextran/dsDNA coacervate, and i PDDA/PAA coacervate, each containing 50 nM GOx and urease, measured before (Blank group) and after adding 5 mM substrates (Glucose group and Urea group). n = 30 independent biological replicates. Data are presented as mean ± SD. j pH difference between the condensate phase and the dilute phase of GOx and urease-loaded 10 mM coacervate solution at varying concentrations of glucose or urea. n = 30 independent biological replicates. Data are presented as mean ± SD. k pH difference between the condensate phase and the dilute phase of GOx and urease-loaded 10 mM coacervate solution at varying coacervate concentrations. n = 30 independent biological replicates. Data are presented as mean ± SD. l pH difference between the condensate phase and the dilute phase of 10 mM R10/D10 coacervate solution containing 50 nM GOx and urease under 5 mM substrate at varying salt concentrations. n = 30 independent biological replicates. Data are presented as mean ± SD.
Adding 5 mM glucose to the GOx-containing coacervate solution causes both the condensate and dilute phases to become more acidic (Fig. 5g). Conversely, adding 5 mM urea to the urease-containing coacervate solution causes both phases to become more alkaline (Fig. 5g). The pH changes triggered by adding glucose or urea are more prominent in the condensed phase than in the dilute phase. This is likely due to the partition of the enzymes and enrichment of H+/OH- in the condensate phase. To figure out whether this effect also applies to other coacervate systems, we test it in DEAE-dextran/dsDNA coacervate (Fig. 5h) and PDDA/PAA coacervate (Fig. 5i). In both cases, we observe similar pH regionalization behavior, confirming that the observed phenomenon is not limited to a single coacervate system.
We then measure the pH difference between the condensate and dilute phases of the coacervate systems after reacting with different concentrations of glucose and urea (Fig. 5j). We observed that, after reacting with glucose or urea, the enzyme-containing coacervates exhibited a pH difference between the condensate and the surrounding dilute phase, which is consistent with the results obtained from the direct addition of H+ or OH-. Additionally, the pH difference between the condensate and dilute phases of the coacervate is also affected by the concentration of the coacervate and salt (Fig. 5k, 5l, and Supplementary Fig. 28). Similar to the effect observed with direct H+ or OH- addition, when the stability of the coacervate droplets is reduced by decreasing the scaffold molecular concentration or increasing the salt concentration, the pH difference between the inside and outside of the coacervate droplets correspondingly decreases.
Controlling cascade reactions in the coacervate protocells
Since we have demonstrated that the D10/R10 coacervates can be used to accommodate biological enzymes and establish pH difference across the interface using enzymatic reactions, we aim to further exploit these properties to develop protocells capable of performing cascading enzymatic reactions (Fig. 6a). First, we examined the stability of R10/D10 coacervates upon mixing with various reagents using bright-field microscopy (as shown in Fig. 6b). Then, we determine the adjustable range of pH of the GOx and urease-loaded R10/D10 coacervate by reacting with the substrate (glucose or urea). We find that the pH of the condensate phase can be regulated within the range of 7-9 by adjusting the substrate type and concentration (Fig. 6c). The feasibility of this approach is also demonstrated using GOx and urease-loaded DEAE-dextran/dsDNA coacervate and PDDA/PAA coacervate (Supplementary Fig. 29). To exclude the potential influence of the added components on the microenvironmental pH of the condensate phase, we measured the pH of the condensate phase of R10/D10 coacervates without enzymes after mixing with different components, and found that the addition of the substrate alone did not significantly affect the internal pH of the coacervates (Supplementary Fig. 30). These results indicate that GOx- and urease-loaded coacervates can regulate the microenvironment’s pH within the normal physiological range, laying a solid foundation for constructing cascade protocells.
