Abstract
β−1,3-glucan synthase is the molecular target for triterpenoid and echinocandin antifungal drugs in clinical. It catalyzes the formation of β−1,3-glucan, which is the primary component of the fungal cell wall. However, the inhibition mechanism of β−1,3-glucan synthase by triterpenoid drugs remains unclear. In this study, we report cryo-electron microscopy (cryo-EM) structures of Saccharomyces cerevisiae β−1,3-glucan synthase Fks1 and Fks2 in the apo state, the triterpenoid drug enfumafungin-bound state, and an open state. Structural analysis along with mutagenesis reveals the enfumafungin binding site, and the mechanism of the clinical drug-resistant mutations of the β−1,3-glucan synthases. Remarkably, the enfumafungin attaches on a single transmembrane helix TM5 of the β−1,3-glucan synthases, reorganizes its nearby lipid environment, and stabilizes the enzyme in a specific basal state with intact active site. Moreover, we elucidate that both the basal state and the open state are essential for FKS’s glycosyltransferase activity. Our research also shows that Fks2 is highly conserved with Fks1 in terms of structure, activity, and drug inhibition. These findings provide deep insights into the fungal cell wall synthesis, and will facilitate the development of antifungal drugs targeting β−1,3-glucan synthase.
Similar content being viewed by others
Introduction
Fungi are among the most diverse groups of organisms on Earth, with an estimated over 1.5 million species1. Among them, approximately 8000 species cause plant diseases, and more than 300 species cause human diseases2. Every year, diseases related to fungal infections affect approximately 300 million people worldwide, resulting in about 1.6 million deaths3,4. Due to the limited availability of clinical antifungal drugs and the continuous emergence of resistant pathogens, the expansion of the fungal infections necessitates urgent attention to developing distinct therapeutic strategies4,5,6.
β-1,3-glucan synthase (FKS) is a pivotal integral membrane enzyme in fungal cells, catalyzing the synthesis of β-1,3-glucan, a fundamental component of the fungal cell wall that contributes to cell integrity and viability7,8. β-1,3-glucan synthase belongs to the glycosyltransferase 48 (GT48) family (http://www.cazy.org). Previous studies have also shown that Rho1 is essential for activity of the 1,3-β-glucan synthase both in vivo and in vitro9,10,11,12,13,14. Using uridine di-phosphate-activated glucose (UDP-Glc) as the sugar donor, β-1,3-glucan synthase facilitates the creation of β(1 → 3) glycosidic linkages in β-1,3-glucan and transports the glucan across the membrane (Fig. 1a). Fungal β-1,3-glucan synthase is primarily encoded by FKS1 or FKS27. Expression and function of the two genes have been shown to be differential regulated7,15,16,17,18,19. In S. cerevisiae, FKS1 is the predominant isoform during growth, whereas Fks2 is expressed during sporulation or under stress20. S. cerevisiae Fks1 and Fks2 share ~88% sequence identity. In S. cerevisiae, Candida albicans and many other pathogenic fungi, simultaneous knockout of FKS1 and FKS2 is lethal, while knockout of FKS1 alone is alive8,21,22. Mutations in both Fks1 and Fks2 have been reported to be associated with drug resistance10,11,12,13,14. However, whether Fks2 adopts a similar mechanism to Fks1 is unknown yet.
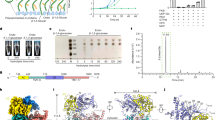
a Cartoon schematic of β-1,3-glucan synthase. b In vitro UDP-Glo glycosyltransferase assay with purified Fks1 and Fks2 with Rho1. The reaction without FKS was used as the control. Data points represent the mean ± SD (n = 3). c Cryo-EM map and atomic model of endogenous Fks1. d Cryo-EM map of endogenous Fks2. e Structural comparison between Fks1-apo (green) and Fks2-apo (blue). f Topological model of Fks1 and Fks2. g Close-up view of the region around HH1A-1B. h Close-up view of the highlighted box region in panel e. The extracellular loop between TM1 and TM2 directly interacts with the extracellular loop between TM7 and TM8 at the outer side of the plasma membrane. Source data are provided as a Source Data file.
So far, structures of S. cerevisiae Fks1 in the apo state or in complex with Rho1, and an echinocandin-resistant mutant Fks1 (S643P) in complex with glucan product have been determined14,23,24. Among them, the cryo-EM structure of Fks1 in complex with Rho1 showed that Rho1 is positioned within a pocket between the glycosyltransferase domain of FKS1 (GT domain) and the transmembrane helix spanning TM7–15. Structural analysis suggested Rho1’s GTP/GDP cycling may promote a dynamic transition between different states of FKS1, inducing the protein as a molecular pump to push the growing glucan chain into FKS1’s transmembrane channel. However, the active sites in all available Fks1 structures were not fully resolved, as many important elements (such as the conserved horizontal interface helices IF1/2/3 that were originally characterized in cellulose synthase25) are partially or totally invisible due to flexibility compared to the AlphaFold2 predicted model (Supplementary Fig. 1a–h). This has significantly hindered our understanding of the mechanism of β-1,3-glucan synthase.
Currently, there are two types of clinical antifungal drugs targeting β-1,3-glucan synthase, including triterpenes (ibrexafungerp) and echinocandins (caspofungin, micafungin, anidulafungin, and rezafungin)22,26,27,28,29. Ibrexafungerp is a semisynthetic analog of the natural triterpene enfumafungin29,30,31. It was also an FDA (U.S. Food and Drug Administration) approved drug in a distinct antifungal class for more than 20 years29. Notably, triterpenes were highly active for many Fks1 mutant based echinocandin resistant strains, indicating its different inhibitory mode comparing to echinocandins32,33,34. Despite its importance, structural insights into the binding mode and mechanism of the triterpene drugs remain unclear.
Here, we determined five different cryo-electron microscopy (cryo-EM) structures of endogenous Fks1 and Fks2 in the apo state, in complex with enfumafungin, and an open state induced by echinocandin incubation. These structures illustrate the enfumafungin binding site and an intact active site of β-1,3-glucan synthase, and depict the conformational change induced by echinocandin incubation. Our results provide deep insights into the pharmacological mechanisms of β-1,3-glucan synthases.
Results
Structure and characterization of endogenous Fks1 and Fks2
Endogenous Fks1 and Fks2 were both purified from yeast cells with C-terminal triple FLAG epitope tags. Because S. cerevisiae predominantly expressed Fks1 in regular condition8, the endogenous Fks2 was purified from a FKS1 knock out strain. The mild detergent N-dodecyl-β-D-maltoside (DDM) and cholesteryl hydrogen succinate (CHS) were used to extract protein from yeast membrane, and lauryl maltose neopentyl glycol (LMNG) and CHS were used in the Cryo-EM analysis. The tagged proteins were purified using an anti-FLAG (M2) affinity column and a size-exclusion chromatography (SEC) (Supplementary Fig. 2a, b). Functional characterization with in vitro UDP-Glo glycosyltransferase assay confirmed that endogenous Fks1 and Fks2 both showed glycosyltransferase activity in the presence of Rho1 and GTPγS (Fig. 1b, Supplementary Fig. 2a, b). It indicates Fks2 also uses Rho1 as its regulatory subunit, like Fks1. To confirm this finding, we incubated purified Fks2 and Rho1 with crosslinker BS3 and separated the crosslinked complex using a size exclusion column. The shifted position of the gel filtration peak indicated the complex of Fks2-Rho1 was successfully purified (Supplementary Fig. 3a). We further performed EM analysis on this crosslinked complex, and the 2D class averages clearly showed additional density corresponding to Rho1 on Fks2 (Supplementary Fig. 3b). The detected binding position of Rho1 is consistent with the published structure of Fks1–Rho1 complex14. Together, these results support that Rho1 is a regulatory subunit for both Fks2 and Fks1.
