Abstract
Adult stem cells and their niches communicate intricately for tissue maintenance and regeneration. However, effectively coordinating these complex interactions is challenging. Here, we demonstrate that transient dedifferentiation of a fraction of epithelial stem cell progenies orchestrates beneficial changes within the entire skin’s cellular networks to favor repair. We achieved this by inducing a mosaic and reversible expression of reprogramming factors (Oct-4, Sox2, Klf4, and c-Myc) in the mouse epidermis. This in vivo partial epidermal reprogramming not only affected the partially reprogrammed cells, but also their microenvironment, including neighboring epithelial cells and T cells, conferring widespread healing characteristics even in the absence of injury. When a wound was introduced, these collective changes accelerated re-epithelialization in both wild-type and a hyperglycemic mouse disease model. Furthermore, the effects extended to dermal healing, leading to reduced scarring and altered angiogenesis. In conclusion, our work reveals that mosaic partial reprogramming of the epidermis influences various cell types within the skin during homeostasis and repair, leading to enhanced cutaneous wound healing.
Similar content being viewed by others
Introduction
Adult stem cells (ASCs) maintain homeostasis by regulating tissue turnover. They are activated upon injury, the physiological damage that disrupts homeostasis, to heal the tissue1. The injury is either completely regenerated; in others, it is repaired with excessive fibrosis, forming scars—collagen-rich nonfunctional tissue2,3,4; or in some cases, the injury remains unrepaired. Previous studies have explored the regulation and development of injury repair processes and identified several factors that determine regenerative capacity. First, the proliferative ability of ASCs and their progeny is crucial5,6,7. In addition, sophisticated spatial and temporal control of communication between ASCs and their niches is critical, because the healing process involves re-establishment of complex organ structures and functions beyond restoration of cell numbers8,9,10. These niches include mesenchymal cells that modulate the extracellular matrix (ECM) for precise cell proliferation and migration, and immune cells that produce inflammatory responses against damage2,11,12,13.
Despite numerous studies highlighting the significance of stem cell-niche interactions in regeneration, understanding the entire network is challenging due to its spatial and temporal complexity. Additionally, manipulation of the network for optimal outcomes has been difficult, as manipulation of individual factors can have unforeseen consequences. For instance, accelerating repair processes, which should be beneficial, can sometimes lead to detrimental effects such as increased fibrosis or scarring14,15,16. Therefore, effective regulation of stem cell-niche communication requires further investigation.
Cellular reprogramming by Oct-4, Sox2, Klf4, and c-Myc (OSKM), known as the Yamanaka factors, has emerged as a valuable tool for the manipulation of cell fate17,18,19. Originally, OSKM were used only in vitro because their forced expression in vivo leads to teratoma formation due to the production of induced pluripotent stem cells20,21. However, it was found that transient and reversible OSKM expression could induce partial reprogramming, which confers cellular plasticity while retaining the original cell identity and functionality. This process did not induce pluripotency, indicating that in vivo partial reprogramming could prevent teratoma formation22,23,24. Partial reprogramming in mice rejuvenated progeria or physiological aging22,23,25,26,27. Furthermore, recent studies have revealed that in vivo partial reprogramming promotes tissue repair in various organs, such as the muscle, liver, and intestine28,29,30. Another study observed a reduction in fibrosis and scarring after injury15. Even the adult heart and nervous system, which are composed of post-mitotic cells and therefore do not heal well, have shown regeneration via partial reprogramming31,32.
To enhance tissue regeneration, short-term OSKM expression partially dedifferentiates resident somatic cells in a cell-autonomous manner at the cellular level29,30,32. However, the influence of partially reprogrammed cells on their in vivo environment is poorly understood. It is hence unclear whether previously reported improvements in tissue regeneration by partial reprogramming are solely mediated by proliferation and dedifferentiation of somatic cells, or if an interplay between the dedifferentiated cells and the surrounding environment plays a crucial role in the process. Understanding such interplay between partially reprogrammed cells and their neighbors/niches in complex in vivo systems will be essential for possible future practical applications of in vivo partial reprogramming to promote tissue homeostasis and repair.
In this study, we addressed this gap in knowledge by establishing a genetically engineered mouse model that allows partial epidermal reprogramming in a mosaic manner. We transiently induced mosaic OSKM expression, coupled with the fluorescent reporter mCherry, specifically in the epithelial cells of the skin—the body’s primary protective barrier against external threats. Using this model, we demonstrate that various types of skin cells enter an injury-responsive-like state upon partial reprogramming of only a small fraction of epidermal cells. Enhanced cutaneous healing was also observed. We conclude that short-term mosaic OSKM expression in the epidermis is sufficient to coordinate the cellular networks in the skin, directing them to be beneficial for healing, and may provide an avenue for the control of skin homeostasis and repair processes.
Results
In vivo mosaic partial epidermal reprogramming through transient OSKM induction
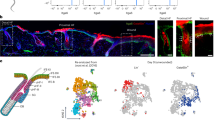
To explore how in vivo partial reprogramming affects the function of skin epithelial stem cells and their progeny during homeostasis and repair, we transiently expressed OSKM in epidermal keratinocytes of mice. We crossed mice carrying the LoxP-STOP-LoxP-rtTA-EGFP and TetO-OSKM-mCherry cassettes33 with a Krt14-CreER transgene that is expressed in the interfollicular epidermal (IFE) cells34,35,36 (Supplementary Fig. 1a–c). In the absence of tamoxifen and doxycycline (Dox), neither EGFP nor mCherry expression was detectable (Supplementary Fig. 1d). After Dox treatment for 3 days, reprogrammable mice (Epi-iOSKM mice) exhibited global GFP and mCherry mosaic expression in the epidermis (Fig. 1a, b; Supplementary Fig. 1b–d) with no histological abnormalities (Supplementary Fig. 1e,f). Mice without the TetO-OSKM-mCherry cassette (control mice) expressed only rtTA-EGFP and not mCherry (Fig. 1b). We also observed the induction of Oct4, Sox2, Klf4, and c-Myc expression in the mCherry+ cells of Epi-iOSKM mice, and quantified that the proportion of Sox2+ cells was 21.91 ± 3.97 % in basal epidermal cells and 25.55 ± 3.43% in suprabasal epidermal cells, confirming effective induction of the reprogramming factors (Fig. 1c; Supplementary Fig. 1g–j). Furthermore, the pluripotency marker Nanog was not detected, indicating that these cells were not fully reprogrammed (Supplementary Fig. 2a). After Dox withdrawal, mCherry was absent (Supplementary Fig. 2b), and no cell death was detected (Supplementary Fig. 2c, d), confirming OSKM was transiently induced only during periods of Dox administration.
a Schematic of epidermal-specific OSKM inducible mouse models. Immunostaining for mCherry with GFP (b) or Sox2 (c) in the control and Epi-iOSKM skin (representative of 5 biological samples). Yellow arrows, mCherry+Sox2+ cells. d UMAP of scRNA-seq from IFE cells. Dashed lines, OSKM-enriched cell types. e Proportion of Basal I, II, Prolif I, II, and Supra I, II cells in IFE cells. Relative expression of Krt10 in IFE cells of each cell type (f), and in Supra IFE cells of each condition (g). Two-tailed Mann-Whitney U test with Bonferroni correction. h Krt10 and mCherry staining in the control and Epi-iOSKM skin with quantifications of Krt10− cells in the suprabasal epidermis. Yellow and gray arrowhead, Krt10− cells in the suprabasal epidermis with and without mCherry expression, respectively. n = 6 from 3 mice/condition. i UMAP with pseudotime and trajectory inference (top) and pseudotime in Supra IFE cells (bottom). Two-tailed Mann-Whitney U test with Benjamini-Hochberg correction. j Proportion of Prolif, Supra, and Basal cells. k Ki67 and mCherry staining in control and Epi-iOSKM skin with quantifications of Ki67+ cells. n = 9 from 3 mice/condition. l–n PAGA graphs showing velocity results in IFE cells. Blue, red, and black arrows illustrate transition patterns from Basal toward Supra cells, from Supra toward Basal/Prolif cells, and others, respectively (l). Bold dashed black arrows illustrate transition patterns from Krt10-low cells to Krt10-high cells (m) and Ki67-high cells to Ki67-low cells (n), while bold black arrows illustrate the reverse transition patterns. Gray and yellow arrows illustrate others (m, n). o Cell-autonomous and non-autonomous effects of mosaic partial epidermal reprogramming. Data are presented as mean ± SEM (one-way ANOVA with Tukey’s and Dunnett’s T3 multiple comparisons test in h, k). Box plots show the median, interquartile range, and 1.5-fold interquartile range whiskers alongside individual outliers in (f, g, i). White dashed lines, the boundary between the epidermis and dermis in (b, c, h, k). Pink arrows, mChneg-enriched cell types (d, e). Scale bars, 50 µm (b, c, h, k).
Mosaic partial reprogramming of epidermal cells induces themselves and their neighbors to undergo lineage-preserved dedifferentiation
Despite recent remarkable findings on partial in vivo reprogramming, the influence of partially reprogrammed cells on their surroundings is unclear. We therefore utilized our mosaic partial reprogramming model to investigate both the effects of partial epidermal reprogramming on neighboring cells, as well as cell-autonomous effects.
Firstly, we performed single-cell RNA sequencing (scRNA-seq) to explore the changes in the molecular and cellular characteristics of skin epithelial cells upon mosaic partial reprogramming. To characterize the respective cellular properties of the OSKM-mCherry-expressing IFE cells (mChpos cells) and neighboring IFE cells without OSKM-mCherry expression (mChneg cells), we sorted mChpos and mChneg epidermal cells from Epi-iOSKM mice (Supplementary Fig. 3a, b). We also collected epidermal cells from control mice and included these in our analysis. To identify the IFE cells, we used uniform manifold approximation and projection (UMAP) plots; this divided the IFE cells into 15 clusters (Supplementary Fig. 3c, d). Based on marker gene expression (Supplementary Fig. 3e), we annotated clusters 0, 1, 3, 4, 5, 6, and 14 as non-proliferative basal cells (Basal), clusters 2, 10, and 13 as proliferating cells (Prolif), and clusters 7, 8, 9, 11, and 12 as non-proliferative suprabasal cells (Supra) (Supplementary Fig. 3f). Interestingly, IFE cells from control mice were rarely found in clusters 2, 3, and 12. However, cells in these clusters were enriched for mChpos populations (Supplementary Fig. 3d). We hence annotated clusters 2, 3, and 12 as Prolif II, Basal II, and Supra II, respectively, potentially describing converted IFE cells upon cell-autonomous OSKM induction, whereas we defined other Prolif, Basal, and Supra cells as Prolif I, Basal I, and Supra I, respectively, representing canonical IFE cells (Fig. 1d).
Surprisingly, we found that the mChneg populations in the Epi-iOSKM epidermis exhibited a higher proportion of Basal II and Supra II cells than control epidermis (Fig. 1d, e). This implies the potential that cell-non-autonomous conversion occurs in cells lacking OSKM expression, causing the mChneg cells to change in a manner similar to mChpos cells. To compare the control-enriched IFE cell types and mChpos and mChneg-enriched IFE cell types, we validated the expression of epidermal differentiation markers. The epidermis comprises a Krt14+ basal layer with epidermal stem/progenitor cells, and a suprabasal layer with Krt10+ differentiated cells (Supplementary Fig. 4a)37. Krt10 expression was lower in Supra II than in Supra I, while the pattern of Krt14 expression showed the opposite trend (Fig. 1f and Supplementary Fig. 4b). Krt10 expression in the suprabasal IFE cells was lower in not only mChpos but also mChneg populations than in the control, demonstrating cell-non-autonomous effects (Fig. 1g). Consistently, immunohistochemical analysis confirmed that short-term OSKM expression generated cells that express less Krt10 in the suprabasal layer in both cell-autonomous and non-autonomous manners (Fig. 1h and Supplementary Fig. 4c). Expression of another suprabasal marker, Dsg1a38, was also downregulated in Epi-iOSKM suprabasal IFE cells in a similar manner (Supplementary Fig. 4d). Together with the reduction of the differentiation markers, partial reprogramming increased the Krt14+ basal layer thickness (Supplementary Fig. 4e). We also comprehensively examined the extent of differentiation in Supra IFE cells using pseudotime computed with Monocle3. The pseudotime values, which are proportional to the expression levels of epidermal differentiation markers (Supplementary Fig. 4f–h), were lower for mChpos/mChneg than controls (Fig. 1i), demonstrating Supra cells in the Epi-iOSKM epidermis exhibit reduced epidermal differentiation characteristics.
We next found that the proportion of Prolif cells is higher in mChpos cells than mChneg cells and control (Fig. 1j). The differentially expressed genes (DEGs) that were only upregulated in mChpos cells—and not in mChneg cells—from the Epi-iOSKM epidermis showed that genes associated with cell division were dramatically upregulated only in autonomously OSKM-expressing cells (Supplementary Fig. 4i). Together, from immunohistochemistry, we observed increased density of Ki67+ cells in the Epi-iOSKM epidermis compared to control (Supplementary Fig. 4j), which was an effect of mChpos cells, not mChneg cells; because the ratio of Ki67+ cells was higher in mCherry-expressing cells than their wildtype neighbors and control epidermal cells (Fig. 1k).
To infer the cell state dynamics of epithelial stem cells and their progeny in the control and Epi-iOSKM epidermis, we performed RNA velocity analysis on our single-cell RNA-seq dataset. In addition to the previously reported dynamics within Basal/Prolif cells and within Supra cells39, control IFE cells exhibited the cell fate transition from Basal to Supra cells, implying epidermal differentiation. In contrast to this, in mChpos and mChneg cells of the Epi-iOSKM epidermis, cell fate transitions from Supra to Basal cells were observed (Fig. 1l and Supplementary Fig. 4k). These transitions, indicative of IFE lineage dedifferentiation, were further supported by gene expression patterns. Cell fate transition trends of mChpos and mChneg cells in Epi-iOSKM epidermis were from high expression of differentiation markers such as Krt10 and Dsg1a to elevated expression of epidermal stem cell markers such as Krt14 and Itga6, reversing the differentiation trends observed in control IFE cells (Fig. 1m and Supplementary Fig. 4l–n). Moreover, only in mChpos cells, the trajectory from Supra to Prolif cells was identified (Fig. 1l). While control and Epi-iOSKM mChneg cells exhibited a trajectory from Mki67-positive to Mki67-negative states, mChpos cells exhibited the reversed pattern (Fig. 1n), indicating induced proliferation in a cell-autonomous manner. Collectively, these observations suggest that mosaic and transient OSKM expression drives IFE cell lineage-preserved partial dedifferentiation in both cell-autonomous and non-autonomous manners, but induces proliferation of IFE cells solely in a cell-autonomous manner (Fig. 1o).