a Schematic illustrating the working mechanism of the coacervate-based protocells: by regulating the pH environment through enzymatic reactions, the downstream enzymatic reaction activity can be further controlled. b Bright-field microscopy images demonstrating the stability of R10/D10 coacervates upon mixing with different reagents. c pH of the condensate phase in GOx and urease-loaded R10/D10 coacervate protocells following the addition of different concentrations of substrates. n = 5 independent biological replicates. Data are presented as mean ± SD. d Enzymatic activity of GOx and urease under different conditions: the “Coac” group represents the activity of GOx or urease within the R10/D10 coacervate alone; the “GOx-Coac” group shows the enzymatic activity of urease in coacervate co-loaded with 50 nM GOx and supplemented with 10 mM glucose; the “Ure-Coac” group shows the activity of GOx in coacervate co-loaded with 50 nM urease and supplemented with 10 mM urea. n = 30 independent biological replicates. Data are presented as mean ± SD. e (i) Gel electrophoresis of PCR products from coacervate protocells at different glucose and urea concentrations. The “No enzyme” group represents reactions lacking both enzymes and coacervates. (ii) Fluorescence intensity indicating the amount of PCR product obtained within R10/D10 coacervates containing GOx or urease, following the addition of varying concentrations of their respective substrates before PCR. n = 30 independent biological replicates. Data are presented as mean ± SD. f (i) Fluorescence images of the coacervate protocells after IVTT reaction. (ii) Profile of the fluorescence intensity of protocells in each group. The lines in (i) denote the region where the data points are taken. (iii) Average fluorescence intensity of coacervate protocells mixed with different substrates after IVTT reactions, indicating the relative transcription and translation rates. n = 20 independent biological replicates. Data are presented as mean ± SD. (iv) Comparison of fluorescence intensity between the condensate and dilute phases after IVTT reactions, representing the relative transcription and translation rates in the two phases. n = 5 independent biological replicates. Data are presented as mean ± SD.
We then investigate the impact of the pH variation of the coacervate protocells caused by one enzymatic reaction on the activity of another enzyme. As shown in Fig. 6d, the activity of GOx (optimal pH 5.5) was significantly suppressed under alkaline conditions created by urease, while urease activity (optimal pH 7.4) was reduced under acidic conditions generated by GOx26,27. These results highlight how enzymatic cross-regulation within coacervates can modulate biochemical processes through pH-dependent mechanisms.
Based on this, we explore whether the coacervate protocells can be leveraged to recapitulate more complex physiological processes, such as gene expression and signal transduction. To investigate the ability of coacervates to conduct and regulate in vitro DNA replication, we use a qPCR kit with a green fluorescent dye to monitor the accumulation of PCR products. By pre-mixing the GOx- and urease-loaded coacervate solution with the qPCR kit and adjusting the pH microenvironment with varying concentrations (0, 2.5, and 5 mM) of glucose or urea, we create different experimental conditions. Following the complete reaction of the substrates, we analyze the PCR products to evaluate how the local pH influences the reaction efficiency. This approach allows us to better understand the role of coacervates in modulating gene expression under varying environmental conditions. After disassembling the coacervate, the PCR products are quantitatively analyzed using gel electrophoresis, and the fluorescence intensity of the generated DNA in the solution is measured using a microplate reader. As shown in Fig. 6e, PCR reactions are significantly inhibited when 2.5 or 5 mM glucose and urea are added to the solution. Because the optimal pH for PCR reactions is about 8.0, which is about the original pH of coacervate, this inhibitory effect may result from the suppression of PCR-related enzyme activity under acidic or alkaline conditions28.
Next, we perform IVTT experiments using the coacervate protocells. We utilize a cell-free transcription and translation kit combined with a GOx- and urease-loaded R10/D10 coacervate solution to create uniform droplets. The addition of 5 mM glucose or urea influences the reaction conditions within the coacervate environment. Following the incubation at 37 °C, we observe the results under a fluorescence microscope. The successful transcription is evidenced by the presence of RNA exhibiting green fluorescence in the coacervate droplets, and successful translation is indicated by proteins displaying red fluorescence (Fig. 6f i). We then proceed with a quantitative analysis of the fluorescence intensity within the coacervate protocells, as shown in Fig. 6f ii and 6f iii. The results reveal that the addition of urea decreases both RNA and protein production. In contrast, the introduction of glucose enhances the rates of transcription and translation within the coacervate droplets compared to the control group. This could be attributed to the fact that the optimal pH conditions for transcription and translation processes are typically neutral, while the initial pH of the coacervate is nearly 8.0. The addition of urea shifts the pH to a more alkaline level (about 9.6), while the addition of glucose brings the pH closer to neutral (about pH 6.8), facilitating the processes of transcription and translation. Furthermore, to evaluate the relative transcription and translation rates in the two phases of the coacervates, the mixtures containing coacervates and the IVTT kit were centrifuged to separate the condensate and dilute phases. Both phases were then individually incubated at 37 °C for the IVTT reactions, and their fluorescence intensities were compared to determine the relative rates of transcription and translation. As shown in Fig. 6f iv, the results indicated that most transcription and translation reactions occurred within the condensate phase of the coacervates. These findings highlight the effective functioning of the coacervate protocells in accommodating and regulating transcription and translation, demonstrating its potential for applications in synthetic biology.