We performed single-particle cryo-EM on purified samples and obtained cryo-EM 3D maps of endogenous Fks1 and Fks2 in apo states at 3.2-Å and 3.6-Å resolution, respectively (Fig. 1c, d, Supplementary Figs. 4 and 5). Atomic models of endogenous Fks1 and Fks2 (named Fks1-apo and Fks2-apo) were built into the maps and refined to good statistics (Supplementary Table 1). The overall structures of Fks1-apo (UniProt code P38631, 1876 residues) and Fks2-apo (UniProt code P40989, 1895 residues) are highly similar, with a root-mean-square deviation (RMSD) of approximately 1.5 Å over 1449 residues (Fig. 1e), which aligns with their ~88% sequence identity. In both Fks1-apo and Fks2-apo structures, FKS is composed of a transmembrane domain (TMD) and a large cytosolic domain (Fig. 1c–e). The TMD comprises 15 transmembrane helices (TM) and four amphipathic horizontal helices (HH1A, 1B, 2A, 2B), which can be grouped into two parts (TMD1 and TMD2) with a sizable cavity between them. The cytosolic domain consists of two subdomains including an N-terminal domain (NTD; Fks1: Asn145 to Thr444; Fks2: Asn164 to Ser463) and a glycosyltransferase (GT) domain inserted between TM6 and TM7 (Fks1: Ile713 to Gln1281; Fks2: Ile732 to Gln1300) (Fig. 1f). The active site locates between GT and TMD, and the glucan transport channel is likely right below the active site and corresponds to the cavity between TMD1 and TMD2.
Furthermore, structures of Fks1-apo and Fks2-apo display several features that were not resolved before. We identified an element, HH1B, that forms extensive hydrophobic interactions with TM9, TM14, and HH2A (Fig. 1g). These interactions help to improve the TMD2 to be more rigid. HH1B is connected with HH1A by a short loop (HH1-loop) (Fks1: Arg1512 to Ser1521; Fks2: Arg1531 to Ser1540), which is partially flexible in Fks1 but ordered in Fks2. The HH1-loop in Fks2 is stabilized by a short helix right before HH2A. In addition, the exoplasmic loop connecting TM1 and TM2 was modeled in Fks2, and it interacts directly with the exoplasmic loop connecting TM7 and TM8, resulting in a more ordered exoplasmic structure (Fig. 1h).
Enfumafungin binding site and resistant mutations of FKS
Enfumafungin (EFU) is a triterpene glycoside that was produced from the fungus Hormonema sp30,31. It features four six-membered rings with four additional specific groups: glucose unit, acetate group, hemiacetal moiety, and carboxylic acid group (Fig. 2a)35. Previous studies showed EFU displayed high antifungal activity with <1 μg/ml minimal inhibitory concentrations (MIC) against various fungal pathogens, and its development has led to the semi-synthetic clinical drug, ibrexafungerp22,29. We determined the MIC for EFU in S. cerevisiae to be ~0.07 μg/ml (Fig. 2b). To understand how EFU interacts and inhibits FKS, we determined the cryo-EM structures of Fks1 in complex with EFU (Fks1–EFU) and Fks2 in complex with EFU (Fks2–EFU) at a resolution of 3.0-Å and 3.3-Å, respectively (Fig. 2c, d, Supplementary Figs. 6 and 7, Table 1). The map revealed an obvious elongated extra density around the TM5 of Fks1 and Fks2 in the transmembrane region, which aligns well with the EFU molecule (Fig. 2c, d).
a Molecular formula of enfumafungin. b MIC of enfumafungin for S. cerevisiae. Cell viability assay of yeast in medium containing enfumafungin in a series of concentrations, indicating the MIC of enfumafungin at 0.072 μg/ml. c Cryo-EM maps of Fks1 in complex with enfumafungin (Fks1–EFU). The electron density of enfumafungin is shown as a mesh, and the structure is represented as sticks. d Cryo-EM maps of Fks2 in complex with enfumafungin (Fks2–EFU). The electron density of enfumafungin is shown as a mesh, and the structure is represented as sticks. e Interaction network between EFU and Fks1. Key residues involved in the interaction are shown as sticks. f Interaction network between EFU and Fks2. Key residues involved in the interaction are shown as sticks. g Relative position of the EFU to the active site and glucan transport channel in a model of Fks1–EFU in complex with Rho1 and β-1,3-glucan. Structural coordinates for Rho1 were obtained from PDB 8WLA, and the 1,3-β-glucan product model was derived from PDB 7YUY. The EFU-binding pocket is positioned approximately 30 Å from the catalytic site of Fks1 and roughly 20 Å from the predicted glucan translocation channel. The top panel presents a cartoon and semi-transparent surface representation of the Fks1–EFU structural model. The bottom panel offers a zoom-in view of the EFU-binding site, the active site, and the 1,3-β-glucan chain. h UDP-Glc hydrolysis activity of wild type (WT) and mutant Fks1 with or without enfumafungin. The reaction without Fks1 was used as the control (Ctr). Data points represent the mean ± SD (n = 3). i MIC50 for EFU in wild-type S. cerevisiae cells (Ctl) and fks1Δ cells with a plasmid carrying either wild-type Fks1 (WT) or mutants. Data points represent the mean ± SD (n = 3). Source data are provided as a Source Data file.
In structures of both Fks1–EFU and Fks2–EFU, EFU was found stacking against the TM5 and embedding in the exoplasmic leaflet of the lipid bilayer in a manner of ergosterol (Fig. 2e, f). The EFU molecule locates ~30 Å from the active site, and ~20 Å from the putative β-1,3-glucan-transporting channel of Fks1 (Fig. 2g). The hydrophobic six-membered rings are mainly stabilized by Y638 and F639 in Fks1 (F657 and F658 in Fks2). Besides, the carboxylic acid function of EFU interacts directly with E635 and W695 of Fks1 (E654 and W714 of Fks2). In contrast, the hydrophilic glucose and acetate group of EFU has limited contact with FKS. Specifically, the glucose group is relatively more flexible according to its low density. It is consistent with previous findings that various EFU analogs with modification on these two groups still highly effective36,37.