Mosaic partial reprogramming triggers an epidermis-wide repair response in the absence of injury
To investigate how partial reprogramming and particularly its cell-non-autonomous effects influence the properties of Epi-iOSKM epidermis, we performed additional scRNA-seq by collecting all epidermal cells from control and Epi-iOSKM mice without fluorescence sorting. After identifying IFE cells (Supplementary Fig. 5a), we found altered cell subtype composition between control and Epi-iOSKM epidermis (Supplementary Fig. 5b). Notably, genes related to the cellular response to external stimuli were highly upregulated in the Epi-iOSKM epidermis (Supplementary Fig. 5c). Other DEGs—those associated with protein synthesis, programmed cell death, and differentiation—were also related to the response of the tissue against the external stimulant (Supplementary Fig. 5c)40,41. This suggests that short-term OSKM expression recapitulates the response of the epidermis to injury, which disrupts the epidermal barrier and exposes keratinocytes to external stimuli. We then confirmed these main changes of the Epi-iOSKM epidermis in our initial scRNA-seq data, in which mChpos and mChneg cells were separated. Most alterations in the response of cells to external stimuli were confirmed in the pure mChneg and mChpos populations, albeit to different degrees (Fig. 2a).
a Heatmap of the gene module scores (associated with gene sets upregulated in the epidermis of Epi-iOSKM mice) in IFE cells of each condition. Pink boxes, cell-non-autonomous effects. b, c scRNA-seq data from IFE cells of control and Epi-iOSKM mice in our study and reference data from unwounded and wounded IFE cells were integrated. UMAP with unbiased clustering (b), and proportion of each cluster in reference (c, top) and our study (c, bottom). Red boxes, clusters enriched in wounded, mChneg, and mChpos IFE cells (c). UMAP of IFE cells from reference (d, left) and our study (d, right) with annotated cell types, and proportion of each cell type in cluster 6 (e). f UMAP with relative expression of Krt17 (top) and Krt6a (bottom). g Relative expression of Krt17 (top) and Krt6a (bottom) in IFE cells of each condition. Immunostaining for mCherry with Krt17 (h) or Krt6a (i) with MFI quantification in control and Epi-iOSKM epidermis. Data are mean ± SEM (n = 6 from 3 mice/condition, one-way ANOVA with Tukey’s multiple comparisons test). Module score of genes associated with cellular response to hypoxia (j) or cytokine signaling in immune system (k) by REACTOME. Box plots show the median, interquartile range, and 1.5-fold IQR whiskers with outliers (n = 3013 from 3 mice (control), 1612 (mChneg) or 2101 (mChpos) from 6 mice). Immunostaining of HIF-1α (l), phospho-STAT3 (m) and mCherry in skin from control and Epi-iOSKM mice (representative of 3 biological samples). n Diagram illustrating partial epidermal reprogramming conferring healing features and cellular plasticity within epidermal lineages compared to full reprogramming. Bold dashed lines, cluster 6 in integrated UMAP (b, d, f). White dashed lines, boundary between the epidermis and dermis (h, i, l, m). Pink arrows, the cell non-autonomous effects of partial epidermal reprogramming (h, i, l, m). Asterisks, autofluorescence in the skin (l, m). Two-tailed Mann–Whitney U test with Bonferroni correction (g, j, k). Scale bars, 50 µm (h, i, l, m). iPSCs, induced pluripotent stem cells; HF hair follicle, MFI mean fluorescence intensity.
To compare mChneg and mChpos cells with epithelial cells under actual wounding conditions, we integrated our data with a publicly available reference scRNA-seq dataset containing IFE cells from wounded and unwounded skin39 using the anchor-based reciprocal principal components analysis (RPCA) (Fig. 2b). In addition to the cell types annotated in the original study, we identified a cluster of wound-responsive (W.R) cells unique to wounded conditions (Supplementary Fig. 6a). In the integrated map, nine distinct clusters were identified in an unbiased manner (clusters 0–8, Fig. 2b). The trend of proportional changes in each cluster was similar across the two experimental designs: transition from control to mChneg or mChpos populations, and from unwounded to wounded conditions. Cells in clusters 0, 1, 3, and 7 exhibited higher proportions in the control and unwounded epidermis than those in the other conditions, whereas the opposite trend was observed for clusters 2, 4, and 6 (Fig. 2c). Among them, cluster 6 was highly enriched with W.R cells from the reference scRNA-seq data, and Basal II and Supra II from our scRNA-seq data, indicating that the three cell types share common features (Fig. 2d, e). Notably, Krt17, Krt16, and Krt6a, which are markers of active keratinocytes induced in injured skin42, were highly expressed in cluster 6 (Fig. 2f and Supplementary Fig. 6b, c). Their expression was upregulated in mChneg and mChpos cells from the Epi-iOSKM epidermis compared to control epidermis (Fig. 2g and Supplementary Fig. 6d). Krt17 and Krt6a protein expression was also increased in mChpos and mChneg cells of the Epi-iOSKM epidermis (Fig. 2h, i).
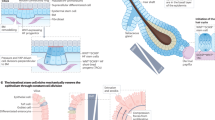
Wound repair progresses in several stages: the early response (including hemostasis and inflammation), proliferation (including re-epithelialization), and remodeling (Supplementary Fig. 6e)2,43. In the early response phase, the skin epidermis recognizes external stressors via hypoxia and inflammatory pathways (Supplementary Fig. 6f)44,45,46. Interestingly, even without external stressors, partial epidermal reprogramming enhanced hypoxia-associated pathways and cytokine-mediated inflammatory pathways in both cell-autonomous and non-autonomous manners (Fig. 2j, k). Consistently, the levels of HIF-1α and phospho-STAT3, hallmarks for hypoxia and inflammatory pathways respectively44,45,47, were increased in partially reprogrammed epidermis (Fig. 2l, m). Our interpretation of these findings is that the conversion of epidermal cells by partial reprogramming recapitulates the early response phase of wound repair, in addition to the following proliferation stage, in which Krt17+ and Krt6+ active keratinocyte function. Collectively, molecular alterations upon partial reprogramming mimic the skin repair processes despite the absence of injury in both cell-autonomous and non-autonomous manners.
Partial reprogramming confers plasticity within epidermal cell lineages
During repair, injury-responsive IFE cells acquire plasticity—the ability of cells to change their properties in response to the environmental cues (Supplementary Fig. 7a). After regeneration, the plastic cells lose their characteristics, such as Krt17/6a expression, and return to the original stable state (Supplementary Fig. 7b–d). Similarly, we confirmed that Epi-iOSKM epidermis acquired increased differentiation potency in comparison to control epidermis (Supplementary Fig. 7e). In addition, after Dox withdrawal for 10 days, Krt17 and Krt6a upregulation by transient OSKM expression was reversed in the Epi-iOSKM epidermis (Supplementary Fig. 7f, g), with the epidermis returning to a homeostatic state (Supplementary Fig. 7h). The fact that conversion under partial reprogramming is reversible and the tissue reacquires specific epidermal properties, as in the repair process, indicates that partial reprogramming imposes the potential only to become epidermal cells, and no other cell types in the in vivo microenvironment, unlike complete reprogramming-conferred plasticity to become multiple kinds of cells (Fig. 2n). The expression of IFE cell markers, and lack of expression of the pluripotent cell marker Nanog in the Epi-iOSKM epidermis after 3 days Dox treatment (Supplementary Figs. 2a and 7i–k) support our partial reprogramming model proposing induced cellular plasticity only within IFE cell lineages.
Cell-autonomous PI3K activation is necessary for the partial epidermal reprogramming
We then aimed to elucidate how partial reprogramming results in injury-responsive-like conversion of IFE cells. Firstly, we focused on the transition of OSKM-expressing mChpos cells. Because increased proliferation was only observed in these, and not in mChneg cells (Fig. 1j, k), we investigated Prolif cells from the scRNA-seq data by analyzing enriched pathways from DEGs that changed only in mChpos cells of the Epi-iOSKM epidermis. We found that the expression of genes involved in signaling pathways regulating pluripotency of stem cells was altered (Fig. 3a). We focused on these genes as candidates to elucidate cell-autonomous effects because induction of pluripotency is a major direct result of prolonged OSKM expression17,18,19, and it can explain the observed increase in proliferation in a cell-autonomous manner. Besides Pou5f1, Sox2, and c-Myc upregulation, which were directly induced as part of OSKM, we observed the downregulation of six genes associated with signaling pathways that induce cell proliferation (Fig. 3b). Of these, only Pik3r1, encoding the phosphoinositide 3-kinase (PI3K) subunit P85α, could account for the activation of the relevant signaling, because P85α inhibits PI3K activity (Fig. 3c)48,49. Pik3r1 gene expression in mChpos cells from the Epi-iOSKM epidermis was downregulated not only in Prolif, but also in Basal and Supra IFE cells (Fig. 3d). Besides Pik3r1, we investigated the expression of other genes that regulate PI3K activity and found that the expression of Shb, which activates PI3K (Fig. 3c)50,51, was upregulated in mChpos IFE cells. The expression of Pik3r1 and Shb was not altered in mChneg cells of the Epi-iOSKM epidermis (Fig. 3d). Next, we examined the changes induced in well-known cellular pathways upon partial reprogramming52 and observed a higher activation of the PI3K pathway in mChpos cells (Fig. 3e). Specifically, unlike other pathways activated in the Epi-iOSKM epidermis, such as the JAK-STAT, epidermal growth factor receptor (EGFR), and MAPK pathways, the PI3K pathway was not affected in mChneg cells (Fig. 3e; Supplementary Fig. 8a). Further, using immunohistochemistry, we observed increased AKT phosphorylation, a hallmark of PI3K pathway activation53, only in mChpos cells of Epi-iOSKM epidermis (Fig. 3f), confirming that PI3K activation was a cell-autonomous effect of OSKM expression. Well-known target genes of FOXO transcription factors, inhibited by the PI3K-AKT pathway, such as a cell cycle-related gene, Ccng2 (ref.54), and a modulator of the negative feedback loop of the pathway, Irs1 (ref.55), were downregulated in mChpos IFE cells, supporting PI3K downstream activation (Supplementary Fig. 8b).
a Enriched KEGG pathway of the DEGs from Prolif cells, upregulated in mChpos but not in mChneg of Epi-iOSKM epidermis compared to control epidermis. Gene set enrichment analysis based on the hypergeometric test (one-sided, Benjamini-Hochberg correction). DEGs are defined as adj p <0.05 and |log2FC| > 1 (two-sided Mann-Whitney U test with Bonferroni correction). b Expression of DEGs involved in signaling pathways regulating pluripotency of stem cells from (a). c Schematic of PI3K subunits and related cellular pathways. The P85α regulatory subunit inhibits the activity of PI3K. SHB adaptor protein increases PI3K activity. d Gene expression of Pik3r1 or Shb. Two-tailed Mann–Whitney U test with Bonferroni correction. e Heatmap of pathway activity in IFE cells of each condition based on PROGENy. f Phospho-AKT and mCherry staining in the control and Epi-iOSKM skin with quantifications of the proportion of phospho-AKT+ cells. n = 6 from 3 mice/condition. Yellow arrows, phospho-AKT+mCherry+ cells. Immunostaining of HIF-1α and mCherry (g), Krt17 and mCherry (h), or Ki67 and mCherry (i) in the control and Epi-iOSKM skin treated by vehicle (left side) or LY294002 (right side). Quantification of Krt17 MFI in the epidermis (h, right) and the proportion of Ki67+ cells (i, right). Representative images from 3 individual biological samples (g). n = 9 in (h) or 12 in (i) from 3 mice/condition. j Diagram showing injury-responsive-like transition of epidermal keratinocytes via cell-autonomous PI3K pathway activation upon partial reprogramming. Data are represented as mean ± SEM (Welch’s ANOVA with Dunnett’s T3 multiple comparisons test in (f), two-way ANOVA with Tukey’s multiple comparisons test in (h), or two-tailed unpaired Student’s t-test in (i)). White dashed lines, the boundary between the epidermis and dermis in (f–i). Asterisks, autofluorescence in the skin in (f). Scale bars, 50 µm (f–i).
Subsequently, we assessed whether the PI3K pathway functions upstream of injury-responsive-like molecular changes induced by transient OSKM activation. The dorsal skin of Epi-iOSKM mice was topically treated with the PI3K inhibitor LY29400256,57. This reversed some effects of the partial epidermal reprogramming: HIF-1α activation as part of the wound-responsive-like response in the epidermis of Epi-iOSKM mice was attenuated upon LY294002 treatment (Fig. 3g and Supplementary Fig. 8c). The upregulation of Krt17 and Krt6a, epithelial cell markers in wound repair, was also reversed (Fig. 3h and Supplementary Fig. 8d). Moreover, an increase in cell proliferation, which is a cell-autonomous effect of partial reprogramming, was attenuated by PI3K inhibition (Fig. 3i).
A previous study reported that the PI3K pathway is activated during wound healing58. We performed immunohistochemistry at several time points during wound healing, and observed increased AKT phosphorylation at 3- and 7-days post-wounding (dpw) (Supplementary Fig. 8e). However, phospho-AKT was not detected until 1 dpw (Supplementary Fig. 8f), corresponding to the beginning of the early response phase of wound repair. In contrast, HIF-1α expression was induced immediately upon wounding within 3 hours (Supplementary Fig. 8g) even in the absence of detectable phospho-AKT (Supplementary Fig. 8f). This suggests that the early response to the wounding via the HIF-1α pathway is independent of PI3K activation (Supplementary Fig. 8h). Thus, we conclude that partial reprogramming via cell-autonomous PI3K activation promotes wound-responsive-like pathways (Fig. 3j) but is distinct from the early response of keratinocytes to wounding.
EGFR signaling propagates the effects of partially reprogrammed cells to the neighbors
We next moved our focus to the conversion of neighboring cells. To test whether the changes in mChneg populations in the Epi-iOSKM epidermis were caused by direct effects of partially reprogrammed keratinocytes, we performed cell-cell interaction (CCI) analysis on our scRNA-seq data from IFE cells. Ligand-receptor interactions between IFE cells were modified by partial reprogramming (Supplementary Fig. 9a). Notably, signaling pathways associated with the ECM, such as FN1 and KLK, those activating EGFR, such as EPGN and EGF, and those promoting EGFR/ErbB3 complex signaling, such as NRG59, were significantly activated in the Epi-iOSKM epidermis (Fig. 4a and Supplementary Fig. 9b).
a Relative information flow for Epi-iOSKM-enriched signaling pathways in control and Epi-iOSKM IFE cells. b Volcano plot showing DEGs in mChpos IFE cells versus controls. Red and blue dots, upregulated and downregulated DEGs. Green dots, upregulated genes termed extracellular space (GO-CC). DEGs are defined as adj p <0.05 and |log2FC | > 1.2 (two-sided Mann-Whitney U test with Bonferroni correction). c Gene set enrichment of the extracellular space DEGs based on the REACTOME database (top) and protein domain analysis results of the same DEGs using SMART (bottom). Gene set enrichment based on the hypergeometric test (one-sided, Benjamini-Hochberg correction). d Expression of genes known as upregulated by wounding to contribute healing among extracellular space-associated DEGs in (b). e EGFR pathway activity (two-tailed Mann–Whitney U test with Bonferroni correction). Box plots show median, interquartile range, and 1.5-fold IQR whiskers with outliers (n = 1849 (Basal-control), 328 (Prolif-control), or 836 (Supra-control) from 3 mice or 1053 (Basal-mChneg), 218 (Prolif-mChneg), 341 (Supra-mChneg), 1030 (Basal-mChpos), 550 (Prolif-mChpos), or 521 (Supra-mChpos) from 6 mice). f Phospho-EGFR and mCherry staining in control and Epi-iOSKM skin with quantification of phospho-EGFR+ area/total epidermis area (%). n = 7 from 3 mice/condition. Immunostaining of HIF-1α and mCherry (g) or Krt17 and mCherry (h) in control and Epi-iOSKM skin treated with vehicle or AG1478, with quantification of epidermal Krt17 MFI (h). Representative images from 3 biological samples (g). n = 8 (OSKM-AG1478), 9 (Control-Vehicle and Control-AG1478), or 10 (OSKM-Vehicle) from 3 mice (h). i Phospho-EGFR and mCherry immunostaining in control and Epi-iOSKM skin treated with vehicle or LY294002, with quantification of epidermal phospho-EGFR MFI. n = 9 from 3 mice/condition. j Diagram showing molecular mechanisms underlying healing features in the Epi-iOSKM epidermis. Data are mean ± SEM (two-tailed Mann-Whitney test (f) or two-way ANOVA with Tukey’s multiple comparisons test (h, i)). Dashed lines, boundary between the epidermis and dermis (f–i). Pink arrows, cell non-autonomous effects (f). Pink wording, EGFR-associated expressions (a, b, d). Scale bars, 50 µm (f–i). GO-CC Gene Ontology-Cellular Component.
