Abstract
Corals form their reef-building aragonite (CaCO3) skeletons via transient precursor phases yet understanding of the dynamics of these early-stage transformations remains incomplete. Using time-independent myriad mapping (MM) at 50 nm resolution, we map five mineral phases near the skeleton surface of Stylophora pistillata corals grown in varying seawater pH. All precursors, crystalline and amorphous, exhibit a consistent exponential decay from the growth front, with a shared decay length of 0.7 ± 0.1 μm, independent of time, phase, or pH. This spatial decay, paired with the constant growth rate of the skeleton, reveals a decay time of 5.1 ± 0.5 minutes. The dominant precursor is not amorphous but crystalline: calcium carbonate hemihydrate (CCHH, CaCO₃·½H₂O). These results suggest that exponential crystallization kinetics govern coral biomineralization and may be a widespread feature in biogenic, geologic, and synthetic systems—traceable long after initial mineral deposition.
Similar content being viewed by others
Introduction
Coral reefs, a conglomerate of the intricate skeletal structures of corals, are among the most ecologically diverse systems on the planet1. However, the complex biomineralization processes by which corals form their skeletons are poorly understood2. Calcium carbonate is the main component in multiple biomineral phases on the surface of coral skeletons during their formation. Some of the attaching phases are amorphous3, others crystalline4, but all are metastable and therefore transient, eventually maturing into crystalline aragonite (CaCO3), which makes up 99% by weight of all mature scleractinian or stony coral skeletons, along with 1% organic molecules1,5. Growth occurs by both particle attachment3,6 and ion attachment7. The current understanding of skeleton growth is that amorphous calcium carbonate (ACCH2O) particles attach to the growing surface and gradually crystallize into mature aragonite, through a series of phases and phase transitions2,3,4,7. The known phases are ACCH2O, anhydrous amorphous calcium carbonate (ACC), calcium carbonate hemihydrate (CCHH)8, and monohydrocalcite (MHC)4. All these amorphous and crystalline phases are localized at and near the surface of forming coral skeletons; therefore, they are precursor phases4.
Corals form crystalline aragonite from a calcifying fluid between the skeleton and the organism7,9,10. Key questions remain about coral biomineralization mechanisms, such as how precursor phases are deposited onto the growth front, or how and when they crystallize into mature aragonite. In principle, they could start crystallizing before, during, or after attachment to the growth front. Observing the metastable precursor phases with high spatial resolution on the forming surface of coral skeletons is key to answering these questions.
Recently, using PhotoEmission Electron Microscopy (PEEM)11,12, with ~50 nm pixel resolution13, a new method was developed to enable the observation of the forming coral skeleton surface at a length scale unachievable with other imaging tools, such as confocal microscopy, used to observe the calcifying fluid and calicoblastic cells14. This method was named myriad mapping (MM)4 to highlight that it can quantitatively display many phases. In coral skeletons, myriad maps (also MMs) display the five mineral phases present on the coral skeleton surface detected by PEEM.
Expanding on recent work by Schmidt et al.4, for this work we used the MM method to explore how and where the known CaCO3 precursors and mature phase, ACCH2O, ACC, CCHH, MHC, and aragonite, are distributed in space near the surface of Stylophora pistillata coral skeletons, from corals grown at pH 8.05 and pH 7.2 seawater. Stylophora pistillata is shown in Fig. 1. The proportion of each precursor was measured quantitatively as a function of distance from the forming edge, starting from the coral skeleton’s surface and deep into the skeleton’s bulk, to investigate how precursor phases transform into mature aragonite in space and time.
This coral is widely studied because of its resilience and its fast growth in aquaria. A Optical micrograph of one nubbin from a living Stylophora pistillata coral, showing the polyps with 6, 12, or 18 tentacles, some extended, others retracted. Each polyp resides in a hole called a corallite. Tissue joins all polyps and thus covers the skeleton completely. Near the top of the nubbin, the skeleton is visible through transparent tissue. At this stage, in fact, there are very few zooxanthellae, which progressively colonize the tissue and render it darker (bottom). The forming skeleton subject of this study is at the top of the nubbin, its growth is most active, and it is brightest with this illumination. The photograph was acquired using a stereomicroscope (Zeiss Axio Zoom V16), with side LED illumination to better reveal the transparency of the skeleton and tissue. B Scanning electron microscope (SEM) image of one corallite, subdivided into 6 sectors by 6 walls termed septa. The width of a corallite ranges in size between 0.5 and 1.5 mm. Notice the nanoparticulate surface of the skeleton, revealed by tissue removal, and partial etching in water of the forming surface, where the most metastable and therefore most soluble phases are removed preferentially. Both the image and micrograph are typical of thousands of repeat observations.
Results and discussion
Profile of phase transformation near the edge
Eight areas of skeleton surfaces were acquired from corals grown in seawater at pH 8.05 (5 areas) or pH 7.2 (3 areas). Minimal radiation damage was done to these areas during PEEM data acquisition for each area and its repeat15.
Myriad maps (MMs) of the eight ~50 µm square areas were acquired near the tips of coral nubbins where growth velocities are maximum5 and precursor phases are expected to be more abundant in forming coral skeletons. To generate an MM, the unknown spectrum at each pixel is fitted by a linear combination of five known component spectra. The resulting coefficients, constrained between 0 and 1, are the proportion of each component spectrum. This procedure yields five “proportion maps (pMaps),” one per spectral component, where pixel values correspond to the local abundance of that phase. These maps are then merged into a single MM using five distinct colors. To ensure quantitative fidelity, we restrict the display to dominant phases by omitting proportions below 0.5. Each pixel is assigned a single color, with brightness encoding the proportion (e.g., dark to bright cyan corresponds to 0.5–1.0). While information on mixed phases is not retained, the displayed pixels are fully quantitative.
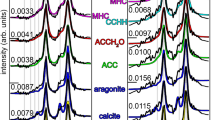
Representative MMs constructed from the five component spectra are shown in Fig. 2A, B. The component spectra are also shown in Fig. 2B, acquired from synthetic ACCH2O, ACC, CCHH, MHC, and aragonite. These are small regions from 2 of 8 total areas analyzed. Figure 2A, B shows only 200 × 200 pixel regions. Supplementary Fig. 1 presents the 8 areas in their entirety, seven of which are at the forming edge of skeletons, sometimes termed thickening deposits (TDs)16. One area (area 5) on a spine contains several centers of calcification (CoCs), sometimes termed rapid accretion deposits16,17,18 or centers of rapid accretion, as well as TDs19, more than 4 μm from the fresh forming surface of the skeleton. The CoCs in area 5 are far enough away from the surface that no pixels from CoCs are included in this work. This is a relevant distinction, as CoCs and fibers are known to form with different amounts of organics, and different kinetics17,19,20.
A MM of a portion of a Stylophora pistillata area of coral skeleton grown at pH 8.05 and B pH 7.2. Pixels are 56 nm in (A) and 45 nm in (B). Pixels with only proportions below 0.5 are displayed in black in both MMs, whereas a pixel where one phase has a proportion of 0.5–1.0 is a colored pixel, as indicated by the color legend in (B). The X-ray absorption near-edge structure spectra (345–355 eV) of 4 precursors and aragonite are also shown in (B), color coded as MM pixels, except for the aragonite spectrum, which is plotted in white rather than blue. C, D Proportion profiles as a function of distance from the edge for each precursor and mature aragonite. In (C, D), each colored datapoint is the average proportion of each phase at each distance, calculated from 106 pixels. The colored curve is the best-fitting exponential decay or exponential rise (Supplementary Fig. 2), and the R2 indicates the goodness of fit. The color legend in (B) identifies biomineral precursors and mature aragonite phases by color and extends to all precursor maps and plots in this work. Magenta data points are open circles to distinguish them from red. The precursor proportions for each of the five components in each of the eight areas are shown in Supplementary Fig. 3.