Discussion
This study employs coacervate droplet-based protocells as a minimal model to explore how phase separation can give rise to and utilize local pH differences for biochemical reaction regulation. We first identified a pH difference across the nucleolus in several tested cell lines and found that treatment with CX-5461 or Act D altered nucleolar structure and function, accompanied by changes in the pH difference between the nucleolus and nucleoplasm. We then developed an in vitro R10/D10 coacervate system to mirror such a pH difference and examine its response to environmental changes. Notably, uniform coacervate droplets are generated through the microfluidic technique, which allows precise control of the droplet composition as well as environmental parameters. We observe that a pH difference can be established between the condensate phase and the dilute phase of coacervates with the presence of acid or base added to the system. Through molecular dynamics simulations, we conclude that these pH differences likely arise from the enrichment of H+/OH- in the condensate phase rich in charged polymers. Intriguingly, we find that the pH difference no longer exists after the disassembly of the coacervate by adding salt, which mirrors the same phenomenon observed after the disassembly of the nucleolus. Based on the pH-regionalized feature of the coacervate droplets, we further demonstrate a way of finely tuning the local pH by incorporating GOx- and urease-catalyzed reactions by leveraging the proteins’ partitioning affinity behavior. The acidic and alkaline byproducts of the enzymatic reactions can alter the local pH in dependence of the substrate concentration, which further impacts the activity of another, downstream enzymatic reaction. With this, the coacervate droplets can serve as protocells accommodating various biological-relevant processes such as PCR and IVTT.
In conclusion, our work emphasizes how pH heterogeneity in coacervate-based protocells can be harnessed to regulate biochemical reactivity. Our findings also highlight the potential of coacervate droplets as pH-regionalized protocells to simulate cellular environments, study phase separation-related cellular activities, and perform complex bio(chemical) reactions.
Methods
Materials
Arginine (R10) and aspartic acid (D10) polypeptides are obtained from Anhui Guoping Pharmaceuticals. The OC1 (FH0777), 3T3 (FH0380), HELA (FH0314), and AML12 (FH0338) cell lines are sourced from Fuheng Biology. DMEM, fetal bovine serum, PBS, penicillin/streptomycin, SNARF-4F, 0.25% trypsin-ethylenediaminetetraacetic acid, intracellular pH calibration buffer kit, live cell imaging solution, urease, and urea are purchased from Thermo Fisher Scientific. Actinomycin D (Act D), and CX-5461 are obtained from Med Chem Express. Diethylaminoethyl dextran hydrochloride (DEAE-dextran), low molecular weight deoxyribonucleic acid (derived from salmon sperm), and poly (dimethyl diallyl ammonium chloride) (PDDA, average Mw 200,000-350,000) are sourced from Sigma-Aldrich. Poly (sodium acrylate) (PAA, average Mw 5000) is obtained from J&K Scientific. Glucose oxidase is ordered from Sangon Biotech. Cell-free gene expression kits and qPCR kits are ordered from Mini PCR Bio. The Glucose Oxidase Activity Assay Kit is ordered from Solarsbio. Urease Activity Assay Kit is ordered from Boxbio. FITC-GOx and RBITC-urease are ordered from AGOMA.
Nucleolar pH quantification using SNARF-4F
SNARF-4F is dissolved in high-grade DMSO to prepare a 5 mM stock solution. For cell co-culture, the concentration of SNARF-4F is adjusted to 7.5 μM. Intracellular pH calibration is performed using a calibration buffer kit following the manufacturer’s instructions. After staining with SNARF-4F, fluorescence images are captured using confocal microscopy with excitation at 488 nm and emissions detected at 580 nm and 640 nm. The intracellular pH is determined based on the emission intensity ratio of at 580 nm and 640 nm (Em580/Em640). The detailed calculation method for the standard curve is provided in the Supplementary Note.
