Abstract
Cross-allergies affect a significant proportion of the population, and contribute to detrimental health and socioeconomic impacts, yet allergen immunotherapies often target a single allergen source disregarding cross-reactive allergens from other sources. Here we introduce an immunization approach developed for improved desensitization in cross-allergic patients using a consensus allergen (cnsLTP1), which contains orthologous non-specific lipid transfer proteins (nsLTP) derived from relevant fruit and pollen allergens. In BALB/c mice, vaccination via either mRNA-lipid nanoparticle (LNP) vehicle or traditional protein formulation induces cnsLTP1-specific IgGs capable of recognizing and binding to multiple nsLTPs. These IgGs block allergen binding by patient serum IgEs and prevent humanized rat basophil degranulation in vitro. Meanwhile, in an allergic mouse model, the mRNA-LNP formulation is tolerated and induces allergen-specific IgG responses but does not ameliorate subsequent allergen challenge responses. Regardless, this cross-allergen mRNA-LNP-based immunotherapy may have translation value once route of administration, formulation and/or dosing are optimized.
Similar content being viewed by others
Introduction
Allergy is a chronic condition affecting 10–30% of the global population, with increasing incidence1. Cross-reactivity occurs when structurally similar proteins from different sources share epitopes, leading to immune recognition and reactions induced by multiple allergens2,3,4. The management of food allergy is complicated by an abundance of homologous, cross-reactive proteins in edible foods and aeroallergens5,6. This results in patients having allergic sensitization to many biologically related foods or cross-reactivity between pollen and plant foods4.
The short-term treatment of food allergy involves either avoiding the allergen source or using nonspecific acute treatments, such as antihistamines for milder cases and adrenaline for severe systemic cases (anaphylaxis)7. Besides avoidance measures, the only approach for mitigating the effects of food allergy is food allergy immunotherapy (AIT) (through oral (OIT), sublingual (SLIT), or epicutaneous (EPIT) routes) either alone or in combination with an anti-IgE monoclonal antibody, such as omalizumab8. However, these therapies are often associated with frequent and potentially severe adverse reactions. Pollen AIT has proven effective for the treatment of respiratory symptoms, but prospective studies are needed to establish its role in the prevention and moderation of symptoms upon food exposure in cases of pollen-food allergy syndrome (PFAS)2. Over time, AIT modulates the immune response, shifting the production of allergen-specific antibodies from IgE to IgG4 subtypes, while inducing a tolerant TH1- and regulatory T (Treg)-cell phenotype9,10. These allergen-specific IgG4 antibodies protect against allergic reactions by blocking the allergen, thereby reducing allergen sensitivity11,12,13,14.
Even though AIT may effectively reduce the frequency and risk of severe food allergic reactions, its main limitations include the risk of severe adverse reactions and the duration of treatment. In the case of OIT, long-term tolerance may require continued ingestion of a maintenance dose. For pollen AIT, the treatment lasts from three to five years with frequent clinical follow-ups and variable efficacy among patients11. Moreover, traditional AIT relies on protein extracts where the specific allergens crucial for desensitization are combined with other proteins at undetermined concentrations. Each mixture therefore not only contains the target allergen, but also other unrelated proteins that might restrict the immune response against the triggering allergen15. Particularly for patients with multiple food allergies due to genuine and cross-reactive allergens, desensitization to a specific allergen isoform in the extract might not induce protection against other related allergens found in other food and pollen sources16,17,18. This lack of paraspecificity, coupled with lengthy treatment durations contributes to high discontinuation rates observed in AIT for respiratory allergens like grass pollen19,20,21. Specifically, studies report discontinuation rates of up to 77% for SCIT and 36% for SLIT22.
In this study, we explore a new therapeutic approach for mitigating the effect of extensive cross-reactive food allergies by combining mRNA-based immunization with a designed consensus allergen - a single engineered protein that closely resembles multiple related natural allergens. Successful delivery of natural allergens encoded by mRNA has already been achieved in the context of AIT23,24,25,26. Therefore, we hypothesized that an approach relying on a consensus allergen would enable simultaneous treatment of multiple allergies. Further, delivery via mRNA encapsulated in a lipid nanoparticle (mRNA-LNP) can ensure predominantly intracellular expression, thereby reducing extracellular allergen presence and availability for binding by preexisting IgEs. With this approach, we aim to provide a treatment that alleviates many of the adverse effects associated with conventional AIT.
To explore the utilization of the abovementioned strategy for cross-protection against different allergies, we focused on widely cross-reactive allergens: non-specific lipid transfer proteins (nsLTPs). nsLTPs are allergens belonging to a highly conserved protein family found in multiple plant-derived foods, such as peaches, apples, and nuts, as well as in pollen from olive tree, plane tree, pellitory, and mugwort. LTP syndrome is more prevalent in Mediterranean countries and is often linked to peach as the primary sensitizing food. Due to the broad cross-reactivity of nsLTPs, patients develop symptoms upon consumption of fruits, nuts, and vegetables, and symptoms can range from mild oral reactions to severe anaphylaxis18,25. Commonly, patients with nsLTP allergies are polysensitized to several nsLTPs, potentially hampering the clinical efficacy of traditional AIT protocols.
To evaluate our immunization approach, we immunise naive mice with either mRNA or protein-based formulations of the consensus allergen and characterize the cross-binding properties of serum antibodies to a panel of recombinant and natural nsLTPs, as well as their capacity to prevent IgE binding to allergens and basophil degranulation, demonstrating the IgG blocking capacity. In addition, we test our formulation in an allergic mouse model. The mRNA-LNP formulation is well tolerated upon injection, while inducing allergen-specific IgG responses. Taken together, our results provide proof of concept for a new treatment strategy based on mRNA-LNP technology and consensus allergens, as well as showcase the utility of this approach on nsLTPs involved in LTP syndrome.
Results
Consensus allergen retains nsLTP folding and binds IgE
For the design of the consensus allergen (cnsLTP1), nsLTP amino acid sequences from different plant botanical families, constituting food and pollen allergens, were included to broadly represent the ubiquitous nsLTP family. While the designed consensus allergen did not mirror any specific natural sequence, it exhibited the highest sequence identity to food nsLTPs, with Mal d 3 (apple) being the closest related, showing over 81% sequence identity, followed by Pru p 3 (peach) with 75% sequence identity. In contrast, the least identity was observed with pellitory pollen nsLTPs, to the consensus allergen sharing less than 30% identity with Par j 2 (Supplementary Fig. 1). This broad range of sequence diversity, from high identity with Mal d 3 to low identity with Par j 2, highlights the ability of the cnsLTP1 to encompass epitopic features across diverse nsLTP variants, supporting its utility in eliciting cross-reactive immune responses.
For characterization and further comparison of the cnsLTP1 to other nsLTPs, we designed a panel of recombinant allergens. This panel included a variety of nsLTP allergens from both food (like fruit, nuts, and legumes) and pollen sources. These allergens were chosen for their significant differences in amino acid sequences, as shown in the sequence identity matrix (Fig. 1c). The selected allergens have varying degrees of identity to the cnsLTP1, ranging from 29.35% (Par j 2 and Ole e 7) to 74.73% (Pru p 3), as detailed in Fig. 1c. Despite considerable sequence variability across the nsLTP family, structural comparison revealed high identity between pollen and food nsLTPs, as well as with the consensus allergen (Fig. 1a), suggesting that all nsLTP variants and the consensus allergen share a common structural scaffold. The far-UV circular dichroism (CD) spectrum of the cnsLTP1 (Fig. 1b) indicated a predominant α-helical structure, fitting well with its predicted three-dimensional conformation. When heated to 85 °C, the protein did not fully denature but retained a spectrum indicative of an α-helix-rich structure. The original three-dimensional structure was restored after cooling the sample to 25 °C.
a Superimposed structures of nsLTP allergens and the consensus allergen, visualized with ChimeraX. Structures were retrieved from RCSB.org (Pru p 3: 2B5S; Cor a 8: 4XUW) or homology modeled (Par j 2, Ara h 9, Ole e 7, Tri a 14, Pla a 3, and cnsLTP1). b Circular dichroism spectroscopy of the consensus allergen before and after heating to 85 °C. c Sequence identity matrix (%) including all the recombinant allergens used in the study (Clustal Omega).
