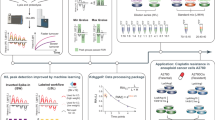
Abstract
Mass spectrometry (MS) is indispensable for high-throughput quantitation of protein expression. But protein function is regulated by factors beyond abundance alone. Here, we evaluate two supercharging reagents, dimethyl sulfoxide (DMSO) and m-nitrobenzyl alcohol (mNBA), in narrow-window data-independent acquisition (nDIA)-MS. DMSO markedly enhances MS signal and protein identification, whereas mNBA primarily increases peptide identifications. Optimizating nDIA-MS with 3% DMSO boosts signal intensity by up to 56%, enabling identification of ~9,600 proteins from 1 µg HeLa digest in 15 min. Using this methodology, we quantify solubility and abundance changes in 8,694 proteins across three cell lines following short-term treatment with the proteasome inhibitor MG132 and the SUMO-activating enzyme inhibitor ML-792. MG132 affects the solubility of 1,723 proteins and the abundance of 374, and ML-792 affects 1,294 and 288, respectively. The drugs elicit distinct and sometimes opposing solubility shifts; for instance, MG132 insolubilizes HSF1, ML-792 solubilizes SP100 and insolubilizes PLOR3G, and SMAD2 shows opposite responses to those two treatments. These results reveal widespread, drug-induced remodeling of the protein solubility landscape and establish solubility profiling by nDIA-MS as a broadly applicable platform for uncovering protein state transitions and cellular responses to perturbation.
Similar content being viewed by others
Introduction
Mass spectrometry (MS)-based shotgun proteomics has advanced significantly over the past decade, firmly establishing itself as the gold standard for high-throughput protein quantitation1,2. However, protein function is dictated by a complex interplay of factors beyond abundance. Dynamic protein states including subcellular localization, chromatin association, and membrane association play critical roles in cellular regulation but remain largely invisible to conventional proteomic workflows. Despite the importance of elucidating and measuring these intracellular protein states, high-throughput strategies to systematically track them remain scarce and technically challenging3,4.
The proteasome inhibitor MG132 is a well-established tool for studying the ubiquitin-proteasome system (UPS), a critical regulator of cellular proteostasis and protein degradation5. MG132 exerts its effects by inhibiting the proteolytic activity of the 26S proteasome, leading to the accumulation of polyubiquitinated proteins in cells6. This accumulation can modulate a host of cellular processes such as gene expression, cell cycle, and programmed cell death5. The SUMO-activating enzyme inhibitor ML-792 has gained attention as a powerful reagent for investigating the role of SUMOylation in cellular regulation. ML-792 inhibits the activation of SUMO proteins, thereby decreasing the overall level of SUMOylation in cells7. That inhibition disturbs the delicate balance between SUMOylation and deSUMOylation, leading to changes in the stability, function, and localization of many proteins8,9. Notably, ML-792 has been shown to inhibit the formation of promyelocytic leukemia nuclear bodies (PML-NBs) in the cell nucleus and to influence chromatin remodeling, protein phase transitions, and the cellular response to stress10.
Efforts to identify proteins and pathways affected by inhibitors of proteasome function and SUMO-activation have been extensively pursued11,12,13,14,15. However, most studies have focused on profiling the ubiquitin- or SUMO-modified proteomes and assessing changes in overall protein abundance16,17,18. We recently discovered that treatment with MG132 and/or ML-792 alters the chromatin association of RAD54L219, suggesting that the proteasome and SUMOylation pathways modulate protein localization. Within cells, proteins can exist in either soluble states—freely diffusing in the cytoplasm or nucleoplasm—or insoluble states, in which they are tightly bound to membrane, chromatin, or other cellular structures20,21,22. Perturbations including drug treatments and various forms of cellular stress can drive shifts in protein solubility23,24,25,26. Motivated by our findings, we aimed to globally investigate how inhibition of the proteasome or SUMO pathway influences protein solubility across the proteome.
Supercharging in electrospray ionization (ESI)-based MS refers to the use of chemical additives that increase the charge states of analyte ions, thereby improving ionization efficiency, signal intensity, and sequence coverage. Supercharging reagents such as dimethyl sulfoxide (DMSO) and m-nitrobenzyl alcohol (mNBA) modify droplet surface tension and solvent evaporation dynamics during ionization, promoting the formation of higher-charged ions27,28,29. These effects enhance detection of low-abundance peptides and proteins and increase overall proteome coverage, making supercharging particularly useful for high-throughput quantitative workflows.
In the current study, we present a workflow to uncover protein targets that undergo solubility change following MG132 and ML-792 treatment. To enhance the method’s sensitivity, we integrate supercharging mobile phase modifiers with narrow-window data-independent acquisition (nDIA) MS analysis30,31,32. The optimized method enables identification of ~9600 human proteins in just 15 min. We analyze whole cell extracts (WCE) and Triton X-100 extracts (insoluble fractions) from three cancer cell lines—U2OS, HeLa, and HEK293A—treated with MG132 and ML-792. The resulting data provides a comprehensive, quantitative overview of solubility shifts for 8694 proteins. In addition to the significant enrichment of chromatin-associated proteins in the insoluble fraction, we also identify proteins annotated to other cellular components, demonstrating the versatility of the workflow for characterizing a broad range of cellular proteins beyond those associated with chromatin. Overall, this study highlights the potential of solubility-shift proteomics as a powerful, high-throughput tool for identifying functional protein changes that extend beyond simple alterations in protein expression.
Results
Supercharging mobile phase modifiers enhance nDIA-MS
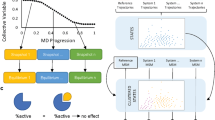
The enhancing effect of 5% DMSO and other supercharging reagents on electrospray ionization and peptide detection in proteomics (data-dependent acquisition, DDA) has been previously reported27,28,29. Inspired by those prior studies, we tested two commonly applied supercharging mobile phase modifiers, dimethylsulfoxide (DMSO) and m-nitrobenzyl alcohol (m-NBA), in combination with nDIA-MS. In testing of DMSO concentrations ranging from 1% to 7%, 3% DMSO produced the highest total ion current (TIC), approximately 56% greater than without DMSO (Fig. 1A, B, and Supplementary Data 1). TIC values were increased by 33%, 28%, and 42% at DMSO concentrations of 1%, 5%, and 7%, respectively. The greatest number of proteins was also identified with 3% DMSO (Fig. 1C). Interestingly, the highest peptide identification was achieved with only 1% DMSO (Fig. 1D). Importantly, DMSO had minimal impact on peptide length and charge state distributions (Fig. 1E–J). These findings suggest that, beyond the previously reported charge state coalescence, additional mechanisms may contribute to DMSO-mediated signal enhancement33.
A The TIC intensities obtained with different composition of LC solvents. B Example of TIC profiles obtained with or without 3% DMSO. (C) Identified proteins, (D) identified peptides, (E) average peptide length, (F) average peptide charge, (G) number of peptides with 2+ charge state, (H) number of peptides with 3+ charge state, and (I) number of peptides with 4+ charge state obtained with different composition of LC solvents, (J) distribution of peptide charge states obtained with different composition of LC solvents, (K, L) number of protein groups (K) and peptides (L) identified with the optimized LC-MS method from 1 µg of HeLa digest using in-silico predicted and experimental spectral library, respectively. All error bars and scatter points represent n = 3 technical replicates from a single HeLa digest sample. Data are presented as mean ± SD. Source data are provided as a Source Data file.
Although m-NBA is known to exert a stronger supercharging effect than DMSO, prior studies did not find clear improvement in peptide or protein identification rates using m-NBA. Consistent with those reports, our test of m-NBA concentrations from 0.001% to 0.1% revealed no obvious increase in TIC intensities (Fig. 1A). At higher concentrations, m-NBA even led to a decrease in TIC, correlating with decreased protein identification (Fig. 1C). Nevertheless, peptide identification increased with m-NBA, with 0.1% yielding the highest number of peptides—surpassing the effect observed with DMSO (Fig. 1D). In addition, m-NBA increased both the average peptide length and the charge state, with 0.1% having the strongest effect (Fig. 1E, F). Analysis of peptide charge distribution revealed a dose-dependent decrease in doubly charged peptides and a corresponding increase in triply and quadruply charged peptides (Fig. 1G–J).