CX-5461 and Actinomycin D inhibitors treatment
For CX-5461-treated cells, 1 mM CX-5461 in high-grade DMSO is added to the culture media and incubated with living cells at a concentration of 1 μg/mL for 1 hour. For low concentration Act D-treated cells, Act D stock solution in DMSO is diluted to 0.063 μg/mL and applied to the cells for 1 hour. For high concentration Act D-treated cells, an Act D stock solution in high-grade DMSO is diluted to 5 μg/mL and applied to the cells for 2 hours. After these treatments, the cells are stained with SNARF-4F, and the intracellular pH is measured with confocal fluorescence.
Coacervate generation
D10/R10 coacervates are produced by directly mixing a 10 mM stock solution of R10 (the pH is adjusted to 8.0 using 1 M NaOH or HCl, and other solutions are adjusted similarly unless otherwise specified) and D10 (pH 8.0) in water at a 1:1 volume ratio. The coacervate samples are freshly prepared for characterization and equilibrated at room temperature for 15 minutes before analysis.
For the GOx-loaded coacervate protocells, 50 nM GOx is added to the 10 mM R10/D10 coacervate solution. Similarly, urease-loaded coacervate protocells are prepared by adding 50 nM urease to the 10 mM R10/D10 coacervate solution.
To create DEAE-dextran/dsDNA coacervates, 10 mg/mL DEAE-dextran (pH 8.0) stock solution and 10 mg/mL dsDNA (pH 8.0) stock solution are mixed directly at a volume ratio of 1:1. To prepare the PDDA/PAA coacervates, 10 mM PDDA (pH 8.0) and 10 mM PAA (pH 8.0) stock solutions are mixed at a volume ratio of 1:4.
Measuring the turbidity of the coacervate solution
To evaluate the stability of the coacervate solution across different pH values, 10 mM R10 and 10 mM D10 solutions are first adjusted to target pH levels ranging from 6.0 to 9.0 with 1 M HCl or 1 M NaOH. The R10 and D10 solutions were then thoroughly mixed at the corresponding pH values and sit for 15 minutes to equilibrate. Next, the turbidity of the coacervate solutions was measured at 600 nm (λ600) using UV-vis spectroscopy, and the turbidity (T) is calculated using the following equation:
Determining the pH value of coacervate in condensate phase and dilute phase
Varying volumes (0, 0.4, 0.8, 1.2, 1.6, and 2.0 μL) of 1 M NaOH or 1 M HCl are mixed with 1 mL of 10 mM R10 solution and 1 mL of 10 mM D10 solution. After mixing, the coacervate solutions are left to stand for 15 minutes. Then, the solutions are centrifugated to separate the condensate phase. The pH values of the condensate and dilute phases are measured using a micro-pH meter.
To investigate the effect of coacervate concentration on the pH difference between the two phases, 1 μM HCl or NaOH is mixed with R10 and D10 at different concentrations (1, 2, 4, 6, 8, and 10 mM), respectively. After 15 minutes, the condensate and dilute phases of the coacervate solution are separated by centrifugation at 2.4 g for 20 minutes, and the pH of each phase is measured using a micro-pH meter.
To examine the impact of salt concentration on the pH difference between the two phases, varying concentrations (0, 10, 20, 30, 40, and 50 mM) of NaCl are added to a 10 mM R10 and D10 solution mixed with 1 μM HCl or NaOH. The pH of both phases is then measured using the method described above.
When measuring the pH of the coacervate solution using SNARF-4F, 10 mM coacervates were mixed with different concentrations of NaOH or HCl and thoroughly vortexed. Subsequently, 7.5 μM SNARF-4F was added to the solution. Fluorescence images were then captured using confocal microscopy with excitation at 488 nm and emissions collected at 580 nm and 640 nm. The pH was determined based on the emission intensity ratio at 580 nm to 640 nm (Em580/Em640).