The panel of recombinant allergens, appearing with a molecular weight between 10 and 15 kDa on SDS-PAGE, were all recognized by a commercial Mal d 3 polyclonal antibody in their reduced but not in their native form, indicating a change in structure upon reduction and the presence of hidden epitopes (Supplementary Fig. 2). We hypothesize that the anti-Mal d 3 antibody has been raised against an unfolded Mal d 3 and, therefore, is restricted to linear epitopes that are hidden when the protein is not completely reduced.
To ascertain the clinical relevance of the designed cnsLTP1, we assessed its recognition by IgEs present in the serum of patients diagnosed with nsLTP cross-allergies. Our analysis included 10 patients presenting severe oral allergy syndrome, who were recruited after systemic reactions to nsLTP-containing foods and were sensitized to various nsLTPs but exhibited distinct sensitization patterns (Supplementary Table 2). The patients did not show any other relevant co-sensitization beyond nsLTPs. Western blotting demonstrated that IgE antibodies from all tested patients recognized the cnsLTP1 (Supplementary Fig. 3). These findings substantiate that the cnsLTP1 contains relevant IgE epitopes, underscoring its clinical relevance in the context of nsLTP cross-allergies.
cnsLTP1 IgG induction by mRNA-LNP and protein immunization
To evaluate the immunogenicity of the cnsLTP1, we introduced cnsLTP1, Pru p 3, and Par j 2 by intramuscular administration to naive BALB/c mice through an mRNA-LNP. The cnsLTP1 was also tested by a conventional protein immunization with incomplete Freund’s adjuvant. Additionally, we evaluated the effect of the route of administration by testing the cnsLTP1 mRNA formulation and the protein formulation with poly(I:C) or Freund’s adjuvant via subcutaneous injection. All formulations were administered three times at three-week intervals (Fig. 2a). No signs of discomfort or fever were observed in the animals throughout the study.
a Injection and bleeding schedule for immunized mice. b Total antibody titers were measured by ELISA on day 63 for selected isotypes in mice immunised with PBS (black dots), cnsLTP1-protein (pink squares), cnsLTP1-mRNA (teal triangles), Pru p 3-mRNA (orange upward triangles) or Par j 2-mRNA (green rhombus) c cnsLTP1 allergen-specific IgG1 antibody responses measured by ELISA following injection with no doses (day 0), 1 dose (day 21), 2 doses (day 42), or 3 doses (day 63) of PBS (black dots), cnsLTP1-protein (pink squares), cnsLTP1-mRNA (teal triangles), Pru p 3-mRNA (orange upward triangles) or Par j 2-mRNA (green rhombus) d cnsLTP1 allergen-specific IgG2a antibody responses measured by ELISA following injection with no doses (day 0), 1 dose (day 21), 2 doses (day 42), or 3 doses (day 63) PBS (black dots), cnsLTP1-protein (pink squares), cnsLTP1-mRNA (teal triangles), Pru p 3-mRNA (orange upward triangles) or Par j 2-mRNA (green rhombus). Bar heights are means ± SD. Statistical significance was determined using a two-way ANOVA with Dunnett’s multiple comparisons test comparing each sample in a group to the respective PBS control: * P ≤ 0.05, ** P ≤ 0.01, *** P ≤ 0.001, **** P ≤ 0.0001. Figures were created in BioRender. Rivera de Torre, E. (2026) https://BioRender.com/8olw5jx, and graphs were plotted using GraphPad Prism version 10.
Both methods of immunization increased antibody titers across all isotypes, except IgA, compared to PBS-treated controls (Fig. 2b). Protein immunization resulted in significantly increased IgG1, whereas mRNA immunization significantly increased IgG2a and IgM compared to the PBS-treated controls, indicating a more IgG2a- and IgM-driven response in the mRNA-immunized mice (Fig. 2b). The mice immunized with either the cnsLTP1 or the Pru p 3 mRNA-LNP formulation produced low levels of cnsLTP1-binding antibodies after a single dose (day 21) (Fig. 2c). In contrast, the protein formulation necessitated a booster dose to elicit a comparable response. Both mRNA and protein formulation demonstrated increased allergen-specific antibody titers after the first (day 42) and second (day 63) booster doses, with increased IgG1 for both protein and mRNA immunization, whereas only mRNA immunization significantly increased cnsLTP1-binding IgG2a titers (Figs. 2d and 3). Equivalent results were found using subcutaneous administration (Supplementary Fig. 4a–c).
a Serum IgG1 responses measured on day 63 (after three immunisations) in mice immunised with PBS (black dots), cnsLTP1-mRNA (teal upward triangles), or cnsLTP1-protein (pink squares). Each data point represents an individual mouse (technical triplicates), with five mice per group. Data are presented as mean ± SD. b IgG2 titres. Serum IgG2 responses measured on day 63 (after three immunisations) in mice immunised with PBS (black dots), cnsLTP1 mRNA (teal upward triangles), or cnsLTP1 protein (pink squares). Each data point represents an individual mouse (technical triplicates), with five mice per group. Data are presented as mean ± SD. Statistical significance was determined using two-way ANOVA with Dunnett’s multiple comparisons test, comparing each immunised group to the respective PBS control (***P ≤ 0.0001). Graphs were generated using GraphPad Prism version 10.
Thus, a distinct trend emerged, where cnsLTP1-binding IgG2a titers in mRNA-immunized mice were higher than IgG1 titers, while the opposite was observed in protein-immunized mice, where the titers of IgG1 were higher than those of IgG2a (Figs. 2c and 3). In the context of allergen neutralization, the human isotype IgG4 is responsible for neutralizing allergens and preventing the binding to IgE27. Such a role is performed by IgG1 and IgG2a isotypes in mice28.
To evaluate the induction of TH1/TH2 cells in response to mRNA or protein immunization, we restimulated T cells from the spleen and draining inguinal lymph nodes (iLNs) (Supplementary Figs. 5, 6). Following intramuscular immunization, spleens of mice subjected to mRNA immunization had slightly lower percentages of IFNgamma (IFNg)-producing TH1 cells and higher IL4-producing TH2 cells, resulting in a lower TH1/TH2 ratio, whereas the cnsLTP1 protein formulation was similar to PBS (Supplementary Fig. 5a, b). The draining iLNs, however, had similar or higher percentages of TH1 cells following mRNA immunization compared to protein immunization or PBS, with no significant differences observed for TH2 cells and TH1/TH2 ratios (Supplementary Fig. 5c, d). Following subcutaneous immunization, no significant differences were observed across neither the spleen nor iLNs (Supplementary Fig. 5e–h).
We also assessed the induction of IgE to compare the potential of the two immunization formulations for triggering an allergic reaction. No allergen-specific IgE was detected in mice immunized intramuscularly with either the mRNA-LNP or protein formulation. However, in the subcutaneous administration of the protein formulation, the inclusion of an adjuvant, specifically poly(I:C), significantly increased the production of allergen-specific IgE compared to PBS-treated mice (Supplementary Fig. 4d, e).
Cross-reactivity of IgGs with recombinant nsLTP
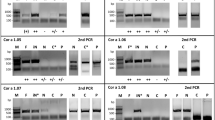
To assess whether antibodies produced in response to immunization with the cnsLTP1 recognized natural allergens, we conducted an IgG1 and IgG2a ELISA with a panel of recombinantly expressed allergens. Antibodies elicited against the cnsLTP1 demonstrated binding capabilities across all tested allergens except Ole e 7, with significantly stronger recognition than antibodies derived from Pru p 3 or Par j 2 immunizations. Specifically, mice immunized with the cnsLTP1 mRNA formulation showed broader cross-reactivity, recognizing all allergens tested, including Pru p 3, Cor a 8, Ara h 9, Par j 2, Pla a 3, and Tri a 14. In contrast, mice immunized with Pru p 3 mRNA recognized Pru p 3, Cor a 8, Ara h 9, and Pla a 3, but not Tri a 14 or Par j 2. Similarly, mice immunized with Par j 2 mRNA primarily recognized Par j 2 and Pla a 3, but showed no significant recognition of the other allergens (Fig. 4).
a IgG1 titres. Serum IgG1 responses measured on day 63 (after three immunisations) in mice immunised with PBS (black dots), Pru p 3-mRNA (orange downward triangles), Par j 2-mRNA (green rhombus), or cnsLTP1-mRNA (teal upward triangles) against specific allergens. Each data point represents an individual mouse (technical triplicates), with five mice per group. Data are presented as mean ± SD. b IgG2 titres. Serum IgG2 responses measured on day 63 (after three immunisations) in mice immunised with PBS (black dots), Pru p 3-mRNA (orange downward triangles), Par j 2-mRNA (green rhombus), or cnsLTP1-mRNA (teal upward triangles). Each data point represents an individual mouse (technical triplicates), with five mice per group. Data are presented as mean ± SD. Statistical significance was determined using two-way ANOVA with Dunnett’s multiple comparisons test, comparing each immunised group to the respective PBS control (*P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001). Graphs were generated using GraphPad Prism version 10.