Next, we examined how OSKM-expressing cells modify cellular communication within the epidermis. We identified upregulated DEGs in mChpos IFE cells compared to controls, focusing on genes encoding proteins in the extracellular space that could potentially mediate CCIs (Fig. 4b). Analysis using both the REACTOME pathway database and the SMART protein domain database revealed a significant upregulation of EGFR-related genes (Fig. 4c). The expression of Epgn, Tgfa, and Areg, which encode ligands that activate EGFR, was upregulated in the Epi-iOSKM epidermis (Fig. 4d). We confirmed both cell-autonomous and non-autonomous activation of EGFR signaling in the epidermis of Epi-iOSKM mice by pathway analysis of scRNA-seq data (Fig. 4e) and immunohistochemistry of phospho-EGFR (Fig. 4f). The MAPK and JAK-STAT pathways, which are downstream signaling pathways of EGFR60, were similarly activated (Supplementary Fig. 9c).
EGFR signaling is well known as a key mechanism activated during skin repair to induce epidermal proliferation and migration61,62,63. Additionally, EGFR activation is known to upregulate HIF-1α and Krt1764,65,66,67. To assess whether EGFR signaling activation in mChpos and mChneg cells contributed to healing features induced by mosaic partial epidermal reprogramming, we topically treated the dorsal skin of Epi-iOSKM mice with AG1478, an EGFR inhibitor (Supplementary Fig. 9d). Upregulation of HIF-1α and Krt17, representative markers for wound-responsive-like activation, was significantly reversed by EGFR inhibition in the Epi-iOSKM epidermis (Fig. 4g, h and Supplementary Fig. 9e), demonstrating that EGFR signaling is essential to acquire healing features. For the next question, we investigated how EGFR signaling was activated in the Epi-iOSKM epidermis. We found that PI3K activation in mChpos cells is necessary for EGFR activation in the Epi-iOSKM epidermis (Fig. 4i). Furthermore, treatment of cultured keratinocytes with a PI3K activator induced EGFR activation (Supplementary Fig. 9f). These demonstrate that PI3K activation in OSKM-expressing IFE cells conferred healing characteristics to themselves, and activated EGFR signaling to induce healing features not only to themselves but also to their neighbors (Fig. 4j).
EGFR activation in concert with WNT inhibition accelerates repair without altering proliferation
The cellular pathways identified in the Epi-iOSKM epidermis were associated with increased proliferation; EGF signaling, a key modulator for keratinocyte proliferation, as well as IL1 and VEGF signaling that can indirectly contribute, were significantly enriched60,62,63,68,69 (Fig. 4b, d). However, we did not observe any changes in the proliferation of mChneg cells (Fig. 1j, k). We hypothesized that an additional factor might regulate cell proliferation of mChneg cells. PROGENy pathway analysis revealed that WNT signaling was the only pathway showing significantly different activity levels in mChneg cells, but not in mChpos cells, of Epi-iOSKM mice compared to control (Fig. 3e). WNT signaling was downregulated in mChneg cells of Epi-iOSKM epidermis compared to both control epidermal cells and mChpos cells of Epi-iOSKM epidermis (Supplementary Fig. 10a).
WNT signaling, known to activate cell proliferation70, emerged as a promising candidate regulating the proliferation of mChneg cells. To investigate how WNT signaling was inhibited in wildtype cells of Epi-iOSKM epidermis, we examined the RNA levels of secreted WNT regulatory molecules, which can broadly modulate the WNT pathway activity in the epidermis. First, we found that the expression of Wnt4, Wnt6, Wnt16, and Rspo3, a Wnt signaling enhancer70,71, was lower in IFE cells of the Epi-iOSKM epidermis than in control epidermis (Supplementary Fig. 10b). Next, we examined the expression of well-known WNT inhibitory molecules. Among these, the number of cells expressing Sfrp1 was much higher than those expressing other inhibitory genes in the control epidermis, suggesting that Sfrp1 is the major inhibitory modulator of WNT ligand activity in the epidermis (Supplementary Fig. 10c)72,73. The expression of Sfrp1, encoding the key repressor protein, significantly increased in mChpos cells (Supplementary Fig. 10c). Collectively, the gene expression of secreted proteins that modulate WNT signaling activity was altered toward a tendency to suppress the signal (Supplementary Fig. 10d).
To examine the effects of EGF activation in combination with WNT inhibition, we treated HaCaT human keratinocytes with EGF and the WNT secretion inhibitor LGK97474. This condition recapitulates the in vivo microenvironment of mChneg cells in the Epi-iOSKM epidermis, consisting of high EGF and low WNT ligand concentrations. Keratinocyte proliferation was increased by treatment with EGF alone but was not affected by treatment with both EGF and LGK974 (Supplementary Fig. 10e). Conversely, the wound healing rates of in vitro scratch wounds were promoted by the treatment of EGF with or without LGK974 (Supplementary Fig. 10f). These findings indicate that simultaneous EGF treatment and WNT inhibition switch keratinocytes to a state beneficial for healing while avoiding an increase in cell proliferation.
In conclusion, unlike mChpos cells, which acquired healing features and increased proliferative ability via activation of PI3K and EGFR signaling pathway, mChneg cells exhibited: (1) healing features via upregulation of the EGFR pathway, (2) downregulation of the WNT pathway, and (3) no changes in proliferation (Supplementary Fig. 10g). These findings suggest that EGFR pathway activation, combined with WNT pathway inhibition in mChneg epidermal cells of Epi-iOSKM mice converted them into a wound-responsive-like, but non-proliferative state.
Partial epidermal reprogramming activates immune niches
Having shown that partial reprogramming of IFE cells can influence the surrounding environment by inducing cellular plasticity in neighboring keratinocytes, similar to injury responses, we next determined whether and how partial reprogramming affects other cell types. We specifically focused on immune cells, as our REACTOME analysis showed that chemokines known to modulate skin immunity were significantly upregulated in mChpos cells of the Epi-iOSKM epidermis (Fig. 4c and Supplementary Fig. 11a). Also, keratinocytes activated by skin injury are known to promote T cell immunity75,76,77. T-cell responses, in turn, reciprocally activate epidermal keratinocytes, contributing to wound re-epithelialization (Fig. 5a)75,76,77,78,79.
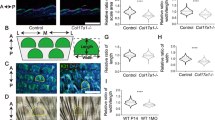
a Reciprocal interactions between epidermal keratinocytes and T cells during wound repair (blue) and skin inflammation (brown). b Proportion of each cell type and each immune cell subtype. c CD3 and Krt14 staining in control and Epi-iOSKM skin (left) with quantification of CD3+ cell numbers/length (middle left), the ratio of rounded cells among CD3+ cells in the epidermis (middle right), and CD3+ cell numbers/area in the dermis (right). n = 15 from 3 mice/condition. Volcano plot showing DEGs in wounded IFE cells versus that of unwounded (d), or Epi-iOSKM IFE cells versus that of control (e). Red and blue dots represent upregulated and downregulated DEGs in wounded (d) or Epi-iOSKM (e) IFE cells, respectively. f RORγt and Krt14 staining in the control and Epi-iOSKM skin (left) with quantifications of RORγt+ cell numbers/area in the skin (right). Yellow arrows, RORγt+ cells. n = 12 from 3 mice/condition. g Molecular mechanisms that recruit RORγt+ cells in the Epi-iOSKM skin. h Gross photos of control mice after 9 days Dox or Epi-iOSKM mice after 3 or 9 days Dox (representative of 5 biological samples). Face (i), plucked dorsal back (ii), enlarged dorsal back (iii), and ventral parts (iv) of the back skin (ii). Hind paws (v), and front paws (vi). i H&E staining in the control and Epi-iOSKM skin after 9 days Dox with quantifications of relative epidermal thickness. n = 9 from 3 mice/condition. j CD3 and Krt14 staining in control and Epi-iOSKM skin after 9 days Dox with quantification of CD3+ area/total dermis area (%). n = 12 from 3 mice/condition. Data are mean ± SEM (two-tailed unpaired Student’s t-test or Mann-Whitney test (c, i, j) or two-way ANOVA with Tukey’s multiple comparisons test (f)). DEGs were defined as adj p <0.05 and |log2FC| > 1 (two-sided Mann-Whitney U test with Bonferroni correction), and among genes encoding cytokines, upregulated DEGs in (d) were presented as black or gray colors either DEGs or not in (e), respectively (d, e). Scale bars, 50 µm (c, f, i, j).
We found that partial epidermal reprogramming, which we propose to resemble the wound repair process, promotes T-cell immunity in the skin. Specifically, the proportion of T cells in the epidermis and the numbers of both epidermal and dermal T cells increased in the skin of Epi-iOSKM mice (Fig. 5b, c and Supplementary Fig. 11b) without a corresponding increase in T cell proliferation (Supplementary Fig. 11c, d), suggesting that partial epidermal reprogramming primarily promotes T cell recruitment. Furthermore, the ratio of rounded T cells in the epidermis—activated epidermal T cells known to be crucial for wound re-epithelialization—47,75,76,80, also increased (Fig. 5c). To identify how the partially reprogrammed epithelial cells acted on immune cells, we analyzed increased cytokines after wounding using the reference scRNA-seq data containing IFE cells from wounded and unwounded skin (Fig. 5d)39. We then assessed how their expression was altered by partial reprogramming of IFE cells. Notably, genes encoding ligands reported to activate dendritic epidermal T cells (DETCs) and dermal γδ T cells during injury repair, including Ccl8, Ccl20, Cxcl10, Cxcl16, Il1b, and Tnf81,82,83,84,85,86,87,88,89, were upregulated upon partial epidermal reprogramming (Fig. 5e and Supplementary Fig. 11e–g).
Moreover, the skin of Epi-iOSKM mice mimicked aspects of the immune niche in the repair state. Among the immune cytokines, Ccl20 exhibited the greatest increase in expression (Fig. 5e and Supplementary Fig. 11e–h). CCL20 is a chemokine that attracts T cells expressing its sole receptor, CCR6. During repair, the CCL20-CCR6 axis is activated, recruiting RORγt+ T cells near the wound edge82,84,90,91, to induce re-epithelialization79. Similarly, in the skin of Epi-iOSKM mice, the number of RORγt+ cells increased (Fig. 5f). Furthermore, intradermal injection of a CCR6 antagonist reversed the increase of RORγt+ cells in Epi-iOSKM skin (Fig. 5f). This indicates that partial epidermal reprogramming increased RORγt+ T cells in the skin via the CCL20-CCR6 axis (Fig. 5g). Collectively, partially reprogrammed epithelial cells not only had similar molecular and cellular identities to the injured epithelium (Figs. 2–4) but also functioned similarly in epithelial-immune crosstalk (Fig. 5).
Prolonged OSKM expression leads to epidermal hyperplasia accompanied by skin inflammation
Previous studies have reported that in vivo OSKM induction leads to detrimental effects including premature death and teratoma formation20,21, but we did not observe any deleterious effects, such as mortality and tumorigenesis, in our Epi-iOSKM mice after Dox treatment for 3 days (Fig. 5h and Supplementary Fig. 12a). To investigate the effects of prolonged OSKM expression in the epidermis, we administered Dox for 9 days in Epi-iOSKM mice. These mice exhibited pathogenic skin morphologies, including swelling of the facial skin, bumps and rashes in the dorsal skin, and exfoliated or swollen paws (Fig. 5h). No significant changes were observed in the other organs, and none of the mice died (Supplementary Fig. 12a). Furthermore, we examined the potential for teratoma development but did not observe tumor formation in the skin or the expression of Nanog, a pluripotent stem cell marker92, or Krt18, an epidermal cancer marker (Supplementary Fig. 12b, c)93.
We have shown that the activation of epithelial-immune crosstalk in the Epi-iOSKM skin resembles processes during wound healing. Notably, these interactions are also excessively activated in certain skin inflammatory diseases (Fig. 5a)4,79,94,95,96. Immunostaining for Krt17, Krt6a, and Ki67, observed in hyperimmune epidermis95,97, revealed a significant increase in the number of IFE cells expressing these proteins in the skin of Epi-iOSKM mice after 9 days of Dox treatment (Supplementary Fig. 12d–f). Although tumors did not develop, prolonged OSKM expression resulted in the thickening of the dorsal epidermis and the stratum corneum (Fig. 5i). Additionally, the number of T cells in the dermis dramatically increased (Fig. 5j). Moreover, following 10 days of Dox withdrawal after a 9-day administration period, the mice exhibited other pathologies, such as dry skin, scaly ears, and hair loss, along with abnormal skin histology characterized by excessive thickening of the stratum corneum and an increased number of CD3+ T cells in both the epidermis and dermis (Supplementary Fig. 12g–i). These persistent inflammatory phenotypes were recovered after 30 days of Dox withdrawal (Supplementary Fig. 12j, k), and pluripotency was still not induced (Supplementary Fig. 12l), suggesting that 9 days of Dox administration caused transient disease phenotypes without fatal consequences. Collectively, prolonged OSKM expression in the epidermis resulted in inflammatory skin pathologies, whereas transient mosaic partial reprogramming induced only regenerative characteristics without adverse effects.