To confirm the EFU binding site, we introduced two mutations in the binding site of Fks1 (E635A and F639S) and subsequently purified the mutant proteins for an in vitro UDP-Glo glycosyltransferase assay. We found that the enzymatic activities of E635A and F639S were unable to be inhibited by the EFU as the wild-type (WT) enzyme (Fig. 2h), indicating their essential role for EFU binding. To further evaluate the function of key residues in the EFU binding site, we generated a group of yeast fks1Δ strains with plasmid containing wild-type Fks1 or mutants, and measured their MIC values of EFU (Fig. 2i). Our results showed E635A, F639S, F639A, and W695L mutants all displayed increased MIC values at >30 folds of that of wild-type Fks1, indicating their crucial role in EFU binding; L642Y and R645F mutants displayed increased MIC values at 2.5–5 folds of that of wild-type Fks1, indicating their moderate role in EFU binding. In contrast, the detected MIC value of A631S, S634A, and F638A closely approximate that of wild-type, suggesting these residues may not be important for EFU’s function. Furthermore, we performed extensive molecular dynamics (MD) simulations of the wild type and mutant Fks1 in complex with enfumafungin embedded in a lipid bilayer (Supplementary Fig. 8). Four EFU resistant mutations E635A, F639A, F639S, and W695L were tested, and the RMSD values of EFU calculated relative to the starting structure were used to explore its binding stability in the active sites of Fks1. The results revealed that the EFU is more dynamic in the mutants compared to WT, while the backbones of all residues in both WT and mutant Fks1 are stable. The differences indicate that these drug resistance mutant sites are crucial for EFU binding. A group of ibrexafungerp-resistant mutations has been identified in previous studies, including F641, S645 of Candida albicans FKS1 and E655, F659, S663, W715 of Candida glabrata FKS2 (Supplementary Table 2)32,34,38,39. Most of these residues or the corresponding residues in FKS1/2 are located around the EFU molecule in Fks1–EFU and Fks2–EFU structures, further confirming our finding of the EFU binding site (Fig. 2e, f, Supplementary Table 2).
Coincidentally, the EFU binding site is largely overlapped with the echinocandin resistance “hot spot” regions (HS1: 635–649, HS2: 1354–1361, and HS3: 690–700)40,41. This is consistent with previous findings that some mutations are resistant to both ibrexafungerp and echinocandins, such as F641, S645 of Candida albicans FKS1 and E655, F659, W715 of Candida glabrata FKS232,34. Our result also confirmed that the most frequently observed echinocandin-resistant mutation, S643P, displayed increased MIC values of enfumafungin at ~15 folds of that of wild-type Fks1 (Fig. 2i). Meanwhile, due to a different binding mode, some echinocandin-resistant mutations, such as R631, D632 of Candida glabrata FKS1, were shown to be sensitive to Ibrexafungerp33.
Remodeling of the lipid environment surrounding FKS by Enfumafungin
To understand the molecular mechanism by which EFU inhibits FKS, we superimposed the Fks1/2 structures in EFU bound state and apo state, and found their overall structures are almost identical (Fig. 3a). But compared to the apo structures, the phenyl groups of F639 in Fks1–EFU and F658 in Fks2–EFU rotate toward EFU molecule and clamp the EFU together with the phenyl groups of Y638 in Fks1 and F657 in Fks2 as a vice (Fig. 3b). Interestingly, movement of F639 of Fks1 or F658 of Fks2 stabilizes a specific ordered lipid molecule in the groove between TM5 and TM8 of Fks1/2-EFU, which was not available in the apo Fks1/2 (Supplementary Fig. 9a). The lipid is in parallel with the EFU and directly interacts with many residues including F639 of TM5, W695 of TM6, L1360 and I1364 of TM8. Among them, F639 and W695 have been shown to affect the function of EFU (Fig. 2e).
a Structural comparison between Fks1–EFU (blue), Fks1–apo (gray) and Fks2-EFU (green). b Zoom in view of the EFU binding site in panel (a). c Ordered lipid densities around EFU, TM8 linker, and HH2B in cryo-EM maps of Fks1–EFU–Lip and Fks2–EFU–Lip. d Interaction network between ordered ergosterols and Fks1. Key residues involved in the interaction are shown as sticks. e Mapping of reported ibrexafungerp-resistant mutations in Fks1–EFU structure. The Cα atoms of ibrexafungerp-resistant mutations summarized in Supplementary Table 2 are shown as salmon spheres. f Growth complementation of fks1Δ cells with empty plasmid (fks1Δ), or plasmid carrying either WT Fks1 (FKS1) or mutants. Cells were serially diluted, spotted onto synthetic Histidine-dropout medium plates (SD-His) and SD-His plates with 0.1 μg/mL FK506, and incubated at 30 °C for 2 days. Source data are provided as a Source Data file.
Furthermore, a group of additional ordered densities corresponding to lipids were identified around TM8–IF3 linker and HH2B in Fks1/2-EFU structures, likely caused by the presence of EFU (Fig. 3c). To better present these densities, we generated refined 3D maps of Fks1 and Fks2 in complex of EFU at 3.3 Å and 3.4 Å resolution respectively by further 3D classification and local refinement (Fks1–EFU–Lip and Fks2–EFU–Lip) (Supplementary Figs. 6 and 7). The long oblate shapes of these lipid densities suggest them to be sterols (Fig. 3c). Remarkably, two sterol molecules interwind with each other, and stack tightly with W1773 of HH2B and Q1388 of TM8-IF3 linker (Fig. 3d). In vivo growth complementation assay of W1773A showed the mutant was unable to rescue fks1Δ yeast growth, supporting its essential functional role (Fig. 3f). Moreover, the sterol molecules also interact directly with several key EFU resistant mutants in the Hot-Spot 2, such as F1366, F1370, and L1378 (Fig. 3d, e, Supplementary Table 2). It probably explains how the mutants in Hot Spot 2 affect the inhibitory function of antifungal drugs while they are at a distance from the EFU binding site.
Intact active site and glucan transport channel in the structures of Fks1/2 in complex with EFU
Previous structural studies revealed that the central catalytic region of Fks1 adopts a cellulose synthase fold with a cytosolic conserved GT-A type glycosyltransferase domain23,24. However, several essential motifs identified in cellulose synthase were either invisible or in different conformations in Fks1/2 structures, such as interface helix IF1/2/3, indicating the active site of FKS to be highly dynamic (Fig. 1, Supplementary Fig. 1). Therefore, it is still unclear whether the FKS catalyzes following a similar mechanism as cellulose synthases25,42,43.
Interestingly, well-ordered IFs were observed in the structures of Fks1–EFU–Lip and Fks2–EFU–lip (Fig. 4a, Supplementary Fig. 9b). A conserved R–T/S–W motif (R443–T444–W445 in Fks1 and R462–S463–W464 in Fks2) and the sterol molecules mentioned above play important role to stabilize IF2 and IF3 (Fig. 4a, b). The R–T/S–W motif is located right before TMH1 (W445–Y469 in Fks1). The side chains of T444 and W445 face the active site on the right in the apo state but turn left in the Fks1–EFU–Lip state, flipping by nearly 180°. Conversely, the side chain of W445 faces leftward in the apo state but switches to the right in the Fks1–EFU–Lip state. Specifically, R443 of Fks1 (R462 in Fks2) interacts directly with D1416 of IF2 and E1268 of IF3 by salt bridges, while the side chain of W445 and a sterol molecule interact with L1414 and L1417 of IF2 by hydrophobic interactions. A horizontal sterol molecule below this motif and IF3 further stabilizes this architecture by forming extensive interactions (Fig. 4b). Notably, we found that R443A, D1416A, and E1268A mutants were all essential for Fks1’s function because they were unable to rescue fks1Δ yeast growth like wild-type Fks1 (Fig. 4d).
a Structure and electron density of IF2/3 and surrounding residues and sterols in Fks1–EFU–Lip. The black boxes mark the positions for cut-in views shown in panels (b) and (c). b Comparison between the RT/SW motif region of Fks1–apo (green) and Fks1–EFU–Lip (blue). Side chains of the motif are flipped nearly 180°. c Close-up view of the channel entry of Fks1 viewed from the cytosol. Key residues are shown in stick representation. d Growth complementation of fks1Δ cells with empty plasmid (fks1Δ), or plasmid carrying either WT Fks1 (FKS1) or mutants. Source data are provided as a Source Data file.