We also tested combinations of 3% DMSO with varying concentrations of m-NBA. Higher concentrations of m-NBA slightly decreased TIC and protein identification. However, the addition of 0.1% m-NBA to 3% DMSO increased peptide number, length, and charge compared with addition of DMSO alone. These observations are consistent with previous reports34.
Taken together, these results suggest that DMSO and m-NBA enhance MS performance through distinct mechanisms, differentially affecting protein and peptide identification. Therefore, the choice of DMSO, m-NBA, or their combination should be tailored to specific experimental goals. Given our aim to maximize protein identification, we selected 3% DMSO for all subsequent experiments.
Data processing was performed using both predicted library and experimental spectral library workflows in DIA-NN. The human protein database was used to generate an in-silico predicted spectral library. For generation of the experimental spectral library, HeLa digests were separated into eight fractions using a C18 column, followed by MS analysis of each fraction. The resulting MS data were used to build a spectral library in DIA-NN. On average, 9392 and 9606 proteins were identified from 1 µg of HeLa digest using predicted and experimental spectral library, respectively (Fig. 1K and Supplementary Data 2), along with 90,885 and 92,276 peptides (Fig. 1L and Supplementary Data 2).
nDIA-MS enables proteome-wide profiling of drug-induced changes in protein solubility
As noted in the introduction, proteins exist within cells in either soluble states—freely diffusing in the cytoplasm or nucleoplasm—or insoluble states, in which they are membrane-associated or tightly bound to cellular structures such as chromatin or the cytoskeleton. Various factors, including drug treatments and cellular stress, can trigger shifts in protein solubility. In our previous study, we used Triton X-100 to pre-extract soluble proteins and successfully detected dynamic chromatin association of RAD54L2 following MG132 and/or ML792 treatment via immunofluorescence staining19. Prompted by those findings, we hypothesized that combining Triton X-100 pre-extraction method with our optimized nDIA-MS proteomic workflow could unveil an extensive landscape of protein solubility shifts and, therefore, protein targets of MG132 and ML-792.
To minimize confounding effects on protein expression and to increase specificity for the detection of solubility shifts, we applied a short-term (1-hour) treatment with MG132 and/or ML-792. Despite the brief exposure, MG132 robustly increased global ubiquitination in both whole cell extracts (WCE) and insoluble fractions (Fig. 2A), and ML-792 effectively suppressed global SUMOylation in both fractions (Fig. 2B).
Western blotting result of Ubiquitin (A) and SUMOs (B). Experiments were repeated at least three times, and similar results were obtained. (C) Schematic of the sample preparation, MS data acquisition, and data processing workflow of solubility shift proteomics. D MS intensity changes of histones and tubulins identified in three cancer cell lines. Only samples without drug treatment are displayed. Three biological replicates were used for each sample group. Scatter points represent the log₂ enrichment ratios of identified histone (n = 14) or tubulin (n = 15) isoforms in the insoluble fraction relative to whole-cell lysate. Box plots show the median (center line), the interquartile range (25th–75th percentiles; box), and the minimum to maximum values (whiskers). E Functional clustering of cellular component categories for proteins that were consistently enriched in insoluble fraction samples. Only proteins with pairwise ratios >2 or <0.5, and p-value < 0.05 are used for analysis. Enrichment analysis was performed using DAVID with default settings. P-values were calculated using the modified Fisher’s exact test (EASE score); exact P-values are indicated on the figure. No adjustments for multiple comparisons were applied. Source data are provided as a Source Data file.
We designed the experimental workflow as shown in Fig. 2C. Briefly, whole cell extract (WCE) samples were prepared by washing cells twice with 1× PBS followed by direct lysis. Insoluble pellet (Insoluble) samples were obtained by pre-extracting soluble proteins in cells with 0.2% Triton X-100 in PBS, followed by a PBS wash and centrifugation to collect the insoluble material. After cell lysis, protein digestion, and sample cleanup, the resulting peptide samples were analyzed by nDIA-MS (Fig. 2C). The MS result provided quantitative information of solubility and abundance changes for 8,694 proteins across three cell lines following short-term treatment with MG132 and ML-792, including 7,439 proteins identified from the WCE samples and 5787 proteins from the insoluble fraction (Supplementary Fig. 1A, and Supplementary Data 3). All samples exhibited Pearson correlation coefficients greater than 0.95 among three biological replicates, demonstrating excellent reproducibility in terms of relative quantitation (Supplementary Fig. 1B).
To verify the effectiveness of the pre-extraction protocol, we checked the levels of all identified tubulins and histones (Fig. 2D). As expected, tubulin levels were significantly decreased in the insoluble fraction, indicating efficient depletion of soluble proteins. In contrast, histones—tightly bound to chromatin—were consistently enriched, confirming successful isolation of chromatin-associated proteins and demonstrating the method’s utility for studying dynamic protein-chromatin interactions.
To further characterize the insoluble proteome, we performed functional clustering analysis of proteins significantly enriched in the insoluble fraction (Fig. 2E). As anticipated, we observed strong enrichment for chromatin-associated Gene Ontology (GO) cellular component categories, including “chromosome, centromeric region,” “chromatin silencing complex,” “MOZ/MORF histone acetyltransferase complex,” and “nucleosome.” Additionally enriched nuclear structures included the small subunit processome, spliceosome, preribosome, telomere, nuclear envelope, kinetochore, and small nuclear ribonucleoproteins (snRNPs).
Short-term inhibition of the proteasome or SUMOylation activates early stress-responsive gene networks
We examined protein abundance changes following a 1 h treatment with MG132 or ML-792, aiming to capture early and sensitive targets of these inhibitors. For MG132 treatment, we compared: i) MG132-treated samples to untreated controls; and ii) combination MG132/ML-792-treated samples to ML-792-treated samples. We then consolidated the results for each cell line (Fig. 3A–F). MG132 treatment led to the upregulation of 130 proteins in U2OS, 40 in HeLa, and 127 in HEK293 A cells (Fig. 3A–C, G). In contrast, 21 proteins in U2OS, 28 in HeLa, and 64 in HEK293A were significantly downregulated—relatively low numbers, consistent with the expected stabilization of proteins upon proteasome inhibition. Altogether, MG132 altered the abundance of 374 proteins across the three cell lines—262 upregulated and 112 downregulated. Comparative analysis revealed 32 proteins were consistently upregulated and 1 downregulated in at least two cell lines (Fig. 3G).
Volcano plots showing the comparisons of protein levels between MG132 vs vehicle, and MG132 + ML-792 vs ML-792 from (A) U2OS, (B) HeLa, and (C) HEK293A cells. Volcano plots showing the comparisons of protein levels between ML-792 vs vehicle, and MG132 + ML-792 vs MG132 from (D) U2OS, (E) HeLa, and (F) HEK293A cells. P-values were calculated using a two-tailed Student’s t-test. No adjustments for multiple comparisons were applied. G Venn diagram showing the overlaps of up- and down-regulated proteins obtained from three cancer cell lines following treatment of MG132. Differentially regulated proteins from two pairwise comparisons of MG132 vs vehicle and MG132 + ML-792 vs ML-792 were combined. H Venn diagram showing the overlaps of up- and down-regulated proteins obtained from three cancer cell lines following treatment of ML-792. I, J Heatmaps showing differential proteins modulated by MG132 (I) and ML792 (J), respectively. Only proteins dysregulated in at least two cell lines are displayed. Western blotting results of representative proteins modulated by MG132 or ML792, including C-FOS (K), EGR1 (K), CCN1/2 (L), HIF1A (M). Experiments were repeated at least three times, and similar results were obtained. Source data are provided as a Source Data file.
For ML-792 treatment, we compared: i) ML-792-treated samples to untreated controls; and ii) MG132 plus ML-792-treated samples to MG132-treated samples. We again consolidated the results for each cell line (Fig. 3D–F, H). The results indicated 46 upregulated proteins in U2OS, 68 in HeLa, and 61 in HEK293A (Fig. 3H). Conversely, 34 proteins in U2OS, 23 in HeLa, and 64 in HEK293A were significantly downregulated. In total, ML-792 altered the abundance of 288 proteins—167 upregulated and 121 downregulated. Six proteins were consistently upregulated in at least two of the three cell lines (Fig. 3H).