From each MM, the proportions of all precursors as a function of distance from the forming coral skeleton surface, hereafter termed “the edge”, were analyzed. All displayed similar profiles: the proportion profiles all decay exponentially within the first 4 μm from the edge. In the same 4 μm region, the aragonite proportion increases exponentially. Several candidate mathematical functions were considered to fit the experimental profiles: linear, quadratic, logarithmic, and exponential functions. Best fits were determined by R2 values. The corresponding equations and fit parameters are presented in Supplementary Table 1, and an example of fitting functions to experimental data is shown in Supplementary Fig. 2. The best-fit, the exponential function, is shown below, where x is the distance from the edge in μm, and a, b, c are fit parameters where a = the precursor proportion at x = 0 μm, b = the decay rate in μm−1, and c = the precursor proportion at x ~ 4 μm. Equation (1) is presented in Supplementary Table 1 as s(x).
The profiles for each precursor and mature aragonite in all pixels of the two represented areas in Fig. 2A, B, and their best fits, are shown in Fig. 2C, D as a function of distance from the edge. The mean proportions for each of the 8 areas are shown in Supplementary Fig. 3. The profiles of Fig. 2C, D, and additionally Supplementary Fig. 3 demonstrate that all profiles are exponential decays. All data fits have R2 = 0.85 or higher in Fig. 2 and 0.5 or higher in Supplementary Fig. 3.
We emphasize that this is the general profile of precursor transformation for all 8 regions, for all precursors across all pH values. Crystallization of all precursor phases in Stylophora pistillata occurs exponentially as a function of distance from the edge.
Constant growth velocity of the edge
The data of Fig. 2 and Supplementary Figs. 1 and 3 capture crystallization in space, that is, as a function of distance from the coral skeleton edge. These spatial data, however, can be linked to crystallization in time by a simple growth model. In this model, the coral skeleton edge grows with velocity v. In each time increment, dt, a layer of precursors with thickness dx is deposited at the edge. The layer, once deposited, begins to transform into a more mature phase. New precursor layers are constantly deposited over time on top of preexisting layers that are already transforming. After deposition, precursors exponentially crystallize into mature-phase aragonite. The edge grows away from older layers with velocity v = dx/dt, as shown in Fig. 3A.
A Model of precursor deposition at the edge. The edge is moving at velocity v away from older layers as newer precursor layers are deposited over time. B The constant growth velocity, in mm per day, measured along the longest axis of Stylophora pistillata nubbins over a 50-day period on 5 nubbins. All growth data are presented in Supplementary Table 3. An average growth velocity of ~210 ± 20 μm per day was observed, where 20 μm is the standard deviation on the calculated mean, 210 μm per day, shown as the gray band. The linear fit of mean experimental growth data (black line) has R2 = 1.
In the model of Fig. 3A, the velocity v function that generates the best fit to the experimental data is assumed constant after experimental confirmation (R2 = 1.00) from an independent experiment measuring the growth velocity at the macro scale in Stylophora pistillata. Five additional coral nubbins were suspended by nylon threads and grown at a constant temperature of 25 °C and pH 8.05. After an initial lag phase, the growth along the longest axis of each coral nubbin was measured. These measurements are shown in Fig. 3B and Supplementary Table 2. We observed a constant growth velocity v of ~210 ± 20 μm per day.
Crucially, constant v means that exponential crystallization can be described as a function of either space or time, interchangeably.
Access to early stages of crystallization
Coral skeleton growth and crystallization are distinct and independent phenomena. Thus, the crystallization rate and growth velocity of the coral skeleton edge are independent of each other. We can, however, use the exponential decays in space and the constant growth velocity in Stylophora pistillata v = 210 ± 20 μm per day to explore the early stages of biomineralization.
In any exponential decay, the rate at which a quantity decays is determined by a decay constant 1/e and is proportional to the current or local amount of material21. The 1/e constant is the best-fit parameter b presented in Eq. (1). Decay constants from each exponential fit as a function of distance x from the edge and time (\(t=\frac{x}{v}\)), including Fig. 2, are presented in Supplementary Fig. 4 and Supplementary Table 3 and the mean length and time constants are presented in Supplementary Fig. 4.
At both pH 7.2 and 8.05, there is a spread of initial precursor proportions at the edge (Supplementary Fig. 3). Unexpectedly, however, the decay constants are consistent across all areas, pH values, and precursors, within the standard deviation error. Supplementary Table 4 shows that even when separating by pH, they are consistent. This is not surprising, because Venn et al. showed that across the same range of seawater pH used here, the calcification rate in Stylophora pistillata corals remains constant as pH decreases, especially during the day when most of the skeleton deposition happens22. Chemically, each precursor is different8,23,24,25,26, thus, precursors could in principle crystallize at their own different rates, but this was not observed in this work. Of note, a transformation rate of 0, that is, instantaneous crystallization, would show total precursor and mature aragonite proportion remaining constant with distance from the edge, which is again not observed in this work.
Since all 1/e constants are consistent, we can represent the exponential transformation of each precursor in each area at each pH by averaging them. The proportions from MMs across all 8 areas, presented in Fig. 4A, were combined to calculate a single mean distribution for each precursor and mature aragonite as a function of distance (Fig. 4B). All fits of the mean have R2 = 0.98 or greater, and all precursor decays are exponential as a function of space. Since v is constant, we can convert these spatial decays into time decays using \(t=\frac{x}{v}\). These data are presented in Fig. 4C and Table 1.
A Mineral phase proportion vs. distance from the edge for 8 million pixels. B The mean proportion for each precursor and mature phase (black or blue proportions on left or right axes) and their exponential fits. The gray data points at the top of (B) are the sum of all proportions at each distance, which is close to 1 (right vertical axis). C Mean precursor proportions decay exponentially as a function of time. The most abundant phase at the surface at t = 0 is CCHH, followed by ACC, and the least abundant are MHC and ACCH2O. Table 1 shows these results numerically. Colors indicate phases as presented in Fig. 2B and extend to all precursor maps and plots in this work. Magenta data points are open circles to distinguish them from red.
The results of Fig. 4 and Table 1 show the profiles and behavior of all areas both as a function of space and time and provide unique new insights into the early stages of biomineralization: (i) crystallization is fast, with 1/e length = ~0.7 ± 0.1 µm and 1/e time τ = 5.1 ± 0.5 min. (ii) The most abundant phase at the surface at t = 0 (when the animal deposited particles and ions on the edge) is not ACCH2O as previously assumed, but CCHH. After CCHH, in order of decreasing abundance ACC, MHC, and ACCH2O are observed. (iii) The mature phase is invariably aragonite. (iv) All decay constants are consistent within the uncertainty, despite dramatic differences in phase transitions: dehydration, and/or crystallization, and/or crystal structure transformation. (v) The bulk of coral skeletons, greater than 4 µm from the edge, is aragonite in agreement with ample previous observations1,5,27,28. (vi) Mass is conserved; thus, the sum of all precursors and mature aragonite is 1 at all distances from the edge.
Fast and uniform crystallization rate
As in all spatial decays, for all time decays the best fits are exponentials, with excellent fits and R2 > 0.98 (Fig. 4C). The consistent crystallization rate of ~5 min for all phases is unexpected and extremely fast, especially considering that all spatial decays were observed a minimum of 10 days and a maximum of 6 months after the death of the animals (Supplementary Table 5). All corals were fixed, dehydrated in ethanol, embedded, and remained embedded (away from air and water) for 10–180 days.
Crystallization of precursor phases is known to occur immediately upon contact with water. During sample cutting, grinding, and polishing, we only used water greatly supersaturated with respect to ACCH2O or any other phases (22 g/L Na2CO3) as a coolant, to prevent premature crystallization. All other methods of sample preparation failed. Even with these precautions, however, minutes and months are very different timescales for crystallization. Why are we even able to detect any precursor phases?