Superposition of Fks1/2–EFU–Lip and cellulose synthase revealed that the position of their IFs and active sites were quite similar to each other, while their putative glucan transport channels were only partially similar (Supplementary Fig. 10a–i). Structure-based sequence alignment also showed low similarity with only a few conserved residues identified, such as the magnesium coordinating residue D1102 and the phosphate coordinating residue R1261 of Fks1 (Supplementary Fig. 10j). This is consistent with the sequence conservation analysis of Fks1 among yeast and major pathogenic fungal species, which showed high conservation in GT but relatively lower conservation in transport channel (Supplementary Fig. 11). The putative glucan transport channel of FKS were mainly composed of TM5-10 and IF1-3. Similar to the structures of cellulose synthases, ordered IF1 and IF2 in Fks1/2–EFU–Lip constitute the entry of the channel (Fig. 4c). Our previous studies have performed mutations for most residues in the active site and confirmed their important roles23. Here, we introduced more mutations in IF2, IF3, and other residues in the glucan transport channel, and performed the growth complementation assay (Fig. 4d). The results showed mutation of key residues of IF2 (F1258A, T1259A, and K1261A) in the channel entry are essential for FKS’s function, while most residues inside of the channel (E1402A, V1403A, Q1407A, Y1409A, S1410A of IF3, Q1305 of TM7) are relatively less important.
Although the corresponding regions of IF3 and IF3–TM8 linker were identified as TM9–10 by inserting into the membrane in the previously reported structure of Fks1 in complex with glucan product24, these two elements in our structures are located in a similar position as many other membrane-integrated polysaccharide synthases, including cellulose synthase, chitin synthase, and hyaluronan synthase25,44,45. The differences may indicate the FKS has a dramatic conformational change during the glucan transport. The exporting path of glucan to extracellular in FKS is likely through a lateral opening, and mainly formed by TM8, TM10 and IF3 in our structures (Fig. 3d). The growth complementation assays for mutations of key residues in the exporting path (F1363, F1366 in TM8 and W1474 and S1478 in TM10) showed each single mutant only partially affects Fks1’s function, and may work together for glucan transporting (Fig. 4d). Furthermore, because the TM8–IF3 linker and top parts of TM8 are amphipathic helices that half embedded in the membrane, the transmembrane region adjacent TM8 in our structures is only half as thick as normal lipid bilayer, which favors the glucan exporting (Supplementary Fig. 9b).
Discovery of an open state of Fks1
Caspofungin(CPF) is the first marketed echinocandin antifungal agent that specifically inhibits β-1,3-glucan synthase46. To gain insights into the inhibitory mechanism of echinocandin, we performed single-particle cryo-EM analysis on Fks1 incubated with 2 mM CPF overnight at 4 °C, and obtained the cryo-EM 3D map of Fks1 in a resolution of 3.7-Å (Supplementary Fig. 12). Unfortunately, no clear extra density corresponding to CPF was identified in the putative echinocandin binding site, which was proposed based on echinocandin-resistant mutations, indicating that the structure remains in an apo state. Interestingly, this structure showed a more open active site (named Fks1–open), whose architecture differs from that of Fks1 in either the apo or EFU-bound states (Fig. 5a).
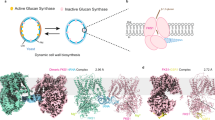
a Structural comparison between Fks1–open (color) and Fks1–apo (gray). The TMD2 shifts in the membrane by ~5 Å. b A close-up view of the channel entry region in Fks1–open viewed from the cytosol. c Superimposed channel entry regions of Fks1–EFU and Fks1–apo in the same view as panel (b). d Growth complementation of fks1Δ cells with empty plasmid (fks1Δ), or plasmid carrying either WT Fks1 (FKS1) or mutants. Cells were serially diluted, spotted onto synthetic Histidine-dropout medium plates (SD-His) and SD-His plates with 0.1 μg/mL FK506, and incubated at 30 °C for 2 days. e In vitro UDP-Glo glycosyltransferase assay of wild-type Fks1 and Fks1 mutants. The reaction without FKS was used as the control. Data points represent the mean ± SD (n = 3). Source data are provided as a Source Data file.
Structural alignment between Fks1–open and Fks1–apo revealed dramatic conformational changes in both TMD2 and the IF1–3 (Fig. 5a). Some important residues in the active site of GTD were also shifted in Fks1–open, such as N1104, E1155, H1195, K1246, and R1248. Compared to the Fks1–apo, the whole TMD2 in Fks1–open rotated and moved by ~5 Å in relative to TMD1, resulting in a more open glucan transport channel between TMD1 and TMD2; IF1 and TM8–IF3 linker rotate ~90° toward outside, while IF3 is shifted by ~20 Å accordingly, generating a more open active site (Fig. 5b, c). Therefore, we named the conformation of Fks1/2–apo and Fks1/2–EFU the “basal state” as they resemble other polysaccharide synthases, such as cellulose synthase25,44,45. In fact, we have further processed a cryo-EM dataset of Fks1 in the apo state by focused 3D classification in cryoSPARC and are able to separate a small subset of particles (9.8%) adopting a conformation alike the open state (Supplementary Fig. 13). This suggests that Fks1 in both the basal and open state are available in solution. While the basal state accounts for the majority, the overnight incubation of Fks1 probably facilitates the conformational change from the basal to the open state. Because these moved elements in Fks1–open have been shown to be directly involved in active site or glucan transporting, the conformational change of Fks1 between basal and open states is clearly linked to its function.
Implications for two different states of FKS
Interestingly, the structure of Fks1–open is similar with the reported structures of Fks1 that was purified in glyco-diosgenin (GDN) (Supplementary Fig. 1). Although their TMD2 are in different positions, IF1/3, part of TM8, and TM8–IF3 linker of these structures are all locating out of the core domain and the whole structures are in the open state (Supplementary Fig. 1). Investigation on all available FKS structures showed that Fks1/2 in basal state was captured in buffer containing LMNG/CHS, while the open state was captured in buffer containing GDN. As a cholesterol analog, CHS closely mimics the property of cholesterol/ergosterol in native membranes, and is commonly used to maintain the native structure of membrane protein during purification47,48. We noticed that a sterol molecule was identified under TM8–IF3 linker of Fks1 in the basal state, but disappeared in the structure of Fks1 in the open state (Fig. 5c). The sterol interacts directly with TM8, TM10 and TM8–IF3 linker, and likely plays an important role in stabilizing the FKS in the basal state. Overnight placement or missing CHS in the solution may release the sterol from the site and thus induce the conformational change from the basal state to the open state.