As anticipated, the number of protein abundance changes identified in this study is relatively modest compared to previous reports due to the short (1-hour) treatment duration. Under these conditions, only the most rapidly responding and sensitive protein targets of MG132 and ML-792 were captured. Fold-changes of proteins altered in at least two cell lines are visualized in Fig. 3I, J, and revealing highly similar expression patterns across treatments and cell types.
Interestingly, both MG132 and ML-792 induced expression of classic immediate-early genes (IEGs), including FOS and EGR1 (Fig. 2I, J)35. IEGs are rapidly activated without the need for de novo protein synthesis and play critical roles in stress responses and cell fate decisions36. Western blot analyses validated these findings: MG132, ML-792, or the combination significantly increased whole-cell expression of FOS and EGR1 in HeLa cells (Fig. 3K). A similar but less pronounced increase was observed in U2OS and HEK293A cells. This trend was also reflected in the insoluble protein fractions. In addition, we observed increased expression of another canonical IEG, NR4A1, in HeLa cells following either treatment (Fig. 3B).
We also detected increased expression of CCN1 (Cyr61) and CCN2 (CTGF), members of the CCN family of matricellular proteins that regulate cell adhesion, migration, angiogenesis, and tissue remodeling (Fig. 3I, J)37,38. These are considered early response genes and have been reported to be rapidly induced by diverse stimuli39. Western blot analysis confirmed increased CCN1 and CCN2 expression in HeLa cells following treatment, with weaker induction in U2OS and HEK293A cells (Fig. 3L). Notably, CCN1 overexpression has been shown to elevate HIF1A levels in gastric cancer cells37. In line with this, we observed elevated HIF1A levels in both MS and Western blot datasets (Fig. 3I, M).
We also detected significant upregulation of CCND1 and other cell cycle regulators—CCND1, CDC25B, and CDCA4—in U2OS and HEK293A cells following MG132 treatment (Fig. 2I, J), supporting previous findings that MG132 blocks TSA-induced degradation of cyclin D1 (CCND1) in MCF-7 cells40. Together, these findings suggest that even short-term MG132 or ML-792 exposure can activate immediate-early gene expression and trigger rapid regulatory changes in protein abundance, particularly involving stress response and cell cycle pathways. Whether other genes identified here have the similar functions in early Stress-Responsive Gene Networks will need further study.
Proteasome inhibition remodels protein solubility across the proteome
To identify changes in protein solubility, we calculated a relative insolubility for each protein by dividing its intensity in the insoluble fraction by its average intensity in the corresponding whole-cell extract (WCE) samples. Proteins with an insolubility ratio >2 and a p-value < 0.05 were considered to become more insoluble upon drug treatment, whereas those with a ratio <0.5 and p-value < 0.05 were considered to become more soluble. We then compared the relative insolubility between MG132-treated and untreated groups, as well as between MG132 plus ML792-treated and ML792-treated groups across all three cell lines (Fig. 4A–C). In U2OS cells, MG132 treatment increased the insolubility of 380 proteins and decreased the insolubility of 260 proteins. In HeLa cells, 402 proteins became more insoluble and 144 more soluble. In HEK293A cells, 448 proteins were more insoluble and 335 more soluble (Fig. 4D, E). Altogether, MG132 treatment led to increased insolubility in 1055 proteins across the three cell lines, with 152 shared across at least two lines. Conversely, 668 proteins became more soluble, with 69 of these shared among at least two cell lines.
Volcano plots showing the comparisons of relative insolubility rates between MG132 vs vehicle, and MG132 + ML-792 vs ML-792 from (A) U2OS, (B) HeLa, and (C) HEK293A cells. P-values were calculated using a two-tailed Student’s t-test. No adjustments for multiple comparisons were applied. Venn diagram showing the overlaps of (D) up- and (E) down-regulated proteins obtained from three cancer cell lines. Differentially regulated proteins from two pairwise comparisons of MG132 vs vehicle and MG132 + ML-792 vs ML-792 were combined. Functional enrichment analysis of the proteins whose insolubility rates were consistently (F) up- and (G) down-regulated in at least two cell lines. Enrichment analysis was performed using DAVID with default settings. P-values were calculated using the modified Fisher’s exact test (EASE score). No adjustments for multiple comparisons were applied. H–J Heatmaps showing proteins modulated by MG132. K–M Western blotting result of representative proteins that modulated by MG132 or ML792, including BAG6, UBL4A, TEX264, HSF1, REV1, and ATR1. Experiments were repeated at least three times, and similar results were obtained. Source data are provided as a Source Data file.
To better understand the biological functions associated with the observed solubility changes, we performed functional enrichment analyses on proteins with altered relative insolubility in at least two cell lines following MG132 treatment (Fig. 4F, G). The most enriched pathways of insolubilized proteins included those involved in ubiquitin-dependent protein catabolism, autophagy, transcriptional regulation, and the DNA damage response (Fig. 4F). The most enriched categories of solubilized proteins included subsets of proteins within the transcription and DNA repair pathways (Fig. 4G).
Proteins involved in the ubiquitin-proteasome system (UPS) were significantly affected. For example, proteasome activator subunits PSME1, PSME2, and PSME3 showed decreased solubility in MG132-treated HeLa cells (Fig. 4B). Eleven proteins involved in ubiquitin-dependent catabolism became less soluble in all three cell lines (Fig. 4H). Highlights include UBL4A and BAG6, components of the BAG6 complex that plays a key role in co-translational protein quality control and degradation41. NDFIP1/2 are endosomal proteins that activate Nedd4 family E3 ligases to regulate receptor signaling42. XBP1, a key transcription factor in the unfolded protein response (UPR), also showed altered solubility43. Other notable proteins include TRIM32 (a TRIM-family E3 ligase44), DTX3L (partnered with PARP9 in immune and DNA damage responses45), and DTL (a substrate adaptor in the DDB1-CUL4 E3 ligase complex involved in cell cycle and DNA repair46).
Autophagy and the ubiquitin-proteasome system (UPS) are two major pathways responsible for the degradation of most cellular proteins in eukaryotic cells47. Whereas the UPS typically handles most of the protein degradation, autophagy is primarily involved in degrading long-lived or aggregated proteins, as well as cellular organelles. Twelve autophagy proteins became less soluble in at least two cell lines following MG132 treatment (Fig. 4I). Five of them—TEX264, AMBRA1, OPTN, ATG9A, and GABARAPL2—are core components of autophagosomes, while CTSD and ATG3 contribute to their assembly48,49. TEX264 is also known as a receptor involved in reticulophagy and DNA-protein crosslink (DPC) repair50,51,52. Our results showed decreased TEX264 solubility across all cell lines (Fig. 4I). TMEM59, which promotes LC3 lipidation through ATG16L1, and SYT11, a regulator of autophagy-lysosome pathways53, also exhibited altered solubility (Fig. 4I). Western blotting validated the decreased solubility of BAG6, UBL4A, and TEX264 in response to MG132 (Fig. 4K, L). These findings suggest that MG132 not only blocks protein degradation but may also trap protein quality control adaptors—such as those in the UPS or autophagy machinery—within subcellular compartments such as chromatin, ER, or endosomes, thereby delaying their recycling.
Interestingly, we found both increased and decreased solubility of proteins in DNA damage response and transcription pathways (Fig. 4J). In total we identify 18 significantly changes in solubility of proteins involved in DNA damage, among which HSF1, UBR5, TP73, PARP9, RAD51AP1, RAD51C, and GNL1 were upregulated in relative insolubility by MG132 whereas H2AX and BCCIP were downregulated in relative insolubility by MG132. HSF1, known for driving chaperone expression, accumulates in nuclear stress bodies under proteotoxic stress54. The notable increase of HSF1’s relative insolubility observed by MS was also detected in Immunofluorescence and western blotting in this study (Fig. 4M, and Supplementary Fig. 2). These findings confirm that MG132 can induce HSF1 condensation and nuclear body formation. Notably, the observed solubility shifts of some DNA damage proteins, such as REV1, were cell line-specific, which was validated by western blotting (Fig. 4J, M).