Partial reprogramming promotes re-epithelialization
Since we had shown that short-term partial reprogramming of epithelial cells induces the skin to exhibit healing characteristics in several ways, we next examined whether this approach improves skin repair upon actual injury. To assess this, we created 6-mm splinted full-thickness excisional wounds, which induce healing kinetics similar to those of humans by preventing the rapid wound contraction commonly observed in mice with loose skin98, during short Dox administration for 3 days (Fig. 6a). This revealed that partial epidermal reprogramming accelerated re-epithelialization, as evidenced by significantly increased lengths of the migrating epithelial tongues at 10 dpw (Fig. 6b). Additionally, wound closure rates were increased in Epi-iOSKM mice, as confirmed by a decrease in the number of days reaching 50% of the wound area (wound half-life) from control = 10.58 to Epi-iOSKM = 9.15 (Supplementary Fig. 13a). Functionally, trans-epidermal water loss, an indicator of epidermal barrier function, was reduced, further supporting enhanced healing outcomes (Fig. 6c).
a Experimental scheme for reversible OSKM expression, splinted 6 mm full-thickness cutaneous wounding, and histological analysis. b Krt14 staining in epithelial migrating tongues at 10 dpw from control and Epi-iOSKM mice with quantifications of tongue lengths. n = 12 from 6 mice/condition. c Quantification of trans-epidermal water loss (gm−2 h−1) at 16 dpw from control and Epi-iOSKM mice. n = 5 mice/condition. d H&E staining of wounds at 20 dpw with quantification of the relative dermal thickness in wounds at 20 dpw or unwounded 2nd telogen skin from control and Epi-iOSKM mice. n = 6 from 3 mice/condition. e t-SNE of ECM ultrastructural parameters quantified from trichrome histology images of wounds at 20 dpw or unwounded 2nd telogen skin from control and Epi-iOSKM mice, with representative images used in the analysis. n = 52 from 3 mice/condition. f Collagen III staining in wounds at 20 dpw or unwounded skin from control and Epi-iOSKM mice with quantification of collagen III+ area/ total dermis area (%). n = 6 from 3 mice/condition. Data are represented as mean ± SEM (two-tailed unpaired Student’s t-test or two-tailed Mann-Whitney test (b, c, d, f). White dashed lines, the boundary between the epidermis and dermis (b, f). yellow dashed lines, the initial wound edges (b). white arrows, the edges of the epithelial tongues (b). Scale bars, 200 µm (b, d, f). dpw days post wounding, TEWL trans-epidermal water loss.
Partial epidermal reprogramming reduces scar formation
During cutaneous wound repair, epidermal stem cells and their niches in the dermis actively interact with each other2,10,12,44,45,75. Interestingly, granulation tissue formation was promoted in Epi-iOSKM mice compared to controls at 3 and 10 dpw (Supplementary Fig. 13b, c), suggesting partial epidermal reprogramming enhances dermal healing. Thus, we next examined how dermal scarring was influenced in Epi-iOSKM skin. After wound closure, at 20 dpw, the increase in dermal thickness associated with scarring was diminished in the skin of the Epi-iOSKM mice (Fig. 6d). Epidermal thickness also decreased (Supplementary Fig. 13d). In addition, the scars formed in Epi-iOSKM mice had intermediate characteristics between unwounded healthy dermal tissues and scars from control mice (Fig. 6e), suggesting enhanced dermal healing. Consistently, the scarred areas of Epi-iOSKM mice contained more mature collagen fibers that demonstrated a complex structure similar to the normal dermis before wounding (Supplementary Fig. 13e)99. Together, the amount of collagen deposition, as detected by aniline blue signals in trichrome staining100,101, decreased in Epi-iOSKM mice, indicating a reduction in scar formation (Supplementary Fig. 13f). Immunohistochemical analysis further revealed that collagen III, the immature and thin collagen102 that only appeared in scars and was not found in unwounded skins, was significantly less present in scarred areas of Epi-iOSKM mice than those of control, supporting less scarring in Epi-iOSKM mice (Fig. 6f and Supplementary Fig. 13g). The percentage of collagen I+ area, the mature and thick collagen103, in the wounded dermal area did not change significantly, but the relative amount of collagen I deposition, normalized to epidermal lengths, was reduced in Epi-iOSKM mice (Supplementary Fig. 13h). Taken together, the ratio of collagen III to collagen I was significantly decreased in Epi-iOSKM mice, suggesting that immature collagen III, typically observed in skin undergoing repair, was converted more rapidly to mature collagen I, which is found in the repaired skin (Supplementary Fig. 13i)104. Collectively, mosaic partial reprogramming of epithelial cells in the skin accelerated re-epithelialization and reduced dermal scarring, thereby broadly enhancing skin repair.
Extensive alterations in wound repair processes induced by partial epidermal reprogramming
We next investigated the mechanistic link between partial epidermal reprogramming and the enhancement of epidermal and dermal repair. Our first question was how transient partial reprogramming until 1 dpw could promote healing at later stages (10–20 dpw). The second question was how partial reprogramming of the epidermis influences dermal scarring. To elucidate how the effects of partial epidermal reprogramming may expand spatiotemporally, we examined changes across multiple cell types in Epi-iOSKM skin during intermediate stages of wound repair (Fig. 7a and Supplementary Fig. 14a).
a Experimental scheme. HIF-1α and mCherry staining in control and Epi-iOSKM wounds at 1 (b) or 7 dpw (c). Yellow arrows, old epithelium with increased HIF-1α signals. d Schematic of HIF-1α levels at the early response phase after wounding. e HIF-1α staining in the control and Epi-iOSKM skin without wounding after 3 days Dox (top) and after 6 days withdrawal (bottom). f HIF-1α and CD31 staining in control and Epi-iOSKM wounds at 10 dpw. g, h Enlarged views of yellow areas in (f) with quantification of relative CD31+ dermal area within 300 × 100 µm2 rectangle near to initial wound sites (g) and red areas in (f) with quantifications within 300 µm near the leading edges (h). n = 5 from 3 mice/condition. i Experimental scheme for HIF-1α inhibition. j Krt14 and CD31 staining in control and Epi-iOSKM wounds at 10 dpw with vehicle or PX-478 treatment. k Quantification of migrating tongue lengths at 10 dpw. n = 6 from 3 mice/condition. l Quantifications of relative CD31+ dermal area of healing area at 10 dpw. n = 6 from 3 mice/condition. m Trichrome staining of control and Epi-iOSKM wounds at 20 dpw with vehicle or PX-478 and quantification of relative dermal thickness. n = 9 from 3 mice/condition. n Collagen III staining in control and Epi-iOSKM wounds at 20 dpw with vehicle or PX-478 and quantification of collagen III+ area/total dermis area (%). n = 9 from 3 mice/condition. Data are mean ± SEM (two-tailed unpaired Student’s t-test (g), two-tailed Mann-Whitney test (h), or two-way ANOVA with Tukey’s multiple comparisons test (k–n)). Images are representative of 4 (b, c) or 3 (e, f, j) biological samples. White dashed lines, boundary between the epidermis and dermis (b, c, e, f, j, m, n); yellow dashed lines, the initial wound edges (b, c, f, j); white arrows, edges of migrating tongues (f, j). Scale bars, 200 µm (b, c, e, f, j, m, n).
First, we examined alterations in the epidermis during healing. While partial epidermal reprogramming induced cell-autonomous proliferation under homeostatic conditions (Fig. 1j, k), the number of proliferating IFE cells was not altered in Epi-iOSKM skin during wound repair (Supplementary Fig. 14b). In contrast, the levels of HIF-1α, which we had found to be increased by partial reprogramming in cell-autonomous and non-autonomous manners (Figs. 2l and 7b), were still elevated near the wound edges at 3 and 7 dpw in Epi-iOSKM mice (Fig. 7c and Supplementary Fig. 14c), even though Dox administration had been stopped. Notably, HIF-1α levels were more increased near old epidermal cells that had experienced partial reprogramming than in regenerated migrating epithelial tongues (Fig. 7c, d). This was surprising because in the absence of wounding, HIF-1α activation in Epi-iOSKM epidermis disappeared after Dox withdrawal for 6 days (Fig. 7e). The different reversibility of HIF-1α activation between no wounding and wounding suggests that partially reprogrammed cells synergize with actual wounds to maintain and enhance the reversible effect observed without wounding.
We next examined dermal alterations in healing Epi-iOSKM skin. During wound healing, three main types of dermal niches interplay with epidermal cells: mesenchymal, immune, and vascular cells. We first assessed the differentiation of mesenchymal cells into myofibroblasts, but detected no notable changes in immunohistochemical analysis for α-SMA, a marker of myofibroblasts (Supplementary Fig. 14d). We then investigated alterations in immune niches. Macrophage infiltrations were unaffected by partial epidermal reprogramming (Supplementary Fig. 14e). However, T cell numbers in wounded area were increased. At 3 dpw, the number of epidermal T cells located proximal to the wound edge was higher in the Epi-iOSKM skin than in the control skin (Supplementary Fig. 14f). At 10 dpw, there were more infiltrated T cells in the wounded area of Epi-iOSKM mice (Supplementary Fig. 14g).
Lastly, we examined the vascular niche by performing immunohistochemical analysis for CD31, a marker for endothelial cells. Interestingly, the pattern of angiogenesis was significantly altered following partial epidermal reprogramming (Fig. 7f–h), correlating with earlier changes in HIF-1α signaling (Fig. 7b–d), which is known to promote neovascularization. In the dermis near the initial wound site, blood vessels in Epi-iOSKM skin were more closely localized to the epidermis at 10 dpw (Fig. 7f, g), consistent with the increase of HIF-1α in the same area at earlier time points (Fig. 7b–d). In the wound bed near the newly generated epithelium, control mice exhibited higher HIF-1α levels compared to the initial wound site during earlier stages of healing (Fig.7b–d). In contrast, in Epi-iOSKM mice, HIF-1α signals were comparable near the initial wound site and the newly generated epithelium, with elevated levels at both locations (Fig. 7b–d). By 10 dpw, Epi-iOSKM skin, which did not exhibit different HIF-1α levels according to the location, showed a dramatic reduction in neovascularization in the dermis near the leading edge of migrating tongues compared with controls (Fig. 7f, h). Despite decreased neovascularization at 10 dpw in the wounded area near leading edges, after wound closure, CD31+ cell areas in the dermis between control and Epi-iOSKM mice became similar (Supplementary Fig. 14h), suggesting that angiogenesis in Epi-iOSKM skin was not completely blocked but instead occurred in an altered pattern.
HIF-1α activation is essential for enhancing wound healing and attenuating scar formation in Epi-iOSKM skin
Previous studies have reported that forced activation of HIF-1α accelerates re-epithelialization105,106,107. In our model, HIF-1α activation induced by partial reprogramming was sustained after wounding (Fig. 7c), suggesting it may continue to promote re-epithelialization even after transient reprogramming has ceased. Moreover, the altered patterns of HIF-1α activation correlated with reorganized angiogenesis (Fig. 7b–h), potentially influencing dermal repair. Notably, appropriate regulation of neovascularization during wound repair is known to reduce scar formation108,109,110,111.
When a wound occurs in Epi-iOSKM mice, HIF-1α is activated twice in the epidermis—first by partial reprogramming and again by the wound itself. To disrupt the synergistic effect of these two stimuli, we transiently inhibited HIF-1α activity by treating the HIF-1α inhibitor PX-478 once at the time of wounding (Fig. 7i). HIF-1α activity decreased at 1 dpw but recovered by 10 dpw as expected (Supplementary Fig. 15a, b). Interestingly, transient HIF-1α inhibition during the early repair phase reversed the accelerated re-epithelialization in Epi-iOSKM mice, bringing it to levels comparable to controls (Fig. 7j, k). Moreover, temporal inhibition of HIF-1α markedly reduced angiogenesis. Without inhibition, control mice displayed widespread angiogenesis across the wound bed, whereas Epi-iOSKM mice showed angiogenesis concentrated near the original wound site (Fig. 7f–h). Following PX-478 treatment, however, both groups exhibited a pronounced reduction in angiogenesis, with low CD31 expression at newly formed dermis near both the initial wound sites and leading edges (Fig. 7j, I and Supplementary Fig. 15c, d). These findings indicate that HIF-1α is a key regulator of neovascularization patterns during wound repair in both control and Epi-iOSKM mice.
We next examined whether HIF-1α, which is essential for the alterations in re-epithelization and angiogenesis in Epi-iOSKM mice, also contributes to scar reduction. At 20 dpw, dermal thickness in the scarred area of PX-478 treated Epi-iOSKM mice was significantly thicker than that that of untreated Epi-iOSKM mice and was comparable to that of PX-478 treated control mice (Fig. 7m). Consistently, the alterations in collagen I deposition observed in Epi-iOSKM skin were restored by PX-478 treatment (Supplementary Fig. 15e). In addition, the reduced proportion of collagen III+ area observed in Epi-iOSKM mice was reversed by HIF-1α inhibition during partial epidermal reprogramming (Fig. 7n), indicating that elevated HIF-1α levels are essential for the reduction in scarring. Collectively, increased HIF-1α activation upon partial epidermal reprogramming—particularly the atypical pattern observed in the old epidermis near the initial wound site rather than in the newly generated epithelium (Fig. 7b–d)—substantially contributes to the widespread changes that promote wound repair.
Partial epidermal reprogramming enhances impaired wound healing under hyperglycemic conditions
We next investigated potential clinical implications by generating a mouse model of hyperglycemia, the most widely used model for impaired wound healing, through streptozotocin injection into male mice112 (Fig. 8a and Supplementary Fig. 16a). This disease model exhibited decelerated wound closure (Fig. 8b). Consistent with our findings in wild-type mice, short-term mosaic OSKM induction in diabetic mice accelerated wound closure compared with diabetic controls. At 11 dpw, wound areas were significantly smaller in diabetic Epi-iOSKM mice than in diabetic controls, bringing them to levels comparable to those of vehicle-treated controls (Fig. 8b).
a, Experimental scheme for STZ injection for modeling of hyperglycemia mice, reversible OSKM expression, and splinted 6 mm full-thickness cutaneous wounding. b Gross photos of splinted full-thickness cutaneous wounds at 0 and 11 dpw from vehicle control, STZ control and STZ Epi-iOSKM mice with quantification of wound area (%) at 11 dpw in the same mice. n = 7 (Veh Control and STZ Control) or 8 (STZ Epi-iOSKM) mice. c Ki67 staining in epithelial migrating tongues at 3 dpw from vehicle control, STZ control and STZ Epi-iOSKM mice with quantifications of the percentage of Ki67-positive cells. n = 6 from 3 mice/condition. d Krt14 and CD31 staining in epithelial migrating tongues at 11 dpw from vehicle control, STZ control and STZ Epi-iOSKM mice (representative of 3 individual biological samples). e Quantifications of epithelial migrating tongue lengths at 11 dpw. n = 6 from 3 mice/condition. f Quantifications of relative CD31+ area in the dermis of the healing area at 11 dpw. n = 6 from 3 mice/condition. g Table describing how partial epidermal reprogramming affects wound repair processes in wildtype and hyperglycemic mice. Data are represented as mean ± SEM (one-way ANOVA with Tukey’s multiple comparisons test (b, e) or Welch’s ANOVA with Dunnett’s T3 multiple comparisons test (c, f). Red and white dashed lines, the boundary between the epidermis and dermis (c, d), respectively; yellow dashed lines, the initial wound edges (c, d); white arrows, the edges of the epithelial tongues in (d). Scale bars, 200 µm (c, d). STZ streptozotocin, Veh vehicle.