Notably, although the FKS purified in LMNG-CHS or GDN were captured in different states, the protein in both detergents was proved to be active23,24. Therefore, the basal and open states of Fks1 are both likely to be intermediate states that occur during catalysis. In other words, a flexible and dynamic active site identified in the above structures may be essential for the catalysis of FKS. To confirm this hypothesis, we generated two Fks1 double mutants: A1264C–D1416C and S1415C–A1485C, which are supposed to fix Fks1 in basal or open architectures, respectively, by forming a disulfide bond (Fig. 5b, c). In vivo growth complementation assays of the Fks1 double mutants showed that neither of them was able to rescue fks1Δ yeast growth on plates containing FK506 like the single mutants or wild type Fks1 (Fig. 5d). We also purified A1264C–D1416C and S1415C–A1485C, and measured their in vitro activities (Fig. 5e). The results showed the enzymatic activities of both double mutants were dramatically reduced when compared with wild type Fks1. Taken together, these results support that both basal and open states of Fks1 are important for its function.
Discussion
Although the structures of Fks1 in apo and glucan binding states have been determined before, the catalytic and inhibitory mechanisms of Fks1 and Fks2 remain unclear. Here we have determined five cryo-EM structures of Fks1 and Fks2 in the apo state, in complex with the triterpenoid drug enfumafungin, and an essential open state for FKS’s function at 2.97–3.72 Å resolutions. It should be noted that side-chain placement is well supported by our map densities at current resolutions, whereas specific atomic interactions mentioned above are mainly inferred.
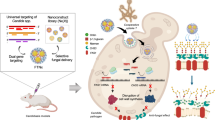
Based on the structures and functional studies, we proposed a working model of FKS and its inhibitors (Fig. 6). With the help of Rho1, FKS catalyzes the formation of glucan by conformational change between the basal and open states. In the basal state, the active site of FKS with flexible IFs is dynamic for the initiation of glucan synthesis. A sterol molecule helps FKS stabilize in this state. The glucan elongation triggers the movement of the IF1/3 out of the core domain and leads to the thinning of the membrane and the opening of the lateral channel exit. EFU fixes the enzyme specifically in the basal state by stabilizing sterols around the channel exit and IFs. Capturing FKS in complex with substrates and product will undoubtedly provide more insights into this model.
With the help of Rho, FKS catalyzes the initiation and elongation of glucan by conformational change between the basal and open states. EFU inhibits FKS by fixing the enzyme specifically in the basal state. More details are provided in the text.
Sterols were previously shown to help the stabilization of the overall Fks1 structure23. Here, we further proposed that sterols are involved in the function of enfumafungin targeting Fks1 and Fks2. Considering the antifungal azole drugs work by inhibiting the sterol biosynthesis, it would be interesting to investigate the effect of the combined use of azole and echinocandins or ibrexafungerp. Most previous studies showed azoles have neither synergistic nor antagonistic interactions with echinocandins or ibrexafungerp in fungal pathogens49,50,51, while some literature showed that the combination of ibrexafungerp plus azoles was synergistic for the tested Aspergillus fumigatus strains in vitro and in animal models52,53. Although sterols are involved in the structural stabilization of the Fks1/2 and triterpenes, whether the azole-mediated inhibition of sterol biosynthesis directly affects the function of FKS and triterpene drugs still awaits future research.
Identification of the EFU binding site in this work provides a distinct pocket for virtual screening of inhibitors targeting FKS. In the predicted structures of S. cerevisiae and C. albicans Fks1 in complex with EFU by AlphaFold3, EFU was wrongly placed in the active site, indicating the EFU binding site is different from the known binding mode in the AlphaFold training database (Supplementary Fig. 14). Typical small-molecule drug binding sites of the target protein usually resemble pockets or grooves. However, the triterpenoid drug binding site of FKS identified in this work is mainly on the surface of one transmembrane helix of the membrane-embedded enzyme. Instead of inhibiting the activity of FKS, binding of the drug affects the function of FKS by disturbing the lipid environment. Therefore, our work probably indicates a mode of drug action that can be used for drug development on “undruggable” target proteins lacking deep pockets for ligand interaction.
In S. cerevisiae, C. albicans and many other pathogenic fungi, FKS1 or FKS2 alone is not essential, but their simultaneous knockout leads to lethal8,21,22. Furthermore, mutations in both Fks1 and Fks2 have been reported to be associated with echinocandin and triterpene drug resistance. These suggest the drugs work by inhibiting both Fks1 and Fks2 in the cells. This study provided the structural insights into Fks2. The conserved structures between Fks1 and Fks2 indicated that the echinocandin can inhibit both enzymes in a similar mechanism, and explained the overlap of echinocandin-resistant mutations in Fks1 and Fks2. The structures of enfumafungin-bound Fks1 and Fks2 also support that the two enzymes share the same inhibition and drug-resistant mechanisms of triterpenes.
In summary, our findings shed light on the molecular basis of the catalytic cycle, activation by Rho1, and inhibition by triterpenoids and echinocandins of β-1,3-glucan synthase. Our work provides a platform for optimization of current triterpenoid and echinocandin antifungal drugs, and rational design of potent drugs that target β-1,3-glucan synthase.
Methods
Expression and purification of endogenous Fks1 and Fks2
The C-terminal of FKS1 in the yeast strain W303 (EUROSCARF) was engineered with a triple-FLAG and SpHis5 using a PCR-based genomic epitope-tagging method. Using the same method, FKS2 was engineered in an FKS1 knockout W303 strain. The yeast strains were cultured in YPD medium (10 g/L yeast extract, 20 g/L peptone, and 20 g/L glucose) for approximately 20 h before harvesting. Cell pellets were resuspended in lysis buffer containing 20 mM Tris-HCl (pH 7.4), 0.2 M sorbitol, 50 mM potassium acetate, 2 mM EDTA, and 1 mM phenylmethylsulfonyl fluoride (PMSF). Cells were lysed using a French press at 15,000 psi. The lysate was subjected to centrifugation at 10,000×g for 30 min at 4 °C, and the supernatant was further centrifuged at 100,000×g for 60 min at 4 °C. The membrane pellet was collected and resuspended in buffer A, which consisted of 10% glycerol, 20 mM Tris-HCl (pH 7.4), 1% n-dodecyl-β-D-maltoside (DDM), 0.1% cholesteryl hydrogen succinate (CHS), 500 mM NaCl, 1 mM MgCl2, 1 mM EDTA, and 1 mM PMSF. After incubation at 4 °C for 30 min, the sample was centrifuged at 100,000×g for 30 min to remove insoluble material. The solubilized membrane fraction was applied to a pre-equilibrated anti-FLAG M2 affinity column (GenScript) at 4 °C. After extensive washing with buffer B (20 mM HEPES, pH 7.4, 150 mM NaCl, 0.01% lauryl maltose neopentyl glycol (LMNG), 0.001% CHS, and 1 mM MgCl2), the bound proteins were eluted using buffer B supplemented with 0.15 mg/mL 3× FLAG peptide. The eluted fraction was further purified via size-exclusion chromatography using a Superose 6 10/300 Increase column equilibrated in buffer C (20 mM HEPES, pH 7.4, 150 mM NaCl, 0.003% LMNG, 0.0003% CHS, and 1 mM MgCl2). The purity of the final protein preparation was assessed by SDS-PAGE, and the protein was concentrated for subsequent cryo-electron microscopy (cryo-EM) analysis.