A total of 56 transcriptional regulators showed solubility shifts upon MG132 treatment (Fig. 4J). Given that many DNA repair and transcription factors operate on chromatin, the observed solubility shifts may reflect altered chromatin association. This implies a regulatory mechanism in which proteasome activity modulates the chromatin engagement and function of these factors. This is consistent with reports of ubiquitin-mediated regulation of transcription and repair proteins55.
In summary, our data reveal widespread remodeling of protein solubility following proteasome inhibition, particularly affecting proteins involved in DNA damage response, transcription, UPS, and autophagy. These results highlight a complex, proteasome-dependent regulatory layer that governs protein localization and function during early stress responses.
ML-792 modulates proteome solubility
To examine how SUMOylation inhibition influences protein solubility, we calculated relative insolubility as in the previous section and compared: i) ML-792-treated versus non-treated; and ii) combination MG132/ML-792-treated versus MG132-treated groups across three cell lines (Fig. 5A–C). Proteins exhibiting more than a 2-fold increase or a decrease below 0.5 in insolubility, with a p-value < 0.05, were considered significantly modulated by ML792. In U2OS cells, ML792 treatment led to increased insolubility of 175 proteins and solubilization of 229 proteins (Fig. 5D, E). In HeLa cells, 145 proteins became more insoluble and 256 more soluble. HEK293A cells showed 323 proteins with increased insolubility and 279 with increased solubility. Across all three cell lines, a total of 597 proteins were insolubilized and 697 proteins were solubilized following ML-792 treatment, with 44 and 63 of these proteins, respectively, shared across at least two cell lines.
Volcano plots showing the comparisons of relative insolubility rates between ML-792 vs vehicle, and MG132 + ML-792 vs MG132 from (A) U2OS, (B) HeLa, and (C) HEK293A cells. P-values were calculated using a two-tailed Student’s t-test. No adjustments for multiple comparisons were applied. Venn diagram showing the overlaps of (D) up- and (E) down-regulated proteins obtained from three cancer cell lines. Differentially regulated proteins from two pairwise comparisons of ML-792 vs vehicle and MG132 + ML-792 vs MG132 were combined. F Functional enrichment analysis of the proteins whose insolubility rates were consistently up- and down-regulated in at least two cell lines. Enrichment analysis was performed using DAVID with default settings. P-values were calculated using the modified Fisher’s exact test (EASE score). No adjustments for multiple comparisons were applied. G–I Heatmaps showing proteins modulated by ML-792. J–L Western blotting result of representative proteins that modulated by MG132 or ML792, including FANCD2, POLR3G, SP100, and DAXX. Experiments were repeated at least three times, and similar results were obtained. Source data are provided as a Source Data file.
Functional enrichment analysis of proteins with consistent solubility shifts across two or more cell lines revealed significant enrichment in transcriptional regulation pathways (Fig. 5F). Additionally, components of the SUMOylation machinery and the Fanconi anemia pathway were solubilized following ML-792 treatment (Fig. 5F–H). As a validation of SUMOylation inhibition, all three SUMO isoforms—SUMO1, SUMO2, and SUMO3—were solubilized in all three cell lines upon ML-792 treatment (Fig. 5G).
ML-792 predominantly affected solubility of transcription-related proteins, increasing solubility in some while decreasing it in others (Fig. 5I). For example, RNA polymerase III subunits POLR3G and POLR3K consistently showed increased relative insolubility across all three cell lines (Fig. 5I). NACC1/2, TTLL5, and ZNF219 also became less soluble following ML-792 treatment. RNA polymerase III is assembled in the cytoplasm, with part of it remaining cytoplasmic for roles in innate immunity and the rest imported into the nucleus56. The nuclear import and function of RNA polymerase III are known to be regulated by SUMOylation57,58. Our findings support that SUMOylation may inhibit nuclear import or chromatin association of POLR3G and POLR3K, leading to the observed changes of their solubility (Fig. 5I). The solubility change of POLR3G was further confirmed by western blot and immunofluorescence (Fig. 5I, K; Supplementary Fig. 2). Notably, POLR3G’s relative insolubility was also increased by MG132 (Fig. 5K), indicating that both ubiquitination and SUMOylation contribute to its regulation.
We also identified significant solubility changes in SP100 and DAXX, two well-characterized transcriptional repressors59. SP100 is a key component of PML nuclear bodies and interacts with chromatin through SUMOylation and HP1 binding60. DAXX, similarly, is recruited to nuclear substructures like PML bodies via SUMO-dependent interactions61. ML-792-induced solubilization of SP100 and DAXX was confirmed by western blot and immunofluorescence (Fig. 5A–C, L; Supplementary Fig. 2), suggesting that SUMOylation is essential for their stable sequestration into nuclear bodies.
In addition to those described above, our findings uncovered ML-792–induced solubility changes of numerous other transcription-associated proteins that have not previously been linked to SUMOylation, suggesting additional SUMO-dependent regulatory mechanisms.
Common targets of MG132 and ML-792 suggest overlap between ubiquitination and SUMOylation pathways
The analyses above identified numerous proteins whose solubility is regulated by either MG132 or ML-792 alone. Interestingly, we also observed a subset of proteins whose solubility was affected by both treatments which may deepen our understanding of the biological interplay between ubiquitination and SUMOylation.
We checked protein targets that are co-regulated by both MG132 and ML-792 (Fig. 6A–C). The majority of overlapping targets exhibited the same directional effect (solubilization or insolubilization) by both drugs (Fig. 6A). For example, in U2OS cells, MG132 and ML-792 decreased the solubility of NAB2 and RARS2 while increasing the solubility of UVRAG and GUK1. In HeLa cells, MG132 and ML-792 insolubilized NHLRC2 and solubilized CERS5. In HEK293A cells, both drugs insolubilized FBXO22 and EPHB2 and solubilized PAN2. Exceptionally, several proteins exhibited opposing effects; in HeLa cells, USP3 was solubilized by MG132 but insolubilized by ML-792 (Fig. 6A). UBXN7 in U2OS cells and LIFR and TSEN34 in HeLa cells were insolubilized by MG132 but solubilized by ML-792.
A Scatter plots showing the proteins modulated by sole treatment of both MG132 (x-axis) and ML-792 (y-axis) from three cancer cell lines. Only proteins with p-values below 0.05 are displayed. B Scatter plots showing the proteins modulated by sole treatment of MG132 (x-axis) and subsequent treatment of ML-792 (y-axis) from three cell lines. Only proteins with p-values below 0.05 are displayed. (C) Scatter plots showing the proteins modulated by sole treatment of ML-792 (x-axis) and subsequent treatment of MG-132 (y-axis) from three cell lines. Only proteins with p-values below 0.05 are displayed. D–G Western blotting results of representative proteins that modulated by MG132 or ML792, including NAB2, RAD54L2, SMAD2, and RB1. Experiments were repeated at least three times, and similar results were obtained. Source data are provided as a Source Data file.
NAB2 is essential for the nuclear export of mRNA62. Previous studies have also shown that NAB2 forms RNA-containing condensate-like structures in the nucleus under glucose stress62. The state of NAB2 condensation influences the extent of nuclear mRNA accumulation and can be recapitulated in vitro, where NAB2 forms RNA-containing liquid droplets. Our observation of substantially decreased NAB2 solubility with both MG132 and ML-792 treatments was validated by western blotting and immunofluorescence microscopy (Fig. 6A, D, and Supplementary Fig. 2), suggesting that both ubiquitination and SUMOylation can independently promote the formation of nuclear NAB2 condensates.