Molecular dynamics simulation of R10/D10 coacervate formation and ion distribution
CG molecular dynamics simulations are conducted with the Martini 3.0 coarse-grained force field, generating R10 and D10 CG peptides using the martinize.py script in Martini 3.023,29. In all simulations, equal numbers of peptides are randomly distributed in a 20×20×20 nm³ simulation box and solvated with Martini water beads, where one bead represents four water molecules. Subsequently, 100 H3O⁺ or OH⁻ beads are introduced into the simulation box, replacing some water beads. Specifically, H₃O⁺ was modeled as a TQ5 bead with a net charge of +1 e, and OH⁻ as a TQ5 bead with a net charge of −1 e. All other parameters were set according to the default MARTINI 3.0 force field30. Na⁺ and Cl⁻ ions are also added for electrostatic balance.
To investigate the effect of coacervate concentration on H⁺ and OH⁻ distribution, simulations are carried out with 100, 200, and 300 pairs of R10 and D10 CG polypeptides. Following steepest descent minimization, the systems are equilibrated for 10 ns with a 10 fs time step, followed by a 5 μs production run at the same time step. To explore the impact of salt concentration on ion distribution, 0, 1000, and 4000 pairs of Na⁺ and Cl⁻ are added to the simulation box with 300 pairs of R10 and D10. All other parameters are maintained consistently throughout the experiments.
Post-simulation snapshots of the simulation box are captured using the VMD software. For data analysis, normalized RDFs are calculated using the GROMACS tool gmx rdf. The contact number is calculated using gmx mindist with a cut-off of 0.6 nm. The density profile is computed using gmx energy, and the PMF is determined through umbrella sampling (US) simulations.
Microfluidic generation of uniform coacervate droplets
Uniform coacervate droplets are generated using a previously developed microfluidic device integrated with mechanical vibration25. The pre-formed R10/D10, DEAE-dextran/dsDNA, and PDDA/PAA coacervate solutions are centrifuged at 2.4 g for 20 minutes. The precipitated coacervate phase, serving as the dispersed phase, is then injected into the microfluidic device via a syringe, while the dilute phase flows into the collection tube as the continuous phase. The typical flow rates are 20 μL/h for the discrete phase and 5 mL/h for the continuous phase. A sinusoidal perturbation with a 500 Hz actuation frequency is applied to the dispersed phase using a mechanical vibrator to facilitate active droplet generation. The microfluidic process for preparing uniform coacervates is consistent throughout the study.
Microscale thermophoresis analysis
The binding affinity between FITC-GOx and DEAE-dextran/PDDA/R10/dsDNA/PAA/D10 is measured using MST methods31,32. Binding reactions are conducted in water. Ligands (DEAE-dextran/PDDA/R10/dsDNA/PAA/D10) are initially prepared at a concentration of 20 μM and dilution series at 0.6104, 1.2207, 2.4414, 4.8828, 9.7656, 19.5313, 39.0625, 78.125, 156.25, 312.5, 625, 1250, 2500, 5000, 10000, and 20000 nM, respectively, as prepared. The resulting ligand solutions are mixed with 200 nM FITC-GOx in a 1:1 volume ratio. To prevent non-specific adsorption, the samples are loaded into NanoTemper monolithic capillaries immediately after preparation.
Molecular dynamics simulation of protein partitioning affinity into R10/D10 coacervate
To investigate the partitioning affinity of GOx and urease for the coacervates, the files of GOx (1GAL) and urease (4G7E) are downloaded from the Protein Data Bank, and their CG models are generated using the martinize.py script. For the partitioning affinity simulations, 300 pairs of polypeptides and 2 GOx or urease molecules are randomly distributed in a 20×20×20 nm³ simulation box and individually solvated with Martini water beads. Na⁺ and Cl⁻ ions are introduced to maintain electrostatic balance. After minimization, the systems are equilibrated for 10 ns with a 10 fs time step, followed by a 5 μs production run with the same time step.
In simulations examining the interaction between R10/D10 and either GOx or urease, 100 R10/D10 molecules and 1 GOx or urease molecule are randomly distributed in a 20×20×20 nm³ simulation box and solvated with Martini water beads. Na⁺ and Cl⁻ ions are also added for electrostatic balance. After the steepest descent minimization, the systems are equilibrated for 10 ns with a 10 fs time step, then a 5 μs production run is performed using the same time step. The RDFs are calculated using the gmx RDF tool, and the PMF is determined through umbrella sampling simulations.