It is known that hyperglycemia impairs cell proliferative capacity113, and consistently, our diabetic model exhibited reduced epidermal cell proliferation. Interestingly, this defect was restored in Epi-iOSKM mice (Fig. 8c). These suggests that the effects of partial epidermal reprogramming on diabetic wound healing differ from those observed in the wildtype healing model: in the normal healing, partial reprogramming did not alter epidermal proliferation during repair (Supplementary Fig. 14b). We next evaluated HIF-1α activity, another key factor for epithelial activation. HIF-1α levels were elevated in diabetic Epi-iOSKM mice compared with diabetic controls, similar to the effects observed in the non-diabetic condition (Supplementary Fig. 16b). Collectively, these results indicate that partial epidermal reprogramming can counteract diabetes-induced impairments in epidermal repair.
We additionally found that hyperglycemia impaired re-epithelialization, neovascularization, and granulation tissue formation after wounding, consistent with previous reports113,114,115. These defects were also restored by partial epidermal reprogramming (Fig. 8d–f and Supplementary Fig. 16c, d). Diabetic Epi-iOSKM mice exhibited faster re-epithelialization than diabetic controls (Fig. 8d, e), demonstrating that the beneficial effects observed in normal mice (Fig. 6b) can also be applied in a disease context. In contrast, in the case of angiogenesis, the effects of partial epidermal reprogramming differed under hyperglycemic conditions, which are characterized by a marked reduction in vascularization. The total CD31+ area in the dermis increased in diabetic Epi-iOSKM mice compared to diabetic controls, partially rescuing the vascularization defect (Fig. 8d, f), in contrast to the non-diabetic condition, where Epi-iOSKM mice exhibited an altered pattern of neovascularization (Fig. 7f–h) but no changes in the total amount (Fig. 7l). We next examined collagen deposition after wound closure under hyperglycemic conditions but did not detect significant differences between Epi-iOSKM and control skin (Supplementary Fig. 16e–g). However, decreased granulation tissue formation observed in diabetes was recovered by partial epidermal reprogramming (Supplementary Fig. 16d). Collectively, under impaired healing conditions by diabetes, partial epidermal reprogramming exhibited distinct effects compared to normal conditions, ultimately improving the defective healing processes (Fig. 8g).
Discussion
In this study, we developed a mouse model of in vivo mosaic partial epidermal reprogramming. We showed that the skin of these mice switched to a state ready for healing by mimicking injury repair processes. To be specific, we demonstrated that the partially reprogrammed epithelial cells orchestrate cell-cell communications in the skin during both homeostasis and repair, to convert their neighbors and niches to be more regenerative, in addition to contributing to repair themselves. To distinguish the effects of partially reprogrammed cells from those of non-reprogrammed neighboring cells, we used the mCherry reporter system33. This enabled us to reveal distinct cell-autonomous and cell-non-autonomous responses of mChpos and mChneg cells within the epidermis of Epi-iOSKM. In addition, we observed changes in other cell types relevant for wound healing, including T cell recruitment and activation within the skin (Fig. 9a). Notably, during the repair process, Epi-iOSKM mice exhibited accelerated re-epithelialization, altered angiogenesis, and diminished scarring (Fig. 9b). These beneficial effects occurred without any observable detrimental effects to the organism, making our approach a promising potential therapeutic avenue to enhance wound healing.
a Effects of partial epidermal reprogramming on skin homeostasis. b Effects of partial epidermal reprogramming on skin wound repair. Images are reproduced with permission from Bioinsight under the CC BY license (a, b).
To clinically apply in vivo partial reprogramming, it is essential to understand how the partially reprogrammed cells influence the complex networks in the tissue and the organism as a whole. Our mosaic partial epidermal reprogramming model reveals that the beneficial effects are not restricted to the partially reprogrammed cells but extend to their surroundings. This finding raises non-cell-autonomous effects of partial reprogramming as a previously unrecognized mechanism to enhance tissue repair. Moreover, it appears advantageous for safety that not all cells are partially reprogrammed. This is because neighboring cells are endowed with healing features but are not proliferating, thus avoiding adverse consequences via uncontrollable cell division. Additionally, in the practical application of general in vivo partial reprogramming, it is technically unlikely that OSKM or other regeneration factors will be expressed in all cells. Thus, our mosaic model may mimic the real application more precisely, and this realistic model suggests that a minimal and safe level of partial reprogramming efficiency is sufficient to promote regeneration. Further studies on the effects of partial reprogramming rates could provide deeper insights into the safe and effective levels required for clinical use.
We explored the molecular mechanisms to induce the Epi-iOSKM epidermis to favor repair, and identified the contribution of cell-autonomous PI3K and EGFR pathway activation, along with cell-non-autonomous EGFR pathway activation and WNT pathway inhibition (Supplementary Fig. 10g). Our experimental results, together with previous studies, suggest that these pathways interact with each other in complex ways for partial epidermal reprogramming. Notably, we found that PI3K pathway activation in mChpos cells is necessary for EGFR activation (Fig. 4i). This finding implies that injury-responsive-like conversion of mChpos cells via the PI3K pathway may represent an initial step, after which the converted OSKM-expressing cells then activate EGFR signaling in both autocrine and paracrine manners, to confer healing features broadly in the epidermis. Additionally, we found the transcriptional downregulation of WNT ligands and upregulation of SFRP1, a WNT inhibitor, in mChpos cells, which could suppress WNT pathway in mChneg cells. The downregulation of Wnt4 under in vivo OSKM expression has already been reported28. Sfrp1 can be upregulated upon PI3K pathway activation116,117, and Sox2 is able to directly bind the Sfrp1 promoters118. Despite these changes in gene expression suppressing the WNT pathway, we observed that the activity of WNT pathway was not altered in mChpos cells (Supplementary Fig. 10a). This may be the involvement of mChpos cells-specific PI3K pathway activation. PI3K-AKT activation is known to activate WNT-β-catenin pathway by inhibiting GSK-3β, a repressor of β-catenin119,120. Thus, these suggest a dynamic interplay between these mechanisms that modulate PI3K, EGFR, and WNT pathways to confer healing features in the Epi-OSKM epidermis.
Mosaic partial reprogramming in IFE cells promoted not only epidermal repair but also dermal repair. In particular, accelerated re-epithelialization was accompanied by altered angiogenesis patterns and reduced scar formation. Mechanistically, we identified epidermal HIF-1α activation as a key factor contributing to enhanced dermal healing in Epi-iOSKM skin. However, both enhanced re-epithelialization and HIF-1α activation are generally associated with increased angiogenesis, contrary to the angiogenic phenotype observed in Epi-iOSKM skin. One possible explanation for this unexpected angiogenesis pattern is the spatially biased hyperactivation of HIF-1α. Specifically, HIF-1α signaling was increased in the old epithelium that had undergone partial reprogramming, rather than in the newly regenerated epithelium (Fig. 7b–d). Because HIF-1α regulates angiogenic and chemotactic signaling pathways121,122, relatively stronger HIF-1α induction near the initial wound site compared to the leading edge may contribute to a spatial bias in neovascularization, with enhanced vascularization in the initial wound-adjacent dermis and reduced vessel extension toward the leading edge. Although this model is plausible, a direct causal relationship has not been demonstrated. To assess the contribution of HIF-1α to the repair phenotypes of Epi-iOSKM skin, we inhibited HIF-1α activity, which reversed the enhanced dermal healing phenotype. However, similar effects were also observed in control skin (Fig. 7k–n), indicating that HIF-1α does not function exclusively in the context of partial reprogramming. Indeed, HIF-1α is a stress-responsive transcription factor that is rapidly activated after wounding and coordinates multiple repair processes45,107. Taken together, our data identify HIF-1α as one of the key factors linking the effects of partial epidermal reprogramming to altered dermal healing. However, the precise mechanisms by which the transient activation of IFE cells leads to reduced scar formation remain to be elucidated.
A previous study activated OSKM in all skin cells, including both the epidermis and dermis15; however, their phenotypes were quite different from the effects of mosaic partial epidermal reprogramming we found: Fibrosis decreased, but the wound closure rate was decelerated. These differences can be attributed to two factors: when and where OSKM was expressed. First, in the previous study, OSKM was expressed from 5 days before wounding until 10 dpw; however, in our study, we stopped OSKM expression at 1 dpw. Second, the target cell types of partial reprogramming were different. The previous study expressed OSKM in all cell types in the skin and suggested that partial reprogramming of dermal fibroblasts underlay the observed phenotypes15, unlike our study, which induced only epidermal-specific partial reprogramming. Taken together, our findings indicate that OSKM expression in an epidermal-specific manner and transiently around wound development is notably beneficial for enhancing overall skin repair processes. This suggests that epithelial stem cell progenies act as a key player regulating the overall skin repair process, unlike most previous studies that focused on how these cells were regulated by their niches. Furthermore, the timing of OSKM induction in our model highlights the significance of early-stage regulation during wound healing. In conclusion, the mosaic partial reprogramming model reveals that the effects of epithelial cells at early healing stages on their neighbors and niches are crucial determinants of the regenerative ability of the skin. From a translational perspective, our findings possess the potential that not only OSKM expression but also other ways that results in epidermal plasticity in a mosaic and transient manner could enhance wound healing in a similar manner.
In conclusion, the insights we have uncovered here through mosaic partial reprogramming strongly suggest that epithelial stem cells and their progeny serve as regulatory cell types for complex cell-cell communication in skin maintenance and repair processes. Overall, our findings hence have broad implications for in vivo partial reprogramming in tissue homeostasis and damage conditions.
Methods
Animals
K14-CreERT (JAX Strain #005107)123, ROSA26-rtTA(neo);OKMSCh250 (JAX Strain #031012)33 mice were obtained from Jackson Laboratory. Epi-iOSKM mice and their littermates as controls, used for each experiment, were obtained by crossing K14-CreERT and ROSA26-rtTA(neo); OKMSCh250 mice. All experiments were performed on 7–9-week-old mice in the 2nd telogen phase of the hair growth cycle. All control mice received the same doses of tamoxifen and doxycycline as the Epi-iOSKM mice. Both male and female mice were used for all experiments except the scRNA-seq experiments and hyperglycemic disease modeling, which were performed on male mice. All mice were bred and maintained in the animal care facility of the Pohang University of Science and Technology (POSTECH) Biotech Center per the institutional ethical guidelines. Mice were group-housed with bedding and a red hut under specific pathogen-free conditions, maintained on a 12-h light-dark cycle at an ambient temperature of 22 °C and 45–65% relative humidity. All animal protocols and experiments were approved by the POSTECH Institutional Animal Care and Use Committee (POSTECH-2021-0115, POSTECH-2022-0080, POSTECH-2023-0067, POSTECH-2023-0156 and POSTECH-2024-0140).
Cell lines
HaCaT human keratinocytes124 were donated by Dr. Kyong-Tai Kim (POSTECH). HaCaT cells were cultured in Dulbecco’s modified Eagle medium (DMEM) with high glucose (4.5 g/L glucose, 4 mM L-glutamine, sodium pyruvate; Cytiva, SH30243.01) supplemented with 10% fetal bovine serum (FBS, Corning, 35-015-CV) and 100 U/ml penicillin-streptomycin (Gibco, 15140122) using a cell culture incubator maintaining at 37 °C in a 5% CO2 atmosphere. For the PI3K activation experiment, cells were serum-starved in DMEM with high glucose containing 1% FBS for 5 h, then treated with 10 µM 740 Y-P (MedChemExpress, HY-P0175) or 0.1% DMSO for 24 h at 37 °C. For EdU cell proliferation assay, cells were serum-starved in DMEM with high glucose containing 1% FBS for 12 h, then treated with 20 ng/mL EGF (Gibco, PMG8043) or 20 μM LGK974 (MedChemExpress, HY-17545) for 24 h at 37 °C.
Chemical treatment
Tamoxifen (Sigma-Aldrich, T5648) was dissolved in corn oil to a final concentration of 20 mg/ml and used to induce CreER. Tamoxifen was intraperitoneally injected into Epi-iOSKM mice and sex-matched littermate controls once daily for 6 days. To induce OSKM, control and Epi-iOSKM mice were fed 2 mg/ml doxycycline (Dox, Sigma-Aldrich, D9891) in drinking water (supplemented with 5% sucrose) and administered 2 mg/ml Dox dissolved in water by oral gavage every 2 days. For the short-term induction of OSKM, mice were treated with Dox in drinking water for 3 days and orally gavaged with Dox in water on days 0 and 2 after starting Dox treatment. For prolonged induction of OSKM, mice were administered Dox in drinking water for 9 days and orally treated with Dox in water on days 0, 2, 4, 6, and 8. To inhibit PI3K pathway, control and Epi-iOSKM mice were topically treated with 80 nmol of LY294002 (MedChemExpress, HY-10108) in acetone (Sigma-Aldrich, 179124) on the right side of the dorsal skin, and vehicle (acetone) alone on the left side of the dorsal skin once a day on days −1, 0, and 2 of Dox treatment. To inhibit EGFR pathway, control and Epi-iOSKM mice were topically treated with 50 μg of AG1478 (Sigma-Aldrich, 658552) in acetone on the right side of the dorsal skin, and vehicle (acetone) alone on the left side of the dorsal skin twice on days 0 and 2 of Dox treatment. The CCR6 antagonist 1 (MedChemExpress, HY-151435) was dissolved to a final concentration of 125 mg/ml in 1% DMSO in saline and injected intradermally into the dorsal skin of control and Epi-iOSKM mice twice a day for 4 days during Dox treatment. To inhibit HIF-1α, control and Epi-iOSKM mice were topically treated at the wounded site with 10 mg/kg and intraperitoneally injected with 40 mg/kg of PX-478 (MedChemExpress, HY-10231) in saline, 20 min prior to wounding.