A comparison of cnsLTP1 mRNA and protein formulations revealed that the former induced significant IgG1 recognition of Pru p 3, Cor a 8, and Par j 2 (Fig. 3a). Regarding IgG2a, the antibodies induced using the mRNA formulation demonstrated significant binding to all the allergens compared to the antibodies derived using the protein formulation (Fig. 3b). This enhanced recognition was accompanied by higher titers of allergen-specific IgG2a. Notably, the intensity of recognition observed in the ELISA correlated with the percentage sequence identity shared with the cnsLTP1, with the strongest signal observed for Pru p 3 (74.73% sequence identity) and the weakest for Par j 2 and Ole e 7 (29.35% sequence identity).
Murine antibodies block allergen-IgE interaction
To investigate the ability of vaccine-induced antibodies to block relevant human IgE epitopes, we evaluated whether murine antibodies induced by immunization with the cnsLTP1 could block the interaction between allergens and human IgEs from patient sera using two complementary assays: an IgE ELISA (Fig. 5) and a degranulation assay with hRBL-2H3 cells (Fig. 6). Initial tests revealed that the patient sera exhibited varied sensitization patterns across the panel of recombinant allergens, including the cnsLTP1, Pru p 3, Cor a 8, Ara h 9, Par j 2, Tri a 14, and Pla a 3. These patterns were consistent between the IgE ELISA and the degranulation assay, confirming the utility of both approaches for evaluating allergen-IgE interactions (Figs. 5a and 6a).
a Recognition of recombinant nsLTP allergens by human serum IgE from selected patients. The experiment was performed twice with technical triplicates. c Inhibition of human serum IgE binding to nsLTP allergens by mouse serum antibodies measured by ELISA after immunization with cnsLTP1 in mRNA (teal) and protein (pink) formulations. Mouse serum was added in increasing dilutions (1:10 to 1:10,000), while human serum was used at a 1:10 dilution, as described in the Methods. Dotted lines indicate non-linear regression fits used to calculate IC₅₀ values. Data were normalized to the maximum signal of technical triplicates. c IC₅₀ values calculated from the inhibition curves shown in (b) for immunizations with cnsLTP1 in mRNA (teal) and protein (pink) formulations. Data are presented as mean ± SD. Individual IC₅₀ values are reported in Supplementary Table 3. Statistical significance was determined using two-way ANOVA with Dunnett’s multiple comparisons test, comparing each sample to the respective PBS control (*P ≤ 0.05). Graphs were plotted using GraphPad Prism version 10.
a Percentage of β-hexosaminidase release from hRBL-2H3 cells sensitized with human serum IgE from different patients and stimulated with nsLTP allergens. Human serum samples were used at a 1:5 dilution b Inhibition of β-hexosaminidase release from hRBL-2H3 cells by preincubation of nsLTP allergens with mouse serum antibodies after immunization with cnsLTP1 in mRNA (teal) and protein (pink) formulations. Mouse serum was added in increasing dilutions (1:10 to 1:10,000), and human serum was used at a 1:5 dilution. Data were normalized to the maximum signal after subtraction of the negative control. c IC₅₀ values calculated as a non-linear regression from the inhibition curves shown in (b) for immunizations with cnsLTP1 in mRNA (teal) and protein (pink) formulations. Data are presented as mean ± SD of three individually calculated values IC₅₀ values. Individual IC₅₀ values are reported in Supplementary Table 3. Statistical significance was determined using two-way ANOVA with Dunnett’s multiple comparisons test, comparing each sample to the respective PBS control (*P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001). Graphs were plotted using GraphPad Prism version 10.
Blocking experiments demonstrated that antibodies from both mRNA-LNP and protein-immunized mice could effectively inhibit IgE binding to all recombinant allergens in a dose-dependent manner (Fig. 5c). Furthermore, this blocking translated into functional inhibition, as reduced degranulation of hRBL-2H3 cells was observed when the allergens were preincubated with mouse serum prior to being introduced to the cells pre-sensitized with patient serum (Fig. 6c). Even though serum from mRNA-LNP-immunized mice showed stronger blocking potency compared to serum from protein-immunized mice, as evidenced by lower IC50 values, the differences were not statistically significant (Figs. 5b, 6b; Supplementary Table 3). These findings indicate that the cnsLTP1 in the mRNA-LNP and protein formulations induced a robust and effective antibody response capable of disrupting allergen-IgE interactions and strongly reducing cellular degranulation in response to a wide range of nsLTPs originating from pollen and food sources.
mRNA-LNP cnsLTP1 induces allergen-specific antibodies in an allergic mouse model
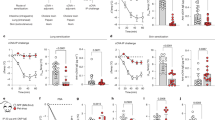
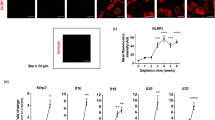
To assess the therapeutic potential of the cnsLTP1 mRNA-LNP formulation in a sensitized setting, we established an allergic anaphylactic mouse model using intranasal Pru p 3 and LPS (Fig. 7a). After sensitization, mice were monitored after each injection of the mRNA-LNP formulation, and no clinical signs of allergy or anaphylaxis were observed. Following intraperitoneal challenge with Pru p 3, body temperature in the treated animals did not differ significantly from that of sensitized but untreated mice. Thus, with the employed experimental setup and small cohorts of mice, the utility of the immunization approach for reducing the level of anaphylaxis could not be established. Measurement of mMCP-1, a marker of mast cell degranulation, did not reveal significant differences between groups (Fig. 7c). Analysis of Pru p 3–specific antibodies in the serum of the mice revealed that levels of IgE were comparable between treated and untreated sensitized groups (Fig. 7g). Allergen-specific IgG1 levels were comparable for non-treated and treated mice for Pru p 3, Cor a 8, and Ara h 9 (Fig. 7d). However, the levels of Par j 2, Tri a 14, and Pla a 3-specific IgGs were significantly higher in treated mice, demonstrating the induction of cross-reactive antibody populations caused by the cnsLTP1-mRNA treatment. Following the same tendency, mice immunized with cnsLTP1 mRNA-LNP exhibited a significant increase in IgG2a and IgG2b levels for all the nsLTPs tested (Fig. 7e-f).
a Schematic overview of the sensitization, treatment, and allergen challenge protocol, including timing of intramuscular immunization, blood collection, intraperitoneal challenge, and body temperature monitoring. b Change in core body temperature following intraperitoneal challenge with Pru p 3.in sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. Statistical significance was determined using the two-sided Mann–Whitney test. c Serum mouse mast cell protease-1 (mMCP-1) concentrations measured by ELISA in non-sensitized healthy control (grey), sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. d Allergen-specific IgG1 antibody levels measured by ELISA in non-sensitized healthy control (grey), sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. e Allergen-specific IgG2a antibody levels measured by ELISA in non-sensitized healthy control (grey), sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. f Allergen-specific IgG2b antibody levels measured by ELISA non-sensitized healthy control (grey), sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. g Pru p 3–specific IgE levels determined by ELISA in non-sensitized healthy control (grey), sensitized anaphylactic control (black), and sensitized mice treated with cnsLTP1 mRNA-LNP (teal) mice. All data are presented as mean ± standard SD, with eight mice per group. Statistical significance was determined using one-way ANOVA with two-sided Dunnett’s multiple comparisons test (unless otherwise stated), comparing each group to the anaphylactic control: *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. Exact P values are indicated where P > 0.05. Schematics were created in BioRender. Rivera de torre, E. (2026) https://BioRender.com/q5fxgfj, and graphs were plotted using GraphPad Prism version 10.
Although the therapeutic utilization of induced antibodies in the allergic mouse model could not be established in the current setup (Fig. 7b), their induction demonstrates the capacity of the consensus allergen to elicit an antigen-specific antibody response. We therefore speculate that changing the route of administration and further optimizing formulation and/or dosing could lead to protective efficacy.