Prompted by the drug-induced solubility shift and relocalization of RAD54L2 reported previously, we next examined proteins exhibiting changes upon sequential treatment with MG132 and ML-792 (Fig. 6B, C)19. As shown, our MS results confirmed that RAD54L2 solubility decreased after initial MG132 treatment and increased following subsequent ML-792 treatment in U2OS cells (Fig. 6B), which is consistent with our previously reported results19 and the current western blotting result (Fig. 6E). Several other proteins, like SMAD2 and ARFIP1, exhibited effects similar to those observed with RAD54L2 in U2OS cells, with SMAD2 exhibiting solubility shift by the drug combination in both U2OS and HEK293A cells. Similar effects were observed with RB1, FBXO3, ASB8, WWC2, and RTCA in HeLa cells, and with FBXW8 in HEK293A cells (Fig. 6B). In contrast, some proteins underwent solubilization following initial MG132 treatment and insolubilization upon subsequent ML-792 treatment--SLC25A25 (U2OS), USP3 (HeLa), and RARS2 (HEK293A). Changing the sequence of drug treatment, several targets exhibited insolubilization following initial ML-792-treatment and solubilization upon subsequent MG132 treatment—BTF3L4 (U2OS), USP3 (HeLa), ZNF143 (HeLa), ATP6AP2 (HEK293A), and EPHB2 (HEK293A) (Fig. 6C). With that same sequence, several targets exhibited solubilization following initial ML-792 treatment and insolubilization upon subsequent MG132 treatment—KATNA1 (U2OS), NFU1 (HeLa), AEBP2 (HEK293A), and RANGRF (HEK293A).
SMAD2 is a key intracellular mediator of TGF-β signaling pathway, which is known to induce its nuclear translocation63,64. Our observation of SMAD2 solubility shift following sequential treatment with MG132 and ML-792 implies potential effects of ubiquitination and SUMOylation in mediating nucleoplasm shuttling of SMAD2 (Fig. 6B). The solubility shift of SMAD2 was confirmed by western blotting (Fig. 6F). ARFIP1 is a member of the BAR domain-containing protein family involved in sensing and generating membrane curvature, and it is recruited by ARF to Golgi membranes65. The observed solubility shifts in ARFIP1 following combination MG132/ML-792 treatment prompts the hypothesis that ubiquitination and SUMOylation influence the subcellular localization and membrane-binding properties of ARFIP1 (Fig. 6B). RB1 is primarily localized to the nucleus and is chromatin-associated66. Changes in cell cycle progression alter RB1 affinity for chromatin, modulating its role in transcriptional regulation. Previous studies have shown that MG132 decreases the levels of unphosphorylated RB1, leading to sequestration of E2FA and inhibition of its transcriptional activity. SUMOylation of RB1 enhances its interaction with CDK2 and promotes its phosphorylation during early G1 phase. Our observed solubility shifts of RB1 in HeLa cells induced by MG132 and ML-792 suggest that ubiquitination and SUMOylation co-regulate the chromatin affinity of RB1, likely through changes in its phosphorylation66 (Fig. 6B). Interestingly, both MS and western blotting showed cell line-specific solubility shifts of RB1 (Fig. 6B, G).
Discussion
MS-based proteomics is widely employed to monitor protein abundance, map post-translational modifications, and investigate protein–protein interactions. In this study, we sought to extend its use to the systematic profiling of protein solubility—a versatile readout of protein state and changes in cellular organization. To enhance the sensitivity of this approach, we evaluated the effect of two supercharging mobile phase modifiers, DMSO and mNBA, in nDIA-MS analysis. We optimized a method incorporating 3% DMSO in the MS workflow that enabled identification of ~9600 proteins in 15 min. Using the optimized approach, we quantified solubility changes in 8694 proteins in response to short-term treatment with MG132 and ML-792.
Notably, this workflow uses a restricted 495–745 Th precursor mass range, which was selected based on preliminary optimization to maximize protein identifications; however, it may exclude peptides outside this window, representing a trade-off between peptide coverage and protein-level sensitivity (Supplementary Fig. 3 and Supplementary Data 5). Narrowing the mass range progressively restricted peptide length distributions without affecting charge states or protein molecular weights (Supplementary Fig. 4). Comparative analyses across multiple mass ranges and isolation windows showed broader ranges favor peptide identifications, while 495–745 Th with a 1 Th window maximized protein identifications (Supplementary Fig. 5). Protein loading experiments (100–1 μg) indicated that identifications increased with input, and the 1 Th window mitigated overloading effects at higher loads (Supplementary Fig. 6).
As mNBA has been reported to influence peptide retention in the mobile phase67, we examined its impact on chromatographic behavior. Both DMSO and mNBA preserved chromatographic performance, with stable peak shapes, no systematic retention shifts, and only minor, localized differences.
To further assess the reliability of data quality, we performed an empirical validation using a two-species entrapment analysis68. By combining human and maize protein databases while excluding shared peptides, this approach enabled direct estimation of false-positive assignments. Since all samples contained only human proteins, maize identifications reflected empirical FDR. Across both benchmark HeLa digests and solubility-proteomics datasets, the empirical FDRs were approximately 1.0–1.3% at the protein level and 0.15–0.2% at the peptide level (Supplementary Fig. 7). This confirms the robustness of data processing and supports the reliability of the proteomic identifications underlying the solubility profiling analyses.
Our work builds significantly upon previously reported use of supercharging reagents27,28,29. The primary contribution of this work lies in the systematic optimization of nDIA-MS workflows and the generation of a comprehensive proteome-scale dataset, rather than the characterization of biological mechanisms. Key innovations of this study are as follows: 1) We tested the use of two supercharging reagents (DMSO and mNBA) and found that DMSO significantly enhances MS signals in nDIA-MS. Protein identification was optimal with 3% DMSO, and peptide identification was optimal with 1% DMSO. DMSO did not alter the overall charge state of peptides, suggesting that signal enhancement was due to mechanisms other than charge state coalescence. mNBA notably increased peptide identification, particularly the identification of peptides with 3+ and 4+ charge states, suggesting the potential value of mNBA in peptide-level studies, such as immunopeptidomics and post-translational modification (PTM) analyses. DMSO and m-NBA enhance nDIA-MS performance through complementary mechanisms: DMSO increases overall signal intensity, likely by improving desolvation and protonation, whereas m-NBA acts as a supercharging reagent, increasing peptide charge states and fragmentation efficiency27,28,69,70,71,72. Their combination further improves peptide coverage, allowing flexible optimization of protein and peptide detection depending on experimental goals. These findings highlight how mobile phase modifiers can be tailored to enhance sensitivity and depth in high-throughput proteomics. 2) The optimized nDIA-MS method is ultra-fast, requiring only 15 min per sample to achieve identification of ~10,000 proteins in a single experiment. 3) The method combines “Triton pre-extraction” of samples with nDIA-MS to enable examination of solubility shifts across a broad range of proteins throughout the cell. 4) The resulting proteome-wide map of solubility remodeling provides an additional layer of information and insights to complement data on protein expression, chemical modifications, and protein-protein interactions. Specifically, we identified 294 proteins exhibiting drug-induced solubility shifts across at least two cell lines. Among these, 54 are previously known to be associated with ubiquitination, proteasome inhibition, or SUMOylation, while the remaining 240 proteins are not previously linked to these pathways (Supplementary Data 4). The limited overlap in solubility changes across cell lines reflects a combination of factors, including our study design with short-term drug treatment to preserve native proteome states, and the fact that MG132 primarily stabilizes proteins through proteasome inhibition. Therefore, observed down-regulation arises from indirect or cell-specific secondary responses, which are expected to be variable. Differences in growth conditions, inhibitor exposure, and confluency further contribute to this variability. Furthermore, intrinsic differences in baseline proteomes and cellular responses across these biologically distinct cell lines likely influence the observed outcomes, highlighting that both experimental design and biological context shape drug-induced proteome remodeling. 5) The combination of MS, western blotting, and immunofluorescence microscopy offers a powerful approach for identifying and validating unknown protein phase transitions, such as chromatin binding, nucleocytoplasmic shuttling, RNA-binding alterations, membrane binding, condensation, and changes in subcellular localization, setting the stage for mechanistic elucidation of uncharacterized cellular functions.
To further contextualize our solubility-profiling approach, we compared our dataset with results from Thermal Proteome Profiling (TPP), an established method for probing protein stability and target engagement. Using both full-curve TPP and two-temperature (2T-TPP) analyses73, we identified proteins whose thermal stability was altered by MG132 or ML-792 treatment. Overlap analysis revealed a partial intersection with solubility-profiling hits (40 and 23 proteins for MG132 and ML-792, respectively), with functional clustering showing enrichment in mitosis-related processes for MG132 and kinases/proteases for ML-792 (Supplementary Fig. 8 and Supplementary Data 6). These findings highlight that solubility profiling and TPP capture complementary biophysical properties of proteins—solubility changes versus thermal stability—emphasizing that our method provides unique and orthogonal insights into drug-induced proteome dynamics.