Expression and purification of Rho1
The cDNA encoding RHO1 was amplified from the genome of Saccharomyces cerevisiae strain W303 and subcloned into the pFastBac vector with an N-terminal 6×His tag. A mutation (Q68H) was introduced to generate an active form of Rho1 using a PCR-based site-directed mutagenesis strategy. The Bac-to-Bac baculovirus expression system in Spodoptera frugiperda cells (Gibco Sf9 cells, cat. no. 11496015) was used for protein expression. SF9 cells were infected at a density of 2 × 106 cells/mL and harvested at 60 h post-infection. Collected cells were resuspended in Buffer A (20 mM Tris-HCl, pH 7.4, 500 mM NaCl, 10% glycerol, and 1 mM MgCl₂) and lysed on ice by sonication. Cell debris was removed by centrifugation at 15,000×g for 30 min at 4 °C, and the supernatant was applied to a 2 mL Nickel–NTA column. The column was washed with buffer containing 20 mM HEPES (pH 7.4), 20 mM imidazole, and 150 mM NaCl, and the bound protein was eluted using the wash buffer supplemented with 200 mM imidazole. The eluted protein was concentrated and further purified by size-exclusion chromatography using a Superose 6 10/300 GL column (GE Healthcare). Peak fractions were collected and used for subsequent functional assays.
Cryo-electron microscopy
Aliquots (2.5 μL) of Fks1 and Fks2 alone or in complex with EFU at a concentration of approximately 3 mg/mL were applied to glow-discharged holey carbon grids (Quantifoil Au R1.2/1.3, 300 mesh) and flash-frozen in liquid ethane using a FEI Vitrobot Mark IV. Grid screening was performed on a 300-kV FEI TF30 electron microscope, and Cryo-EM data were automatically collected using EPU on either a 200-kV Thermo Fisher Scientific Glacios2 or a 300-kV Thermo Fisher Scientific Krios G4 electron microscope. Data acquisition was carried out with defocus values ranging from −1.0 to −2.0 μm. The microscope was operated with a K3 direct electron detector at a nominal magnification of 130,000×. The total electron dose was set to 40–60 electrons per Ų at the sample level.
Cryo-EM image processing
All data processing steps were performed using CryoSPARC. The collected movies were motion-corrected, and contrast transfer function (CTF) parameters were estimated. Micrographs were then filtered using exposure curation. Particle picking was conducted with a template-based picker using manually selected 2D class averages as templates, followed by particle extraction. After 2D classification and Ab-initio reconstruction, selected particles were further refined using non-uniform refinement.
For the structure determination of apo-state Fks1, a total of 6816 raw movie micrographs were collected. Automated particle picking yielded 401,620 particles, which were subjected to 2D classification and heterogeneous refinement. Based on the quality assessment of three heterogeneously refined 3D maps, 248,992 particles were selected for subsequent refinement and post-processing. The final reconstruction, performed with C1 symmetry, resulted in a 3.23 Å resolution 3D map.
For the structure determination of EFU-bound state Fks1, a total of 6457 raw movie micrographs were collected. Automated particle picking yielded 479,938 particles, which were subjected to 2D classification and heterogeneous refinement. Based on the quality assessment of three heterogeneously refined 3D maps, 301,005 particles were selected for subsequent refinement and post-processing. The final reconstruction, performed with C1 symmetry, resulted in a 2.97 Å resolution 3D map. After further 3D classification and local refinement, improved 3D maps of Fks1with ordered IFs were obtained at 3.3 Å resolution (Fks1–EFU–Lip).
For the structure determination of open-state Fks1, a total of 4402 raw movie micrographs were collected. Automated particle picking yielded 616,794 particles, which were subjected to 2D classification and heterogeneous refinement. Based on the quality assessment of three heterogeneously refined 3D maps, 142,777 particles were selected for subsequent refinement and post-processing. The final reconstruction, performed with C1 symmetry, resulted in a 3.72 Å resolution 3D map.
For the structure determination of apo state Fks2, a total of 6588 raw movie micrographs were collected. Automated particle picking yielded 621,453 particles, which were subjected to 2D classification and heterogeneous refinement. Based on the quality assessment of three heterogeneously refined 3D maps, 97,369 particles were selected for subsequent refinement and post-processing. The final reconstruction, performed with C1 symmetry, resulted in a 3.64 Å resolution 3D map.
For the structure determination of EFU-state Fks2, a total of 9020 raw movie micrographs were collected. Automated particle picking yielded 510,461 particles, which were subjected to 2D classification and heterogeneous refinement. Based on the quality assessment of three heterogeneously refined 3D maps, 250,126 particles were selected for subsequent refinement and post-processing. The final reconstruction, performed with C1 symmetry, resulted in a 3.32 Å resolution 3D map. After further 3D classification and local refinement, improved 3D maps of Fks2 with ordered IFs were obtained at 3.4 Å resolution (Fks2–EFU–Lip).
Structural modeling, refinement, and validation
The predicted structures of Fks1 and Fks2, generated by AlphaFold2, were used as initial models. These models were fitted into the cryo-EM density maps and manually adjusted using COOT and Chimera. The complete Fks1 and Fks2 models were refined through real-space refinement in the PHENIX program, followed by further manual adjustments in COOT. Model validation was performed using MolProbity. Structural figures were prepared using Chimera and PyMOL (https://pymol.org/2/).
UDP-Glo glycosyltransferase assay
The enzymatic activity of wild-type (WT) FKS and its mutants was assessed using the UDP-Glo™ Glycosyltransferase Assay (Promega, Catalog No. V6961), which quantifies the activity of glycosyltransferases utilizing UDP-sugars as substrates. Each reaction was performed in a total volume of 5 μL, containing 0.12 μg of purified proteins, 20 mM HEPES (pH 7.4), 150 mM NaCl, 0.025% n-dodecyl-β-D-maltoside (DDM), 0.0025% cholesteryl hydrogen succinate (CHS), 1 mM MgCl2, 1 mM GTP-γS, 1 mM UDP-glucose, and 1 mM glucose. The reaction mixtures were incubated at 30 °C for 30 min, followed by the addition of 5 μL UDP Detection Reagent. After an additional 60 min incubation at room temperature, luminescence was measured using a Synergy H1 Hybrid Multi-Mode Microplate Reader (BioTek).
Colony growth assay
We prepared the Fks1 knockout strain (ΔFks1) in the W303 strain (EUROSCARF). Fks1 mutants and truncations were constructed using plasmid pRS423–His3 (Addgene) and transformed into the ΔFks1 strain. Wild-type yeast strains and FKS1 knockout strains were initially cultured in SD medium at 30 °C overnight to reach the same optical density (OD). Serial 10-fold dilutions of the cells were then prepared and spotted onto SD-His plates with or without 0.1 μg/mL FK506. The plates were incubated at 30 °C for 2 days, after which colony growth was assessed.