Discussion
For patients suffering from multiple allergies, diagnosis and treatment are often complex, especially in the context of allergy cross-reactivity. Patients undergoing existing methods of AIT are generally desensitized only against the major sensitized allergen, which does not ensure protection against cross-reactive allergens, leading to long and ineffective treatment regimens2,29,30,31. For food allergies, patients are advised to avoid allergen sources, leading to dietary restrictions and reduced quality of life, especially for those suffering from cross-reactive allergies32,33,34. To address these challenges, we present a new strategy for inducing broadly neutralizing IgG antibodies via immunization based on the design of a consensus allergen delivered using mRNA-LNP or protein format, offering potential simultaneous desensitization against multiple allergies.
Central to our approach is the structural conservation of nsLTPs, rendering them ideal candidates for consensus allergen representation. Our design combined sequences from diverse food- and pollen-derived nsLTPs, resulting in the cnsLTP1. Remarkably, the cnsLTP1 demonstrated renaturation capabilities post-thermal denaturation, underscoring the indispensable role of its intramolecular disulfide bonds in structural integrity, as observed before in natural nsLTPs35, validating the structural integrity of the designed allergen.
Further validating the clinical relevance of the cnsLTP1, its recognition by patient-derived IgEs supports that the designed protein shares epitopes with native allergens. This recognition indirectly points toward the potential of the cnsLTP1 for eliciting IgG-blocking antibodies capable of inhibiting IgE epitope binding, which is one of the key features to consider for its use as a potential immunization approach36,37.
Using the cnsLTP1 for immunization, we observed distinct outcomes based on the formulation (mRNA-LNP or protein), adjuvant, and route of administration, as well as the use of natural versus consensus allergens. The mRNA-LNP formulation consistently induced higher IgG2a responses than protein formulations, eliciting comparable levels of IgG1 and IgG2a (Fig. 2, Fig. 4; Supplementary Fig. 4). Substitution of poly(I:C) with Freund’s adjuvant in protein formulations demonstrated that Freund’s adjuvant avoids the induction of allergen-specific IgE while maintaining IgG1 and IgG2a responses (Supplementary Fig. 4)38. This highlights the significant impact of adjuvant selection on the immunological outcomes of protein immunizations.
Comparisons between natural and consensus allergens revealed that the cnsLTP1 induces broader cross-reactive antibodies, with the ability to recognize conformational epitopes of a diverse panel of allergens, including both food- and pollen-derived nsLTPs, with the exception of Ole e 7. Interestingly, although Ole e 7 shares structural similarity with other nsLTPs in terms of secondary and tertiary structure, it is part of a different nsLTP family and exhibits significant divergence in its primary amino acid sequence24,25. This likely accounts for the lack of recognition by antibodies raised against the cnsLTP1. In contrast, immunizations with natural allergens, such as Pru p 3 or Par j 2, produced restricted cross-reactivity profiles, with Pru p 3 eliciting antibodies that predominantly recognized food nsLTPs, while Par j 2 immunization induced antibodies binding only to pollen Par j 2 and Pla a 3. These findings highlight the effectiveness of consensus designs in addressing cross-reactive allergies comprehensively, particularly when epitopic diversity across allergen families is a critical consideration. Furthermore, while the observed IgG1/IgG2a ratios post-mRNA-LNP immunization suggested a potential TH1 bias in previous studies, our results did not yield conclusive IgG1/IgG2a ratios indicative of a clear TH1- or TH2-skewed response39,40,41.
To further evaluate the immune profile induced by mRNA-LNP immunization, we analyzed T-cell populations. Flow cytometry analysis of T-cell populations in the spleen and lymph nodes did not reveal significant polarization of a specific T-cell subset. These findings suggest that while the mRNA-LNP formulation induces robust antibody responses, its effects on T cell–mediated immunity remain unclear and warrant further investigation. Clarifying these aspects will be essential to better understand the immune response and safety of the treatment38,42.
Functionally, the cross-reactive antibodies raised upon immunization with the cnsLTP1 effectively hindered human serum IgE binding and demonstrated functional relevance by preventing the degranulation of hRBL-2H3 cells. Therefore, the antibodies raised upon immunization with the cnsLTP1 can bind to conformational IgE epitopes. This underscores their blocking potential toward several clinically relevant epitopes across both pollen and food allergens within the nsLTP family. Such attributes enhance the potential of consensus design, particularly for patients affected by cross-reactive allergies.
In addition to studies in naïve mice, we evaluated the mRNA-LNP consensus allergen in a Pru p 3–sensitized anaphylactic mouse model to gain insights into its performance under allergic conditions. The absence of symptoms in the sensitized mice upon immunization suggests that leakage of the expressed consensus allergen is minimal or insufficient to elicit immediate reactions, indicating that the treatment was well tolerated in allergic patients. However, upon allergen challenge, body temperature was not significantly different between treated and non-treated anaphylactic mice. The therapeutic utility of our immunization approach could thus not be established, and it is likely that a higher titer or Treg support might be necessary to fully block anaphylaxis43,44. Nevertheless, the immunized sensitized mice displayed a significant increase in allergen specific–IgG1, IgG2a, and IgG2b, consistent with a broad cross-reactive humoral response. These results suggest that optimization of the route of administration, formulation, or immunization schedule could be worth pursuing in the aim to achieve clinically meaningful protection in sensitized individuals43,45.
Together, these findings highlight both the promise and current limitations of the consensus mRNA-LNP approach. While effective induction of cross-reactive antibodies was achieved in both naïve and allergic contexts, further optimization and testing in sensitized models (with larger cohorts) is essential before the approach can be translated into clinical settings.
Beyond the broad applications of the technology platform and the immunization approach described here, our findings may also find immediate utility for management of other allergies where cross-reactivity is prevalent, including profilins and storage proteins (2S albumins, 11S globulins, and 7S vicilins). The successful management of cross-reactive syndromes has the potential to improve the quality of life of millions of patients and may offer significant cost savings for their associated healthcare, as well as it could help reduce absenteeism from work43,45. Additionally, the presented approach could find broad utilization beyond these specific allergies, extending to areas such as vaccination against viral and bacterial pathogens with high mutational capacity and antigenic diversity.
Methods
This study complies with all relevant ethical regulations and is approved by the Danish Animal Experiments Inspectorate, Institutional Animal Care and Use Committee of IBIMA and by the Consejería de Agricultura, Ganadería, Pesca y Desarrollo Sostenible of the Junta de Andalucía. A summary table with all the antibodies, catalogue numbers and dilutions can be found in Supplementary Table 4.
Design of consensus allergen
nsLTPs were selected based on phylogenetic clustering of sequences and different allergen families26. For each cluster or allergen family, at least a single nsLTP was chosen. Amino acid sequences were retrieved from UniProt and aligned using the Clustal Omega algorithm, accessed through the European Bioinformatics Institute (EMBL-EBI) web server (https://www.ebi.ac.uk/Tools/msa/clustalo/) with default parameters46,47. Only sequences of ~90 amino acids in length were selected to be included in the consensus design. The consensus sequence was designed based on the selection of the most conserved or physicochemical-relevant amino acid for each position. For each position, the most frequent amino acid was kept, and in case of a tie, the most abundant amino acid within the most present physicochemical property was used: acid (E, D), basic (K, R), polar (S, T, Y, N, Q), and nonpolar (G, A, V, L, I, M, W, F). P and C formed their own group. The final sequence is in Supplementary Table 5.
Expression of nsLTP allergens
Recombinant allergens were generated by cloning and heterologous expression. Codon-optimized consensus allergen cDNA (Eurofins) was subcloned into the pPICZαA expression vector (#V19520, Invitrogen) and introduced into Komagataella phaffii KM71H (#C18200, Invitrogen) by electroporation47. Colonies were selected by their ability to grow in the presence of YPDS (20 g/L peptone, 10 g/L yeast extract, 20% (w/v) dextrose, 182.2 g/L sorbitol, 20 g/L agar) supplemented with Zeocin 1000 µg/mL plates. Individual colonies grew at 30 °C and 220 rpm for 24 h in BMGY (10 g/L yeast extract, 20 g/L peptone, 0.1 M potassium phosphate pH 6.0, 1.34% (w/v) yeast nitrogen base (YNB), 0.04 μg/mL biotin, 1% (v/v) glycerol), and expression was induced over 72 h in BMMY (10 g/L yeast extract, 20 g/L peptone, 0.1 M potassium phosphate pH 6.0, 1.34% (w/v) YNB, 0.04 μg/mL biotin, 0.5% (v/v) methanol) with addition of 1% (v/v) methanol every 24 h. The supernatant was collected and filtered through a 0.22 µm membrane and dialyzed against Bis-Tris 20 mM pH 6.0 in a 3.5 K MWCO nitrocellulose membrane. Dialyzed samples were purified by cation exchange on a UNOSphereS (BioRad) column. Elutions corresponding to the correct molecular weight acoording to SDS-PAGE analysis were pooled, dialyzed against 50 mM ammonium bicarbonate pH 7.0, lyophilized, and stored at −80 °C.