Overall, our findings reveal widespread, drug-induced protein solubility shifts and highlight solubility proteomic profiling by nDIA-MS as a broadly applicable approach for unveiling protein state transitions and cellular responses to perturbation.
Methods
Cells and cell culture
HeLa cells were purchased from the American Type Culture Collection (ATCC) (Manassas, VA) and maintained in Dulbecco’s modified Eagle’s medium containing 10% fetal calf serum at 37 °C with 5% CO2. HEK293A cells were purchased from Thermo Fisher Scientific (R70507) and maintained in Dulbecco’s modified Eagle’s medium containing 10% fetal calf serum at 37 °C with 5% CO2. U2OS cells were purchased from the ATCC (Manassas, VA) and maintained in McCoy’s 5 A (Modified) medium containing 10% fetal calf serum at 37 °C with 5% CO2.
Chemicals, reagents, and antibodies
The chemicals used in this study included MG132 (S2619; Selleck Chemicals), ML-792 (HY-108702; MedChem Express), Water (10509404; Thermo Fisher Scientific, LC–MS grade, Optima), acetonitrile (10001334; Thermo Fisher Scientific, LC–MS grade, Optima), formic acid (13454279; Thermo Scientific Pierce, LC–MS grade), m-Nitrobenzyl alcohol (73148; Sigma-Aldrich), HeLa protein digest standard (88328; Thermo Scientific Pierce), and sequence grade trypsin (V5113, Promega).
Antibodies used in this study included SUMO-2/3 (4971S; Cell Signaling Technology), SMAD2 (3103 T; Cell Signaling Technology), RB1 (9309 T; Cell Signaling Technology), EGR1 (4153 T; Cell Signaling Technology), NAB2 (sc-23867, Santa Cruz Biotechnology), SP100 (GTX131569, GeneTex), DAXX (4533 T; Cell Signaling Technology), c-Fos (2250 T; Cell Signaling Technology), HSF1 (4356 T; Cell Signaling Technology), CYR61 (14479 T; Cell Signaling Technology), CTGF (86641 T; Cell Signaling Technology), BAG6 (8523S; Cell Signaling Technology), UBL4A (14253-1-AP, Proteintech), TEX264 (NBP1-89866, Novus Biologicals), POLR3G (24701-1-AP, Proteintech), HIF1A (14179 T; Cell Signaling Technology), ATR (2790S; Cell Signaling Technology), Ubiquitin (58295S; Cell Signaling Technology), REV1 (sc-393022, Santa Cruz).
Western blotting
For whole cell extract (WCE) preparation, cells were washed with PBS and resuspended in 1× Laemmli buffer for lysis. The lysates were boiled at 95 °C for 10 minutes and analyzed by Western blotting. For pre-extraction samples, cells were permeabilized with 0.2% Triton X-100 in PBS for 10 min in a cold room, followed by centrifugation at 16,800 × g for 10 min at 4 °C. The resulting pellet was washed once with ice-cold PBS and then resuspended in 1× Laemmli buffer. The resuspended pellet was boiled at 95 °C for 10 min and analyzed by Western blotting.
Immunofluorescence staining
For immunofluorescence staining of the pre-extraction samples, cells were cultured on Falcon 8 well culture slide (354118; Corning). After indicated chemical treatment, cells were permeabilized with 0.2% Triton X-100/PBS for 2 min on ice, followed by fixation with 4% paraformaldehyde for 20 min at room temperature. Cells were then permeabilized with 0.5% Triton X-100/PBS for 10 min and blocked with 4% BSA/PBS for 1 h at room temperature. Diluted primary antibodies in 4% BSA/PBS were then added and incubated at 4 °C overnight. At the following day, cells were then incubated with Alexa Fluor Plus 488 goat anti-mouse secondary antibody (A32723; Thermo Fisher Scientific; 1:1000 dilution) or Alexa Fluor Plus 488 goat anti-rabbit secondary antibody (A32731; Thermo Fisher Scientific; 1:1000 dilution) for 1 hour at room temperature. Slides were mounted with Prolong Diamond antifade mountant with 4′,6-diamidino-2-phenylindole (P36971; Thermo Fisher Scientific). Images were acquired on a Zeiss LSM880 confocal microscope.
MS sample preparation
For the WCE samples, 5 × 10⁶ cells from each sample were washed twice with 1x PBS, then centrifuged at 150 × g for 3 min at 4 °C. The cell pellets were collected for further processing. For insoluble pellet samples, cells were permeabilized with 0.2% Triton X-100 in PBS for 10 min in a cold room, followed by centrifugation at 16,800 × g for 10 min at 4 °C. The resulting pellet was washed once with ice-cold PBS and then collected for further analysis. The collected pellets were resuspended in ice-cold extraction buffer containing 4 M urea and 50 mM NH₄HCO₃. The samples were then sonicated using a microtip sonicator at 35 W for 2 × 10 s pulses. Homogenates were clarified by centrifugation at 10,000 × g for 10 min at 4 °C. Protein concentrations were determined using the BCA assay. Proteins in the supernatant were denatured by boiling at 95 °C for 5 min. For mass spectrometry (MS) sample preparation, 100 µg of each protein sample was diluted to a final volume of 100 µL with 50 mM NH₄HCO₃. The samples were then reduced with 5 mM DTT at 37 °C for 1 h, followed by alkylation with 15 mM iodoacetamide at room temperature in the dark for 30 min. The reaction was quenched with an additional 15 mM DTT. Proteolytic digestion was performed by adding 5 µL of 400 ng/µL trypsin (Promega) and incubating the samples at 37 °C overnight. The samples were acidified with 1 µL of 10% formic acid (final concentration ~0.1% v/v) and centrifuged at 10,000 × g for 10 min at 4 °C. The supernatant was desalted using a BioPureSPN Mini PROTO 300 C18 column (The Nest Group, Cat. No. HUM S18V), dried in a vacuum, and stored at -80 °C until further analysis.
LC–MS/MS data acquisition
LC–MS/MS analysis was performed using an Orbitrap Astral MS coupled with a Vanquish Neo UHPLC system (Thermo Fisher Scientific). Vacuum dried peptide samples were resuspended with 0.1% of formic acid. Peptides were separated using a C18 column (CoAnn Technologies, Cat. No. HEB07502001718I, 75 μm × 20 cm) at a flow rate of 400 nL/min. Peptides were chromatographically separated using a linear gradient of solvent B (0.1% formic acid in ACN) and solvent A (0.1% formic acid in water) unless otherwise specified. Supercharging reagents (DMSO or m-NBA) were added to both mobile phase A and mobile phase B at the indicated concentrations (DMSO: 1% to 7%; m-NBA: 0.001% to 0.1%, v/v). Mobile phases were prepared immediately prior to each analytical run to ensure consistent reagent delivery throughout the analysis. Linear gradients were as follows: from 2% to 8% of solvent B 1.5 min, 8% to 38% of solvent B from 1.6 to 12.6 min, 38% to 100% of solvent B from 12.6 to 13 min, 100% B from 13 to 15 min. In the optimized nDIA method, MS1 spectra were collected in the Orbitrap every 0.6 s at a resolution of 240,000. The full scan range was 495-745 m/z unless specified otherwise. The MS1 normalized AGC target was set to 250% with a maximum injection time of 50 ms. DIA MS2 scans were acquired in the Astral analyzer over a range of 495–745 m/z with a normalized AGC target of 1000% and a maximum injection time of 2.5 ms and an HCD collision energy setting of 25%. Window placement optimization was turned on. The isolation window was set at 1 Th without window overlap.