Antifungal susceptibility testing
Antifungal agents were prepared as stock solutions at a concentration of 10 mg/mL in dimethyl sulfoxide (DMSO) and stored at −20 °C, except for caspofungin, which was dissolved in water due to its high solubility. The final concentrations used in the assays were as indicated in the text and figures. For micro-culture assays involving multiple samples, late logarithmic-phase cultures grown in SD-His medium were diluted to a final cell density of 2 × 106 cells/mL in 200 μL of SD-His medium containing serially increasing concentrations of the antifungal agents or an equivalent volume of solvent as a control. Cultures were incubated at 30 °C with continuous agitation on an orbital roller, and the turbidity was measured following 24 and 48 h of incubation. The minimum inhibitory concentration (MIC50) was defined as the lowest antifungal concentration required to inhibit 50% of microbial growth after 24 h of incubation. All reported values represent the mean of at least three independent experiments.
Molecular dynamics simulations
All-atom molecular dynamics (MD) simulations, were performed using GROMACS 2025.1 with the CHARMM36m force field for the protein and CGenFF parameters for the ligand. The simulation systems were constructed using CHARMM-GUI, embedding the protein-ligand complex in a POPC bilayer solvated with TIP3P water and 0.15 M NaCl; detailed system parameters are provided in Supplementary Table 3. Following energy minimization, the systems were equilibrated using the standard CHARMM-GUI six-step protocol with gradually released position restraints. Four independent 100 ns production runs were performed for each system with a 2 fs time step. The temperature and pressure were maintained at 303.15 K and 1.0 bar using the V-rescale thermostat and C-rescale barostat, respectively, while long-range electrostatic interactions were treated using the PME method and hydrogen bonds were constrained using the LINCS algorithm.
Structure prediction and analysis
Structural comparisons of the apo-Fks1 (Supplementary Fig. 1) utilized the predicted model of S. cerevisiae Fks1 retrieved from the AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk; Entry ID: AF-P38631-F1, model version 4). For the protein-ligand complex predictions (Supplementary Fig. 14), the structures of S. cerevisiae Fks1 (UniProt ID: P38631) and C. albicans Fks1 (UniProt ID: A0A1D8PCT0) in complex with enfumafungin (EFU) were generated using a locally deployed version of AlphaFold 3. The predictions were performed in April 2025. The SMILES string of enfumafungin was provided as the ligand input. The confidence of the predicted models was evaluated using the predicted local distance difference test (pLDDT) for local residue quality and the predicted aligned error (PAE) for domain packing and ligand placement reliability. Visualizations of the models colored by pLDDT scores and the corresponding PAE plots are provided in Supplementary Fig. 14.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM 3D maps and the corresponding atomic models of the Fks1 and Fks2 have been deposited at the EMDB database and the RCSB PDB with the respective accession codes of EMD-66359 and 9WY1 (Fks1 in apo state), EMD-66409 and 9WZU (Fks1 in complex with enfumafungin), EMD-66410 and 9WZV (Fks1 with intact active site), EMD-66411 and 9WZX (Fks1 in open state), EMD-66408 and 9WZT (Fks2 in apo state), EMD-66407 and 9WZS (Fks2 in complex with enfumafungin), EMD-66419 and 9X04 (Fks2 with intact active site). PDB codes of previously published structures used in this study are 8WLA and 7YUY. MD simulation input, parameter, and output files are provided in Supplementary Data 1. Source data are provided as a Source Data file. Source data are provided with this paper.
References
Hawksworth, D. L. The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol. Res. 105, 1422–1432 (2001).
Nature Microbiology. Stop neglecting fungi. Nat. Microbiol. 2, 17120 (2017).
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv113 (2012).
Almeida, F., Rodrigues, M. L. & Coelho, C. The still underestimated problem of fungal diseases worldwide. Front. Microbiol. 10, 214 (2019).
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194 (2012).
Lima, S. L., Colombo, A. L. & de Almeida, J. N. Fungal cell wall: emerging antifungals and drug resistance. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.02573 (2019).
Douglas, C. M. et al. The Saccharomyces cerevisiae Fks1 (Etg1) gene encodes an integral membrane-protein which is a subunit of 1,3-beta-D-glucan synthase. Proc. Natl. Acad. Sci. USA 91, 12907–12911 (1994).
Mazur, P. et al. Differential expression and function of two homologous subunits of yeast 1,3-beta-D-glucan synthase. Mol. Cell Biol. 15, 5671–5681 (1995).
Qadota, H. et al. Identification of yeast Rho1p GTPase as a regulatory subunit of 1,3-beta-glucan synthase. Science 272, 279–281 (1996).
Arellano, M., Duran, A. & Perez, P. Rho1 GTPase activates the (1-3)beta-D-glucan synthase and is involved in Schizosaccharomyces pombe morphogenesis. Embo J. 15, 4584–4591 (1996).
Diaz, M. et al. The Schizosaccharomyces pombe Cwg2+ gene codes for the beta-subunit of a geranylgeranyltransferase type-I required for beta-glucan synthesis. Embo J. 12, 5245–5254 (1993).
Mazur, P. & Baginsky, W. In vitro activity of 1,3-beta-D-glucan synthase requires the GTP-binding protein Rho1. J. Biol. Chem. 271, 14604–14609 (1996).
Drgonova, J. et al. Rho1p, a yeast protein at the interface between cell polarization and morphogenesis. Science 272, 277–279 (1996).
Li, J. et al. Cryo-EM structure of the beta−1,3-glucan synthase FKS1-Rho1 complex. Nat. Commun. 16, 2054 (2025).
Healey, K. R. et al. Differential regulation of echinocandin targets Fks1 and Fks2 in Candida glabrata by the post-transcriptional regulator Ssd1. J. Fungi (Basel) https://doi.org/10.3390/jof6030143 (2020).
Katiyar, S. K. et al. Fks1 and Fks2 are functionally redundant but differentially regulated in Candida glabrata: implications for echinocandin resistance. Antimicrob. Agents Chemother. 56, 6304–6309 (2012).
Garcia-Effron, G., Lee, S., Park, S., Cleary, J. D. & Perlin, D. S. Effect of Candida glabrata FKS1 and FKS2 mutations on echinocandin sensitivity and kinetics of 1,3-beta-D-glucan synthase: implication for the existing susceptibility breakpoint. Antimicrob. Agents Chemother. 53, 3690–3699 (2009).
Gonzalez-Jimenez, I. et al. Expression of 1,3-beta-glucan synthase subunits in Candida glabrata is regulated by the cell cycle and growth conditions and at both transcriptional and post-transcriptional levels. Antimicrob. Agents Chemother. 69, e0050025 (2025).
Suwunnakorn, S., Wakabayashi, H., Kordalewska, M., Perlin, D. S. & Rustchenko, E. FKS2 and FKS3 genes of opportunistic human pathogen Candida albicans influence echinocandin susceptibility. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.02299-17 (2018).
Douglas, C. M. et al. Identification of the FKS1 gene of Candida albicans as the essential target of 1,3-beta-D-glucan synthase inhibitors. Antimicrob. Agents Chemother. 41, 2471–2479 (1997).
Jallow, S. & Govender, N. P. Ibrexafungerp: a first-in-class oral triterpenoid glucan synthase inhibitor. J. Fungi https://doi.org/10.3390/jof7030163 (2021).