The exponential decays provide a plausible explanation for this observation. In any exponential decay, it doesn’t matter if one detects the phenomenon at its beginning (e.g., during deposition by the animal and fast crystallization) or long after it (e.g., after days or months, at a synchrotron, when crystallization happens slowly), one can fit an exponential and obtain the 1/e time constant, which is constant at any point in time along the exponential decay21. The 1/e time τ is time-independent. Thus, the spatial resolution of MMs, which are also time-independent, enables an exponential fit in space. Constant v makes it possible to convert space into time, and therefore calculate the 1/e time, as shown in Fig. 4.
Similarly, in radiocarbon dating, exponential decay of 14C into 12C makes it possible to reconstruct the age of the last naturally abundant carbon consumption by animals up to 50,000 years into the past29. In dating of zircons, the exponential decay of U into Pb isotopes enables dating of rocks as old as 4.4 billion years30. Here, dating is uninteresting because we know precisely when the animals were fixed. But we can reconstruct the relative abundances of precursors when Stylophora pistillata corals deposited their skeletons during the life of the corals, from the first moment precursors were deposited onto the edge. This was done here using a time-independent method of data taking, MM. Thus, from the slow-stage of crystallization in space, we reconstructed the fast-stage of early crystallization in the living corals.
Importantly, having now demonstrated that fixed, dehydrated, and embedded coral samples can be as old as 6 months and still have some precursor phases, other methods can be used to analyze precursors, including high-resolution infrared4 or Raman spectroscopies, transmission electron microscopy, or nanoscale X-ray diffraction (nano-XRD).
CCHH is the most abundant precursor at deposition
What happens to precursors over time during biomineralization? The data of Fig. 4 and Table 1 answer precisely this question: the most abundant phase on average deposited by Stylophora pistillata corals onto their growing skeletons is aragonite, at more than 80%. Among the early precursors, the most abundant is not, as expected in Stylophora pistillata, the most metastable ACCH2O3,4. The most abundant precursor is CCHH, which is deposited with an ~8% abundance by the animals. CCHH is a new calcium carbonate phase discovered only 7 years ago, which at that time was only prepared synthetically8. Two years ago, CCHH was observed on the surface of coral skeletons from six species, including Stylophora pistillata4. Here we find that it is the most abundant in the first 5 min of skeleton deposition, and resiliently so at all pH values (Fig. 2, Supplementary Fig. 3, Fig. 4, Table 1). CCHH is also abundantly found on the surface of forming nacre4; thus, it is a ubiquitous precursor in aragonite biominerals.
The second most abundant precursor is anhydrous ACC, at 4% abundance. This phase was first observed by Politi et al. in sea urchin spicules31, then by Killian et al. in sea urchin teeth32, then in nacre by DeVol et al.33, in the coral skeletons of Stylophora pistillata by Mass et al.3, and in other coral species by Sun et al.7 and Schmidt et al.7,34. Thus, the surface abundance of ACC was not a surprising observation. The relative paucity of ACCH2O, expected to be the first deposited and most abundant surface phase for Stylophora pistillata3, has only 2% abundance. This is surprising, as ACCH2O was expected to be the first deposited and most abundant surface phase by Mass et al.3.
The Mass et al.3 paper discusses only ACC and ACCH2O precursors because, with component mapping or myriad mapping, one can only find phases for which one has the X-ray absorption near-edge structure spectrum (Fig. 2B) and fits the unknown spectrum from each pixel with a linear combination of known spectra. CCHH was not discovered by Zou et al., until 20198; thus, its existence was unknown in 2017. Once we noticed that some spectra from coral skeletons were not well fit by either ACCH2O, ACC, or any combinations thereof, we broadened the search, included CCHH, MHC, and vaterite, then excluded vaterite because the fits are not significantly improved by fitting with six rather than five component spectra, and vaterite was the worst fitting of the three metastable crystalline precursors4.
Kinetic model
A kinetic model was designed to understand if the exponential decay of precursors into aragonite is sufficient to generate the results observed in Stylophora pistillata skeletons, or if other phenomena must be taken into account. Figure 5A presents a simulated MM built using the initial precursor proportions N0 reported in Table 1 and assuming a general exponential decay of the form \(f\left({dt}\right)={e}^{\frac{-{dt}}{t}}\), where τ = 5.1 min is the average decay constant (1/e time, or lifetime) listed in Table 1 and dt is the time increment during which a new layer of precursor with thickness dx is deposited at the growing edge, as shown in Fig. 3A.
A A portion of MM (80 × 80 pixels) simulated using the initial concentration N0, a in Eq. (1), for each precursor phase and the average τ = 5.1 min reported in Table 1, dx = 50 nm, and dt = 24 s. The full 200 × 200 pixel simulation is presented in Supplementary Fig. 6. On the horizontal axis is a 4 µm distance from the edge. B The proportion for each precursor and mature phase extracted from 10 simulated MMs, compared to the mean experimental data displayed in Fig. 4B. The simulated precursor proportion is the average ± StDev of 10 simulations. Colors indicate phases as presented in Fig. 2B and extend to all precursor maps and plots in this work. Magenta data points are open circles to distinguish them from red.
The deposited layer thickness, dx, in this experiment corresponds to the spatial resolution of the data acquired by PEEM and the resulting MM, with dx = 1 pixel ≈50 nm. The time increment dt was optimized during model construction to fit the experimental data, and the best-fit value was found to be dt = 24 s. This value corresponds to a growth rate of ~180 µm per day, which is similar to the measured growth rate of 210 ± 20 µm per day shown in Fig. 3B.
The consistent growth rate and close match of simulated and experimental data near the edge in Fig. 5B confirms that, to first-order approximation, a simple kinetic model assuming only exponential decay of phases into aragonite is sufficient to describe how all precursor phases transform into aragonite at the growing edge of Stylophora pistillata coral skeletons.
However, small deviations between simulated and experimental data indicate that second-order processes also impact phase transformation. In Fig. 5A, precursors occur mostly as single pixels, not clusters of clumped precursor pixels as seen in experimental MMs (Fig. 2A, B, Supplementary Fig. 1). In Fig. 5B, 2–4 µm into the skeleton, the model deviates from experimental data. This deviation indicates that long-lived precursor phases exist in the natural skeleton, possibly kinetically trapped, which were not included in the model. These are small, but nonzero proportions of all precursor phases present deep in the skeleton. Additionally, individual phase characteristics of ACCH2O, ACC, CCHH, and MHC, release of water, diffusion, etc., are not accounted for in the model.
Implications of exponential crystallization in coral skeletons
The observed exponential decay of precursor phases into aragonite suggests that the transformation of precursors to aragonite is not a nucleation-limited phenomenon. Since nucleation happens within the already deposited solid, it must be heterogeneous nucleation of aragonite on the surface of the precursor phases35. If nucleation were rate-limiting for transformation to aragonite, one would expect sigmoidal kinetics, with an induction period, then acceleration, then slowdown36. Similarly, cooperative or autocatalytic phenomena, such as one already crystallized particle or pixel inducing the transformation of another, would resemble sigmoidal functions, not exponentials37.
Diffusion-limited processes are also ruled out by exponential decays. Diffusion-limited phenomena give power laws that follow the square root of time or the inverse square root of time (t 1/2 or t −1/2), not exponentials38.
The observed exponential decays are diagnostic of a memoryless phenomenon. Thus, the transition to aragonite as approximated by the kinetic model is, to first order, a simple Poisson process, a physical mechanism that is a counting process in which every pixel of the precursor phase has a stochastic probability of converting to aragonite.