MS data analysis
Raw files from DIA experiments were analyzed in DIA-NN 1.8.1. The in-silico spectral library was predicted from a human reference database (UniProt 2024 release, 20,598 entries) allowing N-term M excision and 1 missed cleavage. The DIA-NN search included the following settings: Protein inference = ‘Genes’, Neural network classifier = ‘Single-pass mode’, Quantification strategy = ‘Robust LC (high precision)’, Cross-run normalization = ‘RT-dependent’, Library Generation = ‘Smart Profiling’ and Speed and RAM usage = ‘Optimal results’. Mass accuracy and MS1 accuracy were set to 0 for automatic inference. ‘No share spectra’, ‘Heuristic protein inference’ and ‘MBR’ were checked. False discovery rates (FDRs) were controlled within DIA-NN using a target–decoy competition (TDC) strategy. The output results from DIA-NN were filtered with ≤ 1% FDR at both the peptide and protein levels (Q.Value ≤ 0.01 for precursors and PG.Q.Value ≤ 0.01 for protein groups). All thresholds were applied using the DIA-NN R package (https://github.com/vdemichev/diann-rpackage) to ensure transparent and reproducible data processing. All identified proteins had at least one unique peptide. Protein quantification was carried out using the MaxLFQ algorithm implemented in the DIA-NN R package.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are available in the main text or the supplementary materials. Source Data are provided with this paper. The mass spectrometry proteomics data for supercharging reagents evaluation and solubility proteomics have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD064180 (supercharging reagents evaluation) and PXD064185 (solubility proteomics). The mass spectrometry data for preliminary instrument parameter optimization, evaluation, and Thermal Proteome Profiling (TPP) have been deposited to the MassIVE repository (https://massive.ucsd.edu) with the dataset identifier MSV000099787 (PXD070462) [https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID = PXD070462] (Thermal Proteome Profiling), MSV000099789 (PXD070463) [https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID = PXD070463] (instrument parameter optimizatiom), MSV000099791 (PXD070464) [https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID = PXD070464] (instrument parameter evaluation). Source data are provided with this paper.
References
Angel, T. E. et al. Mass spectrometry-based proteomics: existing capabilities and future directions. Chem. Soc. Rev. 41, 3912–3928 (2012).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Ortega, J. A. et al. Nucleocytoplasmic Proteomic Analysis Uncovers eRF1 and Nonsense-Mediated Decay as Modifiers of ALS/FTD Toxicity. Neuron 106, 90 (2020).
Molzahn, C. et al. Shift of the insoluble content of the proteome in the aging mouse brain. Proc. Natl. Acad. Sci. USA 120, e2310057120 (2023).
Thibaudeau, T. A. & Smith, D. M. A Practical Review of Proteasome Pharmacology. Pharm. Rev. 71, 170–197 (2019).
Han, Y. H., Moon, H. J., You, B. R. & Park, W. H. The effect of MG132, a proteasome inhibitor on HeLa cells in relation to cell growth, reactive oxygen species and GSH. Oncol. Rep. 22, 215–221 (2009).
He, X. Y. et al. Probing the roles of SUMOylation in cancer cell biology by using a selective SAE inhibitor. Nat. Chem. Biol. 13, 1164 (2017).
Gu, Y. R. et al. The emerging roles of SUMOylation in the tumor microenvironment and therapeutic implications. Exp. Hematol. Oncol. 12, 58 (2023).
Wilkinson, K. A., Nakamura, Y. & Henley, J. M. Targets and consequences of protein SUMOylation in neurons. Brain Res Rev. 64, 195–212 (2010).
Gutierrez-Morton, E. & Wang, Y. The role of SUMOylation in biomolecular condensate dynamics and protein localization. Cell Insight 3, 100199 (2024).
Cwilichowska, N., Swiderska, K. W., Dobrzyn, A., Drag, M. & Poreba, M. Diagnostic and therapeutic potential of protease inhibition. Mol. Asp. Med 88, 101144 (2022).
Osei-Amponsa, V. & Walters, K. J. Proteasome substrate receptors and their therapeutic potential. Trends Biochem Sci. 47, 950–964 (2022).
Liu, C., Choe, V. & Rao, H. Genome-wide approaches to systematically identify substrates of the ubiquitin-proteasome pathway. Trends Biotechnol. 28, 461–467 (2010).
Claessens, L. A. & Vertegaal, A. C. O. SUMO proteases: from cellular functions to disease. Trends Cell Biol. 34, 901–912 (2024).
Da Silva-Ferrada, E., Lopitz-Otsoa, F., Lang, V., Rodriguez, M. S. & Matthiesen, R. Strategies to Identify Recognition Signals and Targets of SUMOylation. Biochem Res Int 2012, 875148 (2012).
Svozil, J., Hirsch-Hoffmann, M., Dudler, R., Gruissem, W. & Baerenfaller, K. Protein abundance changes and ubiquitylation targets identified after inhibition of the proteasome with syringolin A. Mol. Cell Proteom. 13, 1523–1536 (2014).
Lohraseb, I. et al. Global ubiquitinome profiling identifies NEDD4 as a regulator of Profilin 1 and actin remodelling in neural crest cells. Nat. Commun. 13, 2018 (2022).
Tatham, M. H., Matic, I., Mann, M. & Hay, R. T. Comparative Proteomic Analysis Identifies a Role for SUMO in Protein Quality Control. Sci. Signal 4, rs4 (2011).
Zhang, H. M. et al. RAD54L2-mediated DNA damage avoidance pathway specifically preserves genome integrity in response to topoisomerase 2 poisons. Sci. Adv. 9, eadi6681 (2023).
van der Zanden, S. Y., Jongsma, M. L. M., Neefjes, A. C. M., Berlin, I. & Neefjes, J. Maintaining soluble protein homeostasis between nuclear and cytoplasmic compartments across mitosis. Trends Cell Biol. 33, 18–29 (2023).
van Tartwijk, F. W. & Kaminski, C. F. Protein Condensation, Cellular Organization, and Spatiotemporal Regulation of Cytoplasmic Properties. Adv. Biol. (Weinh.) 6, e2101328 (2022).
Rawlings, A. E. Membrane proteins: always an insoluble problem?. Biochem Soc. T 44, 790–795 (2016).
Sui, X. et al. Widespread remodeling of proteome solubility in response to different protein homeostasis stresses. Proc. Natl. Acad. Sci. USA 117, 2422–2431 (2020).
Grossmann, L. & McClements, D. J. Current insights into protein solubility: A review of its importance for alternative proteins. Food Hydrocolloid 137, 108416 (2023).
Franzmann, T. M. & Alberti, S. Protein Phase Separation as a Stress Survival Strategy. Cold Spring Harb. Perspect. Med. 9, a034058 (2019).
Sui, X. J., Cox, D., Nie, S., Reid, G. E. & Hatters, D. M. A Census of Hsp70-Mediated Proteome Solubility Changes upon Recovery from Heat Stress. J. Proteome Res 21, 1251–1261 (2022).
Hahne, H. et al. DMSO enhances electrospray response, boosting sensitivity of proteomic experiments. Nat. Methods 10, 989–991 (2013).
Strzelecka, D., Holman, S. W. & Eyers, C. E. Evaluation of dimethyl sulfoxide (DMSO) as a mobile phase additive during top 3 label-free quantitative proteomics. Int J. Mass Spectrom. 391, 157–160 (2015).
Iavarone, A. T. & Williams, E. R. Supercharging in electrospray ionization: effects on signal and charge. Int J. Mass Spectrom. 219, 63–72 (2002).
Abaye, D. A., Agbo, I. A. & Nielsen, B. V. Current perspectives on supercharging reagents in electrospray ionization mass spectrometry. Rsc Adv. 11, 20355–20369 (2021).
Guzman, U. H. et al. Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition. Nat. Biotechnol. 42, 1855–1866 (2024).
Lou, R. H. & Shui, W. Q. Acquisition and Analysis of DIA-Based Proteomic Data: A Comprehensive Survey in 2023. Mol. Cell Proteomics 23, 100712 (2024).
Meyer, J. G. & Komives, E. A. Charge State Coalescence During Electrospray Ionization Improves Peptide Identification by Tandem Mass Spectrometry. J. Am. Soc. Mass Spectr. 23, 1390–1399 (2012).
Van Wanseele, Y. et al. Assessing mixtures of supercharging agents to increase the abundance of a specific charge state of Neuromedin U. Talanta 198, 206–214 (2019).
Gallo, F. T., Katche, C., Morici, J. F., Medina, J. H. & Weisstaub, N. V. Immediate Early Genes, Memory and Psychiatric Disorders: Focus on c-Fos, Egr1 and Arc. Front Behav. Neurosci. 12, 79 (2018).