Detection of enzymatic activities in coacervates
To assess the catalytic activity of GOx within coacervates, a 50 nM GOx-loaded R10/D10 coacervate solution and a bulk aqueous solution (pH 8.0) containing 50 nM GOx were each mixed with glucose solutions at various concentrations (0, 1, 2, 3, 4, 5, 10, 15, and 20 mM). After different reaction times (5 min, 10 min, 15 min, 30 min, 45 min, and 60 min), to minimize turbidity interference from the protocell-containing samples, the mixtures were centrifuged, and the supernatants were collected for GOx activity measurement using a Glucose Oxidase Activity Assay Kit. The initial reaction rate at each substrate concentration was determined by analyzing the linear region of the product formation curve over a 60-minute period (Supplementary Figs. 23 and 24).
Similarly, to evaluate urease activity in coacervates, a 50 nM urease-loaded R10/D10 coacervate solution and a bulk aqueous solution (pH 8.0) containing 50 nM urease are mixed with urea solutions at concentrations of 0, 1, 2, 3, 4, 5, 10, 15, and 20 mM. After different reaction times, to minimize turbidity interference from the protocell-containing samples, the mixtures were centrifuged and the supernatants collected for urease activity measurement using a Urease Activity Assay Kit. The initial reaction rate at each substrate concentration was determined by analyzing the linear region of the product formation curve over a 60-minute period (Supplementary Fig. 25 and 26).
Enzymatic reaction-induced pH difference across the two phases of coacervate protocells
To generate a pH difference across the two phases of R10/D10 coacervate, glucose and urea at concentrations ranging from 0 to 5 mM are added to a 50 nM GOx- or urease-loaded 10 mM R10/D10 coacervate solution. After incubating for 1 hour, the condensate and dilute phases are separated by centrifugation at 2.4 g for 20 minutes, and the pH of each phase is measured using a micro-pH meter.
To examine the effect of coacervate concentration on the pH difference between the two phases, glucose or urea (5 mM) is added to R10/D10 coacervates at concentrations of 1, 2, 4, 6, 8, and 10 mM, each containing 50 nM GOx or urease. After 1 h of incubation, the two phases were separated by centrifugation, and their pH values were measured using a micro-pH meter.
To explore the impact of salt concentration on the pH difference across the phases, salts at concentrations of 0, 10, 20, 30, 40, and 50 mM are added to a 10 mM R10/D10 coacervate solution containing 50 nM GOx or urease, together with 5 mM substrates. After 1 h of incubation, the two phases were separated by centrifugation, and their pH values were measured using a micro-pH meter.
The pH difference between the two phases of DEAE-dextran/dsDNA and PDDA/PAA coacervates is generated by adding 5 mM substrates (glucose or urea) to coacervate solutions containing 50 nM GOx or urease and incubating for 1 h. The condensate and dilute phases are then separated by centrifugation, and their pH values are measured using a micro-pH meter.
Regulating enzymatic activities of GOx and urease in coacervate protocells
To establish acidic conditions, 10 mM glucose is added to a 10 mM R10/D10 coacervate solution co-loaded with 50 nM GOx and 50 nM urease. After the glucose reaction for 1 hour, 1 mM urea is introduced to evaluate urease activity. Following a 15-minute reaction, the solution is centrifuged, and the supernatant is collected for urease activity measurement using the urease activity assay kit.
For alkaline conditions, 10 mM urea is added to a 10 mM R10/D10 coacervate solution co-loaded with 50 nM urease and 50 nM GOx. After the urea reaction for 1 hour, 1 mM glucose is introduced to assess the relative activity of GOx. Similarly, the solution undergoes a 15-minute reaction, centrifugation, and collection of the supernatant, followed by GOx activity measurement using the glucose oxidase activity assay kit.
Regulating PCR efficiency in coacervate protocells
GOx- or urease-loaded R10/D10 coacervate protocells are prepared as previously described. The resulting coacervate solutions are then mixed with a commercial PCR reagent solution at a 10:1 volume ratio. After reacting with glucose or urea at concentrations of 0, 2.5, or 5 mM, the PCR process is performed using a PCR machine (Arhat 96-Deep Well MP60401). The specific reaction parameters are set as per the manufacturer’s instructions. After the reaction, the products are collected and treated with a 1 M NaCl solution with a volume equal to that of the mixture for further study.