Histology and immunostaining
Freshly harvested skin tissues were fixed in 4% paraformaldehyde (PFA, Electron Microscopy Sciences, 50-980-487) for 15 min and washed in PBS for at least 3 h at room temperature. For cryo-embedding, skin tissues were put into 30% sucrose/PBS overnight at 4 °C, then embedded in the OCT compound (Sakura Finetek, 4583) and stored at −80 °C until cryo-sectioning step. Cryosections of 14–35 μm in thickness were collected on positively charged glass slides (Globe Scientific Inc., 1358 W) and stored at −80 °C until use. For immunostaining, sections were fixed in 4% PFA for 2 min and washed with PBS and 0.1% Triton X-100 in PBS. The slides were then blocked in a blocking buffer (1% bovine serum albumin, 5% donkey serum, and 2% cold-water fish gelatin in 0.3% Triton X-100 in PBS) for 1 h at room temperature. Then the sections were incubated with primary antibodies overnight at 4 °C, and incubated with secondary antibodies for 2–4 h at room temperature. After washing at least three times for 30 min with PBS, the sections were mounted in DAPI Fluoromount-G (SouthenBiotech, 0100-20). For phospho-EGFR staining, sections were permeabilized with 0.3% H2O2 in methanol at −20 °C for 30 min. The following antibodies were used for immunofluorescence in this study at the dilutions indicated: rabbit anti-GFP (Abcam, ab290, 1:500), rat anti-RFP (ChromoTek, 5f8-150, 1:100), rabbit anti-Sox2 (Abcam, ab97959, 1:500), rabbit anti-Oct4 (Abcam, ab181557, 1:200), rabbit anti-Klf4 (Abcam, ab129473, 1:200), rabbit anti-c-Myc (Abcam, ab32072, 1:100), rabbit anti-Cleaved Caspase-3 (Cell Signaling Technology, 9661S, 1:200), rabbit anti-Krt10 (Biolegend, 905404, 1:500), rabbit anti-Krt14 (Biolegend, 905304, 1:500), chicken anti-Krt14 (Biolegend, 906004, 1:500), rabbit anti-Ki-67 (Cell Signaling Technology, 12202S, 1:500), rabbit anti-Krt17 (Abcam, ab53707, 1:500), rabbit anti-Krt6a (Biolegend, 905701, 1:500), rabbit anti-HIF-1α (Cell Signaling Technology, 36169S, 1:100), rabbit anti-phospho-STAT3 (Cell Signaling Technology, 9145S, 1:100), rabbit anti-phospho-AKT (Cell Signaling Technology, 4060S, 1:200), rabbit anti-phospho-EGFR (Cell Signaling Technology, 3777S, 1:200), rat anti-CD3 (Thermo Fisher Scientific, 14-0032-82, 1:100), rat anti-RORγt (Thermo Fisher Scientific, 14-6988-82, 1:100), rabbit anti-Krt18 (Sigma-Aldrich, HPA001605, 1:200), rabbit anti-Nanog (Cell Signaling Technology, 8822S, 1:200), rabbit anti-α-SMA (Abcam, ab5694, 1:500), rat anti-F4/80 (Biorad, MCA497R, 1:200), rat anti-CD31 (Biolegend, 102502, 1:200), rabbit anti-Collagen I (Abcam, ab21286, 1:100), rabbit anti-Collagen III (Abcam, ab7778, 1:50), donkey polyclonal Cy3 anti-Rat IgG antibody (Jackson ImmunoResearch Labs, 712-165-153, 1:400), donkey polyclonal Alexa Fluor 488 anti-Rabbit IgG antibody (Jackson ImmunoResearch Labs, 711-545-152), donkey polyclonal Alexa Fluor 488 anti-Chicken IgG antibody (Jackson ImmunoResearch Labs, 703-545-155), donkey polyclonal Alexa Fluor 647 anti-Rabbit IgG antibody (Jackson ImmunoResearch Labs, 711-605-152), and donkey polyclonal Alexa Fluor 647 anti-Chicken IgG antibody (Jackson ImmunoResearch Labs, 703-605-155). The nucleus was labeled with DAPI. H&E staining was performed using standard methods. Masson’s Trichrome staining was performed using the NovaUltra™ Masson Trichrome Stain kit (IHC World, IW-3006) per the manufacturer’s instructions on the OCT-frozen section. The sections were stained with Weigert’s Iron Hematoxylin solution for 5 min, followed by staining with Biebrich scarlet acid fushin solution for 2 min. These sections were then differentiated in PP Acid solution for 6 min, stained with 16-fold diluted Aniline Blue solution for 1 min, and differentiated with Acetic Acid solution for 2 min. Following dehydration with 95% and 100% ethanol, sections were cleared with Histo-Clear solution and mounted with Histomount solution.
Imaging and image analysis
Fluorescent images were obtained using a Nikon CSU-W1 SoRa spinning disk confocal microscope with a ×20 or ×40 objective (Nikon Instruments Inc.) or an EVOS M7000 microscope with a ×10 objective (Invitrogen). Images from a spinning disk confocal microscope were presented as maximum-intensity z-projection images. Bright-field images were obtained using a Leica Mica imaging system with a ×20 objective (Leica Microsystems), an EVOS M5000 microscope with a ×10 objective (Invitrogen), or a Zeiss Axioscan 7 slide scanner with a ×10 objective (Carl Zeiss Co. Ltd.). Images from Mica were tiled and merged using the Leica software. The images were assembled and processed using Adobe Photoshop (ver. 24.7.4, Adobe, Inc.) and Adobe Illustrator (ver. 27.9.4, Adobe, Inc.). The ImageJ software was used for image analysis and quantification. Trichrome images were used to quantify the dermal collagen ultrastructure. Collagen deposition was measured from the aniline blue-stained area assessed using the Color deconvolution2 plugin in Fiji101,125. To compare its ultrastructure, machine learning analysis was performed using the fiber quantification pipeline in MATLAB126,127. The MATLAB scripts for the fiber quantification workflow are available at the GitHub repository (https://github.com/shamikmascharak/Mascharak-et-al-ENF).
Western blot
HaCaT cells were washed with ice-cold PBS and lysed on ice in 250 µL RIPA lysis and extraction buffer (Thermo Fisher Scientific, 89901) freshly supplemented with a protease inhibitor cocktail (Roche, 04693132001). Protein concentration was determined using the BCA assay (Thermo Fisher Scientific, 23225). Samples were heated in 1× sodium dodecyl sulfate (SDS)-PAGE Loading Buffer (LPS Solution, CBS002) at 70 °C for 10 min, separated by SDS–PAGE, and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, IPVH00010). Membranes were blocked in 5% skim milk in TBS with 0.1% Tween-20 (TBS-T, LPS Solution, CBT007) for 1 h at room temperature, incubated with primary antibodies overnight at 4 °C. After washing with TBS-T, membranes were incubated with the secondary antibodies for 2 h at room temperature and washed again. The following antibodies were used in this study at the indicated dilutions: rabbit anti-EGFR (Cell Signaling Technology, 4267S, 1:1000), rabbit anti-phospho-EGFR (Cell Signaling Technology, 3777S, 1:1000), rabbit anti-AKT (Cell Signaling Technology, 4685S, 1:1000), rabbit anti-phospho-AKT (Cell Signaling Technology, 4060S, 1:1000), rabbit anti-GAPDH (Cell Signaling Technology, 2118S, 1:1000), and goat anti-rabbit IgG HRP conjugate (Bio-Rad 170–6515, 1:5000). The signal was detected using Clarity Western ECL Substrate (Bio-Rad, 1705061) and imaged with an Amersham Imager 680 (GE HealthCare). Unprocessed, full-length blots are provided in the Source Data.
In vitro scratch wound healing assay
HaCaT cells were cultured to 95% confluence in 12-well plates, and then the center of the well was scratched using a sterile 1000 μL pipette tip. The scratch areas were measured at 0, 6, 12, 18, and 24 h after scratching and quantified using the ImageJ software. 20 ng/mL EGF (Gibco, PMG8043) and 20 μM LGK974 (MedChemExpress, HY-17545) were treated 12 h before scratching.
Click-iTTM EdU cell proliferation assay
For labeling of newly synthesized DNA by EdU incorporation, HaCaT cells were incubated with 10 μM 5-ethynyl-2′-deoxyuridine (EdU, Sigma-Aldrich, 900584) for 2 h and then processed using the Click-iTTM EdU Cell Proliferation Alexa Fluor™ 488 Imaging Kit (Thermo Fisher Scientific, C10337), per the manufacturer’s protocol.
Splinted full-thickness cutaneous wounding
Punch biopsies were conducted on the center of the backs of anesthetized mice in the 2nd telogen phase of the hair growth cycle (7–9 weeks old). To wound the dorsal back skin, the hair on the dorsal area was shaved using an electric clipper and swabbed with a sterile alcohol preparation pad. A full-thickness wound was created inside each splint using a 6 mm sterile skin biopsy punch (Kai Medical, BP-60F) and surgical scissors. Six interrupted sutures of 6-0 silk (Ailee, SK617) were used to secure a silicone splint, cut from a 0.5 mm thick silicone sheet (Sigma-Aldrich, GBL664581), to the surrounding skin. The wounds and splints were covered with sterile semi-occlusive Tegaderm Film dressing (3 M Healthcare, 1624 W). Wound progression was documented using standardized photographs.
Trans-epidermal water loss (TEWL) measurement
TEWL of wounded control and Epi-iOSKM skin was analyzed using a GPSkin Barrier Pro-II device (GPOWER Inc., South Korea). TEWL measurements were performed using a 5 mm diameter device probe for about 10 s, and measurements were recorded using the GPskin research program (ver 1.0.0, GPOWER Inc.)128.
STZ and blood glucose assessment
To induce hyperglycemia, male mice were used, as they are more susceptible to STZ-induced diabetes than female mice129,130,131. Both control and Epi-iOSKM male mice received daily intraperitoneal injections of STZ (Sigma-Aldrich, S0130) dissolved in 50 mM sodium citrate buffer (Sigma-Aldrich, S4641) or vehicle alone for 4 days. Blood glucose levels were measured before injection and 12 days later using a Bayer Contour Plus glucometer (Panasonic Healthcare Co., Japan)132 on tail vein blood samples.
Epidermal cell isolation for scRNA-seq
The dorsal skin tissues of mice were collected and dissociated with the gentleMACS™ Dissociator (Miltenyi Biotec) and Epidermis Dissociation Kit (Miltenyi Biotec, 130-095-928) according to the manufacturer’s instructions. The cell suspension was sequentially filtered through 70 μm (Falcon, 352350) and 40 μm (Falcon, 352340) strainers. Centrifugation was performed at 400 × g for 10 min at 4 °C and the cell pellets were resuspended in PBS containing 5% FBS. The resuspended cells were incubated with Red Blood Cell Lysis Buffer (Roche, 11814389001) for 3 min at room temperature and centrifuged at 400 × g for 10 min at 4 °C. The final cell pellets were resuspended in PBS containing 5% FBS and the resuspended cells were sorted using a CytoFLEX SRT cell sorter (Beckman Coulter) and the CytExpert SRT software (Beckman Coulter). Live singlet cells were identified using DAPI staining (Sigma-Aldrich, D9542) and sorted. Further gating was used to isolate mCherry-positive and mCherry-negative cells. For the scRNA-seq using mCherry-sorted epidermal cells (1st dataset), mCherry-negative cells from three control mice and mCherry-negative and mCherry-positive cells from six Epi-iOSKM mice were sorted. For the scRNA-seq using total epidermal cells from control and Epi-iOSKM mice regardless of OSKM-mCherry expression (2nd dataset), all sorted live cells from one mouse per group were used. All cells within a condition were pooled.
scRNA-seq library preparation
Libraries for scRNA-seq were generated using the Chromium Next GEM Single Cell 3′ Reagent Kits v3.1 for dual indexing including Chromium Next GEM Single Cell 3′ Kit v3.1 (10X Genomics, PN-1000269), Chromium Next GEM Chip G Single Cell 3′ Kit (10X Genomics, PN-1000127), 3′ Feature Barcode Kit (10X Genomics, PN-1000262), Dual Index Kit TT Set A (10X Genomics, PN-1000215), and Dual Index Kit NN Set A (10X Genomics, PN-1000243). For multiplexing, samples were labeled with cell multiplexing oligo (CMO) from the 3′ CellPlex Kit Set A (10X Genomics, PN-1000261) and pooled in equal ratios. The pooled cell suspension was diluted to 1.6 × 106 cells/ml, and 30.9 µl was mixed with reverse transcription (RT) master mix to capture a total of 30,000 cells. To generate the gel bead-in-emulsions (GEMs), the mixture was loaded onto the Chromium Next GEM chip G for microfluidic processing on a Chromium controller (10X Genomics). Following incubation in a thermal cycler, the GEMs were broken, and pooled cDNA was purified using silane magnetic beads. During cDNA size selection using 0.6X SPRIselect beads (Beckman Coulter, B23318), the supernatant and bead pellet fractions were treated independently in order to extract DNA from the CMO feature barcode and cDNA from poly-adenylated mRNA, respectively. To generate cell multiplexing libraries, 75 µl of the supernatant was retained and further purified. Indexing primers (10X Genomics, PN-1000243) and cell multiplexing barcode DNA were then added to the PCR reaction mixture for amplification. For cDNA library generation, bead pellets were purified, and cDNA was eluted. The barcoded cDNAs were amplified using a thermal cycler. Subsequent steps for library construction, including enzymatic fragmentation, end repair, A-tailing, adaptor ligation, and sample index PCR, were performed according to the manufacturer’s protocol (10X Genomics, CG000388 Rev A). The size and concentration of cDNA and libraries were analyzed using a High Sensitivity DNA ScreenTape Analysis Kit with the TapeStation system (Agilent Technologies). Libraries were pooled and sequenced on a Novaseq S4 platform (Illumina) using a paired-end 100 bp protocol to generate a minimum of 20,000 read pairs per cell for the 3′ gene expression library and 5000 read pairs per cell for the cell multiplexing library.
scRNA-seq data pre-processing
An identical pipeline was used to pre-process the scRNA-seq data from the 1st dataset with mCherry-sorted cells and the 2nd dataset with total epidermal cells. For all data, generated raw reads were mapped to the mouse reference genome (GRCm39) and demultiplexed based on barcodes specific to each sample using the Cell Ranger multi (v7.0.1, 10X Genomics). Based on quality control metrics calculated by the addPerCellQC function from the scater (v1.26.1) R package133, the cells were considered low-quality if the detected unique molecular modifiers (UMI) were less than 500 or if the mitochondrial RNA content of the total UMI exceeded 20% or 10% for the 1st and 2nd datasets, respectively. Cells were clustered using the quickCluster function from the scran (v1.26.2) R package134. Cell-specific size factors were calculated using the computeSumFactors function from the same package. Raw counts were then normalized by these size factors and log-transformed with a pseudocount of 1. Highly variable genes (HVGs) were identified based on variance modeling computed using the modelGeneVar from the scran R package. The false discovery rate thresholds were 0.05 for the 1st dataset and 0.005 for the 2nd dataset. The top 15 principal components (PCs) were computed based on the HVGs and used for clustering and UMAP dimension reduction for the 1st dataset. The top 20 PCs were used in the case of the 2nd dataset. Cell types were annotated based on cluster marker genes identified using the FindMarkers function from the Seurat (v5.0.1) R package135. IFE cells were identified and subsetted based on expression patterns of canonical gene markers (Krt5, Krt14, Krt10, and Krt1) across cell clusters, then underwent the pre-processing steps described above.