Furthermore, having observed consistent τ for all phases suggests that many precursor phases, despite their chemical differences, share the same first-order kinetics and therefore the same activation barrier. Speculative explanations for consistent decay rates include physical constraints on coral skeleton growth, such as, for example, the effect of confinement on crystal formation39. Confinement changes the kinetics or thermodynamics of crystallization by restricting the dimensions of the system in one, two, or three directions39. Speculative organic examples include generalist catalysts or inhibitors, such as protons doing acid/base catalysis, or water, or ions or molecules that regulate reaction rates for multiple, diverse reactions or phase transitions. Generalist enzymes are well-known in bacteria40, but have rarely been reported in corals41.
Lastly, and most puzzling, is the fact that we observed exponential decays up to 6 months after the death of the animal. Six months is 6 × 30 × 24 × 20 = 86,400 = ~105 times longer than the 5-min decay time τ of all precursors. Other exponentially decaying phenomena (14C, U/Pb, etc.) are usually detected after ~10 decay times τ, not 105 29,30. Kinetic trapping cannot explain this puzzle, as kinetically trapped phases would be randomly distributed throughout the mature skeleton and have flat concentrations, not decay exponentially from the edge.
The consistent time constant τ ~ 5 min was observed for all precursor phases (Table 1). Consistent τ across all phases could suggest some precursor-stabilizing molecules in corals, known to exist in other biominerals24,42,43,44,45, but why would different molecules stabilize different phases similarly? A physical phenomenon is more likely, but this is currently not understood. If it is a physical phenomenon that leads to exponential crystallization with similar 1/e constants for all phases, this must also take place in other biogenic, geologic, or synthetic systems. And it can be detected not only with PEEM and MM as done here, but with a variety of spectroscopic or diffraction methods in a variety of forms of natural minerals, biominerals, or lab-grown materials.
Furthermore, fast crystallization may explain why Stylophora pistillata is resilient to seawater acidity: ACCH2O is 100x more soluble than aragonite25,46, and even more soluble at lower pH. Fast crystallization to aragonite could make the skeleton insoluble even when seawater and the extracellular calcifying fluid are more acidic. Future investigations may reveal that the spatial decay profile and crystallization time constants for Stylophora pistillata are general for all corals, or that each species has its own constants. The deep-sea corals that form their skeletons at lower seawater pH values are of particular interest. How does their crystallization rate compare to surface corals?
Directly capturing crystallization as it unfolds—with both spatial resolution and phase identification—remains a major technical challenge across disciplines. Consequently, the earliest stages of biomineralization have largely eluded direct observation. Here, through indirect yet spatially resolved myriad mapping, we approach a near-real-time view of coral skeleton formation, revealing an unexpectedly rapid crystallization process in the slow-forming biogenic minerals. These results provide a rare glimpse into early-stage crystallization processes that are otherwise difficult to capture, particularly in non-biogenic systems. Whether or not the exponential crystallization behavior observed here reflects a more general mechanism, occurring in other biominerals and geologic minerals, remains unknown. Our findings, however, provide the methods and the software to broaden spatial decay analysis to reveal exponential crystallization decays in other biogenic, geologic, and synthetic materials, if they exist.
Methods
Samples
Stylophora pistillata corals used in this analysis were grown in aquaria at the Centre Scientifique de Monaco (CSM). The aquaria are constantly supplied with fresh seawater directly from the Mediterranean at pH 8.05, controlled at a temperature of 25 °C and salinity of 38 ppt, exposed to 10,800 Lux, and given Artemia salina nauplii as marine food twice a week4. Rotifers are also provided as food daily14,45.
Before fresh seawater from the Mediterranean reaches the aquaria, it goes through an intermediate header tank, which is bubbled with CO2-free air to achieve a constant pH level of 8.05 for the control tank. Another intermediate header tank was bubbled with CO2 to achieve a constant pH level of pH 7.2 for the acidified tank. Diligent monitoring of pH with instantaneous pH monitoring and CO2 adjustment is used to precisely maintain pH in the intermediate tanks and the aquaria. While these growing conditions cannot perfectly recreate environmental conditions of coral reefs, CO2 bubbling allows replication of the changing seawater chemistry corals face in more acidic environments45.
Preparation
The 4 + 5 = 9 nubbins were used in this analysis, two at pH 8.05 and two at pH 7.2, 2 cm in length, for PEEM analysis and 5 at pH 8.05 for the growth rate measurements in Fig. 3B. The 4 nubbins for PEEM analysis were removed from the aquaria and immediately fixed, dehydrated, and embedded in EpoFix in 1-inch round molds at CSM before shipment to the Berkeley, CA34. The preparation at CSM was optimized to protect precursors from transforming into mature aragonite.
At Lawrence Berkeley National Lab (LBNL), samples were cut to expose the tip of each coral nubbin prior to experiments using PhotoEmission Electron Microscope (PEEM-3) on beamline 11.0.1.1 at the Advanced Light Source (ALS) at LBNL4,13. They were cut, ground, polished, and trimmed using only 22 g/L Na2CO3 in DI water as a coolant, to avoid dissolution of soluble phases or crystallization of precursor phases. They were then dehydrated in ultra-high vacuum, then coated with 1 nm Pt in the central portion containing coral skeleton and 40 nm Pt elsewhere47 to make the sample a necessary conductor for high voltage acquisitions in PEEM11. Time of exposure to air was minimized to increase the probability of observing precursor phases near the edges of coral skeletons using PEEM. No samples were analyzed with more than 28.5 h from cutting to analysis, as presented in Supplementary Table 5. The samples used for this analysis and their harvesting and preparation procedures are also described in previous publications4,7,34.
Acquisition with PEEM
Data from the coral samples were acquired using PEEM-3 on beamline 11.0.1.1 at the ALS, at LBNL. PEEM-3 can obtain higher spatial resolution, but for these experiments, we used ~50 nm pixel resolution11,12,13. All acquisitions were performed at the Ca-L edge, where various phases of CaCO3 near the coral skeleton edge are the most distinguished4.
Data were acquired in the form of Stacks, which are arrays of 121 PEEM-3 images, each containing 1030 × 1054 pixels, acquired while scanning across the Ca-L edge. Each Stack is a Ca movie, over photon energy from 340 to 360 eV with circular polarization to reduce the effect of crystal orientations in PEEM-3 images. Energy increments were 0.1 eV between 345 and 355 eV where all CaCO3 phases have distinct peaks4, and 0.5 eV increments outside this range. Two Stacks for each area were taken to ensure data quality and reproducibility during analysis, and to observe if phases tended to transform between movies. All movies used for this analysis were taken in virgin areas, with no prior exposure of the coral skeleton area to the soft-X-ray beam.
Myriad Mapping (MM)
Myriad mapping, or making myriad maps MM is a data processing and visualization strategy to quantitatively map the spatial distribution of multiple components with distinct, known characteristics. MM was first introduced in 2024 by the Gilbert group and published by Schmidt et al.4. MM can be used for any number of phases and at any length scale, but in the present work it was used to display quantitatively 5 mineral phases, with known spectra at the Ca-L edge, from Stacks acquired at and near the forming edge of Stylophora pistillata coral skeletons. MM is superior to component mapping34 because MMs allow for mapping of any number of phases, instead of the traditional maximum of 3 in RGB or CMY imaging. In MMs different biomineral phases are displayed in different colors, and only phases with proportion >50% are displayed with brightness representing the precise proportion of each colored phase, between 50% (darker), intermediate, and 100% (brighter color), as previously done by Schmidt et al.4. Pixels with mixed phases are not shown in color but displayed as white pixels.