For quantitative analysis through nucleic acid gel electrophoresis, 15 μL of the product solution is mixed with 4 μL of loading dye. The samples are loaded onto a gel and subjected to electrophoresis for approximately 15 minutes. Finally, the results are analyzed using a gel imaging system. For quantitative analysis through UV-vis spectroscopy, this solution is added to a 96-well plate. Then, the fluorescence intensity of the solution is quantitatively analyzed using a microplate reader.
Regulating IVTT efficiency in coacervate protocells
Cell-free transcription and translation reaction kits are mixed with 50 nM GOx- or urease-loaded R10/D10 coacervate protocells (10 mM) at a 10:1 volume ratio, followed by incubation with 5 mM urea, water, or 5 mM glucose. After 1 hour of reaction with glucose or urea, the coacervate solution is centrifuged at 2.4 g for 20 minutes to separate the condensate phase. The condensate is then injected into a microfluidic device, as previously described, to generate uniform coacervate microdroplets. These uniform coacervate droplets are subsequently incubated at 37 °C for 30 minutes to initiate transcription and translation reactions. Green fluorescence signals (indicating RNA synthesis) and red fluorescence signals (indicating protein synthesis) are observed in the uniform coacervate microdroplets. The gray values of the droplets are analyzed and recorded.
Statistics and reproducibility
All confocal and other microscopy images represent experiments independently repeated at least 3 times, yielding similar result. For all graphs and statistical analyses, the number of independent replicates (n) is provided. Statistical tests are specified in the legends or on the figure panels, and exact P values are provided whenever possible.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper.
Code availability
The molecular dynamics simulation codes supporting the findings of this study are available in the source data.
References
Li, X., Egervari, G., Wang, Y., Berger, S. L. & Lu, Z. Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat. Rev. Mol. Cell Biol. 19, 563–578 (2018).
Ravindran, R. et al. Peroxisome biogenesis initiated by protein phase separation. Nature 617, 608–615 (2023).
Stegen, S. & Carmeliet, G. Metabolic regulation of skeletal cell fate and function. Nat. Rev. Endocrinol. 20, 399–413 (2024).
Baker-Austin, C. & Dopson, M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 15, 165–171 (2007).
King, M. R. et al. Macromolecular condensation organizes nucleolar sub-phases to set up a pH gradient. Cell 187, 1889–1906.e1824 (2024).
Dai, Y. et al. Biomolecular condensates regulate cellular electrochemical equilibria. Cell 187, 5951–5966 e5918 (2024).
Lenne, P.-F. & Trivedi, V. Sculpting tissues by phase transitions. Nat. Commun. 13, 664 (2022).
Cook, A. B., Novosedlik, S. & van Hest, J. C. M. Complex Coacervate Materials as Artificial Cells. Acc. Mater. Res. 4, 287–298 (2023).
Wang, Y. et al. Biomolecular condensates mediate bending and scission of endosome membranes. Nature 634, 1204–1210 (2024).
Galvanetto, N. et al. Extreme dynamics in a biomolecular condensate. Nature 619, 876–883 (2023).
Xu, C., Martin, N., Li, M. & Mann, S. Living material assembly of bacteriogenic protocells. Nature 609, 1029–1037 (2022).
Jambon-Puillet, E. et al. Phase-separated droplets swim to their dissolution. Nat. Commun. 15, 3919 (2024).
Testa, A. et al. Sustained enzymatic activity and flow in crowded protein droplets. Nat. Commun. 12, 6293 (2021).
Wang, Z. et al. Coacervate Microdroplets as Synthetic Protocells for Cell Mimicking and Signaling Communications. Small Methods 7, e2300042 (2023).
Chen, H., Guo, J., Bian, F. & Zhao, Y. Microfluidic technologies for cell deformability cytometry. Smart Med 1, e20220001 (2022).
Shang, L., Cheng, Y. & Zhao, Y. Emerging Droplet Microfluidics. Chem. Rev. 117, 7964–8040 (2017).
Yuan, Y. et al. Digital droplet RT-LAMP increases speed of SARS-CoV-2 viral RNA detection. Smart Med. 3, e20240008 (2024).