Kumar, V., Huang, J., Dong, Y. W. & Hao, G. F. Targeting Fks1 proteins for novel antifungal drug discovery. Trends Pharm. Sci. 45, 366–384 (2024).
Zhao, C. R. et al. Structure of a fungal 1,3-beta-glucan synthase. Sci. Adv. 9, eadh7820 (2023).
Hu, X. L. et al. Structural and mechanistic insights into fungal ß−1,3-glucan synthase FKS1. Nature 616, 190 (2023).
Morgan, J. L., Strumillo, J. & Zimmer, J. Crystallographic snapshot of cellulose synthesis and membrane translocation. Nature 493, 181–186 (2013).
Datry, A. & Thellier, M. Echinocandins: a new class of antifungal agents, a new mechanism of action. J. Mycol. Med. 12, S5–S9 (2002).
Pound, M. W., Townsend, M. L. & Drew, R. H. Echinocandin pharmacodynamics: review and clinical implications. J. Antimicrob. Chemother. 65, 1108–1118 (2010).
Wiederhold, N. P. Pharmacology, mechanisms of action, and unique characteristics of antifungal agents. Mycoses 54, 36–36 (2011).
McCarthy, M. W. Pharmacokinetics and pharmacodynamics of ibrexafungerp. Drugs RD 22, 9–13 (2022).
Pelaez, F. et al. The discovery of enfumafungin, a novel antifungal compound produced by an endophytic Hormonema species biological activity and taxonomy of the producing organisms. Syst. Appl. Microbiol. 23, 333–343 (2000).
Onishi, J. et al. Discovery of novel antifungal (1,3)-beta-D-glucan synthase inhibitors. Antimicrob. Agents Chemother. 44, 368–377 (2000).
Jimenez-Ortigosa, C., Perez, W. B., Angulo, D., Borroto-Esoda, K. & Perlin, D. S. De novo acquisition of resistance to SCY-078 in Candida glabrata involves FKS mutations that both overlap and are distinct from those conferring echinocandin resistance. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.00833-17 (2017).
Nunnally, N. S., Etienne, K. A., Angulo, D., Lockhart, S. R. & Berkow, E. L. In vitro activity of ibrexafungerp, a novel glucan synthase inhibitor against Candida glabrata isolates with FKS mutations. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.01692-19 (2019).
Aldejohann, A. M. et al. In vitro activity of ibrexafungerp against clinically relevant echinocandin-resistant Candida strains. Antimicrob. Agents Chemother. 68, e0132423 (2024).
Schwartz, R. E. et al. Isolation and structural determination of enfumafungin, a triterpene glycoside antifungal agent that is a specific inhibitor of glucan synthesis. J. Am. Chem. Soc. 122, 4882–4886 (2000).
Zhong, Y. L. et al. Synthesis of antifungal glucan synthase inhibitors from enfumafungin. J. Org. Chem. 77, 3297–3310 (2012).
Apgar, J. M. et al. Novel orally active inhibitors of beta−1,3-glucan synthesis derived from enfumafungin. Bioorg. Med. Chem. Lett. 25, 5813–5818 (2015).
Jorgensen, K. M., Astvad, K. M. T., Hare, R. K. & Arendrup, M. C. EUCAST ibrexafungerp MICs and wild-type upper limits for contemporary danish yeast isolates. J. Fungi (Basel) https://doi.org/10.3390/jof8101106 (2022).
Mesquida, A. et al. DeltaF659 and F659S substitutions at the HS1 of FKS2 gene, along with E655A and W715L upstream and downstream substitutions, correlate with high ibrexafungerp MICs against Candidaglabrata. Clin. Microbiol. Infect. 28, 1154 e1155–1154 e1158 (2022).
Perlin, D. S. Resistance to echinocandin-class antifungal drugs. Drug Resist. Updat. 10, 121–130 (2007).
Perlin, D. S. Mechanisms of echinocandin antifungal drug resistance. Ann. NY Acad. Sci. 1354, 1–11 (2015).
Morgan, J. L., McNamara, J. T. & Zimmer, J. Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP. Nat. Struct. Mol. Biol. 21, 489–496 (2014).
Morgan, J. L. W. et al. Observing cellulose biosynthesis and membrane translocation in crystallo. Nature 531, 329 (2016).
Chen, D. D. et al. Structure, catalysis, chitin transport, and selective inhibition of chitin synthase. Nat. Commun. 14, 4776 (2023).
Maloney, F. P. et al. Structure, substrate recognition and initiation of hyaluronan synthase. Nature 604, 195–201 (2022).
Denning, D. W. Echinocandin antifungal drugs. Lancet 362, 1142–1151 (2003).
Saitô, H. Dynamic pictures of membrane proteins in two-dimensional crystal, lipid bilayer and detergent as revealed by site-directed solid-state C NMR. Chem. Phys. Lipids 132, 101–112 (2004).
Zocher, M., Zhang, C., Rasmussen, S. G. F., Kobilka, B. K. & Müller, D. J. Cholesterol increases kinetic, energetic, and mechanical stability of the human β-adrenergic receptor. Proc. Natl. Acad. Sci. USA 109, E3463–E3472 (2012).
Fioriti, S. et al. Antifungal combinations against Candida species: from bench to bedside. J. Fungi (Basel) https://doi.org/10.3390/jof8101077 (2022).
Chandrasekar, P. H. & Sobel, J. D. Micafungin: a new echinocandin. Clin. Infect. Dis. 42, 1171–1178 (2006).
Roling, E. E. et al. Antifungal activities of fluconazole, caspofungin (MK0991), and anidulafungin (LY 303366) alone and in combination against Candida spp. and Crytococcus neoformans via time-kill methods. Diagn. Microbiol Infect. Dis. 43, 13–17 (2002).
Petraitis, V. et al. Combination therapy with ibrexafungerp (formerly SCY-078), a first-in-class triterpenoid inhibitor of (1–>3)-beta-d-glucan synthesis, and isavuconazole for treatment of experimental invasive pulmonary aspergillosis. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.02429-19 (2020).
Ghannoum, M. et al. Evaluation of the antifungal activity of the novel oral glucan synthase inhibitor SCY-078, singly and in combination, for the treatment of invasive aspergillosis. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.00244-18 (2018).
Acknowledgments
Cryo-EM data were collected in the Peking University Health Science Center Cryo-Electron Microscopy Facility. We thank Dandan Chen and Lihong Chen for facilitating data collection. We thank Yong Wang for facilitating the MD analysis. This work was supported by grants from the National Natural Science Foundation of China (32571451 and 92578126 to L.B.), Beijing Natural Science Foundation (7252080 to L.B.), and Peking University (to L.B.).
Author information
Authors and Affiliations
Contributions
Z.Y. and L.B. conceived and designed the experiments. Z.Y., L.S., L.W., Y.N., R.L., D.C., T.S., and L.B. performed the experiments. Z.Y., L.S., L.W., C.Y., Y.S., and L.B. analyzed the data. Z.Y. and L.B. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks David Perlin and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
You, ZL., Sun, L., Wang, LX. et al. Inhibition mechanism of the fungal β−1,3-glucan synthases by triterpenoid antifungal drugs. Nat Commun 17, 2347 (2026). https://doi.org/10.1038/s41467-026-69114-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69114-8