In this work, the five component phases were amorphous calcium carbonate hydrated (ACCH2O), amorphous calcium carbonate (ACC), calcium carbonate hemihydrate (CCHH), monohydrocalcite (MHC), and aragonite (CaCO3). Stacks acquired on PEEM-3 across the Ca-L edge underwent rigorous spectral processing to produce MMs for the 8 areas used in this analysis (Supplementary Fig. 1)4,7,34. A summary of the processing includes aligning and averaging the Stacks, thus producing an average image of each Stack. The open-source package GG Macros software was developed by our group to produce average images of each Stack, and is distributed free of charge on GitHub and Zenodo48. GG Macros v1.0.0 run in Igor Pro 8 was used for this analysis (Wavemetrics Inc., Lake Oswego, OR, USA).
Briefly, the unknown spectrum in each pixel of the Stack was best fit with a linear combination of the 5 known spectra termed Cni164, and the precise proportion (from 0 to 1) of each component phase was saved as a proportion map (pMap). For each area, 5 pMaps were produced, 4 for precursors and 1 for mature aragonite. The superposition of the five pMaps results in an MM for the area. The MMs were used for quantitative visualization of precursor phases on the forming coral skeleton surface (Fig. 2, Supplementary Fig. 1), while pMaps were used for all the statistical analysis in this work (Figs. 2,4, Supplementary Figs. 1–4, Supplementary Tables 1, 3–5).
Selection of areas analyzed
Eight 50 × 50 µm square areas from 4 forming coral skeleton samples were selected from a much larger set of PEEM data. All areas of the PEEM data were examined in a multistep process to determine precursor quantity and quality, and those that did not meet the most criteria were excluded. The first step was to produce MMs for all areas and confirm that each area exhibited a presence of precursor phases. Many areas only contained aragonite, presumably because they had already crystallized fully at the time of analysis; thus, they had to be excluded. Secondly, two stacks and therefore 2 MMs were acquired and processed for each area, to determine the energy landscape of all pixels in that area. If any phase transitions from the 1st to the 2nd Stack went thermodynamically uphill, in any pixels of an area, that area was excluded from this analysis. A third and final step to select final areas for this analysis was to examine the precursor abundance deeper (>4 µm) into the coral skeleton. Any areas where one or more precursors increased as a function of distance from the edge were excluded. This only occurred in a few areas, and in those areas, the reason for the precursor increase may have been the presence of CoCs near the edge. CoCs are known to contain amorphous precursors for a decade post-mortem3,33,49,50.
The eight areas selected passed all three criteria49. These eight areas were analyzed to reveal the profiles of biomineral phase proportions as a function of distance from the forming surface of the fresh skeleton, termed “the edge”. A total of five areas at pH 8.05 and three areas at pH 7.2 were analyzed. In each of these eight areas, the proportion of biomineral precursors, ACCH2O, ACC, CCHH, or MHC, and mature aragonite present was measured in each pixel and displayed quantitatively in MMs. The proportion of each phase in an area can range between 0 and 1, and the sum of all precursors and mature aragonite must equal 1. In this experiment, proportions were collected from pMaps of each phase, each containing 106 pixels, and each pixel was ~50 nm in size in all 8 areas × 5 pMaps = 40 pMaps, thus 40 million pixels in total.
Proportion profiles
A coding script called Precursor Phase Distance (PPD) was developed in MATLAB to analyze spectromicroscopy data, specifically to collect the proportion of each precursor and mature aragonite as a function of distance from the fresh forming coral edge to 40 µm deeper into the skeleton bulk, using their respective pMaps resulting from PEEM acquisition and MM processing. Several inputs are used for the analysis of an area, but defining the edge is the most important decision. To do this, we used two images, both produced by GG Macros v1.0.048 for each area, and refined in Photoshop: the Calcium Mask and the Edge Mask.
Difference masking
The difference map (DifMap) is obtained by averaging 5 images on-peak, acquired around peak 1 at 352.6 eV, and 9 images off-peak, acquired where there are no Ca peaks, around 344 eV, and doing a digital subtraction of on-peak minus off-peak images. Averaging is done beforehand to reduce noise. This DifMap produces a Ca distribution map, which is then thresholded to a minimum value that still enables spectra to be recognizably one phase or another. Where the threshold is placed, numerically, on each DifMap is decided by looking at spectra near the edge, by assessing their signal-to-noise ratio in each area. The thresholded DifMap is the DifMask. The precursors and mature aragonite in this work are Ca-rich phases; thus, they have a high signal-to-noise ratio, and this drops abruptly at the edge. Flat, noisy spectra are excluded by the DifMask, and good spectra are retained. Once exported, the DifMask is applied to the MM to exclude non-skeleton pixels and mask them as black.
χ 2 masking
The goodness of the fit in each pixel is calculated in Igor during MM and saved as a χ2 map (X2Map). Each X2Map is thresholded to the maximum acceptable value of χ2 = 0.01. Lower values (better fits) are retained, higher values (bad fits) are excluded. The thresholded X2Map is the X2Mask. In all X2Masks, all pixels with χ2 > 0.01 are excluded and displayed as black pixels.
Calcium mask
The χ2 and difference masks (e.g., X2Mask0.01+DifMask) are merged into a single mask for each area in Photoshop and called the Calcium Mask. The Calcium Mask identifies all good Ca spectra in each area, and therefore in each of the five pMaps for that area, thus including all pixels from the skeleton, but also those from particles or tissues outside the skeleton, or CoCs within the skeleton. In all pMaps used for the analysis of proportion profiles, pixels with good Ca spectra are white pixels, and those with noisy or flat spectra are black pixels.
Edge mask
The calcium mask defines the edge well, but still includes Ca spectra outside of the skeleton, and retains CoCs inside the skeleton. The particles outside the skeleton must be removed with a hand mask, which retains only the skeleton as white pixels and non-skeleton as black pixels, as shown in Supplementary Fig. 5A.
Precursor Phase Distance (PPD) code
Input
A complete example pMap used by the PPD code is shown in Supplementary Fig. 5. Supplementary Fig. 5A shows the Edge Mask for the MM of S49 at pH 8.05, also in Supplementary Fig. 1, and Supplementary Fig. 5B is one out of five pMaps used for PPD analysis in that area with the Calcium Mask and Edge Mask applied. All data for the analysis were collected from Edge Masked-pMaps, such as the one presented in Supplementary Fig. 5B.
In summary, the inputs needed to run PPD successfully, with defined quality, edge, and dimension, are listed below.
pMap: the proportion map for ACCH2O, ACC, CCHH, MHC, or aragonite. Proportions range from 0 to 1 for each pixel.
Calcium Mask: resulting mask from DifMask and X2Mask, overlapped, to separate Ca-rich skeleton from epoxy in the area analyzed.
Edge Mask: resulting mask from excluding particles and tissues outside the coral skeleton.
Width of the pMap: the width of the pMap (same as the width of the area) in pixels, which is always precisely 1030 pixels in PEEM-3 images.
FoV: the field of view in µm of the area as collected in PEEM. Areas acquired at pH 8.05 have FoV 56 µm, areas acquired at pH 7.2 have FoV 45 µm.
Depth: the distance from the edge at which proportions are collected, chosen to be 4 µm for this analysis.
Output
Upon running the open software program MATLAB51, the PPD first and foremost identifies the pixels from which to extract proportions. PPD is identified by combining the pMap, the Calcium Mask, and the Edge Mask into a masked pMap. A masked pMap is shown in Supplementary Fig. 5B; brighter pixels have a higher proportion, while dimmer pixels have a lower proportion. Then, the dimensions to define the area and the desired depth from the edge to collect information are used to start proportion collection from the identified pixels. The depth for this analysis was chosen to be 4 µm after testing up to 10 µm from the edge. The PPD does not have a method to determine the center of a septum, which leads to duplicate collection of pixels as the PPD moves from one side of the area to the other. Thus, 4 µm was chosen since the centers of all septa in this analysis are at least 4 µm away from the edge.