Lafontaine, D. L. J., Riback, J. A., Bascetin, R. & Brangwynne, C. P. The nucleolus as a multiphase liquid condensate. Nat. Rev. Mol. Cell Biol. 22, 165–182 (2020).
Tafech, A., Beaujean, C., Usson, Y. & Stéphanou, A. Generalization of the Ratiometric Method to Extend pH Range Measurements of the BCECF Probe. Biomolecules 13, 442 (2023).
Aryan, F. et al. Nucleolus activity-dependent recruitment and biomolecular condensation by pH sensing. Mol. Cell 83, 4413–4423.e4410 (2023).
Mars, J. C. et al. The chemotherapeutic agent CX-5461 irreversibly blocks RNA polymerase I initiation and promoter release to cause nucleolar disruption, DNA damage and cell inviability. NAR Cancer 2, zcaa032 (2020).
Choi, S., Meyer, M. O., Bevilacqua, P. C. & Keating, C. D. Phase-specific RNA accumulation and duplex thermodynamics in multiphase coacervate models for membraneless organelles. Nat. Chem. 14, 1110–1117 (2022).
Souza, P. C. T. et al. Martini 3: a general purpose force field for coarse-grained molecular dynamics. Nat. Methods 18, 382–388 (2021).
Liu, Y., Wang, X., Wan, Z., Ngai, T. & Tse, Y. S. Capturing coacervate formation and protein partition by molecular dynamics simulation. Chem. Sci. 14, 1168–1175 (2023).
Wang, C., Yang, X., Wang, Q., Zhang, L. & Shang, L. Glucose Responsive Coacervate Protocells from Microfluidics for Diabetic Wound Healing. Adv. Sci. 11, 2400712 (2024).
Onosakponome Iruoghene, E. A. L., Eze, S. O., Onyebuchi & Chilaka, F. Chiemeka Kinetics and Thermodynamic Properties of Glucose Oxidase Obtained from Aspergillus fumigatus ASF4. Trop. J. Nat. Prod. Res. 6, 438–445 (2022).
Cesareo, S. D. & Langton, S. R. Kinetic properties of Helicobacter pylori urease compared with jack bean urease. FEMS Microbiol. Lett. 78, 15–21 (1992).
Gao, H., Kong, J., Li, Z., Xiao, G. & Meng, X. Quantitative analysis of temperature, salinity and pH on WSSV proliferation in Chinese shrimp Fenneropenaeus chinensis by real-time PCR. Aquaculture 312, 26–31 (2011).
Tsanai, M., Frederix, P. W. J. M., Schroer, C. F. E., Souza, P. C. T. & Marrink, S. J. Coacervate formation studied by explicit solvent coarse-grain molecular dynamics with the Martini model. Chem. Sci. 12, 8521–8530 (2021).
Guo, Y., Huang, S.-F., Mabuchi, T. & Tokumasu, T. Analysis of structural and water diffusional properties of ionomer thin film by coarse-grained molecular dynamics simulation. J. Mol. Liq. 391, 123190 (2023).
Al-Jubair, T. et al. Characterization of human aquaporin protein-protein interactions using microscale thermophoresis (MST). STAR Protoc. 3, 101316 (2022).
Jerabek-Willemsen, M., Wienken, C. J., Braun, D., Baaske, P. & Duhr, S. Molecular Interaction Studies Using Microscale Thermophoresis. ASSAY Drug Dev. Technol. 9, 342–353 (2011).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2024YFA0919100), the National Natural Science Foundation of China (32271383), and the Shenzhen Medical Research Fund (B2401006). We are particularly thankful to the Core Facility of Shanghai Medical College, Fudan University.
Author information
Authors and Affiliations
Contributions
C.W. designed and performed the majority of the experiments, analyzed most of the data, and wrote the manuscript. Z.F. contributed to the analysis of turbidity data. L.Z. carried out part of the cell experiments. L.S. supervised the project, contributed to the experimental design, and co-wrote the manuscript. Y.Z. contributed to the experimental design and manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dhwanit Dave and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, C., Fang, Z., Zhang, L. et al. Coacervate droplets as pH-regionalized protocells. Nat Commun 17, 2252 (2026). https://doi.org/10.1038/s41467-026-68980-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68980-6