Characterization of consensus allergen
Purified proteins (2 μg/well) were mixed with 3X loading dye, with or without 1 mM DTT, and boiled for 10 min at 95 °C47. Proteins were transferred to a pre-activated PVDF membrane (#LC2002, Novex) using a SureLock Minicell at 30 V for 1 h. Subsequently, the membrane was blocked overnight in a solution of PBS containing 0.1% Tween 20 (v/v) (PBST) and 5% (w/v) skim milk. After blocking, the membrane underwent three washes with PBST, followed by incubation with a primary rabbit anti-Mal d 3 polyclonal antibody (Catalog #abx300086, Abbexa) for 1 h. This step was succeeded by another series of washes. The membrane was then incubated with a secondary HRP-conjugated goat anti-rabbit antibody (#31460, Invitrogen) for 1 h, concluded by a final wash. Detection of bound antibodies was achieved using Clarity ECL Substrate (#1705061, BioRad), with chemiluminescence signals measured thereafter.
The secondary structure of recombinant proteins and thermal stability was analyzed with far-UV CD spectroscopy using a Jasco J-715 spectropolarimeter equipped with a Neslab RTE-111 thermostat using a 0.1 cm pathlength thermostated cuvette. Consensus allergens were dissolved in 10 mM HEPES, 0.1 M NaCl to avoid pH changes during heating. CD spectra were recorded at 25 °C and 85 °C, after 0.5 °C/min heating while registering changes in ellipticity at 208 nm. Reverse cooling was performed at the same rate, and final spectra were recorded after cooling down to 25 °C.
Sera from allergic patients
Patients were selected due to their history of cross-reactivity to multiple nsLTPs confirmed by skin prick test and ImmunoCAP (Thermo Fisher). All studies involving human material were reviewed and approved prior to initiation by the Comitè d’Ètica de la Investigació amb Medicaments (CEIm) de l’Hospital Universitari Vall d’Hebron and by the Comitè d’Ètica de la Investigació amb Medicaments (CEIm) de l’Hospital Clínic de Barcelona, as applicable. All participants were recruited in accordance with the approved protocols and provided written informed consent prior to sample collection.
IgE recognition of consensus allergen in patients
IgE recognition of the cnsLTP1 was tested with human sera from 10 food nsLTP allergic patients. Purified protein was transferred onto a nitrocellulose membrane after SDS-PAGE (200 ng/strip) and an indirect Western blot was performed with patients’ sera to test for IgE recognition. After 1 h of incubation with blocking buffer, strips were incubated with individual patient sera (1/5 diluted in blocking buffer) for 2 h at room temperature (RT) and later washed three times with PBST. Peroxidase-labeled anti-IgE antibody (#9160-05, Southern Biotech) was incubated (1/1000) for 1 h, and after the final three washes, bound IgE was detected with Clarity ECL Substrate (Bio-Rad) using chemiluminescence detector Fujifilm LAS3000.
mRNA design
The cnsLTP1 mRNA sequence was designed for MHC-II presentation through targeted lysosomal delivery mechanisms by fusing the cnsLTP1 coding sequence to a lysosomal transmembrane domain (LAMP1) and a C-terminal lysosomal translocation sequence48. The mRNA encoding for cnsLTP1 was codon-optimized for humans and single-point synonymous modifications were included to avoid secondary structure formation in the mRNA. The mRNA encoding the consensus allergen was purchased from RiboPro, including a 150nt poly-A-tail and a Cap1 sequence.
Consensus allergen mRNA-LNP preparations
The mRNA was encapsulated in LNP based on the Pfizer/BioNTech vaccine Comirnaty (BNT162b2), but with the PEGylated lipid ALC-0159 exchanged with DMG-PEG2000, resulting in a formulation consisting of ALC0315:Cholesterol:DSPC:DMG-PEG2000 (molar ratio 46.3:42.7:9.4:1.6, N/P ratio 6 corresponding to 39 nmol lipid per µg mRNA)49,50. All lipids were acquired from Avanti Polar Lipids. Lipids were dissolved in ethanol and mRNA in 100 mM sodium acetate (pH 4), and particles were formed by mixing in 1:3 ratios using the NanoAssemblr Ignite microfluidic mixer (Precision Nanosystems) at a final concentration of 100 µg/mL mRNA. Particles were buffer exchanged to HBS (25 mM HEPES, 150 mM NaCl, pH 7.4) by discontinuous diafiltration using Amicon Ultracel spin filters (100 kDa cutoff) and upconcentrated to 0.35 µg/µL. Characterization of mRNA-LNPs was done with dynamic light scattering (DLS) and M3-PALS using a Zetasizer Nano ZS (Malvern Instruments) to determine size, polydispersity, and zeta potential and found to have a size (Z-Ave) of 76.4 ± 5.6 nm and a PDI of 0.11 ± 0.04 (mean values for four batches prepared). Encapsulation efficiency and final yield of mRNA were determined with the RiboGreen Assay, by comparing the RiboGreen emission for LNPs in Tris buffer to LNPs disassembled by incubating with 0.1% Triton X-100 at 60 °C for 30 min. The particles were slightly smaller and more monodisperse than those in BNT162b250. The zeta potential of −12.0 ± 5.0 mV is comparable to what has been reported for BNT162b250. The mRNA encapsulation efficiency was high, reaching 91.7 ± 3.6%. Particles were diluted in 5% sucrose to the desired concentrations and volumes and stored at −80 °C until use. The size (Z-Ave), polydispersity, zeta potential, and encapsulation efficiency were determined after thaw to confirm LNP stability (Supplementary Table 1).
Consensus allergen protein preparations
The consensus allergen protein was dissolved in sterile PBS containing 50 µg/mL of polyinosinic–polycytidylic acid sodium salt (poly(I:C) #P1530, Sigma-Aldrich) or mixed 50% (v/v) with incomplete Freund’s adjuvant (#F5506, Sigma-Aldrich).
Immunization schedule for naïve mice
The studies were approved by the Danish Animal Experiments Inspectorate (approval #2020-15-0201-00748). The animals were co-housed in the DTU BioFacility which includes SPF-status barrier breeding unit, a laboratory for sample processing and facilities for surgical procedures. Animals were maintained on a 12 h light/12 h dark cycle, with ambient temperature kept at 20–22 °C and relative humidity maintained at 45–65%. Food and water were provided ad libitum, and animals were group-housed with environmental enrichment. The immunization was performed in two independent groups of mice, subcutaneously or intramuscularly. For the subcutaneous immunization, 4-to-5-week-old BALB/cAnNRj female mice (#0003 Janvier Labs) were immunized by subcutaneous injection three times at three-week intervals with 9 µg mRNA-LNP (n = 6), 24 µg protein-poly(I:C) (n = 3), 24 µg protein-incomplete Freund’s adjuvant (n = 3), or PBS (n = 7). For the intramuscular immunization, four-to-five-week-old BALB/cAnNRj female mice were immunized by intramuscular injection three times at three-week intervals with 7.5 µg mRNA-LNP (n = 5), 20 µg protein-incomplete Freund’s adjuvant (n = 5), or PBS (n = 5). Mice were euthanised by cervical dislocation. The specific amounts of mRNA-LNP or protein were determined in titration experiments comprising three mice each, and the responses were evaluated by measuring the presence of total Igs and allergen-specific IgG1 and IgG2a. The specific doses were determined in previous titration experiments, and the range was established42,51,52,53.