For each incremental distance up to 4 µm, the proportions of the phase defined in the pMap were collected for each pixel in the given incremental distance. The output is organized as a dictionary, where each key corresponds to the distance increment and the key values are the phase proportions of all the pixels in the key52. The dictionary for each pMap was exported as a csv file, containing the collected proportions for a given phase. Analysis of these csv files was performed using Python 3.11.3 and associated Python libraries and packages, Scipy 1.10.1, Pandas 2.0.1, and Numpy 1.24.352,53,54.
Fitting methodology
Python 3.11.3, Numpy 1.24.352, SciPy 1.10.153, and Pandas 2.0.154 packages were used to analyze all 40 csv files from proportion data collection with PPD.
Formatting data
From each csv file, the distance values and their associated precursor or mature aragonite proportions were extracted and represented as NumPy arrays, 1 Dimensional (1D) and nDimensional (nD), respectively, for efficient numerical operations. The mean precursor proportion for each distance value, 1D arrays, was then calculated from the nD arrays, visualized in Supplementary Fig. 3, so that the mean precursor proportion versus distance, 1D vs 1D array, from the edge for each file was used for analysis.
Fitting data
Extensive comparisons were performed to determine the best-fitting model for the observed profiles of precursor proportion as a function of distance from the edge. The curve fit operation from the SciPy optimize package was used to determine which function best describes the data. Several candidate functions were utilized to fit the profiles observed in data because of their broad ability in describing biological, chemical, and physical mechanisms, including linear, quadratic, exponential, and logarithmic decay, as shown in Supplementary Table 1. Of note, previously, logarithmic decay was seen to best describe the trend in crystallization rates as a function of distance from the edge across several coral species in ref. 34 using Kaleidagraph® (Synergy Software, Reading, PA, USA). A bug was identified in the Kaleidagraph exponential decay fitting macro, which led to that misinterpretation. Once discovered with SciPy that the best fits are exponential, we tested if exponential and all other decays with precisely the same formulae produce the same R2 values in Kaleidagraph, and they do. Thus, the best fits are exponential curves, independent of which software is used. Exponential decay in this publication best describes all precursor decays, while exponential rise best describes the mature aragonite phase.
The SciPy curve fit operation requires several arguments: the data, a candidate function with estimated fit parameters, and an optimization method. These are:
Data: the mean precursor or mature aragonite proportion vs. distance from the edge, 1D array vs. 1D array, from 1 csv file. This is what is being fitted by candidate functions.
Candidate Function: a candidate function, linear, quadratic, exponential decay, or logarithmic decay from Supplementary Table 1, with assumed fit parameters a0, b0, and c0 as needed.
Fitting Method: an optimization method determines the parameters as needed in this analysis, such that the candidate function best fits the data. Optimization methods start with initial conditions in the form of a candidate function with assumed parameters a0, b0, and c0 as needed, and proceed to determine the best combination of parameters a, b, and c as needed using a defined method.
The SciPy optimizer operation allows for different types of optimization methods. For this analysis, the curve fit function was used, which implements the Levenberg-Marquardt algorithm55. This algorithm is suitable for a wide range of data distributions, including data that may be noisy or non-uniform, such as the data used in this experiment, and uses the nonlinear least squares method to best fit a defined function to data.
Each of these arguments, data for one proportion, candidate function with assumed parameters, and fitting method is needed to run one fit. A systematic approach was implemented to run many fits for each dataset across a range of initial conditions. A large range of initial guesses were generated, represented as 1D arrays, and iterated through in increasing order for each parameter of each function. For example, the linear function in Supplementary Table 1 has two fit parameters, a and b. Two 1D arrays of initial guesses, one array for a0 and one array for b0, were generated and iterated for each csv file. Similar 1D arrays were generated and iterated for the initial fit parameters of each function. The number of fits run for each function, for each set of data, is as follows: 100 fits for linear, 1000 fits for quadratic, exponential decay, and logarithmic decay, respectively. A total of 3100 fits for each set of data was performed.
Determining best fits
The best fit of each candidate function out of 100 for linear, 1000 fits for quadratic, exponential decay, and logarithmic decay, respectively, was determined by calculating the coefficient of determination (R2). R2 is a comprehensive method to assess the goodness of fit in general modeling systems and is described in detail here56. The R2 formulation used in this analysis is defined as 1 – the total sum of squares divided by the residual sum of squares56. Values of R2 range from 0 to 1, with values closer to 1 indicating better fits.
For the data in each csv file, 33 out of 40 resulted in the best characterization assuming an exponential decay function, and the remaining 7 resulted in a logarithmic decay or growth function, 5 of which are of the mature aragonite phase distributions. In all 7 cases, the R2 were excellent with both logarithmic and exponential curves; thus, to maintain consistency, the best exponential decay and rise function from each precursor and mature aragonite phase was used for this analysis, including 1/e length and precursor quantity calculations. A different fitting strategy to determine exponential fits of the 7 logarithmic decays/growths is described below.
Aragonite fits
Logarithmic best fits for the mature aragonite phase were reevaluated using the same exponential function but using the SciPy minimizer operation instead of the optimizer operation. Additionally, all precursor phases were reevaluated using this operation to determine the inconsistency in mature phase fits. We emphasize that all precursor phases had consistent exponential fits across both fitting operations, and using the minimizer operation resulted in exponential best fits for the mature aragonite phase.
The SciPy minimizer operation allows for the use of loss functions to place priority on fitting certain characteristics of a distribution, in this case, the region closest to the skeleton edge.
The standard loss function for the SciPy minimizer operation used in this analysis determines the quality of the minimized candidate function using a standard method, the mean squared error of the predicted proportion and the actual proportion53. In most cases, the standard loss function uses the predicted proportion calculated from the minimized candidate function. In several cases, the standard loss function failed, resulting in the best fit being a logarithmic fit for aragonite as seen using the SciPy optimizer operation.
A weighted predicted proportion was used instead, only in the case of 7 failed standard minimizations. The minimized candidate function, if the first lost function failed, included a weight to place more priority on fitting the proportion closer to the edge. The weight is given as a function \({{{\mathcal{w}}}}(x)={e}^{-x}\) where x is the distance from the edge. This was done to maximize available data for the analysis while maintaining focus on the proportion closest to the edge, the focus of this analysis. Shown below are examples of the standard \({{{\mathcal{L}}}}\) and weighted \({{{{\mathcal{L}}}}}_{{{\mathcal{w}}}}\) loss functions, Eq. (2) and Eq. (3) respectively, for a minimized candidate exponential decay, where N is the number of iterations in x, yi, and a, b, and c are fit parameters as presented in Eq. (1), defined in Supplementary Table 1 as s(x).
Implementing Eq. (3) resulted in exponential best fits for the 7 logarithmic fits whose standard loss function minimization, Eq. (2), failed. Again, we emphasize that this minimizer operation was tested on all data to ensure consistency in fits. The only inconsistent fits were the 7 fits whose standard loss function minimization failed, which were found to be exponential when prioritizing the data closer to the coral skeleton edge. All results in this analysis were determined using the best exponential fits for all precursors and the mature aragonite phase.
Preventing numerical errors
nD arrays could not be used for fitting, as each distance value contained n duplicate precursor or aragonite proportions. The mean precursor proportion, 1D array, was used for all fitting purposes to avoid numerical errors caused by duplicate distance values. The Std Deviation, 1D array, was used to represent the margin of error from measured data contained in the nD arrays, and for subsequent calculations.
The distance value closest to the edge, actually 0 µm and the first element of the 1D array, was adjusted to be 0.01 µm to avoid numerical errors in logarithmic fitting calculations described above.
Kinetic model
Framework
A simple kinetic model was developed to simulate the transformation of precursor phases into aragonite during skeleton deposition and uploaded on Zenodo and GitHub57. The goal is to build a simulated MM, one column of pixels at a time. The domain of the model MM was chosen to be 200 columns, each of height 200 pixels. Each pixel corresponds to 50 nm, the spatial resolution of data acquired by PEEM. Each column represents one discrete deposition event of thickness dx = 1 pixel ~50 nm, occurring at a time interval dt (shown in Fig. 3). For this analysis, dt is the only parameter not imposed but best-fit to the data through model testing. The best-fitting value was found to be dt = 24 s.