Bahrami, S. & Drablos, F. Gene regulation in the immediate-early response process. Adv. Biol. Regul. 62, 37–49 (2016).
Jia, Q. A., Xu, B. H., Zhang, Y. Y., Ali, A. & Liao, X. CCN Family Proteins in Cancer: Insight Into Their Structures and Coordination Role in Tumor Microenvironment. Front. Genet. 12, 649387 (2021).
Yeger, H. & Perbal, B. CCN family of proteins: critical modulators of the tumor cell microenvironment. J. Cell Commun. Signal 10, 229–240 (2016).
Kular, L., Pakradouni, J., Kitabgi, P., Laurent, M. & Martinerie, C. The CCN family: a new class of inflammation modulators?. Biochimie 93, 377–388 (2011).
Alao, J. P., Stavropoulou, A. V., Lam, E. W., Coombes, R. C. & Vigushin, D. M. Histone deacetylase inhibitor, trichostatin A induces ubiquitin-dependent cyclin D1 degradation in MCF-7 breast cancer cells. Mol. Cancer 5, 8 (2006).
Krenciute, G. et al. Nuclear BAG6-UBL4A-GET4 Complex Mediates DNA Damage Signaling and Cell Death. J. Biol. Chem. 288, 20547–20557 (2013).
O’Leary, C. E. et al. Ndfip-mediated degradation of Jak1 tunes cytokine signalling to limit expansion of CD4+ effector T cells. Nat. Commun. 7, 11226 (2016).
Glimcher, L. H., Lee, A. H. & Iwakoshi, N. N. XBP-1 and the unfolded protein response (UPR). Nat. Immunol. 21, 963–965 (2020).
Locke, M., Tinsley, C. L., Benson, M. A. & Blake, D. J. TRIM32 is an E3 ubiquitin ligase for dysbindin. Hum. Mol. Genet. 18, 2344–2358 (2009).
Yan, Q. S., Ding, J. Y., Khan, S. J., Lawton, L. N. & Shipp, M. A. DTX3L E3 ligase targets p53 for degradation at poly ADP-ribose polymerase-associated DNA damage sites. iScience 26, 106444 (2024).
Gao, J. et al. The CUL4-DDB1 ubiquitin ligase complex controls adult and embryonic stem cell differentiation and homeostasis. Elife 4, e07539 (2015).
Raffeiner, M., Zhu, S. S., González-Fuente, M. & Üstün, S. Interplay between autophagy and proteasome during protein turnover. Trends Plant Sci. 28, 698–714 (2023).
Nakatogawa, H. Mechanisms governing autophagosome biogenesis. Nat. Rev. Mol. Cell Bio 21, 439–458 (2020).
Melia, T. J., Lystad, A. H. & Simonsen, A. Autophagosome biogenesis: From membrane growth to closure. J. Cell Biol. 219, e202002085 (2020).
Fielden, J., Popovic, M. & Ramadan, K. TEX264 at the intersection of autophagy and DNA repair. Autophagy 18, 40–49 (2022).
Lascaux, P. et al. TEX264 drives selective autophagy of DNA lesions to promote DNA repair and cell survival. Cell 187 (2024).
Delorme-Axford, E., Popelka, H. & Klionsky, D. J. TEX264 is a major receptor for mammalian reticulophagy. Autophagy 15, 1677–1681 (2019).
Boada-Romero, E. et al. TMEM59 defines a novel ATG16L1-binding motif that promotes local activation of LC3. Embo J. 32, 566–582 (2013).
Gaglia, G. et al. HSF1 phase transition mediates stress adaptation and cell fate decisions. Nat. Cell Biol. 22, 151 (2020).
Hammond-Martel, I., Yu, H. & Affar, E. B. Roles of ubiquitin signaling in transcription regulation. Cell Signal 24, 410–421 (2012).
Tian, K., Wang, R., Huang, J., Wang, H. & Ji, X. Subcellular localization shapes the fate of RNA polymerase III. Cell Rep. 42, 112941 (2023).
Kessler, A. C. & Maraia, R. J. The nuclear and cytoplasmic activities of RNA polymerase III, and an evolving transcriptome for surveillance. Nucleic Acids Res. 49, 12017–12034 (2021).
Wang, Z., Wu, C., Aslanian, A., Yates, J. R. & Hunter, T. Defective RNA polymerase III is negatively regulated by the SUMO-Ubiquitin-Cdc48 pathway. Elife 7, e35447 (2018).
Li, H. et al. Sequestration and inhibition of Daxx-mediated transcriptional repression by PML. Mol. Cell Biol. 20, 1784–1796 (2000).
Sternsdorf, T., Jensen, K., Reich, B. & Will, H. The nuclear dot protein sp100, characterization of domains necessary for dimerization, subcellular localization, and modification by small ubiquitin-like modifiers. J. Biol. Chem. 274, 12555–12566 (1999).
Shih, H. M., Chang, C. C., Kuo, H. Y. & Lin, D. Y. Daxx mediates SUMO-dependent transcriptional control and subnuclear compartmentalization. Biochem Soc. Trans. 35, 1397–1400 (2007).
Heinrich, S. et al. Glucose stress causes mRNA retention in nuclear Nab2 condensates. Cell Rep. 43, 113593 (2024).
Nakao, A. et al. TGF-beta receptor-mediated signalling through Smad2, Smad3 and Smad4. EMBO J. 16, 5353–5362 (1997).
Fink, S. P., Mikkola, D., Willson, J. K. & Markowitz, S. TGF-beta-induced nuclear localization of Smad2 and Smad3 in Smad4 null cancer cell lines. Oncogene 22, 1317–1323 (2003).
Kanoh, H., Williger, B. T. & Exton, J. H. Arfaptin 1, a putative cytosolic target protein of ADP-ribosylation factor, is recruited to Golgi membranes. J. Biol. Chem. 272, 5421–5429 (1997).
Sanidas, I., Lawrence, M. S. & Dyson, N. J. Patterns in the tapestry of chromatin-bound RB. Trends Cell Biol. 34, 288–298 (2024).
Lin, C. W. et al. m-nitrobenzyl alcohol supercharging reagent enhances the chromatographic separation and the charging of disulfide bond linked and His-tag peptides. J. Chromatogr. A 1722, 464828 (2024).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Sterling, H. J., Prell, J. S., Cassou, C. A. & Williams, E. R. Protein conformation and supercharging with DMSO from aqueous solution. J. Am. Soc. Mass Spectrom. 22, 1178–1186 (2011).
Sterling, H. J. & Williams, E. R. Origin of supercharging in electrospray ionization of noncovalent complexes from aqueous solution. J. Am. Soc. Mass Spectrom. 20, 1933–1943 (2009).
Iavarone, A. T. & Williams, E. R. Mechanism of charging and supercharging molecules in electrospray ionization. J. Am. Chem. Soc. 125, 2319–2327 (2003).
Going, C. C. & Williams, E. R. Supercharging with m-nitrobenzyl alcohol and propylene carbonate: forming highly charged ions with extended, near-linear conformations. Anal. Chem. 87, 3973–3980 (2015).
Zhang, Y. et al. Thermal proteome profiling reveals fructose-1,6-bisphosphate as a phosphate donor to activate phosphoglycerate mutase 1. Nat. Commun. 15, 8936 (2024).
Acknowledgements
We thank all members of MD Anderson Proteomics Core Facility and Metabolomics Core Facility for their help and constructive discussions. This work was supported by NIH grant number 1S10OD012304-01, NIH/NCI grant number P30CA016672, and the University of Texas MD Anderson Cancer Center. This research was supported in part by the National Cancer Institute of the National Institutes of Health grant number P30CA033572.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.X., H.Z., P.L.L. Methodology: Y.X., and H.Z. Investigation: Y.X., H.Z., L.T. and B.W. Visualization: Y.X., and H.Z. Supervision: P.L.L. and J.N.W. Writing—original draft: Y.X., and H.Z. Writing—review and editing: Y.X., H.Z., P.L.L. and J.N.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, Y., Zhang, H., Tan, L. et al. Supercharging-enhanced nDIA-MS enables global profiling of drug-induced proteome solubility shifts. Nat Commun 17, 2350 (2026). https://doi.org/10.1038/s41467-026-69025-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69025-8