scRNA-seq data analysis
Module scores to analyze the aggregated expression of a gene set were computed using the addModuleScore function from the Seurat R package. The curated gene sets based on the Gene Ontology Biological Process terms were obtained from MSigDB136 and used to evaluate how key biological pathways were regulated across different cell types upon OSKM induction. To examine the regulation of pathways related to hypoxia signaling and cytokine-mediated inflammatory response in epithelial cells under different conditions, module scores were calculated using gene sets associated with the REACTOME database terms cellular response to hypoxia and cytokine signaling in immune system. Monocle3 (v1.3.4) was used to assess differentiation in IFE cells. Cells were clustered using the cluster_cells function with UMAP as the dimensionality reduction method. A pseudotime trajectory was constructed with the learn_graph function using default parameters. To establish a biologically relevant root for the pseudotime trajectory, we selected cells with high Krt14 expression and low Krt10 expression, indicative of undifferentiated IFE cells. Using this, we ordered the cells along the inferred pseudotime using the order_cells function. scVelo (v0.2.5)137 was used to analyze dynamic transcriptional changes and infer RNA velocity. The annotated spliced and unspliced reads required for the analysis were obtained using Velocyto (v0.17.17)138. We then computed RNA velocities using the dynamical model implemented in scVelo, which allows for a more flexible, dynamic representation of RNA velocity that accounts for the temporal changes in expression across cell states. This was conducted separately for each condition. Following the computation of RNA velocities, we performed trajectory analysis and connectivity mapping using partition-based graph abstraction (PAGA) also implemented in scVelo. Directed graphs representing the connectivity and progression between transcriptionally distinct clusters were constructed by leveraging the velocity-inferred cell-state transitions. To visualize the RNA velocity results and PAGA trajectories in an embedding that optimally represents trajectory information, we constructed a diffusion map representation that preserves the global topology and directionality of transitions. Diffusion maps were computed using the palantir.utils.run_diffusion_maps function implemented in the Palantir package (v1.3.3)139. We employed the CytoTRACE2 (v1.0.0) R package140 to evaluate cellular potency across IFE cells. Predictions of developmental potential were generated using the cytotrace2 function with the full_model parameter set to TRUE, which enables an ensemble approach that utilizes all available models to maximize robustness and accuracy. For comparison and visualization, we focused on the CytoTrace2_Relative output, which provides a ranked relative order of cells according to their absolute potency scores. The scores are normalized to a range from 0 (most differentiated) to 1 (least differentiated). Pathway activity scores were inferred using the progeny (v1.20.0) R package52 using the top 500 responsive genes for each pathway curated in the package. We used the CellChat (v2.1.2) R package141 to investigate how OSKM induction altered IFE cell communication. Pre-processed data were separated by condition (control versus Epi-iOSKM), processed according to the CellChat pipeline, and then merged again for analysis. Over-expressed ligand-receptor (LR) pairs were identified using the identifyOverExpressedGenes function, followed by the identifyOverExpressedInteractions function for LR interactions curated within the CellChatDB.mouse function. Communication probabilities between interacting cell groups were calculated using the computeCommunProb and computeCommunProbPathway functions, employing the trimean method to compute the average gene expression per cell type. Finally, the rankNet function was used to visualize and compare the relative information flow after merging the processed CellChat objects from each condition. To functionally interpret the gene set upregulated in mChpos IFE cells compared to controls and included in the Gene Ontology Cellular Component term extracellular space, we performed enrichment analysis using the REACTOME database. This was conducted using the gost function from the g:Profiler2 (v0.2.3) R package142. Additionally, we assessed enrichment on the SMART protein domain database curated in the STRING database version 11.5 using the STRINGdb (v2.10.1) R package143. The hypergeometric test was computed with the get_enrichment function, using the Benjamini-Hochberg procedure for multiple testing correction.
Public scRNA-seq data analysis
To compare the molecular changes caused by OSKM induction with those caused by skin wounding, we analyzed publicly available scRNA-seq data (accession code GEO: GSE142471)39 from two wounded and three unwounded mouse skin samples. The individual count matrices were merged and pre-processed using the same pipeline described earlier in the scRNA-seq data pre-processing subsection. Sample details such as origin, condition, and cell type were obtained from the original study39. We specifically focused on basal and spinous cell populations by subsetting such cells from the entire dataset. These subsets were pre-processed following the established methodology. To identify cytokines upregulated upon wounding, we compared gene expression profiles of cells from wounded and unwounded conditions using the FindMarkers function from the Seurat R package. Next, we integrated the public IFE scRNA-seq data with IFE cells from 2nd dataset using an anchor-based integration method144. Integration anchors were identified based on reciprocal PCA using the FindIntegrationAnchors function of the Seurat R package. The two IFE datasets were integrated using the IntegrateData function of the same R package. Finally, a two-dimensional UMAP visualization was generated using the top 10 PCs of the integrated data.
Statistics and reproducibility
Details for statistical analyses, including the number of replicates and the statistical tests used, are presented in the figure legends and method sections where applicable. Statistical analyses were performed by using Prism v.9.4.1 (Graphpad) and R software (R4.2.3). Two-tailed Student’s t-test, two-tailed Mann-Whitney test, one-way ANOVA with Tukey’s or Dunnett’s T3 multiple comparison test, or two-way ANOVA with Tukey’s multiple comparison test were applied to analyze the experimental data. Two-tailed Mann-Whitney U test with Bonferroni correction or Benjamini-Hochberg correction was applied to analyze the scRNA-seq data. To analyze the wound closure rates, asymmetric sigmoidal interpolation (5PL, where X is log(concentration)) with two-way repeated measures ANOVA was conducted. Statistical analysis was set at P < 0.05. DEGs were defined as genes with adjusted P-values less than 0.05.
The sample size was not predetermined using any statistical method, and no data were excluded from the analyses. Experimental and control mice were determined by genotyping and were randomly selected for the experiments. Data collection and analysis were not performed blind to the conditions of the experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All scRNA-seq data are deposited in the SRA database and are publicly available under accession code PRJNA1139235. All other data supporting the findings of this study are provided in the Source data. Source data are provided with this paper.
Code availability
All original codes generated during scRNA-seq analysis are available at the following GitHub repository: https://github.com/CB-postech/scRNA-seq_mouse_Epi-iOSKM.
References
Arwert, E. N., Hoste, E. & Watt, F. M. Epithelial stem cells, wound healing and cancer. Nat. Rev. Cancer 12, 170–180 (2012).
Gurtner, G. C., Werner, S., Barrandon, Y. & Longaker, M. T. Wound repair and regeneration. Nature 453, 314–321 (2008).
Kumar, A., Godwin, J. W., Gates, P. B., Garza-Garcia, A. A. & Brockes, J. P. Molecular basis for the nerve dependence of limb regeneration in an adult vertebrate. Science 318, 772–777 (2007).
Martin, P. Wound healing-aiming for perfect skin regeneration. Science 276, 75–81 (1997).
Plikus, M. V. et al. Epithelial stem cells and implications for wound repair. Semin. Cell Dev. Biol. 23, 946–953 (2012).
Aragona, M. et al. Defining stem cell dynamics and migration during wound healing in mouse skin epidermis. Nat. Commun. 8, 14684 (2017).
Saxena, S., Vekaria, H., Sullivan, P. G. & Seifert, A. W. Connective tissue fibroblasts from highly regenerative mammals are refractory to ROS-induced cellular senescence. Nat. Commun. 10, 4400 (2019).
Mesa, K. R., Rompolas, P. & Greco, V. The Dynamic Duo: Niche/Stem Cell Interdependency. Stem Cell Rep. 4, 961–966 (2015).
Xin, T., Greco, V. & Myung, P. Hardwiring stem cell communication through tissue structure. Cell 164, 1212–1225 (2016).
Park, S. et al. Tissue-scale coordination of cellular behaviour promotes epidermal wound repair in live mice. Nat. Cell Biol. 19, 155–163 (2017).
Xue, M. & Jackson, C. J. Extracellular matrix reorganization during wound healing and its impact on abnormal scarring. Adv. Wound Care 4, 119–136 (2015).
Talbott, H. E., Mascharak, S., Griffin, M., Wan, D. C. & Longaker, M. T. Wound healing, fibroblast heterogeneity, and fibrosis. Cell Stem Cell 29, 1161–1180 (2022).
Mack, K. L. et al. Allele-specific expression reveals genetic drivers of tissue regeneration in mice. Cell Stem Cell 30, 1368–1381.e1366 (2023).
Ring, N. A. R. et al. The p-rpS6-zone delineates wounding responses and the healing process. Dev. Cell 58, 981–992 e986 (2023).
Doeser, M. C., Scholer, H. R. & Wu, G. Reduction of fibrosis and scar formation by partial reprogramming in vivo. Stem Cells 36, 1216–1225 (2018).
Reynolds, L. E. et al. Accelerated re-epithelialization in beta3-integrin-deficient- mice is associated with enhanced TGF-beta1 signaling. Nat. Med. 11, 167–174 (2005).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502, 65–70 (2013).
Maza, I. et al. Transient acquisition of pluripotency during somatic cell transdifferentiation with iPSC reprogramming factors. Nat. Biotechnol. 33, 769–774 (2015).
Abad, M. et al. Reprogramming in vivo produces teratomas and iPS cells with totipotency features. Nature 502, 340–345 (2013).
Ohnishi, K. et al. Premature termination of reprogramming in vivo leads to cancer development through altered epigenetic regulation. Cell 156, 663–677 (2014).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733.e1712 (2016).
Browder, K. C. et al. In vivo partial reprogramming alters age-associated molecular changes during physiological aging in mice. Nat. Aging 2, 243–253 (2022).
Ha, J. et al. Intermediate cells of in vitro cellular reprogramming and in vivo tissue regeneration require desmoplakin. Sci. Adv. 8, eabk1239 (2022).
Yang, J. H. et al. Loss of epigenetic information as a cause of mammalian aging. Cell 186, 305–326.e327 (2023).
Rodriguez-Matellan, A., Alcazar, N., Hernandez, F., Serrano, M. & Avila, J. In vivo reprogramming ameliorates aging features in dentate gyrus cells and improves memory in mice. Stem Cell Rep. 15, 1056–1066 (2020).
Chondronasiou, D. et al. Multi-omic rejuvenation of naturally aged tissues by a single cycle of transient reprogramming. Aging Cell 21, e13578 (2022).
Wang, C. et al. In vivo partial reprogramming of myofibers promotes muscle regeneration by remodeling the stem cell niche. Nat. Commun. 12, 3094 (2021).
Hishida, T. et al. In vivo partial cellular reprogramming enhances liver plasticity and regeneration. Cell Rep. 39, 110730 (2022).
Kim, J. et al. Partial in vivo reprogramming enables injury-free intestinal regeneration via autonomous Ptgs1 induction. Sci. Adv. 9, eadi8454 (2023).
Lu, Y. et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129 (2020).
Chen, Y. et al. Reversible reprogramming of cardiomyocytes to a fetal state drives heart regeneration in mice. Science 373, 1537–1540 (2021).
Elbaz, J. et al. Highly efficient reprogrammable mouse lines with integrated reporters to track the route to pluripotency. Proc. Natl. Acad. Sci. USA 119, e2207824119 (2022).
Ichijo, R. et al. Tbx3-dependent amplifying stem cell progeny drives interfollicular epidermal expansion during pregnancy and regeneration. Nat. Commun. 8, 508 (2017).
Zhang, Y. V., White, B. S., Shalloway, D. I. & Tumbar, T. Stem cell dynamics in mouse hair follicles: a story from cell division counting and single cell lineage tracing. Cell Cycle 9, 1504–1510 (2010).
Mascre, G. et al. Distinct contribution of stem and progenitor cells to epidermal maintenance. Nature 489, 257–262 (2012).
Wang, S. et al. Single cell transcriptomics of human epidermis identifies basal stem cell transition states. Nat. Commun. 11, 4239 (2020).
Cao, X. et al. RIPK4 promotes epidermal differentiation through phase separation and activation of LATS1/2. Dev. Cell 60, 2761–2776.e2711 (2025).
Haensel, D. et al. Defining epidermal basal cell states during skin homeostasis and wound healing using single-cell transcriptomics. Cell Rep. 30, 3932–3947 e3936 (2020).
Zhulyn, O. et al. Evolutionarily divergent mTOR remodels translatome for tissue regeneration. Nature 620, 163–171 (2023).
Greenhalgh, D. G. The role of apoptosis in wound healing. Int. J. Biochem. Cell Biol. 30, 1019–1030 (1998).
Paladini, R. D., Takahashi, K., Bravo, N. S. & Coulombe, P. A. Onset of re-epithelialization after skin injury correlates with a reorganization of keratin filaments in wound edge keratinocytes: defining a potential role for keratin 16. J. Cell Biol. 132, 381–397 (1996).
Lim, C., Lim, J. & Choi, S. Wound-induced hair follicle neogenesis as a promising approach for hair regeneration. Mol. Cells 46, 573–578 (2023).
Hong, W. X. et al. The role of hypoxia-inducible factor in wound healing. Adv. Wound Care 3, 390–399 (2014).
Liu, S. et al. A tissue injury sensing and repair pathway distinct from host pathogen defense. Cell 186, 2127–2143.e2122 (2023).
Koh, T. J. & DiPietro, L. A. Inflammation and wound healing: the role of the macrophage. Expert Rev. Mol. Med. 13, e23 (2011).
Keyes, B. E. et al. Impaired epidermal to dendritic T cell signaling slows wound repair in aged skin. Cell 167, 1323–1338.e1314 (2016).
Lin, Y. et al. PIK3R1 negatively regulates the epithelial-mesenchymal transition and stem-like phenotype of renal cancer cells through the AKT/GSK3beta/CTNNB1 signaling pathway. Sci. Rep. 5, 8997 (2015).
Liu, Y. et al. Pan-cancer analysis on the role of PIK3R1 and PIK3R2 in human tumors. Sci. Rep. 12, 5924 (2022).
Welsh, N., Makeeva, N. & Welsh, M. Overexpression of the Shb SH2 domain-protein in insulin-producing cells leads to altered signaling through the IRS-1 and IRS-2 proteins. Mol. Med. 8, 695–704 (2002).
Holmqvist, K. et al. The adaptor protein shb binds to tyrosine 1175 in vascular endothelial growth factor (VEGF) receptor-2 and regulates VEGF-dependent cellular migration. J. Biol. Chem. 279, 22267–22275 (2004).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Ersahin, T., Tuncbag, N. & Cetin-Atalay, R. The PI3K/AKT/mTOR interactive pathway. Mol. Biosyst. 11, 1946–1954 (2015).
Zhang, X., Tang, N., Hadden, T. J. & Rishi, A. K. Akt, FoxO and regulation of apoptosis. Biochim Biophys. Acta 1813, 1978–1986 (2011).
Hannenhalli, S. & Kaestner, K. H. The evolution of Fox genes and their role in development and disease. Nat. Rev. Genet. 10, 233–240 (2009).
Gharbi, S. I. et al. Exploring the specificity of the PI3K family inhibitor LY294002. Biochem. J. 404, 15–21 (2007).
Agarwal, S., Mirzoeva, S., Readhead, B., Dudley, J. T. & Budunova, I. PI3K inhibitors protect against glucocorticoid-induced skin atrophy. EBioMedicine 41, 526–537 (2019).
Squarize, C. H., Castilho, R. M., Bugge, T. H. & Gutkind, J. S. Accelerated wound healing by mTOR activation in genetically defined mouse models. PLoS ONE 5, e10643 (2010).
van Lengerich, B., Agnew, C., Puchner, E. M., Huang, B. & Jura, N. EGF and NRG induce phosphorylation of HER3/ERBB3 by EGFR using distinct oligomeric mechanisms. Proc. Natl. Acad. Sci. USA 114, E2836–E2845 (2017).
El-Rayes, B. F. & LoRusso, P. M. Targeting the epidermal growth factor receptor. Br. J. Cancer 91, 418–424 (2004).
Barrientos, S., Stojadinovic, O., Golinko, M. S., Brem, H. & Tomic-Canic, M. Growth factors and cytokines in wound healing. Wound Repair Regen. 16, 585–601 (2008).
Gallini, S. et al. Injury prevents Ras mutant cell expansion in mosaic skin. Nature 619, 167–175 (2023).
Gibbs, S. et al. Epidermal growth factor and keratinocyte growth factor differentially regulate epidermal migration, growth, and differentiation. Wound Repair Regen. 8, 192–203 (2000).