The initial composition of each column was defined by the experimentally observed phase proportions N0 = [2%, 4%, 8%, 2%, 82%] in Table 1, for ACCH2O, ACC, CCHH, MHC, and aragonite, respectively. Each pixel was randomly assigned a phase value, such that the proportion of each phase over all 200 pixels in each column matches N0. A new column is added at each time step, dt, up until a total of 200 columns exist.
At each time step, dt, existing columns are updated to simulate the decay of precursors into aragonite. Precursor pixels are converted to aragonite according to a Bernoulli trial with probability \({P}_{{{\mathrm{aragonite}}}}=1-{e}^{\frac{-{dt}}{{{{\rm{\tau }}}}}}\), where the exponential decay of the precursor is \({e}^{\frac{-{dt}}{\tau }}\), and τ is the characteristic decay constant 5 min measured from data and reported in Table 1. The Bernoulli trial was chosen to represent that each pixel (reaction site) has an independent probability of transforming into aragonite within a given dt. The approach captures the intrinsically probabilistic nature of phase transformation at the nanoscale while preserving the observed macroscopic exponential decay. The first deposited column, column 0 on the right, has been exponentially decaying for 200 × dt total time, and the last deposited column, column 200 on the left, has been exponentially decaying for 0 × dt total time.
Simulated MM generation
During the simulation, the total proportion of each phase at every spatial position is collected. The majority phase at each position is then determined as the phase with the greatest cumulative proportion. The majority of phases at each position are a resulting model, MM. The model MM displays the spatial distribution of all majority phases across the modeled growth region. A completed model MM is shown in Fig. 5A.
Phase proportions
Using a final model MM, the proportion of each phase as a function of distance from the growing edge is calculated. For each column, pixel counts per phase were collected and normalized by the total column height of 200 pixels. The results are proportions for each phase, for each of the 200 columns. The proportions for each phase can be plotted as a function of distance from the edge in µm.
Comparing to experimental data
The averaged results of running the simulation 10 times are compared to experimental data in Fig. 5B. The uncertainty of the simulated data is the standard deviation of the precursor proportion across the 10 trials at each column or distance.
Reproducibility
All simulations were performed using Python 3.11.3, NumPy 1.24.3, and Scipy 1.10.1 for numerical operations, and seeds for the 10 simulation trials are seed numbers 0-9.
Plotting data
All plotting shown in this work was performed using the Matplotlib 3.7.158 package in Python.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The precursor proportions generated in this work from MMs and Stylophora pistillata coral nubbin growth data have been deposited in the data.xlsx file available on Zenodo and GitHub57 with https://doi.org/10.5281/zenodo.18175786 and are source data to reproduce Figs. 2C, D, 3B, 4B, C and 5B, Supplementary Figs. 2 and 4–6, Table 1, and Supplementary Tables 2–4. The data used to produce MMs is available with no restricted access and can be obtained by contacting the corresponding author of this work at any period after publication of this work.
Code availability
GG Macros v1.0.0, Igor Pro 8, MATLAB R2023b, Python 3.11.3, Numpy 1.24.3, Pandas 2.0.1, Scipy 1.10.1, and Matplotlib 3.7.1 are used in available code, demsontrations, and software. Interactive demonstrations of the PPD code, performing exponential fits, plotting data, and the kinetic model, are publicly accessible at the following Zenodo57 and GitHub with https://doi.org/10.5281/zenodo.18175786. From available code demonstrations, all results presented in this work can be reproduced by any interested readers. Additionally, the software to produce MMs from PEEM data is available on Zenodo48 and GitHub with https://doi.org/10.5281/zenodo.17314121.
References
Cohen, A. L. McConnaughey TA. Geochemical perspectives on coral mineralization. Rev. Miner. Geochem 54, 151–187 (2003).
Von Euw, S. et al. Biological control of aragonite formation in stony corals. Science 356, 933–938 (2017).
Mass, T. et al. Amorphous calcium carbonate particles form coral skeletons. Proc. Natl. Acad. Sci. USA 114, E7670–E7678 (2017).
Schmidt, C. A. et al. Myriad Mapping of nanoscale minerals reveals calcium carbonate hemihydrate in forming nacre and coral biominerals. Nat. Commun. 15, 1812 (2024).
Gladfelter, E. H. Skeletal Development in Acropora-Cervicornis .3. A comparison of monthly rates of linear extension and calcium-carbonate accretion measured over a year. Coral Reefs 3, 51–57 (1984).
De Yoreo, J. J. et al. Crystallization by particle attachment in synthetic, biogenic, and geologic environments. Science 349, aaa6760 (2015).
Sun, C. Y. et al. From particle attachment to space-filling coral skeletons. Proc. Natl. Acad. Sci. USA 117, 30159–30170 (2020).
Zou, Z. Y. et al. A hydrated crystalline calcium carbonate phase: calcium carbonate hemihydrate. Science 363, 396 (2019).
Tambutté, S. et al. Coral biomineralization: from the gene to the environment. J. Exp. Mar. Biol. Ecol. 408, 58–78 (2011).
Gilbert, P. et al. Biomineralization: Integrating mechanism and evolutionary history. Sci. Adv. 8, eabl9653 (2022).
De Stasio, G. et al. MEPHISTO: performance tests of a novel synchrotron imaging photoelectron spectromicroscope. Rev. Sci. Instrum. 69, 2062–2066 (1998).
Scholl, A., Ohldag, H., Nolting, F., Stöhr, J. & Padmore, H. A. X-ray photoemission electron microscopy, a tool for the investigation of complex magnetic structures (invited). Rev. Sci. Instrum. 73, 1362–1366 (2002).
De Stasio, G. et al. MEPHISTO spectromicroscope reaches 20 nm lateral resolution. Rev. Sci. Instrum. 70, 1740–1742 (1999).
Venn, A. A. et al. Impact of seawater acidification on pH at the tissue-skeleton interface and calcification in reef corals. Proc. Natl. Acad. Sci. USA 110, 1634–1639 (2013).
Parasassi, T., Sapora, O., Giusti, A. M., De Stasio, G. & Ravagnan, G. Alterations in erythrocyte-membrane lipids induced by low-doses of ionizing-radiation as revealed by 1,6-diphenyl-1,3,5-hexatriene fluorescence lifetime. Int. J. Rad. Biol. 59, 59–69 (1991).
Scucchia, F., Sauer, K., Fara, S., Mass, T. & Zaslansky, P. 4D insights into coral biomineralization: effects of ocean acidification on the early skeleton development of a stony coral. Adv. Sci. 12, e73149 (2025).
Benzerara, K. et al. Study of the crystallographic architecture of corals at the nanoscale by scanning transmission X-ray microscopy and transmission electron microscopy. Ultramicroscopy 111, 1268–1275 (2011).
Malik, A. et al. Molecular and skeletal fingerprints of scleractinian coral biomineralization: from the sea surface to mesophotic depths. Acta Biomater. 120, 263–276 (2021).
Stolarski, J. Three-dimensional micro-and nanostructural characteristics of the scleractinian coral skeleton: a biocalcification proxy. Acta Palaeontol. Pol. 48, 497–530 (2003).
Meibom, A. et al. Vital effects in coral skeletal composition display strict three-dimensional control. Geophys. Res. Lett. 33, L11608 (2006).
Hobbie, R. K. & Roth, B. J. Exponential growth and decay. In Intermediate Physics for Medicine and Biology (eds Hobbie R. K. & Roth, B. J.) 31–47 (Springer International Publishing, 2007).