Establishment of allergic anaphylactic mouse model and immunization schedule
Four- to five-week-old BALB/cAnNRj mice were obtained from Janvier Labs (Le Genest-Saint-Isle, France) and housed at the IBIMA Plataforma BIONAND animal facility, which includes SPF-status barrier breeding unit, a laboratory for sample processing and facilities for surgical procedures (Registration No. ES 290670001687). All animal experiments were formally reviewed and authorized prior to initiation by the Institutional Animal Care and Use Committee of IBIMA and by the Consejería de Agricultura, Ganadería, Pesca y Desarrollo Sostenible of the Junta de Andalucía, under approved protocol number 14/05/2025/051. All procedures were conducted in full compliance with Spanish and European legislation regulating animal experimentation (RD1201/2005, Law 32/2007, Directive 2010/63/EU, and RD53/2013) and adhered to the principles of Replacement, Reduction, and Refinement (3Rs). Animals were maintained on a 12 h light/12 h dark cycle, with ambient temperature kept at 20–22 °C and relative humidity maintained at 45–65%. Food and water were provided ad libitum. Mice were divided into three co-housed groups: non-sensitized controls (n = 8), sensitized non-treated (n = 8), and sensitized mRNA-LNP treated (n = 8). In all groups 3 male and 5 female mice were included. Sensitization was performed intranasally under isoflurane anesthesia using 20 µg of natural Pru p 3 (Bial Laboratory, Zamudio, Spain) combined with 20 ng of LPS (InvivoGen, San Diego, CA) in a final volume of 12 µL PBS. Control mice received 12 µL PBS following the same schedule54,55. Starting at week 6, mice in the treatment group received intramuscular injections of 7.5 µg cnsLTP1 mRNA-LNP in 30 µL once weekly for four consecutive weeks, while control and sensitized non-treated groups received 30 µL PBS according to the same schedule. One week after the last treatment dose, sensitized mice were challenged intraperitoneally with 100 µg of Pru p 3 to assess systemic reactions. Anaphylaxis was evaluated within 30–40 min after challenge by monitoring rectal temperature and recording clinical symptoms according to an established scoring system56. Immediately after the last rectal temperature measurement, mice were bled from the retro-orbital plexus under ketamine-xylazine intraperitoneal anesthesia, and sera were stored at –20 °C for humoral analyses. Following blood extraction, mice were euthanized by cervical dislocation, and spleens were aseptically removed to prepare single-cell suspensions for cellular assays.
Total antibodies evaluation by ELISA
Total antibodies were measured according to the manufacturer’s instructions (#88-50630-88, Invitrogen), using 1/5000 diluted mouse serum.
Mouse Igs recognition of cnsLTP1 and natural nsLTPs by ELISA
Two types of ELISAs were performed to quantify mouse antibody responses against cnsLTP1, recombinant nsLTPs, and Pru p 3. Unless otherwise stated, all assays were performed in duplicate and washed thoroughly between steps using PBS containing 0.05% (v/v) Tween-20 (PBST).
Assays were carried out using the Mouse Ig Isotyping Uncoated ELISA Kit (#88-50630-88, Invitrogen) with minor modifications. Half-area 96-well plates (#675061, Greiner) were coated overnight at 4 °C with 500 ng/well of either cnsLTP1 or recombinant nsLTPs. Plates were washed with PBST and PBS and blocked with 125 µL blocking buffer for 2 h at room temperature (RT). A mixture of 25 µL assay buffer (PBS containing 0.05% Tween-20 and 0.5% BSA) and 25 µL of diluted mouse serum (1:1000 or 1:5000) was added and incubated for 1 h at RT. After washing, wells were incubated with anti-mouse isotype-specific antibodies for 1 h, followed by a goat anti-rat IgG (H + L) HRP-conjugated secondary antibody (#A10549, Invitrogen; 1:2000, 1 h at RT). Bound antibodies were detected using 50 µL/well TMB substrate, and the reaction was stopped with 1 M H₂SO₄ before reading absorbance at 450 nm.
To quantify Pru p 3–specific antibodies (IgG1, IgG2a, IgG2b, and IgE), high-binding 96-well ELISA plates (#439454, Thermo Scientific) were coated overnight at 4 °C with purified Pru p 3. After blocking with BSA-containing buffer, mouse sera were added at 1:8 dilution for IgE detection and 1:50 dilution for IgG isotypes and incubated overnight at 4 °C. Plates were washed and incubated with biotinylated rat anti-mouse isotype-specific antibodies, followed by avidin–horseradish peroxidase (HRP). Detection was performed with TMB substrate, and the reaction was stopped with H₂SO₄ prior to reading absorbance at 450 nm.
Mouse IgE recognition of cnsLTP1 and natural nsLTPs by DELFIA
Half-area 96-well plates were coated overnight at 4 °C with 2.5 μg/mL anti-mouse IgE antibody (MA5-16779, Invitrogen). Following coating, the plates were washed three times with PBST and three times with PBS before being blocked with 125 μL of blocking buffer for 2 h at RT. To each well, 25 μL of assay buffer (PBS with 0.05% Tween-20 and 0.5% BSA) and 25 μL of diluted mouse serum (1/1000 or 1/5000) were added and incubated for 1 h at RT. The plates were then washed, and 50 μL of biotinylated cnsLTP1 (10 µg/mL) was added to each well. After five washes with PBST and three washes with PBS, europium-labeled streptavidin (#1244-360, Revvity) was added at a 1/500 dilution in assay buffer and incubated at 19–21 °C for 30 min. The plates were washed again with PBST and PBS, followed by the addition of DELFIA enhancement solution (#4001-0010, Revvity) for 5 min. Finally, time-resolved fluorescence (TRF) was measured at 615 nm after excitation at 340 nm using a VICTOR Nivo plate reader (PerkinElmer).
Inhibition of IgE binding to natural nsLTPs by ELISA
ELISA plates were coated, blocked, and washed as described. Following the addition of mouse serum in increasing serial dilutions, the plate was incubated for 1 h at RT and washed three times with PBST and three times with PBS. Human serum from four allergic patients with proven IgE-mediated peach allergies was pooled and added in 1/10 dilutions to wells following 1 h incubation. goat antihuman IgE secondary antibody, HRP (#A18793, Invitrogen) was diluted (1/500) and added to wells and incubated for 1 h. Bound antibodies were detected with 50 µL/well TMB substrate, the reaction was stopped with 1 M H2SO4, and the plates were read at 450 nm.
mMCP-1 assay
To obtain an additional readout of systemic anaphylaxis, serum levels of mouse mast cell protease-1 (mMCP-1) were measured following allergen challenge. Blood was collected from the retro-orbital plexus 1 h after intraperitoneal administration of Pru p 3, allowed to clot at room temperature, and centrifuged at 10,000 rpm g for 10 min at 4 °C. Sera were stored at –20 °C until analysis. Quantification of mMCP-1 was performed using a commercial ELISA kit (88-7503-88, Invitrogen) according to the manufacturer’s instructions. Absorbance was read at 450 nm with wavelength correction at 570 nm using a microplate reader. Concentrations of mMCP-1 were calculated from a standard curve generated with recombinant mMCP-1 supplied in the kit.
TH1/TH2 cell polarization following immunization
The right iLN and spleen were harvested from the mice on the day of takedown and kept in PBS. Cells were released from the tissues by mashing through a 70 µm cell strainer, and the spleen cells were subjected to RBC lysis buffer (#00-4333-57, eBioscience) to remove red blood cells. Cells were activated in RPMI 1640 (#61870044, Thermo Fisher Scientific), 10% fetal bovine serum (FBS), and 1% penicillin/streptomycin, with 1X Leukocyte Activation Cocktail, with GolgiPlug (# 550583, BD Bioscience) for 6 h in a humidified 37 °C incubator. Cells were washed in PBS and 2% FBS (FACS buffer) and stained at 4 °C for 25 min in staining buffer containing anti-B220-BV605 (#563708, BD Horizon), anti-CD3e-PerCP (#561089, BD Horizon), anti-CD44-BV480 (#566200, BD Horizon), anti-CD62L-PECy7 (#560516, BD Horizon), anti-CD4-FITC (#100406, Biolegend), and LD-near infrared (#L34976A, Invitrogen). Cells were washed twice in FACS buffer and fixed for 15 min at RT in BD Cytofix (#554714, BD Bioscience). Cells were washed twice and incubated for 15 min in 1X Cytoperm buffer (#554714, BD Bioscience) at RT. Cells were spun down and resuspended in Cytoperm containing anti-IFN-γ-PE (#554412, BD Pharmingen), anti-IL4-BV421 (#562915, BD Horizon), and anti-IL13-APC (#159405, Biolegend) and stained for 30 min at RT. Cells were washed twice in Cytoperm, resuspended in FACS buffer, and acquired on a FACSymphony S6 (BD Bioscience). For gating strategy, see Supplementary Fig. 6.