Zhong, H. et al. Modulation of hypoxia-inducible factor 1alpha expression by the epidermal growth factor/phosphatidylinositol 3-kinase/PTEN/AKT/FRAP pathway in human prostate cancer cells: implications for tumor angiogenesis and therapeutics. Cancer Res. 60, 1541–1545 (2000).
Peng, X. H. et al. Cross-talk between epidermal growth factor receptor and hypoxia-inducible factor-1alpha signal pathways increases resistance to apoptosis by up-regulating survivin gene expression. J. Biol. Chem. 281, 25903–25914 (2006).
Ma, S., Rao, L., Freedberg, I. M. & Blumenberg, M. Transcriptional control of K5, K6, K14, and K17 keratin genes by AP-1 and NF-kappaB family members. Gene Expr. 6, 361–370 (1997).
Chung, B. M., Murray, C. I., Van Eyk, J. E. & Coulombe, P. A. Identification of novel interaction between annexin A2 and keratin 17: evidence for reciprocal regulation. J. Biol. Chem. 287, 7573–7581 (2012).
Fuchs, E. & Raghavan, S. Getting under the skin of epidermal morphogenesis. Nat. Rev. Genet. 3, 199–209 (2002).
Chedid, M., Rubin, J. S., Csaky, K. G. & Aaronson, S. A. Regulation of keratinocyte growth factor gene expression by interleukin 1. J. Biol. Chem. 269, 10753–10757 (1994).
MacDonald, B. T., Tamai, K. & He, X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev. Cell 17, 9–26 (2009).
Sigal, M. et al. Stromal R-spondin orchestrates gastric epithelial stem cells and gland homeostasis. Nature 548, 451–455 (2017).
Sunkara, R. R. et al. SFRP1 in skin tumor initiation and cancer stem cell regulation with potential implications in epithelial cancers. Stem Cell Rep. 14, 271–284 (2020).
Tumbar, T. et al. Defining the epithelial stem cell niche in skin. Science 303, 359–363 (2004).
Liu, J. et al. Targeting Wnt-driven cancer through the inhibition of Porcupine by LGK974. Proc. Natl. Acad. Sci. USA 110, 20224–20229 (2013).
Jameson, J. et al. A role for skin gammadelta T cells in wound repair. Science 296, 747–749 (2002).
Ramirez, K., Witherden, D. A. & Havran, W. L. All hands on DE(T)C: Epithelial-resident gammadelta T cells respond to tissue injury. Cell Immunol. 296, 57–61 (2015).
Boothby, I. C., Cohen, J. N. & Rosenblum, M. D. Regulatory T cells in skin injury: at the crossroads of tolerance and tissue repair. Sci. Immunol. 5, eaaz9631 (2020).
Sharp, L. L., Jameson, J. M., Cauvi, G. & Havran, W. L. Dendritic epidermal T cells regulate skin homeostasis through local production of insulin-like growth factor 1. Nat. Immunol. 6, 73–79 (2005).
Konieczny, P. et al. Interleukin-17 governs hypoxic adaptation of injured epithelium. Science 377, eabg9302 (2022).
Witherden, D. A. et al. The CD100 receptor interacts with its plexin B2 ligand to regulate epidermal gammadelta T cell function. Immunity 37, 314–325 (2012).
Cai, Y., Fleming, C. & Yan, J. New insights of T cells in the pathogenesis of psoriasis. Cell Mol. Immunol. 9, 302–309 (2012).
Sutton, C. E., Mielke, L. A. & Mills, K. H. IL-17-producing gammadelta T cells and innate lymphoid cells. Eur. J. Immunol. 42, 2221–2231 (2012).
Islam, S. A. et al. Mouse CCL8, a CCR8 agonist, promotes atopic dermatitis by recruiting IL-5+ T(H)2 cells. Nat. Immunol. 12, 167–177 (2011).
Hedrick, M. N. et al. CCR6 is required for IL-23-induced psoriasis-like inflammation in mice. J. Clin. Investig. 119, 2317–2329 (2009).
Mabuchi, T. et al. CCR6 is required for epidermal trafficking of gammadelta-T cells in an IL-23-induced model of psoriasiform dermatitis. J. Investig. Dermatol. 133, 164–171 (2013).
Richmond, J. M., Strassner, J. P., Essien, K. I. & Harris, J. E. T-cell positioning by chemokines in autoimmune skin diseases. Immunol. Rev. 289, 186–204 (2019).
Yates, C. C. et al. Delayed reepithelialization and basement membrane regeneration after wounding in mice lacking CXCR3. Wound Repair Regen. 17, 34–41 (2009).
Sutton, C. E. et al. Interleukin-1 and IL-23 induce innate IL-17 production from gammadelta T cells, amplifying Th17 responses and autoimmunity. Immunity 31, 331–341 (2009).
Alam, M. S. et al. TNF plays a crucial role in inflammation by signaling via T cell TNFR2. Proc. Natl. Acad. Sci. USA 118, e2109972118 (2021).
Kennedy-Crispin, M. et al. Human keratinocytes’ response to injury upregulates CCL20 and other genes linking innate and adaptive immunity. J. Investig. Dermatol. 132, 105–113 (2012).
Martin, B., Hirota, K., Cua, D. J., Stockinger, B. & Veldhoen, M. Interleukin-17-producing gammadelta T cells selectively expand in response to pathogen products and environmental signals. Immunity 31, 321–330 (2009).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Yamashiro, Y. et al. Ectopic coexpression of keratin 8 and 18 promotes invasion of transformed keratinocytes and is induced in patients with cutaneous squamous cell carcinoma. Biochem. Biophys. Res. Commun. 399, 365–372 (2010).
Lowes, M. A., Suarez-Farinas, M. & Krueger, J. G. Immunology of psoriasis. Annu. Rev. Immunol. 32, 227–255 (2014).
Depianto, D., Kerns, M. L., Dlugosz, A. A. & Coulombe, P. A. Keratin 17 promotes epithelial proliferation and tumor growth by polarizing the immune response in skin. Nat. Genet. 42, 910–914 (2010).
Subudhi, I. et al. Metabolic coordination between skin epithelium and type 17 immunity sustains chronic skin inflammation. Immunity 57, 1665–1680.e7 (2024).
Krueger, J. et al. Tofacitinib attenuates pathologic immune pathways in patients with psoriasis: a randomized phase 2 study. J. Allergy Clin. Immunol. 137, 1079–1090 (2016).
Zomer, H. D. & Trentin, A. G. Skin wound healing in humans and mice: Challenges in translational research. J. Dermatol. Sci. 90, 3–12 (2018).
Sidgwick, G. P. & Bayat, A. Extracellular matrix molecules implicated in hypertrophic and keloid scarring. J. Eur. Acad. Dermatol. Venereol. 26, 141–152 (2012).
Xie, Z. et al. Dual growth factor releasing multi-functional nanofibers for wound healing. Acta Biomater. 9, 9351–9359 (2013).
Lintel, H. et al. Transdermal deferoxamine administration improves excisional wound healing in chronically irradiated murine skin. J. Transl. Med. 20, 274 (2022).
Holm Nielsen, S. et al. Exploring the role of extracellular matrix proteins to develop biomarkers of plaque vulnerability and outcome. J. Intern. Med. 287, 493–513 (2020).
Tang, V. W. Collagen, stiffness, and adhesion: the evolutionary basis of vertebrate mechanobiology. Mol. Biol. Cell 31, 1823–1834 (2020).
Rodrigues, M., Kosaric, N., Bonham, C. A. & Gurtner, G. C. Wound healing: a cellular perspective. Physiol. Rev. 99, 665–706 (2019).
Botusan, I. R. et al. Stabilization of HIF-1alpha is critical to improve wound healing in diabetic mice. Proc. Natl. Acad. Sci. USA 105, 19426–19431 (2008).
Xu, J., Liu, X., Zhao, F., Zhang, Y. & Wang, Z. HIF1alpha overexpression enhances diabetic wound closure in high glucose and low oxygen conditions by promoting adipose-derived stem cell paracrine function and survival. Stem Cell Res. Ther. 11, 148 (2020).
Li, G. et al. A small molecule HIF-1alpha stabilizer that accelerates diabetic wound healing. Nat. Commun. 12, 3363 (2021).
DiPietro, L. A. Angiogenesis and scar formation in healing wounds. Curr. Opin. Rheumatol. 25, 87–91 (2013).
DiPietro, L. A. Angiogenesis and wound repair: when enough is enough. J. Leukoc. Biol. 100, 979–984 (2016).
Wilgus, T. A., Ferreira, A. M., Oberyszyn, T. M., Bergdall, V. K. & Dipietro, L. A. Regulation of scar formation by vascular endothelial growth factor. Lab. Investig. 88, 579–590 (2008).
Diao, J. S., Xia, W. S. & Guo, S. Z. Bevacizumab: a potential agent for prevention and treatment of hypertrophic scar. Burns 36, 1136–1137 (2010).
Ma, J. et al. Single-cell RNA-Seq analysis of diabetic wound macrophages in STZ-induced mice. J. Cell Commun. Signal 17, 103–120 (2023).
Avitabile, S. et al. Interleukin-22 promotes wound repair in diabetes by improving keratinocyte pro-healing functions. J. Investig. Dermatol. 135, 2862–2870 (2015).
Sun, Y., Song, L., Zhang, Y., Wang, H. & Dong, X. Adipose stem cells from type 2 diabetic mice exhibit therapeutic potential in wound healing. Stem Cell Res. Ther. 11, 298 (2020).
Okizaki, S. et al. Vascular endothelial growth factor receptor type 1 signaling prevents delayed wound healing in diabetes by attenuating the production of IL-1beta by recruited macrophages. Am. J. Pathol. 186, 1481–1498 (2016).
Saidak, Z. et al. Wnt/beta-catenin signaling mediates osteoblast differentiation triggered by peptide-induced alpha5beta1 integrin priming in mesenchymal skeletal cells. J. Biol. Chem. 290, 6903–6912 (2015).
Mirotsou, M. et al. Secreted frizzled related protein 2 (Sfrp2) is the key Akt-mesenchymal stem cell-released paracrine factor mediating myocardial survival and repair. Proc. Natl. Acad. Sci. USA 104, 1643–1648 (2007).
Kasashima, H. et al. Stromal SOX2 upregulation promotes tumorigenesis through the generation of a SFRP1/2-expressing cancer-associated fibroblast population. Dev. Cell 56, 95–110.e110 (2021).
Sharma, M., Chuang, W. W. & Sun, Z. Phosphatidylinositol 3-kinase/Akt stimulates androgen pathway through GSK3beta inhibition and nuclear beta-catenin accumulation. J. Biol. Chem. 277, 30935–30941 (2002).
He, Y. et al. Targeting PI3K/Akt signal transduction for cancer therapy. Signal Transduct. Target Ther. 6, 425 (2021).
Pugh, C. W. & Ratcliffe, P. J. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat. Med 9, 677–684 (2003).
Forsythe, J. A. et al. Activation of vascular endothelial growth factor gene transcription by hypoxia-inducible factor 1. Mol. Cell Biol. 16, 4604–4613 (1996).
Vasioukhin, V., Degenstein, L., Wise, B. & Fuchs, E. The magical touch: genome targeting in epidermal stem cells induced by tamoxifen application to mouse skin. Proc. Natl. Acad. Sci. USA 96, 8551–8556 (1999).
Lee, D., Lim, J., Woo, K. C. & Kim, K. T. Piperonylic acid stimulates keratinocyte growth and survival by activating epidermal growth factor receptor (EGFR). Sci. Rep. 8, 162 (2018).
Landini, G., Martinelli, G. & Piccinini, F. Colour deconvolution: stain unmixing in histological imaging. Bioinformatics 37, 1485–1487 (2021).
Mascharak, S. et al. Preventing engrailed-1 activation in fibroblasts yields wound regeneration without scarring. Science 372, eaba2374 (2021).
Mascharak, S. et al. Multi-omic analysis reveals divergent molecular events in scarring and regenerative wound healing. Cell Stem Cell 29, 315–327.e316 (2022).
Ye, L., Wang, Z., Li, Z., Lv, C. & Man, M. Q. Validation of GPSkin Barrier((R)) for assessing epidermal permeability barrier function and stratum corneum hydration in humans. Ski. Res. Technol. 25, 25–29 (2019).
Deeds, M. C. et al. Single dose streptozotocin-induced diabetes: considerations for study design in islet transplantation models. Lab. Anim. 45, 131–140 (2011).
Le May, C. et al. Estrogens protect pancreatic beta-cells from apoptosis and prevent insulin-deficient diabetes mellitus in mice. Proc. Natl. Acad. Sci. USA 103, 9232–9237 (2006).
Graham, M. L., Janecek, J. L., Kittredge, J. A., Hering, B. J. & Schuurman, H. J. The streptozotocin-induced diabetic nude mouse model: differences between animals from different sources. Comp. Med. 61, 356–360 (2011).
Lee, D. et al. Diabetic sensory neuropathy and insulin resistance are induced by loss of UCHL1 in Drosophila. Nat. Commun. 15, 468 (2024).
McCarthy, D. J., Campbell, K. R., Lun, A. T. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Lun, A. T., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res 5, 2122 (2016).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Setty, M. et al. Characterization of cell fate probabilities in single-cell data with Palantir. Nat. Biotechnol. 37, 451–460 (2019).
Kang, M. et al. Improved reconstruction of single-cell developmental potential with CytoTRACE 2. Nat. Methods 22, 2258–2263 (2025).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Kolberg, L. et al. g:Profiler-interoperable web service for functional enrichment analysis and gene identifier mapping (2023 update). Nucleic Acids Res. 51, W207–W212 (2023).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Acknowledgements
We thank the members of Choi Laboratory for their helpful discussions and comments on this manuscript. This work was supported by National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIT) (NRF-RS-2025-00560319, NRF-RS-2022-NR072134, NRF-RS-2022-NR067309, NRF-RS-2023-00223298, and NRF-RS-2024-00433755 to S.C.; NRF-RS-2024-00426031 to J.K.K.; NRF-RS-2023-00221112 to J.K.K. and S.C.). S.C. has been supported by a POSCO Science Fellowship from the POSCO TJ Park Foundation. M.K., E.C., Y.J., B.K., C.L., J.L., J.K.K. and S.C. were supported by a BK21 FOUR Research Fellowship funded by the Ministry of Education, Republic of Korea.
Author information
Authors and Affiliations
Contributions
S.C., M.K., and Y.J. conceived the project. M.K., Y.J., and B.K. performed most of the experiments and quantified and analyzed the data. E.C. and J.K.K. performed the bioinformatic analysis. Y.H.C. generated RNA-seq libraries. C.L. performed flow cytometry. J.L. performed splinted full-thickness cutaneous wounding. J.H.L, K.-P.K., and B.-K.K. provided intellectual input. J.K.K. and S.C. acquired funding. M.K., Y.J., J.K.K., and S.C. wrote the manuscript, with discussion and feedback from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kwak, M., Choi, E., Jo, Y. et al. Mosaic partial epidermal reprogramming remodels neighbors and niches to refine skin homeostasis and repair. Nat Commun 17, 2191 (2026). https://doi.org/10.1038/s41467-026-69047-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69047-2