Venn, A. A. et al. Effects of light and darkness on pH regulation in three coral species exposed to seawater acidification. Sci. Rep. 9, 2201 (2019).
Bots, P., Benning, L. G., Rodriguez-Blanco, J.-D., Roncal-Herrero, T. & Shaw, S. Mechanistic insights into the crystallization of amorphous calcium carbonate (ACC). Cryst. Growth Des. 12, 3806–3814 (2012).
Gong, Y. U. T. et al. Phase transitions in biogenic amorphous calcium carbonate. Proc. Natl. Acad. Sci. USA 109, 6088–6093 (2012).
Radha, A. V., Forbes, T. Z., Killian, C. E., Gilbert, P. U. P. A. & Navrotsky, A. Transformation and crystallization energetics of synthetic and biogenic amorphous calcium carbonate. Proc. Natl. Acad. Sci. USA 107, 16438–16443 (2010).
Rodriguez-Blanco, J. D., Shaw, S., Bots, P., Roncal-Herrero, T. & Benning, L. G. The role of pH and Mg on the stability and crystallization of amorphous calcium carbonate. J. Alloy. Compd. 536, S477–S479 (2012).
Cohen, A. L. & Holcomb, M. Why corals care about ocean acidification: uncovering the mechanism. Oceanography 22, 118–127 (2009).
Cohen, A. L., McCorkle, D. C., de Putron, S., Gaetani, G. A. & Rose, K. A. Morphological and compositional changes in the skeletons of new coral recruits reared in acidified seawater: insights into the biomineralization response to ocean acidification. Geochem. Geophys. Geosys. 10, Q07005 (2009).
Godwin, H. Half-life of radiocarbon. Nature 195, 984–984 (1962).
Wilde, S. A., Valley, J. W., Peck, W. H. & Graham, C. M. Evidence from detrital zircons for the existence of continental crust and oceans on the Earth 4.4 Gyr ago. Nature 409, 175–178 (2001).
Politi, Y. et al. Transformation mechanism of amorphous calcium carbonate into calcite in the sea urchin larval spicule. Proc. Natl. Acad. Sci. USA 105, 17362–17366 (2008).
Killian, C. E. et al. Mechanism of calcite co-orientation in the sea urchin tooth. J. Am. Chem. Soc. 131, 18404–18409 (2009).
DeVol, R. T. et al. Nanoscale transforming mineral phases in fresh nacre. J. Am. Chem. Soc. 137, 13325–13333 (2015).
Schmidt, C. A. et al. Faster crystallization during coral skeleton formation correlates with resilience to ocean acidification. J. Am. Chem. Soc. 144, 1332–1341 (2022).
Turnbull, D. Kinetics of heterogeneous nucleation. J. Chem. Phys. 18, 198–203 (1950).
Avrami, M. Kinetics of phase change. I General theory. J. Chem. Phys. 7, 1103–1112 (1939).
Brown, M. E. The Prout-Tompkins rate equation in solid-state kinetics. Thermochim. Acta 300, 93–106 (1997).
Crank, J. The Mathematics of Diffusion (Oxford University Press, 1979).
Meldrum, F. C. & O’Shaughnessy, C. Crystallization in confinement. Adv. Mater. 32, 2001068 (2020).
Nam, H. et al. Network context and selection in the evolution to enzyme specificity. Science 337, 1101–1104 (2012).
Bezsudnova, E. Y. et al. Probing the role of the residues in the active site of the transaminase from Thermobaculum terrenum. PLoS ONE 16, e0255098 (2021).
Akiva-Tal, A. et al. In situ molecular NMR picture of bioavailable calcium stabilized as amorphous CaCO3 biomineral in crayfish gastroliths. Proc. Natl. Acad. Sci. USA 108, 14763–14768 (2011).
Al-Sawalmih, A., Li, C. H., Siegel, S., Fratzl, P. & Paris, O. On the Stability Of Amorphous Minerals In Lobster Cuticle. Adv. Mater. 21, 4011 (2009).
Stephens, C. J., Ladden, S. F., Meldrum, F. C. & Christenson, H. K. Amorphous calcium carbonate is stabilized in confinement. Adv. Funct. Mater. 20, 2108–2115 (2010).
Tambutté, E. et al. Morphological plasticity of the coral skeleton under CO-driven seawater acidification. Nat. Commun. 6, 7368 (2015).
Mergelsberg, S. T. et al. Metastable solubility and local structure of amorphous calcium carbonate (ACC). Geochim. Cosmochim. Acta 289, 196–206 (2020).
De Stasio, G., Frazer, B. H., Gilbert, B., Richter, K. L. & Valley, J. W. Compensation of charging in X-PEEM: a successful test on mineral inclusions in 4.4 Ga old zircon. Ultramicroscopy 98, 57–62 (2003).
Gilbert B. Gilbert PUPA. GG Macros. https://doi.org/10.5281/zenodo.17314121 (2025).
Coronado, I., Fine, M., Bosellini, F. R. & Stolarski, J. Impact of ocean acidification on crystallographic vital effect of the coral skeleton. Nat. Commun. 10, 2896 (2019).
Stolarski, J. et al. A modern scleractinian coral with a two-component calcite–aragonite skeleton. Proc. Natl. Acad. Sci. USA 118, e2013316117 (2021).
Moler, C. & Little, J. A history of MATLAB. Proc. ACM Program. Lang. 4, 1–67 (2020).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python (vol 33, pg 219, 2020). Nat. Methods 17, 352–352 (2020).
McKinney, W. Data structures fro statistical computing in Python. SciPy. https://doi.org/10.25080/Majora-92bf1922-00a (2010).
Marquardt, D. W. Citation Classic—algorithm for least-squares estimation of non-linear parameters. Contents/Eng. Technol. Appl. Sci. (1979).
Nakagawa, S. & Schielzeth, H. A general and simple method for obtaining R2 from generalized linear mixed-effects models. Methods Ecol. Evol. 4, 133–142 (2013).
Rechav, Z. & LeCloux, I. M. Zenodo, https://doi.org/10.5281/zenodo.18175786 (2026).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
The authors thank Aiden Gustafson and James J. De Yoreo for scientific discussions, M. Cristina Castillo Alvarez and Connor A. Schmidt for assistance during sample preparation and PEEM data acquisition, and Andreas Scholl for technical help during PEEM measurements. This work was supported by the National Science Foundation Graduate Research Fellowship Program (grant DGE-1747503) (Z.R.). This material is based upon work supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE-1747503. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation. This work was supported by the U.S. Department of Energy, Office of Science, Basic Energy Sciences, Chemical Sciences, Geosciences, and Biosciences Division at the University of Wisconsin–Madison (grant DE-FG02-07ER15899) (P.G.) and at Lawrence Berkeley National Laboratory (grant FWP-FP00011135) (P.G.), and by the National Science Foundation Biomaterials Program (grant DMR-2220274) (P.G.). This research used resources of the Advanced Light Source, a U.S. Department of Energy Office of Science User Facility under Contract No. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
P.G., I.L., and Z.R. conceptualized the study and carried out the investigation. E.T., S.T., and A.V. provided samples. P.G., I.L., Z.R., and B.A. performed PEEM data acquisition. M.M. production was carried out by I.L., S.A., N.B., N.C., B.D.-K., J.D., A.L., S.L., R.R., L.S., J.L.S., J.S.S., C.W., J.Y., Z.R., and P.G. I.L. and Z.R. performed the data analysis. P.G. and Z.R. acquired funding. P.G. and Z.R. wrote the original draft of the manuscript. P.G., Z.R., and all co-authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Gabriela Farfan and the other anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rechav, Z., Tambutté, E., LeCloux, I.M. et al. Exponential crystallization in corals. Nat Commun 17, 2870 (2026). https://doi.org/10.1038/s41467-026-69215-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69215-4