Inhibition of allergen-induced degranulation in RBL-2H3
RBL-2H3 cells (#ACC312, DSMZ), originating from rat basophilic leukemia, were transfected using TurboFectin 8.0 (#TF81001, OriGene) with DNA encoding the human high-affinity IgE receptor α-unit (hFcεRIα) (RC203321, OriGene), resulting in their humanization (hRBL-2H3)57. These cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (#36253, Stemcell) supplemented with 5% (v/v) FBS (# F4135, Sigma-Aldrich), 300 µg/ml L-glutamine, 50 U/mL penicillin, and 50 µg/mL streptomycin. Cultures were grown in 25 cm² flasks (#156367, Thermo Fisher) at 37 °C within a humidified environment containing 5% (v/v) CO₂. Cells were subcultured twice weekly using 0.25% (w/v) trypsin with 10 mM EDTA (#25200056, Fisher Scientific). To evaluate allergen-induced degranulation, the release of N-acetyl-β-D-hexosaminidase was measured. Cells were first IgE-coated overnight in complete medium containing 5% (v/v) serum from allergic patients. The next day, the medium was discarded, and the cells were stimulated with 50 nM of allergen diluted in Tyrode’s solution (#J67607-AP, Alpha Aesar). The enzymatic activity was assessed using a colorimetric assay, as described by Vogel et al53. 1 h after stimulation, 30 µL of cell supernatant was transferred to a 96-well plate and combined with 50 µL of a substrate solution containing 40 mM citric acid, 0.35% (w/v) p-nitrophenyl-N-acetyl-β-D-glucosaminide, pH 4.5. The mixture was incubated for 1 h at 37 °C. The reaction was terminated by adding 100 µL of 0.4 M glycine (pH 10.7) to each well, and absorbance was measured at 405 nm using a VICTOR Nivo plate reader (PerkinElmer). For total β-hexosaminidase activity, cells in unstimulated control wells were lysed with compound C48/80 (C2313, Sigma-Aldrich) at 100 µg/mL58. A 30 µL aliquot of the lysate was used for the enzymatic reaction. For the inhibition experiment, mouse serum in increasing serial dilutions was preincubated for 1 h at 37 °C with the allergen prior to addition to the cells preincubated with 5% (v/v) serum from allergic patients.
Data and statistical analysis
We analyzed data and performed statistics in GraphPad Prism (Graphpad, v10.4.1). Flow cytometry analysis was performed using FlowJo (BD, v10.10.0). When comparing multiple groups to control, we conducted two-way ANOVA analyses with Dunnett’s multiple comparison tests comparing the groups to the PBS controls. When comparing two groups, we used unpaired two-tailed Student’s T-tests. For the anaphylactic model we conducted one-way ANOVA analyses assuming normal (Gaussian) distribution. We considered P ≤ 0.05 to be statistically significant and indicated P values with * for P ≤ 0.05, ** for P ≤ 0.01, *** for P ≤ 0.001, and **** for P ≤ 0.0001. For the murine serum-induced blocking of human IgEs, we fitted non-linear regressions using a three-parameter inhibition versus response equation (Y = Bottom + (Top−Bottom)/(X/IC50)) using least squares regression.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Any further requests can be addressed to the corresponding author. Source data are provided with this paper.
References
Pawankar, R. Allergic diseases and asthma: a global public health concern and a call to action. World Allergy Organ. J. 7, 1–3 (2014).
Carlson, G. & Coop, C. Pollen food allergy syndrome (PFAS): A review of current available literature. Ann. Allergy, Asthma Immunol. 123, 359–365 (2019).
Mastrorilli, C., Cardinale, F., Giannetti, A. & Caffarelli, C. Pollen-food allergy syndrome: a not so rare disease in childhood. Medicine 55, 641 (2019).
Cox, A. L., Eigenmann, P. A. & Sicherer, S. H. Clinical relevance of cross-reactivity in food allergy. J. Allergy Clin. Immunol. Pract. 9, 82–99 (2021).
Zuberbier, T., Lötvall, J., Simoens, S., Subramanian, S. V. & Church, M. K. Economic burden of inadequate management of allergic diseases in the European Union: a GA2LEN review. Allergy 69, 1275–1279 (2014).
Kleine-Tebbe, J. et al. Is allergy immunotherapy with birch sufficient to treat patients allergic to pollen of tree species of the birch homologous group? Allergy 75, 1327–1336 (2020).
Randall, K. L. & Hawkins, C. A. Antihistamines and allergy. Aust. Prescr. 41, 41–45 (2018).
Baker, D. L., Nakamura, G. R., Lowman, H. B. & Fischer, S. K. Evaluation of IgE antibodies to omalizumab (Xolair®) and their potential correlation to anaphylaxis. AAPS J. 18, 115–123 (2015).
Romagnani, S. Immunologic influences on allergy and the TH1/TH2 balance. J. Allergy Clin. Immunol. 113, 395–400 (2004).
Zissler, U. M. et al. Early IL-10 producing B-cells and coinciding Th/Tr17 shifts during three year grass-pollen AIT. eBioMedicine 36, 475–488 (2018).
Akdis, M. et al. Immune responses in healthy and allergic individuals are characterized by a fine balance between allergen-specific T regulatory 1 and T helper 2 cells. J. Exp. Med. 199, 1567–1575 (2004).
Akdis, C. A. & Akdis, M. Mechanisms of allergen-specific immunotherapy and immune tolerance to allergens. World Allergy Organ J. 8, 17 (2015).
Anvari, S. & Anagnostou, K. The nuts and bolts of food immunotherapy: the future of food allergy. Children 5, 47 (2018).
Moote, W., Kim, H. & Ellis, A. K. Allergen-specific immunotherapy. Allergy, Asthma Clin. Immunol. 14, 53 (2018).
Calderón, M. A., Cox, L., Casale, T. B., Moingeon, P. & Demoly, P. Multiple-allergen and single-allergen immunotherapy strategies in polysensitized patients: Looking at the published evidence. J. Allergy Clin. Immunol. 129, 929–934 (2012).
Hamada, K., Horiike, T., Ota, H., Mizuno, K. & Shinozawa, T. Presence of isochore structures in reptile genomes suggested by the relationship between GC contents of intron regions and those of coding regions. Genes Genet. Syst. 78, 195–198 (2003).
Finley, A. & Atkinson, E. Subcutaneous immunotherapy for pollen food allergy syndrome: a systematic review. J. Allergy Clin. Immunol. 147, AB107 (2021).
González Pérez, A., Carbonell Martínez, A., Escudero Pastor, A. I., Navarro Garrido, C. & Miralles López, J. C. Pru p 3 oral immunotherapy efficacy, induced immunological changes and quality of life improvement in patients with LTP syndrome. Clin. Transl. Allergy 10, 20 (2020).
Musa, F., Al-Ahmad, M., Arifhodzic, N. & Al-Herz, W. Compliance with allergen immunotherapy and factors affecting compliance among patients with respiratory allergies. Hum. Vaccines Immunother. 13, 514–517 (2017).
Penagos, M., Eifan, A. O., Durham, S. R. & Scadding, G. W. Duration of allergen immunotherapy for long-term efficacy in allergic rhinoconjunctivitis. Curr. Treat. Options Allergy 5, 275–290 (2018).
Gehrt, F., Xu, Q., Baiardini, I., Canonica, G. W. & Pfaar, O. Adherence in allergen immunotherapy: current situation and future implications. Allergol. Sel. 6, 276–284 (2022).
Liu, D. et al. Clinical response to subcutaneous immunotherapy at 3 years in allergic rhinitis patients is predicted by short-term treatment effectiveness. Clin. Transl. Allergy 13, e12223 (2023).
Skypala, I. J. et al. Non-specific lipid-transfer proteins: Allergen structure and function, cross-reactivity, sensitization, and epidemiology. Clin. Transl. Allergy 11, e12010 (2021).
Scheurer, S., van Ree, R. & Vieths, S. The role of lipid transfer proteins as food and pollen allergens outside the Mediterranean area. Curr. Allergy Asthma Rep. 21, 7 (2021).
Oeo-Santos, C. et al. A recombinant isoform of the Ole e 7 olive pollen allergen assembled by de novo mass spectrometry retains the allergenic ability of the natural allergen. J. Proteom. 187, 39–46 (2018).
Rivera-de-Torre, E. et al. Discovery of broadly-neutralizing antibodies against brown recluse spider and Gadim scorpion sphingomyelinases using consensus toxins as antigens. Protein Sci. 33, e4901 (2024).
James, L. K. & Till, S. J. Potential mechanisms for IgG4 inhibition of immediate hypersensitivity reactions. Curr. Allergy Asthma Rep. 16, 23 (2016).
Castan, L. et al. Overview of in vivo and ex vivo endpoints in murine food allergy models: Suitable for evaluation of the sensitizing capacity of novel proteins? Allergy 75, 289–301 (2020).
Nelson, H. S. Allergen immunotherapy (AIT) for the multiple-pollen sensitive patient. Expert Rev. Clin. Pharmacol. 9, 1443–1451 (2016).
Pajno, G. B. et al. Clinical practice recommendations for allergen-specific immunotherapy in children: the Italian consensus report. Ital. J. Pediatrics 43, 13 (2017).
Zemelka-Wiacek, M. et al. Hot topics in allergen immunotherapy, 2023: Current status and future perspective. Allergy n/a
Ferreira, F., Hawranek, T., Gruber, P., Wopfner, N. & Mari, A. Allergic cross-reactivity: from gene to the clinic. Allergy 59, 243–267 (2004).
Weber, R. W. Cross-reactivity of pollen allergens: impact on allergen immunotherapy. Ann. Allergy Asthma Immunol. 99, 203–212 (2007).
Biedermann, T. et al. Birch pollen allergy in Europe. Allergy 74, 1237–1248 (2019).
Scheurer, S. & Schülke, S. Interaction of non-specific lipid-transfer proteins with plant-derived lipids and its impact on allergic sensitization. Front. Immunol. 9, 1389 (2018).
Satitsuksanoa, P., Angelina, A., Palomares, O. & Akdis, M. Mechanisms in AIT: Insights 2021. Allergol. Sel. 6, 259–266 (2022).
Huber, S. et al. Does clinical outcome of birch pollen immunotherapy relate to induction of blocking antibodies preventing IgE from allergen binding? A pilot study monitoring responses during first year of AIT. Clin. Transl. Allergy 8, 39 (2018).
Sabbaghi, A. et al. A formulated poly (I:C)/CCL21 as an effective mucosal adjuvant for gamma-irradiated influenza vaccine. Virol. J. 18, 201 (2021).
Firacative, C. et al. Identification of T helper (Th)1- and Th2-associated antigens of Cryptococcus neoformans in a murine model of pulmonary infection. Sci. Rep. 8, 2681 (2018).
Mountford, A. P., Fisher, A. & Wilson, R. A. The profile of IgG1 and IgG2a antibody responses in mice exposed to Schistosoma mansoni. Parasite Immunol. 16, 521–527 (1994).
Xu, X. et al. Use of a Liver-Targeting Immune-Tolerogenic mRNA Lipid Nanoparticle Platform to Treat Peanut-Induced Anaphylaxis by Single- and Multiple-Epitope Nucleotide Sequence Delivery. ACS Nano 17, 4942–4957 (2023).
Jitthamstaporn, S. et al. Nucleoside-modified mRNA vaccines yield robust blocking antibody responses against major house dust mite allergens. Allergy 78, 315–318 (2023).
Li, J., Li, X., Guan, K. & Yin, J. Allergen-specific immunotherapy with mRNA vaccines reduces allergic airway inflammation in mice. J. Allergy Clin. Immunol. 155, AB288 (2025).
Kanjarawi, R. et al. Regulatory CD4+Foxp3+ T cells control the severity of anaphylaxis. PLoS ONE 8, e69183 (2013).
Shao, X. et al. Leveraging an mRNA platform for the development of vaccines against egg allergy. Vaccines 13, 448 (2025).
Madeira, F. et al. Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res. 50, W276–W279 (2022).
Damsbo, A. et al. A comparative study of the performance of E. coli and K. phaffii for expressing α-cobratoxin. Toxicon 239, 107613 (2024).
Wilke, S., Krausze, J. & Büssow, K. Crystal structure of the conserved domain of the DC lysosomal associated membrane protein: implications for the lysosomal glycocalyx. BMC Biol. 10, 1–15 (2012).
Polack, F. P. et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Lee, Y., Jeong, M., Park, J., Jung, H. & Lee, H. Immunogenicity of lipid nanoparticles and its impact on the efficacy of mRNA vaccines and therapeutics. Exp. Mol. Med 55, 2085–2096 (2023).
Szebeni, J. et al. Insights into the structure of comirnaty Covid-19 vaccine: a theory on soft, partially bilayer-covered nanoparticles with hydrogen bond-stabilized mRNA–lipid complexes. ACS Nano 17, 13147–13157 (2023).
Münter, R., Larsen, J. B. & Andresen, T. L. The vast majority of nucleic acid-loaded lipid nanoparticles contain cargo. J. Colloid Interface Sci. 674, 139–144 (2024).
Roesler, E. et al. Immunize and disappear—Safety-optimized mRNA vaccination with a panel of 29 allergens. J. Allergy Clin. Immunol. 124, 1070–1077.e11 (2009).
Rodriguez, M. J. et al. Pru p 3-Epitope-based sublingual immunotherapy in a murine model for the treatment of peach allergy. Mol. Nutr. Food Res. 61, 1700110 (2017).
Rodriguez, M. J. et al. LPS promotes Th2 dependent sensitisation leading to anaphylaxis in a Pru p 3 mouse model. Sci. Rep. 7, 40449 (2017).
Li, X. M. et al. A murine model of peanut anaphylaxis: T- and B-cell responses to a major peanut allergen mimic human responses. J. Allergy Clin. Immunol. 106, 150–158 (2000).
Vogel, L., Lüttkopf, D., Hatahet, L., Haustein, D. & Vieths, S. Development of a functional in vitro assay as a novel tool for the standardization of allergen extracts in the human system. Allergy 60, 1021–1028 (2005).
Hernández-Aguilar, I., Vizuet-de-Rueda, J. C., Galván-Morales, M. Á, Montero-Vargas, J. M. & Teran, L. M. Rapid generation of an RBL cellular model to study proteins that cause allergenic reactions in vitro. Immunol. Res. 72, 874–879 (2024).
Acknowledgements
E.R.dT acknowledges support from DTU Discovery Grant, DTU Proof of Concept, Innovation Fund Denmark InnoExplorer [2071-00021] and DFF Inge Lehmann program [4306-00008B]. A.H.L. acknowledges support from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme [850974 and 101112851] and the Villum Foundation [00025302]. K.H.J. acknowledges support from Lundbeckfonden [R347-2020-2174] and Arvid Nilssons Foundation. M.T.V. acknowledges support from the Spanish Ministry of Science and Education [PID2020-116692RB-I00]. CM holds a “Nicolas Monardes” research contract by the Andalusian Regional Ministry Health (RC0004-2021). JLP acknowledges grants RYC2021-034536-I and CNS2023-145619 funded by MICIU/AEI/10.13039/501100011033 and by European Union NextGenerationEU/PRTR. CJA acknowledges grant RYC2023-043687-I funded by MICIU/AEI/10.13039/501100011033 and by ESF + .
Author information
Authors and Affiliations
Contributions
E.R.dT., K.H.J., A.H.L., and T.P.J. conceptualized the study. E.R.dT., K.H.J., M.V., T.P.J., M.M., R.M., C.M. and J.K.C. developed the methodology. E.R.dT., M.M., J.P.B., J.T., R.M., J.K.C., K.H.J., L.F.V., H.H.E., U.F.F., J.L.P., C.J.A., and B.M. carried out the investigation. K.H.J., E.R.dT., J.V.K., M.M., and J.P.B. handled data analysis and visualization. E.R.dT., A.H.L., K.H.J., and M.V. were responsible for funding acquisition. E.R.dT took the lead in project administration, while E.R.dT, A.H.L., O.L., V.C., J.B., M.P., S.R.H., and M.V. provided resources. E.R.dT., K.H.J., and A.H.L. supervised. E.R.dT., A.H.L., M.M., and K.H.J. wrote the original draft, and all authors contributed to the review and editing process.
Corresponding authors
Ethics declarations
Competing interests
E.R.dT., A.H.L., and T.P.J. are named inventors on a patent application (WO2023242436) based on the work presented in this paper. The rest of the authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ronald Van Ree and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Møiniche, M., Johansen, K.H., Parrón-Ballesteros, J. et al. An mRNA-delivered consensus allergen induces a neutralizing IgG response against food and pollen allergens. Nat Commun 17, 2402 (2026). https://doi.org/10.1038/s41467-026-69134-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69134-4